PERSPECTIVES. SMOCs: supramolecular organizing centres that control innate immunity
|
|
|
- Bennett Wiggins
- 6 years ago
- Views:
Transcription
1 Nature Reviews Immunology AOP, published online 31 October 2014; doi: /nri3757 PERSPECTIVES OPINION SMOCs: supramolecular organizing centres that control innate immunity Jonathan C. Kagan, Venkat Giri Magupalli and Hao Wu Abstract The diverse receptor families of the innate immune system activate signal transduction pathways that are important for host defence, but common themes to explain the operation of these pathways remain undefined. In this Opinion article, we propose on the basis of recent structural and cell biological studies the concept of supramolecular organizing centres (SMOCs) as location-specific higher-order signalling complexes in which increased local concentrations of signalling components promote the intrinsically weak allosteric interactions that are required for enzyme activation. We suggest that SMOCs are assembled on various membrane-bound organelles or other intracellular sites, which may assist signal amplification to reach a response threshold and potentially define the specificity of cellular responses that are induced in response to infectious and non-infectious insults. Perhaps no area of immunology has benefitted more from the sequencing of the human (and mouse) genome than that of innate immunity. Modern studies of innate immunity received widespread attention with the discovery in the late 1990s that Toll-like receptors (TLRs) link microbial detection with the induction of adaptive immunity 1. Because TLRs and their associated families of signalling proteins have sequence homology, surveying the human and mouse genomes for uncharacterized orthologous proteins became a common approach to study these biological processes. Thus, within a few years of the discovery of cell-surface and endosomal TLRs 1 3, more than 100 genes had been identified that regulate the signalling pathways induced by these receptors 4 8, as well as the functionally related, cytosolic NOD-like receptors (NLRs), RIG I-like receptors (RLRs) and others 9 (FIG. 1; TABLE 1). Individual members of these pattern recognition receptor (PRR) families detect conserved pathogenassociated molecular patterns (PAMPs) that are present on bacteria, viruses and fungi, or recognize intrinsic damage-associated molecular patterns (DAMPs) that are elicited by cellular injury. Upon ligand binding, these receptors activate numerous cellular responses to fight infection and restore homeostasis. The success of using bioinformatics and reverse genetics to study innate immune signalling pathways came at the expense of alternative strategies to address these areas. As such, studies of the biochemistry, cell biology and dynamics of these signalling pathways have been much less common. In fact, most early studies of TLRs and their associated signalling proteins did not include any analysis of the subcellular localization of the newly identified protein(s). Thus, although we know the identity of many genes that are involved in innate immunity, the functional mechanisms of the proteins encoded by these genes, and how they interact in space and time, are poorly understood. The lack of knowledge on the specific activities of the proteins that control innate immunity has given rise to biological models that do not address many aspects of the signalling process, such as the subcellular site where a given signalling event occurs, or the dynamics of putative protein protein interactions. Current models of TLR, NLR or RLR signalling rather depict a series of arrows connecting receptors with downstream signalling proteins, yet we have little understanding of what these arrows actually represent. Do they represent direct protein protein interactions? If so, are these interactions constitutive or are they induced upon microbial encounter? How are these inter actions regulated and where in the cell do they occur? As described below, recent biochemical and cell biological studies have provided important insight into these questions. These new studies indicate that numerous protein regulators of innate immunity are organized into higher-order signalling complexes that define the subcellular sites and specificity of innate immune signal transduction. In this Opinion article, we propose that these higher-order signalling complexes function as supramolecular organizing centres (SMOCs) that control cellular responses induced by specific families of upstream receptors. We discuss how SMOCs can operate from various locations within the cell, and describe how they consist of proteins that either sense the activation of upstream receptors or elicit specific downstream effector responses. We further propose that these complexes include context-dependent components, which may be cell type-specific or organelle-specific regulators, such that a given SMOC can elicit diverse cellular responses depending on the stimulus. A benefit of coordinating innate immune signalling pathways around a set of organizing centres may be the modularity of the system, whereby numerous upstream stimuli can be directed into a common downstream module. Indeed, this is the case when considering the operation of other nonmembranous organizing centres in mammalian cells, such as the microtubule organizing centre (MTOC) and the proteasome. In these examples, a large protein complex coordinates an entire biological process that may be needed to address diverse cellular needs. SMOCs for PRRs The classical view of signal transduction involving a serial reaction in which ligands induce conformational changes in receptors followed by the activation of enzymes and the generation of second messengers required NATURE REVIEWS IMMUNOLOGY ADVANCE ONLINE PUBLICATION 1
2 TLR4 Cytoplasm Viral RNA RIG-I or MDA5 IRAK1 or IRAK2 TIRAP MYD88 Myddosome PtdIns(4,5)P 2 IRAK4 Endosome TLR9 Mitochondrial dysfunction leading to cardiolipin exposure MAVS NLRP3 (autoinhibited) Cardiolipin RLR signalling complex (filamentous MAVS) PtdIns3P NOD PYD LRR CARD ASC NLRP3 inflammasome Pro-caspase 1 TRAF TBK1 IRF TRAF6 TAK1 TAB1 IκBα p50 p65 Myddosome Pro-IL-1β IL-1β IRF NF-κB p50 p65 Type I IFN Pro-inflammatory cytokines Nucleus Figure 1 SMOC formation for TLRs, RLRs and NLRs. Depicted are the best-studied supramolecular organizing centres (SMOCs), including the ligands and regulatory proteins that promote their assembly, and the downstream biological activities induced by these protein complexes. The figure does not show the exact stoichiometry of the protein components in each signalling complex. Binding of lipopolysaccharide (not shown) activates Toll-like receptor 4 (TLR4), leading to assembly of a Myddosome on the plasma membrane. By contrast, unmethylated CpG-containing DNA oligonucleotides (not shown) promote TLR9 to assemble an endosomal Myddosome. The TLR-specific sorting adaptor Toll/IL 1R domain-containing adaptor protein (TIRAP) facilitates Myddosome assembly. TIRAP has an amino terminal lipid-binding domain that interacts promiscuously with acidic phosphoinositides and phosphatidylserine. For example, TIRAP is depicted as binding phosphatidylinositol 3 phosphate (PtdIns3P) and phosphatidylinositol 4,5 bisphosphate (PtdIns(4,5)P 2 ) on the endosomal membrane and the plasma membrane, respectively. Binding of cardiolipin, which translocates to the outer mitochondrial membrane upon mitochondrial dysfunction, relieves the autoinhibited state of NOD-, LRR- and pyrin domain-containing 3 (NLRP3). This in turn may promote NLRP3 inflammasome assembly through downstream pyrin domain (PYD) PYD and caspase activation and recruitment domain (CARD) CARD interactions. Activation of RIG I like receptors retinoic acid-inducible gene I (RIG I) or melanoma differentiation-associated protein 5 (MDA5) leads to the formation of higher-order oligomers of mitochondrial antiviral signalling protein (MAVS). IFN, interferon; IκBα, NF-κB inhibitor-α; IL, interleukin; IRAK, IL 1 receptor-associated kinase; IRF, IFN-regulatory factor; LRR, leucine-rich repeat; MYD88, myeloid differentiation primary response protein 88; NOD, nucleotide-binding oligomerization domain; NF-κB, nuclear factor-κb; TAB1, TAK1-binding protein 1; TAK1, TGFβ-associated kinase 1; TRAF, tumour necrosis factor receptor-associated factor. serious modifications in the case of PRRs. The emerging concept for PRRs supports the formation of higher-order signalling complexes as the mode of signal transduction and amplification. Indeed, with the exception of the signalling pathway involving cyclic GMP AMP synthase (cgas) and stimulator of interferon genes protein (STING) described below, there is little or no role of second messengers in the earliest events associated with PRR signal transduction. Early studies. Cellular studies using light microscopy imaging provided the first evidence for the existence of higher-order signalling complexes in innate immune pathways. The tumour necrosis factor (TNF) receptor superfamily comprises some of the earliest discovered members of the innate immune system, such as TNF receptor 1 (TNFR1) and FAS (also known as CD95 and TNFRSF6). These receptors do not have intrinsic enzymatic activity, but aggregate upon stimulation by trimeric ligands of the corresponding TNF superfamily. As an example, FAS as part of the death-inducing signalling complex (DISC) forms aggregated clusters or puncta of hundreds of nanometres to micrometres in diameter 10,11, which are much larger than ligand receptor trimers. The formation of sizable clusters that are visible by light microscopy has become a recurrent observation for activated 2 ADVANCE ONLINE PUBLICATION
3 innate immune receptors, including TLRs 12 14, RLRs 15,16, and NLR or absent in melanoma 2 (AIM2) inflammasomes 17,18 (FIG. 1; TABLE 1). Biochemical and structural studies of innate immune signalling complexes indicated that such protein clusters are not random aggregates but instead have a defined molecular basis of assembly. An almost ubiquitous feature of innate immune pathways is the participation of signal transduction proteins with death domains, or the related death effector domains (DEDs), caspase activation and recruitment domains (CARDs) and pyrin domains (PYDs) 19. Crystal structures of oligo meric death domain complexes revealed an ordered helical assembly mechanism that underlies the oligomerization of several proteins 19 including the 5:7 complex of p53 inducible protein with a death domain (PIDD) and RIP-associated ICH1/CED3 homologous protein with a death domain (RAIDD; also known as CRADD) in the core of the PIDDosome for caspase 2 activation 20 ; the 5:5 complex of FAS and FAS-associated death domain protein (FADD) in the DISC for caspase 8 activation 21 ; and the 6:4:4 complex of myeloid differentiation primary response protein 88 (MYD88), interleukin 1 receptor (IL 1R)-associated kinase 4 (IRAK4) and IRAK2 in the Myddosome for kinase activation in the TLR pathway 22 (TABLE 1). The common helical symmetry indicates that death domains and their related domains might be able to form large helical filaments and even larger filamentous signalling complexes to generate microscopically visible SMOCs. Recent advances. Exciting new studies have now confirmed the structural predictions, as well as further extended our mechanistic understanding of innate immune signalling. The RLR signalling adaptor mitochondrial antiviral signalling protein (MAVS), which contains an amino terminal CARD, forms helical filaments that activate the interferon pathway 16,23,24 (FIG. 2). The CARD-containing adaptor B cell lymphoma 10 (BCL 10) which activates mucosa-associated lymphoid tissue lymphoma translocation protein 1 (MALT1) for nuclear factor-κb (NF κb) signalling downstream of several innate and adaptive immune receptors assembles into similar helical filaments 25. Recent studies of cytosolic inflammasomes, which activate caspase 1 to induce interleukin 1β (IL 1β) maturation and pyroptosis, show Table 1 SMOCs of the innate immune response SMOC Triggers or ligands Receptors or sensors that filaments of the adaptor protein ASC (which contains both a PYD and a CARD) and caspase 1 (which contains a CARD) form star-shaped structures that mediate signal transduction and proximityinduced caspase 1 activation (FIG. 2). Electron microscopy-based structural studies showed that these filaments are assembled through three conserved types of interactions that were initially observed in the crystal structures of death domain complexes 24,25,27. These new advances indicate that nucleated polymerization may be a general principle in promoting the assembly of filamentous complexes upon stimulation to mediate innate immune signal transduction (FIG. 2). A sensor protein (receptor) in a pathway first becomes activated in the presence of ligands by overcoming autoinhibition and subsequently oligomerizes. The oligomerized sensor then provides the platform to nucleate a downstream protein to polymerize into filaments. For example, recognition of cytosolic double-stranded DNA (dsdna) by the AIM2 inflammasome or the response to various infectious and danger signals by NLR inflammasomes disrupts intra molecular domain interactions 29 to enable oligomerization of the PYDs of inflamma some sensor proteins 26,27. These oligomerized PYDs then function as a platform to nucleate the polymerization of the PYDs in the adaptor protein ASC. The ASC PYD filaments bring the CARDs of ASC into proximity to nucleate the polymerization of the CARDs of caspase 1, which in turn promotes caspase 1 dimerization and activation. Similarly, the polyubiquitin-stabilized double CARD of retinoic Adaptors Effectors Functions FAS DISC FAS ligand FAS FADD Caspase 8 Apoptosis PIDDosome DNA damage PIDD RAIDD Caspase 2 Apoptosis Myddosome PAMPs TLRs TIRAP and MYD88 RLR complex Viral RNAs RIG I and MDA5 Inflammasome PAMPs and DAMPs NLRs and ALRs IRAKs NF κb activation MAVS TRAFs NF κb activation and interferon response ASC Caspase 1 Pyroptosis and IL 1β maturation ALR, AIM2 like receptor; DAMP, damage-associated molecular pattern; DISC, death-inducing signalling complex; FADD, FAS-associated death domain protein; IL 1β, interleukin 1β; IRAK, IL 1 receptorassociated kinase; MAVS, mitochondrial antiviral signalling protein; MDA5, melanoma differentiationassociated protein 5; MYD88, myeloid differentiation primary response protein 88; NF-κB, nuclear factor-κb; NLR, NOD-like receptor; PAMP, pathogen-associated molecular pattern; PIDD, p53 inducible protein with a death domain; RAIDD, RIP-associated ICH1/CED3 homologous protein with a death domain; RIG-I, retinoic acid-inducible gene I; RLR, RIG-I-like receptor; SMOC, supramolecular organizing centre; TIRAP, Toll/IL 1R domain-containing adaptor protein; TLR, Toll-like receptor; TRAF, tumour necrosis factor receptor-associated factor. acid-inducible gene I (RIG I) forms a helical tetramer to imprint the polymerization of MAVS through its CARDs to promote downstream signalling 24, Oligomerization mechanisms that do not involve the death domain superfamily have also been discovered in innate immunity 14,33. A common molecular mechanism of signal transduction seems to be the tight coupling between oligomerization regardless of the type and allostery. On the one hand, ligand-induced allosteric changes in the respective receptors promote oligomerization. On the other hand, oligomerization of effector enzymes, such as kinases and caspases, enhances the allosteric changes that are presumed to be required for enzyme activation. This oligomerizationfacilitated allostery is illustrated by the recently reported crystal structure of the unphosphorylated IRAK4 kinase domain asymmetric dimer captured in the conformation of the Myddosome-induced IRAK4 trans-autophosphorylation reaction 34. In the crystal structure, one IRAK4 monomer exists in the active kinase conformation despite a lack of phosphorylation and it precisely binds the activation loop phosphosite of the other IRAK4 monomer for phosphotransfer. In solution, unphosphorylated IRAK4 forms weak dimers. By markedly increasing the local concentration of IRAK4, the Myddosome promotes dimerization to drive allosteric autoactivation of IRAK4. Oligomerization-driven allosteric changes probably have crucial roles in the higherorder signalling complexes of other innate immune receptors; it is the challenge of structural biologists to characterize these often weak and transient conformations. NATURE REVIEWS IMMUNOLOGY ADVANCE ONLINE PUBLICATION 3
4 a b c Modelled AIM2 PYD filament RIG-I 2CARD tetramer Ligand stimulation Receptor monomer Receptor oligomerization and nucleus formation ASC PYD filament structure MAVS CARD filament structure Adaptor monomer Nucleated adaptor polymerization Adaptor oligomer Figure 2 Structures of SMOCs that are formed by the mechanism of nucleated polymerization. a A ribbon diagram of the electron cryomicroscopic structure of polymerized ASC pyrin domain (ASC PYD ) filaments in inflammasomes (shown in red, purple and orange for each of the three helical strands), in complex with polymerized absent in melanoma 2 PYD (AIM2 PYD ) filaments (shown in blue) formed upon double-stranded DNA (dsdna) stimulation. b A ribbon diagram of the electron cryomicroscopic structure of polymerized mitochondrial antiviral signalling protein caspase activation and recruitment domain (MAVS CARD ) in the RIG I-like receptor (RLR) pathway (shown in purple), in complex with a retinoic acid-inducible gene I (RIG-I) double CARD (RIG-I 2CARD ) tetramer (shown in blue, cyan, green and yellow) upon viral RNA stimulation. c Proposed mechanism of nucleated polymerization for the formation of ASC PYD and MAVS CARD SMOCs. Receptors (for example, AIM2 and RIG I) are shown in red wedges for monomers and in red disks for oligomerized forms. Adaptors (for example, ASC and MAVS) are shown in blue wedges for monomers and in blue cylinders for filaments. Mechanistic implications. The assembly of higher-order signalling complexes not only provides a phenomenological explanation for the observation of punctate structures in cells, but also implicates an elegant mechanism of signal amplification, preceding enzyme activation, that enables a response threshold to be reached. Nucleated polymerization ensures that a small number of sensor proteins can activate many downstream signalling proteins. For example, in the AIM2 inflammasome, a substoichiometric amount of AIM2 can polymerize many more ASC molecules 27. In turn, activated ASC further oligomerizes many more caspase 1 molecules to amplify signal transduction. Within a single cell, it is likely that once the signalling cascade is successfully initiated, almost all caspase 1 molecules are recruited to inflammasomes so that a maximal response is generated. Signal amplification and the cooperativity in the assembly of higher-order signalling complexes may both contribute to the threshold, all or none host defence programmes that have been observed at the single-cell level for many innate immune pathways 35,36. In addition, signal amplification may explain the sensitivity of mammalian cells to incredibly small numbers of bacteria. For example, it has been estimated that a single Escherichia coli bacterium can activate 1,000 macrophages through TLR4 (REF. 37), suggesting that very few receptors on any individual cell are engaged during infection. We suggest that SMOC formation around the cytosolic tail of TLR4 may explain the all or none response that macrophages often exhibit to a wide range of lipopolysaccharide (LPS) concentrations 37. In addition, the role of SMOCs in signal amplification may be fundamentally analogous to that of second messengers, such as camp in signal transduction by G protein-coupled receptors. However, we believe that camp-mediated responses may be more graded than those mediated by SMOCs, with the strength of the response being determined by the amount, half-life and deactivation kinetics of the second messengers. If signal amplification to reach a response threshold is the mechanism by which innate immune pathways are turned on, then once formed, how are SMOCs turned off to terminate signalling? In the case of BCL 10 containing signalosomes, they are recruited to autophagosomes through an interaction between ubiquitylated BCL 10 and the autophagy adaptor p62 (also known as SQSTM1), leading to degradation 38. Deficiency of ATG16L1 a protein that is required for autophagosome formation has been shown to enhance endotoxininduced IL 1β and IL 18 production, probably through a TLR-mediated pathway 39. Given that SMOCs may be too large to be degraded efficiently by proteasomes, autophagosome-mediated degradation seems to make sense. It is interesting to note that the size distribution of puncta can be quite different for different SMOCs. For example, TLRs form relatively small clusters, which may account for the smaller size of the Myddosome compared with inflammasome filaments. Some inflammasomes, as well as STING (which mediates the interferon response upon stimulation of cgas by cytosolic dsdna), have gigantic perinuclear puncta 40,41. Whether the distinct size of puncta is reflective of the threshold of activation and/or degradation mechanisms remains to be addressed. Consistent with the ability of many death domain superfamily members to polymerize, several of these domains including the CARD of MAVS and the PYD of ASC have prion-like activities in yeast 16,27,28. In mammalian cells, it seems that ASC-containing specks of inflammasomes are not degraded rapidly but are released upon pyroptosis into the extracellular space, where they promote further 4 ADVANCE ONLINE PUBLICATION
5 IL 1β processing and can be engulfed by macrophages to induce inflammasome activation in the recipient cells 42,43. Therefore, understanding the biophysical principles of SMOC assembly and degradation not only provides insights into unique signalling properties within each cell, but also implicates mechanisms of unexpected signal transduction between cells. The subcellular localization of SMOCs Cell biological and biochemical analyses of individual components of SMOCs have shown that these complexes are usually assembled on the cytosolic surface of membranous organelles of mammalian cells. This trend of SMOC assembly on membranes can be observed for TLRs, as well as for RLRs and NLRs. Because the RLRs and NLRs are not transmembrane proteins, their need to induce SMOC assembly on organelles is intriguing, and it indicates that membranebased SMOC assembly is not simply a consequence of signal transduction being initiated by a transmembrane receptor such as a TLR. Toll-like receptors. Although the Myddosome has been defined structurally 22, there is an incomplete understanding of the composition, dynamics and regulation of the endogenous TLR-induced Myddosome in mammalian cells. Recent work has provided insight into the behaviour of the Myddosome in macrophages, through the demonstration that it can form either at the plasma membrane or on endosomes 44. For example, LPS treatment of macrophages activates TLR4 to assemble a Myddosome at the cell surface, whereas unmethylated CpG-containing DNA oligonucleotides activate TLR9 to assemble an endosomal Myddosome. In both cases, an activated receptor is not sufficient to induce Myddosome assembly; rather, the TLRspecific Toll/IL 1R (TIR) domain-containing adaptor protein (TIRAP; also known as MAL) is required for Myddosome assembly 44. Thus, TIRAP is the first natural regulator of TLR-induced Myddosome formation to be identified. TIRAP is a peripheral membrane protein that contains an N terminal lipid-binding domain that interacts promiscuously with acidic phosphoinositides and phosphatidylserine 45. This promiscuity of lipid binding enables TIRAP to survey several plasma membrane and endosomal subdomains for the presence of an activated (ligand-bound) TLR. When TIRAP detects a ligand-bound TLR through its carboxyterminal TIR domain 46,47, it recruits the core component of the Myddosome, MYD88, which seeds the formation of this SMOC through interactions with IRAKs 22,44,48. Altering the lipid specificity of TIRAP such that it can only interact with either plasma membrane-localized or endosome-localized lipids restricts the ability of the cell to assemble Myddosomes from the location in which TIRAP resides 44. Thus, although TIRAP contains a promiscuous lipid-binding domain, the individual targets of this domain enable location-specific assembly of a SMOC. TIRAP was originally thought to be a regulator that defines the specificity of signalling pathway activation induced by plasma membrane-localized TLRs, as compared with endosomal TLRs 8. The observation that TIRAP regulates Myddosome formation from both subcellular locations reignites the question of how organelle-specific innate immune responses are achieved. We suggest that whereas TIRAP and the other components of the Myddosome participate in TLR signalling from multiple locations within the cell, organelle-specific regulators may exist to determine the specificity of signal transduction. Identifying these putative regulators may provide tools to understand how the composition and function of SMOCs can be modulated naturally, and perhaps therapeutically. RIG I like receptors. The RLR system assembles a SMOC on membranes, despite the fact that the upstream receptors are cytosolic proteins. The core of the RLRinduced SMOC is MAVS, which is localized to membranes though a C terminal transmembrane domain 15,16. Although the localization domain of MAVS is not structurally similar to that of TIRAP, these domains are similar in that they can be targeted to multiple organelles. In the case of MAVS, its transmembrane domain enables it to localize to mitochondria, peroxisomes and the mitochondria-associated membranes (MAMs) of the endoplasmic reticulum 15,49,50. Also, similar to TIRAP, MAVS can induce signalling responses from each of these organelles 15,49,50, although biochemical evidence for the formation of higher-order oligomers is only available for mitochondria-localized MAVS 16. It remains to be determined whether similar MAVS-containing SMOCs are assembled on peroxisomes and MAMs during viral infections. Recent studies have shown that functional TIRAP or MAVS proteins can be generated by replacing the localization domains of either of these proteins with other domains that have similar membranetargeting activities 15,44,50. These data provide strong evidence that the sole function of these domains is to direct TIRAP or MAVS to the membranes that are necessary for PRR-induced SMOC assembly. Although it is clear that MAVS must localize to specific organelles to initiate SMOC assembly and signalling, it remains unclear why mitochondria, peroxisomes and MAMs have evolved as the preferred sites of RLR signal transduction. One possibility is that the metabolic functions of these organelles control MAVS activation, a theory that is supported by recent studies indicating that dysfunctional mitochondria cannot promote efficient RLR-dependent innate immune responses 51,52. NOD-like receptors. The NLR system provides yet another example of SMOC assembly on membranes, at least in the case of NOD-, LRR- and PYD-containing 3 (NLRP3). Although NLRP3 is generally thought to be a cytosolic protein, it translocates to mitochondria when cells are stimulated with several types of inflammasome activators 53,54. The translocation of NLRP3 to mitochondria has been reported to occur through interactions with the lipid cardiolipin which is only displayed on the outer membrane of damaged mitochondria 55 or with MAVS 56. In this regard, NLRP3 is similar to TIRAP in that its ability to promote SMOC assembly is linked to its ability to interact with both membrane-bound lipids (phosphoinositides) and proteins (TLRs) in a specific region of the cell. It is unclear whether other NLR family members also interact with cardiolipin or any other lipid to promote SMOC assembly, but these studies provide a strong mandate to consider this possibility. Conclusions and future perspectives Examples now exist for the three major families of PRRs TLRs, RLRs and NLRs that SMOC assembly occurs on membranes, even when the upstream receptors are cytosolic proteins. In each documented example, a membrane protein seeds the formation of a higher-order signalling complex that activates specific innate immune responses. A biophysical explanation for the apparent membrane localization of SMOCs may be the marked energetic enhancement of protein protein interactions on a two-dimensional membrane surface compared with those in a three-dimensional cellular milieu 57. For this reason, we propose that future studies of the biochemical mechanisms of SMOC assembly and function would benefit from NATURE REVIEWS IMMUNOLOGY ADVANCE ONLINE PUBLICATION 5
6 a greater consideration of the subcellular sites where this assembly occurs. This analysis may also help to address the question of why SMOCs have evolved to operate from specific organelles. Additional studies of the relationship between organelle function and SMOC assembly should address the possibility that organelles are not solely needed as a scaffold for SMOC assembly, but rather that a metabolic or biochemical activity of the organelle may contribute as well. This cell biological analysis may reveal important means of controlling and manipulating SMOC assembly and subsequent inflammatory responses. Jonathan C. Kagan, Venkat Giri Magupalli and Hao Wu are at the Boston Children s Hospital and Harvard Medical School, Boston, Massachusetts 02446, USA. Correspondence to J.C.K. and H.W. s: jonathan.kagan@childrens.harvard.edu; hao.wu@childrens.harvard.edu doi: /nri3757 Published online 31 October 2014; corrected online 3 November Medzhitov, R., Preston-Hurlburt, P. & Janeway, C. A. Jr. A human homologue of the Drosophila Toll protein signals activation of adaptive immunity. Nature 388, (1997). 2. Medzhitov, R. et al. MyD88 is an adaptor protein in the htoll/il 1 receptor family signaling pathways. Mol. Cell 2, (1998). 3. Poltorak, A. et al. Defective LPS signaling in C3H/HeJ and C57BL/10ScCr mice: mutations in Tlr4 gene. Science 282, (1998). 4. Yamamoto, M. et al. TRAM is specifically involved in the Toll-like receptor 4 mediated MyD88 independent signaling pathway. Nature Immunol. 4, (2003). 5. Yamamoto, M. et al. Role of adaptor TRIF in the MyD88 independent Toll-like receptor signaling pathway. Science 301, (2003). 6. Oshiumi, H. et al. TIR-containing adapter molecule (TICAM)-2, a bridging adapter recruiting to Toll-like receptor 4 TICAM 1 that induces interferon-β. J. Biol. Chem. 278, (2003). 7. Oshiumi, H., Matsumoto, M., Funami, K., Akazawa, T. & Seya, T. TICAM 1, an adaptor molecule that participates in Toll-like receptor 3 mediated interferon-β induction. Nature Immunol. 4, (2003). 8. Horng, T., Barton, G. M., Flavell, R. A. & Medzhitov, R. The adaptor molecule TIRAP provides signalling specificity for Toll-like receptors. Nature 420, (2002). 9. Kumar, H., Kawai, T. & Akira, S. Pathogen recognition by the innate immune system. Int. Rev. Immunol. 30, (2011). 10. Algeciras-Schimnich, A. et al. Molecular ordering of the initial signaling events of CD95. Mol. Cell. Biol. 22, (2002). 11. Siegel, R. M. et al. SPOTS: signaling protein oligomeric transduction structures are early mediators of death receptor-induced apoptosis at the plasma membrane. J. Cell Biol. 167, (2004). 12. Latz, E. et al. Ligand-induced conformational changes allosterically activate Toll-like receptor 9. Nature Immunol. 8, (2007). 13. Visintin, A., Latz, E., Monks, B. G., Espevik, T. & Golenbock, D. T. Lysines 128 and 132 enable lipopolysaccharide binding to MD 2, leading to Tolllike receptor 4 aggregation and signal transduction. J. Biol. Chem. 278, (2003). 14. Yin, Q. et al. E2 interaction and dimerization in the crystal structure of TRAF6 Nature Struct. Mol. Biol. 16, (2009). 15. Seth, R. B., Sun, L., Ea, C. K. & Chen, Z. J. Identification and characterization of MAVS, a mitochondrial antiviral signaling protein that activates NF κb and IRF 3. Cell 122, (2005). 16. Hou, F. et al. MAVS forms functional prion-like aggregates to activate and propagate antiviral innate immune response. Cell 146, (2011). 17. Hornung, V. et al. AIM2 recognizes cytosolic dsdna and forms a caspase 1 activating inflammasome with ASC. Nature 458, (2009). 18. Wu, J., Fernandes-Alnemri, T. & Alnemri, E. S. Involvement of the AIM2, NLRC4, and NLRP3 inflammasomes in caspase 1 activation by Listeria monocytogenes. J. Clin. Immunol. 30, (2010). 19. Ferrao, R. & Wu, H. Helical assembly in the death domain (DD) superfamily. Curr. Opin. Struct. Biol. 22, (2012). 20. Park, H. H. et al. Death domain assembly mechanism revealed by crystal structure of the oligomeric PIDDosome core complex. Cell 128, (2007). 21. Wang, L. et al. The Fas FADD death domain complex structure reveals the basis of DISC assembly and disease mutations. Nature Struct. Mol. Biol. 17, (2010). 22. Lin, S. C., Lo, Y. C. & Wu, H. Helical assembly in the MyD88 IRAK4 IRAK2 complex in TLR/IL 1R signalling. Nature 465, (2010). 23. Xu, H. et al. Structural basis for the prion-like MAVS filaments in antiviral innate immunity. Elife 3, e Wu, B. et al. Molecular imprinting as a signalactivation mechanism of the viral RNA sensor RIG I. Mol. Cell 55, Qiao, Q. et al. Structural architecture of the CARMA1/Bcl10/MALT1 signalosome: nucleationinduced filamentous assembly. Mol. Cell 51, (2013). 26. Lu, A., Kabaleeswaran, V., Fu, T., Magupalli, V. G. & Wu, H. Crystal structure of the F27G AIM2 PYD mutant and similarities of its self-association to DED/DED interactions. J. Mol. Biol. 426, Lu, A. et al. Unified polymerization mechanism for the assembly of ASC-dependent inflammasomes. Cell 156, Cai, X. et al. Prion-like polymerization underlies signal transduction in antiviral immune defense and inflammasome activation. Cell 156, Hu, Z. et al. Crystal structure of NLRC4 reveals its autoinhibition mechanism. Science 341, (2013). 30. Peisley, A., Wu, B., Yao, H., Walz, T. & Hur, S. RIG I forms signaling-competent filaments in an ATP-dependent, ubiquitin-independent manner. Mol. Cell 51, (2013). 31. Zeng, W. et al. Reconstitution of the RIG I pathway reveals a signaling role of unanchored polyubiquitin chains in innate immunity. Cell 141, (2010). 32. Jiang, X. et al. Ubiquitin-induced oligomerization of the RNA sensors RIG I and MDA5 activates antiviral innate immune response. Immunity 36, (2012). 33. Li, J. et al. The RIP1/RIP3 necrosome forms a functional amyloid signaling complex required for programmed necrosis. Cell 150, (2012). 34. Ferrao, R. et al. IRAK4 dimerization and transautophosphorylation are induced by myddosome assembly. Mol. Cell 55, Tay, S. et al. Single-cell NF κb dynamics reveal digital activation and analogue information processing. Nature 466, (2010). 36. Wu, H. Higher-order assemblies in a new paradigm of signal transduction. Cell 153, (2013). 37. Gioannini, T. L. & Weiss, J. P. Regulation of interactions of Gram-negative bacterial endotoxins with mammalian cells. Immunol. Res. 39, (2007). 38. Paul, S., Kashyap, A. K., Jia, W., He, Y. W. & Schaefer, B. C. Selective autophagy of the adaptor protein Bcl10 modulates T cell receptor activation of NF κb. Immunity 36, (2012). 39. Saitoh, T. et al. Loss of the autophagy protein Atg16L1 enhances endotoxin-induced IL 1β production. Nature 456, (2008). 40. Ishikawa, H. & Barber, G. N. STING is an endoplasmic reticulum adaptor that facilitates innate immune signalling. Nature 455, (2008). 41. Gao, D. et al. Cyclic GMP-AMP synthase is an innate immune sensor of HIV and other retroviruses. Science 341, (2013). 42. Franklin, B. S. et al. The adaptor ASC has extracellular and prionoid activities that propagate inflammation. Nature Immunol. 15, Baroja-Mazo, A. et al. The NLRP3 inflammasome is released as a particulate danger signal that amplifies the inflammatory response. Nature Immunol. 15, Bonham, K. S. et al. A promiscuous lipid-binding protein diversifies the subcellular sites of Toll-like receptor signal transduction. Cell 156, Kagan, J. C. & Medzhitov, R. Phosphoinositidemediated adaptor recruitment controls Toll-like receptor signaling. Cell 125, (2006). 46. Horng, T., Barton, G. M. & Medzhitov, R. TIRAP: an adapter molecule in the Toll signaling pathway. Nature Immunol. 2, (2001). 47. Fitzgerald, K. A. et al. Mal (MyD88 adapter-like) is required for Toll-like receptor 4 signal transduction. Nature 413, (2001). 48. Motshwene, P. G. et al. An oligomeric signaling platform formed by the Toll-like receptor signal transducers MyD88 and IRAK 4. J. Biol. Chem. 284, (2009). 49. Horner, S. M., Liu, H. M., Park, H. S., Briley, J. & Gale, M. Jr. Mitochondrial-associated endoplasmic reticulum membranes (MAM) form innate immune synapses and are targeted by hepatitis C virus. Proc. Natl Acad. Sci. USA 108, (2011). 50. Dixit, E. et al. Peroxisomes are signaling platforms for antiviral innate immunity. Cell 141, (2010). 51. Koshiba, T., Yasukawa, K., Yanagi, Y. & Kawabata, S. Mitochondrial membrane potential is required for MAVS-mediated antiviral signaling. Sci. Signal. 4, ra7 (2011). 52. Odendall, C. et al. Diverse intracellular pathogens activate type III interferon expression from peroxisomes. Nature Immunol. 15, Zhou, R., Yazdi, A. S., Menu, P. & Tschopp, J. A role for mitochondria in NLRP3 inflammasome activation. Nature 469, (2011). 54. Misawa, T. et al. Microtubule-driven spatial arrangement of mitochondria promotes activation of the NLRP3 inflammasome. Nature Immunol. 14, (2013). 55. Iyer, S. S. et al. Mitochondrial cardiolipin is required for Nlrp3 inflammasome activation. Immunity 39, (2013). 56. Subramanian, N., Natarajan, K., Clatworthy, M. R., Wang, Z. & Germain, R. N. The adaptor MAVS promotes NLRP3 mitochondrial localization and inflammasome activation. Cell 153, (2013). 57. Wu, Y., Vendome, J., Shapiro, L., Ben-Shaul, A. & Honig, B. Transforming binding affinities from three dimensions to two with application to cadherin clustering. Nature 475, (2011). Acknowledgements J.C.K. is supported by the US National Institutes of Health (NIH; grants AI093589, AI and AI ) and an unrestricted gift from Mead Johnson & Company. J.C.K. holds an Investigators in the Pathogenesis of Infectious Disease Award from the Burroughs Wellcome Fund. H.W. is supported by NIH and the Asa and Patricia Springer Professorship of the Harvard Medical School. Competing interests statement The authors declare no competing interests. 6 ADVANCE ONLINE PUBLICATION
7 Correction In Figure 1 of the original version of this article, the Toll-like receptor within the endosome was incorrectly labelled as TLR4. This should have been labelled as TLR9 and has now been corrected online. Nature Reviews Immunology apologizes for this error.
Toll-like Receptor Signaling
 Toll-like Receptor Signaling 1 Professor of Medicine University of Massachusetts Medical School, Worcester, MA, USA Why do we need innate immunity? Pathogens multiply very fast We literally swim in viruses
Toll-like Receptor Signaling 1 Professor of Medicine University of Massachusetts Medical School, Worcester, MA, USA Why do we need innate immunity? Pathogens multiply very fast We literally swim in viruses
Innate Immunity & Inflammation
 Innate Immunity & Inflammation The innate immune system is an evolutionally conserved mechanism that provides an early and effective response against invading microbial pathogens. It relies on a limited
Innate Immunity & Inflammation The innate immune system is an evolutionally conserved mechanism that provides an early and effective response against invading microbial pathogens. It relies on a limited
Newly Recognized Components of the Innate Immune System
 Newly Recognized Components of the Innate Immune System NOD Proteins: Intracellular Peptidoglycan Sensors NOD-1 NOD-2 Nod Protein LRR; Ligand Recognition CARD RICK I-κB p50 p65 NF-κB Polymorphisms in Nod-2
Newly Recognized Components of the Innate Immune System NOD Proteins: Intracellular Peptidoglycan Sensors NOD-1 NOD-2 Nod Protein LRR; Ligand Recognition CARD RICK I-κB p50 p65 NF-κB Polymorphisms in Nod-2
Innate immunity. Abul K. Abbas University of California San Francisco. FOCiS
 1 Innate immunity Abul K. Abbas University of California San Francisco FOCiS 2 Lecture outline Components of innate immunity Recognition of microbes and dead cells Toll Like Receptors NOD Like Receptors/Inflammasome
1 Innate immunity Abul K. Abbas University of California San Francisco FOCiS 2 Lecture outline Components of innate immunity Recognition of microbes and dead cells Toll Like Receptors NOD Like Receptors/Inflammasome
Intrinsic cellular defenses against virus infection
 Intrinsic cellular defenses against virus infection Detection of virus infection Host cell response to virus infection Interferons: structure and synthesis Induction of antiviral activity Viral defenses
Intrinsic cellular defenses against virus infection Detection of virus infection Host cell response to virus infection Interferons: structure and synthesis Induction of antiviral activity Viral defenses
2. Innate immunity 2013
 1 Innate Immune Responses 3 Innate immunity Abul K. Abbas University of California San Francisco The initial responses to: 1. Microbes: essential early mechanisms to prevent, control, or eliminate infection;
1 Innate Immune Responses 3 Innate immunity Abul K. Abbas University of California San Francisco The initial responses to: 1. Microbes: essential early mechanisms to prevent, control, or eliminate infection;
SCIENCE CHINA Life Sciences. Supramolecular organizing centers (SMOCs) as signaling machines in innate immune activation
 SCIENCE CHINA Life Sciences Peking University Biological Science Education & Research (1925 2015) November 2015 Vol.58 No.11: 1067 1072 REVIEW doi: 10.1007/s11427-015-4951-z Supramolecular organizing centers
SCIENCE CHINA Life Sciences Peking University Biological Science Education & Research (1925 2015) November 2015 Vol.58 No.11: 1067 1072 REVIEW doi: 10.1007/s11427-015-4951-z Supramolecular organizing centers
The Innate Immune Response is Conserved Throughout Evolution and is Triggered by Pattern Recognition. Lipopolysaccharide = Lipid + Polysaccharide
 The Innate Immune Response is Conserved Throughout Evolution and is Triggered by Pattern Recognition Lipopolysaccharide = Lipid + Polysaccharide E.coli Cell wall organization Lipopolysaccharide Outer membrane
The Innate Immune Response is Conserved Throughout Evolution and is Triggered by Pattern Recognition Lipopolysaccharide = Lipid + Polysaccharide E.coli Cell wall organization Lipopolysaccharide Outer membrane
Toll-like Receptors (TLRs): Biology, Pathology and Therapeutics
 Toll-like Receptors (TLRs): Biology, Pathology and Therapeutics Dr Sarah Sasson SydPATH Registrar 23 rd June 2014 TLRs: Introduction Discovered in 1990s Recognise conserved structures in pathogens Rely
Toll-like Receptors (TLRs): Biology, Pathology and Therapeutics Dr Sarah Sasson SydPATH Registrar 23 rd June 2014 TLRs: Introduction Discovered in 1990s Recognise conserved structures in pathogens Rely
Innate Immunity. Chapter 3. Connection Between Innate and Adaptive Immunity. Know Differences and Provide Examples. Antimicrobial peptide psoriasin
 Chapter Know Differences and Provide Examples Innate Immunity kin and Epithelial Barriers Antimicrobial peptide psoriasin -Activity against Gram (-) E. coli Connection Between Innate and Adaptive Immunity
Chapter Know Differences and Provide Examples Innate Immunity kin and Epithelial Barriers Antimicrobial peptide psoriasin -Activity against Gram (-) E. coli Connection Between Innate and Adaptive Immunity
Role of Innate Immunity in Control of Adaptive Immunity
 Role of Innate Immunity in Control of Adaptive Immunity Innate Immunity The burden of pathogen sensing is placed on the innate immune system Danger hypothesis Missing Self Based on the detection of molecular
Role of Innate Immunity in Control of Adaptive Immunity Innate Immunity The burden of pathogen sensing is placed on the innate immune system Danger hypothesis Missing Self Based on the detection of molecular
Identification of Microbes
 Identification of Microbes Recognition by PRR (pattern recognition receptors) Recognize conserved molecular patterns on microbes called pathogen associated molecular patterns (PAMPs) which are not present
Identification of Microbes Recognition by PRR (pattern recognition receptors) Recognize conserved molecular patterns on microbes called pathogen associated molecular patterns (PAMPs) which are not present
Journal club. Lama Nazzal
 Journal club Lama Nazzal Background Kidney stone disease affects about 12% of men and 5% of women during their lifetimes in the United States Intrarenal nephrocalcinosis is often asymptomatic, but can
Journal club Lama Nazzal Background Kidney stone disease affects about 12% of men and 5% of women during their lifetimes in the United States Intrarenal nephrocalcinosis is often asymptomatic, but can
Chapter 3 The Induced Responses of Innate Immunity
 Chapter 3 The Induced Responses of Innate Immunity Pattern recognition by cells of the innate immune system Pattern recognition by cells of the innate immune system 4 main pattern recognition receptors
Chapter 3 The Induced Responses of Innate Immunity Pattern recognition by cells of the innate immune system Pattern recognition by cells of the innate immune system 4 main pattern recognition receptors
1. TLR. TLR Toll-like receptors. Toll Toll-like receptor, TLR TLR TLR TLR. type I TLR TLR. Toll
 54pp.145 152 2004 1. TLR T B TLR Toll-like receptors TLR TLR I IFN TLR T B B T Toll NF- B 1996 565-0871 3-1 TEL 06-6879-8303 FAX 06-6879-8305 E-mail uemattsu@biken.osaka-u.ac.jp Toll Toll-like receptor,
54pp.145 152 2004 1. TLR T B TLR Toll-like receptors TLR TLR I IFN TLR T B B T Toll NF- B 1996 565-0871 3-1 TEL 06-6879-8303 FAX 06-6879-8305 E-mail uemattsu@biken.osaka-u.ac.jp Toll Toll-like receptor,
Innate Immunity: (I) Molecules & (II) Cells
 Innate Immunity: (I) Molecules & (II) Cells Stephanie Eisenbarth, M.D., Ph.D. FOCIS Advanced Course 2/19/18 Department of Laboratory Medicine Yale School of Medicine Department of Immunobiology Yale School
Innate Immunity: (I) Molecules & (II) Cells Stephanie Eisenbarth, M.D., Ph.D. FOCIS Advanced Course 2/19/18 Department of Laboratory Medicine Yale School of Medicine Department of Immunobiology Yale School
TOLL-LIKE RECEPTORS AND CYTOKINES IN SEPSIS
 TOLL-LIKE RECEPTORS AND CYTOKINES IN SEPSIS A/PROF WILLIAM SEWELL ST VINCENT S CLINICAL SCHOOL, UNSW SYDPATH, ST VINCENT S HOSPITAL SYDNEY GARVAN INSTITUTE INNATE VERSUS ADAPTIVE IMMUNE RESPONSES INNATE
TOLL-LIKE RECEPTORS AND CYTOKINES IN SEPSIS A/PROF WILLIAM SEWELL ST VINCENT S CLINICAL SCHOOL, UNSW SYDPATH, ST VINCENT S HOSPITAL SYDNEY GARVAN INSTITUTE INNATE VERSUS ADAPTIVE IMMUNE RESPONSES INNATE
Gout and Nucleic Acid Metabolism Vol.33 No
 Gout and Nucleic Acid Metabolism Vol.33 No.1 2009 1 1 2 3 in vitro 14 IgM 1 IgM IgM 1 PAMPs Pattern recognition receptors PRRs PRRs PRRs PAMPs Toll Toll-like receptor TLR PAMPs Nod Nod-like receptor NLR
Gout and Nucleic Acid Metabolism Vol.33 No.1 2009 1 1 2 3 in vitro 14 IgM 1 IgM IgM 1 PAMPs Pattern recognition receptors PRRs PRRs PRRs PAMPs Toll Toll-like receptor TLR PAMPs Nod Nod-like receptor NLR
Innate Immunity. Connection Between Innate and Adaptive Immunity. Know Differences and Provide Examples Chapter 3. Antimicrobial peptide psoriasin
 Know Differences and Provide Examples Chapter * Innate Immunity * kin and Epithelial Barriers * Antimicrobial peptide psoriasin -Activity against Gram (-) E. coli Connection Between Innate and Adaptive
Know Differences and Provide Examples Chapter * Innate Immunity * kin and Epithelial Barriers * Antimicrobial peptide psoriasin -Activity against Gram (-) E. coli Connection Between Innate and Adaptive
Innate Immunity. Hathairat Thananchai, DPhil Department of Microbiology Faculty of Medicine Chiang Mai University 2 August 2016
 Innate Immunity Hathairat Thananchai, DPhil Department of Microbiology Faculty of Medicine Chiang Mai University 2 August 2016 Objectives: Explain how innate immune system recognizes foreign substances
Innate Immunity Hathairat Thananchai, DPhil Department of Microbiology Faculty of Medicine Chiang Mai University 2 August 2016 Objectives: Explain how innate immune system recognizes foreign substances
Lecture on Innate Immunity and Inflammation
 Lecture on Innate Immunity and Inflammation Evolutionary View Epithelial barriers to infection Four main types of innate recognition molecules:tlrs, CLRs, NLRs, RLRs NF-κB, the master transcriptional regulator
Lecture on Innate Immunity and Inflammation Evolutionary View Epithelial barriers to infection Four main types of innate recognition molecules:tlrs, CLRs, NLRs, RLRs NF-κB, the master transcriptional regulator
Signal Transduction: G-Protein Coupled Receptors
 Signal Transduction: G-Protein Coupled Receptors Federle, M. (2017). Lectures 4-5: Signal Transduction parts 1&2: nuclear receptors and GPCRs. Lecture presented at PHAR 423 Lecture in UIC College of Pharmacy,
Signal Transduction: G-Protein Coupled Receptors Federle, M. (2017). Lectures 4-5: Signal Transduction parts 1&2: nuclear receptors and GPCRs. Lecture presented at PHAR 423 Lecture in UIC College of Pharmacy,
Innate Immunity. Hathairat Thananchai, DPhil Department of Microbiology Faculty of Medicine Chiang Mai University 25 July 2017
 Innate Immunity Hathairat Thananchai, DPhil Department of Microbiology Faculty of Medicine Chiang Mai University 25 July 2017 Objectives: Explain how innate immune system recognizes foreign substances
Innate Immunity Hathairat Thananchai, DPhil Department of Microbiology Faculty of Medicine Chiang Mai University 25 July 2017 Objectives: Explain how innate immune system recognizes foreign substances
The Innate Immune Response is Conserved Throughout Evolution and is Triggered by Pattern Recognition. Lipopolysaccharide = Lipid + Polysaccharide
 The Innate Immune Response is onserved Throughout Evolution and is Triggered by Pattern Recognition Lipopolysaccharide = Lipid + Polysaccharide E.coli ell wall organization Lipopolysaccharide Outer membrane
The Innate Immune Response is onserved Throughout Evolution and is Triggered by Pattern Recognition Lipopolysaccharide = Lipid + Polysaccharide E.coli ell wall organization Lipopolysaccharide Outer membrane
Cell Signaling part 2
 15 Cell Signaling part 2 Functions of Cell Surface Receptors Other cell surface receptors are directly linked to intracellular enzymes. The largest family of these is the receptor protein tyrosine kinases,
15 Cell Signaling part 2 Functions of Cell Surface Receptors Other cell surface receptors are directly linked to intracellular enzymes. The largest family of these is the receptor protein tyrosine kinases,
SUPPORTING INFORMATION. The Architecture of the TIR Domain Signalosome in the Toll-like Receptor-4. Signaling Pathway
 SUPPORTING INFORMATION The Architecture of the TIR Domain Signalosome in the Toll-like Receptor-4 Signaling Pathway Emine Guven-Maiorov 1, Ozlem Keskin 1, Attila Gursoy 2*, Carter VanWaes 3, Zhong Chen
SUPPORTING INFORMATION The Architecture of the TIR Domain Signalosome in the Toll-like Receptor-4 Signaling Pathway Emine Guven-Maiorov 1, Ozlem Keskin 1, Attila Gursoy 2*, Carter VanWaes 3, Zhong Chen
Lecture on Innate Immunity and Inflammation. Innate Immunity: An Evolutionary View
 Lecture on Innate Immunity and Inflammation Evolutionary View Epithelial barriers to infection Four main types of innate recognition molecules:tlrs, CLRs, NLRs, RLRs NF-κB, the master transcriptional regulator
Lecture on Innate Immunity and Inflammation Evolutionary View Epithelial barriers to infection Four main types of innate recognition molecules:tlrs, CLRs, NLRs, RLRs NF-κB, the master transcriptional regulator
Immunology Part II. Innate Immunity. 18. April 2018, Ruhr-Universität Bochum Marcus Peters,
 Immunology Part II Innate Immunity 18. April 2018, Ruhr-Universität Bochum Marcus Peters, marcus.peters@rub.de Conserved structures of pathogens PAMPs are detected by Pattern Recognition Receptors PRRs
Immunology Part II Innate Immunity 18. April 2018, Ruhr-Universität Bochum Marcus Peters, marcus.peters@rub.de Conserved structures of pathogens PAMPs are detected by Pattern Recognition Receptors PRRs
The death receptors: signaling and modulation
 The death receptors: signaling and modulation 1 1 The extrinsic cell death pathway 2 Nat Rev Drug Discov. 2008 Dec;7(12):1001-12. 2 Death receptors Belong to the tumor necrosis factor (TNF) receptor gene
The death receptors: signaling and modulation 1 1 The extrinsic cell death pathway 2 Nat Rev Drug Discov. 2008 Dec;7(12):1001-12. 2 Death receptors Belong to the tumor necrosis factor (TNF) receptor gene
Alcoholic hepatitis is a drug-induced disorder
 Alcoholic hepatitis is a drug-induced disorder Gyongyi Szabo, MD, PhD Professor of Medicine University of Massachusetts Medical School Source: 2 Sobernation.com Clinical Progression of ALD Mortality Acute
Alcoholic hepatitis is a drug-induced disorder Gyongyi Szabo, MD, PhD Professor of Medicine University of Massachusetts Medical School Source: 2 Sobernation.com Clinical Progression of ALD Mortality Acute
Receptor mediated Signal Transduction
 Receptor mediated Signal Transduction G-protein-linked receptors adenylyl cyclase camp PKA Organization of receptor protein-tyrosine kinases From G.M. Cooper, The Cell. A molecular approach, 2004, third
Receptor mediated Signal Transduction G-protein-linked receptors adenylyl cyclase camp PKA Organization of receptor protein-tyrosine kinases From G.M. Cooper, The Cell. A molecular approach, 2004, third
Scott Abrams, Ph.D. Professor of Oncology, x4375 Kuby Immunology SEVENTH EDITION
 Scott Abrams, Ph.D. Professor of Oncology, x4375 scott.abrams@roswellpark.org Kuby Immunology SEVENTH EDITION CHAPTER 11 T-Cell Activation, Differentiation, and Memory Copyright 2013 by W. H. Freeman and
Scott Abrams, Ph.D. Professor of Oncology, x4375 scott.abrams@roswellpark.org Kuby Immunology SEVENTH EDITION CHAPTER 11 T-Cell Activation, Differentiation, and Memory Copyright 2013 by W. H. Freeman and
Lipopolysaccharide = Lipid + Polysaccharide. The Innate Immune Response is Conserved Throughout Evolution and is Triggered by Pattern Recognition
 Lipopolysaccharide = Lipid + Polysaccharide The Innate Immune Response is onserved Throughout volution and is Triggered by Pattern Recognition.coli ell wall organization Lipid A Lipopolysaccharide Outer
Lipopolysaccharide = Lipid + Polysaccharide The Innate Immune Response is onserved Throughout volution and is Triggered by Pattern Recognition.coli ell wall organization Lipid A Lipopolysaccharide Outer
CYTOKINE RECEPTORS AND SIGNAL TRANSDUCTION
 CYTOKINE RECEPTORS AND SIGNAL TRANSDUCTION What is Cytokine? Secreted popypeptide (protein) involved in cell-to-cell signaling. Acts in paracrine or autocrine fashion through specific cellular receptors.
CYTOKINE RECEPTORS AND SIGNAL TRANSDUCTION What is Cytokine? Secreted popypeptide (protein) involved in cell-to-cell signaling. Acts in paracrine or autocrine fashion through specific cellular receptors.
Structure and Function of Antigen Recognition Molecules
 MICR2209 Structure and Function of Antigen Recognition Molecules Dr Allison Imrie allison.imrie@uwa.edu.au 1 Synopsis: In this lecture we will examine the major receptors used by cells of the innate and
MICR2209 Structure and Function of Antigen Recognition Molecules Dr Allison Imrie allison.imrie@uwa.edu.au 1 Synopsis: In this lecture we will examine the major receptors used by cells of the innate and
Chapter 9. Cellular Signaling
 Chapter 9 Cellular Signaling Cellular Messaging Page 215 Cells can signal to each other and interpret the signals they receive from other cells and the environment Signals are most often chemicals The
Chapter 9 Cellular Signaling Cellular Messaging Page 215 Cells can signal to each other and interpret the signals they receive from other cells and the environment Signals are most often chemicals The
KEY CONCEPT QUESTIONS IN SIGNAL TRANSDUCTION
 Signal Transduction - Part 2 Key Concepts - Receptor tyrosine kinases control cell metabolism and proliferation Growth factor signaling through Ras Mutated cell signaling genes in cancer cells are called
Signal Transduction - Part 2 Key Concepts - Receptor tyrosine kinases control cell metabolism and proliferation Growth factor signaling through Ras Mutated cell signaling genes in cancer cells are called
Signaling Through Immune System Receptors (Ch. 7)
 Signaling Through Immune System Receptors (Ch. 7) 1. General principles of signal transduction and propagation. 2. Antigen receptor signaling and lymphocyte activation. 3. Other receptors and signaling
Signaling Through Immune System Receptors (Ch. 7) 1. General principles of signal transduction and propagation. 2. Antigen receptor signaling and lymphocyte activation. 3. Other receptors and signaling
Biol403 MAP kinase signalling
 Biol403 MAP kinase signalling The mitogen activated protein kinase (MAPK) pathway is a signalling cascade activated by a diverse range of effectors. The cascade regulates many cellular activities including
Biol403 MAP kinase signalling The mitogen activated protein kinase (MAPK) pathway is a signalling cascade activated by a diverse range of effectors. The cascade regulates many cellular activities including
INFLAMMASOME IN INTESTINAL INFLAMMATION AND CANCER. Laura Stronati ENEA - Roma
 INFLAMMASOME IN INTESTINAL INFLAMMATION AND CANCER Laura Stronati ENEA - Roma Inflammasome: definition, components and activation TLRs NODs RLRs CLRs PRRs LPS, Flagellin, DNA, RNA, etc) PAMPs DAMPs ATP,
INFLAMMASOME IN INTESTINAL INFLAMMATION AND CANCER Laura Stronati ENEA - Roma Inflammasome: definition, components and activation TLRs NODs RLRs CLRs PRRs LPS, Flagellin, DNA, RNA, etc) PAMPs DAMPs ATP,
Lipopolysaccharide = Lipid + Polysaccharide. The Innate Immune Response is Conserved Throughout Evolution and is Triggered by Pattern Recognition
 Lipopolysaccharide = Lipid + Polysaccharide The Innate Immune Response is onserved Throughout Evolution and is Triggered by Pattern Recognition E.coli ell wall organization Lipopolysaccharide Outer membrane
Lipopolysaccharide = Lipid + Polysaccharide The Innate Immune Response is onserved Throughout Evolution and is Triggered by Pattern Recognition E.coli ell wall organization Lipopolysaccharide Outer membrane
Integrated Innate Immunity Combining Activation and Effector Functions
 Section 4 Integrated Innate Immunity Combining Activation and Effector Functions 4.1 THE DETECTION OF BACTERIAL LIPOPOLYSACCHARIDE LPS is one of the most potent immune stimuli known. It is found in the
Section 4 Integrated Innate Immunity Combining Activation and Effector Functions 4.1 THE DETECTION OF BACTERIAL LIPOPOLYSACCHARIDE LPS is one of the most potent immune stimuli known. It is found in the
Identification of an Antiviral Signaling Variant Demonstrates Immune Regulation Through Alternative Translation
 Identification of an Antiviral Signaling Variant Demonstrates Immune Regulation Through Alternative Translation The Harvard community has made this article openly available. Please share how this access
Identification of an Antiviral Signaling Variant Demonstrates Immune Regulation Through Alternative Translation The Harvard community has made this article openly available. Please share how this access
3/10/14. Ultrastructural organization. Gram Stain. Infection leads to production of inducers of inflammation. Gram negative.
 Infection leads to production of inducers of inflammation or dendritic cell Inflammatory mediators: Complex and many, but include: Lipids and Proteins (cytokines/chemokines) TNF Others Ultrastructural
Infection leads to production of inducers of inflammation or dendritic cell Inflammatory mediators: Complex and many, but include: Lipids and Proteins (cytokines/chemokines) TNF Others Ultrastructural
Cell Biology Lecture 9 Notes Basic Principles of cell signaling and GPCR system
 Cell Biology Lecture 9 Notes Basic Principles of cell signaling and GPCR system Basic Elements of cell signaling: Signal or signaling molecule (ligand, first messenger) o Small molecules (epinephrine,
Cell Biology Lecture 9 Notes Basic Principles of cell signaling and GPCR system Basic Elements of cell signaling: Signal or signaling molecule (ligand, first messenger) o Small molecules (epinephrine,
Pathogen Recognition and Inflammatory Signaling in Innate Immune Defenses
 CLINICAL MICROBIOLOGY REVIEWS, Apr. 2009, p. 240 273 Vol. 22, No. 2 0893-8512/09/$08.00 0 doi:10.1128/cmr.00046-08 Copyright 2009, American Society for Microbiology. All Rights Reserved. Pathogen Recognition
CLINICAL MICROBIOLOGY REVIEWS, Apr. 2009, p. 240 273 Vol. 22, No. 2 0893-8512/09/$08.00 0 doi:10.1128/cmr.00046-08 Copyright 2009, American Society for Microbiology. All Rights Reserved. Pathogen Recognition
Enzyme-coupled Receptors. Cell-surface receptors 1. Ion-channel-coupled receptors 2. G-protein-coupled receptors 3. Enzyme-coupled receptors
 Enzyme-coupled Receptors Cell-surface receptors 1. Ion-channel-coupled receptors 2. G-protein-coupled receptors 3. Enzyme-coupled receptors Cell-surface receptors allow a flow of ions across the plasma
Enzyme-coupled Receptors Cell-surface receptors 1. Ion-channel-coupled receptors 2. G-protein-coupled receptors 3. Enzyme-coupled receptors Cell-surface receptors allow a flow of ions across the plasma
Cell Quality Control. Peter Takizawa Department of Cell Biology
 Cell Quality Control Peter Takizawa Department of Cell Biology Cellular quality control reduces production of defective proteins. Cells have many quality control systems to ensure that cell does not build
Cell Quality Control Peter Takizawa Department of Cell Biology Cellular quality control reduces production of defective proteins. Cells have many quality control systems to ensure that cell does not build
Inflammation: How to Cool the Fire Inside your Gut? REINVENTING DIAGNOSTICS
 Inflammation: How to Cool the Fire Inside your Gut? REINVENTING DIAGNOSTICS Future of Healthcare REINVENTING DIAGNOSTICS Inflammation Gut Inflammation Basis of a Healthy
Inflammation: How to Cool the Fire Inside your Gut? REINVENTING DIAGNOSTICS Future of Healthcare REINVENTING DIAGNOSTICS Inflammation Gut Inflammation Basis of a Healthy
Under the Radar Screen: How Bugs Trick Our Immune Defenses
 Under the Radar Screen: How Bugs Trick Our Immune Defenses Session 3: Toll-like receptors (TLRs) Marie-Eve Paquet and Gijsbert Grotenbreg Whitehead Institute for Biomedical Research Introduction to Toll-like
Under the Radar Screen: How Bugs Trick Our Immune Defenses Session 3: Toll-like receptors (TLRs) Marie-Eve Paquet and Gijsbert Grotenbreg Whitehead Institute for Biomedical Research Introduction to Toll-like
Insulin Resistance. Biol 405 Molecular Medicine
 Insulin Resistance Biol 405 Molecular Medicine Insulin resistance: a subnormal biological response to insulin. Defects of either insulin secretion or insulin action can cause diabetes mellitus. Insulin-dependent
Insulin Resistance Biol 405 Molecular Medicine Insulin resistance: a subnormal biological response to insulin. Defects of either insulin secretion or insulin action can cause diabetes mellitus. Insulin-dependent
Lipids and Membranes
 Lipids and Membranes Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry I Philadelphia University Faculty of pharmacy Membrane transport D. Endocytosis and Exocytosis
Lipids and Membranes Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry I Philadelphia University Faculty of pharmacy Membrane transport D. Endocytosis and Exocytosis
Innate immune regulation of T-helper (Th) cell homeostasis in the intestine
 Innate immune regulation of T-helper (Th) cell homeostasis in the intestine Masayuki Fukata, MD, Ph.D. Research Scientist II Division of Gastroenterology, Department of Medicine, F. Widjaja Foundation,
Innate immune regulation of T-helper (Th) cell homeostasis in the intestine Masayuki Fukata, MD, Ph.D. Research Scientist II Division of Gastroenterology, Department of Medicine, F. Widjaja Foundation,
CELL BIOLOGY - CLUTCH CH THE IMMUNE SYSTEM.
 !! www.clutchprep.com CONCEPT: OVERVIEW OF HOST DEFENSES The human body contains three lines of against infectious agents (pathogens) 1. Mechanical and chemical boundaries (part of the innate immune system)
!! www.clutchprep.com CONCEPT: OVERVIEW OF HOST DEFENSES The human body contains three lines of against infectious agents (pathogens) 1. Mechanical and chemical boundaries (part of the innate immune system)
The T cell receptor for MHC-associated peptide antigens
 1 The T cell receptor for MHC-associated peptide antigens T lymphocytes have a dual specificity: they recognize polymporphic residues of self MHC molecules, and they also recognize residues of peptide
1 The T cell receptor for MHC-associated peptide antigens T lymphocytes have a dual specificity: they recognize polymporphic residues of self MHC molecules, and they also recognize residues of peptide
Allergy and Immunology Review Corner: Chapter 13 of Immunology IV: Clinical Applications in Health and Disease, by Joseph A. Bellanti, MD.
 Allergy and Immunology Review Corner: Chapter 13 of Immunology IV: Clinical Applications in Health and Disease, by Joseph A. Bellanti, MD. Chapter 13: Mechanisms of Immunity to Viral Disease Prepared by
Allergy and Immunology Review Corner: Chapter 13 of Immunology IV: Clinical Applications in Health and Disease, by Joseph A. Bellanti, MD. Chapter 13: Mechanisms of Immunity to Viral Disease Prepared by
G-Protein Signaling. Introduction to intracellular signaling. Dr. SARRAY Sameh, Ph.D
 G-Protein Signaling Introduction to intracellular signaling Dr. SARRAY Sameh, Ph.D Cell signaling Cells communicate via extracellular signaling molecules (Hormones, growth factors and neurotransmitters
G-Protein Signaling Introduction to intracellular signaling Dr. SARRAY Sameh, Ph.D Cell signaling Cells communicate via extracellular signaling molecules (Hormones, growth factors and neurotransmitters
Test Bank for The Immune System 4th Edition by Parham
 Test Bank for The Immune System 4th Edition by Parham CHAPTER 3: INNATE IMMUNITY: THE INDUCED RESPONSE TO INFECTION 3 1 C-type lectins are so called because of the role of in facilitating receptor:ligand
Test Bank for The Immune System 4th Edition by Parham CHAPTER 3: INNATE IMMUNITY: THE INDUCED RESPONSE TO INFECTION 3 1 C-type lectins are so called because of the role of in facilitating receptor:ligand
King s Research Portal
 King s Research Portal DOI: 10.1042/BCJ20170714 Document Version Peer reviewed version Link to publication record in King's Research Portal Citation for published version (APA): Otsu, K., & Nakayama, H.
King s Research Portal DOI: 10.1042/BCJ20170714 Document Version Peer reviewed version Link to publication record in King's Research Portal Citation for published version (APA): Otsu, K., & Nakayama, H.
TD-BF01: Innate immunity to microorganisms
 TD-BF01: Innate immunity to microorganisms I. Toll receptors (adapted from Takeuchi, O. et al. (1999) Immunity 11:443; Kawai, T. et al. (1999) Immunity 11:115; Hemmi, H. et al. (2000) Nature 408:740; Muzio,
TD-BF01: Innate immunity to microorganisms I. Toll receptors (adapted from Takeuchi, O. et al. (1999) Immunity 11:443; Kawai, T. et al. (1999) Immunity 11:115; Hemmi, H. et al. (2000) Nature 408:740; Muzio,
JPEMS Nantes, Basic Immunology INNATE IMMUNITY
 JPEMS Nantes, 2014- Basic Immunology INNATE IMMUNITY Teacher: Pr. Régis Josien, Laboratoire d Immunologie and INSERM U1064, CHU Nantes Regis.Josien@univ-nantes.fr 1 Contents 1. General features and specificity
JPEMS Nantes, 2014- Basic Immunology INNATE IMMUNITY Teacher: Pr. Régis Josien, Laboratoire d Immunologie and INSERM U1064, CHU Nantes Regis.Josien@univ-nantes.fr 1 Contents 1. General features and specificity
General information. Cell mediated immunity. 455 LSA, Tuesday 11 to noon. Anytime after class.
 General information Cell mediated immunity 455 LSA, Tuesday 11 to noon Anytime after class T-cell precursors Thymus Naive T-cells (CD8 or CD4) email: lcoscoy@berkeley.edu edu Use MCB150 as subject line
General information Cell mediated immunity 455 LSA, Tuesday 11 to noon Anytime after class T-cell precursors Thymus Naive T-cells (CD8 or CD4) email: lcoscoy@berkeley.edu edu Use MCB150 as subject line
TRAF molecules in cell signaling and in human diseases
 Xie Journal of Molecular Signaling 2013, 8:7 REVIEW Open Access TRAF molecules in cell signaling and in human diseases Ping Xie Abstract The tumor necrosis factor receptor (TNF-R)-associated factor (TRAF)
Xie Journal of Molecular Signaling 2013, 8:7 REVIEW Open Access TRAF molecules in cell signaling and in human diseases Ping Xie Abstract The tumor necrosis factor receptor (TNF-R)-associated factor (TRAF)
Summary and Discussion antigen presentation
 Summary and Discussion antigen presentation 247 248 Summary & Discussion Summary and discussion: antigen presentation For a cell to communicate information about its internal health and status to the immune
Summary and Discussion antigen presentation 247 248 Summary & Discussion Summary and discussion: antigen presentation For a cell to communicate information about its internal health and status to the immune
Signal Transduction Cascades
 Signal Transduction Cascades Contents of this page: Kinases & phosphatases Protein Kinase A (camp-dependent protein kinase) G-protein signal cascade Structure of G-proteins Small GTP-binding proteins,
Signal Transduction Cascades Contents of this page: Kinases & phosphatases Protein Kinase A (camp-dependent protein kinase) G-protein signal cascade Structure of G-proteins Small GTP-binding proteins,
Interferon Induction by RNA Viruses and Antagonism by Viral Pathogens
 Viruses 2014, 6, 4999-5027; doi:10.3390/v6124999 Review OPEN ACCESS viruses ISSN 1999-4915 www.mdpi.com/journal/viruses Interferon Induction by RNA Viruses and Antagonism by Viral Pathogens Yuchen Nan
Viruses 2014, 6, 4999-5027; doi:10.3390/v6124999 Review OPEN ACCESS viruses ISSN 1999-4915 www.mdpi.com/journal/viruses Interferon Induction by RNA Viruses and Antagonism by Viral Pathogens Yuchen Nan
A particular set of insults induces apoptosis (part 1), which, if inhibited, can switch to autophagy. At least in some cellular settings, autophagy se
 A particular set of insults induces apoptosis (part 1), which, if inhibited, can switch to autophagy. At least in some cellular settings, autophagy serves as a defence mechanism that prevents or retards
A particular set of insults induces apoptosis (part 1), which, if inhibited, can switch to autophagy. At least in some cellular settings, autophagy serves as a defence mechanism that prevents or retards
October 26, Lecture Readings. Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell
 October 26, 2006 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
October 26, 2006 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
Supplementary information for: Community detection for networks with. unipartite and bipartite structure. Chang Chang 1, 2, Chao Tang 2
 Supplementary information for: Community detection for networks with unipartite and bipartite structure Chang Chang 1, 2, Chao Tang 2 1 School of Life Sciences, Peking University, Beiing 100871, China
Supplementary information for: Community detection for networks with unipartite and bipartite structure Chang Chang 1, 2, Chao Tang 2 1 School of Life Sciences, Peking University, Beiing 100871, China
The Regulatory Role of Interleukin-1 Receptor Associated Kinase M in Toll-Like Receptor Signaling
 Cleveland State University EngagedScholarship@CSU ETD Archive 2016 The Regulatory Role of Interleukin-1 Receptor Associated Kinase M in Toll-Like Receptor Signaling Hao Zhou Cleveland State University
Cleveland State University EngagedScholarship@CSU ETD Archive 2016 The Regulatory Role of Interleukin-1 Receptor Associated Kinase M in Toll-Like Receptor Signaling Hao Zhou Cleveland State University
SUPPLEMENTARY INFORMATION. Divergent TLR7/9 signaling and type I interferon production distinguish
 SUPPLEMENTARY INFOATION Divergent TLR7/9 signaling and type I interferon production distinguish pathogenic and non-pathogenic AIDS-virus infections Judith N. Mandl, Ashley P. Barry, Thomas H. Vanderford,
SUPPLEMENTARY INFOATION Divergent TLR7/9 signaling and type I interferon production distinguish pathogenic and non-pathogenic AIDS-virus infections Judith N. Mandl, Ashley P. Barry, Thomas H. Vanderford,
Intracellular MHC class II molecules promote TLR-triggered innate. immune responses by maintaining Btk activation
 Intracellular MHC class II molecules promote TLR-triggered innate immune responses by maintaining Btk activation Xingguang Liu, Zhenzhen Zhan, Dong Li, Li Xu, Feng Ma, Peng Zhang, Hangping Yao and Xuetao
Intracellular MHC class II molecules promote TLR-triggered innate immune responses by maintaining Btk activation Xingguang Liu, Zhenzhen Zhan, Dong Li, Li Xu, Feng Ma, Peng Zhang, Hangping Yao and Xuetao
TCR, MHC and coreceptors
 Cooperation In Immune Responses Antigen processing how peptides get into MHC Antigen processing involves the intracellular proteolytic generation of MHC binding proteins Protein antigens may be processed
Cooperation In Immune Responses Antigen processing how peptides get into MHC Antigen processing involves the intracellular proteolytic generation of MHC binding proteins Protein antigens may be processed
Bruce A. Beutler and Jules A. Hoffmann. Ralph M. Steinman
 PRESS RELEASE 20-0-03 The Nobel Assembly at Karolinska Institutet has today decided that The 20 Nobel Prize in Physiology or Medicine shall be divided, with one half jointly to Bruce A. Beutler and Jules
PRESS RELEASE 20-0-03 The Nobel Assembly at Karolinska Institutet has today decided that The 20 Nobel Prize in Physiology or Medicine shall be divided, with one half jointly to Bruce A. Beutler and Jules
Page 32 AP Biology: 2013 Exam Review CONCEPT 6 REGULATION
 Page 32 AP Biology: 2013 Exam Review CONCEPT 6 REGULATION 1. Feedback a. Negative feedback mechanisms maintain dynamic homeostasis for a particular condition (variable) by regulating physiological processes,
Page 32 AP Biology: 2013 Exam Review CONCEPT 6 REGULATION 1. Feedback a. Negative feedback mechanisms maintain dynamic homeostasis for a particular condition (variable) by regulating physiological processes,
Inflammation - Molecular events
 Academic lectures for students of medical schools 3rd Year updated 2004-2015 GENERAL PATHOPHYSIOLOGY Inflammation - Molecular events R. A. Benacka, MD, PhD Department of Pathophysiology Medical faculty
Academic lectures for students of medical schools 3rd Year updated 2004-2015 GENERAL PATHOPHYSIOLOGY Inflammation - Molecular events R. A. Benacka, MD, PhD Department of Pathophysiology Medical faculty
Signal Transduction: Information Metabolism. Chem 454: Regulatory Mechanisms in Biochemistry University of Wisconsin-Eau Claire
 Signal Transduction: Information Metabolism Chem 454: Regulatory Mechanisms in Biochemistry University of Wisconsin-Eau Claire Introduction Information Metabolism How cells receive, process and respond
Signal Transduction: Information Metabolism Chem 454: Regulatory Mechanisms in Biochemistry University of Wisconsin-Eau Claire Introduction Information Metabolism How cells receive, process and respond
Relative sizes of infectious agents
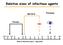 Relative sizes of infectious agents Bacteria Protozoa Viruses RBC 0.005 0.01 0.03 01 03 05 1 3 5 10 30 50 100 300 Size in microns ( µm ) - log scale Immunity to Infection Principle 1 Every clinical infection
Relative sizes of infectious agents Bacteria Protozoa Viruses RBC 0.005 0.01 0.03 01 03 05 1 3 5 10 30 50 100 300 Size in microns ( µm ) - log scale Immunity to Infection Principle 1 Every clinical infection
T-cell activation T cells migrate to secondary lymphoid tissues where they interact with antigen, antigen-presenting cells, and other lymphocytes:
 Interactions between innate immunity & adaptive immunity What happens to T cells after they leave the thymus? Naïve T cells exit the thymus and enter the bloodstream. If they remain in the bloodstream,
Interactions between innate immunity & adaptive immunity What happens to T cells after they leave the thymus? Naïve T cells exit the thymus and enter the bloodstream. If they remain in the bloodstream,
T-cell activation T cells migrate to secondary lymphoid tissues where they interact with antigen, antigen-presenting cells, and other lymphocytes:
 Interactions between innate immunity & adaptive immunity What happens to T cells after they leave the thymus? Naïve T cells exit the thymus and enter the bloodstream. If they remain in the bloodstream,
Interactions between innate immunity & adaptive immunity What happens to T cells after they leave the thymus? Naïve T cells exit the thymus and enter the bloodstream. If they remain in the bloodstream,
The role of pattern-recognition receptors in innate immunity: update on Toll-like receptors
 The role of pattern-recognition receptors in innate immunity: update on Toll-like receptors Taro Kawai 1,2 & Shizuo Akira 1,2 The discovery of Toll-like receptors (TLRs) as components that recognize conserved
The role of pattern-recognition receptors in innate immunity: update on Toll-like receptors Taro Kawai 1,2 & Shizuo Akira 1,2 The discovery of Toll-like receptors (TLRs) as components that recognize conserved
REGULATION OF ENZYME ACTIVITY. Medical Biochemistry, Lecture 25
 REGULATION OF ENZYME ACTIVITY Medical Biochemistry, Lecture 25 Lecture 25, Outline General properties of enzyme regulation Regulation of enzyme concentrations Allosteric enzymes and feedback inhibition
REGULATION OF ENZYME ACTIVITY Medical Biochemistry, Lecture 25 Lecture 25, Outline General properties of enzyme regulation Regulation of enzyme concentrations Allosteric enzymes and feedback inhibition
Pattern Recognition Receptors and the Innate Immune Response to Viral Infection
 Viruses 2011, 3, 920-940; doi:10.3390/v3060920 OPEN ACCESS viruses ISSN 1999-4915 www.mdpi.com/journal/viruses Review Pattern Recognition Receptors and the Innate Immune Response to Viral Infection Mikayla
Viruses 2011, 3, 920-940; doi:10.3390/v3060920 OPEN ACCESS viruses ISSN 1999-4915 www.mdpi.com/journal/viruses Review Pattern Recognition Receptors and the Innate Immune Response to Viral Infection Mikayla
Animal Models to Understand Immunity
 Animal Models to Understand Immunity Hussein El Saghire hesaghir@sckcen.be Innate Adaptive immunity Immunity MAPK and NF-kB TLR pathways receptors Fast Slow Non-specific Specific NOD-like receptors T-cell
Animal Models to Understand Immunity Hussein El Saghire hesaghir@sckcen.be Innate Adaptive immunity Immunity MAPK and NF-kB TLR pathways receptors Fast Slow Non-specific Specific NOD-like receptors T-cell
BIOLOGY. Cell Communication CAMPBELL. Reece Urry Cain Wasserman Minorsky Jackson. Lecture Presentation by Nicole Tunbridge and Kathleen Fitzpatrick
 CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson 11 Cell Communication Lecture Presentation by Nicole Tunbridge and Kathleen Fitzpatrick Cellular Messaging Cells can signal to
CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson 11 Cell Communication Lecture Presentation by Nicole Tunbridge and Kathleen Fitzpatrick Cellular Messaging Cells can signal to
MHC class I MHC class II Structure of MHC antigens:
 MHC class I MHC class II Structure of MHC antigens: MHC class I antigens consist of a transmembrane heavy chain (α chain) that is non-covalently associated with β2- microglobulin. Membrane proximal domain
MHC class I MHC class II Structure of MHC antigens: MHC class I antigens consist of a transmembrane heavy chain (α chain) that is non-covalently associated with β2- microglobulin. Membrane proximal domain
The roles of TLRs, RLRs and NLRs in pathogen recognition
 International Immunology, Vol. 21, No. 4, pp. 317 337 doi:10.1093/intimm/dxp017 ª The Japanese Society for Immunology. 2009. All rights reserved. For permissions, please e-mail: journals.permissions@oxfordjournals.org
International Immunology, Vol. 21, No. 4, pp. 317 337 doi:10.1093/intimm/dxp017 ª The Japanese Society for Immunology. 2009. All rights reserved. For permissions, please e-mail: journals.permissions@oxfordjournals.org
ACTIVATION OF T LYMPHOCYTES AND CELL MEDIATED IMMUNITY
 ACTIVATION OF T LYMPHOCYTES AND CELL MEDIATED IMMUNITY The recognition of specific antigen by naïve T cell induces its own activation and effector phases. T helper cells recognize peptide antigens through
ACTIVATION OF T LYMPHOCYTES AND CELL MEDIATED IMMUNITY The recognition of specific antigen by naïve T cell induces its own activation and effector phases. T helper cells recognize peptide antigens through
Cellular stress response and innate immune signaling: integrating pathways in host defense and inflammation
 Review Cellular stress response and innate immune signaling: integrating pathways in host defense and inflammation Sujatha Muralidharan and Pranoti Mandrekar 1 Department of Medicine, University of Massachusetts
Review Cellular stress response and innate immune signaling: integrating pathways in host defense and inflammation Sujatha Muralidharan and Pranoti Mandrekar 1 Department of Medicine, University of Massachusetts
Innate Immunity. Jan 8 th Prof. dr. sc. Ivana Novak Nakir 1
 Innate Immunity Jan 8 th 2018. Prof. dr. sc. Ivana Novak Nakir 1 Adaptive Innate 2 Immune system overview 1 st line of defense skin (2m 2 ) and mucosal membranes (~400m 2 ): physical barrier, lymphoid
Innate Immunity Jan 8 th 2018. Prof. dr. sc. Ivana Novak Nakir 1 Adaptive Innate 2 Immune system overview 1 st line of defense skin (2m 2 ) and mucosal membranes (~400m 2 ): physical barrier, lymphoid
3rd International Conference on Neurology & Therapeutics.
 3rd International Conference on Neurology & Therapeutics www.neuroimmunology.ca Multiple sclerosis is a devastating disease The first description of the disease was mentioned in 14th century In 1838 Dr.
3rd International Conference on Neurology & Therapeutics www.neuroimmunology.ca Multiple sclerosis is a devastating disease The first description of the disease was mentioned in 14th century In 1838 Dr.
Propagation of the Signal
 OpenStax-CNX module: m44452 1 Propagation of the Signal OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section,
OpenStax-CNX module: m44452 1 Propagation of the Signal OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section,
Scott Abrams, Ph.D. Professor of Oncology, x4375 Kuby Immunology SEVENTH EDITION
 Scott Abrams, Ph.D. Professor of Oncology, x4375 scott.abrams@roswellpark.org Kuby Immunology SEVENTH EDITION CHAPTER 13 Effector Responses: Cell- and Antibody-Mediated Immunity Copyright 2013 by W. H.
Scott Abrams, Ph.D. Professor of Oncology, x4375 scott.abrams@roswellpark.org Kuby Immunology SEVENTH EDITION CHAPTER 13 Effector Responses: Cell- and Antibody-Mediated Immunity Copyright 2013 by W. H.
Introduction to pathology lecture 5/ Cell injury apoptosis. Dr H Awad 2017/18
 Introduction to pathology lecture 5/ Cell injury apoptosis Dr H Awad 2017/18 Apoptosis = programmed cell death = cell suicide= individual cell death Apoptosis cell death induced by a tightly regulated
Introduction to pathology lecture 5/ Cell injury apoptosis Dr H Awad 2017/18 Apoptosis = programmed cell death = cell suicide= individual cell death Apoptosis cell death induced by a tightly regulated
Principles of Genetics and Molecular Biology
 Cell signaling Dr. Diala Abu-Hassan, DDS, PhD School of Medicine Dr.abuhassand@gmail.com Principles of Genetics and Molecular Biology www.cs.montana.edu Modes of cell signaling Direct interaction of a
Cell signaling Dr. Diala Abu-Hassan, DDS, PhD School of Medicine Dr.abuhassand@gmail.com Principles of Genetics and Molecular Biology www.cs.montana.edu Modes of cell signaling Direct interaction of a
Pattern recognition receptors and control of adaptive immunity
 Noah W. Palm Ruslan Medzhitov Pattern recognition receptors and control of adaptive immunity Authors address Noah W. Palm 1, Ruslan Medzhitov 1 1 Howard Hughes Medical Institute and Department of Immunobiology,
Noah W. Palm Ruslan Medzhitov Pattern recognition receptors and control of adaptive immunity Authors address Noah W. Palm 1, Ruslan Medzhitov 1 1 Howard Hughes Medical Institute and Department of Immunobiology,
Chapter 11. Cell Communication
 Chapter 11 Cell Communication Overview: The Cellular Internet Cell-to-cell communication Is absolutely essential for multicellular organisms Concept 11.1: External signals are converted into responses
Chapter 11 Cell Communication Overview: The Cellular Internet Cell-to-cell communication Is absolutely essential for multicellular organisms Concept 11.1: External signals are converted into responses
MAVS coordination of antiviral innate immunity. Christine Vazquez a and Stacy M. Horner a,b,* Duke University Medical Center Durham, NC 27710
 JVI Accepted Manuscript Posted Online 6 May 2015 J. Virol. doi:10.1128/jvi.01918-14 Copyright 2015, American Society for Microbiology. All Rights Reserved. 1 2 3 MAVS coordination of antiviral innate immunity
JVI Accepted Manuscript Posted Online 6 May 2015 J. Virol. doi:10.1128/jvi.01918-14 Copyright 2015, American Society for Microbiology. All Rights Reserved. 1 2 3 MAVS coordination of antiviral innate immunity
Lecture 15. Signal Transduction Pathways - Introduction
 Lecture 15 Signal Transduction Pathways - Introduction So far.. Regulation of mrna synthesis Regulation of rrna synthesis Regulation of trna & 5S rrna synthesis Regulation of gene expression by signals
Lecture 15 Signal Transduction Pathways - Introduction So far.. Regulation of mrna synthesis Regulation of rrna synthesis Regulation of trna & 5S rrna synthesis Regulation of gene expression by signals
Signaling. Dr. Sujata Persad Katz Group Centre for Pharmacy & Health research
 Signaling Dr. Sujata Persad 3-020 Katz Group Centre for Pharmacy & Health research E-mail:sujata.persad@ualberta.ca 1 Growth Factor Receptors and Other Signaling Pathways What we will cover today: How
Signaling Dr. Sujata Persad 3-020 Katz Group Centre for Pharmacy & Health research E-mail:sujata.persad@ualberta.ca 1 Growth Factor Receptors and Other Signaling Pathways What we will cover today: How
