Immunogenetics Laboratory, Immunology Program, Memorial Sloan-Kettering Cancer Center, New York, NY
|
|
|
- Christopher Derrick Howard
- 5 years ago
- Views:
Transcription
1 Biology of Blood and Marrow Trans lantation 5:77 85 (1999) 1999 American Society for Blood and Marrow Transplantation ASBMT HLA-C disparity between patients and unrelated donors matched for HLA-A, -B, and -DRB1 alleles: impact of serological vs. DNA typing for HLA-A and -B loci Vinod K. Prasad, 1,3 Nancy A. Kernan, 1 Glenn Heller, 2 Richard J. O Reilly, 1 Soo Young Yang 3 1 Bone Marrow Transplantation Service, Department of Pediatrics; 2 Department of Biostatistics; and 3 Biochemical Immunogenetics Laboratory, Immunology Program, Memorial Sloan-Kettering Cancer Center, New York, NY Offprint requests: Vinod K. Prasad, MD, MRCP, Box 379, Memorial Sloan-Kettering Cancer Center, 1275 York Avenue, New York, NY (Received 11 October 1998; accepted 15 February 1999) ABSTRACT High incidences of graft failure, graft-vs.-host disease (GVHD), and serious infections following unrelated donor (URD) marrow transplantation, despite apparent human leukocyte antigen (HLA) identity, may reflect the presence of molecular disparities, including those for HLA-C alleles between the patient and the URD. The level of these disparities could be significant, because as many as 42 alleles are currently known for HLA-C locus. We studied 84 patients and 251 potential URDs to evaluate 1) the extent of HLA-C disparity between the patient and the URD identified by serology for HLA-A and -B and by DNA typing for -DRB1 and 2) the level of HLA-C disparity between patients and URDs matched by high-resolution DNA typing for HLA-A, -B, and -DRB1. The DNA typing was performed at the Memorial Sloan Kettering Cancer Center, and the serotyping was provided by the registries. Of 251 URDs matched by HLA-A and -B serology and -DRB1 (sa_sb_dnadrb1); 94, 75, and 82 were 6/6, 5/6, and 4/6 matches, respectively. Of 94 sa_sb_dnadrb1 6/6 URDs, 51 (54.3%) were matched for both HLA-C alleles. In contrast, 31 (41.3%) 5/6 (p 0.12) and 15 (18.3%) 4/6 (p 0.01) sa_sb_dnadrb1 URDs were matched for both HLA-C alleles. Following DNA typing for HLA-A and -B, 52 (55.3%) of 94 6/6, 30 (40%) of 75 5/6, and 25 (30.5%) of 82 4/6 sa_sb_dnadrb1 URDs remained 6/6, 5/6, and 4/6 matches at the DNA level (dnaa_b_drb1). HLA-C disparities continued to exist in the dnaa_b_drb1 URD group. Of 54 dnaa_b_drb1 6/6 URDs, 41 (75.9%) were matched for both HLA-C alleles. Only 45.3% of the 5/6 (p 0.01) and 22.2% of the 4/6 (p 0.01) dnaa_b_drb1 URDs were matched for both HLA-C alleles. In the 6/6 category, the frequency of HLA-C matching improved (75.9 vs. 54.3%; p 0.01) following DNA matching for HLA-A and -B. In comparison to mismatching for HLA-B locus, mismatching for either HLA-DRB1 or -A resulted in a lower odds ratio for HLA-C disparity. The presence of a common haplotype in the sa_sb_dnadrb1 (p 0.06) URD category improved the level of HLA-C matching. We identified alleles that are associated with high (B*1501, B*4402, B*5101, DRB1*0101, A*0201, A*1101, A*2301, and A*3201) or low (B*0702, B*0801, B*1302, B*3502, DRB1*0301, DRB1*1104, A*0101, A*3001, and A*6801) probability of HLA-C disparity. Overall, sa_sb_dnadrb1 as well as dnaa_b_drb1 matched URDs for non-caucasian patients were more likely to have HLA-C disparity in comparison to the matched URDs of Caucasian patients. However, a high incidence of HLA-C disparities was identified even in the URDs for Caucasian patients. Whether the disparities demonstrated by this study contribute to the higher immunological complications noted following URD bone marrow transplantation is unclear. Outcome analysis and studies aimed at understanding the functional role of HLA-C may provide an answer. KEY WORDS Molecular disparity Bone marrow transplantation Unrelated donor DNA typing This study was supported in part by grants from the National Cancer Institute, National Institutes of Health (N.A.K., G.H., R.J.O., S.Y.Y.; PO1 CA23766), the Butler Foundation (N.A.K.), the Vincent Astor Chair in Clinical Research (R.J.O.), and Navy Medical Research (S.Y.Y.; N ). INTRODUCTION In the last decade, a matched unrelated donor (URD) has become an increasingly effective source of stem cells for transplantation for patients without a family donor [1,2]. In
2 addition, because of the tremendous growth of national and international donor registries, the number of URD stem cell transplants has increased during this period [1 5]. Concurrently, there has also been a consistent improvement in the outcome of URD transplants [6 10]. Despite these gains, the incidences of immunological complications (including graft-vs.-host disease [GVHD], graft rejection, and delayed immune reconstitution, leading to potentially life-threatening infections) continue to be far more severe and frequent following URD bone marrow transplantation (BMT) [8,11 13]. The difference in outcomes between sibling donor and URD transplantations may reflect the presence of genotypic disparities between the patients and the URDs that remain undetected by conventional HLA typing using serology for HLA-A and -B and DNA typing for -DRB1. The siblings who match for HLA-A, -B, and -DR by conventional testing (and who are, therefore, phenotypically identical), are very likely to be genotypically identical because of the inheritance of haplotypes comprising the whole HLA complex from the parents [14]. Because recombination within the HLA complex is rare [14], the family donors are also likely to have allelic identity within the HLA complex at loci other than HLA-A, -B, and -DR. In contrast, a patient URD pair that matches for HLA-A and -B by serology and for HLA-DRB1 by the DNA typing may be genotypically disparate as a result of the presence of molecular variant disparities at HLA-A and/or HLA-B loci at the allelic level; and disparities at HLA loci other than HLA-A, -B, and -DRB; or both. We have recently shown that genotypic disparities at the HLA-A and -B loci do exist between serologically matched patient URD pairs and that DNA typing can elucidate these disparities [15]. However, DNA typing for HLA-A and -B may not assure genotypic matching for other alleles within the HLA complex. Recent laboratory and clinical data indicate that HLA-C plays a more significant role in alloreactions and BMT than previously suspected [16 23]. With 42 known alleles [24], HLA-C is a highly polymorphic locus. Therefore, it would not be surprising if significant disparity for HLA-C were to be found between patients and otherwise matched URDs. In the past, the significance of HLA-C in alloreactions affecting the outcome of marrow transplants was not studied. Analyses were hindered by technical difficulties, including the existence of multiple alleles constituting as many as 30 45% of the total that could not be typed by serology and were reported as blanks [25]. Evaluation of HLA-C disparities and their significance has now become possible with the development of DNA typing methods for class I loci [17,26 28]. In this study we have applied novel, highly discriminatory molecular techniques in a large cohort to identify at the DNA level the HLA-C disparities between patients and their potential URDs who were also molecularly typed for HLA-A, -B, and -DRB1. Our aims were as follows: a) to identify the extent and the pattern of HLA-C genotypic disparities existing between patient URD pairs matched by conventional typing (HLA-A and -B serology and DNA typing for HLA-DRB1); b) to examine if the selection of URDs based on matching for HLA-A, -B, and -DRB1 by DNA typing has any impact on the incidence or degree of HLA-C disparity; c) to analyze the influence of individual matching for HLA-A, -B, and -DRB1 loci on the level of HLA-C genotypic disparity; d) to determine if the level of HLA-C disparity in patient URD pairs expressing one of the commonly observed haplotypes is different from pairs without these common haplotypes; and e) to determine the influence of the race/ethnicity of the patients on the frequency of HLA-C disparity. MATERIALS AND METHODS The study was conducted between July 1996 and February 1998, at the Memorial Sloan Kettering Cancer Center (MSKCC), New York. During the study period, 84 patients and 251 potential URDs were identified and typed by high-resolution DNA typing for HLA-A, -B, -C, and - DRB1. By HLA-A and HLA-B serology and sequence-specific oligonucleotide probe (SSOP) analysis of HLA-DRB1 (sa_sb_dnadrb1), 94 URDs were 6/6, 75 were 5/6, and 82 were 4/6 matches. Following DNA typing for HLA-A and - B (dnaa_b_drb1), 52 of 94 (55.3%) originally 6/6 URDs remained 6/6. Of 75 5/6 and 82 4/6 sa_sb_dnadrb1 URDs, 30 (40%) and 25 (30.5%) remained 5/6 and 4/6 DNA matches, respectively. Fifty-seven of 84 patients selfreported as Caucasians. A total of 182 URDs were selected for 57 Caucasian patients. The remaining 69 URDs were selected for 27 non-caucasian patients (Hispanic, 10; African-American, 8; Asian, 8; others, 1). The cohort of URDs selected for Caucasian patients will be called ptcauc and those selected for non-caucasian patients will be called ptnoncauc. Typing technique The DNA typing for HLA-A, -B, -C, and -DRB1 alleles for the patients and URDs were conducted at our center. The serological typing on the URDs was provided by the registries. Our pilot studies to evaluate the need for reserotyping all samples from URDs indicated that this expensive and time-consuming testing was unnecessary. Level of discrepancy was very low. In addition, we have recently demonstrated that after DNA typing, less than 2% of URDs were identified as having HLA-A and -B alleles outside of their registry-defined serotypes [29]. URDs who matched for World Health Organization (WHO) defined split antigens [30] were considered serological matches. DNA typing for HLA-A, -B, and -C was performed by SSOP, as previously described [29,31], and HLA-DRB1 typing was performed according to the well-established protocol [32]. Alleles were assigned according to the DNA sequences published by the WHO committee [30]. The nucleated cells from the blood samples were collected, and genomic DNA was isolated [28]. PCR amplification of the full length of exons 2 and 3 of the HLA-A, -B, and -C genes from the genomic DNA was performed using primers derived from introns 1 and 3. The PCR product was dot-blotted on nylon membrane and then cross-linked by ultraviolet light. The membranes were hybridized with digoxigeninddutp labeled oligonucleotide probes. After washing, the membranes were treated with anti-digoxigenin-fab antibody conjugated to alkaline phosphatase and were then sprayed with Lumiphos 480 (Boehringer Mannheim, Germany) and exposed to photographic film. We used a panel of 44, 52, and 44 probes for the HLA-A, -B, and -C loci, 78
3 yp g Figure 1. Frequency of the occurrence of HLA-C alleles in the patient population depicted as percentages Each column represents one HLA-C allele. Alleles belonging to each serogroup are placed together. The alleles that give a blank reaction on serological testing are grouped at the right end of the graph. respectively. The probes were designed from the sequences of exons 2 and 3. A locally developed computer program analyzed the hybridization pattern for each sample and assigned alleles. The results were checked and confirmed manually. Heterozygotes were identified by the presence of two different hybridization score patterns. Data analysis and statistics URDs were stratified according to the degree of matching by HLA-A, -B serology, and HLA-DRB1 (sa_sb_dnadrb1) into 6/6, 5/6, and 4/6 groups. A subgroup of URDs was found to be matched for HLA-A, -B, and -DRB1 by DNA typing (dnaa_b_drb1). Following DNA typing for HLA-C, URDs were classified as HLA-C matched (matched for both HLA-C alleles) or HLA-C mismatched (mismatched for one or both HLA-C alleles). All tests of no association between categorical variables, including the frequency of HLA-C disparity, were computed using Fisher s exact test. RESULTS HLA characteristics of patients and URDs A total of 22 HLA-C alleles were identified in our patient cohort, of which Cw*0401 was the most common (Fig. 1). In comparison to the two published reports on the Caucasoid population of Europe [33,34], the frequency of the HLA-C alleles in our patients revealed one notable difference. Cw*0401 occurred with a frequency of 18.5% in our patients, compared with 10 12% in the European studies [33,34]. Of the 84 patients, 8 (9.5%) were found to be homozygous for HLA-C locus following family studies. One allele in 30 of 84 patients (35.7%) and both alleles in two patients (2.4%) belonged to the serologically blank category. Of the serologically blank HLA-C alleles, Cw*1203 was the most common. A total of 29 HLA-A, 45 HLA B, and 37 HLA-DRB1 alleles were observed (Table 1). A*0201, B*0702, and DRB1*0701 were the most common alleles within the respective loci. Following DNA typing, 2.1% of HLA-A and 2.8% of HLA-B alleles were found to be outside the serological group they were assigned by the National Marrow Donor Program or other registry. HLA-C matching: sa_sb_dnadrb1 vs. dnaa_b_drb1 HLA-C disparity for one allele between the patient and URD was seen in 37 of the 94 (39.4%) 6/6 sa_sb_dnadrb1 URDs. Six (6.4%) URDs were disparate for both HLA-C alleles. The remaining 51 (54.3%) 6 of 6 sa_sb_dnadrb1 URDs were matched at both HLA-C alleles. Compared with the 6/6 sa_sb_dnadrb1 category, the level of HLA-C disparity was significantly higher (Fig. 2a) within the 4/6 (p 0.01) sa_sb_dnadrb1 URD category. In addition, the level of HLA-C disparity was higher within the 4/6 compared with the 5/6 (p 0.01) sa_sb_dnadrb1 URDs. However, the difference in the level of HLA-C disparity between 6/6 and 5/6 BB&MT 79
4 Table 1. Frequency of HLA-A, -B, and DRB1 alleles in the patient population HLA-A allele Frequency (%) HLA-B allele Frequency (%) HLA-DRB1 allele Frequency (%) A* B* DRB1* A* B* DRB1* A* B* DRB1* A* B* DRB1* A* B* DRB1* A* B* DRB1* A* B* DRB1* A* B* DRB1* A* B* DRB1* A* B* DRB1* A* B* DRB1* A* B* DRB1* A* B* DRB1* A* B* DRB1* A* B* DRB1* A* B* DRB1* A* B* DRB1* A* B* DRB1* A* B* DRB1* A* B* DRB1* A* B* DRB1* A* B* DRB1* A* B* DRB1* A* B* DRB1* A* B* DRB1* A* B* DRB1* A* B* DRB1* A* B* DRB1* A* B* DRB1* Total B* DRB1* B* DRB1* B* DRB1* B*15N 0.60 DRB1* B* DRB1* B* DRB1* B* DRB1* B* DRB1* B* Total B* B* B* B* B* B* B* B*51N 0.60 B* Total Patient number, 84; allele number, sa_sb_dnadrb1 URDs (45.7 vs. 58.7%) was not as strong (p 0.12). HLA-C disparities continued to exist despite matching for HLA-A and -B by DNA typing. Of 54 6/6 dnaa_b_drb1 URDs, 12 (22.2%) and one (1.9%) were found to be mismatched for one or both HLA-C alleles, respectively. The level of HLA-C disparity was higher in 5/6 (p 0.08) and 4/6 (p 0.01) dnaa_b_drb1 subsets (Fig. 2b). In addition, the level of HLA-C disparity was higher within the 4/6 compared with the 5/6 (p 0.01) dnaa_b_drb1 URDs. The level of HLA-C disparity between sa_sb_dnadrb1 and dnaa_b_drb1 within each matching category (6/6, 5/6, and 4/6) was compared to determine if URD matching by DNA typing for HLA-A and -B improves the level of HLA-C matching (Table 2). A higher percentage of 6/6 dnaa_b_drb1 URD were matched for both HLA-C alleles in comparison with 6/6 sabdnadrb1 URDs (75.9 vs. 54.3%; p 0.01). Overall, dnaa_b_drb1 URDs were found to have significantly less (p 0.01) HLA-C disparity than the sa_sb_dnadrb1 URDs. The differences 80
5 yp g A B Figure 2. Level of HLA-C genotype identified in (A) conventionally matched (sa_sb_dnadrb1) and (B) DNA matched (dnaa_b_drb1) URDS Each column represents the HLA-A, -B, and -DRB1 matching status of the unrelated donor (URD) (6/6, 5/6, and 4/6). The blocks within each column represent the level of HLA-C matching. The number of URDs in each category is shown within the columns and the percentage values are next to the column. (A) Compared with the 6/6 sa_sb_dnadrb1 category, the level of HLA-C disparity was significantly higher within the 4/6 (p 0.01) sa_sb_dnadrb1 URD category. In addition, the level of HLA-C disparity was higher within the 4/6 compared with the 5/6 (p 0.01) sa_sb_dnadrb1 URD. However, the difference in the level of HLA-C matching between 6/6 and 5/6 sa_sb_dnadrb1 URDs (54.3 vs. 41.3%; p 0.12) was not as strong. (B) Compared with the 6/6 dnaa_b_drb1 category, the level of HLA-C disparity was higher in the 5/6 (p 0.08) and the 4/6 (p 0.01) dnaa_b_drb1 subsets. In addition, the level of HLA-C disparity was higher within the 4/6 compared with the 5/6 (p 0.01) dnaa_b_drb1 category. within the 5/6 and 4/6 categories were not significant (p 0.19 for 5/6 and p 1.0 for 4/6). Effect of matching for HLA-A, -B, and -DRB1 loci individually The effect of genotypic matching for HLA-A, -B, and - DRB1 loci individually on the level of HLA-C disparity was determined (Table 3). The URDs (n 251) were stratified for HLA-A, -B, and -DRB1 matching as follows: MA, matched for both HLA-A alleles (n 171); MMA, mismatched for HLA-A alleles (n 80); MB, matched for both HLA-B alleles (n 121); MMB, mismatched for HLA-B alleles (n 130); MDRB1, matched for both HLA-DRB1 alleles (n 114); and MMDRB1, mismatched for HLA-DRB1 alleles (n 137). Within the MA group, 77 of 171 (45.1%) were matched for both HLA-C alleles, compared with 20 of 80 (25%) URDs within the MMA group (p 0.01; odds ratio 2.46). In the MB group, 67 of 121 (55.4%) were matched for HLA-C, whereas in the MMB group, 30 of 130 (23.1%) URDs were matched for both HLA-C alleles (p 0.01; odds ratio 4.14). Within the MDRB1 group, 56 of 114 (49.1%), and in the MMDRB1 group, 41 of 137 (29.9%) URDs were matched for both HLA-C alleles (p 0.01; odds ratio 2.26). Further analysis of the patients and their matched URDs led to the identification of HLA-B, -DRB1, and -A alleles that are associated with high risk and others with low risk of HLA-C disparity. We define high risk and low risk as observing greater than 2:1 odds for HLA-C mismatching and matching, respectively. For example, all HLA- A matched URDs were stratified into HLA-C matched and HLA-C mismatched subgroups. The frequency of occurrence of various HLA-A alleles was then compared between the two subgroups. The HLA-A alleles observing greater than 2:1 odds for HLA-C matching (low risk) and HLA-C mismatching (high risk) were identified. The matched URDs expressing B*0702, B*0801, B*1302, B*3502, DRB1*0301, DRB1*1104, A*0101, A*3001, and A*6801 were at low risk for HLA-C disparity. In contrast, matched URDs with Table 2. Comparison of the level of HLA-C disparity between sa_sb_dnadrb1 and dnaa_b_drb1 within each matching category a Match criteria for Number of HLA-C match HLA-C mismatch HLA-A, -B, and -DRB1 URDs n (%) n (%) p 6/6 sa_sb_dnadrb (54.3) 43 (45.7) dnaa_b_drb (75.9) 13 (24.1) 5/6 sa_sb_dnadrb (41.3) 44 (58.7) dnaa_b_drb (56.7) 13 (43.3) 4/6 sa_sb_dnadrb (18.3) 67 (81.7) dnaa_b_drb (20.0) 20 (80.0) All sa_sb_dnadrb (38.6) 154 (61.4) dnaa_b_drb (57.8) 46 (42.2) a Matching categories include 6/6, 5/6, and 4/6. BB&MT 81
6 Table 3. Effect of DNA matching for HLA-A, -B, and -DRB1 alleles on the level of HLA-C matching Both HLA-C allele One HLA-C allele Both HLA-C allele Total number Comparison Allele match/mismatch match [n (%)] mismatch [n (%)] mismatch [n (%)] of URDs parameter p Odds ratio Matched for both -A (MA) 77 (45.0) 76 (44.4) 18 (10.5) 171 Mismatched for -A (MMA) 20 (25.0) 45 (56.25) 15 (18.75) 80 Matched for both -B (MB) 67 (55.4) 51 (42.1) 3 (2.5) 121 Mismatched for -B (MMB) 30 (23.1) 70 (53.8) 30 (23.1) 130 Matched for both -DRB1 (MDRB1) 56 (49.12) 47 (41.23) 11 (9.65) 114 Mismatched for -DRB1 (MMDRB1) 41 (29.9) 74 (54.0) 22 (16.1) 137 MA vs. MMA MB vs. MMB MDRB1 vs. MMDRB B*1501, B*4402, B*5101, DRB1*0101, A*0201, A*1101, A*2301, and A*3201 were at high risk. The influence of common haplotypes Haplotypes were assigned for 59 of 84 patients by family typings. Of 251 URDs, 188 had been selected for these 59 patients. Fifteen patients carried one of the 10 most common serologically defined haplotypes (A1B8DR3, A2B7DR15, A2B15DR4, A2B44DR4, A2B50DR7, A2B51DR13, A3B7DR15, A24B35DR11, A29B44DR7, and A30B13DR7) [25]. None of the patients had two of the above haplotypes. A total of 55 URDs had been selected for these 15 patients by original typing (sa_sb_dnadrb1). The URDs for these patients made up the common haplotype (CH) sa_sb_dnadrb1 subgroup, and the URDs (n 133) identified for the rest of the patients (n 44) constituted the NoCH sa_sb_dnadrb1 group. Both of the groups were further stratified according their original typing status into 6/6, 5/6, and 4/6 subgroups. The level of donor recipient genotypic disparity was lower in the CH sa_sb_dnadrb1 group in comparison to the NoCH sa_sb_dnadrb1 group (n 0.06). The interpretation of the differences in the level of HLA-C disparity between CH sa_sb_dnadrb1 and NoCH sa_sb_dnadrb1 URDs within 6/6, 5/6, and 4/6 subgroups was limited because of small numbers within individual subgroups. Effect of patient ethnicity/race Of 182 ptcauc URDs, 80, 56, and 46 were 6/6, 5/6, and 4/6 matches by original typing (sa_sb_dnadrb1; Table 4). Of 69 ptnoncauc URDs, 14, 19, and 36 were 6/6, 5/6, and 4/6 matches by original typing. The percentage of URDs with HLA-C disparity was significantly higher in the ptnoncauc compared with the ptcauc URDs (75.4 vs. 56.0%; p 0.01). Because of the small sample size, differences between ptcauc and ptnoncauc sa_sb_dnadrb1 URDs were not significant in the 6/6 (p 0.39), 5/6 (p 0.78), and 4/6 (p 0.16) matching subcategories. Similar analysis was carried out for the URDs matched by DNA typing for HLA-A, -B, and -DRB1 (dnaa_b_drb1). The ptcauc group contained 50 6/6, 49 5/6, and 48 4/6 and the ptnoncauc group had 4 6 /6, 15 5/6, and 24 4/6 dnaa_b_drb1 matched URDs. Again, in the dnaa_b_drb1 group, the percentage of URDs with HLA-C disparity was significantly higher in the ptnon- CAUC compared with the ptcauc URDs (72.1 vs. 49.7%; p 0.01). The differences between ptcauc and ptnon- CAUC dnaa_b_drb1 URDs were not significant in the 6/6 (p 0.56) and 5/6 (p 0.56) subgroups, potentially due to small numbers of ptnoncauc URDs within the subgroups. DISCUSSION We have recently shown that the use of serologic typing for HLA-A and -B results in HLA-A and -B disparities in 47% of 6/6 matched patient URD pairs [15]. In this study we demonstrate that HLA-C disparities exist between many patients and their conventionally matched (sa_sb_dnadrb1) or DNA matched (dnaa_b_drb1) URD. Although HLA-C Table 4. Effect of ethnicity/race of the patients on the level of HLA-C disparity between patient URD pairs matched by serology for HLA-A and -B and DNA typing for HLA-DRB1(sA_sB_dnaDRB1) as well as between pairs matched by DNA typing for HLA-A, -B, and -DRB1 (dnaa_b_drb1) HLA-A, -B, and -DRB1 sa_sb_dnadrb1 URD dnaabdrb1 URD matching HLA-C match Mismatch HLA-C match Mismatch status [n (%)] [n (%)] Total p [n (%)] [n (%)] Total p 6 of 6 ptcauc 45 (56.3) 35 (43.8) (74.0) 13 (26.0) ptnoncauc 6 (42.9) 8 (57.1) 14 4 (100) 0 (0.0) 4 5 of 6 ptcauc 24 (42.9) 32 (57.1) (42.9) 28 (57.1) ptnoncauc 7 (36.8) 12 (63.2) 19 8 (53.3) 7 (46.7) 15 4 of 6 ptcauc 11 (23.9) 35 (76.1) (33.3) 32 (66.7) ptnoncauc 4 (11.1) 32 (88.9) 36 0 (0) 24 (100) 24 All URDs ptcauc 80 (44.0) 102 (56.0) (50.3) 73 (49.7) ptnoncauc 17 (24.6) 52 (75.4) (27.9) 31 (72.1)
7 yp g matching is not a part of current criteria for selecting stem cell donors, the guidelines may change in view of growing evidence of the importance of HLA-C in clinical stem cell transplantation. Recent clinical reports suggest that a donor recipient mismatch at the HLA-C locus may be associated with GVHD [20,23], marrow graft rejection [20,22], and renal transplant rejection [35]. In a recent Japanese report, HLA-C mismatching was associated with a high risk (p 0.001) of grade III IV acute GVHD [23]. In addition, the authors observed a slightly higher incidence of relapse in HLA-C matched recipients (p 0.06). Although the findings are noteworthy, they should not be extrapolated to a much more ethno-racially mixed U.S. population. The level of HLA-C disparity identified in our study confirms and extends that reported by a European group who examined 73 patients and 184 potential matched URDs selected by serology for HLA-A and -B and DNA typing for -DRB1 [17]. The level of HLA-C disparity in the Japanese study was lower than that seen by us. In the present study, we additionally determined that HLA-C disparities continue to exist, although to a lesser extent, despite DNA typing for HLA-A and -B. In addition, fewer than 6 of 6 HLA-A, -B, and -DRB1 match either by serology or by DNA typing results in a tendency towards worse HLA-C matching. The indicators of the probability of HLA-C matching that we have identified in this study may be of use until HLA-C matching by DNA typing is accepted as a criteria for URD selection. Because the odds ratio of HLA-C matching in B-mismatched URDs is higher in comparison to either A- or DRB1- mismatched URDs, it is likely that when selecting a partially mismatched URD, matching for HLA-B may be preferable to matching at either of the other two loci. In a separate study, we not only established the presence of a strong linkage disequilibrium (LD) between HLA-B and -C loci at the allelic level, but also determined the probability of HLA-C matching between two unrelated individuals who match for both -B alleles (36). The determinations were made for most of the common HLA-B alleles seen in our population. With the above approach, one is less likely to select an HLA-C mismatched URD. Furthermore, information about the high-risk alleles like B*1501, B*4402, B*5101, DRB1*0101, A*0201, A*1101, A*2301, and A*3201; and low-risk alleles like B*0702, B*0801, B*1302, B*3502, DRB1*0301, DRB1*1104, A*0101, A*3001, and A*6801 may be of value when DNA typing for HLA-C is unavailable for all patients and URDs. HLA-C typing should be directed toward the patients and URDs who carry one or more high-risk HLA-A, -B, or -DRB1 allele. An important finding in our study is that patients who carry a commonly occurring haplotype are less likely to have HLA-C disparity. This observation was made in the serologically matched URD and reflects LD between HLA-C and other loci at the allelic level. It is interesting that HLA-A1B8DR3, the most common haplotype in the United States, is strikingly unique. Following DNA typing for HLA-A and -B, all HLA-A1 and HLA-B8 serotypes in the present and in a previous study [15] remained A*0101 and B*0801, respectively. Therefore, all sero-matched URDs with these serotypes remained DNA matches at HLA-A and -B. Furthermore, B*0801 demonstrates a very strong LD with Cw*0701. In a study of 849 individuals with known haplotypes, all B*0801 were seen in association with Cw*0701 [36]. It is therefore very likely that a seromatched HLA-A1B8DR3 URD will be DNA matched for HLA-A and -B, as well as -C alleles. A large cohort of patients and URDs with alleles from a majority of the known HLA-A, -B, -C, and -DR serological groups suggests that these data represent a fair cross-section of the population. Although most of the HLA-A, -B, and -DR haplotypes common in the Caucasian population were represented (A1B8DR3, A3B7DR2, A2B62DR4, and more), haplotypes that are comparatively more common in Asian (A24B52DR2) and African-Americans (A30B42DR3) [25,37] were also seen in our cohort, suggesting a mixed ethnic background of the study population. Our analysis of the patient s ethnic/racial background on the level of HLA- C disparity revealed two important points. First, there was a high incidence of HLA-C disparity in URDs for Caucasian as well as non-caucasian patients. As many as 43.8% of 6/6 sa_sb_dnadrb1 and 25.4% of dnaa_b_drb1 URDs for Caucasian patients showed HLA-C disparities. Second, the overall level of HLA-C disparities was higher if patients were non-caucasians rather than Caucasians, irrespective of the typing methodology used for HLA-A and -B (p 0.01 for sa_sb_dnadrb1; p 0.01 for dnaa_b_drb1). The difference in the level of HLA-C disparity between ptcauc and ptnoncauc URDs was not significant when the 6/6 or 5/6 subgroups were analyzed separately. High HLA-C disparity in Caucasians reflects wide ethno-geographic origin of the Caucasian population of the United States. However, it must be remembered that these observations may be influenced by bias arising from voluntary and self-reported racial designations. Although we clearly demonstrate the presence of a high incidence of HLA-C disparities at the DNA level in HLAmatched patient URD pairs, the clinical significance of these disparities has as yet not been conclusively proven. We hope that future clinical studies will further illustrate the importance of HLA-C in BMT. If such studies confirm a correlation between HLA-C matching and transplant outcome, HLA-C matching by DNA typing may be recommended for URD selection, especially if multiple URDs are available. One may argue that inclusion of HLA-C as a matching criterion will limit the number of available donors. Our own analysis offers encouraging results. For 40.9% (9 of 22) patients with more than one 6/6 URD, 40.9% (9 of 22) with more than one 5/6 URDs, and 56.5% (13 of 23) with more than one 4/6 URD, a better HLA-C match was available within our cohort of URDs. Our observation suggests that finding a better HLA-C match within the HLA- A, -B, and -DRB1 matched URD identified by current registries is possible. In the last few years, HLA-C has been increasingly recognized by a number of laboratory investigators to play an important immunological role in regulating the cytotoxic activity of natural killer (NK) cells in their interactions with hemopoietic cells. The engagement of killer inhibitory receptors on NK cells by self-type HLA-C gene products on the target cells can inhibit NK cell-mediated cytotoxicity. Indeed, HLA-C has been identified as the critical self-identifying ligand that protects targets from lysis by allodiscriminatory NK cells [38 44]. Given the role played by NK cells as the mediators of GVHD and graft rejection [45,46], it is BB&MT 83
8 likely that HLA-C may be an important determinant of BMT outcome. Analyses of the results of transplants between HLA-C disparate individuals coupled with an improved understanding of the role of HLA-C in donor host cell interactions will define whether and to what degree the routine use of HLA-C typing by DNA methods improves selection of an appropriate URD. ACKNOWLEDGMENTS The authors thank Carol Landrey for immense help with sample and serology data collection on the URDs. We thank the staff members of the histocompatibility laboratory and deeply appreciate their technical support. The authors thank Frank Lewis of Information Service at MSKCC for his help with ethnicity/race data collection on the patients. REFERENCES 1 Dupont B: Immunology of hematopoietic stem cell transplantation: a brief review of its history. Immunol Rev 157:5, Hansen JA, Petersdorf E, Martin PJ, Anasetti C: Hematopoietic stem cell transplants from unrelated donors. Immunol Rev 157:141, Freytes CO, Beatty PG: Representation of Hispanics in the National Marrow Donor Program. Bone Marrow Transplant 17:323, Perkins HA, Hansen JA: The U.S. National Marrow Donor Program. Am J Pediatr Hematol Oncol 16:30, Downie TR, Hows JM, Gore SM, Bradley BA, Howard MR: A survey of use of unrelated volunteer donor bone marrow transplantation at 46 centres worldwide, International Marrow Unrelated Search and Transplant (IMUST) Study. Bone Marrow Transplant 15:499, Anasetti C, Etzioni R, Petersdorf EW, Martin PJ, Hansen JA: Marrow transplantation from unrelated volunteer donors. Annu Rev Med 46:169, Hansen JA, Anasetti C, Petersdorf E, Clift RA, Martin PJ: Marrow transplants from unrelated donors. Transplant Proc 26:1710, Kernan NA, Bartsch G, Ash RC, Beatty PG, Champlin R, Filipovich A, Gajewski J, Hansen JA, Henslee-Downey J, McCullough J, McGlave P, Perkins HA, Phillips GL, Sanders J, Stroncek D, Thomas ED, Blume KG: Analysis of 462 transplantations from unrelated donors facilitated by the National Marrow Donor Program. N Engl J Med 328:593, Hansen JA, Gooley TA, Martin PJ, Appelbaum F, Chauncey TR, Clift RA, Petersdorf EW, Radich J, Sanders JE, Storb R, Sullivan KM, Anasetti C: Bone marrow transplants from unrelated donors for patients with chronic myeloid leukemia. N Engl J Med 338:962, Marks DI, Bird JM, Cornish JM, Goulden NJ, Jones CG, Knechtli CJ, Pamphilon DH, Steward CG, Oakhill A: Unrelated donor bone marrow transplantation for children and adolescents with Philadelphia-positive acute lymphoblastic leukemia. J Clin Oncol 16:931, Szydlo R, Goldman JM, Klein JP, Gale RP, Ash RC, Bach FH, Bradley BA, Casper JT, Flomenberg N, Gajewski JL, Gluckman E, Henslee-Downey PJ, Hows JM, Jacobsen N, Kolb HJ, Lowenberg B, Masaoka T, Rowlings PA, Sondel PM, van Bekkum DW, van Rood JJ, Vowels MR, Zhang MJ, Horowitz MM: Results of allogeneic bone marrow transplants for leukemia using donors other than HLA-identical siblings. J Clin Oncol 15:1767, Anasetti C, Beatty PG, Storb R, Martin PJ, Mori M, Sanders JE, Thomas ED, Hansen JA: Effect of HLA incompatibility on graft-versushost disease, relapse, and survival after marrow transplantation for patients with leukemia or lymphoma. Hum Immunol 29:79, Ochs L, Shu XO, Miller J, Enright H, Wagner J, Filipovich A, Miller W, Weisdorf D: Late infections after allogeneic bone marrow transplantations: comparison of incidence in related and unrelated donor transplant recipients. Blood 86:3979, Hansen TH, Carreno BM, Sachs DH: The major histocompatibility complex. In: WE Paul (ed) Fundamental Immunology. New York: Raven Press, 577, Prasad VK, Kernan NA, Heller G, O Reilly RJ, Yang SY: DNA typing for HLA-A and -B identifies disparities between patients and unrelated donors matched by HLA-A and -B serology and DRB1. Blood 90:562a, [abstr] 16 Falk CS, Schendel DJ: HLA-C revisited. Ten years of change. Immunol Res 16:203, Grundschober C, Rufer N, Sanchez-Mazas A, Madrigal A, Jeannet M, Roosnek E, Tiercy JM: Molecular characterization of HLA-C incompatibilities in HLA-ABDR-matched unrelated bone marrow donor-recipient pairs. Sequence of two new Cw alleles (Cw*02023 and Cw*0707) and recognition by cytotoxic T lymphocytes. Tissue Antigens 49:612, Petersdorf EW, Stanley JF, Martin PJ, Hansen JA: Molecular diversity of the HLA-C locus in unrelated marrow transplantation. Tissue Antigens 44:93, Nagler A, Brautbar C, Slavin S, Bishara A: Bone marrow transplantation using unrelated and family related donors: the impact of HLA-C disparity. Bone Marrow Transplant 18:891, Bishara A, Amar A, Brautbar C, Condiotti R, Lazarovitz V, Nagler A: The putative role of HLA-C recognition in graft versus host disease (GVHD) and graft rejection after unrelated bone marrow transplantation (BMT). Exp Hematol 23:1667, Barnardo MC, Davey NJ, Bunce M, Brookes PA, Lechler RI, Welsh KI, Batchelor JR: A correlation between HLA-C matching and donor antirecipient CTL precursor frequency in bone marrow transplantation. Transplantation 61:1420, Petersdorf EW, Longton GM, Anasetti C, Mickelson EM, McKinney SK, Smith AG, Martin PJ, Hansen JA: Association of HLA-C disparity with graft failure after marrow transplantation from unrelated donors. Blood 89:1818, Sasazuki T, Juji T, Morishima Y, Kinukawa N, Kashiwabara H, Inoko H, Yoshida T, Kimura A, Akaza T, Kamikawaji N, Kodera Y, Takaku F, Nose Y, Ono T, Sakamaki T, Kato S, Akiyama Y, Okamoto S, Dohy H, Harada M, Asano S: Effect of matching of class I HLA alleles on clinical outcome after transplantation of hematopoietic stem cells from an unrelated donor. N Engl J Med 339:1177, Bodmer JG, Marsh SG, Albert ED, Bodmer WF, Bontrop RE, Charron D, Dupont B, Erlich HA, Fauchet R, Mach B, Mayr WR, Perham P, Sasazuki T, Schreuder GMT, Strominger JL, Svejgaard A, Terasaki P: Nomenclature for factors of the HLA system, Hum Immunol 53:98, Imanishi T, Akaza T, Kimura A, Tokunaga K, Gojobori T: Allele and haplotype frequencies of HLA and complement loci in various ethnic groups. Eleventh Histocompatibility Workshop and Conference. Yokohoma, Japan, 1991, p Bunce M, Welsh KI: Rapid DNA typing for HLA-C using sequence-specific primers (PCR-SSP): identification of serological and non-serologically defined HLA-C alleles including several new alleles. Tissue Antigens 43:7, Kennedy LJ, Poulton KV, Dyer PA, Ollier WE, Thomson W: Definition of HLA-C alleles using sequence-specific oligonucleotide probes (PCR-SSOP). Tissue Antigens 46:187, Prasad VK, Yang SY: Allele assignment for HLA-A, -B, and -C genes to the Tenth International Histocompatibility Workshop cell 84
9 yp g lines. Tissue Antigens 47:538, Prasad VK, Kernan NA, Heller G, O Reilly RJ, Yang SY: DNA typing for HLA-A and -B identifies disparities between patients and unrelated donors matched by HLA-A and -B serology and DRB1. Blood 93:399, Bodmer JG, Marsh SG, Albert ED, Bodmer WF, Bontrop RE, Charron D, Dupont B, Erlich HA, Mach B, Mayr WR: Nomenclature for factors of the HLA system, Tissue Antigens 46:1, Cereb N, Yang SY: Dimorphic primers derived from intron 1 for use in the molecular typing of HLA-B alleles. Tissue Antigens 50:74, Ng J, Hurley CK, Carter C, Baxter-Lowe LA, Bing D, Chopek M, Hegland J, Lee TD, Li TC, Hsu S, KuKuruga D, Mason JM, Monos D, Noreen H, Rosner G, Schmeckpeper B, Dupont B, Hartzman RJ: Largescale DRB and DQB1 oligonucleotide typing for the NMDP registry: progress report from year 2. Tissue Antigens 47:21, Bunce M, Barnardo MC, Procter J, Marsh SG, Vilches C, Welsh KI: High resolution HLA-C typing by PCR-SSP: identification of allelic frequencies and linkage disequilibria in 604 unrelated random UK Caucasoids and a comparison with serology. Tissue Antigens 50:100, [Corrected and republished article printed in Tissue Antigens 48:680, 1996.] 34 Schlesiger BC, Laumbacher B, Wank R: Sequence based typing (SBT) of HLA-C alleles in 214 Caucasians using the automated sequencer ALF. 12th International Histocompatibility Workshop and Conference, Paris, 1997, p Chapman JR, Taylor C, Ting A, Morris PJ: Hyperacute rejection of a renal allograft in the presence of anti-hla-cw5 antibody. Transplantation 42:91, Prasad VK, Heller G, Kernan NA, O Reilly RJ, Yang SY: The probability of HLA-C matching between patient and unrelated donor at the molecular level: estimations based on the linkage disequilibrium between DNA typed HLA-B and HLA-C alleles. Transplantation, In press. 37 Dupont B, Yang SY: Histocompatibility. In: SJ Forman, KG Blume, ED Thomas (eds): Bone Marrow Transplantation, 1st edition. Cambridge, MA, Blackwell Scientific Publications, 1993, p Bottino C, Vitale M, Pende D, Biassoni R, Moretta A: Receptors for HLA class I molecules in human NK cells. Semin Immunol 7:67, Colonna M, Borsellino G, Falco M, Ferrara GB, Strominger JL: HLA- C is the inhibitory ligand that determines dominant resistance to lysis by NK1- and NK2-specific natural killer cells. Proc Natl Acad Sci U S A 90:12000, Falk CS, Steinle A, Schendel DJ: Expression of HLA-C molecules confers target cell resistance to some non-major histocompatibility complex-restricted T cells in a manner analogous to allospecific natural killer cells. J Exp Med 182:1005, Lanier LL, Phillips JH: NK cell recognition of major histocompatibility complex class I molecules. Semin Immunol 7:75, Mandelboim O, Reyburn HT, Vales-Gomez M, Pazmany L, Colonna M, Borsellino G, Strominger JL: Protection from lysis by natural killer cells of group 1 and 2 specificity is mediated by residue 80 in human histocompatibility leukocyte antigen C alleles and also occurs with empty major histocompatibility complex molecules. J Exp Med 184:913, Mandelboim O, Reyburn HT, Sheu EG, Vales-Gomez M, Davis DM, Pazmany L, Strominger JL: The binding site of NK receptors on HLA- C molecules. Immunity 6:341, Kim J, Chwae YJ, Kim MY, Choi IH, Park JH, Kim SJ: Molecular basis of HLA-C recognition by p58 natural killer cell inhibitory receptors. J Immunol 159:3875, Ferrara JL, Guillen FJ, van Dijken PJ, Marion A, Murphy GF, Burakoff SJ: Evidence that large granular lymphocytes of donor origin mediate acute graft-versus-host disease. Transplantation 47:50, Xun C, Brown SA, Jennings CD, Henslee-Downey PJ, Thompson JS: Acute graft-versus-host-like disease induced by transplantation of human activated natural killer cells into SCID mice. Transplantation 56:409, BB&MT 85
National Marrow Donor Program HLA-Matching Guidelines for Unrelated Marrow Transplants
 Biology of Blood and Marrow Transplantation 9:610-615 (2003) 2003 American Society for Blood and Marrow Transplantation 1083-8791/03/0910-0003$30.00/0 doi:10.1016/s1083-8791(03)00329-x National Marrow
Biology of Blood and Marrow Transplantation 9:610-615 (2003) 2003 American Society for Blood and Marrow Transplantation 1083-8791/03/0910-0003$30.00/0 doi:10.1016/s1083-8791(03)00329-x National Marrow
Plenary paper. Introduction
 Plenary paper Impact of HLA class I and class II high-resolution matching on outcomes of unrelated donor bone marrow transplantation: HLA-C mismatching is associated with a strong adverse effect on transplantation
Plenary paper Impact of HLA class I and class II high-resolution matching on outcomes of unrelated donor bone marrow transplantation: HLA-C mismatching is associated with a strong adverse effect on transplantation
Histocompatibility Evaluations for HSCT at JHMI. M. Sue Leffell, PhD. Professor of Medicine Laboratory Director
 Histocompatibility Evaluations for HSCT at JHMI M. Sue Leffell, PhD Professor of Medicine Laboratory Director JHMI Patient Population Adults Peds NMDP data >20,000 HSCT JHMI HSCT Protocols Bone marrow
Histocompatibility Evaluations for HSCT at JHMI M. Sue Leffell, PhD Professor of Medicine Laboratory Director JHMI Patient Population Adults Peds NMDP data >20,000 HSCT JHMI HSCT Protocols Bone marrow
Haplo vs Cord vs URD Debate
 3rd Annual ASBMT Regional Conference for NPs, PAs and Fellows Haplo vs Cord vs URD Debate Claudio G. Brunstein Associate Professor University of Minnesota Medical School Take home message Finding a donor
3rd Annual ASBMT Regional Conference for NPs, PAs and Fellows Haplo vs Cord vs URD Debate Claudio G. Brunstein Associate Professor University of Minnesota Medical School Take home message Finding a donor
Allogeneic hematopoietic stem cell transplantation from family members other than HLA-identical siblings over the last decade ( )
 TRANSPLANTATION Allogeneic hematopoietic stem cell transplantation from family members other than -identical siblings over the last decade (1991-2000) Yoshinobu Kanda, Shigeru Chiba, Hisamaru Hirai, Hisashi
TRANSPLANTATION Allogeneic hematopoietic stem cell transplantation from family members other than -identical siblings over the last decade (1991-2000) Yoshinobu Kanda, Shigeru Chiba, Hisamaru Hirai, Hisashi
The Human Major Histocompatibility Complex
 The Human Major Histocompatibility Complex 1 Location and Organization of the HLA Complex on Chromosome 6 NEJM 343(10):702-9 2 Inheritance of the HLA Complex Haplotype Inheritance (Family Study) 3 Structure
The Human Major Histocompatibility Complex 1 Location and Organization of the HLA Complex on Chromosome 6 NEJM 343(10):702-9 2 Inheritance of the HLA Complex Haplotype Inheritance (Family Study) 3 Structure
EBMT2008_1_21:EBMT :06 Pagina 46 * CHAPTER 3. Immunogenetics of allogeneic HSCT * 3.1. The role of HLA in HSCT. J.M.
 EBMT2008_1_21:EBMT2008 6-11-2008 9:06 Pagina 46 * CHAPTER 3 Immunogenetics of allogeneic HSCT * 3.1 The role of HLA in HSCT J.M. Tiercy EBMT2008_1_21:EBMT2008 6-11-2008 9:06 Pagina 47 CHAPTER 3.1 HLA and
EBMT2008_1_21:EBMT2008 6-11-2008 9:06 Pagina 46 * CHAPTER 3 Immunogenetics of allogeneic HSCT * 3.1 The role of HLA in HSCT J.M. Tiercy EBMT2008_1_21:EBMT2008 6-11-2008 9:06 Pagina 47 CHAPTER 3.1 HLA and
Basel - 6 September J.-M. Tiercy National Reference Laboratory for Histocompatibility (LNRH) University Hospital Geneva
 Basel - 6 eptember 2012 J.-M. Tiercy National Reference Laboratory for Histocompatibility (LNRH) University Hospital Geneva Outline the HLA system is (a) complex anti-hla immunisation and alloreactivity
Basel - 6 eptember 2012 J.-M. Tiercy National Reference Laboratory for Histocompatibility (LNRH) University Hospital Geneva Outline the HLA system is (a) complex anti-hla immunisation and alloreactivity
Role of NMDP Repository in the Evolution of HLA Matching and Typing for Unrelated Donor HCT
 Role of NMDP Repository in the Evolution of HLA Matching and Typing for Unrelated Donor HCT Stephen Spellman, MBS Director, Immunobiology and Observational Research Assistant Scientific Director CIBMTR,
Role of NMDP Repository in the Evolution of HLA Matching and Typing for Unrelated Donor HCT Stephen Spellman, MBS Director, Immunobiology and Observational Research Assistant Scientific Director CIBMTR,
Cover Page. The handle holds various files of this Leiden University dissertation.
 Cover Page The handle http://hdl.handle.net/1887/20898 holds various files of this Leiden University dissertation. Author: Jöris, Monique Maria Title: Challenges in unrelated hematopoietic stem cell transplantation.
Cover Page The handle http://hdl.handle.net/1887/20898 holds various files of this Leiden University dissertation. Author: Jöris, Monique Maria Title: Challenges in unrelated hematopoietic stem cell transplantation.
The impact of HLA matching on unrelated donor hematopoietic stem cell transplantation in Korean children
 VOLUME 46 ㆍ NUMBER ㆍ March 0 THE KOREAN JOURNAL OF HEMATOLOGY ORIGINAL ARTICLE The impact of HLA matching on unrelated donor hematopoietic stem cell transplantation in Korean children Meerim Park, Kyung
VOLUME 46 ㆍ NUMBER ㆍ March 0 THE KOREAN JOURNAL OF HEMATOLOGY ORIGINAL ARTICLE The impact of HLA matching on unrelated donor hematopoietic stem cell transplantation in Korean children Meerim Park, Kyung
Indian Journal of Nephrology Indian J Nephrol 2001;11: 88-97
 88 Indian Journal of Nephrology Indian J Nephrol 2001;11: 88-97 ARTICLE HLA gene and haplotype frequency in renal transplant recipients and donors of Uttar Pradesh (North India) S Agrawal, AK Singh, RK
88 Indian Journal of Nephrology Indian J Nephrol 2001;11: 88-97 ARTICLE HLA gene and haplotype frequency in renal transplant recipients and donors of Uttar Pradesh (North India) S Agrawal, AK Singh, RK
Completing the CIBMTR Confirmation of HLA Typing Form (Form 2005)
 Completing the CIBMTR Confirmation of HLA Typing Form (Form 2005) Stephen Spellman Research Manager NMDP Scientific Services Maria Brown Scientific Services Specialist Data Management Conference 2007 1
Completing the CIBMTR Confirmation of HLA Typing Form (Form 2005) Stephen Spellman Research Manager NMDP Scientific Services Maria Brown Scientific Services Specialist Data Management Conference 2007 1
HHS Public Access Author manuscript Bone Marrow Transplant. Author manuscript; available in PMC 2015 July 01.
 KIR and HLA genotypes have no identifiable role in single unit dominance following double unit umbilical cord blood transplantation Nidale Tarek 1,6, Meighan M. Gallagher 2,7, Joanne F. Chou 3, Marissa
KIR and HLA genotypes have no identifiable role in single unit dominance following double unit umbilical cord blood transplantation Nidale Tarek 1,6, Meighan M. Gallagher 2,7, Joanne F. Chou 3, Marissa
Allele and Haplotype Frequencies of Human Leukocyte Antigen-A, -B, -C, -DRB1, and -DQB1 From Sequence- Based DNA Typing Data in Koreans
 Original Article Diagnostic Immunology Ann Lab Med 2015;35:429-435 http://dx.doi.org/10.3343/alm.2015.35.4.429 ISSN 2234-3806 eissn 2234-3814 Allele and Haplotype Frequencies of Human Leukocyte Antigen-A,
Original Article Diagnostic Immunology Ann Lab Med 2015;35:429-435 http://dx.doi.org/10.3343/alm.2015.35.4.429 ISSN 2234-3806 eissn 2234-3814 Allele and Haplotype Frequencies of Human Leukocyte Antigen-A,
The New England Journal of Medicine BONE MARROW TRANSPLANTS FROM UNRELATED DONORS FOR PATIENTS WITH CHRONIC MYELOID LEUKEMIA
 BONE MARROW TRANSPLANTS FROM UNRELATED DONORS FOR PATIENTS WITH CHRONIC MYELOID LEUKEMIA JOHN A. HANSEN, M.D., THEODORE A. GOOLEY, PH.D., PAUL J. MARTIN, M.D., FREDERICK APPELBAUM, M.D., THOMAS R. CHAUNCEY,
BONE MARROW TRANSPLANTS FROM UNRELATED DONORS FOR PATIENTS WITH CHRONIC MYELOID LEUKEMIA JOHN A. HANSEN, M.D., THEODORE A. GOOLEY, PH.D., PAUL J. MARTIN, M.D., FREDERICK APPELBAUM, M.D., THOMAS R. CHAUNCEY,
Unrelated Donor Hematopoietic Cell Transplantation: Factors Associated with a Better HLA Match
 CLINICAL RESEARCH Unrelated Donor Hematopoietic Cell Transplantation: Factors Associated with a Better HLA Match Jason Dehn, 1 Mukta Arora, 2 Stephen Spellman, 1 Michelle Setterholm, 1 Mary Horowitz, 3
CLINICAL RESEARCH Unrelated Donor Hematopoietic Cell Transplantation: Factors Associated with a Better HLA Match Jason Dehn, 1 Mukta Arora, 2 Stephen Spellman, 1 Michelle Setterholm, 1 Mary Horowitz, 3
The National Marrow Donor Program. Graft Sources for Hematopoietic Cell Transplantation. Simon Bostic, URD Transplant Recipient
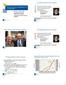 1988 199 1992 1994 1996 1998 2 22 24 26 28 21 212 214 216 218 Adult Donors Cord Blood Units The National Donor Program Graft Sources for Hematopoietic Cell Transplantation Dennis L. Confer, MD Chief Medical
1988 199 1992 1994 1996 1998 2 22 24 26 28 21 212 214 216 218 Adult Donors Cord Blood Units The National Donor Program Graft Sources for Hematopoietic Cell Transplantation Dennis L. Confer, MD Chief Medical
IMMUNOLOGY. Elementary Knowledge of Major Histocompatibility Complex and HLA Typing
 IMMUNOLOGY Elementary Knowledge of Major Histocompatibility Complex and HLA Typing Tapasya Srivastava and Subrata Sinha Department of Biochemistry All India Institute of Medical Sciences New Delhi - 110029
IMMUNOLOGY Elementary Knowledge of Major Histocompatibility Complex and HLA Typing Tapasya Srivastava and Subrata Sinha Department of Biochemistry All India Institute of Medical Sciences New Delhi - 110029
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Gragert L, Eapen M, Williams E, et al. HLA match likelihoods
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Gragert L, Eapen M, Williams E, et al. HLA match likelihoods
HLA-DR-matched Parental Donors for Allogeneic Hematopoietic Stem Cell Transplantation in Patients with High-risk Acute Leukemia
 BRIEF COMMUNICATION HLA-DR-matched Parental Donors for Allogeneic Hematopoietic Stem Cell Transplantation in Patients with High-risk Acute Leukemia Shang-Ju Wu, Ming Yao,* Jih-Luh Tang, Bo-Sheng Ko, Hwei-Fang
BRIEF COMMUNICATION HLA-DR-matched Parental Donors for Allogeneic Hematopoietic Stem Cell Transplantation in Patients with High-risk Acute Leukemia Shang-Ju Wu, Ming Yao,* Jih-Luh Tang, Bo-Sheng Ko, Hwei-Fang
Effect of HLA mismatch on acute graft-versus-host disease
 Int J Hematol (2013) 98:300 308 DOI 10.1007/s12185-013-1405-x PROGRESS IN HEMATOLOGY New clinical and basic aspects of graft-versus-host disease Effect of HLA mismatch on acute graft-versus-host disease
Int J Hematol (2013) 98:300 308 DOI 10.1007/s12185-013-1405-x PROGRESS IN HEMATOLOGY New clinical and basic aspects of graft-versus-host disease Effect of HLA mismatch on acute graft-versus-host disease
Documentation of Changes to EFI Standards: v 5.6.1
 Modified Standard B - PERSONNEL QUALIFICATIONS B1.000 The laboratory must employ one or more individuals who meet the qualifications and fulfil the responsibilities of the Director/Co-Director, Technical
Modified Standard B - PERSONNEL QUALIFICATIONS B1.000 The laboratory must employ one or more individuals who meet the qualifications and fulfil the responsibilities of the Director/Co-Director, Technical
HLA and new technologies. Vicky Van Sandt
 HLA and new technologies. Vicky Van Sandt Life-threatning malignant and non malignant blood disorders can be cured by hematopoetic stem cell transplantation (HSCT). GVHD is the 2nd most prevalent cause
HLA and new technologies. Vicky Van Sandt Life-threatning malignant and non malignant blood disorders can be cured by hematopoetic stem cell transplantation (HSCT). GVHD is the 2nd most prevalent cause
CHAPTER 3 LABORATORY PROCEDURES
 CHAPTER 3 LABORATORY PROCEDURES CHAPTER 3 LABORATORY PROCEDURES 3.1 HLA TYPING Molecular HLA typing will be performed for all donor cord blood units and patients in the three reference laboratories identified
CHAPTER 3 LABORATORY PROCEDURES CHAPTER 3 LABORATORY PROCEDURES 3.1 HLA TYPING Molecular HLA typing will be performed for all donor cord blood units and patients in the three reference laboratories identified
MATCHMAKER, MATCHMAKER, MAKE ME A MATCH, FIND ME A MISMATCHED TRANSPLANT TO CATCH
 MATCHMAKER, MATCHMAKER, MAKE ME A MATCH, FIND ME A MISMATCHED TRANSPLANT TO CATCH TEJASWINI M. DHAWALE, M.D. HEME FELLOWS CONFERENCE NOVEMBER 08, 2013 CASE PRESENTATION 51 yo M with history of MDS (unilinear
MATCHMAKER, MATCHMAKER, MAKE ME A MATCH, FIND ME A MISMATCHED TRANSPLANT TO CATCH TEJASWINI M. DHAWALE, M.D. HEME FELLOWS CONFERENCE NOVEMBER 08, 2013 CASE PRESENTATION 51 yo M with history of MDS (unilinear
21/05/2018. Continuing Education. Presentation Recording. learn.immucor.com
 Transplant Webinar Series: Ep. 6 Donor Selection for Haematopoietic Stem Cell Transplantation Future Webinars The Role of NGS in the Transplant Setting Featuring Dr Sujatha Krishnakumar Sirona Genomics,
Transplant Webinar Series: Ep. 6 Donor Selection for Haematopoietic Stem Cell Transplantation Future Webinars The Role of NGS in the Transplant Setting Featuring Dr Sujatha Krishnakumar Sirona Genomics,
DEFINITIONS OF HISTOCOMPATIBILITY TYPING TERMS
 DEFINITIONS OF HISTOCOMPATIBILITY TYPING TERMS The definitions below are intended as general concepts. There will be exceptions to these general definitions. These definitions do not imply any specific
DEFINITIONS OF HISTOCOMPATIBILITY TYPING TERMS The definitions below are intended as general concepts. There will be exceptions to these general definitions. These definitions do not imply any specific
Outcomes of pediatric bone marrow transplantation for leukemia and myelodysplasia using matched. unrelated donors
 Outcomes of pediatric bone marrow transplantation for leukemia and myelodysplasia using matched sibling, mismatched related or matched unrelated donors Immunobiology Working Committee PIs: Peter Shaw and
Outcomes of pediatric bone marrow transplantation for leukemia and myelodysplasia using matched sibling, mismatched related or matched unrelated donors Immunobiology Working Committee PIs: Peter Shaw and
Dr. Yi-chi M. Kong August 8, 2001 Benjamini. Ch. 19, Pgs Page 1 of 10 TRANSPLANTATION
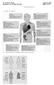 Benjamini. Ch. 19, Pgs 379-399 Page 1 of 10 TRANSPLANTATION I. KINDS OF GRAFTS II. RELATIONSHIPS BETWEEN DONOR AND RECIPIENT Benjamini. Ch. 19, Pgs 379-399 Page 2 of 10 II.GRAFT REJECTION IS IMMUNOLOGIC
Benjamini. Ch. 19, Pgs 379-399 Page 1 of 10 TRANSPLANTATION I. KINDS OF GRAFTS II. RELATIONSHIPS BETWEEN DONOR AND RECIPIENT Benjamini. Ch. 19, Pgs 379-399 Page 2 of 10 II.GRAFT REJECTION IS IMMUNOLOGIC
BE THE MATCH. The Role of HLA in Finding a Match for Bone Marrow or Peripheral Blood Stem Cell Transplantation
 BE THE MATCH The Role of HLA in Finding a Match for Bone Marrow or Peripheral Blood Stem Cell Transplantation Leukemia Leukemia is a type of cancer of the blood or bone marrow. It is characterized by an
BE THE MATCH The Role of HLA in Finding a Match for Bone Marrow or Peripheral Blood Stem Cell Transplantation Leukemia Leukemia is a type of cancer of the blood or bone marrow. It is characterized by an
New trends in donor selection in Europe: "best match" versus haploidentical. Prof Jakob R Passweg
 New trends in donor selection in Europe: "best match" versus haploidentical Prof Jakob R Passweg HSCT change in donor type: 1990-2015 9000 H S C T 8000 7000 6000 5000 4000 HLA identical sibling/twin Haplo-identical
New trends in donor selection in Europe: "best match" versus haploidentical Prof Jakob R Passweg HSCT change in donor type: 1990-2015 9000 H S C T 8000 7000 6000 5000 4000 HLA identical sibling/twin Haplo-identical
Human Leukocyte Antigens and donor selection
 Human Leukocyte Antigens and donor selection Duangtawan Thammanichanond, MD. PhD. Histocompatibility and Immunogenetics Laboratory, Faculty of Medicine Ramathibodi Hospital, Mahidol University Outline
Human Leukocyte Antigens and donor selection Duangtawan Thammanichanond, MD. PhD. Histocompatibility and Immunogenetics Laboratory, Faculty of Medicine Ramathibodi Hospital, Mahidol University Outline
HLA-A*26 and Susceptibility of Iranian Patients with Non-Hodgkin Lymphoma
 HLA-A*26 and Susceptibility of Iranian Patients with Non-Hodgkin Lymphoma Arezou Sayad 1, Mohammad Taghi Akbari 2**, Mahshid Mehdizadeh 3,4, Mohammad Taheri 1, Abbas Hajifathali 3* 1 Department of Medical
HLA-A*26 and Susceptibility of Iranian Patients with Non-Hodgkin Lymphoma Arezou Sayad 1, Mohammad Taghi Akbari 2**, Mahshid Mehdizadeh 3,4, Mohammad Taheri 1, Abbas Hajifathali 3* 1 Department of Medical
ASSESSMENT OF THE RISK FOR TYPE 1 DIABETES MELLITUS CONFERRED BY HLA CLASS II GENES. Irina Durbală
 ASSESSMENT OF THE RISK FOR TYPE 1 DIABETES MELLITUS CONFERRED BY HLA CLASS II GENES Summary Irina Durbală CELL AND MOLECULAR BIOLOGY DEPARTMENT FACULTY OF MEDICINE, OVIDIUS UNIVERSITY CONSTANŢA Class II
ASSESSMENT OF THE RISK FOR TYPE 1 DIABETES MELLITUS CONFERRED BY HLA CLASS II GENES Summary Irina Durbală CELL AND MOLECULAR BIOLOGY DEPARTMENT FACULTY OF MEDICINE, OVIDIUS UNIVERSITY CONSTANŢA Class II
HLA Mismatches. Professor Steven GE Marsh. Anthony Nolan Research Institute EBMT Anthony Nolan Research Institute
 HLA Mismatches Professor Steven GE Marsh HLA Mismatches HLA Genes, Structure, Polymorphism HLA Nomenclature HLA Mismatches in HSCT Defining a mismatch HLA Mismatches HLA Genes, Structure, Polymorphism
HLA Mismatches Professor Steven GE Marsh HLA Mismatches HLA Genes, Structure, Polymorphism HLA Nomenclature HLA Mismatches in HSCT Defining a mismatch HLA Mismatches HLA Genes, Structure, Polymorphism
Risk assessment in haematopoietic stem cell transplantation: Histocompatibility
 Best Practice & Research Clinical Haematology Vol. 20, No. 2, pp. 155e170, 2007 doi:10.1016/j.beha.2006.09.001 available online at http://www.sciencedirect.com 2 Risk assessment in haematopoietic stem
Best Practice & Research Clinical Haematology Vol. 20, No. 2, pp. 155e170, 2007 doi:10.1016/j.beha.2006.09.001 available online at http://www.sciencedirect.com 2 Risk assessment in haematopoietic stem
HLA Amino Acid Polymorphisms and Kidney Allograft Survival. Supplemental Digital Content
 HLA Amino Acid Polymorphisms and Kidney Allograft Survival Supplemental Digital Content Malek Kamoun, MD, PhD a, Keith P. McCullough, MS b, Martin Maiers, MS c, Marcelo A. Fernandez Vina, MD, PhD d, Hongzhe
HLA Amino Acid Polymorphisms and Kidney Allograft Survival Supplemental Digital Content Malek Kamoun, MD, PhD a, Keith P. McCullough, MS b, Martin Maiers, MS c, Marcelo A. Fernandez Vina, MD, PhD d, Hongzhe
Biol Blood Marrow Transplant 17: (2011) Ó 2011 American Society for Blood and Marrow Transplantation
 Outcomes of Patients with Myeloid Malignancies Treated with Allogeneic Hematopoietic Stem Cell Transplantation from Matched Unrelated Donors Compared with One Human Leukocyte Antigen Mismatched Related
Outcomes of Patients with Myeloid Malignancies Treated with Allogeneic Hematopoietic Stem Cell Transplantation from Matched Unrelated Donors Compared with One Human Leukocyte Antigen Mismatched Related
Unrelated donor transplants DPB1 disparities contribute to severe GVHD and reduced patient survival after unrelated donor bone marrow transplantation
 (2002) 30, 497 502 2002 Nature Publishing Group All rights reserved 0268 3369/02 $25.00 www.nature.com/bmt Unrelated donor transplants DPB1 disparities contribute to severe GVHD and reduced patient survival
(2002) 30, 497 502 2002 Nature Publishing Group All rights reserved 0268 3369/02 $25.00 www.nature.com/bmt Unrelated donor transplants DPB1 disparities contribute to severe GVHD and reduced patient survival
The role of HLA in Allogeneic Hematopoietic Stem Cell Transplantation and Platelet Refractoriness.
 The role of HLA in Allogeneic Hematopoietic Stem Cell Transplantation and Platelet Refractoriness. Robert Liwski, MD, PhD, FRCPC Medical Director HLA Typing Laboratory Department of Pathology Dalhousie
The role of HLA in Allogeneic Hematopoietic Stem Cell Transplantation and Platelet Refractoriness. Robert Liwski, MD, PhD, FRCPC Medical Director HLA Typing Laboratory Department of Pathology Dalhousie
The Relationship between Minor Histocompatibility Antigens and Graft Versus Host Disease in Unrelated Peripheral Blood Stem Cell Transplants
 mhags and GVHD Page 1 The Relationship between Minor Histocompatibility Antigens and Graft Versus Host Disease in Unrelated Peripheral Blood Stem Cell Transplants Running Title: mhags and GVHD Conflict
mhags and GVHD Page 1 The Relationship between Minor Histocompatibility Antigens and Graft Versus Host Disease in Unrelated Peripheral Blood Stem Cell Transplants Running Title: mhags and GVHD Conflict
Hugo Castro-Malaspina, Richard E. Harris, James Gajewski, Norma Ramsay, Robert Collins, Bernie Dharan, Roberta King, and H.
 CLINICAL OBSERVATIONS, INTERVENTIONS, AND THERAPEUTIC TRIALS Unrelated donor marrow transplantation for myelodysplastic syndromes: outcome analysis in 510 transplants facilitated by the National Marrow
CLINICAL OBSERVATIONS, INTERVENTIONS, AND THERAPEUTIC TRIALS Unrelated donor marrow transplantation for myelodysplastic syndromes: outcome analysis in 510 transplants facilitated by the National Marrow
Clinical outcomes of HLA- DPB1 mismatches in 10/10 HLA- matched unrelated donor- recipient pairs undergoing allogeneic stem cell transplant
 Accepted: 13 June 2017 DOI: 10.1111/ejh.12916 ORIGINAL ARTICLE Clinical outcomes of HLA- DPB1 mismatches in 10/10 HLA- matched unrelated donor- recipient pairs undergoing allogeneic stem cell transplant
Accepted: 13 June 2017 DOI: 10.1111/ejh.12916 ORIGINAL ARTICLE Clinical outcomes of HLA- DPB1 mismatches in 10/10 HLA- matched unrelated donor- recipient pairs undergoing allogeneic stem cell transplant
Curriculum Vitae. Office Address: Histogenetics Tel.#: (914) Executive Blvd. histogenetics.com Ossining, NY 10562
 Curriculum Vitae Name: Nezih Cereb Office Address: Histogenetics Tel.#: (914) 762-030 300 Executive Blvd E-mail: nezih@ histogenetics.com Ossining, NY 10562 Fax#: (914) 762-4441 Cerificates Licensed Physician:
Curriculum Vitae Name: Nezih Cereb Office Address: Histogenetics Tel.#: (914) 762-030 300 Executive Blvd E-mail: nezih@ histogenetics.com Ossining, NY 10562 Fax#: (914) 762-4441 Cerificates Licensed Physician:
HLA AND KIR GENE POLYMORPHISM IN HEMATOPOIETIC STEM CELL TRANSPLANTATION
 FROM THE DEPARTMENT OF LABORATORY MEDICINE, DIVISION OF CLINICAL IMMUNOLOGY KAROLINSKA INSTITUTET, STOCKHOLM, SWEDEN HLA AND KIR GENE POLYMORPHISM IN HEMATOPOIETIC STEM CELL TRANSPLANTATION Marie Schaffer
FROM THE DEPARTMENT OF LABORATORY MEDICINE, DIVISION OF CLINICAL IMMUNOLOGY KAROLINSKA INSTITUTET, STOCKHOLM, SWEDEN HLA AND KIR GENE POLYMORPHISM IN HEMATOPOIETIC STEM CELL TRANSPLANTATION Marie Schaffer
Problems and possible solutions in finding an unrelated bone marrow donor. Results of consecutive searches for 240 Dutch patients
 Bone Marrow Transplantation, (1997) 20, 1011 1017 1997 Stockton Press All rights reserved 0268 3369/97 $12.00 Problems and possible solutions in finding an unrelated bone marrow donor. Results of consecutive
Bone Marrow Transplantation, (1997) 20, 1011 1017 1997 Stockton Press All rights reserved 0268 3369/97 $12.00 Problems and possible solutions in finding an unrelated bone marrow donor. Results of consecutive
Support of Unrelated Stem Cell Donor Searches by Donor Center-Initiated HLA Typing of Potentially Matching Donors
 Support of Unrelated Stem Cell Donor Searches by Donor Center-Initiated HLA Typing of Potentially Matching Donors Alexander H. Schmidt 1 *, Ute V. Solloch 1, Daniel Baier 1, Alois Grathwohl 1, Jan Hofmann
Support of Unrelated Stem Cell Donor Searches by Donor Center-Initiated HLA Typing of Potentially Matching Donors Alexander H. Schmidt 1 *, Ute V. Solloch 1, Daniel Baier 1, Alois Grathwohl 1, Jan Hofmann
journal of medicine The new england Outcomes after Transplantation of Cord Blood or Bone Marrow from Unrelated Donors in Adults with Leukemia abstract
 The new england journal of medicine established in 1812 november 25, 2004 vol. 351 no. 22 Outcomes after Transplantation of Cord Blood or Bone Marrow from Unrelated Donors in Adults with Leukemia Mary
The new england journal of medicine established in 1812 november 25, 2004 vol. 351 no. 22 Outcomes after Transplantation of Cord Blood or Bone Marrow from Unrelated Donors in Adults with Leukemia Mary
Review Article Role of HLA in Hematopoietic Stem Cell Transplantation
 Bone Marrow Research Volume 2012, Article ID 680841, 7 pages doi:10.1155/2012/680841 Review Article Role of HLA in Hematopoietic Stem Cell Transplantation Meerim Park 1 and Jong Jin Seo 2 1 Department
Bone Marrow Research Volume 2012, Article ID 680841, 7 pages doi:10.1155/2012/680841 Review Article Role of HLA in Hematopoietic Stem Cell Transplantation Meerim Park 1 and Jong Jin Seo 2 1 Department
How to Find an Unrelated Donor Theory & Technology
 How to Find an Unrelated Donor Theory & Technology Carlheinz R. Müller Zentrales Knochenmarkspender-Register für die Bundesrepublik Deutschland (ZKRD) Ulm, Germany How to Find an Unrelated Donor HLA-Basics
How to Find an Unrelated Donor Theory & Technology Carlheinz R. Müller Zentrales Knochenmarkspender-Register für die Bundesrepublik Deutschland (ZKRD) Ulm, Germany How to Find an Unrelated Donor HLA-Basics
Stem Cell Transplantation for Severe Aplastic Anemia
 Number of Transplants 10/24/2011 Stem Cell Transplantation for Severe Aplastic Anemia Claudio Anasetti, MD Professor of Oncology and Medicine Chair, Blood and Marrow Transplant Dpt Moffitt Cancer Center
Number of Transplants 10/24/2011 Stem Cell Transplantation for Severe Aplastic Anemia Claudio Anasetti, MD Professor of Oncology and Medicine Chair, Blood and Marrow Transplant Dpt Moffitt Cancer Center
HLA Match Likelihoods for Hematopoietic Stem-Cell Grafts in the U.S. Registry
 The new england journal of medicine special article HLA Likelihoods for Hematopoietic Stem-Cell Grafts in the U.S. Registry Loren Gragert, B.S., B.A., Mary Eapen, M.B., B.S., Eric Williams, Ph.D., John
The new england journal of medicine special article HLA Likelihoods for Hematopoietic Stem-Cell Grafts in the U.S. Registry Loren Gragert, B.S., B.A., Mary Eapen, M.B., B.S., Eric Williams, Ph.D., John
Unrelated Donor StemCell Transplantation: The Role of the National Marrow Donor Program
 Unrelated Donor StemCell Transplantation: The Role of the National Marrow Donor Program August 01, 2003 Palliative and Supportive Care [1], Oncology Journal [2] By Chatchada Karanes, MD [3], Dennis Confer,
Unrelated Donor StemCell Transplantation: The Role of the National Marrow Donor Program August 01, 2003 Palliative and Supportive Care [1], Oncology Journal [2] By Chatchada Karanes, MD [3], Dennis Confer,
25/10/2017. Clinical Relevance of the HLA System in Blood Transfusion. Outline of talk. Major Histocompatibility Complex
 Clinical Relevance of the HLA System in Blood Transfusion Dr Colin J Brown PhD FRCPath. October 2017 Outline of talk HLA genes, structure and function HLA and immune complications of transfusion TA-GVHD
Clinical Relevance of the HLA System in Blood Transfusion Dr Colin J Brown PhD FRCPath. October 2017 Outline of talk HLA genes, structure and function HLA and immune complications of transfusion TA-GVHD
Minimal Requirements for Histocompatibility & Immunogenetics Laboratory
 Minimal Requirements for Histocompatibility & Immunogenetics Laboratory The 4 th WBMT Congress and Workshop Riyadh, KSA - January 15-17, 2017 HLA Discovery, 1958 The Nobel Prize in Physiology or Medicine
Minimal Requirements for Histocompatibility & Immunogenetics Laboratory The 4 th WBMT Congress and Workshop Riyadh, KSA - January 15-17, 2017 HLA Discovery, 1958 The Nobel Prize in Physiology or Medicine
Manipulation of T Cells in the Thnsplant Inoculum
 International Journal of Cell Cloning 4: 122-126 Suppl 1 (1986) Manipulation of T Cells in the Thnsplant Inoculum J. Kersey Bone Marrow Transplantation Program, University of Minnesota, Minneapolis, MN,
International Journal of Cell Cloning 4: 122-126 Suppl 1 (1986) Manipulation of T Cells in the Thnsplant Inoculum J. Kersey Bone Marrow Transplantation Program, University of Minnesota, Minneapolis, MN,
KEY WORDS KIR ligand incompatibility HLA Leukemia Unrelated bone marrow transplantation
 Biology of Blood and Marrow Transplantation 13:315-328 (2007) 2007 American Society for Blood and Marrow Transplantation 1083-8791/07/1303-0001$32.00/0 doi:10.1016/j.bbmt.2006.10.027 Effects of HLA Allele
Biology of Blood and Marrow Transplantation 13:315-328 (2007) 2007 American Society for Blood and Marrow Transplantation 1083-8791/07/1303-0001$32.00/0 doi:10.1016/j.bbmt.2006.10.027 Effects of HLA Allele
KEY WORDS: Allogeneic, Hematopoietic cell transplantation, Graft-versus-host disease, Immunosuppressants, Cyclosporine, Tacrolimus
 A Retrospective Comparison of Tacrolimus versus Cyclosporine with Methotrexate for Immunosuppression after Allogeneic Hematopoietic Cell Transplantation with Mobilized Blood Cells Yoshihiro Inamoto, 1
A Retrospective Comparison of Tacrolimus versus Cyclosporine with Methotrexate for Immunosuppression after Allogeneic Hematopoietic Cell Transplantation with Mobilized Blood Cells Yoshihiro Inamoto, 1
Factors Influencing Haematopoietic Progenitor cell transplant outcome Optimising donor selection
 Factors Influencing Haematopoietic Progenitor cell transplant outcome Optimising donor selection Alison Logan Transplantation Laboratory Manchester Royal Infirmary Haematopoietic progenitor cell transplants
Factors Influencing Haematopoietic Progenitor cell transplant outcome Optimising donor selection Alison Logan Transplantation Laboratory Manchester Royal Infirmary Haematopoietic progenitor cell transplants
Cord Blood Stem Cell Banking and Transplantation
 Cord Blood Stem Cell Banking and Transplantation JOHN W. ADAMSON New York Blood Center, New York, New York, USA Key Words. Cord blood Stem cells Cord blood banking Cord blood transplantation. Cord blood.stern
Cord Blood Stem Cell Banking and Transplantation JOHN W. ADAMSON New York Blood Center, New York, New York, USA Key Words. Cord blood Stem cells Cord blood banking Cord blood transplantation. Cord blood.stern
HLA class II DRB1 and DQB1 allelic polymorphism and sclerosing lymphocytic lobulitis of the breast
 J Clin Pathol 1999;52:445 449 445 Department of Histopathology, Southampton University Hospitals, Southampton, UK A C Bateman J M Theaker Wessex Histocompatibility Laboratory, Southampton University Hospitals
J Clin Pathol 1999;52:445 449 445 Department of Histopathology, Southampton University Hospitals, Southampton, UK A C Bateman J M Theaker Wessex Histocompatibility Laboratory, Southampton University Hospitals
10/18/2012. A primer in HLA: The who, what, how and why. What?
 A primer in HLA: The who, what, how and why What? 1 First recognized in mice during 1930 s and 1940 s. Mouse (murine) experiments with tumors Independent observations were made in humans with leukoagglutinating
A primer in HLA: The who, what, how and why What? 1 First recognized in mice during 1930 s and 1940 s. Mouse (murine) experiments with tumors Independent observations were made in humans with leukoagglutinating
Transplantation. Immunology Unit College of Medicine King Saud University
 Transplantation Immunology Unit College of Medicine King Saud University Objectives To understand the diversity among human leukocyte antigens (HLA) or major histocompatibility complex (MHC) To know the
Transplantation Immunology Unit College of Medicine King Saud University Objectives To understand the diversity among human leukocyte antigens (HLA) or major histocompatibility complex (MHC) To know the
The Use of HLA /HPA Selected Platelets
 The Use of HLA /HPA Selected Platelets Dr Colin Brown PhD FRCPath. Consultant Clinical Scientist Histocompatibility and Immunogenetics HLA and Transfusion Human Leucocyte Antigens HLA Human Platelet Antigens
The Use of HLA /HPA Selected Platelets Dr Colin Brown PhD FRCPath. Consultant Clinical Scientist Histocompatibility and Immunogenetics HLA and Transfusion Human Leucocyte Antigens HLA Human Platelet Antigens
Transplant Booklet D Page 1
 Booklet D Pretest Correct Answers 4. (A) is correct. Technically, performing a hematopoietic stem cell transplant is one of the simplest transplantation procedures. The hematopoietic stem cells are infused
Booklet D Pretest Correct Answers 4. (A) is correct. Technically, performing a hematopoietic stem cell transplant is one of the simplest transplantation procedures. The hematopoietic stem cells are infused
Transplantation with unrelated bone marrow in leukaemic patients above 40 years of age
 Bone Marrow Transplantation, (1998) 21, 43 49 1998 Stockton Press All rights reserved 0268 3369/98 $12.00 Transplantation with unrelated bone marrow in leukaemic patients above 40 years of age O Ringdén
Bone Marrow Transplantation, (1998) 21, 43 49 1998 Stockton Press All rights reserved 0268 3369/98 $12.00 Transplantation with unrelated bone marrow in leukaemic patients above 40 years of age O Ringdén
Significance of the MHC
 CHAPTER 7 Major Histocompatibility Complex (MHC) What is is MHC? HLA H-2 Minor histocompatibility antigens Peter Gorer & George Sneell (1940) Significance of the MHC role in immune response role in organ
CHAPTER 7 Major Histocompatibility Complex (MHC) What is is MHC? HLA H-2 Minor histocompatibility antigens Peter Gorer & George Sneell (1940) Significance of the MHC role in immune response role in organ
Comparable Outcome with T-Cell Depleted Unrelated-Donor versus Related-Donor Allogeneic Bone Marrow Transplantation
 Biology of Blood and Marrow Transplantation 8:601-607 (2002) 2002 American Society for Blood and Marrow Transplantation ASBMT Comparable Outcome with T-Cell Depleted Unrelated-Donor versus Related-Donor
Biology of Blood and Marrow Transplantation 8:601-607 (2002) 2002 American Society for Blood and Marrow Transplantation ASBMT Comparable Outcome with T-Cell Depleted Unrelated-Donor versus Related-Donor
Haploidentical Transplantation today: and the alternatives
 Haploidentical Transplantation today: and the alternatives Daniel Weisdorf MD University of Minnesota February, 2013 No matched sib: where to look? URD donor requires close HLA matching and 3-12 weeks
Haploidentical Transplantation today: and the alternatives Daniel Weisdorf MD University of Minnesota February, 2013 No matched sib: where to look? URD donor requires close HLA matching and 3-12 weeks
Telephone: ; Fax: ; E mail:
 MINUTES AND OVERVIEW PLAN CIBMTR WORKING COMMITTEE FOR GRAFT SOURCES & MANIPULATION Grapevine, TX Thursday, February 27, 2014, 2:45 4:45 pm Co Chair: Co Chair: Co Chair: Statisticians: Scientific Director:
MINUTES AND OVERVIEW PLAN CIBMTR WORKING COMMITTEE FOR GRAFT SOURCES & MANIPULATION Grapevine, TX Thursday, February 27, 2014, 2:45 4:45 pm Co Chair: Co Chair: Co Chair: Statisticians: Scientific Director:
Clinical Relevance of the HLA System in Blood Transfusion. Dr Colin J Brown PhD FRCPath. October 2017
 Clinical Relevance of the HLA System in Blood Transfusion Dr Colin J Brown PhD FRCPath. October 2017 Outline of talk HLA genes, structure and function HLA and immune complications of transfusion TA-GVHD
Clinical Relevance of the HLA System in Blood Transfusion Dr Colin J Brown PhD FRCPath. October 2017 Outline of talk HLA genes, structure and function HLA and immune complications of transfusion TA-GVHD
Samples Available for Recipient Only. Samples Available for Recipient and Donor
 Unrelated HCT Research Sample Inventory - Summary for First Allogeneic Transplants in CRF and TED with biospecimens available through the CIBMTR Repository stratified by availability of paired samples,
Unrelated HCT Research Sample Inventory - Summary for First Allogeneic Transplants in CRF and TED with biospecimens available through the CIBMTR Repository stratified by availability of paired samples,
Sylwia Mizia, 1 Dorota Dera-Joachimiak, 1 Malgorzata Polak, 1 Katarzyna Koscinska, 1 Mariola Sedzimirska, 1 and Andrzej Lange 1, 2. 1.
 Bone Marrow Research Volume 2012, Article ID 873695, 5 pages doi:10.1155/2012/873695 Clinical Study Both Optimal Matching and Procedure Duration Influence Survival of Patients after Unrelated Donor Hematopoietic
Bone Marrow Research Volume 2012, Article ID 873695, 5 pages doi:10.1155/2012/873695 Clinical Study Both Optimal Matching and Procedure Duration Influence Survival of Patients after Unrelated Donor Hematopoietic
Paediatric bone marrow transplantation
 546 Archives ofdise in Childhood 1991; 66: 546-550 CURRENT TOPIC Department of Haematology Royal Postgraduate Medical School, Hammeramith Hospital, London W12 ONN Correspondence to: Dr Hows. Paediatric
546 Archives ofdise in Childhood 1991; 66: 546-550 CURRENT TOPIC Department of Haematology Royal Postgraduate Medical School, Hammeramith Hospital, London W12 ONN Correspondence to: Dr Hows. Paediatric
Disease stage stratified effects of cell dose in unrelated BMT for hematological malignancies: a report from Japan marrow donor program
 (2010), 1 11 & 2010 Macmillan Publishers Limited All rights reserved 0268-3369/10 www.nature.com/bmt ORIGINAL ARTICLE Disease stage stratified effects of cell dose in unrelated BMT for hematological malignancies:
(2010), 1 11 & 2010 Macmillan Publishers Limited All rights reserved 0268-3369/10 www.nature.com/bmt ORIGINAL ARTICLE Disease stage stratified effects of cell dose in unrelated BMT for hematological malignancies:
High Resolution Donor HLA-Matching: Saving Lives
 Newsletter CONTENTS High resolution donor HLA- matching... 1 New Website: cibmtr.org... 1 Perspectives... 3 Annual BMT meeting news... 4 2005 Summary Slides (pull-out section)... 5-8 AGNIS: Help for Data
Newsletter CONTENTS High resolution donor HLA- matching... 1 New Website: cibmtr.org... 1 Perspectives... 3 Annual BMT meeting news... 4 2005 Summary Slides (pull-out section)... 5-8 AGNIS: Help for Data
Samples Available for Recipient Only. Samples Available for Recipient and Donor
 Unrelated HCT Research Sample Inventory - Summary for First Allogeneic Transplants in CRF and TED with biospecimens available through the CIBMTR Repository stratified by availability of paired samples,
Unrelated HCT Research Sample Inventory - Summary for First Allogeneic Transplants in CRF and TED with biospecimens available through the CIBMTR Repository stratified by availability of paired samples,
Samples Available for Recipient and Donor
 Unrelated HCT Research Sample Inventory - Summary for First Allogeneic Transplants in CRF and TED with biospecimens available through the CIBMTR Repository stratified by availability of paired samples,
Unrelated HCT Research Sample Inventory - Summary for First Allogeneic Transplants in CRF and TED with biospecimens available through the CIBMTR Repository stratified by availability of paired samples,
Antigen Presentation to T lymphocytes
 Antigen Presentation to T lymphocytes Immunology 441 Lectures 6 & 7 Chapter 6 October 10 & 12, 2016 Jessica Hamerman jhamerman@benaroyaresearch.org Office hours by arrangement Antigen processing: How are
Antigen Presentation to T lymphocytes Immunology 441 Lectures 6 & 7 Chapter 6 October 10 & 12, 2016 Jessica Hamerman jhamerman@benaroyaresearch.org Office hours by arrangement Antigen processing: How are
Seventy percent of people who are candidates for allogeneic hematopoietic
 274 7th Biennial Symposium on Minorities, the Medically Underserved and Cancer Supplement to Cancer The National Marrow Donor Program Meeting the Needs of the Medically Underserved Dennis L. Confer, M.D.
274 7th Biennial Symposium on Minorities, the Medically Underserved and Cancer Supplement to Cancer The National Marrow Donor Program Meeting the Needs of the Medically Underserved Dennis L. Confer, M.D.
IMMUNOGENETICS AND TRANSPLANTATION
 IMMUNOGENETICS AND TRANSPLANTATION WHAT S THE MHC GOT TO DO WITH TRANSPLANTATION? We have learned that the molecules of Class I and II MHC are involved in presenting antigenic peptides to the receptors
IMMUNOGENETICS AND TRANSPLANTATION WHAT S THE MHC GOT TO DO WITH TRANSPLANTATION? We have learned that the molecules of Class I and II MHC are involved in presenting antigenic peptides to the receptors
XIV. HLA AND TRANSPLANTATION MEDICINE
 XIV. HLA AND TRANSPLANTATION MEDICINE A. Introduction 1. The HLA system includes a complex array of genes and their molecular products that are involved in immune regulation and cellular differentiation.
XIV. HLA AND TRANSPLANTATION MEDICINE A. Introduction 1. The HLA system includes a complex array of genes and their molecular products that are involved in immune regulation and cellular differentiation.
Haploidentical Transplantation: The Answer to our Donor Problems? Mary M. Horowitz, MD, MS CIBMTR, Medical College of Wisconsin January 2017
 Haploidentical Transplantation: The Answer to our Donor Problems? Mary M. Horowitz, MD, MS CIBMTR, Medical College of Wisconsin January 2017 Allogeneic Transplant Recipients in the US, by Donor Type 9000
Haploidentical Transplantation: The Answer to our Donor Problems? Mary M. Horowitz, MD, MS CIBMTR, Medical College of Wisconsin January 2017 Allogeneic Transplant Recipients in the US, by Donor Type 9000
The MHC and Transplantation Brendan Clark. Transplant Immunology, St James s University Hospital, Leeds, UK
 The MHC and Transplantation Brendan Clark Transplant Immunology, St James s University Hospital, Leeds, UK Blood Groups Immunofluorescent staining has revealed blood group substance in the cell membranes
The MHC and Transplantation Brendan Clark Transplant Immunology, St James s University Hospital, Leeds, UK Blood Groups Immunofluorescent staining has revealed blood group substance in the cell membranes
An Introduction to Bone Marrow Transplant
 Introduction to Blood Cancers An Introduction to Bone Marrow Transplant Rushang Patel, MD, PhD, FACP Florida Hospital Medical Group S My RBC Plt Gran Polycythemia Vera Essential Thrombocythemia AML, CML,
Introduction to Blood Cancers An Introduction to Bone Marrow Transplant Rushang Patel, MD, PhD, FACP Florida Hospital Medical Group S My RBC Plt Gran Polycythemia Vera Essential Thrombocythemia AML, CML,
Determination of HLA-A, -C, -B, -DRB1 allele and haplotype frequency in Japanese population based on family study
 Tissue Antigens ISSN 0001-2815 Determination of HLA-A, -C, -B, -DRB1 allele and haplotype frequency in Japanese population based on family study N. Ikeda 1,H.Kojima 1,M.Nishikawa 1, K. Hayashi 1, T. Futagami
Tissue Antigens ISSN 0001-2815 Determination of HLA-A, -C, -B, -DRB1 allele and haplotype frequency in Japanese population based on family study N. Ikeda 1,H.Kojima 1,M.Nishikawa 1, K. Hayashi 1, T. Futagami
Transplantation Immunology Booklet C
 Booklet C Pretest Correct Answers 3. (C) is correct. One of the major barriers to successful hematopoietic stem cell transplantation is the antigens coded for by the major histocompatibility gene complex.
Booklet C Pretest Correct Answers 3. (C) is correct. One of the major barriers to successful hematopoietic stem cell transplantation is the antigens coded for by the major histocompatibility gene complex.
Available online at , 2(1):27-37
 Available online at www.apjbg.com Asia-Pacific Journal of Blood Types and Genes 2018, 2(1):27-37 APJBG Allele and haplotype frequencies of human leukocyte antigen-a, -B, -C, -DRB1, and -DQB1 in Chinese
Available online at www.apjbg.com Asia-Pacific Journal of Blood Types and Genes 2018, 2(1):27-37 APJBG Allele and haplotype frequencies of human leukocyte antigen-a, -B, -C, -DRB1, and -DQB1 in Chinese
A glow of HLA typing in organ transplantation
 Mahdi Clinical and Translational Medicine 2013, 2:6 REVIEW Open Access A glow of HLA typing in organ transplantation Batool Mutar Mahdi Abstract The transplant of organs and tissues is one of the greatest
Mahdi Clinical and Translational Medicine 2013, 2:6 REVIEW Open Access A glow of HLA typing in organ transplantation Batool Mutar Mahdi Abstract The transplant of organs and tissues is one of the greatest
Umbilical Cord Blood Transplantation
 Umbilical Cord Blood Transplantation Current Results John E. Wagner, M.D. Blood and Marrow Transplant Program and Stem Cell Institute University of Minnesota Donor Choices Unrelated Marrow/PBSC Results
Umbilical Cord Blood Transplantation Current Results John E. Wagner, M.D. Blood and Marrow Transplant Program and Stem Cell Institute University of Minnesota Donor Choices Unrelated Marrow/PBSC Results
ORIGINAL ARTICLE. Bone Marrow Transplantation (2015) 50, ; doi: /bmt ; published online 8 June 2015
 Bone Marrow Transplantation (2015) 50, 1201 1205 2015 Macmillan Publishers Limited All rights reserved 0268-3369/15 www.nature.com/bmt ORIGINAL ARTICLE High-resolution HLA matching in unrelated donor transplantation
Bone Marrow Transplantation (2015) 50, 1201 1205 2015 Macmillan Publishers Limited All rights reserved 0268-3369/15 www.nature.com/bmt ORIGINAL ARTICLE High-resolution HLA matching in unrelated donor transplantation
Use of a five-agent GVHD prevention regimen in recipients of unrelated donor marrow
 Bone Marrow Transplantation, (1999) 23, 889 893 1999 Stockton Press All rights reserved 0268 3369/99 $12.00 http://www.stockton-press.co.uk/bmt Use of a five-agent GVHD prevention regimen in recipients
Bone Marrow Transplantation, (1999) 23, 889 893 1999 Stockton Press All rights reserved 0268 3369/99 $12.00 http://www.stockton-press.co.uk/bmt Use of a five-agent GVHD prevention regimen in recipients
Marrow transplantation from unrelated donors for patients with severe aplastic anemia who have failed immunosuppressive therapy
 Biology of Blood and Marrow Transplantation 5:243 252 (1999) 1999 American Society for Blood and Marrow Transplantation ASBMT #48 Marrow transplantation from unrelated donors for patients with severe aplastic
Biology of Blood and Marrow Transplantation 5:243 252 (1999) 1999 American Society for Blood and Marrow Transplantation ASBMT #48 Marrow transplantation from unrelated donors for patients with severe aplastic
KEY WORDS: Unrelated SCT, HLA-mismatch, ATG, Graft-versus-host disease
 HLA-Mismatched Unrelated Donors as an Alternative Graft Source for Allogeneic Stem Cell Transplantation after Antithymocyte Globulin-Containing Conditioning Regimen Nicolaus Kröger, 1 Tatjana Zabelina,
HLA-Mismatched Unrelated Donors as an Alternative Graft Source for Allogeneic Stem Cell Transplantation after Antithymocyte Globulin-Containing Conditioning Regimen Nicolaus Kröger, 1 Tatjana Zabelina,
Predicting HLA Class II Alloantigen Immunogenicity From the Number and Physiochemical Properties of Amino Acid Polymorphisms
 CLINICAL AND TRANSLATIONAL RESEARCH Predicting HLA Class II Alloantigen Immunogenicity From the Number and Physiochemical Properties of Amino Acid Polymorphisms Vasilis Kosmoliaptsis, 1,2,6 Linda D. Sharples,
CLINICAL AND TRANSLATIONAL RESEARCH Predicting HLA Class II Alloantigen Immunogenicity From the Number and Physiochemical Properties of Amino Acid Polymorphisms Vasilis Kosmoliaptsis, 1,2,6 Linda D. Sharples,
Medhat Askar, 1 Ronald Sobecks, 2 Yasuo Morishima, 3 Takakazu Kawase, 3 Amy Nowacki, 4 Hideki Makishima, 5 Jaroslaw Maciejewski 5
 Biol Blood Marrow Transplant 17:1404-1415, 2011 HistoCheck versus High-Risk HLA Allele Mismatch Combinations 1409 8. Kiyoi H, Yanada M, Ozekia K. Clinical significance of FLT3 in leukemia. Int J Hematol.
Biol Blood Marrow Transplant 17:1404-1415, 2011 HistoCheck versus High-Risk HLA Allele Mismatch Combinations 1409 8. Kiyoi H, Yanada M, Ozekia K. Clinical significance of FLT3 in leukemia. Int J Hematol.
REVIEW ARTICLE. Umbilical Cord Blood Transplantation: Where Do We Stand? ASBMT BB&MT. Raymond C. Wadlow, David L. Porter
 Biology of Blood and Marrow Transplantation 8:637-647 (2002) 2002 American Society for Blood and Marrow Transplantation ASBMT REVIEW ARTICLE Umbilical Cord Blood Transplantation: Where Do We Stand? Raymond
Biology of Blood and Marrow Transplantation 8:637-647 (2002) 2002 American Society for Blood and Marrow Transplantation ASBMT REVIEW ARTICLE Umbilical Cord Blood Transplantation: Where Do We Stand? Raymond
Handling Immunogenetic Data Managing and Validating HLA Data
 Handling Immunogenetic Data Managing and Validating HLA Data Steven J. Mack PhD Children s Hospital Oakland Research Institute 16 th IHIW & Joint Conference Sunday 3 June, 2012 Overview 1. Master Analytical
Handling Immunogenetic Data Managing and Validating HLA Data Steven J. Mack PhD Children s Hospital Oakland Research Institute 16 th IHIW & Joint Conference Sunday 3 June, 2012 Overview 1. Master Analytical
Trends in Hematopoietic Cell Transplantation. AAMAC Patient Education Day Oct 2014
 Trends in Hematopoietic Cell Transplantation AAMAC Patient Education Day Oct 2014 Objectives Review the principles behind allogeneic stem cell transplantation Outline the process of transplant, some of
Trends in Hematopoietic Cell Transplantation AAMAC Patient Education Day Oct 2014 Objectives Review the principles behind allogeneic stem cell transplantation Outline the process of transplant, some of
