Enterococcal-type glycopeptide resistance genes in non-enterococcal organisms
|
|
|
- Edward Perkins
- 6 years ago
- Views:
Transcription
1 FEMS Microbiology Letters 185 (1999) 1^7 MiniReview Enterococcal-type glycopeptide resistance genes in non-enterococcal organisms Robin Patel * Division of Infectious Diseases and Infectious Diseases Research Laboratory, Mayo Clinic and Foundation, Rochester, MN, USA Received 8 October 1999; received in revised form 14 January 2000; accepted 18 January Abstract Although the emergence of vancomycin-resistant enterococci can be attributed, in part, to the increasing use of vancomycin in clinical practice, and glycopeptide use in animal husbandry, the origins of the enterococcal vancomycin resistance genes are not clear. The vancomycin resistance-associated genes in Enterococcus gallinarum, Enterococcus casseliflavus/flavescens, Lactobacillus spp., Leuconostoc spp., Pediococcus spp., and Erysipelothrix rhusiopathiae, are not the source of the high-level vancomycin resistance-associated genes in enterococci. There are, however, environmental organisms which have been found to have gene clusters homologous to the enterococcal vana, vanb and vanc gene clusters; these include the biopesticide Paenibacillus popilliae, and, to a lesser extent, the glycopeptide-producing organisms Amycolatopsis orientalis and Streptomyces toyocaensis. Still, the exact sources of the enterococcal vancomycin resistance genes remain a mystery. ß 1999 Federation of European Microbiological Societies. Published by Elsevier Science B.V. All rights reserved. Keywords: Enterococcus; Glycopeptide resistance; Vancomycin; Paenibacillus popilliae Among the most dramatic and clinically worrisome examples of resistance to antimicrobial agents in recent years has been the emergence and spread of vancomycin resistance in enterococci. This increase poses important problems, including the death of available antimicrobial agents active against these organisms, in many cases, and the possibility that vancomycin resistance genes can be transferred to other Gram-positive bacteria, especially Staphylococcus aureus, coagulase-negative staphylococci (CNS), Streptococcus pneumoniae, Bacillus spp. and Corynebacterium spp. The transfer of vancomycin resistance to methicillin-resistant S. aureus (MRSA), methicillin-resistant CNS (MR-CNS), high-level penicillin-resistant S. pneumoniae and/or penicillin-resistant Corynebacterium spp. (e.g. Corynebacterium jeikeium) might compromise successful therapy for infections caused by these organisms. Enterococci with VanA- or VanB-type high-level vancomycin resistance have acquired genetic material which leads to the manufacture of an abnormal ligase (in addition to the normal ligase described below) responsible for the synthesis of the depsipeptide D-alanyl-D-lactate (D-Ala- * Tel.: +1 (507) ; Fax: +1 (507) ; patel.robin@mayo.edu D-Lac) which is incorporated into a pentapeptide peptidoglycan cell wall precursor (UDP-MurNAc-L-Ala-D-Glu-L- Lys-D-Ala-D-Lac) to which vancomycin binds poorly. In contrast, in vancomycin-susceptible cells, vancomycin complexes with the D-Ala-D-Ala termini of normal pentapeptide peptidoglycan cell wall precursor (UDP-MurNAc- L-Ala-D-Glu-L-Lys-D-Ala-D-Ala) thereby inhibiting cell wall synthesis. Although this normal pentapeptide peptidoglycan cell wall precursor pathway is preserved in enterococci with VanA- or VanB-type vancomycin resistance, there is hydrolysis of some of its components. VanA-type glycopeptide resistance is characterized by acquired inducible resistance to both vancomycin and teicoplanin. It is mediated by Tn1546 or closely related elements [1]. Tn1546 encodes nine polypeptides that can be assigned to di erent functional groups: Transposition functions, regulatory proteins (VanR and VanS), synthesis of depsipeptide D-Ala-D-Lac (VanH and VanA) and hydrolysis of precursors of normal peptidoglycan, at least in vitro (VanX and VanY); the function of VanZ is unknown [1]. The vanr, vans, vanh, vana and vanx genes are necessary and su cient for the inducible expression of resistance to glycopeptides [1]. VanY and VanZ are accessory peptides and are not required for resistance. Genetic heterogeneity has been described in the vana gene clusters of vancomycin-resistant enterococci (VRE) [2], and the / 99 / $20.00 ß 1999 Federation of European Microbiological Societies. Published by Elsevier Science B.V. All rights reserved. PII: S (00)
2 2 R. Patel / FEMS Microbiology Letters 185 (1999) 1^7 vana gene cluster has been found on the chromosome as well as on plasmids [3]. VanB-type glycopeptide resistance is characterized by acquired inducible resistance to various concentrations of vancomycin, but not to teicoplanin. The vanb gene cluster has homology to the vana gene cluster, but has been less well studied. It consists of genes encoding polypeptides assigned to regulation of vancomycin resistance genes (vanr B and vans B ), synthesis of the depsipeptide D-Ala- D-Lac (vanh B and vanb), and hydrolysis of precursors of normal peptidoglycan, at least in vitro (vanx B and vany B ) [4]. An open reading frame vanw is also present and encodes a polypeptide of unknown function [4]. Our group has identi ed sequence variability in the vanb gene of different isolates of enterococci with VanB resistance [5]. The sequence variability observed in these isolates is consistent with the hypothesis that the spread of vancomycin resistance amongst Enterococcus spp. results not just from dissemination of a single clone, but also as the result of horizontal transfer of resistance genes from as yet unde- ned organisms [6]. High-level enterococcal vancomycin resistance is readily transferred from one enterococcus to another in the laboratory. Additionally, laboratory experiments have achieved the transfer of high-level vancomycin resistance from enterococci to S. aureus [7]. Vancomycin resistance has also been transferred in vitro by conjugation or transformation from enterococci to Streptococcus sanguis, Lactococcus lactis, Streptococcus pyogenes, and Listeria monocytogenes [8,9]. The origins of the vancomycin resistance-associated genes in enterococci are unknown, but the selective pressures favoring the emergence of VRE are more clear. Vancomycin was rst licensed in the United States in 1958; however, it took until 1986 to document high-level vancomycin resistance in enterococci. Given the complexity of the mechanism of vancomycin resistance in enterococci, it is possible that it took time to gather together multiple individual genes required for the expression of resistance. This hypothesis is not supported by the multicentric nature of the emergence of VRE in the United States and Europe. On the contrary, the emergence of VRE in the late 1980s suggests that preassembled operons were transferred to enterococci, before or at that time, and selected for under the pressure of dramatically increased vancomycin use in clinical practice and glycopeptide use (e.g. avoparcin and orienticin) in animal husbandry. Prior to the late 1970s, there was little clinical use of vancomycin in the United States; thereafter, however, with the identi cation of Clostridium di cile as a cause of diarrhea and with the emergence of MRSA and MR-CNS, oral and intravenous vancomycin use, respectively, increased dramatically, likely contributing to the emergence of VRE, especially in the United States. In the United States, the acquisition of vancomycin resistance by multiply drug-resistant (e.g. ampicillin- and aminoglycoside-resistant) enterococci has resulted in VRE which are `very resistant enterococci', which have (unfortunately) spread nosocomially, and have, in some instances, resulted in untreatable infections. In Europe, on the other hand, the use of avoparcin in animal husbandry has created a hospitable, but not necessarily a hospital, environment for the emergence of VRE. In contrast to VRE in the United States, European VRE are less likely to be multiply drug resistant and are less frequently cultured from hospitalized patients, presumably relating to their distinct origin, and perhaps to the absence of as yet uncharacterized human intestinal colonization-promoting factors present in VRE in the United States. There are several reports of the enterococcal vana and vanb genes being found in non-enterococcal bacteria isolated from humans. The vana gene has been found in vancomycin-resistant clinical isolates of Cellulomonas turbata and Arcanobacterium haemolyticum (typically these organisms are vancomycin susceptible) isolated from the stools of two patients during an outbreak of VRE infection in London, England [9]. The vanb gene has been found in a vancomycin-resistant isolate of Streptococcus bovis isolated from a stool swab collected on admission from a patient as surveillance for VRE [10]. The vana gene cluster has been identi ed in a Bacillus circulans isolate in a case of catheter-related infection [11]. In the aforementioned cases, the enterococcal vancomycin resistance genes appear to have been transferred into these isolates, and these organisms themselves are unlikely to have been the original source for the vancomycin resistance-associated genes in enterococci. Two species of enterococci are intrinsically resistant to vancomycin: Enterococcus gallinarum and Enterococcus casseli avus/ avescens. The vanc-1 gene in E. gallinarum and the vanc-2/3 genes in E. casseli avus/ avescens encode a D-Ala:D-Ser ligase, which is associated with low-level vancomycin resistance (a D-Ala:D-Ala ligase gene is also present) [12]. In E. gallinarum, three enzymes are su cient for vancomycin resistance: the VanC-1 ligase for production of D-Ala-D-Ser, the VanXY C protein for hydrolysis of D-Ala-D-Ala and for removal of D-Ala from UDP-Mur- NAc-pentapeptide ending in D-Ala, and VanT for D-serine synthesis [13]. There is only 37^39% deduced (partial) amino acid sequence identity between the VanC enzymes and VanA/B [14]; therefore the VanC enzymes are not the source of high-level, vana- and vanb-associated vancomycin resistance in enterococci. As an aside, the origin of and the rationale behind the development of D-Ala:D-Ser ligases are not clear as this pathway does not appear to be metabolically advantageous for the bacteria and seems too ancient to be explained by selective pressure due to the use of glycopeptides in clinical or agricultural practice [15]. Two novel types of vancomycin resistance have recently been described in enterococci. VanD-type vancomycin resistance is distinct from VanA- and VanB-type vancomycin resistance and although associated with synthesis of
3 R. Patel / FEMS Microbiology Letters 185 (1999) 1^7 3 the depsipeptide D-Ala-D-Lac, does not appear to be transferable [16,17]. VanE-type vancomycin resistance is lowlevel vancomycin resistance recently described in E. faecalis and associated with a probable D-Ala:D-Ser ligase similar to that present in VanC-type enterococci [18]. Both of these new types of vancomycin resistance are not frequently seen amongst current clinical isolates of VRE and are not the source of glycopeptide resistance in the more common VanA- and VanB-type VRE isolates. Several bacteria have been known to be resistant to vancomycin since before the identi cation of vancomycin resistance in enterococci. These include Lactobacillus spp., Leuconostoc spp., Pediococcus spp., and Erysipelothrix rhusiopathiae. Vancomycin resistance in Lactobacillus casei, Pediococcus pentosaceus, Lactobacillus plantarum and Leuconostoc mesenteroides is associated with the presence of a pentapeptide peptidoglycan precursor terminating in D-Ala-D-Lac [12,19]. The peptidoglycan precursor structure in L. mesenteroides (MurNAc-L-Ala-D-Glu-L- Lys-[L-Ala]-D-Ala-D-Lac) [19], and that in L. plantarum (MurNAc-L-Ala-D-Glu-m-Dpm-D-Ala-D-Lac) [20], di er from that in enterococci, but still terminate in D-Ala-D- Lac. It is not clear whether one or two ligases are present in such organisms. A L. plantarum strain de cient for D- and L-lactate dehydrogenase activities, and which therefore produced only trace amounts of D- and L-lactate, has been constructed [20]. These changes drastically affected the peptidoglycan synthesis pathway [20], and a new precursor, terminating in D-alanine was identi able, in addition to that terminating in D-lactate [20]. In addition, the organism had highly enhanced susceptibility to vancomycin [20]. Whether the origin of the D-alanine-ending precursor in the mutant was a result of a second ligase or mixed substrate speci city of a single ligase which preferentially uses D-lactate is not completely clear. However, since a comparison of the ligase genes from constitutively vancomycin-resistant lactobacilli (including L. plantarum) and L. mesenteroides, has shown that their deduced amino acid sequences are more closely related to each other than to that of a vancomycin-susceptible strain of Lactobacillus leichmannii, the latter is probably the case [21]. Vancomycin resistance in Lactobacillus confusus, Leuconostoc spp., and Pediococcus spp. is not transferable to enterococci using lter mating [22]. Although Leuconostoc spp. generally harbor a number of plasmids, plasmid-free strains of Leuconostoc spp. obtained by curing with novobiocin and ethidium bromide have been shown by some investigators to retain resistance to glycopeptides [19]. There has been one report of a clinical isolate of Lactobacillus viridescens in which glycopeptide resistance was associated with the presence of plasmid DNA and variations in cell wall protein patterns were noted following exposure to vancomycin [23], and another report that ethidium bromide curing of multiple plasmids was associated with concomitant loss of vancomycin resistance in Lactobacillus acidophilus [24]. These observations would suggest that two ligases are present in these organisms and that vancomycin resistance from lactobacilli could be transferable. This issue is controversial. Although resistance to glycopeptides in Leuconostoc spp., Lactobacillus spp., and Pediococcus spp. is associated with the presence of peptidoglycan precursors terminating in D-Ala-D-Lac, as is the situation for enterococci with VanA- and VanB-mediated vancomycin resistance [12,19], the D-Ala:D-Lac ligase and D-lactate dehydrogenase enzymes involved in the synthesis of the altered precursors in Leuconostoc spp. and Lactobacillus spp. are not closely related to VanA or VanB and VanH or VanH B, respectively [21,26^28]. In one study, no polymerase chain reaction (PCR) product was ampli ed in L. rhamnosus GG with any of three sets of vana primers used, and enterococcal vana, vanb, vanh, vanx, vanz, vany, vans and vanr genetic material did not hybridize with DNA of L. rhamnosus GG [29]. More detailed studies have revealed that the levels of identity between the D-Ala:D-Lac ligase in Leuconostoc spp. and Lactobacillus spp. and VanA/B are 26^35% [21]. The D-Ala:D-Lac ligases of the vancomycin-resistant lactic acid bacteria di er in their primary sequence from VanA, VanB and VanD D-Ala:D-Lac ligases, VanC D-Ala:D-Ser ligases, and D-Ala:D-Ala ligases, notably in the g-loop region, which is critical to catalysis [15]. This suggests that the evolution toward D-Ala-D-Lac formation occurred independently in ligases from lactic acid bacteria and VanA, VanB and VanD enterococci [15]. In lactic acid bacteria, lactate is abundant and the use of D- lactate instead of D-alanine could simply re ect metabolic parsimony [15]. Selective pressure due to the presence of glycopeptide-producing organisms in the environment could also have played a role in the development of D- Ala:D-Lac ligases in lactic acid bacteria [15]. Interestingly, it has recently been demonstrated that insertional inactivation of the rrp-31 gene, which encodes a response regulator of Lactobacillus sakei, increases the vancomycin MIC of this isolate, which is otherwise vancomycin susceptible [30]. Whether this relates to loss of autolysis, related to an altered two-component sensor-regulator system, as has recently been described in S. pneumoniae [31], or altered peptidoglycan precursor synthesis, or another mechanism, remains to be determined [30]. E. rhusiopathiae is resistant to vancomycin with a MIC 90 of v64 Wg ml 31 (range, 4^v64 mg ml 31 ) [32], however, to the best of my knowledge, the mechanism of vancomycin resistance in E. rhusiopathiae remains to be described. PCR assays designed to detect the vana and vanb genes in enterococci have not detected these genes in E. rhusiopathiae (Patel, unpublished results). For the aforementioned reasons, Lactobacillus spp., Leuconostoc spp., Pediococcus spp., E. rhusiopathiae, and the intrinsically vancomycin-resistant enterococci are not the source of the glycopeptide resistance genes in VRE. There are, however, environmental organisms which have been found to have gene clusters resembling the
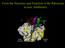
4 4 R. Patel / FEMS Microbiology Letters 185 (1999) 1^7 Fig. 1. Alignment of glycopeptide resistance gene clusters of VanA, VanB, and VanD enterococci, P. popilliae, and the glycopeptide-producing organisms A. orientalis and S. toyocaensis [4,17,27,49^51]. The percents amino acid identity to vany, vanz, vanh, vana, and vanx, are shown below the respective genes (below the arrows). The percent G and C content of each gene is shown above the genes (above the arrows). Reproduced with permission from Patel et al. [34]. vana and vanb gene clusters; these include Paenibacillus popilliae, and, to a lesser extent, Amycolatopsis orientalis and Streptomyces toyocaensis. Biopesticidal powders containing spores of P. popilliae have been introduced into turf in the Eastern US for the suppression of Japanese beetle populations since the late 1930s. The type strain of P. popilliae (American Type Culture Collection 14706) has a vancomycin MIC of 800 Wg ml 31 [33]. The putative ligase gene in P. popilliae has 77% nucleotide identity to the sequence of the vana gene, and was originally designated vane [33], however since another vane gene was described [18], the putative ligase gene in P. popilliae was renamed vanf [34]. Putative genes, designated `vany F ', `vanz F ', `vanh F ', and `vanx F ', which resemble the enterococcal vancomycin resistance genes, vany, vanz, vanh, and vanx, respectively, have been identi ed in P. popilliae [34]. The predicted amino acid sequences of the putative proteins are similar to those found in VanA VRE; 61% identity for VanY F, 21% identity for VanZ F, 74% identity for VanH F, 77% identity for VanF, and 79% identity for VanX F (Fig. 1). The putative ligase vanf gene has been ampli ed from a P. popilliae isolate held in dried Japanese beetle hemolymph since 1945 and from a collection of 32 P. popilliae isolates [33]. This provides compelling evidence that the ligase gene present in P. popilliae was not transferred to this organism from an enterococcus. P. popilliae has recently been reported to be a human pathogen [35]; in addition, an organism resembling Paenibacillus alvei, which is intermediately susceptible to vancomycin (vancomycin MIC, 8 Wg ml 31 ), has been reported as a cause of pneumonia and empyema in a patient [36]. The mechanism of resistance to vancomycin in this organism is unknown, however no vana- or vanb-type gene was present in this organism using PCR (Patel, unpublished results). Recently, Marshall et al. cloned three genes encoding homologues of VanH, VanA, and VanX from the vancomycin producer A. orientalis C329.2 and the A47934 producer S. toyocaensis NRRL [27]. The predicted amino acid sequences of these proteins are similar to those found in VanA VRE: 54 to 61% identity for VanH, 59 to 63% identity for VanA, and 61 to 64% identity for VanX (Fig. 1) [27]. Phylogenetic analysis demonstrates that the gene products from S. toyocaensis, A. orientalis, P. popilliae, and VRE form distinct subfamilies of D-Ala:D-Lac ligases and D-lactate dehydrogenases [27,33]. The orientations of vanh, vana, and vanx, in P. popilliae, S. toyocaensis, and A. orientalis, are identical to the orientations found in VRE, and the signature overlap of the 5P end of the vana and homologous genes with the 3P end of the vanh genes is conserved [27,33]. Southern analysis of total DNA from other glycopeptide-producing organisms, including, A. orientalis (chloro-eremomycin producer), A. orientalis subsp. lurida (ristocetin producer), and Amycolatopsis coloradensis subsp. labeda (teicoplanin and avoparcin producer), with a probe derived from the vanh, vana, and vanx cluster from A. orientalis C329.2 has demonstrated cross-hybridizing DNA in all strains [27]. In addition, the vanh, vana, vanx cluster can be ampli ed from these other glycopeptide-producing organisms by PCR with degenerate primers complementary to conserved regions of VanH and VanX [27]. In S. toyocaensis there are two D-:D-ligases: a D-Ala:D-Ala ligase and a D-Ala:D-Lac ligase [37]. Marshall et al. hypothesize that the origin of clinically relevant vancomycin resistance lies within the glycopeptide-producing organisms [27]. However, the amino acid identities identi ed by these authors between the glycopeptide-producing organisms S. toyocaensis and A. orientalis and VanA and VanB VRE are substantially less than the identities reported between these genes in the vana gene cluster and in P. popilliae (Fig. 1). Furthermore, the G+C content of the P. popilliae vanh F, vanf and vanx F genes is virtually identical to that of the homologous genes in VRE, and signi cantly di erent from the homologous
5 R. Patel / FEMS Microbiology Letters 185 (1999) 1^7 5 genes in S. toyocaensis and A. orientalis (Fig. 1). The vancomycin resistance gene cluster in P. popilliae is more similar to that in VRE than are the gene clusters in S. toyocaensis and A. orientalis. Vancomycin resistance is not transferable from A. orientalis (or the teicoplanin producer A. teichomyceticus) to enterococci [22]. It is possible that the enterococcal vancomycin resistance-associated genes originated in the glycopeptide-producing organisms (where they would presumably be required to prevent such organisms from committing suicide), were then transferred into organisms of similar G+C content to enterococci (like P. popilliae), and thereafter into enterococci. Our group has not, however, been able to transfer vancomycin resistance from P. popilliae to enterococci (Patel, unpublished results). The incomplete identity of the P. popilliae vancomycin resistance genes and the enterococcal vancomycin resistance genes suggests that there exist other organisms which may be the direct sources of vancomycin resistance genes in enterococci. Given that the G+C contents of the VRE vanh, vana and vanx genes are 5^10% higher than those of the adjacent vanr, vans, vany, and vanz genes (Fig. 1), it is plausible that the vanh, vana, and vanx genes have been mobilized as a unit from another source and positioned into contact with appropriate control elements. Transfer of vancomycin resistance from an organism like P. popilliae to an enterococcus may rst require transfer of a conjugative transposon from an enterococcus (or another Gram-positive bacterium) to this organism, followed by insertion of vancomycin resistance determinants into the conjugative transposon, and thereafter transfer of the transposon and vancomycin resistance back into an enterococcus [38]. This may account for our di culty in trying to transfer vancomycin resistance from P. popilliae into an enterococcus. Interestingly, although all North American isolates of P. popilliae we have examined are vancomycin resistant, most Latin American P. popilliae isolates are vancomycin susceptible suggesting that vancomycin resistance has been acquired by P. popilliae (Yousten, unpublished results). Given that we have been able to detect the vanf gene in a P. popilliae isolate from 1945, transfer of vancomycin resistance into P. popilliae would have occurred over ve decades ago [33]. The vanh/vana/vanx gene system may have evolved by mutations of genes encoding proteins involved in normal cell wall biosynthesis. For example, in the Gram-negative bacterium Escherichia coli, which is never challenged by glycopeptide antibiotics (because they cannot penetrate the outer membrane permeability barrier), ddpx (a vanx homologue) is part of a dipeptide transport and degradation system which presumably functions to recapture D-Ala-D- Ala [39]. The catalytic e ciency of D-Ala-D-Ala hydrolysis for DdpX is 25-fold lower than for enterococcal VanX, suggesting that DdpX functions in cell wall turnover, rather than protection [40]. The vanh/vana/vanx gene system may have been built using a more e cient VanX homologue and mutating the gene that synthesizes D- Ala-D-Ala to synthesize D-Ala-D-Lac. For example, it has been shown that single-residue changes in the active-site regions of D-,D-ligases can cause substantial changes in recognition and activation of hydroxy or amino acids [41]. Other bacteria with known elevated vancomycin MICs include Clostridium innocuum (vancomycin MIC Wg ml 31, range 8^16 Wg ml 31 ), Nocardia spp. (vancomycin MIC Wg ml 31, range 16^128 Wg ml 31 ), and single isolates of Gemella haemolysans (vancomycin MIC s 32 Wg ml 31 ) and Corynebacterium spp. (vancomycin MIC 32 Wg ml 31 ) [25,42^44]. The mechanism of resistance to vancomycin in these isolates in unknown, although no enterococcal vana or vanb genes were identi ed in C. innocuum using a polymerase chain reaction assay (Patel, unpublished results). Vancomycin intermediately susceptible S. aureus isolates have been recently described in Japan and in the United States [45]. The mechanism of vancomycin resistance in vancomycin intermediately susceptible S. aureus is postulated to be a result of thickened cell wall and resultant increased vancomycin-binding capacity e ectively preventing vancomycin molecules from reaching the cytoplasmic membrane by trapping them within the existing cell wall [46]. The vana and vanb genes have not been detected in these isolates. Recently, emergence of vancomycin tolerance in S. pneumoniae as a result of loss of function of the VncS histidine kinase of a two-component sensor-regulator system thought to regulate a basic pathway triggering autolysis, has been reported [31]. Although there is 38% identity between the deduced amino acid sequences of VncS and the response regulator, VcnR, and the VanS B -VanR B twocomponent regulatory system in enterococci, scanning of the pneumococcal genome reveals no homologues of vanh, vana, vanb or vanx [31]. Although similar twocomponent systems a ect susceptibility to vancomycin in pneumococci and enterococci, their underlying roles in resistance and tolerance appear to be distinct, presumably re ecting di erent gene products controlled by the response regulator [31]. In conclusion, although the emergence of VRE can be attributed, in part, to the increasing use of oral and parenteral vancomycin in humans since the late 1970s for the treatment of C. di cile and MRSA (and MR-CNS) infections, respectively, and glycopeptide use in animal husbandry [47,48], the exact source of the enterococcal vancomycin resistance genes remains a mystery. Glycopeptide-producing organisms are potential candidates in this regard, especially as the original source of vancomycinresistant genes. Given the ndings in P. popilliae, antimicrobial resistance in glycopeptide-producing organisms may have been transferred to other organisms with a G+C content more similar to enterococci, with vancomycin resistance then being transferred from these organisms into enterococci.
6 6 R. Patel / FEMS Microbiology Letters 185 (1999) 1^7 References [1] Arthur, M., Reynolds, P.E., Depardieu, F., Evers, S., Dutka-Malen, S., Quintiliani, R. and Courvalin, P. (1996) Mechanisms of glycopeptide resistance in enterococci. J. Infect. 32, 11^16. [2] Handwerger, S., Skoble, J., Discotto, L.F. and Pucci, M.J. (1995) Heterogeneity of the vana gene cluster in clinical isolates of enterococci from the northeastern United States. Antimicrob. Agents Chemother. 39, 362^368. [3] Handwerger, S. and Skoble, J. (1995) Identi cation of chromosomal mobile element conferring high-level vancomycin resistance in Enterococcus faecium. Antimicrob. Agents Chemother. 39, 2446^2453. [4] Evers, S. and Courvalin, P. (1996) Regulation of VanB-type vancomycin resistance gene expression by the VanS(B)-VanR(B) two-component regulatory system in Enterococcus faecalis V583. J. Bacteriol. 178, 1302^1309. [5] Patel, R., Uhl, J.R., Kohner, P., Hopkins, M.K., Steckelberg, J.M., Kline, B. and Cockerill, F.R. (1998) DNA sequence variation within vana, vanb, vanc-1, and vanc-2/3 genes of clinical Enterococcus spp. isolates. Antimicrob. Agents Chemother. 42, 202^205. [6] Patel, R., Uhl, J.R., Kohner, P., Hopkins, M.K. and Cockerill, F.R. (1997) Multiplex PCR detection of vana, vanb, vanc-1 and vanc-2/3 genes in enterococci. J. Clin. Microbiol. 35, 703^707. [7] Noble, W.C., Virani, Z. and Cree, R.G. (1992) Co-transfer of vancomycin and other resistance genes from Enterococcus faecalis NCTC to Staphylococcus aureus. FEMS Microbiol. Lett. 72, 195^ 198. [8] Biavasco, F., Giovanetti, E., Miele, A., Vignaroli, C., Facinelli, B. and Varaldo, P.E. (1996) In vitro conjugative transfer of VanA vancomycin resistance between Enterococci and Listeriae of di erent species. Eur. J. Clin. Microbiol. Infect. Dis. 15, 50^59. [9] Power, E.G.M., Abdulla, Y.H., Talsania, H.G., Spice, W., Aathithan, S. and French, G.L. (1995) vana genes in vancomycin-resistant isolates of Oerskovia turbata and Arcanobacterium (Corynebacterium) haemolyticum. J. Antimicrob. Chemother. 36, 595^606. [10] Poyart, C., Pierre, C., Quesne, G., Pron, B., Berche, P. and Trieu- Cuot, P. (1997) Emergence of vancomycin resistance in the genus Streptococcus: Characterization of a vanb transferable determinant in Streptococcus bovis. Antimicrob. Agents Chemother. 41, 24^29. [11] Fontana, R., Ligozzi, M., Pedrotti, C., Padovani, E.M. and Cornaglia, G. (1997) Vancomycin-resistant Bacillus circulans carrying the vana gene responsible for vancomycin resistance in enterococci. Eur. J. Clin. Microbiol. Infect. Dis. 16, 473^474. [12] Billot-Klein, D., Gutmann, L., Sable, S., Guittet, E. and van Heijenoort, J. (1994) Modi cation of peptidoglycan precursors is a common feature of the low-level vancomycin-resistant VANB-type Enterococcus D366 and of the naturally glycopeptide-resistant species Lactobacillus casei, Pediococcus pentosaceus, Leuconostoc mesenteroides, and Enterococcus gallinarum. J. Bacteriol. 176, 2398^2405. [13] Arias, C.A., Martin-Martinez, M., Blundell, T.L., Arthur, M., Courvalin, P. and Reynolds, P.E. (1999) Characterization and modelling of VanT: a novel, membrane-bound, serine racemase from vancomycin-resistant Enterococcus gallinarum BM4174. Mol. Microbiol. 31, 1653^1664. [14] Navarro, F. and Courvalin, P. (1994) Analysis of genes encoding D- alanine-d-alanine ligase-related enzymes in Enterococcus casseli avus and Enterococcus avescens. Antimicrob. Agents Chemother. 38, 1788^1793. [15] Evers, S., Casadewall, B., Charles, M., Dutka-Malen, S., Galimand, M. and Courvalin, P. (1996) Evolution of structure and substrate speci city in D-alanine:D-alanine ligases and related enzymes. J. Mol. Evol. 42, 706^712. [16] Perichon, B., Reynolds, P. and Courvalin, P. (1997) VanD-type glycopeptide-resistant Enterococcus faecium BM4339. Antimicrob. Agents Chemother. 41, 2016^2018. [17] Casadewall, B. and Courvalin, P. (1999) Characterization of the vand glycopeptide resistance gene cluster from Enterococcus faecium BM4339. J. Bacteriol. 181, 3644^3648. [18] Fines, M., Perichon, B., Reynolds, P., Sahm, D. and Courvalin, P. (1999) VanE, a new type of acquired glycopeptide resistance in Enterococcus faecalis BM4405. Antimicrob. Agents Chemother. 43, 2161^2164. [19] Handwerger, S., Pucci, M.J., Volk, K.J., Liu, J. and Lee, M.S. (1994) Vancomycin-resistant Leuconostoc mesenteroides and Lactobacillus casei synthesize cytoplasmic peptidoglycan precursors that terminate in lactate. J. Bacteriol. 176, 260^264. [20] Ferain, T., Hobbs Jr., J.N., Richardson, J., Bernard, N., Garmyn, D., Hols, P., Allen, N.E. and Delcour, J. (1996) Knockout of the two ldh genes has a major impact on peptidoglycan precursor synthesis in Lactobacillus plantarum. J. Bacteriol. 178, 5431^5437. [21] Elisha, B.G. and Courvalin, P. (1995) Analysis of genes encoding D- alanine:d-alanine ligase-related enzymes in Leuconostoc mesenteroides and Lactobacillus spp.. Gene 152, 79^83. [22] Dutka-Malen, S., Molinas, C., Arthur, M. and Courvalin, P. (1990) The VANA glycopeptide resistance protein is related to D-analyl-Dalanine ligase cell wall biosynthesis enzymes. Mol. Gen. Genet. 224, 364^372. [23] Huygens, F. (1993) Vancomycin binding to cell walls of non-streptococcal vancomycin-resistant bacteria. J. Antimicrob. Chemother. 32, 551^558. [24] Vescovo, M., Morelli, L. and Bottazzi, V. (1982) Drug resistance plasmids in Lactobacillus acidophilus and Lactobacillus reuteri. Appl. Environ. Microbiol. 43, 50^56. [25] Reed, C., Efstratiou, A., Morrison, D. and Woodford, N. (1993) Glycopeptide-resistant Gemella haemolysans from blood. Lancet 342, 927^928. [26] Park, I.S. and Walsh, C.T. (1997) D-Alanyl-D-lactate and D-alanyl- D-alanine synthesis by D-alanyl-D-alanine ligase from vancomycinresistant Leuconostoc mesenteroides. E ects of a phenylalanine 261 to tyrosine mutation. J. Biol. Chem. 272, 9210^9214. [27] Marshall, C.G., Lessard, I.A., Park, I. and Wright, G.D. (1998) Glycopeptide antibiotic resistance genes in glycopeptide-producing organisms. Antimicrob. Agents Chemother. 42, 2215^2220. [28] Dartois, V., Phalip, V., Schmitt, P. and Divies, C. (1995) Puri cation, properties and DNA sequence of the D-lactate dehydrogenase from Leuconostoc mesenteroides subsp. cremoris. Res. Microbiol. 146, 291^ 302. [29] Tynkkynen, S., Singh, K.V. and Varmanen, P. (1998) Vancomycin resistance factor of Lactobacillus rhamnosus GG in relation to enterococcal vancomycin resistance (van) genes. Int. J. Food Microbiol. 41, 195^204. [30] Morel-Deville, F., Fauvel, F. and Morel, P. (1998) Two-component signal-transducing systems involved in stress responses and vancomycin susceptibility in Lactobacillus sakei. Microbiology 144, 2873^ [31] Novak, R., Henriques, B., Charpentier, E., Normark, S. and Tuomanen, E. (1999) Emergence of vancomycin tolerance in Streptococcus pneumoniae. Nature 399, 590^593. [32] Venditti, M., Gelfusa, V., Tarasi, A., Brandimarte, C. and Serra, P. (1990) Antimicrobial susceptibilities of Erysipelothrix rhusiopathiae. Antimicrob. Agents Chemother. 34, 2038^2040. [33] Rippere, K., Patel, R., Uhl, J.R., Piper, K.E., Steckelberg, J.M., Kline, B.C., Cockerill, F.R. and Yousten, A.A. (1998) DNA sequence resembling vana and vanb in the vancomycin-resistant biopesticide Bacillus popilliae. J. Infect. Dis. 178, 584^588. [34] Patel, R., Piper, K., Cockerill, F.C., Steckelberg, J.M. and Yousten, A.A. (2000) The biopesticide Paenibacillus popilliae has a vancomycin resistance gene cluster homologous to the enterococcal VanA vancomycin resistance gene cluster. Antimicrob. Agents Chemother. 44, 705^709. [35] Wu, Y.J., Hong, T.C., Hou, C.J.Y., Chou, Y.S., Tsai, C.H. and Yang, D.I. (1999) Bacillus popilliae endocarditis with prolonged complete heart block. Am. J. Med. Sci. 317, 263^265.
7 R. Patel / FEMS Microbiology Letters 185 (1999) 1^7 7 [36] Coudron, P.E., Payne, J.M. and Markowitz, S.M. (1991) Pneumonia and empyema infection associated with a Bacillus species that resembles B. alvei. J. Clin. Microbiol. 29, 1777^1779. [37] Marshall, C.G. and Wright, G.D. (1997) The glycopeptide antibiotic producer Streptomyces toyocaensis NRRL has both D-alanyl- D-alanine and D-alanyl-D-lactate ligases. FEMS Microbiol. Lett. 157, 295^299. [38] Dingman, D.W. (1999) Conjugative transposition of Tn916 and Tn925 in Bacillus popilliae. Can. J. Microbiol. 45, 530^535. [39] Lessard, I.A.D. and Walsh, C.T. (1999) VanX, a bacterial D-alanyl- D-alanine dipeptidase: Resistance, immunity, or survival function? Proc. Natl. Acad. Sci. USA 96, 11028^ [40] Lessard, I.A., Pratt, S.D., McCa erty, D.G., Bussiere, D.E., Hutchins, C., Wanner, B.L., Katz, L. and Walsh, C.T. (1998) Homologs of the vancomycin resistance D-Ala-D-Ala dipeptidase VanX in Streptomyces toyocaensis, Escherichia coli and Synechocystis: attributes of catalytic e ciency, stereoselectivity and regulation with implications for function. Chem. Biol. 5, 489^504. [41] Healy, V.L., Park, I.S. and Walsh, C.T. (1998) Active-site mutants of the VanC2 D-alanyl-D-serine ligase, characteristic of one vancomycin-resistant bacterial phenotype, revert towards wild-type D-alanyl- D-alanine ligases. Chem. Biol. 5, 197^207. [42] Barnass, S., Holland, K. and Tabaqchali, S. (1991) Vancomycin-resistant Corynebacterium species causing prosthetic valve endocarditis successfully treated with imipenem and cipro oxacin. J. Infect. 22, 161^169. [43] Mory, F., Lozniewski, A., David, V., Carlier, J.P., Dubreuil, L. and Leclercq, R. (1998) Low-level vancomycin resistance in Clostridium innocuum. J. Clin. Microbiol. 36, 1767^1768. [44] Jensen, K.T., Schonheyder, H., Pers, C. and Thomsen, V.F. (1992) In vitro activity of teicoplanin and vancomycin against gram-positive bacteria from human clinical and veterinary sources. Apmis 100, 543^552. [45] Centers for Disease Control (1997) Update: Staphylococcus aureus with reduced susceptibility to vancomycin - United States, MMWR Morbidity and Mortality Weekly Report 46, 813^815. [46] Hanaki, H., Inaba, Y., Sasaki, K. and Hiramatsu, K. (1998) A novel method of detecting Staphylococcus aureus heterogeneously resistant to vancomycin (hetero-vrsa). Jpn. J. Antibiot. 51, 521^530. [47] Yoshimura, H., Ishimaru, M., Endoh, Y.S., Suginaka, M. and Yamatani, S. (1998) Isolation of glycopeptide-resistant enterococci from chickens in Japan. Antimicrob. Agents Chemother. 42, [48] Bager, F., Madsen, M., Christensen, J. and Aarestrup, F.M. (1997) Avoparcin used as a growth promoter is associated with the occurrence of vancomycin-resistant Enterococcus faecium on Danish poultry and pig farms. Prev. Vet. Med. 31, 95^112. [49] Ostrowsky, B.E. et al. (1999) A cluster of VanD vancomycin-resistant Enterococcus faecium: molecular characterization and epidemiology. J. Infect. Dis. 180, 1177^1185. [50] Arthur, M., Depardieu, F., Molinas, C., Reynolds, P. and Courvalin, P. (1995) The vanz gene of Tn1546 from Enterococcus faecium BM4147 confers resistance to teicoplanin. Gene 154, 87^92. [51] Tatusova, T.A. and Madden, T.L. (1999) Blast 2 sequences - a new tool for comparing protein and nucleotide sequences. FEMS Microbiol Lett. 174, 247^250.
Spectrum of vancomycin and susceptibility testing
 Spectrum of vancomycin and susceptibility testing Olivier Denis Service de Microbiologie Laboratoire de bactériologie Service de Microbiologie Hôpital Erasme Glycopeptides Vancomycin 1958 < Amycolatopsis
Spectrum of vancomycin and susceptibility testing Olivier Denis Service de Microbiologie Laboratoire de bactériologie Service de Microbiologie Hôpital Erasme Glycopeptides Vancomycin 1958 < Amycolatopsis
Moderate-Level Resistance to Glycopeptide LY Mediated by Genes of the vana and vanb Clusters in Enterococci
 ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, Aug. 1999, p. 1875 1880 Vol. 43, No. 8 0066-4804/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. Moderate-Level Resistance to
ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, Aug. 1999, p. 1875 1880 Vol. 43, No. 8 0066-4804/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. Moderate-Level Resistance to
Received 21 April 1997/Returned for modification 30 June 1997/Accepted 28 August 1997
 JOURNAL OF CLINICAL MICROBIOLOGY, Dec. 1997, p. 3258 3263 Vol. 35, No. 12 0095-1137/97/$04.00 0 Copyright 1997, American Society for Microbiology Comparison of Agar Dilution, Broth Microdilution, E-Test,
JOURNAL OF CLINICAL MICROBIOLOGY, Dec. 1997, p. 3258 3263 Vol. 35, No. 12 0095-1137/97/$04.00 0 Copyright 1997, American Society for Microbiology Comparison of Agar Dilution, Broth Microdilution, E-Test,
Vancomycin is a member of the glycopeptide class of antibiotics. Vancomycin resistance
 Chapter # Vancomycin Resistance VanS/VanR Two-Component omponent Systems Hee Jeon Hong, Matthew I. Hutchings and Mark J. Buttner* Abstract Vancomycin is a member of the glycopeptide class of antibiotics.
Chapter # Vancomycin Resistance VanS/VanR Two-Component omponent Systems Hee Jeon Hong, Matthew I. Hutchings and Mark J. Buttner* Abstract Vancomycin is a member of the glycopeptide class of antibiotics.
Peptidoglycan Structure of Lactobacillus casei, a Species Highly Resistant to Glycopeptide Antibiotics
 JOURNAL OF BACTERIOLOGY, Oct. 1997, p. 6208 6212 Vol. 179, No. 19 0021-9193/97/$04.00 0 Copyright 1997, American Society for Microbiology Peptidoglycan Structure of Lactobacillus casei, a Species Highly
JOURNAL OF BACTERIOLOGY, Oct. 1997, p. 6208 6212 Vol. 179, No. 19 0021-9193/97/$04.00 0 Copyright 1997, American Society for Microbiology Peptidoglycan Structure of Lactobacillus casei, a Species Highly
A Novel Lantibiotic Acting on Bacterial Cell Wall Synthesis Produced by the Uncommon FLAVIA MARINELLI. DBSM, University of Insubria, Varese Italy
 A Novel Lantibiotic Acting on Bacterial Cell Wall Synthesis Produced by the Uncommon Actinomycete Planomonospora sp. FLAVIA MARINELLI DBSM, University of Insubria, Varese Italy Vicuron Pharmaceuticals,
A Novel Lantibiotic Acting on Bacterial Cell Wall Synthesis Produced by the Uncommon Actinomycete Planomonospora sp. FLAVIA MARINELLI DBSM, University of Insubria, Varese Italy Vicuron Pharmaceuticals,
Resistance to commonly used antibiotics is a major clinical
 The molecular basis of vancomycin resistance in clinically relevant Enterococci: Crystal structure of D-alanyl-D-lactate ligase (VanA) David I. Roper*, Trevor Huyton, Alexei Vagin, and Guy Dodson York
The molecular basis of vancomycin resistance in clinically relevant Enterococci: Crystal structure of D-alanyl-D-lactate ligase (VanA) David I. Roper*, Trevor Huyton, Alexei Vagin, and Guy Dodson York
ORIGINAL ARTICLE. Italy
 ORIGINAL ARTICLE Proficiency of Italian clinical laboratories in detecting reduced glycopeptide susceptibility in Enterococcus and Staphylococcus spp. using routine laboratory methodologies M. E. Jones
ORIGINAL ARTICLE Proficiency of Italian clinical laboratories in detecting reduced glycopeptide susceptibility in Enterococcus and Staphylococcus spp. using routine laboratory methodologies M. E. Jones
Resistance to new anti-grampositive. Roland Leclercq, Microbiology, CHU Cote de Nacre, Caen, France
 Resistance to new anti-grampositive agents Roland Leclercq, Microbiology, CHU Cote de Nacre, Caen, France Recently available antimicrobials against MDR Gram-positive infections Cyclic lipopeptide: daptomycin
Resistance to new anti-grampositive agents Roland Leclercq, Microbiology, CHU Cote de Nacre, Caen, France Recently available antimicrobials against MDR Gram-positive infections Cyclic lipopeptide: daptomycin
Surveillance of Enterococci in Belgium. M. Ieven, K. Loens, B. Jans and H. Goossens
 Surveillance of Enterococci in Belgium M. Ieven, K. Loens, B. Jans and H. Goossens Surveillance of Enterococci in Belgium Overview Introduction and epidemiological surveillance Results of isolates received
Surveillance of Enterococci in Belgium M. Ieven, K. Loens, B. Jans and H. Goossens Surveillance of Enterococci in Belgium Overview Introduction and epidemiological surveillance Results of isolates received
SURVEILLANCE FOR GLYCOPEPTIDE-RESISTANT ENTEROCOCCI
 SURVEILLANCE FOR GLYCOPEPTIDE-RESISTANT ENTEROCOCCI Drs N Bosman, T Nana & C Sriruttan CMID NHLS Dr Charlotte Sriruttan SASCM 3/11 GLOBAL DATA VRE first isolated in Europe in 1987, (Leclercq R., et al
SURVEILLANCE FOR GLYCOPEPTIDE-RESISTANT ENTEROCOCCI Drs N Bosman, T Nana & C Sriruttan CMID NHLS Dr Charlotte Sriruttan SASCM 3/11 GLOBAL DATA VRE first isolated in Europe in 1987, (Leclercq R., et al
The Activity of Glycopeptide Antibiotics Against Resistant Bacteria Correlates with their Ability to
 AAC Accepts, published online ahead of print on August 1 Antimicrob. Agents Chemother. doi:1.118/aac.38-1 Copyright 1, American Society for Microbiology. All Rights Reserved. 1 3 The Activity of Glycopeptide
AAC Accepts, published online ahead of print on August 1 Antimicrob. Agents Chemother. doi:1.118/aac.38-1 Copyright 1, American Society for Microbiology. All Rights Reserved. 1 3 The Activity of Glycopeptide
Resistance to linezolid in enterococci and staphylococci referred to the national reference laboratory
 Resistance to linezolid in enterococci and staphylococci referred to the national reference laboratory Dr Danièle Meunier, AMRHAI BSAC - Antimicrobial Susceptibility Testing User Days Oxazolidinone Linezolid
Resistance to linezolid in enterococci and staphylococci referred to the national reference laboratory Dr Danièle Meunier, AMRHAI BSAC - Antimicrobial Susceptibility Testing User Days Oxazolidinone Linezolid
Antibiotic treatment of streptococcal and enterococcal endocarditis: an overview
 European Heart Journal (1995) 16 {Supplement B), 75-79 Antibiotic treatment of streptococcal and enterococcal endocarditis: an overview P. FRANCIOLI Division of Hospital Preventative Medicine and Department
European Heart Journal (1995) 16 {Supplement B), 75-79 Antibiotic treatment of streptococcal and enterococcal endocarditis: an overview P. FRANCIOLI Division of Hospital Preventative Medicine and Department
Development of C sporins. Beta-lactam antibiotics - Cephalosporins. Second generation C sporins. Targets - PBP s
 Beta-lactam antibiotics - Cephalosporins Development of C sporins Targets - PBP s Activity - Cidal - growing organisms (like the penicillins) Principles of action - Affinity for PBP s Permeability properties
Beta-lactam antibiotics - Cephalosporins Development of C sporins Targets - PBP s Activity - Cidal - growing organisms (like the penicillins) Principles of action - Affinity for PBP s Permeability properties
New Insights into Peptidoglycan Biosynthesis. Louis B. Rice Warren Alpert Medical School of Brown University Providence, RI
 New Insights into Peptidoglycan Biosynthesis Louis B. Rice Warren Alpert Medical School of Brown University Providence, RI Outline of Presentation Review role of penicillin-binding proteins in cell wall
New Insights into Peptidoglycan Biosynthesis Louis B. Rice Warren Alpert Medical School of Brown University Providence, RI Outline of Presentation Review role of penicillin-binding proteins in cell wall
ResFinder 4.0: towards in silico antibiograms
 ResFinder 4.0: towards in silico antibiograms Valeria Bortolaia, DVM, PhD Research group for Genomic Epidemiology National Food Institute Technical University of Denmark The path to WGS-based AMR prediction
ResFinder 4.0: towards in silico antibiograms Valeria Bortolaia, DVM, PhD Research group for Genomic Epidemiology National Food Institute Technical University of Denmark The path to WGS-based AMR prediction
Home Page Table of Contents Reviews Editorial Board Contact Us
 of 1 http://www.antimicrobe.org/ 31/01/2010 09:32 AM Home Page Table of Contents Reviews Editorial Board Contact Us Staphylococcus aureus Review of C. difficile Therapy Chagas Disease What's New Review
of 1 http://www.antimicrobe.org/ 31/01/2010 09:32 AM Home Page Table of Contents Reviews Editorial Board Contact Us Staphylococcus aureus Review of C. difficile Therapy Chagas Disease What's New Review
JAC Increase in glutamine-non-amidated muropeptides in the peptidoglycan of vancomycin-resistant Staphylococcus aureus strain Mu50
 Journal of Antimicrobial Chemotherapy (1998) 42, 315 320 JAC Increase in glutamine-non-amidated muropeptides in the peptidoglycan of vancomycin-resistant Staphylococcus aureus strain Mu50 H. Hanaki a,
Journal of Antimicrobial Chemotherapy (1998) 42, 315 320 JAC Increase in glutamine-non-amidated muropeptides in the peptidoglycan of vancomycin-resistant Staphylococcus aureus strain Mu50 H. Hanaki a,
In Vitro Susceptibilities of Aerobic and Facultative Non-Spore- Forming Gram-Positive Bacilli to HMR 3647 (RU 66647) and 14 Other Antimicrobials
 ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, May 1998, p. 1028 1033 Vol. 42, No. 5 0066-4804/98/$04.00 0 Copyright 1998, American Society for Microbiology In Vitro Susceptibilities of Aerobic and Facultative
ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, May 1998, p. 1028 1033 Vol. 42, No. 5 0066-4804/98/$04.00 0 Copyright 1998, American Society for Microbiology In Vitro Susceptibilities of Aerobic and Facultative
Rapid identification and resistance assessment: The future is mass spectrometry
 Rapid identification and resistance assessment: The future is mass spectrometry Dr Sanmarié Schlebusch Director of Microbiology Mater Pathology Brisbane Outline Introduction Plug and play Pre-prep and
Rapid identification and resistance assessment: The future is mass spectrometry Dr Sanmarié Schlebusch Director of Microbiology Mater Pathology Brisbane Outline Introduction Plug and play Pre-prep and
CLINICAL USE OF GLYCOPEPTIDES. Herbert Spapen Intensive Care Department University Hospital Vrije Universiteit Brussel
 CLINICAL USE OF GLYCOPEPTIDES Herbert Spapen Intensive Care Department University Hospital Vrije Universiteit Brussel Glycopeptides Natural Vancomycin introduced in 1958 Teicoplanin introduced in Europe
CLINICAL USE OF GLYCOPEPTIDES Herbert Spapen Intensive Care Department University Hospital Vrije Universiteit Brussel Glycopeptides Natural Vancomycin introduced in 1958 Teicoplanin introduced in Europe
Steven D. Brown* and Maria M. Traczewski. The Clinical Microbiology Institute, 9725 SW Commerce Circle, Wilsonville, Oregon 97070
 ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, May 2010, p. 2063 2069 Vol. 54, No. 5 0066-4804/10/$12.00 doi:10.1128/aac.01569-09 Copyright 2010, American Society for Microbiology. All Rights Reserved. Comparative
ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, May 2010, p. 2063 2069 Vol. 54, No. 5 0066-4804/10/$12.00 doi:10.1128/aac.01569-09 Copyright 2010, American Society for Microbiology. All Rights Reserved. Comparative
השפעת חיידקים פרוביוטיים
 השפעת חיידקים פרוביוטיים החיים בחלל )המעי(... על רון שאול יחידת גסטרו ילדים מרכז רפואי רמב"ם Introduction The intestinal microflora primarily in the large bowel consists mostly on benign bacterial species
השפעת חיידקים פרוביוטיים החיים בחלל )המעי(... על רון שאול יחידת גסטרו ילדים מרכז רפואי רמב"ם Introduction The intestinal microflora primarily in the large bowel consists mostly on benign bacterial species
Antimicrobial resistance and virulence genes in bacteria
 Antimicrobial resistance and virulence genes in bacteria Johanne Ahrenfeldt PhD Student BSU course - November 2016 1 How to analyse WGS data part 2 an introduction to CGEs methods KmerFinder SpeciesFinder
Antimicrobial resistance and virulence genes in bacteria Johanne Ahrenfeldt PhD Student BSU course - November 2016 1 How to analyse WGS data part 2 an introduction to CGEs methods KmerFinder SpeciesFinder
AMR prediction based on WGS data
 AMR prediction based on WGS data Valeria Bortolaia, DVM, PhD Research Group for Genomic Epidemiology National Food Institute Technical University of Denmark This lecture How to use WGS for AMR surveillance?
AMR prediction based on WGS data Valeria Bortolaia, DVM, PhD Research Group for Genomic Epidemiology National Food Institute Technical University of Denmark This lecture How to use WGS for AMR surveillance?
First detection of the mcr-1 gene in Escherichia coli isolated from livestock between 2013
 AAC Accepted Manuscript Posted Online 29 August 2016 Antimicrob. Agents Chemother. doi:10.1128/aac.01472-16 Copyright 2016, American Society for Microbiology. All Rights Reserved. 1 2 First detection of
AAC Accepted Manuscript Posted Online 29 August 2016 Antimicrob. Agents Chemother. doi:10.1128/aac.01472-16 Copyright 2016, American Society for Microbiology. All Rights Reserved. 1 2 First detection of
THE INCIDENCE of infections caused by
 Vancomycin Prescribing Practices in Hospitalized Chronic Hemodialysis Patients Kevin Green, MD, Gerald Schulman, MD, David W. Haas, MD, William Schaffner, MD, and Erika M.C. D Agata, MD, MPH To determine
Vancomycin Prescribing Practices in Hospitalized Chronic Hemodialysis Patients Kevin Green, MD, Gerald Schulman, MD, David W. Haas, MD, William Schaffner, MD, and Erika M.C. D Agata, MD, MPH To determine
Clinical Pharmacokinetics and Pharmacodynamics of Telavancin Compared with the Other Glycopeptides
 Clin Pharmacokinet (2018) 57:797 816 https://doi.org/10.1007/s40262-017-0623-4 REVIEW ARTICLE Clinical Pharmacokinetics and Pharmacodynamics of Telavancin Compared with the Other Glycopeptides Valentin
Clin Pharmacokinet (2018) 57:797 816 https://doi.org/10.1007/s40262-017-0623-4 REVIEW ARTICLE Clinical Pharmacokinetics and Pharmacodynamics of Telavancin Compared with the Other Glycopeptides Valentin
ESCMID Online Lecture Library. by author
 Hospital Universitario Virgen Macarena, Seville New drugs against MRSA and VRE L. Eduardo López Cortés Seville, 8th July Tedizolid Oxazolidinone Ceftaroline // Ceftobiprole 5 th gen cephalosporin Overview
Hospital Universitario Virgen Macarena, Seville New drugs against MRSA and VRE L. Eduardo López Cortés Seville, 8th July Tedizolid Oxazolidinone Ceftaroline // Ceftobiprole 5 th gen cephalosporin Overview
Vancomycin Rationale for the EUCAST clinical breakpoints, version June 2010
 Vancomycin Rationale for the EUCAST clinical breakpoints, version 2.1 17 June 2010 Introduction The glycopeptides are a class of agents composed of amino acid residues and attached sugars. Glycopeptides
Vancomycin Rationale for the EUCAST clinical breakpoints, version 2.1 17 June 2010 Introduction The glycopeptides are a class of agents composed of amino acid residues and attached sugars. Glycopeptides
Molecular Characterization of Vancomycin-Resistant Enterococci from Hospitalized Patients and Poultry Products in The Netherlands
 JOURNAL OF CLINICAL MICROBIOLOGY, July 1998, p. 1927 1932 Vol. 36, No. 7 0095-1137/98/$04.00 0 Copyright 1998, American Society for Microbiology. All Rights Reserved. Molecular Characterization of Vancomycin-Resistant
JOURNAL OF CLINICAL MICROBIOLOGY, July 1998, p. 1927 1932 Vol. 36, No. 7 0095-1137/98/$04.00 0 Copyright 1998, American Society for Microbiology. All Rights Reserved. Molecular Characterization of Vancomycin-Resistant
Cubicin (Daptomycin) Priv.Doz. Dr. med. Markus Rothenburger
 Cubicin (Daptomycin) Priv.Doz. Dr. med. Markus Rothenburger Cubicin (Daptomycin): A new generation of antibiotics First-in-class cyclic natural lipopeptide Activity against major gram positive pathogens
Cubicin (Daptomycin) Priv.Doz. Dr. med. Markus Rothenburger Cubicin (Daptomycin): A new generation of antibiotics First-in-class cyclic natural lipopeptide Activity against major gram positive pathogens
Cases in employees. Cases. Day of onset (July)
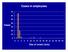 Cases in employees 70 60 50 Cases 40 30 20 10 0 0 2 4 6 8 10 12 14 16 18 20 22 24 26 28 30 Day of onset (July) 70 60 50 40 Cases 30 Cases in employees and visitors Employees Visitors 20 10 0 0 2 4 6 8
Cases in employees 70 60 50 Cases 40 30 20 10 0 0 2 4 6 8 10 12 14 16 18 20 22 24 26 28 30 Day of onset (July) 70 60 50 40 Cases 30 Cases in employees and visitors Employees Visitors 20 10 0 0 2 4 6 8
ALERT. Clinical microbiology considerations related to the emergence of. New Delhi metallo beta lactamases (NDM 1) and Klebsiella
 ALERT Clinical microbiology considerations related to the emergence of New Delhi metallo beta lactamases (NDM 1) and Klebsiella pneumoniae carbapenemases (KPC) amongst hospitalized patients in South Africa
ALERT Clinical microbiology considerations related to the emergence of New Delhi metallo beta lactamases (NDM 1) and Klebsiella pneumoniae carbapenemases (KPC) amongst hospitalized patients in South Africa
TRANSIENT INTESTINAL CARRIAGE AFTER INGESTION OF ANTIBIOTIC-RESISTANT E. FAECIUM FROM MEAT
 TRANSIENT INTESTINAL CARRIAGE AFTER INGESTION OF ANTIBIOTIC-RESISTANT E. FAECIUM FROM MEAT TRANSIENT INTESTINAL CARRIAGE AFTER INGESTION OF ANTIBIOTIC-RESISTANT ENTEROCOCCUS FAECIUM FROM CHICKEN AND PORK
TRANSIENT INTESTINAL CARRIAGE AFTER INGESTION OF ANTIBIOTIC-RESISTANT E. FAECIUM FROM MEAT TRANSIENT INTESTINAL CARRIAGE AFTER INGESTION OF ANTIBIOTIC-RESISTANT ENTEROCOCCUS FAECIUM FROM CHICKEN AND PORK
(multidrug-resistant Pseudomonas aeruginosa; MDRP)
 220 2009 (multidrug-resistant Pseudomonas aeruginosa; MDRP) 21 4 1 21 10 4 amikacin (AMK), imipenem/cilastatin (IPM), ciprofloxacin (CPFX) multidrug-resistant Pseudomonas aeruginosa (MDRP) CHROMagar TM
220 2009 (multidrug-resistant Pseudomonas aeruginosa; MDRP) 21 4 1 21 10 4 amikacin (AMK), imipenem/cilastatin (IPM), ciprofloxacin (CPFX) multidrug-resistant Pseudomonas aeruginosa (MDRP) CHROMagar TM
Industrial biotechnology agustin krisna wardani
 Industrial biotechnology agustin krisna wardani Nutraceuticals The term Nutraceuticals, launched by Stephen De-Felici in the 1980s A food or part of a food that may provide medicinal or health benefits,
Industrial biotechnology agustin krisna wardani Nutraceuticals The term Nutraceuticals, launched by Stephen De-Felici in the 1980s A food or part of a food that may provide medicinal or health benefits,
Enhanced EARS-Net Surveillance 2017 First Half
 1 Enhanced EARS-Net Surveillance 2017 First Half In this report Main results for 2017, first half Breakdown of factors by organism and resistance subtype Device-association Data quality assessment Key
1 Enhanced EARS-Net Surveillance 2017 First Half In this report Main results for 2017, first half Breakdown of factors by organism and resistance subtype Device-association Data quality assessment Key
The microbial diagnosis of infective endocarditis (IE)
 The microbial diagnosis of infective endocarditis (IE) Pierrette Melin Medical Microbiology pm-chulg sbimc 10.05.2007 1 Introduction for diagnosis Review of microbiological investigation of IE and perspectives
The microbial diagnosis of infective endocarditis (IE) Pierrette Melin Medical Microbiology pm-chulg sbimc 10.05.2007 1 Introduction for diagnosis Review of microbiological investigation of IE and perspectives
RNA Processing in Eukaryotes *
 OpenStax-CNX module: m44532 1 RNA Processing in Eukaryotes * OpenStax This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section, you
OpenStax-CNX module: m44532 1 RNA Processing in Eukaryotes * OpenStax This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section, you
Macrolides, Clindamycin & Ketolides Polymyxins
 Macrolides, Clindamycin & Ketolides Polymyxins Kwan Soo Ko Macrolides - Erythromycin - Azithromycin - Clarithromycin Lincosamides - Lincomycin - Clindamycin Unrelated chemically But, many similar biological
Macrolides, Clindamycin & Ketolides Polymyxins Kwan Soo Ko Macrolides - Erythromycin - Azithromycin - Clarithromycin Lincosamides - Lincomycin - Clindamycin Unrelated chemically But, many similar biological
Shirin Abadi, B.Sc.(Pharm.), ACPR, Pharm.D. Clinical Pharmacy Specialist & Pharmacy Education Coordinator, BC Cancer Agency Clinical Associate
 Shirin Abadi, B.Sc.(Pharm.), ACPR, Pharm.D. Clinical Pharmacy Specialist & Pharmacy Education Coordinator, BC Cancer Agency Clinical Associate Professor of Pharmacy & Associate Member of Medicine, UBC
Shirin Abadi, B.Sc.(Pharm.), ACPR, Pharm.D. Clinical Pharmacy Specialist & Pharmacy Education Coordinator, BC Cancer Agency Clinical Associate Professor of Pharmacy & Associate Member of Medicine, UBC
10/4/16. mcr-1. Emerging Resistance Updates. Objectives. National Center for Emerging and Zoonotic Infectious Diseases. Alex Kallen, MD, MPH, FACP
 National Center for Emerging and Zoonotic Infectious Diseases Emerging Resistance Updates Alex Kallen, MD, MPH, FACP Lead Antimicrobial Resistance and Emerging Pathogens Team Prevention and Response Branch
National Center for Emerging and Zoonotic Infectious Diseases Emerging Resistance Updates Alex Kallen, MD, MPH, FACP Lead Antimicrobial Resistance and Emerging Pathogens Team Prevention and Response Branch
From the Structure and Function of the Ribosome to new Antibiotics
 From the Structure and Function of the Ribosome to new Antibiotics Crick s central dogma of molecular biology: DNA makes DNA makes RNA makes protein Jim Watson, 1964 J.A. Lake, 1976 (J.M.B. 105, 131)
From the Structure and Function of the Ribosome to new Antibiotics Crick s central dogma of molecular biology: DNA makes DNA makes RNA makes protein Jim Watson, 1964 J.A. Lake, 1976 (J.M.B. 105, 131)
Infections Amenable to OPAT. (Nabin Shrestha + Ajay Mathur)
 3 Infections Amenable to OPAT (Nabin Shrestha + Ajay Mathur) Decisions regarding outpatient treatment of infections vary with the institution, the prescribing physician, the individual patient s condition
3 Infections Amenable to OPAT (Nabin Shrestha + Ajay Mathur) Decisions regarding outpatient treatment of infections vary with the institution, the prescribing physician, the individual patient s condition
Practice Problems 8. a) What do we define as a beneficial or advantageous mutation to the virus? Why?
 Life Sciences 1a Practice Problems 8 1. You have two strains of HIV one is a wild type strain of HIV and the second has acquired a mutation in the gene encoding the protease. This mutation has a dual effect
Life Sciences 1a Practice Problems 8 1. You have two strains of HIV one is a wild type strain of HIV and the second has acquired a mutation in the gene encoding the protease. This mutation has a dual effect
Resistance to Polymyxins in France
 Resistance to Polymyxins in France Paris Prof. Patrice Nordmann NDM producers in Enterobacteriaceae The polymyxins; colistin and polymyxin B Colistin - Synthesis by Bacillus polymyxa spp colistinus -
Resistance to Polymyxins in France Paris Prof. Patrice Nordmann NDM producers in Enterobacteriaceae The polymyxins; colistin and polymyxin B Colistin - Synthesis by Bacillus polymyxa spp colistinus -
biological mechanisms Streptococcus pneumoniae virulence factors Mycobacterium tuberculosis pathogenesis Bacillus anthracis vaccine development
 http://www.microbio.uab.edu/ Bacteriology at UAB Streptococcus pneumoniae Mycobacterium tuberculosis Bacillus anthracis Mycoplasma biological mechanisms virulence factors pathogenesis vaccine development
http://www.microbio.uab.edu/ Bacteriology at UAB Streptococcus pneumoniae Mycobacterium tuberculosis Bacillus anthracis Mycoplasma biological mechanisms virulence factors pathogenesis vaccine development
Bio Microbiology - Spring 2013 Study Guide 20 Normal Flora - Normal Responses - Disease
 Bio 230 - Microbiology - Spring 2013 Study Guide 20 Normal Flora - Normal Responses - Disease It has been calculated that the normal human houses about 10 12 bacteria on the skin, 10 10 in the mouth, and
Bio 230 - Microbiology - Spring 2013 Study Guide 20 Normal Flora - Normal Responses - Disease It has been calculated that the normal human houses about 10 12 bacteria on the skin, 10 10 in the mouth, and
Antibiotic treatment comparison in patients with diarrhea
 Original Research Article Antibiotic treatment comparison in patients with diarrhea Deva Lal Kast * Senior Consultant Physician, Department of General Medicine, Krishna Hospital, Ex senior Specialist and
Original Research Article Antibiotic treatment comparison in patients with diarrhea Deva Lal Kast * Senior Consultant Physician, Department of General Medicine, Krishna Hospital, Ex senior Specialist and
In Vitro Susceptibility of Staphylococci and Enterococci to Vancomycin and Teicoplanin
 In Vitro Susceptibility of Staphylococci and Enterococci to Vancomycin and Teicoplanin 19 A. Guzek, K. Korzeniewski, Aneta Nitsch-Osuch, Z. Rybicki, and E. Prokop Abstract Hospital-acquired infections
In Vitro Susceptibility of Staphylococci and Enterococci to Vancomycin and Teicoplanin 19 A. Guzek, K. Korzeniewski, Aneta Nitsch-Osuch, Z. Rybicki, and E. Prokop Abstract Hospital-acquired infections
of glycopeptide-resistant enterococci
 ~ ORIGINAL ARTICLE Comparison of agar-based media for primary isolation of glycopeptide-resistant enterococci l? R. Chadwickl, D. E]. Brown2, M. H. Wilcox2, T A. Collynsl, E. Walpo1e2, J. Dillon, R. Smith2,
~ ORIGINAL ARTICLE Comparison of agar-based media for primary isolation of glycopeptide-resistant enterococci l? R. Chadwickl, D. E]. Brown2, M. H. Wilcox2, T A. Collynsl, E. Walpo1e2, J. Dillon, R. Smith2,
Gentilucci, Amino Acids, Peptides, and Proteins. Peptides and proteins are polymers of amino acids linked together by amide bonds CH 3
 Amino Acids Peptides and proteins are polymers of amino acids linked together by amide bonds Aliphatic Side-Chain Amino Acids - - H CH glycine alanine 3 proline valine CH CH 3 - leucine - isoleucine CH
Amino Acids Peptides and proteins are polymers of amino acids linked together by amide bonds Aliphatic Side-Chain Amino Acids - - H CH glycine alanine 3 proline valine CH CH 3 - leucine - isoleucine CH
What is new in EUCAST South Africa, May, Gunnar Kahlmeter EUCAST Technical Data Coordinator and Webmaster Sweden
 What is new in EUCAST 2016 17 South Africa, May, 2016 Gunnar Kahlmeter EUCAST Technical Data Coordinator and Webmaster Sweden What is new in EUCAST 2016/17? New organisms with breakpoints (Addendum 2016)
What is new in EUCAST 2016 17 South Africa, May, 2016 Gunnar Kahlmeter EUCAST Technical Data Coordinator and Webmaster Sweden What is new in EUCAST 2016/17? New organisms with breakpoints (Addendum 2016)
2018 CNISP HAI Surveillance Case definitions
 2018 CNISP HAI Surveillance Case definitions The following case definitions for the surveillance of healthcare-associated infections (HAIs) are used by all acute-care hospitals that participate in the
2018 CNISP HAI Surveillance Case definitions The following case definitions for the surveillance of healthcare-associated infections (HAIs) are used by all acute-care hospitals that participate in the
GLYCOPEPTIDES : how can a structural modification bring to a new life an old family of antibiotics?
 GLYCPEPTIDES : how can a structural modification bring to a new life an old family of antibiotics? F. Van Bambeke Pharmacologie cellulaire et moléculaire Université catholique de Louvain Brussels - Belgium
GLYCPEPTIDES : how can a structural modification bring to a new life an old family of antibiotics? F. Van Bambeke Pharmacologie cellulaire et moléculaire Université catholique de Louvain Brussels - Belgium
Received 31 July 1992/Accepted 12 November 1992
 ANTMCROBAL AGENTS AND CHEMOTHERAPY, Feb 1993, p 32-36 66-8/93/232-5$2/ Copyright 1993, American Society for Microbiology Vol 37, No 2 Abnormal Peptidoglycan Produced in a Methicillin-Resistant Strain of
ANTMCROBAL AGENTS AND CHEMOTHERAPY, Feb 1993, p 32-36 66-8/93/232-5$2/ Copyright 1993, American Society for Microbiology Vol 37, No 2 Abnormal Peptidoglycan Produced in a Methicillin-Resistant Strain of
Multi-clonal origin of macrolide-resistant Mycoplasma pneumoniae isolates. determined by multiple-locus variable-number tandem-repeat analysis
 JCM Accepts, published online ahead of print on 30 May 2012 J. Clin. Microbiol. doi:10.1128/jcm.00678-12 Copyright 2012, American Society for Microbiology. All Rights Reserved. 1 2 Multi-clonal origin
JCM Accepts, published online ahead of print on 30 May 2012 J. Clin. Microbiol. doi:10.1128/jcm.00678-12 Copyright 2012, American Society for Microbiology. All Rights Reserved. 1 2 Multi-clonal origin
Supporting Information
 Supporting Information Discovery of linear low-cationic peptides to target methicillin-resistant Staphylococcus aureus in vivo Yuan Liu, Meirong Song, Shuangyang Ding,, Kui Zhu*,,, Beijing Advanced Innovation
Supporting Information Discovery of linear low-cationic peptides to target methicillin-resistant Staphylococcus aureus in vivo Yuan Liu, Meirong Song, Shuangyang Ding,, Kui Zhu*,,, Beijing Advanced Innovation
42 Metabolic engineering of lactic acid bacteria for the improvement of fermented dairy products
 42 Metabolic engineering of lactic acid bacteria for the improvement of fermented dairy products J. Hugenholtz, M. Starrenburg, I. Boels, W. Sybesma, A.C. Chaves, A. Mertens and M. Kleerebezem Wageningen
42 Metabolic engineering of lactic acid bacteria for the improvement of fermented dairy products J. Hugenholtz, M. Starrenburg, I. Boels, W. Sybesma, A.C. Chaves, A. Mertens and M. Kleerebezem Wageningen
h g o Klare, Dietlitide Badstiibner, Carola Koristabel atid WoEfgang Witte
 ORIGINAL ARTICLE Identification of enterococci and determination of their glycopeptide resistance in German and Austrian clinical microbiology laboratories Clin Microbial Infect 1999; 5: 535-539 h g o
ORIGINAL ARTICLE Identification of enterococci and determination of their glycopeptide resistance in German and Austrian clinical microbiology laboratories Clin Microbial Infect 1999; 5: 535-539 h g o
Probiotics for Primary Prevention of Clostridium difficile Infection
 Probiotics for Primary Prevention of Clostridium difficile Infection Objectives Review risk factors for Clostridium difficile infection (CDI) Describe guideline recommendations for CDI prevention Discuss
Probiotics for Primary Prevention of Clostridium difficile Infection Objectives Review risk factors for Clostridium difficile infection (CDI) Describe guideline recommendations for CDI prevention Discuss
Mupirocin Rationale for the EUCAST clinical breakpoints, version th July 2010
 Mupirocin Rationale for the EUCAST clinical breakpoints, version 1.0 6 th July 2010 Foreword EUCAST The European Committee on Antimicrobial Susceptibility Testing (EUCAST) is organised by the European
Mupirocin Rationale for the EUCAST clinical breakpoints, version 1.0 6 th July 2010 Foreword EUCAST The European Committee on Antimicrobial Susceptibility Testing (EUCAST) is organised by the European
Vancomycin resistance in Staphylococcus aureus
 INTERNATIONAL NEWSLETTER no 3 SEPTEMBER 2002 STATE-OF-THE-ART THE BIOMERIEUX SOLUTION DID YOU KNOW? PRACTICAL ADVICE Vancomycin resistance in Staphylococcus aureus VITEK 2 performance Web site 10 th ISSSI
INTERNATIONAL NEWSLETTER no 3 SEPTEMBER 2002 STATE-OF-THE-ART THE BIOMERIEUX SOLUTION DID YOU KNOW? PRACTICAL ADVICE Vancomycin resistance in Staphylococcus aureus VITEK 2 performance Web site 10 th ISSSI
Application of real-time PCR for the detection of macrolide resistance in Belgian M. pneumoniae strains
 Application of real-time PCR for the detection of macrolide resistance in Belgian M. pneumoniae strains K. Loens, S. C. Masha,, K. Lagrou, H. Goossens and M. Ieven Introduction (1) One of the smallest
Application of real-time PCR for the detection of macrolide resistance in Belgian M. pneumoniae strains K. Loens, S. C. Masha,, K. Lagrou, H. Goossens and M. Ieven Introduction (1) One of the smallest
Heavy metals as alternatives to antibiotics: Panacea or Pandora s Box?
 Heavy metals as alternatives to antibiotics: Panacea or Pandora s Box? International Symposium on Alternatives to Antibiotics Challenges and Solutions in Animal Production Headquarters of the OIE Paris,
Heavy metals as alternatives to antibiotics: Panacea or Pandora s Box? International Symposium on Alternatives to Antibiotics Challenges and Solutions in Animal Production Headquarters of the OIE Paris,
INTESTINAL MICROBIOTA EXAMPLES OF INDIVIDUAL ANALYSES
 EXAMPLES OF INDIVIDUAL ANALYSES INTESTINAL MICROBIOTA Microbiota in the animal or human intestine has evolved together with the host. Consequently, the gastrointestinal tract could be considered a metacommunity,
EXAMPLES OF INDIVIDUAL ANALYSES INTESTINAL MICROBIOTA Microbiota in the animal or human intestine has evolved together with the host. Consequently, the gastrointestinal tract could be considered a metacommunity,
Carbapenemase Producing Enterobacteriaceae: Screening
 Carbapenemase Producing Enterobacteriaceae: Screening Dr David Harvey Consultant Microbiology and Infection Prevention and Control Nov 2015 Aims Is CPE a problem? Does screening have the potential to help?
Carbapenemase Producing Enterobacteriaceae: Screening Dr David Harvey Consultant Microbiology and Infection Prevention and Control Nov 2015 Aims Is CPE a problem? Does screening have the potential to help?
Endocardite infectieuse
 Endocardite infectieuse 1. Raccourcir le traitement: jusqu où? 2. Proposer un traitement ambulatoire: à partir de quand? Endocardite infectieuse A B 90 P = 0.014 20 P = 0.0005 % infective endocarditis
Endocardite infectieuse 1. Raccourcir le traitement: jusqu où? 2. Proposer un traitement ambulatoire: à partir de quand? Endocardite infectieuse A B 90 P = 0.014 20 P = 0.0005 % infective endocarditis
FIRST BIOCHEMISTRY EXAM Tuesday 25/10/ MCQs. Location : 102, 105, 106, 301, 302
 FIRST BIOCHEMISTRY EXAM Tuesday 25/10/2016 10-11 40 MCQs. Location : 102, 105, 106, 301, 302 The Behavior of Proteins: Enzymes, Mechanisms, and Control General theory of enzyme action, by Leonor Michaelis
FIRST BIOCHEMISTRY EXAM Tuesday 25/10/2016 10-11 40 MCQs. Location : 102, 105, 106, 301, 302 The Behavior of Proteins: Enzymes, Mechanisms, and Control General theory of enzyme action, by Leonor Michaelis
SUMMARY OF PRODUCT CHARACTERISTICS
 SUMMARY OF PRODUCT CHARACTERISTICS 1. NAME OF THE VETERINARY MEDICINAL PRODUCT Doxylin 50% WSP, powder for oral solution in pre-ruminant calves, pigs and chickens (BE, BG, NL, RO) Doxylin 50%, powder for
SUMMARY OF PRODUCT CHARACTERISTICS 1. NAME OF THE VETERINARY MEDICINAL PRODUCT Doxylin 50% WSP, powder for oral solution in pre-ruminant calves, pigs and chickens (BE, BG, NL, RO) Doxylin 50%, powder for
Modulation of bacterial community and metabolome in whole crop corn silage by inoculating homo- or heterofermenters
 The 18th International Silage Conference Modulation of bacterial community and metabolome in whole crop corn silage by inoculating homo- or heterofermenters Guo X.S., Xu D.M., Ke W.C., Ding W.R., Zhang
The 18th International Silage Conference Modulation of bacterial community and metabolome in whole crop corn silage by inoculating homo- or heterofermenters Guo X.S., Xu D.M., Ke W.C., Ding W.R., Zhang
Clinical Failure of Vancomycin Treatment of Staphylococcus aureus Infection in a Tertiary Care Hospital in Southern Brazil
 224 BJID 2003; 7 (June) Clinical Failure of Vancomycin Treatment of Staphylococcus aureus Infection in a Tertiary Care Hospital in Southern Brazil Larissa Lutz, Adão Machado, Nadia Kuplich and Afonso Luís
224 BJID 2003; 7 (June) Clinical Failure of Vancomycin Treatment of Staphylococcus aureus Infection in a Tertiary Care Hospital in Southern Brazil Larissa Lutz, Adão Machado, Nadia Kuplich and Afonso Luís
Probiotics: Their Role in Medicine Today. Objectives. Probiotics: What Are They? 11/3/2017
 Probiotics: Their Role in Medicine Today Viki Barr Pharm.D., BCPS AQ ID Assistant Professor, Pharmacy Practice Rosalind Franklin University of Medicine and Science Clinical Pharmacist, Infectious Diseases
Probiotics: Their Role in Medicine Today Viki Barr Pharm.D., BCPS AQ ID Assistant Professor, Pharmacy Practice Rosalind Franklin University of Medicine and Science Clinical Pharmacist, Infectious Diseases
Affinity of Doripenem and Comparators to Penicillin-Binding Proteins in Escherichia coli and ACCEPTED
 AAC Accepts, published online ahead of print on February 00 Antimicrob. Agents Chemother. doi:./aac.01-0 Copyright 00, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights
AAC Accepts, published online ahead of print on February 00 Antimicrob. Agents Chemother. doi:./aac.01-0 Copyright 00, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights
Chapter 14. Bugs that Resist Drugs
 Chapter 14 Bugs that Resist Drugs See website Learning Objectives Important Terminology Power point- posted after chapter is completed What happened to Carlos Don, Rebecca Lohsen, Ricky Lannetti? Carlos
Chapter 14 Bugs that Resist Drugs See website Learning Objectives Important Terminology Power point- posted after chapter is completed What happened to Carlos Don, Rebecca Lohsen, Ricky Lannetti? Carlos
Lecture 2: Virology. I. Background
 Lecture 2: Virology I. Background A. Properties 1. Simple biological systems a. Aggregates of nucleic acids and protein 2. Non-living a. Cannot reproduce or carry out metabolic activities outside of a
Lecture 2: Virology I. Background A. Properties 1. Simple biological systems a. Aggregates of nucleic acids and protein 2. Non-living a. Cannot reproduce or carry out metabolic activities outside of a
Report: antimicrobial resistance in commensal Enterococcus spp. from poultry, pigs, cows and veal calves
 Veterinary and Agrochemical Research Centre Report: antimicrobial resistance in commensal Enterococcus spp. from poultry, pigs, cows and veal calves P. Butaye 1 Introduction Enterococci are regarded as
Veterinary and Agrochemical Research Centre Report: antimicrobial resistance in commensal Enterococcus spp. from poultry, pigs, cows and veal calves P. Butaye 1 Introduction Enterococci are regarded as
THE USE OF THE PENICILLINASE-RESISTANT
 Therapeutic problems THE USE OF THE PENICILLINASE-RESISTANT PENICILLIN IN THE PNEUMONIAS OF CHILDREN MARTHA D. Yow, MARY A. SOUTH AND CHARLES G. HESS From the Department of Pediatrics, Baylor University
Therapeutic problems THE USE OF THE PENICILLINASE-RESISTANT PENICILLIN IN THE PNEUMONIAS OF CHILDREN MARTHA D. Yow, MARY A. SOUTH AND CHARLES G. HESS From the Department of Pediatrics, Baylor University
Identification of vat(e) in Enterococcus faecalis Isolates from Retail Poultry and Its Transferability to Enterococcus faecium
 ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, Dec. 2002, p. 3823 3828 Vol. 46, No. 12 0066-4804/02/$04.00 0 DOI: 10.1128/AAC.46.12.3823 3828.2002 Identification of vat(e) in Enterococcus faecalis Isolates from
ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, Dec. 2002, p. 3823 3828 Vol. 46, No. 12 0066-4804/02/$04.00 0 DOI: 10.1128/AAC.46.12.3823 3828.2002 Identification of vat(e) in Enterococcus faecalis Isolates from
Guidance on screening and confirmation of carbapenem resistant Enterobacteriacae (CRE) December 12, 2011
 Guidance on screening and confirmation of carbapenem resistant Enterobacteriacae (CRE) December 12, 2011 Objectives: To discuss the guidelines for detection of CRE in the laboratory setting. To review
Guidance on screening and confirmation of carbapenem resistant Enterobacteriacae (CRE) December 12, 2011 Objectives: To discuss the guidelines for detection of CRE in the laboratory setting. To review
Molecular mechanisms of antibiotic resistance in Neisseria gonorrhoeae
 medicaldaily.com http://en.vircell.com/diseases ppcorn.com Molecular mechanisms of antibiotic resistance in Neisseria gonorrhoeae Robert Nicholas University of North Carolina at Chapel Hill cdc.gov Timeline
medicaldaily.com http://en.vircell.com/diseases ppcorn.com Molecular mechanisms of antibiotic resistance in Neisseria gonorrhoeae Robert Nicholas University of North Carolina at Chapel Hill cdc.gov Timeline
#Corresponding author: Pathology Department, Singapore General Hospital, 20 College. Road, Academia, Level 7, Diagnostics Tower, , Singapore
 AAC Accepts, published online ahead of print on 21 October 2013 Antimicrob. Agents Chemother. doi:10.1128/aac.01754-13 Copyright 2013, American Society for Microbiology. All Rights Reserved. 1 Title: Escherichia
AAC Accepts, published online ahead of print on 21 October 2013 Antimicrob. Agents Chemother. doi:10.1128/aac.01754-13 Copyright 2013, American Society for Microbiology. All Rights Reserved. 1 Title: Escherichia
New genomic typing method MLST
 New genomic typing method MLST Bon KIMURA fingerprinting PFGE DNA multilocus sequence typingmlst alleles PFGE MLST 1990 PCR 1 PCR DNA PFGE 1 PFGE RAPDrandomly amplified polymorphic DNA 3 AFLPAmplified
New genomic typing method MLST Bon KIMURA fingerprinting PFGE DNA multilocus sequence typingmlst alleles PFGE MLST 1990 PCR 1 PCR DNA PFGE 1 PFGE RAPDrandomly amplified polymorphic DNA 3 AFLPAmplified
Proposals for E.coli surveillance Informatics- proposal for the Infection. VRE molecular epidemiology AMR alerts HPA repatriation Rotavirus
 HPS Update Proposals for E.coli surveillance Informatics- proposal for the Infection Intelligence Platform (IIP) VRE molecular epidemiology AMR alerts HPA repatriation Rotavirus IIP Strategic intent.to
HPS Update Proposals for E.coli surveillance Informatics- proposal for the Infection Intelligence Platform (IIP) VRE molecular epidemiology AMR alerts HPA repatriation Rotavirus IIP Strategic intent.to
Staphylococci. What s to be Covered. Clinical Scenario #1
 Staphylococci Micrococcus, which, when limited in its extent and activity, causes acute suppurative inflammation (phlegmon), produces, when more extensive and intense in its action on the human system,
Staphylococci Micrococcus, which, when limited in its extent and activity, causes acute suppurative inflammation (phlegmon), produces, when more extensive and intense in its action on the human system,
Chem 135: First Midterm
 Chem 135: First Midterm September 28 th, 2007 Please provide all answers in the space provided. Extra paper is available if needed. You may not use calculators for this exam, but you are free to use (previously
Chem 135: First Midterm September 28 th, 2007 Please provide all answers in the space provided. Extra paper is available if needed. You may not use calculators for this exam, but you are free to use (previously
EDUCATIONAL COMMENTARY CLOSTRIDIUM DIFFICILE UPDATE
 EDUCATIONAL COMMENTARY CLOSTRIDIUM DIFFICILE UPDATE Educational commentary is provided through our affiliation with the American Society for Clinical Pathology (ASCP). To obtain FREE CME/CMLE credits click
EDUCATIONAL COMMENTARY CLOSTRIDIUM DIFFICILE UPDATE Educational commentary is provided through our affiliation with the American Society for Clinical Pathology (ASCP). To obtain FREE CME/CMLE credits click
Introduction. Crowded environments where the air is re-circulated can often be heavily infected with unseen germs and viruses.
 Introduction The continued increase in urbanisation,, population growth and global travel means germs and viruses can spread faster and further than ever before. Air, sea, rail and road travel have never
Introduction The continued increase in urbanisation,, population growth and global travel means germs and viruses can spread faster and further than ever before. Air, sea, rail and road travel have never
Open Access. Keywords: Enterococcus faecalis, Enterococcus faecium, Vancomycin resistance, Virulence gene.
 34 The Open Microbiology Journal, 2012, 6, 34-39 Open Access Survey of Virulence Determinants among Vancomycin Resistant Enterococcus faecalis and Enterococcus faecium Isolated from Clinical Specimens
34 The Open Microbiology Journal, 2012, 6, 34-39 Open Access Survey of Virulence Determinants among Vancomycin Resistant Enterococcus faecalis and Enterococcus faecium Isolated from Clinical Specimens
Whole genome sequencing & new strain typing methods in IPC. Lyn Gilbert ACIPC conference Hobart, November 2015
 Whole genome sequencing & new strain typing methods in IPC Lyn Gilbert ACIPC conference Hobart, November 2015 Why do strain typing? Evolution, population genetics, geographic distribution 2 Why strain
Whole genome sequencing & new strain typing methods in IPC Lyn Gilbert ACIPC conference Hobart, November 2015 Why do strain typing? Evolution, population genetics, geographic distribution 2 Why strain
Report on the Japanese Veterinary Antimicrobial Resistance Monitoring System
 Report on the Japanese Veterinary Antimicrobial Resistance Monitoring System 2014 2015 National Veterinary Assay Laboratory Ministry of Agriculture, Forestry and Fisheries 2018 Contents Introduction...
Report on the Japanese Veterinary Antimicrobial Resistance Monitoring System 2014 2015 National Veterinary Assay Laboratory Ministry of Agriculture, Forestry and Fisheries 2018 Contents Introduction...
Renal Unit. Catheter Related Bacteraemia Guidelines
 Renal Unit Policy Manager Drew Henderson Policy Group Renal Unit Policy Established 21/01/2014 Policy Review Period/Expiry 21/01/2015 Last Updated 21/01/2014 This policy does apply to Medical/Dental Staff
Renal Unit Policy Manager Drew Henderson Policy Group Renal Unit Policy Established 21/01/2014 Policy Review Period/Expiry 21/01/2015 Last Updated 21/01/2014 This policy does apply to Medical/Dental Staff
Use of linezolid susceptibility test results as a surrogate for the susceptibility of Gram-positive pathogens to tedizolid, a novel oxazolidinone
 Zurenko et al. Annals of Clinical Microbiology and Antimicrobials 2014, 13:46 RESEARCH Open Access Use of linezolid susceptibility test results as a surrogate for the susceptibility of Gram-positive pathogens
Zurenko et al. Annals of Clinical Microbiology and Antimicrobials 2014, 13:46 RESEARCH Open Access Use of linezolid susceptibility test results as a surrogate for the susceptibility of Gram-positive pathogens
pentapeptide PG intermediate terminating in
 Research Article 463 Enzymes of vancomycin resistance: the structure of D-alanine Dlactate ligase of naturally resistant Leuconostoc mesenteroides Alexandre P Kuzin 1, Tao Sun 1, Jodi Jorczak-Baillass
Research Article 463 Enzymes of vancomycin resistance: the structure of D-alanine Dlactate ligase of naturally resistant Leuconostoc mesenteroides Alexandre P Kuzin 1, Tao Sun 1, Jodi Jorczak-Baillass
Chapter 19. Pathogenic Gram-Positive Bacteria. Staphylococcus & Streptococcus
 Chapter 19 Pathogenic Gram-Positive Bacteria Staphylococcus & Streptococcus Staphylococcus Normal members of every human's microbiota Can be opportunistic pathogens Facultative anaerobes Cells occur in
Chapter 19 Pathogenic Gram-Positive Bacteria Staphylococcus & Streptococcus Staphylococcus Normal members of every human's microbiota Can be opportunistic pathogens Facultative anaerobes Cells occur in
THE EFFECT OF RATION WITH ANTIBIOTICS (Virginiamycin) and TEMULAWAK (Curcuma xanthorriza Roxb.) TO BROILER PERFORMANCES
 THE EFFECT OF RATION WITH ANTIBIOTICS (Virginiamycin) and TEMULAWAK (Curcuma xanthorriza Roxb.) TO BROILER PERFORMANCES Lovita Adriani 1, Endang Sujana 2, Andi Mushawwir 3 1,3 Laboratorium Fisiologi Ternak
THE EFFECT OF RATION WITH ANTIBIOTICS (Virginiamycin) and TEMULAWAK (Curcuma xanthorriza Roxb.) TO BROILER PERFORMANCES Lovita Adriani 1, Endang Sujana 2, Andi Mushawwir 3 1,3 Laboratorium Fisiologi Ternak
Sri Lankan Journal of Infectious Diseases 2018 Vol.8(2): DOI: /sljid.v8i2.8221
 115 Research article Epidemiology of invasive infections caused by vancomycin sensitive and resistant enterococcal strains among oncology patients at the National Cancer Institute of Sri Lanka from 1st
115 Research article Epidemiology of invasive infections caused by vancomycin sensitive and resistant enterococcal strains among oncology patients at the National Cancer Institute of Sri Lanka from 1st
KILLS FOUR. Watch Informative Video
 Watch Informative Video 1 KILLS FOUR Virus Bacteria Mold Fungus After a decade of testing Inspired TEC s Cluster Ion Technology emerges as a formidable strategy to combat many of the biological, chemical
Watch Informative Video 1 KILLS FOUR Virus Bacteria Mold Fungus After a decade of testing Inspired TEC s Cluster Ion Technology emerges as a formidable strategy to combat many of the biological, chemical
