In vitro analysis of factors involved in the disassembly of Sindbis virus cores by 60S ribosomal subunits identifies a possible role of low ph
|
|
|
- Delphia Parker
- 5 years ago
- Views:
Transcription
1 Journal of General Virology (2002), 83, Printed in Great Britain... In vitro analysis of factors involved in the disassembly of Sindbis virus cores by 60S ribosomal subunits identifies a possible role of low ph Gerd Wengler and Gisela Wengler Institut fu r Virologie der Veterina rmedizin, Justus-Liebig-Universita t Giessen, Giessen, Germany Disassembly of alphavirus cores early in infection involves interaction of the core with 60S ribosomal subunits. This interaction might be subjected to regulatory processes. We have established an in vitro system of core disassembly in order to identify cellular proteins involved in the regulation of disassembly. No evidence for the existence of such proteins was found, but it became apparent that certain organic solvents and detergents or a high proton concentration (ph 6 0) stimulated core disassembly. Alphaviruses infect cells by an endosomal pathway. The low ph in the endosome activates a fusion activity of the viral surface protein E1 and leads to fusion of the viral membrane with the endosomal membrane, followed by release of the core into the cytoplasm. Since the presence of the E1 protein in the plasma membrane of infected cells leads to increased membrane permeability at low ph, our findings indicate that disassembly of alphavirus cores could be regulated by the proton concentration. We propose that the viral membrane proteins present in the endosomal membrane after fusion form a pore, which allows the flow of protons from the endosome into the cytoplasm. This process would generate a region of low ph in the cytoplasm at the correct time and place to allow the efficient disassembly of the incoming viral core by 60S subunits. Introduction Alphaviruses are enveloped positive-strand RNA viruses, which constitute a genus in the Togaviridae family (van Regenmortel et al., 2000). The structure and replication of alphaviruses have been recently reviewed (Strauss & Strauss, 1994; Schlesinger & Schlesinger, 2001). The core of alphaviruses consists of the viral genome and 240 molecules of the viral core protein, which are organized into a T 4 icosahedral complex of about 40 nm in diameter (Choi et al., 1991; Paredes et al., 1992). The core accumulates in large amounts in the cytoplasm of infected cells. Mature virus is formed by a budding process in which the cytoplasmic core is enveloped by a modified cellular membrane containing the viral spike proteins (see Simons & Garoff, 1980, for a review). Alphaviruses infect cells by an endosomal pathway (see Marsh & Helenius, 1989, for a review). At ph 5 8, a reorganization of the spike proteins in the endosome leads to fusion of the viral Author for correspondence: Gerd Wengler. Fax gerd.wengler gmx.de and endosomal membranes, followed by exposure of the core to the cytoplasm (Wahlberg et al., 1992; Bron et al., 1993; Justmann et al., 1993). This description leads to the question of how alphavirus cores are disassembled early in infection. Alphavirus core protein associates with the large ribosomal subunit during synthesis prior to incorporation into viral cores (Ulmanen et al., 1976, 1979) and purified core protein binds to the large ribosomal subunit in vitro (Wengler et al., 1984). Exogeneously added cores are disassembled in vitro in cell lysates with a concomitant transfer of core protein to the large ribosomal subunit (Wengler et al., 1992), and a similar transfer of viral core protein to 60S ribosomal subunits has been detected in vivo (Wengler & Wengler, 1984; Singh & Helenius, 1992). These data have led us to propose that alphavirus cores are unstable in the presence of ribosomes early in infection, because a ribosome binding site is exposed on the core surface, leading to an interaction of the cores with ribosomes, followed by transfer of core protein to ribosomes, whereas during virus multiplication, newly synthesized core protein binds to ribosomes and inactivates their ability to disassemble cores (Wengler, 1987) SGM CEBH
2 G. Wengler and G. Wengler However, experiments showing that the saturation of ribosomes with core protein during alphavirus replication cannot represent the sole mechanism of regulation of core disassembly have been reported (Singh et al., 1997). BHK cells were infected with a recombinant Semliki Forest (SF) virus, which did not contain the coding region for the viral structural proteins, followed 6 h later by superinfection with [ S]methionine-labelled SF virus. Radioactively labelled cores were detected in the cytoplasm of these cells without disassembly. These results might be explained by the assumption that cellular factors are involved in the reaction between the incoming viral core and ribosomes, and that these factors are inactivated during infection. We have therefore made an attempt to identify such factors. We have established in vitro systems in which the interaction of the cores of the alphaviruses Sindbis (SIN) virus and SF virus with ribosomes have been analysed in a systematic manner. We found no evidence for the involvement of cellular proteins in the regulation of disassembly. On the other hand, the data obtained indicated that in the presence of appropriate solvents or detergents, or at low ph, the SIN virus core rapidly disassembled by interaction with the 60S ribosomal subunit. These experiments and their possible implications for the regulation of disassembly of alphavirus cores are described. Methods Preparation of viral cores. Cores of SIN or SF virus were isolated from virus labelled with [ S]methionine and [ S]cysteine and purified as described by Boege et al. (1980). For the isolation of cores, virus was suspended in TN buffer (20 mm triethanolamine, ph 7 4, 100 mm NaCl) and incubated in the presence of 1% NP-40 at 20 C for 10 min, followed by centrifugation at 4 C on 5 20% (w v) sucrose density gradients in TN buffer for 25 min at r.p.m. in an SW55 Beckman rotor. The distribution of radioactivity and optical density was determined, and fractions containing cores were identified. These fractions were stored on ice and the cores were used within 48 h. Preparation of the S30 supernatant and cytoplasmic proteins. One vol. of packed BHK-21 cells was suspended in 1 5 vols of RSB buffer (10 mm Tris HCl, ph 7 4, 10 mm NaCl, 1 mm MgCl )on ice. After incubation for 5 min at 0 C, glass beads (0 3 mm diameter) were added and the cells were disrupted by vortexing. The homogenate was subjected to centrifugation for 10 min at r.p.m. in an SS34 Sorvall rotor at 4 C. The S30 supernatant obtained was stored in aliquots at 160 C. Cytoplasmic proteins were isolated from the S30 supernatant after removal of the ribosomes by centrifugation. Aliquots of the resulting S100 material were subjected to ammonium sulphate precipitation by addition of 100% ammonium sulphate to 40, 60 or 80% saturation. After incubation for 2 h at 0 C, the precipitated protein was recovered by centrifugation for 20 min at g, dialysed into TN buffer and stored in aliquots at 160 C. Preparation of ribosomal subunits and rrna. For preparation of ribosomal subunits, the S30 supernatant was adjusted to 2 mm EDTA, ph 7 4, and subjected to gradient centrifugation using the conditions described above for the preparation of cores. Typically, cores and ribosomal subunits for each experiment were prepared in parallel in the same centrifugation. Ribosomal subunits were localized from the optical density profile of the gradient, stored on ice and used within the next 48 h. Ribosomal RNA was purified from a ribosome pellet by phenol extraction, followed by sucrose density-gradient centrifugation. Gradients used for the preparation of cores and ribosomes were also used for the fractionation of 28S and 18S rrna, but the time of centrifugation was extended to 4 h. Preparation of protein synthesis initiation factors. Initiation factors for protein synthesis were prepared by extraction of ribosomes with 500 mm KCl, as described by Barton et al. (1996). The minimal disassembly reaction and its use to analyse the effect of solvents or detergents. In a typical minimal disassembly reaction, 10 µlof S-labelled viral cores, 90 µl of ribosomal subunits and 100 µl of BSA (10 mg ml) in 300 mm NaCl, 20 mm triethanolamine, ph 7 4, were prepared at 0 C. The final reaction contained 20 mm triethanolamine, ph 7 4, 200 mm NaCl, 5 mg ml BSA and approximately 5% sucrose. No detectable disassembly occurred under these conditions at 0 C. Core disassembly was performed by incubation at 30 C. The reaction was stopped by the addition of 1 vol. of 10 mm Tris HCl, ph 8 1, 2 mm EDTA, on ice. The reaction volume was loaded on to a 15 40% (w v) sucrose density gradient in TN buffer and subjected to centrifugation for 80 min at r.p.m. at 4 C inan SW55 Beckman rotor, followed by analysis of the optical density and radioactivity profile. In experiments in which the increase in the reaction volume inherent in this procedure was undesirable, the same salt conditions were obtained in a volume of 100 µl by assembling 10 µl of cores, 80 µl of ribosomal subunits, 5 µl of 2 M NaCl and 5 µl of BSA (100 mg ml). For analysis of the effect of solvents or detergents, 80 µlof ribosomal subunits, 10 µl of viral cores and 100 µl of BSA (10 mg ml) in 300 mm NaCl, 20 mm triethanolamine, ph 7 4, were mixed on ice. Ten µl of solvent, or 10 µl of a solution of 10% (w v) detergent in water, or 10 µl of water as a control, were added at 0 C, and the reactions were incubated and analysed as described above. Analysis of the effect of proteins on the minimal disassembly reaction. Reactions were carried out as follows: 76 µl of ribosomal subunits, 10 µl of cores, 5 µl of 2 M NaCl, 1 µl of 100 mm ATP, 1 µl of 100 mm GTP, 2 µl of 100 mm MgCl and 5 µl of the protein concentrate to be analysed were prepared at 0 C. The control reaction contained 5 µl of BSA (10 mg ml) in the buffer used for the protein concentrate. Reactions were analysed as described above. Analysis of the effect of ph on the minimal disassembly reaction. A standard minimal reaction containing 1 mm triethanolamine, ph 7 4, instead of 20 mm, was assembled. To this end, ribosomes and cores were isolated from sucrose gradients containing 100 mm NaCl and 1 mm triethanolamine, ph 7 4. The reactions were then carried out as described above: 10 µl of S-labelled cores in approximately 10% sucrose in 100 mm NaCl, 1 mm triethanolamine, ph 7 4, and 80 µl of ribosomes in the same buffer were mixed at 0 C and 5 µl of 2 M NaCl and 5 µl of BSA (100 mg ml) in water were added to obtain concentrations of 200 mm NaCl and 5 mg ml BSA. The ph was modified by the addition of 4 µl of 500 mm MES phs of 5 8, 6 0, 6 2, 6 4, 6 6, 6 8 or7 0. The final ph of the reaction did not differ significantly from the ph of the MES buffer. The reaction was incubated at 25 C and stopped by addition of 300 µl of 80 mm Tris HCl, ph 8 1, 1 mm EDTA, at 0 C. The reaction products were analysed as described for the minimal reaction performed at neutral ph. Results Disassembly of cores in the minimal reaction Disassembly of alphavirus cores occurs in vitro in a system containing cores and purified 60S ribosomal subunits in the CEBI
3 Disassembly of Sindbis virus cores at low ph Fig. 1. Disassembly of cores in the minimal reaction. The standard minimal reaction containing 35 S-labelled SIN virus cores and 60S ribosomal subunits was prepared at 0 C and incubated at 30 C, followed by analysis of the reaction products by sucrose-density-gradient centrifugation (see Methods). The optical density profile of the gradients allowed localization of the ribosomal subunits, and the radioactivity profile allowed localization of the 35 S-labelled core protein, as shown. The bottom of the gradient is to the left. The positions of the 150S intact viral core and of the 60S and 40S ribosomal subunits are indicated in (A). The products generated in a standard minimal reaction, incubated at 0 C for 30 min or at 30 C for 6, 12 and 24 min are presented in (A) (D), respectively. The products generated from a minimal reaction incubated for 12 min at 37 C and from a control reaction without 60S ribosomal subunits incubated at 37 C for 24 min are presented in (E) and (F), respectively. The products generated during a 12 min incubation at 30 C in a modified minimal reaction containing either an increased NaCl concentration of 300 mm, or MgCl 2 (2 mm) and ATP (1 mm) as additional components, are presented in (G) and (H), respectively. absence of additional components (Wengler et al., 1996). We decided to analyse this system, which we called the minimal reaction system, in order to evaluate a possible stimulating activity of cellular molecules. In the minimal reaction system chosen, [ S]methionine-labelled cores and 60S ribosomal subunits were incubated together as described in Methods, and the reaction products were analysed by gradient centrifugation. Some of these analyses are presented in Fig. 1. It can be seen that at 0 C no disassembly of cores occurred (Fig. 1A), whereas at 30 C (Fig. 1B D) after 24 min incubation, about one-third of the cores had disassembled. Increasing the temperature to 37 C (Fig. 1E) or the salt concentration to 300 mm NaCl (Fig. 1G) greatly accelerated the reaction. Addition of MgCl (2 mm) and ATP (1 mm) led to a small stimulation of the reaction (Fig. 1H). In many of the assays for a possible stimulating activity of cellular proteins, MgCl and nucleoside triphosphates had to be present. An important control (Fig. 1F) showed that in the absence of 60S ribosomal subunits the cores were stable in the minimal reaction for 24 min, even at 37 C. Kinetic analysis of core disassembly in the minimal reaction allowed determination of the influence of the concentration of 60S ribosomal subunits or cores on the rate of disassembly. In the assays presented in Fig. 1, 20 µg of 28S RNA were present in each reaction as acceptor, as part of the 60S subunit. It can be seen that the sensitivity of the gradient analysis allowed analysis of reactions containing between 4 µg and 80 µg of RNA in subunits and a similar 20-fold variation in the concentration of radioactively labelled cores. These experiments showed that the rate of disassembly varied linearly with the concentration of both the 60S ribosomal subunits and the cores (data not shown). The disassembly of cores in the minimal reaction is therefore a first-order reaction for both the ribosomal subunits and the cores. The rrna is exposed extensively on the surface of the ribosomal subunits (Ban et al., 2000). We therefore analysed the CEBJ
4 G. Wengler and G. Wengler Fig. 2. Disassembly of SIN virus cores in a minimal reaction containing rrna as acceptor. Minimal reactions containing RNA instead of ribosomes were prepared at 0 C (see Methods). The reactions were incubated at 30 C and the reaction products analysed by sucrose-density-gradient centrifugation, as shown. The products generated in reactions containing 28S rrna as acceptor after incubation at 0 C for 24 min or at 30 C for 12 min or 24 min are shown in (A) (C), respectively. In these analyses 10 45% (w/v) sucrose density gradients were centrifuged for 2 h at r.p.m. in an SW55 Beckman rotor. The positions of the 150S intact viral core and the 28S rrna are indicated in (A). The products generated in a reaction containing both 28S and 18S rrna as acceptor after incubation at 30 C for 24 min are presented in (D). In this analysis the 10 45% sucrose density gradient was centrifuged for 4 h at r.p.m. in order to allow separation of the two RNA molecules. Intact cores were centrifuged into the pellet under these conditions. ability of 28S and 18S rrna to disassemble cores in the minimal reaction. Some of the data are presented in Fig. 2. The results can be compared directly with those presented in Fig. 1, because the experiments were performed in parallel using the same core preparation and identical reaction conditions. At 0 C, no disassembly was observed (Fig. 2A), whereas at 30 C, the cores disassembled with a concomitant transfer of core protein to RNA (Fig. 2B, C). A comparison of the data with the corresponding results obtained from the reaction containing ribosomal subunits (Fig. 1A, C, D) showed that no significant difference existed between the 60S subunits and the 28S rrna in their ability to disassemble the cores. A reaction containing both 28S and 18S rrna was analysed in the gradient shown in Fig. 2(D). It can be seen that the core protein associated preferentially with the 28S rrna, but that the 18S rrna also bound some core protein molecules. SIN virus cores were used in the experiments presented above, but similar results were obtained with SF virus cores (data not shown). Analysis of a possible role of cytoplasmic factors in the disassembly of cores In order to identify cytoplasmic factors that might stimulate the disassembly of cores, a small volume of concentrated cellular proteins was included in the minimal reaction. The following three protein preparations were used: (i) initiation factors of protein synthesis; (ii) casein kinase II, since the core protein of alphaviruses contains a predicted conserved phosphorylation site for casein kinase II; and (iii) a post-ribosomal supernatant derived from BHK-21 cells, which was subjected to a series of ammonium sulphate precipitations; the precipitated proteins were dialysed into the reaction buffer of the minimal reaction. The preparation of these proteins and the methods used to assess their influence on the disassembly are described in Methods. None of these protein preparations had an effect on core disassembly (data not shown). As a next step in the search for proteins that stimulate core disassembly, we analysed more complex in vitro systems. A comparison of the core disassembly in a system containing a complete post-mitochondrial BHK-21 S30 fraction and the corresponding minimal system is shown in Fig. 3. Cores were added either to the S30 fraction containing ribosomes and the cellular cytoplasmic proteins prepared in 10 mm NaCl, 10 mm Tris HCl, ph 7 4, 1 mm MgCl, or to purified 60S ribosomal subunits in the same buffer. Both samples were adjusted to the final reaction conditions as described in Fig. 3. It should be noted that the disassembly activity of this modified minimal reaction system (Fig. 3A C) did not differ significantly from the standard minimal system (Fig. 1, B D). The data presented in Fig. 3 show that the S30-containing system (Fig. 3A C ) was slightly more active in the disassembly than the reaction containing purified subunits (Fig. 3A C). The small increase in activity might result either from a biochemical process or from a physico-chemical difference between the two systems. Besides the cytoplasmic proteins, the S30 fraction contains, for example, trna molecules, which might influence core disassembly. To analyse this question, we pretreated the S30 fraction with either trypsin or the SH-group reagent N- ethylmaleimide and compared the in vitro disassembly of cores in reactions containing pretreated and untreated S30 fractions. The same rate of disassembly that was detected in Fig. 3(A C ) was found, whether the S30 fraction was pretreated CECA
5 Disassembly of Sindbis virus cores at low ph or not (data not shown). These results were obtained using both SIN virus- and SF virus-derived cores (data not shown). The use of the S30 material therefore also gave no evidence for a cellular protein that might be involved in the regulation of core disassembly. Fig. 3. Comparison of core disassembly in a minimal reaction and a reaction containing a complete S30 fraction. A post-mitochondrial S30 fraction was prepared from BHK-21 cells by mechanical lysis and centrifugation (see Methods). For each reaction containing a complete S30 fraction, 10 µl of 35 S-labelled SIN virus cores in TN buffer and 70 µl of S30 supernatant in 10 mm NaCl, 10 mm Tris HCl, ph 7 4, and 1 mm MgCl 2 were mixed on ice, and a final concentration of 200 mm NaCl, 10 mm Tris HCl, ph 7 4, 1 mm triethanolamine, 2 mm ATP, 2 mm GTP and 5 mm MgCl 2 in a reaction volume of 100 µl was established. The corresponding minimal reaction was assembled in the same way but contained 70 µl purified ribosomal subunits instead of 70 µl S30. In this experiment, ribosomal subunits were purified by gradient centrifugation in the buffer used to prepare the S30 fraction (10 mm Tris HCl, ph 7 4, 10 mm NaCl, 1 mm MgCl 2 ) and both ribosomal subunits were included in the minimal reaction. Sucrose-density-gradient analyses were performed as described in Fig. 1. The position of the 150S intact cores and the 60S and 40S ribosomal subunits are indicated in (A). The products generated in reactions containing purified ribosomal subunits incubated for 10, 20 and 40 min at 30 C are shown in (A) (C), respectively. The corresponding analyses of the reactions containing the S30 supernatant are presented in (A ) (C ), respectively. Analysis of the influence of organic solvents, detergents and low ph on the disassembly of cores In the experiments described above, S30 proteins were digested by protease. In some of these experiments, protease inhibitors solubilized in DMSO were used. The results obtained indicated that DMSO might stimulate the disassembly. We therefore analysed the effect of DMSO at 5% (v v) concentration in the minimal reaction on the disassembly of SIN cores. The results showed that the presence of DMSO greatly stimulated the core disassembly. In the absence of 60S ribosomal subunits, the cores were stable in the presence of DMSO (Fig. 4A, A, B,B ). Similar data were obtained in reactions containing 5% (v v) acetonitrile (data not shown). The S30 fraction used in the experiment reported in Fig. 3 was prepared from mechanically disrupted cells. Similar experiments were also performed using cytoplasmic proteins extracted in the presence of octylglucoside. In these analyses, the octylglucoside-extracted material was highly active in core disassembly. These findings led us to analyse the effect of detergents on disassembly in the minimal reaction. SIN cores were used in these analyses and all detergents were present at 0 5% final concentration. Representative analyses are shown in Fig. 4. It can be seen that NP-40 had no significant influence on the reaction (Fig. 4C) but that octylglucoside greatly stimulated core disassembly (Fig. 4D). Neither detergent dissociated the cores in the absence of 60S ribosomes (Fig. 4C,D ). The results of our analyses can be summarized as follows: the non-ionic detergents NP-40, Triton X-100 and reduced Triton X-100 had no significant influence on the core disassembly in the minimal reaction, whereas the non-ionic detergents hexylglucoside, octylglucoside, decylglucoside, octylmaltoside and dodecylmaltoside greatly stimulated disassembly. These detergents left the sedimentation behaviour of the core unaltered during a disassembly reaction in the absence of ribosomes. The effect of the zwitterionic sulfobetaine detergents depended on the length of the hydrophobic chain of the detergent: sulfobetaine 3-8 only slightly stimulated the reaction, whereas the sulfobetaines 3-10, 3-12 and 3-14 led to rapid disassembly in the standard assay (data not shown). In contrast to the non-ionic detergents, an effect of sulfobetaines 3-12 and 3-14 on the cores in the control reaction in the absence of ribosomes could be seen: the cores aggregated in the presence of 0 5% sulfobetaine 3-12 and were converted into a slowly sedimenting structure in the presence of 0 5% sulfobetaine 3-14 (data not shown). The data reported above show that the presence of DMSO, acetonitrile and a number of detergents greatly stimulated the disassembly of SIN cores in the minimal reaction but had no CECB
6 G. Wengler and G. Wengler Fig. 4. Influence of solvents or detergents on core disassembly in the minimal reaction. The disassembly of 35 S-labelled SIN virus cores was analysed in a minimal reaction containing solvents at 5% (v/v) concentration or detergents at 0 5% concentration. Reactions were assembled as described in Methods. All reactions were performed in parallel in the presence and absence of ribosomes. Purified 60S and 40S ribosomal subunits were included in the minimal reaction in order to identify a possible transfer of core protein to the small subunit. The reaction products were analysed by sucrose gradient centrifugation. The products generated during incubation at 30 C for 15 min in the control reaction, which contained no solvent or detergent, are shown for the complete reaction and the reaction without ribosomes in (A) and (A ), respectively. The products generated in the presence of 5% DMSO in a complete reaction and in a reaction without ribosomes are presented in (B) and (B ), respectively. The corresponding analyses of the reactions containing 0 5% NP-40 are shown in (C) and (C ), respectively, and the corresponding analyses of reactions containing 0 5% octylglucoside (OG) are shown in (D) and (D ), respectively. detectable effects on the cores in the control reaction in the absence of ribosomes. These data lead to the question of whether the cores, isolated by gradient centrifugation from the control reactions, were functionally modified by the incubation. The cores might have the same reactivity in the minimal reaction as the untreated cores, they might be activated and rapidly dissociated even in the absence of the stimulating molecule, or they might be converted into an inactive conformation that is no longer susceptible to the stimulating activity of the organic solvent or detergent used in their preparation. The results of experiments analysing this question are shown in Fig. 5. It can be seen that cores pretreated with NP-40 were not functionally modified: the NP-40-treated cores could not be activated by NP-40. This was to be expected since NP-40 does not stimulate the disassembly. This reaction can be regarded as a control reaction showing that the experiment does not lead to a non-specific activation of cores. The data presented in Fig. 5 showed that the octylglucosidetreated cores were not stably altered either structurally or functionally: they were not converted into a reactive conformation since they were not disassembled in the standard minimal reaction in the absence of detergent (Fig. 5B) but they remained sensitive to stimulation by octylglucoside (Fig. 5B ). The detergents Zwittergent 3-10 and dodecylmaltoside, which had a strong stimulating activity in the minimal reaction, had the same properties in these reactions (data not shown). The data in Fig. 5 also showed that DMSO had a different effect: the DMSO-treated cores were not significantly disassembled, either in the standard minimal reaction (Fig. 5C) or in the presence of DMSO (Fig. 5C ). DMSO treatment therefore converts the SIN core into a stable inactive conformation, which cannot be activated for disassembly by DMSO. In none of these experiments was an activated core isolated that reacted rapidly in the minimal reaction in the absence of organic solvent or detergent. The results reported above indicate that non-enzymatic processes might be involved in the regulation of core disassembly. In this situation, the finding that the presence of CECC
7 Disassembly of Sindbis virus cores at low ph which a low ph was present during the reaction between cores and 60S subunits. The results of a representative experiment are presented in Fig. 6. It can be seen that a small stimulation was observed at ph 6 4, and that at ph 6 2 the disassembly was strongly stimulated. Cores were stable in all of these reactions in the absence of 60S ribosomes: the analyses of these control reactions incubated at ph 7 0 and ph 6 0 are shown in Fig. 6(G and H) of the figure. As discussed above, these data lead to the question of whether the SIN virus cores, treated at ph 6 0 in the absence of 60S ribosomes, have an altered reactivity. Such cores can be recovered by density gradient centrifugation (Fig. 6H). The results of these analyses show that these cores have the same reactivity as the authentic viral cores that were analysed in the experiment presented in Fig. 6 (data not shown). These experiments showed that the low ph treatment had no detectable permanent effect on the structure or reactivity of the cores. The data indicate that, in order to stimulate core disassembly, the low ph has to be present during the reaction between cores and 60S ribosomes. The influence of DMSO, octylglucoside, NP-40 and low ph on the disassembly of SIN cores was also analysed using an in vitro system containing a complete rabbit reticulocyte S30 extract instead of purified 60S subunits. In this complete system, which comes much closer to a physiological situation, the same effects were observed that were described above for the minimal reaction system: DMSO, octylgucoside and low ph greatly stimulated core disassembly, whereas NP-40 had no effect (data not shown). The experiments reported in Figs 4, 5 and 6 contained SIN virus cores. SF virus cores behaved differently in reactions containing organic solvents and detergents, or at low ph. The differences will be discussed in the following section. Fig. 5. Analyses of the reactivity of cores pretreated with solvents or detergents. Pretreatment of cores was performed as follows: cores were subjected to a standard minimal reaction in the absence of ribosomes at 30 C for 12 min in the presence of the solvent or detergent being analysed (see Methods). The cores were then isolated by sucrose-densitygradient centrifugation as shown in Fig. 4(B D ). These cores were used in two parallel minimal reactions containing ribosomal subunits either with or without solvent or detergent. Gradient analyses of the reaction products generated in these two parallel reactions after 12 min incubation at 30 C are presented. Products generated when NP-40-pretreated cores were disassembled in reactions containing no NP-40 or 0 5% NP-40 are shown in (A) and (A ), respectively. Products generated when octylglucosidepretreated cores were disassembled in reactions containing no octylglucoside or 0 5% octylglucoside are shown in (B) and (B ), respectively. Products generated when DMSO-pretreated cores were disassembled in reactions containing no DMSO or 5% DMSO are shown in (C) and (C ), respectively. viral structural proteins in the membrane of alphavirus-infected cells leads to a great increase in membrane permeability at low ph (Lanzrein et al., 1993) prompted us to analyse conditions in Discussion In a search for cellular factors involved in core disassembly, the minimal reaction was used to assay three protein preparations: protein synthesis initiation factors, casein kinase II and soluble cytoplasmic proteins. The alphavirus genome is translated as mrna and the viral core does not contain a completely closed protein shell. Initiation factors of protein synthesis therefore might bind to the genome packed in the core, and this interaction might stimulate disassembly. Protein phosphorylation is used extensively in the regulation of cellular processes. Therefore the Prosite program (Hofmann et al., 1999) was used to identify possible conserved phosphorylation sites in the core protein of SIN and SF viruses. A casein kinase II acceptor site, present in both proteins, comprising the sequence S (160) AYD in the SIN virus core protein was identified. The effect of proteins precipitated with ammonium sulphate from an S100 fraction of BHK cells on core disassembly was also analysed. None of these protein preparations had a detectable effect on core disassembly; none of the reactions was significantly different from the minimal CECD
8 G. Wengler and G. Wengler Fig. 6. Influence of low ph on core disassembly in a minimal reaction. A minimal reaction containing 35 S-labelled SIN virus cores was adjusted to low ph values by addition of appropriate MES buffer, incubated at 25 C for 10 min and terminated by neutralization at 0 C as described in Methods. Purified 60S and 40S ribosomal subunits were included in the minimal reaction in order to identify a possible transfer of core protein to the small subunit. Gradient analyses of the reaction products were performed as described in Fig. 1. The products generated in reactions adjusted to phs of 7 0, 6 8, 6 6, 6 4, 6 2 and 6 0 are presented in (A) (F), respectively. The products generated in reactions at ph 7 0 and at ph 6 0 containing no ribosomes are presented in (G) and (H), respectively. reaction in the presence of ATP and Mg + (Fig. 1H and data not shown). A further search for the existence of cellular factors involved in core disassembly was performed by using a complete post-mitochondrial S30 fraction in the in vitro analyses. Core disassembly was found to be slightly faster in the reaction containing the S30 fraction than in the control reaction containing purified 60S subunits (Fig. 3). This finding might be explained either by the presence in the S30 of cellular factors involved in disassembly or by the fact that the physicochemical properties in the two reactions are different: the S30 fraction, for example, also contains trna molecules, which might have an activity in core disassembly. Further experiments were performed in which the activity of two in vitro disassembly reactions containing either untreated S30 or S30 digested with trypsin were compared. The difference between these reactions is restricted to the protein component. This experiment is possible because the 28S rrna, which is not destroyed by protease-treatment, is the active part of the ribosome in core disassembly. Trypsin treatment did not reduce the activity of the S30 fraction and similar results were obtained using an S30 fraction treated with the SH-groupspecific reagent N-ethylmaleimide. These experiments were also performed using rabbit reticulocyte S30 and similar results were obtained (data not shown). SIN virus cores were used in the experiments presented above, but many of these experiments were performed in parallel with SF virus cores and principally the same results were obtained (data not shown). In these experiments, therefore, no evidence could be found for the involvement of cellular factors in the disassembly of alphavirus cores. From the outset, we used cores derived from both SF and SIN viruses. The evolutionary distance between these viruses is rather large (Powers et al., 2001). Our assumption that in the in vitro systems the stability and or reactivity of the cores might be different, and that therefore the chances of detecting a stimulating activity might be better if both cores were used, were confirmed in our experiments. In the minimal disassembly reaction and in the search for stimulating proteins, no significant difference was observed between SF and SIN virus cores. The effect of organic solvents, detergents and low ph on the disassembly of SF virus cores was, however, very different from the corresponding effects on SIN virus cores described in CECE
9 Disassembly of Sindbis virus cores at low ph Figs 4, 5 and 6. The effect of DMSO, NP-40, octylglucoside and Zwittergent 3-12 on the disassembly of SF cores in the minimal reaction was analysed. None of these molecules had a stimulating effect (data not shown). The effect of low ph on the disassembly of SF cores was analysed, as shown for SIN virus cores in Fig. 6. No stimulation was detected, and the decrease in ph led to increasing aggregation of the SF cores in control reactions in the absence of ribosomes (data not shown). These data are in accordance with earlier findings that purified cores of SF and SIN virus react quite differently if exposed to ribonuclease, EDTA or low ph (So derlund et al., 1972, 1979). That the stimulating effects on core disassembly were observed in the minimal reaction indicated that they represented direct effects on the disassembly. The finding that the stimulating effects were also observed in a reticulocyte S30 lysate system showed that they were not an artefact of the conditions present in the minimal reaction system. The experiments reported above may therefore be used as a starting point for in vitro analysis of the mechanism of core disassembly. In this context, the following points are of importance: (i) In the minimal disassembly reaction, core protein can also be transferred to 18S rrna (Fig. 2) or to the small ribosomal subunit (Fig. 3). Furthermore, in a minimal disassembly reaction that contains only purified 18S rrna as acceptor, the cores are disassembled with a concomitant transfer of core protein to the 18S rrna (data not shown). (ii) The rate of core disassembly in the minimal reaction is linearly dependent on the concentration of cores and 60S ribosomal subunits. (iii) Treatment of SIN virus cores with activating agents in the absence of 60S subunits does not generate a stable reactive conformation of cores. These data indicate that interactions of viral cores and an RNA molecule, which is not necessarily 28S rrna, can generate a complex that is the target for the activating molecule: the presence of this molecule favours transfer of core protein on to the RNA within this complex, whereas the dissociation of the complex into the reactants is favoured in the absence of the activating molecule. Further analysis of the role of solvents, detergents and a low ph on the disassembly of SIN virus cores in the minimal reaction will probably allow characterization of the processes involved in the disassembly of alphavirus cores in more detail. What are the implications of our findings that the disassembly of SIN virus cores is stimulated by detergents, organic solvents and low ph on our attempt to identify a mechanism that might regulate the disassembly of cores by 60S subunits? The most direct interpretation is that these conditions are all non-physiological, and that therefore these data cannot be used to identify such a mechanism. The finding that the disassembly of SF virus cores is not stimulated in vitro would be in line with this interpretation. On the other hand, it is possible that the stimulation of disassembly of SIN virus cores by low ph detected above also occurs in vivo. In a series of publications, Kempf and co-workers have shown that the accumulation of alphavirus structural proteins in cellular membranes alters their permeability at low ph (see Lanzrein et al., 1993; Nyfeler et al., 2001, and references cited therein). Considering these data, we propose that, during alphavirus infection, the membrane proteins of the infecting virus particles form a pore in the endosomal membrane after fusion. The resulting flow of protons through this pore from the endosome into the cytoplasm would generate a region of low ph at the appropriate time and place to stimulate the disassembly of alphavirus cores. As indicated in the Introduction, we have analysed the mechanism of core disassembly and assembly in a number of earlier experiments. All data obtained in these experiments remain valid, but the identification of the possible role of low ph in core disassembly leads to this new proposal. The involvement of a low ph step in the disassembly of alphavirus cores does not explain why this disassembly is inhibited in alphavirus-infected cells that are superinfected 6 h post-infection, as described by Singh et al. (1997). Further experiments will be necessary to clarify this point. If the above hypothesis is correct, it seems likely that a low ph step is not only involved in the disassembly of SIN virus cores but that, in vivo, the proton flow into the cytoplasm would be involved in the disassembly of alphavirus cores in general. In the in vitro assay, viral cores purified by detergent treatment of virus particles and gradient centrifugation were used. It is not unlikely that the cores of some alphaviruses, such as SIN virus, remain in a native state during these procedures, whereas the cores of other alphaviruses, such as SF virus, are structurally altered and are converted into a non-reactive state. That alphavirus cores can adopt a non-reactive conformation is indicated by our finding that SIN virus cores, when treated with DMSO, are no longer susceptible to stimulation in vitro. These considerations lead to the prediction that the insertion of the membrane proteins of both SF and SIN virus particles into the target membrane during the early stages of virus infection generates a pore. Experimental evidence for the formation of these pores has been recently obtained by us (unpublished). This study was supported by the Deutsche Forschungsgemeinschaft (Projekt We ). We thank Mrs C. Reitz for artwork. References Ban, N., Nissen, P., Hansen, J., Moore, P. B. & Steitz, T. A. (2000). The complete atomic structure of the large ribosomal subunit at 2 4 A resolution. Science 289, Barton, D. J., Morasco, B. J. & Flanegan, J. B. (1996). Assays for poliovirus polymerase, 3D (Pol), and authentic RNA replication in HeLa S10 extracts. Methods in Enzymology 275, Boege, U., Wengler, G., Wengler, G. & Wittmann-Liebold, B. (1980). Partial amino acid sequences of Sindbis and Semliki Forest virus-specific core proteins. Virology 103, Bron, R., Wahlberg, J. M., Garoff, H. & Wilschut, J. (1993). Membrane fusion of Semliki Forest virus in a model system: correlation between fusion kinetics and structural changes in the envelope glycoprotein. EMBO Journal 12, CECF
10 G. Wengler and G. Wengler Choi, H.-K., Tong, L., Minor, W., Dumas, P., Boege, U., Rossmann, M. G. & Wengler, G. (1991). Structure of Sindbis virus core protein reveals a chymotrypsin-like serine proteinase and the organization of the virion. Nature 354, Dick, M., Barth, B. U. & Kempf, C. (1996). The E1 protein is mandatory for pore formation by Semliki Forest virus spikes. Virology 220, Hofmann, K., Bucher, P., Falquet, L. & Bairoch, A. (1999). The PROSITE database, its status in Nucleic Acids Research 27, Justmann, J., Klimjack, M. R. & Kielian, M. (1993). Role of spike protein conformational changes in fusion of Semliki Forest virus. Journal of Virology 67, Lanzrein, M., Weingart, R. & Kempf, C. (1993). ph-dependent pore formation in Semliki Forest virus-infected Aedes albopictus cells. Virology 193, Marsh, M. & Helenius, A. (1989). Virus entry into animal cells. Advances in Virus Research 36, Nyfeler, S., Senn, K. & Kempf, C. (2001). Expression of Semliki Forest virus E1 protein in Escherichia coli: low ph-induced pore formation. Journal of Biological Chemistry 276, Paredes, A. M., Simon, M. L. & Brown, D. T. (1992). The mass of the Sindbis virus nucleocapsid suggests it has T 4 icosahedral symmetry. Virology 187, Powers, A. M., Brault, A. C., Shirako, Y., Strauss, E. G., Kang, W., Strauss, J. H. & Weaver, S. C. (2001). Evolutionary relationships and systematics of the alphaviruses. Journal of Virology 75, Schlesinger, S. & Schlesinger, M. J. (2001). Togaviridae: the viruses and their replication. In Fields Virology, 4th edn, pp Edited by D. M. Knipe & P. M. Howley. Lippincott Williams & Wilkins. Simons, K. & Garoff, H. (1980). The budding mechanisms of enveloped animal viruses. Journal of General Virology 50, Singh, I. R. & Helenius, A. (1992). Role of ribosomes in Semliki Forest virus nucleocapsid uncoating. Journal of Virology 66, Singh, I. R., Suomalainen, M., Varadarajan, S., Garoff, H. & Helenius, A. (1997). Multiple mechanisms for the inhibition of entry and uncoating of superinfecting Semliki Forest virus. Virology 231, So derlund, H., Ka a ria inen, L., von Bondsdorff, C.-H. & Weckstro m, P. (1972). Properties of Semliki Forest virus nucleocapsid. II. An irreversible contraction by acid ph. Virology 47, So derlund, H., von Bondsdorff, C.-H. & Ulmanen, I. (1979). Comparison of the structural properties of Sindbis and Semliki Forest virus nucleocapsids. Journal of General Virology 45, Strauss, J. H. & Strauss, E. G. (1994). The alphaviruses: gene expression, replication, evolution. Microbiological Reviews 58, Ulmanen, I., So derlund, H. & Ka a ria inen, L. (1976). Semliki Forest virus capsid associates with the 60S ribosomal subunit in infected cells. Journal of Virology 20, Ulmanen, I., So derlund, H. & Ka a ria inen, L. (1979). Role of protein synthesis in the assembly of Semliki Forest virus nucleocapsid. Virology 99, Van Regenmortel, M. H. V., Fauquet, C. M., Bishop, D. H. L., Carstens, E. B., Estes, M. K., Lemon, S. M., Maniloff, J., Mayo, M. A., McGeoch, D. J., Pringle, C. R. & Wickner, R. B. (editors) (2000). Virus Taxonomy. Seventh Report of the International Committee on Taxonomy of Viruses. San Diego: Academic Press. Wahlberg, J. M., Bron, R., Wilschut, J. & Garoff, H. (1992). Membrane fusion of Semliki Forest virus involves homotrimers of the fusion protein. Journal of Virology 66, Wengler, G. (1987). The mode of assembly of alphavirus cores implies a mechanism for the disassembly of the cores in the early stages of infection. Archives of Virology 94, Wengler, G. & Wengler, G. (1984). Identification of a transfer of viral core protein to cellular ribosomes during the early stages of alphavirus infection. Virology 134, Wengler, G., Wengler, G., Boege, U. & Wahn, K. (1984). Establishment and analysis of a system which allows assembly and disassembly of alphavirus core-like particles under physiological conditions in vitro. Virology 132, Wengler, G., Wu rkner, D. & Wengler, G. (1992). Identification of a sequence element in the alphavirus core protein which mediates interaction of cores with ribosomes and the disassembly of cores. Virology 191, Wengler, G., Gros, C. & Wengler, G. (1996). Analyses of the role of structural changes in the regulation of uncoating and assembly of alphavirus cores. Virology 222, Received 21 March 2002; Accepted 23 May 2002 CECG
Role of Ribosomes in Semliki Forest Virus Nucleocapsid Uncoating
 JOURNAL OF VIROLOGY, Dec. 1992, p. 749-758 22-538X/92/12749-1$2./ Copyright 1992, American Society for Microbiology Vol. 66, No. 12 Role of Ribosomes in Semliki Forest Virus Nucleocapsid Uncoating ILA
JOURNAL OF VIROLOGY, Dec. 1992, p. 749-758 22-538X/92/12749-1$2./ Copyright 1992, American Society for Microbiology Vol. 66, No. 12 Role of Ribosomes in Semliki Forest Virus Nucleocapsid Uncoating ILA
Viral structure م.م رنا مشعل
 Viral structure م.م رنا مشعل Viruses must reproduce (replicate) within cells, because they cannot generate energy or synthesize proteins. Because they can reproduce only within cells, viruses are obligate
Viral structure م.م رنا مشعل Viruses must reproduce (replicate) within cells, because they cannot generate energy or synthesize proteins. Because they can reproduce only within cells, viruses are obligate
Translation. Host Cell Shutoff 1) Initiation of eukaryotic translation involves many initiation factors
 Translation Questions? 1) How does poliovirus shutoff eukaryotic translation? 2) If eukaryotic messages are not translated how can poliovirus get its message translated? Host Cell Shutoff 1) Initiation
Translation Questions? 1) How does poliovirus shutoff eukaryotic translation? 2) If eukaryotic messages are not translated how can poliovirus get its message translated? Host Cell Shutoff 1) Initiation
Lecture 2: Virology. I. Background
 Lecture 2: Virology I. Background A. Properties 1. Simple biological systems a. Aggregates of nucleic acids and protein 2. Non-living a. Cannot reproduce or carry out metabolic activities outside of a
Lecture 2: Virology I. Background A. Properties 1. Simple biological systems a. Aggregates of nucleic acids and protein 2. Non-living a. Cannot reproduce or carry out metabolic activities outside of a
Chapter 19: Viruses. 1. Viral Structure & Reproduction. What exactly is a Virus? 11/7/ Viral Structure & Reproduction. 2.
 Chapter 19: Viruses 1. Viral Structure & Reproduction 2. Bacteriophages 3. Animal Viruses 4. Viroids & Prions 1. Viral Structure & Reproduction Chapter Reading pp. 393-396 What exactly is a Virus? Viruses
Chapter 19: Viruses 1. Viral Structure & Reproduction 2. Bacteriophages 3. Animal Viruses 4. Viroids & Prions 1. Viral Structure & Reproduction Chapter Reading pp. 393-396 What exactly is a Virus? Viruses
Problem-solving Test: The Mechanism of Protein Synthesis
 Q 2009 by The International Union of Biochemistry and Molecular Biology BIOCHEMISTRY AND MOLECULAR BIOLOGY EDUCATION Vol. 37, No. 1, pp. 58 62, 2009 Problem-based Learning Problem-solving Test: The Mechanism
Q 2009 by The International Union of Biochemistry and Molecular Biology BIOCHEMISTRY AND MOLECULAR BIOLOGY EDUCATION Vol. 37, No. 1, pp. 58 62, 2009 Problem-based Learning Problem-solving Test: The Mechanism
Replication of Sindbis Virus V. Polyribosomes and mrna in Infected Cells
 JOURNAL OF VIROLOGY, Sept. 1974, p. 552-559 Vol. 14, No. 3 Copyright @ 1974 American Society for Microbiology Printed in U.S.A. Replication of Sindbis Virus V. Polyribosomes and mrna in Infected Cells
JOURNAL OF VIROLOGY, Sept. 1974, p. 552-559 Vol. 14, No. 3 Copyright @ 1974 American Society for Microbiology Printed in U.S.A. Replication of Sindbis Virus V. Polyribosomes and mrna in Infected Cells
Chapter 19: Viruses. 1. Viral Structure & Reproduction. 2. Bacteriophages. 3. Animal Viruses. 4. Viroids & Prions
 Chapter 19: Viruses 1. Viral Structure & Reproduction 2. Bacteriophages 3. Animal Viruses 4. Viroids & Prions 1. Viral Structure & Reproduction Chapter Reading pp. 393-396 What exactly is a Virus? Viruses
Chapter 19: Viruses 1. Viral Structure & Reproduction 2. Bacteriophages 3. Animal Viruses 4. Viroids & Prions 1. Viral Structure & Reproduction Chapter Reading pp. 393-396 What exactly is a Virus? Viruses
VIRUSES. 1. Describe the structure of a virus by completing the following chart.
 AP BIOLOGY MOLECULAR GENETICS ACTIVITY #3 NAME DATE HOUR VIRUSES 1. Describe the structure of a virus by completing the following chart. Viral Part Description of Part 2. Some viruses have an envelope
AP BIOLOGY MOLECULAR GENETICS ACTIVITY #3 NAME DATE HOUR VIRUSES 1. Describe the structure of a virus by completing the following chart. Viral Part Description of Part 2. Some viruses have an envelope
Identification of a Region in the Sindbis Virus Nucleocapsid Protein That Is Involved in Specificity of RNA Encapsidation
 JOURNAL OF VIROLOGY, May 1996, p. 2757 2763 Vol. 70, No. 5 0022-538X/96/$04.00 0 Copyright 1996, American Society for Microbiology Identification of a Region in the Sindbis Virus Nucleocapsid Protein That
JOURNAL OF VIROLOGY, May 1996, p. 2757 2763 Vol. 70, No. 5 0022-538X/96/$04.00 0 Copyright 1996, American Society for Microbiology Identification of a Region in the Sindbis Virus Nucleocapsid Protein That
In Vitro-Assembled Alphavirus Core-Like Particles Maintain a Structure Similar to That of Nucleocapsid Cores in Mature Virus
 JOURNAL OF VIROLOGY, Nov. 2002, p. 11128 11132 Vol. 76, No. 21 0022-538X/02/$04.00 0 DOI: 10.1128/JVI.76.21.11128 11132.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved. In Vitro-Assembled
JOURNAL OF VIROLOGY, Nov. 2002, p. 11128 11132 Vol. 76, No. 21 0022-538X/02/$04.00 0 DOI: 10.1128/JVI.76.21.11128 11132.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved. In Vitro-Assembled
Chapter 13B: Animal Viruses
 Chapter 13B: Animal Viruses 1. Overview of Animal Viruses 2. DNA Viruses 3. RNA Viruses 4. Prions 1. Overview of Animal Viruses Life Cycle of Animal Viruses The basic life cycle stages of animal viruses
Chapter 13B: Animal Viruses 1. Overview of Animal Viruses 2. DNA Viruses 3. RNA Viruses 4. Prions 1. Overview of Animal Viruses Life Cycle of Animal Viruses The basic life cycle stages of animal viruses
Dr. Ahmed K. Ali Attachment and entry of viruses into cells
 Lec. 6 Dr. Ahmed K. Ali Attachment and entry of viruses into cells The aim of a virus is to replicate itself, and in order to achieve this aim it needs to enter a host cell, make copies of itself and
Lec. 6 Dr. Ahmed K. Ali Attachment and entry of viruses into cells The aim of a virus is to replicate itself, and in order to achieve this aim it needs to enter a host cell, make copies of itself and
Research Article 531. *Corresponding authors. Key words: assembly, capsid structure, mutational analysis, Sindbis virus, virus budding
 Research Article 531 Identification of a protein binding site on the surface of the alphavirus nucleocapsid and its implication in virus assembly Sukyeong Lee 1, Katherine E Owen 1, Hok-Kin Choi 1, Heuiran
Research Article 531 Identification of a protein binding site on the surface of the alphavirus nucleocapsid and its implication in virus assembly Sukyeong Lee 1, Katherine E Owen 1, Hok-Kin Choi 1, Heuiran
STRUCTURE, GENERAL CHARACTERISTICS AND REPRODUCTION OF VIRUSES
 STRUCTURE, GENERAL CHARACTERISTICS AND REPRODUCTION OF VIRUSES Introduction Viruses are noncellular genetic elements that use a living cell for their replication and have an extracellular state. Viruses
STRUCTURE, GENERAL CHARACTERISTICS AND REPRODUCTION OF VIRUSES Introduction Viruses are noncellular genetic elements that use a living cell for their replication and have an extracellular state. Viruses
Trident Membrane Protein Extraction Kit
 Cat. No. Size Shelf life GTX16373 5/ 20 tests 12 months at the appropriate storage temperatures (see below) Contents Component Storage Amount for 5 tests Amount for 20 tests Buffer A -20 o C 2.5 ml 10
Cat. No. Size Shelf life GTX16373 5/ 20 tests 12 months at the appropriate storage temperatures (see below) Contents Component Storage Amount for 5 tests Amount for 20 tests Buffer A -20 o C 2.5 ml 10
Coronaviruses. Virion. Genome. Genes and proteins. Viruses and hosts. Diseases. Distinctive characteristics
 Coronaviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Spherical enveloped particles studded with clubbed spikes Diameter 120-160 nm Coiled helical
Coronaviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Spherical enveloped particles studded with clubbed spikes Diameter 120-160 nm Coiled helical
number Done by Corrected by Doctor Ashraf
 number 4 Done by Nedaa Bani Ata Corrected by Rama Nada Doctor Ashraf Genome replication and gene expression Remember the steps of viral replication from the last lecture: Attachment, Adsorption, Penetration,
number 4 Done by Nedaa Bani Ata Corrected by Rama Nada Doctor Ashraf Genome replication and gene expression Remember the steps of viral replication from the last lecture: Attachment, Adsorption, Penetration,
Reoviruses. Virion. Genome. Genes and proteins. Viruses and hosts. Diseases. Distinctive characteristics
 Reoviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Naked icosahedral capsid (T=13), diameter 60-85 nm Capsid consists of two or three concentric protein
Reoviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Naked icosahedral capsid (T=13), diameter 60-85 nm Capsid consists of two or three concentric protein
Differences in the Postfusion Conformations of Full-Length and Truncated Class II Fusion Protein E of Tick-Borne Encephalitis Virus
 JOURNAL OF VIROLOGY, May 2005, p. 6511 6515 Vol. 79, No. 10 0022-538X/05/$08.00 0 doi:10.1128/jvi.79.10.6511 6515.2005 Copyright 2005, American Society for Microbiology. All Rights Reserved. Differences
JOURNAL OF VIROLOGY, May 2005, p. 6511 6515 Vol. 79, No. 10 0022-538X/05/$08.00 0 doi:10.1128/jvi.79.10.6511 6515.2005 Copyright 2005, American Society for Microbiology. All Rights Reserved. Differences
Overview of virus life cycle
 Overview of virus life cycle cell recognition and internalization release from cells progeny virus assembly membrane breaching nucleus capsid disassembly and genome release replication and translation
Overview of virus life cycle cell recognition and internalization release from cells progeny virus assembly membrane breaching nucleus capsid disassembly and genome release replication and translation
Recombinant Protein Expression Retroviral system
 Recombinant Protein Expression Retroviral system Viruses Contains genome DNA or RNA Genome encased in a protein coat or capsid. Some viruses have membrane covering protein coat enveloped virus Ø Essential
Recombinant Protein Expression Retroviral system Viruses Contains genome DNA or RNA Genome encased in a protein coat or capsid. Some viruses have membrane covering protein coat enveloped virus Ø Essential
Wilmington, Delaware cells were harvested in the cold and pelleted. The cell. pellet was suspended in 2 ml of cold buffer consisting
 JOURNAL OF VIROLOGY, June 1969, p. 599-64 Vol. 3, No. 6 Copyright 1969 American Society for Microbiology Printed in U.S.A. Sindbis Virus-induced Viral Ribonucleic Acid Polymerasel T. SREEVALSAN' AND FAY
JOURNAL OF VIROLOGY, June 1969, p. 599-64 Vol. 3, No. 6 Copyright 1969 American Society for Microbiology Printed in U.S.A. Sindbis Virus-induced Viral Ribonucleic Acid Polymerasel T. SREEVALSAN' AND FAY
10 mm KCl in a Ti-15 zonal rotor at 35,000 rpm for 16 hr at
 Proc. Nat. Acad. SCi. USA Vol. 68, No. 11, pp. 2752-2756, November 1971 Translation of Exogenous Messenger RNA for Hemoglobin on Reticulocyte and Liver Ribosomes (initiation factors/9s RNA/liver factors/reticulocyte
Proc. Nat. Acad. SCi. USA Vol. 68, No. 11, pp. 2752-2756, November 1971 Translation of Exogenous Messenger RNA for Hemoglobin on Reticulocyte and Liver Ribosomes (initiation factors/9s RNA/liver factors/reticulocyte
Preparation and properties of a novel influenza subunit vaccine G. SCHMIDT* H. BACHMAYER E. LIEHL. guinea-pigs, s.c. and i.m. for rabbits.
 Postgraduate Medical Journal (June 1976), 52, 360-367. Preparation and properties of a novel influenza subunit vaccine H. BACHMAYER E. LIEHL Ph.D. Ph.D. G. SCHMIDT Ph.D. Summary Haemagglutinin and neuraminidase
Postgraduate Medical Journal (June 1976), 52, 360-367. Preparation and properties of a novel influenza subunit vaccine H. BACHMAYER E. LIEHL Ph.D. Ph.D. G. SCHMIDT Ph.D. Summary Haemagglutinin and neuraminidase
Chapter13 Characterizing and Classifying Viruses, Viroids, and Prions
 Chapter13 Characterizing and Classifying Viruses, Viroids, and Prions 11/20/2017 MDufilho 1 Characteristics of Viruses Viruses Minuscule, acellular, infectious agent having either DNA or RNA Cause infections
Chapter13 Characterizing and Classifying Viruses, Viroids, and Prions 11/20/2017 MDufilho 1 Characteristics of Viruses Viruses Minuscule, acellular, infectious agent having either DNA or RNA Cause infections
Human Genome Complexity, Viruses & Genetic Variability
 Human Genome Complexity, Viruses & Genetic Variability (Learning Objectives) Learn the types of DNA sequences present in the Human Genome other than genes coding for functional proteins. Review what you
Human Genome Complexity, Viruses & Genetic Variability (Learning Objectives) Learn the types of DNA sequences present in the Human Genome other than genes coding for functional proteins. Review what you
Encapsidation of Sendai Virus Genome RNAs by Purified
 JOURNAL OF VIROLOGY, Mar. 1988, p. 834-838 22-538X/88/3834-5$2./ Copyright C) 1988, American Society for Microbiology Vol. 62, No. 3 Encapsidation of Sendai Virus Genome RNAs by Purified NP Protein during
JOURNAL OF VIROLOGY, Mar. 1988, p. 834-838 22-538X/88/3834-5$2./ Copyright C) 1988, American Society for Microbiology Vol. 62, No. 3 Encapsidation of Sendai Virus Genome RNAs by Purified NP Protein during
October 26, Lecture Readings. Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell
 October 26, 2006 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
October 26, 2006 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
RESCUE OF INFECTIOUS PARTICLES FROM PRE-ASSEMBLED ALPHAVIRUS NUCLEOCAPSID CORES. Richard J. Kuhn 1,2 *
 JVI Accepts, published online ahead of print on 6 April 2011 J. Virol. doi:10.1128/jvi.00039-11 Copyright 2011, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved.
JVI Accepts, published online ahead of print on 6 April 2011 J. Virol. doi:10.1128/jvi.00039-11 Copyright 2011, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved.
Introduction to viruses. BIO 370 Ramos
 Introduction to viruses BIO 370 Ramos 1 2 General Structure of Viruses Size range most
Introduction to viruses BIO 370 Ramos 1 2 General Structure of Viruses Size range most
Virus Entry/Uncoating
 Virus Entry/Uncoating Delivery of genome to inside of a cell Genome must be available for first step of replication The Problem--barriers to infection Virion Barriers: Non-enveloped viruses capsid Enveloped
Virus Entry/Uncoating Delivery of genome to inside of a cell Genome must be available for first step of replication The Problem--barriers to infection Virion Barriers: Non-enveloped viruses capsid Enveloped
Picornaviruses. Virion. Genome. Genes and proteins. Viruses and hosts. Diseases. Distinctive characteristics
 Picornaviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Naked icosahedral capsid (T=1) Diameter of 30 nm Genome Linear single-stranded RNA, positive
Picornaviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Naked icosahedral capsid (T=1) Diameter of 30 nm Genome Linear single-stranded RNA, positive
There are approximately 30,000 proteasomes in a typical human cell Each proteasome is approximately 700 kda in size The proteasome is made up of 3
 Proteasomes Proteasomes Proteasomes are responsible for degrading proteins that have been damaged, assembled improperly, or that are of no profitable use to the cell. The unwanted protein is literally
Proteasomes Proteasomes Proteasomes are responsible for degrading proteins that have been damaged, assembled improperly, or that are of no profitable use to the cell. The unwanted protein is literally
Structural biology of viruses
 Structural biology of viruses Biophysical Chemistry 1, Fall 2010 Coat proteins DNA/RNA packaging Reading assignment: Chap. 15 Virus particles self-assemble from coat monomers Virus Structure and Function
Structural biology of viruses Biophysical Chemistry 1, Fall 2010 Coat proteins DNA/RNA packaging Reading assignment: Chap. 15 Virus particles self-assemble from coat monomers Virus Structure and Function
11/15/2011. Outline. Structural Features and Characteristics. The Good the Bad and the Ugly. Viral Genomes. Structural Features and Characteristics
 Chapter 19 - Viruses Outline I. Viruses A. Structure of viruses B. Common Characteristics of Viruses C. Viral replication D. HIV II. Prions The Good the Bad and the Ugly Viruses fit into the bad category
Chapter 19 - Viruses Outline I. Viruses A. Structure of viruses B. Common Characteristics of Viruses C. Viral replication D. HIV II. Prions The Good the Bad and the Ugly Viruses fit into the bad category
Biochemical Techniques 06 Salt Fractionation of Proteins. Biochemistry
 . 1 Description of Module Subject Name Paper Name 12 Module Name/Title 2 1. Objectives Understanding the concept of protein fractionation Understanding protein fractionation with salt 2. Concept Map 3.
. 1 Description of Module Subject Name Paper Name 12 Module Name/Title 2 1. Objectives Understanding the concept of protein fractionation Understanding protein fractionation with salt 2. Concept Map 3.
Supplementary Material
 Supplementary Material Nuclear import of purified HIV-1 Integrase. Integrase remains associated to the RTC throughout the infection process until provirus integration occurs and is therefore one likely
Supplementary Material Nuclear import of purified HIV-1 Integrase. Integrase remains associated to the RTC throughout the infection process until provirus integration occurs and is therefore one likely
MagCapture Exosome Isolation Kit PS Q&A
 MagCapture Exosome Isolation Kit PS Q&A Specifications and performance P.1 Comparison of the conventional method P.2 Operation methods and composition P.4 Amount of starting sample P.5 Analysis after exosomes
MagCapture Exosome Isolation Kit PS Q&A Specifications and performance P.1 Comparison of the conventional method P.2 Operation methods and composition P.4 Amount of starting sample P.5 Analysis after exosomes
Materials and Methods , The two-hybrid principle.
 The enzymatic activity of an unknown protein which cleaves the phosphodiester bond between the tyrosine residue of a viral protein and the 5 terminus of the picornavirus RNA Introduction Every day there
The enzymatic activity of an unknown protein which cleaves the phosphodiester bond between the tyrosine residue of a viral protein and the 5 terminus of the picornavirus RNA Introduction Every day there
Last time we talked about the few steps in viral replication cycle and the un-coating stage:
 Zeina Al-Momani Last time we talked about the few steps in viral replication cycle and the un-coating stage: Un-coating: is a general term for the events which occur after penetration, we talked about
Zeina Al-Momani Last time we talked about the few steps in viral replication cycle and the un-coating stage: Un-coating: is a general term for the events which occur after penetration, we talked about
5/6/17. Diseases. Disease. Pathogens. Domain Bacteria Characteristics. Bacteria Viruses (including HIV) Pathogens are disease-causing organisms
 5/6/17 Disease Diseases I. II. Bacteria Viruses (including HIV) Biol 105 Chapter 13a Pathogens Pathogens are disease-causing organisms Domain Bacteria Characteristics 1. Domain Bacteria are prokaryotic.
5/6/17 Disease Diseases I. II. Bacteria Viruses (including HIV) Biol 105 Chapter 13a Pathogens Pathogens are disease-causing organisms Domain Bacteria Characteristics 1. Domain Bacteria are prokaryotic.
D. J. Dargan,* C. B. Gait and J. H. Subak-Sharpe
 Journal of General Virology (1992), 73, 407-411. Printed in Great Britain 407 The effect of cicloxolone sodium on the replication in cultured cells of adenovirus type 5, reovirus type 3, poliovirus type
Journal of General Virology (1992), 73, 407-411. Printed in Great Britain 407 The effect of cicloxolone sodium on the replication in cultured cells of adenovirus type 5, reovirus type 3, poliovirus type
Summary of Endomembrane-system
 Summary of Endomembrane-system 1. Endomembrane System: The structural and functional relationship organelles including ER,Golgi complex, lysosome, endosomes, secretory vesicles. 2. Membrane-bound structures
Summary of Endomembrane-system 1. Endomembrane System: The structural and functional relationship organelles including ER,Golgi complex, lysosome, endosomes, secretory vesicles. 2. Membrane-bound structures
Life Sciences 1A Midterm Exam 2. November 13, 2006
 Name: TF: Section Time Life Sciences 1A Midterm Exam 2 November 13, 2006 Please write legibly in the space provided below each question. You may not use calculators on this exam. We prefer that you use
Name: TF: Section Time Life Sciences 1A Midterm Exam 2 November 13, 2006 Please write legibly in the space provided below each question. You may not use calculators on this exam. We prefer that you use
Hepatitis B Antiviral Drug Development Multi-Marker Screening Assay
 Hepatitis B Antiviral Drug Development Multi-Marker Screening Assay Background ImQuest BioSciences has developed and qualified a single-plate method to expedite the screening of antiviral agents against
Hepatitis B Antiviral Drug Development Multi-Marker Screening Assay Background ImQuest BioSciences has developed and qualified a single-plate method to expedite the screening of antiviral agents against
Chapter 6- An Introduction to Viruses*
 Chapter 6- An Introduction to Viruses* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. 6.1 Overview of Viruses
Chapter 6- An Introduction to Viruses* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. 6.1 Overview of Viruses
Viral reproductive cycle
 Lecture 29: Viruses Lecture outline 11/11/05 Types of viruses Bacteriophage Lytic and lysogenic life cycles viruses viruses Influenza Prions Mad cow disease 0.5 µm Figure 18.4 Viral structure of capsid
Lecture 29: Viruses Lecture outline 11/11/05 Types of viruses Bacteriophage Lytic and lysogenic life cycles viruses viruses Influenza Prions Mad cow disease 0.5 µm Figure 18.4 Viral structure of capsid
Virus Structure. Characteristics of capsids. Virus envelopes. Virion assembly John Wiley & Sons, Inc. All rights reserved.
 Virus Structure Characteristics of capsids Virus envelopes Virion assembly Capsids package viral genomes and transmit them to a new host cell Capsid rigid, symmetrical container composed of viral protein
Virus Structure Characteristics of capsids Virus envelopes Virion assembly Capsids package viral genomes and transmit them to a new host cell Capsid rigid, symmetrical container composed of viral protein
Introductory Virology. Ibrahim Jamfaru School of Medicine UHAS
 Introductory Virology Ibrahim Jamfaru School of Medicine UHAS Lecture outline Definition of viruses and general characteristics Structure of virus (virion) Chemical composition of viruses Virus morphology
Introductory Virology Ibrahim Jamfaru School of Medicine UHAS Lecture outline Definition of viruses and general characteristics Structure of virus (virion) Chemical composition of viruses Virus morphology
Overview: Chapter 19 Viruses: A Borrowed Life
 Overview: Chapter 19 Viruses: A Borrowed Life Viruses called bacteriophages can infect and set in motion a genetic takeover of bacteria, such as Escherichia coli Viruses lead a kind of borrowed life between
Overview: Chapter 19 Viruses: A Borrowed Life Viruses called bacteriophages can infect and set in motion a genetic takeover of bacteria, such as Escherichia coli Viruses lead a kind of borrowed life between
Supplementary Information
 Supplementary Information Structural aspects of messenger RNA maintenance by the ribosome Lasse B. Jenner 1,2, Natalia Demeshkina 1,3, Gulnara Yusupova 1,3* and Marat Yusupov 1,3*. 1 Institut de Génétique
Supplementary Information Structural aspects of messenger RNA maintenance by the ribosome Lasse B. Jenner 1,2, Natalia Demeshkina 1,3, Gulnara Yusupova 1,3* and Marat Yusupov 1,3*. 1 Institut de Génétique
number Done by Corrected by Doctor Ashraf Khasawneh
 number 3 Done by Mahdi Sharawi Corrected by Doctor Ashraf Khasawneh *Note: Please go back to the slides to view the information that the doctor didn t mention. Prions Definition: Prions are rather ill-defined
number 3 Done by Mahdi Sharawi Corrected by Doctor Ashraf Khasawneh *Note: Please go back to the slides to view the information that the doctor didn t mention. Prions Definition: Prions are rather ill-defined
ATTACHMENT OF RIBOSOMES TO MEMBRANES DURING POLYSOME FORMATION IN MOUSE SARCOMA 180 CELLS
 Published Online: 1 June, 1971 Supp Info: http://doi.org/10.1083/jcb.49.3.683 Downloaded from jcb.rupress.org on November 2, 2018 ATTACHMENT OF RIBOSOMES TO MEMBRANES DURING POLYSOME FORMATION IN MOUSE
Published Online: 1 June, 1971 Supp Info: http://doi.org/10.1083/jcb.49.3.683 Downloaded from jcb.rupress.org on November 2, 2018 ATTACHMENT OF RIBOSOMES TO MEMBRANES DURING POLYSOME FORMATION IN MOUSE
Virology. *Viruses can be only observed by electron microscope never by light microscope. The size of the virus: nm in diameter.
 Virology We are going to start with general introduction about viruses, they are everywhere around us; in food; within the environment; in direct contact to etc.. They may cause viral infection by itself
Virology We are going to start with general introduction about viruses, they are everywhere around us; in food; within the environment; in direct contact to etc.. They may cause viral infection by itself
Section 6. Junaid Malek, M.D.
 Section 6 Junaid Malek, M.D. The Golgi and gp160 gp160 transported from ER to the Golgi in coated vesicles These coated vesicles fuse to the cis portion of the Golgi and deposit their cargo in the cisternae
Section 6 Junaid Malek, M.D. The Golgi and gp160 gp160 transported from ER to the Golgi in coated vesicles These coated vesicles fuse to the cis portion of the Golgi and deposit their cargo in the cisternae
PRODUCT: RNAzol BD for Blood May 2014 Catalog No: RB 192 Storage: Store at room temperature
 PRODUCT: RNAzol BD for Blood May 2014 Catalog No: RB 192 Storage: Store at room temperature PRODUCT DESCRIPTION. RNAzol BD is a reagent for isolation of total RNA from whole blood, plasma or serum of human
PRODUCT: RNAzol BD for Blood May 2014 Catalog No: RB 192 Storage: Store at room temperature PRODUCT DESCRIPTION. RNAzol BD is a reagent for isolation of total RNA from whole blood, plasma or serum of human
Universal sample preparation method for proteome analysis
 nature methods Universal sample preparation method for proteome analysis Jacek R Wi niewski, Alexandre Zougman, Nagarjuna Nagaraj & Matthias Mann Supplementary figures and text: Supplementary Figure 1
nature methods Universal sample preparation method for proteome analysis Jacek R Wi niewski, Alexandre Zougman, Nagarjuna Nagaraj & Matthias Mann Supplementary figures and text: Supplementary Figure 1
Virology Journal. Open Access. Abstract. BioMed Central
 Virology Journal BioMed Central Research Stimulation of poliovirus RNA synthesis and virus maturation in a HeLa cell-free in vitro translation-rna replication system by viral protein 3CD pro David Franco
Virology Journal BioMed Central Research Stimulation of poliovirus RNA synthesis and virus maturation in a HeLa cell-free in vitro translation-rna replication system by viral protein 3CD pro David Franco
Structural Requirements for Low-pH-Induced Rearrangements in the Envelope Glycoprotein of Tick-Borne Encephalitis Virus
 JOURNAL OF VIROLOGY, Nov. 1996, p. 8142 8147 Vol. 70, No. 11 0022-538X/96/$04.00 0 Copyright 1996, American Society for Microbiology Structural Requirements for Low-pH-Induced Rearrangements in the Envelope
JOURNAL OF VIROLOGY, Nov. 1996, p. 8142 8147 Vol. 70, No. 11 0022-538X/96/$04.00 0 Copyright 1996, American Society for Microbiology Structural Requirements for Low-pH-Induced Rearrangements in the Envelope
8/13/2009. Diseases. Disease. Pathogens. Domain Bacteria Characteristics. Bacteria Shapes. Domain Bacteria Characteristics
 Disease Diseases I. Bacteria II. Viruses including Biol 105 Lecture 17 Chapter 13a are disease-causing organisms Domain Bacteria Characteristics 1. Domain Bacteria are prokaryotic 2. Lack a membrane-bound
Disease Diseases I. Bacteria II. Viruses including Biol 105 Lecture 17 Chapter 13a are disease-causing organisms Domain Bacteria Characteristics 1. Domain Bacteria are prokaryotic 2. Lack a membrane-bound
Enzymes: The Catalysts of Life
 Chapter 6 Enzymes: The Catalysts of Life Lectures by Kathleen Fitzpatrick Simon Fraser University Activation Energy and the Metastable State Many thermodynamically feasible reactions in a cell that could
Chapter 6 Enzymes: The Catalysts of Life Lectures by Kathleen Fitzpatrick Simon Fraser University Activation Energy and the Metastable State Many thermodynamically feasible reactions in a cell that could
SUPPLEMENTARY MATERIAL
 SUPPLEMENTARY MATERIAL Purification and biochemical properties of SDS-stable low molecular weight alkaline serine protease from Citrullus Colocynthis Muhammad Bashir Khan, 1,3 Hidayatullah khan, 2 Muhammad
SUPPLEMENTARY MATERIAL Purification and biochemical properties of SDS-stable low molecular weight alkaline serine protease from Citrullus Colocynthis Muhammad Bashir Khan, 1,3 Hidayatullah khan, 2 Muhammad
Three-dimensional structure of a membrane-containing virus
 Proc. Natl. Acad. Sci. USA Vol. 90, pp. 9095-9099, October 1993 Microbiology Three-dimensional structure of a membrane-containing virus ANGEL M. PAREDES*, DENNIS T. BROWN*t, ROSALBA ROTHNAGELt, WAH CHIU*,
Proc. Natl. Acad. Sci. USA Vol. 90, pp. 9095-9099, October 1993 Microbiology Three-dimensional structure of a membrane-containing virus ANGEL M. PAREDES*, DENNIS T. BROWN*t, ROSALBA ROTHNAGELt, WAH CHIU*,
Molecular Cell Biology Problem Drill 16: Intracellular Compartment and Protein Sorting
 Molecular Cell Biology Problem Drill 16: Intracellular Compartment and Protein Sorting Question No. 1 of 10 Question 1. Which of the following statements about the nucleus is correct? Question #01 A. The
Molecular Cell Biology Problem Drill 16: Intracellular Compartment and Protein Sorting Question No. 1 of 10 Question 1. Which of the following statements about the nucleus is correct? Question #01 A. The
LESSON 4.6 WORKBOOK. Designing an antiviral drug The challenge of HIV
 LESSON 4.6 WORKBOOK Designing an antiviral drug The challenge of HIV In the last two lessons we discussed the how the viral life cycle causes host cell damage. But is there anything we can do to prevent
LESSON 4.6 WORKBOOK Designing an antiviral drug The challenge of HIV In the last two lessons we discussed the how the viral life cycle causes host cell damage. But is there anything we can do to prevent
Chapter 13 Viruses, Viroids, and Prions. Biology 1009 Microbiology Johnson-Summer 2003
 Chapter 13 Viruses, Viroids, and Prions Biology 1009 Microbiology Johnson-Summer 2003 Viruses Virology-study of viruses Characteristics: acellular obligate intracellular parasites no ribosomes or means
Chapter 13 Viruses, Viroids, and Prions Biology 1009 Microbiology Johnson-Summer 2003 Viruses Virology-study of viruses Characteristics: acellular obligate intracellular parasites no ribosomes or means
ABSTRACT. WEST, JOHN ALLEN. Investigation of the Role of the E2 Endodomain in Sindbis Virus
 ABSTRACT WEST, JOHN ALLEN. Investigation of the Role of the E2 Endodomain in Sindbis Virus Assembly. (Under the direction of Dr. Dennis T. Brown) Sindbis virus (SV) is the prototype member of the alphavirus
ABSTRACT WEST, JOHN ALLEN. Investigation of the Role of the E2 Endodomain in Sindbis Virus Assembly. (Under the direction of Dr. Dennis T. Brown) Sindbis virus (SV) is the prototype member of the alphavirus
In Vitro Protein-Synthesizing Activity of Vesicular Stomatitis Virus-Infected Cell Extracts
 JOURNAL OF VIROLOGY, Aug. 1973, p. 265-274 Copyright 1973 American Society for Microbiology Vol. 12, No. 2 Printed in U.S.A. In Vitro Protein-Synthesizing Activity of Vesicular Stomatitis Virus-Infected
JOURNAL OF VIROLOGY, Aug. 1973, p. 265-274 Copyright 1973 American Society for Microbiology Vol. 12, No. 2 Printed in U.S.A. In Vitro Protein-Synthesizing Activity of Vesicular Stomatitis Virus-Infected
Size nm m m
 1 Viral size and organization Size 20-250nm 0.000000002m-0.000000025m Virion structure Capsid Core Acellular obligate intracellular parasites Lack organelles, metabolic activities, and reproduction Replicated
1 Viral size and organization Size 20-250nm 0.000000002m-0.000000025m Virion structure Capsid Core Acellular obligate intracellular parasites Lack organelles, metabolic activities, and reproduction Replicated
Fayth K. Yoshimura, Ph.D. September 7, of 7 RETROVIRUSES. 2. HTLV-II causes hairy T-cell leukemia
 1 of 7 I. Diseases Caused by Retroviruses RETROVIRUSES A. Human retroviruses that cause cancers 1. HTLV-I causes adult T-cell leukemia and tropical spastic paraparesis 2. HTLV-II causes hairy T-cell leukemia
1 of 7 I. Diseases Caused by Retroviruses RETROVIRUSES A. Human retroviruses that cause cancers 1. HTLV-I causes adult T-cell leukemia and tropical spastic paraparesis 2. HTLV-II causes hairy T-cell leukemia
Medical Virology. Herpesviruses, Orthomyxoviruses, and Retro virus. - Herpesviruses Structure & Composition: Herpesviruses
 Medical Virology Lecture 2 Asst. Prof. Dr. Dalya Basil Herpesviruses, Orthomyxoviruses, and Retro virus - Herpesviruses Structure & Composition: Herpesviruses Enveloped DNA viruses. All herpesviruses have
Medical Virology Lecture 2 Asst. Prof. Dr. Dalya Basil Herpesviruses, Orthomyxoviruses, and Retro virus - Herpesviruses Structure & Composition: Herpesviruses Enveloped DNA viruses. All herpesviruses have
Sindbis virus-induced inhibition of protein synthesis is partially reversed by medium containing an elevated potassium concentration
 Journal of General Virology (1994), 75, 411~415. Printed in Great Britain 411 Sindbis virus-induced inhibition of protein synthesis is partially reversed by medium containing an elevated potassium concentration
Journal of General Virology (1994), 75, 411~415. Printed in Great Britain 411 Sindbis virus-induced inhibition of protein synthesis is partially reversed by medium containing an elevated potassium concentration
HIV-1 p24 ELISA Pair Set Cat#: orb54951 (ELISA Manual)
 HIV-1 p24 ELISA Pair Set Cat#: orb54951 (ELISA Manual) BACKGROUND Human Immunodeficiency Virus ( HIV ) can be divided into two major types, HIV type 1 (HIV-1) and HIV type 2 (HIV-2). HIV-1 is related to
HIV-1 p24 ELISA Pair Set Cat#: orb54951 (ELISA Manual) BACKGROUND Human Immunodeficiency Virus ( HIV ) can be divided into two major types, HIV type 1 (HIV-1) and HIV type 2 (HIV-2). HIV-1 is related to
19 Viruses BIOLOGY. Outline. Structural Features and Characteristics. The Good the Bad and the Ugly. Structural Features and Characteristics
 9 Viruses CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson Outline I. Viruses A. Structure of viruses B. Common Characteristics of Viruses C. Viral replication D. HIV Lecture Presentation
9 Viruses CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson Outline I. Viruses A. Structure of viruses B. Common Characteristics of Viruses C. Viral replication D. HIV Lecture Presentation
ab Histone Deacetylase (HDAC) Activity Assay Kit (Fluorometric)
 ab156064 Histone Deacetylase (HDAC) Activity Assay Kit (Fluorometric) Instructions for Use For the quantitative measurement of Histone Deacetylase activity in cell lysates This product is for research
ab156064 Histone Deacetylase (HDAC) Activity Assay Kit (Fluorometric) Instructions for Use For the quantitative measurement of Histone Deacetylase activity in cell lysates This product is for research
Application of μmacs Streptavidin MicroBeads for the analysis of HIV-1 directly from patient plasma
 Excerpt from MACS&more Vol 8 1/2004 Application of μmacs Streptavidin MicroBeads for the analysis of HIV-1 directly from patient plasma L. Davis Lupo and Salvatore T. Butera HIV and Retrovirology Branch,
Excerpt from MACS&more Vol 8 1/2004 Application of μmacs Streptavidin MicroBeads for the analysis of HIV-1 directly from patient plasma L. Davis Lupo and Salvatore T. Butera HIV and Retrovirology Branch,
DNA codes for RNA, which guides protein synthesis.
 Section 3: DNA codes for RNA, which guides protein synthesis. K What I Know W What I Want to Find Out L What I Learned Vocabulary Review synthesis New RNA messenger RNA ribosomal RNA transfer RNA transcription
Section 3: DNA codes for RNA, which guides protein synthesis. K What I Know W What I Want to Find Out L What I Learned Vocabulary Review synthesis New RNA messenger RNA ribosomal RNA transfer RNA transcription
I. Bacteria II. Viruses including HIV. Domain Bacteria Characteristics. 5. Cell wall present in many species. 6. Reproduction by binary fission
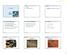 Disease Diseases I. Bacteria II. Viruses including are disease-causing organisms Biol 105 Lecture 17 Chapter 13a Domain Bacteria Characteristics 1. Domain Bacteria are prokaryotic 2. Lack a membrane-bound
Disease Diseases I. Bacteria II. Viruses including are disease-causing organisms Biol 105 Lecture 17 Chapter 13a Domain Bacteria Characteristics 1. Domain Bacteria are prokaryotic 2. Lack a membrane-bound
MedChem 401~ Retroviridae. Retroviridae
 MedChem 401~ Retroviridae Retroviruses plus-sense RNA genome (!8-10 kb) protein capsid lipid envelop envelope glycoproteins reverse transcriptase enzyme integrase enzyme protease enzyme Retroviridae The
MedChem 401~ Retroviridae Retroviruses plus-sense RNA genome (!8-10 kb) protein capsid lipid envelop envelope glycoproteins reverse transcriptase enzyme integrase enzyme protease enzyme Retroviridae The
Instructions. Fuse-It-Color. Overview. Specifications
 Membrane fusion is a novel and highly superior method for incorporating various molecules and particles into mammalian cells. Cargo-specific liposomal carriers are able to attach and rapidly fuse with
Membrane fusion is a novel and highly superior method for incorporating various molecules and particles into mammalian cells. Cargo-specific liposomal carriers are able to attach and rapidly fuse with
Translation of Sindbis virus mrna: analysis of sequences downstream of the initiating AUG codon that enhance translation.
 Translation of Sindbis virus mrna: analysis of sequences downstream of the initiating AUG codon that enhance translation. I Frolov and S Schlesinger J. Virol. 1996, 70(2):1182. CONTENT ALERTS Updated information
Translation of Sindbis virus mrna: analysis of sequences downstream of the initiating AUG codon that enhance translation. I Frolov and S Schlesinger J. Virol. 1996, 70(2):1182. CONTENT ALERTS Updated information
AP Biology Reading Guide. Concept 19.1 A virus consists of a nucleic acid surrounded by a protein coat
 AP Biology Reading Guide Name Chapter 19: Viruses Overview Experimental work with viruses has provided important evidence that genes are made of nucleic acids. Viruses were also important in working out
AP Biology Reading Guide Name Chapter 19: Viruses Overview Experimental work with viruses has provided important evidence that genes are made of nucleic acids. Viruses were also important in working out
PREPARATION OF IF- ENRICHED CYTOSKELETAL PROTEINS
 TMM,5-2011 PREPARATION OF IF- ENRICHED CYTOSKELETAL PROTEINS Ice-cold means cooled in ice water. In order to prevent proteolysis, make sure to perform all steps on ice. Pre-cool glass homogenizers, buffers
TMM,5-2011 PREPARATION OF IF- ENRICHED CYTOSKELETAL PROTEINS Ice-cold means cooled in ice water. In order to prevent proteolysis, make sure to perform all steps on ice. Pre-cool glass homogenizers, buffers
General Virology I. Dr Esam Ibraheem Azhar (BSc, MSc, Ph.D Molecular Medical Virology) Asst. Prof. Medical Laboratory Technology Department
 General Virology I Dr Esam Ibraheem Azhar (BSc, MSc, Ph.D Molecular Medical Virology) Asst. Prof. Medical Laboratory Technology Department ١ General Virology I Lecture Outline Introduction istory Definition
General Virology I Dr Esam Ibraheem Azhar (BSc, MSc, Ph.D Molecular Medical Virology) Asst. Prof. Medical Laboratory Technology Department ١ General Virology I Lecture Outline Introduction istory Definition
The Zombies of the Scientific Community Viruses and Other Acellular Infectious Agents. Acellular Agents
 viruses protein and nucleic acid viroids RNA virusoids RNA prions proteins The Zombies of the Scientific Community Viruses and Other Acellular Infectious Agents Acellular Agents Viruses major cause of
viruses protein and nucleic acid viroids RNA virusoids RNA prions proteins The Zombies of the Scientific Community Viruses and Other Acellular Infectious Agents Acellular Agents Viruses major cause of
Aura Virus Structure Suggests that the T 4 Organization Is a Fundamental Property of Viral Structural Proteins
 JOURNAL OF VIROLOGY, July 2002, p. 7239 7246 Vol. 76, No. 14 0022-538X/02/$04.00 0 DOI: 10.1128/JVI.76.14.7239 7246.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved. Aura Virus
JOURNAL OF VIROLOGY, July 2002, p. 7239 7246 Vol. 76, No. 14 0022-538X/02/$04.00 0 DOI: 10.1128/JVI.76.14.7239 7246.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved. Aura Virus
Polyomaviridae. Spring
 Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
Supplementary Information. Sonorensin: A new bacteriocin with potential of an anti-biofilm agent and a food
 Supplementary Information Sonorensin: A new bacteriocin with potential of an anti-biofilm agent and a food biopreservative Lipsy Chopra, Gurdeep Singh, Kautilya Kumar Jena and Debendra K. Sahoo* Biochemical
Supplementary Information Sonorensin: A new bacteriocin with potential of an anti-biofilm agent and a food biopreservative Lipsy Chopra, Gurdeep Singh, Kautilya Kumar Jena and Debendra K. Sahoo* Biochemical
Spike Protein-Nucleocapsid Interactions Drive
 JOURNAL OF VIROLOGY, Aug. 1992, P. 4737-4747 0022-538X/92/084737-11$02.00/0 Copyright 1992, American Society for Microbiology Vol. 66, No. 8 Spike Protein-Nucleocapsid Interactions Drive the Budding of
JOURNAL OF VIROLOGY, Aug. 1992, P. 4737-4747 0022-538X/92/084737-11$02.00/0 Copyright 1992, American Society for Microbiology Vol. 66, No. 8 Spike Protein-Nucleocapsid Interactions Drive the Budding of
Synthesis of Proteins in Cells Infected with Herpesvirus,
 Proceedings of the National Academy of Science8 Vol. 66, No. 3, pp. 799-806, July 1970 Synthesis of Proteins in Cells Infected with Herpesvirus, VI. Characterization of the Proteins of the Viral Membrane*
Proceedings of the National Academy of Science8 Vol. 66, No. 3, pp. 799-806, July 1970 Synthesis of Proteins in Cells Infected with Herpesvirus, VI. Characterization of the Proteins of the Viral Membrane*
TRANSLATION. Translation is a process where proteins are made by the ribosomes on the mrna strand.
 TRANSLATION Dr. Mahesha H B, Yuvaraja s College, University of Mysore, Mysuru. Translation is a process where proteins are made by the ribosomes on the mrna strand. Or The process in the ribosomes of a
TRANSLATION Dr. Mahesha H B, Yuvaraja s College, University of Mysore, Mysuru. Translation is a process where proteins are made by the ribosomes on the mrna strand. Or The process in the ribosomes of a
Probing the Potential Glycoprotein Binding Site of Sindbis Virus Capsid Protein With Dioxane and Model Building
 PROTEINS: Structure, Function, and Genetics 33:311 317 (1998) Probing the Potential Glycoprotein Binding Site of Sindbis Virus Capsid Protein With Dioxane and Model Building Sukyeong Lee, Richard J. Kuhn,
PROTEINS: Structure, Function, and Genetics 33:311 317 (1998) Probing the Potential Glycoprotein Binding Site of Sindbis Virus Capsid Protein With Dioxane and Model Building Sukyeong Lee, Richard J. Kuhn,
Proteins and symmetry
 Proteins and symmetry Viruses (symmetry) Viruses come in many shapes, sizes and compositions All carry genomic nucleic acid (RNA or DNA) Structurally and genetically the simplest are the spherical viruses
Proteins and symmetry Viruses (symmetry) Viruses come in many shapes, sizes and compositions All carry genomic nucleic acid (RNA or DNA) Structurally and genetically the simplest are the spherical viruses
Chapter 19: The Genetics of Viruses and Bacteria
 Chapter 19: The Genetics of Viruses and Bacteria What is Microbiology? Microbiology is the science that studies microorganisms = living things that are too small to be seen with the naked eye Microorganisms
Chapter 19: The Genetics of Viruses and Bacteria What is Microbiology? Microbiology is the science that studies microorganisms = living things that are too small to be seen with the naked eye Microorganisms
Unique Features of Hepatitis C Virus Capsid Formation Revealed by De Novo Cell-Free Assembly
 JOURNAL OF VIROLOGY, Sept. 2004, p. 9257 9269 Vol. 78, No. 17 0022-538X/04/$08.00 0 DOI: 10.1128/JVI.78.17.9257 9269.2004 Copyright 2004, American Society for Microbiology. All Rights Reserved. Unique
JOURNAL OF VIROLOGY, Sept. 2004, p. 9257 9269 Vol. 78, No. 17 0022-538X/04/$08.00 0 DOI: 10.1128/JVI.78.17.9257 9269.2004 Copyright 2004, American Society for Microbiology. All Rights Reserved. Unique
Estimations of the Molecular Weight of the Influenza Virus Genome
 o r. gem Viral. &97I), H, Io3-Io9 103 Printed in Great Britain Estimations of the Molecular Weight of the Influenza Virus Genome By J. J. SKEHEL National Institute for Medical Research, Mill Hill, London
o r. gem Viral. &97I), H, Io3-Io9 103 Printed in Great Britain Estimations of the Molecular Weight of the Influenza Virus Genome By J. J. SKEHEL National Institute for Medical Research, Mill Hill, London
N-Glycosidase F Deglycosylation Kit
 For life science research only. Not for use in diagnostic procedures. FOR IN VITRO USE ONLY. N-Glycosidase F Deglycosylation Kit Kit for the deglycosylation of asparagine-linked glycan chains on glycoproteins.
For life science research only. Not for use in diagnostic procedures. FOR IN VITRO USE ONLY. N-Glycosidase F Deglycosylation Kit Kit for the deglycosylation of asparagine-linked glycan chains on glycoproteins.
3. on T helper {cells / lymphocytes} ; 3. ACCEPT macrophages / dendritic cells / CD4 cells
 1(a) 1. (structure G is {glycoprotein / gp120} ; 2. used for {attachment / eq} to CD4 (molecules / receptors /antigens) ; 1. IGNORE gp 41 and gp 160 and other wrong numbers 3. on T helper {cells / lymphocytes}
1(a) 1. (structure G is {glycoprotein / gp120} ; 2. used for {attachment / eq} to CD4 (molecules / receptors /antigens) ; 1. IGNORE gp 41 and gp 160 and other wrong numbers 3. on T helper {cells / lymphocytes}
ARV Mode of Action. Mode of Action. Mode of Action NRTI. Immunopaedia.org.za
 ARV Mode of Action Mode of Action Mode of Action - NRTI Mode of Action - NNRTI Mode of Action - Protease Inhibitors Mode of Action - Integrase inhibitor Mode of Action - Entry Inhibitors Mode of Action
ARV Mode of Action Mode of Action Mode of Action - NRTI Mode of Action - NNRTI Mode of Action - Protease Inhibitors Mode of Action - Integrase inhibitor Mode of Action - Entry Inhibitors Mode of Action
