INSIGHT THE BINDING SITE OF HUMAN MELATONIN MTR1A RECEPTOR
|
|
|
- Joy Alicia Warren
- 5 years ago
- Views:
Transcription
1 ACADEMIA ROMÂNĂ Revue Roumaine de Chimie Rev. Roum. Chim., 2015, 60(2-3), Dedicated to Professor Zeno Simon on the occasion of his 80 th anniversary INSIGHT THE BINDING SITE OF HUMAN MELATONIN MTR1A RECEPTOR Liliana HALIP, * Ramona CURPĂN, Ana BOROTA and Alina BORA Timişoara Institute of the Roumanian Academy Chemistry, 24 Mihai Viteazul Avenue, RO , Timişoara, Roumania Received August 25, 2014 Besides the known applications of melatonin which include circadian rhythm s regulation, sleep disorders and depression, recent studies have reported the melatonin implication in cardiovascular regulation, neurodegenerative diseases or cancer treatment. In this light, the melatonin receptors have gained much attention in the last few years, and many drug discovery programs have been focused on the identification of new small molecules which selectively bind to the two melatonin receptor subtypes. In this study, we report the homology modeling of the human melatonin MTR1A based on the crystal structure of the beta1-adrenergic receptor (beta1-ar) receptor, the characterization of the binding site and the most relevant ligand receptor interactions. Docking simulations revealed an agonist binding mode characterized by interactions with two residues from helix 3, Ser3.35 and Ser 3.39, respectively and a residue from helix 5, His5.46. INTRODUCTION * Melatonin is a hormone primarily synthesized and secreted by pineal glad following the light-dark cycle. Besides its important role in the regulation of circadian rhythm and the treatment of sleep disorders or depression, melatonin was reported to influence several other important physiological functions like modulation of the cardiovascular response, tumor suppression or diabetes. 1 5 The most important actions of melatonin are exerted via two membrane receptors, melatonin MTR1A (MT1) and MTR1B (MT2), members of G-protein coupled receptor family (GPCRs). Currently, all marketed melatonergic drugs target on both melatonin receptor subtypes showing no significant selectivity. In order to develop new therapeutic agents with improved selectivity, it is necessary to understand and determine all the aspects regarding ligand binding and functional efficacy for each melatonin subtype. In the present work, the full characterization of the melatonin MTR1A binding site and the primary hot spots for agonist binding are provided in order to facilitate the generation of pharmacophore models or to setup the most appropriate conditions and parameters for successful virtual screening studies. METHODS Sequence alignment and template selection The sequence of the human melatonin MTR1A receptor was extracted from the UniProt database 6 * Corresponding author: lili.ostopovici@acad-icht.tm.edu.ro
2 214 Liliana Halip et al. (accession code: P48039) in Fasta format and was automatically aligned using the T-coffee server 7 with the sequences of the all class A GPCRs receptors for which experimental the three-dimensional (3D) structure are available. The alignment was manually adjusted in order to preserve the highly-conserved amino acid motif specific for each transmembrane 8 and to avoid deletions or insertions in the transmembrane domain (TMD). When necessary, the deletions and insertions were pasted into a single piece per loop and placed in the most adequate point according to the 3D structure of the template. The disulfide bridge between the second extracellular loop (XL2) and the extracellular part of the third helix, conserved in all GPCRs, was taken in consideration. Amino acid numbering Specific amino acids from the transmembrane helices were named using the numbering system proposed by Ballesteros and Weinstein in Briefly, for the most highly conserved residue in each transmembrane 8 a value of 50 was assigned, preceded by the number of the transmembrane helix. The other residues on the same helix will have a value corresponding to their position relative to the most conserved residue in transmembrane. Model building and loop modeling The homology modeling package Modeller 10 (version 9v11) was used to generate the threedimensional model for melatonin MTR1A receptor based on the crystal structure of beta1-ar. The resulted model was geometrically minimized to reduce sterical clashes. The final model has over 90% of residues in the favorable regions of the Ramachandran map and all the main-chain and side-chain parameters are in the normal range. Binding site detection The location of the primary binding site in the MTR1A receptor was determined by using results from mutagenesis studies 11,12 and from computational predictions with SiteMap application implemented in Schrödinger software package. Molecular Docking Ligand setup. Ligands were prepared for docking using LigPrep 2.9 application 16 from Schrödinger. The polar and non-polar hydrogen atoms were added to ligands and all possible tautomers at physiological conditions (ph=7.4) were generated. Also, the ionization states were set and the final geometries were minimized using OPLS_2005 force field. Receptor preparation. Protein Preparation Workflow 17,18 was used to add all the hydrogen atoms to the MTR1A receptor, to optimize the charge state of Asp, Glu, Arg, Lys and His residues and the orientation of hydroxyl and amide groups. Docking Protocol. Glide receptor grids were generated from the prepared receptor model, with the docking grids centered on the binding site. The binding site was defined by a cubic docking box with the dimensions large enough to accommodate ligands with a length 20 Å, while to the ligandmidpoint box a side of 14 Å was given. The docking runs were performed with Glide in SP mode using default settings for parameters. RESULTS AND DISCUSSION Template selection and model building. Selection of the most appropriate template structure represents one of the most critical step in the comparative modeling technique since it can affect significantly the models final accuracy. The typical method for template identification consists in finding the closest neighbor or a given sequence protein, called target, by serial pairwise sequence alignments. For melatonin MTR1A receptor, the available sequence protein space is given by a number of 28 GPCRs for which structural information is available and deposited in Protein Data Bank. 23,24 Out of these, 24 sequences belong to class A GPCRs same as melatonin MTR1A receptor and therefore were further considered as potential templates for homology modeling. Since all the class A GPCRs share an orthosteric binding cavity, with different sizes and shapes, located in the extracellular side of the transmembrane bundle, the binding site similarity is also an important parameter to consider, when new therapeutic compounds are pursued. Consequently, the finding of the most appropriate template for melatonin MT1A receptor was conducted based on two main criteria: receptor and binding site similarity. Binding site pocket was restricted to a reference set of 44 residues previously defined by Gloriam. 25 Classification of MTR1A and those 24 class A GPCRs was based on receptor and binding site similarity output results with minor or moderate differences regarding the number of clusters and their components (e.g. H1 receptor). For MTR1A receptor, the minimum distance in the class A
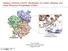
3 Human melatonin receptor 215 sequences space is given by a cluster of three receptors: beta1-adrenergic receptor (AR), beta2-ar and human dopamine D3 receptor (Fig. 1). For each applied criteria, the minimum distances between melatonin MTR1A receptor and the three potential templates are comparable, with beta1-ar being the closest relative. Moreover, when sequence identity was calculated, beta1-ar showed also the highest value with respect to MTR1A receptor. The sequence alignment between template (beta1- AR) and target (MTR1A) was manually refined according to 3D coordinates of the beta1-ar (Fig. 2). The homology models for melatonin MTR1A receptor was generated based on the alignment shown in Fig. 2 and the resulted model, including the main backbone, was energetically minimized. Binding site characterization. The location of the primary binding site identified by SiteMap application is in agreement with the available experimental data. 11,12 Therefore, the following residues, Ser3.35, Ser3.39, Val5.42, His5.46 and Gly6.55 reported to be critical or having relevant involvements in the agonist binding at MTR1A receptor 11,12 were located towards the extracellular side of helices 3, 5 and 6. The sidechains of these residues are oriented inside the transmembrane bundle. Noteworthy, compared to other class A GPCRs, binding cavity of melatonin MTR1A receptor goes deeply inside the transmembrane bundle (Fig. 3) and it has a strong hydrophobic character in several positions, wellknown as being important for GPCRs agonist binding pocket. For example, positions 3.32 and 3.33 which are known as important anchor points of the biogenic amines in the GPCRs binding site, are occupied in the MTR1A receptor by a methionine and a glycine residue. The potential hot spot residues for the melatonin binding a high lipophilic compound are represented by polar or charged amino acids like Ser3.39, Tyr5.38, His5.46, Asn6.52, Tyr7.39 and Tyr7.43 (Fig. 4). Fig. 1 Dendrogram of melatonin MTR1A receptor and class A GPCRs with known 3D structure based on receptor (A) and binding site (B) similarity. Melatonin MTR1A receptor is highlighted in dark gray and its closest neighbors in light gray. Fig. 2 Sequence alignment of melatonin MTR1A receptor and beta1-ar.
4 216 Liliana Halip et al. Fig. 3 MTR1A binding site, detected by SiteMap3.0 14,15. The hydrophobic regions are represented with black, while the hydrophilic regions are represented with gray surfaces. Fig. 4 The potential hot spot residues (black surface) for melatonin interaction at the MTR1A binding site, upper- (A) and lateral (B) view. Molecular docking of the endogenous ligand, melatonin, revealed a binding mode characterized by two types of ligand-receptor interactions: hydrogen bonds and π-π interactions (Fig.5). The amide moiety is positioned in the close neighborhood of Ser3.35 and Ser3.39 located on helix 3 and has a favorable orientation to establish hydrogen bonds with their side-chains. The indole
5 Human melatonin receptor 217 ring is stabilized in the situs by π-π interactions established with the side-chains of several aromatic amino acids like His5.46, Phe5.47 and Trp6.48. Moreover, the nitrogen indole acts as a hydrogen bond donor to the hydroxyl group of Tyr7.39. The ethyl linker is accommodated in a hydrophobic space defined by amino acids like Val3.36 and Ile3.37, from helix 3 and Ala7.42 from helix 7. The 5-methoxy substituent is faced towards aliphatic amino acids like Ile3.40 from helix 3 and Val5.42 and Leu5.48 from helix 5 (Fig. 5). In a similar manner, other MTR1A agonists ramelteon, 2-iodo-melatonin and agomelatine have a similar binding mode in which the amide function is interacting with the two serine residues from helix 3 and the aromatic ring is involved in a stacking interaction with His5.46 (Fig. 6). Fig. 5 Binding mode of melatonin in human MTR1A binding site. Fig. 6 Agonist binding mode at MTR1A receptor, illustrated on melatonin.
6 218 Liliana Halip et al. CONCLUSIONS The homology model of the MTR1A receptor based on the crystal structure of the beta1- adrenergic receptor was generated using homology modeling. The final model has all steric and topologic parameters within the normal range and it has allowed the recapitulation of experimental data by docking simulations using the endogenous ligand and several known agonists. The agonist binding site was mapped and the most relevant ligand receptor interactions were described. The agonist binding mode is characterized by the formation of hydrogen bonds with two polar residues from helix 3, Ser3.35 and Ser 3.36 and a stacking interaction with His5.46 from helix 5. Acknowledgements: This work was supported by CNCSIS- UEFISCDI project PN-II-RU-PD , agreement no 119/ to LH and to Institute of Chemistry of Roumanian Academy, Timişoara, Project no 1.2/2014 to RC and AB. REFERENCES 1. A. Cavallo, S. R. Daniels, L. M. Dolan, J. A. Bean and J. C. Khoury, J. Pineal Res., 2004, 36, S. R. Pandi-Perumal, V. Srinivasan, G. J. M. Maestroni, D. P. Cardinali, B. Poeggeler and R. Hardeland, FEBS J., 2006, 273, A. Carrillo-Vico, J. M. Guerrero, P. J. Lardone and R. J. Reiter, Endocrine, 2005, 27, A. Dominguez-Rodriguez, Cardiovasc. Pharmacol., 2013, 2, e C. Ekmekcioglu, Biomed. Pharmacother., 2006, 60, T. U. Consortium, Nucleic Acids Res., 2012, 40, D C. Notredame, D. G. Higgins and J. Heringa, J. Mol. Biol., 2000, 302, J. M. Baldwin, G. F. Schertler and V. M. Unger, J. Mol. Biol., 1997, 272, J. A. Ballesteros and H. Weinstein, in Methods Neurosci (Ed.: S.C.S.B.T.-M. in Neurosciences), Academic Press, 1995, pp A. Sali and T. L. Blundell, J. Mol. Biol., 1993, 234, T. Kokkola, M. A. Watson, J. White, S. Dowell, S. M. Foord and J. T. Laitinen, Biochem. Biophys. Res. Commun., 1998, 249, S. Conway, E. S. Mowat, J. E. Drew, P. Barrett, P. Delagrange and P. J. Morgan, Biochem. Biophys. Res. Commun., 2001, 282, SiteMap, version 3.0, Schrödinger, LLC, New York, NY, T. A. Halgren, Chem. Biol. Drug Des., 2007, 69, T. A. Halgren, J. Chem. Inf. Model., 2009, 49, Schrödinger Release : LigPrep, version 2.9, Schrödinger, LLC, New York, NY, G. M. Sastry, M. Adzhigirey, T. Day, R. Annabhimoju and W. Sherman, J. Comput. Aided. Mol. Des., 2013, 27, Schrödinger Suite Protein Preparation Wizard; Epik version 2.7, Schrödinger, LLC, New York, NY, 2013; Impact version 6.2, Schrödinger, LLC, New York, NY, 2014; Prime version 3.4, Schrödinger, LLC, New York, NY, R. A. Friesner, R. B. Murphy, M. P. Repasky, L. L. Frye, J. R. Greenwood, T. A. Halgren, P. C. Sanschagrin and D. T. Mainz, J. Med. Chem., 2006, 49, R. A. Friesner, J. L. Banks, R. B. Murphy, T. A. Halgren, J. J. Klicic, D. T. Mainz, M. P. Repasky, E. H. Knoll, M. Shelley, J. K. Perry, et al., J. Med. Chem., 2004, 47, T. A. Halgren, R. B. Murphy, R. A. Friesner, H. S. Beard, L. L. Frye, W. T. Pollard and J. L. Banks, J. Med. Chem., 2004, 47, Small-Molecule Drug Discovery Suite : Glide, version 6.2, Schrödinger, LLC, New York, NY, H. Berman, K. Henrick and H. Nakamura, Nat Struct Mol Biol, 2003, 10, D. E. Gloriam, S. M. Foord, F. E. Blaney and S. L. Garland, J. Med. Chem., 2009, 52,
Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels. Supplementary Information
 Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels Craig T. Armstrong, Philip E. Mason, J. L. Ross Anderson and Christopher E. Dempsey
Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels Craig T. Armstrong, Philip E. Mason, J. L. Ross Anderson and Christopher E. Dempsey
CS612 - Algorithms in Bioinformatics
 Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
IDENTIFICATION OF POTENTIAL INHIBITORS FOR LOWERING CHOLESTEROL LEVEL BY INHIBITING PROPROTEIN CONVERTASE SUBTILISIN KEXIN 9
 Vol 9, Issue 6, 2016 Online - 2455-3891 Print - 0974-2441 Research Article IDENTIFICATION OF POTENTIAL INHIBITORS FOR LOWERING CHOLESTEROL LEVEL BY INHIBITING PROPROTEIN CONVERTASE SUBTILISIN KEXIN 9 ABSTRACT
Vol 9, Issue 6, 2016 Online - 2455-3891 Print - 0974-2441 Research Article IDENTIFICATION OF POTENTIAL INHIBITORS FOR LOWERING CHOLESTEROL LEVEL BY INHIBITING PROPROTEIN CONVERTASE SUBTILISIN KEXIN 9 ABSTRACT
Page 8/6: The cell. Where to start: Proteins (control a cell) (start/end products)
 Page 8/6: The cell Where to start: Proteins (control a cell) (start/end products) Page 11/10: Structural hierarchy Proteins Phenotype of organism 3 Dimensional structure Function by interaction THE PROTEIN
Page 8/6: The cell Where to start: Proteins (control a cell) (start/end products) Page 11/10: Structural hierarchy Proteins Phenotype of organism 3 Dimensional structure Function by interaction THE PROTEIN
3.2 Ligand-Binding at Nicotinic Acid Receptor Subtypes GPR109A/B
 3.2 Ligand-Binding at Nicotinic Acid Receptor Subtypes GPR109A/B 3.2.1 Characterization of the Ligand Binding Site at GPR109A Receptor Ligands of GPR109A Receptor are Carboxylic Acids Nicotinic acid (pyridine-3-carboxylic
3.2 Ligand-Binding at Nicotinic Acid Receptor Subtypes GPR109A/B 3.2.1 Characterization of the Ligand Binding Site at GPR109A Receptor Ligands of GPR109A Receptor are Carboxylic Acids Nicotinic acid (pyridine-3-carboxylic
Amino Acids. Review I: Protein Structure. Amino Acids: Structures. Amino Acids (contd.) Rajan Munshi
 Review I: Protein Structure Rajan Munshi BBSI @ Pitt 2005 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2005 Amino Acids Building blocks of proteins 20 amino acids
Review I: Protein Structure Rajan Munshi BBSI @ Pitt 2005 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2005 Amino Acids Building blocks of proteins 20 amino acids
SUPPORTING INFORMATION FOR. A Computational Approach to Enzyme Design: Using Docking and MM- GBSA Scoring
 SUPPRTING INFRMATIN FR A Computational Approach to Enzyme Design: Predicting ω- Aminotransferase Catalytic Activity Using Docking and MM- GBSA Scoring Sarah Sirin, 1 Rajesh Kumar, 2 Carlos Martinez, 2
SUPPRTING INFRMATIN FR A Computational Approach to Enzyme Design: Predicting ω- Aminotransferase Catalytic Activity Using Docking and MM- GBSA Scoring Sarah Sirin, 1 Rajesh Kumar, 2 Carlos Martinez, 2
This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is worth 2 points.
 MBB 407/511 Molecular Biology and Biochemistry First Examination - October 1, 2002 Name Social Security Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is
MBB 407/511 Molecular Biology and Biochemistry First Examination - October 1, 2002 Name Social Security Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is
Molecular Biology. general transfer: occurs normally in cells. special transfer: occurs only in the laboratory in specific conditions.
 Chapter 9: Proteins Molecular Biology replication general transfer: occurs normally in cells transcription special transfer: occurs only in the laboratory in specific conditions translation unknown transfer:
Chapter 9: Proteins Molecular Biology replication general transfer: occurs normally in cells transcription special transfer: occurs only in the laboratory in specific conditions translation unknown transfer:
Chemical Nature of the Amino Acids. Table of a-amino Acids Found in Proteins
 Chemical Nature of the Amino Acids All peptides and polypeptides are polymers of alpha-amino acids. There are 20 a- amino acids that are relevant to the make-up of mammalian proteins (see below). Several
Chemical Nature of the Amino Acids All peptides and polypeptides are polymers of alpha-amino acids. There are 20 a- amino acids that are relevant to the make-up of mammalian proteins (see below). Several
Biomolecules: amino acids
 Biomolecules: amino acids Amino acids Amino acids are the building blocks of proteins They are also part of hormones, neurotransmitters and metabolic intermediates There are 20 different amino acids in
Biomolecules: amino acids Amino acids Amino acids are the building blocks of proteins They are also part of hormones, neurotransmitters and metabolic intermediates There are 20 different amino acids in
Introduction to proteins and protein structure
 Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature22394 Supplementary Table 1 Observed intermolecular interactions within the GLP-1:GLP-1R TMD interface. Superscripts refer to the Wootten residue numbering system
SUPPLEMENTARY INFORMATION doi:10.1038/nature22394 Supplementary Table 1 Observed intermolecular interactions within the GLP-1:GLP-1R TMD interface. Superscripts refer to the Wootten residue numbering system
Proteins are sometimes only produced in one cell type or cell compartment (brain has 15,000 expressed proteins, gut has 2,000).
 Lecture 2: Principles of Protein Structure: Amino Acids Why study proteins? Proteins underpin every aspect of biological activity and therefore are targets for drug design and medicinal therapy, and in
Lecture 2: Principles of Protein Structure: Amino Acids Why study proteins? Proteins underpin every aspect of biological activity and therefore are targets for drug design and medicinal therapy, and in
Review II: The Molecules of Life
 Review II: The Molecules of Life Judy Wieber BBSI @ Pitt 2007 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2007 Outline Introduction Proteins Carbohydrates Lipids
Review II: The Molecules of Life Judy Wieber BBSI @ Pitt 2007 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2007 Outline Introduction Proteins Carbohydrates Lipids
PROTEINS. Amino acids are the building blocks of proteins. Acid L-form * * Lecture 6 Macromolecules #2 O = N -C -C-O.
 Proteins: Linear polymers of amino acids workhorses of the cell tools, machines & scaffolds Lecture 6 Macromolecules #2 PRTEINS 1 Enzymes catalysts that mediate reactions, increase reaction rate Structural
Proteins: Linear polymers of amino acids workhorses of the cell tools, machines & scaffolds Lecture 6 Macromolecules #2 PRTEINS 1 Enzymes catalysts that mediate reactions, increase reaction rate Structural
Maha AbuAjamieh. Tamara Wahbeh. Mamoon Ahram
 12 Maha AbuAjamieh Tamara Wahbeh Mamoon Ahram - - Go to this sheet s last page for definitions of the words with an asterisk above them (*) - You should memorise the 3-letter abbreviations, of all the
12 Maha AbuAjamieh Tamara Wahbeh Mamoon Ahram - - Go to this sheet s last page for definitions of the words with an asterisk above them (*) - You should memorise the 3-letter abbreviations, of all the
Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase. Biol 405 Molecular Medicine
 Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase Biol 405 Molecular Medicine 1998 Crystal structure of phenylalanine hydroxylase solved. The polypeptide consists of three regions: Regulatory
Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase Biol 405 Molecular Medicine 1998 Crystal structure of phenylalanine hydroxylase solved. The polypeptide consists of three regions: Regulatory
Objective: You will be able to explain how the subcomponents of
 Objective: You will be able to explain how the subcomponents of nucleic acids determine the properties of that polymer. Do Now: Read the first two paragraphs from enduring understanding 4.A Essential knowledge:
Objective: You will be able to explain how the subcomponents of nucleic acids determine the properties of that polymer. Do Now: Read the first two paragraphs from enduring understanding 4.A Essential knowledge:
Secondary Structure. by hydrogen bonds
 Secondary Structure In the previous protein folding activity, you created a hypothetical 15-amino acid protein and learned that basic principles of chemistry determine how each protein spontaneously folds
Secondary Structure In the previous protein folding activity, you created a hypothetical 15-amino acid protein and learned that basic principles of chemistry determine how each protein spontaneously folds
Chapter 3: Amino Acids and Peptides
 Chapter 3: Amino Acids and Peptides BINF 6101/8101, Spring 2018 Outline 1. Overall amino acid structure 2. Amino acid stereochemistry 3. Amino acid sidechain structure & classification 4. Non-standard
Chapter 3: Amino Acids and Peptides BINF 6101/8101, Spring 2018 Outline 1. Overall amino acid structure 2. Amino acid stereochemistry 3. Amino acid sidechain structure & classification 4. Non-standard
Biological systems interact, and these systems and their interactions possess complex properties. STOP at enduring understanding 4A
 Biological systems interact, and these systems and their interactions possess complex properties. STOP at enduring understanding 4A Homework Watch the Bozeman video called, Biological Molecules Objective:
Biological systems interact, and these systems and their interactions possess complex properties. STOP at enduring understanding 4A Homework Watch the Bozeman video called, Biological Molecules Objective:
Biomolecules Amino Acids & Protein Chemistry
 Biochemistry Department Date: 17/9/ 2017 Biomolecules Amino Acids & Protein Chemistry Prof.Dr./ FAYDA Elazazy Professor of Biochemistry and Molecular Biology Intended Learning Outcomes ILOs By the end
Biochemistry Department Date: 17/9/ 2017 Biomolecules Amino Acids & Protein Chemistry Prof.Dr./ FAYDA Elazazy Professor of Biochemistry and Molecular Biology Intended Learning Outcomes ILOs By the end
Practice Problems 3. a. What is the name of the bond formed between two amino acids? Are these bonds free to rotate?
 Life Sciences 1a Practice Problems 3 1. Draw the oligopeptide for Ala-Phe-Gly-Thr-Asp. You do not need to indicate the stereochemistry of the sidechains. Denote with arrows the bonds formed between the
Life Sciences 1a Practice Problems 3 1. Draw the oligopeptide for Ala-Phe-Gly-Thr-Asp. You do not need to indicate the stereochemistry of the sidechains. Denote with arrows the bonds formed between the
Judy Wieber. Department of Computational Biology. May 27, 2008
 Review II: The Molecules of Life Judy Wieber BBSI @ Pitt 2008 Department of Computational Biology University it of Pittsburgh School of Medicine i May 27, 2008 Outline Introduction Proteins Carbohydrates
Review II: The Molecules of Life Judy Wieber BBSI @ Pitt 2008 Department of Computational Biology University it of Pittsburgh School of Medicine i May 27, 2008 Outline Introduction Proteins Carbohydrates
Secondary Structure North 72nd Street, Wauwatosa, WI Phone: (414) Fax: (414) dmoleculardesigns.com
 Secondary Structure In the previous protein folding activity, you created a generic or hypothetical 15-amino acid protein and learned that basic principles of chemistry determine how each protein spontaneously
Secondary Structure In the previous protein folding activity, you created a generic or hypothetical 15-amino acid protein and learned that basic principles of chemistry determine how each protein spontaneously
Macromolecules of Life -3 Amino Acids & Proteins
 Macromolecules of Life -3 Amino Acids & Proteins Shu-Ping Lin, Ph.D. Institute of Biomedical Engineering E-mail: splin@dragon.nchu.edu.tw Website: http://web.nchu.edu.tw/pweb/users/splin/ Amino Acids Proteins
Macromolecules of Life -3 Amino Acids & Proteins Shu-Ping Lin, Ph.D. Institute of Biomedical Engineering E-mail: splin@dragon.nchu.edu.tw Website: http://web.nchu.edu.tw/pweb/users/splin/ Amino Acids Proteins
9/6/2011. Amino Acids. C α. Nonpolar, aliphatic R groups
 Amino Acids Side chains (R groups) vary in: size shape charge hydrogen-bonding capacity hydrophobic character chemical reactivity C α Nonpolar, aliphatic R groups Glycine (Gly, G) Alanine (Ala, A) Valine
Amino Acids Side chains (R groups) vary in: size shape charge hydrogen-bonding capacity hydrophobic character chemical reactivity C α Nonpolar, aliphatic R groups Glycine (Gly, G) Alanine (Ala, A) Valine
Chemistry 121 Winter 17
 Chemistry 121 Winter 17 Introduction to Organic Chemistry and Biochemistry Instructor Dr. Upali Siriwardane (Ph.D. Ohio State) E-mail: upali@latech.edu Office: 311 Carson Taylor Hall ; Phone: 318-257-4941;
Chemistry 121 Winter 17 Introduction to Organic Chemistry and Biochemistry Instructor Dr. Upali Siriwardane (Ph.D. Ohio State) E-mail: upali@latech.edu Office: 311 Carson Taylor Hall ; Phone: 318-257-4941;
CHM333 LECTURE 6: 1/25/12 SPRING 2012 Professor Christine Hrycyna AMINO ACIDS II: CLASSIFICATION AND CHEMICAL CHARACTERISTICS OF EACH AMINO ACID:
 AMINO ACIDS II: CLASSIFICATION AND CHEMICAL CHARACTERISTICS OF EACH AMINO ACID: - The R group side chains on amino acids are VERY important. o Determine the properties of the amino acid itself o Determine
AMINO ACIDS II: CLASSIFICATION AND CHEMICAL CHARACTERISTICS OF EACH AMINO ACID: - The R group side chains on amino acids are VERY important. o Determine the properties of the amino acid itself o Determine
Amino Acids. Lecture 4: Margaret A. Daugherty. Fall Swiss-prot database: How many proteins? From where?
 Lecture 4: Amino Acids Margaret A. Daugherty Fall 2004 Swiss-prot database: How many proteins? From where? 1986 Use http://us.expasy.org to get to swiss-prot database Proteins are the workhorses of the
Lecture 4: Amino Acids Margaret A. Daugherty Fall 2004 Swiss-prot database: How many proteins? From where? 1986 Use http://us.expasy.org to get to swiss-prot database Proteins are the workhorses of the
PAPER No. : 16, Bioorganic and biophysical chemistry MODULE No. : 22, Mechanism of enzyme catalyst reaction (I) Chymotrypsin
 Subject Paper No and Title 16 Bio-organic and Biophysical Module No and Title 22 Mechanism of Enzyme Catalyzed reactions I Module Tag CHE_P16_M22 Chymotrypsin TABLE OF CONTENTS 1. Learning outcomes 2.
Subject Paper No and Title 16 Bio-organic and Biophysical Module No and Title 22 Mechanism of Enzyme Catalyzed reactions I Module Tag CHE_P16_M22 Chymotrypsin TABLE OF CONTENTS 1. Learning outcomes 2.
BIRKBECK COLLEGE (University of London)
 BIRKBECK COLLEGE (University of London) SCHOOL OF BIOLOGICAL SCIENCES M.Sc. EXAMINATION FOR INTERNAL STUDENTS ON: Postgraduate Certificate in Principles of Protein Structure MSc Structural Molecular Biology
BIRKBECK COLLEGE (University of London) SCHOOL OF BIOLOGICAL SCIENCES M.Sc. EXAMINATION FOR INTERNAL STUDENTS ON: Postgraduate Certificate in Principles of Protein Structure MSc Structural Molecular Biology
Classification of amino acids: -
 Page 1 of 8 P roteinogenic amino acids, also known as standard, normal or primary amino acids are 20 amino acids that are incorporated in proteins and that are coded in the standard genetic code (subunit
Page 1 of 8 P roteinogenic amino acids, also known as standard, normal or primary amino acids are 20 amino acids that are incorporated in proteins and that are coded in the standard genetic code (subunit
paper and beads don t fall off. Then, place the beads in the following order on the pipe cleaner:
 Beady Pipe Cleaner Proteins Background: Proteins are the molecules that carry out most of the cell s dayto-day functions. While the DNA in the nucleus is "the boss" and controls the activities of the cell,
Beady Pipe Cleaner Proteins Background: Proteins are the molecules that carry out most of the cell s dayto-day functions. While the DNA in the nucleus is "the boss" and controls the activities of the cell,
Methionine (Met or M)
 Fig. 5-17 Nonpolar Fig. 5-17a Nonpolar Glycine (Gly or G) Alanine (Ala or A) Valine (Val or V) Leucine (Leu or L) Isoleucine (Ile or I) Methionine (Met or M) Phenylalanine (Phe or F) Polar Trypotphan (Trp
Fig. 5-17 Nonpolar Fig. 5-17a Nonpolar Glycine (Gly or G) Alanine (Ala or A) Valine (Val or V) Leucine (Leu or L) Isoleucine (Ile or I) Methionine (Met or M) Phenylalanine (Phe or F) Polar Trypotphan (Trp
PHARMACOPHORE MODELING
 CHAPTER 9 PHARMACOPHORE MODELING 9.1 PHARMACOPHORES AS VIRTUAL SCREENING TOOLS The pharmacophore concept has been widely used over the past decades for rational drug design approaches and has now been
CHAPTER 9 PHARMACOPHORE MODELING 9.1 PHARMACOPHORES AS VIRTUAL SCREENING TOOLS The pharmacophore concept has been widely used over the past decades for rational drug design approaches and has now been
A molecular docking study of anticancer drug paclitaxel and its analogues
 Indian Journal of Biochemistry & Biophysics Vol. 48, April 2011, pp. 101-105 A molecular docking study of anticancer drug paclitaxel and its analogues Ruma Sinha, Ambarish Sharan Vidyarthi and Shankaracharya*
Indian Journal of Biochemistry & Biophysics Vol. 48, April 2011, pp. 101-105 A molecular docking study of anticancer drug paclitaxel and its analogues Ruma Sinha, Ambarish Sharan Vidyarthi and Shankaracharya*
Four melanocyte-stimulating hormones have the following amino acid sequences:
 Assignment 14: Melanocyte-stimulating hormone belongs to a group called the melanocortins. This group includes ACTH, alpha-msh, beta-msh and gamma-msh; these peptides are all cleavage products of a large
Assignment 14: Melanocyte-stimulating hormone belongs to a group called the melanocortins. This group includes ACTH, alpha-msh, beta-msh and gamma-msh; these peptides are all cleavage products of a large
Properties of amino acids in proteins
 Properties of amino acids in proteins one of the primary roles of DNA (but far from the only one!!!) is to code for proteins A typical bacterium builds thousands types of proteins, all from ~20 amino acids
Properties of amino acids in proteins one of the primary roles of DNA (but far from the only one!!!) is to code for proteins A typical bacterium builds thousands types of proteins, all from ~20 amino acids
The Structure and Function of Macromolecules
 The Structure and Function of Macromolecules Macromolecules are polymers Polymer long molecule consisting of many similar building blocks. Monomer the small building block molecules. Carbohydrates, proteins
The Structure and Function of Macromolecules Macromolecules are polymers Polymer long molecule consisting of many similar building blocks. Monomer the small building block molecules. Carbohydrates, proteins
Structure and activation of the TSH receptor transmembrane domain
 Autoimmun Highlights (2017) 8:2 https://doi.org/10.1007/s13317-016-0090-1 ORIGINAL ARTICLE Structure and activation of the TSH receptor transmembrane domain Ricardo Núñez Miguel 1 Jane Sanders 1 Jadwiga
Autoimmun Highlights (2017) 8:2 https://doi.org/10.1007/s13317-016-0090-1 ORIGINAL ARTICLE Structure and activation of the TSH receptor transmembrane domain Ricardo Núñez Miguel 1 Jane Sanders 1 Jadwiga
For questions 1-4, match the carbohydrate with its size/functional group name:
 Chemistry 11 Fall 2013 Examination #5 PRACTICE 1 ANSWERS For the first portion of this exam, select the best answer choice for the questions below and mark the answers on your scantron. Then answer the
Chemistry 11 Fall 2013 Examination #5 PRACTICE 1 ANSWERS For the first portion of this exam, select the best answer choice for the questions below and mark the answers on your scantron. Then answer the
Protein Structure Monday, March
 Michael Morales ffice: 204/202 ary; knock hard if you visit as my office is some door. Lab: 238 & 205 ary ffice hours Either make an appointment or just drop by Phone: 829-3965 E-Mail: moralesm@buffalo.edu
Michael Morales ffice: 204/202 ary; knock hard if you visit as my office is some door. Lab: 238 & 205 ary ffice hours Either make an appointment or just drop by Phone: 829-3965 E-Mail: moralesm@buffalo.edu
Introduction to Protein Structure Collection
 Introduction to Protein Structure Collection Teaching Points This collection is designed to introduce students to the concepts of protein structure and biochemistry. Different activities guide students
Introduction to Protein Structure Collection Teaching Points This collection is designed to introduce students to the concepts of protein structure and biochemistry. Different activities guide students
Copyright 2008 Pearson Education, Inc., publishing as Pearson Benjamin Cummings
 Concept 5.4: Proteins have many structures, resulting in a wide range of functions Proteins account for more than 50% of the dry mass of most cells Protein functions include structural support, storage,
Concept 5.4: Proteins have many structures, resulting in a wide range of functions Proteins account for more than 50% of the dry mass of most cells Protein functions include structural support, storage,
2. Which of the following amino acids is most likely to be found on the outer surface of a properly folded protein?
 Name: WHITE Student Number: Answer the following questions on the computer scoring sheet. 1 mark each 1. Which of the following amino acids would have the highest relative mobility R f in normal thin layer
Name: WHITE Student Number: Answer the following questions on the computer scoring sheet. 1 mark each 1. Which of the following amino acids would have the highest relative mobility R f in normal thin layer
CHAPTER 21: Amino Acids, Proteins, & Enzymes. General, Organic, & Biological Chemistry Janice Gorzynski Smith
 CHAPTER 21: Amino Acids, Proteins, & Enzymes General, Organic, & Biological Chemistry Janice Gorzynski Smith CHAPTER 21: Amino Acids, Proteins, Enzymes Learning Objectives: q The 20 common, naturally occurring
CHAPTER 21: Amino Acids, Proteins, & Enzymes General, Organic, & Biological Chemistry Janice Gorzynski Smith CHAPTER 21: Amino Acids, Proteins, Enzymes Learning Objectives: q The 20 common, naturally occurring
The Basics: A general review of molecular biology:
 The Basics: A general review of molecular biology: DNA Transcription RNA Translation Proteins DNA (deoxy-ribonucleic acid) is the genetic material It is an informational super polymer -think of it as the
The Basics: A general review of molecular biology: DNA Transcription RNA Translation Proteins DNA (deoxy-ribonucleic acid) is the genetic material It is an informational super polymer -think of it as the
For questions 1-4, match the carbohydrate with its size/functional group name:
 Chemistry 11 Fall 2013 Examination #5 PRACTICE 1 For the first portion of this exam, select the best answer choice for the questions below and mark the answers on your scantron. Then answer the free response
Chemistry 11 Fall 2013 Examination #5 PRACTICE 1 For the first portion of this exam, select the best answer choice for the questions below and mark the answers on your scantron. Then answer the free response
Reactions and amino acids structure & properties
 Lecture 2: Reactions and amino acids structure & properties Dr. Sameh Sarray Hlaoui Common Functional Groups Common Biochemical Reactions AH + B A + BH Oxidation-Reduction A-H + B-OH + energy ª A-B + H
Lecture 2: Reactions and amino acids structure & properties Dr. Sameh Sarray Hlaoui Common Functional Groups Common Biochemical Reactions AH + B A + BH Oxidation-Reduction A-H + B-OH + energy ª A-B + H
Protein Structure Klemens Wild, BZH,
 Protein Structure recommended books Proteins protein definition From gr. proteios (superior, erstrangig) 1836 JJ Berzelius Functions: structural, enzymes, muscle, transport immune system, Linear polymer
Protein Structure recommended books Proteins protein definition From gr. proteios (superior, erstrangig) 1836 JJ Berzelius Functions: structural, enzymes, muscle, transport immune system, Linear polymer
BIO 311C Spring Lecture 15 Friday 26 Feb. 1
 BIO 311C Spring 2010 Lecture 15 Friday 26 Feb. 1 Illustration of a Polypeptide amino acids peptide bonds Review Polypeptide (chain) See textbook, Fig 5.21, p. 82 for a more clear illustration Folding and
BIO 311C Spring 2010 Lecture 15 Friday 26 Feb. 1 Illustration of a Polypeptide amino acids peptide bonds Review Polypeptide (chain) See textbook, Fig 5.21, p. 82 for a more clear illustration Folding and
Allosteric Inhibition of SHP2: Identification of a Potent, Selective, and Orally Efficacious Phosphatase Inhibitor!
 Allosteric Inhibition of SHP2: Identification of a Potent, Selective, and Orally Efficacious Phosphatase Inhibitor Allosteric pocket SHP2 Phosphatase ovel allosteric Phosphatase inhibitor Evan Carder Wipf
Allosteric Inhibition of SHP2: Identification of a Potent, Selective, and Orally Efficacious Phosphatase Inhibitor Allosteric pocket SHP2 Phosphatase ovel allosteric Phosphatase inhibitor Evan Carder Wipf
Ionization of amino acids
 Amino Acids 20 common amino acids there are others found naturally but much less frequently Common structure for amino acid COOH, -NH 2, H and R functional groups all attached to the a carbon Ionization
Amino Acids 20 common amino acids there are others found naturally but much less frequently Common structure for amino acid COOH, -NH 2, H and R functional groups all attached to the a carbon Ionization
Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site.
 Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
The Structure and Function of Large Biological Molecules Part 4: Proteins Chapter 5
 Key Concepts: The Structure and Function of Large Biological Molecules Part 4: Proteins Chapter 5 Proteins include a diversity of structures, resulting in a wide range of functions Proteins Enzymatic s
Key Concepts: The Structure and Function of Large Biological Molecules Part 4: Proteins Chapter 5 Proteins include a diversity of structures, resulting in a wide range of functions Proteins Enzymatic s
Chemical Mechanism of Enzymes
 Chemical Mechanism of Enzymes Enzyme Engineering 5.2 Definition of the mechanism 1. The sequence from substrate(s) to product(s) : Reaction steps 2. The rates at which the complex are interconverted 3.
Chemical Mechanism of Enzymes Enzyme Engineering 5.2 Definition of the mechanism 1. The sequence from substrate(s) to product(s) : Reaction steps 2. The rates at which the complex are interconverted 3.
A Chemical Look at Proteins: Workhorses of the Cell
 A Chemical Look at Proteins: Workhorses of the Cell A A Life ciences 1a Lecture otes et 4 pring 2006 Prof. Daniel Kahne Life requires chemistry 2 amino acid monomer and it is proteins that make the chemistry
A Chemical Look at Proteins: Workhorses of the Cell A A Life ciences 1a Lecture otes et 4 pring 2006 Prof. Daniel Kahne Life requires chemistry 2 amino acid monomer and it is proteins that make the chemistry
Biochemistry - I. Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture 1 Amino Acids I
 Biochemistry - I Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture 1 Amino Acids I Hello, welcome to the course Biochemistry 1 conducted by me Dr. S Dasgupta,
Biochemistry - I Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture 1 Amino Acids I Hello, welcome to the course Biochemistry 1 conducted by me Dr. S Dasgupta,
Lipids: diverse group of hydrophobic molecules
 Lipids: diverse group of hydrophobic molecules Lipids only macromolecules that do not form polymers li3le or no affinity for water hydrophobic consist mostly of hydrocarbons nonpolar covalent bonds fats
Lipids: diverse group of hydrophobic molecules Lipids only macromolecules that do not form polymers li3le or no affinity for water hydrophobic consist mostly of hydrocarbons nonpolar covalent bonds fats
PROTEINS. Building blocks, structure and function. Aim: You will have a clear picture of protein construction and their general properties
 PROTEINS Building blocks, structure and function Aim: You will have a clear picture of protein construction and their general properties Reading materials: Compendium in Biochemistry, page 13-49. Microbiology,
PROTEINS Building blocks, structure and function Aim: You will have a clear picture of protein construction and their general properties Reading materials: Compendium in Biochemistry, page 13-49. Microbiology,
استاذ الكيمياءالحيوية
 قسم الكيمياء الحيوية د.دولت على سالمه استاذ الكيمياءالحيوية ٢٠١٥-٢٠١٤ الرمز الكودي : ٥١٢ المحاضرة األولى ١ Content : Definition of proteins Definition of amino acids Definition of peptide bond General
قسم الكيمياء الحيوية د.دولت على سالمه استاذ الكيمياءالحيوية ٢٠١٥-٢٠١٤ الرمز الكودي : ٥١٢ المحاضرة األولى ١ Content : Definition of proteins Definition of amino acids Definition of peptide bond General
G protein coupled receptor interactions with cholesterol deep in the membrane
 G protein coupled receptor interactions with cholesterol deep in the membrane Samuel Genheden a,b, Jonathan W. Essex a,c, & Anthony G. Lee c,d * a School of Chemistry, University of Southampton, Southampton,
G protein coupled receptor interactions with cholesterol deep in the membrane Samuel Genheden a,b, Jonathan W. Essex a,c, & Anthony G. Lee c,d * a School of Chemistry, University of Southampton, Southampton,
Raghad Abu Jebbeh. Amani Nofal. Mamoon Ahram
 ... 14 Raghad Abu Jebbeh Amani Nofal Mamoon Ahram This sheet includes part of lec.13 + lec.14. Amino acid peptide protein Terminology: 1- Residue: a subunit that is a part of a large molecule. 2- Dipeptide:
... 14 Raghad Abu Jebbeh Amani Nofal Mamoon Ahram This sheet includes part of lec.13 + lec.14. Amino acid peptide protein Terminology: 1- Residue: a subunit that is a part of a large molecule. 2- Dipeptide:
Amino Acids االحماض األمينية
 Amino Acids االحماض األمينية Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry I Philadelphia University Faculty of pharmacy Cell Structure Amino Acids The study of
Amino Acids االحماض األمينية Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry I Philadelphia University Faculty of pharmacy Cell Structure Amino Acids The study of
Protein Investigator. Protein Investigator - 3
 Protein Investigator Objectives To learn more about the interactions that govern protein structure. To test hypotheses regarding protein structure and function. To design proteins with specific shapes.
Protein Investigator Objectives To learn more about the interactions that govern protein structure. To test hypotheses regarding protein structure and function. To design proteins with specific shapes.
INSILICO DOCKING AND INTERACTION ANALYSIS OF ELLAGIC ACID AND CURCUMIN DERIVATIVES AGAINST HUMAN CANCER
 ISSN: 0976-2876 (Print) ISSN: 2250-0138(Online) INSILICO DOCKING AND INTERACTION ANALYSIS OF ELLAGIC ACID AND CURCUMIN DERIVATIVES AGAINST HUMAN CANCER NAYANA MOHAN a AND M.S. LATHA b1 ab Department of
ISSN: 0976-2876 (Print) ISSN: 2250-0138(Online) INSILICO DOCKING AND INTERACTION ANALYSIS OF ELLAGIC ACID AND CURCUMIN DERIVATIVES AGAINST HUMAN CANCER NAYANA MOHAN a AND M.S. LATHA b1 ab Department of
Introduction. Basic Structural Principles PDB
 BCHS 6229 Protein Structure and Function Lecture 1 (October 11, 2011) Introduction Basic Structural Principles PDB 1 Overview Main Goals: Carry out a rapid review of the essentials of protein structure
BCHS 6229 Protein Structure and Function Lecture 1 (October 11, 2011) Introduction Basic Structural Principles PDB 1 Overview Main Goals: Carry out a rapid review of the essentials of protein structure
Short polymer. Dehydration removes a water molecule, forming a new bond. Longer polymer (a) Dehydration reaction in the synthesis of a polymer
 HO 1 2 3 H HO H Short polymer Dehydration removes a water molecule, forming a new bond Unlinked monomer H 2 O HO 1 2 3 4 H Longer polymer (a) Dehydration reaction in the synthesis of a polymer HO 1 2 3
HO 1 2 3 H HO H Short polymer Dehydration removes a water molecule, forming a new bond Unlinked monomer H 2 O HO 1 2 3 4 H Longer polymer (a) Dehydration reaction in the synthesis of a polymer HO 1 2 3
Moorpark College Chemistry 11 Fall Instructor: Professor Gopal. Examination # 5: Section Five May 7, Name: (print)
 Moorpark College Chemistry 11 Fall 2013 Instructor: Professor Gopal Examination # 5: Section Five May 7, 2013 Name: (print) Directions: Make sure your examination contains TEN total pages (including this
Moorpark College Chemistry 11 Fall 2013 Instructor: Professor Gopal Examination # 5: Section Five May 7, 2013 Name: (print) Directions: Make sure your examination contains TEN total pages (including this
1. Describe the relationship of dietary protein and the health of major body systems.
 Food Explorations Lab I: The Building Blocks STUDENT LAB INVESTIGATIONS Name: Lab Overview In this investigation, you will be constructing animal and plant proteins using beads to represent the amino acids.
Food Explorations Lab I: The Building Blocks STUDENT LAB INVESTIGATIONS Name: Lab Overview In this investigation, you will be constructing animal and plant proteins using beads to represent the amino acids.
University of Groningen. The medicinal chemistry of 2-aminotetralin-derived benzamides Homan, Evert Jan
 University of Groningen The medicinal chemistry of 2-aminotetralin-derived benzamides Homan, Evert Jan IMPORTAT OTE: You are advised to consult the publisher's version (publisher's PDF) if you wish to
University of Groningen The medicinal chemistry of 2-aminotetralin-derived benzamides Homan, Evert Jan IMPORTAT OTE: You are advised to consult the publisher's version (publisher's PDF) if you wish to
Catalysis & specificity: Proteins at work
 Catalysis & specificity: Proteins at work Introduction Having spent some time looking at the elements of structure of proteins and DNA, as well as their ability to form intermolecular interactions, it
Catalysis & specificity: Proteins at work Introduction Having spent some time looking at the elements of structure of proteins and DNA, as well as their ability to form intermolecular interactions, it
Organic molecules are molecules that contain carbon and hydrogen.
 Organic Chemistry, Biochemistry Introduction Organic molecules are molecules that contain carbon and hydrogen. All living things contain these organic molecules: carbohydrates, lipids, proteins, and nucleic
Organic Chemistry, Biochemistry Introduction Organic molecules are molecules that contain carbon and hydrogen. All living things contain these organic molecules: carbohydrates, lipids, proteins, and nucleic
Green Segment Contents
 Green Segment Contents Parts Reference Guide Green Segment 1 8 2 6 3 4 5 7 1. Amino Acid Side Chain Chart shows the properties and atomic structure of side chains. 2. Amino Acid Side Chains affect protein
Green Segment Contents Parts Reference Guide Green Segment 1 8 2 6 3 4 5 7 1. Amino Acid Side Chain Chart shows the properties and atomic structure of side chains. 2. Amino Acid Side Chains affect protein
Bioinformatics for molecular biology
 Bioinformatics for molecular biology Structural bioinformatics tools, predictors, and 3D modeling Structural Biology Review Dr Research Scientist Department of Microbiology, Oslo University Hospital -
Bioinformatics for molecular biology Structural bioinformatics tools, predictors, and 3D modeling Structural Biology Review Dr Research Scientist Department of Microbiology, Oslo University Hospital -
4. THE THREE-DIMENSIONAL STRUCTURE OF PROTEINS
 4. THE THREE-DIMENSIONAL STRUCTURE OF PROTEINS 4.1 Proteins Structures and Function Levels of Structure in Proteins Native conformation - Biological activity - Random structure: no obvious regular repeating
4. THE THREE-DIMENSIONAL STRUCTURE OF PROTEINS 4.1 Proteins Structures and Function Levels of Structure in Proteins Native conformation - Biological activity - Random structure: no obvious regular repeating
GPCR Structure-Based Virtual Screening Approach for CB2 Antagonist Search
 Article GPCR Structure-Based Virtual Screening Approach for CB2 Antagonist Search Jian-Zhong Chen, Junmei Wang, and Xiang-Qun Xie J. Chem. Inf. Model., 2007, 47 (4), 1626-1637 DOI: 10.1021/ci7000814 Publication
Article GPCR Structure-Based Virtual Screening Approach for CB2 Antagonist Search Jian-Zhong Chen, Junmei Wang, and Xiang-Qun Xie J. Chem. Inf. Model., 2007, 47 (4), 1626-1637 DOI: 10.1021/ci7000814 Publication
Supplementary Information
 Supplementary Information Two common structural motifs for TCR recognition by staphylococcal enterotoxins Karin Erica Johanna Rödström 1, Paulina Regenthal 1, Christopher Bahl 2, Alex Ford 2, David Baker
Supplementary Information Two common structural motifs for TCR recognition by staphylococcal enterotoxins Karin Erica Johanna Rödström 1, Paulina Regenthal 1, Christopher Bahl 2, Alex Ford 2, David Baker
Insight into the mechanism of lipids binding and uptake by CD36 receptor
 www.bioinformation.net Hypothesis Volume 11(6) Insight into the mechanism of lipids binding and uptake by CD36 receptor Zineb Tarhda*, Azeddine Ibrahimi Biotechnology lab (MedBiotech), Faculté de Médecine
www.bioinformation.net Hypothesis Volume 11(6) Insight into the mechanism of lipids binding and uptake by CD36 receptor Zineb Tarhda*, Azeddine Ibrahimi Biotechnology lab (MedBiotech), Faculté de Médecine
Chapter 2 Structural Insights into Activation and Allosteric Modulation of G Protein-Coupled Receptors
 Chapter 2 Structural Insights into Activation and Allosteric Modulation of G Protein-Coupled Receptors Andrew C. Kruse Abstract G protein-coupled receptors (GPCRs) are cell-surface receptors that regulate
Chapter 2 Structural Insights into Activation and Allosteric Modulation of G Protein-Coupled Receptors Andrew C. Kruse Abstract G protein-coupled receptors (GPCRs) are cell-surface receptors that regulate
AP Bio. Protiens Chapter 5 1
 Concept.4: Proteins have many structures, resulting in a wide range of functions Proteins account for more than 0% of the dry mass of most cells Protein functions include structural support, storage, transport,
Concept.4: Proteins have many structures, resulting in a wide range of functions Proteins account for more than 0% of the dry mass of most cells Protein functions include structural support, storage, transport,
If you like us, please share us on social media. The latest UCD Hyperlibrary newsletter is now complete, check it out.
 Sign In Forgot Password Register username username password password Sign In If you like us, please share us on social media. The latest UCD Hyperlibrary newsletter is now complete, check it out. ChemWiki
Sign In Forgot Password Register username username password password Sign In If you like us, please share us on social media. The latest UCD Hyperlibrary newsletter is now complete, check it out. ChemWiki
Structure of the measles virus hemagglutinin bound to the CD46 receptor. César Santiago, María L. Celma, Thilo Stehle and José M.
 Supporting Figures and Table for Structure of the measles virus hemagglutinin bound to the CD46 receptor César Santiago, María L. Celma, Thilo Stehle and José M. Casasnovas This PDF file includes: Supplementary
Supporting Figures and Table for Structure of the measles virus hemagglutinin bound to the CD46 receptor César Santiago, María L. Celma, Thilo Stehle and José M. Casasnovas This PDF file includes: Supplementary
1-To know what is protein 2-To identify Types of protein 3- To Know amino acids 4- To be differentiate between essential and nonessential amino acids
 Amino acids 1-To know what is protein 2-To identify Types of protein 3- To Know amino acids 4- To be differentiate between essential and nonessential amino acids 5-To understand amino acids synthesis Amino
Amino acids 1-To know what is protein 2-To identify Types of protein 3- To Know amino acids 4- To be differentiate between essential and nonessential amino acids 5-To understand amino acids synthesis Amino
Lecture 15. Membrane Proteins I
 Lecture 15 Membrane Proteins I Introduction What are membrane proteins and where do they exist? Proteins consist of three main classes which are classified as globular, fibrous and membrane proteins. A
Lecture 15 Membrane Proteins I Introduction What are membrane proteins and where do they exist? Proteins consist of three main classes which are classified as globular, fibrous and membrane proteins. A
Term Definition Example Amino Acids
 Name 1. What are some of the functions that proteins have in a living organism. 2. Define the following and list two amino acids that fit each description. Term Definition Example Amino Acids Hydrophobic
Name 1. What are some of the functions that proteins have in a living organism. 2. Define the following and list two amino acids that fit each description. Term Definition Example Amino Acids Hydrophobic
Gentilucci, Amino Acids, Peptides, and Proteins. Peptides and proteins are polymers of amino acids linked together by amide bonds CH 3
 Amino Acids Peptides and proteins are polymers of amino acids linked together by amide bonds Aliphatic Side-Chain Amino Acids - - H CH glycine alanine 3 proline valine CH CH 3 - leucine - isoleucine CH
Amino Acids Peptides and proteins are polymers of amino acids linked together by amide bonds Aliphatic Side-Chain Amino Acids - - H CH glycine alanine 3 proline valine CH CH 3 - leucine - isoleucine CH
Lecture 4: 8/26. CHAPTER 4 Protein Three Dimensional Structure
 Lecture 4: 8/26 CHAPTER 4 Protein Three Dimensional Structure Summary of the Lecture 3 There are 20 amino acids and only the L isomer amino acid exist in proteins Each amino acid consists of a central
Lecture 4: 8/26 CHAPTER 4 Protein Three Dimensional Structure Summary of the Lecture 3 There are 20 amino acids and only the L isomer amino acid exist in proteins Each amino acid consists of a central
Midterm 1 Last, First
 Midterm 1 BIS 105 Prof. T. Murphy April 23, 2014 There should be 6 pages in this exam. Exam instructions (1) Please write your name on the top of every page of the exam (2) Show all work for full credit
Midterm 1 BIS 105 Prof. T. Murphy April 23, 2014 There should be 6 pages in this exam. Exam instructions (1) Please write your name on the top of every page of the exam (2) Show all work for full credit
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10913 Supplementary Figure 1 2F o -F c electron density maps of cognate and near-cognate trna Leu 2 in the A site of the 70S ribosome. The maps are contoured at 1.2 sigma and some of
doi:10.1038/nature10913 Supplementary Figure 1 2F o -F c electron density maps of cognate and near-cognate trna Leu 2 in the A site of the 70S ribosome. The maps are contoured at 1.2 sigma and some of
Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular
 Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular weight markers (M). Supplementary Figure-2. Overlay of
Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular weight markers (M). Supplementary Figure-2. Overlay of
Sheet #1 Dr. Mamoun Ahram 01/07/2014. Amino Acids
 Amino Acids Dr s office in physiology department in floor #1 Office hours: every day after 1 *How to study biochemistry for Dr. Mamoun s material: Slides will be sent before the lecture, try to read it
Amino Acids Dr s office in physiology department in floor #1 Office hours: every day after 1 *How to study biochemistry for Dr. Mamoun s material: Slides will be sent before the lecture, try to read it
Levels of Protein Structure:
 Levels of Protein Structure: PRIMARY STRUCTURE (1 ) - Defined, non-random sequence of amino acids along the peptide backbone o Described in two ways: Amino acid composition Amino acid sequence M-L-D-G-C-G
Levels of Protein Structure: PRIMARY STRUCTURE (1 ) - Defined, non-random sequence of amino acids along the peptide backbone o Described in two ways: Amino acid composition Amino acid sequence M-L-D-G-C-G
Biology 2E- Zimmer Protein structure- amino acid kit
 Biology 2E- Zimmer Protein structure- amino acid kit Name: This activity will use a physical model to investigate protein shape and develop key concepts that govern how proteins fold into their final three-dimensional
Biology 2E- Zimmer Protein structure- amino acid kit Name: This activity will use a physical model to investigate protein shape and develop key concepts that govern how proteins fold into their final three-dimensional
Structure and Activation of the Visual Pigment Rhodopsin
 Seminar 02.12.2011 Structure and Activation of the Visual Pigment Rhodopsin by Steven O.Smith, 2010 Constantin Schneider FB Physik, FU Berlin GPCR: Overview Rhodopsin is a G protein-coupled receptor (GPCR)
Seminar 02.12.2011 Structure and Activation of the Visual Pigment Rhodopsin by Steven O.Smith, 2010 Constantin Schneider FB Physik, FU Berlin GPCR: Overview Rhodopsin is a G protein-coupled receptor (GPCR)
Amino Acids. Amino Acids. Fundamentals. While their name implies that amino acids are compounds that contain an NH. 3 and CO NH 3
 Fundamentals While their name implies that amino acids are compounds that contain an 2 group and a 2 group, these groups are actually present as 3 and 2 respectively. They are classified as α, β, γ, etc..
Fundamentals While their name implies that amino acids are compounds that contain an 2 group and a 2 group, these groups are actually present as 3 and 2 respectively. They are classified as α, β, γ, etc..
AA s are the building blocks of proteins
 Chamras Chemistry 106 Lecture otes Chapter 24: Amino Acids, Peptides, and Proteins General Formula: () n (') α-amino Acids: (n = 1) Example: Amino Acids and Proteins: Glycine Alanine Valine AA s are the
Chamras Chemistry 106 Lecture otes Chapter 24: Amino Acids, Peptides, and Proteins General Formula: () n (') α-amino Acids: (n = 1) Example: Amino Acids and Proteins: Glycine Alanine Valine AA s are the
