Objective. Background
|
|
|
- Jemimah Kristina Sims
- 6 years ago
- Views:
Transcription
1 Section E. Host Factor Data Objective Upon completion of this exercise, you will be able to use the Influenza Research Database (IRD; and Virus Pathogen Resource (ViPR; to: Search for host factor experiments using the faceted search function Investigate detailed metadata describing individual experiments Explore pre-computed host response patterns within a given experiment Compare biosets using Boolean Operations Save searches to the Workbench Save results to the Workbench Download data and use it in downstream analyses Perform host factor GO and pathway enrichment analysis Background From its inception in March 2012, the Host Factor data component has been shared between the Influenza Research Database (IRD) and Virus Pathogen Resource (ViPR) in order to support analysis of host responses to human pathogen infection. Currently available host factor data is derived from multiple experiments involving influenza A H1N1 pandemic or seasonal virus, avian influenza A H5N1 virus, SARS-CoV, and MERS-CoV infections in human cell lines and in experimental animals. The data and associated metadata captured in IRD/ViPR fall into three levels: Study: study metadata includes description of the study, PI, point of contact, publications, etc. Experiment: each study is comprised of one or more experiments. Experiment associated metadata includes experiment name, type, measurement technique, experimenters, conditional variables, host species, protocols, etc. Bioset: each bioset contains a list of host factors typically generated by comparing host responses in virus-infected samples with mock-infected samples under different conditions (strain, time point, dose, etc.). 43
2 Workflow Search for Host Factor Experiments: - Use faceted search to find experiments of interest - Explore information about individual experiments Explore Host Response Patterns: - Explore pre-compiled host response patterns on the Experiment page - Search patterns for host factors of interest - Conduct GO enrichment analysis on a pattern - Build working sets from patterns on the Experiment page Use Workbench to Manage Data and Conduct Analyses: - Save host factor data to the Workbench - Perform Boolean operations on host factor working sets Visualize and Conduct Enrichment Analyses: - Output data for downstream visualization and analysis - Conduct GO and pathway enrichment analysis in an external resource 44
3 I. Search for Host Factor Experiments and Compare Biosets 1. Search for host factor experiments using faceted search terms a. To access the host factor experiments, mouse-over the Search Data tab and click the Host Factor Experiments option from the pull down menu. b. The Host Factor Experiment landing page is organized into two panels: a faceted navigation menu on the left, and a table of filtered host factor experiments on the right. The facets allow for quick and efficient data filtering using various structured metadata. c. Now search for host factor experiments using one or more facets. Select the checkbox(es) next to the facet(s) of your interest. In this exercise, we are going to select Orthomyxoviridae under the Family header. This will trigger the table to dynamically update and display the experiments available for this family. Next, select a specific Viral Agent VN1203 (H5N1). The table on the right will now display all experiments that use this strain. 2. Access detailed metadata describing individual experiments a. In the returned table, click ICL004-R in the Experiment ID column. b. You will be taken to the Host Factor Experiment [IRD_SV_ICL004-R] page. Here you can view the Experiment Information, Experiment Sample Summary, Host Factor Bioset Information, Host Factor Bioset Summary, Host Factor Bioset Patterns, and Host Factor Results. 45
4 3. Explore pre-computed host factor bioset patterns a. From the Host Factor Experiment page, scroll down to Host Factor Bioset Patterns, which contains a summary table of statistically significant host factors grouped by expression patterns. Note the patterns that contain the largest numbers of host factors. Click the hyperlinked number in the first row/column to view the list of factors having the selected pattern, together with the fold change and statistical support values. b. Now search for the interferon beta gene and find its expression pattern in experiment ICL004-R. To do so, return to the Host Factor Experiment [IRD_SV_ICL004-R] page by clicking the breadcrumb, type IFNb in the Symbol search box within the Host Factor Bioset Patterns section, and click the Find button. 46
5 c. The expression patterns containing IFNb will be returned in the table below the Find box. What is the expression pattern for this gene? Does it match your expectation? To explain the pattern, we will need to look at the experimental results. d. Now click on the hyperlinked pattern (0,0,+,+,+,+), which will take you to the Experiment Results Details page. e. The Experiment Result Details page displays host factors and associated experimental results. You can select host factors and save them to a working set by clicking Add to, save the search to your Workbench so that you can rerun the same search at a later time, download data and associated metadata, or perform enrichment or pathway analysis. f. Next, click the select all box above the table and then the Download button to download the experiment results. g. In the Download Options box, select Special Tab Delimited - Displayed Columns Only, name the file ICL004-R IFNb pattern, and then click Download. 47
6 4. Perform host factor GO enrichment analysis a. From the Experiment Result Details page for expression pattern (0,0,+,+,+,+), make sure the Select all box above the table is checked, mouse over Run and click Enrichment. b. The Enrichment Algorithm and Gene-Annotation Collection page will be displayed. Select the following options and then click Run. Enrichment Algorithm: CLASSIFI; Gene-Annotation Collection: Gene Ontology (GO); Gene- Annotation Background: From Experiment. 48
7 c. The Enrichment Result page gives the terms, the collections (GO domains in this case), and the p-values calculated by the CLASSIFI algorithm using a hypergeometric distribution function. A selection of results is shown below. d. Select the first term by ticking the corresponding checkbox and click View Term Contents. e. A list of host factors annotated with this GO term will be displayed. 49
8 II. Compare Host Responses to MERS-CoV and SARS-CoV Infections This exercise is based on: Josset, L., et al. (2013) Cell Host Response to Infection with Novel Human Coronavirus EMC Predicts Potential Antivirals and Important Differences with SARS Coronavirus. mbio, 4(3): e MERS-CoV and SARS-CoV are members of the Coronaviridae, a family of enveloped, positive-stranded RNA viruses. Both viruses belong to the Betacoronavirus genus; however, MERS is closer to Bat HKU 4 or 5 (Lineage 2C) than SARS-CoV: they share 50% amino acid identity in the conserved replicase domains. Both viruses cause severe respiratory infections in humans with relatively high mortality frequencies: MERS at ~30%, and SARS at ~10%. These viruses bind to different receptors, specifically, SARS binds to ACE2, while MERS binds to Dipeptidyl peptidase 4 (DPP4). Although these receptors are conserved across many species, most Coronaviridae viruses have limited host range. Now we will compare the host responses to MERS-CoV and SARS-CoV infections to determine which host pathways show a shared response to these two severe respiratory pathogens. a. Navigate back to the Host Factor Experiments page. (Note: If using breadcrumb, prior selections will be remembered. Please deselect ICL004-R before proceeding.) b. Search for experiments containing MERS-CoV or SARS-CoV by selecting the following facets: Family: Coronaviridae; Viral Agent: MERS-CoV, icsars-cov Urbani; Analyte Type: Transcript Quantification c. Next, right-click (Command-click for Macs) on the experiment IDs for ECL001-R and SCL005-R, which were performed under similar conditions, and open them in new tabs. d. On the individual Host Factor Experiment pages, scroll down to the Host Factor Bioset Summary section. How are the host responses to these viruses different at the bioset level? Are the early responses different? Do the peak responses occur at similar times and/or to a similar magnitude? e. Next, we will compare the pattern trends between peak responses for MERS-CoV and SARS-CoV infections. Return to the Host Factor Biosets page, select ECL001_EMC_24h_array_DE and click Patterns above the table. f. The Host Factor Patterns page displays the pattern of the selected bioset. Click on the hyperlinked number corresponding with (+), which stands for up-regulation. g. This will take you to the Experiment Results Details page where it lists all host factors for the selected criteria. 50
9 Virus Pathogen Database and Resource (ViPR) - Flaviv... h. In order to compare the host factors from this pattern to those from another experimental pattern, we will need Flaviviridae to save the host factors to a working set. To do so, check the Select all checkbox at the top of the page, click Add to About Us Community Announcements located Links above Resources the Support table, then in the light SEARCH box, DATA name ANALYZE it MERS & VISUALIZE peak WORKBENCH + and click SUBMIT Save. DATA VIRUS FAMILIES You are logged in as yun.zhang@jcvi.org ViPR Home Flaviviridae Home Host Factor... Host Factor... Host Factor Patterns Host Factor Results Experiment Result Details For each host factor that exceeds the differential expression criteria defined in the experimental protocol, the Fold Change value and adjusted p-value or q-value is shown in the corresponding table cell. Otherwise, the cell is blank. Displaying 500 of 7,741 records. Select all 7,741 records (including those not displayed) Data sorted by Symbol, Host Factor ID in ascend order Your Selected Items: 7,741 items selected Deselect All Add to Save Search Download Download Reactome Data Pathway View HOST FACTOR ID OR SYMBOL Find data for Use comma to separate multiple entries. Ex. DDX58 Reset Associated Bioset Information 1 Study Name ECL001: MERS-CoV infection in Calu3 cells: A time course Experiment Name ECL001-R: MERS-CoV infection in Calu3 cells: A time course Bioset Criteria passes Agilent QC flag and FC > 1.5 and q-value <.05 ECL001-R MERS-CoV, 24 hours, 5 MOI, Calu3 cells Symbol Name Immport Host Factor ID Entrez Gene ID Public Identifier Bioset Information Key Log2 FC P-Value A1CF apobec1 complementation factor A_23_P NM_ A2BP1 ataxin 2-binding protein 1 A_24_P NM_ A2ML1 alpha-2-macroglobulin-like 1 A_23_P NM_ AADAC arylacetamide deacetylase (esterase) A_23_P NM_ i. Repeat AADACL1 the above arylacetamide steps deacetylase-like for bioset 1 SCL005-R_48_WT A_23_P and NM_ save the host 1 factors into 3.2a working set AADAT aminoadipate aminotransferase A_23_P NM_ named SARS peak +. AANAT arylalkylamine n-acetyltransferase A_23_P NM_ AASDH aminoadipate-semialdehyde A_24_P NM_ j. Click Workbench dehydrogenase in the gray navigation bar to access items saved in the Workbench area. AASDHPPT A_23_P NM_ aminoadipate-semialdehyde dehydrogenase- k. In the Workbench phosphopantetheinyl table, select transferasethe two working sets you just created, and then click on More AASDHPPT A_24_P NM_ Actions. aminoadipate-semialdehyde dehydrogenasephosphopantetheinyl transferase AATK apoptosis-associated tyrosine A_23_P NM_ l. This will bring kinase up the More Actions light box, click Intersect and name the new working set ABCA1 atp-binding cassette, sub-family a A_24_P NM_ intersect + MERS SARS. ABCA10 (abc1), member 1 atp-binding cassette, sub-family a (abc1), member 10 A_23_P NM_ ABCA11P ABCA12 ABCA13 atp-binding cassette, sub-family a (abc1), member 11 (pseudogene) atp-binding cassette, sub-family a (abc1), member 12 atp-binding cassette, sub-family a (abc1), member 13 A_23_P NR_ A_23_P NM_ A_23_P NM_
10 Virus Pathogen Database and Resource (ViPR) - Flaviv... SEARCH DATA ANALYZE & VISUALIZE WORKBENCH SUBMIT DATA VIRUS FAMILIES You are logged in as yun.zhang@jcvi.org ViPR Home Flaviviridae Home My Workbench My Workbench View workbench tutorial: Video Tutorial Tutorial View workbench tutorial: Download Video Tutorial Download Tutorial data of selected working set The workbench lets you store and retrieve various content types in private workspaces, including: working sets (e.g. segments, prot Copy Copy selected working sets uploaded files. Workbench items can be shared with selected collaborators with workbench accounts. Custom workbench The folders workbench ar lets you store and retrieve various content types in private workspaces, including: working sets (e.g. segments, proteins), saved searches, analysis results, and Combine Combine selected working sets into a new working set Display of workbench items can be restricted by content type, access privileges, and organizational folder using the checkboxes uploaded files. on Workbench items can be shared with selected collaborators with workbench accounts. Custom workbench folders are made available to organize content. Intersect Create intersection of selected working sets into a new working set Display of workbench items can be restricted by content type, access privileges, and organizational folder using the checkboxes on the left side of the page. Workbench tutorials, in either video or pdf formats, provide more information on how to use these features. Workbench tutorials, in either Create video Group or pdf formats, provide Add a more collaborator information group on how to use these features. Content s Searches Results Uploaded Files Access Private Shared by me Shared with me Public Flaviviridae Your Selected Items: 2 items selected Deselect All s Sharing Folders Move To Trash More Actions Sharing Folders Move To Trash More Actions UNAVAILABLE ACTIONS Searches Only certain actions are available to you for the items selected. To utilize the disabled actions, please Displaying go back 20 records and select per different page items. Display tings Displaying 20 Results records per page Display tings Uploaded Files Subtract Select all 57 items Create a new working set by subtract one working set from another Next > Page: 1 of 3 Select all 57 items About Us Community Announcements Links Resources Support Name ECL001-R 24h peak response + SCL005-R 48hr peak response + DENV 4 serotype groups Asia Content Type Convert Convert the selected working set into a new working set with different type Access Next > Page: 1 of 3 Content Name Data Type Items Folder Access Date Modified Combine with Uploaded File Combine the selected Type working set with an uploaded sequence Private Combine with ECL001-R 24h Combine peak the selected sequence Host set with Factor a working 7741 set -N/A- Private 2/23/2014 Data Type Items Shared Folder by me Access Date Modified response + 6:56 PM EST Unsubscribe Searches Unsubscribe selected searches from your workbench Shared with me SCL005-R 48hr peak Host Factor N/A- Private 2/23/2014 Host Factor 7741 Public -N/A- Save Private Unsaved Searches 2/23/2014Save selected unsaved searches to your workbench response + 6:53 PM EST Save Unsaved 6:56 PM EST Save selected unsaved analysis result(s) to the workbench Pending Public DENV 4 serotype groups Metadata -N/A- -N/A- Private 2/21/2014 Host Factor N/A- Private Asia 2/23/2014 Special Close 6:53 PM EST Unsaved DENV 4 serotype groups Metadata -N/A- -N/A- Private 2/21/2014 Metadata -N/A- Starred -N/A- Only Private America 2/21/2014 Trash Virus Protein 291 -N/A- Private 2/21/2014 DENV1-4 NS1 America Folders m. After the intersected working set is created, click View next to the working set name in the DENV 4 serotype groups Metadata -N/A- -N/A- Private 2/21/2014 Home Folder DENV1-4 NS1 Asia America Workbench table. Workbench Tutorial WNV-Chimera Uploaded DENV1-4 NS1 America Virus Protein 291 -N/A- Private 2/21/2014 WNV US Africa n. On Details DENV1-4 NS1 page, Asia select all Virus records Protein 411 by -N/A-checking Private the checkbox 2/21/2014 above the table, click Download. WNV-Chimera Nucleotide -N/A- -N/A- Private 2/20/2014 DENV 2 genome WNV US Africa Uploaded File Genome 43 -N/A- Private Dengue 2/20/ human Nov2013 DENV2 polyprotein o. The resulting file is WNV Excel compatible US Africa Tree and will -N/A- be -N/A- used to Private demonstrate /14/2014humandownstream off-site Dengue 2 poly Metadata DENV2 polyprotein MSA JalView -N/A- -N/A- Private Brazilian 12/3/2013 vs Nicaraguan analyses that can be performed. DENV 2 genome tree Tree -N/A- -N/A- Private 11/25/2013 Dengue 2 poly human Dengue 2 human Genome 34 -N/A- Shared By Me Dengue 11/8/2013 NS1 protein Metadata p. Next we will use the downloaded file to conduct an enrichment analysis. betwen 4 serotypes Open Nov2013 the downloaded DENV2 polyprotein Virus Protein 26 -N/A- Private Dengue 6/5/2013 NS1 protein file in Excel, sort the file human on Column C (Entrez gene ID), copy all IDs to the clipboard (don t Dengue Dengue 2 poly Metadata -N/A- -N/A- Private 6/4/2013 worry about duplicates), and then go to Brazilian vs Nicaraguan genome Dengue 2 poly Virus Protein 34 -N/A- Shared By Me 6/4/2013 human q. On the DAVID site, select the Functional Annotation tool on the left bar. Then proceed to upload Dengue NS1 protein Metadata -N/A- -N/A- Private 6/3/2013 betwen 4 serotypes data by choosing the tab Upload. Next, paste your list in the A: Paste a list field, select Dengue NS1 protein Virus Protein 728 -N/A- Private 6/3/2013 ENTREZ_GENE_ID from the Select Identifier pull down, and then select Gene List From Dengue Genome 75 -N/A- Private 5/1/2013 the List Type. Finally, click Submit and DAVID will run an enrichment analysis. genome Dengue 1-4 full genome human MSA Virus Pathogen Database and Resource (ViPR) - Flaviv... SEARCH DATA ANALYZE & VISUALIZE WORKBENCH SUBMIT DATA VIRUS FAMILIES You are logged in as yun.zhang@jcvi.org More Actions My Workbench AVAILABLE ACTIONS ViPR Home Flaviviridae Home My Workbench Muscle -N/A- -N/A- Private 4/30/2013 r. The pathway enrichment shown below demonstrates that the shared overlap between peak responses to SARS-CoV and MERS-CoV is comprised of immune response signaling pathways. 1 of 2 2/23/14 4:51 PM Content Flaviviridae Upload File Edit Group Upload a file About Us Community Announcements Links Resources Support Edit a collaborator group Your Selected Items: 2 items selected Deselect All File Search Workbench Virus Protein 411 -N/A- Private 2/21/2014 Nucleotide -N/A- -N/A- Private 2/20/2014 Genome 43 -N/A- Private 2/20/2014 WNV US Africa Tree -N/A- -N/A- Private 2/14/2014 DENV2 polyprotein MSA JalView -N/A- -N/A- Private 12/3/ tree Dengue 1-4 full genome human MSA Tree -N/A- -N/A- Private 11/25/ Genome 34 -N/A- Shared By Me 11/8/2013 Virus Protein 26 -N/A- Private 6/5/2013 -N/A- -N/A- Private 6/4/2013 Virus Protein 34 -N/A- Shared By Me 6/4/2013 -N/A- -N/A- Private 6/3/2013 Virus Protein 728 -N/A- Private 6/3/2013 Genome 75 -N/A- Private 5/1/2013 Muscle -N/A- -N/A- Private 4/30/ of 2 2/23/14 4:10 PM 52
11 References Li C, et al Host Regulatory Network Response to Infection with Highly Pathogenic H5N1 Avian Influenza Virus. J. Virol. 85: doi: /jvi Huang DW, et al Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 37:1-13. Huang DW, et al Systematic and integrative analysis of large gene lists using DAVID Bioinformatics Resources. Nature Protoc. 4: Josset L, et al Cell Host Response to Infection with Novel Human Coronavirus EMC Predicts Potential Antivirals and Important Differences with SARS Coronavirus. mbio 4:e doi: /mbio Lee JA, et al Components of the antigen processing and presentation pathway revealed by gene expression microarray analysis following B cell antigen receptor (BCR) stimulation. BMC Bioinformatics. 7:237. doi: / Müller MA, et al Human coronavirus EMC does not require the SARS-coronavirus receptor and maintains broad replicative capability in mammalian cell lines. mbio 3:e doi: /mbio Raj VS, et al Dipeptidyl peptidase 4 is a functional receptor for the emerging human coronavirus-emc. Nature 495: van Boheemen S, et al Genomic characterization of a newly discovered coronavirus associated with acute respiratory distress syndrome in humans. mbio 3:e doi: /mbio Zaki AM, et al Isolation of a novel coronavirus from a man with pneumonia in Saudi Arabia. N. Engl. J. Med. 367:
Section D. Identification of serotype-specific amino acid positions in DENV NS1. Objective
 Section D. Identification of serotype-specific amino acid positions in DENV NS1 Objective Upon completion of this exercise, you will be able to use the Virus Pathogen Resource (ViPR; http://www.viprbrc.org/)
Section D. Identification of serotype-specific amino acid positions in DENV NS1 Objective Upon completion of this exercise, you will be able to use the Virus Pathogen Resource (ViPR; http://www.viprbrc.org/)
Influenza H3N2 Virus Variation Analysis
 Influenza H3N2 Virus Variation Analysis Objective Upon completion of this exercise, you will be able to: search for virus sequences and view detailed information about these sequences in IRD, build a phylogenetic
Influenza H3N2 Virus Variation Analysis Objective Upon completion of this exercise, you will be able to: search for virus sequences and view detailed information about these sequences in IRD, build a phylogenetic
Rotavirus Genotyping and Enhanced Annotation in the Virus Pathogen Resource (ViPR) Yun Zhang J. Craig Venter Institute ASV 2016 June 19, 2016
 Rotavirus Genotyping and Enhanced Annotation in the Virus Pathogen Resource (ViPR) Yun Zhang J. Craig Venter Institute ASV 2016 June 19, 2016 Loading Virus Pathogen Database and Analysis About Resource
Rotavirus Genotyping and Enhanced Annotation in the Virus Pathogen Resource (ViPR) Yun Zhang J. Craig Venter Institute ASV 2016 June 19, 2016 Loading Virus Pathogen Database and Analysis About Resource
Section B. Comparative Genomics Analysis of Influenza H5N2 Viruses. Objective
 Section B. Comparative Genomics Analysis of Influenza H5N2 Viruses Objective Upon completion of this exercise, you will be able to use the Influenza Research Database (IRD; http://www.fludb.org/) to: Search
Section B. Comparative Genomics Analysis of Influenza H5N2 Viruses Objective Upon completion of this exercise, you will be able to use the Influenza Research Database (IRD; http://www.fludb.org/) to: Search
a. From the grey navigation bar, mouse over Analyze & Visualize and click Annotate Nucleotide Sequences.
 Section D. Custom sequence annotation After this exercise you should be able to use the annotation pipelines provided by the Influenza Research Database (IRD) and Virus Pathogen Resource (ViPR) to annotate
Section D. Custom sequence annotation After this exercise you should be able to use the annotation pipelines provided by the Influenza Research Database (IRD) and Virus Pathogen Resource (ViPR) to annotate
Module 3. Genomic data and annotations in public databases Exercises Custom sequence annotation
 Module 3. Genomic data and annotations in public databases Exercises Custom sequence annotation Objectives Upon completion of this exercise, you will be able to use the annotation pipelines provided by
Module 3. Genomic data and annotations in public databases Exercises Custom sequence annotation Objectives Upon completion of this exercise, you will be able to use the annotation pipelines provided by
SEQUENCE FEATURE VARIANT TYPES
 SEQUENCE FEATURE VARIANT TYPES DEFINITION OF SFVT: The Sequence Feature Variant Type (SFVT) component in IRD (http://www.fludb.org) is a relatively novel approach that delineates specific regions, called
SEQUENCE FEATURE VARIANT TYPES DEFINITION OF SFVT: The Sequence Feature Variant Type (SFVT) component in IRD (http://www.fludb.org) is a relatively novel approach that delineates specific regions, called
PROTOCOL FOR INFLUENZA A VIRUS GLOBAL SWINE H1 CLADE CLASSIFICATION
 PROTOCOL FOR INFLUENZA A VIRUS GLOBAL SWINE H1 CLADE CLASSIFICATION January 23, 2017 1. Background Swine H1 viruses have diversified into three major genetic lineages over time. Recently, Anderson et al.
PROTOCOL FOR INFLUENZA A VIRUS GLOBAL SWINE H1 CLADE CLASSIFICATION January 23, 2017 1. Background Swine H1 viruses have diversified into three major genetic lineages over time. Recently, Anderson et al.
Fondation Merieux J Craig Venter Institute Bioinformatics Workshop. December 5 8, 2017
 Fondation Merieux J Craig Venter Institute Bioinformatics Workshop December 5 8, 2017 Module 5: Comparative Genomics Analysis Outline Definition of comparative genomics Applications in studying human pathogens
Fondation Merieux J Craig Venter Institute Bioinformatics Workshop December 5 8, 2017 Module 5: Comparative Genomics Analysis Outline Definition of comparative genomics Applications in studying human pathogens
Module 3: Pathway and Drug Development
 Module 3: Pathway and Drug Development Table of Contents 1.1 Getting Started... 6 1.2 Identifying a Dasatinib sensitive cancer signature... 7 1.2.1 Identifying and validating a Dasatinib Signature... 7
Module 3: Pathway and Drug Development Table of Contents 1.1 Getting Started... 6 1.2 Identifying a Dasatinib sensitive cancer signature... 7 1.2.1 Identifying and validating a Dasatinib Signature... 7
Micro-RNA web tools. Introduction. UBio Training Courses. mirnas, target prediction, biology. Gonzalo
 Micro-RNA web tools UBio Training Courses Gonzalo Gómez//ggomez@cnio.es Introduction mirnas, target prediction, biology Experimental data Network Filtering Pathway interpretation mirs-pathways network
Micro-RNA web tools UBio Training Courses Gonzalo Gómez//ggomez@cnio.es Introduction mirnas, target prediction, biology Experimental data Network Filtering Pathway interpretation mirs-pathways network
Coronaviruses. Virion. Genome. Genes and proteins. Viruses and hosts. Diseases. Distinctive characteristics
 Coronaviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Spherical enveloped particles studded with clubbed spikes Diameter 120-160 nm Coiled helical
Coronaviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Spherical enveloped particles studded with clubbed spikes Diameter 120-160 nm Coiled helical
38 Int'l Conf. Bioinformatics and Computational Biology BIOCOMP'16
 38 Int'l Conf. Bioinformatics and Computational Biology BIOCOMP'16 PGAR: ASD Candidate Gene Prioritization System Using Expression Patterns Steven Cogill and Liangjiang Wang Department of Genetics and
38 Int'l Conf. Bioinformatics and Computational Biology BIOCOMP'16 PGAR: ASD Candidate Gene Prioritization System Using Expression Patterns Steven Cogill and Liangjiang Wang Department of Genetics and
Hands-On Ten The BRCA1 Gene and Protein
 Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
IMPaLA tutorial.
 IMPaLA tutorial http://impala.molgen.mpg.de/ 1. Introduction IMPaLA is a web tool, developed for integrated pathway analysis of metabolomics data alongside gene expression or protein abundance data. It
IMPaLA tutorial http://impala.molgen.mpg.de/ 1. Introduction IMPaLA is a web tool, developed for integrated pathway analysis of metabolomics data alongside gene expression or protein abundance data. It
BlueBayCT - Warfarin User Guide
 BlueBayCT - Warfarin User Guide December 2012 Help Desk 0845 5211241 Contents Getting Started... 1 Before you start... 1 About this guide... 1 Conventions... 1 Notes... 1 Warfarin Management... 2 New INR/Warfarin
BlueBayCT - Warfarin User Guide December 2012 Help Desk 0845 5211241 Contents Getting Started... 1 Before you start... 1 About this guide... 1 Conventions... 1 Notes... 1 Warfarin Management... 2 New INR/Warfarin
To open a CMA file > Download and Save file Start CMA Open file from within CMA
 Example name Effect size Analysis type Level Tamiflu Hospitalized Risk ratio Basic Basic Synopsis The US government has spent 1.4 billion dollars to stockpile Tamiflu, in anticipation of a possible flu
Example name Effect size Analysis type Level Tamiflu Hospitalized Risk ratio Basic Basic Synopsis The US government has spent 1.4 billion dollars to stockpile Tamiflu, in anticipation of a possible flu
To begin using the Nutrients feature, visibility of the Modules must be turned on by a MICROS Account Manager.
 Nutrients A feature has been introduced that will manage Nutrient information for Items and Recipes in myinventory. This feature will benefit Organizations that are required to disclose Nutritional information
Nutrients A feature has been introduced that will manage Nutrient information for Items and Recipes in myinventory. This feature will benefit Organizations that are required to disclose Nutritional information
Data Management, Data Management PLUS User Guide
 Data Management, Data Management PLUS User Guide Table of Contents Introduction 3 SHOEBOX Data Management and Data Management PLUS (DM+) for Individual Users 4 Portal Login 4 Working With Your Data 5 Manually
Data Management, Data Management PLUS User Guide Table of Contents Introduction 3 SHOEBOX Data Management and Data Management PLUS (DM+) for Individual Users 4 Portal Login 4 Working With Your Data 5 Manually
OneTouch Reveal Web Application. User Manual for Healthcare Professionals Instructions for Use
 OneTouch Reveal Web Application User Manual for Healthcare Professionals Instructions for Use Contents 2 Contents Chapter 1: Introduction...4 Product Overview...4 Intended Use...4 System Requirements...
OneTouch Reveal Web Application User Manual for Healthcare Professionals Instructions for Use Contents 2 Contents Chapter 1: Introduction...4 Product Overview...4 Intended Use...4 System Requirements...
Data Management System (DMS) User Guide
 Data Management System (DMS) User Guide Eversense and the Eversense logo are trademarks of Senseonics, Incorporated. Other brands and their products are trademarks or registered trademarks of their respective
Data Management System (DMS) User Guide Eversense and the Eversense logo are trademarks of Senseonics, Incorporated. Other brands and their products are trademarks or registered trademarks of their respective
The Hospital Anxiety and Depression Scale Guidance and Information
 The Hospital Anxiety and Depression Scale Guidance and Information About Testwise Testwise is the powerful online testing platform developed by GL Assessment to host its digital tests. Many of GL Assessment
The Hospital Anxiety and Depression Scale Guidance and Information About Testwise Testwise is the powerful online testing platform developed by GL Assessment to host its digital tests. Many of GL Assessment
Part I. Content: History of Viruses. General properties of viruses. Viral structure. Viral classifications. Virus-like agents.
 Viruses Part I Content: History of Viruses. General properties of viruses. Viral structure. Viral classifications. Virus-like agents. History Through the 1800s, many scientists discovered that something
Viruses Part I Content: History of Viruses. General properties of viruses. Viral structure. Viral classifications. Virus-like agents. History Through the 1800s, many scientists discovered that something
Exploring HIV Evolution: An Opportunity for Research Sam Donovan and Anton E. Weisstein
 Microbes Count! 137 Video IV: Reading the Code of Life Human Immunodeficiency Virus (HIV), like other retroviruses, has a much higher mutation rate than is typically found in organisms that do not go through
Microbes Count! 137 Video IV: Reading the Code of Life Human Immunodeficiency Virus (HIV), like other retroviruses, has a much higher mutation rate than is typically found in organisms that do not go through
Novel Coronavirus 2012
 Novel Coronavirus 2012 Susan I. Gerber, MD Respiratory Virus Program Division of Viral Diseases National Center for Immunization and Respiratory Diseases Centers for Disease Control and Prevention Jan.
Novel Coronavirus 2012 Susan I. Gerber, MD Respiratory Virus Program Division of Viral Diseases National Center for Immunization and Respiratory Diseases Centers for Disease Control and Prevention Jan.
Importància clínica dels nous virus emergents: grip i coronavirus
 Importància clínica dels nous virus emergents: grip i coronavirus Carolina Garcia-Vidal Servei de Malaties Infeccioses Hospital Universitari de Bellvitge Barcelona, Spain Neumonía viral: no sólo afecta
Importància clínica dels nous virus emergents: grip i coronavirus Carolina Garcia-Vidal Servei de Malaties Infeccioses Hospital Universitari de Bellvitge Barcelona, Spain Neumonía viral: no sólo afecta
Tutorial: RNA-Seq Analysis Part II: Non-Specific Matches and Expression Measures
 : RNA-Seq Analysis Part II: Non-Specific Matches and Expression Measures March 15, 2013 CLC bio Finlandsgade 10-12 8200 Aarhus N Denmark Telephone: +45 70 22 55 09 Fax: +45 70 22 55 19 www.clcbio.com support@clcbio.com
: RNA-Seq Analysis Part II: Non-Specific Matches and Expression Measures March 15, 2013 CLC bio Finlandsgade 10-12 8200 Aarhus N Denmark Telephone: +45 70 22 55 09 Fax: +45 70 22 55 19 www.clcbio.com support@clcbio.com
CaseBuilder - Quick Reference Guide
 ADP UNEMPLOYMENT COMPENSATION MANAGEMENT CaseBuilder - Quick Reference Guide After signing into CaseBuilder, the first screen the user will see is called the Dashboard. The user can then navigate to any
ADP UNEMPLOYMENT COMPENSATION MANAGEMENT CaseBuilder - Quick Reference Guide After signing into CaseBuilder, the first screen the user will see is called the Dashboard. The user can then navigate to any
Data mining with Ensembl Biomart. Stéphanie Le Gras
 Data mining with Ensembl Biomart Stéphanie Le Gras (slegras@igbmc.fr) Guidelines Genome data Genome browsers Getting access to genomic data: Ensembl/BioMart 2 Genome Sequencing Example: Human genome 2000:
Data mining with Ensembl Biomart Stéphanie Le Gras (slegras@igbmc.fr) Guidelines Genome data Genome browsers Getting access to genomic data: Ensembl/BioMart 2 Genome Sequencing Example: Human genome 2000:
Meta-analysis using RevMan. Yemisi Takwoingi October 2015
 Yemisi Takwoingi October 2015 Contents 1 Introduction... 1 2 Dataset 1 PART I..2 3 Starting RevMan... 2 4 Data and analyses in RevMan... 2 5 RevMan calculator tool... 2 Table 1. Data for derivation of
Yemisi Takwoingi October 2015 Contents 1 Introduction... 1 2 Dataset 1 PART I..2 3 Starting RevMan... 2 4 Data and analyses in RevMan... 2 5 RevMan calculator tool... 2 Table 1. Data for derivation of
CNV PCA Search Tutorial
 CNV PCA Search Tutorial Release 8.1 Golden Helix, Inc. March 18, 2014 Contents 1. Data Preparation 2 A. Join Log Ratio Data with Phenotype Information.............................. 2 B. Activate only
CNV PCA Search Tutorial Release 8.1 Golden Helix, Inc. March 18, 2014 Contents 1. Data Preparation 2 A. Join Log Ratio Data with Phenotype Information.............................. 2 B. Activate only
LESSON 4.4 WORKBOOK. How viruses make us sick: Viral Replication
 DEFINITIONS OF TERMS Eukaryotic: Non-bacterial cell type (bacteria are prokaryotes).. LESSON 4.4 WORKBOOK How viruses make us sick: Viral Replication This lesson extends the principles we learned in Unit
DEFINITIONS OF TERMS Eukaryotic: Non-bacterial cell type (bacteria are prokaryotes).. LESSON 4.4 WORKBOOK How viruses make us sick: Viral Replication This lesson extends the principles we learned in Unit
Content Part 2 Users manual... 4
 Content Part 2 Users manual... 4 Introduction. What is Kleos... 4 Case management... 5 Identity management... 9 Document management... 11 Document generation... 15 e-mail management... 15 Installation
Content Part 2 Users manual... 4 Introduction. What is Kleos... 4 Case management... 5 Identity management... 9 Document management... 11 Document generation... 15 e-mail management... 15 Installation
Clay Tablet Connector for hybris. User Guide. Version 1.5.0
 Clay Tablet Connector for hybris User Guide Version 1.5.0 August 4, 2016 Copyright Copyright 2005-2016 Clay Tablet Technologies Inc. All rights reserved. All rights reserved. This document and its content
Clay Tablet Connector for hybris User Guide Version 1.5.0 August 4, 2016 Copyright Copyright 2005-2016 Clay Tablet Technologies Inc. All rights reserved. All rights reserved. This document and its content
Of Camels, Bats and Coronaviruses: the (beginning of the) story of MERS-CoV
 Of Camels, Bats and Coronaviruses: the (beginning of the) story of MERS-CoV Allison McGeer, MSc, MD, FRCPC Mount Sinai Hospital University of Toronto Objectives Discuss the epidemiology, clinical presentation,
Of Camels, Bats and Coronaviruses: the (beginning of the) story of MERS-CoV Allison McGeer, MSc, MD, FRCPC Mount Sinai Hospital University of Toronto Objectives Discuss the epidemiology, clinical presentation,
Overview: Chapter 19 Viruses: A Borrowed Life
 Overview: Chapter 19 Viruses: A Borrowed Life Viruses called bacteriophages can infect and set in motion a genetic takeover of bacteria, such as Escherichia coli Viruses lead a kind of borrowed life between
Overview: Chapter 19 Viruses: A Borrowed Life Viruses called bacteriophages can infect and set in motion a genetic takeover of bacteria, such as Escherichia coli Viruses lead a kind of borrowed life between
NFT User Guide Release Date 01/09/2018
 NFT User Guide Release Date 01/09/2018 Center for Information Management, Inc. Table of Contents NFT List... 4 NFT Process Overview and Features... 10 NFT Process Steps... 12 Step 1: Agent clicks [Add]
NFT User Guide Release Date 01/09/2018 Center for Information Management, Inc. Table of Contents NFT List... 4 NFT Process Overview and Features... 10 NFT Process Steps... 12 Step 1: Agent clicks [Add]
Agile Product Lifecycle Management for Process
 Nutrition Surveillance Management User Guide Release 5.2.1 Part No. E13901-01 September 2008 Copyrights and Trademarks Copyright 1995, 2008, Oracle Corporation and/or its affiliates. All rights reserved.
Nutrition Surveillance Management User Guide Release 5.2.1 Part No. E13901-01 September 2008 Copyrights and Trademarks Copyright 1995, 2008, Oracle Corporation and/or its affiliates. All rights reserved.
SMPD 287 Spring 2015 Bioinformatics in Medical Product Development. Final Examination
 Final Examination You have a choice between A, B, or C. Please email your solutions, as a pdf attachment, by May 13, 2015. In the subject of the email, please use the following format: firstname_lastname_x
Final Examination You have a choice between A, B, or C. Please email your solutions, as a pdf attachment, by May 13, 2015. In the subject of the email, please use the following format: firstname_lastname_x
Quick Start Guide for the CPI Web Training Modules and Assessment FOR NEW USERS
 FOR NEW USERS Access to PT and PTA CPI Web will only be provided if you complete the training session and complete the PT and PTA CPI/WEB Assessment (CPI Assessment). You will only have to complete the
FOR NEW USERS Access to PT and PTA CPI Web will only be provided if you complete the training session and complete the PT and PTA CPI/WEB Assessment (CPI Assessment). You will only have to complete the
Variant Classification. Author: Mike Thiesen, Golden Helix, Inc.
 Variant Classification Author: Mike Thiesen, Golden Helix, Inc. Overview Sequencing pipelines are able to identify rare variants not found in catalogs such as dbsnp. As a result, variants in these datasets
Variant Classification Author: Mike Thiesen, Golden Helix, Inc. Overview Sequencing pipelines are able to identify rare variants not found in catalogs such as dbsnp. As a result, variants in these datasets
BREEAM In-Use International 2015 Client User Guide
 BREEAM In-Use International 2015 Client User Guide BREEAM In-Use International 2015 - Client User Guide V1.3.1 1 Contents Contents... 2 Log In and Registration... 3 Accessing the Online Tool... 3 Registration
BREEAM In-Use International 2015 Client User Guide BREEAM In-Use International 2015 - Client User Guide V1.3.1 1 Contents Contents... 2 Log In and Registration... 3 Accessing the Online Tool... 3 Registration
User Instruction Guide
 User Instruction Guide Table of Contents Logging In and Logging Out of MMSx 1 Creating a TPN (Terminal Profile Number) 2 Single Merchant 2 From Navigation Bar 2 From Home Page Link 4 Multiple Merchants
User Instruction Guide Table of Contents Logging In and Logging Out of MMSx 1 Creating a TPN (Terminal Profile Number) 2 Single Merchant 2 From Navigation Bar 2 From Home Page Link 4 Multiple Merchants
1. Automatically create Flu Shot encounters in AHLTA in 2 mouse clicks. 2. Ensure accurate DX and CPT codes used for every encounter, every time.
 In clinics around the MHS, upwards of 70% of all flu shot workload credit is lost because the encounters are not documented within AHLTA. Let the Immunization KAT s PASBA approved coding engine do the
In clinics around the MHS, upwards of 70% of all flu shot workload credit is lost because the encounters are not documented within AHLTA. Let the Immunization KAT s PASBA approved coding engine do the
Chronic Pain Management Workflow Getting Started: Wrenching In Assessments into Favorites (do once!)
 Chronic Pain Management Workflow Getting Started: Wrenching In Assessments into Favorites (do once!) 1. Click More Activities to star flowsheets into your chunky button screen. 3. Use the search function
Chronic Pain Management Workflow Getting Started: Wrenching In Assessments into Favorites (do once!) 1. Click More Activities to star flowsheets into your chunky button screen. 3. Use the search function
Envelope e_ :------, Envelope glycoproteins e_ ~ Single-stranded RNA ----, Nucleocapsid
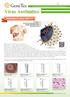 Virus Antib(tdies Your Expertise, Our Antibodies, Accelerated Discovery. Envelope e_---------:------, Envelope glycoproteins e_-------1~ Single-stranded RNA ----, Nucleocapsid First identified in 1989,
Virus Antib(tdies Your Expertise, Our Antibodies, Accelerated Discovery. Envelope e_---------:------, Envelope glycoproteins e_-------1~ Single-stranded RNA ----, Nucleocapsid First identified in 1989,
Influenza viruses. Virion. Genome. Genes and proteins. Viruses and hosts. Diseases. Distinctive characteristics
 Influenza viruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Enveloped particles, quasi-spherical or filamentous Diameter 80-120 nm Envelope is derived
Influenza viruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Enveloped particles, quasi-spherical or filamentous Diameter 80-120 nm Envelope is derived
User Guide. Association analysis. Input
 User Guide TFEA.ChIP is a tool to estimate transcription factor enrichment in a set of differentially expressed genes using data from ChIP-Seq experiments performed in different tissues and conditions.
User Guide TFEA.ChIP is a tool to estimate transcription factor enrichment in a set of differentially expressed genes using data from ChIP-Seq experiments performed in different tissues and conditions.
DNA Sequence Bioinformatics Analysis with the Galaxy Platform
 DNA Sequence Bioinformatics Analysis with the Galaxy Platform University of São Paulo, Brazil 28 July - 1 August 2014 Dave Clements Johns Hopkins University Robson Francisco de Souza University of São
DNA Sequence Bioinformatics Analysis with the Galaxy Platform University of São Paulo, Brazil 28 July - 1 August 2014 Dave Clements Johns Hopkins University Robson Francisco de Souza University of São
v Feature Stamping SMS 12.0 Tutorial Prerequisites Requirements TABS model Map Module Mesh Module Scatter Module Time minutes
 v. 12.0 SMS 12.0 Tutorial Objectives In this lesson will teach how to use conceptual modeling techniques to create numerical models that incorporate flow control structures into existing bathymetry. The
v. 12.0 SMS 12.0 Tutorial Objectives In this lesson will teach how to use conceptual modeling techniques to create numerical models that incorporate flow control structures into existing bathymetry. The
Quick-Start Guide TeamUnify, LLC
 Quick-Start Guide Setup Basics System Settings 1 When you initially sign up for MainSet, you need to set up Roster Group colors and designate coaches. first, Click settings 1. Navigate to http://mainset.com
Quick-Start Guide Setup Basics System Settings 1 When you initially sign up for MainSet, you need to set up Roster Group colors and designate coaches. first, Click settings 1. Navigate to http://mainset.com
CIViC Clinical Interpretations of Variants in Cancer. rnal/v49/n2/full/ng.3774.html
 CIViC Clinical Interpretations of Variants in Cancer www.civicdb.org http://www.nature.com/ng/jou rnal/v49/n2/full/ng.3774.html CIViC Homepage Aim Statement Curation Stats Recent Activity Links at Bottom
CIViC Clinical Interpretations of Variants in Cancer www.civicdb.org http://www.nature.com/ng/jou rnal/v49/n2/full/ng.3774.html CIViC Homepage Aim Statement Curation Stats Recent Activity Links at Bottom
CS 312: Algorithms Analysis. Gene Sequence Alignment. Overview: Objectives: Code:
 CS 312: Algorithms Analysis Gene Sequence Alignment Overview: In this project, you will implement dynamic programming algorithms for computing the minimal cost of aligning gene sequences and for extracting
CS 312: Algorithms Analysis Gene Sequence Alignment Overview: In this project, you will implement dynamic programming algorithms for computing the minimal cost of aligning gene sequences and for extracting
User Guide for Classification of Diabetes: A search tool for identifying miscoded, misclassified or misdiagnosed patients
 User Guide for Classification of Diabetes: A search tool for identifying miscoded, misclassified or misdiagnosed patients For use with isoft Premiere Synergy Produced by André Ring 1 Table of Contents
User Guide for Classification of Diabetes: A search tool for identifying miscoded, misclassified or misdiagnosed patients For use with isoft Premiere Synergy Produced by André Ring 1 Table of Contents
Bioinformatics Laboratory Exercise
 Bioinformatics Laboratory Exercise Biology is in the midst of the genomics revolution, the application of robotic technology to generate huge amounts of molecular biology data. Genomics has led to an explosion
Bioinformatics Laboratory Exercise Biology is in the midst of the genomics revolution, the application of robotic technology to generate huge amounts of molecular biology data. Genomics has led to an explosion
MERS H7N9. coronavirus. influenza virus. MERS: Since September 2012: 160 cases, including 68 deaths (42%)
 MERS coronavirus H7N9 influenza virus Marc Van Ranst, KU Leuven MERS: Since September 2012: 160 cases, including 68 deaths (42%) MERS coronavirus patient, 83 years, Saudi-Arabia H7N9: Since February 2013:
MERS coronavirus H7N9 influenza virus Marc Van Ranst, KU Leuven MERS: Since September 2012: 160 cases, including 68 deaths (42%) MERS coronavirus patient, 83 years, Saudi-Arabia H7N9: Since February 2013:
Influenza Virus HA Subtype Numbering Conversion Tool and the Identification of Candidate Cross-Reactive Immune Epitopes
 Influenza Virus HA Subtype Numbering Conversion Tool and the Identification of Candidate Cross-Reactive Immune Epitopes Brian J. Reardon, Ph.D. J. Craig Venter Institute breardon@jcvi.org Introduction:
Influenza Virus HA Subtype Numbering Conversion Tool and the Identification of Candidate Cross-Reactive Immune Epitopes Brian J. Reardon, Ph.D. J. Craig Venter Institute breardon@jcvi.org Introduction:
Add_A_Class_with_Class_Number_Revised Thursday, March 18, 2010
 Slide 1 Text Captions: PAWS Tutorial "Add a Class using Class Number" Created for: Version 9.0 Date: March, 2010 Slide 2 Text Captions: Objective In this tutorial you will learn how to add a class to your
Slide 1 Text Captions: PAWS Tutorial "Add a Class using Class Number" Created for: Version 9.0 Date: March, 2010 Slide 2 Text Captions: Objective In this tutorial you will learn how to add a class to your
Cancer Informatics Lecture
 Cancer Informatics Lecture Mayo-UIUC Computational Genomics Course June 22, 2018 Krishna Rani Kalari Ph.D. Associate Professor 2017 MFMER 3702274-1 Outline The Cancer Genome Atlas (TCGA) Genomic Data Commons
Cancer Informatics Lecture Mayo-UIUC Computational Genomics Course June 22, 2018 Krishna Rani Kalari Ph.D. Associate Professor 2017 MFMER 3702274-1 Outline The Cancer Genome Atlas (TCGA) Genomic Data Commons
Update: Porcine epidemic diarrhoea and swine influenza. Eric Neumann DVM PhD Epi-Insight Limited Palmerston North, New Zealand
 Update: Porcine epidemic diarrhoea and swine influenza Eric Neumann DVM PhD Epi-Insight Limited Palmerston North, New Zealand Approach Review of the pathogen Review of the outbreak Research outcomes in
Update: Porcine epidemic diarrhoea and swine influenza Eric Neumann DVM PhD Epi-Insight Limited Palmerston North, New Zealand Approach Review of the pathogen Review of the outbreak Research outcomes in
Data Management System (DMS) User Guide
 Data Management System (DMS) User Guide Eversense and the Eversense logo are trademarks of Senseonics, Incorporated. Other brands and their products are trademarks or registered trademarks of their respective
Data Management System (DMS) User Guide Eversense and the Eversense logo are trademarks of Senseonics, Incorporated. Other brands and their products are trademarks or registered trademarks of their respective
Severe Acute Respiratory Syndrome (SARS) Coronavirus
 Severe Acute Respiratory Syndrome (SARS) Coronavirus Coronaviruses Coronaviruses are single stranded enveloped RNA viruses that have a helical geometry. Coronaviruses are the largest of RNA viruses with
Severe Acute Respiratory Syndrome (SARS) Coronavirus Coronaviruses Coronaviruses are single stranded enveloped RNA viruses that have a helical geometry. Coronaviruses are the largest of RNA viruses with
بسم هللا الرحمن الرحيم
 - 1 - - - 1 P a g e بسم هللا الرحمن الرحيم This sheet was made from record section 1 all information are included - Introduction Our respiratory tract is divided anatomically to upper (URT),middle and
- 1 - - - 1 P a g e بسم هللا الرحمن الرحيم This sheet was made from record section 1 all information are included - Introduction Our respiratory tract is divided anatomically to upper (URT),middle and
Coronaviruses cause acute, mild upper respiratory infection (common cold).
 Coronaviruses David A. J. Tyrrell Steven H. Myint GENERAL CONCEPTS Clinical Presentation Coronaviruses cause acute, mild upper respiratory infection (common cold). Structure Spherical or pleomorphic enveloped
Coronaviruses David A. J. Tyrrell Steven H. Myint GENERAL CONCEPTS Clinical Presentation Coronaviruses cause acute, mild upper respiratory infection (common cold). Structure Spherical or pleomorphic enveloped
OECD QSAR Toolbox v.4.2. An example illustrating RAAF scenario 6 and related assessment elements
 OECD QSAR Toolbox v.4.2 An example illustrating RAAF scenario 6 and related assessment elements Outlook Background Objectives Specific Aims Read Across Assessment Framework (RAAF) The exercise Workflow
OECD QSAR Toolbox v.4.2 An example illustrating RAAF scenario 6 and related assessment elements Outlook Background Objectives Specific Aims Read Across Assessment Framework (RAAF) The exercise Workflow
Section B. Comparative Genomics Analysis of 2013 H7N9 Influenza A Viruses. Objective
 Section B. Comparative Genomics Analysis of 2013 H7N9 Influenza A Viruses Objective Upon completion of this exercise, you will be able to use the Influenza Research Database (IRD; http://www.fludb.org/)
Section B. Comparative Genomics Analysis of 2013 H7N9 Influenza A Viruses Objective Upon completion of this exercise, you will be able to use the Influenza Research Database (IRD; http://www.fludb.org/)
NYSIIS. Immunization Evaluator and Manage Schedule Manual. October 16, Release 1.0
 NYSIIS Immunization Evaluator and Manage Schedule Manual October 16, 2007 Release 1.0 Revision History Release Modifications Author Revision Date 1.0 Initial Draft R Savage 10/16/2007 INTRODUCTION: 1 BUILDING
NYSIIS Immunization Evaluator and Manage Schedule Manual October 16, 2007 Release 1.0 Revision History Release Modifications Author Revision Date 1.0 Initial Draft R Savage 10/16/2007 INTRODUCTION: 1 BUILDING
MS/MS Library Creation of Q-TOF LC/MS Data for MassHunter PCDL Manager
 MS/MS Library Creation of Q-TOF LC/MS Data for MassHunter PCDL Manager Quick Start Guide Step 1. Calibrate the Q-TOF LC/MS for low m/z ratios 2 Step 2. Set up a Flow Injection Analysis (FIA) method for
MS/MS Library Creation of Q-TOF LC/MS Data for MassHunter PCDL Manager Quick Start Guide Step 1. Calibrate the Q-TOF LC/MS for low m/z ratios 2 Step 2. Set up a Flow Injection Analysis (FIA) method for
Instructor Guide to EHR Go
 Instructor Guide to EHR Go Introduction... 1 Quick Facts... 1 Creating your Account... 1 Logging in to EHR Go... 5 Adding Faculty Users to EHR Go... 6 Adding Student Users to EHR Go... 8 Library... 9 Patients
Instructor Guide to EHR Go Introduction... 1 Quick Facts... 1 Creating your Account... 1 Logging in to EHR Go... 5 Adding Faculty Users to EHR Go... 6 Adding Student Users to EHR Go... 8 Library... 9 Patients
Bjoern Peters La Jolla Institute for Allergy and Immunology Buenos Aires, Oct 31, 2012
 www.iedb.org Bjoern Peters bpeters@liai.org La Jolla Institute for Allergy and Immunology Buenos Aires, Oct 31, 2012 Overview 1. Introduction to the IEDB 2. Application: 2009 Swine-origin influenza virus
www.iedb.org Bjoern Peters bpeters@liai.org La Jolla Institute for Allergy and Immunology Buenos Aires, Oct 31, 2012 Overview 1. Introduction to the IEDB 2. Application: 2009 Swine-origin influenza virus
Core Technology Development Team Meeting
 Core Technology Development Team Meeting To hear the meeting, you must call in Toll-free phone number: 1-866-740-1260 Access Code: 2201876 For international call in numbers, please visit: https://www.readytalk.com/account-administration/international-numbers
Core Technology Development Team Meeting To hear the meeting, you must call in Toll-free phone number: 1-866-740-1260 Access Code: 2201876 For international call in numbers, please visit: https://www.readytalk.com/account-administration/international-numbers
Bioinformation by Biomedical Informatics Publishing Group
 Predicted RNA secondary structures for the conserved regions in dengue virus Pallavi Somvanshi*, Prahlad Kishore Seth Bioinformatics Centre, Biotech Park, Sector G, Jankipuram, Lucknow 226021, Uttar Pradesh,
Predicted RNA secondary structures for the conserved regions in dengue virus Pallavi Somvanshi*, Prahlad Kishore Seth Bioinformatics Centre, Biotech Park, Sector G, Jankipuram, Lucknow 226021, Uttar Pradesh,
Technical Bulletin. Technical Information for Quidel Molecular Influenza A+B Assay on the Bio-Rad CFX96 Touch
 Technical Bulletin Technical Information for Quidel Molecular Influenza A+B Assay on the Bio-Rad CFX96 Touch Quidel Corporation has verified the performance of the Quidel Molecular Influenza A+B Assay
Technical Bulletin Technical Information for Quidel Molecular Influenza A+B Assay on the Bio-Rad CFX96 Touch Quidel Corporation has verified the performance of the Quidel Molecular Influenza A+B Assay
User Guide. Protein Clpper. Statistical scoring of protease cleavage sites. 1. Introduction Protein Clpper Analysis Procedure...
 User Guide Protein Clpper Statistical scoring of protease cleavage sites Content 1. Introduction... 2 2. Protein Clpper Analysis Procedure... 3 3. Input and Output Files... 9 4. Contact Information...
User Guide Protein Clpper Statistical scoring of protease cleavage sites Content 1. Introduction... 2 2. Protein Clpper Analysis Procedure... 3 3. Input and Output Files... 9 4. Contact Information...
Supplemental Information Dose Response Parameters for Gain of Function Pathogens
 Supplemental Information Dose Response Parameters for Gain of Function Pathogens Infection Dose-Response To quantify the likelihood of an individual or animal becoming infected from exposure to virus,
Supplemental Information Dose Response Parameters for Gain of Function Pathogens Infection Dose-Response To quantify the likelihood of an individual or animal becoming infected from exposure to virus,
Data Management System (DMS) User Guide
 Data Management System (DMS) User Guide Eversense and the Eversense logo are trademarks of Senseonics, Incorporated. Other brands and their products are trademarks or registered trademarks of their respective
Data Management System (DMS) User Guide Eversense and the Eversense logo are trademarks of Senseonics, Incorporated. Other brands and their products are trademarks or registered trademarks of their respective
Evolution of influenza
 Evolution of influenza Today: 1. Global health impact of flu - why should we care? 2. - what are the components of the virus and how do they change? 3. Where does influenza come from? - are there animal
Evolution of influenza Today: 1. Global health impact of flu - why should we care? 2. - what are the components of the virus and how do they change? 3. Where does influenza come from? - are there animal
ShadeVision v Color Map
 Kevin Aamodt Page 1 7/1/2004 ShadeVision v.3.01 Color Map The ShadeVision v.3.01 color map will allow the user to custom select the lookup regions of the Gingival, Middle, and Incisal areas of a tooth.
Kevin Aamodt Page 1 7/1/2004 ShadeVision v.3.01 Color Map The ShadeVision v.3.01 color map will allow the user to custom select the lookup regions of the Gingival, Middle, and Incisal areas of a tooth.
ProScript User Guide. Pharmacy Access Medicines Manager
 User Guide Pharmacy Access Medicines Manager Version 3.0.0 Release Date 01/03/2014 Last Reviewed 11/04/2014 Author Rx Systems Service Desk (T): 01923 474 600 Service Desk (E): servicedesk@rxsystems.co.uk
User Guide Pharmacy Access Medicines Manager Version 3.0.0 Release Date 01/03/2014 Last Reviewed 11/04/2014 Author Rx Systems Service Desk (T): 01923 474 600 Service Desk (E): servicedesk@rxsystems.co.uk
IPA Advanced Training Course
 IPA Advanced Training Course October 2013 Academia sinica Gene (Kuan Wen Chen) IPA Certified Analyst Agenda I. Data Upload and How to Run a Core Analysis II. Functional Interpretation in IPA Hands-on Exercises
IPA Advanced Training Course October 2013 Academia sinica Gene (Kuan Wen Chen) IPA Certified Analyst Agenda I. Data Upload and How to Run a Core Analysis II. Functional Interpretation in IPA Hands-on Exercises
PBSI-EHR Off the Charts!
 PBSI-EHR Off the Charts! Enhancement Release 3.2.1 TABLE OF CONTENTS Description of enhancement change Page Encounter 2 Patient Chart 3 Meds/Allergies/Problems 4 Faxing 4 ICD 10 Posting Overview 5 Master
PBSI-EHR Off the Charts! Enhancement Release 3.2.1 TABLE OF CONTENTS Description of enhancement change Page Encounter 2 Patient Chart 3 Meds/Allergies/Problems 4 Faxing 4 ICD 10 Posting Overview 5 Master
Introduced ICD Changes in Charting... 2
 Table of Contents InSync Product Release Notes January 2014 Introduced ICD-10... 2 Changes in Charting... 2 Configuring ICD-10 at Practice Level... 2 ICD-10 Master List... 3 Practice Favorite List... 4
Table of Contents InSync Product Release Notes January 2014 Introduced ICD-10... 2 Changes in Charting... 2 Configuring ICD-10 at Practice Level... 2 ICD-10 Master List... 3 Practice Favorite List... 4
LESSON 4.5 WORKBOOK. How do viruses adapt Antigenic shift and drift and the flu pandemic
 DEFINITIONS OF TERMS Gene a particular sequence of DNA or RNA that contains information for the synthesis of a protien or RNA molecule. For a complete list of defined terms, see the Glossary. LESSON 4.5
DEFINITIONS OF TERMS Gene a particular sequence of DNA or RNA that contains information for the synthesis of a protien or RNA molecule. For a complete list of defined terms, see the Glossary. LESSON 4.5
Self Assessment 8.3 to 8.4.x
 Self Assessment 8.3 to 8.4.x User Guide November 30, 2009 Contents What Self Assessments does Managing self assessments Creating self assessments Adding questions to your self assessment Grading and answers
Self Assessment 8.3 to 8.4.x User Guide November 30, 2009 Contents What Self Assessments does Managing self assessments Creating self assessments Adding questions to your self assessment Grading and answers
Creating a User Defined Pneumonia-Specific Syndrome in ESSENCE. Preventive Medicine Directorate September 2016
 Creating a User Defined Pneumonia-Specific Syndrome in ESSENCE Preventive Medicine Directorate September 2016 0 Pneumonia-Specific Syndrome NMCPHC retrospective analyses suggest that surveillance using
Creating a User Defined Pneumonia-Specific Syndrome in ESSENCE Preventive Medicine Directorate September 2016 0 Pneumonia-Specific Syndrome NMCPHC retrospective analyses suggest that surveillance using
Titrations in Cytobank
 The Premier Platform for Single Cell Analysis (1) Titrations in Cytobank To analyze data from a titration in Cytobank, the first step is to upload your own FCS files or clone an experiment you already
The Premier Platform for Single Cell Analysis (1) Titrations in Cytobank To analyze data from a titration in Cytobank, the first step is to upload your own FCS files or clone an experiment you already
Regulation of cell signaling cascades by influenza A virus
 The Second International Symposium on Optimization and Systems Biology (OSB 08) Lijiang, China, October 31 November 3, 2008 Copyright 2008 ORSC & APORC, pp. 389 394 Regulation of cell signaling cascades
The Second International Symposium on Optimization and Systems Biology (OSB 08) Lijiang, China, October 31 November 3, 2008 Copyright 2008 ORSC & APORC, pp. 389 394 Regulation of cell signaling cascades
LEFT VENTRICLE SEGMENTATION AND MEASUREMENT Using Analyze
 LEFT VENTRICLE SEGMENTATION AND MEASUREMENT Using Analyze 2 Table of Contents 1. Introduction page 3 2. Segmentation page 4 3. Measurement Instructions page 11 4. Calculation Instructions page 14 5. References
LEFT VENTRICLE SEGMENTATION AND MEASUREMENT Using Analyze 2 Table of Contents 1. Introduction page 3 2. Segmentation page 4 3. Measurement Instructions page 11 4. Calculation Instructions page 14 5. References
PedCath IMPACT User s Guide
 PedCath IMPACT User s Guide Contents Overview... 3 IMPACT Overview... 3 PedCath IMPACT Registry Module... 3 More on Work Flow... 4 Case Complete Checkoff... 4 PedCath Cath Report/IMPACT Shared Data...
PedCath IMPACT User s Guide Contents Overview... 3 IMPACT Overview... 3 PedCath IMPACT Registry Module... 3 More on Work Flow... 4 Case Complete Checkoff... 4 PedCath Cath Report/IMPACT Shared Data...
Training Peaks P90X and P90X2 Workout Schedule Instructions
 Training Peaks P90X and P90X2 Workout Schedule Instructions Go to www.trainingpeaks.com and click on Products Click on Plans Click on Training Plans. Don t worry about the options to browse by type or
Training Peaks P90X and P90X2 Workout Schedule Instructions Go to www.trainingpeaks.com and click on Products Click on Plans Click on Training Plans. Don t worry about the options to browse by type or
Pathway Exercises Metabolism and Pathways
 1. Find the metabolic pathway for glycolysis. For this exercise use PlasmoDB.org Pathway Exercises Metabolism and Pathways a. Navigate to the search page for Identify Metabolic Pathways based on Pathway
1. Find the metabolic pathway for glycolysis. For this exercise use PlasmoDB.org Pathway Exercises Metabolism and Pathways a. Navigate to the search page for Identify Metabolic Pathways based on Pathway
From Birds to People: The West Nile Virus Story
 Howard Hughes Medical Institute From Birds to People: The West Nile Virus Story About This Worksheet This worksheet complements the Click and Learn From Birds to People: The West Nile Virus Story. Author:
Howard Hughes Medical Institute From Birds to People: The West Nile Virus Story About This Worksheet This worksheet complements the Click and Learn From Birds to People: The West Nile Virus Story. Author:
Entering HIV Testing Data into EvaluationWeb
 Entering HIV Testing Data into EvaluationWeb User Guide Luther Consulting, LLC July, 2014/v2.2 All rights reserved. Table of Contents Introduction... 3 Accessing the CTR Form... 4 Overview of the CTR Form...
Entering HIV Testing Data into EvaluationWeb User Guide Luther Consulting, LLC July, 2014/v2.2 All rights reserved. Table of Contents Introduction... 3 Accessing the CTR Form... 4 Overview of the CTR Form...
Documenting Patient Immunization. New Brunswick 2018/19
 Documenting Patient Immunization New Brunswick 2018/19 Table of Contents Documenting Patient Immunization New Brunswick...3 Immunization Module Features...4 Configuration...5 Marketing Message Setup...6
Documenting Patient Immunization New Brunswick 2018/19 Table of Contents Documenting Patient Immunization New Brunswick...3 Immunization Module Features...4 Configuration...5 Marketing Message Setup...6
Add_A_Class_with_Class_Search_Revised Thursday, March 18, 2010
 Slide 1 Text Captions: PAWS Tutorial "Add a Class using Class Search" Created for: Version 9.0 Date: March, 2010 Slide 2 Text Captions: Objective In this tutorial you will learn how to add a class to your
Slide 1 Text Captions: PAWS Tutorial "Add a Class using Class Search" Created for: Version 9.0 Date: March, 2010 Slide 2 Text Captions: Objective In this tutorial you will learn how to add a class to your
Aggregate Report Instructions
 Version 2018_v4 Workplace Health Solutions Center for Workplace Health Research & Evaluation Version 2018_v4 Table of Contents Purpose.....3 Data Privacy....3 About Life's Simple 7....4 Table 1. Life's
Version 2018_v4 Workplace Health Solutions Center for Workplace Health Research & Evaluation Version 2018_v4 Table of Contents Purpose.....3 Data Privacy....3 About Life's Simple 7....4 Table 1. Life's
RESULTS REPORTING MANUAL. Hospital Births Newborn Screening Program June 2016
 RESULTS REPORTING MANUAL Hospital Births Newborn Screening Program June 2016 CONTENTS GETTING STARTED... 1 Summary... 1 Logging In... 1 Access For New Hires... 2 Reporting Parental Refusals... 3 Adding
RESULTS REPORTING MANUAL Hospital Births Newborn Screening Program June 2016 CONTENTS GETTING STARTED... 1 Summary... 1 Logging In... 1 Access For New Hires... 2 Reporting Parental Refusals... 3 Adding
ISR Process for Internal Service Providers
 ISR Process for Internal Service Providers Course Outline 1) Internal Service Request Process Overview Internal Service Requests (ISR) are created by the end user via the BUworks Central Portal Procurement
ISR Process for Internal Service Providers Course Outline 1) Internal Service Request Process Overview Internal Service Requests (ISR) are created by the end user via the BUworks Central Portal Procurement
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature14008 Supplementary Figure 1. Sequence alignment of A/little yellow-shouldered bat/guatemala/060/2010 (H17N10) polymerase with that of human strain A/Victoria/3/75(H3N2). The secondary
doi:10.1038/nature14008 Supplementary Figure 1. Sequence alignment of A/little yellow-shouldered bat/guatemala/060/2010 (H17N10) polymerase with that of human strain A/Victoria/3/75(H3N2). The secondary
IMPORTANT!!! Please read the FAQ document BEFORE you step through this tutorial.
 IMPORTANT!!! Please read the FAQ document BEFORE you step through this tutorial. If you choose to participate in the CrossFit Viral Nutrition Challenge, please purchase the Premium version of My Fitness
IMPORTANT!!! Please read the FAQ document BEFORE you step through this tutorial. If you choose to participate in the CrossFit Viral Nutrition Challenge, please purchase the Premium version of My Fitness
