REPORT Expression of GJB2 and GJB6 Is Reduced in a Novel DFNB1 Allele
|
|
|
- Clarissa Paula Adams
- 6 years ago
- Views:
Transcription
1 REPORT Expression of GJB2 and GJB6 Is Reduced in a Novel DFNB1 Allele Ellen Wilch, Mei Zhu, Kirk B. Burkhart, * Martha Regier, and Karen H. Friderici Jill L. Elfenbein, Rachel A. Fisher, In a large kindred of German descent, we found a novel allele that segregates with deafness when present in trans with the 35delG allele of GJB2. Qualitative polymerase chain reaction based allele-specific expression assays showed that expression of both GJB2 and GJB6 from the novel allele is dramatically reduced. This is the first evidence of a deafnessassociated regulatory mutation of GJB2 and of potential coregulation of GJB2 and GJB6. Mutations in GJB2, the gene encoding gap junction protein connexin 26 (Cx26), are the most common cause of prelingual-onset, recessively inherited, nonsyndromic, sensorineural hearing loss (SNHL) in humans. GJB2 and GJB6, which encodes connexin 30 (Cx30), comprise the DFNB1/A3 locus at 13q12 (fig. 1). In the vertebrate cochlea, Cx26 and Cx30 are coexpressed as heteromeric connexons in nonsensory cells of the organ of Corti as well as in subsets of cells of the spiral ligament and stria vascularis 1 3 and are implicated in the maintenance of cochlear K gradients necessary for proper hair-cell function. 4,5 More than 80 recessive mutations of GJB2, nearly all affecting proper translation of the Cx26 protein, and several dominant missense mutations are implicated in DFNB1/A3 hearing loss. Others cause skin disease with accompanying SNHL, including Vohwinkel syndrome (MIM ), Bart-Pumphrey syndrome (MIM ), palmoplantar keratoderma, with deafness (PPK [MIM ]), and keratitis-ichthyosis-deafness (KID [MIM ]). Three dominant missense mutations of GJB6 have been shown to cause the skin disorder Clouston syndrome (hidrotic ectodermal dysplasia [MIM ]). In contrast to the abundance and diversity of GJB2 hearing-loss mutations, only a single dominant mutation of GJB6 causing nonsyndromic hearing loss had been published 6 before the identification of two large deletions of 309 kb and 232 kb: del(gjb6-d13s1830) and del(gjb6- D13S1854), respectively truncating the 5 end of GJB6 (fig. 1). These deletions segregate with hearing loss when present homozygously or heterozygously with each other or in trans with a recessive GJB2 mutation Investigators have suggested, but have not demonstrated, that loss of appropriate regulation of GJB2 from chromosomes bearing these deletions may underlie the hearing loss in these individuals, 7 11 which may be exacerbated by loss of one GJB6 allele. Common et al. 12 showed immunohistochemical evidence that Cx26 expression is disrupted in certain skin-cell types in an individual bearing the larger deletion in trans with GJB2 35delG, suggesting disruption of a GJB2 cis-regulatory element located within this deleted interval. Mutation screening of GJB2 in individuals whose hearing loss and family history are consistent with GJB2 deafness reveals a significant number of subjects with only one identified mutation. Additional screening for GJB6 deletions explains only some of these heterozygotes. We report here evidence of a novel pathogenic DFNB1 allele for which we demonstrate reduction in or loss of detectable expression of message from both GJB2 and GJB6. Figure 2 shows a small portion of the pedigree of MSU-DF5, a large American kindred of mainly German descent. After obtaining written informed consent, we collected DNA and performed audiological testing on 1200 family members. We screened all samples for GJB2 35delG and for SLC26A4 L445W. Ten MSU-DF5 family members are hearing impaired due to homozygosity of 35delG (one, DF5-68, is shown in fig. 2); one deaf family member has received a diagnosis of Pendred syndrome (MIM ) and is homozygous for SLC26A4 L445W (not shown in pedigree). Microsatellite and SNP genotyping across the GJB2/GJB6 genomic region (fig. 1 and table 1) allowed us to define a number of haplotypes. Three distinct haplotypes bearing 35delG exist within the family, indicating more than one founder for 35delG in this community (two 35delG haplotypes appear in fig. 2). Notably, four deaf family members, DF5-20, -70, -122, and -194, are heterozygous for 35delG and share a common haplotype (indicated in fig. 2 by a black bar with a star) that is longer than 600 kb and shorter than 3.1 Mb on their non-35delg chromosome. These individuals are profoundly hearing impaired, in contrast to DF5-68, who is homozygous for 35delG and the father of DF5-70 (fig. 3). Of the 14 family members who carry the novel allele, all those with 35delG on their other chromosome (4 individuals) have hearing loss, whereas all those with any other allele (10 individuals) have normal hearing ( P!.001, by Fisher s exact test). We also performed a linkage analysis (easylinkage Plus From the Genetics Program (E.W.; K.H.F.) and Departments of Microbiology & Molecular Genetics (M.Z.; K.B.B.; K.H.F.), Epidemiology (M.R.), Audiology & Speech Sciences (J.L.E.), and Pediatrics & Human Development (R.A.F.; K.H.F.), Michigan State University, East Lansing Received March 6, 2006; accepted for publication April 20, 2006; electronically published May 17, Address for correspondence and reprints: Dr. Karen Friderici, Microbiology & Molecular Genetics, 5163 Biomedical Physical Sciences Building, Michigan State University, East Lansing, MI frideric@msu.edu * Present affiliation: Life Sciences Institute, University of Michigan, Ann Arbor. Present affiliation: Bioport Corporation, Lansing, MI. Am. J. Hum. Genet. 2006;79: by The American Society of Human Genetics. All rights reserved /2006/ $ The American Journal of Human Genetics Volume 79 July
2 Figure 1. Map encompassing haplotyped region of 13q11-12, showing locations of GJB2, GJB6, genotyped microsatellite and SNP markers, and breakpoints of del(gjb6-d13s1830) and del(gjb6-d13s1854), all shown approximately to scale. The locations of the variants used for allele-specific expression assays GJB2 35delG, GJB2 94, and rs are also indicated. Boxes indicate exons, and the hatched boxes indicate the coding regions of the two genes. Vertical lines show approximate locations of genotyped microsatellites (in bold) and SNPs. D13S232 is the most proximal microsatellite showing recombination of the novel haplotype in affected individuals. The transcriptional start sites of GJB2 and GJB6 are indicated by right-angled arrows. v.3.01rc1, SuperLink v1.4) at the DFNB1 locus, using 28 individuals (shown in the boxed areas in fig. 2). We assigned the same mutant status to both the novel haplotype and the 35delG allele, and all other haplotypes were considered nonmutant. The resulting LOD score of 2.08 indicates that there is a good likelihood that the novel haplotype harbors a DFNB1 mutation. We sequenced the entire coding regions and 5 and 3 UTRs of both GJB2 and GJB6 in three of our affected probands, including splice sites, alternative exons and promoter regions, the entire single intron of GJB2, and several kilobases of sequence around each gene, and found no Figure 2. A small portion of the MSU-DF5 pedigree, with DFNB1 haplotypes indicated for selected individuals. Five individuals with congenital SNHL are represented by blackened symbols. One of these, DF5-68, is hearing impaired because of homozygosity of GJB2 35delG (DG). The other four are heterozygous for GJB2 35delG and bear the same haplotype on their non-35delg chromosome (black bar with star). Individuals contained within the boxed areas were included in the linkage analysis. sequence variants unique to the novel haplotype. Heterozygosities found in sequencing also showed that GJB2 and GJB6 are intact on both chromosomes and that the affected individuals do not bear either del(gjb6-d13s1830) or del(gjb6-d13s1854). Southern blotting indicated no rearrangements or unusual methylation around GJB2 (fig. 4). We hypothesized that an unidentified cis-regulatory element of GJB2 exists within the DFNB1 locus and is disrupted on our novel pathogenic allele. Given the imprac- Table 1. Selected Haplotype Assignments Based on Genotypes at Microsatellites and SNPs in a 620-kb Region of Chromosome 13 Including GJB2 and GJB6 Marker Location (bp) Novel DFNB1 Allele A B C D E D13S ,580, rs ,660,956 C T T C C C 35delG 19,661,685 DG DG rs ,662,895 C T T C C C rs ,664,130 T C C T C T GJB ,664,944 C C C A C C SNP ,665,308 T C C T C T rs ,665,957 C A ND C A ND rs ,668,529 T A ND T T ND A21445T 19,673,593 A A A A T A rs ,673,997 C G ND C C C A19277T 19,675,761 A ND A ND T A rs ,682,018 A G G A G A rs ,694,497 C C C C A C D13S175 19,746, rs ,795,790 T C C T C C rs ,795,911 C G G C G G rs ,795,932 A C C A C C rs ,796,279 T C C T C C rs ,796,316 A G G A G G rs ,796,330 A T T A T T rs ,837,040 G A G G G G rs ,025,569 ND ND G C G G rs ,198,712 T T ND A A A NOTE. The A and B columns indicate delg (DG) alleles carried by DF5-20 and DF5-70, respectively. The C column corresponds to the haplotype shown by a box with stippling, D to the haplotype shown by a box with horizontal lines, and E to the haplotype shown by a white box in figures 2, 5, and 6. Locations are based on UCSC Genome Browser, May 2004 assembly. p wild type; ND p not determined. The American Journal of Human Genetics Volume 79 July
3 Figure 3. Audiograms of four MSU-DF5 family members (identified by ID/sex/age). db HLp decibels hearing level; ANSI p American National Standards Institute. DF5-20, -70, and -122 are GJB2 35delG (35DG) heterozygotes bearing the novel pathogenic allele. Note that all three have profound SNHL at all tested frequencies. By contrast, DF5-68, homozygous for GJB2 35delG and the father of DF5-70, has significantly more residual hearing, particularly in his left ear. Although only pure-tone air-conduction thresholds are shown (circle p right ear; # p left ear; arrows p threshold beyond tested limit), bone conduction thresholds and immittance measures (not shown) indicate that the hearing loss in all four family members is sensorineural. ticality of searching for candidate variants across the entire locus, we first sought to acquire evidence of loss of GJB2 expression from this allele. We developed three allele-specific PCR assays to assess the relative abundance of transcript from each allele, using tissue both easily available and expected to express Cx26. To isolate RNA, buccal cells from several family members were collected on Cytosoft brushes and were immediately stored in 1 ml Trizol (Gibco/Life Technologies) on ice. Some samples were stored at 20 C for one to several days. RNA was isolated after the Trizol protocol, was resuspended in 20 ml H 2 O, and was stored at 80 C. We followed a modified Superscript II (Invitrogen) reverse-transcription protocol for Figure 4. Southern blot of region around GJB2, in DF5-20 and an unrelated control. The legend is available in its entirety in the online edition of The American Journal of Human Genetics. cdna synthesis; 5 8 ml template RNA, 1 ml of 0.5 mg/ml oligo(dt) primer, 1 ml of 10-mM dntp, and 2 ml H 2 O were heated to 70 C for 10 min, were chilled on ice, and then were mixed and incubated at 42 C for 2 min after 4 ml of 5# first-strand buffer and 2 ml of 0.1-M dithiothreitol was added. Addition of 1 ml Superscript II reverse transcriptase was followed by incubation at 42 C for 50 min and then incubation at 70 C for 15 min. A final incubation at 37 C for 20 min followed the addition of 1 ml RNaseH. A negative control to test for genomic DNA contamination was generated for each sample by omitting the Superscript II and replacing it with 1 ml H 2 O. These negative controls were carried through the assays side by side with the corresponding sample. In no instance was product observed for these controls (data not shown). We designed a cdna-specific PCR assay to assess the relative abundance of 35delG and non-35delg product amplified from GJB2 cdna synthesized from 35delG heterozygotes (fig. 5A). A standard PCR-based assay for the identification of 35delG in genomic DNA 13 is based on the introduction of a restriction site for BstNI by a 3 mismatch primer (5 -GCTGGTGGAGTGTTTGTTCACACCCGC-3 ) 176 The American Journal of Human Genetics Volume 79 July
4 Figure 5. Allele-specific expression assays for GJB2. A, Schematic showing design of BstNI (which assays for 35delG [35DG] genotype) and BseYI (which assays for 94 genotype) digestion assay performed using 139-bp PCR product amplified from cdna only. A forward primer (F) located in the first, noncoding exon of GJB2 is separated on genomic DNA from the reverse mismatch primer (R), located in the coding region of GJB2, by a 13-kb intron (F p forward primer for genomic 35delG assay). The BstNI site is introduced into product amplified from wild-type (non-35delg [indicated by ]) template. B and D, Pedigrees including DF5-70 and DF5-20, heterozygous for both GJB2 35delG and the novel pathogenic allele (black bar with star). Also indicated are the 35delG and 94 genotypes. C and E, Results of cdna-specific assay for GJB2 expression based on 35delG genotype. Products of 139 bp were amplified from all tested family members (ud p undigested, no enzyme added; d p addition of restriction enzyme). DF5-20, -70, and -71 (normalhearing sibling of DF5-70) are all heterozygous for 35delG; however, after digestion with BstNI, only DF5-71 clearly shows both 139- bp and 110-bp products, indicating representation of two different alleles at the cdna level. DF5-20 and DF5-70, who both bear the novel pathogenic non-35delg allele, show either no or barely detectable 110-bp product, indicating underrepresentation of this allele in the pool of amplified product (M p Marker V [Roche]). F, Results of cdna-specific assay for GJB2 expression based on genotype at GJB2 94. Products of 139 bp (ud p undigested) were amplified from both tested family members, who are both heterozygous (A/C) at 94. Although the digested product (d) amplified from DF5-72 cdna clearly indicates heterozygosity at the cdna level, lack of any detectable 139-bp product in the digested DF5-67 sample indicates underrepresentation of the novel (C) allele among the amplified product. that overlaps the run of Gs between nt 30 and nt 35. The BstNI restriction site is present in PCR product amplified from wild-type (non-35delg) alleles only and is absent in PCR product amplified from 35delG alleles. We paired a 5 primer located in exon 1 (5 -CGCAGAGACCCCAACGC- CGAGA-3 )withthe3 mismatch primer to generate a 139- bp PCR product from cdna template only, which, when incubated with BstNI, will yield digestion products of 110 bp and 29 bp only if the 35delG mutation is not present. For each individual we assayed, we amplified from cdna template and from the corresponding negative control. A 20-ml volume PCR (1 5 ml template cdna, 2 ml10# Qiagen buffer, 4 ml Q solution (Qiagen), 0.16 ml of 25-mM dntp (Invitrogen), 1 ml of each 20-mM primer, 5 U Invitrogen Taq DNA polymerase, and remainder H 2 O) was denatured for 3 min at 94 C, followed by 40 cycles at 94 C for 30 s, 66 C for 30 s, and 72 C for 30 s, with a final extension at 72 C for 5 min. Of each PCR, 10 ml was digested with 10 U BstNI (New England Biolabs) for 12 h,and3ml of each reaction was run on a 3.5% NuSieve 3:1 agarose gel containing 0.3 mg/ml ethidium bromide. The results (fig. 5B 5E) for the two deaf 35delG heterozygotes that we assayed, DF5-20 and DF5-70, indicate that, although they are heterozygous for 35delG at the genomic DNA level, the barely detectable or absent 110- bp product indicates underrepresentation of GJB2 transcript from the novel (non-35delg) allele. Results from the parents and sibling of DF5-70 are consistent with their genotypes and provide appropriate controls. Unbiased amplification and digestion of both 35delG and non- 35delG alleles is shown by the result for DF5-71, the sibling of DF5-70, who is also heterozygous for 35delG but carries a normal non-35delg chromosome. Sequencing of genomic DNA across the 139-bp target sequence (not shown) confirmed that there are no differences between these two individuals that would give rise to a bias in either amplification or digestion. To demonstrate that loss of expression of this allele is The American Journal of Human Genetics Volume 79 July
5 not unique to individuals carrying the 35delG mutation on their other chromosome 13, we developed a second assay to determine allele-specific GJB2 expression in a normal-hearing adult offspring of DF5-20 bearing the novel haplotype (fig. 5A and 5D). DF5-67 carries a previously unreported exon 1 SNP at position 94. BseYI digestion of the 139-bp product amplified with the same primers as described above will yield 102-bp and 27-bp products only if an A is present at position 94 (10 ml of each PCR product was digested with 5 U BseYI enzyme [New England Biolabs] for 12 h,5ml of 1% SDS was added, and electrophoresis was performed as above). Although both DF5-67 and -72 are heterozygous for 94 at the genomic level, all of the product amplified from DF5-67 cdna was digested (fig. 5F), indicating that no or very little of the PCR product had been amplified from template derived from the novel allele that bears the common variant, C. Again, sequencing of genomic DNA across the target sequence shows no differences between DF5-67 and DF5-72 that would yield biased amplification or digestion of the resulting amplimers. We hypothesized that loss of expression of GJB2 might be accompanied by loss of expression of GJB6, since these two genes lie within 30 kb of each other and their products are coexpressed in the cochlea. DF5-65, a normal-hearing adult offspring of DF5-20, bears the novel haplotype and is heterozygous for rs in the 3 UTR of GJB6, allowing us to test this hypothesis (fig. 6A and 6B). Using a forward primer located in the fourth noncoding exon (5 -CACCATTGGCTTCTAGGCAC-3 ) and a reverse primer located close to the 3 end of the 3 UTR (5 -CCACACTGTT- CCGTCTACAT-3 ), we amplified a 1,550-bp product from cdna template only, in a 20-ml PCR under the following conditions: 2 ml of10# Qiagen buffer, 4 ml Q solution (Qiagen), 0.16 ml of 25-mM dntp (Invitrogen), 1 ml of each 20-mM primer, 5 U Taq DNA polymerase (Invitrogen), and remainder H 2 O were denatured for 5 min at 94 C, followed by 50 cycles at 94 C for 30 s, 57 C for 30 s, and 72 C for 2 min, with a final extension at 72 C for 5 min. Digestion of this product for 2 hat37 C with 10 U HpyCH4IV (New England Biolabs) yields digestion products of 537, 460, 338, and 214 bp if an A (the rare variant) is present at rs ; the 460-bp product is digested to 292- and 168-bp products if a C is present at rs (the common variant, carried, in this case, on the novel allele). Two microliters of each reaction was run on a 1.5% agarose gel (10 mg ethidium bromide/30 ml) in 0.5# Tris-borate- EDTA buffer. Comparison of the result for DF5-65 with that for a control (DF5-72, his mother) who is also heterozygous for rs (fig. 6B) and whose DNA is identical across the target sequence, shows that the 292- and 168-bp products, derived from amplification of the novel allele, are significantly underrepresented in DF5-65 (fig. 6C). This is consistent with the interpretation that expression of GJB6 as well as that of GJB2 is diminished from the novel allele. Additionally, since the cdna specificity of each assay depends on the amplification of PCR product Figure 6. Allele-specific expression assay for GJB6. A, Schematic of design of HpyCH4IV digestion assay for allele-specific expression of GJB6 mrna performed using 1,550-bp PCR product amplified from cdna only. A forward primer (F) located in exon 5 of GJB6 is separated on genomic DNA from the reverse primer (R), located in the GJB6 3 UTR, by a 16-kb intron in addition to 1,500 bp of the final exon. B, Pedigree showing rs genotypes of assayed subjects. DF5-65 bears the novel pathogenic allele (black bar with star) and is heterozygous at rs C, Results of cdnaspecific assay for GJB6 expression. Digestion of product amplified from DF5-72 yields 460-bp as well as 292- and 168-bp products, indicating that both GJB6 alleles are expressed. Digestion of product amplified from DF5-65 yields a robust 460-bp band, but the band at 292 bp is only barely visible. This indicates underrepresentation of the novel allele among the amplified product. from only cdna from which a long intron has been properly spliced out, failure to accumulate PCR product might also be expected from an allele that is mutant for proper splicing. It is highly unlikely that GJB2 and GJB6 on this chromosome both contain unidentified splice mutations. This study provides evidence that the MSU-DF5 family is segregating a novel DFNB1 allele that is characterized by significant reduction in expression of both GJB2 and GJB6. This loss of expression is heritable, cis-acting, and not due to a simple parent-of-origin epigenetic modification. We sought and found no coding-region mutations, splice-site mutations, or any other sequence variant unique to the novel allele within or close to GJB2 or GJB6, suggesting that loss of function of an as-yet-unidentified cis-regulatory element(s) is responsible for our observations. It is possible that a locus-control region (LCR) regulates the coexpression of GJB2 and GJB6. An LCR was initially described in the b-globin locus, where it is absolutely required for expression of any of the genes in this cluster. Other examples include the growth hormone and T H 2 cytokine loci. 14 Such an element(s) may be located within the common interval deleted on chromosomes bearing del(gjb6-d13s1830) or del(gjb6-d13s1854). The severity of hearing loss in the individuals in our kindred who are heterozygous for 35delG and this novel allele is similar to that in individuals who are del(gjb6- D13S1830)/35delG, who as a group have been shown to have more severe hearing loss than 35delG homozy- 178 The American Journal of Human Genetics Volume 79 July
6 gotes. 15 Beyond the identification of the basal promoters of both genes, nothing is known of cis-acting elements that regulate GJB2 and GJB6. The common observation of an excess of deaf individuals bearing only a single identified GJB2 mutation strongly suggests that GJB2 regulatory mutations remain to be identified. Acknowledgments We thank the members of the MSU-DF5 family, for their generous participation, and Dr. Thomas Friedman, Dr. Robert Morell, and Dr. Patrick Venta, for helpful discussion and critical reading of the manuscript. This research was supported by the Michigan State University Foundation and by National Institute on Deafness and Other Communication Disorders grant DC (to K.H.F.). Web Resource The URL for data presented herein is as follows: Online Mendelian Inheritance in Man (OMIM), (for Vohwinkel syndrome, Bart-Pumphrey syndrome, PPK, KID, hidrotic ectodermal dysplasia, and Pendred syndrome) References 1. Forge A, Becker D, Casalotti S, Edwards J, Marziano N, Nevill G (2003) Gap junctions in the inner ear: comparison of distribution patterns in different vertebrates and assessment of connexin composition in mammals. J Comp Neurol 467: Forge A, Marziano N, Casalotti SO, Becker DL, Jagger D (2003) The inner ear contains heteromeric channels composed of cx26 and cx30 and deafness-related mutations in cx26 have a dominant negative effect on cx30. Cell Commun Adhes 10: Ahmad S, Chen S, Sun J, Lin X (2003) Connexins 26 and 30 are co-assembled to form gap junctions in the cochlea of mice. Biochem Biophys Res Commun 307: Kikuchi T, Kimura RS, Paul DL, Takasaka T, Adams JC (2000) Gap junction systems in the mammalian cochlea. Brain Res Rev 32: Wangemann P (2002) K cycling and the endocochlear potential. Hear Res 165: Grifa A, Wagner CA, D Ambrosio L, Melchionda S, Bernardi F, Lopez-Bigas N, Rabionet R, Arbones ML, Monica MD, Estivill X, Zelante L, Lang F, Gasparini P (1999) Mutations in GJB6 cause nonsyndromic autosomal dominant deafness at DFNA3 locus. Nat Genet 23: Lerer I, Sagi M, Ben-Neriah Z, Wang T, Levi H, Abeliovich D (2001) A deletion mutation in GJB6 cooperating with a GJB2 mutation in trans in non-syndromic deafness: a novel founder mutation in Ashkenazi Jews. Hum Mutat 18: del Castillo I, Villamar M, Moreno-Pelayo MA, del Castillo FJ, Alvarez A, Telleria D, Menendez I, Moreno F (2002) A deletion involving the connexin 30 gene in nonsyndromic hearing impairment. N Engl J Med 346: Pallares-Ruiz N, Blanchet P, Mondain M, Claustres M, Roux AF (2002) A large deletion including most of GJB6 in recessive non syndromic deafness: a digenic effect? Eur J Hum Genet 10: del Castillo FJ, Rodriguez-Ballesteros M, Alvarez A, Hutchin T, Leonardi E, de Oliveira CA, Azaiez H, et al (2005) A novel deletion involving the connexin-30 gene, del(gjb6- D13S1854), found in trans with mutations in the GJB2 gene (connexin-26) in subjects with DFNB1 non-syndromic hearing impairment. J Med Genet 42: del Castillo I, Moreno-Pelayo MA, del Castillo FJ, Brownstein Z, Marlin S, Adina Q, Cockburn DJ, et al (2003) Prevalence and evolutionary origins of the del(gjb6-d13s1830) mutation in the DFNB1 locus in hearing-impaired subjects: a multicenter study. Am J Hum Genet 73: Common JE, Bitner-Glindzicz M, O Toole EA, Barnes MR, Jenkins L, Forge A, Kelsell DP (2005) Specific loss of connexin 26 expression in ductal sweat gland epithelium associated with the deletion mutation del(gjb6-d13s1830). Clin Exp Dermatol 30: Wilcox SA, Osborn AH, Dahl HH (2000) A simple PCR test to detect the common 35delG mutation in the connexin 26 gene. Mol Diagn 5: Dean A (2006) On a chromosome far, far away: LCRs and gene expression. Trends Genet 22: Snoeckx RL, Huygen PLM, Feldmann D, Marlin S, Denoyelle F, Waligora J, Mueller-Malesinska M, et al (2005) GJB2 mutations and degree of hearing loss: a multicenter study. Am J Hum Genet 77: Tu ZJ, Kiang DT (1998) Mapping and characterization of the basal promoter of the human connexin26 gene. Biochim Biophys Acta 1443: Kiang DT, Jin N, Tu ZJ, Lin HH (1997) Upstream genomic sequence of the human connexin26 gene. Gene 199: Essenfelder GM, Larderet G, Waksman G, Lamartine J (2005) Gene structure and promoter analysis of the human GJB6 gene encoding connexin30. Gene 350: The American Journal of Human Genetics Volume 79 July
GJB2. Downloaded from jssu.ssu.ac.ir at 16:32 IRDT on Friday March 22nd delG. Direct Sequencing DHPLC . V153I, V27I, E114G, R127H
 6-708 GJB 8 7 6 5 * 0 Richard J.H. Smith 000 - :. GJB. 80 6. 0.. GJB 5delG. 0 0 : 5delG. ARMS-PCR 5delG Direct Sequencing DHPLC 67delT 5delG :. (). (%7/5) GJB :. V5I, V7I, EG, R7H.del del. GJB : 5delG.
6-708 GJB 8 7 6 5 * 0 Richard J.H. Smith 000 - :. GJB. 80 6. 0.. GJB 5delG. 0 0 : 5delG. ARMS-PCR 5delG Direct Sequencing DHPLC 67delT 5delG :. (). (%7/5) GJB :. V5I, V7I, EG, R7H.del del. GJB : 5delG.
Cell & Molecular Biology, Islamic Azad University, Marand Branch, Iran Marand, Iran. ABSTRACT
 1 Human and Animal Health Vol. 59: e16160046, January-December 2016 http://dx.doi.org/10.1590/1678-4324-2016150046 ISSN 1678-4324 Online Edition BRAZILIAN ARCHIVES OF BIOLOGY AND TECHNOLOGY A N I N T E
1 Human and Animal Health Vol. 59: e16160046, January-December 2016 http://dx.doi.org/10.1590/1678-4324-2016150046 ISSN 1678-4324 Online Edition BRAZILIAN ARCHIVES OF BIOLOGY AND TECHNOLOGY A N I N T E
Investigating Seven Recently Identified Genes in 100 Iranian Families with Autosomal Recessive Non-syndromic Hearing Loss
 Iranian Rehabilitation Journal, Vol. 13, Issue 3, Autumn 2015 Original Article Investigating Seven Recently Identified Genes in 100 Iranian Families with Autosomal Recessive Non-syndromic Hearing Loss
Iranian Rehabilitation Journal, Vol. 13, Issue 3, Autumn 2015 Original Article Investigating Seven Recently Identified Genes in 100 Iranian Families with Autosomal Recessive Non-syndromic Hearing Loss
The Turkish Journal of Pediatrics 2005; 47:
 The Turkish Journal of Pediatrics 2005; 47: 213-221 Original Identification of an ancestral haplotype of the 35delG mutation in the GJB2 (connexin 26) gene responsible for autosomal recessive non-syndromic
The Turkish Journal of Pediatrics 2005; 47: 213-221 Original Identification of an ancestral haplotype of the 35delG mutation in the GJB2 (connexin 26) gene responsible for autosomal recessive non-syndromic
Prevalence of the connexin 26 mutation 35delG in nonsyndromic hearing loss in Egypt
 Prevalence of the connexin 26 mutation 35delG in nonsyndromic hearing loss in Egypt M. W. M. Mustafa Audiology Unit, Sohag University Hospitals, Sohag 82524, Egypt. Correspondence to: Dr. Mohamed Wael
Prevalence of the connexin 26 mutation 35delG in nonsyndromic hearing loss in Egypt M. W. M. Mustafa Audiology Unit, Sohag University Hospitals, Sohag 82524, Egypt. Correspondence to: Dr. Mohamed Wael
STUDY OF RECESSIVE DEAFNESS LOCUS (DFNB1) BY LINKAGE ANALYSIS IN SOME FAMILIES FROM BALOCHISTAN
 STUDY OF RECESSIVE DEAFNESS LOCUS (DFNB1) BY LINKAGE ANALYSIS IN SOME FAMILIES FROM BALOCHISTAN A synopsis submitted to BALOCHISTAN UNIVERSITY OF INFORMATION TECHNOLOGY ENGINEERING & MANAGEMENT SCIENCES
STUDY OF RECESSIVE DEAFNESS LOCUS (DFNB1) BY LINKAGE ANALYSIS IN SOME FAMILIES FROM BALOCHISTAN A synopsis submitted to BALOCHISTAN UNIVERSITY OF INFORMATION TECHNOLOGY ENGINEERING & MANAGEMENT SCIENCES
The Association Between GJB2 Mutation and GJB6 Gene in Non Syndromic Hearing Loss School Children
 ORIGINAL ARTICLE The Association Between GJB2 Mutation and GJB6 Gene in Non Syndromic Hearing Loss School Children A Asma*, A Ashwaq**, A G Norzana****, A Maizaton Atmadini*****, B H I Ruszymah******,
ORIGINAL ARTICLE The Association Between GJB2 Mutation and GJB6 Gene in Non Syndromic Hearing Loss School Children A Asma*, A Ashwaq**, A G Norzana****, A Maizaton Atmadini*****, B H I Ruszymah******,
article MATERIALS AND METHODS
 May/June 2003 Vol. 5 No. 3 article Mutation spectrum of the connexin 26 (GJB2) gene in Taiwanese patients with prelingual deafness Hsiao-Lin Hwa, MD 1, Tsang-Ming Ko, MD, PhD 1,2, Chuan-Jen Hsu, MD 3,
May/June 2003 Vol. 5 No. 3 article Mutation spectrum of the connexin 26 (GJB2) gene in Taiwanese patients with prelingual deafness Hsiao-Lin Hwa, MD 1, Tsang-Ming Ko, MD, PhD 1,2, Chuan-Jen Hsu, MD 3,
Case Report Two Portuguese Cochlear Implanted Dizygotic Twins: ACaseReport
 Case Reports in Genetics Volume, Article ID 686, 5 pages doi:.55//686 Case Report Two Portuguese Cochlear Implanted Dizygotic Twins: ACaseReport JoanaRitaChora, Helena Simões-Teixeira, Tiago Daniel Matos,
Case Reports in Genetics Volume, Article ID 686, 5 pages doi:.55//686 Case Report Two Portuguese Cochlear Implanted Dizygotic Twins: ACaseReport JoanaRitaChora, Helena Simões-Teixeira, Tiago Daniel Matos,
Corporate Medical Policy
 Corporate Medical Policy Genetic Testing for Hereditary Hearing Loss File Name: Origination: Last CAP Review: Next CAP Review: Last Review: genetic_testing_for_hereditary_hearing_loss 10/2013 7/2018 7/2019
Corporate Medical Policy Genetic Testing for Hereditary Hearing Loss File Name: Origination: Last CAP Review: Next CAP Review: Last Review: genetic_testing_for_hereditary_hearing_loss 10/2013 7/2018 7/2019
ABSTRACT Background Inherited hearing impairment affects about 1 in 2000 newborns. Up to 50 percent of all patients
 DELETION INVOLVING THE CONNEXIN 30 GENE IN NONSYNDROMIC HERING IMPIRMENT IGNCIO DEL CSTILLO, PH.D., MNUEL VILLMR, PH.D., MIGUEL. MORENO-PELYO, PH.D., FRNCISCO J. DEL CSTILLO, PH.D., RCELI ÁLVREZ, M.SC.,
DELETION INVOLVING THE CONNEXIN 30 GENE IN NONSYNDROMIC HERING IMPIRMENT IGNCIO DEL CSTILLO, PH.D., MNUEL VILLMR, PH.D., MIGUEL. MORENO-PELYO, PH.D., FRNCISCO J. DEL CSTILLO, PH.D., RCELI ÁLVREZ, M.SC.,
Prevalence of Connexin 26 Mutations in Patients from Jordan with Non Syndromic Hearing Loss
 Kamla-Raj 2006 Int J Hum Genet, 6(2): 119-124 (2006) Prevalence of Connexin 26 Mutations in Patients from Jordan with Non Syndromic Hearing Loss A. A. Mahasneh* and R. M. Battah Department of Biotechnology
Kamla-Raj 2006 Int J Hum Genet, 6(2): 119-124 (2006) Prevalence of Connexin 26 Mutations in Patients from Jordan with Non Syndromic Hearing Loss A. A. Mahasneh* and R. M. Battah Department of Biotechnology
Spectrum of GJB2 mutations in Cypriot nonsyndromic hearing loss subjects
 c Indian Academy of Sciences RESEARCH NOTE Spectrum of GJB2 mutations in Cypriot nonsyndromic hearing loss subjects VASSOS NEOCLEOUS 1, CONSTANTINA COSTI 1, CHRISTOS SHAMMAS 1, ELENA SPANOU 2,3, VIOLETTA
c Indian Academy of Sciences RESEARCH NOTE Spectrum of GJB2 mutations in Cypriot nonsyndromic hearing loss subjects VASSOS NEOCLEOUS 1, CONSTANTINA COSTI 1, CHRISTOS SHAMMAS 1, ELENA SPANOU 2,3, VIOLETTA
CONGENITAL sensorineural. Connexin 26 Studies in Patients With Sensorineural Hearing Loss ORIGINAL ARTICLE
 ORIGINAL ARTICLE Connexin 26 Studies in Patients With Sensorineural Hearing Loss Margaret A. Kenna, MD; Bai-Lin Wu, PhD; Douglas A. Cotanche, PhD; Bruce R. Korf, MD, PhD; Heidi L. Rehm, PhD Objective:
ORIGINAL ARTICLE Connexin 26 Studies in Patients With Sensorineural Hearing Loss Margaret A. Kenna, MD; Bai-Lin Wu, PhD; Douglas A. Cotanche, PhD; Bruce R. Korf, MD, PhD; Heidi L. Rehm, PhD Objective:
Introduction. IAPA: June 04 1
 Introduction Conflicting views on the prevalence and nature of otoacoustic emission [OAE] abnormalities in ARNSHL families (Morell et al, 1998; Cohn & Kelley, 1999). Detailed study of OAEs in greater number
Introduction Conflicting views on the prevalence and nature of otoacoustic emission [OAE] abnormalities in ARNSHL families (Morell et al, 1998; Cohn & Kelley, 1999). Detailed study of OAEs in greater number
Pedigree Construction Notes
 Name Date Pedigree Construction Notes GO TO à Mendelian Inheritance (http://www.uic.edu/classes/bms/bms655/lesson3.html) When human geneticists first began to publish family studies, they used a variety
Name Date Pedigree Construction Notes GO TO à Mendelian Inheritance (http://www.uic.edu/classes/bms/bms655/lesson3.html) When human geneticists first began to publish family studies, they used a variety
Role of Paired Box9 (PAX9) (rs ) and Muscle Segment Homeobox1 (MSX1) (581C>T) Gene Polymorphisms in Tooth Agenesis
 EC Dental Science Special Issue - 2017 Role of Paired Box9 (PAX9) (rs2073245) and Muscle Segment Homeobox1 (MSX1) (581C>T) Gene Polymorphisms in Tooth Agenesis Research Article Dr. Sonam Sethi 1, Dr. Anmol
EC Dental Science Special Issue - 2017 Role of Paired Box9 (PAX9) (rs2073245) and Muscle Segment Homeobox1 (MSX1) (581C>T) Gene Polymorphisms in Tooth Agenesis Research Article Dr. Sonam Sethi 1, Dr. Anmol
Proposal form for the evaluation of a genetic test for NHS Service Gene Dossier/Additional Provider
 Proposal form for the evaluation of a genetic test for NHS Service Gene Dossier/Additional Provider TEST DISEASE/CONDITION POPULATION TRIAD Submitting laboratory: London North East RGC GOSH Approved: September
Proposal form for the evaluation of a genetic test for NHS Service Gene Dossier/Additional Provider TEST DISEASE/CONDITION POPULATION TRIAD Submitting laboratory: London North East RGC GOSH Approved: September
2. stereocilia make contact with membrane, feel vibration. Tiplink is deflected, allows ions to go inside cell body and chemical signal is generated.
 Hearing Loss 1. Most common sensory deficit in human 2. 3 in ten people over age 60 have hearing loss 3. At least 1.4 million children have hearing problems 4. Estimated that 3 in 1,000 infants are born
Hearing Loss 1. Most common sensory deficit in human 2. 3 in ten people over age 60 have hearing loss 3. At least 1.4 million children have hearing problems 4. Estimated that 3 in 1,000 infants are born
Genome 371, Autumn 2018 Quiz Section 9: Genetics of Cancer Worksheet
 Genome 371, Autumn 2018 Quiz Section 9: Genetics of Cancer Worksheet All cancer is due to genetic mutations. However, in cancer that clusters in families (familial cancer) at least one of these mutations
Genome 371, Autumn 2018 Quiz Section 9: Genetics of Cancer Worksheet All cancer is due to genetic mutations. However, in cancer that clusters in families (familial cancer) at least one of these mutations
H earing impairment is a common and highly heterogeneous
 588 LETTER TO JMG A novel deletion involving the connexin-30 gene, del(gjb6- d13s1854), found in trans with mutations in the GJB2 gene (connexin-26) in subjects with DFNB1 non-syndromic hearing impairment
588 LETTER TO JMG A novel deletion involving the connexin-30 gene, del(gjb6- d13s1854), found in trans with mutations in the GJB2 gene (connexin-26) in subjects with DFNB1 non-syndromic hearing impairment
Prevalence of Hearing Impairment
 Prevalence of Hearing Impairment 28 million Americans 2 million profoundly deaf 1/1000 congenitally deaf 1/3 impaired by age 65 1/2 impaired by age 80 NIDCD National Strategic Research Plan, 1989 Genetic
Prevalence of Hearing Impairment 28 million Americans 2 million profoundly deaf 1/1000 congenitally deaf 1/3 impaired by age 65 1/2 impaired by age 80 NIDCD National Strategic Research Plan, 1989 Genetic
SALSA MLPA KIT P050-B2 CAH
 SALSA MLPA KIT P050-B2 CAH Lot 0510, 0909, 0408: Compared to lot 0107, extra control fragments have been added at 88, 96, 100 and 105 nt. The 274 nt probe gives a higher signal in lot 0510 compared to
SALSA MLPA KIT P050-B2 CAH Lot 0510, 0909, 0408: Compared to lot 0107, extra control fragments have been added at 88, 96, 100 and 105 nt. The 274 nt probe gives a higher signal in lot 0510 compared to
SALSA MLPA KIT P060-B2 SMA
 SALSA MLPA KIT P6-B2 SMA Lot 111, 511: As compared to the previous version B1 (lot 11), the 88 and 96 nt DNA Denaturation control fragments have been replaced (QDX2). Please note that, in contrast to the
SALSA MLPA KIT P6-B2 SMA Lot 111, 511: As compared to the previous version B1 (lot 11), the 88 and 96 nt DNA Denaturation control fragments have been replaced (QDX2). Please note that, in contrast to the
IVF Michigan, Rochester Hills, Michigan, and Reproductive Genetics Institute, Chicago, Illinois
 FERTILITY AND STERILITY VOL. 80, NO. 4, OCTOBER 2003 Copyright 2003 American Society for Reproductive Medicine Published by Elsevier Inc. Printed on acid-free paper in U.S.A. CASE REPORTS Preimplantation
FERTILITY AND STERILITY VOL. 80, NO. 4, OCTOBER 2003 Copyright 2003 American Society for Reproductive Medicine Published by Elsevier Inc. Printed on acid-free paper in U.S.A. CASE REPORTS Preimplantation
Genetics of Hearing Loss
 Genetics of Hearing Loss Daryl A. Scott MD/PhD Molecular & Human Genetics 1/20/2015 Why do we care? 1 100% 75% Hearing Loss 500:1000 50% 314:1000 25% 1:1000 17:1000 Newborn 18 yrs 65 yrs 75 yrs 60% Members
Genetics of Hearing Loss Daryl A. Scott MD/PhD Molecular & Human Genetics 1/20/2015 Why do we care? 1 100% 75% Hearing Loss 500:1000 50% 314:1000 25% 1:1000 17:1000 Newborn 18 yrs 65 yrs 75 yrs 60% Members
A gene is a sequence of DNA that resides at a particular site on a chromosome the locus (plural loci). Genetic linkage of genes on a single
 8.3 A gene is a sequence of DNA that resides at a particular site on a chromosome the locus (plural loci). Genetic linkage of genes on a single chromosome can alter their pattern of inheritance from those
8.3 A gene is a sequence of DNA that resides at a particular site on a chromosome the locus (plural loci). Genetic linkage of genes on a single chromosome can alter their pattern of inheritance from those
Prevalence of Cx26 (GJB2) Gene Mutations Causing Recessive Nonsyndromic Hearing Impairment in India
 Kamla-Raj 2005 Int J Hum Genet, 5(4): 241-246 (2005) Prevalence of Cx26 (GJB2) Gene Mutations Causing Recessive Nonsyndromic Hearing Impairment in India P.V. Ramchander 1, V.U. Nandur 2, K. Dwarakanath
Kamla-Raj 2005 Int J Hum Genet, 5(4): 241-246 (2005) Prevalence of Cx26 (GJB2) Gene Mutations Causing Recessive Nonsyndromic Hearing Impairment in India P.V. Ramchander 1, V.U. Nandur 2, K. Dwarakanath
Dan Koller, Ph.D. Medical and Molecular Genetics
 Design of Genetic Studies Dan Koller, Ph.D. Research Assistant Professor Medical and Molecular Genetics Genetics and Medicine Over the past decade, advances from genetics have permeated medicine Identification
Design of Genetic Studies Dan Koller, Ph.D. Research Assistant Professor Medical and Molecular Genetics Genetics and Medicine Over the past decade, advances from genetics have permeated medicine Identification
MRC-Holland MLPA. Description version 19;
 SALSA MLPA probemix P6-B2 SMA Lot B2-712, B2-312, B2-111, B2-511: As compared to the previous version B1 (lot B1-11), the 88 and 96 nt DNA Denaturation control fragments have been replaced (QDX2). SPINAL
SALSA MLPA probemix P6-B2 SMA Lot B2-712, B2-312, B2-111, B2-511: As compared to the previous version B1 (lot B1-11), the 88 and 96 nt DNA Denaturation control fragments have been replaced (QDX2). SPINAL
Clinical evidence of the nonpathogenic nature of the M34T variant in the connexin 26 gene
 (), & Nature Publishing Group All rights reserved -/ $. www.nature.com/ejhg ARTICLE Clinical evidence of the nonpathogenic nature of the MT variant in the connexin gene Delphine Feldmann, Franc oise Denoyelle,
(), & Nature Publishing Group All rights reserved -/ $. www.nature.com/ejhg ARTICLE Clinical evidence of the nonpathogenic nature of the MT variant in the connexin gene Delphine Feldmann, Franc oise Denoyelle,
ORIGINAL ARTICLE. Connexin 26 Gene Mutations in Congenitally Deaf Children
 Connexin 26 Gene Mutations in Congenitally Deaf Children Pitfalls for Genetic Counseling ORIGINAL ARTICLE Sandrine Marlin, MD, PhD; Éréa-Noël Garabédian, MD; Gilles Roger, MD; Lucien Moatti, MD; Nicole
Connexin 26 Gene Mutations in Congenitally Deaf Children Pitfalls for Genetic Counseling ORIGINAL ARTICLE Sandrine Marlin, MD, PhD; Éréa-Noël Garabédian, MD; Gilles Roger, MD; Lucien Moatti, MD; Nicole
Abstract. Introduction. RBMOnline - Vol 8. No Reproductive BioMedicine Online; on web 10 December 2003
 RBMOnline - Vol 8. No 2. 224-228 Reproductive BioMedicine Online; www.rbmonline.com/article/1133 on web 10 December 2003 Article Preimplantation genetic diagnosis for early-onset torsion dystonia Dr Svetlana
RBMOnline - Vol 8. No 2. 224-228 Reproductive BioMedicine Online; www.rbmonline.com/article/1133 on web 10 December 2003 Article Preimplantation genetic diagnosis for early-onset torsion dystonia Dr Svetlana
Genetic stories behind village sign languages
 Genetic stories behind village sign languages the co-evolution of deafness with sign language June, 2013 Minerva-Gentner Symposium on Emergent Languages and Cultural Evolution Berg en Dal, The Netherlands
Genetic stories behind village sign languages the co-evolution of deafness with sign language June, 2013 Minerva-Gentner Symposium on Emergent Languages and Cultural Evolution Berg en Dal, The Netherlands
Computational Systems Biology: Biology X
 Bud Mishra Room 1002, 715 Broadway, Courant Institute, NYU, New York, USA L#4:(October-0-4-2010) Cancer and Signals 1 2 1 2 Evidence in Favor Somatic mutations, Aneuploidy, Copy-number changes and LOH
Bud Mishra Room 1002, 715 Broadway, Courant Institute, NYU, New York, USA L#4:(October-0-4-2010) Cancer and Signals 1 2 1 2 Evidence in Favor Somatic mutations, Aneuploidy, Copy-number changes and LOH
recessive mode of action. Molecular Medicine Unit, St James s University Hospital, Leeds LS9 7TF, UK M J Houseman
 20 Molecular Medicine Unit, St James s University Hospital, Leeds LS9 7TF, UK M J Houseman Yorkshire Regional DNA Laboratory, St James s University Hospital, Leeds LS9 7TF, UK L A Ellis G R Taylor Clinical
20 Molecular Medicine Unit, St James s University Hospital, Leeds LS9 7TF, UK M J Houseman Yorkshire Regional DNA Laboratory, St James s University Hospital, Leeds LS9 7TF, UK L A Ellis G R Taylor Clinical
GJB2 MUTATIONS IN NON SYNDROMIC HEARING LOSS IN THE REPUBLIC OF MACEDONIA
 BJMG 12/2 (2009) 11-16 10.2478/v10034-010-0004-x ORIGINAL ARTICLE GJB2 MUTATIONS IN NON SYNDROMIC HEARING LOSS IN THE REPUBLIC OF MACEDONIA Sukarova Stefanovska E 1, Momirovska, A 2,3, Cakar M 4, Efremov
BJMG 12/2 (2009) 11-16 10.2478/v10034-010-0004-x ORIGINAL ARTICLE GJB2 MUTATIONS IN NON SYNDROMIC HEARING LOSS IN THE REPUBLIC OF MACEDONIA Sukarova Stefanovska E 1, Momirovska, A 2,3, Cakar M 4, Efremov
The New England Journal of Medicine
 MUTATIONS IN THE CONNEXIN 6 GENE (GJB) AMONG ASHKENAZI JEWS WITH NONSYNDROMIC RECESSIVE DEAFNESS ROBERT J. MORELL, PH.D., HUNG JEFF KIM, M.D., LINDA J. HOOD, PH.D., LEAH GOFORTH, M.S., KAREN FRIDERICI,
MUTATIONS IN THE CONNEXIN 6 GENE (GJB) AMONG ASHKENAZI JEWS WITH NONSYNDROMIC RECESSIVE DEAFNESS ROBERT J. MORELL, PH.D., HUNG JEFF KIM, M.D., LINDA J. HOOD, PH.D., LEAH GOFORTH, M.S., KAREN FRIDERICI,
/07/ PEDIATRIC RESEARCH Vol. 61, No. 6, 2007 Copyright 2007 International Pediatric Research Foundation, Inc.
 0031-3998/07/6106-0687 PEDIATRIC RESEARCH Vol. 61, No. 6, 2007 Copyright 2007 International Pediatric Research Foundation, Inc. Printed in U.S.A. GJB2 and GJB6 Mutations in Children with Congenital Cytomegalovirus
0031-3998/07/6106-0687 PEDIATRIC RESEARCH Vol. 61, No. 6, 2007 Copyright 2007 International Pediatric Research Foundation, Inc. Printed in U.S.A. GJB2 and GJB6 Mutations in Children with Congenital Cytomegalovirus
Single Gene (Monogenic) Disorders. Mendelian Inheritance: Definitions. Mendelian Inheritance: Definitions
 Single Gene (Monogenic) Disorders Mendelian Inheritance: Definitions A genetic locus is a specific position or location on a chromosome. Frequently, locus is used to refer to a specific gene. Alleles are
Single Gene (Monogenic) Disorders Mendelian Inheritance: Definitions A genetic locus is a specific position or location on a chromosome. Frequently, locus is used to refer to a specific gene. Alleles are
Genome - Wide Linkage Mapping
 Biological Sciences Initiative HHMI Genome - Wide Linkage Mapping Introduction This activity is based on the work of Dr. Christine Seidman et al that was published in Circulation, 1998, vol 97, pgs 2043-2048.
Biological Sciences Initiative HHMI Genome - Wide Linkage Mapping Introduction This activity is based on the work of Dr. Christine Seidman et al that was published in Circulation, 1998, vol 97, pgs 2043-2048.
Sensorineural hearing loss and the incidence of Cx26 mutations in Austria
 (2001) 9, 226 ± 230 ã 2001 Nature Publishing Group All rights reserved 1018-4813/01 $15.00 www.nature.com/ejhg SHORT REPORT Sensorineural hearing loss and the incidence of Cx26 mutations in Austria Judith
(2001) 9, 226 ± 230 ã 2001 Nature Publishing Group All rights reserved 1018-4813/01 $15.00 www.nature.com/ejhg SHORT REPORT Sensorineural hearing loss and the incidence of Cx26 mutations in Austria Judith
Non-syndromic, autosomal-recessive deafness
 Clin Genet 2006: 69: 371 392 Printed in Singapore. All rights reserved Review Non-syndromic, autosomal-recessive deafness # 2006 The Authors Journal compilation # 2006BlackwellMunksgaard CLINICAL GENETICS
Clin Genet 2006: 69: 371 392 Printed in Singapore. All rights reserved Review Non-syndromic, autosomal-recessive deafness # 2006 The Authors Journal compilation # 2006BlackwellMunksgaard CLINICAL GENETICS
Problem set questions from Final Exam Human Genetics, Nondisjunction, and Cancer
 Problem set questions from Final Exam Human Genetics, Nondisjunction, and ancer Mapping in humans using SSRs and LOD scores 1. You set out to genetically map the locus for color blindness with respect
Problem set questions from Final Exam Human Genetics, Nondisjunction, and ancer Mapping in humans using SSRs and LOD scores 1. You set out to genetically map the locus for color blindness with respect
Pedigree Analysis Why do Pedigrees? Goals of Pedigree Analysis Basic Symbols More Symbols Y-Linked Inheritance
 Pedigree Analysis Why do Pedigrees? Punnett squares and chi-square tests work well for organisms that have large numbers of offspring and controlled mating, but humans are quite different: Small families.
Pedigree Analysis Why do Pedigrees? Punnett squares and chi-square tests work well for organisms that have large numbers of offspring and controlled mating, but humans are quite different: Small families.
DOES THE BRCAX GENE EXIST? FUTURE OUTLOOK
 CHAPTER 6 DOES THE BRCAX GENE EXIST? FUTURE OUTLOOK Genetic research aimed at the identification of new breast cancer susceptibility genes is at an interesting crossroad. On the one hand, the existence
CHAPTER 6 DOES THE BRCAX GENE EXIST? FUTURE OUTLOOK Genetic research aimed at the identification of new breast cancer susceptibility genes is at an interesting crossroad. On the one hand, the existence
article Genetics IN Medicine 517
 July 2008 Vol. 10 No. 7 article Infant hearing loss and connexin testing in a diverse population Lisa A. Schimmenti, MD 1, Ariadna Martinez, MS, MS 2, Milhan Telatar, PhD 3, Chih-Hung Lai, PhD 3, Nina
July 2008 Vol. 10 No. 7 article Infant hearing loss and connexin testing in a diverse population Lisa A. Schimmenti, MD 1, Ariadna Martinez, MS, MS 2, Milhan Telatar, PhD 3, Chih-Hung Lai, PhD 3, Nina
HST.161 Molecular Biology and Genetics in Modern Medicine Fall 2007
 MIT OpenCourseWare http://ocw.mit.edu HST.161 Molecular Biology and Genetics in Modern Medicine Fall 2007 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
MIT OpenCourseWare http://ocw.mit.edu HST.161 Molecular Biology and Genetics in Modern Medicine Fall 2007 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
Lecture 17: Human Genetics. I. Types of Genetic Disorders. A. Single gene disorders
 Lecture 17: Human Genetics I. Types of Genetic Disorders A. Single gene disorders B. Multifactorial traits 1. Mutant alleles at several loci acting in concert C. Chromosomal abnormalities 1. Physical changes
Lecture 17: Human Genetics I. Types of Genetic Disorders A. Single gene disorders B. Multifactorial traits 1. Mutant alleles at several loci acting in concert C. Chromosomal abnormalities 1. Physical changes
IVS8+1 DelG, a Novel Splice Site Mutation Causing DFNA5 Deafness in a Chinese Family
 Original Article IVS8+1 DelG, a Novel Splice Site Mutation Causing DFNA5 Deafness in a Chinese Family Mei Na Li Yang 1, Xiao Fei Shen 1, Qin Jun Wei 2, Jun Yao 2, Ya Jie Lu 2, Xin Cao 2, Guang Qian Xing
Original Article IVS8+1 DelG, a Novel Splice Site Mutation Causing DFNA5 Deafness in a Chinese Family Mei Na Li Yang 1, Xiao Fei Shen 1, Qin Jun Wei 2, Jun Yao 2, Ya Jie Lu 2, Xin Cao 2, Guang Qian Xing
CANCER GENETICS PROVIDER SURVEY
 Dear Participant, Previously you agreed to participate in an evaluation of an education program we developed for primary care providers on the topic of cancer genetics. This is an IRB-approved, CDCfunded
Dear Participant, Previously you agreed to participate in an evaluation of an education program we developed for primary care providers on the topic of cancer genetics. This is an IRB-approved, CDCfunded
BIOL2005 WORKSHEET 2008
 BIOL2005 WORKSHEET 2008 Answer all 6 questions in the space provided using additional sheets where necessary. Hand your completed answers in to the Biology office by 3 p.m. Friday 8th February. 1. Your
BIOL2005 WORKSHEET 2008 Answer all 6 questions in the space provided using additional sheets where necessary. Hand your completed answers in to the Biology office by 3 p.m. Friday 8th February. 1. Your
Lack of Association between Endoplasmic Reticulum Stress Response Genes and Suicidal Victims
 Kobe J. Med. Sci., Vol. 53, No. 4, pp. 151-155, 2007 Lack of Association between Endoplasmic Reticulum Stress Response Genes and Suicidal Victims KAORU SAKURAI 1, NAOKI NISHIGUCHI 2, OSAMU SHIRAKAWA 2,
Kobe J. Med. Sci., Vol. 53, No. 4, pp. 151-155, 2007 Lack of Association between Endoplasmic Reticulum Stress Response Genes and Suicidal Victims KAORU SAKURAI 1, NAOKI NISHIGUCHI 2, OSAMU SHIRAKAWA 2,
A. Incorrect! Cells contain the units of genetic they are not the unit of heredity.
 MCAT Biology Problem Drill PS07: Mendelian Genetics Question No. 1 of 10 Question 1. The smallest unit of heredity is. Question #01 (A) Cell (B) Gene (C) Chromosome (D) Allele Cells contain the units of
MCAT Biology Problem Drill PS07: Mendelian Genetics Question No. 1 of 10 Question 1. The smallest unit of heredity is. Question #01 (A) Cell (B) Gene (C) Chromosome (D) Allele Cells contain the units of
Systems of Mating: Systems of Mating:
 8/29/2 Systems of Mating: the rules by which pairs of gametes are chosen from the local gene pool to be united in a zygote with respect to a particular locus or genetic system. Systems of Mating: A deme
8/29/2 Systems of Mating: the rules by which pairs of gametes are chosen from the local gene pool to be united in a zygote with respect to a particular locus or genetic system. Systems of Mating: A deme
What favorite organism of geneticists is described in the right-hand column?
 What favorite organism of geneticists is described in the right-hand column? Model Organism fruit fly?? Generation time 12 days ~ 5000 days Size 2 mm 1500-1800mm Brood size hundreds a couple dozen would
What favorite organism of geneticists is described in the right-hand column? Model Organism fruit fly?? Generation time 12 days ~ 5000 days Size 2 mm 1500-1800mm Brood size hundreds a couple dozen would
Mendelian & Complex Traits. Quantitative Imaging Genomics. Genetics Terminology 2. Genetics Terminology 1. Human Genome. Genetics Terminology 3
 Mendelian & Complex Traits Quantitative Imaging Genomics David C. Glahn, PhD Olin Neuropsychiatry Research Center & Department of Psychiatry, Yale University July, 010 Mendelian Trait A trait influenced
Mendelian & Complex Traits Quantitative Imaging Genomics David C. Glahn, PhD Olin Neuropsychiatry Research Center & Department of Psychiatry, Yale University July, 010 Mendelian Trait A trait influenced
impairment Corresponding author: Prof Igor Medica, MD PhD, Outpatient Paediatric Clinic, 13
 GJB2 and GJB6 mutations in Croatians with prelingual non-syndromic hearing impairment Igor Medica 1,2, Gorazd Rudolf 1, Manuela Balaban 1,2, Borut Peterlin 1 1 Division of Medical Genetics, Department
GJB2 and GJB6 mutations in Croatians with prelingual non-syndromic hearing impairment Igor Medica 1,2, Gorazd Rudolf 1, Manuela Balaban 1,2, Borut Peterlin 1 1 Division of Medical Genetics, Department
Ch 8 Practice Questions
 Ch 8 Practice Questions Multiple Choice Identify the choice that best completes the statement or answers the question. 1. What fraction of offspring of the cross Aa Aa is homozygous for the dominant allele?
Ch 8 Practice Questions Multiple Choice Identify the choice that best completes the statement or answers the question. 1. What fraction of offspring of the cross Aa Aa is homozygous for the dominant allele?
Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells.
 SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
SUPPLEMENTAL FIGURE AND TABLE LEGENDS Supplemental Figure S1. Expression of Cirbp mrna in mouse tissues and NIH3T3 cells. A) Cirbp mrna expression levels in various mouse tissues collected around the clock
MODULE NO.14: Y-Chromosome Testing
 SUBJECT Paper No. and Title Module No. and Title Module Tag FORENSIC SIENCE PAPER No.13: DNA Forensics MODULE No.21: Y-Chromosome Testing FSC_P13_M21 TABLE OF CONTENTS 1. Learning Outcome 2. Introduction:
SUBJECT Paper No. and Title Module No. and Title Module Tag FORENSIC SIENCE PAPER No.13: DNA Forensics MODULE No.21: Y-Chromosome Testing FSC_P13_M21 TABLE OF CONTENTS 1. Learning Outcome 2. Introduction:
MRC-Holland MLPA. Description version 29;
 SALSA MLPA KIT P003-B1 MLH1/MSH2 Lot 1209, 0109. As compared to the previous lots 0307 and 1006, one MLH1 probe (exon 19) and four MSH2 probes have been replaced. In addition, one extra MSH2 exon 1 probe,
SALSA MLPA KIT P003-B1 MLH1/MSH2 Lot 1209, 0109. As compared to the previous lots 0307 and 1006, one MLH1 probe (exon 19) and four MSH2 probes have been replaced. In addition, one extra MSH2 exon 1 probe,
SALSA MLPA probemix P185-C2 Intersex Lot C2-1015: As compared to the previous version C1 (lot C1-0611), the lengths of four probes have been adjusted.
 mix P185-C2 Intersex Lot C2-1015: As compared to the previous version C1 (lot C1-0611), the lengths of four s have been adjusted. The sex-determining region on chromosome Y (SRY) is the most important
mix P185-C2 Intersex Lot C2-1015: As compared to the previous version C1 (lot C1-0611), the lengths of four s have been adjusted. The sex-determining region on chromosome Y (SRY) is the most important
LTA Analysis of HapMap Genotype Data
 LTA Analysis of HapMap Genotype Data Introduction. This supplement to Global variation in copy number in the human genome, by Redon et al., describes the details of the LTA analysis used to screen HapMap
LTA Analysis of HapMap Genotype Data Introduction. This supplement to Global variation in copy number in the human genome, by Redon et al., describes the details of the LTA analysis used to screen HapMap
SNP Array NOTE: THIS IS A SAMPLE REPORT AND MAY NOT REFLECT ACTUAL PATIENT DATA. FORMAT AND/OR CONTENT MAY BE UPDATED PERIODICALLY.
 SAMPLE REPORT SNP Array NOTE: THIS IS A SAMPLE REPORT AND MAY NOT REFLECT ACTUAL PATIENT DATA. FORMAT AND/OR CONTENT MAY BE UPDATED PERIODICALLY. RESULTS SNP Array Copy Number Variations Result: GAIN,
SAMPLE REPORT SNP Array NOTE: THIS IS A SAMPLE REPORT AND MAY NOT REFLECT ACTUAL PATIENT DATA. FORMAT AND/OR CONTENT MAY BE UPDATED PERIODICALLY. RESULTS SNP Array Copy Number Variations Result: GAIN,
SEX. Genetic Variation: The genetic substrate for natural selection. Sex: Sources of Genotypic Variation. Genetic Variation
 Genetic Variation: The genetic substrate for natural selection Sex: Sources of Genotypic Variation Dr. Carol E. Lee, University of Wisconsin Genetic Variation If there is no genetic variation, neither
Genetic Variation: The genetic substrate for natural selection Sex: Sources of Genotypic Variation Dr. Carol E. Lee, University of Wisconsin Genetic Variation If there is no genetic variation, neither
Aim: To develop a screening in order to determine
 Rev Bras Otorrinolaringol 2007;73(3):412-7. REVIEW ARTICLE Diagnosis routine and approach in genetic sensorineural hearing loss Fatima Regina Abreu Alves 1, Fernando de Andrade Quintanilha Ribeiro 2 Keywords:
Rev Bras Otorrinolaringol 2007;73(3):412-7. REVIEW ARTICLE Diagnosis routine and approach in genetic sensorineural hearing loss Fatima Regina Abreu Alves 1, Fernando de Andrade Quintanilha Ribeiro 2 Keywords:
Genetics and Genomics in Medicine Chapter 8 Questions
 Genetics and Genomics in Medicine Chapter 8 Questions Linkage Analysis Question Question 8.1 Affected members of the pedigree above have an autosomal dominant disorder, and cytogenetic analyses using conventional
Genetics and Genomics in Medicine Chapter 8 Questions Linkage Analysis Question Question 8.1 Affected members of the pedigree above have an autosomal dominant disorder, and cytogenetic analyses using conventional
Name: PS#: Biol 3301 Midterm 1 Spring 2012
 Name: PS#: Biol 3301 Midterm 1 Spring 2012 Multiple Choice. Circle the single best answer. (4 pts each) 1. Which of the following changes in the DNA sequence of a gene will produce a new allele? a) base
Name: PS#: Biol 3301 Midterm 1 Spring 2012 Multiple Choice. Circle the single best answer. (4 pts each) 1. Which of the following changes in the DNA sequence of a gene will produce a new allele? a) base
Connexin 26 (GJB2) gene-related deafness and speech intelligibility after cochlear implantation
 Connexin 26 (GJB2) gene-related deafness and speech intelligibility after cochlear implantation Sinnathuray, A. R., Toner, J. G., Clarke-lyttle, J., Geddis, A., Patterson, C., & Hughes, A. (2004). Connexin
Connexin 26 (GJB2) gene-related deafness and speech intelligibility after cochlear implantation Sinnathuray, A. R., Toner, J. G., Clarke-lyttle, J., Geddis, A., Patterson, C., & Hughes, A. (2004). Connexin
MRC-Holland MLPA. Description version 29; 31 July 2015
 SALSA MLPA probemix P081-C1/P082-C1 NF1 P081 Lot C1-0114. As compared to the previous B2 version (lot 0813 and 0912), 11 target probes are replaced or added, and 10 new reference probes are included. P082
SALSA MLPA probemix P081-C1/P082-C1 NF1 P081 Lot C1-0114. As compared to the previous B2 version (lot 0813 and 0912), 11 target probes are replaced or added, and 10 new reference probes are included. P082
Generating Mouse Models of Pancreatic Cancer
 Generating Mouse Models of Pancreatic Cancer Aom Isbell http://www2.massgeneral.org/cancerresourceroom/types/gi/index.asp Spring/Summer 1, 2012 Alexandros Tzatsos, MD PhD Bardeesy Lab: Goals and Objectives
Generating Mouse Models of Pancreatic Cancer Aom Isbell http://www2.massgeneral.org/cancerresourceroom/types/gi/index.asp Spring/Summer 1, 2012 Alexandros Tzatsos, MD PhD Bardeesy Lab: Goals and Objectives
Surgical and Non-Surgical Causes of Progressive Hearing Loss in Children: What can be done about it?
 Surgical and Non-Surgical Causes of Progressive Hearing Loss in Children: What can be done about it? Daniela Carvalho, MD, MMM, FAAP Professor, Surgery Department UCSD Pediatric Otolaryngology Rady Children
Surgical and Non-Surgical Causes of Progressive Hearing Loss in Children: What can be done about it? Daniela Carvalho, MD, MMM, FAAP Professor, Surgery Department UCSD Pediatric Otolaryngology Rady Children
High Frequency of GJB2 Mutation W24X among Slovak Romany (Gypsy) Patients with Non-Syndromic Hearing Loss (NSHL)
 Gen. Physiol. Biophys. (2003), 22, 549 556 549 High Frequency of GJB2 Mutation W24X among Slovak Romany (Gypsy) Patients with Non-Syndromic Hearing Loss (NSHL) G. Minárik 1,2,V.Ferák 1,E.Feráková 1,A.Ficek
Gen. Physiol. Biophys. (2003), 22, 549 556 549 High Frequency of GJB2 Mutation W24X among Slovak Romany (Gypsy) Patients with Non-Syndromic Hearing Loss (NSHL) G. Minárik 1,2,V.Ferák 1,E.Feráková 1,A.Ficek
Psych 3102 Lecture 3. Mendelian Genetics
 Psych 3102 Lecture 3 Mendelian Genetics Gregor Mendel 1822 1884, paper read 1865-66 Augustinian monk genotype alleles present at a locus can we identify this? phenotype expressed trait/characteristic can
Psych 3102 Lecture 3 Mendelian Genetics Gregor Mendel 1822 1884, paper read 1865-66 Augustinian monk genotype alleles present at a locus can we identify this? phenotype expressed trait/characteristic can
A Deletion Mutation in GJB6 Cooperating With a GJB2 Mutation in trans in Non-syndromic Deafness: A Novel Founder Mutation in Ashkenazi Jews
 HUMA MUTATIO Mutation in Brief #58 (00) Online MUTATIO I BRIEF A Deletion Mutation in GJB Cooperating With a GJB Mutation in trans in on-syndromic Deafness: A ovel Founder Mutation in Ashkenazi Jews Israela
HUMA MUTATIO Mutation in Brief #58 (00) Online MUTATIO I BRIEF A Deletion Mutation in GJB Cooperating With a GJB Mutation in trans in on-syndromic Deafness: A ovel Founder Mutation in Ashkenazi Jews Israela
Introduction to genetic variation. He Zhang Bioinformatics Core Facility 6/22/2016
 Introduction to genetic variation He Zhang Bioinformatics Core Facility 6/22/2016 Outline Basic concepts of genetic variation Genetic variation in human populations Variation and genetic disorders Databases
Introduction to genetic variation He Zhang Bioinformatics Core Facility 6/22/2016 Outline Basic concepts of genetic variation Genetic variation in human populations Variation and genetic disorders Databases
Summary. Introduction. Atypical and Duplicated Samples. Atypical Samples. Noah A. Rosenberg
 doi: 10.1111/j.1469-1809.2006.00285.x Standardized Subsets of the HGDP-CEPH Human Genome Diversity Cell Line Panel, Accounting for Atypical and Duplicated Samples and Pairs of Close Relatives Noah A. Rosenberg
doi: 10.1111/j.1469-1809.2006.00285.x Standardized Subsets of the HGDP-CEPH Human Genome Diversity Cell Line Panel, Accounting for Atypical and Duplicated Samples and Pairs of Close Relatives Noah A. Rosenberg
Genetic Hearing Loss in Children
 Genetic Hearing Loss in Children José Faibes Lubianca & Ricardo Godinho The prevalence of genetic hearing loss reaches very high numbers. In developed countries, about 50% of the cases of pre-lingual severe
Genetic Hearing Loss in Children José Faibes Lubianca & Ricardo Godinho The prevalence of genetic hearing loss reaches very high numbers. In developed countries, about 50% of the cases of pre-lingual severe
Welcome to the Genetic Code: An Overview of Basic Genetics. October 24, :00pm 3:00pm
 Welcome to the Genetic Code: An Overview of Basic Genetics October 24, 2016 12:00pm 3:00pm Course Schedule 12:00 pm 2:00 pm Principles of Mendelian Genetics Introduction to Genetics of Complex Disease
Welcome to the Genetic Code: An Overview of Basic Genetics October 24, 2016 12:00pm 3:00pm Course Schedule 12:00 pm 2:00 pm Principles of Mendelian Genetics Introduction to Genetics of Complex Disease
MRC-Holland MLPA. Description version 08; 30 March 2015
 SALSA MLPA probemix P351-C1 / P352-D1 PKD1-PKD2 P351-C1 lot C1-0914: as compared to the previous version B2 lot B2-0511 one target probe has been removed and three reference probes have been replaced.
SALSA MLPA probemix P351-C1 / P352-D1 PKD1-PKD2 P351-C1 lot C1-0914: as compared to the previous version B2 lot B2-0511 one target probe has been removed and three reference probes have been replaced.
The Meaning of Genetic Variation
 Activity 2 The Meaning of Genetic Variation Focus: Students investigate variation in the beta globin gene by identifying base changes that do and do not alter function, and by using several CD-ROM-based
Activity 2 The Meaning of Genetic Variation Focus: Students investigate variation in the beta globin gene by identifying base changes that do and do not alter function, and by using several CD-ROM-based
Published in: Otology & Neurotology
 Auditory perception and speech discrimination after cochlear implantation in patients with connexin 26 (gjb2) gene-related Deafness Sinnathuray, A. R., Toner, J. G., Geddis, A., Clarke-Lyttle, J., Patterson,
Auditory perception and speech discrimination after cochlear implantation in patients with connexin 26 (gjb2) gene-related Deafness Sinnathuray, A. R., Toner, J. G., Geddis, A., Clarke-Lyttle, J., Patterson,
ASSESSMENT OF THE RISK FOR TYPE 1 DIABETES MELLITUS CONFERRED BY HLA CLASS II GENES. Irina Durbală
 ASSESSMENT OF THE RISK FOR TYPE 1 DIABETES MELLITUS CONFERRED BY HLA CLASS II GENES Summary Irina Durbală CELL AND MOLECULAR BIOLOGY DEPARTMENT FACULTY OF MEDICINE, OVIDIUS UNIVERSITY CONSTANŢA Class II
ASSESSMENT OF THE RISK FOR TYPE 1 DIABETES MELLITUS CONFERRED BY HLA CLASS II GENES Summary Irina Durbală CELL AND MOLECULAR BIOLOGY DEPARTMENT FACULTY OF MEDICINE, OVIDIUS UNIVERSITY CONSTANŢA Class II
MRC-Holland MLPA. Description version 30; 06 June 2017
 SALSA MLPA probemix P081-C1/P082-C1 NF1 P081 Lot C1-0517, C1-0114. As compared to the previous B2 version (lot B2-0813, B2-0912), 11 target probes are replaced or added, and 10 new reference probes are
SALSA MLPA probemix P081-C1/P082-C1 NF1 P081 Lot C1-0517, C1-0114. As compared to the previous B2 version (lot B2-0813, B2-0912), 11 target probes are replaced or added, and 10 new reference probes are
Silent mutations in the phenylalanine hydroxylase
 6866 Med Genet 1991; 28: 686-690 Silent mutations in the phenylalanine hydroxylase gene as an aid to the diagnosis of phenylketonuria L Kalaydjieva, B Dworniczak, C Aulehla-Scholz,M Devoto, G Romeo, M
6866 Med Genet 1991; 28: 686-690 Silent mutations in the phenylalanine hydroxylase gene as an aid to the diagnosis of phenylketonuria L Kalaydjieva, B Dworniczak, C Aulehla-Scholz,M Devoto, G Romeo, M
DFNB93, a novel locus for autosomal recessive moderate-to-severe hearing impairment
 Clin Genet 2011: 79: 594 598 Printed in Singapore. All rights reserved 2011 John Wiley & Sons A/S CLINICAL GENETICS doi: 10.1111/j.1399-0004.2010.01593.x DFNB93, a novel locus for autosomal recessive moderate-to-severe
Clin Genet 2011: 79: 594 598 Printed in Singapore. All rights reserved 2011 John Wiley & Sons A/S CLINICAL GENETICS doi: 10.1111/j.1399-0004.2010.01593.x DFNB93, a novel locus for autosomal recessive moderate-to-severe
DIAGNOSIS Causes/Etiology of Hearing Loss
 DIAGNOSIS Causes/Etiology of Hearing Loss DIAGNOSIS Causes/Etiology of Hearing Loss VI. How Do We Hear? Sound waves enter our ears and are amplified by the ear drum and middle ear bones (ossicles), allowing
DIAGNOSIS Causes/Etiology of Hearing Loss DIAGNOSIS Causes/Etiology of Hearing Loss VI. How Do We Hear? Sound waves enter our ears and are amplified by the ear drum and middle ear bones (ossicles), allowing
Genetics Review. Alleles. The Punnett Square. Genotype and Phenotype. Codominance. Incomplete Dominance
 Genetics Review Alleles These two different versions of gene A create a condition known as heterozygous. Only the dominant allele (A) will be expressed. When both chromosomes have identical copies of the
Genetics Review Alleles These two different versions of gene A create a condition known as heterozygous. Only the dominant allele (A) will be expressed. When both chromosomes have identical copies of the
Review. Hereditary Non-Syndromic Sensorineural Hearing Loss. Transforming Silence to Sound. Iris Schrijver
 Review Journal of Molecular Diagnostics, Vol. 6, No. 4, November 2004 Copyright American Society for Investigative Pathology and the Association for Molecular Pathology Hereditary Non-Syndromic Sensorineural
Review Journal of Molecular Diagnostics, Vol. 6, No. 4, November 2004 Copyright American Society for Investigative Pathology and the Association for Molecular Pathology Hereditary Non-Syndromic Sensorineural
HEREDITY SAMPLE TOURNAMENT
 HEREDITY SAMPLE TOURNAMENT PART 1 - BACKGROUND: 1. Heterozygous means. A. Information about heritable traits B. Unique/ different molecular forms of a gene that are possible at a given locus C. Having
HEREDITY SAMPLE TOURNAMENT PART 1 - BACKGROUND: 1. Heterozygous means. A. Information about heritable traits B. Unique/ different molecular forms of a gene that are possible at a given locus C. Having
Hearing Function in Heterozygous Carriers of a Pathogenic GJB2 Gene Mutation
 Physiol. Res. 62: 323-330, 2013 Hearing Function in Heterozygous Carriers of a Pathogenic GJB2 Gene Mutation D. GROH 1,2, P. SEEMAN 3, M. JILEK 1, J. POPELÁŘ 1, Z. KABELKA 2, J. SYKA 1 1 Department of
Physiol. Res. 62: 323-330, 2013 Hearing Function in Heterozygous Carriers of a Pathogenic GJB2 Gene Mutation D. GROH 1,2, P. SEEMAN 3, M. JILEK 1, J. POPELÁŘ 1, Z. KABELKA 2, J. SYKA 1 1 Department of
Nature Genetics: doi: /ng Supplementary Figure 1. Assessment of sample purity and quality.
 Supplementary Figure 1 Assessment of sample purity and quality. (a) Hematoxylin and eosin staining of formaldehyde-fixed, paraffin-embedded sections from a human testis biopsy collected concurrently with
Supplementary Figure 1 Assessment of sample purity and quality. (a) Hematoxylin and eosin staining of formaldehyde-fixed, paraffin-embedded sections from a human testis biopsy collected concurrently with
Figure 1: Transmission of Wing Shape & Body Color Alleles: F0 Mating. Figure 1.1: Transmission of Wing Shape & Body Color Alleles: Expected F1 Outcome
 I. Chromosomal Theory of Inheritance As early cytologists worked out the mechanism of cell division in the late 1800 s, they began to notice similarities in the behavior of BOTH chromosomes & Mendel s
I. Chromosomal Theory of Inheritance As early cytologists worked out the mechanism of cell division in the late 1800 s, they began to notice similarities in the behavior of BOTH chromosomes & Mendel s
Class XII Chapter 5 Principles of Inheritance and Variation Biology
 Question 1: Mention the advantages of selecting pea plant for experiment by Mendel. Mendel selected pea plants to carry out his study on the inheritance of characters from parents to offspring. He selected
Question 1: Mention the advantages of selecting pea plant for experiment by Mendel. Mendel selected pea plants to carry out his study on the inheritance of characters from parents to offspring. He selected
Lecture 13: May 24, 2004
 Lecture 13: May 24, 2004 CH14: Mendel and the gene idea *particulate inheritance parents pass on discrete heritable units *gene- unit of inheritance which occupies a specific chromosomal location (locus)
Lecture 13: May 24, 2004 CH14: Mendel and the gene idea *particulate inheritance parents pass on discrete heritable units *gene- unit of inheritance which occupies a specific chromosomal location (locus)
GENETICS - NOTES-
 GENETICS - NOTES- Warm Up Exercise Using your previous knowledge of genetics, determine what maternal genotype would most likely yield offspring with such characteristics. Use the genotype that you came
GENETICS - NOTES- Warm Up Exercise Using your previous knowledge of genetics, determine what maternal genotype would most likely yield offspring with such characteristics. Use the genotype that you came
CURRENT GENETIC TESTING TOOLS IN NEONATAL MEDICINE. Dr. Bahar Naghavi
 2 CURRENT GENETIC TESTING TOOLS IN NEONATAL MEDICINE Dr. Bahar Naghavi Assistant professor of Basic Science Department, Shahid Beheshti University of Medical Sciences, Tehran,Iran 3 Introduction Over 4000
2 CURRENT GENETIC TESTING TOOLS IN NEONATAL MEDICINE Dr. Bahar Naghavi Assistant professor of Basic Science Department, Shahid Beheshti University of Medical Sciences, Tehran,Iran 3 Introduction Over 4000
UNIT IV. Chapter 14 The Human Genome
 UNIT IV Chapter 14 The Human Genome UNIT 2: GENETICS Chapter 7: Extending Medelian Genetics I. Chromosomes and Phenotype (7.1) A. Two copies of each autosomal gene affect phenotype 1. Most human traits
UNIT IV Chapter 14 The Human Genome UNIT 2: GENETICS Chapter 7: Extending Medelian Genetics I. Chromosomes and Phenotype (7.1) A. Two copies of each autosomal gene affect phenotype 1. Most human traits
The coiled-coil domain containing protein CCDC40 is essential for motile cilia function and left-right axis formation
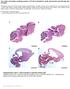 The coiled-coil domain containing protein CCDC40 is essential for motile cilia function and left-right axis formation Anita Becker-Heck#, Irene Zohn#, Noriko Okabe#, Andrew Pollock#, Kari Baker Lenhart,
The coiled-coil domain containing protein CCDC40 is essential for motile cilia function and left-right axis formation Anita Becker-Heck#, Irene Zohn#, Noriko Okabe#, Andrew Pollock#, Kari Baker Lenhart,
