Supporting Online Material for
|
|
|
- Meghan Jacobs
- 6 years ago
- Views:
Transcription
1 A Partial Retraction was published on 22 September See last page. Supporting Online Material for Detection of an Infectious Retrovirus, XMRV, in Blood Cells of Patients with Chronic Fatigue Syndrome Vincent C. Lombardi, Francis W. Ruscetti, Jaydip Das Gupta, Max A. Pfost, Kathryn S. Hagen, Daniel L. Peterson, Sandra K. Ruscetti, Rachel K. Bagni, Cari Petrow-Sadowski, Bert Gold, Michael Dean, Robert H. Silverman, Judy A. Mikovits* This PDF file includes: *To whom correspondence should be addressed. Materials and Methods Figs. S1 to S6 Tables S1 and S2 References Published 8 October 2009 on Science Express DOI: /science
2 1 Detection of an Infectious Retrovirus, XMRV, in Blood Cells of Patients with Chronic Fatigue Syndrome Vincent C Lombardi, Francis W Ruscetti, Jaydip Das Gupta, Max A Pfost, Kathryn S Hagen, Daniel L Peterson, Sandra K Ruscetti, Rachel K Bagni, Cari Petrow-Sadowski, Bert Gold, Michael Dean, Robert H Silverman, Judy A Mikovits* Supporting Online Material * judym@wpinstitute.org, Telephone: , Fax: This PDF contains: Materials and Methods Figs S1 to S6 Table S1 and S2 References
3 2 Supporting Online Materials and Methods Patient samples. Banked samples were selected for this study from patients fulfilling the 1994 CDC Fukuda Criteria for Chronic Fatigue Syndrome (S1) and the 2003 Canadian Consensus Criteria for Chronic Fatigue Syndrome/myalgic encephalomyelitis (CFS/ME) and presenting with severe disability. Samples were selected from several regions of the United States where outbreaks of CFS had been documented (S2). These are patients that have been seen in private medical practices, and their diagnosis of CFS is based upon prolonged disabling fatigue and the presence of cognitive deficits and reproducible immunological abnormalities. These included but were not limited to perturbations of the 2-5A synthetase/rnase L antiviral pathway, low natural killer cell cytotoxicity (as measured by standard diagnostic assays), and elevated cytokines particularly interleukin-6 and interleukin-8. In addition to these immunological abnormalities, the patients characteristically demonstrated impaired exercise performance with extremely low VO2 max measured on stress testing. The patients had been seen over a prolonged period of time and multiple longitudinal observations of the clinical and laboratory abnormalities had been documented. DNA and RNA isolation. Whole blood was drawn from subjects by venipuncture using standardized phlebotomy procedures into 8-mL greencapped Vacutainers containing the anti-coagulant sodium heparin (Becton Dickinson, Franklin Lakes, NJ). Plasma was collected by centrifugation, aspirated and stored at -80 ºC for later use. The plasma was replaced with PBS and the blood resuspended and further diluted with an equal volume of PBS. PBMC were isolated by layering the diluted blood onto Ficoll-Paque PLUS (GE Healthcare, Waukesha, WI), centrifuging for 22 min at 800 g, aspirating the PBMC layer and washing it once in PBS. The PBMC (approximately 2 x 10 7 cells) were centrifuged at 500x g for 7 min and either stored as unactivated cells in 90% FBS and 10% DMSO at -80 ºC for further culture and analysis or resuspended in TRIzol (Invitrogen, Carlsbad, CA) and stored at -80 ºC for DNA and RNA extraction and analysis. DNA was isolated from TRIzol preps according the to manufacturer s protocol and also isolated from frozen PBMC pellets using the QIAamp DNA Mini purification kit (QIAGEN, Valencia, CA) according to the manufacturer s protocol, and the final DNA was resuspended in RNase/DNasefree water and quantified using the Quant-iT Pico Green dsdna Kit (Invitrogen, Carlsbad, CA). RNA was isolated from TRIzol preps according to the manufacturer s protocol and quantified using the Quant-iT Ribo Green RNA kit (Invitrogen, Carlsbad, CA). cdna was made from RNA using the iscript Select cdna synthesis kit (Bio-Rad, Hercules, CA) according to the manufacturer s protocol. PCR. To avoid potential problems with laboratory DNA contamination, nested PCR was performed with separate reagents in a separate laboratory room designated to be free of high copy amplicon or plasmid DNA. Negative controls in the absence of added DNA were included in every experiment. Identification
4 of XMRV gag and env genes was performed by PCR in separate reactions. Reactions were performed as follows: 100 to 250 ng DNA, 2 µl of 25 mm MgCl 2, 25 µl of HotStart-IT FideliTaq Master Mix (USB Corporation, Cleveland, OH), 0.75 µl of each of 20 µm forward and reverse oligonucleotide primers in reaction volumes of 50 µl. For identification of gag, 419F (5 - ATCAGTTAACCTACCCGAGTCGGAC-3 ) and 1154R (5 - GCCGCCTCTTCTTCATTGTTCTC-3 ) were used as forward and reverse primers. For env, 5922F (5 - GCTAATGCTACCTCCCTCCTGG-3 ) and 6273R (5 -GGAGCCCACTGAGGAATCAAAACAGG-3 ) were used. For both gag and env PCR, 94 C for 4 min initial denaturation was performed for every reaction followed by 94 C for 30 seconds, 57 C for 30 seconds and 72 C for 1 minute. The cycle was repeated 45 times followed by final extension at 72 C for 2 minutes. Six microliters of each reaction product was loaded onto 2% agarose gels in TBE buffer with 1 kb+ DNA ladder (Invitrogen, Carlsbad, CA) as markers. PCR products were purified using Wizard SV Gel and PCR Clean-Up kit (Promega, Madison, WI) and sequenced. PCR amplification for sequencing full-length XMRV genomes was performed on DNA amplified by nested or semi-nested PCR from overlapping regions from PBMC DNA. For 5 end amplification of R-U5 region, 4F (5 - CCAGTCATCCGATAGACTGAGTCGC-3 ) and 1154R was used for first round and 4F and 770R (5 -TACCATCCTGAGGCCATCCTACATTG-3 ) was used for second round. For regions including gag-pro and partial pol, 350F(5 - GAGTTCGTATTCCCGGCCGCAGC-3 ) and 5135R (5 - CCTGCGGCATTCCAAATCTCG-3 ) was used for first round followed by second round with 419F and 4789R (5 -GGGTGAGTCTGTGTAGGGAGTCTAA-3 ). For regions including partial pol and env region, 4166F (5 - CAAGAAGGACAACGGAGAGCTGGAG-3 ) and 7622R (5 - GGCCTGCACTACCGAAAT TCTGTC-3 ) were used for first round followed by 4672F (5 GAGCCACCTACAATCAGACAAAAGGAT-3 ) and 7590R (5 - CTGGACCAAGCGGTTGAGAATACAG-3 ) for second round. For the 3 end including the U3-R region, 7472F (5 -TCAGGACAAGGGTGGTTTGAG-3 ) and 8182R (5 -CAAACAGCAAAAGGCTTTATTGG-3 ) were used for first round followed by 7472F and 8147R (5 -CCGGGCGACTCAGTCTATC-3 ) for second round. The reaction mixtures and conditions were as described above except for the following: For larger fragments, the final extension was done at 68 C for 10 min instead of 72 C for 2 min. All second round PCR products were column purified as described above and overlapping sequences were determined with internal primers. Nested RT-PCR for gag sequences was done as described (5) with modifications. GAG-O-R primer was used for 1st strand synthesis; cycle conditions were 52 o C annealing, for 35 cycles. For second round PCR, annealing was at 54 o C for 35 cycles. PCR analysis performed on 20 of the identical patient PBMC DNA specimens stored at the NCI (Frederick, MD) since 2007 confirmed nearly identical gag sequences, thereby diminishing the possibility of laboratory contamination as a source of XMRV. 3
5 4 Phylogenetic Analysis. Sequences were aligned using ClustalX (S3). Clustal alignments were imported into MEGA4 to generate neighbor-joining trees using the Kimura 2-parameter plus Γ distribution (K80+Γ) distance model (S4). Free parameters were reduced to the K80 model, and α values were estimated from the data set using a maximum likelihood approach in PAUP*4.0 (Sinauer Associates, Inc. Publishers, Sunderland, MA, USA). The bootstrap consensus tree inferred from 1000 replicates is taken to represent the evolutionary history of the taxa analyzed. Accession numbers were acquired from GenBank. ( FLV (NC_001940), MoMLV (NC_001501), XMRV VP35 (DQ241301), XMRV VP42 (DQ241302) XMRV VP62 (EF185282). Genomic Nonecotropic MLV Provirus Sequences were downloaded from PLOS Genetics (S5). Isolation, separation and culture of primary cells. Leukopaks of peripheral blood from healthy donors were collected according to a NIH approved IRB #99- CC-0168 protocol. Patients peripheral blood and plasma samples were from frozen banked samples obtained under NIH exempt status. Mononuclear leukocytes from both normal and patients cells were isolated by Ficoll-Hypaque gradient centrifugation. The light density fraction (buffy coat) was collected and washed twice with PBS. PBMC were activated by 1 µg/ml PHA (Abbott Diagnostics, Abbott Park, IL) and after 72 hours the cells were cultured with 20 units/ml of IL-2 (Zeptometrix, Buffalo, NY) and subcultured every 3-5 days. For isolation of CD4+T cells, CD8, CD11b, CD14, CD19, CD33 and CD56 positive cells were removed using magnetic activated cell sorting (MACs) methods according to the manufacturer s instructions (Miltenyi Biotec, Inc., Auburn, CA). After isolation, the CD3+, CD4+ T cells (>95% pure) were cultured in RPMI-1640 medium supplemented with 10% fetal calf serum (FCS), 2 mm glutamine, 1 mm sodium pyruvate and antibiotics. CD4 + T cells were activated by culturing with 20 units/ml of IL-2 and 1 µg/ml PHA. In vitro expansion of primary B-cells. NIH 3T3 cells transduced with a retroviral vector expressing CD40L (gift of Eugene Barsov, NCI-Frederick) were maintained in Dulbecco s Modified Eagle s Medium (DMEM) (Invitrogen, Carlsbad, CA) supplemented with 10% calf serum (CS) (Lonza, Basel, Switzerland) and 1% penicillin, streptomycin and L-glutamine (Invitrogen, Carlsbad, CA) at 37 C with 5% CO 2. To stimulate B cell expansion, ~3.5 x 10 6 NIH3T3-CD40L cells were trypsinized (0.25% trypsin with EDTA )(Invitrogen, Carlsbad, CA), resuspended in 3 ml medium and irradiated with an absorbed radiation dose (rad) of 9600 using a Cesium 137 irradiator. Cells plus 7 ml medium were added to a T25 cell culture flask (Corning, Corning, NY) and allowed to adhere (2-3 h) to the flask surface (optimal density ~50%). CD19+ B cells were isolated from PBMC using immunomagnetic bead technology (Miltenyi Biotec, Auburn, CA). CD19+ cells were separated from 10 8 freshly isolated PBMC by positive selection according to the manufacturer s protocol. After magnetic separation, CD19+ B cells (>95% pure) were added to
6 5 an irradiated NIH3T3-CD40L monolayer and incubated at 37 C with 5% CO 2. Cultures were monitored for B cell proliferation and split 1:5 every hr onto freshly irradiated NIH 3T3-CD40L monolayer. CD19+ primary B cells were cultured and expanded in primary B cell expansion media: Iscove s Modified Dulbecco s Medium (IMDM) (Invitrogen, Carlsbad, CA) + 10% FCS (Atlanta Biologicals, Lawrenceville, GA), 1% penicillin, streptomycin and L-glutamine (Invitrogen, Carlsbad, CA), 40 ng/ml interleukin 4 (IL-4) (PeproTech, Inc., Rocky Hill, NJ), 50 µg/ml holo-transferrin (Sigma, St. Louis, MO) and 5 µg/ml insulin (Invitrogen, Carlsbad, CA). Cell culture and reagents. Raji, SupT1 and LNCaP were obtained from American Type Culture Collection (ATCC, Manassas, VA). The cells were maintained in RPMI-1640 supplemented with L-glutamine (2 mm), penicillin (100 U/mL), streptomycin (100 ng/ml), and FCS (10%) and subcultured 1:5 every 4-5 days. HCD-57 cells are a mouse erythroleukemia cell line that expresses both ecotropic and polytropic MLVs; HCD-57/SFFV are HCD-57 cells infected with SFFV. BaF3-ER cells are a murine pro B cell line engineered to express the erythropoietin receptor. BaF3ER-SFFV Env cells were derived and maintained as described (S6). Flow cytometry for viral proteins. Adherent cells were incubated in trypsin for 10 minutes at 37 o C. After additional washes, adherent and suspension cells were incubated for 15 min at RT in 1 ml of paraformaldehyde (4% w/v in PBS), washed in permeabilization wash buffer (0.5% saponin 0.1%, sodium azide, 2% human AB sera in PBS) (PWB), and resuspended in 300 µl of permeabilization buffer (PBS with 2.5% saponin) (PB). After incubating at 22 o C for 20 min, 5 ml of human AB sera and either rat anti-mlv p30 mab, rat anti-sffv Env mab, goat anti-rauscher MLV gp70 Env, p30 Gag, or p10 Gag, or the appropriate isotype control (anti-rat IgG, rat myeloma supernatant, or preimmune goat serum) were added, and the cells incubated at 4 o C for an additional 30 min. Cells were then washed in PWB, resuspended in 100 µl of PB with 3 µl (0.6 µg) of FITCconjugated goat anti-rat IgG or rabbit anti-goat antibody (BD PharMingen, San Jose, CA) and incubated for 20 min at 4 o C. The efficiency of permeabilization was determined using a FITC-conjugated anti-actin antibody. Cells were then washed twice in PB, resuspended in 500 µl of sheath fluid (BD PharMingen, San Jose, CA) to prevent clumping and analyzed by flow cytometry. For experiments in which purified cell populations were examined, cells were stained with an anti- CD3 or anti-cd19 antibody prior to permeabilization, and analyzed by gating on the CD3 + or CD19+ subsets. Western Blot (WB) analysis. Cells were pelleted, washed twice with PBS, and lysed for 30 min on ice in RIPA lysis buffer (50 mm Tris, ph 7.4, 150 mm NaCl, 0.25% deoxycholate,1% NP-40, and protease inhibitor cocktail (Sigma, St. Louis, MO). Debris was removed by centrifugation for 15 min at 21,000x g at 4 o C. Protein concentration was determined with the Bio-Rad Protein Assay reagent
7 6 and equal amounts of protein ( ug) were separated by SDS-PAGE electrophoresis on 4-20% Tris-Glycine gels (Invitrogen, Carlsbad, CA) and then transferred to Immobilon-P membranes (Millipore, Billerica, MA). The membranes were blocked with 5% non-fat dry milk/1x TBST (Tris-buffered saline with 0.1% Triton X-100) for 1 h at room temperature, hybridized with the appropriate antiserum diluted in 5% non-fat dry milk/1xtbst for 2 h at room temperature or overnight at 4 o C, washed twice with 1xTBST, hybridized with the appropriate horseradish-peroxidase conjugated antibody diluted 1:5000 for 1 h at room temperature, and washed three times with 1xTBST. Hybridized bands were visualized using HyGlo chemiluminescent HRP antibody detection reagent (Denville Scientific, Metuchen, NJ) and exposed to film (Kodak, Rochester, NY). Antibodies used were a rat monoclonal Ab to SFFV gp55 Env (7C10), diluted 1:100 and detected with peroxidase-labeled anti-rat secondary antibody (Amersham, Waukesha, WI); goat anti-rauscher MLV gp70 Env, p30 Gag and p10 Gag (provided by NCI); and goat anti-nzb xenotropic MLV (provided by NCI), all diluted 1:2500 and detected with peroxidase labeled anti-goat secondary antibody (Santa Cruz Biotechnology, Santa Cruz, CA). Viral transmission. Frozen cell-free plasma and 0.22 µm filtered cell free supernatants from PBMC and T cell cultures were diluted 1:1 with tissue culture media and 600 µl aliquots were added to a six-well culture plate with the LNCaP cell line (50% confluent) or a million primary activated CD4+ T cells isolated from healthy donors. The plates were centrifuged for 5 min at 1500 RPM, rotated 180 o and centrifuged again for 5 min. The entire cycle was repeated once and cells were then diluted in their growth media. For cell-cell transmission, 1 x 10 6 T cells or PBMC without any IL-2 in the growth media were added to a six-well culture plate with the LNCaP cell line (50% confluent) in 1 ml of media for 3 h. After 1 hr, T cells in suspension were removed and the LNCaP cells were grown for several passages in the absence of IL-2 which caused any remaining T cells to die. At the times after transmission indicated, protein analysis was done by western blot and flow cytometry. Genotyping. The rs R462Q SNP was genotyped using Applied Biosystems Taqman 5' nucleotidase assays, Taqman Universal PCR Master Mix: No AmpErase UNG, and 5 ng of genomic DNA. The thermal cycling conditions consisted of an initial hold at 95 o C for 10 minutes followed by 50 cycles of a 15 second 95 o C denaturation step and a one minute 60 o C annealing and extension step. A 7900HT instrument was used to detect fluorescent probes, and Sequence Detection Software (SDS) v2.2 was used to discriminate alleles and call genotypes (Applied Biosystems, Foster City, CA). The variant is in Hardy-Weinberg equilibrium in both cases and controls. A Chi square test was performed for both genotypes and alleles of RNASEL comparing XMRV negative and XMRV positive controls. Both tests were not significant and the allele test is displayed in Table S2. Homozygous R462Q variant of RNASEL is represented in approximately 13% of the human population (S6, S7).
8 7 Flow cytometry for detection of antiviral antibodies in CFS plasma. The murine cell lines BaF3ER and BaF3ER-SFFV Env (S8) were grown in 2 units/ml of Epo in RPMI 1640 plus 7% FCS. 500,000 cells per sample in log phase were used as targets for direct staining. Cell lines were first washed in wash buffer (2% FBS, 0.02% Na Azide, PBS) and resuspended in 200 µl of BSA staining buffer (BD PharMingen, San Jose, CA). Patient plasma was thawed rapidly and used at 20 µl or 2 µl per tube (1:10 and 1:100 respectively) and incubated at 4 o C or on ice for 30 minutes. Cells were then washed with 0.5mL of wash buffer. Tubes were centrifuged at 800 rpm for 5 minutes, the supernatant was removed and tubes blotted on a towel. Next, 100 µl of the following working solution was added: 5 µl human A/B sera, 1 µl biotin labeled anti-human IgG (for human plasma) or biotin-labeled anti-rat IgG (for SFFV Env mab)(ebioscience, San Diego, CA), 1 µl of strep/avidin phycoerythrin (PE), 94 µl cold staining buffer. Samples were then incubated at 4 o C for 20 minutes, washed with 0.5 ml of wash buffer, and spun at 800 rpm for 5 minutes before being analyzed by flow cytometry. For the competition experiments, 100 µl of cold staining buffer and 10 µl of human plasma were added to each tube prior to addition of either anti- SFFV Env mab (7C10) or Y3 myeloma supernatant (control). Samples were incubated at 4 o C or on ice for 20 minutes, washed with 0.5 ml of wash buffer and spun at 800 rpm for 5 minutes before being analyzed by flow cytometry. Legends: Figure S1. Gag sequences of XMRV in CFS patients. Partial sequences (nt ) from CFS XMRV strains WPI-1130, WPI-1138 and WPI-1169 in comparison to XMRV strains VP35, VP42 and VP62 derived from prostate cancer patients. The yellow highlighting denotes the differences from the reference strain (VP62). Figure S2. Phylogenetic Analysis of XMRV in CFS patients. Neighbor-joining analysis of CFS XMRV strains WPI-1104, WPI-1106 and WPI-1178 with previously identified XMRVs and the nonecotropic MLVs xenotropic (Xmv), polytropic (Pmv) and modified polytropic (Mmpv) (S5). Ecotropic MLVs FLV and MoMLV were included to root the tree as outgroups in the phylogenetic reconstruction. Bootstrap supports of >70% are shown next to branch nodes. The scale measures evolutionary distance in substitutions per nucleotide. Subgroup designations are marked with arrows. XMRVs (WPI and VPs) form a distinct clade clustering together and separately from the xenotropic (Xmv) proviruses (shown in red). Figure S3. Detection of cloned XMRV-VP62 using a rat mab to SFFV Env and a goat antiserum to mouse NZB xenotropic MLV. A. Lysates were prepared from XMRV-VP62-infected Raji (lane1), LNCaP (lane 2) or Sup-T1 (lane 3). Positive
9 8 controls used were HCD-57 cells, a mouse erythroleukemia cell line expressing polytropic MLV gp70 Env (lane 4), and HCD-57 cells infected with SFFV, which also express SFFV gp55 Env (lane 5). WB analysis was carried out using rat anti-sffv Env mab 7C10. Molecular weight markers in kd are shown on the left. B. Lysates were prepared from XMRV-VP62-infected Raji (lane 1), LNCaP (lane 2) or Sup-T1 (lane 3). Lysates from SFFV-infected mouse HCD-57 cells (lane 4) and from uninfected Raji, LNCaP and Sup-T1 are shown in lanes 5-7, respectively. WB was carried out using goat antiserum to mouse NZB xenotropic MLV. Molecular weight markers in kd are shown on the left. Figure S4. Expression of XMRV proteins in PBMC from CFS patients. A. Activated B cells from CFS patient WPI-1125, activated T cells from CFS patient WPI-1105 or normal activatedt cells were incubated with goat antisera (black area) against Rauscher MLV gp70 Env (top), p30 Gag (middle) and p10 Gag (bottom) and analyzed by IFC. Preimmune goat serum (light area) was used as a control. B. A B cell line from a CFS patient was incubated with rat anti-sffv Env mab (right panel) or control myeloma supernatant (left panel) and then analyzed by IFC. C. Lysates were prepared from B cells (lane 1) or T cells (lanes 2 and 3) from CFS patients that had been grown for 42 days on CD40L or IL-2 respectively were analyzed by WB using rat anti-sffv Env mab (top panel) or goat anti-nzb xenotropic MLV serum (bottom panel). Lane 4: normal T cells; Lane 5: mouse HCD-57 cells; Lane 7: SFFV-infected HCD-57 cells. Molecular weight markers in kd are shown on the left. Figure S5: Infectious XMRV in CFS patients PBMC and plasma. A. The indicated T-cell cultures from CFS patients were co-cultured with LNCaP as described in the Methods. XMRV p30 Gag expression was detected in the LNCaP cells using a rat anti-mlv p30 Gag mab and IFC. Bottom panel: LNCaP co-cultured with normal T cells. B. Plasma from the indicated CFS patients was co-cultured with LNCaP. At the second passage, XMRV p30 Gag expression in the LNCaP cells was detected by flow cytometry using a rat anti-mlv p30 Gag monoclonal Ab. Co-culture with plasma from a normal healthy donor is shown in the bottom panel. Figure S6: Presence of antibodies in CFS plasma that recognize the cell surface of SFFV Env expressing BAF3ER cells. A. Plasma from CFS patients or normal healthy controls was diluted 1:10, reacted with BaF3-ER or BaF3ER- SFFV Env cells and analyzed by IFC. Shown is the difference in mean fluorescence intensity (MFI) between CFS and control plasma direct binding to BaF3ER-SFFV Env cells versus BaF3ER (control) cells. B. Competition experiment, carried out as described in the Methods, showing that plasma from a CFS patient can block binding of a rat anti-sffv Env mab to BaF3ER-SFFV Env cells. Left panel: CFS plasma diluted 1:10 (white area) eliminates most of the anti-sffv Env binding (striped area) and overlaps with the negative control (black area). Right panel: CFS plasma diluted 1:100 (white area) eliminates less
10 9 of the anti-sffv Env binding (striped area) and overlaps much more with the positive than the negative control (black area). Supporting References S1. Fukuda, K., S. E. Straus, et al. Ann, Intern. Med., 121, 953(1994). S2. E. DeFreitas et al., Proc Natl Acad Sci U S A 88, 2922 (1991). S3. JD Thompson et al., Nucleic Acids Res 25,4876 (1997). S4. K. Tamura et al., Molecular Biology and Evolution 24,1596 (2007). S5. P. Jern, J.P. Stoye, J.M. Coffin, PLOS Genetics 3, e183 (2007). S6. A. Rokman et al., Am J Hum Genet 70, 1299 (May, 2002). S7. L. Wang et al., Am J Hum Genet 71, 116 (2002). S8. K. Nishigaki et al., J Virol 75, 7893 ( 2001)
11
12
13
14
15
16
17
18
19 Partial Retraction Robert H. Silverman, 1 * Jaydip Das Gupta, 1 Vincent C. Lombardi, 2 Francis W. Ruscetti, 3 Max A. Pfost, 2 Kathryn S. Hagen, 2 Daniel L. Peterson, 2 Sandra K. Ruscetti, 4 Rachel K. Bagni, 5 Cari Petrow-Sadowski, 6 Bert Gold, 3 Michael Dean, 3 Judy A. Mikovits 2 1 Department of Cancer Biology, The Lerner Research Institute, The Cleveland Clinic Foundation, Cleveland, OH 44195, USA. 2 Whittemore Peterson Institute, Reno, NV 89557, USA. 3 Laboratory of Experimental Immunology, National Cancer Institute Frederick, Frederick, MD 21701, USA. 4 Laboratory of Cancer Prevention, National Cancer Institute Frederick, Frederick, MD 21701, USA. 5 Advanced Technology Program, National Cancer Institute Frederick, Frederick, MD 21701, USA. 6 Basic Research Program, Scientific Applications International Corporation, National Cancer Institute Frederick, Frederick, MD 21701, USA. *To whom correspondence should be addressed. silverr@ccf.org Present address: Sierra Internal Medicine, 865 Tahoe Boulevard No. 306, Incline Village, NV 89451, USA. In our 23 October 2009 Report, Detection of an infectious retrovirus, XMRV, in blood cells of patients with chronic fatigue syndrome (1), two of the coauthors, Silverman and Das Gupta, analyzed DNA samples from chronic fatigue syndrome (CFS) patients and healthy controls. A reexamination by Silverman and Das Gupta of the samples they used shows that some of the CFS peripheral blood mononuclear cell (PBMC) DNA preparations are contaminated with XMRV plasmid DNA (2). The following figures and table were based on the contaminated data: Figure 1, single-round PCR detection of XMRV sequences in CFS PBMC DNA samples; table S1, XMRV sequences previously attributed to CFS patients; and figure S2, the phylogenetic analysis of those sequences. Therefore, we are retracting those figures and table. References 1. V. C. Lombardi, F. W. Ruscetti, J. Das Gupta, M. A. Pfost, K. S. Hagen, D. L. Peterson, S. K. Ruscetti, R. K. Bagni, C. Petrow-Sadowski, B. Gold, M. Dean, R. H. Silverman, J. A. Mikovits, Science 326, 585 (2009). 2. Supporting online material showing the reanalysis is available at 1 Supporting Online Material Materials and Methods SOM Text Figs. S1 to S4 Tables S1 and S2 References 3 August 2011; accepted 16 September 2011 Published online 22 September 2011; /science Include this information when citing this paper. / / 22 September 2011 / Page 1/ /science
*To whom correspondence should be addressed. This PDF file includes:
 www.sciencemag.org/cgi/content/full/science.1212182/dc1 Supporting Online Material for Partial Retraction to Detection of an Infectious Retrovirus, XMRV, in Blood Cells of Patients with Chronic Fatigue
www.sciencemag.org/cgi/content/full/science.1212182/dc1 Supporting Online Material for Partial Retraction to Detection of an Infectious Retrovirus, XMRV, in Blood Cells of Patients with Chronic Fatigue
Picture of cell art Picture of cell under a microscope Daniel L. Peterson, MD Whittemore Peterson Institute
 XMRV, A New Human Pathogenic Retrovirus: Detection In Chronic Diseases Picture of cell art Picture of cell under a microscope Daniel L. Peterson, MD Whittemore Peterson Institute Daniel L. Peterson, MD
XMRV, A New Human Pathogenic Retrovirus: Detection In Chronic Diseases Picture of cell art Picture of cell under a microscope Daniel L. Peterson, MD Whittemore Peterson Institute Daniel L. Peterson, MD
Islet viability assay and Glucose Stimulated Insulin Secretion assay RT-PCR and Western Blot
 Islet viability assay and Glucose Stimulated Insulin Secretion assay Islet cell viability was determined by colorimetric (3-(4,5-dimethylthiazol-2-yl)-2,5- diphenyltetrazolium bromide assay using CellTiter
Islet viability assay and Glucose Stimulated Insulin Secretion assay Islet cell viability was determined by colorimetric (3-(4,5-dimethylthiazol-2-yl)-2,5- diphenyltetrazolium bromide assay using CellTiter
Supplementary Information
 Supplementary Information Supplementary Figure 1. CD4 + T cell activation and lack of apoptosis after crosslinking with anti-cd3 + anti-cd28 + anti-cd160. (a) Flow cytometry of anti-cd160 (5D.10A11) binding
Supplementary Information Supplementary Figure 1. CD4 + T cell activation and lack of apoptosis after crosslinking with anti-cd3 + anti-cd28 + anti-cd160. (a) Flow cytometry of anti-cd160 (5D.10A11) binding
Primary Adult Naïve CD4+ CD45RA+ Cells. Prepared by: David Randolph at University of Alabama, Birmingham
 Primary Adult Naïve CD4+ CD45RA+ Cells Prepared by: David Randolph (drdrdr@uab.edu) at University of Alabama, Birmingham Goal: To obtain large numbers of highly pure primary CD4+ CD45RO- CD25- cells from
Primary Adult Naïve CD4+ CD45RA+ Cells Prepared by: David Randolph (drdrdr@uab.edu) at University of Alabama, Birmingham Goal: To obtain large numbers of highly pure primary CD4+ CD45RO- CD25- cells from
Supplementary Materials for
 immunology.sciencemag.org/cgi/content/full/2/16/eaan6049/dc1 Supplementary Materials for Enzymatic synthesis of core 2 O-glycans governs the tissue-trafficking potential of memory CD8 + T cells Jossef
immunology.sciencemag.org/cgi/content/full/2/16/eaan6049/dc1 Supplementary Materials for Enzymatic synthesis of core 2 O-glycans governs the tissue-trafficking potential of memory CD8 + T cells Jossef
Protocol for Gene Transfection & Western Blotting
 The schedule and the manual of basic techniques for cell culture Advanced Protocol for Gene Transfection & Western Blotting Schedule Day 1 26/07/2008 Transfection Day 3 28/07/2008 Cell lysis Immunoprecipitation
The schedule and the manual of basic techniques for cell culture Advanced Protocol for Gene Transfection & Western Blotting Schedule Day 1 26/07/2008 Transfection Day 3 28/07/2008 Cell lysis Immunoprecipitation
MTC-TT and TPC-1 cell lines were cultured in RPMI medium (Gibco, Breda, The Netherlands)
 Supplemental data Materials and Methods Cell culture MTC-TT and TPC-1 cell lines were cultured in RPMI medium (Gibco, Breda, The Netherlands) supplemented with 15% or 10% (for TPC-1) fetal bovine serum
Supplemental data Materials and Methods Cell culture MTC-TT and TPC-1 cell lines were cultured in RPMI medium (Gibco, Breda, The Netherlands) supplemented with 15% or 10% (for TPC-1) fetal bovine serum
Gladstone Institutes, University of California (UCSF), San Francisco, USA
 Fluorescence-linked Antigen Quantification (FLAQ) Assay for Fast Quantification of HIV-1 p24 Gag Marianne Gesner, Mekhala Maiti, Robert Grant and Marielle Cavrois * Gladstone Institutes, University of
Fluorescence-linked Antigen Quantification (FLAQ) Assay for Fast Quantification of HIV-1 p24 Gag Marianne Gesner, Mekhala Maiti, Robert Grant and Marielle Cavrois * Gladstone Institutes, University of
No Evidence of Murine-Like Gammaretroviruses in CFS Patients Previously Identified as XMRV-Infected
 No Evidence of Murine-Like Gammaretroviruses in CFS Patients Previously Identified as XMRV-Infected Konstance Knox, 1,2 Donald Carrigan, 1,2 Graham Simmons, 3,4 Fernando Teque, 5 Yanchen Zhou, 3,4 John
No Evidence of Murine-Like Gammaretroviruses in CFS Patients Previously Identified as XMRV-Infected Konstance Knox, 1,2 Donald Carrigan, 1,2 Graham Simmons, 3,4 Fernando Teque, 5 Yanchen Zhou, 3,4 John
Detailed step-by-step operating procedures for NK cell and CTL degranulation assays
 Supplemental methods Detailed step-by-step operating procedures for NK cell and CTL degranulation assays Materials PBMC isolated from patients, relatives and healthy donors as control K562 cells (ATCC,
Supplemental methods Detailed step-by-step operating procedures for NK cell and CTL degranulation assays Materials PBMC isolated from patients, relatives and healthy donors as control K562 cells (ATCC,
In vitro human regulatory T cell expansion
 - 1 - Human CD4 + CD25 + CD127 dim/- regulatory T cell Workflow isolation, in vitro expansion and analysis In vitro human regulatory T cell expansion Introduction Regulatory T (Treg) cells are a subpopulation
- 1 - Human CD4 + CD25 + CD127 dim/- regulatory T cell Workflow isolation, in vitro expansion and analysis In vitro human regulatory T cell expansion Introduction Regulatory T (Treg) cells are a subpopulation
The Schedule and the Manual of Basic Techniques for Cell Culture
 The Schedule and the Manual of Basic Techniques for Cell Culture 1 Materials Calcium Phosphate Transfection Kit: Invitrogen Cat.No.K2780-01 Falcon tube (Cat No.35-2054:12 x 75 mm, 5 ml tube) Cell: 293
The Schedule and the Manual of Basic Techniques for Cell Culture 1 Materials Calcium Phosphate Transfection Kit: Invitrogen Cat.No.K2780-01 Falcon tube (Cat No.35-2054:12 x 75 mm, 5 ml tube) Cell: 293
Plasmids Western blot analysis and immunostaining Flow Cytometry Cell surface biotinylation RNA isolation and cdna synthesis
 Plasmids psuper-retro-s100a10 shrna1 was constructed by cloning the dsdna oligo 5 -GAT CCC CGT GGG CTT CCA GAG CTT CTT TCA AGA GAA GAA GCT CTG GAA GCC CAC TTT TTA-3 and 5 -AGC TTA AAA AGT GGG CTT CCA GAG
Plasmids psuper-retro-s100a10 shrna1 was constructed by cloning the dsdna oligo 5 -GAT CCC CGT GGG CTT CCA GAG CTT CTT TCA AGA GAA GAA GCT CTG GAA GCC CAC TTT TTA-3 and 5 -AGC TTA AAA AGT GGG CTT CCA GAG
In vitro human regulatory T cell expansion
 - 1 - Human CD4 + CD25 + regulatory T cell isolation, Workflow in vitro expansion and analysis In vitro human regulatory T cell expansion Introduction Regulatory T (Treg) cells are a subpopulation of T
- 1 - Human CD4 + CD25 + regulatory T cell isolation, Workflow in vitro expansion and analysis In vitro human regulatory T cell expansion Introduction Regulatory T (Treg) cells are a subpopulation of T
HIV-1 Virus-like Particle Budding Assay Nathan H Vande Burgt, Luis J Cocka * and Paul Bates
 HIV-1 Virus-like Particle Budding Assay Nathan H Vande Burgt, Luis J Cocka * and Paul Bates Department of Microbiology, Perelman School of Medicine at the University of Pennsylvania, Philadelphia, USA
HIV-1 Virus-like Particle Budding Assay Nathan H Vande Burgt, Luis J Cocka * and Paul Bates Department of Microbiology, Perelman School of Medicine at the University of Pennsylvania, Philadelphia, USA
Phosphate buffered saline (PBS) for washing the cells TE buffer (nuclease-free) ph 7.5 for use with the PrimePCR Reverse Transcription Control Assay
 Catalog # Description 172-5080 SingleShot Cell Lysis Kit, 100 x 50 µl reactions 172-5081 SingleShot Cell Lysis Kit, 500 x 50 µl reactions For research purposes only. Introduction The SingleShot Cell Lysis
Catalog # Description 172-5080 SingleShot Cell Lysis Kit, 100 x 50 µl reactions 172-5081 SingleShot Cell Lysis Kit, 500 x 50 µl reactions For research purposes only. Introduction The SingleShot Cell Lysis
Supplementary data Supplementary Figure 1 Supplementary Figure 2
 Supplementary data Supplementary Figure 1 SPHK1 sirna increases RANKL-induced osteoclastogenesis in RAW264.7 cell culture. (A) RAW264.7 cells were transfected with oligocassettes containing SPHK1 sirna
Supplementary data Supplementary Figure 1 SPHK1 sirna increases RANKL-induced osteoclastogenesis in RAW264.7 cell culture. (A) RAW264.7 cells were transfected with oligocassettes containing SPHK1 sirna
TITLE: Examination of a Newly Discovered Human Retrovirus, XMRV, in Breast Cancer
 AWARD NUMBER: W81XWH-10-1-0464 TITLE: Examination of a Newly Discovered Human Retrovirus, XMRV, in Breast Cancer PRINCIPAL INVESTIGATOR: Fayth K. Yoshimura, Ph.D. CONTRACTING ORGANIZATION: Wayne State
AWARD NUMBER: W81XWH-10-1-0464 TITLE: Examination of a Newly Discovered Human Retrovirus, XMRV, in Breast Cancer PRINCIPAL INVESTIGATOR: Fayth K. Yoshimura, Ph.D. CONTRACTING ORGANIZATION: Wayne State
E.Z.N.A. SQ Blood DNA Kit II. Table of Contents
 E.Z.N.A. SQ Blood DNA Kit II Table of Contents Introduction and Overview...2 Kit Contents/Storage and Stability...3 Blood Storage and DNA Yield...4 Preparing Reagents...5 100-500 μl Whole Blood Protocol...6
E.Z.N.A. SQ Blood DNA Kit II Table of Contents Introduction and Overview...2 Kit Contents/Storage and Stability...3 Blood Storage and DNA Yield...4 Preparing Reagents...5 100-500 μl Whole Blood Protocol...6
Direct ex vivo characterization of human antigen-specific CD154 + CD4 + T cells Rapid antigen-reactive T cell enrichment (Rapid ARTE)
 Direct ex vivo characterization of human antigen-specific CD154 + CD4 + T cells Rapid antigen-reactive T cell enrichment (Rapid ARTE) Introduction Workflow Antigen (ag)-specific T cells play a central
Direct ex vivo characterization of human antigen-specific CD154 + CD4 + T cells Rapid antigen-reactive T cell enrichment (Rapid ARTE) Introduction Workflow Antigen (ag)-specific T cells play a central
Supporting Information
 Supporting Information Pang et al. 10.1073/pnas.1322009111 SI Materials and Methods ELISAs. These assays were performed as previously described (1). ELISA plates (MaxiSorp Nunc; Thermo Fisher Scientific)
Supporting Information Pang et al. 10.1073/pnas.1322009111 SI Materials and Methods ELISAs. These assays were performed as previously described (1). ELISA plates (MaxiSorp Nunc; Thermo Fisher Scientific)
Rapid antigen-specific T cell enrichment (Rapid ARTE)
 Direct ex vivo characterization of human antigen-specific CD154+CD4+ T cell Rapid antigen-specific T cell enrichment (Rapid ARTE) Introduction Workflow Antigen (ag)-specific T cells play a central role
Direct ex vivo characterization of human antigen-specific CD154+CD4+ T cell Rapid antigen-specific T cell enrichment (Rapid ARTE) Introduction Workflow Antigen (ag)-specific T cells play a central role
Figure S1. Schematic presentation of genomic replication of idsiv after transfection and infection. After transfection of idsiv plasmid DNA into 293T
 Figure S1. Schematic presentation of genomic replication of idsiv after transfection and infection. After transfection of idsiv plasmid DNA into 293T cells, the RNA genomes with all modifications are generated
Figure S1. Schematic presentation of genomic replication of idsiv after transfection and infection. After transfection of idsiv plasmid DNA into 293T cells, the RNA genomes with all modifications are generated
SUPPLEMENTARY INFORMATION. Involvement of IL-21 in the epidermal hyperplasia of psoriasis
 SUPPLEMENTARY INFORMATION Involvement of IL-21 in the epidermal hyperplasia of psoriasis Roberta Caruso 1, Elisabetta Botti 2, Massimiliano Sarra 1, Maria Esposito 2, Carmine Stolfi 1, Laura Diluvio 2,
SUPPLEMENTARY INFORMATION Involvement of IL-21 in the epidermal hyperplasia of psoriasis Roberta Caruso 1, Elisabetta Botti 2, Massimiliano Sarra 1, Maria Esposito 2, Carmine Stolfi 1, Laura Diluvio 2,
CHAPTER 4 RESULTS. showed that all three replicates had similar growth trends (Figure 4.1) (p<0.05; p=0.0000)
 CHAPTER 4 RESULTS 4.1 Growth Characterization of C. vulgaris 4.1.1 Optical Density Growth study of Chlorella vulgaris based on optical density at 620 nm (OD 620 ) showed that all three replicates had similar
CHAPTER 4 RESULTS 4.1 Growth Characterization of C. vulgaris 4.1.1 Optical Density Growth study of Chlorella vulgaris based on optical density at 620 nm (OD 620 ) showed that all three replicates had similar
For the 5 GATC-overhang two-oligo adaptors set up the following reactions in 96-well plate format:
 Supplementary Protocol 1. Adaptor preparation: For the 5 GATC-overhang two-oligo adaptors set up the following reactions in 96-well plate format: Per reaction X96 10X NEBuffer 2 10 µl 10 µl x 96 5 -GATC
Supplementary Protocol 1. Adaptor preparation: For the 5 GATC-overhang two-oligo adaptors set up the following reactions in 96-well plate format: Per reaction X96 10X NEBuffer 2 10 µl 10 µl x 96 5 -GATC
Chromatin IP (Isw2) Fix soln: 11% formaldehyde, 0.1 M NaCl, 1 mm EDTA, 50 mm Hepes-KOH ph 7.6. Freshly prepared. Do not store in glass bottles.
 Chromatin IP (Isw2) 7/01 Toshi last update: 06/15 Reagents Fix soln: 11% formaldehyde, 0.1 M NaCl, 1 mm EDTA, 50 mm Hepes-KOH ph 7.6. Freshly prepared. Do not store in glass bottles. 2.5 M glycine. TBS:
Chromatin IP (Isw2) 7/01 Toshi last update: 06/15 Reagents Fix soln: 11% formaldehyde, 0.1 M NaCl, 1 mm EDTA, 50 mm Hepes-KOH ph 7.6. Freshly prepared. Do not store in glass bottles. 2.5 M glycine. TBS:
Situation of XMRV and Blood Transfusion. Celso Bianco, MD ISBT Working Party on TTID Lisbon, June 19, 2011
 Situation of XMRV and Blood Transfusion Celso Bianco, MD ISBT Working Party on TTID Lisbon, June 19, 2011 Lombardi et al. Science 326, 585 (2009) Conclusions: CFS and XMRV XMRV found in 67% of CFS patients
Situation of XMRV and Blood Transfusion Celso Bianco, MD ISBT Working Party on TTID Lisbon, June 19, 2011 Lombardi et al. Science 326, 585 (2009) Conclusions: CFS and XMRV XMRV found in 67% of CFS patients
Essential Medium, containing 10% fetal bovine serum, 100 U/ml penicillin and 100 µg/ml streptomycin. Huvec were cultured in
 Supplemental data Methods Cell culture media formulations A-431 and U-87 MG cells were maintained in Dulbecco s Modified Eagle s Medium. FaDu cells were cultured in Eagle's Minimum Essential Medium, containing
Supplemental data Methods Cell culture media formulations A-431 and U-87 MG cells were maintained in Dulbecco s Modified Eagle s Medium. FaDu cells were cultured in Eagle's Minimum Essential Medium, containing
TSH Receptor Monoclonal Antibody (49) Catalog Number MA3-218 Product data sheet
 Website: thermofisher.com Customer Service (US): 1 800 955 6288 ext. 1 Technical Support (US): 1 800 955 6288 ext. 441 TSH Receptor Monoclonal Antibody (49) Catalog Number MA3-218 Product data sheet Details
Website: thermofisher.com Customer Service (US): 1 800 955 6288 ext. 1 Technical Support (US): 1 800 955 6288 ext. 441 TSH Receptor Monoclonal Antibody (49) Catalog Number MA3-218 Product data sheet Details
Online Data Supplement. Anti-aging Gene Klotho Enhances Glucose-induced Insulin Secretion by Upregulating Plasma Membrane Retention of TRPV2
 Online Data Supplement Anti-aging Gene Klotho Enhances Glucose-induced Insulin Secretion by Upregulating Plasma Membrane Retention of TRPV2 Yi Lin and Zhongjie Sun Department of physiology, college of
Online Data Supplement Anti-aging Gene Klotho Enhances Glucose-induced Insulin Secretion by Upregulating Plasma Membrane Retention of TRPV2 Yi Lin and Zhongjie Sun Department of physiology, college of
In vitro human regulatory T cell suppression assay
 Human CD4 + CD25 + regulatory T cell isolation, in vitro suppression assay and analysis In vitro human regulatory T cell suppression assay Introduction Regulatory T (Treg) cells are a subpopulation of
Human CD4 + CD25 + regulatory T cell isolation, in vitro suppression assay and analysis In vitro human regulatory T cell suppression assay Introduction Regulatory T (Treg) cells are a subpopulation of
Recombinant Protein Expression Retroviral system
 Recombinant Protein Expression Retroviral system Viruses Contains genome DNA or RNA Genome encased in a protein coat or capsid. Some viruses have membrane covering protein coat enveloped virus Ø Essential
Recombinant Protein Expression Retroviral system Viruses Contains genome DNA or RNA Genome encased in a protein coat or capsid. Some viruses have membrane covering protein coat enveloped virus Ø Essential
CD14 + S100A9 + Monocytic Myeloid-Derived Suppressor Cells and Their Clinical Relevance in Non-Small Cell Lung Cancer
 CD14 + S1A9 + Monocytic Myeloid-Derived Suppressor Cells and Their Clinical Relevance in Non-Small Cell Lung Cancer Po-Hao, Feng M.D., Kang-Yun, Lee, M.D. Ph.D., Ya-Ling Chang, Yao-Fei Chan, Lu- Wei, Kuo,Ting-Yu
CD14 + S1A9 + Monocytic Myeloid-Derived Suppressor Cells and Their Clinical Relevance in Non-Small Cell Lung Cancer Po-Hao, Feng M.D., Kang-Yun, Lee, M.D. Ph.D., Ya-Ling Chang, Yao-Fei Chan, Lu- Wei, Kuo,Ting-Yu
Supplementary Data Table of Contents:
 Supplementary Data Table of Contents: - Supplementary Methods - Supplementary Figures S1(A-B) - Supplementary Figures S2 (A-B) - Supplementary Figures S3 - Supplementary Figures S4(A-B) - Supplementary
Supplementary Data Table of Contents: - Supplementary Methods - Supplementary Figures S1(A-B) - Supplementary Figures S2 (A-B) - Supplementary Figures S3 - Supplementary Figures S4(A-B) - Supplementary
T H 1, T H 2 and T H 17 polarization of naïve CD4 + mouse T cells
 A complete workflow for cell preparation, isolation, polarization and analysis T H 1, T H 2 and T H 17 polarization of naïve CD4 + mouse T cells Introduction Workflow CD4 + T helper (T H) cells play a
A complete workflow for cell preparation, isolation, polarization and analysis T H 1, T H 2 and T H 17 polarization of naïve CD4 + mouse T cells Introduction Workflow CD4 + T helper (T H) cells play a
Product Contents. 1 Specifications 1 Product Description. 2 Buffer Preparation... 3 Protocol. 3 Ordering Information 4
 INSTRUCTION MANUAL Quick-RNA Midiprep Kit Catalog No. R1056 Highlights 10 minute method for isolating RNA (up to 1 mg) from a wide range of cell types and tissue samples. Clean-Spin column technology allows
INSTRUCTION MANUAL Quick-RNA Midiprep Kit Catalog No. R1056 Highlights 10 minute method for isolating RNA (up to 1 mg) from a wide range of cell types and tissue samples. Clean-Spin column technology allows
RNA extraction, RT-PCR and real-time PCR. Total RNA were extracted using
 Supplementary Information Materials and Methods RNA extraction, RT-PCR and real-time PCR. Total RNA were extracted using Trizol reagent (Invitrogen,Carlsbad, CA) according to the manufacturer's instructions.
Supplementary Information Materials and Methods RNA extraction, RT-PCR and real-time PCR. Total RNA were extracted using Trizol reagent (Invitrogen,Carlsbad, CA) according to the manufacturer's instructions.
PBMC from each patient were suspended in AIM V medium (Invitrogen) with 5% human
 Anti-CD19-CAR transduced T-cell preparation PBMC from each patient were suspended in AIM V medium (Invitrogen) with 5% human AB serum (Gemini) and 300 international units/ml IL-2 (Novartis). T cell proliferation
Anti-CD19-CAR transduced T-cell preparation PBMC from each patient were suspended in AIM V medium (Invitrogen) with 5% human AB serum (Gemini) and 300 international units/ml IL-2 (Novartis). T cell proliferation
Supplementary Data 1. Alanine substitutions and position variants of APNCYGNIPL. Applied in
 Supplementary Data 1. Alanine substitutions and position variants of APNCYGNIPL. Applied in Supplementary Fig. 2 Substitution Sequence Position variant Sequence original APNCYGNIPL original APNCYGNIPL
Supplementary Data 1. Alanine substitutions and position variants of APNCYGNIPL. Applied in Supplementary Fig. 2 Substitution Sequence Position variant Sequence original APNCYGNIPL original APNCYGNIPL
Supplemental Materials and Methods Plasmids and viruses Quantitative Reverse Transcription PCR Generation of molecular standard for quantitative PCR
 Supplemental Materials and Methods Plasmids and viruses To generate pseudotyped viruses, the previously described recombinant plasmids pnl4-3-δnef-gfp or pnl4-3-δ6-drgfp and a vector expressing HIV-1 X4
Supplemental Materials and Methods Plasmids and viruses To generate pseudotyped viruses, the previously described recombinant plasmids pnl4-3-δnef-gfp or pnl4-3-δ6-drgfp and a vector expressing HIV-1 X4
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Figure 1: Cryopreservation alters CD62L expression by CD4 T cells. Freshly isolated (left) or cryopreserved PBMCs (right) were stained with the mix of antibodies described
Supplementary Figure 1 Supplementary Figure 1: Cryopreservation alters CD62L expression by CD4 T cells. Freshly isolated (left) or cryopreserved PBMCs (right) were stained with the mix of antibodies described
Data Sheet TIGIT / NFAT Reporter - Jurkat Cell Line Catalog #60538
 Data Sheet TIGIT / NFAT Reporter - Jurkat Cell Line Catalog #60538 Background: TIGIT is a co-inhibitory receptor that is highly expressed in Natural Killer (NK) cells, activated CD4+, CD8+ and regulatory
Data Sheet TIGIT / NFAT Reporter - Jurkat Cell Line Catalog #60538 Background: TIGIT is a co-inhibitory receptor that is highly expressed in Natural Killer (NK) cells, activated CD4+, CD8+ and regulatory
Figure S1. Generation of inducible PTEN deficient mice and the BMMCs (A) B6.129 Pten loxp/loxp mice were mated with B6.
 Figure S1. Generation of inducible PTEN deficient mice and the BMMCs (A) B6.129 Pten loxp/loxp mice were mated with B6.129-Gt(ROSA)26Sor tm1(cre/ert2)tyj /J mice. To induce deletion of the Pten locus,
Figure S1. Generation of inducible PTEN deficient mice and the BMMCs (A) B6.129 Pten loxp/loxp mice were mated with B6.129-Gt(ROSA)26Sor tm1(cre/ert2)tyj /J mice. To induce deletion of the Pten locus,
Supplemental Experimental Procedures
 Cell Stem Cell, Volume 2 Supplemental Data A Temporal Switch from Notch to Wnt Signaling in Muscle Stem Cells Is Necessary for Normal Adult Myogenesis Andrew S. Brack, Irina M. Conboy, Michael J. Conboy,
Cell Stem Cell, Volume 2 Supplemental Data A Temporal Switch from Notch to Wnt Signaling in Muscle Stem Cells Is Necessary for Normal Adult Myogenesis Andrew S. Brack, Irina M. Conboy, Michael J. Conboy,
Ex vivo Human Antigen-specific T Cell Proliferation and Degranulation Willemijn Hobo 1, Wieger Norde 1 and Harry Dolstra 2*
 Ex vivo Human Antigen-specific T Cell Proliferation and Degranulation Willemijn Hobo 1, Wieger Norde 1 and Harry Dolstra 2* 1 Department of Laboratory Medicine - Laboratory of Hematology, Radboud University
Ex vivo Human Antigen-specific T Cell Proliferation and Degranulation Willemijn Hobo 1, Wieger Norde 1 and Harry Dolstra 2* 1 Department of Laboratory Medicine - Laboratory of Hematology, Radboud University
Figure S1. PMVs from THP-1 cells expose phosphatidylserine and carry actin. A) Flow
 SUPPLEMENTARY DATA Supplementary Figure Legends Figure S1. PMVs from THP-1 cells expose phosphatidylserine and carry actin. A) Flow cytometry analysis of PMVs labelled with annexin-v-pe (Guava technologies)
SUPPLEMENTARY DATA Supplementary Figure Legends Figure S1. PMVs from THP-1 cells expose phosphatidylserine and carry actin. A) Flow cytometry analysis of PMVs labelled with annexin-v-pe (Guava technologies)
Antibodies: LB1 buffer For 50 ml For 10ml For 30 ml Final 1 M HEPES, ph 2.5 ml 0.5 ml 1.5 ml 50mM. 5 M NaCl 1.4 ml 280 µl 0.
 Experiment: Date: Tissue: Purpose: ChIP-Seq Antibodies: 11x cross-link buffer: Regent Stock Solution Final Vol for 10 ml of 11xstock concentration 5 M NaCl 0.1M 0.2 ml 0.5 M EDTA 1 mm 20 ul 0.5 M EGTA,
Experiment: Date: Tissue: Purpose: ChIP-Seq Antibodies: 11x cross-link buffer: Regent Stock Solution Final Vol for 10 ml of 11xstock concentration 5 M NaCl 0.1M 0.2 ml 0.5 M EDTA 1 mm 20 ul 0.5 M EGTA,
TFEB-mediated increase in peripheral lysosomes regulates. Store Operated Calcium Entry
 TFEB-mediated increase in peripheral lysosomes regulates Store Operated Calcium Entry Luigi Sbano, Massimo Bonora, Saverio Marchi, Federica Baldassari, Diego L. Medina, Andrea Ballabio, Carlotta Giorgi
TFEB-mediated increase in peripheral lysosomes regulates Store Operated Calcium Entry Luigi Sbano, Massimo Bonora, Saverio Marchi, Federica Baldassari, Diego L. Medina, Andrea Ballabio, Carlotta Giorgi
SUPPORTING MATREALS. Methods and Materials
 SUPPORTING MATREALS Methods and Materials Cell Culture MC3T3-E1 (subclone 4) cells were maintained in -MEM with 10% FBS, 1% Pen/Strep at 37ºC in a humidified incubator with 5% CO2. MC3T3 cell differentiation
SUPPORTING MATREALS Methods and Materials Cell Culture MC3T3-E1 (subclone 4) cells were maintained in -MEM with 10% FBS, 1% Pen/Strep at 37ºC in a humidified incubator with 5% CO2. MC3T3 cell differentiation
Technical Resources. BD Immunocytometry Systems. FastImmune Intracellular Cytokine Staining Procedures
 FastImmune Intracellular Cytokine Staining Procedures BD has developed protocols for the detection of intracellular cytokines in activated lymphocytes and in activated monocytes. The procedures have been
FastImmune Intracellular Cytokine Staining Procedures BD has developed protocols for the detection of intracellular cytokines in activated lymphocytes and in activated monocytes. The procedures have been
Serum Amyloid A3 Gene Expression in Adipocytes is an Indicator. of the Interaction with Macrophages
 Serum Amyloid A3 Gene Expression in Adipocytes is an Indicator of the Interaction with Macrophages Yohei Sanada, Takafumi Yamamoto, Rika Satake, Akiko Yamashita, Sumire Kanai, Norihisa Kato, Fons AJ van
Serum Amyloid A3 Gene Expression in Adipocytes is an Indicator of the Interaction with Macrophages Yohei Sanada, Takafumi Yamamoto, Rika Satake, Akiko Yamashita, Sumire Kanai, Norihisa Kato, Fons AJ van
Product Manual. Omni-Array Sense Strand mrna Amplification Kit, 2 ng to 100 ng Version Catalog No.: Reactions
 Genetic Tools and Reagents Universal mrna amplification, sense strand amplification, antisense amplification, cdna synthesis, micro arrays, gene expression, human, mouse, rat, guinea pig, cloning Omni-Array
Genetic Tools and Reagents Universal mrna amplification, sense strand amplification, antisense amplification, cdna synthesis, micro arrays, gene expression, human, mouse, rat, guinea pig, cloning Omni-Array
RayBio KinaseSTAR TM Akt Activity Assay Kit
 Activity Assay Kit User Manual Version 1.0 March 13, 2015 RayBio KinaseSTAR TM Akt Activity Kit Protocol (Cat#: 68AT-Akt-S40) RayBiotech, Inc. We Provide You With Excellent Support And Service Tel:(Toll
Activity Assay Kit User Manual Version 1.0 March 13, 2015 RayBio KinaseSTAR TM Akt Activity Kit Protocol (Cat#: 68AT-Akt-S40) RayBiotech, Inc. We Provide You With Excellent Support And Service Tel:(Toll
Mammalian Membrane Protein Extraction Kit
 Mammalian Membrane Protein Extraction Kit Catalog number: AR0155 Boster s Mammalian Membrane Protein Extraction Kit is a simple, rapid and reproducible method to prepare cellular protein fractions highly
Mammalian Membrane Protein Extraction Kit Catalog number: AR0155 Boster s Mammalian Membrane Protein Extraction Kit is a simple, rapid and reproducible method to prepare cellular protein fractions highly
90 min 18 min. 45 min. 14 d
 Isolation, cultivation, and expansion of Pan T cells from human PBMCs In vitro expansion of human Pan T cells Introduction T lymphocytes (T cells) play a central role in the adaptive immune system by controlling
Isolation, cultivation, and expansion of Pan T cells from human PBMCs In vitro expansion of human Pan T cells Introduction T lymphocytes (T cells) play a central role in the adaptive immune system by controlling
XMRV among Prostate Cancer Patients from the Southern United States and Analysis of Possible Correlates of Infection
 XMRV among Prostate Cancer Patients from the Southern United States and Analysis of Possible Correlates of Infection Bryan Danielson Baylor College of Medicine Department of Molecular Virology and Microbiology
XMRV among Prostate Cancer Patients from the Southern United States and Analysis of Possible Correlates of Infection Bryan Danielson Baylor College of Medicine Department of Molecular Virology and Microbiology
Product Contents. 1 Specifications 1 Product Description. 2 Buffer Preparation... 3 Protocol. 3 Ordering Information 4 Related Products..
 INSTRUCTION MANUAL Quick-RNA MidiPrep Catalog No. R1056 Highlights 10 minute method for isolating RNA (up to 1 mg) from a wide range of cell types and tissue samples. Clean-Spin column technology allows
INSTRUCTION MANUAL Quick-RNA MidiPrep Catalog No. R1056 Highlights 10 minute method for isolating RNA (up to 1 mg) from a wide range of cell types and tissue samples. Clean-Spin column technology allows
Luminescent platforms for monitoring changes in the solubility of amylin and huntingtin in living cells
 Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2016 Contents Supporting Information Luminescent platforms for monitoring changes in the
Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2016 Contents Supporting Information Luminescent platforms for monitoring changes in the
SUPPLEMENTARY MATERIAL
 SUPPLEMENTARY MATERIAL Table S1. Primers and fluorescent probes used for qrt-pcr analysis of relative expression levels of PPP family phosphatases. gene name forward primer, 5-3 probe, 5-3 reverse primer,
SUPPLEMENTARY MATERIAL Table S1. Primers and fluorescent probes used for qrt-pcr analysis of relative expression levels of PPP family phosphatases. gene name forward primer, 5-3 probe, 5-3 reverse primer,
SUPPLEMENTAL INFORMATION
 SUPPLEMENTAL INFORMATION EXPERIMENTAL PROCEDURES Tryptic digestion protection experiments - PCSK9 with Ab-3D5 (1:1 molar ratio) in 50 mm Tris, ph 8.0, 150 mm NaCl was incubated overnight at 4 o C. The
SUPPLEMENTAL INFORMATION EXPERIMENTAL PROCEDURES Tryptic digestion protection experiments - PCSK9 with Ab-3D5 (1:1 molar ratio) in 50 mm Tris, ph 8.0, 150 mm NaCl was incubated overnight at 4 o C. The
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells
 MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
Hepatitis B Virus Genemer
 Product Manual Hepatitis B Virus Genemer Primer Pair for amplification of HBV Viral Specific Fragment Catalog No.: 60-2007-10 Store at 20 o C For research use only. Not for use in diagnostic procedures
Product Manual Hepatitis B Virus Genemer Primer Pair for amplification of HBV Viral Specific Fragment Catalog No.: 60-2007-10 Store at 20 o C For research use only. Not for use in diagnostic procedures
VEGFR2-Mediated Vascular Dilation as a Mechanism of VEGF-Induced Anemia and Bone Marrow Cell Mobilization
 Cell Reports, Volume 9 Supplemental Information VEGFR2-Mediated Vascular Dilation as a Mechanism of VEGF-Induced Anemia and Bone Marrow Cell Mobilization Sharon Lim, Yin Zhang, Danfang Zhang, Fang Chen,
Cell Reports, Volume 9 Supplemental Information VEGFR2-Mediated Vascular Dilation as a Mechanism of VEGF-Induced Anemia and Bone Marrow Cell Mobilization Sharon Lim, Yin Zhang, Danfang Zhang, Fang Chen,
Development of sero-assays to screen for XMRV antibodies in clinical samples
 National Cancer Institute Development of sero-assays to screen for XMRV antibodies in clinical samples Rachel K. Bagni, Ph.D. September 8, 2010 Xentropic murine leukaemia virus-related virus - XMRV The
National Cancer Institute Development of sero-assays to screen for XMRV antibodies in clinical samples Rachel K. Bagni, Ph.D. September 8, 2010 Xentropic murine leukaemia virus-related virus - XMRV The
p47 negatively regulates IKK activation by inducing the lysosomal degradation of polyubiquitinated NEMO
 Supplementary Information p47 negatively regulates IKK activation by inducing the lysosomal degradation of polyubiquitinated NEMO Yuri Shibata, Masaaki Oyama, Hiroko Kozuka-Hata, Xiao Han, Yuetsu Tanaka,
Supplementary Information p47 negatively regulates IKK activation by inducing the lysosomal degradation of polyubiquitinated NEMO Yuri Shibata, Masaaki Oyama, Hiroko Kozuka-Hata, Xiao Han, Yuetsu Tanaka,
Anti-Lamin B1/LMNB1 Picoband Antibody
 Anti-Lamin B1/LMNB1 Picoband Antibody Catalog Number:PB9611 About LMNB1 Lamin-B1 is a protein that in humans is encoded by the LMNB1 gene. The nuclear lamina consists of a two-dimensional matrix of proteins
Anti-Lamin B1/LMNB1 Picoband Antibody Catalog Number:PB9611 About LMNB1 Lamin-B1 is a protein that in humans is encoded by the LMNB1 gene. The nuclear lamina consists of a two-dimensional matrix of proteins
Human Apolipoprotein A1 EIA Kit
 A helping hand for your research Product Manual Human Apolipoprotein A1 EIA Kit Catalog Number: 83901 96 assays 1 Table of Content Product Description 3 Assay Principle 3 Kit Components 3 Storage 4 Reagent
A helping hand for your research Product Manual Human Apolipoprotein A1 EIA Kit Catalog Number: 83901 96 assays 1 Table of Content Product Description 3 Assay Principle 3 Kit Components 3 Storage 4 Reagent
Graveley Lab shrna knockdown followed by RNA-seq Biosample Preparation and Characterization Document
 Graveley Lab shrna knockdown followed by RNA-seq Biosample Preparation and Characterization Document Wet Lab: Sara Olson and Lijun Zhan Computational Lab: Xintao Wei and Michael Duff PI: Brenton Graveley
Graveley Lab shrna knockdown followed by RNA-seq Biosample Preparation and Characterization Document Wet Lab: Sara Olson and Lijun Zhan Computational Lab: Xintao Wei and Michael Duff PI: Brenton Graveley
Supplementary Figure 1. Method development.
 Supplementary Figure 1 Method development. Titration experiments to determine standard antibody:lysate concentration. Lysates (~2 mg of total proteins) were prepared from cells expressing FLAG- tagged
Supplementary Figure 1 Method development. Titration experiments to determine standard antibody:lysate concentration. Lysates (~2 mg of total proteins) were prepared from cells expressing FLAG- tagged
Instructions for Use. RealStar Influenza Screen & Type RT-PCR Kit /2017 EN
 Instructions for Use RealStar Influenza Screen & Type RT-PCR Kit 4.0 05/2017 EN RealStar Influenza Screen & Type RT-PCR Kit 4.0 For research use only! (RUO) 164003 INS-164000-EN-S01 96 05 2017 altona
Instructions for Use RealStar Influenza Screen & Type RT-PCR Kit 4.0 05/2017 EN RealStar Influenza Screen & Type RT-PCR Kit 4.0 For research use only! (RUO) 164003 INS-164000-EN-S01 96 05 2017 altona
Supplementary Figures
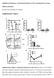 Inhibition of Pulmonary Anti Bacterial Defense by IFN γ During Recovery from Influenza Infection By Keer Sun and Dennis W. Metzger Supplementary Figures d a Ly6G Percentage survival f 1 75 5 1 25 1 5 1
Inhibition of Pulmonary Anti Bacterial Defense by IFN γ During Recovery from Influenza Infection By Keer Sun and Dennis W. Metzger Supplementary Figures d a Ly6G Percentage survival f 1 75 5 1 25 1 5 1
For Research Use Only Ver
 INSTRUCTION MANUAL Quick-RNA Miniprep Kit Catalog Nos. R1054 & R1055 Highlights High-quality total RNA (including small RNAs) from a wide range of samples. You can opt to isolate small and large RNAs in
INSTRUCTION MANUAL Quick-RNA Miniprep Kit Catalog Nos. R1054 & R1055 Highlights High-quality total RNA (including small RNAs) from a wide range of samples. You can opt to isolate small and large RNAs in
Sipper BK Experimental Animal Co. (Shanghai, China) and bred in a specific. pathogen-free environment. The animal study protocol was approved by the
 Supplementary information, Data S1 Materials and Methods Mice, Ad vectors and reagents Female C57BL/6 mice, 8-10 weeks of age, were purchased from Joint Ventures Sipper BK Experimental Animal Co. (Shanghai,
Supplementary information, Data S1 Materials and Methods Mice, Ad vectors and reagents Female C57BL/6 mice, 8-10 weeks of age, were purchased from Joint Ventures Sipper BK Experimental Animal Co. (Shanghai,
For Research Use Only Ver
 INSTRUCTION MANUAL Quick-RNA Miniprep Kit Catalog Nos. R1054 & R1055 Highlights High-quality total RNA (including small RNAs) from a wide range of samples. You can opt to isolate small and large RNAs in
INSTRUCTION MANUAL Quick-RNA Miniprep Kit Catalog Nos. R1054 & R1055 Highlights High-quality total RNA (including small RNAs) from a wide range of samples. You can opt to isolate small and large RNAs in
Supplemental Information. T Cells Enhance Autoimmunity by Restraining Regulatory T Cell Responses via an Interleukin-23-Dependent Mechanism
 Immunity, Volume 33 Supplemental Information T Cells Enhance Autoimmunity by Restraining Regulatory T Cell Responses via an Interleukin-23-Dependent Mechanism Franziska Petermann, Veit Rothhammer, Malte
Immunity, Volume 33 Supplemental Information T Cells Enhance Autoimmunity by Restraining Regulatory T Cell Responses via an Interleukin-23-Dependent Mechanism Franziska Petermann, Veit Rothhammer, Malte
Cell isolation. Spleen and lymph nodes (axillary, inguinal) were removed from mice
 Supplementary Methods: Cell isolation. Spleen and lymph nodes (axillary, inguinal) were removed from mice and gently meshed in DMEM containing 10% FBS to prepare for single cell suspensions. CD4 + CD25
Supplementary Methods: Cell isolation. Spleen and lymph nodes (axillary, inguinal) were removed from mice and gently meshed in DMEM containing 10% FBS to prepare for single cell suspensions. CD4 + CD25
Transfection of Sf9 cells with recombinant Bacmid DNA
 Transposition Bacmid DNA Mini Culturing baculo cells Transfection of Sf9 cells with recombinant Bacmid DNA Amplification of the virus Titration of baculo stocks Testing the expression Transposition 1.
Transposition Bacmid DNA Mini Culturing baculo cells Transfection of Sf9 cells with recombinant Bacmid DNA Amplification of the virus Titration of baculo stocks Testing the expression Transposition 1.
MagniSort Human CD4 Memory T cell Enrichment Kit Catalog Number: RUO: For Research Use Only. Not for use in diagnostic procedures.
 Page 1 of 2 MagniSort Human CD4 Memory T cell Enrichment Kit RUO: For Research Use Only. Not for use in diagnostic procedures. Normal human peripheral blood mononuclear cells were unsorted (left) or sorted
Page 1 of 2 MagniSort Human CD4 Memory T cell Enrichment Kit RUO: For Research Use Only. Not for use in diagnostic procedures. Normal human peripheral blood mononuclear cells were unsorted (left) or sorted
Commercially available HLA Class II tetramers (Beckman Coulter) conjugated to
 Class II tetramer staining Commercially available HLA Class II tetramers (Beckman Coulter) conjugated to PE were combined with dominant HIV epitopes (DRB1*0101-DRFYKTLRAEQASQEV, DRB1*0301- PEKEVLVWKFDSRLAFHH,
Class II tetramer staining Commercially available HLA Class II tetramers (Beckman Coulter) conjugated to PE were combined with dominant HIV epitopes (DRB1*0101-DRFYKTLRAEQASQEV, DRB1*0301- PEKEVLVWKFDSRLAFHH,
Tel: ; Fax: ;
 Tel.: +98 216 696 9291; Fax: +98 216 696 9291; E-mail: mrasadeghi@pasteur.ac.ir Tel: +98 916 113 7679; Fax: +98 613 333 6380; E-mail: abakhshi_e@ajums.ac.ir A Soluble Chromatin-bound MOI 0 1 5 0 1 5 HDAC2
Tel.: +98 216 696 9291; Fax: +98 216 696 9291; E-mail: mrasadeghi@pasteur.ac.ir Tel: +98 916 113 7679; Fax: +98 613 333 6380; E-mail: abakhshi_e@ajums.ac.ir A Soluble Chromatin-bound MOI 0 1 5 0 1 5 HDAC2
Peptide stimulation and Intracellular Cytokine Staining
 v Peptide stimulation and Intracellular Cytokine Staining Authors A. Cosma, S. Allgayer MATERIALS Date 01-02-2007 REAGENTS: Version 1.0 - PBMCs - Culture medium - Costimulating antibodies - Peptide pools
v Peptide stimulation and Intracellular Cytokine Staining Authors A. Cosma, S. Allgayer MATERIALS Date 01-02-2007 REAGENTS: Version 1.0 - PBMCs - Culture medium - Costimulating antibodies - Peptide pools
HCC1937 is the HCC1937-pcDNA3 cell line, which was derived from a breast cancer with a mutation
 SUPPLEMENTARY INFORMATION Materials and Methods Human cell lines and culture conditions HCC1937 is the HCC1937-pcDNA3 cell line, which was derived from a breast cancer with a mutation in exon 20 of BRCA1
SUPPLEMENTARY INFORMATION Materials and Methods Human cell lines and culture conditions HCC1937 is the HCC1937-pcDNA3 cell line, which was derived from a breast cancer with a mutation in exon 20 of BRCA1
Application of μmacs Streptavidin MicroBeads for the analysis of HIV-1 directly from patient plasma
 Excerpt from MACS&more Vol 8 1/2004 Application of μmacs Streptavidin MicroBeads for the analysis of HIV-1 directly from patient plasma L. Davis Lupo and Salvatore T. Butera HIV and Retrovirology Branch,
Excerpt from MACS&more Vol 8 1/2004 Application of μmacs Streptavidin MicroBeads for the analysis of HIV-1 directly from patient plasma L. Davis Lupo and Salvatore T. Butera HIV and Retrovirology Branch,
Procaspase-3. Cleaved caspase-3. actin. Cytochrome C (10 M) Z-VAD-fmk. Procaspase-3. Cleaved caspase-3. actin. Z-VAD-fmk
 A HeLa actin - + + - - + Cytochrome C (1 M) Z-VAD-fmk PMN - + + - - + actin Cytochrome C (1 M) Z-VAD-fmk Figure S1. (A) Pan-caspase inhibitor z-vad-fmk inhibits cytochrome c- mediated procaspase-3 cleavage.
A HeLa actin - + + - - + Cytochrome C (1 M) Z-VAD-fmk PMN - + + - - + actin Cytochrome C (1 M) Z-VAD-fmk Figure S1. (A) Pan-caspase inhibitor z-vad-fmk inhibits cytochrome c- mediated procaspase-3 cleavage.
2,6,9-Triazabicyclo[3.3.1]nonanes as overlooked. amino-modification products by acrolein
![2,6,9-Triazabicyclo[3.3.1]nonanes as overlooked. amino-modification products by acrolein 2,6,9-Triazabicyclo[3.3.1]nonanes as overlooked. amino-modification products by acrolein](/thumbs/86/94743397.jpg) Supplementary Information 2,6,9-Triazabicyclo[3.3.1]nonanes as overlooked amino-modification products by acrolein Ayumi Tsutsui and Katsunori Tanaka* Biofunctional Synthetic Chemistry Laboratory, RIKEN
Supplementary Information 2,6,9-Triazabicyclo[3.3.1]nonanes as overlooked amino-modification products by acrolein Ayumi Tsutsui and Katsunori Tanaka* Biofunctional Synthetic Chemistry Laboratory, RIKEN
Instructions for Use. APO-AB Annexin V-Biotin Apoptosis Detection Kit 100 tests
 3URGXFW,QIRUPDWLRQ Sigma TACS Annexin V Apoptosis Detection Kits Instructions for Use APO-AB Annexin V-Biotin Apoptosis Detection Kit 100 tests For Research Use Only. Not for use in diagnostic procedures.
3URGXFW,QIRUPDWLRQ Sigma TACS Annexin V Apoptosis Detection Kits Instructions for Use APO-AB Annexin V-Biotin Apoptosis Detection Kit 100 tests For Research Use Only. Not for use in diagnostic procedures.
Norgen s HIV proviral DNA PCR Kit was developed and validated to be used with the following PCR instruments: Qiagen Rotor-Gene Q BioRad icycler
 3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com HIV Proviral DNA PCR Kit Product # 33840 Product Insert Background Information
3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com HIV Proviral DNA PCR Kit Product # 33840 Product Insert Background Information
ADVL0411 study. (temozolomide in patients with leukemia)
 ADVL0411 study (temozolomide in patients with leukemia) I. Overview of study: We are interested in determining the level of MGMT activity and MRS mutations in leukemic samples (plasma and lymphoblasts)
ADVL0411 study (temozolomide in patients with leukemia) I. Overview of study: We are interested in determining the level of MGMT activity and MRS mutations in leukemic samples (plasma and lymphoblasts)
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/1175194/dc1 Supporting Online Material for A Vital Role for Interleukin-21 in the Control of a Chronic Viral Infection John S. Yi, Ming Du, Allan J. Zajac* *To whom
www.sciencemag.org/cgi/content/full/1175194/dc1 Supporting Online Material for A Vital Role for Interleukin-21 in the Control of a Chronic Viral Infection John S. Yi, Ming Du, Allan J. Zajac* *To whom
Isolation, Propagation, and Titration of Human Immunodeficiency Virus Type 1 From Peripheral Blood of Infected Individuals
 Isolation of HIV-1 From PBMC of Infected Individuals 17 2 Isolation, Propagation, and Titration of Human Immunodeficiency Virus Type 1 From Peripheral Blood of Infected Individuals Hanneke Schuitemaker
Isolation of HIV-1 From PBMC of Infected Individuals 17 2 Isolation, Propagation, and Titration of Human Immunodeficiency Virus Type 1 From Peripheral Blood of Infected Individuals Hanneke Schuitemaker
Purification and Fluorescent Labeling of Exosomes Asuka Nanbo 1*, Eri Kawanishi 2, Ryuji Yoshida 2 and Hironori Yoshiyama 3
 Purification and Fluorescent Labeling of Exosomes Asuka Nanbo 1*, Eri Kawanishi 2, Ryuji Yoshida 2 and Hironori Yoshiyama 3 1 Graduate School of Medicine, Hokkaido University, Sapporo, Japan; 2 Graduate
Purification and Fluorescent Labeling of Exosomes Asuka Nanbo 1*, Eri Kawanishi 2, Ryuji Yoshida 2 and Hironori Yoshiyama 3 1 Graduate School of Medicine, Hokkaido University, Sapporo, Japan; 2 Graduate
Western Immunoblotting Preparation of Samples:
 Western Immunoblotting Preparation of Samples: Total Protein Extraction from Culture Cells: Take off the medium Wash culture with 1 x PBS 1 ml hot Cell-lysis Solution into T75 flask Scrap out the cells
Western Immunoblotting Preparation of Samples: Total Protein Extraction from Culture Cells: Take off the medium Wash culture with 1 x PBS 1 ml hot Cell-lysis Solution into T75 flask Scrap out the cells
COMPONENT NAME COMPONENT # QUANTITY STORAGE SHELF LIFE FORMAT. Store at 2-8 C. Do not freeze. Store at 2-8 C. Do not freeze.
 This document is available at www.stemcell.com/pis Catalog #18765 EasySep Mouse CD4+CD62L+ T Cell Isolation Kit For processing 1x 10^9 cells Description Isolate highly purified naïve CD4+ T cells (CD4+CD62L+)
This document is available at www.stemcell.com/pis Catalog #18765 EasySep Mouse CD4+CD62L+ T Cell Isolation Kit For processing 1x 10^9 cells Description Isolate highly purified naïve CD4+ T cells (CD4+CD62L+)
Supporting Online Material Material and Methods References Supplemental Figures S1, S2, and S3
 Supporting Online Material Material and Methods References Supplemental Figures S1, S2, and S3 Sarbassov et al. 1 Material and Methods Materials Reagents were obtained from the following sources: protein
Supporting Online Material Material and Methods References Supplemental Figures S1, S2, and S3 Sarbassov et al. 1 Material and Methods Materials Reagents were obtained from the following sources: protein
Supporting Information
 Supporting Information Valkenburg et al. 10.1073/pnas.1403684111 SI Materials and Methods ELISA and Microneutralization. Sera were treated with Receptor Destroying Enzyme II (RDE II, Accurate) before ELISA
Supporting Information Valkenburg et al. 10.1073/pnas.1403684111 SI Materials and Methods ELISA and Microneutralization. Sera were treated with Receptor Destroying Enzyme II (RDE II, Accurate) before ELISA
