... Department of Biological Sciences, John Tabor Laboratories, University of Essex, Colchester CO4 3SQ, UK
|
|
|
- Britney McGee
- 5 years ago
- Views:
Transcription
1 Journal of General Virology (2000), 81, Printed in Great Britain... The 2A proteins of three diverse picornaviruses are related to each other and to the H-rev107 family of proteins involved in the control of cell proliferation Pamela J. Hughes and Glyn Stanway Department of Biological Sciences, John Tabor Laboratories, University of Essex, Colchester CO4 3SQ, UK The 2A protein appears to be diverse among picornaviruses, in contrast to the other non-structural proteins, which have homologous structures and functions. In enteroviruses and rhinoviruses, 2A is a trypsin-like protease involved in protein processing and in shut-off of host-cell macromolecular synthesis. The aphthovirus and cardiovirus 2A is associated with an unusual processing event at the 2A/2B junction. It is shown here that the 2A protein of several diverse picornaviruses, the human parechoviruses, Aichi virus and avian encephalomyelitis virus, possess previously unrecognized conserved motifs and are likely to have a common function. Moreover, these motifs, a conserved histidine and flanking amino acids, an asparagine cysteine dipeptide and a putative transmembrane domain, are characteristic of a family of cellular proteins, at least two of which are involved in the control of cell growth. These observations have important implications for an understanding of picornavirus genome structure and evolution, as well as pointing to possible functions of 2A in these viruses. Introduction The family Picornaviridae, which contains several important human and animal pathogens, is a large and diverse family of positive-sense RNA viruses (Stanway, 1990; Rueckert, 1996). More than 230 serotypes have been recognized and are currently grouped into six genera: Aphthovirus, Cardiovirus, Enterovirus, Hepatovirus, Parechovirus and Rhinovirus (King et al., 1999; Stanway & Hyypia, 1999). Another three viruses, Aichi virus, porcine enterovirus 1 [to be renamed porcine teschovirus 1 (PTV1)] and equine rhinotracheitis virus B (ERBV), are likely to be classified as type members of new genera in the future, as they exhibit significant diversity from members of existing genera (Wutz et al., 1996; Yamashita et al., 1998; Doherty et al., 1999; King et al., 1999). The picornavirus genome is nucleotides long and is enclosed within a naked particle, made up of 60 copies of each of the capsid proteins. The genome encodes a single polyprotein, which undergoes a cleavage cascade performed by virus-encoded activities, to give the final virus proteins. The genomes of the various genera encode 10, 11 or 12 final Author for correspondence: Glyn Stanway. Fax stanwg essex.ac.uk proteins, but intermediates in the cascade can have a significant half-life and may have distinct functions (Rueckert, 1996). All picornaviruses share essentially the same genome organization. The capsid proteins (1A, 1B, 1C and 1D, commonly known as VP4, VP2, VP3 and VP1, respectively) are encoded towards the N terminus of the polyprotein and the non-structural proteins (2A, 2B, 2C, 3A, 3B, 3C and 3D) are encoded downstream of these. VP4 and VP2 are assembled into the particle in the form of a precursor, VP0, the final step of assembly being the cleavage of this precursor, which appears to be required for virion infectivity and stability. However, in the human parechoviruses and in Aichi virus, this cleavage appears not to occur and consequently these particles have only three proteins, VP0, VP3 and VP1 (Hyypia et al., 1992; Stanway et al., 1994; Yamashita et al., 1998). For most picornavirus proteins, there is clear homology between the corresponding protein-encoding region of different genera. One exception is L, the protein encoded at the extreme N terminus of the polyprotein in aphthoviruses, cardioviruses, ERBV and Aichi virus (Rueckert, 1996; Wutz et al., 1996; Yamashita et al., 1998). With the exception of aphthoviruses and ERBV, where they are homologues, picornavirus L proteins are structurally and functionally distinct (Rueckert, 1996). The other variable picornavirus locus is the SGM CAB
2 P. J. Hughes and G. Stanway region encoding 2A. In rhinoviruses and enteroviruses, 2A is a trypsin-like, cysteine protease involved in polyprotein processing, while in cardioviruses, aphthoviruses, ERBV and PTV1, it is associated with an unusual processing activity, involving an Asn Pro Gly Pro (NPGP) motif (Ryan & Flint, 1997). The 2A proteins of parechoviruses, hepatoviruses and Aichi virus are not believed to be involved in processing (Schultheiss et al., 1995; Jia et al., 1993; Yamashita et al., 1998). In addition, no similarity between the 2A protein of parechoviruses, hepatoviruses and Aichi virus has previously been noted. We show here that the 2A proteins of parechoviruses are, in fact, homologous to the corresponding region of the polyprotein of the hepato-like virus, avian encephalomyelitis virus (AEV), and, to a lesser extent, to the 2A protein of Aichi virus (Marvil et al., 1999; Yamashita et al., 1998). Moreover, these proteins are also related to a recently identified family of cellular proteins, possibly involved in the control of cell proliferation. These observations have important implications for an understanding of the replication of these viruses and of picornavirus evolution. Methods Sequences. The sequences used in this work are in the EMBL database, under the following accession numbers: human parechovirus 1 (HPeV1), strain Harris (L02971); human parechovirus 2 (HPeV2), strain Williamson (AJ005695); human parechovirus 2, strain (Connecticut 86) (AF055846); AEV, Calnek vaccine strain (AJ225173); Aichi virus, strain A (AB010145); human H-rev107 (X92814); rat H-rev107 (X76453); lecithin retinol acyltransferase (LRAT) (AF071510); Tazarotene-induced gene protein (TIG3) (AF060228); Southampton virus (L07418); Norwalk virus (M87661); and Lordsdale virus (X86557). Sequence comparisons. Database searches were performed by using the EBI FASTA3 (http: www2.ebi.ac.uk fasta3 ) or BLAST2 (http: www2.ebi.ac.uk blast2 ) search facilities (Pearson & Lipman, 1988; Altschul et al., 1990). A range of FASTA3 parameters was used to analyse the HPeV1 sequence and a match with AEV was found by using the blosum50 matrix, Ktup 2, GapPenOpen 12 and GapPen- Extend 2. These parameters were used for all other searches. Clustal W (http: www2.ebi.ac.uk clustalw ) was used to align multiple sequences (Thompson et al., 1994). Sequence analysis. Sequences were analysed for the presence of likely transmembrane domains by using the program TMpred (http: software TMPRED form.html; Hofmann & Stoffel, 1993). The program ppsearch (http: www2.ebi.ac.uk ppsearch ) was used to perform PROSITE pattern searches. Results and Discussion It has been shown previously that HPeV1 (originally known as echovirus 22) is a typical picornavirus, but has a low degree of identity to most other members of this family (Hyypia et al., 1992; Stanway et al., 1994). Consequently, HPeV1 has been assigned as the prototype of a new genus, Parechovirus, which also includes the closely related HPeV2, two strains of which have now been sequenced (Ghazi et al., 1998; Oberste et al., 1998; King et al., 1999; Stanway & Hyypia, 1999). The 2A protein of parechoviruses was believed to lack homology to the corresponding proteins from other picornaviruses and its function is not clear. Previous attempts to identify related proteins, using FASTA3 or BLAST searches of protein databases, in order to gain insights into possible functions, yielded no significant matches (Hyypia et al., 1992; Ghazi et al., 1998). More extensive comparisons were therefore undertaken, employing a range of parameters to search the SWISS-PROT database with the HPeV1 2A sequence as the search string. These revealed a significant match with the corresponding region of the polyprotein of the recently sequenced picornavirus AEV (Marvil et al., 1999) (Fig. 1). The published sequence of AEV predicts that, in this virus, 2A is short and therefore the observed region of homology extends into the predicted 2B-encoding region. However, it is clear that the genomic position, immediately downstream of the capsid protein-encoding region, is the same in both cases. The protein sequences are relatively diverse, but there are two regions of clear similarity. One is centred on amino acid 24 (His) of the parechovirus 2A and the other is a Lys Asn Cys Glu Thr (KNCET) motif (positions in HPeV1 2A). Moreover, the latter motif is almost equidistant from a long (18 amino acids) hydrophobic domain in the AEV and HPeV1 proteins. In view of these similarities, FASTA3 and BLAST database searches were performed, with the homologous region from AEV as the search string. These revealed a significant match with a human protein, H-rev107, and its rat homologue (Hajnal et al., 1994; Sers et al., 1997; Husmann et al., 1998). Further searches with H-rev107 as the search string confirmed an already observed similarity to the Tazarotene-induced gene protein 3 (TIG3), also called retinoic acid-responder 3 protein (DiSepio et al., 1998), and identified a previously unreported match with the enzyme lecithin retinol acyltransferase (LRAT) (Ruiz et al., 1999). All these cellular proteins have a long, hydrophobic domain and an Asn Cys Glu (NCE) motif resembling the conserved KNCET motif of HPeV1 and AEV. In addition, they exhibit similarity to the virus proteins in the HPeV1 position 24 region. Finally, inspection of the 2A proteins of other picornaviruses indicated that the recently described Aichi virus, which constitutes a distinct genetic lineage among picornaviruses, also has these features, in a weaker form, but that they are absent from other members of this family, including hepatitis A virus, the closest relative of AEV (Yamashita et al., 1998; Marvil et al., 1999). It can be seen from the alignment of the homologous region of the three available HPeV sequences, AEV and Aichi virus, together with the cellular proteins, that there is conservation of all the sequence features described above (Fig. 1). In addition, the HPeV1 2 and Aichi 2A proteins are virtually co-n-terminal with H-rev107 and TIG3 and have a CAC
3 Picornavirus 2A homology to cellular proteins Fig. 1. Amino acid sequence alignment of the four cellular H-box/NC proteins and predicted 2A proteins of five picornaviruses. Residues seen in at least seven of the sequences are indicated by shading and the highly conserved H-box and NC regions are boxed. The transmembrane domain, predicted in all the sequences, is in white on black and the basic amino acid that follows this in most of the sequences is highlighted in dark grey. The sequence ALTXKAXXXKXXL, common to H-rev107 and human parechoviruses, is in bold. The two sequenced strains of HPeV2 are designated HPeV2W (Williamson) and HPeV2C [ (Connecticut/86)]. H-rev107r and H-rev107h are the rat and human homologues of this protein. The actual C-terminal boundary of the AEV 2A protein predicted here is not known. The level of significance of a match observed when comparing a sequence with a database can be assessed by the E value. This is the number of times the sequence is expected to match equally well as the observed match with a database of the same size containing random sequences. Close matches therefore have low E values. The E values of FASTA3-derived matches with the SWALL database for each protein subsequently introduced into this alignment were: AEV with TIG3 3 1, H-rev107h 4 3, HPeV1 4 3; HPeV1 with HPeV2C , HPeV2C 10 52, AEV 5 6; HPeV2W with HPeV2C , HPeV ; HPeV2C with HPeV , HPeV2W ; H-rev107r with H-rev107h , TIG , LRAT 9 9; H-rev107h with H-rev107r , TIG , LRAT 9 7; LRAT with TIG3 0 13, H-rev107r 6 1, H-rev107h 8 4. None of the other matches fell below the cut-off level of E 10, but there is a clear relationship between the proteins shown in the highlighted areas, which define the H-box/NC proteins. similar overall length. The HPeV1 position 24 region, hereafter termed the H-box, is seen in two forms among these proteins: HWA(I L) in human and rat H-rev107, TIG3 and Aichi virus 2A; and H(Y F)G(I V) in HPeV1 2 and AEV 2A, together with LRAT. In addition, although there were no other absolutely conserved amino acids, the nature of some flanking residues is conserved and a Gly Asp (GD) dipeptide is seen in most of the proteins. The NCE region is less well conserved in Aichi virus 2A and only Asn Cys (hereafter termed the NC motif) is seen. The residues flanking the Aichi virus H-box are also less closely related to those of the other proteins. However, even in this virus, the significance of the similarity is emphasized by the presence of the hydrophobic region downstream of the NC motif. Moreover, there is clear primary sequence identity between the Aichi virus 2A and all the cellular proteins in the NC motif region, where the common sequence NCXHFV is seen. The other characteristic feature of the virus and cellular proteins is the long (18 24 amino acids) hydrophobic region. According to the predictive program TMpred, this is highly likely to be a transmembrane domain (Hofmann & Stoffel, 1993). Deletion of this region has been shown to reduce the activity of H-rev107 severely, suggesting that it plays a key role in the function of this protein (Sers et al., 1997). In all the proteins, except LRAT, this putative transmembrane domain is followed immediately by a basic amino acid. A further notable similarity is that seen between the H-rev107 proteins and HPeV 2A, where the motif ALTXKAXXXKXXL (positions in HPeV1) can be seen in analogous positions. In terms of overall amino acid identity, the H-rev107 and TIG3 proteins are closely related, while the other cellular protein, LRAT, is relatively distant from these. This is also evident from the alignment, which shows that LRAT has a longer N-terminal region and two relative insertions compared with the other cellular proteins (Fig. 1). If these are removed from comparisons, LRAT clearly clusters with the cellular CAD
4 P. J. Hughes and G. Stanway Fig. 2. Dendrogram, based on amino acid identities, expressing the relationship between the cellular H-box/NC proteins and picornavirus 2A proteins. proteins, but remains a distant member (Fig. 2). Among the virus proteins, as expected, those from the human parechoviruses form a tight cluster. If, as seems likely, the sequence identities between these proteins are indicative of shared functional properties, the strong conservation of the H-box, NC motif and putative transmembrane domain, within relatively poorly conserved proteins, leads to the prediction that these are critical functional elements within this class of protein. These features, which do not correlate with those of any other known proteins when subjected to PROSITE analysis, are therefore interesting targets for future work on both the virus proteins and their cellular homologues. His and Cys are two of the three amino acids involved in the catalytic triad of the 3C-like proteases seen in several viruses, including the 2A protein of rhinoviruses and enteroviruses (Ryan & Flint, 1997). Moreover, the conserved His and Cys residues within the H-box NC proteins are similarly located to their counterparts in these proteases (Fig. 3). However, the amino acid sequence flanking the active site nucleophile of trypsin-like proteases (Cys in rhinovirus and enterovirus 2A) is characteristically Gly X Cys Gly (GXCG), and the H-box NC proteins do not conform to this pattern. His and Cys are also critical amino acids of papain-like proteases, such as the aphthovirus L, and of pestivirus p20 proteases, but once again there is no similarity of flanking amino acids in the H-box NC proteins (Ryan & Flint, 1997; Ryan et al., 1998). These observations suggest that the H-box NC proteins are not proteases of a known type; indeed, it has already been shown that the HPeV1 2A has no proteolytic activity directed to either its own N or C terminus (Schultheiss et al., 1995). The identification of LRAT, an enzyme responsible for the conversion of all-trans-retinol into retinyl esters, as a putative member of the H-box NC proteins Fig. 3. Protein 2A types among the six picornavirus genera, together with Aichi virus and AEV. Aichi virus has been proposed as the type member of a new picornavirus genus, while in other parts of the genome the closest relatives of AEV are the hepatoviruses. Members of two other proposed new picornavirus genera (ERBV and PTV1) are aphthovirus-like in terms of 2A (King et al., 1999). H, D and C refer to the catalytic triad of trypsin-like proteases, while Hbox, NC and tm (transmembrane domain) are the characteristic motifs of the H-box/NC proteins defined here. The hepatovirus 2A is of unknown function, while in addition to sequences associated with NPGP processing (shaded), the cardiovirus 2A has an additional region of unknown function. The actual C-terminal boundary of the AEV 2A protein predicted here is not known. further suggests a non-proteolytic role for the virus proteins (Ruiz et al., 1999). However, a proteolytic function cannot be ruled out. The function of the cellular H-box NC proteins may give a clue to the role of virus proteins in replication, but unfortunately there is little information on how these proteins exert an effect at the mechanistic level. This is compounded by the fact that the H-box and NC motifs have not been recognized previously and are not seen in any well-characterized proteins. It is known, however, that H-rev107 is downregulated in a number of tumour cells and that overexpression leads to inhibition of cell proliferation and resistance to transformation by the ras oncogene homologue (Hajnal et al., 1994; Sers et al., 1997; Husmann et al., 1998). This suggests that H-rev107 may act as a negative regulator of oncogenic ras signals, possibly by binding to and inhibiting an effector molecule (Husmann et al., 1998). Expression of TIG3, which is induced by retinoic acid, likewise reduces cell proliferation (DiSepio et al., 1998). Accordingly, it may contribute to the known inhibition of cell growth by retinoic acid. Both H- rev107 and TIG3 have been described as class II tumour CAE
5 Picornavirus 2A homology to cellular proteins Fig. 4. Alignment of part of the cellular and picornavirus H-box/NC proteins with part of the non-structural region (ORF1) of caliciviruses of genetic group II (Southampton, Norwalk and Lordsdale viruses). The inclusion of the caliciviruses in the alignment is based on the observation, by FASTA3 analysis of the SWALL database, of a match between H-rev107r and Norwalk virus (E 2 0). A subsequent search, using the region of Norwalk virus sequence shown in the figure, indicated, as expected, a close relationship with two other caliciviruses, Southampton virus (E ) and Lordsdale virus (E ). The whole non-structural region of the picornavirus HPeV1 is also aligned schematically with that of ORF1 of the calicivirus Lordsdale virus, in order to demonstrate the similar arrangement of non-structural proteins. suppressors. These are proteins that are expressed at very low levels in cell lines or tumours, permitting cell proliferation, even though their genes are intact (Husmann et al., 1998). As already discussed, LRAT was identified on the basis of enzymic activity, the conversion of all-trans-retinol into retinyl esters, which is an important pathway in the metabolism and storage of retinol (Ruiz et al., 1999). It appears to be unrelated to known retinol-binding proteins and to other acyltransferases, suggesting that it functions in a distinct manner. However, the association with retinoids may again suggest a role in the control of cell growth. If the virus H-box NC proteins have a similar effect on cell proliferation, this may be a mechanism, for example, for reducing host-cell macromolecular synthesis and may therefore serve as an alternative to the host-cell protein synthesis shut-off induced by some picornaviruses. On the other hand, the virus proteins may be involved in phenomena such as the regulation of apoptosis, as proteins capable of inducing or suppressing this process have been identified in several viruses. The implicit identification of their critical features, through the recognition of the H-box NC protein family, gives pointers to how these questions may be addressed. From its earliest isolation, it was observed that HPeV1 causes a distinctive cellular pathology, involving characteristic nuclear changes (Wigand & Sabin, 1961; Shaver et al., 1961; Jamison, 1974). The report that H-rev107 associates with the nuclear membrane, together with the relationships described here, raises the intriguing possibility that this pathology results from the action of the 2A protein (Sers et al., 1997). It is assumed that many virus proteins originated by acquisition from the host genome. In the case of picornaviruses, the structural similarity of 3C (and that of the related entero rhinovirus 2A protein) to cellular trypsin-like proteases is cited as an example (Gorbalenya, 1992). The papain-like, aphthovirus L protein may also have arisen from capture of a host RNA. The presence in coxsackievirus A9 of an Arg Gly Asp (RGD)-containing, C-terminal VP1 extension (relative to other enteroviruses), with similarity to transforming growth factor β1, is a more speculative example of a region of a picornavirus protein with a putative cellular origin (Chang et al., 1989, 1992; Hughes et al., 1995). Apart from these cases, the observation reported here, that some picornavirus 2A proteins have cellular homologues, is the clearest evidence in picornaviruses for such a mechanism. The demonstration that the 2A proteins of three picornaviruses are related reduces the apparent diversity of this locus and, as shown in Fig. 3, picornaviruses fall essentially into four CAF
6 P. J. Hughes and G. Stanway groups. These are viruses with: a trypsin-like protease (rhinoviruses enteroviruses); an NPGP motif involved in 2A 2B cleavage (cardioviruses aphthoviruses); a 2A unrelated to known proteins (hepatoviruses); or an H-box NC protein (parechoviruses Aichi virus AEV). Although the C terminus of the cardiovirus 2A is homologous to the aphthovirus protein, it has an additional 130 N-terminal amino acids. The occurrence of an H-box NC protein at the same locus in three diverse picornavirus groups suggests that it was acquired by a single event in a common ancestor and that it has been maintained through subsequent divergence. Since AEV is related to hepatoviruses, which lack a corresponding protein, it is likely that hepatoviruses lost this protein after divergence from the AEV lineage. This may have taken place through drift, or through deletion and subsequent capture of another proteinencoding sequence at this locus. In view of the distinct nature of the 2A proteins of AEV and hepatoviruses, it may be preferable to consider them as members of separate genera, notwithstanding the fact that they are more closely related to one another than to other picornaviruses in other parts of the genome (Marvil et al., 1999). Recently, a parechovirus-like virus has been isolated from rodents (Nicklasson et al., 1998, 1999). It will be interesting to ascertain whether this virus has an H-box NC protein and, if so, how closely related it is to that of HPeVs. Relatively diverse picornaviruses have H-box NC proteins and it is possible that they occur in other viruses. It was therefore interesting to find similar features in one genetic group of the caliciviruses (group II), containing Norwalk, Southampton and Lordsdale viruses, by comparison with H-rev107 sequences (Lambden et al., 1993; Dingle et al., 1995; Hardy & Estes, 1996). These viruses each have sequences reminiscent of the H-box and also possess an NC motif (Fig. 4). Caliciviruses have a similar relative arrangement of nonstructural proteins to picornaviruses, which can be summarized: NTP-binding protein (possibly helicase); 3C-like protease; polymerase (2C; 3C; 3D in picornaviruses). The calicivirus H-box NC protein is upstream of the NTP-binding protein and the relative positions of the motifs are similar to those of the picornavirus protein (Fig. 4). The calicivirus protein is distinguished from the picornavirus 2A by the absence of a significant transmembrane domain, suggesting that it may function in a different manner. However, it seems that H-box NC proteins may occur in several RNA viruses, suggesting that they may contribute an important, but as yet unknown function to the virus life-cycle. This work was supported by The Wellcome Trust, grant number We would like to thank Jane Stanway and Fiona Thompson for their invaluable assistance in the preparation of the manuscript. References Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. (1990). Basic local alignment search tool. Journal of Molecular Biology 215, Chang, K. H., Auvinen, P., Hyypia, T. & Stanway, G. (1989). The nucleotide sequence of coxsackievirus A9: implications for receptor binding and enterovirus classification. Journal of General Virology 70, Chang, K. H., Day, C., Walker, J., Hyypia, T. & Stanway, G. (1992). The nucleotide sequences of wild-type coxsackievirus A9 strains imply that an RGD motif in VP1 is functionally significant. Journal of General Virology 73, Dingle, K. E., Lambden, P. R., Caul, E. O. & Clarke, I. N. (1995). Human enteric Caliciviridae: the complete genome sequence and expression of virus-like particles from a genetic group II small round structured virus. Journal of General Virology 76, DiSepio, D., Ghosn, C., Eckert, R. L., Deucher, A., Robinson, N., Duvic, M., Chandraratna, R. A. S. & Nagpal, S. (1998). Identification and characterization of a retinoid-induced class II tumor suppressor growth regulatory gene. Proceedings of the National Academy of Sciences, USA 95, Doherty, M., Todd, D., McFerran, N. & Hoey, E. M. (1999). Sequence analysis of a porcine enterovirus serotype 1 isolate: relationships with other picornaviruses. Journal of General Virology 80, Ghazi, F., Hughes, P. J., Hyypia, T. & Stanway, G. (1998). Molecular analysis of human parechovirus type 2 (formerly echovirus 23). Journal of General Virology 79, Gorbalenya, A. E. (1992). Host-related sequences in RNA viral genomes. Seminars in Virology 3, Hajnal, A., Klemenz, R. & Schafer, R. (1994). Subtraction cloning of H-rev107, a gene specifically expressed in H-ras resistant fibroblasts. Oncogene 9, Hardy, M. E. & Estes, M. K. (1996). Completion of the Norwalk virus genome sequence. Virus Genes 12, Hofmann, K. & Stoffel, W. (1993). TMbase a database of membrane spanning protein segments. Biological Chemistry Hoppe-Seyler 374, 166. Hughes, P. J., Horsnell, C., Hyypia, T. & Stanway, G. (1995). The coxsackievirus A9 RGD motif is not essential for virus viability. Journal of Virology 69, Husmann, K., Sers, C., Fietze, E., Mincheva, A., Lichter, P. & Schafer, R. (1998). Transcriptional and translational downregulation of H- REV107, a class II tumour suppressor gene located on human chromosome 11q Oncogene 17, Hyypia, T., Horsnell, C., Maaronen, M., Khan, M., Kalkkinen, N., Auvinen, P., Kinnunen, L. & Stanway, G. (1992). A distinct picornavirus group identified by sequence analysis. Proceedings of the National Academy of Sciences, USA 89, Jamison, R. M. (1974). An electron microscopic study of the intracellular development of echovirus 22. Archiv fu r die gesamte Virusforschung 44, Jia, X. Y., Summers, D. F. & Ehrenfeld, E. (1993). Primary cleavage of the HAV capsid protein precursor in the middle of the proposed 2A coding region. Virology 193, King, A. M. Q., Brown, F., Christian, P., Hovi, T., Hyypia, T., Knowles, N. J., Lemon, S. M., Minor, P. D., Palmenberg, A. C., Skern, T. & Stanway, G. (1999). Picornaviridae. In Virus Taxonomy. Seventh Report of the International Committee on Taxonomy of Viruses. Edited by M. H. V. Van Regenmortel, C. M. Fauquet, D. H. L. Bishop, C. H. Calisher, E. B. Carsten, M. K. Estes, S. M. Lemon, J. Maniloff, M. A. Mayo, D. J. McGeoch, C. R. Pringle & R. B. Wickner. New York & San Diego: Academic Press (in press). Lambden, P. R., Caul, E. O., Ashley, C. R. & Clarke, I. N. (1993). Sequence and genome organization of a human small round-structured (Norwalk-like) virus. Science 259, CAG
7 Picornavirus 2A homology to cellular proteins Marvil, P., Knowles, N. J., Mockett, A. P. A., Britton, P., Brown, T. D. K. & Cavanagh, D. (1999). Avian encephalomyelitis virus is a picornavirus and is most closely related to hepatitis A virus. Journal of General Virology 80, Niklasson, B., Ho rnfeldt, B. & Lundman, B. (1998). Could myocarditis, insulin-dependent diabetes mellitus, and Guillain Barre syndrome be caused by one or more infectious agents carried by rodents? Emerging Infectious Diseases 4, Niklasson, B., Kinnunen, L., Ho rnfeldt, B., Ho rling, J., Benemar, C., Hedlund, K. O., Matskova, L., Hyypia, T. & Winberg, G. (1999). A new picornavirus isolated from bank voles (Clethrionomys glareolus). Virology 255, Oberste, M. S., Maher, K. & Pallansch, M. A. (1998). Complete sequence of echovirus 23 and its relationship to echovirus 22 and other human enteroviruses. Virus Research 56, Pearson, W. R. & Lipman, D. J. (1988). Improved tools for biological sequence comparison. Proceedings of the National Academy of Sciences, USA 85, Rueckert, R. R. (1996). Picornaviridae: the viruses and their replication. In Fields Virology, 3rd edn, pp Edited by B. N. Fields, D. M. Knipe & P. M. Howley. Philadelphia: Lippincott Raven. Ruiz, A., Winston, A., Lim, Y.-H., Gilbert, B. A., Rando, R. R. & Bok, D. (1999). Molecular and biochemical characterization of lecithin retinol acyltransferase. Journal of Biological Chemistry 274, Ryan, M. D. & Flint, M. (1997). Virus-encoded proteinases of the picornavirus super-group. Journal of General Virology 78, Ryan, M. D., Monaghan, S. & Flint, M. (1998). Virus-encoded proteinases of the Flaviviridae. Journal of General Virology 79, Schultheiss, T., Emerson, S. U., Purcell, R. H. & Gauss-Muller, V. (1995). Polyprotein processing in echovirus 22: a first assessment. Biochemical and Biophysical Research Communications 217, Sers, C., Emmenegger, U., Husmann, K., Bucher, K., Andres, A. C. & Schafer, R. (1997). Growth-inhibitory activity and downregulation of the class II tumor-suppressor gene H-rev107 in tumor cell lines and experimental tumors. Journal of Cell Biology 136, Shaver, D. N., Barron, A. L. & Karzon, D. T. (1961). Distinctive cytopathology of ECHO viruses types 22 and 23. Proceedings of the Society for Experimental Biology and Medicine 106, Stanway, G. (1990). Structure, function and evolution of picornaviruses. Journal of General Virology 71, Stanway, G. & Hyypia, T. (1999). Parechoviruses. Journal of Virology 73, Stanway, G., Kalkkinen, N., Roivainen, M., Ghazi, F., Khan, M., Smyth, M., Meurman, O. & Hyypia, T. (1994). Molecular and biological characteristics of echovirus 22, a representative of a new picornavirus group. Journal of Virology 68, Thompson, J. D., Higgins, D. G. & Gibson, T. J. (1994). CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Research 22, Wigand, R. & Sabin, A. B. (1961). Properties of ECHO types 22, 23 and 24 viruses. Archiv fu r die gesamte Virusforschung 11, Wutz, G., Auer, H., Nowotny, N., Grosse, B., Skern, T. & Kuechler, E. (1996). Equine rhinovirus serotypes 1 and 2: relationship to each other and to aphthoviruses and cardioviruses. Journal of General Virology 77, Yamashita, T., Sakae, K., Tsuzuki, H., Suzuki, Y., Ishikawa, N., Takeda, N., Miyamura, T. & Yamazaki, S. (1998). Complete nucleotide sequence and genetic organization of Aichi virus, a distinct member of the Picornaviridae associated with acute gastroenteritis in humans. Journal of Virology 72, Received 1 June 1999; Accepted 20 September 1999 CAH
Picornaviruses. Virion. Genome. Genes and proteins. Viruses and hosts. Diseases. Distinctive characteristics
 Picornaviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Naked icosahedral capsid (T=1) Diameter of 30 nm Genome Linear single-stranded RNA, positive
Picornaviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Naked icosahedral capsid (T=1) Diameter of 30 nm Genome Linear single-stranded RNA, positive
ICTVdB Virus Descriptions
 [Home] [Index of Viruses ] [Descriptions] [ Character List ] [ Picture Gallery ] [ Interactive Key ] [ Data Entry] [ 2002 ICTV] ICTVdB Virus Descriptions Descriptions are generated automatically from the
[Home] [Index of Viruses ] [Descriptions] [ Character List ] [ Picture Gallery ] [ Interactive Key ] [ Data Entry] [ 2002 ICTV] ICTVdB Virus Descriptions Descriptions are generated automatically from the
Human parechovirus type 1, 3, 4, 5 and 6 detection in picornavirus cultures ACCEPTED. Running title: HPeV type 1, 3, 4, 5 and 6 in virus cultures
 JCM Accepts, published online ahead of print on 12 December 2007 J. Clin. Microbiol. doi:10.1128/jcm.02009-07 Copyright 2007, American Society for Microbiology and/or the Listed Authors/Institutions. All
JCM Accepts, published online ahead of print on 12 December 2007 J. Clin. Microbiol. doi:10.1128/jcm.02009-07 Copyright 2007, American Society for Microbiology and/or the Listed Authors/Institutions. All
Molecular analysis of duck hepatitis virus type 1 reveals a novel lineage close to the genus Parechovirus in the family Picornaviridae
 Journal of General Virology (2006), 87, 3307 3316 DOI 10.1099/vir.0.81804-0 Molecular analysis of duck hepatitis virus type 1 reveals a novel lineage close to the genus Parechovirus in the family Picornaviridae
Journal of General Virology (2006), 87, 3307 3316 DOI 10.1099/vir.0.81804-0 Molecular analysis of duck hepatitis virus type 1 reveals a novel lineage close to the genus Parechovirus in the family Picornaviridae
Susanne Johansson, 1 Bo Niklasson, 2 Jacob Maizel, 3 Alexander E. Gorbalenya, 4 * and A. Michael Lindberg 1 *
 JOURNAL OF VIROLOGY, Sept. 2002, p. 8920 8930 Vol. 76, No. 17 0022-538X/02/$04.00 0 DOI: 10.1128/JVI.76.17.8920 8930.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved. Molecular
JOURNAL OF VIROLOGY, Sept. 2002, p. 8920 8930 Vol. 76, No. 17 0022-538X/02/$04.00 0 DOI: 10.1128/JVI.76.17.8920 8930.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved. Molecular
Sequences in the 5" Non-coding Region of Human Rhinovims 14 RNA that Affect in vitro Translation
 J. gen. Virol. (1989), 70, 2799-2804. Printed in Great Britain 2799 Key words: rhinovirus, human type 14/5' non-coding region~in vitro translation Sequences in the 5" Non-coding Region of Human Rhinovims
J. gen. Virol. (1989), 70, 2799-2804. Printed in Great Britain 2799 Key words: rhinovirus, human type 14/5' non-coding region~in vitro translation Sequences in the 5" Non-coding Region of Human Rhinovims
Sequencing of Porcine Enterovirus Groups II and III Reveals Unique Features of Both Virus Groups
 REFERENCES CONTENT ALERTS Sequencing of Porcine Enterovirus Groups II and III Reveals Unique Features of Both Virus Groups Andi Krumbholz, Malte Dauber, Andreas Henke, Eckhard Birch-Hirschfeld, Nick J.
REFERENCES CONTENT ALERTS Sequencing of Porcine Enterovirus Groups II and III Reveals Unique Features of Both Virus Groups Andi Krumbholz, Malte Dauber, Andreas Henke, Eckhard Birch-Hirschfeld, Nick J.
Review article. Structure, function and evolution of picornaviruses. Glyn Stanway. Introduction. Structure of picornaviruses
 Journal of General Virology (1990), 71, 2483-2501. Printed in Great Britain 2483 Review article Structure, function and evolution of picornaviruses Glyn Stanway Department of Biology, University of Essex,
Journal of General Virology (1990), 71, 2483-2501. Printed in Great Britain 2483 Review article Structure, function and evolution of picornaviruses Glyn Stanway Department of Biology, University of Essex,
number Done by Corrected by Doctor Ashraf
 number 4 Done by Nedaa Bani Ata Corrected by Rama Nada Doctor Ashraf Genome replication and gene expression Remember the steps of viral replication from the last lecture: Attachment, Adsorption, Penetration,
number 4 Done by Nedaa Bani Ata Corrected by Rama Nada Doctor Ashraf Genome replication and gene expression Remember the steps of viral replication from the last lecture: Attachment, Adsorption, Penetration,
Coronaviruses. Virion. Genome. Genes and proteins. Viruses and hosts. Diseases. Distinctive characteristics
 Coronaviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Spherical enveloped particles studded with clubbed spikes Diameter 120-160 nm Coiled helical
Coronaviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Spherical enveloped particles studded with clubbed spikes Diameter 120-160 nm Coiled helical
HAV HBV HCV HDV HEV HGV
 Viral Hepatitis HAV HBV HCV HDV HEV HGV Additional well-characterized viruses that can cause sporadic hepatitis, such as yellow fever virus, cytomegalovirus, Epstein-Barr virus, herpes simplex virus, rubella
Viral Hepatitis HAV HBV HCV HDV HEV HGV Additional well-characterized viruses that can cause sporadic hepatitis, such as yellow fever virus, cytomegalovirus, Epstein-Barr virus, herpes simplex virus, rubella
Translation. Host Cell Shutoff 1) Initiation of eukaryotic translation involves many initiation factors
 Translation Questions? 1) How does poliovirus shutoff eukaryotic translation? 2) If eukaryotic messages are not translated how can poliovirus get its message translated? Host Cell Shutoff 1) Initiation
Translation Questions? 1) How does poliovirus shutoff eukaryotic translation? 2) If eukaryotic messages are not translated how can poliovirus get its message translated? Host Cell Shutoff 1) Initiation
on June 19, 2018 by guest
 JOURNAL OF VIROLOGY, Sept. 2001, p. 8021 8030 Vol. 75, No. 17 0022-538X/01/$04.00 0 DOI: 10.1128/JVI.75.17.8021 8030.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved. Construction
JOURNAL OF VIROLOGY, Sept. 2001, p. 8021 8030 Vol. 75, No. 17 0022-538X/01/$04.00 0 DOI: 10.1128/JVI.75.17.8021 8030.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved. Construction
Viral Taxonomic Classification
 Viruses Part I Viral Taxonomic Classification Order>> -virales Family>> - viridae Subfamily>> -virinae Genus>> -virus Species Order>> Picornavirales Family>> Picornaviridae Subfamily>> Picornavirinae Genus>>
Viruses Part I Viral Taxonomic Classification Order>> -virales Family>> - viridae Subfamily>> -virinae Genus>> -virus Species Order>> Picornavirales Family>> Picornaviridae Subfamily>> Picornavirinae Genus>>
aV. Code assigned:
 This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the
Peptide hydrolysis uncatalyzed half-life = ~450 years HIV protease-catalyzed half-life = ~3 seconds
 Uncatalyzed half-life Peptide hydrolysis uncatalyzed half-life = ~450 years IV protease-catalyzed half-life = ~3 seconds Life Sciences 1a Lecture Slides Set 9 Fall 2006-2007 Prof. David R. Liu In the absence
Uncatalyzed half-life Peptide hydrolysis uncatalyzed half-life = ~450 years IV protease-catalyzed half-life = ~3 seconds Life Sciences 1a Lecture Slides Set 9 Fall 2006-2007 Prof. David R. Liu In the absence
Foot-and-Mouth Disease
 CLINICAL MICROBIOLOGY REVIEWS, Apr. 2004, p. 465 493 Vol. 17, No. 2 0893-8512/04/$08.00 0 DOI: 10.1128/CMR.17.2.465 493.2004 Foot-and-Mouth Disease Marvin J. Grubman* and Barry Baxt Plum Island Animal
CLINICAL MICROBIOLOGY REVIEWS, Apr. 2004, p. 465 493 Vol. 17, No. 2 0893-8512/04/$08.00 0 DOI: 10.1128/CMR.17.2.465 493.2004 Foot-and-Mouth Disease Marvin J. Grubman* and Barry Baxt Plum Island Animal
numbe r Done by Corrected by Doctor
 numbe r 5 Done by Mustafa Khader Corrected by Mahdi Sharawi Doctor Ashraf Khasawneh Viral Replication Mechanisms: (Protein Synthesis) 1. Monocistronic Method: All human cells practice the monocistronic
numbe r 5 Done by Mustafa Khader Corrected by Mahdi Sharawi Doctor Ashraf Khasawneh Viral Replication Mechanisms: (Protein Synthesis) 1. Monocistronic Method: All human cells practice the monocistronic
Replication Defective Enterovirus Infections: Implications for Type I Diabetes
 Replication Defective Enterovirus Infections: Implications for Type I Diabetes N. M. Chapman Department of Pathology & Microbiology University of Nebraska Medical Center Enterovirus Genome and 2 Capsid
Replication Defective Enterovirus Infections: Implications for Type I Diabetes N. M. Chapman Department of Pathology & Microbiology University of Nebraska Medical Center Enterovirus Genome and 2 Capsid
Polyomaviridae. Spring
 Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
Investigation of the subcellular localization of HPeV 3A and 2A proteins.
 Biomedical Research 2017; 28 (20): 8639-8648 ISSN 0970-938X www.biomedres.info Investigation of the subcellular localization of HPeV 3A and 2A proteins. Arsalan Salimi *, Siamak Amirfakhri School of Biological
Biomedical Research 2017; 28 (20): 8639-8648 ISSN 0970-938X www.biomedres.info Investigation of the subcellular localization of HPeV 3A and 2A proteins. Arsalan Salimi *, Siamak Amirfakhri School of Biological
Evidence for enteroviral persistence in humans
 Journal of General Virology (1997), 78, 307 312. Printed in Great Britain...... SHORT COMMUNICATION Evidence for enteroviral persistence in humans Daniel N. Galbraith, Carron Nairn and Geoffrey B. Clements
Journal of General Virology (1997), 78, 307 312. Printed in Great Britain...... SHORT COMMUNICATION Evidence for enteroviral persistence in humans Daniel N. Galbraith, Carron Nairn and Geoffrey B. Clements
Diversity of human parechoviruses isolated in stool samples collected from. Thai children with acute gastroenteritis
 JCM Accepts, published online ahead of print on 28 October 2009 J. Clin. Microbiol. doi:10.1128/jcm.01015-09 Copyright 2009, American Society for Microbiology and/or the Listed Authors/Institutions. All
JCM Accepts, published online ahead of print on 28 October 2009 J. Clin. Microbiol. doi:10.1128/jcm.01015-09 Copyright 2009, American Society for Microbiology and/or the Listed Authors/Institutions. All
Polymerase Chain Reaction for Human Picornaviruses
 J. gen. Virol. (1989), 70, 3261-3268. Printed in Great Britain 3261 Key words: PCRihuman picornaviruses/hybridization Polymerase Chain Reaction for Human Picornaviruses By TIMO HYYPIA,* PETRI AUVINEN AND
J. gen. Virol. (1989), 70, 3261-3268. Printed in Great Britain 3261 Key words: PCRihuman picornaviruses/hybridization Polymerase Chain Reaction for Human Picornaviruses By TIMO HYYPIA,* PETRI AUVINEN AND
Severe Acute Respiratory Syndrome (SARS) Coronavirus
 Severe Acute Respiratory Syndrome (SARS) Coronavirus Coronaviruses Coronaviruses are single stranded enveloped RNA viruses that have a helical geometry. Coronaviruses are the largest of RNA viruses with
Severe Acute Respiratory Syndrome (SARS) Coronavirus Coronaviruses Coronaviruses are single stranded enveloped RNA viruses that have a helical geometry. Coronaviruses are the largest of RNA viruses with
Detection of All Known Parechoviruses by Real-Time PCR
 JOURNAL OF CLINICAL MICROBIOLOGY, Aug. 2008, p. 2519 2524 Vol. 46, No. 8 0095-1137/08/$08.00 0 doi:1128/jcm.00277-08 Copyright 2008, American Society for Microbiology. All Rights Reserved. Detection of
JOURNAL OF CLINICAL MICROBIOLOGY, Aug. 2008, p. 2519 2524 Vol. 46, No. 8 0095-1137/08/$08.00 0 doi:1128/jcm.00277-08 Copyright 2008, American Society for Microbiology. All Rights Reserved. Detection of
Vincent Racaniello
 Vincent Racaniello vrr1@columbia.edu www.virology.ws Poliomyelitis Polio (grey), myelon (marrow) = Greek itis (inflammation of) = Latin A common, acute viral disease characterized clinically by a brief
Vincent Racaniello vrr1@columbia.edu www.virology.ws Poliomyelitis Polio (grey), myelon (marrow) = Greek itis (inflammation of) = Latin A common, acute viral disease characterized clinically by a brief
ALIGNMENTS AND COMPARATIVE PROFILESOF PICORNAVIRUS GENERA
 In: Molecular Biology of Picornaviruses. Eds: B. Semler & E. Wimmer, ASM Press, pp 149-155 (2002) ALIGNMENTS AND COMPARATIVE PROFILESOF PICORNAVIRUS GENERA Ann C. Palmenberg* and Jean-Yves Sgro Insti.
In: Molecular Biology of Picornaviruses. Eds: B. Semler & E. Wimmer, ASM Press, pp 149-155 (2002) ALIGNMENTS AND COMPARATIVE PROFILESOF PICORNAVIRUS GENERA Ann C. Palmenberg* and Jean-Yves Sgro Insti.
Rajesh Kannangai Phone: ; Fax: ; *Corresponding author
 Amino acid sequence divergence of Tat protein (exon1) of subtype B and C HIV-1 strains: Does it have implications for vaccine development? Abraham Joseph Kandathil 1, Rajesh Kannangai 1, *, Oriapadickal
Amino acid sequence divergence of Tat protein (exon1) of subtype B and C HIV-1 strains: Does it have implications for vaccine development? Abraham Joseph Kandathil 1, Rajesh Kannangai 1, *, Oriapadickal
the world and viruses
 More than 5,450 viruses belonging to more than 2,000 species, 287 genera, 73 families and 3 orders are recognized in the 8th ICTVreport report. the world and viruses 1 1889 H2N2 Emerging viruses in the
More than 5,450 viruses belonging to more than 2,000 species, 287 genera, 73 families and 3 orders are recognized in the 8th ICTVreport report. the world and viruses 1 1889 H2N2 Emerging viruses in the
Recombination and Selection in the Evolution of Picornaviruses and Other Mammalian Positive-Stranded RNA Viruses
 JOURNAL OF VIROLOGY, Nov. 2006, p. 11124 11140 Vol. 80, No. 22 0022-538X/06/$08.00 0 doi:10.1128/jvi.01076-06 Copyright 2006, American Society for Microbiology. All Rights Reserved. Recombination and Selection
JOURNAL OF VIROLOGY, Nov. 2006, p. 11124 11140 Vol. 80, No. 22 0022-538X/06/$08.00 0 doi:10.1128/jvi.01076-06 Copyright 2006, American Society for Microbiology. All Rights Reserved. Recombination and Selection
Fayth K. Yoshimura, Ph.D. September 7, of 7 RETROVIRUSES. 2. HTLV-II causes hairy T-cell leukemia
 1 of 7 I. Diseases Caused by Retroviruses RETROVIRUSES A. Human retroviruses that cause cancers 1. HTLV-I causes adult T-cell leukemia and tropical spastic paraparesis 2. HTLV-II causes hairy T-cell leukemia
1 of 7 I. Diseases Caused by Retroviruses RETROVIRUSES A. Human retroviruses that cause cancers 1. HTLV-I causes adult T-cell leukemia and tropical spastic paraparesis 2. HTLV-II causes hairy T-cell leukemia
Translational Control of Viral Gene Expression in Eukaryotes
 MICROBIOLOGY AND MOLECULAR BIOLOGY REVIEWS, June 2000, p. 239 280 Vol. 64, No. 2 1092-2172/00/$04.00 0 Copyright 2000, American Society for Microbiology. All Rights Reserved. Translational Control of Viral
MICROBIOLOGY AND MOLECULAR BIOLOGY REVIEWS, June 2000, p. 239 280 Vol. 64, No. 2 1092-2172/00/$04.00 0 Copyright 2000, American Society for Microbiology. All Rights Reserved. Translational Control of Viral
What causes cancer? Physical factors (radiation, ionization) Chemical factors (carcinogens) Biological factors (virus, bacteria, parasite)
 Oncogenes What causes cancer? Chemical factors (carcinogens) Physical factors (radiation, ionization) Biological factors (virus, bacteria, parasite) DNA Mutation or damage Oncogenes Tumor suppressor genes
Oncogenes What causes cancer? Chemical factors (carcinogens) Physical factors (radiation, ionization) Biological factors (virus, bacteria, parasite) DNA Mutation or damage Oncogenes Tumor suppressor genes
Prevalence of human parechovirus in Chinese children hospitalized for acute gastroenteritis
 ORIGINAL ARTICLE VIROLOGY Prevalence of human parechovirus in Chinese children hospitalized for acute gastroenteritis D.-L. Zhang 1,2, Y. Jin 3, D.-D. Li 1, W.-X. Cheng 1,3, Z.-Q. Xu 1, J.-M. Yu 1, M.
ORIGINAL ARTICLE VIROLOGY Prevalence of human parechovirus in Chinese children hospitalized for acute gastroenteritis D.-L. Zhang 1,2, Y. Jin 3, D.-D. Li 1, W.-X. Cheng 1,3, Z.-Q. Xu 1, J.-M. Yu 1, M.
Overview of virus life cycle
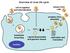 Overview of virus life cycle cell recognition and internalization release from cells progeny virus assembly membrane breaching nucleus capsid disassembly and genome release replication and translation
Overview of virus life cycle cell recognition and internalization release from cells progeny virus assembly membrane breaching nucleus capsid disassembly and genome release replication and translation
Cahn - Ingold - Prelog system. Proteins: Evolution, and Analysis Lecture 7 9/15/2009. The Fischer Convention (1) G (2) (3)
 Chapter 4 (1) G Proteins: Evolution, and Analysis Lecture 7 9/15/2009 A V L I M P F W Chapter 4 (2) S (3) T N Q Y C K R H D E The Fischer Convention Absolute configuration about an asymmetric carbon related
Chapter 4 (1) G Proteins: Evolution, and Analysis Lecture 7 9/15/2009 A V L I M P F W Chapter 4 (2) S (3) T N Q Y C K R H D E The Fischer Convention Absolute configuration about an asymmetric carbon related
Human Rhinovirus 87 and Enterovirus 68 Represent a Unique Serotype with Rhinovirus and Enterovirus Features
 JOURNAL OF CLINICAL MICROBIOLOGY, Nov. 2002, p. 4218 4223 Vol. 40, No. 11 0095-1137/02/$04.00 0 DOI: 10.1128/JCM.40.11.4218 4223.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved.
JOURNAL OF CLINICAL MICROBIOLOGY, Nov. 2002, p. 4218 4223 Vol. 40, No. 11 0095-1137/02/$04.00 0 DOI: 10.1128/JCM.40.11.4218 4223.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved.
Discovery of a Novel Murine Type C Retrovirus by Data Mining
 JOURNAL OF VIROLOGY, Mar. 2001, p. 3053 3057 Vol. 75, No. 6 0022-538X/01/$04.00 0 DOI: 10.1128/JVI.75.6.3053 3057.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved. Discovery
JOURNAL OF VIROLOGY, Mar. 2001, p. 3053 3057 Vol. 75, No. 6 0022-538X/01/$04.00 0 DOI: 10.1128/JVI.75.6.3053 3057.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved. Discovery
Last time we talked about the few steps in viral replication cycle and the un-coating stage:
 Zeina Al-Momani Last time we talked about the few steps in viral replication cycle and the un-coating stage: Un-coating: is a general term for the events which occur after penetration, we talked about
Zeina Al-Momani Last time we talked about the few steps in viral replication cycle and the un-coating stage: Un-coating: is a general term for the events which occur after penetration, we talked about
Reoviruses. Virion. Genome. Genes and proteins. Viruses and hosts. Diseases. Distinctive characteristics
 Reoviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Naked icosahedral capsid (T=13), diameter 60-85 nm Capsid consists of two or three concentric protein
Reoviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Naked icosahedral capsid (T=13), diameter 60-85 nm Capsid consists of two or three concentric protein
Objective: You will be able to explain how the subcomponents of
 Objective: You will be able to explain how the subcomponents of nucleic acids determine the properties of that polymer. Do Now: Read the first two paragraphs from enduring understanding 4.A Essential knowledge:
Objective: You will be able to explain how the subcomponents of nucleic acids determine the properties of that polymer. Do Now: Read the first two paragraphs from enduring understanding 4.A Essential knowledge:
Biological systems interact, and these systems and their interactions possess complex properties. STOP at enduring understanding 4A
 Biological systems interact, and these systems and their interactions possess complex properties. STOP at enduring understanding 4A Homework Watch the Bozeman video called, Biological Molecules Objective:
Biological systems interact, and these systems and their interactions possess complex properties. STOP at enduring understanding 4A Homework Watch the Bozeman video called, Biological Molecules Objective:
Introduction retroposon
 17.1 - Introduction A retrovirus is an RNA virus able to convert its sequence into DNA by reverse transcription A retroposon (retrotransposon) is a transposon that mobilizes via an RNA form; the DNA element
17.1 - Introduction A retrovirus is an RNA virus able to convert its sequence into DNA by reverse transcription A retroposon (retrotransposon) is a transposon that mobilizes via an RNA form; the DNA element
Fayth K. Yoshimura, Ph.D. September 7, of 7 HIV - BASIC PROPERTIES
 1 of 7 I. Viral Origin. A. Retrovirus - animal lentiviruses. HIV - BASIC PROPERTIES 1. HIV is a member of the Retrovirus family and more specifically it is a member of the Lentivirus genus of this family.
1 of 7 I. Viral Origin. A. Retrovirus - animal lentiviruses. HIV - BASIC PROPERTIES 1. HIV is a member of the Retrovirus family and more specifically it is a member of the Lentivirus genus of this family.
Fine Mapping of a cis-acting Sequence Element in Yellow Fever Virus RNA That Is Required for RNA Replication and Cyclization
 JOURNAL OF VIROLOGY, Feb. 2003, p. 2265 2270 Vol. 77, No. 3 0022-538X/03/$08.00 0 DOI: 10.1128/JVI.77.3.2265 2270.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved. Fine Mapping
JOURNAL OF VIROLOGY, Feb. 2003, p. 2265 2270 Vol. 77, No. 3 0022-538X/03/$08.00 0 DOI: 10.1128/JVI.77.3.2265 2270.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved. Fine Mapping
Complete Nucleotide Sequence of RNA1 of Cucumber Mosaic Virus Y Strain and Evolutionary Relationships among Genome RNAs of the Virus Strains
 Complete Nucleotide Sequence of RNA1 of Cucumber Mosaic Virus Y Strain and Evolutionary Relationships among Genome RNAs of the Virus Strains Jiro KATAOKA*, Chikara MASUTA* and Yoichi TAKANAMI* Abstract
Complete Nucleotide Sequence of RNA1 of Cucumber Mosaic Virus Y Strain and Evolutionary Relationships among Genome RNAs of the Virus Strains Jiro KATAOKA*, Chikara MASUTA* and Yoichi TAKANAMI* Abstract
CHARACTERIZATION OF VIRAL PROTEASES FROM NORWALK VIRUS, POLIOVIRUS, AND TRANSMISSIBLE GASTROENTERITIS VIRUS USING A
 CHARACTERIZATION OF VIRAL PROTEASES FROM NORWALK VIRUS, POLIOVIRUS, AND TRANSMISSIBLE GASTROENTERITIS VIRUS USING A FLUORESCENCE RESONANCE ENERGY TRANSFER ASSAY by VENKATA KIRAN PASUPULLETI B.S., ANDHRA
CHARACTERIZATION OF VIRAL PROTEASES FROM NORWALK VIRUS, POLIOVIRUS, AND TRANSMISSIBLE GASTROENTERITIS VIRUS USING A FLUORESCENCE RESONANCE ENERGY TRANSFER ASSAY by VENKATA KIRAN PASUPULLETI B.S., ANDHRA
Determination of the temporal pattern and importance of BALF1 expression in Epstein-Barr viral infection
 Determination of the temporal pattern and importance of BALF1 expression in Epstein-Barr viral infection Melissa Mihelidakis May 6, 2004 7.340 Research Proposal Introduction Apoptosis, or programmed cell
Determination of the temporal pattern and importance of BALF1 expression in Epstein-Barr viral infection Melissa Mihelidakis May 6, 2004 7.340 Research Proposal Introduction Apoptosis, or programmed cell
Amino acids. Side chain. -Carbon atom. Carboxyl group. Amino group
 PROTEINS Amino acids Side chain -Carbon atom Amino group Carboxyl group Amino acids Primary structure Amino acid monomers Peptide bond Peptide bond Amino group Carboxyl group Peptide bond N-terminal (
PROTEINS Amino acids Side chain -Carbon atom Amino group Carboxyl group Amino acids Primary structure Amino acid monomers Peptide bond Peptide bond Amino group Carboxyl group Peptide bond N-terminal (
Mutants and HBV vaccination. Dr. Ulus Salih Akarca Ege University, Izmir, Turkey
 Mutants and HBV vaccination Dr. Ulus Salih Akarca Ege University, Izmir, Turkey Geographic Distribution of Chronic HBV Infection 400 million people are carrier of HBV Leading cause of cirrhosis and HCC
Mutants and HBV vaccination Dr. Ulus Salih Akarca Ege University, Izmir, Turkey Geographic Distribution of Chronic HBV Infection 400 million people are carrier of HBV Leading cause of cirrhosis and HCC
Virus-encoded proteinases of the picornavirus super-group
 Journal of General Virology (1997), 78, 699 723. Printed in Great Britain...... Virus-encoded proteinases of the picornavirus super-group Martin D. Ryan and Mike Flint Department of Biochemistry, University
Journal of General Virology (1997), 78, 699 723. Printed in Great Britain...... Virus-encoded proteinases of the picornavirus super-group Martin D. Ryan and Mike Flint Department of Biochemistry, University
Introduction: RNA viruses
 SECTION I Introduction: RNA viruses Carol Shoshkes Reiss Viruses that infect the central nervous system may cause acute, chronic, or latent infections. In some cases, the diseases manifested are attributable
SECTION I Introduction: RNA viruses Carol Shoshkes Reiss Viruses that infect the central nervous system may cause acute, chronic, or latent infections. In some cases, the diseases manifested are attributable
Complete Genomic Sequencing Shows that Polioviruses and Members of Human Enterovirus Species C Are Closely Related in the Noncapsid Coding Region
 JOURNAL OF VIROLOGY, Aug. 2003, p. 8973 8984 Vol. 77, No. 16 0022-538X/03/$08.00 0 DOI: 10.1128/JVI.77.16.8973 8984.2003 Complete Genomic Sequencing Shows that Polioviruses and Members of Human Enterovirus
JOURNAL OF VIROLOGY, Aug. 2003, p. 8973 8984 Vol. 77, No. 16 0022-538X/03/$08.00 0 DOI: 10.1128/JVI.77.16.8973 8984.2003 Complete Genomic Sequencing Shows that Polioviruses and Members of Human Enterovirus
Influenza viruses. Virion. Genome. Genes and proteins. Viruses and hosts. Diseases. Distinctive characteristics
 Influenza viruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Enveloped particles, quasi-spherical or filamentous Diameter 80-120 nm Envelope is derived
Influenza viruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Enveloped particles, quasi-spherical or filamentous Diameter 80-120 nm Envelope is derived
Norwalk Virus N-Terminal Nonstructural Protein Is Associated with Disassembly of the Golgi Complex in Transfected Cells
 JOURNAL OF VIROLOGY, May 2004, p. 4827 4837 Vol. 78, No. 9 0022-538X/04/$08.00 0 DOI: 10.1128/JVI.78.9.4827 4837.2004 Norwalk Virus N-Terminal Nonstructural Protein Is Associated with Disassembly of the
JOURNAL OF VIROLOGY, May 2004, p. 4827 4837 Vol. 78, No. 9 0022-538X/04/$08.00 0 DOI: 10.1128/JVI.78.9.4827 4837.2004 Norwalk Virus N-Terminal Nonstructural Protein Is Associated with Disassembly of the
Lecture 2: Virology. I. Background
 Lecture 2: Virology I. Background A. Properties 1. Simple biological systems a. Aggregates of nucleic acids and protein 2. Non-living a. Cannot reproduce or carry out metabolic activities outside of a
Lecture 2: Virology I. Background A. Properties 1. Simple biological systems a. Aggregates of nucleic acids and protein 2. Non-living a. Cannot reproduce or carry out metabolic activities outside of a
R E V I E W SUMMARY. Reviews in Medical Virology
 Reviews in Medical Virology R E V I E W Rev. Med. Virol. 2010; 20: 327 337. Published online 14 July 2010 in Wiley Online Library (wileyonlinelibrary.com)..660 Recombination among picornaviruses A.N. Lukashev
Reviews in Medical Virology R E V I E W Rev. Med. Virol. 2010; 20: 327 337. Published online 14 July 2010 in Wiley Online Library (wileyonlinelibrary.com)..660 Recombination among picornaviruses A.N. Lukashev
Biol115 The Thread of Life"
 Biol115 The Thread of Life" Lecture 9" Gene expression and the Central Dogma"... once (sequential) information has passed into protein it cannot get out again. " ~Francis Crick, 1958! Principles of Biology
Biol115 The Thread of Life" Lecture 9" Gene expression and the Central Dogma"... once (sequential) information has passed into protein it cannot get out again. " ~Francis Crick, 1958! Principles of Biology
A highly divergent picornavirus in a marine mammal. ACCEPTED
 JVI Accepts, published online ahead of print on 17 October 2007 J. Virol. doi:10.1128/jvi.01240-07 Copyright 2007, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved.
JVI Accepts, published online ahead of print on 17 October 2007 J. Virol. doi:10.1128/jvi.01240-07 Copyright 2007, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved.
Identification and characterization of novel picornaviruses by molecular biological methods. Ph.D. Thesis. Péter Pankovics
 Identification and characterization of novel picornaviruses by molecular biological methods Ph.D. Thesis Péter Pankovics Supervisor: 1 Gábor Reuter, M.D., Ph.D., Med. Habil. Program leader: 2 Levente Emődy,
Identification and characterization of novel picornaviruses by molecular biological methods Ph.D. Thesis Péter Pankovics Supervisor: 1 Gábor Reuter, M.D., Ph.D., Med. Habil. Program leader: 2 Levente Emődy,
Viruses. CLS 212: Medical Microbiology Miss Zeina Alkudmani
 Viruses CLS 212: Medical Microbiology Miss Zeina Alkudmani History Through the 1800s, many scientists discovered that something smaller than bacteria could cause disease and they called it virion (Latin
Viruses CLS 212: Medical Microbiology Miss Zeina Alkudmani History Through the 1800s, many scientists discovered that something smaller than bacteria could cause disease and they called it virion (Latin
Coronaviruses cause acute, mild upper respiratory infection (common cold).
 Coronaviruses David A. J. Tyrrell Steven H. Myint GENERAL CONCEPTS Clinical Presentation Coronaviruses cause acute, mild upper respiratory infection (common cold). Structure Spherical or pleomorphic enveloped
Coronaviruses David A. J. Tyrrell Steven H. Myint GENERAL CONCEPTS Clinical Presentation Coronaviruses cause acute, mild upper respiratory infection (common cold). Structure Spherical or pleomorphic enveloped
Protein Modeling Event
 Protein Modeling Event School Name: School Number: Team Member 1: Team Member 2: : Pre-Build Score: On-Site Build Score: Test Score: Tie Breaker: Total: Final Rank: Part I: Pre-Build (40% of total score)
Protein Modeling Event School Name: School Number: Team Member 1: Team Member 2: : Pre-Build Score: On-Site Build Score: Test Score: Tie Breaker: Total: Final Rank: Part I: Pre-Build (40% of total score)
The 3-dimensional play of human parechovirus infection; Cell, virus and antibody Westerhuis, Brenda
 UvA-DARE (Digital Academic Repository) The 3-dimensional play of human parechovirus infection; Cell, virus and antibody Westerhuis, Brenda Link to publication Citation for published version (APA): Westerhuis,
UvA-DARE (Digital Academic Repository) The 3-dimensional play of human parechovirus infection; Cell, virus and antibody Westerhuis, Brenda Link to publication Citation for published version (APA): Westerhuis,
The Basics: A general review of molecular biology:
 The Basics: A general review of molecular biology: DNA Transcription RNA Translation Proteins DNA (deoxy-ribonucleic acid) is the genetic material It is an informational super polymer -think of it as the
The Basics: A general review of molecular biology: DNA Transcription RNA Translation Proteins DNA (deoxy-ribonucleic acid) is the genetic material It is an informational super polymer -think of it as the
Chapter 6- An Introduction to Viruses*
 Chapter 6- An Introduction to Viruses* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. 6.1 Overview of Viruses
Chapter 6- An Introduction to Viruses* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. 6.1 Overview of Viruses
T he family Picornaviridae is comprised of
 214 REVIEW Picornavirus uncoating M S Smyth, J H Martin... Recently, much has been learned about the molecular mechanisms involved in the pathogenesis of picornaviruses. This has been accelerated by the
214 REVIEW Picornavirus uncoating M S Smyth, J H Martin... Recently, much has been learned about the molecular mechanisms involved in the pathogenesis of picornaviruses. This has been accelerated by the
CS612 - Algorithms in Bioinformatics
 Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
Genome of Hepatitis B Virus. VIRAL ONCOGENE Dr. Yahwardiah Siregar, PhD Dr. Sry Suryani Widjaja, Mkes Biochemistry Department
 Genome of Hepatitis B Virus VIRAL ONCOGENE Dr. Yahwardiah Siregar, PhD Dr. Sry Suryani Widjaja, Mkes Biochemistry Department Proto Oncogen and Oncogen Oncogen Proteins that possess the ability to cause
Genome of Hepatitis B Virus VIRAL ONCOGENE Dr. Yahwardiah Siregar, PhD Dr. Sry Suryani Widjaja, Mkes Biochemistry Department Proto Oncogen and Oncogen Oncogen Proteins that possess the ability to cause
For all of the following, you will have to use this website to determine the answers:
 For all of the following, you will have to use this website to determine the answers: http://blast.ncbi.nlm.nih.gov/blast.cgi We are going to be using the programs under this heading: Answer the following
For all of the following, you will have to use this website to determine the answers: http://blast.ncbi.nlm.nih.gov/blast.cgi We are going to be using the programs under this heading: Answer the following
Bioinformation by Biomedical Informatics Publishing Group
 Predicted RNA secondary structures for the conserved regions in dengue virus Pallavi Somvanshi*, Prahlad Kishore Seth Bioinformatics Centre, Biotech Park, Sector G, Jankipuram, Lucknow 226021, Uttar Pradesh,
Predicted RNA secondary structures for the conserved regions in dengue virus Pallavi Somvanshi*, Prahlad Kishore Seth Bioinformatics Centre, Biotech Park, Sector G, Jankipuram, Lucknow 226021, Uttar Pradesh,
7.012 Quiz 3 Answers
 MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel Friday 11/12/04 7.012 Quiz 3 Answers A > 85 B 72-84
MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel Friday 11/12/04 7.012 Quiz 3 Answers A > 85 B 72-84
Envelope e_ :------, Envelope glycoproteins e_ ~ Single-stranded RNA ----, Nucleocapsid
 Virus Antib(tdies Your Expertise, Our Antibodies, Accelerated Discovery. Envelope e_---------:------, Envelope glycoproteins e_-------1~ Single-stranded RNA ----, Nucleocapsid First identified in 1989,
Virus Antib(tdies Your Expertise, Our Antibodies, Accelerated Discovery. Envelope e_---------:------, Envelope glycoproteins e_-------1~ Single-stranded RNA ----, Nucleocapsid First identified in 1989,
COMPARATIVE STUDY OF ANTIBODY LEVELS DEVELOPED BY VACCINATION AGAINST POLIO VIRUS IN POPULATION AFTER VACCINE TYPE ALTERATION
 Acta Microbiologica et Immunologica Hungarica, 62 (1), pp. 75 85 (2015) DOI: 10.1556/AMicr.62.2015.1.6 COMPARATIVE STUDY OF ANTIBODY LEVELS DEVELOPED BY VACCINATION AGAINST POLIO VIRUS IN POPULATION AFTER
Acta Microbiologica et Immunologica Hungarica, 62 (1), pp. 75 85 (2015) DOI: 10.1556/AMicr.62.2015.1.6 COMPARATIVE STUDY OF ANTIBODY LEVELS DEVELOPED BY VACCINATION AGAINST POLIO VIRUS IN POPULATION AFTER
Rapid Simultaneous Detection of Enterovirus and Parechovirus RNAs in Clinical Samples by One-Step Real-Time Reverse Transcription-PCR Assay
 JOURNAL OF CLINICAL MICROBIOLOGY, July 2011, p. 2620 2624 Vol. 49, No. 7 0095-1137/11/$12.00 doi:10.1128/jcm.02445-10 Copyright 2011, American Society for Microbiology. All Rights Reserved. Rapid Simultaneous
JOURNAL OF CLINICAL MICROBIOLOGY, July 2011, p. 2620 2624 Vol. 49, No. 7 0095-1137/11/$12.00 doi:10.1128/jcm.02445-10 Copyright 2011, American Society for Microbiology. All Rights Reserved. Rapid Simultaneous
LESSON 4.4 WORKBOOK. How viruses make us sick: Viral Replication
 DEFINITIONS OF TERMS Eukaryotic: Non-bacterial cell type (bacteria are prokaryotes).. LESSON 4.4 WORKBOOK How viruses make us sick: Viral Replication This lesson extends the principles we learned in Unit
DEFINITIONS OF TERMS Eukaryotic: Non-bacterial cell type (bacteria are prokaryotes).. LESSON 4.4 WORKBOOK How viruses make us sick: Viral Replication This lesson extends the principles we learned in Unit
Analysis of the interaction between the 2C protein of parechoviruses and lipid droplets. Ali Aidarous Khrid
 Analysis of the interaction between the 2C protein of parechoviruses and lipid droplets Ali Aidarous Khrid A thesis submitted for the degree of Doctor of Philosophy School of Biological Sciences University
Analysis of the interaction between the 2C protein of parechoviruses and lipid droplets Ali Aidarous Khrid A thesis submitted for the degree of Doctor of Philosophy School of Biological Sciences University
REGULATED SPLICING AND THE UNSOLVED MYSTERY OF SPLICEOSOME MUTATIONS IN CANCER
 REGULATED SPLICING AND THE UNSOLVED MYSTERY OF SPLICEOSOME MUTATIONS IN CANCER RNA Splicing Lecture 3, Biological Regulatory Mechanisms, H. Madhani Dept. of Biochemistry and Biophysics MAJOR MESSAGES Splice
REGULATED SPLICING AND THE UNSOLVED MYSTERY OF SPLICEOSOME MUTATIONS IN CANCER RNA Splicing Lecture 3, Biological Regulatory Mechanisms, H. Madhani Dept. of Biochemistry and Biophysics MAJOR MESSAGES Splice
MOLECULAR CELL BIOLOGY
 1 Lodish Berk Kaiser Krieger scott Bretscher Ploegh Matsudaira MOLECULAR CELL BIOLOGY SEVENTH EDITION CHAPTER 13 Moving Proteins into Membranes and Organelles Copyright 2013 by W. H. Freeman and Company
1 Lodish Berk Kaiser Krieger scott Bretscher Ploegh Matsudaira MOLECULAR CELL BIOLOGY SEVENTH EDITION CHAPTER 13 Moving Proteins into Membranes and Organelles Copyright 2013 by W. H. Freeman and Company
7.012 Problem Set 6 Solutions
 Name Section 7.012 Problem Set 6 Solutions Question 1 The viral family Orthomyxoviridae contains the influenza A, B and C viruses. These viruses have a (-)ss RNA genome surrounded by a capsid composed
Name Section 7.012 Problem Set 6 Solutions Question 1 The viral family Orthomyxoviridae contains the influenza A, B and C viruses. These viruses have a (-)ss RNA genome surrounded by a capsid composed
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature14008 Supplementary Figure 1. Sequence alignment of A/little yellow-shouldered bat/guatemala/060/2010 (H17N10) polymerase with that of human strain A/Victoria/3/75(H3N2). The secondary
doi:10.1038/nature14008 Supplementary Figure 1. Sequence alignment of A/little yellow-shouldered bat/guatemala/060/2010 (H17N10) polymerase with that of human strain A/Victoria/3/75(H3N2). The secondary
Molecular Evolution of the Human Enteroviruses: Correlation of Serotype with VP1 Sequence and Application to Picornavirus Classification
 JOURNAL OF VIROLOGY, Mar. 1999, p. 1941 1948 Vol. 73, No. 3 0022-538X/99/$04.00 0 Molecular Evolution of the Human Enteroviruses: Correlation of Serotype with VP1 Sequence and Application to Picornavirus
JOURNAL OF VIROLOGY, Mar. 1999, p. 1941 1948 Vol. 73, No. 3 0022-538X/99/$04.00 0 Molecular Evolution of the Human Enteroviruses: Correlation of Serotype with VP1 Sequence and Application to Picornavirus
VIRUSES. 1. Describe the structure of a virus by completing the following chart.
 AP BIOLOGY MOLECULAR GENETICS ACTIVITY #3 NAME DATE HOUR VIRUSES 1. Describe the structure of a virus by completing the following chart. Viral Part Description of Part 2. Some viruses have an envelope
AP BIOLOGY MOLECULAR GENETICS ACTIVITY #3 NAME DATE HOUR VIRUSES 1. Describe the structure of a virus by completing the following chart. Viral Part Description of Part 2. Some viruses have an envelope
Enterovirus D68-US Outbreak 2014 Laboratory Perspective
 Enterovirus D68-US Outbreak 2014 Laboratory Perspective Allan Nix Team Lead Picornavirus Laboratory Polio and Picornavirus Laboratory Branch PAHO / SARInet Webinar November 6, 2014 National Center for
Enterovirus D68-US Outbreak 2014 Laboratory Perspective Allan Nix Team Lead Picornavirus Laboratory Polio and Picornavirus Laboratory Branch PAHO / SARInet Webinar November 6, 2014 National Center for
7.014 Problem Set 7 Solutions
 MIT Department of Biology 7.014 Introductory Biology, Spring 2005 7.014 Problem Set 7 Solutions Question 1 Part A Antigen binding site Antigen binding site Variable region Light chain Light chain Variable
MIT Department of Biology 7.014 Introductory Biology, Spring 2005 7.014 Problem Set 7 Solutions Question 1 Part A Antigen binding site Antigen binding site Variable region Light chain Light chain Variable
Viral structure م.م رنا مشعل
 Viral structure م.م رنا مشعل Viruses must reproduce (replicate) within cells, because they cannot generate energy or synthesize proteins. Because they can reproduce only within cells, viruses are obligate
Viral structure م.م رنا مشعل Viruses must reproduce (replicate) within cells, because they cannot generate energy or synthesize proteins. Because they can reproduce only within cells, viruses are obligate
Virus Entry/Uncoating
 Virus Entry/Uncoating Delivery of genome to inside of a cell Genome must be available for first step of replication The Problem--barriers to infection Virion Barriers: Non-enveloped viruses capsid Enveloped
Virus Entry/Uncoating Delivery of genome to inside of a cell Genome must be available for first step of replication The Problem--barriers to infection Virion Barriers: Non-enveloped viruses capsid Enveloped
B19, see Parvovirus B19 Bone marrow, gene transfer with parvovirus. Erythrovirus, see Parvovirus B19, Simian parvovirus
 ... Subject Index Adeno-associated virus Cap and genome encapsidation 87 DNA integration homologous recombination 90, 91 latency vs replication 77, 78 mechanism 79 requirements 78, 79 site in human genome
... Subject Index Adeno-associated virus Cap and genome encapsidation 87 DNA integration homologous recombination 90, 91 latency vs replication 77, 78 mechanism 79 requirements 78, 79 site in human genome
Week 5 Section. Junaid Malek, M.D.
 Week 5 Section Junaid Malek, M.D. HIV: Anatomy Membrane (partiallystolen from host cell) 2 Glycoproteins (proteins modified by added sugar) 2 copies of RNA Capsid HIV Genome Encodes: Structural Proteins
Week 5 Section Junaid Malek, M.D. HIV: Anatomy Membrane (partiallystolen from host cell) 2 Glycoproteins (proteins modified by added sugar) 2 copies of RNA Capsid HIV Genome Encodes: Structural Proteins
Chair of Medical Biology, Microbiology, Virology, and Immunology STRUCTURE, CLASSIFICATION AND PHYSIOLOGY OF VIRUSES
 Chair of Medical Biology, Microbiology, Virology, and Immunology STRUCTURE, CLASSIFICATION AND PHYSIOLOGY OF VIRUSES Viruses are small obligate intracellular parasites, which by definition contain either
Chair of Medical Biology, Microbiology, Virology, and Immunology STRUCTURE, CLASSIFICATION AND PHYSIOLOGY OF VIRUSES Viruses are small obligate intracellular parasites, which by definition contain either
Multiple sequence alignment
 Multiple sequence alignment Bas. Dutilh Systems Biology: Bioinformatic Data Analysis Utrecht University, February 18 th 2016 Protein alignments We have seen how to create a pairwise alignment of two sequences
Multiple sequence alignment Bas. Dutilh Systems Biology: Bioinformatic Data Analysis Utrecht University, February 18 th 2016 Protein alignments We have seen how to create a pairwise alignment of two sequences
RAS Genes. The ras superfamily of genes encodes small GTP binding proteins that are responsible for the regulation of many cellular processes.
 ۱ RAS Genes The ras superfamily of genes encodes small GTP binding proteins that are responsible for the regulation of many cellular processes. Oncogenic ras genes in human cells include H ras, N ras,
۱ RAS Genes The ras superfamily of genes encodes small GTP binding proteins that are responsible for the regulation of many cellular processes. Oncogenic ras genes in human cells include H ras, N ras,
Viruses. Rotavirus (causes stomach flu) HIV virus
 Viruses Rotavirus (causes stomach flu) HIV virus What is a virus? A virus is a microscopic, infectious agent that may infect any type of living cell. Viruses must infect living cells in order to make more
Viruses Rotavirus (causes stomach flu) HIV virus What is a virus? A virus is a microscopic, infectious agent that may infect any type of living cell. Viruses must infect living cells in order to make more
trans-encapsidation of a Poliovirus Replicon by Different Picornavirus Capsid Proteins
 JOURNAL OF VIROLOGY, October 1998, p. 7972 7977 Vol. 72, No. 10 0022-538X/98/$04.00 0 Copyright 1998, American Society for Microbiology. All Rights Reserved. trans-encapsidation of a Poliovirus Replicon
JOURNAL OF VIROLOGY, October 1998, p. 7972 7977 Vol. 72, No. 10 0022-538X/98/$04.00 0 Copyright 1998, American Society for Microbiology. All Rights Reserved. trans-encapsidation of a Poliovirus Replicon
Detection of Human Parechoviruses from clinical stool samples in Aichi, Japan
 JCM Accepts, published online ahead of print on June 1 J. Clin. Microbiol. doi:1./jcm.-1 Copyright 1, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved. 1 Detection
JCM Accepts, published online ahead of print on June 1 J. Clin. Microbiol. doi:1./jcm.-1 Copyright 1, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved. 1 Detection
Virus and Prokaryotic Gene Regulation - 1
 Virus and Prokaryotic Gene Regulation - 1 We have discussed the molecular structure of DNA and its function in DNA duplication and in transcription and protein synthesis. We now turn to how cells regulate
Virus and Prokaryotic Gene Regulation - 1 We have discussed the molecular structure of DNA and its function in DNA duplication and in transcription and protein synthesis. We now turn to how cells regulate
E E Hepatitis E SARS 29, Lancet. E A B Enterically-Transmitted Non-A, Hepatitis E. Virus HEV nm. 1.35g/cm s ALT AST HEV HEV
 7850 2004 Hepatitis E Tian-Cheng LI Naokazu TAKEDA Tatsuo MIYAMURA SARS 8 Lancet E E E Hepatitis E VirusHEV E E HEV HEV E 1955 29,000 E E A A B Enterically-Transmitted Non-A, Non-B Hepatitis 1983 Balayan
7850 2004 Hepatitis E Tian-Cheng LI Naokazu TAKEDA Tatsuo MIYAMURA SARS 8 Lancet E E E Hepatitis E VirusHEV E E HEV HEV E 1955 29,000 E E A A B Enterically-Transmitted Non-A, Non-B Hepatitis 1983 Balayan
CDC website:
 Hepatitis B virus CDC website: http://www.cdc.gov/ncidod/diseases/hepatitis/slideset/hep_b/slide_1.htm Relevance Key Features es of Hepatitis t B Virus 250 million people infected worldwide. In areas of
Hepatitis B virus CDC website: http://www.cdc.gov/ncidod/diseases/hepatitis/slideset/hep_b/slide_1.htm Relevance Key Features es of Hepatitis t B Virus 250 million people infected worldwide. In areas of
