TetX is a Flavin-Dependent Monooxygenase Conferring. Resistance to Tetracycline Antibiotics*
|
|
|
- Alaina Parks
- 5 years ago
- Views:
Transcription
1 JBC Papers in Press. Published on September 27, 2004 as Manuscript M TetX is a Flavin-Dependent Monooxygenase Conferring Resistance to Tetracycline Antibiotics* Wangrong Yang, Ian F. Moore, Kalinka P. Koteva, David C. Bareich, Donald W. Hughes, and Gerard D. Wright Antimicrobial Research Centre, Department of Biochemistry and Biomedical Sciences, and Department of Chemistry, McMaster University, 1200 Main St. W., Ontario L8N 3Z5, Canada Running Title: Tetracycline degradation by TetX To whom correspondence should be addressed. Tel.: (ext ); Fax: ; wrightge@mcmaster.ca. Copyright 2004 by The American Society for Biochemistry and Molecular Biology, Inc.
2 SUMMARY The tetracycline antibiotics block microbial translation and constitute an important group of antimicrobial agents that find broad clinical utility. Resistance to this class of antibiotics is primarily the result of active efflux or ribosomal protection; however a novel mechanism of resistance has been reported to be oxygen-dependent destruction of the drugs catalyzed by the enzyme TetX. Paradoxically, the tetx genes have been identified on transposable elements found in anaerobic bacteria of the genus Bacteroides. Overexpression of recombinant TetX in Escherichia coli followed by protein purification revealed a stoichiometric complex with flavin adenine dinucleotide. Reconstitution of in vitro enzyme activity demonstrated a broad tetracycline antibiotic spectrum and a requirement for molecular oxygen and NADPH in antibiotic degradation. The tetracycline products of TetX activity were unstable at neutral ph, but mass spectral and NMR characterization under acidic conditions supported initial mono-hydroxylation at position 11a followed by intramolecular cyclization and non-enzymatic breakdown to other undefined products. TetX is therefore an FAD-dependent monooxygenase. The enzyme not only catalyzed efficient degradation of a broad range of tetracycline analogues, but also conferred resistance to these antibiotics in vivo. This is the first molecular characterization of an antibiotic inactivating monooxygenase, the origins of which may lie in environmental bacteria. 2
3 (Introduction) Tetracyclines represent one of the most successful classes of antibiotics used in the past 50 years. Since the first identification of chlortetracycline in 1948 from extracts of Streptomyces aureofaciens, numerous analogues, both natural and semisynthetic, have found clinical use (Fig. 1). This class of antibiotic has been a mainstay in the treatment of bacterial infections, being highly prized for their broad spectrum of antimicrobial activity, oral availability, and low cost. Tetracyclines are also widely used in agriculture to treat bacterial infections of plants (1), in aquaculture (2), and in animal growth (3). Given this intensive use, it is not surprising that tetracycline resistance has emerged over the past decades as a significant issue in the use of this class of antibiotic (for reviews see refs. (4-7). Tetracycline resistance in the clinic manifests itself through two primary mechanisms: active efflux and ribosomal protection. Efflux proteins in particular are major sources of resistance, responsible for the bulk of clinical failures of this class. These proteins belong to the major facilitator family of integral membrane efflux enzymes that couple the energetically unfavorable movement of tetracycline with the proton motive force to pump tetracyclines out of the cell against the concentration gradient. Ribosomal protection mechanisms on the other hand harness soluble structural homologues of elongation factors to destabilize the interaction between tetracyclines and their cellular target the ribosome. In neither efflux nor ribosomal protection mechanisms however, is the concentration of tetracycline in the environment altered. In contrast, resistance to other antibiotics such as the β-lactams and the aminoglycosides occurs primarily via the destruction or covalent modification of the 3
4 antibiotics, which effectively decreases the local concentration of antibiotic. Such a mechanism was unknown for the tetracyclines except for a series of reports over a decade ago describing a gene, tetx, that encoded a putative 388 amino acid NADPH 1 -requiring enzyme that was associated with tetracycline resistance (8-10). Amino acid sequence analysis revealed putative FAD-binding and monooxygenase fold domains (Fig. 2). The tetx gene was identified in transposons Tn4351 (10) and Tn4400 (9) harbored by the obligate anaerobe Bacteroides fragilis. Transfer of this gene to aerobically growing Escherichia coli uncovered a cryptic tetracycline resistance activity that was associated with destruction of the antibiotic and a commensurate darkening of the growth medium (9,10). Preliminary biochemical studies using crude cell free extracts revealed that both oxygen and NADPH were required for tetracycline resistance activity (11). More recently, two orthologues of the original gene, tetx1 and tetx2, were identified in another Bacteroides transposon, CTnDOT (12). The predicted TetX2 is 99% identical to the original TetX while TetX1 is an N-terminal truncate (359 amino acids) with 66% identity to the other proteins lacking the FAD-binding domain (Fig. 2). We report here the overexpression of TetX in E. coli and characterize it as a broad spectrum tetracycline degrading enzyme operating by an unprecedented mechanism. TetX is a flavin-dependent monooxygenase that regioselectively hydroxylates the tetracycline substrate resulting in an unstable compound that undergoes non-enzymatic decomposition. 4
5 EXPERIMENTAL PROCEDURES Reagents- Tetracyclines, NADP +, NADPH, and glucose-6-phosphate were from Sigma (Oakville, ON). Glucose-6-phosphate dehydrogenase was from Roche Diagnostics (Laval, PQ). Expression of recombinant TetX proteins- Plasmids encoding various tetx genes were the generous gifts of A. Salyers and N. Shoemaker, University of Wisconsin. Plasmid pdb1 was created by digestion of plasmid pbs2 (11) with Hind III and EcoR I followed by ligation of the tetx containing fragment into puc18. Overexpression constructs of the tetx, tetx1, and tetx2 genes were amplified by PCR using the oligonucleotide primers listed in Table 1 of Supplementary Information and using the appropriate plasmids as templates. PCR was performed on a Progeny 96 well themocycler with 95 C 1 minute, 52 C 1 minute, 72 C 1.5 minutes and was repeated for 30 cycles. The PCR products were excised from a 0.8% agarose gel, extracted by Qiagen QIAquick Gel extraction kit, digested with Nde I and Hind III, and ligated into plasmid pet28 (Novagen) digested with the same restriction enzymes generating fusion constructs that give an N-terminal hexa-his tagged protein for ease of purification. Plasmids were used to transform E. coli BL21(DE3) and were selected by kanamycin resistance. The absence of adventitious mutations during amplification was confirmed by complete gene sequencing. A single colony of E. coli BL21 (DE3) containing the appropriate plasmid constructs was used to inoculate 25 ml of Luria Broth supplemented with 50 µg/ml kanamycin and incubated at 37 C and 250 rpm for hours. Ten ml of this culture were used to inoculate 1 L of Luria Bertani medium supplemented with 50 µg/ml 5
6 kanamycin. The cultures were grown at 37 C, 250 rpm to an OD600 of approximately 0.6, followed by the addition of sterile isopropyl-β-d-thiogalactopyrandoside to 1mM. The culture was then incubated overnight with 250 rpm shaking at 16 C. Cells were collected by centrifugation at 8000 rpm for 5 minutes, resuspended in 10 ml of 0.1 mm dithiothreitol, 1 mm phenylmethanesulfonylfluoride, 1 mm EDTA, 20 mm HEPES ph 8.0 and lyzed by 3 passes through a French pressure cell at a maximum pressure of 20,000 psi. The cell lysate was clarified by centrifugation at 15,000 rpm for 15 minutes. The supernatant was applied to a 1 ml Ni-agarose column equilibrated with 20 mm HEPES, ph 8.0. Enzyme was eluted by application of a linear gradient with 20 mm HEPES mm imidazole. Fractions were analyzed by electrophoresis through 11% sodium dodecylsulfate polyacrylamide gels to assess purity of the protein. If required, an additional chromatographic step consisting of application of the pooled fractions onto a Mono Q column, equilibrated with 20 mm Tris HCl ph 8.0 and elution with a gradient to 1 M NaCl. Purified TetX and TetX2 (but not TetX1) were yellow in color and all proteins were stored in 20 mm HEPES ph 8.0. A 1L culture yielded 6 mg of pure protein (Figure S20). Analysis of flavin cofactor content- Purified TetX2 was boiled to denature the protein and briefly centrifuged to remove the precipitate. A sample of the supernatant was applied to a C18 column (Alltech Econosil, 10U, 250 x 22 mm) equilibrated with 5 mm ammonium acetate (ph 6.0) and the bound flavin was separated by a linear gradient of 5 mm ammonium acetate (ph 6.0) to 100% methanol in 20 minutes with a flow rate of 6
7 1ml/ minute while monitoring the absorbance at 451 nm. Commercial FMN, FAD, and riboflavin served as standards. Spectrophotometric assay of TetX activity- Each 100 µl reaction in a 96 well microtitre plate included 1 mm NADPH and up to 3 mm tetracycline substrate in 25 mm TAPS ph 8.5. The decrease in absorbance at 340 nm (NADPH oxidation) upon antibiotic inactivation or the change in the absorbance of oxytetracycline at 400 nm (ε 400 = 1080 M -1 cm -1 ) was monitored using a Molecular Devices SpectraMax Plus microtitre plate reader. Steady state kinetic parameters were determined by fitting initial rate (v) data to the standard Michaelis-Menten equation using the Grafit 4 software(13): v = k cat [E o ][S]/([S] + K m ) where E o is the total enzyme concentration. Microbiological assay- The effects of TetX activity on the antimicrobial properties of tetracyclines were assessed by a microbiological disk assay. Inactivation reactions contained 3 mm oxytetracycline, 1 mm NADP +, 40 mm glucose-6-phosphate, 0.3 U glucose-6-phosphate dehydrogenase, and 10 µg of purified TetX2, 25 mm TAPS ph 8.5 in a total volume of 0.1 ml. A 15 µl aliquot was applied to a sterile filter paper disc (5 mm), and air dried for minutes. The disk was then placed on a tryptic soy agar plate inoculated with an overnight culture of tetracycline-sensitive Micrococcus luteus diluted to an OD nm of and incubated at 30 C for 48 hours. HPLC separation of products of tetracycline inactivation- The products of tetracycline inactivation were separated by reverse phase HPLC using a Dionex Acclaim 120 C18 column (3 µm 120 Å 4.6 mm x 150 mm). The column was equilibrated with 7
8 H 2 O plus 0.05% trifluoroacetic acid and tetracyclines and the products of inactivation were eluted using a linear gradient to 95% CH 3 CN plus 0.05 % trifluoroacetic acid over 14 minutes at a flow rate of 1 ml/minute. NMR analysis of oxytetracycline and inactivation products- A solution of 4 mm NADP +, 40 mm glucose-6- phosphate and 20 units of glucose-6-phosphate dehydrogenase in 25 mm TAPS ph 8.5 was incubated at 37 C for 20 minutes to generate NADPH. MgCl 2 (1 mm) was added followed by 1.7 mg of TetX2 and 5 mg of oxytetracycline. The total reaction volume was 5 ml and the mixture was incubated at room temperature. The progress of the reaction was monitored by reversed-phase HPLC. Following completion of the reaction, concentrated HCl was added to a final concentration of 0.1 M. The crude reaction mixture was applied to a C-18 Sep-Pak mini column equilibrated in water. The product of TetX2 catalysis, P1 was eluted with water, and the purity of the sample verified by HPLC. A total of four reactions were run and purified P1 was combined and lyophilized. A final purification by preparative scale HPLC was performed prior to NMR analysis. The lyophilized product was dissolved in 0.1 M DCl/D 2 O and the 1 H and 13 C NMR spectra were recorded on a Brucker AV 600 instrument. Mass spectrometry- Mass spectrometry of the products of enzymatic reactions was performed on an Applied Biosystems Q Trap LC-MS system. High resolution mass spectrometry by Dr. K. Green at the McMaster Regional Centre for Mass Spectrometry on a Waters-Micromass Global Ultima Quadrupole Time of Flight mass spectrometer. RESULTS 8
9 TetX is a flavoprotein- We first subcloned tetx into puc18 creating the plasmid pdb1. This gene conferred tetracycline resistance to E. coli W3110 and resulted in discoloration of the medium associated with antibiotic inactivation (Fig. 3). To obtain purified proteins in sufficient quantity for molecular studies, we prepared separate constructs of N-terminal hexa-his tagged TetX, TetX1, and TetX2 in E. coli under control of a T7 promoter. Only the constructs expressing TetX and TetX2 conferred tetracycline resistance. Purified TetX and TetX2, but not the truncated TetX1, were visibly yellow in solution, indicative of bound flavin cofactor (see below). TetX1 is therefore an inactive protein and its presence in the CTnDOT transposon may be a relic of an incomplete gene duplication event. We selected the TetX2 expressing construct for all further studies. Purification of TetX2 gave a protein of the predicted 44 kda by SDS polyacrylamide gel electrophoresis, and analytical gel filtration was consistent with a monomeric protein of the correct size. Solutions of purified TetX2 were yellow in color and the UV-visible spectrum was consistent with the presence of a flavin cofactor, with absorbance maxima at 366 nm and 445 nm (Fig. 4). Denaturation of the enzyme followed by reverse phase HPLC of the soluble material identified FAD as the bound cofactor. The FAD-TetX2 stoichiometry was found to vary between batches (~60->80%) and addition of exogenous FAD to 2 µm was found to stabilize activity, therefore this coenzyme was typically added to purified protein during storage in the dark. TetX requires NADPH and O 2 for enzyme activity- Primary sequence analysis and preliminary studies by the Salyers group using cell extracts predicted that TetX might be an NADP + -requiring oxidoreductase (11). We confirmed with purified enzyme that 9
10 NADPH was essential for oxytetracycline degradation activity and could not be substituted by NADH. Furthermore, degassing of solutions followed by incubation of the reaction in an N 2 -only atmosphere resulted in no inactivation of oxytetracycline (not shown) demonstrating that O 2 was also essential for oxytetracycline inactivation, and consistent with the observation that only aerobically grown cultures showed tetracycline resistance (10). A survey of the influence of initial rate on ph demonstrated that the enzyme showed maximal activity at ph 8.5. TetX catalyzes the inactivation of a broad spectrum of tetracycline antibiotics- The tetracycline inactivation activity of TetX2 was established by a series of biochemical assays including UV-visible spectroscopy and reverse phase HPLC. Tetracyclines show two absorption maxima, one at 260 nm and another at 363 nm. The β-tricarbonyl chromophore (ring A) is responsible for the 260 nm absorbance while the aryl betadiketone chromophore (rings B, C, D) is responsible for the 363 nm absorbance and the yellow color of tetracyclines. Incubation of tetracyclines and TetX2 under assay conditions (presence of NADPH and O 2, ph 8.5) resulted in the disappearance of the absorbance maximum at 363 nm, a more modest decrease in the absorbance maximum at 260 nm, and a broad low intensity increase in the absorbance at wavelengths greater than 430 nm (Fig. 5). These changes in absorbance as a result of enzyme action provided a means to continuously monitor the progress of the reaction. Since NADPH has a maximum absorbance at 340 nm, the absorbance of both substrates overlap in the 360 nm region. Therefore, the absorbance at 400 nm (ε 400 of oxytetracycline is 1080 M -1 cm -1 ) (See Fig 5 10
11 for spectrum) was chosen to monitor the enzyme activity in continuous assays in 96 well microtitre plates. The progress of the TetX2 catalyzed reaction could also be monitored by the disappearance of the antibiotic substrate (S) and the appearance of product peaks by reverse phase HPLC (Fig. 6). Using oxytetracycline (Fig. 1) as a model substrate, the antibiotic was converted to two products P1 and P2 (Fig. 6). The temporal separation between the appearance of P1 and P2 implied that P2 was derived from P1 (Fig 6). Further analysis of the conversion of P1 to P2 revealed that this process was enzyme independent and accelerated at neutral ph, but was slower at acidic ph (data not shown). Therefore, the exclusive product of TetX2 activity is P1. TetX2-catalyzed modification of oxytetracycline was associated with loss of antibiotic activity, therefore TetX has been rigorously established as a tetracycline inactivating enzyme. Purified P1 did not show any antimicrobial activity in our assays, however since we demonstrated that this compound is unstable at neutral ph, we cannot conclusively state that P1 is devoid of antimicrobial activity itself, or if downstream products such as P2 are the inactive compounds. We do know that initial modification of antibiotic catalyzed by TetX2 is essential for conversion of tetracycline into the inactive metabolites that eventually result in the black pigment seen in cell free extracts that are characteristic of the presence of this enzyme. Preliminary studies indicate that this black pigment is a high molecular weight polymer of undefined structure (unpublished results). TetX2 catalyzed the efficient inactivation of a variety of tetracycline antibiotics (Table 1 and Table 2). The enzyme showed a maximum of 5-fold discrimination between these substrates in the steady state, including natural products such as 11
12 tetracycline, oxytetracycline, and demeclocycline as well as semi-synthetic compounds such as minocycline and doxycycline (Table 1). TetX2 is a monooxygenase that oxidizes oxytetracycline- LC-MS analysis of the first product peak of oxytetracycline degradation, P1 (Fig. 6B) gave an m/z value of 477 in positive ion mode, equivalent to the addition of 16 Da to oxytetracycline (mass 461). This is consistent with monooxidation of the tetracycle. The oxidized product P1 is unstable under the conditions (ph 8.5) of the enzymatic reaction. However, it was found that decomposition of P1 could be prevented by acidifying the product reaction mixture to ph 1. Therefore purified P1 was kept in 0.1M HCl and characterization by NMR was performed in 0.1M DCl/D 2 O. High resolution positive ion ESI-MS performed on purified P1 gave an m/z of The expected mass of positively charged P1 corresponding to C 22 H 25 N 2 O + 10 is The structure of P1- To determine the identity of P1 the 1 H, 13 C, COSY, HSQC, HMBC and NOE spectra of P1 were determined and compared with the corresponding spectra of oxytetracycline determined under identical conditions (see Supplementary information for complete spectra, Figures S1 to S19 and Tables S2 to S5 for complete assignments). The 1D 1 H NMR spectra of oxytetracycline and P1 are shown in Figure 7. There are the same number of proton resonances observed in the 1 H NMR spectrum of both oxytetracycline and P1. The aromatic region of the 1 H NMR spectrum of P1 (H 8 (7.60 ppm), H 7 (7.12 ppm), H 9 (7.04 ppm)) shows no change in the observed coupling pattern or significant changes in chemical shift when compared with oxytetracycline. This is significant since a large number of flavin monooxygenases hydroxylate activated aromatic rings similar to ring D of oxytetracycline. The methine protons of P1 show 12
13 significant changes in chemical shift and coupling compared with oxytetracycline. In oxytetracycline (0.1M DCl/D 2 O) the four non-aromatic methine protons are observable as: a doublet at 4.30 ppm, (H 4, J 4,4a = 1.4 Hz), a doublet of doublets at 3.87 ppm (H 5, J 5,4a = 11.4 Hz, J 5,5a = 8.3 Hz), a doublet at 2.89 ppm (H 5a ) and a doublet of doublets at 2.87 ppm (H 4a ). In P1 four methine proton signals are observed as a doublet at 4.16 ppm (J = 8.9 Hz), a doublet at 3.97 ppm (J = 1.4 Hz), a doublet of doublets at 3.67 ppm (J = 8.9, 1.4 Hz) and a singlet at 2.85 ppm. The COSY spectrum of P1 confirms the coupling interaction between the doublet of doublets at 3.67 ppm and the doublets at 4.16 ppm and 3.97 ppm (Figures S5-S7). The collapse of one doublet to a singlet and the loss of one of the doublet of doublets from the oxytetracycline spectrum means the coupling interaction of either H 5a or H 4 has been removed from the system of methine protons in P1. Either the C 5a -H 5a or C 4 -H 4 bond is broken and a new C-H bond is formed somewhere else in the molecule during the conversion of oxytetracycline to P1 or the dihedral angle between H 5a or H 4 and their adjacent proton is 90. An NOE difference spectrum of oxytetracycline obtained by saturating the protons of the methyl group at carbon 6 (6- CH 3, 1.72 ppm) reveals enhancement of the resonances of H 7 (7.14 ppm), H 5 (3.876 ppm) and H 5a (2.90 ppm) (Figure S8). By saturating the protons of 6-CH 3 (1.45 ppm) in P1 enhancements of H 7 (7.12 ppm) and the methine signals at 4.16 ppm and 2.85 ppm are observed (Figure S9). Since the same number of enhancements is observed in both oxytetracycline and P1 a change in bonding around 6-CH 3 is unlikely and suggests the resonance for H 5a is the singlet at 2.85 ppm. The 13 C NMR spectrum of P1 contains 21 signals, the same number observed for oxytetracycline (Figures S10-S12). In the 13 C NMR spectrum of oxytetracycline there 13
14 are five carbonyl resonances assigned to C11, C12, C1, C3 (shown as enol tautomers) and CONH 2 (14). In P1 there are only four, suggesting one of the keto/enol carbons has undergone a hybridization change. Concomitant with the loss of a carbonyl resonance is the appearance of a new signal at ppm in the 13 C NMR spectrum of P1. The 2-D HSQC and HMBC proton-carbon correlation spectra help to assign the 13 C resonances and identify the site of hydroxylation. As expected the HSQC spectrum of P1 shows ten proton-carbon correlations (Figure S13). These correlations fix 3 of the 6 aromatic carbons: C8 ( ppm), C9 ( ppm),c7 ( ppm) (Figure S14). The 6-CH 3 carbon resonance is ppm and the two carbon resonances of the dimethyl amine group are ppm and ppm (Figure S15). The 4 methine carbons resonances are ppm, ppm, ppm and ppm. The HMBC spectrum of P1 contains several 2 and 3 bond proton-carbon correlations (Figures S16 to S19). The protons of 6-CH 3 are expected to show three correlations, a 2 bond coupling to C6 and two 3 bond couplings to C6a and C5a. All three couplings are observed in the HMBC spectra allowing the following assignments, C6a ( ppm), C6 (79.63 ppm) and C5a (59.77 ppm) (Figure S16). The singlet at 2.85 ppm in the 1 H NMR spectrum can now be assigned to H 5a since the HSQC spectrum shows a correlation between C5a (59.77 ppm) and the singlet at 2.85 ppm. Taken with the NOE data this indicates that in P1 the dihedral angle between H5a and H5 is close to 90 explaining the lack of observed coupling between them. The remaining proton resonances can then be assigned as follows, H 5, 4.16 ppm, (J H5,H4a = 8.9 Hz), H 4, 3.97 ppm, (J H4,H4a = 1.8 Hz) and H 4a, 3.67 ppm (J H4a,H5 = 8.9 Hz, J H4aH4 = 1.8 Hz). Using the 14
15 one bond correlations in the HSQC spectra the remaining methine carbons can be assigned as follows: C4 (69.27 ppm), C5 (66.33 ppm) and C4a (43.37 ppm). Proton H5a is expected to show three 2 bond 1 H- 13 C couplings to C11a, C5 and C6 and four 3 bond couplings to C6a, C4a, C11 and C12. Five correlations are observed in the HMBC spectrum, ppm, ppm, ppm, and the already assigned ppm (C5) and ppm (C4a) (Figures S17, S18). Correlations to the already assigned C6a ( ppm) and C6 (79.63 ppm) are not observed. The three resonances at ppm, ppm and ppm therefore correspond to C11, C12 and C11a. These are significant chemical shift changes compared to oxytetracycline, where C11 is ppm, C ppm and C11a ppm. C11a and C12 have undergone a hybridization change since their signals have shifted upfield. Flavin dependent hydroxylases act on activated double bonds. The most widely studied reactions are those where hydroxylation of an aromatic substrate occurs. A typical example is phenol hydroxylase where the substrate is hydroxylated ortho to the hydroxyl group. An intermediate α-hydroxy ketone is formed during the reaction that loses a proton in a subsequent step to regenerate the aromatic ring. The aromatic ring of oxytetracycline is unchanged upon transformation to P1. The enol tautomer of a β- diketone can be considered an activated double bond. Hydroxylation of C11a of the β- diketone would generate an α-hydroxy ketone at C11a-C12. Unlike the substrates for aromatic hydroxylases there is no proton α to the carbonyl group at C11a to deprotonate and reform the double bond of the enol. C11a changes the hybridization from sp2 to sp3 during hydroxylation and is consistent with the change in chemical shift from ppm to ppm. The formation of an α-hydroxy ketone is expected to shift the C12 15
16 resonance downfield not upfield from ppm to ppm. However, there is a unique property of 6-hydroxytetracyclines that explains the shift of the C12 resonance upfield. Non-enzymatic oxidation reactions of tetracyclines at C11a are known. In 1963 Blackwood et. al prepared several C11a halogenated derivatives of various tetracyclines (15). These included: 11a-fluorotetracycline, 11a-fluoro-6-oxytetracycline, 11achlorotetracycline and 11a-chloro-6-oxytetracycline. They observed that the 11ahalogenated tetracycline derivatives with a hydroxyl group at C6 (like oxytetracycline) formed an intramolecular 6,12-hemiketal with the carbonyl group at C12. The 11ahalogenated tetracycline derivatives without a hydroxyl group at C6, 11a-fluoro-6- demethyl-6-deoxytetracycline and 11a-fluoro-6-deoxy-oxytetracycline did not form an intramolecular hemiketal and a free ketone group at C12 was observed in the IR spectrum of these compounds. We therefore propose that TetX2 hydroxylates C11a of oxytetracycline and that the ketone initially formed at C12 undergoes rapid conversion to the 6,12-hemiketal. This is consistent with a chemical shift value of ppm for C12. Formation of the intramolecular hemi-ketal explains the acid stability of P1 and accounts for the loss of a carbonyl signal in the NMR. Also, formation of the intramolecular bridge adds a degree of rigidity to the molecule. Comparison of MM2 energy minimized structures of 11a-hydroxy-oxytetracycline with and without the 6,12-hemiketal shows that formation of the intramolecular bond changes the dihedral angle between H 5a,H 5 from 145º to 87º. This is consistent with the lack of coupling observed between H 5a and H 5 in the 1 H NMR spectrum. The remaining 13 C assignments are as follows: C11 ( ppm), C1 ( ppm), C3 ( ppm), CONH 2 ( ppm), C10 ( ppm), C10a ( ppm), 16
17 C2 (98.96 ppm) and C12a (74.10 ppm). These carbons do not show any significant chemical shift difference from oxytetracycline. DISCUSSION The enzymes TetX and TetX2 are FAD-requiring monooxygenases that inactivate a broad selection of tetracycline antibiotics. The presence of this prosthetic group suggested that tetracycline inactivation could be the result of reductive electron transfer or hydroxylation reactions. We demonstrated that TetX requires both NADPH and molecular oxygen to inactivate tetracycline consistent with a role as a monooxygenase. These results predict NADPH reduction of the flavin cofactor to FADH 2, followed by reaction of O 2 with the resulting electron rich isoaloxazine to form a reactive FAD-4ahydroperoxide. Our work demonstrates that tetracycline inactivation is triggered by regiospecific hydroxylation at C11a (Scheme 1), which could proceed via initial epoxidation of the C11a-C12 enol. The absence of the predicted carbonyl signature of 11a-hydroxy-oxytetracycline in the 13 C NMR spectrum suggests that the acid stabilized product P1 is the 6-12-hemiketal. We were unable to directly assess the antibiotic activity of P1 as this compound spontaneously decomposed to downstream inactive products. However, hydroxylation at C11a would likely impact the Mg 2+ chelating properties of tetracycline, which are a requirement for ribosome binding (16). LC-MS analysis of the HPLC peak P2 revealed loss of a molecule of water, however we were unable to purify and study this compound as this peak contained more than one unstable compound (not shown). The expression of the hydroxylase TetX in E. coli aerobically grown in the presence of tetracycline, results in the formation an amorphous black pigment, indicative of a complex polymeric 17
18 structure. Therefore, initial hydroxylation catalyzed by this enzyme precipitates a cascade of molecular events that result in tetracycline deactivation. This is the first example to our knowledge of a flavin monooxygenase catalyzing antibiotic inactivation. TetX is capable of recognizing and inactivating a broad range of tetracycline antibiotics of both natural and semi-synthetic origin. In vitro analysis of specificity as measured by k cat /K m using purified enzymes showed only a 6-fold difference between the best substrate (oxytetracycline) and the poorest (minocycline). Correspondingly, minocycline is also the poorest substrate for the enzyme in vivo with an MIC of 8 µg/ml while the presence of the enzyme confers resistance to 256 µg/ml oxytetracycline. The correlation is however not complete as tetracycline, which is comparable to minocycline as a substrate of the purified enzyme, nonetheless is robustly resisted in vivo. The disparity in MIC is therefore not reflected adequately in disparity of similar magnitude in steady state kinetic parameters. The molecular basis for this observation is obscure at present but the MIC data may also reflect additional downstream processing and inactivation of tetracycline into multiple products. The paradoxical discovery of the tetx genes, which we have unambiguously shown encode oxygen-requiring monooxygenases, in the obligate anaerobe B. fragilis speaks to the extensive exchange of genetic material in the microbial world. The tetx genes were all localized on Bacteroides transposons that mobilize genes for exchange. The G+C content of the tetx genes is marginally lower (37%) than the B. fragilis genome (42%), however it is within the reported values of other bacteria of the same genus (11). BLAST search reveals the presence of highly homologous gene products (E values < ) of TetX in the sequenced genomes of the aerobic soil bacteria Cytophaga hutchinsonii 18
19 (phylogenetically related to the Bacteroides), Streptomyces coelicolor and Streptomyces avermitilis, suggesting that this family of enzymes is widespread in the environment. Aromatic polyketide natural products that resemble tetracyclines, for example the anthracyclines such as daunorubicin, actinorhodin, granaticin, and many others, are produced by numerous Streptomyces. TetX-like monooxygenases that are present in these bacteria may reflect the density of such molecules in the environment and the requirement to oxidatively modify them, perhaps not always as a means of detoxification, but also in biosynthesis. While the G+C content of tetx genes isolated from Bacteroides do not approach that of the Streptomyces (>70%), they are similar to the G+C content of Cytophaga (33-42%). Like other antibiotics such as the aminoglycosides (17) and the glycopeptides (18), the origins of this mechanism of tetracycline resistance may be the environment. ACKNOWLEDGMENTS We thank A. Salyers and N. Shoemaker for gifts of tetx plasmids. This article is dedicated to Professor Christopher Walsh on the occasion of his 60 th birthday. 19
20 REFERENCES 1. McManus, P. S., Stockwell, V. O., Sundin, G. W., and Jones, A. L. (2002) Annu Rev Phytopathol 40, Schmidt, A. S., Bruun, M. S., Dalsgaard, I., and Larsen, J. L. (2001) Appl Environ Microbiol 67, Sengelov, G., Halling-Sorensen, B., and Aarestrup, F. M. (2003) Vet Microbiol 95, Chopra, I., and Roberts, M. (2001) Microbiol Mol Biol Rev 65, McMurray, L. M., and Levy, S. B. (2000) in Gram-positive pathogens (Fischetti, V. A., Novick, R. P., Ferretti, J. J., Portnoy, D. A., and Rood, J. I., eds), pp , ASM Press, Washington, D.C. 6. Schnappinger, D., and Hillen, W. (1996) Arch. Microbiol. 165, Sum, P. E., and Petersen, P. (1999) Bioorg Med Chem Lett 9, Guiney, D. G., Jr., Hasegawa, P., and Davis, C. E. (1984) Plasmid 11, Park, B. H., and Levy, S. B. (1988) Antimicrobial Agents and Chemotherapy 32, Speer, B. S., and Salyers, A. A. (1988) Journal of Bacteriology 170, Speer, B. S., Bedzyk, L., and Salyers, A. A. (1991) Journal of Bacteriology 173, Whittle, G., Hund, B. D., Shoemaker, N. B., and Salyers, A. A. (2001) Appl Environ Microbiol 67, Leatherbarrow, R. J. (2000), 4.0 Ed., Erithacus Software Ltd, Staines, UK 20
21 14. Asleson, G. L., and Frank, C. W. (1975) J. Am. Chem. Soc. 97, Blackwood, R. K., Beereboom, J. J., Rennhard, H. H., Schach von Wittenau, M., and Stephens, C. R. (1963) J. Am. Chem. Soc. 85, Brodersen, D. E., Clemons, W. M., Jr., Carter, A. P., Morgan-Warren, R. J., Wimberly, B. T., and Ramakrishnan, V. (2000) Cell 103, Benveniste, R., and Davies, J. (1973) Proc. Natl. Acad. Sci. U S A 70, Marshall, C. G., Lessard, I. A., Park, I., and Wright, G. D. (1998) Antimicrob. Agents Chemother. 42, Footnotes: * This work was supported by the Natural Sciences and Engineering Council of Canada and by a Canada Research Chair in Antibiotic Biochemistry (to G.D.W.). 1 The abbreviations used are: NADPH, nicotinamide adenine dinucleotide phosphate (reduced), FAD, flavin adenine dinucleotide,, PCR, polymerase chain reaction, EDTA, ethylenedinitrilo-tetraacetic acid, HEPES, N-2-hydroxyethylpiperazine-N -2- ethanesulfonic acid, FMN, flavin mononucleotide, TAPS, N-tris(hydroxymethyl)methyl- 3-amino-propanesulfonic acid, HPLC, high-performance liquid chromatography, SDS, sodium dodecyl sulfate, ESI-MS, electrospray ionization mass spectrometry, NMR, nuclear magnetic resonance, COSY, correlated spectroscopy, HSQC, heteronuclear single quantum coherence, HMBC, heteronuclear multiple bond coherence, NOE, nuclear Overhauser effect, LC-MS, liquid chromatography mass spectrometry, MIC, minimum inhibitory concentration. 21
22 FIGURE LEGENDS Figure 1. Tetracycline structure and nomenclature. A. Structures of tetracycline antibiotics used in this study. B. Numbering scheme of tetracycline antibiotics. Figure 2. Architecture of TetX orthologues from Bacteroides fragilis. The N-terminal domain 1, which is lacking in TetX1, is a predicted flavin binding domain (Pfam FAD3), while the C-terminal domain 2 is a signature Pfam monooxygenase fold. Figure 3. Effect of TetX on tetracycline in liquid culture. E. coli W3110 bearing the tetx gene on plasmid pdb1 when grown in the presence of tetracycline turns the medium black and is associated with tetracycline degradation. Figure 4. UV-visible spectrum of purified TetX2. The absorbance maxima at 366 and 445 nm shown in the inset are indicative of flavin content. Figure 5. UV-visible spectrum of oxytetracycline degradation catalyzed by TetX2. The sample cuvette contained purified TetX2, oxytetracycline and an NADPH regenerating system while the reference cuvette did not contain TetX2 (see Experimental Procedures for details). Each scan was taken at 20 s intervals. Figure 6. Reverse phase HPLC separation of oxytetracycline and TetX2 catalyzed products. A) Decrease in the oxytetracycline peak at 363 nm resulting from disruption of the aryl beta-diketone chromophore upon TetX2-catalyzed hydroxylation. B) Absorbance at 260 nm reflecting the aromatic (Ring D) region of the substrates and products. Figure 7. The 600 MHz 1 H NMR spectrum of P1 and oxytetracycline in 0.1 M DCl/D 2 O. The aromatic region of P1 shows no change compared to oxytetracycline. The nonaromatic region shows significant changes in chemical shift and coupling of the methine protons of P1. 22
23 TABLES Table 1. (page 11) Steady state kinetic parameters for the inactivation of tetracycline analogues by TetX2. Substrate K m k cat k cat /K m (µm) (s -1 ) (M -1 s -1 ) Oxytetracycline ± ± x 10 4 Tetracycline 54.0 ± ± x 10 3 Chlortetracycline 110 ± ± x 10 3 Demeclocycline 19.9 ± ± x 10 4 Doxycycline 83.7 ± ± x 10 3 Minocycline 28.4 ± ± x 10 3 NADPH ± ± x 10 3 Kinetic values are shown ± standard error of fit of the data to the Michealis-Menten equation. 1-1 mm of NADPH, 25 mm of TAPS buffer, ph 8.5, were used in the reactions for all tetracycline antibiotics mm oxytetracycline, 25 mm of TAPS buffer, ph
24 Table 2. (page 11) MIC Values of Tetracycline Antibiotics with E. coli W3110/pDB1. 1 Antibiotic MIC (µg/ml) Oxytetracycline 256 Tetracycline 256 Chlortetracycline 128 Demeclocycline 64 Doxycycline 32 Minocycline 8 1- Plasmid pdb1 contains the tetx gene in a puc18 background. Values for puc18 alone were < 2 µg/ml. 24
25 FIGURES Figure 1. (page 3) A. R 1 R R 2 R 4 3 N(CH 3 ) 2 OH H H OH NH 2 B. 9 OH OH O O R 1 R 2 R 3 R 4 Oxytetracycline H OH CH 3 OH Tetracycline H OH CH 3 H Chlortetracycline Cl OH CH 3 H Demeclocycline Cl OH H H Doxycycline H H CH 3 OH Minocycline N(CH 3 ) 2 H H H 8 7 D 6a 6 5 5a C 10 10a 11a 11 OH OH B O O H 3 C CH 3 N 4a 12a 12 4 A O OH O NH 2 25
26 Figure 2. (page 4) 1 2 TetX TetX1 TetX2 26
27 Figure 3. (page 8) 27
28 Figure 4. (page 9) 28
29 Figure 5. (page 10) 29
30 Figure 6. (page 10) A S S S A363 S S S S B A260 S P1 S P1 S P1 S P2 P1 S P2 P1 S P2 P1 S P2 Omin 2 min 5 min 10min 15min 20min 30 min 30
31 Figure 7. (page 12) 31
32 Scheme 1. (page 15) HO CH 3 OH N(CH 3 ) 2 H H OH OH NH 2 OH O O - O O Oxytetracycline HB HO O H O N NH HO CH 3 OH N(CH 3 ) 2 OH H H OH NH 2 OH OH O O O O 11a-hydroxy-oxytetracycline H O OH N NH N O NH N N O R FAD-4a-hydroperoxide N R N O N R N O P2 etc. H 2 O CH 3 OH N(CH 3 ) 2 O H OH OH NH 2 OH OH O OH O O P1 11a-hydroxy-oxytetracycline-6-12-hemiketal 32
33 Supplementary Data for TetX is a Flavin-Dependent Monooxygenase Conferring Resistance to Tetracycline Antibiotics by Wangrong Yang, Ian F. Moore, Kalinka P. Koteva, Donald W. Hughes, David C. Bareich and Gerard D. Wright. Table S1. Oligonucleotide primers for PCR amplification of TetX constructs. Restriction sites are underlined. Protein Primer Sequence TetX 5 -CCG GAA TTC AAG CTT TTA TTA TAC ATT TAA CAA TTG C 5 -CCG GAA TTC CAT ATG ACA ATG CGA ATA GAT ACA GAC TetX1 5 -GCG TCT AGA CAT ATG GCA AAC TTG TTA CAA CAA ACC GG 5 -GGA ATT CAA GCT TTT ATA CAT TCA TTA GCT GTT GAA AAG TAA AGC TetX2 tetx2f 5 -CCG GAA TTC CAT ATG ACA ATG CGA ATA GAT ACA GAC tetx2r 5 - CCG GAA TTC AAG CTT TTA T TA TAC ATT TAA CAA TTG C Table S2. 1 H NMR assignments of oxytetracycline in 0.1M DCl/D 2 O. Proton Chemical Shift (ppm) Coupling Constant (Hz) J 4,4a = N(CH 3 ) , a J 5,4a = a CH J 7,9 = 0.9, J 7,8 = J 8,9 =
34 Table S3. 13 C NMR assignments of oxytetracycline in 0.1M DCl/D 2 O. Carbon Chemical Shift (ppm) CONH N(CH 3 ) , a a CH a a a a Table S4. 1 H NMR Assignments of 11a-hydroxy-oxytetracycline-6,12-hemiketal (P1) in 0.1M DCl/D 2 O. Proton Chemical Shift (ppm) Coupling Constant (Hz) J 4,4a = N(CH 3 ) , a J 5,4a = 8.9 5a CH J 7,8 = J 8,9 =
35 Table S5. 13 C NMR Assignments of 11a-hydroxy-oxytetracycline-6,12-hemiketal (P1) in 0.1M DCl/D 2 O. Carbon Chemical Shift (ppm) CONH N(CH 3 ) a a CH a a a a
36 Figure S1. The 1 H NMR spectrum of P1 in 0.1M DCl/D 2 O. 36
37 Figure S2. Expansion of the aromatic region of the 1 H NMR spectrum of P1. Figure S3. Expansion of the 1 H NMR spectrum of P1 in 0.1M DCl/D 2 O. 37
38 Figure S3. Expansion of the 1 H NMR spectrum of P1 showing the resonances of the four methine protons and the protons of the dimethylamino group. 38
39 Figure S4. Expansion of the 1 H NMR spectrum of P1 showing the resonance of the 6- CH 3 group. 39
40 Figure S5. The COSY spectrum of P1. 40
41 Figure S6. Expansion of the COSY spectrum of P1. The circled peaks indicate the coupling interaction between the doublet of doublets at 3.67 ppm and the doublets at 4.16 ppm and 3.97 ppm. 41
42 Figure S7. Expansion of the aromatic region of the COSY spectrum of P1. 42
43 Figure S8. The NOE difference spectrum of oxytetracycline generated by saturating the 6-CH 3 resonance at 1.72 ppm, enhancement of the resonances at 7.14 ppm, ppm and 2.90 ppm can be seen. 43
44 Figure S9. The NOE difference spectrum of P1 generated by saturating the resonance at 1.45 ppm, enhancement of the resonances at 7.12 ppm, 4.16 ppm and 2.85 ppm can be seen. 44
45 Figure S10. The 13 C NMR spectrum of P1 0.1M DCl/D 2 O. 45
46 Figure S11. Expansion of the 13 C NMR spectrum of P1 0.1M DCl/D 2 O. 46
47 Figure S12. Expansion of the 13 C NMR spectrum of P1 0.1M DCl/D 2 O. 47
48 Figure S13. The HSQC spectrum of P1 0.1M DCl/D 2 O, ten proton-carbon correlations observed. 48
49 Figure S14. Expansion of the HSQC spectrum of P1 showing the correlations of the aromatic protons. 49
50 Figure S15. Expansion of the HSQC spectrum of P1 showing the correlations of the nonaromatic protons. The circled signals show the methine proton-carbon correlations. 50
51 Figure S16. The HMBC spectrum of P1 0.1M DCl/D 2 O. The three circled peaks indicate the 2-bond and 3-bond couplings of the 6-CH 3 protons to C6a, C6 and C5a. 51
52 Figure S17. Expansion of the HMBC spectrum of P1, the circles show the 2-bond and 3- bond correlations of the H5a proton. 52
53 Figure S18. Expansion of the HMBC spectrum of P1, the circle shows the 3-bond correlation to C11. 53
54 Figure S19. Expansion of the aromatic region of the HMBC spectrum of P1. 54
55 Figure S20. Purity of TetX2. SDS-Polyacrylamide gel (11%) of 4 consecutive fractions of TetX2 after purification. The gel was stained with coomassie blue. The mass of the high molecular weight markers (HMW) in kda is shown on the left HMW
56 TetX is a flavin-dependent monooxygenase conferring resistance to tetracycline antibiotics Wangrong Yang, Ian F. Moore, Kalinka P. Koteva, David C. Bareich, Donald W. Hughes and Gerard D. Wright J. Biol. Chem. published online September 27, 2004 Access the most updated version of this article at doi: /jbc.M Alerts: When this article is cited When a correction for this article is posted Click here to choose from all of JBC's alerts Supplemental material:
Resistance to Tetracycline Antibiotics by Wangrong Yang, Ian F. Moore, Kalinka P. Koteva, Donald W. Hughes, David C. Bareich and Gerard D. Wright.
 Supplementary Data for TetX is a Flavin-Dependent Monooxygenase Conferring Resistance to Tetracycline Antibiotics by Wangrong Yang, Ian F. Moore, Kalinka P. Koteva, Donald W. Hughes, David C. Bareich and
Supplementary Data for TetX is a Flavin-Dependent Monooxygenase Conferring Resistance to Tetracycline Antibiotics by Wangrong Yang, Ian F. Moore, Kalinka P. Koteva, Donald W. Hughes, David C. Bareich and
Supporting Information for:
 Supporting Information for: Methylerythritol Cyclodiphosphate (MEcPP) in Deoxyxylulose Phosphate Pathway: Synthesis from an Epoxide and Mechanisms Youli Xiao, a Rodney L. Nyland II, b Caren L. Freel Meyers
Supporting Information for: Methylerythritol Cyclodiphosphate (MEcPP) in Deoxyxylulose Phosphate Pathway: Synthesis from an Epoxide and Mechanisms Youli Xiao, a Rodney L. Nyland II, b Caren L. Freel Meyers
Characterization of the DNA-mediated Oxidation of Dps, a Bacterial Ferritin
 SUPPORTING INFORMATION Characterization of the DNA-mediated Oxidation of Dps, a Bacterial Ferritin Anna R. Arnold, Andy Zhou, and Jacqueline K. Barton Division of Chemistry and Chemical Engineering, California
SUPPORTING INFORMATION Characterization of the DNA-mediated Oxidation of Dps, a Bacterial Ferritin Anna R. Arnold, Andy Zhou, and Jacqueline K. Barton Division of Chemistry and Chemical Engineering, California
Identification of novel endophenaside antibiotics produced by Kitasatospora sp. MBT66
 SUPPORTING INFORMATION belonging to the manuscript: Identification of novel endophenaside antibiotics produced by Kitasatospora sp. MBT66 by Changsheng Wu 1, 2, Gilles P. van Wezel 1, *, and Young Hae
SUPPORTING INFORMATION belonging to the manuscript: Identification of novel endophenaside antibiotics produced by Kitasatospora sp. MBT66 by Changsheng Wu 1, 2, Gilles P. van Wezel 1, *, and Young Hae
SUPPLEMENTARY MATERIAL
 SUPPLEMENTARY MATERIAL Purification and biochemical properties of SDS-stable low molecular weight alkaline serine protease from Citrullus Colocynthis Muhammad Bashir Khan, 1,3 Hidayatullah khan, 2 Muhammad
SUPPLEMENTARY MATERIAL Purification and biochemical properties of SDS-stable low molecular weight alkaline serine protease from Citrullus Colocynthis Muhammad Bashir Khan, 1,3 Hidayatullah khan, 2 Muhammad
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Sherman SI, Wirth LJ, Droz J-P, et al. Motesanib diphosphate
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Sherman SI, Wirth LJ, Droz J-P, et al. Motesanib diphosphate
Supplementary Material (ESI) for Chemical Communications This journal is (c) The Royal Society of Chemistry 2008
 Experimental Details Unless otherwise noted, all chemicals were purchased from Sigma-Aldrich Chemical Company and were used as received. 2-DOS and neamine were kindly provided by Dr. F. Huang. Paromamine
Experimental Details Unless otherwise noted, all chemicals were purchased from Sigma-Aldrich Chemical Company and were used as received. 2-DOS and neamine were kindly provided by Dr. F. Huang. Paromamine
Heparin Sodium ヘパリンナトリウム
 Heparin Sodium ヘパリンナトリウム Add the following next to Description: Identification Dissolve 1 mg each of Heparin Sodium and Heparin Sodium Reference Standard for physicochemical test in 1 ml of water, and
Heparin Sodium ヘパリンナトリウム Add the following next to Description: Identification Dissolve 1 mg each of Heparin Sodium and Heparin Sodium Reference Standard for physicochemical test in 1 ml of water, and
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/2/4/e1500980/dc1 Supplementary Materials for The crystal structure of human dopamine -hydroxylase at 2.9 Å resolution Trine V. Vendelboe, Pernille Harris, Yuguang
advances.sciencemag.org/cgi/content/full/2/4/e1500980/dc1 Supplementary Materials for The crystal structure of human dopamine -hydroxylase at 2.9 Å resolution Trine V. Vendelboe, Pernille Harris, Yuguang
Kit for assay of thioredoxin
 FkTRX-02-V2 Kit for assay of thioredoxin The thioredoxin system is the major protein disulfide reductase in cells and comprises thioredoxin, thioredoxin reductase and NADPH (1). Thioredoxin systems are
FkTRX-02-V2 Kit for assay of thioredoxin The thioredoxin system is the major protein disulfide reductase in cells and comprises thioredoxin, thioredoxin reductase and NADPH (1). Thioredoxin systems are
c Tuj1(-) apoptotic live 1 DIV 2 DIV 1 DIV 2 DIV Tuj1(+) Tuj1/GFP/DAPI Tuj1 DAPI GFP
 Supplementary Figure 1 Establishment of the gain- and loss-of-function experiments and cell survival assays. a Relative expression of mature mir-484 30 20 10 0 **** **** NCP mir- 484P NCP mir- 484P b Relative
Supplementary Figure 1 Establishment of the gain- and loss-of-function experiments and cell survival assays. a Relative expression of mature mir-484 30 20 10 0 **** **** NCP mir- 484P NCP mir- 484P b Relative
Supplementary Materials
 Supplementary Materials 1 Supplementary Table 1. List of primers used for quantitative PCR analysis. Gene name Gene symbol Accession IDs Sequence range Product Primer sequences size (bp) β-actin Actb gi
Supplementary Materials 1 Supplementary Table 1. List of primers used for quantitative PCR analysis. Gene name Gene symbol Accession IDs Sequence range Product Primer sequences size (bp) β-actin Actb gi
A smart acid nanosystem for ultrasensitive. live cell mrna imaging by the target-triggered intracellular self-assembly
 Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2017 A smart ZnO@polydopamine-nucleic acid nanosystem for ultrasensitive live cell mrna imaging
Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2017 A smart ZnO@polydopamine-nucleic acid nanosystem for ultrasensitive live cell mrna imaging
Proteins. Amino acids, structure and function. The Nobel Prize in Chemistry 2012 Robert J. Lefkowitz Brian K. Kobilka
 Proteins Amino acids, structure and function The Nobel Prize in Chemistry 2012 Robert J. Lefkowitz Brian K. Kobilka O O HO N N HN OH Ser65-Tyr66-Gly67 The Nobel prize in chemistry 2008 Osamu Shimomura,
Proteins Amino acids, structure and function The Nobel Prize in Chemistry 2012 Robert J. Lefkowitz Brian K. Kobilka O O HO N N HN OH Ser65-Tyr66-Gly67 The Nobel prize in chemistry 2008 Osamu Shimomura,
Luminescent platforms for monitoring changes in the solubility of amylin and huntingtin in living cells
 Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2016 Contents Supporting Information Luminescent platforms for monitoring changes in the
Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2016 Contents Supporting Information Luminescent platforms for monitoring changes in the
Supporting information (protein purification, kinetic characterization, product isolation, and characterization by NMR and mass spectrometry):
 Supporting Information Mechanistic studies of a novel C-S lyase in ergothioneine biosynthesis: the involvement of a sulfenic acid intermediate Heng Song, 1 Wen Hu, 1,2 Nathchar Naowarojna, 1 Ampon Sae
Supporting Information Mechanistic studies of a novel C-S lyase in ergothioneine biosynthesis: the involvement of a sulfenic acid intermediate Heng Song, 1 Wen Hu, 1,2 Nathchar Naowarojna, 1 Ampon Sae
Europium Labeling Kit
 Europium Labeling Kit Catalog Number KA2096 100ug *1 Version: 03 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Intended Use... 3 Background... 3 Principle of the Assay...
Europium Labeling Kit Catalog Number KA2096 100ug *1 Version: 03 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Intended Use... 3 Background... 3 Principle of the Assay...
A STUDY OF THE METABOLISM OF THEOBROMINE, THEOPHYLLINE, AND CAFFEINE IN MAN* Previous studies (1, 2) have shown that after the ingestion of caffeine
 A STUDY OF THE METABOLISM OF THEOBROMINE, THEOPHYLLINE, AND CAFFEINE IN MAN* BY HERBERT H. CORNISH AND A. A. CHRISTMAN (From the Department of Biological Chemistry, Medical School, University of Michigan,
A STUDY OF THE METABOLISM OF THEOBROMINE, THEOPHYLLINE, AND CAFFEINE IN MAN* BY HERBERT H. CORNISH AND A. A. CHRISTMAN (From the Department of Biological Chemistry, Medical School, University of Michigan,
Culture Density (OD600) 0.1. Culture Density (OD600) Culture Density (OD600) Culture Density (OD600) Culture Density (OD600)
 A. B. C. D. E. PA JSRI JSRI 2 PA DSAM DSAM 2 DSAM 3 PA LNAP LNAP 2 LNAP 3 PAO Fcor Fcor 2 Fcor 3 PAO Wtho Wtho 2 Wtho 3 Wtho 4 DTSB Low Iron 2 4 6 8 2 4 6 8 2 22 DTSB Low Iron 2 4 6 8 2 4 6 8 2 22 DTSB
A. B. C. D. E. PA JSRI JSRI 2 PA DSAM DSAM 2 DSAM 3 PA LNAP LNAP 2 LNAP 3 PAO Fcor Fcor 2 Fcor 3 PAO Wtho Wtho 2 Wtho 3 Wtho 4 DTSB Low Iron 2 4 6 8 2 4 6 8 2 22 DTSB Low Iron 2 4 6 8 2 4 6 8 2 22 DTSB
Supporting information
 Supporting information Figure legends Supplementary Table 1. Specific product ions obtained from fragmentation of lithium adducts in the positive ion mode comparing the different positional isomers of
Supporting information Figure legends Supplementary Table 1. Specific product ions obtained from fragmentation of lithium adducts in the positive ion mode comparing the different positional isomers of
Biologic Oxidation BIOMEDICAL IMPORTAN
 Biologic Oxidation BIOMEDICAL IMPORTAN Chemically, oxidation is defined as the removal of electrons and reduction as the gain of electrons. Thus, oxidation is always accompanied by reduction of an electron
Biologic Oxidation BIOMEDICAL IMPORTAN Chemically, oxidation is defined as the removal of electrons and reduction as the gain of electrons. Thus, oxidation is always accompanied by reduction of an electron
UV Tracer TM Maleimide NHS ester
 UV Tracer TM Maleimide HS ester Product o.: 1020 Product ame: UV-Tracer TM Maleimide-HS ester Chemical Structure: Chemical Composition: C 41 H 67 5 18 Molecular Weight: 1014.08 Appearance: Storage: Yellow
UV Tracer TM Maleimide HS ester Product o.: 1020 Product ame: UV-Tracer TM Maleimide-HS ester Chemical Structure: Chemical Composition: C 41 H 67 5 18 Molecular Weight: 1014.08 Appearance: Storage: Yellow
Chapter 8. An Introduction to Microbial Metabolism
 Chapter 8 An Introduction to Microbial Metabolism The metabolism of microbes Metabolism sum of all chemical reactions that help cells function Two types of chemical reactions: Catabolism -degradative;
Chapter 8 An Introduction to Microbial Metabolism The metabolism of microbes Metabolism sum of all chemical reactions that help cells function Two types of chemical reactions: Catabolism -degradative;
UNIVERSITY OF GUELPH CHEM 4540 ENZYMOLOGY Winter 2005 Quiz #2: March 24, 2005, 11:30 12:50 Instructor: Prof R. Merrill ANSWERS
 UNIVERSITY F GUELPH CHEM 4540 ENZYMLGY Winter 2005 Quiz #2: March 24, 2005, 11:30 12:50 Instructor: Prof R. Merrill ANSWERS Instructions: Time allowed = 80 minutes. Total marks = 30. This quiz represents
UNIVERSITY F GUELPH CHEM 4540 ENZYMLGY Winter 2005 Quiz #2: March 24, 2005, 11:30 12:50 Instructor: Prof R. Merrill ANSWERS Instructions: Time allowed = 80 minutes. Total marks = 30. This quiz represents
<Supplemental information>
 The Structural Basis of Endosomal Anchoring of KIF16B Kinesin Nichole R. Blatner, Michael I. Wilson, Cai Lei, Wanjin Hong, Diana Murray, Roger L. Williams, and Wonhwa Cho Protein
The Structural Basis of Endosomal Anchoring of KIF16B Kinesin Nichole R. Blatner, Michael I. Wilson, Cai Lei, Wanjin Hong, Diana Murray, Roger L. Williams, and Wonhwa Cho Protein
Supporting Information
 Translation of DNA into Synthetic N-Acyloxazolidines Xiaoyu Li, Zev. J. Gartner, Brian N. Tse and David R. Liu* Department of Chemistry and Chemical Biology, Harvard University, Cambridge, Massachusetts
Translation of DNA into Synthetic N-Acyloxazolidines Xiaoyu Li, Zev. J. Gartner, Brian N. Tse and David R. Liu* Department of Chemistry and Chemical Biology, Harvard University, Cambridge, Massachusetts
Lecture Sixteen: METABOLIC ENERGY: [Based on GENERATION Chapter 15
 Lecture Sixteen: METABOLIC ENERGY: [Based on GENERATION Chapter 15 AND STORAGE Berg, (Figures in red are for the 7th Edition) Tymoczko (Figures in Blue are for the 8th Edition) & Stryer] Two major questions
Lecture Sixteen: METABOLIC ENERGY: [Based on GENERATION Chapter 15 AND STORAGE Berg, (Figures in red are for the 7th Edition) Tymoczko (Figures in Blue are for the 8th Edition) & Stryer] Two major questions
Supplementary Table 3. 3 UTR primer sequences. Primer sequences used to amplify and clone the 3 UTR of each indicated gene are listed.
 Supplemental Figure 1. DLKI-DIO3 mirna/mrna complementarity. Complementarity between the indicated DLK1-DIO3 cluster mirnas and the UTR of SOX2, SOX9, HIF1A, ZEB1, ZEB2, STAT3 and CDH1with mirsvr and PhastCons
Supplemental Figure 1. DLKI-DIO3 mirna/mrna complementarity. Complementarity between the indicated DLK1-DIO3 cluster mirnas and the UTR of SOX2, SOX9, HIF1A, ZEB1, ZEB2, STAT3 and CDH1with mirsvr and PhastCons
Foundations in Microbiology Seventh Edition
 Lecture PowerPoint to accompany Foundations in Microbiology Seventh Edition Talaro Chapter 8 An Introduction to Microbial Metabolism Copyright The McGraw-Hill Companies, Inc. Permission required for reproduction
Lecture PowerPoint to accompany Foundations in Microbiology Seventh Edition Talaro Chapter 8 An Introduction to Microbial Metabolism Copyright The McGraw-Hill Companies, Inc. Permission required for reproduction
Supplementary Material
 Supplementary Material Antimicrobial mechanism of theaflavins: They target 1-deoxy-D-xylulose 5-phosphate reductoisomerase, the key enzyme of the MEP terpenoid biosynthetic pathway Xian Hui, Qiao Yue,
Supplementary Material Antimicrobial mechanism of theaflavins: They target 1-deoxy-D-xylulose 5-phosphate reductoisomerase, the key enzyme of the MEP terpenoid biosynthetic pathway Xian Hui, Qiao Yue,
Plasmids Western blot analysis and immunostaining Flow Cytometry Cell surface biotinylation RNA isolation and cdna synthesis
 Plasmids psuper-retro-s100a10 shrna1 was constructed by cloning the dsdna oligo 5 -GAT CCC CGT GGG CTT CCA GAG CTT CTT TCA AGA GAA GAA GCT CTG GAA GCC CAC TTT TTA-3 and 5 -AGC TTA AAA AGT GGG CTT CCA GAG
Plasmids psuper-retro-s100a10 shrna1 was constructed by cloning the dsdna oligo 5 -GAT CCC CGT GGG CTT CCA GAG CTT CTT TCA AGA GAA GAA GCT CTG GAA GCC CAC TTT TTA-3 and 5 -AGC TTA AAA AGT GGG CTT CCA GAG
Oxidative Phosphorylation
 Oxidative Phosphorylation Energy from Reduced Fuels Is Used to Synthesize ATP in Animals Carbohydrates, lipids, and amino acids are the main reduced fuels for the cell. Electrons from reduced fuels are
Oxidative Phosphorylation Energy from Reduced Fuels Is Used to Synthesize ATP in Animals Carbohydrates, lipids, and amino acids are the main reduced fuels for the cell. Electrons from reduced fuels are
CELLULAR METABOLISM. Metabolic pathways can be linear, branched, cyclic or spiral
 CHM333 LECTURE 24 & 25: 3/27 29/13 SPRING 2013 Professor Christine Hrycyna CELLULAR METABOLISM What is metabolism? - How cells acquire, transform, store and use energy - Study reactions in a cell and how
CHM333 LECTURE 24 & 25: 3/27 29/13 SPRING 2013 Professor Christine Hrycyna CELLULAR METABOLISM What is metabolism? - How cells acquire, transform, store and use energy - Study reactions in a cell and how
Supporting Information. were prepared from commercially available ethyl acetoacetate by alkylation with the
 ighly Stereoselective Reductions of α-alkyl-1,3-diketones and α- Alkyl-β-keto esters Catalyzed by Isolated NADP-dependent Ketoreductases Dimitris Kalaitzakis, a David J. Rozzell b, Spiros Kambourakis *b
ighly Stereoselective Reductions of α-alkyl-1,3-diketones and α- Alkyl-β-keto esters Catalyzed by Isolated NADP-dependent Ketoreductases Dimitris Kalaitzakis, a David J. Rozzell b, Spiros Kambourakis *b
An Investigative Study of Reactions Involving Glucosinolates and Isothiocyanates
 An Investigative Study of Reactions Involving Glucosinolates and Isothiocyanates Alzea Chrisel H. Alea 1, Diane Elaine T. Co 2 and Marissa G Noel 3* 1,2,3Chemistry Department, De La Salle University, 2401
An Investigative Study of Reactions Involving Glucosinolates and Isothiocyanates Alzea Chrisel H. Alea 1, Diane Elaine T. Co 2 and Marissa G Noel 3* 1,2,3Chemistry Department, De La Salle University, 2401
New immunomodulators with antitumoral properties; Isolation of active naturally-occurring anti-mitotic components of MR>1KD from pollen extract T60
 I M M U N O M O D U L A T O R S U P P O R T : GRAMINEX Flower Pollen Extract New immunomodulators with antitumoral properties; Isolation of active naturally-occurring anti-mitotic components of MR>1KD
I M M U N O M O D U L A T O R S U P P O R T : GRAMINEX Flower Pollen Extract New immunomodulators with antitumoral properties; Isolation of active naturally-occurring anti-mitotic components of MR>1KD
2,6,9-Triazabicyclo[3.3.1]nonanes as overlooked. amino-modification products by acrolein
![2,6,9-Triazabicyclo[3.3.1]nonanes as overlooked. amino-modification products by acrolein 2,6,9-Triazabicyclo[3.3.1]nonanes as overlooked. amino-modification products by acrolein](/thumbs/86/94743397.jpg) Supplementary Information 2,6,9-Triazabicyclo[3.3.1]nonanes as overlooked amino-modification products by acrolein Ayumi Tsutsui and Katsunori Tanaka* Biofunctional Synthetic Chemistry Laboratory, RIKEN
Supplementary Information 2,6,9-Triazabicyclo[3.3.1]nonanes as overlooked amino-modification products by acrolein Ayumi Tsutsui and Katsunori Tanaka* Biofunctional Synthetic Chemistry Laboratory, RIKEN
SUPPLEMENTARY INFORMATION. Bacterial strains and growth conditions. Streptococcus pneumoniae strain R36A was
 SUPPLEMENTARY INFORMATION Bacterial strains and growth conditions. Streptococcus pneumoniae strain R36A was grown in a casein-based semisynthetic medium (C+Y) supplemented with yeast extract (1 mg/ml of
SUPPLEMENTARY INFORMATION Bacterial strains and growth conditions. Streptococcus pneumoniae strain R36A was grown in a casein-based semisynthetic medium (C+Y) supplemented with yeast extract (1 mg/ml of
THE JOURNAL OF ANTIBIOTICS. Polyketomycin, a New Antibiotic from Streptomyces sp. MK277-AF1. II. Structure Determination
 THE JOURNAL OF ANTIBIOTICS Polyketomycin, a New Antibiotic from Streptomyces sp. MK277-AF1 II. Structure Determination ISAO MOMOSE, WEI CHEN, HIKARU NAKAMURA, HIROSHI NAGANAWA, HIRONOBU IINUMA and TOMIO
THE JOURNAL OF ANTIBIOTICS Polyketomycin, a New Antibiotic from Streptomyces sp. MK277-AF1 II. Structure Determination ISAO MOMOSE, WEI CHEN, HIKARU NAKAMURA, HIROSHI NAGANAWA, HIRONOBU IINUMA and TOMIO
BASIC ENZYMOLOGY 1.1
 BASIC ENZYMOLOGY 1.1 1.2 BASIC ENZYMOLOGY INTRODUCTION Enzymes are synthesized by all living organisms including man. These life essential substances accelerate the numerous metabolic reactions upon which
BASIC ENZYMOLOGY 1.1 1.2 BASIC ENZYMOLOGY INTRODUCTION Enzymes are synthesized by all living organisms including man. These life essential substances accelerate the numerous metabolic reactions upon which
Preparation and Characterization of Cysteine Adducts of Deoxynivalenol
 Preparation and Characterization of Cysteine Adducts of Deoxynivalenol Ana Stanic, Silvio Uhlig, Anita Solhaug, Frode Rise, Alistair L. Wilkins, Christopher O. Miles S1 Figure S1. 1 H spectrum of 1 (DON)
Preparation and Characterization of Cysteine Adducts of Deoxynivalenol Ana Stanic, Silvio Uhlig, Anita Solhaug, Frode Rise, Alistair L. Wilkins, Christopher O. Miles S1 Figure S1. 1 H spectrum of 1 (DON)
Singlet oxygen photosensitisation by the fluorescent probe Singlet Oxygen Sensor Green
 Singlet oxygen photosensitisation by the fluorescent probe Singlet Oxygen Sensor Green Xavier Ragàs, Ana Jiménez-Banzo, David Sánchez-García, Xavier Batllori and Santi Nonell* Grup d Enginyeria Molecular,
Singlet oxygen photosensitisation by the fluorescent probe Singlet Oxygen Sensor Green Xavier Ragàs, Ana Jiménez-Banzo, David Sánchez-García, Xavier Batllori and Santi Nonell* Grup d Enginyeria Molecular,
1/3/2011. Chapter 17 Carboxylic Acids and Their Derivatives. Nucleophilic Addition- Elimination at the Acyl Carbon
 Introduction The carboxyl group (-CO 2 H) is the parent group of a family of compounds called acyl compounds or carboxylic acid derivatives Chapter 17 Carboxylic Acids and Their Derivatives. Nucleophilic
Introduction The carboxyl group (-CO 2 H) is the parent group of a family of compounds called acyl compounds or carboxylic acid derivatives Chapter 17 Carboxylic Acids and Their Derivatives. Nucleophilic
Amylase: a sample enzyme
 Amylase: a sample enzyme Objectives: After completion of this laboratory exercise you will be able to: 1. Explain the importance of enzymes in biology. 2. Explain the basic properties of an enzyme as a
Amylase: a sample enzyme Objectives: After completion of this laboratory exercise you will be able to: 1. Explain the importance of enzymes in biology. 2. Explain the basic properties of an enzyme as a
Stabilization and enhanced reactivity of actinorhodin polyketide synthase minimal complex in polymer/nucleotide coacervate droplets
 Stabilization and enhanced reactivity of actinorhodin polyketide synthase minimal complex in polymer/nucleotide coacervate droplets John Crosby,* Tom Treadwell, Michelle Hammerton, Konstantinos Vasilakis,
Stabilization and enhanced reactivity of actinorhodin polyketide synthase minimal complex in polymer/nucleotide coacervate droplets John Crosby,* Tom Treadwell, Michelle Hammerton, Konstantinos Vasilakis,
CHAPTER-6 IDENTIFICATION, AND CHARACTERISATION OF DEGRADATION IMPURITY IN VALSARTAN TABLETS
 129 CHAPTER-6 IDENTIFICATION, AND CHARACTERISATION OF DEGRADATION IMPURITY IN VALSARTAN TABLETS 130 6.1. Introduction Valsartan is an orally active specific angiotensin II blocker effective in lowering
129 CHAPTER-6 IDENTIFICATION, AND CHARACTERISATION OF DEGRADATION IMPURITY IN VALSARTAN TABLETS 130 6.1. Introduction Valsartan is an orally active specific angiotensin II blocker effective in lowering
Tivadar Orban, Beata Jastrzebska, Sayan Gupta, Benlian Wang, Masaru Miyagi, Mark R. Chance, and Krzysztof Palczewski
 Structure, Volume Supplemental Information Conformational Dynamics of Activation for the Pentameric Complex of Dimeric G Protein-Coupled Receptor and Heterotrimeric G Protein Tivadar Orban, Beata Jastrzebska,
Structure, Volume Supplemental Information Conformational Dynamics of Activation for the Pentameric Complex of Dimeric G Protein-Coupled Receptor and Heterotrimeric G Protein Tivadar Orban, Beata Jastrzebska,
Thiol-Activated gem-dithiols: A New Class of Controllable. Hydrogen Sulfide (H 2 S) Donors
 Thiol-Activated gem-dithiols: A New Class of Controllable Hydrogen Sulfide (H 2 S) Donors Yu Zhao, Jianming Kang, Chung-Min Park, Powell E. Bagdon, Bo Peng, and Ming Xian * Department of Chemistry, Washington
Thiol-Activated gem-dithiols: A New Class of Controllable Hydrogen Sulfide (H 2 S) Donors Yu Zhao, Jianming Kang, Chung-Min Park, Powell E. Bagdon, Bo Peng, and Ming Xian * Department of Chemistry, Washington
Analysis of Uncomplexed and Copper-complexed Methanobactin with UV/Visible Spectrophotometry, Mass Spectrometry and NMR Spectrometry
 Analysis of Uncomplexed and Copper-complexed Methanobactin with UV/Visible Spectrophotometry, Mass Spectrometry and NMR Spectrometry Lee Behling, Alan DiSpirito, Scott Hartsel, Larry Masterson, Gianluigi
Analysis of Uncomplexed and Copper-complexed Methanobactin with UV/Visible Spectrophotometry, Mass Spectrometry and NMR Spectrometry Lee Behling, Alan DiSpirito, Scott Hartsel, Larry Masterson, Gianluigi
Supporting Information for. Boronic Acid Functionalized Aza-Bodipy (azabdpba) based Fluorescence Optodes for the. analysis of Glucose in Whole Blood
 Supporting Information for Boronic Acid Functionalized Aza-Bodipy (azabdpba) based Fluorescence Optodes for the analysis of Glucose in Whole Blood Yueling Liu, Jingwei Zhu, Yanmei Xu, Yu Qin*, Dechen Jiang*
Supporting Information for Boronic Acid Functionalized Aza-Bodipy (azabdpba) based Fluorescence Optodes for the analysis of Glucose in Whole Blood Yueling Liu, Jingwei Zhu, Yanmei Xu, Yu Qin*, Dechen Jiang*
Designer Fentanyls Drugs that kill and how to detect them. Cyclopropylfentanyl
 Designer Fentanyls Drugs that kill and how to detect them Cyclopropylfentanyl Science for a safer world The in vitro metabolism of cyclopropylfentanyl Simon Hudson & Charlotte Cutler, Sport and Specialised
Designer Fentanyls Drugs that kill and how to detect them Cyclopropylfentanyl Science for a safer world The in vitro metabolism of cyclopropylfentanyl Simon Hudson & Charlotte Cutler, Sport and Specialised
130327SCH4U_biochem April 09, 2013
 Option B: B1.1 ENERGY Human Biochemistry If more energy is taken in from food than is used up, weight gain will follow. Similarly if more energy is used than we supply our body with, weight loss will occur.
Option B: B1.1 ENERGY Human Biochemistry If more energy is taken in from food than is used up, weight gain will follow. Similarly if more energy is used than we supply our body with, weight loss will occur.
Masatoshi Shibuya,Takahisa Sato, Masaki Tomizawa, and Yoshiharu Iwabuchi* Department of Organic Chemistry, Graduate School of Pharmaceutical Sciences,
 Oxoammonium ion/naclo 2 : An Expedient, Catalytic System for One-pot Oxidation of Primary Alcohols to Carboxylic Acid with Broad Substrate Applicability Masatoshi Shibuya,Takahisa Sato, Masaki Tomizawa,
Oxoammonium ion/naclo 2 : An Expedient, Catalytic System for One-pot Oxidation of Primary Alcohols to Carboxylic Acid with Broad Substrate Applicability Masatoshi Shibuya,Takahisa Sato, Masaki Tomizawa,
Trypsin Mass Spectrometry Grade
 058PR-03 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Trypsin Mass Spectrometry Grade A Chemically Modified, TPCK treated, Affinity Purified
058PR-03 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Trypsin Mass Spectrometry Grade A Chemically Modified, TPCK treated, Affinity Purified
Thermal shift binding experiments were carried out using Thermofluor 384 ELS system. Protein
 Supplementary Methods Thermal shift assays Thermal shift binding experiments were carried out using Thermofluor 384 ELS system. Protein unfolding was examined by monitoring the fluorescence of ANS (1-anilinonaphthalene-8-
Supplementary Methods Thermal shift assays Thermal shift binding experiments were carried out using Thermofluor 384 ELS system. Protein unfolding was examined by monitoring the fluorescence of ANS (1-anilinonaphthalene-8-
Nature Methods: doi: /nmeth Supplementary Figure 1
 Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
BabyBio IMAC columns DATA SHEET DS
 BabyBio IMAC columns DATA SHEET DS 45 655 010 BabyBio columns for Immobilized Metal Ion Affinity Chromatography (IMAC) are ready-to-use for quick and easy purification of polyhistidine-tagged (His-tagged)
BabyBio IMAC columns DATA SHEET DS 45 655 010 BabyBio columns for Immobilized Metal Ion Affinity Chromatography (IMAC) are ready-to-use for quick and easy purification of polyhistidine-tagged (His-tagged)
Fig.S1 ESI-MS spectrum of reaction of ApA and THPTb after 16 h.
 Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2014 Experiment Cleavage of dinucleotides Dinucleotides (ApA, CpC, GpG, UpU) were purchased from
Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2014 Experiment Cleavage of dinucleotides Dinucleotides (ApA, CpC, GpG, UpU) were purchased from
Overview of the Expressway Cell-Free Expression Systems. Expressway Mini Cell-Free Expression System
 Overview of the Expressway Cell-Free Expression Systems The Expressway Cell-Free Expression Systems use an efficient coupled transcription and translation reaction to produce up to milligram quantities
Overview of the Expressway Cell-Free Expression Systems The Expressway Cell-Free Expression Systems use an efficient coupled transcription and translation reaction to produce up to milligram quantities
Caution: For Laboratory Use. A product for research purposes only. Eu-W1284 Iodoacetamido Chelate & Europium Standard. Product Number: AD0014
 TECHNICAL DATA SHEET Lance Caution: For Laboratory Use. A product for research purposes only. Eu-W1284 Iodoacetamido Chelate & Europium Standard Product Number: AD0014 INTRODUCTION: Iodoacetamido-activated
TECHNICAL DATA SHEET Lance Caution: For Laboratory Use. A product for research purposes only. Eu-W1284 Iodoacetamido Chelate & Europium Standard Product Number: AD0014 INTRODUCTION: Iodoacetamido-activated
Metabolite: substance produced or used during metabolism such as lipids, sugars and amino acids
 Metabolomica Analisi di biofluidi mediante spettroscopia NMR Michael Assfalg Università di Verona Terminology Metabolite: substance produced or used during metabolism such as lipids, sugars and amino acids
Metabolomica Analisi di biofluidi mediante spettroscopia NMR Michael Assfalg Università di Verona Terminology Metabolite: substance produced or used during metabolism such as lipids, sugars and amino acids
Improve Protein Analysis with the New, Mass Spectrometry- Compatible ProteasMAX Surfactant
 Improve Protein Analysis with the New, Mass Spectrometry- Compatible Surfactant ABSTRACT Incomplete solubilization and digestion and poor peptide recovery are frequent limitations in protein sample preparation
Improve Protein Analysis with the New, Mass Spectrometry- Compatible Surfactant ABSTRACT Incomplete solubilization and digestion and poor peptide recovery are frequent limitations in protein sample preparation
PFK Activity Assay Kit (Colorimetric)
 PFK Activity Assay Kit (Colorimetric) Catalog Number KA3761 100 assays Version: 02 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Background... 3 General Information...
PFK Activity Assay Kit (Colorimetric) Catalog Number KA3761 100 assays Version: 02 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Background... 3 General Information...
Metabolism Energy Pathways Biosynthesis. Catabolism Anabolism Enzymes
 Topics Microbial Metabolism Metabolism Energy Pathways Biosynthesis 2 Metabolism Catabolism Catabolism Anabolism Enzymes Breakdown of complex organic molecules in order to extract energy and dform simpler
Topics Microbial Metabolism Metabolism Energy Pathways Biosynthesis 2 Metabolism Catabolism Catabolism Anabolism Enzymes Breakdown of complex organic molecules in order to extract energy and dform simpler
PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System
 Application Note LC/MS PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System Purpose This application note describes an automated workflow
Application Note LC/MS PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System Purpose This application note describes an automated workflow
Application Note # FTMS-46 solarix XR: Analysis of Complex Mixtures
 Application Note # FTMS-46 solarix XR: Analysis of Complex Mixtures Introduction Natural organic matter (NOM) as well as crude oil and crude oil fractions are very complex mixtures of organic compounds
Application Note # FTMS-46 solarix XR: Analysis of Complex Mixtures Introduction Natural organic matter (NOM) as well as crude oil and crude oil fractions are very complex mixtures of organic compounds
Plasmonic blood glucose monitor based on enzymatic. etching of gold nanorods
 Plasmonic blood glucose monitor based on enzymatic etching of gold nanorods Xin Liu, Shuya Zhang, Penglong Tan, Jiang Zhou, Yan Huang, Zhou Nie* and Shouzhuo Yao State Key Laboratory of Chemo/Biosensing
Plasmonic blood glucose monitor based on enzymatic etching of gold nanorods Xin Liu, Shuya Zhang, Penglong Tan, Jiang Zhou, Yan Huang, Zhou Nie* and Shouzhuo Yao State Key Laboratory of Chemo/Biosensing
TECHNICAL BULLETIN. R 2 GlcNAcβ1 4GlcNAcβ1 Asn
 GlycoProfile II Enzymatic In-Solution N-Deglycosylation Kit Product Code PP0201 Storage Temperature 2 8 C TECHNICAL BULLETIN Product Description Glycosylation is one of the most common posttranslational
GlycoProfile II Enzymatic In-Solution N-Deglycosylation Kit Product Code PP0201 Storage Temperature 2 8 C TECHNICAL BULLETIN Product Description Glycosylation is one of the most common posttranslational
Affinity Purification of Photosystem I from Chlamydomonas reinhardtii using a Polyhistidine Tag
 Affinity Purification of Photosystem I from Chlamydomonas reinhardtii using a Polyhistidine Tag Jonathan A. Brain Galina Gulis, Ph.D. 1 Kevin E. Redding, Ph.D. 2 Associate Professor of Chemistry Adjunct
Affinity Purification of Photosystem I from Chlamydomonas reinhardtii using a Polyhistidine Tag Jonathan A. Brain Galina Gulis, Ph.D. 1 Kevin E. Redding, Ph.D. 2 Associate Professor of Chemistry Adjunct
CHAPTER 4 RESULTS. showed that all three replicates had similar growth trends (Figure 4.1) (p<0.05; p=0.0000)
 CHAPTER 4 RESULTS 4.1 Growth Characterization of C. vulgaris 4.1.1 Optical Density Growth study of Chlorella vulgaris based on optical density at 620 nm (OD 620 ) showed that all three replicates had similar
CHAPTER 4 RESULTS 4.1 Growth Characterization of C. vulgaris 4.1.1 Optical Density Growth study of Chlorella vulgaris based on optical density at 620 nm (OD 620 ) showed that all three replicates had similar
BIOLOGICAL DISTRIBUTION. Prokaryotes
 Microbial degradation of aromatic organic pollutants A study of dead-end metabolites including the stereochemical mechanistic studies of cycloisomerase enzyme and their products using deuterium labelled
Microbial degradation of aromatic organic pollutants A study of dead-end metabolites including the stereochemical mechanistic studies of cycloisomerase enzyme and their products using deuterium labelled
Supporting information
 Supporting information Rhabdopeptide/Xenortide-like Peptides from Xenorhabdus innexi with Terminal Amines Showing Potent Anti-protozoal Activity Lei Zhao,, Marcel Kaiser,, Helge B. Bode *,, Molekulare
Supporting information Rhabdopeptide/Xenortide-like Peptides from Xenorhabdus innexi with Terminal Amines Showing Potent Anti-protozoal Activity Lei Zhao,, Marcel Kaiser,, Helge B. Bode *,, Molekulare
Metabolism. Topic 11&12 (ch8) Microbial Metabolism. Metabolic Balancing Act. Topics. Catabolism Anabolism Enzymes
 Topic 11&12 (ch8) Microbial Metabolism Topics Metabolism Energy Pathways Biosynthesis 1 Catabolism Anabolism Enzymes Metabolism 2 Metabolic Balancing Act Catabolism Enzymes involved in breakdown of complex
Topic 11&12 (ch8) Microbial Metabolism Topics Metabolism Energy Pathways Biosynthesis 1 Catabolism Anabolism Enzymes Metabolism 2 Metabolic Balancing Act Catabolism Enzymes involved in breakdown of complex
Introduction to Metabolism Cell Structure and Function
 Introduction to Metabolism Cell Structure and Function Cells can be divided into two primary types prokaryotes - Almost all prokaryotes are bacteria eukaryotes - Eukaryotes include all cells of multicellular
Introduction to Metabolism Cell Structure and Function Cells can be divided into two primary types prokaryotes - Almost all prokaryotes are bacteria eukaryotes - Eukaryotes include all cells of multicellular
Coupled, interconnecting reactions
 Metabolism: Basic concepts Hand-out for the CBT version November 2011 This module is based on 'Biochemistry' by Berg, Tymoczko and Stryer, seventh edition (2011), Chapter 15: Metabolism: Basic Concepts
Metabolism: Basic concepts Hand-out for the CBT version November 2011 This module is based on 'Biochemistry' by Berg, Tymoczko and Stryer, seventh edition (2011), Chapter 15: Metabolism: Basic Concepts
VaTx1 VaTx2 VaTx3. VaTx min Retention Time (min) Retention Time (min)
 a Absorbance (mau) 5 2 5 3 4 5 6 7 8 9 6 2 3 4 5 6 VaTx2 High Ca 2+ Low Ca 2+ b 38.2 min Absorbance (mau) 3 2 3 4 5 3 2 VaTx2 39.3 min 3 4 5 3 2 4. min 3 4 5 Supplementary Figure. Toxin Purification For
a Absorbance (mau) 5 2 5 3 4 5 6 7 8 9 6 2 3 4 5 6 VaTx2 High Ca 2+ Low Ca 2+ b 38.2 min Absorbance (mau) 3 2 3 4 5 3 2 VaTx2 39.3 min 3 4 5 3 2 4. min 3 4 5 Supplementary Figure. Toxin Purification For
بسم هللا الرحمن الرحيم
 بسم هللا الرحمن الرحيم -Please refer to the slides from (4-20) -Slides (4, 5) -Oxidative phosphorylation consists of 2 parts: 1.electron transport chain (series of electron transport proteins much filled
بسم هللا الرحمن الرحيم -Please refer to the slides from (4-20) -Slides (4, 5) -Oxidative phosphorylation consists of 2 parts: 1.electron transport chain (series of electron transport proteins much filled
Key Words: Brassica oleraceae, glucosinolate, liquid chromatography mass spectrometry, FNH-I-003
 IDENTIFICATION OF MAJOR GLUCOSINOLATES IN BROCCOLI (Brassica oleracea var. italica) BY LIQUID CHROMATOGRAPHY MASS SPECTROMETRY (LC-MS) AND DETERMINATION OF ANTICANCER PROPERTIES OF BROCCOLI EXTRACTS Carlos
IDENTIFICATION OF MAJOR GLUCOSINOLATES IN BROCCOLI (Brassica oleracea var. italica) BY LIQUID CHROMATOGRAPHY MASS SPECTROMETRY (LC-MS) AND DETERMINATION OF ANTICANCER PROPERTIES OF BROCCOLI EXTRACTS Carlos
Ch. 9 Cell Respiration. Title: Oct 15 3:24 PM (1 of 53)
 Ch. 9 Cell Respiration Title: Oct 15 3:24 PM (1 of 53) Essential question: How do cells use stored chemical energy in organic molecules and to generate ATP? Title: Oct 15 3:28 PM (2 of 53) Title: Oct 19
Ch. 9 Cell Respiration Title: Oct 15 3:24 PM (1 of 53) Essential question: How do cells use stored chemical energy in organic molecules and to generate ATP? Title: Oct 15 3:28 PM (2 of 53) Title: Oct 19
Characterizing fatty acids with advanced multinuclear NMR methods
 Characterizing fatty acids with advanced multinuclear NMR methods Fatty acids consist of long carbon chains ending with a carboxylic acid on one side and a methyl group on the other. Most naturally occurring
Characterizing fatty acids with advanced multinuclear NMR methods Fatty acids consist of long carbon chains ending with a carboxylic acid on one side and a methyl group on the other. Most naturally occurring
Eszopiclone (Lunesta ): An Analytical Profile
 Eszopiclone (Lunesta ): An Analytical Profile Roxanne E. Franckowski, M.S.* and Robert A. Thompson, Ph.D. U.S. Department of Justice Drug Enforcement Administration Special Testing and Research Laboratory
Eszopiclone (Lunesta ): An Analytical Profile Roxanne E. Franckowski, M.S.* and Robert A. Thompson, Ph.D. U.S. Department of Justice Drug Enforcement Administration Special Testing and Research Laboratory
KE-SIALIQ Sialic Acid Quantitation Kit. SialiQuant Sialic Acid Quantitation Kit
 SialiQuant Sialic Acid Quantitation Kit Part Number KE-SIALIQ Certification of Analysis Lot Number 706.1A Kit Storage Kits should be stored at 4 C. Kit Contents Kit contains all the reagents to quickly
SialiQuant Sialic Acid Quantitation Kit Part Number KE-SIALIQ Certification of Analysis Lot Number 706.1A Kit Storage Kits should be stored at 4 C. Kit Contents Kit contains all the reagents to quickly
Determination of Tetracyclines in Chicken by Solid-Phase Extraction and High-Performance Liquid Chromatography
 Determination of Tetracyclines in Chicken by Solid-Phase Extraction and High-Performance Liquid Chromatography Application ote Food Safety Authors Chen-Hao Zhai and Yun Zou Agilent Technologies Co. Ltd.
Determination of Tetracyclines in Chicken by Solid-Phase Extraction and High-Performance Liquid Chromatography Application ote Food Safety Authors Chen-Hao Zhai and Yun Zou Agilent Technologies Co. Ltd.
Cells extract energy from their environment and use the energy for a host of biological activities including biosynthesis.
 ATP=cellular energy Cells extract energy from their environment and use the energy for a host of biological activities including biosynthesis. The reactions of energy extraction and energy use are called
ATP=cellular energy Cells extract energy from their environment and use the energy for a host of biological activities including biosynthesis. The reactions of energy extraction and energy use are called
TENOFOVIR TABLETS: Final text for addition to The International Pharmacopoeia (June 2010)
 June 2010 TENOFOVIR TABLETS: Final text for addition to The International Pharmacopoeia (June 2010) This monograph was adopted at the Forty-fourth WHO Expert Committee on Specifications for Pharmaceutical
June 2010 TENOFOVIR TABLETS: Final text for addition to The International Pharmacopoeia (June 2010) This monograph was adopted at the Forty-fourth WHO Expert Committee on Specifications for Pharmaceutical
Chapter 5 Microbial Metabolism: The Chemical Crossroads of Life
 Chapter 5 Microbial Metabolism: The Chemical Crossroads of Life Copyright The McGraw-Hill Companies, Inc. Permission required for reproduction or display. The Metabolism of Microbes metabolism all chemical
Chapter 5 Microbial Metabolism: The Chemical Crossroads of Life Copyright The McGraw-Hill Companies, Inc. Permission required for reproduction or display. The Metabolism of Microbes metabolism all chemical
Measuring Lipid Composition LC-MS/MS
 Project: Measuring Lipid Composition LC-MS/MS Verification of expected lipid composition in nanomedical controlled release systems by liquid chromatography tandem mass spectrometry AUTHORED BY: DATE: Sven
Project: Measuring Lipid Composition LC-MS/MS Verification of expected lipid composition in nanomedical controlled release systems by liquid chromatography tandem mass spectrometry AUTHORED BY: DATE: Sven
Identification & Confirmation of Structurally Related Degradation Products of Simvastatin
 Identification & Confirmation of Structurally Related Degradation Products of Simvastatin Power of QTRAP Systems for Identification and Confirmation of Degradation Products Dilip Reddy 1, Chandra Sekar
Identification & Confirmation of Structurally Related Degradation Products of Simvastatin Power of QTRAP Systems for Identification and Confirmation of Degradation Products Dilip Reddy 1, Chandra Sekar
Metabolism. Chapter 8 Microbial Metabolism. Metabolic balancing act. Catabolism Anabolism Enzymes. Topics. Metabolism Energy Pathways Biosynthesis
 Chapter 8 Microbial Metabolism Topics Metabolism Energy Pathways Biosynthesis Catabolism Anabolism Enzymes Metabolism 1 2 Metabolic balancing act Catabolism and anabolism simple model Catabolism Enzymes
Chapter 8 Microbial Metabolism Topics Metabolism Energy Pathways Biosynthesis Catabolism Anabolism Enzymes Metabolism 1 2 Metabolic balancing act Catabolism and anabolism simple model Catabolism Enzymes
Work-flow: protein sample preparation Precipitation methods Removal of interfering substances Specific examples:
 Dr. Sanjeeva Srivastava IIT Bombay Work-flow: protein sample preparation Precipitation methods Removal of interfering substances Specific examples: Sample preparation for serum proteome analysis Sample
Dr. Sanjeeva Srivastava IIT Bombay Work-flow: protein sample preparation Precipitation methods Removal of interfering substances Specific examples: Sample preparation for serum proteome analysis Sample
Carboxylic Acid Derivatives Reading Study Problems Key Concepts and Skills Lecture Topics: Structures and reactivity of carboxylic acid derivatives
 Carboxylic Acid Derivatives Reading: Wade chapter 21, sections 21-1- 21-16 Study Problems: 21-45, 21-46, 21-48, 21-49, 21-50, 21-53, 21-56, 21-58, 21-63 Key Concepts and Skills: Interpret the spectra of
Carboxylic Acid Derivatives Reading: Wade chapter 21, sections 21-1- 21-16 Study Problems: 21-45, 21-46, 21-48, 21-49, 21-50, 21-53, 21-56, 21-58, 21-63 Key Concepts and Skills: Interpret the spectra of
Edgar Naegele. Abstract
 Simultaneous determination of metabolic stability and identification of buspirone metabolites using multiple column fast LC/TOF mass spectrometry Application ote Edgar aegele Abstract A recent trend in
Simultaneous determination of metabolic stability and identification of buspirone metabolites using multiple column fast LC/TOF mass spectrometry Application ote Edgar aegele Abstract A recent trend in
Manja Henze, Dorothee Merker and Lothar Elling. 1. Characteristics of the Recombinant β-glycosidase from Pyrococcus
 S1 of S17 Supplementary Materials: Microwave-Assisted Synthesis of Glycoconjugates by Transgalactosylation with Recombinant Thermostable β-glycosidase from Pyrococcus Manja Henze, Dorothee Merker and Lothar
S1 of S17 Supplementary Materials: Microwave-Assisted Synthesis of Glycoconjugates by Transgalactosylation with Recombinant Thermostable β-glycosidase from Pyrococcus Manja Henze, Dorothee Merker and Lothar
Enzymes what are they?
 Topic 11 (ch8) Microbial Metabolism Topics Metabolism Energy Pathways Biosynthesis 1 Catabolism Anabolism Enzymes Metabolism 2 Metabolic balancing act Catabolism Enzymes involved in breakdown of complex
Topic 11 (ch8) Microbial Metabolism Topics Metabolism Energy Pathways Biosynthesis 1 Catabolism Anabolism Enzymes Metabolism 2 Metabolic balancing act Catabolism Enzymes involved in breakdown of complex
Antoine Bouchoux, Pierre-Emerson Cayemitte, Julien Jardin, Geneviève Gésan-Guiziou, and Bernard Cabane
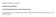 Biophysical Journal, Volume 96 Supplementary Material Casein Micelle Dispersions under Osmotic Stress Antoine Bouchoux, Pierre-Emerson Cayemitte, Julien Jardin, Geneviève Gésan-Guiziou, and Bernard Cabane
Biophysical Journal, Volume 96 Supplementary Material Casein Micelle Dispersions under Osmotic Stress Antoine Bouchoux, Pierre-Emerson Cayemitte, Julien Jardin, Geneviève Gésan-Guiziou, and Bernard Cabane
Metabolomics: quantifying the phenotype
 Metabolomics: quantifying the phenotype Metabolomics Promises Quantitative Phenotyping What can happen GENOME What appears to be happening Bioinformatics TRANSCRIPTOME What makes it happen PROTEOME Systems
Metabolomics: quantifying the phenotype Metabolomics Promises Quantitative Phenotyping What can happen GENOME What appears to be happening Bioinformatics TRANSCRIPTOME What makes it happen PROTEOME Systems
Purdue Institute for Drug Discovery, Purdue University, 720 Clinic Drive, West Lafayette, IN 47907, USA. b
 Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2016 Florescent analogs of cyclic and linear dinucleotides as phosphodiesterase and oligoribonuclease
Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2016 Florescent analogs of cyclic and linear dinucleotides as phosphodiesterase and oligoribonuclease
BIL 256 Cell and Molecular Biology Lab Spring, Tissue-Specific Isoenzymes
 BIL 256 Cell and Molecular Biology Lab Spring, 2007 Background Information Tissue-Specific Isoenzymes A. BIOCHEMISTRY The basic pattern of glucose oxidation is outlined in Figure 3-1. Glucose is split
BIL 256 Cell and Molecular Biology Lab Spring, 2007 Background Information Tissue-Specific Isoenzymes A. BIOCHEMISTRY The basic pattern of glucose oxidation is outlined in Figure 3-1. Glucose is split
Supplementary Information
 Supplementary Information The conifer biomarkers dehydroabietic and abietic acids are widespread in Cyanobacteria Maria Sofia Costa, Adriana Rego, Vitor Ramos, Tiago Afonso, Sara Freitas, Marco Preto,
Supplementary Information The conifer biomarkers dehydroabietic and abietic acids are widespread in Cyanobacteria Maria Sofia Costa, Adriana Rego, Vitor Ramos, Tiago Afonso, Sara Freitas, Marco Preto,
