In Vitro-Assembled Alphavirus Core-Like Particles Maintain a Structure Similar to That of Nucleocapsid Cores in Mature Virus
|
|
|
- Jennifer Lucas
- 6 years ago
- Views:
Transcription
1 JOURNAL OF VIROLOGY, Nov. 2002, p Vol. 76, No X/02/$ DOI: /JVI Copyright 2002, American Society for Microbiology. All Rights Reserved. In Vitro-Assembled Alphavirus Core-Like Particles Maintain a Structure Similar to That of Nucleocapsid Cores in Mature Virus Suchetana Mukhopadhyay,* Paul R. Chipman, Eunmee M. Hong, Richard J. Kuhn, and Michael G. Rossmann Department of Biological Sciences, Purdue University, West Lafayette, Indiana Received 23 May 2002/Accepted 29 July 2002 In vitro-assembled core-like particles produced from alphavirus capsid protein and nucleic acid were studied by cryoelectron microscopy. These particles were found to have a diameter of 420 Å with 240 copies of the capsid protein arranged in a T 4 icosahedral surface lattice, similar to the nucleocapsid core in mature virions. However, when the particles were subjected to gentle purification procedures, they were damaged, preventing generation of reliable structural information. Similarly, purified nucleocapsid cores isolated from virus-infected cells or from mature virus particles were also of poor quality. This suggested that in the absence of membrane and glycoproteins, nucleocapsid core particles are fragile, lacking accurate icosahedral symmetry. Alphavirus virions consist of a nucleocapsid core, a hostderived phospholipid membrane, and an outer glycoprotein layer with 80 trimeric spikes (24). The nucleocapsid core in the mature alphavirus particle has a diameter of 420 Å and consists of 240 copies of the capsid protein surrounding an 11.7-kb viral RNA genome. Each spike on the virus surface is a trimer of E1-E2 glycoprotein heterodimers. Cryoelectron microscopy (cryoem) and three-dimensional image reconstructions of numerous alphaviruses (3, 7, 14, 16, 17, 26, 27) have shown that the capsid protein subunits in the nucleocapsid core and the external glycoprotein spikes have matching T 4 icosahedral symmetry. The assembly of alphaviruses is a multistep process involving nucleocapsid core formation followed by glycoprotein attachment and virus budding (24). Upon translation from a 26S subgenomic mrna, the capsid protein is autocatalytically cleaved from the polyprotein, the viral genome RNA is encapsidated, and nucleocapsid cores are assembled in the cytoplasm. Simultaneously, the glycoproteins are translated and processed through the endoplasmic reticulum and Golgi and, subsequently, they are transported to the plasma membrane. The cytoplasmic nucleocapsid cores interact with the transmembrane E1-E2 glycoproteins at the plasma membrane, allowing the mature virions to bud from the cell. It has been suggested that this interaction is in part accomplished by 2 of the 33 cytoplasmic residues of E2 that bind into a hydrophobic pocket in the capsid protein (11, 21). Forsell et al. (7) have suggested that membrane glycoproteins have a direct role in organizing the structure of the mature alphavirus particle. They suggest that both spike-spike and spike-capsid protein interactions are responsible for proper virus symmetry. More recently, Pletnev et al. (19) and Lescar et al. (12) independently suggested that it is the E1 * Corresponding author. Mailing address: Department of Biological Sciences, Purdue University, West Lafayette, IN Phone: (765) Fax: (765) mukhopas@bilbo.bio.purdue.edu. glycoprotein that forms an icosahedral scaffold on the virus surface during assembly. Pletnev et al. (19) further suggested that cytoplasmic nucleocapsid cores lack accurate icosahedral symmetry and require association with the glycoproteins at the plasma membrane to gain a well-defined icosahedral structure. The nucleocapsid core can be divided into three sections defined by their protein and RNA content. The protein layer is defined by 30 hexamers and 12 pentamers around the periphery of the nucleocapsid core between radii of 180 and 210 Å. The carboxy-terminal domain of the Sindbis virus capsid protein (residues 114 to 264) (4) has been fitted into this layer of the Sindbis virus cryoem density, and the resulting outer surface charge distribution is largely positive, which is important for close contacts with the acidic phospholipid membrane (27). The base of the nucleocapsid core, between radii of 150 and 180 Å, is a mixture of the positively charged amino-terminal domain of the capsid protein and parts of the RNA genome and contributes to the stabilization of the nucleocapsid core (7, 18) during assembly in the cytoplasm. The cryoem density of this protein-rna mixed region is about 80% in height of the density of the outer protein layer, suggesting a slightly lesswell-ordered structure. Nevertheless, alphavirus virions also maintain T 4 icosahedral symmetry in this region (27). Internal to a radius of 150 Å is the core that consists mainly of genomic RNA. Nucleocapsid core-like particles (CLPs) can be assembled in vitro by using purified capsid protein with single-stranded nucleic acid (22, 25) or in vivo using a baculovirus system expressing capsid protein (data not shown). CLPs from either system are the same size, as determined by negative stain and cryoem, and have similar sedimentation properties as cytoplasmic nucleocapsid cores isolated from virus-infected cells (reference 22 and data not shown). However, until now, there has been no three-dimensional structural evidence that has shown that CLPs resemble either cytoplasmic nucleocapsid cores or the nucleocapsid core found in mature virus particles. Here, we present the cryoem reconstructions of in vitroassembled CLPs from Ross River virus (RRV) and western 11128
2 VOL. 76, 2002 NOTES equine encephalitis virus (WEEV) capsid proteins expressed in Escherichia coli and assembled with a synthetic 48-nucleotide oligomer (22, 25). Only after numerous trials and failed attempts at determining the structure of purified CLPs and cytoplasmic nucleocapsid cores was it concluded that extensive centrifugation steps, such as those required for sedimentation through density gradients, were detrimental and consequently were eliminated from the procedure. The cryoem reconstructions of nonpurified CLPs, although only at 30 Å resolution, show that they contain the same T 4 quasi-symmetric arrangement of hexamers and pentamers as nucleocapsid cores found in mature virions. Inability to improve the resolution of the reconstruction suggested that the CLPs vary slightly in structure among themselves and may deviate from exact icosahedral symmetry, consistent with the suggestions of Pletnev et al. (19). Capsid protein expression and purification and CLP assembly. RRV capsid protein (amino acid residues 1 to 270) from the T48 strain (10) was cloned into pet29b (Novagen, Madison, Wis.) as previously described (22) and expressed in BL21(DE3)RIL cells. Cells were grown in Luria broth at 37 C until an optical density at 600 nm (OD 600 ) of 0.5 was reached. At that time, isopropyl- -D-thiogalactopyranoside (IPTG) was added to obtain a final concentration of 1 mm. Cells were allowed to grow for an additional 6 h at 37 C before being harvested by centrifugation at 4,000 g at 4 C for 10 min. Cells were resuspended in 20 mm HEPES (ph 7.5), 0.25 M NaCl, and 5 mm EDTA (30 ml of buffer/liter of cells) and lysed after two passages through a cold French Press cell. The lysate was clarified by centrifugation at 23,000 g at 4 C for 30 min, and then the supernatant was loaded onto a 6-ml Resource S column (Amersham Biosciences, Piscataway, N.J.) that was preequilibrated with lysis buffer. A step gradient was applied, and RRV capsid protein was eluted in two peaks at 0.66 M and 0.94 M NaCl. The protein that eluted without nucleic acid at 0.66 M NaCl was concentrated and exchanged into assembly buffer (25 mm HEPES [ph 7.4], 100 mm potassium acetate, 1.7 mm magnesium acetate) and used for in vitro assembly reactions. WEEV capsid protein (amino acid residues 1 to 259) from strain BFS1703 (9) was cloned into psbet (20) and expressed in BL21(DE3) cells. Cells were grown in Luria broth at 37 C until an OD 600 of 0.4 was reached, at which time IPTG was added to obtain a final concentration of 1 mm; cells continued to grow for an additional 10 h at 37 C. The cells were pelleted at 4,000 g at 4 C for 10 min and resuspended in buffer A (25 mm Tris-Cl [ph 7.4], 5 mm EDTA, 5% glycerol [vol/vol], 5 mm dithiothreitol) and 50 mm NaCl (10 ml of buffer/liter of cells). Cells were lysed using a cold French press, and the soluble fraction was obtained after centrifugation at 23,000 g at 4 C for 30 min. The clarified cell lysate was loaded onto a 20-ml SP Sepharose Fast Flow column (Amersham Biosciences) equilibrated with buffer A. A linear gradient was applied, and the WEEV capsid protein was eluted with 0.6 M NaCl in buffer A. The capsid protein was concentrated with a Centriprep-10 concentrator (Millipore, Bedford, Mass.) and exchanged into buffer A and 0.12 M NaCl. The concentrated protein was applied to a Superdex 75 10/30 column (Amersham Biosciences), and the fractions containing WEEV capsid protein were concentrated and exchanged into assembly buffer. In contrast to Sindbis virus capsid protein (22), full-length RRV and WEEV capsid proteins could be expressed and purified. In vitro assembly of WEEV CLPs was essentially as described for Sindbis virus and RRV CLPs (22). Purification of cytoplasmic nucleocapsid cores. Preassembled nucleocapsid cores were isolated from virus-infected cells. Cells were lysed with detergent (6, 18) to release the nucleocapsid cores from internal membranes (8, 13), and sedimentation centrifugation was used to further purify them from the cell lysate. To eliminate the possibility that nucleocapsid cores were sensitive to the detergent present during their isolation, nucleocapsid cores from the mutant E2-L402Y Sindbis virus, which accumulated in the cytoplasm independent of membrane association (15) and hence did not require detergent for their isolation, were also purified and used for cryoem studies. Furthermore, some of the purified cytoplasmic nucleocapsid cores were cross-linked with dimethyl suberimidate (23) to enhance their stability. CryoEM of purified cytoplasmic nucleocapsid cores and CLPs. CryoEM reconstructions (2) were not successful when the cytoplasmic nucleocapsid cores or CLPs were purified by sedimentation centrifugation (A. E. Hamburger, S. Lee, S. Mukhopadhyay, T. L. Tellinghuisen, and W. Zhang, unpublished data). The failed image reconstruction attempts included efforts using both the common lines method (5) and the model-based approach of the polar Fourier transform method (1). The nucleocapsid core from the previously published whole-virus cryoem structure of RRV (3) and the outer protein layer of the RRV nucleocapsid core were used as starting models. However, in these procedures, the particle orientation and centers varied greatly from cycle to cycle and the correlation coefficients approached zero with further iterations, resulting in a smooth ball of density. It was possible that purified Sindbis virus capsid protein CLPs did not produce reliable reconstructions because an N- terminal truncated Sindbis virus capsid protein (amino acid residues 19 to 264) (22) was used and, as a result, the CLPs were less stable. However, purified CLPs using full-length RRV capsid protein also did not yield satisfactory structural results. Furthermore, cytoplasmic nucleocapsid cores that were cross-linked (23) did not improve the reconstruction, and it was concluded that perhaps the centrifugation step had already damaged the cytoplasmic nucleocapsid cores. In contrast, several high-resolution cryoem structures have been obtained from mature alphavirus particles that were purified by different methods, including sedimentation centrifugation (14, 27; P. R. Chipman, personal communication). These results suggested that cytoplasmic nucleocapsid cores and CLPs, unlike mature virus particles, are fragile and could be damaged during centrifugation, although the damage could only be detected by an inability to produce good three-dimensional reconstructions. CryoEM of nonpurified CLPs. A successful reconstruction of in vitro-assembled RRV and WEEV CLPs was achieved when the particles were not purified by sedimentation gradient centrifugation after being assembled (Table 1). In contrast to the results with purified CLPs, the reconstruction of the nonpurified CLPs showed T 4 quasi-symmetry, consistent with cryoem reconstructions of the whole virus (Fig. 1). The 30 Å resolution limit and the need to reject 70% of all boxed particles for both CLP reconstructions suggests that CLPs, and
3 11130 NOTES J. VIROL. Sample No. of particles in map TABLE 1. CryoEM and three-dimensional image reconstruction data a No. of particles boxed Resolution (Å) Correl. coeff. b Dose (e /Å 2 ) Defocus range ( m) Mature RRV 961 1, Sindbis virus RRV CLP 671 2, RRV core c WEEV CLP 581 2, RRV core c a A Phillips CM300 FEG transmission electron microscope was used to record images on Kodak SO-163 film under low-dose conditions. All micrographs were digitized using a Zeiss-SCAI microdensitometer at 14- m intervals. CryoEM reconstructions were performed as previously described (2). b Correlation coefficient is defined as CMP CC in Baker and Cheng (1). c RRV core refers to the nucleocapsid core from the cryoem reconstruction of the mature RRV particle described above. Starting model most likely cytoplasmic nucleocapsid cores, are fragile and that their structure can be easily damaged during purification (see above). In comparison, reconstructions of mature alphaviruses have at least 20 Å resolution and can readily be extended to 10 Å resolution by increasing the number of particles (14, 27). Presumably, the nucleocapsid core becomes a rigid icosahedral structure only after interactions with the phospholipid membrane and the glycoproteins (7, 19). It was not possible to obtain a successful reconstruction of the cytoplasmic nucleocapsid cores because centrifugation was necessary to produce a sufficiently clean preparation. Biochemical analyses (Table 2) (22) have demonstrated that the in vitro-assembled CLPs have similar properties to cytoplasmic nucleocapsid cores. Thus, it is reasonable to postulate that the cytoplasmic nucleocapsid cores have a similar structure to the CLPs and to the nucleocapsid core of the mature virus. The most obvious difference between the structures of the RRV CLPs and the RRV nucleocapsid core of the mature virus is that the orientation of the pentamers is rotated by about 10 counterclockwise with respect to the icosahedral axial system when viewed from the outside (Fig. 1). In addition, in the RRV CLPs the density around the fivefold axes projects outwards, away from the center of the particle. In contrast, the orientation of the monomers in the hexamers between the RRV CLPs and the RRV nucleocapsid core appears to be the same (Fig. 1). Neither CLPs nor cytoplasmic nucleocapsid cores have yet been in contact with membrane proteins; thus, these particles probably represent an earlier stage of assembly. Hence, the difference at the fivefold axes between CLPs compared to the mature nucleocapsid core might be the result of different stages in the virus assembly process. In contrast to the mature virus, the density in the base of the RRV CLPs has no T 4 quasi-symmetry, and the maximum density in this region is only about 35% in height of the density in the protein layer. This difference in the base of the CLP compared to the nucleocapsid core of the mature virion suggests specific protein-protein and protein-rna interactions in the mature particle that are weaker in the CLPs, possibly because a DNA oligonucleotide rather than viral RNA was used in the assembly reactions. Furthermore, in the protein layer of the RRV CLPs, the volume of density representing each capsid protein monomer is less than its corresponding volume in the RRV nucleocapsid core of the mature virus. In both the RRV and WEEV CLP reconstructions, some additional density appears in the middle of each hexamer (but not pentamer) at a radial distance between 195 and 225 Å. Although this extra density might represent protein or noise, the most probable interpretation is that it represents a DNA oligonucleotide that binds to the external surface of the positively charged hexameric arrangement of capsid proteins (27). Paredes et al. (16) suggested that there is a difference in the nucleocapsid core of Old World (RRV, Sindbis virus) and New World (WEEV, Venezuelan equine encephalomyelitis virus) alphaviruses. Relative to Sindbis virus, the orientations of the hexamers and pentamers in Venezuelan equine encephalomyelitis virus were rotated by approximately 11 and 4 clockwise, respectively (16). In contrast to what has been observed in the mature virus particles, no significant difference in the orientation of the hexamers and pentamers was seen between the RRV CLPs and the WEEV CLPs (Fig. 1). This suggests that CLPs and cytoplasmic nucleocapsid cores are similar in their organization, regardless of the alphavirus. Only after their interaction with the glycoproteins and lipid membrane will they attain a conformation that may be specific to Old and New World viruses. Implications for particle assembly and structure. In vitroassembled CLPs are a reliable representation of the nucleocapsid cores found in the cytoplasm of infected cells. Biochemical analyses showed that CLPs and cytoplasmic nucleocapsid cores were similar in size and stability, and cryoem reconstructions showed that CLPs closely resemble the nucleocapsid core found in the mature virus. Thus, in vitro-assembled CLPs are good models for studying alphavirus nucleocapsid core assembly, permitting the isolation and characterization of assembly intermediates as well as of fully assembled CLPs. FIG. 1. CryoEM density maps of RRV CLPs, WEEV CLPs, and the nucleocapsid core excised from the reconstruction of mature RRV. (A to C) Stereo diagrams, where only the top hemisphere of the map is shown. All maps are at 30 Å resolution and contoured at 2.5. The white triangle represents one icosahedral asymmetric unit at a radius of 170 Å. The top vertex of the triangle is at the fivefold-symmetry axis. (A) The RRV CLP structure with density shown between radii of 150 and 210 Å. (B) The densities of both RRV CLP (blue; radii of 150 to 210 Å) and WEEV CLP (pink; radii of 180 to 210 Å). Note the quasi-symmetry present in both CLP reconstructions. (C) Superimposed densities of RRV CLP (blue; radii of 150 to 210 Å) and the nucleocapsid core from mature RRV (green; radii of 150 to 210 Å). (D) Central cross section along twofold axis of RRV CLP (blue) and WEEV CLP (pink), both from radii of 150 to 210 Å. (E) Same orientation as shown in panel D, with RRV CLP (blue) and the nucleocapsid core from the mature RRV particle (green).
4 VOL. 76, 2002 NOTES 11131
5 11132 NOTES J. VIROL. TABLE 2. Biochemical characterization of in vitro-assembled CLPs and in vivo-assembled nucleocapsid cores Property Cytoplasmic nucleocapsid cores or CLPs Assay method a,b Ionic strength Unstable in assembly buffer Agarose gel, EM plus 0.5 M NaCl ph Stable between ph 5.2 and 8.2 Agarose gel, EM, gradient sedimentation Temperature Unstable above 55 C Agarose gel Nuclease sensitivity Yes Agarose gel, EM Particle size Å CryoEM a All assays were performed as described by Tellinghuisen et al. (22). b EM refers to negative-stain EM. We thank Cheryl Towell and Sharon Wilder for help in preparation of the manuscript, and Rob Ashmore and Chuan Xiao for assistance with RobEM and other programs (see / viruswww/rossmann_home/softwares.shtml). The work was supported by an NIH Program project grant to R.J.K. and M.G.R. and others (AI45976), an NIH grant to R.J.K. (GM56279), a Purdue University reinvestment grant, and a grant from the Keck Foundation. REFERENCES 1. Baker, T. S., and R. H. Cheng A model-based approach for determining orientations of biological macromolecules imaged by cryoelectron microscopy. J. Struct. Biol. 116: Baker, T. S., N. H. Olson, and S. D. Fuller Adding the third dimension to virus life cycles: three-dimensional reconstruction of icosahedral viruses from cryo-electron micrographs. Microbiol. Mol. Biol. Rev. 63: Cheng, R. H., R. J. Kuhn, N. H. Olson, M. G. Rossmann, H. K. Choi, T. J. Smith, and T. S. Baker Nucleocapsid and glycoprotein organization in an enveloped virus. Cell 80: Choi, H. K., L. Tong, W. Minor, P. Dumas, U. Boege, M. G. Rossmann, and G. Wengler Structure of Sindbis virus core protein reveals a chymotrypsin-like serine proteinase and the organization of the virion. Nature (London) 354: Crowther, R. A Procedures for three-dimensional reconstruction of spherical viruses by Fourier synthesis from electron micrographs. Phil. Trans. R. Soc. Lond. B 261: Forsell, K., G. Griffiths, and H. Garoff Preformed cytoplasmic nucleocapsids are not necessary for alphavirus budding. EMBO J. 15: Forsell, K., L. Xing, T. Kozlovska, R. H. Cheng, and H. Garoff Membrane proteins organize a symmetrical virus. EMBO J. 19: Froshauer, S., J. Kartenbeck, and A. Helenius Alphavirus RNA replicase is located on the cytoplasmic surface of endosomes and lysosomes. J. Cell Biol. 107: Hahn, C. S., S. Lustig, E. G. Strauss, and J. H. Strauss Western equine encephalitis virus is a recombinant virus. Proc. Natl. Acad. Sci. USA 85: Kuhn, R. J., H. G. Niesters, Z. Hong, and J. H. Strauss Infectious RNA transcripts from Ross River virus cdna clones and the construction and characterization of defined chimeras with Sindbis virus. Virology 182: Lee, S., K. E. Owen, H. K. Choi, H. Lee, G. Lu, G. Wengler, D. T. Brown, M. G. Rossmann, and R. J. Kuhn Identification of a protein binding site on the surface of the alphavirus nucleocapsid and its implication in virus assembly. Structure 4: Lescar, J., A. Roussel, M. W. Wein, J. Navaza, S. D. Fuller, G. Wengler, G. Wengler, and F. A. Rey The fusion glycoprotein shell of Semliki Forest virus: an icosahedral assembly primed for fusogenic activation at endosomal ph. Cell 105: Lopez, S., J. S. Yao, R. J. Kuhn, E. G. Strauss, and J. H. Strauss Nucleocapsid-glycoprotein interactions required for assembly of alphaviruses. J. Virol. 68: Mancini, E. J., M. Clarke, B. Gowen, T. Rutten, and S. D. Fuller Cryo-electron microscopy reveals the functional anatomy of an enveloped virus, Semliki Forest virus. Mol. Cell 5: Owen, K. E., and R. J. Kuhn Alphavirus budding is dependent on the interaction between the nucleocapsid and hydrophobic amino acids on the cytoplasmic domain of the E2 envelope glycoprotein. Virology 230: Paredes, A., K. Alwell-Warda, S. C. Weaver, W. Chiu, and S. J. Watowich Venezuelan equine encephalomyelitis virus structure and its divergence from Old World alphaviruses. J. Virol. 75: Paredes, A. M., D. T. Brown, R. Rothnagel, W. Chiu, R. J. Schoepp, R. E. Johnston, and B. V. V. Prasad Three-dimensional structure of a membrane-containing virus. Proc. Natl. Acad. Sci. USA 90: Perera, R., K. E. Owen, T. L. Tellinghuisen, A. E. Gorbalenya, and R. J. Kuhn Alphavirus nucleocapsid protein contains a putative coiled-coil alpha-helix important for core assembly. J. Virol. 75: Pletnev, S. V., W. Zhang, S. Mukhopadhyay, B. R. Fisher, R. Hernandez, D. T. Brown, T. S. Baker, M. G. Rossmann, and R. J. Kuhn Locations of carbohydrate sites on Sindbis virus glycoproteins show that E1 forms an icosahedral scaffold. Cell 105: Schenk, P. M., S. Baumann, R. Mattes, and H. H. Steinbiss Improved high-level expression system for eukaryotic genes in Escherichia coli using T7 RNA polymerase and rare Arg trnas. BioTechniques 19: Skoging, U., M. Vihinen, L. Nilsson, and P. Liljeström Aromatic interactions define the binding of the alphavirus spike to its nucleocapsid. Structure 4: Tellinghuisen, T. L., A. E. Hamburger, B. R. Fisher, R. Ostendorp, and R. J. Kuhn In vitro assembly of alphavirus cores by using nucleocapsid protein expressed in Escherichia coli. J. Virol. 73: Tellinghuisen, T. L., and R. J. Kuhn Nucleic acid-dependent crosslinking of the nucleocapsid protein of Sindbis virus. J. Virol. 74: Tellinghuisen, T. L., R. Perera, and R. J. Kuhn Genetic and biochemical studies on the assembly of an enveloped virus. Genet. Eng. 23: Wengler, G., U. Boege, G. Wengler, H. Bischoff, and K. Wahn The core protein of the alphavirus Sindbis virus assembles into core-like nucleoproteins with the viral genome RNA and with other single-stranded nucleic acids in vitro. Virology 118: Zhang, W., B. R. Fisher, N. H. Olson, J. H. Strauss, R. J. Kuhn, and T. S. Baker Aura virus structure suggests that the T 4 organization is a fundamental property of viral structural proteins. J. Virol. 76: Zhang, W., S. Mukhopadhyay, S.V. Pletnev, T. S. Baker, R. J. Kuhn, and M. G. Rossmann. Placement of the structural proteins in Sindbis virus. J. Virol., in press.
Aura Virus Structure Suggests that the T 4 Organization Is a Fundamental Property of Viral Structural Proteins
 JOURNAL OF VIROLOGY, July 2002, p. 7239 7246 Vol. 76, No. 14 0022-538X/02/$04.00 0 DOI: 10.1128/JVI.76.14.7239 7246.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved. Aura Virus
JOURNAL OF VIROLOGY, July 2002, p. 7239 7246 Vol. 76, No. 14 0022-538X/02/$04.00 0 DOI: 10.1128/JVI.76.14.7239 7246.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved. Aura Virus
Virus Structure. Characteristics of capsids. Virus envelopes. Virion assembly John Wiley & Sons, Inc. All rights reserved.
 Virus Structure Characteristics of capsids Virus envelopes Virion assembly Capsids package viral genomes and transmit them to a new host cell Capsid rigid, symmetrical container composed of viral protein
Virus Structure Characteristics of capsids Virus envelopes Virion assembly Capsids package viral genomes and transmit them to a new host cell Capsid rigid, symmetrical container composed of viral protein
Mapping the Structure and Function of the E1 and E2 Glycoproteins in Alphaviruses
 Structure 14, 63 73, January 2006 ª2006 Elsevier Ltd All rights reserved DOI 10.1016/j.str.2005.07.025 Mapping the Structure and Function of the E1 and E2 Glycoproteins in Alphaviruses Suchetana Mukhopadhyay,
Structure 14, 63 73, January 2006 ª2006 Elsevier Ltd All rights reserved DOI 10.1016/j.str.2005.07.025 Mapping the Structure and Function of the E1 and E2 Glycoproteins in Alphaviruses Suchetana Mukhopadhyay,
Coronaviruses. Virion. Genome. Genes and proteins. Viruses and hosts. Diseases. Distinctive characteristics
 Coronaviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Spherical enveloped particles studded with clubbed spikes Diameter 120-160 nm Coiled helical
Coronaviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Spherical enveloped particles studded with clubbed spikes Diameter 120-160 nm Coiled helical
Research Article 531. *Corresponding authors. Key words: assembly, capsid structure, mutational analysis, Sindbis virus, virus budding
 Research Article 531 Identification of a protein binding site on the surface of the alphavirus nucleocapsid and its implication in virus assembly Sukyeong Lee 1, Katherine E Owen 1, Hok-Kin Choi 1, Heuiran
Research Article 531 Identification of a protein binding site on the surface of the alphavirus nucleocapsid and its implication in virus assembly Sukyeong Lee 1, Katherine E Owen 1, Hok-Kin Choi 1, Heuiran
Structural biology of viruses
 Structural biology of viruses Biophysical Chemistry 1, Fall 2010 Coat proteins DNA/RNA packaging Reading assignment: Chap. 15 Virus particles self-assemble from coat monomers Virus Structure and Function
Structural biology of viruses Biophysical Chemistry 1, Fall 2010 Coat proteins DNA/RNA packaging Reading assignment: Chap. 15 Virus particles self-assemble from coat monomers Virus Structure and Function
Three-dimensional structure of a membrane-containing virus
 Proc. Natl. Acad. Sci. USA Vol. 90, pp. 9095-9099, October 1993 Microbiology Three-dimensional structure of a membrane-containing virus ANGEL M. PAREDES*, DENNIS T. BROWN*t, ROSALBA ROTHNAGELt, WAH CHIU*,
Proc. Natl. Acad. Sci. USA Vol. 90, pp. 9095-9099, October 1993 Microbiology Three-dimensional structure of a membrane-containing virus ANGEL M. PAREDES*, DENNIS T. BROWN*t, ROSALBA ROTHNAGELt, WAH CHIU*,
Probing the Potential Glycoprotein Binding Site of Sindbis Virus Capsid Protein With Dioxane and Model Building
 PROTEINS: Structure, Function, and Genetics 33:311 317 (1998) Probing the Potential Glycoprotein Binding Site of Sindbis Virus Capsid Protein With Dioxane and Model Building Sukyeong Lee, Richard J. Kuhn,
PROTEINS: Structure, Function, and Genetics 33:311 317 (1998) Probing the Potential Glycoprotein Binding Site of Sindbis Virus Capsid Protein With Dioxane and Model Building Sukyeong Lee, Richard J. Kuhn,
Overview of virus life cycle
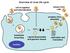 Overview of virus life cycle cell recognition and internalization release from cells progeny virus assembly membrane breaching nucleus capsid disassembly and genome release replication and translation
Overview of virus life cycle cell recognition and internalization release from cells progeny virus assembly membrane breaching nucleus capsid disassembly and genome release replication and translation
NIH Public Access Author Manuscript Nat Struct Biol. Author manuscript; available in PMC 2014 August 28.
 NIH Public Access Author Manuscript Published in final edited form as: Nat Struct Biol. 2003 November ; 10(11): 907 912. doi:10.1038/nsb990. Visualization of membrane protein domains by cryo-electron microscopy
NIH Public Access Author Manuscript Published in final edited form as: Nat Struct Biol. 2003 November ; 10(11): 907 912. doi:10.1038/nsb990. Visualization of membrane protein domains by cryo-electron microscopy
Visualization of membrane protein domains by cryo-electron microscopy of dengue virus
 Visualization of membrane protein domains by cryo-electron microscopy of dengue virus Wei Zhang 1, Paul R Chipman 1, Jeroen Corver 2,3, Peter R Johnson 1, Ying Zhang 1, Suchetana Mukhopadhyay 1, Timothy
Visualization of membrane protein domains by cryo-electron microscopy of dengue virus Wei Zhang 1, Paul R Chipman 1, Jeroen Corver 2,3, Peter R Johnson 1, Ying Zhang 1, Suchetana Mukhopadhyay 1, Timothy
Polyomaviridae. Spring
 Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
Differences in the Postfusion Conformations of Full-Length and Truncated Class II Fusion Protein E of Tick-Borne Encephalitis Virus
 JOURNAL OF VIROLOGY, May 2005, p. 6511 6515 Vol. 79, No. 10 0022-538X/05/$08.00 0 doi:10.1128/jvi.79.10.6511 6515.2005 Copyright 2005, American Society for Microbiology. All Rights Reserved. Differences
JOURNAL OF VIROLOGY, May 2005, p. 6511 6515 Vol. 79, No. 10 0022-538X/05/$08.00 0 doi:10.1128/jvi.79.10.6511 6515.2005 Copyright 2005, American Society for Microbiology. All Rights Reserved. Differences
Science Supporting Online Material.
 - S1 - Science Supporting Online Material. Grünewald et al., Three-dimensional Structure of Herpes Simplex Virus from Cryoelectron Tomography. Materials and Methods Strains, cultivation and purification
- S1 - Science Supporting Online Material. Grünewald et al., Three-dimensional Structure of Herpes Simplex Virus from Cryoelectron Tomography. Materials and Methods Strains, cultivation and purification
Proteins and symmetry
 Proteins and symmetry Viruses (symmetry) Viruses come in many shapes, sizes and compositions All carry genomic nucleic acid (RNA or DNA) Structurally and genetically the simplest are the spherical viruses
Proteins and symmetry Viruses (symmetry) Viruses come in many shapes, sizes and compositions All carry genomic nucleic acid (RNA or DNA) Structurally and genetically the simplest are the spherical viruses
Structure of viruses
 Structure of viruses Lecture 4 Biology 3310/4310 Virology Spring 2018 In order to create something that functions properly - a container, a chair, a house - its essence has to be explored, for it should
Structure of viruses Lecture 4 Biology 3310/4310 Virology Spring 2018 In order to create something that functions properly - a container, a chair, a house - its essence has to be explored, for it should
Structure of viruses
 Structure of viruses Lecture 4 Biology 3310/4310 Virology Spring 2017 In order to create something that functions properly - a container, a chair, a house - its essence has to be explored, for it should
Structure of viruses Lecture 4 Biology 3310/4310 Virology Spring 2017 In order to create something that functions properly - a container, a chair, a house - its essence has to be explored, for it should
Fine Mapping of a cis-acting Sequence Element in Yellow Fever Virus RNA That Is Required for RNA Replication and Cyclization
 JOURNAL OF VIROLOGY, Feb. 2003, p. 2265 2270 Vol. 77, No. 3 0022-538X/03/$08.00 0 DOI: 10.1128/JVI.77.3.2265 2270.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved. Fine Mapping
JOURNAL OF VIROLOGY, Feb. 2003, p. 2265 2270 Vol. 77, No. 3 0022-538X/03/$08.00 0 DOI: 10.1128/JVI.77.3.2265 2270.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved. Fine Mapping
Lecture 2: Virology. I. Background
 Lecture 2: Virology I. Background A. Properties 1. Simple biological systems a. Aggregates of nucleic acids and protein 2. Non-living a. Cannot reproduce or carry out metabolic activities outside of a
Lecture 2: Virology I. Background A. Properties 1. Simple biological systems a. Aggregates of nucleic acids and protein 2. Non-living a. Cannot reproduce or carry out metabolic activities outside of a
Sindbis Virus Glycoproteins Form a Regular Icosahedral Surface Lattice
 JOURNAL OF VIROLOGY, July 1975, p. 141-145 Copyright i 1975 American Society for Microbiology Vol. 16, No. 1 Printed in U.S.A. Sindbis Virus Glycoproteins Form a Regular Icosahedral Surface Lattice C.-H.
JOURNAL OF VIROLOGY, July 1975, p. 141-145 Copyright i 1975 American Society for Microbiology Vol. 16, No. 1 Printed in U.S.A. Sindbis Virus Glycoproteins Form a Regular Icosahedral Surface Lattice C.-H.
Identification of a Region in the Sindbis Virus Nucleocapsid Protein That Is Involved in Specificity of RNA Encapsidation
 JOURNAL OF VIROLOGY, May 1996, p. 2757 2763 Vol. 70, No. 5 0022-538X/96/$04.00 0 Copyright 1996, American Society for Microbiology Identification of a Region in the Sindbis Virus Nucleocapsid Protein That
JOURNAL OF VIROLOGY, May 1996, p. 2757 2763 Vol. 70, No. 5 0022-538X/96/$04.00 0 Copyright 1996, American Society for Microbiology Identification of a Region in the Sindbis Virus Nucleocapsid Protein That
Structure of Immature West Nile Virus
 JOURNAL OF VIROLOGY, June 2007, p. 6141 6145 Vol. 81, No. 11 0022-538X/07/$08.00 0 doi:10.1128/jvi.00037-07 Copyright 2007, American Society for Microbiology. All Rights Reserved. Structure of Immature
JOURNAL OF VIROLOGY, June 2007, p. 6141 6145 Vol. 81, No. 11 0022-538X/07/$08.00 0 doi:10.1128/jvi.00037-07 Copyright 2007, American Society for Microbiology. All Rights Reserved. Structure of Immature
STRUCTURE, GENERAL CHARACTERISTICS AND REPRODUCTION OF VIRUSES
 STRUCTURE, GENERAL CHARACTERISTICS AND REPRODUCTION OF VIRUSES Introduction Viruses are noncellular genetic elements that use a living cell for their replication and have an extracellular state. Viruses
STRUCTURE, GENERAL CHARACTERISTICS AND REPRODUCTION OF VIRUSES Introduction Viruses are noncellular genetic elements that use a living cell for their replication and have an extracellular state. Viruses
Reoviruses. Virion. Genome. Genes and proteins. Viruses and hosts. Diseases. Distinctive characteristics
 Reoviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Naked icosahedral capsid (T=13), diameter 60-85 nm Capsid consists of two or three concentric protein
Reoviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Naked icosahedral capsid (T=13), diameter 60-85 nm Capsid consists of two or three concentric protein
ABSTRACT. WEST, JOHN ALLEN. Investigation of the Role of the E2 Endodomain in Sindbis Virus
 ABSTRACT WEST, JOHN ALLEN. Investigation of the Role of the E2 Endodomain in Sindbis Virus Assembly. (Under the direction of Dr. Dennis T. Brown) Sindbis virus (SV) is the prototype member of the alphavirus
ABSTRACT WEST, JOHN ALLEN. Investigation of the Role of the E2 Endodomain in Sindbis Virus Assembly. (Under the direction of Dr. Dennis T. Brown) Sindbis virus (SV) is the prototype member of the alphavirus
RESCUE OF INFECTIOUS PARTICLES FROM PRE-ASSEMBLED ALPHAVIRUS NUCLEOCAPSID CORES. Richard J. Kuhn 1,2 *
 JVI Accepts, published online ahead of print on 6 April 2011 J. Virol. doi:10.1128/jvi.00039-11 Copyright 2011, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved.
JVI Accepts, published online ahead of print on 6 April 2011 J. Virol. doi:10.1128/jvi.00039-11 Copyright 2011, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved.
Association of Sindbis Virus Capsid Protein with Phospholipid Membranes and the E2 Glycoprotein: Implications for Alphavirus Assembly
 2800 Biochemistry 2005, 44, 2800-2810 Association of Sindbis Virus Capsid Protein with Phospholipid Membranes and the E2 Glycoprotein: Implications for Alphavirus Assembly Thomas A. Wilkinson,, Timothy
2800 Biochemistry 2005, 44, 2800-2810 Association of Sindbis Virus Capsid Protein with Phospholipid Membranes and the E2 Glycoprotein: Implications for Alphavirus Assembly Thomas A. Wilkinson,, Timothy
Translation. Host Cell Shutoff 1) Initiation of eukaryotic translation involves many initiation factors
 Translation Questions? 1) How does poliovirus shutoff eukaryotic translation? 2) If eukaryotic messages are not translated how can poliovirus get its message translated? Host Cell Shutoff 1) Initiation
Translation Questions? 1) How does poliovirus shutoff eukaryotic translation? 2) If eukaryotic messages are not translated how can poliovirus get its message translated? Host Cell Shutoff 1) Initiation
Virology. *Viruses can be only observed by electron microscope never by light microscope. The size of the virus: nm in diameter.
 Virology We are going to start with general introduction about viruses, they are everywhere around us; in food; within the environment; in direct contact to etc.. They may cause viral infection by itself
Virology We are going to start with general introduction about viruses, they are everywhere around us; in food; within the environment; in direct contact to etc.. They may cause viral infection by itself
Viral structure م.م رنا مشعل
 Viral structure م.م رنا مشعل Viruses must reproduce (replicate) within cells, because they cannot generate energy or synthesize proteins. Because they can reproduce only within cells, viruses are obligate
Viral structure م.م رنا مشعل Viruses must reproduce (replicate) within cells, because they cannot generate energy or synthesize proteins. Because they can reproduce only within cells, viruses are obligate
Low ph Induces Swiveling of the Glycoprotein Heterodimers in the Semliki Forest Virus Spike Complex
 Cell, Vol. 81,715-725, June 2, 1995, Copyright 1995 by Cell Press Low ph Induces Swiveling of the Glycoprotein Heterodimers in the Semliki Forest Virus Spike Complex Stephen D. Fuller,* John A. Berriman,t
Cell, Vol. 81,715-725, June 2, 1995, Copyright 1995 by Cell Press Low ph Induces Swiveling of the Glycoprotein Heterodimers in the Semliki Forest Virus Spike Complex Stephen D. Fuller,* John A. Berriman,t
General Virology I. Dr Esam Ibraheem Azhar (BSc, MSc, Ph.D Molecular Medical Virology) Asst. Prof. Medical Laboratory Technology Department
 General Virology I Dr Esam Ibraheem Azhar (BSc, MSc, Ph.D Molecular Medical Virology) Asst. Prof. Medical Laboratory Technology Department ١ General Virology I Lecture Outline Introduction istory Definition
General Virology I Dr Esam Ibraheem Azhar (BSc, MSc, Ph.D Molecular Medical Virology) Asst. Prof. Medical Laboratory Technology Department ١ General Virology I Lecture Outline Introduction istory Definition
Association of the pr Peptides with Dengue Virus at Acidic ph Blocks Membrane Fusion
 JOURNAL OF VIROLOGY, Dec. 2009, p. 12101 12107 Vol. 83, No. 23 0022-538X/09/$12.00 doi:10.1128/jvi.01637-09 Copyright 2009, American Society for Microbiology. All Rights Reserved. Association of the pr
JOURNAL OF VIROLOGY, Dec. 2009, p. 12101 12107 Vol. 83, No. 23 0022-538X/09/$12.00 doi:10.1128/jvi.01637-09 Copyright 2009, American Society for Microbiology. All Rights Reserved. Association of the pr
Chapter 6- An Introduction to Viruses*
 Chapter 6- An Introduction to Viruses* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. 6.1 Overview of Viruses
Chapter 6- An Introduction to Viruses* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. 6.1 Overview of Viruses
CELLS. Cells. Basic unit of life (except virus)
 Basic unit of life (except virus) CELLS Prokaryotic, w/o nucleus, bacteria Eukaryotic, w/ nucleus Various cell types specialized for particular function. Differentiation. Over 200 human cell types 56%
Basic unit of life (except virus) CELLS Prokaryotic, w/o nucleus, bacteria Eukaryotic, w/ nucleus Various cell types specialized for particular function. Differentiation. Over 200 human cell types 56%
In vitro analysis of factors involved in the disassembly of Sindbis virus cores by 60S ribosomal subunits identifies a possible role of low ph
 Journal of General Virology (2002), 83, 2417 2426. Printed in Great Britain... In vitro analysis of factors involved in the disassembly of Sindbis virus cores by 60S ribosomal subunits identifies a possible
Journal of General Virology (2002), 83, 2417 2426. Printed in Great Britain... In vitro analysis of factors involved in the disassembly of Sindbis virus cores by 60S ribosomal subunits identifies a possible
Structural vs. nonstructural proteins
 Why would you want to study proteins associated with viruses or virus infection? Receptors Mechanism of uncoating How is gene expression carried out, exclusively by viral enzymes? Gene expression phases?
Why would you want to study proteins associated with viruses or virus infection? Receptors Mechanism of uncoating How is gene expression carried out, exclusively by viral enzymes? Gene expression phases?
Virus Maturation by Budding
 MICROBIOLOGY AND MOLECULAR BIOLOGY REVIEWS, Dec. 1998, p. 1171 1190 Vol. 62, No. 4 1092-2172/98/$04.00 0 Copyright 1998, American Society for Microbiology. All Rights Reserved. Virus Maturation by Budding
MICROBIOLOGY AND MOLECULAR BIOLOGY REVIEWS, Dec. 1998, p. 1171 1190 Vol. 62, No. 4 1092-2172/98/$04.00 0 Copyright 1998, American Society for Microbiology. All Rights Reserved. Virus Maturation by Budding
Visualization of fusion activation in the Semliki Forest virus spike
 Visualization of fusion activation in the Semliki Forest virus spike John M Kenney', Mathilda Sjoberg 2, Henrik Garoff 2 and Stephen D Fuller'* 1 Biological Structures and Biocomputing Programme, European
Visualization of fusion activation in the Semliki Forest virus spike John M Kenney', Mathilda Sjoberg 2, Henrik Garoff 2 and Stephen D Fuller'* 1 Biological Structures and Biocomputing Programme, European
Viruses defined acellular organisms genomes nucleic acid replicate inside host cells host metabolic machinery ribosomes
 The Viruses Viruses Viruses may be defined as acellular organisms whose genomes consist of nucleic acid, obligately replicate inside host cells using host metabolic machinery and ribosomes to form a pool
The Viruses Viruses Viruses may be defined as acellular organisms whose genomes consist of nucleic acid, obligately replicate inside host cells using host metabolic machinery and ribosomes to form a pool
Julianne Edwards. Retroviruses. Spring 2010
 Retroviruses Spring 2010 A retrovirus can simply be referred to as an infectious particle which replicates backwards even though there are many different types of retroviruses. More specifically, a retrovirus
Retroviruses Spring 2010 A retrovirus can simply be referred to as an infectious particle which replicates backwards even though there are many different types of retroviruses. More specifically, a retrovirus
Isolation of Capsid Protein Dimers from the Tick-Borne Encephalitis Flavivirus and In Vitro Assembly of Capsid-Like Particles
 JOURNAL OF VIROLOGY, Aug. 2004, p. 8078 8084 Vol. 78, No. 15 0022-538X/04/$08.00 0 DOI: 10.1128/JVI.78.15.8078 8084.2004 Copyright 2004, American Society for Microbiology. All Rights Reserved. Isolation
JOURNAL OF VIROLOGY, Aug. 2004, p. 8078 8084 Vol. 78, No. 15 0022-538X/04/$08.00 0 DOI: 10.1128/JVI.78.15.8078 8084.2004 Copyright 2004, American Society for Microbiology. All Rights Reserved. Isolation
Virus Basics. General Characteristics of Viruses. Chapter 13 & 14. Non-living entities. Can infect organisms of every domain
 Virus Basics Chapter 13 & 14 General Characteristics of Viruses Non-living entities Not considered organisms Can infect organisms of every domain All life-forms Commonly referred to by organism they infect
Virus Basics Chapter 13 & 14 General Characteristics of Viruses Non-living entities Not considered organisms Can infect organisms of every domain All life-forms Commonly referred to by organism they infect
Hepatitis B Antiviral Drug Development Multi-Marker Screening Assay
 Hepatitis B Antiviral Drug Development Multi-Marker Screening Assay Background ImQuest BioSciences has developed and qualified a single-plate method to expedite the screening of antiviral agents against
Hepatitis B Antiviral Drug Development Multi-Marker Screening Assay Background ImQuest BioSciences has developed and qualified a single-plate method to expedite the screening of antiviral agents against
Introductory Virology. Ibrahim Jamfaru School of Medicine UHAS
 Introductory Virology Ibrahim Jamfaru School of Medicine UHAS Lecture outline Definition of viruses and general characteristics Structure of virus (virion) Chemical composition of viruses Virus morphology
Introductory Virology Ibrahim Jamfaru School of Medicine UHAS Lecture outline Definition of viruses and general characteristics Structure of virus (virion) Chemical composition of viruses Virus morphology
Ultrastructure of Mycoplasmatales Virus laidlawii x
 J. gen. Virol. (1972), I6, 215-22I Printed in Great Britain 2I 5 Ultrastructure of Mycoplasmatales Virus laidlawii x By JUDY BRUCE, R. N. GOURLAY, AND D. J. GARWES R. HULL* Agricultural Research Council,
J. gen. Virol. (1972), I6, 215-22I Printed in Great Britain 2I 5 Ultrastructure of Mycoplasmatales Virus laidlawii x By JUDY BRUCE, R. N. GOURLAY, AND D. J. GARWES R. HULL* Agricultural Research Council,
VIRUSES. 1. Describe the structure of a virus by completing the following chart.
 AP BIOLOGY MOLECULAR GENETICS ACTIVITY #3 NAME DATE HOUR VIRUSES 1. Describe the structure of a virus by completing the following chart. Viral Part Description of Part 2. Some viruses have an envelope
AP BIOLOGY MOLECULAR GENETICS ACTIVITY #3 NAME DATE HOUR VIRUSES 1. Describe the structure of a virus by completing the following chart. Viral Part Description of Part 2. Some viruses have an envelope
The Zombies of the Scientific Community Viruses and Other Acellular Infectious Agents. Acellular Agents
 viruses protein and nucleic acid viroids RNA virusoids RNA prions proteins The Zombies of the Scientific Community Viruses and Other Acellular Infectious Agents Acellular Agents Viruses major cause of
viruses protein and nucleic acid viroids RNA virusoids RNA prions proteins The Zombies of the Scientific Community Viruses and Other Acellular Infectious Agents Acellular Agents Viruses major cause of
Lecture Readings. Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell
 October 26, 2006 1 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
October 26, 2006 1 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
Materials and Methods , The two-hybrid principle.
 The enzymatic activity of an unknown protein which cleaves the phosphodiester bond between the tyrosine residue of a viral protein and the 5 terminus of the picornavirus RNA Introduction Every day there
The enzymatic activity of an unknown protein which cleaves the phosphodiester bond between the tyrosine residue of a viral protein and the 5 terminus of the picornavirus RNA Introduction Every day there
Virus Basics. General Characteristics of Viruses 5/9/2011. General Characteristics of Viruses. Chapter 13 & 14. Non-living entities
 Virus Basics Chapter 13 & 14 General Characteristics of Viruses Non-living entities Not considered organisms Can infect organisms of every domain All life-formsf Commonly referred to by organism they infect
Virus Basics Chapter 13 & 14 General Characteristics of Viruses Non-living entities Not considered organisms Can infect organisms of every domain All life-formsf Commonly referred to by organism they infect
<Supplemental information>
 The Structural Basis of Endosomal Anchoring of KIF16B Kinesin Nichole R. Blatner, Michael I. Wilson, Cai Lei, Wanjin Hong, Diana Murray, Roger L. Williams, and Wonhwa Cho Protein
The Structural Basis of Endosomal Anchoring of KIF16B Kinesin Nichole R. Blatner, Michael I. Wilson, Cai Lei, Wanjin Hong, Diana Murray, Roger L. Williams, and Wonhwa Cho Protein
Picornaviruses. Virion. Genome. Genes and proteins. Viruses and hosts. Diseases. Distinctive characteristics
 Picornaviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Naked icosahedral capsid (T=1) Diameter of 30 nm Genome Linear single-stranded RNA, positive
Picornaviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Naked icosahedral capsid (T=1) Diameter of 30 nm Genome Linear single-stranded RNA, positive
2) What is the difference between a non-enveloped virion and an enveloped virion? (4 pts)
 Micro 260 SFCC Spring 2010 Name: All diagrams and drawings shall be hand drawn (do not photo-copied from a publication then cut and pasted into work sheet). Do not copy other student s answers. Para phase
Micro 260 SFCC Spring 2010 Name: All diagrams and drawings shall be hand drawn (do not photo-copied from a publication then cut and pasted into work sheet). Do not copy other student s answers. Para phase
Chapter 19: Viruses. 1. Viral Structure & Reproduction. 2. Bacteriophages. 3. Animal Viruses. 4. Viroids & Prions
 Chapter 19: Viruses 1. Viral Structure & Reproduction 2. Bacteriophages 3. Animal Viruses 4. Viroids & Prions 1. Viral Structure & Reproduction Chapter Reading pp. 393-396 What exactly is a Virus? Viruses
Chapter 19: Viruses 1. Viral Structure & Reproduction 2. Bacteriophages 3. Animal Viruses 4. Viroids & Prions 1. Viral Structure & Reproduction Chapter Reading pp. 393-396 What exactly is a Virus? Viruses
Week 5 Section. Junaid Malek, M.D.
 Week 5 Section Junaid Malek, M.D. HIV: Anatomy Membrane (partiallystolen from host cell) 2 Glycoproteins (proteins modified by added sugar) 2 copies of RNA Capsid HIV Genome Encodes: Structural Proteins
Week 5 Section Junaid Malek, M.D. HIV: Anatomy Membrane (partiallystolen from host cell) 2 Glycoproteins (proteins modified by added sugar) 2 copies of RNA Capsid HIV Genome Encodes: Structural Proteins
The protein stoichiometry of viral capsids via tiling theory
 The protein stoichiometry of viral capsids via tiling theory REIDUN TWAROCK Centre for Mathematical Science City University Northampton Square, London EC1V 0HB UNITED KINGDOM Abstract: - A vital part of
The protein stoichiometry of viral capsids via tiling theory REIDUN TWAROCK Centre for Mathematical Science City University Northampton Square, London EC1V 0HB UNITED KINGDOM Abstract: - A vital part of
Introduction. Biochemistry: It is the chemistry of living things (matters).
 Introduction Biochemistry: It is the chemistry of living things (matters). Biochemistry provides fundamental understanding of the molecular basis for the function and malfunction of living things. Biochemistry
Introduction Biochemistry: It is the chemistry of living things (matters). Biochemistry provides fundamental understanding of the molecular basis for the function and malfunction of living things. Biochemistry
Chair of Medical Biology, Microbiology, Virology, and Immunology STRUCTURE, CLASSIFICATION AND PHYSIOLOGY OF VIRUSES
 Chair of Medical Biology, Microbiology, Virology, and Immunology STRUCTURE, CLASSIFICATION AND PHYSIOLOGY OF VIRUSES Viruses are small obligate intracellular parasites, which by definition contain either
Chair of Medical Biology, Microbiology, Virology, and Immunology STRUCTURE, CLASSIFICATION AND PHYSIOLOGY OF VIRUSES Viruses are small obligate intracellular parasites, which by definition contain either
Chapter 19: Viruses. 1. Viral Structure & Reproduction. What exactly is a Virus? 11/7/ Viral Structure & Reproduction. 2.
 Chapter 19: Viruses 1. Viral Structure & Reproduction 2. Bacteriophages 3. Animal Viruses 4. Viroids & Prions 1. Viral Structure & Reproduction Chapter Reading pp. 393-396 What exactly is a Virus? Viruses
Chapter 19: Viruses 1. Viral Structure & Reproduction 2. Bacteriophages 3. Animal Viruses 4. Viroids & Prions 1. Viral Structure & Reproduction Chapter Reading pp. 393-396 What exactly is a Virus? Viruses
October 26, Lecture Readings. Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell
 October 26, 2006 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
October 26, 2006 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
Dr. Ahmed K. Ali Attachment and entry of viruses into cells
 Lec. 6 Dr. Ahmed K. Ali Attachment and entry of viruses into cells The aim of a virus is to replicate itself, and in order to achieve this aim it needs to enter a host cell, make copies of itself and
Lec. 6 Dr. Ahmed K. Ali Attachment and entry of viruses into cells The aim of a virus is to replicate itself, and in order to achieve this aim it needs to enter a host cell, make copies of itself and
Medical Virology. Herpesviruses, Orthomyxoviruses, and Retro virus. - Herpesviruses Structure & Composition: Herpesviruses
 Medical Virology Lecture 2 Asst. Prof. Dr. Dalya Basil Herpesviruses, Orthomyxoviruses, and Retro virus - Herpesviruses Structure & Composition: Herpesviruses Enveloped DNA viruses. All herpesviruses have
Medical Virology Lecture 2 Asst. Prof. Dr. Dalya Basil Herpesviruses, Orthomyxoviruses, and Retro virus - Herpesviruses Structure & Composition: Herpesviruses Enveloped DNA viruses. All herpesviruses have
ELECTRON MICROSCOPIC STUDIES ON EQUINE ENCEPHALOSIS VIRUS
 Onderstepoort]. vet. Res. 40 (2), 53-58 (1973) ELECTRON MICROSCOPIC STUDIES ON EQUINE ENCEPHALOSIS VIRUS G. LECATSAS, B. J. ERASMUS and H. J. ELS, Veterinary Research Institute, Onderstepoort ABSTRACT
Onderstepoort]. vet. Res. 40 (2), 53-58 (1973) ELECTRON MICROSCOPIC STUDIES ON EQUINE ENCEPHALOSIS VIRUS G. LECATSAS, B. J. ERASMUS and H. J. ELS, Veterinary Research Institute, Onderstepoort ABSTRACT
the world and viruses
 More than 5,450 viruses belonging to more than 2,000 species, 287 genera, 73 families and 3 orders are recognized in the 8th ICTVreport report. the world and viruses 1 1889 H2N2 Emerging viruses in the
More than 5,450 viruses belonging to more than 2,000 species, 287 genera, 73 families and 3 orders are recognized in the 8th ICTVreport report. the world and viruses 1 1889 H2N2 Emerging viruses in the
Tivadar Orban, Beata Jastrzebska, Sayan Gupta, Benlian Wang, Masaru Miyagi, Mark R. Chance, and Krzysztof Palczewski
 Structure, Volume Supplemental Information Conformational Dynamics of Activation for the Pentameric Complex of Dimeric G Protein-Coupled Receptor and Heterotrimeric G Protein Tivadar Orban, Beata Jastrzebska,
Structure, Volume Supplemental Information Conformational Dynamics of Activation for the Pentameric Complex of Dimeric G Protein-Coupled Receptor and Heterotrimeric G Protein Tivadar Orban, Beata Jastrzebska,
10/13/11. Cell Theory. Cell Structure
 Cell Structure Grade 12 Biology Cell Theory All organisms are composed of one or more cells. Cells are the smallest living units of all living organisms. Cells arise only by division of a previously existing
Cell Structure Grade 12 Biology Cell Theory All organisms are composed of one or more cells. Cells are the smallest living units of all living organisms. Cells arise only by division of a previously existing
Chapter 13B: Animal Viruses
 Chapter 13B: Animal Viruses 1. Overview of Animal Viruses 2. DNA Viruses 3. RNA Viruses 4. Prions 1. Overview of Animal Viruses Life Cycle of Animal Viruses The basic life cycle stages of animal viruses
Chapter 13B: Animal Viruses 1. Overview of Animal Viruses 2. DNA Viruses 3. RNA Viruses 4. Prions 1. Overview of Animal Viruses Life Cycle of Animal Viruses The basic life cycle stages of animal viruses
number Done by Corrected by Doctor Ashraf
 number 4 Done by Nedaa Bani Ata Corrected by Rama Nada Doctor Ashraf Genome replication and gene expression Remember the steps of viral replication from the last lecture: Attachment, Adsorption, Penetration,
number 4 Done by Nedaa Bani Ata Corrected by Rama Nada Doctor Ashraf Genome replication and gene expression Remember the steps of viral replication from the last lecture: Attachment, Adsorption, Penetration,
Basic Properties of Viruses and Virus Cell Interaction
 WBV5 6/27/03 10:28 PM Page 49 Basic Properties of Viruses and Virus Cell Interaction II PART VIRUS STRUCTURE AND CLASSIFICATION CLASSIFICATION SCHEMES THE BEGINNING AND END OF THE VIRUS REPLICATION CYCLE
WBV5 6/27/03 10:28 PM Page 49 Basic Properties of Viruses and Virus Cell Interaction II PART VIRUS STRUCTURE AND CLASSIFICATION CLASSIFICATION SCHEMES THE BEGINNING AND END OF THE VIRUS REPLICATION CYCLE
WHY? Viruses are considered non-living because they do:
 Viruses What is a Virus? Non-living particle WHY? Viruses are considered non-living because they do: NOT Carry out metabolism NOT Grow or develop NOT Replicate without the help of a living cell (host).
Viruses What is a Virus? Non-living particle WHY? Viruses are considered non-living because they do: NOT Carry out metabolism NOT Grow or develop NOT Replicate without the help of a living cell (host).
PMT. Contains ribosomes attached to the endoplasmic reticulum. Genetic material consists of linear chromosomes. Diameter of the cell is 1 µm
 1. (a) Complete each box in the table, which compares a prokaryotic and a eukaryotic cell, with a tick if the statement is correct or a cross if it is incorrect. Prokaryotic cell Eukaryotic cell Contains
1. (a) Complete each box in the table, which compares a prokaryotic and a eukaryotic cell, with a tick if the statement is correct or a cross if it is incorrect. Prokaryotic cell Eukaryotic cell Contains
Herpesviruses. Virion. Genome. Genes and proteins. Viruses and hosts. Diseases. Distinctive characteristics
 Herpesviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Enveloped icosahedral capsid (T=16), diameter 125 nm Diameter of enveloped virion 200 nm Capsid
Herpesviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Enveloped icosahedral capsid (T=16), diameter 125 nm Diameter of enveloped virion 200 nm Capsid
LESSON 1.4 WORKBOOK. Viral sizes and structures. Workbook Lesson 1.4
 Eukaryotes organisms that contain a membrane bound nucleus and organelles. Prokaryotes organisms that lack a nucleus or other membrane-bound organelles. Viruses small, non-cellular (lacking a cell), infectious
Eukaryotes organisms that contain a membrane bound nucleus and organelles. Prokaryotes organisms that lack a nucleus or other membrane-bound organelles. Viruses small, non-cellular (lacking a cell), infectious
Virology Introduction. Definitions. Introduction. Structure of virus. Virus transmission. Classification of virus. DNA Virus. RNA Virus. Treatment.
 DEVH Virology Introduction Definitions. Introduction. Structure of virus. Virus transmission. Classification of virus. DNA Virus. RNA Virus. Treatment. Definitions Virology: The science which study the
DEVH Virology Introduction Definitions. Introduction. Structure of virus. Virus transmission. Classification of virus. DNA Virus. RNA Virus. Treatment. Definitions Virology: The science which study the
Influenza viruses. Virion. Genome. Genes and proteins. Viruses and hosts. Diseases. Distinctive characteristics
 Influenza viruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Enveloped particles, quasi-spherical or filamentous Diameter 80-120 nm Envelope is derived
Influenza viruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Enveloped particles, quasi-spherical or filamentous Diameter 80-120 nm Envelope is derived
Virology Journal. Open Access. Abstract. BioMed Central
 Virology Journal BioMed Central Research Stimulation of poliovirus RNA synthesis and virus maturation in a HeLa cell-free in vitro translation-rna replication system by viral protein 3CD pro David Franco
Virology Journal BioMed Central Research Stimulation of poliovirus RNA synthesis and virus maturation in a HeLa cell-free in vitro translation-rna replication system by viral protein 3CD pro David Franco
Supplementary Information
 Supplementary Information HBV maintains electrostatic homeostasis by modulating negative charges from phosphoserine and encapsidated nucleic acids Authors: Pei-Yi Su 1,2,3, Ching-Jen Yang 2, Tien-Hua Chu
Supplementary Information HBV maintains electrostatic homeostasis by modulating negative charges from phosphoserine and encapsidated nucleic acids Authors: Pei-Yi Su 1,2,3, Ching-Jen Yang 2, Tien-Hua Chu
Department of Molecular and Structural Biochemistry, North Carolina State University, Raleigh, North Carolina 27695
 JOURNAL OF VIROLOGY, June 2005, p. 7682 7697 Vol. 79, No. 12 0022-538X/05/$08.00 0 doi:10.1128/jvi.79.12.7682 7697.2005 Copyright 2005, American Society for Microbiology. All Rights Reserved. Single Amino
JOURNAL OF VIROLOGY, June 2005, p. 7682 7697 Vol. 79, No. 12 0022-538X/05/$08.00 0 doi:10.1128/jvi.79.12.7682 7697.2005 Copyright 2005, American Society for Microbiology. All Rights Reserved. Single Amino
VIRUS TAXONOMY AND REPLICATION
 VIRUS TAXONOMY AND REPLICATION Paulo Eduardo Brandão, PhD Department of Preventive Veterinary Medicine and Animal Health School of Veterinary Medicine University of São Paulo, Brazil I. VIRUS STRUCTURE
VIRUS TAXONOMY AND REPLICATION Paulo Eduardo Brandão, PhD Department of Preventive Veterinary Medicine and Animal Health School of Veterinary Medicine University of São Paulo, Brazil I. VIRUS STRUCTURE
Size nm m m
 1 Viral size and organization Size 20-250nm 0.000000002m-0.000000025m Virion structure Capsid Core Acellular obligate intracellular parasites Lack organelles, metabolic activities, and reproduction Replicated
1 Viral size and organization Size 20-250nm 0.000000002m-0.000000025m Virion structure Capsid Core Acellular obligate intracellular parasites Lack organelles, metabolic activities, and reproduction Replicated
Supplementary Information
 Supplementary Information Structural basis of improved second generation 3-nitro-tyrosine trna synthetases Richard B. Cooley, Jessica L. Feldman, Camden M. Driggers, Taylor Bundy, Audrey L. Stokes, P.
Supplementary Information Structural basis of improved second generation 3-nitro-tyrosine trna synthetases Richard B. Cooley, Jessica L. Feldman, Camden M. Driggers, Taylor Bundy, Audrey L. Stokes, P.
Microbiology Chapter 7 Viruses
 Microbiology Chapter 7 Viruses 7:1 Viral Structure and Classification VIRUS: a biological particle composed of genetic material (DNA or RNA) encased in a protein coat CAPSID: protein coat surrounding a
Microbiology Chapter 7 Viruses 7:1 Viral Structure and Classification VIRUS: a biological particle composed of genetic material (DNA or RNA) encased in a protein coat CAPSID: protein coat surrounding a
Thursday, October 16 th
 Thursday, October 16 th Good morning. Those of you needing to take the Enzymes and Energy Quiz will start very soon. Students who took the quiz Wednesday: Please QUIETLY work on the chapter 6 reading guide.
Thursday, October 16 th Good morning. Those of you needing to take the Enzymes and Energy Quiz will start very soon. Students who took the quiz Wednesday: Please QUIETLY work on the chapter 6 reading guide.
brought to you by and REFERENCES
 brought to you by www.thebacteriophages.org and www.phage.org REFERENCES 1. Butcher, S. J., T. Dokland, P. M. Ojala, D. H. Bamford, and S. D. Fuller. 1997. Intermediates in the assembly pathway of the
brought to you by www.thebacteriophages.org and www.phage.org REFERENCES 1. Butcher, S. J., T. Dokland, P. M. Ojala, D. H. Bamford, and S. D. Fuller. 1997. Intermediates in the assembly pathway of the
Chapter 25. 바이러스 (The Viruses)
 Chapter 25 바이러스 (The Viruses) Generalized Structure of Viruses 2 2 Virus Classification Classification based on numerous characteristics Nucleic acid type Presence or absence of envelope Capsid symmetry
Chapter 25 바이러스 (The Viruses) Generalized Structure of Viruses 2 2 Virus Classification Classification based on numerous characteristics Nucleic acid type Presence or absence of envelope Capsid symmetry
Introduction to viruses. BIO 370 Ramos
 Introduction to viruses BIO 370 Ramos 1 2 General Structure of Viruses Size range most
Introduction to viruses BIO 370 Ramos 1 2 General Structure of Viruses Size range most
Dr. Gary Mumaugh. Viruses
 Dr. Gary Mumaugh Viruses Viruses in History In 1898, Friedrich Loeffler and Paul Frosch found evidence that the cause of foot-and-mouth disease in livestock was an infectious particle smaller than any
Dr. Gary Mumaugh Viruses Viruses in History In 1898, Friedrich Loeffler and Paul Frosch found evidence that the cause of foot-and-mouth disease in livestock was an infectious particle smaller than any
numbe r Done by Corrected by Doctor
 numbe r 5 Done by Mustafa Khader Corrected by Mahdi Sharawi Doctor Ashraf Khasawneh Viral Replication Mechanisms: (Protein Synthesis) 1. Monocistronic Method: All human cells practice the monocistronic
numbe r 5 Done by Mustafa Khader Corrected by Mahdi Sharawi Doctor Ashraf Khasawneh Viral Replication Mechanisms: (Protein Synthesis) 1. Monocistronic Method: All human cells practice the monocistronic
Some living things are made of ONE cell, and are called. Other organisms are composed of many cells, and are called. (SEE PAGE 6)
 Section: 1.1 Question of the Day: Name: Review of Old Information: N/A New Information: We tend to only think of animals as living. However, there is a great diversity of organisms that we consider living
Section: 1.1 Question of the Day: Name: Review of Old Information: N/A New Information: We tend to only think of animals as living. However, there is a great diversity of organisms that we consider living
Recombinant Protein Expression Retroviral system
 Recombinant Protein Expression Retroviral system Viruses Contains genome DNA or RNA Genome encased in a protein coat or capsid. Some viruses have membrane covering protein coat enveloped virus Ø Essential
Recombinant Protein Expression Retroviral system Viruses Contains genome DNA or RNA Genome encased in a protein coat or capsid. Some viruses have membrane covering protein coat enveloped virus Ø Essential
Last time we talked about the few steps in viral replication cycle and the un-coating stage:
 Zeina Al-Momani Last time we talked about the few steps in viral replication cycle and the un-coating stage: Un-coating: is a general term for the events which occur after penetration, we talked about
Zeina Al-Momani Last time we talked about the few steps in viral replication cycle and the un-coating stage: Un-coating: is a general term for the events which occur after penetration, we talked about
Structural Requirements for Low-pH-Induced Rearrangements in the Envelope Glycoprotein of Tick-Borne Encephalitis Virus
 JOURNAL OF VIROLOGY, Nov. 1996, p. 8142 8147 Vol. 70, No. 11 0022-538X/96/$04.00 0 Copyright 1996, American Society for Microbiology Structural Requirements for Low-pH-Induced Rearrangements in the Envelope
JOURNAL OF VIROLOGY, Nov. 1996, p. 8142 8147 Vol. 70, No. 11 0022-538X/96/$04.00 0 Copyright 1996, American Society for Microbiology Structural Requirements for Low-pH-Induced Rearrangements in the Envelope
Questions in Cell Biology
 Name: Questions in Cell Biology Directions: The following questions are taken from previous IB Final Papers on the subject of cell biology. Answer all questions. This will serve as a study guide for the
Name: Questions in Cell Biology Directions: The following questions are taken from previous IB Final Papers on the subject of cell biology. Answer all questions. This will serve as a study guide for the
Capturing a flavivirus pre-fusion intermediate
 Washington University School of Medicine Digital Commons@Becker Open Access Publications 2009 Capturing a flavivirus pre-fusion intermediate Barbel Kaufmann Purdue University Paul R. Chipman Purdue University
Washington University School of Medicine Digital Commons@Becker Open Access Publications 2009 Capturing a flavivirus pre-fusion intermediate Barbel Kaufmann Purdue University Paul R. Chipman Purdue University
Chapter13 Characterizing and Classifying Viruses, Viroids, and Prions
 Chapter13 Characterizing and Classifying Viruses, Viroids, and Prions 11/20/2017 MDufilho 1 Characteristics of Viruses Viruses Minuscule, acellular, infectious agent having either DNA or RNA Cause infections
Chapter13 Characterizing and Classifying Viruses, Viroids, and Prions 11/20/2017 MDufilho 1 Characteristics of Viruses Viruses Minuscule, acellular, infectious agent having either DNA or RNA Cause infections
An Externalized Polypeptide Partitions between Two Distinct Sites on Genome-Released Poliovirus Particles
 REFERENCES CONTENT ALERTS An Externalized Polypeptide Partitions between Two Distinct Sites on Genome-Released Poliovirus Particles Jun Lin, Naiqian Cheng, Marie Chow, David J. Filman, Alasdair C. Steven,
REFERENCES CONTENT ALERTS An Externalized Polypeptide Partitions between Two Distinct Sites on Genome-Released Poliovirus Particles Jun Lin, Naiqian Cheng, Marie Chow, David J. Filman, Alasdair C. Steven,
Encapsidation of Sendai Virus Genome RNAs by Purified
 JOURNAL OF VIROLOGY, Mar. 1988, p. 834-838 22-538X/88/3834-5$2./ Copyright C) 1988, American Society for Microbiology Vol. 62, No. 3 Encapsidation of Sendai Virus Genome RNAs by Purified NP Protein during
JOURNAL OF VIROLOGY, Mar. 1988, p. 834-838 22-538X/88/3834-5$2./ Copyright C) 1988, American Society for Microbiology Vol. 62, No. 3 Encapsidation of Sendai Virus Genome RNAs by Purified NP Protein during
capsomeres: direct evidence from cryo-electron-microscopy
 The capsid of small papova viruses contains 72 pentameric capsomeres: direct evidence from cryo-electron-microscopy of simian virus 40 Timothy S. Baker,* Jacqueline Drak,* and Minou Bina* *Department of
The capsid of small papova viruses contains 72 pentameric capsomeres: direct evidence from cryo-electron-microscopy of simian virus 40 Timothy S. Baker,* Jacqueline Drak,* and Minou Bina* *Department of
Macromolecule stations. 6 stations
 Macromolecule stations 6 stations 1. Sugar and protein paper pieces to build (with waters) 2. Fatty acid and nucleic acid paper pieces to build with (and water) 3. DNA model with several pieces removed
Macromolecule stations 6 stations 1. Sugar and protein paper pieces to build (with waters) 2. Fatty acid and nucleic acid paper pieces to build with (and water) 3. DNA model with several pieces removed
