Application Note # MT-94 Direct Read-out of Thin Layer Chromatography (TLC) using MALDI-TOF
|
|
|
- Rolf Robertson
- 5 years ago
- Views:
Transcription
1 Bruker Daltonics Application Note # MT-94 Direct Read-out of Thin Layer Chromatography (TLC) using MALDI-TOF Thin Layer Chromatography (TLC) is broadly established to separate, characterize and quantify food and biological ingredients/components. Recently interest in mass spectrometric analysis of TLC-separated compounds has increased significantly. Here we will introduce the direct coupling of TLC with MALDI-TOF mass spectrometry: TLC-MALDI; an adapter target for TLC plates and an extension of the standard flexsoftware that turns the MALDI into a molecular TLC readout platform. Also included are the first results from lipid analysis. Introduction TLC separations remain indispensable for a number of applications such as forensics, herbals, food, cosmetics and clinical/industrial applications. Particular analytes which prove difficult for HPLC separations, such as lipids, are widely characterized by TLC. In addition, TLC remains a quick and simple method to monitor the progress of chemical reactions in the organic synthesis laboratory. For many of these applications, the assignment or confirmation of molecular structures to TLC-separated spots is important, as RF ( ratio of fronts ) values alone are often not highly reproducible and do not allow the unequivocal assignment of a certain compound. However, currently established TLC-MALDI protocols involve scratching the chromatographic phase off the carrier and elution of the analyte prior to spectroscopic or mass spectrometric analysis. This process is not only time consuming and carries the risk of loosing some compounds, but it also relies on previously detected and manually marked spots. Therefore, the structural analysis depends on the ability to visualize the particular analyte by specific staining agents or by unspecific fluorescence quenching. As staining methods do not resolve individual compounds with similar RF values, the achievable analytical resolution is typically limited by the detection method. Here we introduce matrixassisted laser desorption/ionisation time-of-flight mass spectrometry (MALDI-TOF MS) to directly read out TLC traces reproducibly whilst maintaining the chromatographic resolving power. Workflow The hyphenated TLC-MALDI approach requires the uniform coverage of the TLC lanes with MALDI matrix as a preparative step comparable with the effort of staining. The mass spectrometric information is subsequently read out automatically. MALDI plate movement along the chromatographic lanes is driven by the standard movement mechanics of the MALDI ion source without additional robotic requirements. A chromatographic run is generated for each lane. It can then be analyzed using TLC-MS software, providing the extracted ion chromatograms of individual analytes independent of dye staining and at a selectable resolution which largely depends on the chromatography itself.
2 Fig. 1: MTP sized TLC-MALDI Adapter Target (Order No #255595) for 50x75 mm TLC Silica gel (aluminum backed). After TLC separation and matrix coating they are ready for MALDI analysis. Experimental TLC All TLC-MALDI experiments were performed using either aluminum backed mm TLC Silica gel 60 F 254 plates, 200 µm layer thickness (Merck, # ) or mm TLC Silica gel 60 plates on aluminum backs, 200 μm layer thickness) (Merck, # ) and developed in horizontal developing TLC chambers using CHCl 3, ethanol, water, triethylamine (35:35:7:35, v/v/v/v) as the solvent system for lipids and phospholipids, whereas glycolipids were separated under acidic conditions (CHCl 3, CH 3 OH, acetic acid (65:25:10, v/v/v)). Lipids were obtained by extraction of cellular suspensions or biological tissues, prepared and visualized by spraying with a solution of primuline as described [1-3]. ImagePrep allows fully automated matrix preparation with highest spatial resolution and reproducibility. A 100 mg/ ml solution of DHB in acetonitrile/water (1:1, v/v) was used for manual matrix application. 50 mg/ml DHB was used for ImagePrep preparations. Generally, in contrast to conventional MALDI, a larger excess of the matrix over the analyte is needed if spectra are to be recorded directly from the TLC plate. TLC-MALDI software Coupling of TLC with MALDI-TOF MS The mm TLC plates were cut to mm to fit the TLC-MALDI Adapter target (Bruker, Order Nr ); mm TLC plates were unaltered (Fig. 1). MALDI spectra were acquired from the matrix coated TLC plates as described in [1]. Matrix was added either manually using conventional dried droplet preparation on discrete primuline stained spots (marked by pencil) or the entire TLC plate was homogeneously coated with matrix (1-5 mg/cm 2 ) for MALDIimaging or TLC-scanning using the ImagePrep preparation device (Bruker Daltonics) [4]. Fig 2: TLC-MALDI setup dialog. The wizard driven software interface allows for simple analysis setup with multiple chromatographic lanes and defined step rasters along with or orthogonal to the chromatographic axis.
3 MALDI-TOF MS Selected Spots: All MALDI spectra from primuline spots were acquired on an Autoflex MALDI-TOF-MS with 50 Hz nitrogen laser. The extraction voltage was 20 kv and gated matrix suppression was applied to prevent the saturation of the detector by matrix ions. All spectra were acquired in reflector mode using delayed extraction as described in [5]. Spectra were calibrated using a standard lipid mixture desorbed from a standard DHB preparation next to the spots of interest (positive ion mode). For negative ion mode, the characteristic signals of the DHB matrix were used for calibration [6]. MALDI: Imaging and scanning of lanes on TLC plates was performed on an ultraflex III MALDI-TOF/TOF with 200 Hz smartbeam laser in reflector mode. The extraction voltage was set to 25 kv and matrix suppression to m/z <200. Data were acquired under Compass 1.2 (FC 3.0, FA 3.0). TLC-Imaging: A pixel raster of 400 μm 400 μm spots was defined with fleximaging 2.0 on the entire TLC lane area. 200 laser shots were accumulated for each pixel. False colors were assigned to each mass of interest and their spatial distributions across the TLC plate were displayed as a heat map. TLC-Scanning: The position of chromatographic lanes on the TLC plate can be read directly from xy-scales on the adapter target and entered into a new dedicated software tool for TLC-MALDI (Fig. 2), which is available as a TLC- MALDI software patch compatible with Compass 1.2 for chromatographic readout. The Compass 1.3+ software will incorporate TLC-MALDI functionality. The location and dimensions of multiple TLC lanes can be defined, as can raster dimensions (here 400 µm). The intuitive, easy-tohandle TLC-MALDI software controls not only automatic data acquisition but also provides a dedicated TLC data viewer and post-processing tools. Results Erythrocyte membrane lipids cells were extracted using CHCl 3 / CH 3 OH/ H 2 O 2:1:1 (v/v/v) as described in more detail in [2]. The TLC plate was analyzed using three different approaches as shown in Fig. 3. The traditional analysis used specific lipid staining by primuline (Fig. 3A), which forms a non-covalent complex with lipids, therefore, not interfering with subsequent MALDI analysis. Three major spots consisting of PE, PC and SM (see Tab. 1) were detectable using primuline and a minor spot represented LPC. Detailed evaluation of the TLC-MALDI imaging dataset revealed 3 more groups of compounds (LPI, PI, PS) and permitted the differentiation of lipids that could not be distinguished chromatographically Abbr. Lipid name MH + PE Phosphatidylethanolamine 16:0/18: G Ganglioside PS Phosphatidylserine 18:0/20: PC Phosphatidylcholine 16:0/20: SM Sphingomyelin 16: LPC Lyso-Phosphatidylcholine 16: PI Phosphatidylinositol 16:0/18: PA Phosphatidic acid 16:0/18: Tab. 1: Detected and characterized lipids from erythrocyte membrane, stem cells and brain: abbreviation, name, structure and typical protonated molecular ion (for several lipids, e.g., MNa + are actually detected). with the staining-scraping procedure: PE 18:0/20:4 vs. PE 16:0/18:1; LPI 22:5 vs. PI 18:0/18:2 (Figs. 3b, 4). Detection of major lipids Although not shown, all major lipids (phospholipids, apolar lipids such as cholesterol, cholesteryl esters and triacylglycerols) present in biological systems are easily detectable using this approach. Our studies have also provided evidence that even low abundance lipids (< 1% of the total lipids) such as PI are easily detectable within a single dataset providing a dynamic range of ~ 3 orders of magnitude. Even mixtures of polyphosphoinositides (PI, PIP, PIP 2 ) can be easily separated and analyzed (PI peak labeled blue in Fig. 3 and present in Fig. 4 b). Glycolipids require a more acidic solvent system in order to separate them from the common and more abundant phospholipids [7]. Although compounds such as gangliosides are characterized by a much higher polarity resulting in reduced detectability, they are easily detectable using this approach. However, glycolipids are often negatively charged (due to the presence of phosphate or sulphate groups), and require negative mode analysis.
4 Classical Staining vs. Imaging and Chromatographic Readout of TLC-MALDI Data x Mass Spectrum A B C TLC Lane MALDI Image MALDI Chromatogram Extracted Ion Cromatogram m/z Fig 3: A) Classical primuline staining of separated lipids on a TLC plate; B) TLC-MALDI imaging analysis of identical TLC plate with matrix coating; C) chromatographic TLC readout as provided by the TLC-MALDI software permitting direct access to all molecular species (m/z values on the x-axis) along the chromatographic separation on the y-axis. The colored circles correspond to the compounds that are visualized in the TLC- MALDI image. Full readout in minutes Although imaging readouts such as Fig. 3B appear attractive as an analysis format comparing directly and favorably with classical stains (Fig. 3A), they require time consuming, manual data interrogation. Therefore, they do not appear well suited for routine analysis purposes. However, chromatographic treatment of the TLC data from acquisition to visualization and analysis appears to be more efficient and powerful, providing a full readout of the information contained in the TLC-MALDI dataset (Fig. 3C). Acquisition times are reduced from hours to ~ 5 min/lane and the large data file size for an image is reduced to a fraction of the size for the chromatogram. Therefore the TLC-MALDI software has been designed to support the quick and simple definition of lanes in the visual interface for the acquisition of chromatographic traces. Up to 4 lanes can be defined per TLC plate. After acquisition, the entire dataset is visualized using a dedicated DataViewer (Fig. 3C). It provides direct access to all separated compounds through a heat map of all MS peaks along the chromatographic separation (y-axis). For a cursor-selected position in the RF vs. m/z plane, the extracted ion chromatogram is displayed on the right side and the corresponding MS spectrum is shown on the top. This representation of the data allows direct access to all chromatographic mass peaks and a direct distinction can be made between the peaks and background ions which are represented by vertical streaks in the DataView. In addition, distinguishing largely overlapping lipids is straight- forward as their mass separation adds another dimension not available with the mass image. Structure confirmation After the analysis of the separated compounds, structure confirmation typically requires the acquisition of MS/MS spectra. The TLC-MALDI DataViewer facilitates manual pick a peak, which permits the direct manual acquisition of MS/MS spectra, in a few seconds, without the need to find the proper spots (not shown).
5 TLC-MALDI Analysis of Erythrocyte Membrane Lipids 8 PE 18:0/20:4 7 PE 16:0/18:1 8 7 PE PE 18:0/20:4 ( m/z = 812.5) PE 16:0/18:1 ( m/z = 762.5) 6 PC 16:0/20:4 Fragments of LPI 22:5 ( m/z = and 551.4) 5 PC 16:0/18:1 PI 18:0/18:2 ( m/z = 885.6) 16:0 LPC 18: SM 16:0 SM 24: m/z (a) PC SM LPC (b) Fragment of PS 16:0/20:4 ( m/z = 618.5) Unknown (Impurity?) ( m/z = ) PC 16:0/20:4 ( m/z = 804.6) PC 16:0/18:1 ( m/z = 760.6) SM 24:0 ( m/z = 837.7) SM 16:0 ( m/z = 725.6) LPC 18:0 ( m/z = 524.3) LPC 16:0 ( m/z = 496.3) Fig. 4: Spectrum overview of various lipids detected in positive ion mode from erythrocyte membrane. Spectra are numbered according to their elution order. TLC-MALDI imaging data (b) proved to be much more sensitive than common staining techniques (a: primuline stain). MS spectra obtained from (b) provided detailed differences in fatty acyl compositions (1-8). Conclusion TLC-MALDI runs on any Bruker Flex mass spectrometer equipped with a scoutmtp ion source under Compass 1.2 and higher. The ImagePrep is equipped with TLC methods to produce a uniform matrix coating with TLC separated lipids. Previously, TLC-MALDI coupling was expected to be problematic [8] because pieces of the stationary TLC phase might disintegrate and damage the MS. However, no such events were observed even under routine laboratory conditions. It is also remarkable that under TLC-MALDI conditions, fragmentation of analytes was not significantly exaggerated in comparison with standard MALDI. The new technology was applied to the analysis of lipids from various sources such as mesenchymal stem cells [2], human erythrocytes [2] or chicken egg [1] extracts providing evidence that MALDI is a great readout platform for TLC separations of lipids [9]. These analyses demonstrated significantly improved detection limits compared to the classical primuline staining approach. Imaging analysis allowed the visualization of many hitherto undetected lipids and to distinguish several compounds in broad primuline spots. Although imaging mass spectrometry compares favourably with primuline staining analytically, the relatively long acquisition time (~1-2 h) and data evaluation process associated with imaging (~1-2 h), calls for different approaches in routine analysis. Therefore, we developed software to automate the linear scanning of TLC traces with scan times in the 5 min range and use DataViewer to work with TLC-MALDI data in chromatographic format, which drastically reduces manual evaluation times of the analysis to few minutes. All, hardware and software methods are available now for the simple application of TLC-MALDI to routine analysis. Use of optimized matrix protocols with new matrices (such as 9-aminoacridine) are expected to further extend the application range of this new and exciting method. TLC- MALDI is interesting in a great number of applications such as oligosaccharides [10], food, cosmetics, herbals or for quick analytical support in the organic chemical synthesis lab.
6 Bruker Daltonics is continually improving its products and reserves the right to change specifications without notice. Bruker Daltonics , # Acknowledgements We thank Michael Schulz, Merck KGaA, Darmstadt, Germany, for providing various phases and plate formats for protocol optimization. References [1] B. Fuchs, J. Schiller, R. Süß, M. Schürenberg and D. Suckau 2007 A direct and simple method of coupling matrix-assisted laser desorption and ionization time-of-flight mass spectrometry (MALDI-TOF MS) to thin-layer chromatography (TLC) for the analysis of phospholipids from egg yolk Anal. Bioanal. Chem. 389: [2] J. Schiller, B. Fuchs, R. Süß, M. Zscharnack, A. Bader, P. Müller, M. Schürenberg, M. Becker and D. Suckau 2008 Analysis of stem cell lipids by offline TLC-MALDI-TOF MS Anal. Bioanal. Chem. 392: [3] B. Fuchs, J. Schiller, R. Süß, A. Nimptsch, M. Schürenberg and D. Suckau 2008 Capabilities and drawbacks of combined matrix-assisted laser desorption and ionization time-of-flight mass spectrometry (MALDI-TOF MS) and high-performance thin-layer chromatography (TLC): Analysis of egg yolk lipids JPC J. Planar Chromatogr. (in press). [4] Bruker Daltonics Technical Note 18, A New Matrix Application Device for MALDI Tissue Imaging [5] J. Schiller, R. Süß, J. Arnhold, B. Fuchs, J. Leßig, M. Müller, M. Petkovic, H. Spalteholz, O. Zschörnig and K. Arnold 2004 Matrix-assisted laser desorption and ionization time-of-flight (MALDI-TOF) mass spectrometry in lipid and phospholipid research Prog. Lipid Res. 43, [6] J. Schiller, R. Süß, B. Fuchs, M. Müller, M. Petkovic, O. Zschörnig and H. Waschipky 2007 The suitability of different DHB isomers as matrices for the MALDI-TOF MS analysis of phospholipids: which isomer for what purpose? Eur. Biophys. J. 36, [7] Fuchs, B., Nimptsch, A., Süß, R., Schiller, J Analysis of Brain Lipids by Directly Coupled Matrix-Assisted Laser Desorption Ionization Time-of-Flight Mass Spectrometry and High-Performance Thin-Layer Chromatography J. AOAC Int. 91: [8] Fuchs, B., Süß, R., Nimptsch, A., Schiller, J. 200 Matrix-assisted laser desorption and ionization time-of-flight mass spectrometry (MALDI-TOF MS) directly combined with thin-layer chromatography (TLC) - A review of the current state. CHROMATOGRAPHIA. [9] Franz-Josef Mayer-Posner, Jochen Franzen 2002 United States Patent TLC/MALDI carrier plate and method for using the same. [10] Z. Zhang, J. Xie, F. Zhang, and R.J. Linhardt R.J Thin-layer chromatography for the analysis of glycosaminoglycan oligosaccharides. Anal. Biochem. 371, Authors Martin Schürenberg 1, Detlev Suckau 1, Beate Fuchs 2 and Jürgen Schiller 2 1 Bruker Daltonics, Bremen, Germany 2 Inst. Med. Physics & Biophysics, Univ. Leipzig, Germany For research use only. Not for use in diagnostic procedures. Keywords TLC-MALDI lipids Instrumentation & Software autoflex TOF/TOF ultraflex TOF/TOF ImagePrep flexsoftware 1.2 TLC-MALDI Adapter Target # Bruker Daltonik GmbH Bremen Germany Phone +49 (421) Fax +49 (421) sales@bdal.de Bruker Daltonics Inc. Billerica, MA USA Phone +1 (978) Fax +1 (978) ms-sales@bdal.com
Bruker Daltonics. autoflex III smartbeam. The Standard in MALDI-TOF Performance MALDI-TOF/TOF. think forward
 Bruker Daltonics autoflex III smartbeam The Standard in MALDI-TOF Performance think forward MALDI-TOF/TOF Designed for a Routine High Level of Performance The autoflex III smartbeam brings the power of
Bruker Daltonics autoflex III smartbeam The Standard in MALDI-TOF Performance think forward MALDI-TOF/TOF Designed for a Routine High Level of Performance The autoflex III smartbeam brings the power of
Technical Note # TN-31 Redefining MALDI-TOF/TOF Performance
 Bruker Daltonics Technical Note # TN-31 Redefining MALDI-TOF/TOF Performance The new ultraflextreme exceeds all current expectations of MALDI-TOF/TOF technology: A proprietary khz smartbeam-ii TM MALDI
Bruker Daltonics Technical Note # TN-31 Redefining MALDI-TOF/TOF Performance The new ultraflextreme exceeds all current expectations of MALDI-TOF/TOF technology: A proprietary khz smartbeam-ii TM MALDI
Small Molecule Drug Imaging of Mouse Tissue by MALDI-TOF/TOF Mass Spectrometry and FTMS
 Bruker Daltonics Application Note # MT-93/FTMS-38 Small Molecule Drug Imaging of Mouse Tissue by MALDI-TOF/TOF Mass Spectrometry and FTMS Introduction Matrix Assisted Laser Desorption Ionization (MALDI)
Bruker Daltonics Application Note # MT-93/FTMS-38 Small Molecule Drug Imaging of Mouse Tissue by MALDI-TOF/TOF Mass Spectrometry and FTMS Introduction Matrix Assisted Laser Desorption Ionization (MALDI)
Application Note # MT-111 Concise Interpretation of MALDI Imaging Data by Probabilistic Latent Semantic Analysis (plsa)
 Application Note # MT-111 Concise Interpretation of MALDI Imaging Data by Probabilistic Latent Semantic Analysis (plsa) MALDI imaging datasets can be very information-rich, containing hundreds of mass
Application Note # MT-111 Concise Interpretation of MALDI Imaging Data by Probabilistic Latent Semantic Analysis (plsa) MALDI imaging datasets can be very information-rich, containing hundreds of mass
Application Note # MT-91. High Quality MALDI Imaging of Proteins and Peptides in Small Rodent Organ Tissues. Bruker Daltonics.
 Bruker Daltonics Application Note # MT-91 18385 Da 6230 Da 7843 Da High Quality MALDI Imaging of Proteins and Peptides in Small Rodent Organ Tissues New developments in MALDI instrumentation, laser technology
Bruker Daltonics Application Note # MT-91 18385 Da 6230 Da 7843 Da High Quality MALDI Imaging of Proteins and Peptides in Small Rodent Organ Tissues New developments in MALDI instrumentation, laser technology
Glycerolipid Analysis. LC/MS/MS Analytical Services
 Glycerolipid Analysis LC/MS/MS Analytical Services Molecular Characterization and Quantitation of Glycerophospholipids in Commercial Lecithins by High Performance Liquid Chromatography with Mass Spectrometric
Glycerolipid Analysis LC/MS/MS Analytical Services Molecular Characterization and Quantitation of Glycerophospholipids in Commercial Lecithins by High Performance Liquid Chromatography with Mass Spectrometric
MALDI Imaging Drug Imaging Detlev Suckau Head of R&D MALDI Bruker Daltonik GmbH. December 19,
 MALDI Imaging Drug Imaging Detlev Suckau Head of R&D MALDI Bruker Daltonik GmbH December 19, 2014 1 The principle of MALDI imaging Spatially resolved mass spectra are recorded Each mass signal represents
MALDI Imaging Drug Imaging Detlev Suckau Head of R&D MALDI Bruker Daltonik GmbH December 19, 2014 1 The principle of MALDI imaging Spatially resolved mass spectra are recorded Each mass signal represents
Hiroya Hidaka *1), Masaki Takiwaki 2), Mine Yamashita 2), Shinya Otsuki 1), Kenji Kawasaki 3), Mitsutoshi Sugano 3) and Takayuki Honda 4)
 Mild acid hydrolysis of sphingolipids yields lysosphingolipids: a matrix-assisted laser desorption and ionization time-of-flight mass spectrometry study Hiroya Hidaka *1), Masaki Takiwaki 2), Mine Yamashita
Mild acid hydrolysis of sphingolipids yields lysosphingolipids: a matrix-assisted laser desorption and ionization time-of-flight mass spectrometry study Hiroya Hidaka *1), Masaki Takiwaki 2), Mine Yamashita
A Definitive Lipidomics Workflow for Human Plasma Utilizing Off-line Enrichment and Class Specific Separation of Phospholipids
 A Definitive Lipidomics Workflow for Human Plasma Utilizing Off-line Enrichment and Class Specific Separation of Phospholipids Jeremy Netto, 1 Stephen Wong, 1 Federico Torta, 2 Pradeep Narayanaswamy, 2
A Definitive Lipidomics Workflow for Human Plasma Utilizing Off-line Enrichment and Class Specific Separation of Phospholipids Jeremy Netto, 1 Stephen Wong, 1 Federico Torta, 2 Pradeep Narayanaswamy, 2
LC/MS Method for Comprehensive Analysis of Plasma Lipids
 Application Note omics LC/MS Method for Comprehensive Analysis of Plasma s Authors Tomas Cajka and Oliver Fiehn West Coast Metabolomics Center, University of California Davis, 451 Health Sciences Drive,
Application Note omics LC/MS Method for Comprehensive Analysis of Plasma s Authors Tomas Cajka and Oliver Fiehn West Coast Metabolomics Center, University of California Davis, 451 Health Sciences Drive,
Mass-Spectrometric Analysis of Lipids (Lipidomics)
 Mass-Spectrometric Analysis of Lipids (Lipidomics) 1. Identification 2. Quantification 3. Metabolism Why to do lipidomics? Biology: Functions of different lipids? Medicine: Diagnostics and Therapy Industry:
Mass-Spectrometric Analysis of Lipids (Lipidomics) 1. Identification 2. Quantification 3. Metabolism Why to do lipidomics? Biology: Functions of different lipids? Medicine: Diagnostics and Therapy Industry:
The study of phospholipids in single cells using an integrated microfluidic device
 Supporting Information: The study of phospholipids in single cells using an integrated microfluidic device combined with matrix-assisted laser desorption/ionization mass spectrometry Weiyi Xie,, Dan Gao,
Supporting Information: The study of phospholipids in single cells using an integrated microfluidic device combined with matrix-assisted laser desorption/ionization mass spectrometry Weiyi Xie,, Dan Gao,
LOCALISATION, IDENTIFICATION AND SEPARATION OF MOLECULES. Gilles Frache Materials Characterization Day October 14 th 2016
 LOCALISATION, IDENTIFICATION AND SEPARATION OF MOLECULES Gilles Frache Materials Characterization Day October 14 th 2016 1 MOLECULAR ANALYSES Which focus? LOCALIZATION of molecules by Mass Spectrometry
LOCALISATION, IDENTIFICATION AND SEPARATION OF MOLECULES Gilles Frache Materials Characterization Day October 14 th 2016 1 MOLECULAR ANALYSES Which focus? LOCALIZATION of molecules by Mass Spectrometry
Lipids Analysis. Lipids
 Lipids Analysis Stephen Barnes 3 5 15 Lipids Lipids are mostly very hydrophobic Most are conjugates of fatty acids of a variety of chain lengths, which have different degrees of unsaturation, cis trans
Lipids Analysis Stephen Barnes 3 5 15 Lipids Lipids are mostly very hydrophobic Most are conjugates of fatty acids of a variety of chain lengths, which have different degrees of unsaturation, cis trans
Impact of Chromatography on Lipid Profiling of Liver Tissue Extracts
 Impact of Chromatography on Lipid Profiling of Liver Tissue Extracts Application Note Clinical Research Authors Mark Sartain and Theodore Sana Agilent Technologies, Inc. Santa Clara, California, USA Introduction
Impact of Chromatography on Lipid Profiling of Liver Tissue Extracts Application Note Clinical Research Authors Mark Sartain and Theodore Sana Agilent Technologies, Inc. Santa Clara, California, USA Introduction
Sequence Identification And Spatial Distribution of Rat Brain Tryptic Peptides Using MALDI Mass Spectrometric Imaging
 Sequence Identification And Spatial Distribution of Rat Brain Tryptic Peptides Using MALDI Mass Spectrometric Imaging AB SCIEX MALDI TOF/TOF* Systems Patrick Pribil AB SCIEX, Canada MALDI mass spectrometric
Sequence Identification And Spatial Distribution of Rat Brain Tryptic Peptides Using MALDI Mass Spectrometric Imaging AB SCIEX MALDI TOF/TOF* Systems Patrick Pribil AB SCIEX, Canada MALDI mass spectrometric
INTRODUCTION TO MALDI IMAGING
 INTRODUCTION TO MALDI IMAGING Marten F. Snel, Emmanuelle Claude, Thérèse McKenna, and James I. Langridge Waters Corporation, Manchester, UK INT RODUCTION The last few years have seen a rapid increase in
INTRODUCTION TO MALDI IMAGING Marten F. Snel, Emmanuelle Claude, Thérèse McKenna, and James I. Langridge Waters Corporation, Manchester, UK INT RODUCTION The last few years have seen a rapid increase in
[application note] DIRECT TISSUE IMAGING AND CHARACTERIZATION OF PHOSPHOLIPIDS USING A MALDI SYNAPT HDMS SYSTEM
![[application note] DIRECT TISSUE IMAGING AND CHARACTERIZATION OF PHOSPHOLIPIDS USING A MALDI SYNAPT HDMS SYSTEM [application note] DIRECT TISSUE IMAGING AND CHARACTERIZATION OF PHOSPHOLIPIDS USING A MALDI SYNAPT HDMS SYSTEM](/thumbs/93/111965378.jpg) DIRECT TISSUE IMAGING AND CHARACTERIZATION OF PHOSPHOLIPIDS USING A MALDI SYNAPT HDMS SYSTEM Emmanuelle Claude, Marten Snel, Thérèse McKenna, and James Langridge INTRODUCTION The last decade has seen a
DIRECT TISSUE IMAGING AND CHARACTERIZATION OF PHOSPHOLIPIDS USING A MALDI SYNAPT HDMS SYSTEM Emmanuelle Claude, Marten Snel, Thérèse McKenna, and James Langridge INTRODUCTION The last decade has seen a
Three Dimensional Mapping and Imaging of Neuropeptides and Lipids in Crustacean Brain
 Three Dimensional Mapping and Imaging of Neuropeptides and Lipids in Crustacean Brain Using the 4800 MALDI TOF/TOF Analyzer Ruibing Chen and Lingjun Li School of Pharmacy and Department of Chemistry, University
Three Dimensional Mapping and Imaging of Neuropeptides and Lipids in Crustacean Brain Using the 4800 MALDI TOF/TOF Analyzer Ruibing Chen and Lingjun Li School of Pharmacy and Department of Chemistry, University
EXPERIMENT 13: Isolation and Characterization of Erythrocyte
 EXPERIMENT 13: Isolation and Characterization of Erythrocyte Day 1: Isolation of Erythrocyte Steps 1 through 6 of the Switzer & Garrity protocol (pages 220-221) have been performed by the TA. We will be
EXPERIMENT 13: Isolation and Characterization of Erythrocyte Day 1: Isolation of Erythrocyte Steps 1 through 6 of the Switzer & Garrity protocol (pages 220-221) have been performed by the TA. We will be
Mass Spectrometry. Actual Instrumentation
 Mass Spectrometry Actual Instrumentation August 2017 See also http://www.uni-bielefeld.de/chemie/analytik/ms f additional infmation 1. MALDI TOF MASS SPECTROMETRY ON THE ULTRAFLEX 2 2. ESI MASS SPECTROMETRY
Mass Spectrometry Actual Instrumentation August 2017 See also http://www.uni-bielefeld.de/chemie/analytik/ms f additional infmation 1. MALDI TOF MASS SPECTROMETRY ON THE ULTRAFLEX 2 2. ESI MASS SPECTROMETRY
LC/MS/MS SOLUTIONS FOR LIPIDOMICS. Biomarker and Omics Solutions FOR DISCOVERY AND TARGETED LIPIDOMICS
 LC/MS/MS SOLUTIONS FOR LIPIDOMICS Biomarker and Omics Solutions FOR DISCOVERY AND TARGETED LIPIDOMICS Lipids play a key role in many biological processes, such as the formation of cell membranes and signaling
LC/MS/MS SOLUTIONS FOR LIPIDOMICS Biomarker and Omics Solutions FOR DISCOVERY AND TARGETED LIPIDOMICS Lipids play a key role in many biological processes, such as the formation of cell membranes and signaling
Essential Lipidomics Experiments using the LTQ Orbitrap Hybrid Mass Spectrometer
 Application Note: 367 Essential Lipidomics Experiments using the LTQ rbitrap Hybrid Mass Spectrometer Thomas Moehring 1, Michaela Scigelova 2, Christer S. Ejsing 3, Dominik Schwudke 3, Andrej Shevchenko
Application Note: 367 Essential Lipidomics Experiments using the LTQ rbitrap Hybrid Mass Spectrometer Thomas Moehring 1, Michaela Scigelova 2, Christer S. Ejsing 3, Dominik Schwudke 3, Andrej Shevchenko
Solving practical problems. Maria Kuhtinskaja
 Solving practical problems Maria Kuhtinskaja What does a mass spectrometer do? It measures mass better than any other technique. It can give information about chemical structures. What are mass measurements
Solving practical problems Maria Kuhtinskaja What does a mass spectrometer do? It measures mass better than any other technique. It can give information about chemical structures. What are mass measurements
Mass spectrometry Technologies in Lipid chemistry
 Mass spectrometry Technologies in Lipid chemistry Rabah Soliymani University Of Helsinki Protein Chemistry Unit Biomedicum Helsinki Rabah.soliymani@helsinki.fi Complex_&_dynamic_mixtures (few copies to
Mass spectrometry Technologies in Lipid chemistry Rabah Soliymani University Of Helsinki Protein Chemistry Unit Biomedicum Helsinki Rabah.soliymani@helsinki.fi Complex_&_dynamic_mixtures (few copies to
Analysis of Triglycerides in Cooking Oils Using MALDI-TOF Mass Spectrometry and Principal Component Analysis
 Analysis of Triglycerides in Cooking Oils Using MALDI-TOF Mass Spectrometry and Principal Component Analysis Kevin Cooley Chemistry Supervisor: Kingsley Donkor 1. Abstract Triglycerides are composed of
Analysis of Triglycerides in Cooking Oils Using MALDI-TOF Mass Spectrometry and Principal Component Analysis Kevin Cooley Chemistry Supervisor: Kingsley Donkor 1. Abstract Triglycerides are composed of
Application Note # ET-17 / MT-99 Characterization of the N-glycosylation Pattern of Antibodies by ESI - and MALDI mass spectrometry
 Bruker Daltonics Application Note # ET-17 / MT-99 Characterization of the N-glycosylation Pattern of Antibodies by ESI - and MALDI mass spectrometry Abstract Analysis of the N-glycosylation pattern on
Bruker Daltonics Application Note # ET-17 / MT-99 Characterization of the N-glycosylation Pattern of Antibodies by ESI - and MALDI mass spectrometry Abstract Analysis of the N-glycosylation pattern on
MALDI-TOF analysis of whole blood: its usefulness and potential in the assessment of HbA1c levels
 MALDI-TOF analysis of whole blood: its usefulness and potential in the assessment of HbA1c levels Jane Y. Yang, David A. Herold Department of Pathology, University of California San Diego, 9500 Gilman
MALDI-TOF analysis of whole blood: its usefulness and potential in the assessment of HbA1c levels Jane Y. Yang, David A. Herold Department of Pathology, University of California San Diego, 9500 Gilman
The use of mass spectrometry in lipidomics. Outlines
 The use of mass spectrometry in lipidomics Jeevan Prasain jprasain@uab.edu 6-2612 utlines Brief introduction to lipidomics Analytical methodology: MS/MS structure elucidation of phospholipids Phospholipid
The use of mass spectrometry in lipidomics Jeevan Prasain jprasain@uab.edu 6-2612 utlines Brief introduction to lipidomics Analytical methodology: MS/MS structure elucidation of phospholipids Phospholipid
CAMAG TLC-MS INTERFACE
 CAMAG TLC-MS INTERFACE 93.1 249.2 40 30 97.1 20 10 250.2 0 200 400 m/z WORLD LEADER IN PLANAR-CHROMATOGRAPHY Identification and elucidation of unknown substances by hyphenation of TLC / HPTLC and MS The
CAMAG TLC-MS INTERFACE 93.1 249.2 40 30 97.1 20 10 250.2 0 200 400 m/z WORLD LEADER IN PLANAR-CHROMATOGRAPHY Identification and elucidation of unknown substances by hyphenation of TLC / HPTLC and MS The
Matrix Assisted Laser Desorption Ionization Time-of-flight Mass Spectrometry
 Matrix Assisted Laser Desorption Ionization Time-of-flight Mass Spectrometry Time-of-Flight Mass Spectrometry. Basic principles An attractive feature of the time-of-flight (TOF) mass spectrometer is its
Matrix Assisted Laser Desorption Ionization Time-of-flight Mass Spectrometry Time-of-Flight Mass Spectrometry. Basic principles An attractive feature of the time-of-flight (TOF) mass spectrometer is its
An optical dosimeter for the selective detection of gaseous phosgene with ultra-low detection limit
 Supporting information for An optical dosimeter for the selective detection of gaseous phosgene with ultra-low detection limit Alejandro P. Vargas, Francisco Gámez*, Javier Roales, Tània Lopes-Costa and
Supporting information for An optical dosimeter for the selective detection of gaseous phosgene with ultra-low detection limit Alejandro P. Vargas, Francisco Gámez*, Javier Roales, Tània Lopes-Costa and
Mass Spectrometry based metabolomics
 Mass Spectrometry based metabolomics Metabolomics- A realm of small molecules (
Mass Spectrometry based metabolomics Metabolomics- A realm of small molecules (
Application Note # LCMS-89 High quantification efficiency in plasma targeted proteomics with a full-capability discovery Q-TOF platform
 Application Note # LCMS-89 High quantification efficiency in plasma targeted proteomics with a full-capability discovery Q-TOF platform Abstract Targeted proteomics for biomarker verification/validation
Application Note # LCMS-89 High quantification efficiency in plasma targeted proteomics with a full-capability discovery Q-TOF platform Abstract Targeted proteomics for biomarker verification/validation
Application of LC/Electrospray Ion Trap Mass Spectrometry for Identification and Quantification of Pesticides in Complex Matrices
 Application ote #LCMS-2 esquire series Application of LC/Electrospray Ion Trap Mass Spectrometry for Identification and Quantification of Pesticides in Complex Matrices Introduction The simple monitoring
Application ote #LCMS-2 esquire series Application of LC/Electrospray Ion Trap Mass Spectrometry for Identification and Quantification of Pesticides in Complex Matrices Introduction The simple monitoring
Supporting Information
 Supporting Information Wiley-VCH 2006 69451 Weinheim, Germany Tissue Imaging at Atmospheric Pressure using Desorption Electrospray Ionization (DESI) Mass Spectrometry Justin M. Wiseman, Demian R. Ifa,
Supporting Information Wiley-VCH 2006 69451 Weinheim, Germany Tissue Imaging at Atmospheric Pressure using Desorption Electrospray Ionization (DESI) Mass Spectrometry Justin M. Wiseman, Demian R. Ifa,
Data Independent MALDI Imaging HDMS E for Visualization and Identification of Lipids Directly from a Single Tissue Section
 Data Independent MALDI Imaging HDMS E for Visualization and Identification of Lipids Directly from a Single Tissue Section Emmanuelle Claude, Mark Towers, and Kieran Neeson Waters Corporation, Manchester,
Data Independent MALDI Imaging HDMS E for Visualization and Identification of Lipids Directly from a Single Tissue Section Emmanuelle Claude, Mark Towers, and Kieran Neeson Waters Corporation, Manchester,
Supplementary Information
 Supplementary Information Molecular imaging of brain localization of liposomes in mice using MALDI mass spectrometry Annabelle Fülöp 1,2, Denis A. Sammour 1,2, Katrin Erich 1,2, Johanna von Gerichten 4,
Supplementary Information Molecular imaging of brain localization of liposomes in mice using MALDI mass spectrometry Annabelle Fülöp 1,2, Denis A. Sammour 1,2, Katrin Erich 1,2, Johanna von Gerichten 4,
PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System
 Application Note LC/MS PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System Purpose This application note describes an automated workflow
Application Note LC/MS PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System Purpose This application note describes an automated workflow
Phospholipid characterization by a TQ-MS data based identification scheme
 P-CN1716E Phospholipid characterization by a TQ-MS data based identification scheme ASMS 2017 MP-406 Tsuyoshi Nakanishi 1, Masaki Yamada 1, Ningombam Sanjib Meitei 2, 3 1 Shimadzu Corporation, Kyoto, Japan,
P-CN1716E Phospholipid characterization by a TQ-MS data based identification scheme ASMS 2017 MP-406 Tsuyoshi Nakanishi 1, Masaki Yamada 1, Ningombam Sanjib Meitei 2, 3 1 Shimadzu Corporation, Kyoto, Japan,
Ionization Methods. Neutral species Charged species. Removal/addition of electron(s) Removal/addition of proton(s)
 Ionization Methods Neutral species Charged species Removal/addition of electron(s) M + e - (M +. )* + 2e - electron ionization Removal/addition of proton(s) M + (Matrix)-H MH + + (Matrix) - chemical ionization
Ionization Methods Neutral species Charged species Removal/addition of electron(s) M + e - (M +. )* + 2e - electron ionization Removal/addition of proton(s) M + (Matrix)-H MH + + (Matrix) - chemical ionization
Application Note # FTMS-46 solarix XR: Analysis of Complex Mixtures
 Application Note # FTMS-46 solarix XR: Analysis of Complex Mixtures Introduction Natural organic matter (NOM) as well as crude oil and crude oil fractions are very complex mixtures of organic compounds
Application Note # FTMS-46 solarix XR: Analysis of Complex Mixtures Introduction Natural organic matter (NOM) as well as crude oil and crude oil fractions are very complex mixtures of organic compounds
Mass-Based Purification of Natural Product Impurities Using an Agilent 1260 Infinity II Preparative LC/MSD System
 Application Note Food Testing and Agriculture Mass-Based Purification of Natural Product Impurities Using an Agilent 126 Infinity II Preparative LC/MSD System Authors Florian Rieck and Jörg Hippler Agilent
Application Note Food Testing and Agriculture Mass-Based Purification of Natural Product Impurities Using an Agilent 126 Infinity II Preparative LC/MSD System Authors Florian Rieck and Jörg Hippler Agilent
Rapid Lipid Profiling of Serum by Reverse Phase UPLC-Tandem Quadrupole MS
 Rapid Lipid Profiling of Serum by Reverse Phase UPLC-Tandem Quadrupole MS Mark Ritchie and Evelyn Goh Waters Pacific Pte Ltd., Singapore A P P L I C AT ION B E N E F I T S Delivers a rapid 10-min MRM method
Rapid Lipid Profiling of Serum by Reverse Phase UPLC-Tandem Quadrupole MS Mark Ritchie and Evelyn Goh Waters Pacific Pte Ltd., Singapore A P P L I C AT ION B E N E F I T S Delivers a rapid 10-min MRM method
Comprehensive Two-Dimensional HPLC and Informative Data Processing for Pharmaceuticals and Lipids
 PO-CON1576E Comprehensive Two-Dimensional HPLC and Informative Data Processing for Pharmaceuticals and Lipids HPLC 2015 PSB-MULTI-06 Yoshiyuki WATABE, Tetsuo IIDA, Daisuke NAKAYAMA, Kanya TSUJII, Saki
PO-CON1576E Comprehensive Two-Dimensional HPLC and Informative Data Processing for Pharmaceuticals and Lipids HPLC 2015 PSB-MULTI-06 Yoshiyuki WATABE, Tetsuo IIDA, Daisuke NAKAYAMA, Kanya TSUJII, Saki
Automated Sample Preparation/Concentration of Biological Samples Prior to Analysis via MALDI-TOF Mass Spectroscopy Application Note 222
 Automated Sample Preparation/Concentration of Biological Samples Prior to Analysis via MALDI-TOF Mass Spectroscopy Application Note 222 Joan Stevens, Ph.D.; Luke Roenneburg; Tim Hegeman; Kevin Fawcett
Automated Sample Preparation/Concentration of Biological Samples Prior to Analysis via MALDI-TOF Mass Spectroscopy Application Note 222 Joan Stevens, Ph.D.; Luke Roenneburg; Tim Hegeman; Kevin Fawcett
Imaging of lipid species by MALDI mass spectrometry
 Imaging of lipid species by MALDI mass spectrometry Robert C. Murphy, 1 Joseph A. Hankin, and Robert M. Barkley Department of Pharmacology, MS 8303 University of Colorado Denver, Aurora, CO 80045 Abstract
Imaging of lipid species by MALDI mass spectrometry Robert C. Murphy, 1 Joseph A. Hankin, and Robert M. Barkley Department of Pharmacology, MS 8303 University of Colorado Denver, Aurora, CO 80045 Abstract
Analysis of Peptides via Capillary HPLC and Fraction Collection Directly onto a MALDI Plate for Off-line Analysis by MALDI-TOF
 Analysis of Peptides via Capillary HPLC and Fraction Collection Directly onto a MALDI Plate for Off-line Analysis by MALDI-TOF Application Note 219 Joan Stevens, PhD; Luke Roenneburg; Kevin Fawcett (Gilson,
Analysis of Peptides via Capillary HPLC and Fraction Collection Directly onto a MALDI Plate for Off-line Analysis by MALDI-TOF Application Note 219 Joan Stevens, PhD; Luke Roenneburg; Kevin Fawcett (Gilson,
SelexION Technology for Lipid Analysis: Pushing the Boundaries of Lipidomics
 ANSWERS FOR SCIENCE. KNOWLEDGE FOR LIFE. SelexION Technology for Lipid Analysis: Pushing the Boundaries of Lipidomics Baljit Ubhi, Ph.D ASMS Baltimore, June 2014 Lipidomics Profiling Needs and Deliverables
ANSWERS FOR SCIENCE. KNOWLEDGE FOR LIFE. SelexION Technology for Lipid Analysis: Pushing the Boundaries of Lipidomics Baljit Ubhi, Ph.D ASMS Baltimore, June 2014 Lipidomics Profiling Needs and Deliverables
Lipid Class Separation Using UPC 2 /MS
 Michael D. Jones, 1,3 Giorgis Isaac, 1 Giuseppe Astarita, 1 Andrew Aubin, 1 John Shockcor, 1 Vladimir Shulaev, 2 Cristina Legido-Quigley, 3 and Norman Smith 3 1 Waters Corporation, Milford, MA, USA 2 Department
Michael D. Jones, 1,3 Giorgis Isaac, 1 Giuseppe Astarita, 1 Andrew Aubin, 1 John Shockcor, 1 Vladimir Shulaev, 2 Cristina Legido-Quigley, 3 and Norman Smith 3 1 Waters Corporation, Milford, MA, USA 2 Department
Increased Identification Coverage and Throughput for Complex Lipidomes
 Increased Identification Coverage and Throughput for Complex Lipidomes Reiko Kiyonami, David Peake, Yingying Huang, Thermo Fisher Scientific, San Jose, CA USA Application Note 607 Key Words Q Exactive
Increased Identification Coverage and Throughput for Complex Lipidomes Reiko Kiyonami, David Peake, Yingying Huang, Thermo Fisher Scientific, San Jose, CA USA Application Note 607 Key Words Q Exactive
Desorption Electrospray Ionization Coupled with Ultraviolet Photodissociation for Characterization of Phospholipid Isomers in Tissue Sections
 Desorption Electrospray Ionization Coupled with Ultraviolet Photodissociation for Characterization of Phospholipid Isomers in Tissue Sections Dustin R. Klein, Clara L. Feider, Kyana Y. Garza, John Q. Lin,
Desorption Electrospray Ionization Coupled with Ultraviolet Photodissociation for Characterization of Phospholipid Isomers in Tissue Sections Dustin R. Klein, Clara L. Feider, Kyana Y. Garza, John Q. Lin,
Supporting Information
 Supporting Information Development of a High Coverage Pseudotargeted Lipidomics Method Based on Ultra-High Performance Liquid Chromatography-Mass Spectrometry Qiuhui Xuan 1,2#, Chunxiu Hu 1#, Di Yu 1,2,
Supporting Information Development of a High Coverage Pseudotargeted Lipidomics Method Based on Ultra-High Performance Liquid Chromatography-Mass Spectrometry Qiuhui Xuan 1,2#, Chunxiu Hu 1#, Di Yu 1,2,
Metabolomic fingerprinting of serum samples by direct infusion mass spectrometry
 Metabolomic fingerprinting of serum samples by direct infusion mass spectrometry Raúl González-Domínguez * Department of Chemistry, Faculty of Experimental Sciences. University of Huelva, Spain. * Corresponding
Metabolomic fingerprinting of serum samples by direct infusion mass spectrometry Raúl González-Domínguez * Department of Chemistry, Faculty of Experimental Sciences. University of Huelva, Spain. * Corresponding
MALDI-TOF. Introduction. Schematic and Theory of MALDI
 MALDI-TOF Proteins and peptides have been characterized by high pressure liquid chromatography (HPLC) or SDS PAGE by generating peptide maps. These peptide maps have been used as fingerprints of protein
MALDI-TOF Proteins and peptides have been characterized by high pressure liquid chromatography (HPLC) or SDS PAGE by generating peptide maps. These peptide maps have been used as fingerprints of protein
1. Sample Introduction to MS Systems:
 MS Overview: 9.10.08 1. Sample Introduction to MS Systems:...2 1.1. Chromatography Interfaces:...3 1.2. Electron impact: Used mainly in Protein MS hard ionization source...4 1.3. Electrospray Ioniztion:
MS Overview: 9.10.08 1. Sample Introduction to MS Systems:...2 1.1. Chromatography Interfaces:...3 1.2. Electron impact: Used mainly in Protein MS hard ionization source...4 1.3. Electrospray Ioniztion:
Received: 21 September 2009 / Revised: 2 November 2009 / Accepted: 3 November 2009 / Published online: 25 November 2009 # Springer-Verlag 2009
 Anal Bioanal Chem (2010) 396:1273 1280 DOI 10.1007/s00216-009-3292-9 ORIGINAL PAPER Quantitative analysis of urinary phospholipids found in patients with breast cancer by nanoflow liquid chromatography
Anal Bioanal Chem (2010) 396:1273 1280 DOI 10.1007/s00216-009-3292-9 ORIGINAL PAPER Quantitative analysis of urinary phospholipids found in patients with breast cancer by nanoflow liquid chromatography
2D-LC as an Automated Desalting Tool for MSD Analysis
 2D-LC as an Automated Desalting Tool for MSD Analysis Direct Mass Selective Detection of a Pharmaceutical Peptide from an MS-Incompatible USP Method Application Note Biologics and Biosimilars Author Sonja
2D-LC as an Automated Desalting Tool for MSD Analysis Direct Mass Selective Detection of a Pharmaceutical Peptide from an MS-Incompatible USP Method Application Note Biologics and Biosimilars Author Sonja
What You Can t See Can Hurt You. How MS/MS Specificity Can Bite Your Backside
 What You Can t See Can Hurt You How MS/MS Specificity Can Bite Your Backside Johan van den Heever, Tom Thompson, and Don Noot Agri-Food Laboratories Branch Advances in Trace rganic Residue Analysis early
What You Can t See Can Hurt You How MS/MS Specificity Can Bite Your Backside Johan van den Heever, Tom Thompson, and Don Noot Agri-Food Laboratories Branch Advances in Trace rganic Residue Analysis early
Overview on the identification of different classes of. lipids by HPTLC (High Performance Thin Layer. Chromatography) and ITLC (Immuno Thin Layer
 Overview on the identification of different classes of lipids by HPTLC (High Performance Thin Layer Chromatography) and ITLC (Immuno Thin Layer Chromatography) Iuliana Popa 1, Marie-Jeanne David 2, Daniel
Overview on the identification of different classes of lipids by HPTLC (High Performance Thin Layer Chromatography) and ITLC (Immuno Thin Layer Chromatography) Iuliana Popa 1, Marie-Jeanne David 2, Daniel
Applications for MALDI-TOF Imaging Present and Future
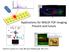 Y Position 8563 / (3000-30000) 12124 / (3000-30000) 14044 / (3000-30000) 10158 35 / (3000-30000) 15012 / (3000-30000) 10894 / (3000-30000) 4964 / (3000-30000) 12414 / (3000-30000) Area Chromatogram - All
Y Position 8563 / (3000-30000) 12124 / (3000-30000) 14044 / (3000-30000) 10158 35 / (3000-30000) 15012 / (3000-30000) 10894 / (3000-30000) 4964 / (3000-30000) 12414 / (3000-30000) Area Chromatogram - All
An Introduction and Overview on Comprehensive Two-Dimensional Gas Chromatography (GCxGC): New Opportunities for Unresolved Complex Mixtures
 An Introduction and Overview on Comprehensive Two-Dimensional Gas Chromatography (GCxGC): New Opportunities for Unresolved Complex Mixtures Daniela Cavagnino GC Product Manager Thermo Fisher Scientific,
An Introduction and Overview on Comprehensive Two-Dimensional Gas Chromatography (GCxGC): New Opportunities for Unresolved Complex Mixtures Daniela Cavagnino GC Product Manager Thermo Fisher Scientific,
A Simple and Accurate Method for the Rapid Quantitation of Drugs of Abuse in Urine Using Liquid Chromatography
 Application Note LCMS-109 A Simple and Accurate Method for the Rapid Quantitation of Drugs of Abuse in Urine Using Liquid Chromatography Time of Flight (LC-TOF) Mass Spectrometry Introduction Many clinical
Application Note LCMS-109 A Simple and Accurate Method for the Rapid Quantitation of Drugs of Abuse in Urine Using Liquid Chromatography Time of Flight (LC-TOF) Mass Spectrometry Introduction Many clinical
Biological Mass Spectrometry. April 30, 2014
 Biological Mass Spectrometry April 30, 2014 Mass Spectrometry Has become the method of choice for precise protein and nucleic acid mass determination in a very wide mass range peptide and nucleotide sequencing
Biological Mass Spectrometry April 30, 2014 Mass Spectrometry Has become the method of choice for precise protein and nucleic acid mass determination in a very wide mass range peptide and nucleotide sequencing
Enrichment of Phospholipids from Biological Matrices with Zirconium Oxide-Modified Silica Sorbents
 Enrichment of Phospholipids from Biological Matrices with Zirconium Oxide-Modified Silica Sorbents Xiaoning Lu, Jennifer E. Claus, and David S. Bell Supelco, Div. of Sigma-Aldrich Bellefonte, PA 16823
Enrichment of Phospholipids from Biological Matrices with Zirconium Oxide-Modified Silica Sorbents Xiaoning Lu, Jennifer E. Claus, and David S. Bell Supelco, Div. of Sigma-Aldrich Bellefonte, PA 16823
Supporting information
 Supporting information A novel lipidomics workflow for improved human plasma identification and quantification using RPLC-MSn methods and isotope dilution strategies Evelyn Rampler 1,2,3, Angela Criscuolo
Supporting information A novel lipidomics workflow for improved human plasma identification and quantification using RPLC-MSn methods and isotope dilution strategies Evelyn Rampler 1,2,3, Angela Criscuolo
Metabolomics: quantifying the phenotype
 Metabolomics: quantifying the phenotype Metabolomics Promises Quantitative Phenotyping What can happen GENOME What appears to be happening Bioinformatics TRANSCRIPTOME What makes it happen PROTEOME Systems
Metabolomics: quantifying the phenotype Metabolomics Promises Quantitative Phenotyping What can happen GENOME What appears to be happening Bioinformatics TRANSCRIPTOME What makes it happen PROTEOME Systems
Supporting information
 Supporting information Figure legends Supplementary Table 1. Specific product ions obtained from fragmentation of lithium adducts in the positive ion mode comparing the different positional isomers of
Supporting information Figure legends Supplementary Table 1. Specific product ions obtained from fragmentation of lithium adducts in the positive ion mode comparing the different positional isomers of
Supplementary Information. Effects of Perfluorooctanoic Acid on Metabolic Profiles in Brain and Liver of Mouse by a
 Supplementary Information Effects of Perfluorooctanoic Acid on Metabolic Profiles in Brain and Liver of Mouse by a High-throughput Targeted Metabolomics Approach Nanyang Yu, Si Wei, *, Meiying Li, Jingping
Supplementary Information Effects of Perfluorooctanoic Acid on Metabolic Profiles in Brain and Liver of Mouse by a High-throughput Targeted Metabolomics Approach Nanyang Yu, Si Wei, *, Meiying Li, Jingping
FPO. Label-Free Molecular Imaging. Innovation with Integrity. Discover, localize and quantify biochemical changes and molecular markers
 FPO Label-Free Molecular Imaging Discover, localize and quantify biochemical changes and molecular markers Innovation with Integrity Mass Spectrometry Bruker, the global leader in MALDI-MS technology,
FPO Label-Free Molecular Imaging Discover, localize and quantify biochemical changes and molecular markers Innovation with Integrity Mass Spectrometry Bruker, the global leader in MALDI-MS technology,
SCS Mass Spectrometry Laboratory
 SCS Mass Spectrometry Laboratory Contact Information Staff 31 Noyes Laboratory (8:00-5:00 M-F) 217-333-2545 http://scs.illinois.edu/massspec/ Furong Sun (frs@illinois.edu) Furong Sun Director Training
SCS Mass Spectrometry Laboratory Contact Information Staff 31 Noyes Laboratory (8:00-5:00 M-F) 217-333-2545 http://scs.illinois.edu/massspec/ Furong Sun (frs@illinois.edu) Furong Sun Director Training
LC-MS/MS Method for the Determination of Tenofovir from Plasma
 LC-MS/MS Method for the Determination of Tenofovir from Plasma Kimberly Phipps, Thermo Fisher Scientific, Runcorn, Cheshire, UK Application Note 687 Key Words SPE, SOLA CX, Hypersil GOLD, tenofovir Abstract
LC-MS/MS Method for the Determination of Tenofovir from Plasma Kimberly Phipps, Thermo Fisher Scientific, Runcorn, Cheshire, UK Application Note 687 Key Words SPE, SOLA CX, Hypersil GOLD, tenofovir Abstract
Applying a Novel Glycan Tagging Reagent, RapiFluor-MS, and an Integrated UPLC-FLR/QTof MS System for Low Abundant N-Glycan Analysis
 Applying a Novel Glycan Tagging Reagent, RapiFluor-MS, and an Integrated UPLC-FLR/QTof MS System for Low Abundant N-Glycan Analysis Ying Qing Yu Waters Corporation, Milford, MA, USA APPLICATION BENEFITS
Applying a Novel Glycan Tagging Reagent, RapiFluor-MS, and an Integrated UPLC-FLR/QTof MS System for Low Abundant N-Glycan Analysis Ying Qing Yu Waters Corporation, Milford, MA, USA APPLICATION BENEFITS
Principles of Shotgun Lipidomics
 Principles of Shotgun Lipidomics Xianlin Han Diabetes and Obesity Research Center Sanford-Burnham Medical Research Institute Lake Nona Orlando, FL 32827 What is shotgun lipidomics? Original definition
Principles of Shotgun Lipidomics Xianlin Han Diabetes and Obesity Research Center Sanford-Burnham Medical Research Institute Lake Nona Orlando, FL 32827 What is shotgun lipidomics? Original definition
Agilent 6410 Triple Quadrupole LC/MS. Sensitivity, Reliability, Value
 Agilent 64 Triple Quadrupole LC/MS Sensitivity, Reliability, Value Sensitivity, Reliability, Value Whether you quantitate drug metabolites, measure herbicide levels in food, or determine contaminant levels
Agilent 64 Triple Quadrupole LC/MS Sensitivity, Reliability, Value Sensitivity, Reliability, Value Whether you quantitate drug metabolites, measure herbicide levels in food, or determine contaminant levels
MALDI Imaging Mass Spectrometry
 MALDI Imaging Mass Spectrometry Nan Kleinholz Mass Spectrometry and Proteomics Facility The Ohio State University Mass Spectrometry and Proteomics Workshop What is MALDI Imaging? MALDI: Matrix Assisted
MALDI Imaging Mass Spectrometry Nan Kleinholz Mass Spectrometry and Proteomics Facility The Ohio State University Mass Spectrometry and Proteomics Workshop What is MALDI Imaging? MALDI: Matrix Assisted
MALDI Mass Spectrometry: A Label-Free Solution for Ultra-High-Throughput Screening
 MALDI Mass Spectrometry: A Label-Free Solution for Ultra-High-Throughput Screening MIPTEC Conference 2016 Meike Hamester, Jens Fuchser Bruker Daltonik, Bremen 1 MALDI PharmaPulse for: Biochemical Screening
MALDI Mass Spectrometry: A Label-Free Solution for Ultra-High-Throughput Screening MIPTEC Conference 2016 Meike Hamester, Jens Fuchser Bruker Daltonik, Bremen 1 MALDI PharmaPulse for: Biochemical Screening
Mass Spectrometry at the Laboratory of Food Chemistry. Edwin Bakx Laboratory of Food Chemistry Wageningen University
 Mass Spectrometry at the Wageningen University Mass Spectrometry at the 3 UPLC/CE - ESI - Ion trap MS systems UPLC Thermo Acella with a Velos or VelosPro CE Beckman PA800 with a Thermo VelosPro 1 UPLC-
Mass Spectrometry at the Wageningen University Mass Spectrometry at the 3 UPLC/CE - ESI - Ion trap MS systems UPLC Thermo Acella with a Velos or VelosPro CE Beckman PA800 with a Thermo VelosPro 1 UPLC-
A pillar[2]arene[3]hydroquinone which can self-assemble to a molecular zipper in the solid state
![A pillar[2]arene[3]hydroquinone which can self-assemble to a molecular zipper in the solid state A pillar[2]arene[3]hydroquinone which can self-assemble to a molecular zipper in the solid state](/thumbs/86/93673402.jpg) A pillar[2]arene[3]hydroquinone which can self-assemble to a molecular zipper in the solid state Mingguang Pan, Min Xue* Department of Chemistry, Zhejiang University, Hangzhou 310027, P. R. China Fax:
A pillar[2]arene[3]hydroquinone which can self-assemble to a molecular zipper in the solid state Mingguang Pan, Min Xue* Department of Chemistry, Zhejiang University, Hangzhou 310027, P. R. China Fax:
Chromatography Vacuum Ultraviolet Spectroscopy
 Application Note Differentiation and Determination Differentiation and Determination of Fatty Acid Methyl of Fatty Esters Acid by Gas Methyl Chromatography Esters by Vacuum Gas Ultraviolet Spectroscopy
Application Note Differentiation and Determination Differentiation and Determination of Fatty Acid Methyl of Fatty Esters Acid by Gas Methyl Chromatography Esters by Vacuum Gas Ultraviolet Spectroscopy
[ APPLICATION NOTE ] High Sensitivity Intact Monoclonal Antibody (mab) HRMS Quantification APPLICATION BENEFITS INTRODUCTION WATERS SOLUTIONS KEYWORDS
![[ APPLICATION NOTE ] High Sensitivity Intact Monoclonal Antibody (mab) HRMS Quantification APPLICATION BENEFITS INTRODUCTION WATERS SOLUTIONS KEYWORDS [ APPLICATION NOTE ] High Sensitivity Intact Monoclonal Antibody (mab) HRMS Quantification APPLICATION BENEFITS INTRODUCTION WATERS SOLUTIONS KEYWORDS](/thumbs/79/80328542.jpg) Yun Wang Alelyunas, Henry Shion, Mark Wrona Waters Corporation, Milford, MA, USA APPLICATION BENEFITS mab LC-MS method which enables users to achieve highly sensitive bioanalysis of intact trastuzumab
Yun Wang Alelyunas, Henry Shion, Mark Wrona Waters Corporation, Milford, MA, USA APPLICATION BENEFITS mab LC-MS method which enables users to achieve highly sensitive bioanalysis of intact trastuzumab
Lipid Analysis. Andréina Laffargue, IRD CRYMCEPT Montpellier workshop, October 17th Introduction to lipid structures
 Lipid Analysis Andréina Laffargue, IRD CRYMCEPT Montpellier workshop, October 17th 2005 Introduction to lipid structures Fatty acids Acylglycerols Glycerophospholipids Sterols Strategies involved in lipid
Lipid Analysis Andréina Laffargue, IRD CRYMCEPT Montpellier workshop, October 17th 2005 Introduction to lipid structures Fatty acids Acylglycerols Glycerophospholipids Sterols Strategies involved in lipid
Components of a Mass Spectrometer
 Components of a Mass Spectrometer Sample Introduction Inlet GC LC Direct Insertion (Syringe/Probe) Ionization Ion Separation Ion Detection Ion Source EI,CI,,, MALDI Mass Analyzer Under vacuum TOF, Quadrupole,
Components of a Mass Spectrometer Sample Introduction Inlet GC LC Direct Insertion (Syringe/Probe) Ionization Ion Separation Ion Detection Ion Source EI,CI,,, MALDI Mass Analyzer Under vacuum TOF, Quadrupole,
Fundamentals of Soft Ionization and MS Instrumentation
 Fundamentals of Soft Ionization and MS Instrumentation Ana Varela Coelho varela@itqb.unl.pt Mass Spectrometry Lab Analytical Services Unit Index Mass spectrometers and its components Ionization methods:
Fundamentals of Soft Ionization and MS Instrumentation Ana Varela Coelho varela@itqb.unl.pt Mass Spectrometry Lab Analytical Services Unit Index Mass spectrometers and its components Ionization methods:
Molecular Cartography:
 Molecular Cartography: Moving Towards Combined Topographical and Chemical Imaging Using AFM and Mass Spectrometry Olga S. Ovchinnikova Organic and Biological Mass Spectrometry Group, Chemical Sciences
Molecular Cartography: Moving Towards Combined Topographical and Chemical Imaging Using AFM and Mass Spectrometry Olga S. Ovchinnikova Organic and Biological Mass Spectrometry Group, Chemical Sciences
SUPPLEMENTARY DATA. Materials and Methods
 SUPPLEMENTARY DATA Materials and Methods HPLC-UV of phospholipid classes and HETE isomer determination. Fractionation of platelet lipid classes was undertaken on a Spherisorb S5W 150 x 4.6 mm column (Waters
SUPPLEMENTARY DATA Materials and Methods HPLC-UV of phospholipid classes and HETE isomer determination. Fractionation of platelet lipid classes was undertaken on a Spherisorb S5W 150 x 4.6 mm column (Waters
Clinical Microbiology
 Clinical Microbiology MALDI Biotyper Fast & Accurate Identification of Microorganisms Innovation with Integrity MALDI-TOF In Microbiology, Every Minute Counts A Powerful Technology for Better Results To
Clinical Microbiology MALDI Biotyper Fast & Accurate Identification of Microorganisms Innovation with Integrity MALDI-TOF In Microbiology, Every Minute Counts A Powerful Technology for Better Results To
Methods in Mass Spectrometry. Dr. Noam Tal Laboratory of Mass Spectrometry School of Chemistry, Tel Aviv University
 Methods in Mass Spectrometry Dr. Noam Tal Laboratory of Mass Spectrometry School of Chemistry, Tel Aviv University Sample Engineering Chemistry Biology Life Science Medicine Industry IDF / Police Sample
Methods in Mass Spectrometry Dr. Noam Tal Laboratory of Mass Spectrometry School of Chemistry, Tel Aviv University Sample Engineering Chemistry Biology Life Science Medicine Industry IDF / Police Sample
Lipid Analysis ISOLATION, SEPARATION, IDENTIFICATION AND. Bridgwater, England LIPIDOMIC ANALYSIS. Fourth Edition. Invergowrie, Dundee, Scotland
 Lipid Analysis ISOLATION, SEPARATION, IDENTIFICATION AND LIPIDOMIC ANALYSIS Fourth Edition WILLIAM W.CHRISTIE MRS Lipid Analysis Unit, Scottish Crop Research Institute, Dundee, Scotland Invergowrie, and
Lipid Analysis ISOLATION, SEPARATION, IDENTIFICATION AND LIPIDOMIC ANALYSIS Fourth Edition WILLIAM W.CHRISTIE MRS Lipid Analysis Unit, Scottish Crop Research Institute, Dundee, Scotland Invergowrie, and
The HPLC Preparative Scale-Up of Soybean Phospholipids Application
 The HPLC Preparative Scale-Up of Soybean Phospholipids Application Food and Flavors Authors Cliff Woodward and Ron Majors Agilent Technologies, Inc. 285 Centerville Road Wilmington, DE 1988-161 USA Abstract
The HPLC Preparative Scale-Up of Soybean Phospholipids Application Food and Flavors Authors Cliff Woodward and Ron Majors Agilent Technologies, Inc. 285 Centerville Road Wilmington, DE 1988-161 USA Abstract
Organic Chemistry Laboratory Fall Lecture 3 Gas Chromatography and Mass Spectrometry June
 344 Organic Chemistry Laboratory Fall 2013 Lecture 3 Gas Chromatography and Mass Spectrometry June 19 2013 Chromatography Chromatography separation of a mixture into individual components Paper, Column,
344 Organic Chemistry Laboratory Fall 2013 Lecture 3 Gas Chromatography and Mass Spectrometry June 19 2013 Chromatography Chromatography separation of a mixture into individual components Paper, Column,
(III) MALDI instrumentation
 Dr. Sanjeeva Srivastava (I) Basics of MALDI-TF (II) Sample preparation In-gel digestion Zip-tip sample clean-up Matrix and sample plating (III) MALDI instrumentation 2 1 (I) Basics of MALDI-TF Analyte
Dr. Sanjeeva Srivastava (I) Basics of MALDI-TF (II) Sample preparation In-gel digestion Zip-tip sample clean-up Matrix and sample plating (III) MALDI instrumentation 2 1 (I) Basics of MALDI-TF Analyte
PesticideScreener. Innovation with Integrity
 PesticideScreener A complete multi-residue solution for pesticide screening using high resolution full scan accurate mass data Innovation with Integrity LC-MS The Challenge of Ever Increasing Number of
PesticideScreener A complete multi-residue solution for pesticide screening using high resolution full scan accurate mass data Innovation with Integrity LC-MS The Challenge of Ever Increasing Number of
SUPPORTING INFORMATION. Multimodal Mass Spectrometry Imaging of N-glycans and Proteins from the Same
 SUPPORTING INFORMATION Multimodal Mass Spectrometry Imaging of N-glycans and Proteins from the Same Tissue Section. Bram Heijs 1, Stephanie Holst 1, Inge H. Briaire-de Bruijn 2, Gabi W. van Pelt 3, Arnoud
SUPPORTING INFORMATION Multimodal Mass Spectrometry Imaging of N-glycans and Proteins from the Same Tissue Section. Bram Heijs 1, Stephanie Holst 1, Inge H. Briaire-de Bruijn 2, Gabi W. van Pelt 3, Arnoud
Measuring Lipid Composition LC-MS/MS
 Project: Measuring Lipid Composition LC-MS/MS Verification of expected lipid composition in nanomedical controlled release systems by liquid chromatography tandem mass spectrometry AUTHORED BY: DATE: Sven
Project: Measuring Lipid Composition LC-MS/MS Verification of expected lipid composition in nanomedical controlled release systems by liquid chromatography tandem mass spectrometry AUTHORED BY: DATE: Sven
MS-IMS (MALDI-IMAGING)?
 MS-IMS (MALDI-IMAGING)? Protein Chemistry/Proteomics and Peptide Synthesis and Array Unit Biomedicum Helsinki and Haartman Institute E-Mail: marc.baumann@helsinki.fi (http://research.med.helsinki.fi/corefacilities/proteinchem)
MS-IMS (MALDI-IMAGING)? Protein Chemistry/Proteomics and Peptide Synthesis and Array Unit Biomedicum Helsinki and Haartman Institute E-Mail: marc.baumann@helsinki.fi (http://research.med.helsinki.fi/corefacilities/proteinchem)
Impurity Identification using a Quadrupole - Time of Flight Mass Spectrometer QTOF
 Impurity Identification using a Quadrupole - Time of Flight Mass Spectrometer QTOF PUSHER TOF DETECTOR ZSPRAY TM Ion Source SAMPLING CONE SKIMMER RF HEXAPOLE RF HEXAPOLE QUADRUPOLE IN NARROW BANDPASS MODE
Impurity Identification using a Quadrupole - Time of Flight Mass Spectrometer QTOF PUSHER TOF DETECTOR ZSPRAY TM Ion Source SAMPLING CONE SKIMMER RF HEXAPOLE RF HEXAPOLE QUADRUPOLE IN NARROW BANDPASS MODE
MALDI Sepsityper Kit
 Instructions for Use MALDI Sepsityper Kit Kit for identification of microorganisms from positive blood cultures using the MALDI Biotyper system CARE products are designed to support our worldwide customers
Instructions for Use MALDI Sepsityper Kit Kit for identification of microorganisms from positive blood cultures using the MALDI Biotyper system CARE products are designed to support our worldwide customers
Rapid Analysis of Water-Soluble Vitamins in Infant Formula by Standard-Addition
 Rapid Analysis of Water-Soluble Vitamins in Infant Formula by Standard-Addition Evelyn Goh Waters Pacific, Singapore APPLICATION BENEFITS This method allows for the simultaneous analysis of 12 water-soluble
Rapid Analysis of Water-Soluble Vitamins in Infant Formula by Standard-Addition Evelyn Goh Waters Pacific, Singapore APPLICATION BENEFITS This method allows for the simultaneous analysis of 12 water-soluble
Lipid Imaging Mass Spectrometry.
 Lipid Imaging Mass Spectrometry. NH 2 P H H H H P H H P H N N Advanced Graduate Course in Metabolomics 3-11-2015 Janusz Kabarowski, Dept. of Microbiology, UAB. Matrix-Assisted Laser Desorption/Ionization
Lipid Imaging Mass Spectrometry. NH 2 P H H H H P H H P H N N Advanced Graduate Course in Metabolomics 3-11-2015 Janusz Kabarowski, Dept. of Microbiology, UAB. Matrix-Assisted Laser Desorption/Ionization
