Research Article Prenatal Diagnosis of Central Nervous System Anomalies by High-Resolution Chromosomal Microarray Analysis
|
|
|
- Gilbert King
- 5 years ago
- Views:
Transcription
1 Hindawi Publishing Corporation BioMed Research International Volume 2015, Article ID , 9 pages Research Article Prenatal Diagnosis of Central Nervous System Anomalies by High-Resolution Chromosomal Microarray Analysis Lijuan Sun, 1 Qingqing Wu, 1 Shi-Wen Jiang, 2 Yani Yan, 1 Xin Wang, 3 Juan Zhang, 1 Yan Liu, 3 Ling Yao, 1 Yuqing Ma, 1 and Li Wang 1 1 Department of Ultrasound, Beijing Obstetrics and Gynecology Hospital, Capital Medical University, Beijing , China 2 Department of Biomedical Science, Mercer University School of Medicine, Department of Obstetrics and Gynecology, Memorial Health Hospital, 4700 Waters Avenue, Savannah, GA , USA 3 Department of Obstetrics, Beijing Obstetrics and Gynecology Hospital, Capital Medical University, Beijing , China Correspondence should be addressed to Qingqing Wu; wuqq2007@163.com and Shi-Wen Jiang; jiang s@mercer.edu Received 15 January 2015; Accepted 7 April 2015 Academic Editor: Marco Fichera Copyright 2015 Lijuan Sun et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. The aims of this study were to evaluate the contribution of chromosomal microarray analysis (CMA) in the prenatal diagnosis of fetuses with central nervous system (CNS) anomalies but normal chromosomal karyotype. A total of 46 fetuses with CNS anomalies with or without other ultrasound anomalies but normal karyotypes were evaluated by array-based comparative genomic hybridisation (acgh) or single-nucleotide polymorphism (SNP) array. The result showed that CNVs were detected in 17 (37.0%) fetuses. Of these, CNVs identified in 5 (5/46, 10.9%) fetuses were considered to be likely pathogenic, and CNVs detected in 3 (3/46, 6.5%) fetuses were defined as being of uncertain clinical significance. Fetuses with CNS malformations plus other ultrasound anomalies had a higher rate of pathogenic CNVs than those with isolated CNS anomalies (13.6% versus 8.3%), but there was no significant difference (Fisher s exact test, P > 0.05). Pathogenic CNVs were detected mostfrequently in fetuses with Dandy-Walker syndrome (2/6, 33.3%) when compared with other types of neural malformations, and holoprosencephaly (2/7, 28.6%) ranked the second. CMA is valuable in prenatal genetic diagnosis of fetuses with CNS anomalies. It should be considered as part of prenatal diagnosis in fetuses with CNS malformations and normal karyotypes. 1. Introduction The prevalence of CNS abnormalities is % in live births and reaches as high as 3 6% in stillbirths [1]. CNS anomalies are a group of serious birth defects associated with high rates of infant death or disability. In addition to theirthreattolife,cnsmalformationscauseenormousdirect and indirect health costs [2]. While the etiology of fetal central nervous system anomalies is highly heterogeneous, genetic conditions are recognized as a major cause [3, 4]. The significance of genetic mutations is also underscored by the fact that many environmental factors lead to CNS malformations through their mutagenic effects. Studies have shown that CNS malformations detected by ultrasonography were strongly associated with chromosomal abnormalities, especially trisomy 13 and 18 [5, 6]. However, there remains a dilemma in prenatal diagnosis of fetuses who have CNS anomalies, either with or without other organ abnormalities, but have normal karyotypes. Conventional chromosome karyotype analysis such as G- Banding has been the standard method for the detection of a wide range of chromosomal abnormalities in the past decades. However, this technique is limited to the detection of chromosomal alterations larger than 5 Mb, and submicroscopic duplications and deletions, which are often associated with mental retardation (MR) and malformations, are not detectable by conventional karyotyping [7]. Chromosomal microarray analysis (CMA) allows the detection of microdeletions and microduplications that are not routinely seen on karyotyping [8]. It is possible to evaluate the entire genome for DNA CNVs, as small as kb, which is equal to a 100-fold magnification in resolution compared with karyotyping [9, 10]. This high-resolution analysis of DNA CNVs, initially applied to cancer studies, has
2 2 BioMed Research International subsequently been extended to postnatal diagnosis of various congenital anomalies and MR. Recent studies have focused specifically on the use of microarray analysis in prenatal diagnosis of fetuses with abnormal ultrasound findings [8, 11, 12], especially on the concerns regarding the relationship between CNVs and congenital heart diseases (CHD) [13, 14]. Application of CMA to identify submicroscopic chromosomal aberrations in fetuses with CNS anomalies has been poorly described in the literature so far. Theaimsofthisstudyweretoevaluatetheutilityof CMA for prenatal diagnosis of fetuses with CNS anomalies detected by ultrasound but normal karyotypes and to explore the CNVs in fetuses carrying different types of CNS malformations. 2. Materials and Methods 2.1. Subject Selection and Ultrasound Findings. From December 2011 to June 2014, 31,802 pregnant women were referred to the Department of Obstetrics and Fetal-Maternal Medicine, Beijing Obstetrics and Gynecology Hospital, Capital Medical University, China, for routine anomaly scan. 46 fetuses were diagnosed for CNS abnormalities, with or without other associated anomalies by transabdominal ultrasonography, showing normal karyotypes in conventional G-band karyotype analysis. Of these 46 primary study subjects, in 24 the CNS anomalies were isolated and 22 fetuses with CNS malformations were associated with other abnormalities. This study was approved by the local Ethics Committee of the Capital Medical University. Written informed consent to participate in the study was obtained from each patient CMA Methods. Umbilical cord blood samples were collected from the 46 pregnant women between 21 and 27 gestational weeks by cordocentesis. Of the 46 samples, 30 cases (65.2%) were examined with array-based comparative genomic hybridisation (acgh), and then additional 16 cases (34.8%) were tested with single-nucleotide polymorphism (SNP) array in consideration of the advantages for SNP detecting copy number aberrations [15]after being approved by Food and Drug Administration (FDA). Genomic DNA was extracted from 2 ml of umbilical cord blood with a commercially available Blood Genomic DNA Extraction kit (BioChain Institute Inc., Newark, CA or Yuanpinghao Biotech Co., Ltd., resp.) according to the manufacturer s instructions. acgh was performed using 8 60K oligonucleotide-based microarray (Agilent), and SNP array was detected using a 750 K microarray (Affymetrix CytoScan 750KArray).Afterhybridization,alaserscannerwasusedfor scanningthearrays,andthedatawasanalyzedwiththeuse of special software package (Workbench and Chromosome Analysis Suite) The Interpretation for CMA Results. To interpret the results, we compared all detected copy number gains or losses with known CNVs listed in publically available databases (DECIPHER Database ( Online Mendelian Inheritance in Man (OMIM, PubMed ( and the Database of Genomic Variants (DGV, For all CNVs, CMA results were interpreted independently of any previous cytogenetic findings Statistical Analysis. Statistical analysis of data was performed using SPSS version Bivariate analysis was carried out using Fisher s exact test (two-tailed). P < 0.05 was considered statistically significant. 3. Results 3.1. CNVs Detection Rates. Of the 46 study subjects, CNVs were detected in 17 (37.0%) cases, whereas no deletion or duplication was found in the remaining 29 cases (63.0%). Pathogenic CNVs were identified in 5 (10.9%) fetuses including 2 cases with isolated CNS malformations and 3 cases associated with other organ abnormalities. In addition, CNVs of uncertain clinical significance were detected in 3 (6.5%) fetuses, and CNVs identified in 9 (19.6%) fetuses were considered to be likely benign and of no clinical significance. Fetuses with CNS malformations plus other ultrasound anomalies had a seemingly higher detection rate than those with isolated CNS anomalies (13.6% versus 8.3%), but the difference did not reach a statistically significant level (Fisher s exact test, P > 0.05)(Table 1) CMA Results for 5 Fetuses with Pathogenic CNVs and 3 Cases with CNVs of Uncertain Clinical Significance. Information for 5 fetuses with pathogenic CNVs and 3 cases with CNVs of uncertain clinical significance was shown in Figures 1 and 2,andTable 2. Of the 24 fetuses with isolated CNS anomalies, pathogenic CNVs were identified in 2 cases (2/24, 8.3%) (Table 2, Cases1and2)asfollows. (1)Case1wasafetuswithholoprosencephalyandsingle nostril on prenatal ultrasound. Sonographic abnormal findings were shown in Figure 3 and confirmed by the autopsy pathology. The microarray analysis result showed a 3.44-Mb deletion within the chromosome 7q36.3, encompassing the sonic hedgehog (SHH) gene (Table 2, Figure 1(a)). Deletions SHHgeneareknowntoberelatedtoholoprosencephaly[16] (OMIM ID: ). (2) Case 2 harbored a 3.15-Mb duplication in chromosome 22q11.21 (Table 2, Figure2(a)) which included 43 OMIMgenessuchasTBX1inafetuswithhydrocephaly.The CMA result has been associated with 22q11.2 microduplication syndrome (OMIM ID: #608363). Dupont et al. reported that 22q11.2 microduplication syndrome was characterized by a highly variable clinical phenotypes, ranging from apparentlynormalorslightlydysmorphicfeaturestoseveremalformations with profound mental retardation [17]. Neurological features of the syndrome included intellectual or learning disability, motor delay, and other neurodevelopmental disorders [18]. In this case, the parents chose termination of pregnancy (TOP) because of fetal hydrocephaly that was confirmed by postnatal imaging.
3 BioMed Research International 3 Table 1: The detection rates of pathogenic CNVs in fetuses with CNS anomalies associated with different ultrasound findings. Ultrasound findings Number of fetuses Number of fetuses with pathogenic CNVs Detection rate (%) Isolated CNS anomaly % Associated with other structural % malformations Cardiovascular system 10 # 1 Urinary system 2 0 Musculoskeletal system 6 1 Digestive system 1 0 Tumors 2 0 Facial anomaly/lip and palate cleft 1 1 Total % # One (Case 7) of 10 fetuses with cardiovascular abnormalities was also associated with facial anomaly, but only in the row of cardiovascular system. The fetus (Case 4) was associated with skeletal dysplasia and lip and palate cleft, but only in the row of musculoskeletal system. Information for clinically significant CNVs detected in 3/22 (13.6%) fetuses with CNS anomalies as well as other abnormalities (Table 2, Cases 3 5) was as follows. (1) Case 3 was a fetus with Dandy-Walker malformation in association with ventricular septal defect and persistent left superior vena cava. The microarray test result revealed a Mb deletion in chromosome 2q13 q14.1 that included four OMIM genes (Table 2, Figure 1(b)): PAX8, IL1B,MERTK,and IL1RN. As reported by Kasai and Narahara, these genes were associated with neurodevelopmental impairment, congenital heart defects, and facial and finger malformations [19]. The parents chose TOP because of the sonographic abnormalities that were confirmed by autopsy. (2) Case 4 carried a 0.17-Mb deletion in chromosome Xq13.3 (Table 2, Figure 2(b)) whichinvolvedtheabc7gene (OMIM ID: ) inamalefetuswithdandy-walker syndrome (cerebellar malformation), absence of septum pellucidum, and arachnoid cyst associated with lip and palate cleft and skeletal dysplasia. Bekri et al. [20]previously identified a missense mutation in the ABC7 gene to be the cause of X-linked sideroblastic anemia with cerebellar ataxia. Although it is not experimentally confirmed, we considered that the microdeletion might be responsible for fetal cerebellar malformation. The parents opted for TOP because of abnormal sonographic findings which were confirmed by autopsy pathology. (3) Case 5 was a fetus with holoprosencephaly (HPE) associated with lip and palate cleft. The microarray test discovereda0.34-mbdeletionwithin2p21(table 2, Figure 2(c)) which harbors SIX3 gene (OMIM ID: ). It has been reported that molecular evaluation of foetuses with holoprosencephaly showed high incidence of microdeletions in four HPE genes, one of which was SIX3 gene [21]. Lacbawan et al. reported that SIX3 mutations could result in relatively severe holoprosencephaly [22]. The parents had chosen TOP because of the severe malformations detected by prenatal ultrasound which were confirmed by autopsy. Our study covered 3 fetuses with the CNVs of uncertain clinical significance (Table 2, Cases 6 8) as the follows. (1) Case 6 was originally referred for prenatal cytogenetic diagnosis because of exencephaly detected by ultrasonography. Following negative findings in fetal karyotype analysis, CMA result revealed a deletion of 4.03 Mb in chromosome 19p12p13.11 (Table 2, Figure 1(c)). The region contained a lot of segmental duplications and many homologous genes, such as ZNF family that probably mediated the recombination giving rise to the deletion. However, extensive literature search failed to identify that any gene in the relevant region might cause known syndromes. Both parents showed negative CMA results, indicating that the deletion was de novo. Thus, the clinical significance of this CNV was uncertain. The family opted for TOP because of the severe ultrasonography results. The patient underwent Rivanol amniocentesis induction of labor with an informed consent, and the autopsy pathology confirmed the ultrasound diagnosis. (2) Case 7 was referred because of holoprosencephaly in association with facial anomaly and ventricular septal defect as ultrasonographic findings. There was a 2.79-Mb deletion in chromosome 4q35.2 (Table 2, Figure 1(d)), with proximal breakpointlocated300kbupstreamtheomimgenefat1. This gene encodes a tumor suppressor essential for controlling cell proliferation during Drosophila development [23]. The gene product is a member of the cadherin superfamily and is expressed at high levels in a number of fetal epithelia. Its product probably functions as an adhesion molecule and/or signaling receptor and is likely to be important in developmental processes and cell communication. We think that it is possible for the microdeletion to influence FAT1 gene expression by position effect because of the near distance. The CMA results from both of parents were negative, indicating a de novo mutation. So the clinical significance of the CNVs in this case remained unclear. The parents chose TOP, and the abnormal ultrasound findings were confirmed by autopsy pathology. (3) Case 8 was a fetus with hydrocephaly associated with sacrococcygeal vertebral anomaly, intrauterine growth retardation (IUGR), and thickened nuchal fold (NF) on prenatal ultrasound. The array test result showed a Mb deletion in chromosome 21q21.1 (Table 2, Figure 2(d)) which contained most of the NCAM2 gene. It has been reported that NCAM2 was a candidate for involvement in certain Down syndrome phenotypes [24]. However, the role of NCAM2 deletion in the pathophysiology of Down syndrome is unknown. CMA on both parents was
4 4 BioMed Research International Table 2: Pathogenic CNVs and variants of unknown significance detected by chromosomal microarray analysis (CMA) in fetuses with CNS anomalies. Case number GA CNS anomalies Associated anomalies CMA result Size (Mb) CNV type Holoprosencephaly, single nostril No Hydrocephaly No Dandy-Walker syndrome Dandy-Walker syndrome VSD PLSVC Skeletal dysplasia lip and palate cleft absence of septum pellucidum arachnoid cyst Holoprosencephaly Lip and palate cleft Exencephaly No Holoprosencephaly Hydrocephaly VSD facial anomaly Thickened NF IUGR sacrococcygeal vertebral anomaly arr7q36. (155,473, ,909,738) 1 arr22q11.21 (18,648,855 21,800,471) 3 arr2q13q14.1 (111,596, ,844,660) 1 arrxq13.3 (74,171,888 74,343,340) 1 arr2p21 (44,749,075 45,098,283) 1 arr19p12p13.11 (19,838,485 23,868,512) 1 arr4q35.2 (187,979, ,767,114) 1 arr 21q21.1 (22,508,434 23,663,338) 1 GA, gestational weeks; VSD, ventricular septal defect; PLSVC, persistent left superior vena cava; NF, nuchal fold; IUGR, intrauterine growth retardation. Human genome build was hg19. OMIM or corresponding disorder 3.44 Loss SHH gene ( ) 3.15 Gain 3.25 Loss 22q11.2 duplication syndrome ( # ) PAX8 ( ) IL1B ( ) MERTK ( ) IL1RN ( ) 0.17 Loss ABC7 ( ) 0.34 Loss SIX 3 gene ( ) 4.03 Loss 2.79 Loss 1.15 Loss
5 BioMed Research International 5 7-log ratio p22.2 p21.3 p21.1 p15.2 p14.3 p14.1 p12.3 p log ratio p25.2 p24.3 p24.1 p23.2 p22.3 p22.1 p16.3 p16.1 p14 p13.2 p log ratio p13.2 p13.12 q11.22 q21.11 q21.13 q21.3 q22.2 q31.1 q31.31 q31.33 q32.2 q33 q35 q36.2 q12.1 q12.3 q14.1 q14.3 q21.2 q22.1 q23.2 q22.3 q24.1 q24.3 q31.2 q32.1 q32.3 q33.2 q34 q36.1 q36.3 q37.2 q13.12 q13.2 q13.32 q13.41 q13.43 (a) (b) (c) 4-log ratio p16.2 p15.33 p15.31 p15.1 p13 q13.1 q13.3 q21.21 q21.23 q22.1 q22.3 q24 q26 q28.1 q28.3 q31.21 q31.23 q32.1 q32.3 q34.1 q34.3 q35.2 (d) Figure 1: Microarray testing results. The CNVs of Cases 1, 3, 6, and 7 were detected by acgh. (a) (d) showed acgh results of Cases 1, 3, 6, and 7, respectively. (a) A 3.44-Mb deletion within chromosome 7q in Case 1. (b) A 3.25-Mb deletion in chromosome 2q in Case 3 harbored four OMIM genes. (c) A 4.03-Mb deletion of chromosome 19p in Case 6. (d) A 2.79-Mb deletion in chromosome 4q35.2 in Case 7. The respective chromosomes are shown and labeled. Signal intensity is plotted on a log 2 scale,suchthatanormalcopynumbergivesavalueof0. Chromosomal deletions are denoted by leftward deviation of the central line (marked by red boxes). negative, pointing to a possibility for de novo mutation. The clinical significance of this CNV was considered to be uncertain. The pregnant woman underwent TOP, and the autopsy pathology confirmed the ultrasonographic findings The Types of Fetal CNS Anomalies and Pathogenic CNVs Incidence. The relationship between the different types of CNS anomalies and the incidence of fetuses with pathogenic CNVs is shown in Table 3. We found that clinically significant CNVs were detected most frequently in fetuses with Dandy- Walker syndrome (2/6, 33.3%), and holoprosencephaly (2/7, 28.6%) ranked the second. 4. Discussion In the past few years, CMA has been extensively used to investigate chromosomal aberrations in the postnatal population with unexplained neurodevelopmental disorders including developmental delay/intellectual disability, autism spectrum disorders, and multiple congenital anomalies [25 27]. A recent meta-analysis [26] of CMA on 13,926 postnatal subjects in whom conventional cytogenetic tests have proven negative reported an overall diagnostic rate of 10% for pathogenic genomic imbalances in such populations. Recently results on application of CMA for prenatal diagnosis have also been published. D Amours et al. [12]
6 6 BioMed Research International (a) (b) (c) (d) Figure 2: Microarray testing results. The CNVs of Cases 2, 4, 5, and 8 were detected by SNP array. (a) (d) showed the SNP results of Cases 2, 4, 5, and 8, respectively. (a) A 3.15-Mb duplication in chromosome 22q11.21 included 43 OMIM genes in Case 2. (b) A 0.17-Mb deletion in chromosome Xq13.3 in Case 4. (c) A 0.34-Mb deletion within chromosome 2p21 in Case 5. (d) A 1.15-Mb deletion of chromosome 21q21.1 in Case 8. The chromosome numbers and cytobands are shown and labeled on the right side. The view on the left side shows the detected segments, regions, and reference annotations in detail. Chromosomal duplication segments are denoted by upward triangle (blue) whereas deletion segments are denoted by downward triangle (red). reported a 8.2% detection rate of pathogenic CNVs in fetuses with major malformations and a normal karyotype by acgh. A meta-analysis conducted by Hillman et al. [28] indicated that CMA had a 5.2% additive value compared with conventional karyotyping. Collectively, these studies suggested that CMA was able to identify additional, clinically significant cytogenetic information in fetuses with various kinds of structural anomalies and a normal karyotype. The efforts to diagnose fetuses with CNS malformations with or without other structural abnormalities in our study represent a further attempt for the usage of this powerful technique in prenatal genetic tests. In our study we focused on fetuses with CNS malformations and an apparently normal karyotype, and the results showed that 10.9% of fetuses had submicroscopic chromosomal abnormalities which were likely to be pathological. This suggests that the microarray technique is able to provide additional information in fetuses with CNS malformations and a normal karyotype. Such information could lead to better prenatal consultation and the assessment of the recurrent risk. Compared with the studies of D Amours et al. [12] and Hillman et al. [28], this project achieved a higher detection rate (10.9%) of pathogenic CNVs. The elevated detection rate could be attributed to the exclusion of congenital noncns anomalies in this study. It should be pointed out that the sample size of this study was relatively small, which could limit the extrapolation of our observations. Further investigations in larger cohorts need to be conducted to validate the potentially benefits of CMA. In this study we found that the detection rates of pathogenic CNVs in various types of CNS anomalies were different. It was noteworthy that fetuses with Dandywalker syndrome (2/6, 33%) emerged to be most frequently associated with submicroscopic chromosomal abnormalities among various congenital neural abnormalities when karyotyping showed normal results. There were a few reports [29, 30] regarding CNVs of Dandy-Walker malformation, such as deletion in chromosomes 13q and 7p, which were different from deletion in chromosomes 2q and Xq in this study. Our results also indicated holoprosencephaly (2/7, 28.6%) as the second common malformation associated with pathogenic CNVs among all types of CNS abnormalities. This observation provides some insights into the pathogenesis of holoprosencephaly in fetuses with a normal karyotype, suggesting that CNVs could be a significant cause of this specific type of CNS abnormalities. Shaffer et al. [31] performed
7 BioMed Research International 7 (a) (b) (c) Figure 3: Sonographic findings in Case 1 at weeks. (a) In two-dimensional coronal plane, single nostril was visualized. (b) Holoprosencephaly was demonstrated in tomographic ultrasound imaging (TUI). ((c) and (d)) Single nostril was further confirmed in threedimensional imaging. (d) Table 3: The types of fetal CNS anomalies and pathogenic CNVs incidence. CNS anomalies classification Number of fetuses Number of fetuses with pathogenic CNVs Anencephaly 1 0 Exencephaly 1 0 Dandy-Walker syndrome 6 2 (33.3%) Holoprosencephaly 7 2 (28.6%) Spinal bifida 9 0 Intracranial tumor (ICT) 1 0 Hydrocephaly 8 1 (12.5%) Schizencephaly 1 0 Agenesis of the corpus callosum (ACC) 2 0 Choroid plexus cyst 2 0 Arachnoid cyst 2 0 Cerebellar hypoplasia 1 0 Subependymal cyst 1 0 Encephalocele/meningoceles 1 0 Other CNS malformation 3 0 Total 46 5 (10.9%) a retrospective analysis of pathogenic CNVs detection rate by CMA for 2858 pregnancies with different organ system abnormalities and normal karyotypes and found that both of posterior fossa defects (including Dandy-Walker syndrome and cerebellar hypoplasia) and holoprosencephaly had the highest detection rates of clinically significant CNVs (14.6% and 10.6%) among various types of CNS anomalies, which was similar to our results. Clinically significant CNVs were detected in 3 fetuses with nervous system anomalies plus other congenital structural malformations, one case with cardiovascular defect, one with lip and palate cleft, and another one with skeletal dysplasia and lip and palate cleft. Fetuses with CNS malformations plus other ultrasound anomalies had a higher detection rate than those with isolated CNS anomalies (13.6% versus 8.3%), but there were no significant differences in the incidence of pathogenic CNVs between them (Fisher s exact test, P > 0.05). Our results are comparable with the study reported by Shaffer et al. [31], who found the detection rate of clinical significant CNVs was 6.5% in fetuses with a single CNS anomaly while 11% in cases with CNS malformations and other abnormalities. Although chromosomal microarray techniques could offer several advantages including high-resolution, whole genome analysis, and a short turnaround time (about 48 h after DNA extraction) in comparison with conventional chromosomal karyotyping, it cannot identify balanced translocations or low level of mosaicism. Therefore, prenatal consultations should not rely on the results of CMA alone, and conventional chromosomal karyotyping and prenatal ultrasound diagnosis should also be required. The present
8 8 BioMed Research International results substantiated that CMA may be especially valuable in routine prenatal diagnosis when fetuses with abnormal ultrasound findings but normal karyotypes. 5. Conclusion Our results have shown that CMA is a valuable diagnostic tool in prenatal genetic diagnosis of CNS anomalies. Our data indicated that assessment of submicroscopic chromosomal aberrations by CMA should be undertaken in fetuses with CNS anomalies and a normal karyotype. This finding not only provides information for clinical consultation but may also allow more accurate genetic diagnosis and a better understanding of the etiology and mechanisms involved in the congenital defects. Conflict of Interests The authors declare that there is no conflict of interests regarding the publication of this paper. Acknowledgments The authors would like to express their great appreciation to Be Creative Lab (Beijing) for technical assistance. This research was supported by Beijing Municipal Administration of Hospitals Clinical medicine Development of special funding support, Code: XMLX201310, Beijing Natural Science Foundation (Grant no ), and Beijing Municipal Science & Technology Commission (no. Z ). References [1] D. Onkar, P. Onkar, and K. Mitra, Evaluation of fetal central nervous system anomalies by ultrasound and its anatomical corelation, Journal of Clinical and Diagnostic Research,vol.8,no. 6, pp. AC05 AC07, [2] Ministry of Health of the People s Republic of China and Disabled Persons Federation of China, Promote birth population quality and reduce birth deficiency and deformity action plan of China ( ), MaternalChildHealthCareofChina,vol. 17, pp , [3] J.Huang,I.Y.M.Wah,R.K.Pooh,andK.W.Choy, Molecular genetics in fetal neurology, Seminars in Fetal and Neonatal Medicine,vol.17,no.6,pp ,2012. [4] F.Petracchi,L.Crespo,C.Michia,L.Igarzabal,andE.Gadow, Holoprosencephaly at prenatal diagnosis: analysis of 28 cases regarding etiopathogenic diagnoses, Prenatal Diagnosis, vol. 31, no. 9, pp , [5] K.R.Goetzinger,D.M.Stamilio,J.M.Dicke,G.A.Macones, and A. O. Odibo, Evaluating the incidence and likelihood ratios for chromosomal abnormalities in fetuses with common central nervous system malformations, The American Journal of Obstetrics and Gynecology, vol.199,no.3,pp.285.e1 285.e6, [6] S. Pitukkijronnakorn, P. Promsonthi, P. Panburana, R. Rangsiprakarn, and A. Chittacharoen, Prenatal ultrasonographic findings in trisomy 13, Journal of the Medical Association of Thailand,vol.91,no.11,pp ,2008. [7] P. Evangelidou, C. Sismani, M. Ioannides et al., Clinical application of whole-genome array CGH during prenatal diagnosis: studyof25selectedpregnancieswithabnormalultrasound findings or apparently balanced structural aberrations, Molecular Cytogenetics,vol.3,article24,2010. [8] R. J. Wapner, C. L. Martin, B. Levy et al., Chromosomal microarray versus karyotyping for prenatal diagnosis, The New England Journal of Medicine, vol.367,no.23,pp , [9] T.-H. Bui, A. Vetro, O. Zuffardi, and L. G. Shaffer, Current controversies in prenatal diagnosis 3: is conventional chromosome analysis necessary in the post-array CGH era? Prenatal Diagnosis,vol.31,no.3,pp ,2011. [10] J. M. Friedman, High-resolution array genomic hybridization in prenatal diagnosis, Prenatal Diagnosis,vol.29,no.1,pp.20 28, [11] E. M. Vestergaard, R. Christensen, O. B. Petersen, and I. Vogel, Prenatal diagnosis: array comparative genomic hybridization in fetuses with abnormal sonographic findings, Acta Obstetricia et Gynecologica Scandinavica,vol.92,no.7,pp ,2013. [12]G.D Amours,Z.Kibar,G.Mathonnetetal., Whole-genome array CGH identifies pathogenic copy number variations in fetuses with major malformations and a normal karyotype, Clinical Genetics,vol.81,no.2,pp ,2012. [13] C.Liao,R.Li,F.Fuetal., Prenataldiagnosisofcongenitalheart defect by genome-wide high-resolution SNP array, Prenatal Diagnosis,vol.34,no.9,pp ,2014. [14] Y. Yan, Q. Wu, and L. Zhang, Detection of submicroscopic chromosomal aberrations by array-based comparative genomic hybridization in fetuses with congenital heart disease, Ultrasound in Obstetrics & Gynecology, vol.43,no.4,pp , [15] X. Xu, E. B. Johnson, L. Leverton et al., The advantage of using SNP array in clinical testing for hematological malignanciesa comparative study of three genetic testing methods, Cancer Genetics,vol.206,no.9-10,pp ,2013. [16] K. Taniguchi, A. E. Anderson, A. E. Sutherland, and D. Wotton, Loss of tgif function causes holoprosencephaly by disrupting the SHH signaling pathway, PLoS Genetics,vol.8,no.2,Article ID e , [17]C.Dupont,F.R.Grati,K.W.Choyetal., Prenataldiagnosis of 24 cases of microduplication 22q11.2: an investigation of phenotype-genotype correlations, Prenatal Diagnosis, vol. 35, no. 1, pp , [18] G. Valvo, F. Novara, P. Brovedani et al., 22q11.2 Microduplication syndrome and epilepsy with continuous spikes and waves during sleep (CSWS). A case report and review of the literature, Epilepsy and Behavior, vol. 25, no. 4, pp , [19] R. Kasai and K. Narahara, Proximal 2q monosomy in a case with mild phenotype, Teratology, vol. 38, pp , [20] S. Bekri, G. Kispal, H. Lange et al., Human ABC7 transporter: gene structure and mutation causing X-linked sideroblastic anemia with ataxia with disruption of cytosolic iron-sulfur protein maturation, Blood, vol. 96, no. 9, pp , [21] C.Bendavid,C.Dubourg,I.Gicqueletal., Molecularevaluation of foetuses with holoprosencephaly shows high incidence of microdeletions in the HPE genes, Human Genetics,vol.119, no. 1-2, pp. 1 8, [22] F. Lacbawan, B. D. Solomon, E. Roessler et al., Clinical spectrum of SIX3-associated mutations in holoprosencephaly: correlation between genotype, phenotype and function, Journal of Medical Genetics, vol. 46, no. 6, pp , 2009.
9 BioMed Research International 9 [23] J. Dunne, A. M. Hanby, R. Poulsom et al., Molecular cloning and tissue expression of FAT, the human homologue of the Drosophila fat gene that is located on chromosome 4q34-q35 and encodes a putative adhesion molecule, Genomics, vol.30, no. 2, pp , [24] A.Paoloni-Giacobino,H.Chen,andS.E.Antonarakis, Cloning of a novel human neural cell adhesion molecule gene (NCAM2) that maps to chromosome region 21q21 and is potentially involved in Down syndrome, Genomics, vol. 43, no. 1,pp.43 51,1997. [25] D.T.Miller,M.P.Adam,S.Aradhyaetal., Consensusstatement: chromosomal microarray is a first-tier clinical diagnostic test for individuals with developmental disabilities or congenital anomalies, TheAmericanJournalofHumanGenetics,vol.86, no. 5, pp , [26] G. S. Sagoo, A. S. Butterworth, S. Sanderson, C. Shaw-Smith, J. P. T. Higgins, and H. Burton, Array CGH in patients with learning disability (mental retardation) and congenital anomalies: updated systematic review and meta-analysis of 19 studies and 13,926 subjects, Genetics in Medicine, vol. 11, no. 3, pp , [27] R. Hochstenbach, E. van Binsbergen, J. Engelen et al., Array analysis and karyotyping: Workflow consequences based on a retrospective study of 36,325 patients with idiopathic developmental delay in the Netherlands, EuropeanJournalofMedical Genetics,vol.52,no.4,pp ,2009. [28] S. C. Hillman, S. Pretlove, A. Coomarasamy et al., Additional information from array comparative genomic hybridization technology over conventional karyotyping in prenatal diagnosis: a systematic review and meta-analysis, Ultrasound in Obstetrics and Gynecology,vol.37,no.1,pp.6 14,2011. [29]M.Mimaki,T.Shiihara,M.Watanabeetal., Holoprosencephaly with cerebellar vermis hypoplasia in 13q deletion syndrome: critical region for cerebellar dysgenesis within 13q32.2q34, Brain & Development,2014. [30] C. Liao, F. Fu, R. Li et al., Dandy-walker syndrome and microdeletions on chromosome 7, Zhonghua Yi Xue Yi Chuan Xue Za Zhi,vol.29,no.1,pp.48 51,2012. [31]L.G.Shaffer,J.A.Rosenfeld,M.P.Dabelletal., Detection rates of clinically significant genomic alterations by microarray analysis for specific anomalies detected by ultrasound, Prenatal Diagnosis,vol.32,no.10,pp ,2012.
Applications of Chromosomal Microarray Analysis (CMA) in pre- and postnatal Diagnostic: advantages, limitations and concerns
 Applications of Chromosomal Microarray Analysis (CMA) in pre- and postnatal Diagnostic: advantages, limitations and concerns جواد کریمزاد حق PhD of Medical Genetics آزمايشگاه پاتوبيولوژي و ژنتيك پارسه
Applications of Chromosomal Microarray Analysis (CMA) in pre- and postnatal Diagnostic: advantages, limitations and concerns جواد کریمزاد حق PhD of Medical Genetics آزمايشگاه پاتوبيولوژي و ژنتيك پارسه
Challenges of CGH array testing in children with developmental delay. Dr Sally Davies 17 th September 2014
 Challenges of CGH array testing in children with developmental delay Dr Sally Davies 17 th September 2014 CGH array What is CGH array? Understanding the test Benefits Results to expect Consent issues Ethical
Challenges of CGH array testing in children with developmental delay Dr Sally Davies 17 th September 2014 CGH array What is CGH array? Understanding the test Benefits Results to expect Consent issues Ethical
%(6/31) [DOI] /j.issn
![%(6/31) [DOI] /j.issn %(6/31) [DOI] /j.issn](/thumbs/92/110077812.jpg) 750 2016 9 1 41 9 [ ] 2012 2 2015 5 31 20~37 21~27 31 G 4 1 26 23 3 11.54%(3/26) 4 / 21 6 19.35%(6/31) [ ] [ ] R714.53 [ ] A [ ] 0577-7402(2016)09-0750-08 [DOI] 10.11855/j.issn.0577-7402.2016.09.10 Application
750 2016 9 1 41 9 [ ] 2012 2 2015 5 31 20~37 21~27 31 G 4 1 26 23 3 11.54%(3/26) 4 / 21 6 19.35%(6/31) [ ] [ ] R714.53 [ ] A [ ] 0577-7402(2016)09-0750-08 [DOI] 10.11855/j.issn.0577-7402.2016.09.10 Application
CURRENT GENETIC TESTING TOOLS IN NEONATAL MEDICINE. Dr. Bahar Naghavi
 2 CURRENT GENETIC TESTING TOOLS IN NEONATAL MEDICINE Dr. Bahar Naghavi Assistant professor of Basic Science Department, Shahid Beheshti University of Medical Sciences, Tehran,Iran 3 Introduction Over 4000
2 CURRENT GENETIC TESTING TOOLS IN NEONATAL MEDICINE Dr. Bahar Naghavi Assistant professor of Basic Science Department, Shahid Beheshti University of Medical Sciences, Tehran,Iran 3 Introduction Over 4000
Prenatal Diagnosis: Are There Microarrays in Your Future?
 Financial Disclosure UCSF Antepartum Intrapartum Management Course June 8 I have no financial relationship with any aspect of private industry Prenatal Diagnosis: Are There Microarrays in Your Future?
Financial Disclosure UCSF Antepartum Intrapartum Management Course June 8 I have no financial relationship with any aspect of private industry Prenatal Diagnosis: Are There Microarrays in Your Future?
SNP Array NOTE: THIS IS A SAMPLE REPORT AND MAY NOT REFLECT ACTUAL PATIENT DATA. FORMAT AND/OR CONTENT MAY BE UPDATED PERIODICALLY.
 SAMPLE REPORT SNP Array NOTE: THIS IS A SAMPLE REPORT AND MAY NOT REFLECT ACTUAL PATIENT DATA. FORMAT AND/OR CONTENT MAY BE UPDATED PERIODICALLY. RESULTS SNP Array Copy Number Variations Result: LOSS,
SAMPLE REPORT SNP Array NOTE: THIS IS A SAMPLE REPORT AND MAY NOT REFLECT ACTUAL PATIENT DATA. FORMAT AND/OR CONTENT MAY BE UPDATED PERIODICALLY. RESULTS SNP Array Copy Number Variations Result: LOSS,
CHROMOSOMAL MICROARRAY (CGH+SNP)
 Chromosome imbalances are a significant cause of developmental delay, mental retardation, autism spectrum disorders, dysmorphic features and/or birth defects. The imbalance of genetic material may be due
Chromosome imbalances are a significant cause of developmental delay, mental retardation, autism spectrum disorders, dysmorphic features and/or birth defects. The imbalance of genetic material may be due
SNP Array NOTE: THIS IS A SAMPLE REPORT AND MAY NOT REFLECT ACTUAL PATIENT DATA. FORMAT AND/OR CONTENT MAY BE UPDATED PERIODICALLY.
 SAMPLE REPORT SNP Array NOTE: THIS IS A SAMPLE REPORT AND MAY NOT REFLECT ACTUAL PATIENT DATA. FORMAT AND/OR CONTENT MAY BE UPDATED PERIODICALLY. RESULTS SNP Array Copy Number Variations Result: GAIN,
SAMPLE REPORT SNP Array NOTE: THIS IS A SAMPLE REPORT AND MAY NOT REFLECT ACTUAL PATIENT DATA. FORMAT AND/OR CONTENT MAY BE UPDATED PERIODICALLY. RESULTS SNP Array Copy Number Variations Result: GAIN,
Approach to Mental Retardation and Developmental Delay. SR Ghaffari MSc MD PhD
 Approach to Mental Retardation and Developmental Delay SR Ghaffari MSc MD PhD Introduction Objectives Definition of MR and DD Classification Epidemiology (prevalence, recurrence risk, ) Etiology Importance
Approach to Mental Retardation and Developmental Delay SR Ghaffari MSc MD PhD Introduction Objectives Definition of MR and DD Classification Epidemiology (prevalence, recurrence risk, ) Etiology Importance
CHROMOSOME MICROARRAY TESTING
 CHROMOSOME MICROARRAY TESTING UnitedHealthcare Commercial Medical Policy Policy Number: 2017T0559J Effective Date: August 1, 2017 Table of Contents Page INSTRUCTIONS FOR USE... 1 BENEFIT CONSIDERATIONS...
CHROMOSOME MICROARRAY TESTING UnitedHealthcare Commercial Medical Policy Policy Number: 2017T0559J Effective Date: August 1, 2017 Table of Contents Page INSTRUCTIONS FOR USE... 1 BENEFIT CONSIDERATIONS...
Case Report Prenatal Diagnosis of Cystic Hygroma related to a Deletion of 16q24.1 with Haploinsufficiency of FOXF1 and FOXC2 Genes
 Case Reports in Genetics Volume 2012, Article ID 490408, 4 pages doi:10.1155/2012/490408 Case Report Prenatal Diagnosis of Cystic Hygroma related to a Deletion of 16q24.1 with Haploinsufficiency of FOXF1
Case Reports in Genetics Volume 2012, Article ID 490408, 4 pages doi:10.1155/2012/490408 Case Report Prenatal Diagnosis of Cystic Hygroma related to a Deletion of 16q24.1 with Haploinsufficiency of FOXF1
PROVIDER POLICIES & PROCEDURES
 PROVIDER POLICIES & PROCEDURES COMPARATIVE GENOMIC HYBRIDIZATION (CGH) MICROARRAY TESTING FOR DEVELOPMENTAL DELAY, AUTISM SPECTRUM DISORDER AND INTELLECTUAL DISABILITY The purpose of this policy is to
PROVIDER POLICIES & PROCEDURES COMPARATIVE GENOMIC HYBRIDIZATION (CGH) MICROARRAY TESTING FOR DEVELOPMENTAL DELAY, AUTISM SPECTRUM DISORDER AND INTELLECTUAL DISABILITY The purpose of this policy is to
Corporate Medical Policy
 Corporate Medical Policy Invasive Prenatal (Fetal) Diagnostic Testing File Name: Origination: Last CAP Review: Next CAP Review: Last Review: invasive_prenatal_(fetal)_diagnostic_testing 12/2014 3/2018
Corporate Medical Policy Invasive Prenatal (Fetal) Diagnostic Testing File Name: Origination: Last CAP Review: Next CAP Review: Last Review: invasive_prenatal_(fetal)_diagnostic_testing 12/2014 3/2018
Clinical Policy Title: Array comparative genomic hybridization testing
 Clinical Policy Title: Array comparative genomic hybridization testing Clinical Policy Number: 02.01.03 Policy contains: Chromosomal microarray analysis. Effective Date: September 1, 2015 Comparative genomic
Clinical Policy Title: Array comparative genomic hybridization testing Clinical Policy Number: 02.01.03 Policy contains: Chromosomal microarray analysis. Effective Date: September 1, 2015 Comparative genomic
Multiple Copy Number Variations in a Patient with Developmental Delay ASCLS- March 31, 2016
 Multiple Copy Number Variations in a Patient with Developmental Delay ASCLS- March 31, 2016 Marwan Tayeh, PhD, FACMG Director, MMGL Molecular Genetics Assistant Professor of Pediatrics Department of Pediatrics
Multiple Copy Number Variations in a Patient with Developmental Delay ASCLS- March 31, 2016 Marwan Tayeh, PhD, FACMG Director, MMGL Molecular Genetics Assistant Professor of Pediatrics Department of Pediatrics
Dr Dipti Deshmukh. Mayu Uemura. 11th September Introduction
 Genetic Findings in Children with Unexplained Developmental Delay Dr Dipti Deshmukh Neurodevelopmental Consultant, St. George's Hospital, London Mayu Uemura 4th year MBBS4, St George s University of London
Genetic Findings in Children with Unexplained Developmental Delay Dr Dipti Deshmukh Neurodevelopmental Consultant, St. George's Hospital, London Mayu Uemura 4th year MBBS4, St George s University of London
Faravareh Khordadpoor (PhD in molecular genetics) 1- Tehran Medical Genetics Laboratory 2- Science and research branch, Islamic Azad University
 Faravareh Khordadpoor (PhD in molecular genetics) 1- Tehran Medical Genetics Laboratory 2- Science and research branch, Islamic Azad University 1395 21 مشاوره ژنتیک و نقش آن در پیش گیری از معلولیت ها 20
Faravareh Khordadpoor (PhD in molecular genetics) 1- Tehran Medical Genetics Laboratory 2- Science and research branch, Islamic Azad University 1395 21 مشاوره ژنتیک و نقش آن در پیش گیری از معلولیت ها 20
Clinical Policy Title: Array comparative genomic hybridization testing
 Clinical Policy Title: Array comparative genomic hybridization testing Clinical Policy Number: 02.01.03 Effective Date: Sept 1, 2015 Initial Review Date: May 13, 2013 Most Recent Review Date: September
Clinical Policy Title: Array comparative genomic hybridization testing Clinical Policy Number: 02.01.03 Effective Date: Sept 1, 2015 Initial Review Date: May 13, 2013 Most Recent Review Date: September
Medical Policy. MP Invasive Prenatal (Fetal) Diagnostic Testing
 Medical Policy MP 2.04.116 BCBSA Ref. Policy: 2.04.116 Last Review: 04/30/2018 Effective Date: 04/30/2018 Section: Medicine Related Policies 2.04.59 Genetic Testing for Developmental Delay/Intellectual
Medical Policy MP 2.04.116 BCBSA Ref. Policy: 2.04.116 Last Review: 04/30/2018 Effective Date: 04/30/2018 Section: Medicine Related Policies 2.04.59 Genetic Testing for Developmental Delay/Intellectual
Spectrum of Cranio-facial anomalies during 2 Ultrasound. trimester on
 Spectrum of Cranio-facial anomalies during 2 Ultrasound nd trimester on Poster No.: C-0378 Congress: ECR 2015 Type: Scientific Exhibit Authors: K. Dave, S. Solanki; Ahmedabad/IN Keywords: Obstetrics (Pregnancy
Spectrum of Cranio-facial anomalies during 2 Ultrasound nd trimester on Poster No.: C-0378 Congress: ECR 2015 Type: Scientific Exhibit Authors: K. Dave, S. Solanki; Ahmedabad/IN Keywords: Obstetrics (Pregnancy
Practical challenges that copy number variation and whole genome sequencing create for genetic diagnostic labs
 Practical challenges that copy number variation and whole genome sequencing create for genetic diagnostic labs Joris Vermeesch, Center for Human Genetics K.U.Leuven, Belgium ESHG June 11, 2010 When and
Practical challenges that copy number variation and whole genome sequencing create for genetic diagnostic labs Joris Vermeesch, Center for Human Genetics K.U.Leuven, Belgium ESHG June 11, 2010 When and
Prenatal Prediction of The Neurologically Impaired Neonate By Ultrasound
 Prenatal Prediction of The Neurologically Impaired Neonate By Ultrasound Robert H. Debbs, D.O.,F.A.C.O.O.G. Professor of OB-GYN Perelman School of Medicine, University of Pennsylvania Director, Pennsylvania
Prenatal Prediction of The Neurologically Impaired Neonate By Ultrasound Robert H. Debbs, D.O.,F.A.C.O.O.G. Professor of OB-GYN Perelman School of Medicine, University of Pennsylvania Director, Pennsylvania
Human Genetic Variation Copy Number Variation
 The Evolution of Laboratory Prenatal Diagnosis Lawrence D. Platt, MD Professor of Obstetrics and Gynecology David Geffen School of Medicine at UCLA Center for Fetal Medicine & Women s Ultrasound Los Angeles,
The Evolution of Laboratory Prenatal Diagnosis Lawrence D. Platt, MD Professor of Obstetrics and Gynecology David Geffen School of Medicine at UCLA Center for Fetal Medicine & Women s Ultrasound Los Angeles,
Sharan Goobie, MD, MSc, FRCPC
 Sharan Goobie, MD, MSc, FRCPC Chromosome testing in 2014 Presenter Disclosure: Sharan Goobie has no potential for conflict of interest with this presentation Objectives Review of standard genetic investigations
Sharan Goobie, MD, MSc, FRCPC Chromosome testing in 2014 Presenter Disclosure: Sharan Goobie has no potential for conflict of interest with this presentation Objectives Review of standard genetic investigations
Chromosome microarray analysis in routine prenatal diagnosis practice: a prospective study on 3000 consecutive clinical cases
 Chromosome microarray analysis in routine prenatal diagnosis practice: a prospective study on 3000 consecutive clinical cases Fiorentino F, Napoletano S, Caiazzo F, Sessa M, Bono S, Spizzichino L, Gordon
Chromosome microarray analysis in routine prenatal diagnosis practice: a prospective study on 3000 consecutive clinical cases Fiorentino F, Napoletano S, Caiazzo F, Sessa M, Bono S, Spizzichino L, Gordon
What s the Human Genome Project Got to Do with Developmental Disabilities?
 What s the Human Genome Project Got to Do with Developmental Disabilities? Disclosures Neither speaker has anything to disclose. Phase Two: Interpretation Officially started in October 1990 Goals of the
What s the Human Genome Project Got to Do with Developmental Disabilities? Disclosures Neither speaker has anything to disclose. Phase Two: Interpretation Officially started in October 1990 Goals of the
CHROMOSOME MICROARRAY TESTING (NON-ONCOLOGY CONDITIONS)
 CHROMOSOME MICROARRAY TESTING (NON-ONCOLOGY CONDITIONS) UnitedHealthcare Oxford Clinical Policy Policy Number: LABORATORY 016.12 T2 Effective Date: June 1, 2018 Table of Contents Page INSTRUCTIONS FOR
CHROMOSOME MICROARRAY TESTING (NON-ONCOLOGY CONDITIONS) UnitedHealthcare Oxford Clinical Policy Policy Number: LABORATORY 016.12 T2 Effective Date: June 1, 2018 Table of Contents Page INSTRUCTIONS FOR
Supplemental Information
 ARTICLE Supplemental Information SUPPLEMENTAL TABLE 6 Mosaic and Partial Trisomies Thirty-eight VLBW infants were identified with T13, of whom 2 had mosaic T13. T18 was reported for 128 infants, of whom
ARTICLE Supplemental Information SUPPLEMENTAL TABLE 6 Mosaic and Partial Trisomies Thirty-eight VLBW infants were identified with T13, of whom 2 had mosaic T13. T18 was reported for 128 infants, of whom
National Medical Policy
 National Medical Policy Subject: Policy Number: Single Nucleotide Polymorphism (SNP) Chromosomal Microarray Analysis for Prenatal Testing and Intellectual Disability, Developmental Delay, and Multiple
National Medical Policy Subject: Policy Number: Single Nucleotide Polymorphism (SNP) Chromosomal Microarray Analysis for Prenatal Testing and Intellectual Disability, Developmental Delay, and Multiple
intracranial anomalies
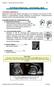 Chapter 5: Fetal Central Nervous System 84 intracranial anomalies Hydrocephaly Dilatation of ventricular system secondary to an increase in the amount of CSF. Effects of hydrocephalus include flattening
Chapter 5: Fetal Central Nervous System 84 intracranial anomalies Hydrocephaly Dilatation of ventricular system secondary to an increase in the amount of CSF. Effects of hydrocephalus include flattening
Clinical evaluation of microarray data
 Clinical evaluation of microarray data David Amor 19 th June 2011 Single base change Microarrays 3-4Mb What is a microarray? Up to 10 6 bits of Information!! Highly multiplexed FISH hybridisations. Microarray
Clinical evaluation of microarray data David Amor 19 th June 2011 Single base change Microarrays 3-4Mb What is a microarray? Up to 10 6 bits of Information!! Highly multiplexed FISH hybridisations. Microarray
SNP Microarray. Prenatal
 SNP Microarray Prenatal SNP Microarray Reveal SNP Microarray is a test that analyzes chromosomes for changes that can explain certain kinds of birth defects. This brochure is designed to answer some of
SNP Microarray Prenatal SNP Microarray Reveal SNP Microarray is a test that analyzes chromosomes for changes that can explain certain kinds of birth defects. This brochure is designed to answer some of
Clinical Interpretation of Cancer Genomes
 IGENZ Ltd, Auckland, New Zealand Clinical Interpretation of Cancer Genomes Dr Amanda Dixon-McIver www.igenz.co.nz 1992 Slovenia and Croatia gain independence USA and Russia declare the Cold War over Steffi
IGENZ Ltd, Auckland, New Zealand Clinical Interpretation of Cancer Genomes Dr Amanda Dixon-McIver www.igenz.co.nz 1992 Slovenia and Croatia gain independence USA and Russia declare the Cold War over Steffi
Global variation in copy number in the human genome
 Global variation in copy number in the human genome Redon et. al. Nature 444:444-454 (2006) 12.03.2007 Tarmo Puurand Study 270 individuals (HapMap collection) Affymetrix 500K Whole Genome TilePath (WGTP)
Global variation in copy number in the human genome Redon et. al. Nature 444:444-454 (2006) 12.03.2007 Tarmo Puurand Study 270 individuals (HapMap collection) Affymetrix 500K Whole Genome TilePath (WGTP)
Prior Authorization Review Panel MCO Policy Submission
 Prior Authorization Review Panel MCO Policy Submission A separate copy of this form must accompany each policy submitted for review. Policies submitted without this form will not be considered for review.
Prior Authorization Review Panel MCO Policy Submission A separate copy of this form must accompany each policy submitted for review. Policies submitted without this form will not be considered for review.
POLICY PRODUCT VARIATIONS DESCRIPTION/BACKGROUND RATIONALE DEFINITIONS BENEFIT VARIATIONS DISCLAIMER CODING INFORMATION REFERENCES POLICY HISTORY
 Original Issue Date (Created): April 26, 2011 Most Recent Review Date (Revised): January 28, 2014 Effective Date: June 1, 2014 POLICY PRODUCT VARIATIONS DESCRIPTION/BACKGROUND RATIONALE DEFINITIONS BENEFIT
Original Issue Date (Created): April 26, 2011 Most Recent Review Date (Revised): January 28, 2014 Effective Date: June 1, 2014 POLICY PRODUCT VARIATIONS DESCRIPTION/BACKGROUND RATIONALE DEFINITIONS BENEFIT
Clinical Policy Title: Genetic testing for primary autosomal recessive microcephaly
 Clinical Policy Title: Genetic testing for primary autosomal recessive microcephaly Clinical Policy Number: (391) 02.01.03 Effective Date: June 1, 2013 Initial Review Date: April 22, 2013 Most Recent Review
Clinical Policy Title: Genetic testing for primary autosomal recessive microcephaly Clinical Policy Number: (391) 02.01.03 Effective Date: June 1, 2013 Initial Review Date: April 22, 2013 Most Recent Review
Genetic Testing for the Developmental Delay/Intellectual Disability, Autism Spectrum Disorder, and Congenital Anomalies
 Genetic Testing for the Developmental Delay/Intellectual Disability, Autism Spectrum Disorder, and Congenital Anomalies Policy Number: 2.04.59 Last Review: 5/2018 Origination: 5/2015 Next Review: 5/2019
Genetic Testing for the Developmental Delay/Intellectual Disability, Autism Spectrum Disorder, and Congenital Anomalies Policy Number: 2.04.59 Last Review: 5/2018 Origination: 5/2015 Next Review: 5/2019
New and Developing Technologies for Genetic Diagnostics National Genetics Reference Laboratory (Wessex) Salisbury, UK - July 2010 BACs on Beads
 New and Developing Technologies for Genetic Diagnostics National Genetics Reference Laboratory (Wessex) Salisbury, UK - July 2010 BACs on Beads Susan Gross, MD Division of Reproductive Genetics Professor
New and Developing Technologies for Genetic Diagnostics National Genetics Reference Laboratory (Wessex) Salisbury, UK - July 2010 BACs on Beads Susan Gross, MD Division of Reproductive Genetics Professor
22q11.2 DELETION SYNDROME. Anna Mª Cueto González Clinical Geneticist Programa de Medicina Molecular y Genética Hospital Vall d Hebrón (Barcelona)
 22q11.2 DELETION SYNDROME Anna Mª Cueto González Clinical Geneticist Programa de Medicina Molecular y Genética Hospital Vall d Hebrón (Barcelona) Genomic disorders GENOMICS DISORDERS refers to those diseases
22q11.2 DELETION SYNDROME Anna Mª Cueto González Clinical Geneticist Programa de Medicina Molecular y Genética Hospital Vall d Hebrón (Barcelona) Genomic disorders GENOMICS DISORDERS refers to those diseases
review Genetics IN Medicine 13
 January 2008 Vol. 10 No. 1 review Pre- and postnatal genetic testing by array-comparative genomic hybridization: genetic counseling perspectives Sandra Darilek, MS, Patricia Ward, MS, Amber Pursley, MS,
January 2008 Vol. 10 No. 1 review Pre- and postnatal genetic testing by array-comparative genomic hybridization: genetic counseling perspectives Sandra Darilek, MS, Patricia Ward, MS, Amber Pursley, MS,
Implementation of the DDD/ClinGen OGT (CytoSure v3) Microarray
 Implementation of the DDD/ClinGen OGT (CytoSure v3) Microarray OGT UGM Birmingham 08/09/2016 Dom McMullan Birmingham Women's NHS Trust WM chromosomal microarray (CMA) testing Population of ~6 million (10%)
Implementation of the DDD/ClinGen OGT (CytoSure v3) Microarray OGT UGM Birmingham 08/09/2016 Dom McMullan Birmingham Women's NHS Trust WM chromosomal microarray (CMA) testing Population of ~6 million (10%)
Whole-Genome SNP Array Analysis Genomic Complexity and Clinical Relevance in Prenatal, Postnatal, and Oncology Testing
 The Evolution of Cytogenetics Testing Whole-Genome SNP Array Analysis Genomic Complexity and Clinical Relevance in Prenatal, Postnatal, and Oncology Testing Mark Micale, Ph.D., F.A.C.M.G. Medical Director,
The Evolution of Cytogenetics Testing Whole-Genome SNP Array Analysis Genomic Complexity and Clinical Relevance in Prenatal, Postnatal, and Oncology Testing Mark Micale, Ph.D., F.A.C.M.G. Medical Director,
Supplementary Online Content
 Supplementary Online Content Honein MA, Dawson AL, Petersen E, et al; US Zika Pregnancy Registry Collaboration. Birth Defects Among Fetuses and Infants of US Women With Laboratory Evidence of Possible
Supplementary Online Content Honein MA, Dawson AL, Petersen E, et al; US Zika Pregnancy Registry Collaboration. Birth Defects Among Fetuses and Infants of US Women With Laboratory Evidence of Possible
Association for Molecular Pathology Promoting Clinical Practice, Basic Research, and Education in Molecular Pathology
 Association for Molecular Pathology Promoting Clinical Practice, Basic Research, and Education in Molecular Pathology 9650 Rockville Pike, Bethesda, Maryland 20814 Tel: 301-634-7939 Fax: 301-634-7990 Email:
Association for Molecular Pathology Promoting Clinical Practice, Basic Research, and Education in Molecular Pathology 9650 Rockville Pike, Bethesda, Maryland 20814 Tel: 301-634-7939 Fax: 301-634-7990 Email:
Genomic structural variation
 Genomic structural variation Mario Cáceres The new genomic variation DNA sequence differs across individuals much more than researchers had suspected through structural changes A huge amount of structural
Genomic structural variation Mario Cáceres The new genomic variation DNA sequence differs across individuals much more than researchers had suspected through structural changes A huge amount of structural
Case 1B. 46,XY,-14,+t(14;21)
 Case 1B 46,XY,-14,+t(14;21) G-banded Chromosome telomere centromere G-dark bands AT-rich few genes G-pale bands GC-rich many genes telomere ideograms ideograms Conventional (light microscopy) p = short
Case 1B 46,XY,-14,+t(14;21) G-banded Chromosome telomere centromere G-dark bands AT-rich few genes G-pale bands GC-rich many genes telomere ideograms ideograms Conventional (light microscopy) p = short
Structural Variation and Medical Genomics
 Structural Variation and Medical Genomics Andrew King Department of Biomedical Informatics July 8, 2014 You already know about small scale genetic mutations Single nucleotide polymorphism (SNPs) Deletions,
Structural Variation and Medical Genomics Andrew King Department of Biomedical Informatics July 8, 2014 You already know about small scale genetic mutations Single nucleotide polymorphism (SNPs) Deletions,
Section: Medicine Effective Date: January15, 2016 Subsection: Pathology/Laboratory Original Policy Date: December 7, 2011 Subject:
 Section: Medicine Effective Date: January15, 2016 Last Review Status/Date: December 2015 Page: 1 of 22 Microarray Analysis and Next-Generation Autism Spectrum Disorder and/or Congenital Description Chromosomal
Section: Medicine Effective Date: January15, 2016 Last Review Status/Date: December 2015 Page: 1 of 22 Microarray Analysis and Next-Generation Autism Spectrum Disorder and/or Congenital Description Chromosomal
Invasive Prenatal (Fetal) Diagnostic Testing
 Protocol Invasive Prenatal (Fetal) Diagnostic Testing (204116) Medical Benefit Effective Date: 04/01/18 Next Review Date: 01/19 Preauthorization Yes Review Dates: 01/15, 01/16, 01/17, 01/18 Preauthorization
Protocol Invasive Prenatal (Fetal) Diagnostic Testing (204116) Medical Benefit Effective Date: 04/01/18 Next Review Date: 01/19 Preauthorization Yes Review Dates: 01/15, 01/16, 01/17, 01/18 Preauthorization
chromosomal anomalies and mental pdf Chapter 8: Chromosomes and Chromosomal Anomalies (PDF) Chromosomal abnormalities -A review - ResearchGate
 DOWNLOAD OR READ : CHROMOSOMAL ANOMALIES AND MENTAL RETARDATION FROM GENOTYPES TO NEUROPSYCHOLOGICAL PHENOTYPES OF GENETIC SYNDROMES AT HIGH INCIDENCEGENOTYPE TO PHENOTYPE PDF EBOOK EPUB MOBI Page 1 Page
DOWNLOAD OR READ : CHROMOSOMAL ANOMALIES AND MENTAL RETARDATION FROM GENOTYPES TO NEUROPSYCHOLOGICAL PHENOTYPES OF GENETIC SYNDROMES AT HIGH INCIDENCEGENOTYPE TO PHENOTYPE PDF EBOOK EPUB MOBI Page 1 Page
Genetic Testing 101: Interpreting the Chromosomes
 Genetic Testing 101: Interpreting the Chromosomes Kristin Lindstrom, MD Division of Genetics and Metabolism Phoenix Children s Hospital AzAAP Pediatrics in the Red Rocks I have no disclosures for this
Genetic Testing 101: Interpreting the Chromosomes Kristin Lindstrom, MD Division of Genetics and Metabolism Phoenix Children s Hospital AzAAP Pediatrics in the Red Rocks I have no disclosures for this
Symposium: OB/GY US (Room B) CNS Anomalies
 82 Symposium: OB/GY US (Room B) 11 : 50 1 2 : 10 CNS Anomalies Brain area Midline structure S u p r a t e n t o r i a l ventricular system Cerebral hemisphere Posterior fossa Head size and shape Image
82 Symposium: OB/GY US (Room B) 11 : 50 1 2 : 10 CNS Anomalies Brain area Midline structure S u p r a t e n t o r i a l ventricular system Cerebral hemisphere Posterior fossa Head size and shape Image
GENETICS ROTATION OBJECTIVES MATERNAL-FETAL MEDICINE FELLOWSHIP
 GENETICS ROTATION OBJECTIVES MATERNAL-FETAL MEDICINE FELLOWSHIP University of New Mexico 1. General Description: UNM MFM fellows rotate through genetics during their PGY5 and PGY7 years. The PGY5 fellow
GENETICS ROTATION OBJECTIVES MATERNAL-FETAL MEDICINE FELLOWSHIP University of New Mexico 1. General Description: UNM MFM fellows rotate through genetics during their PGY5 and PGY7 years. The PGY5 fellow
Complex Hydrocephalus
 2012 Hydrocephalus Association Conference Washington, DC - June 27-July1, 2012 Complex Hydrocephalus Marion L. Walker, MD Professor of Neurosurgery & Pediatrics Primary Children s Medical Center University
2012 Hydrocephalus Association Conference Washington, DC - June 27-July1, 2012 Complex Hydrocephalus Marion L. Walker, MD Professor of Neurosurgery & Pediatrics Primary Children s Medical Center University
Increasing the Sensitivity of Genomic Assays Permits a Higher Resolution Study of the Human Genome. Our Current Knowledge of Genetics and Genomics
 Our Current Knowledge of Genetics and Genomics Unlocking the Diagnostic Potential of Genomic Medicine The Beaumont DNA Array Experience Mark Micale, Ph.D., F.A.C.M.G. Medical Director, Clinical Cytogenomics
Our Current Knowledge of Genetics and Genomics Unlocking the Diagnostic Potential of Genomic Medicine The Beaumont DNA Array Experience Mark Micale, Ph.D., F.A.C.M.G. Medical Director, Clinical Cytogenomics
Case Report Prenatal Diagnosis of 4p and 4q Subtelomeric Microdeletion in De Novo Ring Chromosome 4
 Case Reports in Obstetrics and Gynecology Volume 2013, Article ID 248050, 5 pages http://dx.doi.org/10.1155/2013/248050 Case Report Prenatal Diagnosis of 4p and 4q Subtelomeric Microdeletion in De Novo
Case Reports in Obstetrics and Gynecology Volume 2013, Article ID 248050, 5 pages http://dx.doi.org/10.1155/2013/248050 Case Report Prenatal Diagnosis of 4p and 4q Subtelomeric Microdeletion in De Novo
Genetic Testing for Single-Gene and Multifactorial Conditions
 Clinical Appropriateness Guidelines Genetic Testing for Single-Gene and Multifactorial Conditions EFFECTIVE DECEMBER 1, 2017 Appropriate.Safe.Affordable 2017 AIM Specialty Health 2069-1217 Table of Contents
Clinical Appropriateness Guidelines Genetic Testing for Single-Gene and Multifactorial Conditions EFFECTIVE DECEMBER 1, 2017 Appropriate.Safe.Affordable 2017 AIM Specialty Health 2069-1217 Table of Contents
Canadian College of Medical Geneticists (CCMG) Cytogenetics Examination. May 4, 2010
 Canadian College of Medical Geneticists (CCMG) Cytogenetics Examination May 4, 2010 Examination Length = 3 hours Total Marks = 100 (7 questions) Total Pages = 8 (including cover sheet and 2 pages of prints)
Canadian College of Medical Geneticists (CCMG) Cytogenetics Examination May 4, 2010 Examination Length = 3 hours Total Marks = 100 (7 questions) Total Pages = 8 (including cover sheet and 2 pages of prints)
CNS Embryology 5th Menstrual Week (Dorsal View)
 Imaging of the Fetal Brain; Normal & Abnormal Alfred Abuhamad, M.D. Eastern Virginia Medical School CNS Embryology 5th Menstrual Week (Dorsal View) Day 20 from fertilization Neural plate formed in ectoderm
Imaging of the Fetal Brain; Normal & Abnormal Alfred Abuhamad, M.D. Eastern Virginia Medical School CNS Embryology 5th Menstrual Week (Dorsal View) Day 20 from fertilization Neural plate formed in ectoderm
Understanding the Human Karyotype Colleen Jackson Cook, Ph.D.
 Understanding the Human Karyotype Colleen Jackson Cook, Ph.D. SUPPLEMENTAL READING Nussbaum, RL, McInnes, RR, and Willard HF (2007) Thompson and Thompson Genetics in Medicine, 7th edition. Saunders: Philadelphia.
Understanding the Human Karyotype Colleen Jackson Cook, Ph.D. SUPPLEMENTAL READING Nussbaum, RL, McInnes, RR, and Willard HF (2007) Thompson and Thompson Genetics in Medicine, 7th edition. Saunders: Philadelphia.
Central nervous system. Obstetrics Content Outline Obstetrics - Fetal Abnormalities
 Obstetrics Content Outline Obstetrics - Fetal Abnormalities Many congenital malformations of the CNS result from incomplete closure of the neural tube Effective February 2007 10 16% the most common neural
Obstetrics Content Outline Obstetrics - Fetal Abnormalities Many congenital malformations of the CNS result from incomplete closure of the neural tube Effective February 2007 10 16% the most common neural
Seven cases of intellectual disability analysed by genomewide SNP analysis. Rodney J. Scott
 Seven cases of intellectual disability analysed by genomewide SNP analysis Rodney J. Scott Ability to interrogate Genomic Material ~1930 Crude analyses 2012 Highly precise analyses A (very) brief history
Seven cases of intellectual disability analysed by genomewide SNP analysis Rodney J. Scott Ability to interrogate Genomic Material ~1930 Crude analyses 2012 Highly precise analyses A (very) brief history
Han-Sung Kwon M.D. Department of Obstetrics and Gynecology Konkuk University School of Medicine Seoul, Korea
 Han-Sung Kwon M.D. Department of Obstetrics and Gynecology Konkuk University School of Medicine Seoul, Korea Embryologic features of the developing hindbrain Embryologic features of the developing hindbrain
Han-Sung Kwon M.D. Department of Obstetrics and Gynecology Konkuk University School of Medicine Seoul, Korea Embryologic features of the developing hindbrain Embryologic features of the developing hindbrain
CMA. [DOI] /j.issn
![CMA. [DOI] /j.issn CMA. [DOI] /j.issn](/thumbs/93/111203021.jpg) 902 2017 10 1 42 10 [ ] (CMA) (CNVs) CMA 2015 1 2016 11 70 / 1.0cm G G Affymetrix CytoScan TM 750k CMA CMA 70 CMA 9 CNVs 3 CNVs 1 (LOH) 706 CNVs 2 (33.3% 2/6) 3 CNVs CNVs 1 (33.3% 1/3) 31 CNVs 6 (19.4%
902 2017 10 1 42 10 [ ] (CMA) (CNVs) CMA 2015 1 2016 11 70 / 1.0cm G G Affymetrix CytoScan TM 750k CMA CMA 70 CMA 9 CNVs 3 CNVs 1 (LOH) 706 CNVs 2 (33.3% 2/6) 3 CNVs CNVs 1 (33.3% 1/3) 31 CNVs 6 (19.4%
Section: Genetic Testing Last Reviewed Date: April Policy No: 58 Effective Date: July 1, 2014
 Medical Policy Manual Topic: Genetic Testing, including Chromosomal Microarray Analysis (CMA) and Next Generation Sequencing Panels, for the Genetic Evaluation of Patients with Developmental Delay/Intellectual
Medical Policy Manual Topic: Genetic Testing, including Chromosomal Microarray Analysis (CMA) and Next Generation Sequencing Panels, for the Genetic Evaluation of Patients with Developmental Delay/Intellectual
Chromosome Microarray Analysis (CMA)
 Chromosome Microarray Analysis (CMA) Ina E. Amarillo, PhD FACMG Cytogenomics Lab (Associate Medical Director) Pathology (Assistant Professor) OUTLINE Clinical Indications for Cytogenomics Testing Cytogenomics
Chromosome Microarray Analysis (CMA) Ina E. Amarillo, PhD FACMG Cytogenomics Lab (Associate Medical Director) Pathology (Assistant Professor) OUTLINE Clinical Indications for Cytogenomics Testing Cytogenomics
Genetic Counselling in relation to genetic testing
 Genetic Counselling in relation to genetic testing Dr Julie Vogt Consultant Geneticist West Midlands Regional Genetics Service September 2016 Disclosures for Research Support/P.I. Employee Consultant Major
Genetic Counselling in relation to genetic testing Dr Julie Vogt Consultant Geneticist West Midlands Regional Genetics Service September 2016 Disclosures for Research Support/P.I. Employee Consultant Major
SEAMLESS CGH DIAGNOSTIC TESTING
 SEAMLESS CGH DIAGNOSTIC TESTING GENETISURE DX POSTNATAL ASSAY Informed decisions start with a complete microarray platform for postnatal analysis For In Vitro Diagnostic Use INTENDED USE: GenetiSure Dx
SEAMLESS CGH DIAGNOSTIC TESTING GENETISURE DX POSTNATAL ASSAY Informed decisions start with a complete microarray platform for postnatal analysis For In Vitro Diagnostic Use INTENDED USE: GenetiSure Dx
Copy Number Variants of Uncertain Significance in Prenatal diagnosis Are the Goalposts Moving? Lisa Burvill-Holmes Bristol Genetics Laboratory
 Copy Number Variants of Uncertain Significance in Prenatal diagnosis Are the Goalposts Moving? Lisa Burvill-Holmes Bristol Genetics Laboratory http://www.nbt.nhs.uk/genetics Microarray CGH in Prenatal
Copy Number Variants of Uncertain Significance in Prenatal diagnosis Are the Goalposts Moving? Lisa Burvill-Holmes Bristol Genetics Laboratory http://www.nbt.nhs.uk/genetics Microarray CGH in Prenatal
CLINICAL MEDICAL POLICY
 Policy Name: Policy Number: Approved By: CLINICAL MEDICAL POLICY Chromosomal Microarray Analysis: Comparative Genomic Hybridization (CGH) and Single Nucleotide Polymorphism (SNP) MP-012-MD-WV Medical Management
Policy Name: Policy Number: Approved By: CLINICAL MEDICAL POLICY Chromosomal Microarray Analysis: Comparative Genomic Hybridization (CGH) and Single Nucleotide Polymorphism (SNP) MP-012-MD-WV Medical Management
Application of Array-based Comparative Genome Hybridization in Children with Developmental Delay or Mental Retardation
 Pediatr Neonatol 2008;49(6):213 217 REVIEW ARTICLE Application of Array-based Comparative Genome Hybridization in Children with Developmental Delay or Mental Retardation Jao-Shwann Liang 1,2 *, Keiko Shimojima
Pediatr Neonatol 2008;49(6):213 217 REVIEW ARTICLE Application of Array-based Comparative Genome Hybridization in Children with Developmental Delay or Mental Retardation Jao-Shwann Liang 1,2 *, Keiko Shimojima
THE PENNSYLVANIA STATE UNIVERSITY SCHREYER HONORS COLLEGE DEPARTMENT OF BIOCHEMISTRY AND MOLECULAR BIOLOGY
 THE PENNSYLVANIA STATE UNIVERSITY SCHREYER HONORS COLLEGE DEPARTMENT OF BIOCHEMISTRY AND MOLECULAR BIOLOGY GENOME-WIDE MICROARRAY ANALYSIS IN A CASE-CONTROL STUDY REVEALS EVELATED LEVEL OF GLOBAL COPY
THE PENNSYLVANIA STATE UNIVERSITY SCHREYER HONORS COLLEGE DEPARTMENT OF BIOCHEMISTRY AND MOLECULAR BIOLOGY GENOME-WIDE MICROARRAY ANALYSIS IN A CASE-CONTROL STUDY REVEALS EVELATED LEVEL OF GLOBAL COPY
Case Report Severe Psychomotor Delay in a Severe Presentation of Cat-Eye Syndrome
 Case Reports in Genetics Volume 2015, Article ID 943905, 4 pages http://dx.doi.org/10.1155/2015/943905 Case Report Severe Psychomotor Delay in a Severe Presentation of Cat-Eye Syndrome Guillaume Jedraszak,
Case Reports in Genetics Volume 2015, Article ID 943905, 4 pages http://dx.doi.org/10.1155/2015/943905 Case Report Severe Psychomotor Delay in a Severe Presentation of Cat-Eye Syndrome Guillaume Jedraszak,
November 9, Johns Hopkins School of Medicine, Baltimore, MD,
 Fast detection of de-novo copy number variants from case-parent SNP arrays identifies a deletion on chromosome 7p14.1 associated with non-syndromic isolated cleft lip/palate Samuel G. Younkin 1, Robert
Fast detection of de-novo copy number variants from case-parent SNP arrays identifies a deletion on chromosome 7p14.1 associated with non-syndromic isolated cleft lip/palate Samuel G. Younkin 1, Robert
Approach to the Genetic Diagnosis of Neurological Disorders
 Approach to the Genetic Diagnosis of Neurological Disorders Dr Wendy Jones MBBS MRCP Great Ormond Street Hospital for Children National Hospital for Neurology and Neurosurgery What is a genetic diagnosis?
Approach to the Genetic Diagnosis of Neurological Disorders Dr Wendy Jones MBBS MRCP Great Ormond Street Hospital for Children National Hospital for Neurology and Neurosurgery What is a genetic diagnosis?
Basic Training. ISUOG Basic Training The 20 Planes Approach to the Routine Mid Trimester Scan
 ISUOG The 20 Planes Approach to the Routine Mid Trimester Scan Learning objective At the end of the lecture you will be able to: Explain how to perform a structured routine examination, including measurements,
ISUOG The 20 Planes Approach to the Routine Mid Trimester Scan Learning objective At the end of the lecture you will be able to: Explain how to perform a structured routine examination, including measurements,
AMERICAN BOARD OF MEDICAL GENETICS AND GENOMICS
 AMERICAN BOARD OF MEDICAL GENETICS AND GENOMICS Logbook Guidelines for Certification in Clinical Genetics and Genomics for the 2017 Examination as of 10/5/2015 Purpose: The purpose of the logbook is to
AMERICAN BOARD OF MEDICAL GENETICS AND GENOMICS Logbook Guidelines for Certification in Clinical Genetics and Genomics for the 2017 Examination as of 10/5/2015 Purpose: The purpose of the logbook is to
Generating Spontaneous Copy Number Variants (CNVs) Jennifer Freeman Assistant Professor of Toxicology School of Health Sciences Purdue University
 Role of Chemical lexposure in Generating Spontaneous Copy Number Variants (CNVs) Jennifer Freeman Assistant Professor of Toxicology School of Health Sciences Purdue University CNV Discovery Reference Genetic
Role of Chemical lexposure in Generating Spontaneous Copy Number Variants (CNVs) Jennifer Freeman Assistant Professor of Toxicology School of Health Sciences Purdue University CNV Discovery Reference Genetic
Protocol. Genetic Testing for Developmental Delay and Autism Spectrum Disorder
 Genetic Testing for Developmental Delay and Autism Spectrum (20459, 20483, 20481) Medical Benefit Effective Date: 01/01/18 Next Review Date: 09/18 Preauthorization Yes Review Dates: 09/10, 09/11, 03/12,
Genetic Testing for Developmental Delay and Autism Spectrum (20459, 20483, 20481) Medical Benefit Effective Date: 01/01/18 Next Review Date: 09/18 Preauthorization Yes Review Dates: 09/10, 09/11, 03/12,
Central nervous system
 Chapter 2 Central nervous system NORMAL SONOGRAPHIC ANATOMY The fetal brain undergoes major developmental changes throughout pregnancy. At 7 weeks of gestation, a sonolucent area is seen in the cephalic
Chapter 2 Central nervous system NORMAL SONOGRAPHIC ANATOMY The fetal brain undergoes major developmental changes throughout pregnancy. At 7 weeks of gestation, a sonolucent area is seen in the cephalic
CNV Detection and Interpretation in Genomic Data
 CNV Detection and Interpretation in Genomic Data Benjamin W. Darbro, M.D., Ph.D. Assistant Professor of Pediatrics Director of the Shivanand R. Patil Cytogenetics and Molecular Laboratory Overview What
CNV Detection and Interpretation in Genomic Data Benjamin W. Darbro, M.D., Ph.D. Assistant Professor of Pediatrics Director of the Shivanand R. Patil Cytogenetics and Molecular Laboratory Overview What
Basic Training. ISUOG Basic Training Examining the Upper Lip, Face & Profile
 ISUOG Examining the Upper Lip, Face & Profile Learning objectives At the end of the lecture you will be able to: Describe how to obtain the 3 planes required to assess the anatomy of the fetal face Recognise
ISUOG Examining the Upper Lip, Face & Profile Learning objectives At the end of the lecture you will be able to: Describe how to obtain the 3 planes required to assess the anatomy of the fetal face Recognise
Chromosome 13. Introduction
 Chromosome 13 Chromosome Disorder Outreach Inc. (CDO) Technical genetic content provided by Dr. Iosif Lurie, M.D. Ph.D Medical Geneticist and CDO Medical Consultant/Advisor. Ideogram courtesy of the University
Chromosome 13 Chromosome Disorder Outreach Inc. (CDO) Technical genetic content provided by Dr. Iosif Lurie, M.D. Ph.D Medical Geneticist and CDO Medical Consultant/Advisor. Ideogram courtesy of the University
Clinical Cytogenomics Laboratory. Laboratory. Leading diagnostics for better medicine. Beaumont
 Clinical Cytogenomics Laboratory Leading diagnostics for better medicine Beaumont Laboratory Beaumont Laboratory s Clinical Cytogenomics Lab Genome decoding is essential Knowledge of variations in genome
Clinical Cytogenomics Laboratory Leading diagnostics for better medicine Beaumont Laboratory Beaumont Laboratory s Clinical Cytogenomics Lab Genome decoding is essential Knowledge of variations in genome
Evolution of Genetic Testing. Joan Pellegrino MD Associate Professor of Pediatrics SUNY Upstate Medical University
 Evolution of Genetic Testing Joan Pellegrino MD Associate Professor of Pediatrics SUNY Upstate Medical University Genetic Testing Chromosomal analysis Flourescent in situ hybridization (FISH) Chromosome
Evolution of Genetic Testing Joan Pellegrino MD Associate Professor of Pediatrics SUNY Upstate Medical University Genetic Testing Chromosomal analysis Flourescent in situ hybridization (FISH) Chromosome
Ultrasound Anomaly Details
 Appendix 2. Association of Copy Number Variants With Specific Ultrasonographically Detected Fetal Anomalies Ultrasound Anomaly Details Abdominal wall Bladder exstrophy Body-stalk anomaly Cloacal exstrophy
Appendix 2. Association of Copy Number Variants With Specific Ultrasonographically Detected Fetal Anomalies Ultrasound Anomaly Details Abdominal wall Bladder exstrophy Body-stalk anomaly Cloacal exstrophy
Introduction to Fetal Medicine: Genetics and Embryology
 Introduction to Fetal Medicine: Genetics and Embryology Question: What do cancer biology, molecular biology, neurobiology, cell biology developmental biology and reproductive biology, all have in common?
Introduction to Fetal Medicine: Genetics and Embryology Question: What do cancer biology, molecular biology, neurobiology, cell biology developmental biology and reproductive biology, all have in common?
Determination of Genomic Imbalances by Genome-wide Screening Approaches
 Overview Determination of Genomic Imbalances by Genome-wide Screening Approaches Károly Szuhai Introduction/Methodologies Applications/Results Conclusion Approaches Introduction/Methodologies Chromosome
Overview Determination of Genomic Imbalances by Genome-wide Screening Approaches Károly Szuhai Introduction/Methodologies Applications/Results Conclusion Approaches Introduction/Methodologies Chromosome
Diagnosis of Congenital Cardiac Defects Between 11 and 14 Weeks Gestation in High-Risk Patients
 Article Diagnosis of Congenital Cardiac Defects Between 11 and 14 Weeks Gestation in High-Risk Patients Zeev Weiner, MD, Abraham Lorber, MD, Eliezer Shalev, MD Objective. To examine the feasibility of
Article Diagnosis of Congenital Cardiac Defects Between 11 and 14 Weeks Gestation in High-Risk Patients Zeev Weiner, MD, Abraham Lorber, MD, Eliezer Shalev, MD Objective. To examine the feasibility of
Diagnosis of parental balanced reciprocal translocations by trophectoderm biopsy and comprehensive chromosomal screening
 Diagnosis of parental balanced reciprocal translocations by trophectoderm biopsy and comprehensive chromosomal screening Lian Liu, MD Co-Authors: L. W. Sundheimer1, L. Liu2, R. P. Buyalos1,3, G. Hubert1,3,
Diagnosis of parental balanced reciprocal translocations by trophectoderm biopsy and comprehensive chromosomal screening Lian Liu, MD Co-Authors: L. W. Sundheimer1, L. Liu2, R. P. Buyalos1,3, G. Hubert1,3,
Autism shares features with cerebellar syndromes
 NEWS Autism shares features with cerebellar syndromes BY KELLY RAE CHI 3 DECEMBER 2009 1 / 7 2 / 7 Misshapen structures: MRI scans show that, compared with controls (A), the cerebellum of a child with
NEWS Autism shares features with cerebellar syndromes BY KELLY RAE CHI 3 DECEMBER 2009 1 / 7 2 / 7 Misshapen structures: MRI scans show that, compared with controls (A), the cerebellum of a child with
Addressing the challenges of genomic characterization of hematologic malignancies using microarrays
 Addressing the challenges of genomic characterization of hematologic malignancies using microarrays Sarah South, PhD, FACMG Medical Director, ARUP Laboratories Department of Pediatrics and Pathology University
Addressing the challenges of genomic characterization of hematologic malignancies using microarrays Sarah South, PhD, FACMG Medical Director, ARUP Laboratories Department of Pediatrics and Pathology University
Neuropathology Specialty Conference
 Neuropathology Specialty Conference March 22, 2010 Case 2 Rebecca Folkerth, MD Brigham and Women s Hospital Children s Hospital Harvard Medical School Clinical History 18-gestational-week fetus found on
Neuropathology Specialty Conference March 22, 2010 Case 2 Rebecca Folkerth, MD Brigham and Women s Hospital Children s Hospital Harvard Medical School Clinical History 18-gestational-week fetus found on
UNIT IX: GENETIC DISORDERS
 UNIT IX: GENETIC DISORDERS Younas Masih Lecturer New Life College Of Nursing Karachi 3/4/2016 1 Objectives By the end of this session the Learners will be able to, 1. Know the basic terms related genetics
UNIT IX: GENETIC DISORDERS Younas Masih Lecturer New Life College Of Nursing Karachi 3/4/2016 1 Objectives By the end of this session the Learners will be able to, 1. Know the basic terms related genetics
Neonatal Hypoglycemia. Presented By : Kamlah Olaimat 25\7\2010
 Neonatal Hypoglycemia Presented By : Kamlah Olaimat 25\7\2010 Definition The S.T.A.B.L.E. Program defines hypoglycemia as: Glucose delivery or availability is inadequate to meet glucose demand (Karlsen,
Neonatal Hypoglycemia Presented By : Kamlah Olaimat 25\7\2010 Definition The S.T.A.B.L.E. Program defines hypoglycemia as: Glucose delivery or availability is inadequate to meet glucose demand (Karlsen,
increment in the genome to calculate the statistic of read count. Three billions according to the reference
 SUPPLEMENTAL METHODS Genome-wide Window Determination As described in our previous study 1, we use adjustable sliding windows with increment in the genome to calculate the statistic of read count. Three
SUPPLEMENTAL METHODS Genome-wide Window Determination As described in our previous study 1, we use adjustable sliding windows with increment in the genome to calculate the statistic of read count. Three
Current controversies in prenatal diagnosis 3: is conventional chromosome analysis necessary in the post-array CGH era?
 PRENATAL DIAGNOSIS Prenat Diagn 2011; 31: 235 243. Published online 10 February 2011 in Wiley Online Library (wileyonlinelibrary.com).2722 DEBATE Current controversies in prenatal diagnosis 3: is conventional
PRENATAL DIAGNOSIS Prenat Diagn 2011; 31: 235 243. Published online 10 February 2011 in Wiley Online Library (wileyonlinelibrary.com).2722 DEBATE Current controversies in prenatal diagnosis 3: is conventional
Updating penetrance estimates for syndromes with variable phenotypic manifestation. Adele Corrigan June 27th
 Updating penetrance estimates for syndromes with variable phenotypic manifestation Adele Corrigan June 27th Background Array CGH has led to increased identification of copy number variants (CNVs) Our understanding
Updating penetrance estimates for syndromes with variable phenotypic manifestation Adele Corrigan June 27th Background Array CGH has led to increased identification of copy number variants (CNVs) Our understanding
