ALANINE:GLYOXYLATE AMINOTRANSFERASE (AGXT) GENE (PRIMARY HYPEROXALURIA TYPE 1) MUTATION DATA BASE
|
|
|
- Hugh Blankenship
- 5 years ago
- Views:
Transcription
1 ALANINE:GLYOXYLATE AMINOTRANSFERASE (AGXT) GENE (PRIMARY HYPEROXALURIA TYPE 1) REFERENCE SEQUENCES: c.dna: NM_ g.dna: NG_ MUTATION DATA BASE Nomenclature: HGVS guidelines ( Table 1: Polymorphic variants and those of unknown significance Table 2: Missense and nonsense mutations Table 3: Splice site mutations Table 4: Deletions and insertions Date: 18 August 2017 (future updates will not be made) Curator: Dr Gill Rumsby Disclaimer: While every effort has been made to confirm the accuracy of this database, the author can take no responsibility for errors which have occurred.
2 Table 1: Polymorphic variants and those of unknown significance (VUS) in the AGXT gene Location Sequence variant Codon/effect Major (Pro11) or minor (Leu11) haplotype Exon 1 c.26c>a p.thr9asn major Exon 1 c.27c>a p.thr9thr major n/a Exon 1 c.32c>t p.pro11leu minor Exon 1 c.32c>a p.pro11his major 55% Exon 1 c.35a>g p.lys12arg major In vitro activity (% of normal) or how proven 138% (Williams and 64% (Lumb and Danpure, 2000). Defines the AGXT minor allele Sift predicts tolerated Frequency in controls 0.01 SNP database 0.20 rs Does not track with disease Exon 1 c.65a>g p.asn22ser major 0.05 rs Intron 1 c a>g n/a minor (the A variant occasionally on minor) Intron 1 c c>t n/a minor n/a Intron 1 c c>t n/a minor n/a Intron 1 c _165+92dup74 n/a minor (occasionally major) Intron 1 c t>a n/a minor n/a n/a n/a 0.24 Original Reference (Frishberg et al., 2005; Monico et al., 2005) (Purdue et al., 1990) Williams and (Purdue et al., 1991a)
3 Intron 1 c a>c n/a minor n/a Intron 1 c c>t n/a minor n/a Intron 1 c t>c n/a minor n/a Intron 1 c c>t n/a major n/a Exon 2 c.264c>t p.ala88ala minor n/a 0.21 rs Intron 2 c c>t n/a minor n/a 0.20 rs Intron 2 c _358+64del9 n/a major n/a Intron 3 c c>t n/a n/a Intron 3 c a>c n/a major n/a Intron 3 c t>g n/a n/a Intron 3 c _13dupc n/a major n/a Intron 3 c g>a n/a major n/a Exon 4 c.489g>a p.leu163leu n/a Intron 4 c c>t n/a n/a 0.29 Intron 4 VNTR (29/32)n n/a Exon 5 c.590g>a p.arg197gln major minor/major. Minor typically with largest repeat n/a Mutation taster predicts benign. (Danpure et al., 1994a) (Von Schnakenburg, 1998) Rumsby et al Rumsby et al (Monico et al., (Danpure et al., 1994b) Incorrectly
4 Exon 6 c.654g>a p.ser218ser major n/a 0.06 rs Intron 6 c c>t n/a minor n/a 0.18 rs Intron 6 c g>a n/a n/a Intron 6 c c>t n/a n/a Intron 6 c g>a n/a major n/a Exon 7 c.705g>a p.thr235thr major n/a rs Exon 7 c.742g>t p.ala248ser major 75% Intron 7 c a>g n/a n/a 0.71 rs Intron 7 c c>t n/a n/a 0.43 rs Exon 8 c.836t>c p.ile279thr major /minor b,e 100% on major. On same allele as known mutations (Coulter-Mackie. Et al., Monico. Et al., In 1000 genomes Rs MAF /19 reported as G>T. Found in cis with c.557c>t in PH2 patient (Monico et al., 2007; Von Schnakenburg. 1998) (Monico et al., Rumsby et al Rumsby et al (Coulter- Mackie et al., 2005) (Monico et al., (Monico et al., 2007; Von Schnakenburg, 1998) (Coulter- Mackie et al., 2005) Exon 8 c.839c>t p.ala280val major 100%. On same Absent in 100 (Coulter-
5 allele as c.680+5g>c (Coulter-Mackie. Et al., 2005) controls Intron 8 c g>a n/a major/minor n/a 0.63 rs Exon 9 c.865c>t p.arg289cys minor Exon 9 c.860_861delinscg p.ser287thr Exon 9 c.883g>a p.ala295thr major Intron 9 c c>t n/a major n/a Exon 10 c.976g>a p.val326ile major Exon 10 c.1020a>g p.ile340met minor (occasionally major) Intron 10 c g>a n/a n/a Exon 11 c.1142g>a p.arg381lys No affect on activity or stability (C Tucker, personal communication 20/9/12) No effect on activity, trafficking or stability (Lage et al., 2014) Predicted polymorphism 96%. Found with p.ala112asp (Coulter-Mackie. Et al. 2003) % (Lumb and Danpure. 2000) Lys in other species. Benign by Sift and polyphen 0.03 in South African blacks: always found on African variant of minor allele 0.15 rs Mackie et al., 2005) (Monico et al., (Rinat et al., 1999) Frishberg et al., 2005 Williams and Rumsby Williams and Rumsby (Coulter- Mackie et al ) (Purdue et al., 1992) (Monico et al., Williams and Rumsby
6 Exon 11 c.1174c>t p.leu392leu major n/a 3 -non coding region 3 -non coding region c.*19g>a n/a major/minor n/a c.*41c>a n/a major/minor n/a rs non coding region c.*289c>a n/a major/minor n/a rs found in conjunction with c.33dupc (von Schnakenburg and 1997)
7 Table 2: Variants of unknown significance (VUS) in the AGXT gene Location Sequence variant Codon/effect Major (Pro11) or minor (Leu11) haplotype In vitro activity (% of normal) or how proven Exon 1 c.28c>t p.pro10ser major VUS 38% on minor 36% on major N.B. found in conjunction with homozygosity for 33dupC. Does not affect targeting Exon 2 c.296c>a p.ser99tyr major Exon 3 c.385g>c p.asp129his major Exon 5 c.557c>t p.ala186val major Exon 8 c.837t>g p.ile279met unknown VUS. Not fully conserved. On the same allele as the c.358+2g>t mutation VUS No obvious difference to wild type in yeast system (Lage et al., 2014) VUS. SIFT & mutation taster predict functional change. In SNP database. VUS Normal activity and stability on minor allele (Lage et al., 2014). Prediction=disease Frequency in controls Absent in 80 controls SNP database Original Reference Williams et Williams et al.,2009 Found in cis with 590G>A. Patient had PH2 (Frishberg et al., 2005)
8 causing Exon 8 c.845a>g p.gln282arg major VUS. Prediction disease causing but normal activity and stability on minor allele (Lage et al., 2014) Absent in 50 controls
9 Table 3: Missense and nonsense changes in the AGXT gene Location Sequence variant Codon/effect Major (Pro11) or minor (Leu11) haplotype In vitro activity (% normal control) or how proven Reference Exon 1 c.2t>c p.met1thr unknown Absent in 50 controls (Yuen et al., 2004) Exon 1 c.3g>t p.met1ile major Exon 1 c.22g>c p.val8leu Prediction software; Liver bx proven VUS 38% on minor 36% on major Exon 1 c.28c>t p.pro10ser major N.B. found in conjunction with homozygosity for 33dupC. Does not affect targeting Exon 1 c.32c>g p.pro11arg major <2% on major and minor Exon 1 c.74t>g p.leu25arg unknown Reduced activity and stability on major and (Monico et al., minor (Lage et al., 2014) Exon 1 c.77t>c p.leu26pro major <1% on major and minor a (Monico et al., Exon 1 c.106c>t p.arg36cys minor 0% (Williams and Defective in activity only (Tintillier et al., 2004) on major allele (Lage et al., 2014) Exon 1 c.107g>a p.arg36his major 8.5% on major. 5.4% on minor Exon 1 c.121g>a p.gly41arg major/minor 8% (Danpure et al., 1993) Exon 1 c.122g>a p.gly41glu major <1% (Williams and Exon 1 c.122g>t p.gly41val major 18% on Major (Coulter- Mackie and Lian, 2006) (Pirulli et al., 1999) Exon 1 c.125g>a p.gly42glu major?
10 Exon 1 c.130c>t p.gln44x major n/a (Pirulli et al., 1999) Exon 1 c.139g>a p.gly47arg minor g <3% activity (Pittman et (Monico et al., Absent in 50 controls; al., 2012). Exon 2 c.167t>a p.ile56asn major/minor Unstable on both polymorphic alleles (Lage et al, 2014) Exon 2 c.175g>a p.glu59lys minor Prediction Exon 2 c.187g>c p.gly63arg major Fully conserved Exon 2 c.188g>a p.gly63asp Predicted probably damaging Hopp et al., 2015 Exon 2 c.198c>g p.tyr66x major n/a (Purdue et al., 1991a) Exon 2 c.205c>t p.gln69x major/minor Exon 2 c.209c>a p.thr70asn Biopsy proven. No other Williams and Rumsby mutation found Exon 2 c.242c>a p.ser81x Exon 2 c.242c>t p.ser81leu major Exon 2 c.244g>c p.gly82arg minor Absent in 142 controls; conserved in 17/22 orthologous sequences 2.7% (Lumb and Danpure, 2000) Impaired activity on both alleles, unstable on minor (Lage et al.,2014) Li et al., BMC Nephrology 2014, 15:92 (Van Woerden et al., 2004) Exon 2 c.245g>a p.gly82glu major <2% (Purdue et al., 1992) Exon 2 c.248a>g p.his83arg minor 0% Impaired activity on both alleles, unstable on minor (Lage et al.,2014) Exon 2 c.254c>a p.ala85asp Biopsy proven <5% Exon 2 c.283g>a p.glu95lys minor Decreased activity and stability on both alleles Williams and Rumsby
11 (Lage et al., 2014) Exon 2 c.292g>c p.asp98his Prediction Hopp et al., 2015 Exon 2 c.302t>c p.leu101pro major <1% on major and minor Predominantly Asian ethnicity 3.5% on minor (Coulter- Exon 2 c.322t>c p.trp108arg minor Mackie and Lian, 2006) (Von Schnakenburg and Impaired activity on both 1998) alleles, unstable on minor (Lage et al.,2014) Exon 2 c.323g>a p.trp108x major Mutation Taster predicts disease causing Exon 2 c.324g>t p.trp108cys major Mutation Taster predicts Williams and disease causing Reduced activity and Exon 2 c.326g>t p.gly109val minor stability on major and minor (Lage et al., 2014) Exon 2 c.331c>t p.arg111x major n/a (Coulter-Mackie et al., 2004) Exon 2 c.332g>a p.arg111gln unknown Reduced activity and stability on major and minor (Lage et al., 2014) Exon 2 c.335c>a* p.ala112asp major 5% on major Exon 2 c.346g>a p.gly116arg major (Coulter-Mackie et al., 2003); *erroneously reported as 336C>A <1% on major and minor a (Pirulli et al., 1999) Williams and Exon 2 c.349g>t p.glu117x unknown Nonsense mutation Prediction programmes Exon 2 c.352c>t p.arg118cys minor Negligible AGT activity in liver biopsy Williams and Exon 2 c.353g>a p.arg118his minor Prediction programmes Exon 3 c.364c>t p.arg122x minor n/a Exon 3 c.371a>c p.his124pro major possibly damaging on Williams and Polyphen. Dx confirmed
12 on liver bx. Exon 3 c.409c>t p.gln137x major n/a 100% on minor; 50% on Exon 3 c.423g>t major a.?affects splicing p.glu141asp, may also major (last nucleotide in exon). affect splicing Found in biopsy proven patient (Von Schnakenburg, 1998) Exon 4 c.449t>c p.leu150pro minor <1% on major and minor (Williams and Exon 4 c.454t>a p.phe152ile minor <2% (Danpure et al., 1993) Exon 4 c.457t>g p.leu153val major Normal activity on minor allele. Approx 50% reduction in stability (Lage et al., 2014) (Van Woerden et al., 2004) Exon 4 c.466g>a p.gly156arg major <1% on major and minor (Pirulli et al., 1999; Rinat et al., (Williams and 1999) <1% on major and minor Exon 4 c.473c>t p.ser158leu major (Williams and (Monico et al., 2005) Exon 4 c.473c>a p.ser158x major Nonsense mutation Exon 4 c.481g>t p.gly161cys minor <1% on minor; 9% on major (Williams and Affects activity and stability on major and minor (Lage et al., 2014) (Williams and Exon 4 c.481g>a p.gly161ser minor <1% on minor;15% on major (Williams and. Reduce expression and half-life of AGT; aggregation (Oppici et (Williams and
13 al., 2013) Decreased stability on major and minor (Lage et al., 2014) Exon 4 c.481g>c p.gly161arg major 6.2% on major Affects activity and stability on major and (Coulter-Mackie et al., 2005) minor alleles (Lage et al.,2014) Exon 4 c.482g>a p.gly161asp major Hoyer-Kuhn et al Exon 4 c.497t>c p.leu166pro minor <1% on minor;16% on major (Williams and Affects activity and stability on both major (Williams and and minor but greater affect on minor (Lage et al., 2014) Exon 4 c.508g>a p.gly170arg minor 40% on minor; 68% on major (Lumb and Danpure, 2000) (Purdue et al., 1990) Exon 4 c.518g>a p.cys173tyr major <1% on major and minor (Williams and (Von Schnakenburg and 1998) Exon 4 c.519c>a p.cys173x major n/a (Van Woerden et al., 2004) Exon 5 c.533g>a p.cys178tyr major Exon 5 c.547g>a p.asp183asn major Exon 5 c.560c>t p.ser187phe major Sift + polyphen predict pathological. Biopsy proven 2.9% on major (Coulter- Mackie and Lian, 2006) 2.8% on major (Coulter- Mackie and Lian, 2006) Williams and (Basmaison et al., 2000) (Minatogawa, et al., 1992) Exon 5 c.560c>a p.ser187tyr prediction Hopp et al., 2015
14 Exon 5 c.568g>a p.gly190arg major/minor Exon 5 c.583a>c p.met195leu minor Exon 5 c.584t>g p.met195arg minor Exon 5 c.595g>a p.gly199ser; probable splice change major <1% on major and minor a Affects activity and stability on major and minor alleles (Lage et al. 2014) Reduced activity on major and minor in yeast expression system (Lage et al., 2014) In silico prediction. Tracks with disease in family (Rinat et al., 1999; Von Schnakenburg and 1998) (Frishberg et al., 2005) Rumsby et al., Exon6 c.603c>a p.asp201glu major Exon 6 c.605t>a p.ile202asn <5% on major and <10% on minor (Lage et al., 2014) Prediction: Sift and Polyphen (Monico et al., 2005) Li et al., BMC Nephrology 2014, 15:92 Exon 6 c.612c>a p.tyr204x major n/a (Monico et al., 2005) Exon 6 c.613t>c p.ser205pro major 2.5% on major (Coulter- Mackie and Lian, 2006) (Nishiyama et al., 1991) Exon 6 c.614c>t p.ser205leu major <3% on major and minor Exon 6 c.614c>a p.ser205x major n/a Exon 6 c.628g>c p.ala210pro No other mutation found (liver bx proven) Aquaviva, Exon 6 c.646g>a p.gly216arg major Absent in 50 controls <3% (this paper a ) (Monico et al., Exon 6 c.653c>t p.ser218leu major 10% on Major (Coulter-Mackie et al., 2005) Exon 6 c.661t>c p.ser221pro major <3% on major and minor Exon 7 c.697c>t p.arg233cys minor <1% on minor; 14% on Major (Williams and (von Schnakenburg and 1997)
15 Exon 7 c.698g>a p.arg233his minor <1% on minor; 14% on Major (Williams and (von Schnakenburg and 1997) Exon 7 c.698g>t p.arg233leu unknown Absent in 50 controls (Monico et al., 2005) Exon 7 c.727g>c p.asp243his unknown <5% activity on minor, approx 50% on major. Unstable on both alleles (Laga et al., 2014) (Frishberg et al., 2005) Exon 7 c.731t>c p.ile244thr minor <1% on minor; 50% on (von Schnakenburg and (occasionally major (Lumb and 1997) major) Danpure, 2000) (Monico et al., 2007; Williams Exon 7 c.737g>a p.trp246x minor Nonsense mutation and (von Schnakenburg and Exon 7 c.738g>a p.trp246x major Nonsense mutation 1997) Exon 7 c.753g>a p.trp251x major Nonsense mutation (Coulter-Mackie, et al., 2004) Exon 7 c.757t>c p.cys253arg minor <1% on minor; 7% on Major (Williams and Decreased activity and stability on both alleles (Lage et al., 2014) (Monico et al., 2005; Williams and Exon 8 c.783t>a p.his261gln unknown <3% (Pittman et al., 2012) (Monico et al., Exon 8 c.779a>g p.tyr260cys Prediction Hopp et al., 2015 Exon 8 c.781c>g p.his261asp Prediction Hopp et al., 2015 Exon 8 c.806t>c p.leu269pro major <5% on major and minor alleles and affects stability on both alleles (Lage et al., 2014) Exon 8 c.815t>c p.leu272pro Prediction Hopp et al., 2015 Exon 8 c.822g>c p.glu274asp minor <3% (Pittman et al., 2012) Normal activity on major (Lage et al., 2014)
16 Exon 8 c.823a>c p.ser275arg major/minor <3% on both majorand minor allele (Pittman et al., 2012) Exon 8 c.837t>g p.ile279met Exon 8 c.844c>t p.gln282x unknown Nonsense mutation (Yuen, et al., 2004) Exon 8 c.846g>c p.gln282his major Prediction disease causing. Found in biopsy-proven PH1 Exon 9 c.851t>c p.leu284pro minor Reduced activity and stability on both major and minor (Lage et al., 2014) Exon 9 c.853g>t p.glu285x major Nonsense mutation Exon 9 c.866g>a p.arg289his major Not tested. R conserved in mammals (except rabbit. Williams and Rumsby Exon 9 c.891t>g p.tyr297x major Nonsense mutation Exon 9 C.893T>C p.leu298pro minor Absent in 100 controls; Destabilising (C Tucker, personal communication 20/9/12 and Lage et al., (Rinat et al., 1999) 2014) Variant in cis with p.arg289cys Exon 9 c.907c>t p.gln303x unknown Nonsense mutation (Monico et al., Williams and Rumsby Exon 9 c.922c>t p.gln308x major Nonsense mutation Exon 10 c.947t>c p.leu316pro Biopsy proven. No other Williams and mutation found Exon 10 c.949c>t p.arg317trp Prediction Hopp et al., 2015 Exon 10 c.956c>t p.pro319leu unknown Absent in 50 controls; <5% on minor allele, decreased stability on (Monico, et al.,
17 both alleles (Lage et al., 2014) Exon 10 c.996g>a p.trp332x Nonsense mutation Williams and Biopsy proven PH1 Exon 10 c.997a>t p.arg333x major Nonsense mutation (Rinat et al., 1999) Exon 10 c.1007t>a p.val336asp minor 5.2% on minor; 22.4% (Van Woerden et al., 2004) originally described in cis with on major (Coulter- p.gly170arg; since found on its Mackie and Lian, 2008) own (GR) Williams and Exon 10 c.1014c>g p.tyr338x Nonsense mutation Exon 10 c.1045g>a p.gly349ser Found in biopsy proven Williams and PH1 Exon 10 c.1049g>a p.gly350asp minor 2.9% on major, 7.6% on (Von Schnakenburg and minor (this paper a ) 1998) Exon 10 c.1076t>c p.leu359pro major L conserved in middle of beta sheet. Reduced activity and stability on major and minor alleles (Lage et al., 2014) Williams and Rumsby Exon 11 c.1078c>t p.arg360trp Prediction Hopp et al., 2015 Exon 11 c.1079g>a p.arg360gln major <1% on major and minor a Exon 11 c.1084g>a p.gly362ser Prediction Hopp et al., 2015 Exon 11 c.1102g>a p.ala368thr minor Affects activity, stability and growth on minor allele. Lesser affect on major allele (Laga et al.,2014) Exon 11 c.1148c>a p.ala383asp major Prediction programmes Reduced activity on Exon 11 c.1151t>c p.leu384pro major major and minor (Lage et al., 2014) Willliams and Rumsby
18 Table 4: Splice site mutations in the AGXT gene Mutation Effect Reference Splice donor Intron 2 c.358+1g>t (Van Woerden et al., 2004) Intron 2 c.358+2t>g Intron 6 c.680+1g>c Intron 6 c.680+1g>a Intron 6 c.680+2t>a Intron 6 c.680+5g>c (Coulter-Mackie, et al., 2005) Intron 7 c.776+1g>a p.met259fs inclusion of 24 nucleotides (Williams and (Monico et al., 2007; Williams and Intron 7 c.776+1g>c (Monico et al., Intron 8 c.846+1g>t (Coulter-Mackie et al., 2004) Intron 8 c.846+1g>a Intron 9 c.942+1g>t (Monico et al., 2005) Intron 10 c g>a Splice acceptor Intron 1 c.166-1g>a (Van Woerden et al., 2004) Intron 3 c.424-2a>g p.gly_142gln145del Loss of 12 nucleotides at beginning of (Williams and exon 4 (Williams and Intron 4 c.525-1g>a (Basmaison et al., 2000) Intron 5 c.595+1g>t Hopp et al., 2015 Intron 5 c.596-2a>g Williams and Intron 7 c.777-1g>c (Coulter-Mackie et al., 2004) Intron 7 c.777-2a>g Williams and Intron 8 c.846+1g>t Halbritter et al., 2015 Intron 8 c.847-1g>c (Basmaison et al., 2000) Intron 8 c.847-3c>g (Basmaison et al., 2000) Intron 9 c.943-1g>t Clifford-Mobley & Rumsby Intron 9 c.943-1g>a
19 Table 5: Deletion/insertion mutations in the AGXT gene Location Mutation Codon Major or minor allele Exon 1 c.2_3delinsat p.met1asn (activity <1%) major Exon 1 c.33delc p.lys12fs major Exon 1 c.32_33delcc p.pro11fs major Exon 1 c.33dupc (formerly c.33_34insc) p.lys12fs Exon 1 c.34_35dupaa p.lys12fs Exon 1 c.83delc p.pro28fs major Exon 1 c.116_117dupca p.ala40fs major Exon 1 c.126delg p.leu43fs major major Confirmation Original reference (Williams and (Pirulli et al., 1999) (Williams and (Pirulli et al., 1999) Hopp et al., 2015 (Coulter- Mackie et al., 2005) (Williams and Exon 1 c.126dupg p.leu43fs Hopp et al., 2015 Williams and Exon 2 c.215dupa p.asn72fs Exon 2 c.221_227duptcacact p.val77fs major Exon 2 c.276delt p.asn92fs major Exon 2 c.283_285dupgag p.glu95dup major Exon 2 c.299_307duptcctggttg p.val102_gly103ins Val, Leu,Val Exon 2 c.[299_307dup;c.308g>a] p.[val102insvalleuval;gly103glu] unknown (Basmaison et al., 2000) (Pirulli et al., 1999) Hopp et al, 2015 (Monico, et al.,
20 Exon 2 c.327delg p.gln110fs major (Coulter- Mackie et al., 2005) Exon 3 c.359-4_379del Hopp et al., Exon 3 c.402c>g p.tyr134* 2015 Exon 3 c.416_418deltgg p.val139del major b (Monico et al., Exon 4 c.447_454delgctgctgt p.leu151fs major Exon 4 c.445delg p.val149fs major Exon 4 c.460dela p.thr154fs major Exon 4 c.485delt p.(val162glyfs*50) minor Exon 4 c.496delg p.glu157fs Exon 4 c.519_520delinsga p.[cys173trp;his174asn] major Exon 5 c.557_562delinsatcggt p.[ala186asp;phe187arg] minor Exon 5 c.560_561insggt p.ser187_leu188insval Exon 5 c.570delg p.thr191fs major Exon 5 c.577delc p.leu193fs major Exon 5 c.577dupc p.leu193fs major <10% AGT activity in liver bx aa inserted incorrectly reported as gly. (Williams and Rumsby Hopp et al., 2015 Williams and Williams and Hopp et al., 2015 (Coulter- Mackie et al., 2004)
21 Exon 6 c.642_645deltcca p.pro215fs major Exon 6 c.661_663deltcc p.ser221del Exon6/Intron 6 c.679_680+2delaagt?missplicing or p.lys227glu major Exon 7 c _c del732instgaga NC_ :g _ del732insTGAGA p.lys228_met259del32 (in frame deletion of exon 7) Exon 7 c.725dupt p.asp243fs major Exon 7 c.744delc p.asp249fs major Exon 7 c.751_752delinsaa p.trp251lys (peroxisomal and cytosolic distribution) Exon 8 c.798_802delinsacaatctcag p.ile267fs major Exon 8 c.823_824dupag p.ser275fs unknown Exon8 c.829_830insa,c.830c>a p.ala277fs minor Exon 8 c.832delc p.leu278fs Exon 8 c.834delc p.ile279fs minor Exon 9 c _c del1121 NC_ :g _ del1122 deletion exon 9 Exon 9 c.860_861delinscg p.ser287thr unknown Exon 9 c.886_888delgcg Ala296del major Exon 9 c.919delc p.leu307fs major major (Coulter- Mackie, et al., 2001) (Robbiano et al., 2010) (Williams and (Coulter- Mackie et al., 2004) Kawai et al., 2012 (von Schnakenburg et al., 1998) (Yuen, et al., 2004) Williams and Hopp et al., 2015 (Coulter- Mackie et al., 2005) (Frishberg et al., 2005) (Von Schnakenburg and 1998)
22 Exon 10 c.959_960delca p.thr320fs Exon 10 c deltg p.val324fs major Exon 10 c.973delg p.ala325fs Exon 10 c.976delg p.val326fs major Exon 10 c.983_988delctggct p.ala328_tyr330delinsasp major Exon 11 c.1110_1111delcg p.asn372fs minor Exon 11 c.1125_1126delcg p.val376fs major Exon 11_3 UTRdel truncating Williams and Rumsby (Pirulli et al., 1999) (Coulter- Mackie et al., 2004) Williams and (Coulter- Mackie et al., 2001) (Monico et al., Ex1_5del Ex1_7del Ex5_11del g telomere 5-6 kb deletion 10 kb deletion >6kb deletion Major deletion including HDAC4 (Coulter- Mackie et al., 2001) (Nogueira et al., 2000) Tammachote et al., 2012
23 References Basmaison O, Rolland M-O, Cochat P, Bozon D Identification of 5 novel mutations in the AGXT gene. Hum Mutat.15:577 Coulter-Mackie MB, Applegarth D, Toone JR, Henderson H The major allele of the alanine:glyoxylate aminotransferase gene:seven novel mutations causing primary hyperoxaluria type 1. Mol Genet Metab 82: Coulter-Mackie MB, Lian Q Consequences of missense mutations for dimerisation and turnover of alanine:glyoxylate aminotransferase:study of a spectrum of mutations. Mol Genet Metab. Coulter-Mackie MB, Lian Q, Applegarth D, Toone JR The major allele of the alanine:glyoxylate aminotransferase gene:nine novel mutations and polymorphisms associated with primary hyperoxaluria type 1. Mol Genet Metab 86: Coulter-Mackie MB, Rumsby G, Applegarth DA, Toone JR Three novel deletions in the alanine:glyoxylate aminotransferase gene of three patients with type 1 hyperoxaluria. Mol Genet Metab 74: Coulter-Mackie MB, Tung A, Henderson HE, Toone JR, Applegarth DA The AGT gene in Africa: a distinct minor allele haplotype, a polymorphism (V326I) and a novel PH1 mutation (A112D) in black Africans. Mol Genet Metab 78: Danpure C, Jennings P, Freyer P, Purdue P, Allsop J. 1994a. Primary hyperoxaluria type 1: Genotypic and phenotypic heterogeneity. J Inherit Metab Dis 17: Danpure CJ, Birdsey GM, Rumsby G, Lumb MJ, Purdue PE, Allsop J. 1994b. Molecular characterization and clinical use of a polymorphic tandem repeat in an intron of the human alanine:glyoxylate aminotransferase gene. Hum Genet 94(1): Danpure CJ, Purdue PE, Fryer P, Griffiths S, Allsop J, Lumb MJ, Guttridge KM, Jennings PR, Scheinman JI, Mauer SM and others Enzymological and mutational analysis of a complex primary hyperoxaluria type 1 phenotype involving alanine:glyoxylate aminotransferase peroxisome-to-mitochondrion mistargeting and intraperoxisomal aggregation. Am J Hum Genet 53(2):
24 Frishberg Y, Rinat C, Khatib I, Shalata A, Feinstein S, Becker-Cohen R, Weismann I, Wanders RJA, Rumsby G, Roels F and others Intra-familial clinical heterogeneity: Absence of genotype-phenotype correlation in primary hyperoxaluria type 1 in Israel. Am J Nephrol 25: Hopp K, Cogal A G, Bergstralh E J, Seide B M, Olson J B, Meek A M, Lieske J C, Milliner D S, Harris P C Phenotype-genotype correlations and estimated carrier frequencies of primary hyperoxaluria. J Am Soc Nephrol 26: Feb 2. pii: JASN [Epub ahead of print] Hoyer-Kuhn H, Kohbrok S, Volland R, Franklin J, Hero B, Beck B B Vitamin B6 in primary hyperoxaluria type 1: First prospective trial after 40 years of practice. Clin J Am Soc Nephrol 9: Kawai C, Minatogawa Y, Akiyoshi H, Hirose, S, Suehiro T, Tone S A novel mutation of human liver alanine:glyoxylate aminotransferase causes primary hyperoxaluria type 1: immunohistochemical quantification and subcellular distribution. Acta Histochem.Cytochem 42: Laga M D, Pittman, A M C, Roncador A, Cellini A and Tucker C L Allele specific characterization of alanine:glyoxylate aminotransferase variants associated with primary hyperoxaluria. Plos One in press Lumb MJ, Danpure CJ Functional synergism between the most common polymorphism in human alanine:glyoxylate aminotransferase and four of the most common disease causing mutations. J Biol Chem 275: Minatogawa Y, Tone S, Allsop J, Purdue PE, Takada Y, Danpure CJ, Kido R A serine-to-phenylalanine substitution leads to loss of alanine:glyoxylate aminotransferase catalytic activity and immunoreactivity in a patient with primary hyperoxaluria type 1. Hum Mol Genet 1(8): Monico CG, Olson JB, Milliner DS Implications of genotype and enzyme phenotype in pyridoxine response of patients with type 1 primary hyperoxaluria. Am J Nephrol 25: Monico CG, Rossetti S, Schwanz HA, Olson JB, Lundquist PA, Dawson DB, Harris PC, Milliner DS Comprehensive mutation screening in 55 probands with type 1 primary hyperoxaluria type 1 primary hyperoxaluria shows feasibility of a gene-based diagnosis. J Am Soc Nephrol 18:
25 Nishiyama K, Funai T, Katafuchi R, Hattori F, Onoyama K, Ichiyama A Primary hyperoxaluria type I due to a point mutation of T to C in the coding region of the serine:pyruvate aminotransferase gene. Biochem Biophys Res Commun 176(3): Nogueira PK, Vuong TS, Boutoon O, Maillard A, Marchand M, Rolland MO, Cochat P, Bozon D Partial deletion of the AGXT gene (EX1_EX7del): A new genotype in hyperoxaluria type 1. Hum Mutat. 15: Oppici E, Roncador A, Montioli R, Bianconi S, Cellini B Gly161 mutations associated with primary hyperoxaluria type 1 induce the cytosolic aggregation and the intracellular degradation of the apo-form of alanine:glyoxylate aminotransferase. Biochim Biophys Acta 1832: Pirulli D, Puzzer D, Ferri L, Crovella S, Amoroso A, Ferrettini C, Marangella M, Mazzola G, Florian F Molecular analysis of hyperoxaluria type 1 in Italian patients reveals eight new mutations in the alanine:glyoxylate aminotransferase gene. Hum Genet 104(6): Pittmann A M C, Lage M D, Poltoratsky V, Vrana J D, Paiardini A, Roncador A, Cellini B, Hughes R M, Tucker C L Rapid profiling of disease alleles using a tunable reporter of protein misfolding. Genetics 192: Purdue P, Lumb M, Allsop J, Danpure C. 1991a. An intronic duplication in the alanine: glyoxylate aminotransferase gene facilitates identification of mutations in compound heterozygote patients with primary hyperoxaluria type 1. Hum Genet 87: Purdue P, Lumb M, Allsop J, Minatogawa Y, Danpure C A glycine-to-glutamate substitution abolishes alanine:glyoxylate aminotransferase catalytic activity in a subset of patients with primary hyperoxaluria type 1. Genomics 13: Purdue PE, Takada Y, Danpure CJ Identification of mutations associated with peroxisome-to-mitochondrion mistargeting of alanine/glyoxylate aminotransferase in primary hyperoxaluria type 1. J Cell Biol 111: Rinat C, Wanders RJA, Drukker A, Halle D, Frishberg Y Primary hyperoxaluria type I: A model for multiple mutations in a monogenic disease within a distinct ethnic group. J Am Soc Nephrol 10:
26 Robbiano A, Mandrile G, De Marchi M, Beck B, Baasner A, Murer L, Benetti E, Giachino D Novel human pathological mutations. Gene symbol:agxt.disease:hyperpoxaluria. Hum Genet 127(4):468 Tammachote R, Kingsuwannapong N, Tongkobpetch S, Srichomthong C, Yeetong P, Kingwatanakul P, Monico C G, Suphapeetiporn K, Shotelersuk V Primary hyperoxaluria type 1 and brachydactyly mental retardation caused by a novel mutation in AGXT and a terminal deletion of chromosome 2. Am J Med Genet 158A: Tintillier M, Pochet JM, Cosyns JP, Delgrange E, Donckier J Late onset primary hyperoxaluria triggered by hypothyroidism and presenting as rapidly progressive renal failure - description of a new mutation. Clin Nephrol 62: Van Woerden CS, Groothoff JW, Wijburg FA, Annink C, Wanders RJA, Waterham HR Clinical implications of mutation analysis in primary hyperoxaluria type 1. Kidney Int 66: Von Schnakenburg C Molecular analysis of the AGXT gene and linkage studies in primary hyperoxaluria type 1: PhD Thesis, University of London. von Schnakenburg C, Hulton SA, Milford DV, Roper HP, Rumsby G Variable presentation of primary hyperoxaluria type 1 in 2 patients homozygous for a novel combined deletion and insertion mutation in exon 8 of the AGXT gene. Nephron 78: von Schnakenburg C, Rumsby G Primary hyperoxaluria type 1: a cluster of new mutations in exon 7 of the AGXT gene. J Med Genet 34(6): Von Schnakenburg C, Rumsby G Identification of new mutations in primary hyperoxaluria type 1 (PH1). J Nephrol 11(S1): Williams EL, Rumsby G Selected exonic sequencing of the AGXT gene provides a genetic diagnosis in 50% of patients with primary hyperoxaluria type 1. Clin Chem 53: Williams E L, Acquaviva C, Amoroso A, Chevalier F, Coulter Mackie M, Monico C G, Giachino D, Owen E P, Robbiano A, Salido E, Waterham H, Rumsby G Primary hyperoxaluria type 1: Update and additional mutation analysis of the AGXT gene. Hum Mutat 30,1-8
27 Yuen Y, Lai C, Tong GM, Wong P, Wong FK, Mak S, Lo K, Wong AK, Tong S, Chan Y and others Novel mutations of the AGXT gene causing primary hyperoxaluria type 1. J Nephrol 17:
Names for ABO (ISBT 001) Blood Group Alleles
 Names for ABO (ISBT 001) blood group alleles v1.1 1102 Names for ABO (ISBT 001) Blood Group Alleles General description: The ABO system was discovered as in 1900 and is considered the first and clinically
Names for ABO (ISBT 001) blood group alleles v1.1 1102 Names for ABO (ISBT 001) Blood Group Alleles General description: The ABO system was discovered as in 1900 and is considered the first and clinically
Assessing the phenotypic effects in the general population of rare variants in genes for a dominant Mendelian form of diabetes
 Assessing the phenotypic effects in the general population of rare variants in genes for a dominant Mendelian form of diabetes Supplementary Information Jason Flannick*, Nicola L Beer*, Alexander G Bick,
Assessing the phenotypic effects in the general population of rare variants in genes for a dominant Mendelian form of diabetes Supplementary Information Jason Flannick*, Nicola L Beer*, Alexander G Bick,
Colclough et al., Human Mutation 1
 Region Colclough et al., Human Mutation 1 Supp. Table S1. List of HNF1A mutations Nucleotide Change NM_000545.5 Promoter and 5 UTR Mutations Protein Effect NM_000545.5 Three letter amino acid description
Region Colclough et al., Human Mutation 1 Supp. Table S1. List of HNF1A mutations Nucleotide Change NM_000545.5 Promoter and 5 UTR Mutations Protein Effect NM_000545.5 Three letter amino acid description
Names for H (ISBT 018) Blood Group Alleles
 Names for H (ISBT 018) Blood Group Alleles General description: The H blood group system consists of one antigen, H, that is carried on glycolipids and glycoproteins on the RBC membrane, where it is synthesised
Names for H (ISBT 018) Blood Group Alleles General description: The H blood group system consists of one antigen, H, that is carried on glycolipids and glycoproteins on the RBC membrane, where it is synthesised
Supplementary materials and methods
 Supplementary materials and methods Targeted resequencing RNA probes complementary in sequence to the coding exons (+ 2bp) of target genes were designed using e-array (https://earray.chem.agilent.com/earray/)
Supplementary materials and methods Targeted resequencing RNA probes complementary in sequence to the coding exons (+ 2bp) of target genes were designed using e-array (https://earray.chem.agilent.com/earray/)
Supplementary Table e1. Clinical and genetic data on the 37 participants from the WUSM
 Supplementary Data Supplementary Table e1. Clinical and genetic data on the 37 participants from the WUSM cohort. Supplementary Table e2. Specificity, sensitivity and unadjusted ORs for glioma in participants
Supplementary Data Supplementary Table e1. Clinical and genetic data on the 37 participants from the WUSM cohort. Supplementary Table e2. Specificity, sensitivity and unadjusted ORs for glioma in participants
Targeted sequencing of refractory myeloma reveals a high incidence of mutations in CRBN and Ras pathway genes
 Targeted sequencing of refractory myeloma reveals a high incidence of mutations in CRBN and Ras pathway genes KM Kortüm 1,8 *, EK Mai 3,4 *, NH Hanafiah 3 *, CX Shi 1, YX Zhu 1, L Bruins 1, S Barrio 1,
Targeted sequencing of refractory myeloma reveals a high incidence of mutations in CRBN and Ras pathway genes KM Kortüm 1,8 *, EK Mai 3,4 *, NH Hanafiah 3 *, CX Shi 1, YX Zhu 1, L Bruins 1, S Barrio 1,
Type 1 primary hyperoxaluria (PH1; OMIM ) is a
 JASN Express. Published on April 25, 2007 as doi: 10.1681/ASN.2006111230 Comprehensive Mutation Screening in 55 Probands with Type 1 Primary Hyperoxaluria Shows Feasibility of a Gene- Based Diagnosis Carla
JASN Express. Published on April 25, 2007 as doi: 10.1681/ASN.2006111230 Comprehensive Mutation Screening in 55 Probands with Type 1 Primary Hyperoxaluria Shows Feasibility of a Gene- Based Diagnosis Carla
Supplemental material Mutant DNMT3A: A Marker of Poor Prognosis in Acute Myeloid Leukemia Ana Flávia Tibúrcio Ribeiro, Marta Pratcorona, Claudia
 Supplemental material Mutant DNMT3A: A Marker of Poor Prognosis in Acute Myeloid Leukemia Ana Flávia Tibúrcio Ribeiro, Marta Pratcorona, Claudia Erpelinck-Verschueren, Veronika Rockova, Mathijs Sanders,
Supplemental material Mutant DNMT3A: A Marker of Poor Prognosis in Acute Myeloid Leukemia Ana Flávia Tibúrcio Ribeiro, Marta Pratcorona, Claudia Erpelinck-Verschueren, Veronika Rockova, Mathijs Sanders,
InheriGen Plus Disease Information
 DISEASE INFORMATION & MUTATIONS TESTED 17α-hydroxylase/17,20-lyase Deficiency CYP17A1 Brazilian 87% < 1 in 112 < 1 in 850 p.arg362cys, p.trp406arg, c.1243+5g>a, p.pro48fs, p.asp487_phe489del, p.tyr32x
DISEASE INFORMATION & MUTATIONS TESTED 17α-hydroxylase/17,20-lyase Deficiency CYP17A1 Brazilian 87% < 1 in 112 < 1 in 850 p.arg362cys, p.trp406arg, c.1243+5g>a, p.pro48fs, p.asp487_phe489del, p.tyr32x
Genetic Testing and Analysis. (858) MRN: Specimen: Saliva Received: 07/26/2016 GENETIC ANALYSIS REPORT
 GBinsight Sample Name: GB4411 Race: Gender: Female Reason for Testing: Type 2 diabetes, early onset MRN: 0123456789 Specimen: Saliva Received: 07/26/2016 Test ID: 113-1487118782-4 Test: Type 2 Diabetes
GBinsight Sample Name: GB4411 Race: Gender: Female Reason for Testing: Type 2 diabetes, early onset MRN: 0123456789 Specimen: Saliva Received: 07/26/2016 Test ID: 113-1487118782-4 Test: Type 2 Diabetes
LMM / emerge III Network Reference Sequences October 2016 LMM / emerge III Network Consensus Actionable Gene List *ACMG56 gene
 LMM / emerge III Network Consensus Actionable Gene List *ACMG56 gene ACTA2* Exon 02-09 NM_001613.2 DSG2* Exon 01-15 NM_001943.3 ACTC1* Exon 01-06 NM_005159.4 DSP* Exon 01-24 NM_004415.2 APC* Exon 01-15
LMM / emerge III Network Consensus Actionable Gene List *ACMG56 gene ACTA2* Exon 02-09 NM_001613.2 DSG2* Exon 01-15 NM_001943.3 ACTC1* Exon 01-06 NM_005159.4 DSP* Exon 01-24 NM_004415.2 APC* Exon 01-15
A novel mutation in the AGXT gene causing primary hyperoxaluria type I: genotype phenotype correlation
 c Indian Academy of Sciences RESEARCH ARTICLE A novel mutation in the AGXT gene causing primary hyperoxaluria type I: genotype phenotype correlation SAOUSSEN M DIMEGH 1, CÉCILE AQUAVIVA-BOURDAIN 2, ASMA
c Indian Academy of Sciences RESEARCH ARTICLE A novel mutation in the AGXT gene causing primary hyperoxaluria type I: genotype phenotype correlation SAOUSSEN M DIMEGH 1, CÉCILE AQUAVIVA-BOURDAIN 2, ASMA
Spectrum and frequencies of mutations in MSH2 and MLH1 identified in 1,721 german families suspected of hereditary nonpolyposis colorectal cancer
 Int. J. Cancer: 116, 692 702 (2005) ' 2005 Wiley-Liss, Inc. Spectrum and frequencies of mutations in MSH2 and MLH1 identified in 1,721 german families suspected of hereditary nonpolyposis colorectal cancer
Int. J. Cancer: 116, 692 702 (2005) ' 2005 Wiley-Liss, Inc. Spectrum and frequencies of mutations in MSH2 and MLH1 identified in 1,721 german families suspected of hereditary nonpolyposis colorectal cancer
Supplementary Table I. List of mutations presented by the Portuguese pediatric cohort with FH
 Supplementary Table I. List of mutations presented by the Portuguese pediatric cohort with FH Index patients (n=89) FH children Cascade screening (n=82) cdna Mutation Protein 3 0 c.10580g>a p.arg3527gln
Supplementary Table I. List of mutations presented by the Portuguese pediatric cohort with FH Index patients (n=89) FH children Cascade screening (n=82) cdna Mutation Protein 3 0 c.10580g>a p.arg3527gln
analysis in hereditary breast/ovarian cancer families of Portuguese ancestry
 Clin Genet 2015: 88: 41 48 Printed in Singapore. All rights reserved Short Report 2014 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12441 The role of targeted
Clin Genet 2015: 88: 41 48 Printed in Singapore. All rights reserved Short Report 2014 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12441 The role of targeted
Supplementary Document
 Supplementary Document 1. Supplementary Table legends 2. Supplementary Figure legends 3. Supplementary Tables 4. Supplementary Figures 5. Supplementary References 1. Supplementary Table legends Suppl.
Supplementary Document 1. Supplementary Table legends 2. Supplementary Figure legends 3. Supplementary Tables 4. Supplementary Figures 5. Supplementary References 1. Supplementary Table legends Suppl.
Intron p Hybrid with p.leu245val p. Gly336Cys. p.leu62phe p. Ala137Val p. Asn152Thr hybrid with p.leu245val p.
 RHD negative, i.e. null Phenotype Allele name Nucleotide change Exon Predicted amino acid change D RHD*01N.01 RHD deletion 1-10 p.0 Allele name detail Comments D RHD*08N.01 RHD*Pseudogene RHD* Ψ 37- bp
RHD negative, i.e. null Phenotype Allele name Nucleotide change Exon Predicted amino acid change D RHD*01N.01 RHD deletion 1-10 p.0 Allele name detail Comments D RHD*08N.01 RHD*Pseudogene RHD* Ψ 37- bp
Supplementary material PCNT point mutations and familial intracranial aneurysms
 Supplementary material PCNT point mutations and familial intracranial aneurysms Oswaldo Lorenzo-Betancor, MD, PhD 1, Patrick R. Blackburn, PhD 2,3, Emily Edwards, CCRP 4, Rocío Vázquez-do-Campo, MD 4,
Supplementary material PCNT point mutations and familial intracranial aneurysms Oswaldo Lorenzo-Betancor, MD, PhD 1, Patrick R. Blackburn, PhD 2,3, Emily Edwards, CCRP 4, Rocío Vázquez-do-Campo, MD 4,
De Leeneer et al., Human Mutation 1
 De Leeneer et al., Human Mutation 1 Supp. Table S1A. Overview of the genes included in the workflow Ensembl Gene ID Ensembl Transcript ID Associated Gene Name ENSG00000198691 ENST00000370225 ABCA4 NM_000350.2
De Leeneer et al., Human Mutation 1 Supp. Table S1A. Overview of the genes included in the workflow Ensembl Gene ID Ensembl Transcript ID Associated Gene Name ENSG00000198691 ENST00000370225 ABCA4 NM_000350.2
CHAPTER IV RESULTS Microcephaly General description
 47 CHAPTER IV RESULTS 4.1. Microcephaly 4.1.1. General description This study found that from a previous study of 527 individuals with MR, 48 (23 female and 25 male) unrelated individuals were identified
47 CHAPTER IV RESULTS 4.1. Microcephaly 4.1.1. General description This study found that from a previous study of 527 individuals with MR, 48 (23 female and 25 male) unrelated individuals were identified
Vitamin B6 in Primary Hyperoxaluria I: First Prospective Trial after 40 Years of Practice
 CJASN epress. Published on January 2, 2014 as doi: 10.2215/CJN.06820613 Article Vitamin B6 in Primary Hyperoxaluria I: First Prospective Trial after 40 Years of Practice Heike Hoyer-Kuhn,* Sina Kohbrok,*
CJASN epress. Published on January 2, 2014 as doi: 10.2215/CJN.06820613 Article Vitamin B6 in Primary Hyperoxaluria I: First Prospective Trial after 40 Years of Practice Heike Hoyer-Kuhn,* Sina Kohbrok,*
Hearing Loss Imaging Thyroid function tests TFT (age at test)
 Clinical Information on patients. Unclassified variants are denoted in brackets. EVA is enlarged vestibular aqueducts - Indicates no information. means asian. Patient TFT (age at test) 25215 c.1001+1g>a
Clinical Information on patients. Unclassified variants are denoted in brackets. EVA is enlarged vestibular aqueducts - Indicates no information. means asian. Patient TFT (age at test) 25215 c.1001+1g>a
Supplementary Materials for
 www.sciencemag.org/content/354/6319/aaf7000/suppl/dc1 Supplementary Materials for Genetic identification of familial hypercholesterolemia within a single U.S. health care system Noura S. Abul-Husn, Kandamurugu
www.sciencemag.org/content/354/6319/aaf7000/suppl/dc1 Supplementary Materials for Genetic identification of familial hypercholesterolemia within a single U.S. health care system Noura S. Abul-Husn, Kandamurugu
GALAFOLD (migalastat) capsules, for oral use Initial U.S. Approval: 2018
 HIGHLIGHTS OF PRESCRIBING INFORMATION These highlights do not include all the information needed to use GALAFOLD safely and effectively. See full prescribing information for GALAFOLD. GALAFOLD (migalastat)
HIGHLIGHTS OF PRESCRIBING INFORMATION These highlights do not include all the information needed to use GALAFOLD safely and effectively. See full prescribing information for GALAFOLD. GALAFOLD (migalastat)
Novel NPHS1 gene mutations in a Chinese family with congenital nephrotic syndrome
 c Indian Academy of Sciences RESEARCH NOTE Novel NPHS1 gene mutations in a Chinese family with congenital nephrotic syndrome FENGJIE YANG, YAXIAN CHEN, YU ZHANG, LIRU QIU, YU CHEN and JIANHUA ZHOU Department
c Indian Academy of Sciences RESEARCH NOTE Novel NPHS1 gene mutations in a Chinese family with congenital nephrotic syndrome FENGJIE YANG, YAXIAN CHEN, YU ZHANG, LIRU QIU, YU CHEN and JIANHUA ZHOU Department
Pyridoxine effect in type I primary hyperoxaluria is associated with the most common mutant allele
 Kidney International, Vol. 67 (25), pp. 174 179 GENETIC DISORDERS DEVELOPMENT Pyridoxine effect in type I primary hyperoxaluria is associated with the most common mutant allele CARLA G. MONICO,SANDRO ROSSETTI,JULIE
Kidney International, Vol. 67 (25), pp. 174 179 GENETIC DISORDERS DEVELOPMENT Pyridoxine effect in type I primary hyperoxaluria is associated with the most common mutant allele CARLA G. MONICO,SANDRO ROSSETTI,JULIE
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature10833 Supplementary Table 1: Clinical information for 48 whole exome sequencing samples Sample ID Age Gender tumor location Death OS (months) Recurrence PFS
SUPPLEMENTARY INFORMATION doi:10.1038/nature10833 Supplementary Table 1: Clinical information for 48 whole exome sequencing samples Sample ID Age Gender tumor location Death OS (months) Recurrence PFS
Case Report Primary hyperoxaluria type 1: a case report in a 7-month-old Chinese infant under biopsy and light microscope diagnosis
 Int J Clin Exp Med 2018;11(1):398-402 www.ijcem.com /ISSN:1940-5901/IJCEM0064673 Case Report Primary hyperoxaluria type 1: a case report in a 7-month-old Chinese infant under biopsy and light microscope
Int J Clin Exp Med 2018;11(1):398-402 www.ijcem.com /ISSN:1940-5901/IJCEM0064673 Case Report Primary hyperoxaluria type 1: a case report in a 7-month-old Chinese infant under biopsy and light microscope
Molecular Diagnostic Testing by eyegene: Analysis of Patients With Hereditary Retinal Dystrophy Phenotypes Involving Central Vision Loss
 Genetics Molecular Diagnostic Testing by eyegene: Analysis of Patients With Hereditary Retinal Dystrophy Phenotypes Involving Central Vision Loss Akhila Alapati, 1 Kerry Goetz, 2 John Suk, 1 Mili Navani,
Genetics Molecular Diagnostic Testing by eyegene: Analysis of Patients With Hereditary Retinal Dystrophy Phenotypes Involving Central Vision Loss Akhila Alapati, 1 Kerry Goetz, 2 John Suk, 1 Mili Navani,
Recurrence of Primary Hyperoxaluria After Kidney
 Transplantation Recurrence of Primary Hyperoxaluria After Kidney Transplantation Tahereh Malakoutian, 1 Mojgan Asgari, 2 Massoud Houshmand, 3,4 Ronak Mohammadi, 1 Omid Aryani, 4 Esmaeel Mohammadi Pargoo,
Transplantation Recurrence of Primary Hyperoxaluria After Kidney Transplantation Tahereh Malakoutian, 1 Mojgan Asgari, 2 Massoud Houshmand, 3,4 Ronak Mohammadi, 1 Omid Aryani, 4 Esmaeel Mohammadi Pargoo,
Sample Test Report. Mayo Clinic GeneGuide. Report. Consumer Name DOB: 00/00/0000
 Mayo Clinic GeneGuide Report Consumer Name DOB: // Table Of Contents Mayo Clinic GeneGuide Genetic Test Results Demographics and Ordering Information 3 How to Use This Report 4 Carrier Screening - No variants
Mayo Clinic GeneGuide Report Consumer Name DOB: // Table Of Contents Mayo Clinic GeneGuide Genetic Test Results Demographics and Ordering Information 3 How to Use This Report 4 Carrier Screening - No variants
GALAFOLD (migalastat) capsules, for oral use Initial U.S. Approval: 2018
 HIGHLIGHTS OF PRESCRIBING INFORMATION These highlights do not include all the information needed to use GALAFOLD safely and effectively. See full prescribing information for GALAFOLD. GALAFOLD (migalastat)
HIGHLIGHTS OF PRESCRIBING INFORMATION These highlights do not include all the information needed to use GALAFOLD safely and effectively. See full prescribing information for GALAFOLD. GALAFOLD (migalastat)
Ethnic differences in GRHPR mutations in patients with primary hyperoxaluria type 2
 Clin Genet 2014: 86: 342 348 Printed in Singapore. All rights reserved Short Report 2013 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12292 Ethnic differences
Clin Genet 2014: 86: 342 348 Printed in Singapore. All rights reserved Short Report 2013 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12292 Ethnic differences
The InSiGHT Variant Interpretation Committee: the first ClinGen Expert Panel
 ACGS, June 2018 The InSiGHT Variant Interpretation Committee: the first ClinGen Expert Panel Ian M. Frayling Member of Council & Variant Interpretation Committee (VIC) International Society for Gastrointestinal
ACGS, June 2018 The InSiGHT Variant Interpretation Committee: the first ClinGen Expert Panel Ian M. Frayling Member of Council & Variant Interpretation Committee (VIC) International Society for Gastrointestinal
Genotype to phenotype correlations in cartilage oligomeric matrix protein associated chondrodysplasias
 (2014) 22, 1278 1282 & 2014 Macmillan Publishers Limited All rights reserved 1018-4813/14 www.nature.com/ejhg ARTICLE Genotype to phenotype correlations in cartilage oligomeric matrix protein associated
(2014) 22, 1278 1282 & 2014 Macmillan Publishers Limited All rights reserved 1018-4813/14 www.nature.com/ejhg ARTICLE Genotype to phenotype correlations in cartilage oligomeric matrix protein associated
ATM germline heterozygosity does not play a role in CLL initiation but influences rapid disease progression through loss of the remaining ATM allele
 Published Ahead of Print on September 20, 2011, as doi:10.3324/haematol.2011.048827. Copyright 2011 Ferrata Storti Foundation. Early Release Paper ATM germline heterozygosity does not play a role in CLL
Published Ahead of Print on September 20, 2011, as doi:10.3324/haematol.2011.048827. Copyright 2011 Ferrata Storti Foundation. Early Release Paper ATM germline heterozygosity does not play a role in CLL
Unusual clinical outcome of primary Hyperoxaluria type 1 in Tunisian patients carrying 33_34InsC mutation
 Mbarek et al. BMC Nephrology (2017) 18:195 DOI 10.1186/s12882-017-0612-8 RESEARCH ARTICLE Unusual clinical outcome of primary Hyperoxaluria type 1 in Tunisian patients carrying 33_34InsC mutation Ibtihel
Mbarek et al. BMC Nephrology (2017) 18:195 DOI 10.1186/s12882-017-0612-8 RESEARCH ARTICLE Unusual clinical outcome of primary Hyperoxaluria type 1 in Tunisian patients carrying 33_34InsC mutation Ibtihel
Variant interpretation exercise. ACGS Somatic Variant Interpretation Workshop Joanne Mason 21/09/18
 Variant interpretation exercise ACGS Somatic Variant Interpretation Workshop Joanne Mason 21/09/18 Format of exercise Compile a list of tricky variants across solid cancer and haematological malignancy.
Variant interpretation exercise ACGS Somatic Variant Interpretation Workshop Joanne Mason 21/09/18 Format of exercise Compile a list of tricky variants across solid cancer and haematological malignancy.
Supplementary Online Content
 Supplementary Online Content Ku CA, Hull S, Arno G, et al. Detailed clinical phenotype and molecular genetic findings in CLN3-associated isolated retinal degeneration. JAMA Ophthalmol. Published online
Supplementary Online Content Ku CA, Hull S, Arno G, et al. Detailed clinical phenotype and molecular genetic findings in CLN3-associated isolated retinal degeneration. JAMA Ophthalmol. Published online
Supplemental Table 1. The list of variants with their respective scores for each variant classifier Gene DNA Protein Align-GVGD Polyphen-2 CADD MAPP
 Supplemental Table 1. The list of variants with their respective scores for each variant classifier Gene DNA Protein Align-GVGD Polyphen-2 CADD MAPP Frequency Domain Mammals a 3 S/P b Mammals a 3 S/P b
Supplemental Table 1. The list of variants with their respective scores for each variant classifier Gene DNA Protein Align-GVGD Polyphen-2 CADD MAPP Frequency Domain Mammals a 3 S/P b Mammals a 3 S/P b
JULY 21, Genetics 101: SCN1A. Katie Angione, MS CGC Certified Genetic Counselor CHCO Neurology
 JULY 21, 2018 Genetics 101: SCN1A Katie Angione, MS CGC Certified Genetic Counselor CHCO Neurology Disclosures: I have no financial interests or relationships to disclose. Objectives 1. Review genetic
JULY 21, 2018 Genetics 101: SCN1A Katie Angione, MS CGC Certified Genetic Counselor CHCO Neurology Disclosures: I have no financial interests or relationships to disclose. Objectives 1. Review genetic
Beta Thalassemia Case Study Introduction to Bioinformatics
 Beta Thalassemia Case Study Sami Khuri Department of Computer Science San José State University San José, California, USA sami.khuri@sjsu.edu www.cs.sjsu.edu/faculty/khuri Outline v Hemoglobin v Alpha
Beta Thalassemia Case Study Sami Khuri Department of Computer Science San José State University San José, California, USA sami.khuri@sjsu.edu www.cs.sjsu.edu/faculty/khuri Outline v Hemoglobin v Alpha
chr6: _ del HFE NM_ ,744 bp deletion Deletion of entire HFE gene 2
 Table S1. Details of pathogenic and possibly pathogenic variants identified in the HH genes HFE, HFE2, HAMP, TFR2 and SLC40A1 from published reports. Variants have been classified as either 1, possibly
Table S1. Details of pathogenic and possibly pathogenic variants identified in the HH genes HFE, HFE2, HAMP, TFR2 and SLC40A1 from published reports. Variants have been classified as either 1, possibly
Concurrent Practical Session ACMG Classification
 Variant Effect Prediction Training Course 6-8 November 2017 Prague, Czech Republic Concurrent Practical Session ACMG Classification Andreas Laner / Anna Benet-Pagès 1 Content 1. Background... 3 2. Aim
Variant Effect Prediction Training Course 6-8 November 2017 Prague, Czech Republic Concurrent Practical Session ACMG Classification Andreas Laner / Anna Benet-Pagès 1 Content 1. Background... 3 2. Aim
*Correspondence and offprint requests to: Takashi Sekine; ORIGINAL ARTICLE ABSTRACT INTRODUCTION
 Nephrol Dial Transplant (2014) 29: 376 384 doi: 10.1093/ndt/gft394 Advance Access publication 29 September 2013 Japanese Dent disease has a wider clinical spectrum than Dent disease in Europe/USA: genetic
Nephrol Dial Transplant (2014) 29: 376 384 doi: 10.1093/ndt/gft394 Advance Access publication 29 September 2013 Japanese Dent disease has a wider clinical spectrum than Dent disease in Europe/USA: genetic
Genotype-Phenotype in Egyptian Patients with Nephropathic Cystinosis. (December 2012 report)
 Genotype-Phenotype in Egyptian Patients with Nephropathic Cystinosis (December 2012 report) This is the first study of the genotype of Nephropathic Cystinosis (NC) patients in Egypt and the region of North
Genotype-Phenotype in Egyptian Patients with Nephropathic Cystinosis (December 2012 report) This is the first study of the genotype of Nephropathic Cystinosis (NC) patients in Egypt and the region of North
Spectrum of mutations in CFTR in Finland: 18 years follow-up study and identification of two novel mutations
 Journal of Cystic Fibrosis 4 (2005) 233 237 www.elsevier.com/locate/jcf Spectrum of mutations in CFTR in Finland: 18 years follow-up study and identification of two novel mutations Satu Kinnunen a, Sandra
Journal of Cystic Fibrosis 4 (2005) 233 237 www.elsevier.com/locate/jcf Spectrum of mutations in CFTR in Finland: 18 years follow-up study and identification of two novel mutations Satu Kinnunen a, Sandra
Reverse phenotyping: MCD Detection without clinical diagnosis. Diagnostic: NGS panels versus WES
 Clinical Genetics Leading the way in genetic issues Reverse phenotyping: MCD Detection without clinical diagnosis Diagnostic: NGS panels versus WES Martina Wilke Laboratory Specialist Clinical Genetics
Clinical Genetics Leading the way in genetic issues Reverse phenotyping: MCD Detection without clinical diagnosis Diagnostic: NGS panels versus WES Martina Wilke Laboratory Specialist Clinical Genetics
Reporting TP53 gene analysis results in CLL
 Reporting TP53 gene analysis results in CLL Mutations in TP53 - From discovery to clinical practice in CLL Discovery Validation Clinical practice Variant diversity *Leroy at al, Cancer Research Review
Reporting TP53 gene analysis results in CLL Mutations in TP53 - From discovery to clinical practice in CLL Discovery Validation Clinical practice Variant diversity *Leroy at al, Cancer Research Review
Deep phenotyping of 89 xeroderma pigmentosum patients reveals unexpected heterogeneity dependent on the precise molecular defect
 Deep phenotyping of 89 xeroderma pigmentosum patients reveals unexpected heterogeneity dependent on the precise molecular defect Hiva Fassihi a,1,2, Mieran Sethi b,1, Heather Fawcett c, Jonathan Wing c,
Deep phenotyping of 89 xeroderma pigmentosum patients reveals unexpected heterogeneity dependent on the precise molecular defect Hiva Fassihi a,1,2, Mieran Sethi b,1, Heather Fawcett c, Jonathan Wing c,
Table S2. Putative mutations identified in the CHH cohort
 Table S. Putative utations identified in the CHH cohort Sapl e Phenotyp e Gene nt change aa change Zyg SIF T PPH MaxEn t In vitro ExA C ExAC NFE Previous report ACMG classificatio n ACMG evidence codes
Table S. Putative utations identified in the CHH cohort Sapl e Phenotyp e Gene nt change aa change Zyg SIF T PPH MaxEn t In vitro ExA C ExAC NFE Previous report ACMG classificatio n ACMG evidence codes
Article. Predictors of Incident ESRD among Patients with Primary Hyperoxaluria Presenting Prior to Kidney Failure
 CJASN epress. Published on December 10, 2015 as doi: 10.2215/CJN.02810315 Article Predictors of Incident ESRD among Patients with Primary Hyperoxaluria Presenting Prior to Kidney Failure Fang Zhao,* Eric
CJASN epress. Published on December 10, 2015 as doi: 10.2215/CJN.02810315 Article Predictors of Incident ESRD among Patients with Primary Hyperoxaluria Presenting Prior to Kidney Failure Fang Zhao,* Eric
Laboratory diagnosis of Primary hyperoxaluria. Dr Gill Rumsby UCL Hospitals London, UK
 Laboratory diagnosis of Primary hyperoxaluria Dr Gill Rumsby UCL Hospitals London, UK Steps involved in diagnosis Metabolite Enzyme (protein) Gene (DNA) Tissue/body fluid needed for analysis Metabolite
Laboratory diagnosis of Primary hyperoxaluria Dr Gill Rumsby UCL Hospitals London, UK Steps involved in diagnosis Metabolite Enzyme (protein) Gene (DNA) Tissue/body fluid needed for analysis Metabolite
Genotype phenotype correlation in a cohort of Portuguese patients comprising the entire spectrum of VWD types: impact of NGS
 Coagulation and Fibrinolysis 17 Genotype phenotype correlation in a cohort of Portuguese patients comprising the entire spectrum of VWD types: impact of NGS Teresa Fidalgo 1 ; Ramon Salvado 1 ; Irene Corrales
Coagulation and Fibrinolysis 17 Genotype phenotype correlation in a cohort of Portuguese patients comprising the entire spectrum of VWD types: impact of NGS Teresa Fidalgo 1 ; Ramon Salvado 1 ; Irene Corrales
Prevalence of BRCA1/BRCA2 mutations in a Brazilian population sample at-risk for hereditary breast cancer and characterization of its genetic ancestry
 /, 2016, Vol. 7, (No. 49), pp: 80465-80481 Prevalence of BRCA1/BRCA2 mutations in a Brazilian population sample at- for hereditary breast cancer and characterization of its genetic ancestry Gabriela C.
/, 2016, Vol. 7, (No. 49), pp: 80465-80481 Prevalence of BRCA1/BRCA2 mutations in a Brazilian population sample at- for hereditary breast cancer and characterization of its genetic ancestry Gabriela C.
Novel mutations of PKD genes in Chinese patients suffering from autosomal dominant polycystic kidney disease and seeking assisted reproduction
 He et al. BMC Medical Genetics (2018) 19:186 https://doi.org/10.1186/s12881-018-0693-7 RESEARCH ARTICLE Novel mutations of PKD genes in Chinese patients suffering from autosomal dominant polycystic kidney
He et al. BMC Medical Genetics (2018) 19:186 https://doi.org/10.1186/s12881-018-0693-7 RESEARCH ARTICLE Novel mutations of PKD genes in Chinese patients suffering from autosomal dominant polycystic kidney
SUPPLEMENTARY INFORMATION. Rare independent mutations in renal salt handling genes contribute to blood pressure variation
 SUPPLEMENTARY INFORMATION Rare independent mutations in renal salt handling genes contribute to blood pressure variation Weizhen Ji, Jia Nee Foo, Brian J. O Roak, Hongyu Zhao, Martin G. Larson, David B.
SUPPLEMENTARY INFORMATION Rare independent mutations in renal salt handling genes contribute to blood pressure variation Weizhen Ji, Jia Nee Foo, Brian J. O Roak, Hongyu Zhao, Martin G. Larson, David B.
doi: /brain/awp336 Brain 2010: 133;
 doi:10.1093/brain/awp336 Brain 2010: 133; 655 670 655 BRAIN A JOURNAL OF NEUROLOGY Glucose transporter-1 deficiency syndrome: the expanding clinical and genetic spectrum of a treatable disorder Wilhelmina
doi:10.1093/brain/awp336 Brain 2010: 133; 655 670 655 BRAIN A JOURNAL OF NEUROLOGY Glucose transporter-1 deficiency syndrome: the expanding clinical and genetic spectrum of a treatable disorder Wilhelmina
Re-classification of variants: Implications from the cardiac perspective
 Re-classification of variants: Implications from the cardiac perspective Karen McGuire Oxford Medical Genetics Laboratories, Churchill Hospital, Old Road, Oxford, OX3 7LE. Re-classification New evidence
Re-classification of variants: Implications from the cardiac perspective Karen McGuire Oxford Medical Genetics Laboratories, Churchill Hospital, Old Road, Oxford, OX3 7LE. Re-classification New evidence
Supplementary Figure 1
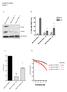 Supplementary Figure 1 A B HEC1A PTEN +/+ HEC1A PTEN -/- 16 HEC1A PTEN -/- 22 PTEN RAD51 % Cells with RAD51 foci 40 -IR +IR 30 20 10 * *! TUBULIN 0 HEC1A PTEN+/+ HEC1A PTEN-/- 16 HEC1A PTEN-/- 22 C D 1.0
Supplementary Figure 1 A B HEC1A PTEN +/+ HEC1A PTEN -/- 16 HEC1A PTEN -/- 22 PTEN RAD51 % Cells with RAD51 foci 40 -IR +IR 30 20 10 * *! TUBULIN 0 HEC1A PTEN+/+ HEC1A PTEN-/- 16 HEC1A PTEN-/- 22 C D 1.0
p.arg119gly p.arg119his p.ala179thr c.540+1g>a c.617_633+6del Prediction basis structure
 a Missense ATG p.arg119gly p.arg119his p.ala179thr p.ala189val p.gly206trp p.gly206arg p.arg251his p.ala257thr TGA 5 UTR 1 2 3 4 5 6 7 3 UTR Splice Site, Frameshift b Mutation p.gly206trp p.gly4fsx50 c.138+1g>a
a Missense ATG p.arg119gly p.arg119his p.ala179thr p.ala189val p.gly206trp p.gly206arg p.arg251his p.ala257thr TGA 5 UTR 1 2 3 4 5 6 7 3 UTR Splice Site, Frameshift b Mutation p.gly206trp p.gly4fsx50 c.138+1g>a
Research Article Identification of Novel ARSA Mutations in Chinese Patients with Metachromatic Leukodystrophy
 International Journal of Genomics Volume 2018, Article ID 2361068, 9 pages https://doi.org/10.1155/2018/2361068 Research Article Identification of Novel ARSA Mutations in Chinese Patients with Metachromatic
International Journal of Genomics Volume 2018, Article ID 2361068, 9 pages https://doi.org/10.1155/2018/2361068 Research Article Identification of Novel ARSA Mutations in Chinese Patients with Metachromatic
Investigating the Patterns in SCN8A Mutations Linked to Early-Onset Seizures
 Investigating the Patterns in SCN8A Mutations Linked to Early-Onset Seizures Item Type text; Electronic Thesis Authors Chen, Debbie Publisher The University of Arizona. Rights Copyright is held by the
Investigating the Patterns in SCN8A Mutations Linked to Early-Onset Seizures Item Type text; Electronic Thesis Authors Chen, Debbie Publisher The University of Arizona. Rights Copyright is held by the
Phenylalanine hydroxylase deficiency in Mexico: genotype phenotype correlations, BH 4 responsiveness and evidence of a founder effect
 Clin Genet 2015: 88: 62 67 Printed in Singapore. All rights reserved Short Report 2014 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12444 Phenylalanine hydroxylase
Clin Genet 2015: 88: 62 67 Printed in Singapore. All rights reserved Short Report 2014 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12444 Phenylalanine hydroxylase
Presentation and role of transplantation in adult patients with type 1 primary hyperoxaluria and the I244T AGXT mutation: Single-center experience
 http://www.kidney-international.org & 2006 International Society of Nephrology original article See commentary on page 984 Presentation and role of transplantation in adult patients with type 1 primary
http://www.kidney-international.org & 2006 International Society of Nephrology original article See commentary on page 984 Presentation and role of transplantation in adult patients with type 1 primary
Mutational and phenotypical spectrum of phenylalanine hydroxylase deficiency in Denmark
 Clin Genet 2016: 90: 247 251 Printed in Singapore. All rights reserved Short Report 2015 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12692 Mutational and
Clin Genet 2016: 90: 247 251 Printed in Singapore. All rights reserved Short Report 2015 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12692 Mutational and
Beta Thalassemia Sami Khuri Department of Computer Science San José State University Spring 2015
 Bioinformatics in Medical Product Development SMPD 287 Three Beta Thalassemia Sami Khuri Department of Computer Science San José State University Hemoglobin Outline Anatomy of a gene Hemoglobinopathies
Bioinformatics in Medical Product Development SMPD 287 Three Beta Thalassemia Sami Khuri Department of Computer Science San José State University Hemoglobin Outline Anatomy of a gene Hemoglobinopathies
Effect of conservative treatment on the renal outcome of children with primary hyperoxaluria type 1
 http://www.kidney-international.org & 2009 International Society of Nephrology Effect of conservative treatment on the renal outcome of children with primary hyperoxaluria type 1 Sonia Fargue 1,Jérôme
http://www.kidney-international.org & 2009 International Society of Nephrology Effect of conservative treatment on the renal outcome of children with primary hyperoxaluria type 1 Sonia Fargue 1,Jérôme
Supplementary Online Content
 Supplementary Online Content Mandelker D, Zhang L, Kemel Y, et al. Mutation detection in patients with advanced cancer by universal sequencing of cancer-related genes in tumor and normal DNA vs guideline-based
Supplementary Online Content Mandelker D, Zhang L, Kemel Y, et al. Mutation detection in patients with advanced cancer by universal sequencing of cancer-related genes in tumor and normal DNA vs guideline-based
UvA-DARE (Digital Academic Repository) Familial hypercholesterolemia: the Dutch approach Huijgen, R. Link to publication
 UvA-DARE (Digital Academic Repository) Familial hypercholesterolemia: the Dutch approach Huijgen, R. Link to publication Citation for published version (APA): Huijgen, R. (2012). Familial hypercholesterolemia:
UvA-DARE (Digital Academic Repository) Familial hypercholesterolemia: the Dutch approach Huijgen, R. Link to publication Citation for published version (APA): Huijgen, R. (2012). Familial hypercholesterolemia:
Somatic mutations in ATP1A1 and ATP2B3 lead to aldosterone-producing adenomas and secondary hypertension
 Somatic mutations in ATP1A1 and ATP2B3 lead to aldosterone-producing adenomas and secondary hypertension Felix Beuschlein, Sheerazed Boulkroun, Andrea Osswald, Thomas Wieland, Hang N. Nielsen, Urs D. Lichtenauer,
Somatic mutations in ATP1A1 and ATP2B3 lead to aldosterone-producing adenomas and secondary hypertension Felix Beuschlein, Sheerazed Boulkroun, Andrea Osswald, Thomas Wieland, Hang N. Nielsen, Urs D. Lichtenauer,
Chromosome 15 Chromosome 17 Chromosome 20
 Chromosome 1 Chromosome 6 Chromosome 11 Chromosome 15 Chromosome 17 Chromosome 20 Supplementary figure 1. Regions of suggestive linkage in PVOD families 1,2 and 3. Six regions were identified with suggestive
Chromosome 1 Chromosome 6 Chromosome 11 Chromosome 15 Chromosome 17 Chromosome 20 Supplementary figure 1. Regions of suggestive linkage in PVOD families 1,2 and 3. Six regions were identified with suggestive
Supplementary Materials for
 www.sciencetranslationalmedicine.org/cgi/content/full/7/270/270ra6/dc1 Supplementary Materials for Integrated allelic, transcriptional, and phenomic dissection of the cardiac effects of titin truncations
www.sciencetranslationalmedicine.org/cgi/content/full/7/270/270ra6/dc1 Supplementary Materials for Integrated allelic, transcriptional, and phenomic dissection of the cardiac effects of titin truncations
Supplementary Material Figure S1. RT-PCR validation and overall growth of pitpnm3, sars, and lemd3 morphants. Table S1. Cohort demographics.
 Supplementary Material Figure S1. RT-PCR validation and overall growth of pitpnm3, sars, and lemd3 morphants. Table S1. Cohort demographics. Table S2. Gene symbols and data regarding potential oligogenic
Supplementary Material Figure S1. RT-PCR validation and overall growth of pitpnm3, sars, and lemd3 morphants. Table S1. Cohort demographics. Table S2. Gene symbols and data regarding potential oligogenic
THE ROLE OF CFTR MUTATIONS IN CAUSING CYSTIC FIBROSIS (CF)
 THE ROLE OF CFTR MUTATIONS IN CAUSING CYSTIC FIBROSIS (CF) Vertex Pharmaceuticals Incorporated, 50 Northern Avenue, Boston, MA 02210. Vertex and the Vertex triangle logo are registered trademarks of Vertex
THE ROLE OF CFTR MUTATIONS IN CAUSING CYSTIC FIBROSIS (CF) Vertex Pharmaceuticals Incorporated, 50 Northern Avenue, Boston, MA 02210. Vertex and the Vertex triangle logo are registered trademarks of Vertex
BIOCHEMICAL AND MOLECULAR HETEROGENEITY IN CARNITINE PALMITOYLTRANSFERASE II DEFICIENCY
 BIOCHEMICAL AND MOLECULAR HETEROGENEITY IN CARNITINE PALMITOYLTRANSFERASE II DEFICIENCY Carmen Sousa, Helena Fonseca, Hugo Rocha, Ana Marcão, Laura Vilarinho, Luísa Diogo, Sílvia Sequeira, Cristina Costa,
BIOCHEMICAL AND MOLECULAR HETEROGENEITY IN CARNITINE PALMITOYLTRANSFERASE II DEFICIENCY Carmen Sousa, Helena Fonseca, Hugo Rocha, Ana Marcão, Laura Vilarinho, Luísa Diogo, Sílvia Sequeira, Cristina Costa,
Supplementary Information. Exome sequencing of serous endometrial tumors identifies recurrent somatic
 Supplementary Information Exome sequencing of serous endometrial tumors identifies recurrent somatic mutations in chromatin-remodeling and ubiquitin ligase complex genes Matthieu Le Gallo 1, Andrea J.
Supplementary Information Exome sequencing of serous endometrial tumors identifies recurrent somatic mutations in chromatin-remodeling and ubiquitin ligase complex genes Matthieu Le Gallo 1, Andrea J.
Supplementary appendix
 Supplementary appendix This appendix formed part of the original submission and has been peer reviewed. We post it as supplied by the authors. Supplement to: Copson ER, Maishman TC, Tapper WJ, et al. Germline
Supplementary appendix This appendix formed part of the original submission and has been peer reviewed. We post it as supplied by the authors. Supplement to: Copson ER, Maishman TC, Tapper WJ, et al. Germline
Laboratory diagnosis of Primary hyperoxaluria. Dr Gill Rumsby UCL Hospitals London, UK
 Laboratory diagnosis of Primary hyperoxaluria Dr Gill Rumsby UCL Hospitals London, UK Steps involved in diagnosis Metabolite Enzyme (protein) Gene (DNA) Tissue/body fluid needed for analysis Metabolite
Laboratory diagnosis of Primary hyperoxaluria Dr Gill Rumsby UCL Hospitals London, UK Steps involved in diagnosis Metabolite Enzyme (protein) Gene (DNA) Tissue/body fluid needed for analysis Metabolite
Genetic and biochemical characteristics of patients with hyperlipidemia who require LDL apheresis
 Genetic and biochemical characteristics of patients with hyperlipidemia who require LDL apheresis Mato Nagel, Ioan Duma, Constantina Vlad, Mandy Benke, Sylvia Nagorka Zentrum für Nephrologie & Stoffwechsel,
Genetic and biochemical characteristics of patients with hyperlipidemia who require LDL apheresis Mato Nagel, Ioan Duma, Constantina Vlad, Mandy Benke, Sylvia Nagorka Zentrum für Nephrologie & Stoffwechsel,
Frequent incidence of BARD1-truncating mutations in germline DNA from triple-negative breast cancer patients
 Clin Genet 2016: 89: 336 340 Printed in Singapore. All rights reserved Short Report 2015 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12620 Frequent incidence
Clin Genet 2016: 89: 336 340 Printed in Singapore. All rights reserved Short Report 2015 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd CLINICAL GENETICS doi: 10.1111/cge.12620 Frequent incidence
Supp. Materials and Methods
 Böhm et al., Human Mutation 1 Supp. Materials and Methods Patients Patient selection was based on the clinical presentation and/or histological findings suggesting centronuclear myopathy: muscle weakness
Böhm et al., Human Mutation 1 Supp. Materials and Methods Patients Patient selection was based on the clinical presentation and/or histological findings suggesting centronuclear myopathy: muscle weakness
CONTRACTING ORGANIZATION: Massachusetts General Hospital Charlestown, MA 02129
 AD Award Number: DAMD17-03-1-0445 TITLE: Molecular Identification of the Schwannomatosis Locus PRINCIPAL INVESTIGATOR: Mia MacCollin, M.D. CONTRACTING ORGANIZATION: Massachusetts General Hospital Charlestown,
AD Award Number: DAMD17-03-1-0445 TITLE: Molecular Identification of the Schwannomatosis Locus PRINCIPAL INVESTIGATOR: Mia MacCollin, M.D. CONTRACTING ORGANIZATION: Massachusetts General Hospital Charlestown,
ANALYSIS OF IL17 AND IL17RA POLYMORPHISMS IN SPANISH PSORIASIS PATIENTS: ASSOCIATION WITH RISK FOR DISEASE.
 ANALYSIS OF IL17 AND IL17RA POLYMORPHISMS IN SPANISH PSORIASIS PATIENTS: ASSOCIATION WITH RISK FOR DISEASE. Batalla A, Coto E*, González-Lara L, González- Fernández D, Maldonado-Seral C, García-García
ANALYSIS OF IL17 AND IL17RA POLYMORPHISMS IN SPANISH PSORIASIS PATIENTS: ASSOCIATION WITH RISK FOR DISEASE. Batalla A, Coto E*, González-Lara L, González- Fernández D, Maldonado-Seral C, García-García
J Clin Oncol 34: by American Society of Clinical Oncology INTRODUCTION
 VOLUME 34 NUMBER 34 DECEMBER 1, 2016 JOURNAL OF CLINICAL ONCOLOGY O R I G I N A L R E P O R T Conflicting Interpretation of Genetic Variants and Cancer Risk by Commercial Laboratories as Assessed by the
VOLUME 34 NUMBER 34 DECEMBER 1, 2016 JOURNAL OF CLINICAL ONCOLOGY O R I G I N A L R E P O R T Conflicting Interpretation of Genetic Variants and Cancer Risk by Commercial Laboratories as Assessed by the
Endocrine Surgery. Characteristics of the Germline MEN1 Mutations in Korea: A Literature Review ORIGINAL ARTICLE. The Korean Journal of INTRODUCTION
 ORIGINAL ARTICLE ISSN 1598-1703 (Print) ISSN 2287-6782 (Online) Korean J Endocrine Surg 2014;14:7-11 The Korean Journal of Endocrine Surgery Characteristics of the Germline MEN1 Mutations in Korea: A Literature
ORIGINAL ARTICLE ISSN 1598-1703 (Print) ISSN 2287-6782 (Online) Korean J Endocrine Surg 2014;14:7-11 The Korean Journal of Endocrine Surgery Characteristics of the Germline MEN1 Mutations in Korea: A Literature
Molecular Diagnostic Laboratory 18 Sequencing St, Gene Town, ZY Tel: Fax:
 Molecular Diagnostic Laboratory 18 Sequencing St, Gene Town, ZY 01234 Tel: 555-920-3333 Fax: 555-920-3334 www.moldxlaboratory.com Patient Name: Jane Doe Specimen type: Blood, peripheral DOB: 04/05/1990
Molecular Diagnostic Laboratory 18 Sequencing St, Gene Town, ZY 01234 Tel: 555-920-3333 Fax: 555-920-3334 www.moldxlaboratory.com Patient Name: Jane Doe Specimen type: Blood, peripheral DOB: 04/05/1990
MUTATION TEST CE IVD. ctnras-braf FEATURES
 ctnras-braf CE IVD TECHNICAL SHEET IDYLLA ctnras-braf MUTATION TEST The Idylla ctnras-braf Mutation Test, performed on the Biocartis Idylla System, is an in vitro diagnostic test for the qualitative detection
ctnras-braf CE IVD TECHNICAL SHEET IDYLLA ctnras-braf MUTATION TEST The Idylla ctnras-braf Mutation Test, performed on the Biocartis Idylla System, is an in vitro diagnostic test for the qualitative detection
Deliverable 2.1 List of relevant genetic variants for pre-emptive PGx testing
 GA N 668353 H2020 Research and Innovation Deliverable 2.1 List of relevant genetic variants for pre-emptive PGx testing WP N and Title: WP2 - Towards shared European Guidelines for PGx Lead beneficiary:
GA N 668353 H2020 Research and Innovation Deliverable 2.1 List of relevant genetic variants for pre-emptive PGx testing WP N and Title: WP2 - Towards shared European Guidelines for PGx Lead beneficiary:
A guide to understanding variant classification
 White paper A guide to understanding variant classification In a diagnostic setting, variant classification forms the basis for clinical judgment, making proper classification of variants critical to your
White paper A guide to understanding variant classification In a diagnostic setting, variant classification forms the basis for clinical judgment, making proper classification of variants critical to your
Short Report. Clin Genet 2009: 76: Printed in Singapore. All rights reserved John Wiley & Sons A/S
 Clin Genet 2009: 76: 242 255 Printed in Singapore. All rights reserved Short Report 2009 John Wiley & Sons A/S CLINICAL GENETICS doi: 10.1111/j.1399-0004.2009.01241.x APC or MUTYH mutations account for
Clin Genet 2009: 76: 242 255 Printed in Singapore. All rights reserved Short Report 2009 John Wiley & Sons A/S CLINICAL GENETICS doi: 10.1111/j.1399-0004.2009.01241.x APC or MUTYH mutations account for
A Comprehensive Study of TP53 Mutations in Chronic Lymphocytic Leukemia: Analysis of 1,287 Diagnostic CLL Samples
 A Comprehensive Study of TP53 Mutations in Chronic Lymphocytic Leukemia: Analysis of 1,287 Diagnostic CLL Samples Sona Pekova, MD., PhD. Chambon Ltd., Laboratory for molecular diagnostics, Prague, Czech
A Comprehensive Study of TP53 Mutations in Chronic Lymphocytic Leukemia: Analysis of 1,287 Diagnostic CLL Samples Sona Pekova, MD., PhD. Chambon Ltd., Laboratory for molecular diagnostics, Prague, Czech
Use of panel tests in place of single gene tests in the cancer genetics clinic
 Clin Genet 2015: 88: 278 282 Printed in Singapore. All rights reserved CLINICAL GENETICS doi: 10.1111/cge.12488 Short Report se of panel tests in place of single gene tests in the cancer genetics clinic
Clin Genet 2015: 88: 278 282 Printed in Singapore. All rights reserved CLINICAL GENETICS doi: 10.1111/cge.12488 Short Report se of panel tests in place of single gene tests in the cancer genetics clinic
Phenylketonuria (PKU) the Biochemical Basis. Biol 405 Molecular Medicine
 Phenylketonuria (PKU) the Biochemical Basis Biol 405 Molecular Medicine PKU a history In 1934 Følling identified a clinical condition - imbecillitas phenylpyruvica. Mental retardation associated with this
Phenylketonuria (PKU) the Biochemical Basis Biol 405 Molecular Medicine PKU a history In 1934 Følling identified a clinical condition - imbecillitas phenylpyruvica. Mental retardation associated with this
Home Brewed Personalized Genomics
 Home Brewed Personalized Genomics The Quest for Meaningful Analysis Results of a 23andMe Exome Pilot Trio of Myself, Wife, and Son February 22, 2013 Gabe Rudy, Vice President of Product Development Exome
Home Brewed Personalized Genomics The Quest for Meaningful Analysis Results of a 23andMe Exome Pilot Trio of Myself, Wife, and Son February 22, 2013 Gabe Rudy, Vice President of Product Development Exome
Integration Solutions
 Integration Solutions (1) a) With no active glycosyltransferase of either type, an ii individual would not be able to add any sugars to the O form of the lipopolysaccharide. Thus, the only lipopolysaccharide
Integration Solutions (1) a) With no active glycosyltransferase of either type, an ii individual would not be able to add any sugars to the O form of the lipopolysaccharide. Thus, the only lipopolysaccharide
A Phase 1/2 Trial of Lumasiran (ALN-GO1), an Investigational RNAi Therapeutic for Primary Hyperoxaluria Type 1
 A Phase 1/2 Trial of Lumasiran (ALN-GO1), an Investigational RNAi Therapeutic for Primary Hyperoxaluria Type 1 OxalEurope Meeting Naples, Italy 8 June 2018 2018 Alnylam Pharmaceuticals, Inc. Background
A Phase 1/2 Trial of Lumasiran (ALN-GO1), an Investigational RNAi Therapeutic for Primary Hyperoxaluria Type 1 OxalEurope Meeting Naples, Italy 8 June 2018 2018 Alnylam Pharmaceuticals, Inc. Background
Molecular genetics of maple syrup urine disease in the Turkish population
 The Turkish Journal of Pediatrics 2009; 51: 97-102 Original Molecular genetics of maple syrup urine disease in the Turkish population Kerstin Gorzelany 1, Ali Dursun 2, Turgay Coşkun 2, Serap H. Kalkanoğlu-Sivri
The Turkish Journal of Pediatrics 2009; 51: 97-102 Original Molecular genetics of maple syrup urine disease in the Turkish population Kerstin Gorzelany 1, Ali Dursun 2, Turgay Coşkun 2, Serap H. Kalkanoğlu-Sivri
