Targeting Human Immunodeficiency Virus (HIV) Type 2 Integrase Protein into HIV Type 1
|
|
|
- Suzanna Lewis
- 5 years ago
- Views:
Transcription
1 JOURNAL OF VIROLOGY, Oct. 1999, p Vol. 73, No X/99/$ Copyright 1999, American Society for Microbiology. All Rights Reserved. Targeting Human Immunodeficiency Virus (HIV) Type 2 Integrase Protein into HIV Type 1 HONGMEI LIU, 1 XIAOYUN WU, 1 HONGLING XIAO, 1 AND JOHN C. KAPPES 1,2,3 * Departments of Medicine 1 and Microbiology, 2 University of Alabama at Birmingham, Birmingham, Alabama 35294, and Birmingham Veterans Affairs Medical Center, Research Service, Birmingham, Alabama Received 17 March 1999/Accepted 2 July 1999 Integrase (IN) is the only retroviral enzyme necessary for the integration of retroviral cdna into the host cell s chromosomes. The structure and function of IN is highly conserved. The human immunodeficiency virus type 2 (HIV-2) IN has been shown to efficiently support 3 processing and strand transfer of HIV-1 DNA substrate in vitro. To determine whether HIV-2 IN protein (IN 2 ) could substitute for HIV-1 IN function in vivo, we used HIV-1 Vpr to deliver the IN 2 into IN mutant HIV-1 virions by expression in trans as a Vpr-IN fusion protein. Trans-complementation with IN 2 markedly increased the infectivity of IN-minus HIV-1. Compared with the homologous trans-in protein, infectivity was increased to a level of 16%. Since IN has been found to play a role in reverse transcription (Wu et al., J. Virol. 73: , 1999), cells infected with IN 2 -complemented HIV-1 were analyzed for DNA products of reverse transcription. DNA levels of approximately 18% of that of wild type were detected. The homologous trans-in protein restored the synthesis of viral cdna to approximately 86% of that of wild-type virus. By complementing integration-defective HIV-1 IN mutant viruses, which were not impaired in cdna synthesis, the trans-in 2 protein was shown to support integration up to a level of 55% compared with that of the homologous trans-in protein. The delivery of heterologous IN protein into HIV-1 particles in trans offers a novel approach to understand IN protein function in vivo. Like all other retroviruses, human immunodeficiency virus type 1 (HIV-1) and HIV-2 integrate a DNA copy of their RNA genome into the host cell s chromosomes. Integration of the viral cdna is catalyzed by the integrase (IN) protein (3). Sequence analysis has shown that many features of the primary structure of IN are highly conserved among retroviruses and retrotransposons. The amino-terminal domain contains an array of histidine and cysteine residues that form a zinc finger motif (5, 6, 18). The central domain contains the catalytic core, which is defined by three acidic residues with stereotyped spacing. This D,D-35-E motif is a universal feature of integrase proteins and is essential for catalytic activity (9, 10, 22, 25, 36). The carboxy-terminal domain is the least conserved region of IN. It binds to the viral DNA ends and also exhibits nonspecific DNA binding properties (19, 24, 30, 31, 37). The sequence conservation of integrase suggests strong structure-function relationships. IN function has been studied extensively in vitro by using purified enzyme and short oligonucleotide substrate that mimics the ends of the viral DNA molecule (8, 21, 32). Such analysis has shown that IN proteins can utilize different retroviral DNA substrates in the in vitro integration reaction. The HIV-2 IN (IN 2 ) has been shown to efficiently support 3 processing and strand transfer of HIV-1 substrate in vitro (37). Similarly, the feline immunodeficiency virus (FIV) IN can cleave and integrate HIV and Moloney murine leukemia virus (MoMLV) DNA ends (38). HIV-1 IN is active on FIV substrate but is barely active on MoMLV substrate (35, 38). While the analysis of the integration reaction in vitro has provided a great deal of information on IN function and has helped to elucidate the molecular mechanism of viral DNA integration, the conditions used to study IN in vitro do not fully duplicate those in vivo. * Corresponding author. Mailing address: University of Alabama at Birmingham, Department of Medicine, 1900 University Blvd., THT 513H, Birmingham, AL Phone: (205) Fax: (205) KappesJC@uab.edu. In the context of a replicating virus, HIV-1 IN is expressed and assembled into virions as the C-terminal component of a larger 160-kDa Gag-Pol polyprotein (Pr160 Gag-Pol ). After proteolytic processing of Pr160 Gag-Pol and entry of the virus core into the host cell, IN exists as a 32-kDa protein together with other viral and cellular proteins that make up the viral nucleoprotein complex (34). Several in vivo studies have suggested that the IN protein may be involved in other step of the virus life cycle. Host cell proteins that bind to IN or the HIV-1 preintegration complex (PIC) and promote integration have been identified (13, 20). Through analysis in nondividing cells, mutations in the C terminus of IN have been shown to disrupt the movement of the viral PIC into the nucleus (15), suggesting a role of IN in the nuclear import machinery pathway (15, 16). In other studies, some IN mutations, including those in highly conserved amino acids residues, were found to have no apparent effect on IN activity in vitro while dramatically reducing the formation of the provirus in infected cells (11, 12, 26, 29). These results were initially explained in part by changes in the structure of the Pr160 Gag-Pol precursor protein, resulting in aberrant virus assembly and maturation. By incorporating IN protein into virions in trans, independently of Pr160 Gag-Pol,we recently demonstrated that the mature IN protein itself promotes the initiation of viral DNA synthesis (40). These results indicate that IN may play important roles in the virus life cycle at several different levels. To examine whether IN 2 could function in place of HIV-1 IN during virus infection, two experimental approaches were undertaken. In the first, we replaced the HIV-1 IN coding region with that of the IN 2, generating a virus that contained an HIV-1 HIV-2 chimeric pol gene (Fig. 1). In the second approach, the IN 2 protein was incorporated into HIV-1 virions in trans by expression as a fusion partner of Vpr (Vpr-IN). This strategy is based on our previously findings and those of others, which have shown that Vpr can be used as a vehicle to deliver enzymatically active IN into HIV-1 virions in trans, independently of Pr160 Gag-Pol (14, 28, 40, 41). 8831
2 8832 NOTES J. VIROL. Downloaded from FIG. 1. Analysis of IN 2 protein function when expressed in cis with the HIV-1 genome. (A) Insertion of IN 2 into the pol gene of HIV-1. The HIV-1 SG3 wt DNA (41) was cut with the BamHI and SalI endonucleases to remove IN, vif, and the 5 half of vpr from the DNA genome. A fragment of psg3 wt DNA encompassing 16 bp of 3 IN sequence, vif and the 5 half of vpr was amplified by PCR, wherein the sense primer contained a MluI restriction site. In a separate reaction, an IN-containing DNA fragment was PCR amplified from the HIV-2 ST clone, wherein the sense and antisense primers contained BamHI and MluI restriction sites, respectively. The two PCR-amplified DNA fragments were ligated into the BamHI-SalI-cut psg3 wt DNA, generating the chimeric virus psg3 IN2. Sequence analysis confirmed an open chimeric pol reading frame (PR-RT-IN 2 ). The 3 18 nucleotides of the HIV-1 IN are retained in the chimeric protein. (B) Infectivity of SG3 IN2 virions. 293T cells were transfected by calcium phosphate DNA precipitation methods with psg3 IN2 DNA. Forty-eight hours later, the culture supernatants were collected, filtered through m-pore-size filters, and analyzed for HIV-1 p24 antigen by enzyme-linked immunosorbent assay (Coulter Inc.). Next, 25, 5, and 1 ng of each virus (p24 antigen equivalents) was used to infect monolayer cultures of P4 indicator cells (7). Two days later, the cells were stained, and infection-positive cells were counted as described earlier (41). The virus infectivity results represent infectious units per 25-ng equivalent of p24 antigen. These results were highly reproducible in three independent experiments. The data shown are from a single representative experiment. Characterization of HIV-1 that encodes an HIV-1 HIV-2 chimeric Pol protein. An IN-containing DNA fragment was PCR amplified from the HIV-2 ST proviral clone (23) and ligated into the BamHI and MluI restriction sites of the HIV-1 psg3 wt molecular clone, generating psg3 IN2 (Fig. 1A). To determine whether the SG3 IN2 virus was infectious, 293T cell monolayers were transfected with psg3 IN2 DNA. For controls, HIV-1 wild-type (SG3 wt ) and IN-minus (psg3 S-IN ; see references 28 and 41) viruses were also transfected. The supernatants of the transfected cultures were collected 72 h later and divided into three aliquots. One aliquot was ultracentrifuged to pellet virus for immunoblot analysis, the second aliquot was analyzed for reverse transcriptase (RT) activity and p24 antigen concentration (Coulter Inc.), and the third was used to infect CD4-LTR- -galactosidase indicator cells (P4) (7). No differences between the wild-type and SG3 IN2 virions with respect to RT activity (relative to p24 antigen concentration) and proteolytic processing of the Gag and Pol proteins were detected (data not shown). However, the SG3 IN2 virions exhibited a marked decrease in infectivity (Fig. 1B). Compared with the IN-minus SG3 IN2 virions, the SG3 IN2 virions were reproducibly more infectious. The HIV Gag-Pol precursor protein not only serves to incorporate the viral enzymes in the virus particle but also plays an important role in virion assembly. Several studies have shown that mutations within IN can cause defects in virus particle production, and virion composition and morphology (1, 2, 4, 11, 33). While our results show that the chimeric SG3 IN2 virus was impaired in infectivity, they do not precisely show at what step(s) in the life cycle the virus was defective. Therefore, it was not possible to understand whether virus infection was blocked because the chimeric Gag-Pol protein was not properly folded and caused a defect in the late stages of the virus life cycle or whether the heterologous IN protein on September 20, 2018 by guest
3 VOL. 73, 1999 NOTES 8833 FIG. 2. Incorporation of the Vpr-IN fusion protein into IN-minus virions. (A) psg3 S-IN and psg3 wt were separately cotransfected into 293T cells with plr2p-vprin 2, plr2p-vpr-in, and plr2p (vector alone), respectively. As described earlier (41), the extracellular progeny virions were concentrated from the supernatants of each culture by ultracentrifugation, lysed, and analyzed by immunoblot analysis using anti HIV-1 IN peptide antibody (A), human anti-hiv-2 antiserum (B), anti-vpr (C), and anti-gag (D) antibodies as probes. itself was unable to mimic the function of the HIV-1 IN protein during the early stages of the virus life cycle. Incorporation of IN 2 protein into HIV-1 virions by expression in trans. To avoid the possible dominant-negative effects of the chimeric Gag-Pol precursor protein, we fused the HIV-1 vpr gene with the HIV-2 IN gene (IN 2 ) and placed the vpr-in 2 gene fusion into the plr2p expression plasmid (plr2p-vprin 2 ) for complementing HIV-1 IN mutant viruses in trans. The plr2p-vprin 2 plasmid was cotransfected into 293T cells with the psg3 S-IN IN-minus clone. As controls, psg3 S-IN was cotransfected with the Vpr HIV-1 IN expression vector (plr2pvprin; references 28, 40, 41) and the plr2p vector. Immunoblot analysis of the progeny virions detected two species of anti-hiv-1 IN-reactive proteins (Fig. 2A), a 47-kDa protein, which is consistent with the combined masses of IN (32 kda) and Vpr (15 kda), and a 32-kDa protein. The detection of the 32-kDa protein in SG3 S-IN virions complemented with Vpr-IN indicated proteolytic processing of the fusion protein and liberation of IN. The Vpr-IN 2 fusion protein was not detected with the HIV-1 anti-in antiserum. However, using antiserum (F0784) obtained from an HIV-2-infected individual as a probe, the 47- and 32-kDa proteins were detected in both SG3 wt and SG3 S-IN virions that were complemented with either the Vpr-IN or Vpr-IN 2 fusion proteins (Fig. 2B). Using anti-vpr antibody, the 47-kDa Vpr-IN and Vpr-IN 2 fusion proteins and their respective Vpr cleavage products were detected (Fig. 2C). While SG3 contains an open vpr reading frame, the virally encoded Vpr protein was not detected under the conditions used. Anti-Gag antibody detected approximately similar amounts of viral Gag proteins (Fig. 2D). These results confirmed that the IN 2 protein can be incorporated into HIV-1 virions when expressed in trans as a fusion partner of Vpr, and subsequently liberated during proteolytic maturation of the virus particle. The IN 2 protein restores the infectivity of HIV-1 IN-minus virus. To analyze whether the IN 2 protein was functional, SG3 S-IN virions complemented with the trans-in 2 protein were analyzed on P4 indicator cells. Our results indicated that virus infectivity was rescued to a level of 16% compared with virions complemented with the homologous HIV-1 trans-in protein (Table 1). This result represents an increase of approximately 100-fold over noncomplemented SG3 S-IN virions. Immunoblot analysis performed on the viral stocks that were used for infection confirmed that the trans-in- and trans-in 2 -complemented virions contained approximately equal amounts of the respective fusion protein (data not shown). These results suggested that the IN 2 protein can function in place of the HIV-1 IN in vivo, albeit with significantly reduced efficiency. IN 2 supports integration of the HIV-1 provirus. To directly examine whether the heterologous trans-in 2 protein could complement the defect in integration of IN-minus HIV-1, a single-cycle integration assay was used. This assay utilized VSV-G-pseudotyped virus, which contains the hygromycin resistance gene within the HIV-1 genome in place of env. In cells infected with pseudotyped virus, where viral DNA is synthesized, integrated, and expressed, the hygromycin resistance gene allows for the outgrowth of cells in the presence of selection medium. VSV-G-pseudotyped, hygromycin-resistant, IN-minus virus (hy-sg3 S-IN ) was complemented by DNA cotransfection with Vpr-IN 2 and Vpr-IN, respectively. Transfection-derived viral stocks were filtered through mpore-size filters, and divided into two aliquot sets. One aliquot set was subjected to ultracentrifugation to pellet virions and was examined by immunoblot analysis. Similar levels (relative to CA protein) of virion associated Vpr-IN and Vpr-IN 2 were detected (data not shown). The second aliquot set was normalized for p24 antigen concentration and was used to infect HeLa CD4 cells in hygromycin selection medium as described earlier (40). Complementation with the IN 2 protein produced Virus TABLE 1. Infectivity of IN-minus HIV-1 complemented with HIV-2 IN protein in trans Result of cotransfection with a : Control Vpr-IN 2 Vpr-IN SG3 wt SG3 S-IN (16) (100) a Four micrograms each of the SG3 wt and SG3 S-IN clones was separately cotransfected into 293T cells with 2 g of the plr2p (control), Vpr-IN 2,or Vpr-IN expression plasmids. Numbers indicate the number of infectious units per 50-ng equivalent (p24 antigen) of input virus, as determined using P4 indicator cells. The numbers in parentheses indicate infectious units relative to the number of Vpr-IN-complemented SG3 S-IN virions, which was arbitrarily set to 100. These results are representative of three independent experiments.
4 8834 NOTES J. VIROL. TABLE 2. Integration of IN mutant HIV-1 complemented with HIV-2 IN protein in trans Hygromycinresistant virus Result of cotransfection with a : Control Vpr-IN 2 Vpr-IN hy-sg3 wt hy-sg3 S-IN (22) (100) hy-sg3 D116A (28) (100) hy-sg3 AA35A (55) (100) a Four micrograms of each of the hy-sg3 wt, hy-sg3 S-IN, hy-sg3 D116A, and hy-sg3 AA35A clones (40) were separately cotransfected into 293T cells with 2 g of the VSV-G env plasmid and 2 g of either the plr2p (control), Vpr-IN 2,or Vpr-IN expression plasmids. Numbers indicate the mean number of hygromycinresistant colonies per 100-ng equivalent (p24 antigen) of input virus. The numbers in parentheses indicate the number of hygromycin-resistant colonies relative to the number produced by Vpr-IN-complemented SG3 S-IN virions, which was arbitrarily set to 100. These results are representative of three independent experiments. 22% of the number of resistant colonies produced by complementation with the homologous trans-in protein (Table 2). Complementation of wild-type virions (hy-sg3 wt ) with either Vpr-IN or Vpr-IN 2 resulted in a slight reduction (approximately twofold) in CFU (data not shown). These result are consistent with the infectivity results (Table 1). We have recently reported that in addition to catalyzing integration, the HIV-1 IN protein plays a nonenzymatic role in the initiation of RT (40). Therefore, to directly analyze the ability of the heterologous IN to catalyze proviral DNA integration, independently of its role in viral DNA synthesis, the Vpr-IN and Vpr-IN 2 fusion proteins were used to complement the hy-sg3 D116A and the hy-sg3 AA35A mutant viruses (40). The hy-sg3 AA35A mutant contains alanine substitutions in each of the three amino acid residues that make up the catalytic triad of IN. While defective in integration, these mutants produce near wild-type levels of viral DNA following infection. Transfection-derived virions were normalized for p24 antigen and used to infect HeLa-CD4 cells. The Vpr-IN 2 -complemented hy-sg3 D116A and hy-sg3 AA35A viruses produced resistant colonies at levels of 28 and 55%, respectively, compared with those complemented with the homologous trans-in protein (Table 2). Taken together, these results clearly show that heterologous IN 2 protein can catalyze integration of the HIV-1 provirus. Moreover, these data confirm reports that the IN 2 protein can efficiently support the integration of HIV-1 DNA substrate in vitro (36 38). IN 2 protein does not efficiently promote HIV-1 DNA synthesis. Several reports have shown that viruses containing certain mutations in IN, including IN deletion mutants, are defective in the synthesis of their viral DNA. We reported that this is predominately due to a role that the IN protein plays in the initiation step of RT. Our published data demonstrated that IN-minus virions (SG3 S-IN ) synthesize reduced levels of viral cdna (5 to 10% of that of wild-type virus) and that this defect can be overcome by providing IN in trans (40). Therefore, the above results (Tables 1 and 2) would suggest that the defect in the infectivity of IN 2 -complemented SG3 S-IN virus was largely due to a block in viral DNA synthesis after infection. To test this directly, we analyzed the synthesis of the nascent viral cdna in infected cells. SG3 S-IN virions containing Vpr-IN 2 were derived by DNA cotransfection and used to infect HeLa- CD4 cells. Eighteen hours later, the cell monolayers were trypsinized, washed extensively, and divided into two aliquot. One aliquot set was lysed and analyzed for intracellular p24 antigen concentration as described earlier (28, 39). DNA was extracted from the other aliquot set, normalized for intercellular p24 antigen concentration, and analyzed for early (R-U5) and late (R-Gag) viral DNA products as described earlier (40). Serial fivefold dilutions of the SG3 wt DNA were prepared and analyzed in parallel as a reference to assess the relative amounts of DNA synthesized by the mutant viruses. The INminus SG3 S-IN virus (lane 7) produced significantly less R-U5 DNA compared with wild-type SG3 wt virions (Fig. 3A). When complemented with the trans-in 2 protein the SG3 S-IN virions produced 18% of the wild-type levels of R-U5 DNA (lane 5). This represents an increase in DNA synthesis of approximately four- to fivefold. Complementation of SG3 S-IN with the homologous trans-in proteins generated 86% the levels of R-U5 DNA compared with SG3 wt. For control, an equivalent (p24 antigen) amount of the SG3 D116A mutant virus was analyzed and found to contain near wild-type levels (89%) of R-U5 DNA. No R-U5 DNA was detected in cells infected with the RT-IN-minus SG3 S-RT virions (40), confirming that the R-U5 DNA detected by PCR was the product of reverse transcription. The absence of this DNA product in cells infected with SG3 S-RT also indicates that the band migrating slightly slower FIG. 3. Reverse transcription of Vpr-IN 2 -complemented viruses. psg3 S-IN was cotransfected into 293T cells by calcium phosphate DNA transfection methods with plr2p-vpr-in 2, plr2p-vprin, and plr2p, respectively. As controls, psg3 wt, psg3 S-RT, and psg3 D116A were also transfected. Forty-eight hours later, culture supernatants were filtered through m-pore-size filters and analyzed by HIV-1 p24 antigen enzyme-linked immunosorbent assay (ELISA) (Coulter Inc.). The virus-containing culture supernatants were normalized to 500 ng of p24 antigen (CA), treated with RNase-free DNase H (20 U/ml for 2 h) (Promega Corporation) and placed on cultures of HeLa-CD4 cells at 37 C. After 4 h, the cell monolayers were washed, trypsinized, resuspended in fetal bovine serum, and divided into two aliquot sets. One aliquot set (which contained one-tenth of the total number of cells) was lysed in phosphate-buffered saline containing 1% Triton X-100 and analyzed by p24 antigen ELISA to quantify intracellular CA protein. The other aliquot set was placed back in culture medium at 37 C for an additional 14 h. The cells were then washed and total DNA was extracted by organic methods. Next, 250-pg equivalents (p24 antigen) of each DNA extract was analyzed by PCR methods for early (R-U5) (A) and late (R-gag) (B) viral DNA products of RT. The amplified products were resolved on 1.5% agarose gels and stained with ethidium bromide. To assess the relative amount of each of the amplified DNA products, four serial fivefold dilutions of the wild-type (SG3 wt ) DNA were analyzed in parallel. The undiluted 250-pg sample was arbitrarily set to 100. As a control for the efficiency of DpnI cleavage of potential carryover plasmid DNA, 6,250 copies of psg3 wt DNA were analyzed after digestion with DpnI as described previously (17, 40). The ethidium bromide staining intensity of each amplified DNA product was measured with a Lynx 5000 molecular biology workstation (Applied Imaging) as described previously (27). The data shown is from a representative experiment that was repeated three times, each time with independent transfection-derived virus preparations.
5 VOL. 73, 1999 NOTES 8835 reverse transcription and preintegration complexes. Moreover, these results suggest that the mere association of the heterologous IN protein with the RT complex is not sufficient to efficiently promote viral DNA synthesis, but rather that specific interactions between the IN protein and other viral components are required. The delivery of heterologous IN protein into HIV-1 particles in trans offers a novel approach to understand IN protein function in vivo. FIG. 4. Complementation of SG3 IN2 virus with homologous trans-in protein. psg3 IN2 and psg3 wt were transfected alone and separately cotransfected into 293T cells with plr2p-vprin. The culture supernatants were collected 48 h later, filtered through m-pore-size filters, and analyzed for HIV-1 p24 antigen by enzyme-linked immunosorbent assay (Coulter Inc.). Next, 25, 5, and 1 ng of each virus (p24 antigen equivalents) was used to infect monolayer cultures of P4 indicator cells. Two days later, the cells were stained and infection-positive cells were counted. The virus infectivity results represent infectious units per 25-ng equivalent of p24 antigen. The data shown are the means from three independent experiments. than the R-Gag band is not a product of viral DNA contamination. Nearly identical results were obtained using the R-Gag primer pair to detect late products of viral DNA synthesis (Fig. 3B). These results show that the IN 2 protein can support the synthesis of HIV-1 DNA, but with significantly reduced efficiency compared with HIV-1 IN. They are also in strong agreement with our results on virus infectivity (Table 1) and together suggest that the factor limiting infectivity was primarily the defect in the synthesis of viral cdna. Complementation of SG3 IN2 virus with trans-in. To better understand the defect in the infectivity of the SG3 IN2 virus (Fig. 1), psg3 IN2 was cotransfected with the HIV-1 trans-in protein (Vpr-IN) and analyzed for infectivity. Figure 4 shows a 27-fold increase in infectivity. This represented an increase that was near that of wild-type virus when complemented with Vpr-IN. This result suggested that the expression of a chimeric Gag-Pol precursor protein did not cause a severe defect in the assembly or maturation of the SG3 IN2 virions. Rather, these data may support our earlier findings for a role of an RT-IN intermediate in the formation of infectious HIV-1 particles (41). In this report, we examined whether the IN 2 protein could mimic the DNA synthesis and integration activities of the HIV-1 IN at the virus replication level. Our data show that while the heterologous IN protein can support HIV-1 DNA synthesis, it is significantly less efficient than the homologous IN protein. The trans-in 2 protein causes an increase in viral DNA synthesis of IN-minus virus of four- to fivefold. However, the HIV-2 trans-in protein appeared to support integration of the provirus with greater efficiency, the integration frequency of the integration-defective D116A mutant virus was increased by almost 200-fold. It was noteworthy that the trans-in 2 protein was consistently less efficient in rescuing the DNA synthesis defect of the SG3 D116A mutant virus compared with that of the SG3 AA35A mutant. It is possible that the D116A mutant IN protein has a dominant-negative effect, perhaps through the formation of nonfunctional heterodimers with the IN 2 protein. The ability of the IN 2 protein to support integration with relatively high efficiency indicates that it associates with the This research was supported by National Institutes of Health grant CA73470 and by the facilities of the AIDS Central Virus and Protein Expression Cores of the Birmingham Center for AIDS Research (P30- AI-27767). This research was also supported by a Merit Review Award funded by the Office of Research and Development, Medical Research Service, U.S. Department of Veterans Affairs. REFERENCES 1. Ansari-Lari, M. A., L. A. Donehower, and R. A. Gibbs Analysis of human immunodeficiency virus type 1 integrase mutants. Virology 211: Ansari-Lari, M. A., and R. A. Gibbs Expression of human immunodeficiency virus type 1 reverse transcriptase in trans during virion release and after infection. J. Virol. 70: Brown, P Integration, p In J. M. Coffin, S. H. Hughes, and H. E. Varmus (ed.), Retroviruses. Cold Spring Harbor Laboratory, Cold Spring Harbor, N.Y. 4. Bukovsky, A., and H. Göttlinger Lack of integrase can markedly affect human immunodeficiency virus type 1 particle production in the presence of an active viral protease. J. Virol. 70: Burke, C. J., G. Sanyal, M. W. Bruner, J. A. Ryan, R. L. LaFemina, H. L. Robbins, A. S. Zeft, C. R. Middaugh, and M. G. Cordingley Structural implication of spectroscopic characterization of a putative zinc-finger peptide from HIV-1 integrase. J. Biol. Chem. 267: Bushman, F. D., A. Engelman, I. Palmer, P. Wingfield, and R. Craigie Domains of the integrase protein of human immunodeficiency virus type 1 responsible for polynucleotidyl transfer and zinc binding. Proc. Natl. Acad. Sci. USA 90: Clavel, F., and P. Charneau Fusion from without directed by human immunodeficiency virus particles. J. Virol. 68: Craigie, R., T. Fujiwara, and F. Bushman The IN protein of Moloney murine leukemia virus processes the viral DNA ends and accomplishes their integration in vitro. Cell 62: Drelich, M., R. Wilhelm, and J. Mous Identification of amino acid residues critical for endonuclease and integration activities of HIV-1 IN in vitro. Virology 188: Engelman, A., and R. Craigie Identification of conserved amino acid residues critical for human immunodeficiency virus type 1 integrase function in vitro. J. Virol. 66: Engelman, A., G. Englund, J. M. Orenstein, M. A. Martin, and R. Craigie Multiple effects of mutations in human immunodeficiency virus type 1 integrase on viral replication. J. Virol. 69: Engelman, A., Y. Liu, H. Chen, M. Farzan, and F. Dyda Structurebased mutagenesis of the catalytic domain of human immunodeficiency virus type 1 integrase. J. Virol. 71: Farnet, C. M., and F. D. Bushman HIV-1 cdna integration: requirement of HMGI(Y) protein for function of preintegration complexes in vitro. Cell 88: Fletcher, T. M., III, M. A. Soares, S. McPhearson, H. Hui, M. Wiskerchen, M. A. Muesing, G. M. Shaw, A. D. Leavitt, J. D. Boeke, and B. H. Hahn Complementation of integrase function in HIV-1 virions. EMBO J. 16: Gallay, P., T. Hope, D. Chin, and D. Trono HIV-1 infection of nondividing cells through the recognition of integrase by the importin/karyopherin pathway. Proc. Natl. Acad. Sci. USA 94: Gallay, P., S. Swingler, J. Song, F. Bushman, and D. Trono HIV nuclear import is governed by the phosphotyrosine-mediated binding of matrix to the core domain of integrase. Cell 83: Heinzinger, N. K., M. I. Bukrinsky, S. A. Haggerty, A. M. Ragland, V. Kewalramani, M.-A. Lee, H. E. Gendelman, L. Ratner, M. Stevenson, and M. Emerman The Vpr protein of human immunodeficiency virus type 1 influences nuclear localization of viral nucleic acids in nondividing host cells. Proc. Natl. Acad. Sci. USA 91: Johnson, M. S., M. A. McClure, D.-F. Feng, J. Gray, and R. F. Doolittle Computer analysis of retroviral pol genes: assignment of enzymatic functions to specific sequences and homologies with nonviral enzymes. Proc. Natl. Acad. Sci. USA 83: Kahn, E., J. P. G. Mack, R. A. Katz, J. Kulkosky, and A. M. Skalka Retroviral integrase domains: DNA binding and the recognition of LTR sequences. Nucleic Acids Res. 19:
6 8836 NOTES J. VIROL. 20. Kalpana, G. V., S. Marmon, W. Wang, G. R. Crabtree, and S. P. Goff Binding and stimulation of HIV-1 integrase by a human homolog of yeast transcription factor SNF5. Science 266: Katzman, M., R. A. Katz, A. M. Skalka, and J. Leis The avian retroviral integration protein cleaves the terminal sequences of linear viral DNA at the in vivo site of integration. J. Virol. 63: Kulkosky, J., K. S. Jones, R. A. Katz, J. P. G. Mack, and A. M. Skalka Residues critical for retroviral integrative recombination in a region that is highly conserved among retroviral/retrotransposon integrases and bacterial insertion sequences transposases. Mol. Cell. Biol. 12: Kumar, P., H. Hui, J. C. Kappes, B. S. Haggarty, J. A. Hoxie, S. K. Arya, G. M. Shaw, and B. H. Hahn Molecular characterization of an attenuated human immunodeficiency virus type 2 isolate. J. Virol. 64: LaFemina, R. L., P. L. Callahan, and M. G. Cordingley Substrate specificity of recombinant human immunodeficiency virus integrase protein. J. Virol. 65: LaFemina, R. L., C. L. Schneider, H. L. Robbins, P. L. Callahan, K. LeGrow, E. Roth, W. A. Schlief, and E. A. Emini Requirement of active human immunodeficiency virus type 1 integrase enzyme for productive infection of human T-lymphoid cells. J. Virol. 66: Leavitt, A. D., G. Robles, N. Alesandro, and H. E. Varmus Human immunodeficiency virus type 1 integrase mutants retain in vitro integrase activity yet fail to integrate viral DNA efficiently during infection. J. Virol. 70: Liu, H., X. Wu, M. Newman, G. M. Shaw, B. H. Hahn, and J. C. Kappes The Vif protein of human and simian immunodeficiency viruses is packaged into virions and associates with viral core structures. J. Virol. 69: Liu, H., X. Wu, H. Xiao, J. A. Conway, and J. C. Kappes Incorporation of functional human immunodeficiency virus type 1 integrase into virions independent of the Gag-Pol precursor protein. J. Virol. 71: Masuda, T., V. Planelles, P. Krogstad, and I. S. Y. Chen Genetic analysis of human immunodeficiency virus type 1 integrase and the U3 att site: unusual phenotype of mutants in the zinc finger-like domain. J. Virol. 69: Miller, M. D., C. M. Farnet, and F. D. Bushman Human immunodeficiency virus type 1 preintegration complexes: studies of organization and composition. J. Virol. 71: Mumm, S. R., and D. P. Grandgenett Defining nucleic acid-binding properties of avian retroviruses integrase by deletion analysis. J. Virol. 65: Sherman, P. A., and J. A. Fyfe Human immunodeficiency virus integration protein expressed in Escherichia coli processes selective DNA cleavage activity. Proc. Natl. Acad. Sci. USA 87: Shin, C.-G., B. Taddeo, W. A. Haseltine, and C. M. Farnet Genetic analysis of the human immunodeficiency virus type 1 integrase protein. J. Virol. 68: Swanstrom, R., and J. W. Wills Synthesis, assembly, and processing of viral proteins, p In J. M. Coffin, S. H. Hughes, and H. E. Varmus (ed.), Retroviruses. Cold Spring Harbor Laboratory, Cold Spring Harbor, N.Y. 35. van Gent, D. C., Y. Elgersma, M. W. J. Bolk, C. Vink, and R. H. A. Plasterk DNA binding properties of the integrase protein of human immunodeficiency viruses types 1 and 2. Nucleic Acids Res. 19: van Gent, D. C., A. A. M. Oude Groeneger, and R. H. A. Plasterk Mutational analysis of the integrase protein of human immunodeficiency virus type 2. Proc. Natl. Acad. Sci. USA 89: van Gent, D. C., C. Vink, A. A. M. Oude Groeneger, and R. H. A. Plasterk Complementation between HIV integrase proteins mutated in different domains. EMBO J. 12: Vink, C., K. H. van der Linden, and R. H. A. Plasterk Activities of the feline immunodeficiency virus integrase protein produced in Escherichia coli. J. Virol. 68: von Schwedler, U., J. Song, C. Aiken, and D. Trono Vif is crucial for human immunodeficiency virus type 1 proviral DNA synthesis in infected cells. J. Virol. 67: Wu, W., H. Liu, H. Xiao, J. A. Conway, E. Hehl, G. V. Kalpana, V. Prasad, and J. C. Kappes Human immunodeficiency virus type 1 integrase protein promotes reverse transcription through specific interactions with the nucleoprotein reverse transcription complex. J. Virol. 73: Wu, X., H. Liu, H. Xiao, J. A. Conway, E. Hunter, and J. C. Kappes Functional RT and IN incorporated into HIV-1 particles independently of the Gag/Pol precursor protein. EMBO J. 16: Downloaded from on September 20, 2018 by guest
Received 8 April 2003/Accepted 16 July 2003
 JOURNAL OF VIROLOGY, Oct. 2003, p. 11050 11059 Vol. 77, No. 20 0022-538X/03/$08.00 0 DOI: 10.1128/JVI.77.20.11050 11059.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved. Replication
JOURNAL OF VIROLOGY, Oct. 2003, p. 11050 11059 Vol. 77, No. 20 0022-538X/03/$08.00 0 DOI: 10.1128/JVI.77.20.11050 11059.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved. Replication
Introduction retroposon
 17.1 - Introduction A retrovirus is an RNA virus able to convert its sequence into DNA by reverse transcription A retroposon (retrotransposon) is a transposon that mobilizes via an RNA form; the DNA element
17.1 - Introduction A retrovirus is an RNA virus able to convert its sequence into DNA by reverse transcription A retroposon (retrotransposon) is a transposon that mobilizes via an RNA form; the DNA element
VIROLOGY. Engineering Viral Genomes: Retrovirus Vectors
 VIROLOGY Engineering Viral Genomes: Retrovirus Vectors Viral vectors Retrovirus replicative cycle Most mammalian retroviruses use trna PRO, trna Lys3, trna Lys1,2 The partially unfolded trna is annealed
VIROLOGY Engineering Viral Genomes: Retrovirus Vectors Viral vectors Retrovirus replicative cycle Most mammalian retroviruses use trna PRO, trna Lys3, trna Lys1,2 The partially unfolded trna is annealed
Fayth K. Yoshimura, Ph.D. September 7, of 7 HIV - BASIC PROPERTIES
 1 of 7 I. Viral Origin. A. Retrovirus - animal lentiviruses. HIV - BASIC PROPERTIES 1. HIV is a member of the Retrovirus family and more specifically it is a member of the Lentivirus genus of this family.
1 of 7 I. Viral Origin. A. Retrovirus - animal lentiviruses. HIV - BASIC PROPERTIES 1. HIV is a member of the Retrovirus family and more specifically it is a member of the Lentivirus genus of this family.
The Karyophilic Properties of Human Immunodeficiency Virus Type 1 Integrase Are Not Required for Nuclear Import of Proviral DNA
 JOURNAL OF VIROLOGY, Aug. 2000, p. 7119 7126 Vol. 74, No. 15 0022-538X/00/$04.00 0 Copyright 2000, American Society for Microbiology. All Rights Reserved. The Karyophilic Properties of Human Immunodeficiency
JOURNAL OF VIROLOGY, Aug. 2000, p. 7119 7126 Vol. 74, No. 15 0022-538X/00/$04.00 0 Copyright 2000, American Society for Microbiology. All Rights Reserved. The Karyophilic Properties of Human Immunodeficiency
Packaging and Abnormal Particle Morphology
 JOURNAL OF VIROLOGY, OCt. 1990, p. 5230-5234 0022-538X/90/105230-05$02.00/0 Copyright 1990, American Society for Microbiology Vol. 64, No. 10 A Mutant of Human Immunodeficiency Virus with Reduced RNA Packaging
JOURNAL OF VIROLOGY, OCt. 1990, p. 5230-5234 0022-538X/90/105230-05$02.00/0 Copyright 1990, American Society for Microbiology Vol. 64, No. 10 A Mutant of Human Immunodeficiency Virus with Reduced RNA Packaging
Fayth K. Yoshimura, Ph.D. September 7, of 7 RETROVIRUSES. 2. HTLV-II causes hairy T-cell leukemia
 1 of 7 I. Diseases Caused by Retroviruses RETROVIRUSES A. Human retroviruses that cause cancers 1. HTLV-I causes adult T-cell leukemia and tropical spastic paraparesis 2. HTLV-II causes hairy T-cell leukemia
1 of 7 I. Diseases Caused by Retroviruses RETROVIRUSES A. Human retroviruses that cause cancers 1. HTLV-I causes adult T-cell leukemia and tropical spastic paraparesis 2. HTLV-II causes hairy T-cell leukemia
Sequences in the 5 and 3 R Elements of Human Immunodeficiency Virus Type 1 Critical for Efficient Reverse Transcription
 JOURNAL OF VIROLOGY, Sept. 2000, p. 8324 8334 Vol. 74, No. 18 0022-538X/00/$04.00 0 Copyright 2000, American Society for Microbiology. All Rights Reserved. Sequences in the 5 and 3 R Elements of Human
JOURNAL OF VIROLOGY, Sept. 2000, p. 8324 8334 Vol. 74, No. 18 0022-538X/00/$04.00 0 Copyright 2000, American Society for Microbiology. All Rights Reserved. Sequences in the 5 and 3 R Elements of Human
Retroviruses. ---The name retrovirus comes from the enzyme, reverse transcriptase.
 Retroviruses ---The name retrovirus comes from the enzyme, reverse transcriptase. ---Reverse transcriptase (RT) converts the RNA genome present in the virus particle into DNA. ---RT discovered in 1970.
Retroviruses ---The name retrovirus comes from the enzyme, reverse transcriptase. ---Reverse transcriptase (RT) converts the RNA genome present in the virus particle into DNA. ---RT discovered in 1970.
Qin Yu and Casey D. Morrow 1. Department of Microbiology, University of Alabama at Birmingham, Birmingham, Alabama 35294
 Virology 254, 160 168 (1999) Article ID viro.1998.9542, available online at http://www.idealibrary.com on Complementarity between 3 Terminal Nucleotides of trna and Primer Binding Site Is a Major Determinant
Virology 254, 160 168 (1999) Article ID viro.1998.9542, available online at http://www.idealibrary.com on Complementarity between 3 Terminal Nucleotides of trna and Primer Binding Site Is a Major Determinant
The HIV life cycle. integration. virus production. entry. transcription. reverse transcription. nuclear import
 The HIV life cycle entry reverse transcription transcription integration virus production nuclear import Hazuda 2012 Integration Insertion of the viral DNA into host chromosomal DNA, essential step in
The HIV life cycle entry reverse transcription transcription integration virus production nuclear import Hazuda 2012 Integration Insertion of the viral DNA into host chromosomal DNA, essential step in
Supplementary information. MARCH8 inhibits HIV-1 infection by reducing virion incorporation of envelope glycoproteins
 Supplementary information inhibits HIV-1 infection by reducing virion incorporation of envelope glycoproteins Takuya Tada, Yanzhao Zhang, Takayoshi Koyama, Minoru Tobiume, Yasuko Tsunetsugu-Yokota, Shoji
Supplementary information inhibits HIV-1 infection by reducing virion incorporation of envelope glycoproteins Takuya Tada, Yanzhao Zhang, Takayoshi Koyama, Minoru Tobiume, Yasuko Tsunetsugu-Yokota, Shoji
The Barrier-to-Autointegration Factor Is a Component of Functional Human Immunodeficiency Virus Type 1 Preintegration Complexes
 JOURNAL OF VIROLOGY, Apr. 2003, p. 5030 5036 Vol. 77, No. 8 0022-538X/03/$08.00 0 DOI: 10.1128/JVI.77.8.5030 5036.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved. The Barrier-to-Autointegration
JOURNAL OF VIROLOGY, Apr. 2003, p. 5030 5036 Vol. 77, No. 8 0022-538X/03/$08.00 0 DOI: 10.1128/JVI.77.8.5030 5036.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved. The Barrier-to-Autointegration
Supplementary Material
 Supplementary Material Nuclear import of purified HIV-1 Integrase. Integrase remains associated to the RTC throughout the infection process until provirus integration occurs and is therefore one likely
Supplementary Material Nuclear import of purified HIV-1 Integrase. Integrase remains associated to the RTC throughout the infection process until provirus integration occurs and is therefore one likely
Materials and Methods , The two-hybrid principle.
 The enzymatic activity of an unknown protein which cleaves the phosphodiester bond between the tyrosine residue of a viral protein and the 5 terminus of the picornavirus RNA Introduction Every day there
The enzymatic activity of an unknown protein which cleaves the phosphodiester bond between the tyrosine residue of a viral protein and the 5 terminus of the picornavirus RNA Introduction Every day there
Differential Effects of C-Terminal Molecular Tagged Integrase on Replication Competent Moloney Murine Leukemia Virus
 Virology 274, 412 419 (2000) doi:10.1006/viro.2000.0481, available online at http://www.idealibrary.com on Differential Effects of C-Terminal Molecular Tagged Integrase on Replication Competent Moloney
Virology 274, 412 419 (2000) doi:10.1006/viro.2000.0481, available online at http://www.idealibrary.com on Differential Effects of C-Terminal Molecular Tagged Integrase on Replication Competent Moloney
Received 26 October 2005/Accepted 1 December 2005
 JOURNAL OF VIROLOGY, Feb. 2006, p. 1939 1948 Vol. 80, No. 4 0022-538X/06/$08.00 0 doi:10.1128/jvi.80.4.1939 1948.2006 Copyright 2006, American Society for Microbiology. All Rights Reserved. Human Immunodeficiency
JOURNAL OF VIROLOGY, Feb. 2006, p. 1939 1948 Vol. 80, No. 4 0022-538X/06/$08.00 0 doi:10.1128/jvi.80.4.1939 1948.2006 Copyright 2006, American Society for Microbiology. All Rights Reserved. Human Immunodeficiency
Viral Vectors In The Research Laboratory: Just How Safe Are They? Dawn P. Wooley, Ph.D., SM(NRM), RBP, CBSP
 Viral Vectors In The Research Laboratory: Just How Safe Are They? Dawn P. Wooley, Ph.D., SM(NRM), RBP, CBSP 1 Learning Objectives Recognize hazards associated with viral vectors in research and animal
Viral Vectors In The Research Laboratory: Just How Safe Are They? Dawn P. Wooley, Ph.D., SM(NRM), RBP, CBSP 1 Learning Objectives Recognize hazards associated with viral vectors in research and animal
Recombinant Protein Expression Retroviral system
 Recombinant Protein Expression Retroviral system Viruses Contains genome DNA or RNA Genome encased in a protein coat or capsid. Some viruses have membrane covering protein coat enveloped virus Ø Essential
Recombinant Protein Expression Retroviral system Viruses Contains genome DNA or RNA Genome encased in a protein coat or capsid. Some viruses have membrane covering protein coat enveloped virus Ø Essential
Protein Produced in Escherichia coli
 JOURNAL OF VIROLOGY, Mar. 1994, P. 1468-1474 Vol. 68, No. 3 0022-538X/94/$04.00+0 Copyright 1994, American Society for Microbiology Activities of the Feline Immunodeficiency Virus Integrase Protein Produced
JOURNAL OF VIROLOGY, Mar. 1994, P. 1468-1474 Vol. 68, No. 3 0022-538X/94/$04.00+0 Copyright 1994, American Society for Microbiology Activities of the Feline Immunodeficiency Virus Integrase Protein Produced
Julianne Edwards. Retroviruses. Spring 2010
 Retroviruses Spring 2010 A retrovirus can simply be referred to as an infectious particle which replicates backwards even though there are many different types of retroviruses. More specifically, a retrovirus
Retroviruses Spring 2010 A retrovirus can simply be referred to as an infectious particle which replicates backwards even though there are many different types of retroviruses. More specifically, a retrovirus
Hepatitis B Antiviral Drug Development Multi-Marker Screening Assay
 Hepatitis B Antiviral Drug Development Multi-Marker Screening Assay Background ImQuest BioSciences has developed and qualified a single-plate method to expedite the screening of antiviral agents against
Hepatitis B Antiviral Drug Development Multi-Marker Screening Assay Background ImQuest BioSciences has developed and qualified a single-plate method to expedite the screening of antiviral agents against
Mark Pandori,* Heather Craig, Laure Moutouh, Jacques Corbeil, and John Guatelli*,,1
 VIROLOGY 251, 302 316 (1998) ARTICLE NO. VY989407 Virological Importance of the Protease-Cleavage Site in Human Immunodeficiency Virus Type 1 Nef Is Independent of both Intravirion Processing and CD4 Down-regulation
VIROLOGY 251, 302 316 (1998) ARTICLE NO. VY989407 Virological Importance of the Protease-Cleavage Site in Human Immunodeficiency Virus Type 1 Nef Is Independent of both Intravirion Processing and CD4 Down-regulation
MedChem 401~ Retroviridae. Retroviridae
 MedChem 401~ Retroviridae Retroviruses plus-sense RNA genome (!8-10 kb) protein capsid lipid envelop envelope glycoproteins reverse transcriptase enzyme integrase enzyme protease enzyme Retroviridae The
MedChem 401~ Retroviridae Retroviruses plus-sense RNA genome (!8-10 kb) protein capsid lipid envelop envelope glycoproteins reverse transcriptase enzyme integrase enzyme protease enzyme Retroviridae The
L I F E S C I E N C E S
 1a L I F E S C I E N C E S 5 -UUA AUA UUC GAA AGC UGC AUC GAA AAC UGU GAA UCA-3 5 -TTA ATA TTC GAA AGC TGC ATC GAA AAC TGT GAA TCA-3 3 -AAT TAT AAG CTT TCG ACG TAG CTT TTG ACA CTT AGT-5 OCTOBER 31, 2006
1a L I F E S C I E N C E S 5 -UUA AUA UUC GAA AGC UGC AUC GAA AAC UGU GAA UCA-3 5 -TTA ATA TTC GAA AGC TGC ATC GAA AAC TGT GAA TCA-3 3 -AAT TAT AAG CTT TCG ACG TAG CTT TTG ACA CTT AGT-5 OCTOBER 31, 2006
Nuclear Localization of Human Immunodeficiency Virus Type 1 Integrase Expressed as a Fusion Protein with Green Fluorescent Protein
 Virology 258, 327 332 (1999) Article ID viro.1999.9727, available online at http://www.idealibrary.com on Nuclear Localization of Human Immunodeficiency Virus Type 1 Integrase Expressed as a Fusion Protein
Virology 258, 327 332 (1999) Article ID viro.1999.9727, available online at http://www.idealibrary.com on Nuclear Localization of Human Immunodeficiency Virus Type 1 Integrase Expressed as a Fusion Protein
Supplemental Materials and Methods Plasmids and viruses Quantitative Reverse Transcription PCR Generation of molecular standard for quantitative PCR
 Supplemental Materials and Methods Plasmids and viruses To generate pseudotyped viruses, the previously described recombinant plasmids pnl4-3-δnef-gfp or pnl4-3-δ6-drgfp and a vector expressing HIV-1 X4
Supplemental Materials and Methods Plasmids and viruses To generate pseudotyped viruses, the previously described recombinant plasmids pnl4-3-δnef-gfp or pnl4-3-δ6-drgfp and a vector expressing HIV-1 X4
Peptide hydrolysis uncatalyzed half-life = ~450 years HIV protease-catalyzed half-life = ~3 seconds
 Uncatalyzed half-life Peptide hydrolysis uncatalyzed half-life = ~450 years IV protease-catalyzed half-life = ~3 seconds Life Sciences 1a Lecture Slides Set 9 Fall 2006-2007 Prof. David R. Liu In the absence
Uncatalyzed half-life Peptide hydrolysis uncatalyzed half-life = ~450 years IV protease-catalyzed half-life = ~3 seconds Life Sciences 1a Lecture Slides Set 9 Fall 2006-2007 Prof. David R. Liu In the absence
Mutations of the Human Immunodeficiency Virus Type 1 p6 Gag Domain Result in Reduced Retention of Pol Proteins during Virus Assembly
 JOURNAL OF VIROLOGY, Apr. 1998, p. 3412 3417 Vol. 72, No. 4 0022-538X/98/$04.00 0 Copyright 1998, American Society for Microbiology Mutations of the Human Immunodeficiency Virus Type 1 p6 Gag Domain Result
JOURNAL OF VIROLOGY, Apr. 1998, p. 3412 3417 Vol. 72, No. 4 0022-538X/98/$04.00 0 Copyright 1998, American Society for Microbiology Mutations of the Human Immunodeficiency Virus Type 1 p6 Gag Domain Result
HIV Nuclear Entry: Clearing the Fog
 Viruses 2010, 2, 1190-1194; doi:10.3390/v2051190 OPEN ACCESS viruses ISSN 1999-4915 www.mdpi.com/journal/viruses Commentary HIV Nuclear Entry: Clearing the Fog Vaibhav B. Shah and Christopher Aiken * Department
Viruses 2010, 2, 1190-1194; doi:10.3390/v2051190 OPEN ACCESS viruses ISSN 1999-4915 www.mdpi.com/journal/viruses Commentary HIV Nuclear Entry: Clearing the Fog Vaibhav B. Shah and Christopher Aiken * Department
Structure-Based Mutagenesis of the Catalytic Domain of Human Immunodeficiency Virus Type 1 Integrase
 JOURNAL OF VIROLOGY, May 1997, p. 3507 3514 Vol. 71, No. 5 0022-538X/97/$04.00 0 Copyright 1997, American Society for Microbiology Structure-Based Mutagenesis of the Catalytic Domain of Human Immunodeficiency
JOURNAL OF VIROLOGY, May 1997, p. 3507 3514 Vol. 71, No. 5 0022-538X/97/$04.00 0 Copyright 1997, American Society for Microbiology Structure-Based Mutagenesis of the Catalytic Domain of Human Immunodeficiency
number Done by Corrected by Doctor Ashraf
 number 4 Done by Nedaa Bani Ata Corrected by Rama Nada Doctor Ashraf Genome replication and gene expression Remember the steps of viral replication from the last lecture: Attachment, Adsorption, Penetration,
number 4 Done by Nedaa Bani Ata Corrected by Rama Nada Doctor Ashraf Genome replication and gene expression Remember the steps of viral replication from the last lecture: Attachment, Adsorption, Penetration,
AND JEREMY LUBAN 1,2 *
 JOURNAL OF VIROLOGY, Oct. 1999, p. 8527 8540 Vol. 73, No. 10 0022-538X/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. Human Immunodeficiency Virus Type 1 Gag Polyprotein
JOURNAL OF VIROLOGY, Oct. 1999, p. 8527 8540 Vol. 73, No. 10 0022-538X/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. Human Immunodeficiency Virus Type 1 Gag Polyprotein
Polyomaviridae. Spring
 Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
Polyomaviridae Spring 2002 331 Antibody Prevalence for BK & JC Viruses Spring 2002 332 Polyoma Viruses General characteristics Papovaviridae: PA - papilloma; PO - polyoma; VA - vacuolating agent a. 45nm
Human Immunodeficiency Virus
 Human Immunodeficiency Virus Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Viruses and hosts Lentivirus from Latin lentis (slow), for slow progression of disease
Human Immunodeficiency Virus Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Viruses and hosts Lentivirus from Latin lentis (slow), for slow progression of disease
DATA SHEET. Provided: 500 µl of 5.6 mm Tris HCl, 4.4 mm Tris base, 0.05% sodium azide 0.1 mm EDTA, 5 mg/liter calf thymus DNA.
 Viral Load DNA >> Standard PCR standard 0 Copies Catalog Number: 1122 Lot Number: 150298 Release Category: A Provided: 500 µl of 5.6 mm Tris HCl, 4.4 mm Tris base, 0.05% sodium azide 0.1 mm EDTA, 5 mg/liter
Viral Load DNA >> Standard PCR standard 0 Copies Catalog Number: 1122 Lot Number: 150298 Release Category: A Provided: 500 µl of 5.6 mm Tris HCl, 4.4 mm Tris base, 0.05% sodium azide 0.1 mm EDTA, 5 mg/liter
Section 6. Junaid Malek, M.D.
 Section 6 Junaid Malek, M.D. The Golgi and gp160 gp160 transported from ER to the Golgi in coated vesicles These coated vesicles fuse to the cis portion of the Golgi and deposit their cargo in the cisternae
Section 6 Junaid Malek, M.D. The Golgi and gp160 gp160 transported from ER to the Golgi in coated vesicles These coated vesicles fuse to the cis portion of the Golgi and deposit their cargo in the cisternae
Lecture 2: Virology. I. Background
 Lecture 2: Virology I. Background A. Properties 1. Simple biological systems a. Aggregates of nucleic acids and protein 2. Non-living a. Cannot reproduce or carry out metabolic activities outside of a
Lecture 2: Virology I. Background A. Properties 1. Simple biological systems a. Aggregates of nucleic acids and protein 2. Non-living a. Cannot reproduce or carry out metabolic activities outside of a
Hsu-Chen Chiu, Szu-Yung Yao, and Chin-Tien Wang*
 JOURNAL OF VIROLOGY, Apr. 2002, p. 3221 3231 Vol. 76, No. 7 0022-538X/02/$04.00 0 DOI: 10.1128/JVI.76.7.3221 3231.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved. Coding Sequences
JOURNAL OF VIROLOGY, Apr. 2002, p. 3221 3231 Vol. 76, No. 7 0022-538X/02/$04.00 0 DOI: 10.1128/JVI.76.7.3221 3231.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved. Coding Sequences
A Novel Approach for Producing Lentiviruses That Are Limited to a Single Cycle of Infection
 JOURNAL OF VIROLOGY, Nov. 2004, p. 11715 11725 Vol. 78, No. 21 0022-538X/04/$08.00 0 DOI: 10.1128/JVI.78.21.11715 11725.2004 Copyright 2004, American Society for Microbiology. All Rights Reserved. A Novel
JOURNAL OF VIROLOGY, Nov. 2004, p. 11715 11725 Vol. 78, No. 21 0022-538X/04/$08.00 0 DOI: 10.1128/JVI.78.21.11715 11725.2004 Copyright 2004, American Society for Microbiology. All Rights Reserved. A Novel
Role of the C terminus Gag protein in human immunodefieieney virus type 1 virion assembly and maturation
 Journal of General Virology (1995), 76, 3171-3179. Printed in Great Britain 3171 Role of the C terminus Gag protein in human immunodefieieney virus type 1 virion assembly and maturation X.-F. Yu, ~ Z.
Journal of General Virology (1995), 76, 3171-3179. Printed in Great Britain 3171 Role of the C terminus Gag protein in human immunodefieieney virus type 1 virion assembly and maturation X.-F. Yu, ~ Z.
INI1/hSNF5-interaction defective HIV-1 IN mutants exhibit impaired particle morphology, reverse transcription and integration in vivo
 Mathew et al. Retrovirology 213, 1:66 RESEARCH Open Access INI1/hSNF5-interaction defective HIV-1 IN mutants exhibit impaired particle morphology, reverse transcription and integration in vivo Sheeba Mathew
Mathew et al. Retrovirology 213, 1:66 RESEARCH Open Access INI1/hSNF5-interaction defective HIV-1 IN mutants exhibit impaired particle morphology, reverse transcription and integration in vivo Sheeba Mathew
JOHN G. JULIAS, ANDREA L. FERRIS, PAUL L. BOYER, AND STEPHEN H. HUGHES* HIV-Drug Resistance Program, NCI-Frederick, Frederick, Maryland
 JOURNAL OF VIROLOGY, July 2001, p. 6537 6546 Vol. 75, No. 14 0022-538X/01/$04.00 0 DOI: 10.1128/JVI.75.14.6537 6546.2001 Replication of Phenotypically Mixed Human Immunodeficiency Virus Type 1 Virions
JOURNAL OF VIROLOGY, July 2001, p. 6537 6546 Vol. 75, No. 14 0022-538X/01/$04.00 0 DOI: 10.1128/JVI.75.14.6537 6546.2001 Replication of Phenotypically Mixed Human Immunodeficiency Virus Type 1 Virions
Modeling the HIV-1 Intasome: A Prototype View of the Target of Integrase Inhibitors
 Viruses 2010, 2, 2777-2781; doi:10.3390/v2122777 OPEN ACCESS viruses ISSN 1999-4915 www.mdpi.com/journal/viruses Commentary Modeling the HIV-1 Intasome: A Prototype View of the Target of Integrase Inhibitors
Viruses 2010, 2, 2777-2781; doi:10.3390/v2122777 OPEN ACCESS viruses ISSN 1999-4915 www.mdpi.com/journal/viruses Commentary Modeling the HIV-1 Intasome: A Prototype View of the Target of Integrase Inhibitors
Choosing Optimal Viral Vector for T-cell Transduction. Viral vectors for blood cells
 Choosing Optimal Viral Vector for T-cell Transduction Max Mamonkin, PhD Center for Cell and Gene Therapy Baylor College of Medicine PACT Webinar Nov 08, 2018 Viral for blood cells Short/long term gene
Choosing Optimal Viral Vector for T-cell Transduction Max Mamonkin, PhD Center for Cell and Gene Therapy Baylor College of Medicine PACT Webinar Nov 08, 2018 Viral for blood cells Short/long term gene
Primate and Feline Lentivirus Vector RNA Packaging and Propagation by Heterologous Lentivirus Virions
 JOURNAL OF VIROLOGY, June 2001, p. 5129 5140 Vol. 75, No. 11 0022-538X/01/$04.00 0 DOI: 10.1128/JVI.75.11.5129 5140.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved. Primate
JOURNAL OF VIROLOGY, June 2001, p. 5129 5140 Vol. 75, No. 11 0022-538X/01/$04.00 0 DOI: 10.1128/JVI.75.11.5129 5140.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved. Primate
Identification of Mutation(s) in. Associated with Neutralization Resistance. Miah Blomquist
 Identification of Mutation(s) in the HIV 1 gp41 Subunit Associated with Neutralization Resistance Miah Blomquist What is HIV 1? HIV-1 is an epidemic that affects over 34 million people worldwide. HIV-1
Identification of Mutation(s) in the HIV 1 gp41 Subunit Associated with Neutralization Resistance Miah Blomquist What is HIV 1? HIV-1 is an epidemic that affects over 34 million people worldwide. HIV-1
Fine Mapping of a cis-acting Sequence Element in Yellow Fever Virus RNA That Is Required for RNA Replication and Cyclization
 JOURNAL OF VIROLOGY, Feb. 2003, p. 2265 2270 Vol. 77, No. 3 0022-538X/03/$08.00 0 DOI: 10.1128/JVI.77.3.2265 2270.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved. Fine Mapping
JOURNAL OF VIROLOGY, Feb. 2003, p. 2265 2270 Vol. 77, No. 3 0022-538X/03/$08.00 0 DOI: 10.1128/JVI.77.3.2265 2270.2003 Copyright 2003, American Society for Microbiology. All Rights Reserved. Fine Mapping
ARV Mode of Action. Mode of Action. Mode of Action NRTI. Immunopaedia.org.za
 ARV Mode of Action Mode of Action Mode of Action - NRTI Mode of Action - NNRTI Mode of Action - Protease Inhibitors Mode of Action - Integrase inhibitor Mode of Action - Entry Inhibitors Mode of Action
ARV Mode of Action Mode of Action Mode of Action - NRTI Mode of Action - NNRTI Mode of Action - Protease Inhibitors Mode of Action - Integrase inhibitor Mode of Action - Entry Inhibitors Mode of Action
Use of Patient-Derived Human Immunodeficiency Virus Type 1 Integrases To Identify a Protein Residue That Affects Target Site Selection
 JOURNAL OF VIROLOGY, Aug. 2001, p. 7756 7762 Vol. 75, No. 16 0022-538X/01/$04.00 0 DOI: 10.1128/JVI.75.16.7756 7762.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved. Use of Patient-Derived
JOURNAL OF VIROLOGY, Aug. 2001, p. 7756 7762 Vol. 75, No. 16 0022-538X/01/$04.00 0 DOI: 10.1128/JVI.75.16.7756 7762.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved. Use of Patient-Derived
Chapter 19: Viruses. 1. Viral Structure & Reproduction. 2. Bacteriophages. 3. Animal Viruses. 4. Viroids & Prions
 Chapter 19: Viruses 1. Viral Structure & Reproduction 2. Bacteriophages 3. Animal Viruses 4. Viroids & Prions 1. Viral Structure & Reproduction Chapter Reading pp. 393-396 What exactly is a Virus? Viruses
Chapter 19: Viruses 1. Viral Structure & Reproduction 2. Bacteriophages 3. Animal Viruses 4. Viroids & Prions 1. Viral Structure & Reproduction Chapter Reading pp. 393-396 What exactly is a Virus? Viruses
Charged Amino Acid Residues of Human Immunodeficiency Virus Type 1 Nucleocapsid p7 Protein Involved in RNA Packaging and Infectivity
 JOURNAL OF VIROLOGY, Oct. 1996, p. 6607 6616 Vol. 70, No. 10 0022-538X/96/$04.00 0 Copyright 1996, American Society for Microbiology Charged Amino Acid Residues of Human Immunodeficiency Virus Type 1 Nucleocapsid
JOURNAL OF VIROLOGY, Oct. 1996, p. 6607 6616 Vol. 70, No. 10 0022-538X/96/$04.00 0 Copyright 1996, American Society for Microbiology Charged Amino Acid Residues of Human Immunodeficiency Virus Type 1 Nucleocapsid
HIV INFECTION: An Overview
 HIV INFECTION: An Overview UNIVERSITY OF PAPUA NEW GUINEA SCHOOL OF MEDICINE AND HEALTH SCIENCES DIVISION OF BASIC MEDICAL SCIENCES DISCIPLINE OF BIOCHEMISTRY & MOLECULAR BIOLOGY PBL MBBS II SEMINAR VJ
HIV INFECTION: An Overview UNIVERSITY OF PAPUA NEW GUINEA SCHOOL OF MEDICINE AND HEALTH SCIENCES DIVISION OF BASIC MEDICAL SCIENCES DISCIPLINE OF BIOCHEMISTRY & MOLECULAR BIOLOGY PBL MBBS II SEMINAR VJ
~Lentivirus production~
 ~Lentivirus production~ May 30, 2008 RNAi core R&D group member Lentivirus Production Session Lentivirus!!! Is it health threatening to lab technician? What s so good about this RNAi library? How to produce
~Lentivirus production~ May 30, 2008 RNAi core R&D group member Lentivirus Production Session Lentivirus!!! Is it health threatening to lab technician? What s so good about this RNAi library? How to produce
Last time we talked about the few steps in viral replication cycle and the un-coating stage:
 Zeina Al-Momani Last time we talked about the few steps in viral replication cycle and the un-coating stage: Un-coating: is a general term for the events which occur after penetration, we talked about
Zeina Al-Momani Last time we talked about the few steps in viral replication cycle and the un-coating stage: Un-coating: is a general term for the events which occur after penetration, we talked about
Inhibition of trna 3 Lys -Primed Reverse Transcription by Human APOBEC3G during Human Immunodeficiency Virus Type 1 Replication
 JOURNAL OF VIROLOGY, Dec. 2006, p. 11710 11722 Vol. 80, No. 23 0022-538X/06/$08.00 0 doi:10.1128/jvi.01038-06 Copyright 2006, American Society for Microbiology. All Rights Reserved. Inhibition of trna
JOURNAL OF VIROLOGY, Dec. 2006, p. 11710 11722 Vol. 80, No. 23 0022-538X/06/$08.00 0 doi:10.1128/jvi.01038-06 Copyright 2006, American Society for Microbiology. All Rights Reserved. Inhibition of trna
Identification of the Nuclear Localization Signal of Human Immunodeficiency Virus Type 2 Vpx
 University of Nebraska - Lincoln DigitalCommons@University of Nebraska - Lincoln Virology Papers Virology, Nebraska Center for June 2003 Identification of the Nuclear Localization Signal of Human Immunodeficiency
University of Nebraska - Lincoln DigitalCommons@University of Nebraska - Lincoln Virology Papers Virology, Nebraska Center for June 2003 Identification of the Nuclear Localization Signal of Human Immunodeficiency
7.012 Quiz 3 Answers
 MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel Friday 11/12/04 7.012 Quiz 3 Answers A > 85 B 72-84
MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel Friday 11/12/04 7.012 Quiz 3 Answers A > 85 B 72-84
Association of Murine Leukemia Virus Pol with Virions, Independent of Gag-Pol Expression
 JOURNAL OF VIROLOGY, Nov. 1999, p. 9632 9637 Vol. 73, No. 11 0022-538X/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. Association of Murine Leukemia Virus Pol with
JOURNAL OF VIROLOGY, Nov. 1999, p. 9632 9637 Vol. 73, No. 11 0022-538X/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. Association of Murine Leukemia Virus Pol with
Structural vs. nonstructural proteins
 Why would you want to study proteins associated with viruses or virus infection? Receptors Mechanism of uncoating How is gene expression carried out, exclusively by viral enzymes? Gene expression phases?
Why would you want to study proteins associated with viruses or virus infection? Receptors Mechanism of uncoating How is gene expression carried out, exclusively by viral enzymes? Gene expression phases?
Chapter 19: Viruses. 1. Viral Structure & Reproduction. What exactly is a Virus? 11/7/ Viral Structure & Reproduction. 2.
 Chapter 19: Viruses 1. Viral Structure & Reproduction 2. Bacteriophages 3. Animal Viruses 4. Viroids & Prions 1. Viral Structure & Reproduction Chapter Reading pp. 393-396 What exactly is a Virus? Viruses
Chapter 19: Viruses 1. Viral Structure & Reproduction 2. Bacteriophages 3. Animal Viruses 4. Viroids & Prions 1. Viral Structure & Reproduction Chapter Reading pp. 393-396 What exactly is a Virus? Viruses
Frequent Segregation of More-Defective Variants from a Rous Sarcoma Virus Packaging Mutant, TK15
 JOURNAL OF VIROLOGY, Oct. 1987, p. 3208-3213 0022-538X/87/103208-06$02.00/0 Copyright 1987, American Society for Microbiology Vol. 61, No. 10 Frequent Segregation of More-Defective Variants from a Rous
JOURNAL OF VIROLOGY, Oct. 1987, p. 3208-3213 0022-538X/87/103208-06$02.00/0 Copyright 1987, American Society for Microbiology Vol. 61, No. 10 Frequent Segregation of More-Defective Variants from a Rous
RAPID COMMUNICATION. Canine Cyclin T1 Rescues Equine Infectious Anemia Virus Tat Trans-Activation in Human Cells
 Virology 268, 7 11 (2000) doi:10.1006/viro.1999.0141, available online at http://www.idealibrary.com on RAPID COMMUNICATION Canine Cyclin T1 Rescues Equine Infectious Anemia Virus Tat Trans-Activation
Virology 268, 7 11 (2000) doi:10.1006/viro.1999.0141, available online at http://www.idealibrary.com on RAPID COMMUNICATION Canine Cyclin T1 Rescues Equine Infectious Anemia Virus Tat Trans-Activation
CURRENT DEVELOMENTS AND FUTURE PROSPECTS FOR HIV GENE THERAPY USING INTERFERING RNA-BASED STRATEGIES
 [Frontiers in Bioscience 5, d527-555, May 1, 2000] CURRENT DEVELOMENTS AND FUTURE PROSPECTS FOR HIV GENE THERAPY USING INTERFERING RNA-BASED STRATEGIES Betty Lamothe, Sadhna Joshi Department of Medical
[Frontiers in Bioscience 5, d527-555, May 1, 2000] CURRENT DEVELOMENTS AND FUTURE PROSPECTS FOR HIV GENE THERAPY USING INTERFERING RNA-BASED STRATEGIES Betty Lamothe, Sadhna Joshi Department of Medical
There are approximately 30,000 proteasomes in a typical human cell Each proteasome is approximately 700 kda in size The proteasome is made up of 3
 Proteasomes Proteasomes Proteasomes are responsible for degrading proteins that have been damaged, assembled improperly, or that are of no profitable use to the cell. The unwanted protein is literally
Proteasomes Proteasomes Proteasomes are responsible for degrading proteins that have been damaged, assembled improperly, or that are of no profitable use to the cell. The unwanted protein is literally
HIV/AIDS. Biology of HIV. Research Feature. Related Links. See Also
 6/1/2011 Biology of HIV Biology of HIV HIV belongs to a class of viruses known as retroviruses. Retroviruses are viruses that contain RNA (ribonucleic acid) as their genetic material. After infecting a
6/1/2011 Biology of HIV Biology of HIV HIV belongs to a class of viruses known as retroviruses. Retroviruses are viruses that contain RNA (ribonucleic acid) as their genetic material. After infecting a
Human Immunodeficiency Virus Type 2 Preintegration Complexes: Activities In Vitro and Response to Inhibitors
 JOURNAL OF VIROLOGY, Apr. 1997, p. 3351 3356 Vol. 71, No. 4 0022-538X/97/$04.00 0 Copyright 1997, American Society for Microbiology Human Immunodeficiency Virus Type 2 Preintegration Complexes: Activities
JOURNAL OF VIROLOGY, Apr. 1997, p. 3351 3356 Vol. 71, No. 4 0022-538X/97/$04.00 0 Copyright 1997, American Society for Microbiology Human Immunodeficiency Virus Type 2 Preintegration Complexes: Activities
Multi-step inhibition explains HIV-1 protease inhibitor pharmacodynamics and resistance
 Research article Related article, page 3704 Multi-step inhibition explains HIV-1 protease inhibitor pharmacodynamics and resistance S. Alireza Rabi, 1 Gregory M. Laird, 1 Christine M. Durand, 1 Sarah Laskey,
Research article Related article, page 3704 Multi-step inhibition explains HIV-1 protease inhibitor pharmacodynamics and resistance S. Alireza Rabi, 1 Gregory M. Laird, 1 Christine M. Durand, 1 Sarah Laskey,
October 26, Lecture Readings. Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell
 October 26, 2006 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
October 26, 2006 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
Structure-Based Mutagenesis of the Human Immunodeficiency Virus Type 1 DNA Attachment Site: Effects on Integration and cdna Synthesis
 JOURNAL OF VIROLOGY, Nov. 1999, p. 9011 9020 Vol. 73, No. 11 0022-538X/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. Structure-Based Mutagenesis of the Human Immunodeficiency
JOURNAL OF VIROLOGY, Nov. 1999, p. 9011 9020 Vol. 73, No. 11 0022-538X/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. Structure-Based Mutagenesis of the Human Immunodeficiency
LESSON 4.4 WORKBOOK. How viruses make us sick: Viral Replication
 DEFINITIONS OF TERMS Eukaryotic: Non-bacterial cell type (bacteria are prokaryotes).. LESSON 4.4 WORKBOOK How viruses make us sick: Viral Replication This lesson extends the principles we learned in Unit
DEFINITIONS OF TERMS Eukaryotic: Non-bacterial cell type (bacteria are prokaryotes).. LESSON 4.4 WORKBOOK How viruses make us sick: Viral Replication This lesson extends the principles we learned in Unit
Oligomerization within Virions and Subcellular Localization of Human Immunodeficiency Virus Type 1 Integrase
 JOURNAL OF VIROLOGY, June 1999, p. 5079 5088 Vol. 73, No. 6 0022-538X/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. Oligomerization within Virions and Subcellular
JOURNAL OF VIROLOGY, June 1999, p. 5079 5088 Vol. 73, No. 6 0022-538X/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. Oligomerization within Virions and Subcellular
Reverse transcription and integration
 Reverse transcription and integration Lecture 9 Biology 3310/4310 Virology Spring 2018 One can t believe impossible things, said Alice. I dare say you haven t had much practice, said the Queen. Why, sometimes
Reverse transcription and integration Lecture 9 Biology 3310/4310 Virology Spring 2018 One can t believe impossible things, said Alice. I dare say you haven t had much practice, said the Queen. Why, sometimes
7.014 Problem Set 7 Solutions
 MIT Department of Biology 7.014 Introductory Biology, Spring 2005 7.014 Problem Set 7 Solutions Question 1 Part A Antigen binding site Antigen binding site Variable region Light chain Light chain Variable
MIT Department of Biology 7.014 Introductory Biology, Spring 2005 7.014 Problem Set 7 Solutions Question 1 Part A Antigen binding site Antigen binding site Variable region Light chain Light chain Variable
numbe r Done by Corrected by Doctor
 numbe r 5 Done by Mustafa Khader Corrected by Mahdi Sharawi Doctor Ashraf Khasawneh Viral Replication Mechanisms: (Protein Synthesis) 1. Monocistronic Method: All human cells practice the monocistronic
numbe r 5 Done by Mustafa Khader Corrected by Mahdi Sharawi Doctor Ashraf Khasawneh Viral Replication Mechanisms: (Protein Synthesis) 1. Monocistronic Method: All human cells practice the monocistronic
Overview of virus life cycle
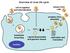 Overview of virus life cycle cell recognition and internalization release from cells progeny virus assembly membrane breaching nucleus capsid disassembly and genome release replication and translation
Overview of virus life cycle cell recognition and internalization release from cells progeny virus assembly membrane breaching nucleus capsid disassembly and genome release replication and translation
HIV 101: Fundamentals of HIV Infection
 HIV 101: Fundamentals of HIV Infection David H. Spach, MD Professor of Medicine University of Washington Seattle, Washington Learning Objectives After attending this presentation, learners will be able
HIV 101: Fundamentals of HIV Infection David H. Spach, MD Professor of Medicine University of Washington Seattle, Washington Learning Objectives After attending this presentation, learners will be able
Viral Genetics. BIT 220 Chapter 16
 Viral Genetics BIT 220 Chapter 16 Details of the Virus Classified According to a. DNA or RNA b. Enveloped or Non-Enveloped c. Single-stranded or double-stranded Viruses contain only a few genes Reverse
Viral Genetics BIT 220 Chapter 16 Details of the Virus Classified According to a. DNA or RNA b. Enveloped or Non-Enveloped c. Single-stranded or double-stranded Viruses contain only a few genes Reverse
III. What are the requirements for taking and passing this course?
 1 Molecular Virology Lecture # 1: Course Introduction I. Instructor and Background Dr. Richard Kuhn rjkuhn@bragg.bio.purdue.edu B-129 Lilly Hall 494-1164 Office Hours - Wednesday 10:30-11:30 II. Objective:
1 Molecular Virology Lecture # 1: Course Introduction I. Instructor and Background Dr. Richard Kuhn rjkuhn@bragg.bio.purdue.edu B-129 Lilly Hall 494-1164 Office Hours - Wednesday 10:30-11:30 II. Objective:
Department of Microbiology, School of Medicine, Box , University of Washington, Seattle, WA 98195, USA
 Virology 366 (2007) 361 376 www.elsevier.com/locate/yviro Substitution of alanine for tyrosine-64 in the fingers subdomain of M-MuLV reverse transcriptase impairs strand displacement synthesis and blocks
Virology 366 (2007) 361 376 www.elsevier.com/locate/yviro Substitution of alanine for tyrosine-64 in the fingers subdomain of M-MuLV reverse transcriptase impairs strand displacement synthesis and blocks
Human Immunodeficiency Virus Type 1 cdnas Produced in the Presence of APOBEC3G Exhibit Defects in Plus-Strand DNA Transfer and Integration
 JOURNAL OF VIROLOGY, July 2007, p. 7099 7110 Vol. 81, No. 13 0022-538X/07/$08.00 0 doi:10.1128/jvi.00272-07 Copyright 2007, American Society for Microbiology. All Rights Reserved. Human Immunodeficiency
JOURNAL OF VIROLOGY, July 2007, p. 7099 7110 Vol. 81, No. 13 0022-538X/07/$08.00 0 doi:10.1128/jvi.00272-07 Copyright 2007, American Society for Microbiology. All Rights Reserved. Human Immunodeficiency
This training module is required for all personnel listed on an IBC protocol that describes work utilizing viral vectors (both replication competent
 This training module is required for all personnel listed on an IBC protocol that describes work utilizing viral vectors (both replication competent and incompetent) regardless of the biosafety level used
This training module is required for all personnel listed on an IBC protocol that describes work utilizing viral vectors (both replication competent and incompetent) regardless of the biosafety level used
Complementation of Human Immunodeficiency Virus Type 1 Replication by Intracellular Selection of Escherichia coli trna 3 Lys Supplied in trans
 JOURNAL OF VIROLOGY, Oct. 2006, p. 9641 9650 Vol. 80, No. 19 0022-538X/06/$08.00 0 doi:10.1128/jvi.00709-06 Copyright 2006, American Society for Microbiology. All Rights Reserved. Complementation of Human
JOURNAL OF VIROLOGY, Oct. 2006, p. 9641 9650 Vol. 80, No. 19 0022-538X/06/$08.00 0 doi:10.1128/jvi.00709-06 Copyright 2006, American Society for Microbiology. All Rights Reserved. Complementation of Human
Helper virus-free transfer of human immunodeficiency virus type 1 vectors
 Journal of General Virology (1995), 76, 691 696. Printed in Great Britabz 691 Helper virus-free transfer of human immunodeficiency virus type 1 vectors Jennifer H. Riehardson,~ Jane F. Kaye, Lisa A. Child
Journal of General Virology (1995), 76, 691 696. Printed in Great Britabz 691 Helper virus-free transfer of human immunodeficiency virus type 1 vectors Jennifer H. Riehardson,~ Jane F. Kaye, Lisa A. Child
Clinical Significance of Human Immunodeficiency Virus Type 1 Replication Fitness
 CLINICAL MICROBIOLOGY REVIEWS, Oct. 2007, p. 550 578 Vol. 20, No. 4 0893-8512/07/$08.00 0 doi:10.1128/cmr.00017-07 Copyright 2007, American Society for Microbiology. All Rights Reserved. Clinical Significance
CLINICAL MICROBIOLOGY REVIEWS, Oct. 2007, p. 550 578 Vol. 20, No. 4 0893-8512/07/$08.00 0 doi:10.1128/cmr.00017-07 Copyright 2007, American Society for Microbiology. All Rights Reserved. Clinical Significance
Cooperation between Reverse Transcriptase and Integrase during Reverse Transcription and Formation of the Preintegrative Complex of Ty1
 EUKARYOTIC CELL, Oct. 2006, p. 1760 1769 Vol. 5, No. 10 1535-9778/06/$08.00 0 doi:10.1128/ec.00159-06 Copyright 2006, American Society for Microbiology. All Rights Reserved. Cooperation between Reverse
EUKARYOTIC CELL, Oct. 2006, p. 1760 1769 Vol. 5, No. 10 1535-9778/06/$08.00 0 doi:10.1128/ec.00159-06 Copyright 2006, American Society for Microbiology. All Rights Reserved. Cooperation between Reverse
Translation. Host Cell Shutoff 1) Initiation of eukaryotic translation involves many initiation factors
 Translation Questions? 1) How does poliovirus shutoff eukaryotic translation? 2) If eukaryotic messages are not translated how can poliovirus get its message translated? Host Cell Shutoff 1) Initiation
Translation Questions? 1) How does poliovirus shutoff eukaryotic translation? 2) If eukaryotic messages are not translated how can poliovirus get its message translated? Host Cell Shutoff 1) Initiation
Virology Introduction. Definitions. Introduction. Structure of virus. Virus transmission. Classification of virus. DNA Virus. RNA Virus. Treatment.
 DEVH Virology Introduction Definitions. Introduction. Structure of virus. Virus transmission. Classification of virus. DNA Virus. RNA Virus. Treatment. Definitions Virology: The science which study the
DEVH Virology Introduction Definitions. Introduction. Structure of virus. Virus transmission. Classification of virus. DNA Virus. RNA Virus. Treatment. Definitions Virology: The science which study the
Micropathology Ltd. University of Warwick Science Park, Venture Centre, Sir William Lyons Road, Coventry CV4 7EZ
 www.micropathology.com info@micropathology.com Micropathology Ltd Tel 24hrs: +44 (0) 24-76 323222 Fax / Ans: +44 (0) 24-76 - 323333 University of Warwick Science Park, Venture Centre, Sir William Lyons
www.micropathology.com info@micropathology.com Micropathology Ltd Tel 24hrs: +44 (0) 24-76 323222 Fax / Ans: +44 (0) 24-76 - 323333 University of Warwick Science Park, Venture Centre, Sir William Lyons
 CDC site UNAIDS Aids Knowledge Base http://www.cdc.gov/hiv/dhap.htm http://hivinsite.ucsf.edu/insite.jsp?page=kb National Institute of Allergy and Infectious Diseases http://www.niaid.nih.gov/default.htm
CDC site UNAIDS Aids Knowledge Base http://www.cdc.gov/hiv/dhap.htm http://hivinsite.ucsf.edu/insite.jsp?page=kb National Institute of Allergy and Infectious Diseases http://www.niaid.nih.gov/default.htm
Induction of APOBEC3G Ubiquitination and Degradation by an HIV-1 Vif-Cul5-SCF Complex
 Induction of APOBEC3G Ubiquitination and Degradation by an HIV-1 Vif-Cul5-SCF Complex Xianghui Yu, 1,2 * Yunkai Yu, 1 * Bindong Liu, 1 * Kun Luo, 1 Wei Kong, 2 Panyong Mao, 1 Xiao-Fang Yu 1,3 1 Department
Induction of APOBEC3G Ubiquitination and Degradation by an HIV-1 Vif-Cul5-SCF Complex Xianghui Yu, 1,2 * Yunkai Yu, 1 * Bindong Liu, 1 * Kun Luo, 1 Wei Kong, 2 Panyong Mao, 1 Xiao-Fang Yu 1,3 1 Department
Molecular Mechanisms by Which Human Immunodeficiency Virus Type 1 Integrase Stimulates the Early Steps of Reverse Transcription
 JOURNAL OF VIROLOGY, Sept. 2007, p. 10037 10046 Vol. 81, No. 18 0022-538X/07/$08.00 0 doi:10.1128/jvi.00519-07 Copyright 2007, American Society for Microbiology. All Rights Reserved. Molecular Mechanisms
JOURNAL OF VIROLOGY, Sept. 2007, p. 10037 10046 Vol. 81, No. 18 0022-538X/07/$08.00 0 doi:10.1128/jvi.00519-07 Copyright 2007, American Society for Microbiology. All Rights Reserved. Molecular Mechanisms
Genetic Dissociation of the Encapsidation and Reverse Transcription Functions in the 5 R Region of Human Immunodeficiency Virus Type 1
 JOURNAL OF VIROLOGY, Jan. 1999, p. 101 109 Vol. 73, No. 1 0022-538X/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. Genetic Dissociation of the Encapsidation and Reverse
JOURNAL OF VIROLOGY, Jan. 1999, p. 101 109 Vol. 73, No. 1 0022-538X/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. Genetic Dissociation of the Encapsidation and Reverse
AIDS - Knowledge and Dogma. Conditions for the Emergence and Decline of Scientific Theories Congress, July 16/ , Vienna, Austria
 AIDS - Knowledge and Dogma Conditions for the Emergence and Decline of Scientific Theories Congress, July 16/17 2010, Vienna, Austria Reliability of PCR to detect genetic sequences from HIV Juan Manuel
AIDS - Knowledge and Dogma Conditions for the Emergence and Decline of Scientific Theories Congress, July 16/17 2010, Vienna, Austria Reliability of PCR to detect genetic sequences from HIV Juan Manuel
are considered to be dead end products of reverse transcription
 JOURNAL OF VIROLOGY, Sept. 2001, p. 7944 7955 Vol. 75, No. 17 0022-538X/01/$04.00 0 DOI: 10.1128/JVI.75.17.7944 7955.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved. Human Immunodeficiency
JOURNAL OF VIROLOGY, Sept. 2001, p. 7944 7955 Vol. 75, No. 17 0022-538X/01/$04.00 0 DOI: 10.1128/JVI.75.17.7944 7955.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved. Human Immunodeficiency
VIRUSES. 1. Describe the structure of a virus by completing the following chart.
 AP BIOLOGY MOLECULAR GENETICS ACTIVITY #3 NAME DATE HOUR VIRUSES 1. Describe the structure of a virus by completing the following chart. Viral Part Description of Part 2. Some viruses have an envelope
AP BIOLOGY MOLECULAR GENETICS ACTIVITY #3 NAME DATE HOUR VIRUSES 1. Describe the structure of a virus by completing the following chart. Viral Part Description of Part 2. Some viruses have an envelope
Received 18 October 1999/Accepted 11 February 2000
 JOURNAL OF VIROLOGY, May 2000, p. 4795 4806 Vol. 74, No. 10 0022-538X/00/$04.00 0 Copyright 2000, American Society for Microbiology. All Rights Reserved. Identification of Critical Amino Acid Residues
JOURNAL OF VIROLOGY, May 2000, p. 4795 4806 Vol. 74, No. 10 0022-538X/00/$04.00 0 Copyright 2000, American Society for Microbiology. All Rights Reserved. Identification of Critical Amino Acid Residues
VIRAL TITER COUNTS. The best methods of measuring infectious lentiviral titer
 VIRAL TITER COUNTS The best methods of measuring infectious lentiviral titer FLUORESCENCE CYCLES qpcr of Viral RNA SUMMARY Viral vectors are now routinely used for gene transduction in a wide variety of
VIRAL TITER COUNTS The best methods of measuring infectious lentiviral titer FLUORESCENCE CYCLES qpcr of Viral RNA SUMMARY Viral vectors are now routinely used for gene transduction in a wide variety of
