Low pass sequencing for CNV detection in hemato/onco routine diagnostics?
|
|
|
- Bartholomew Miles
- 5 years ago
- Views:
Transcription
1 Low pass sequencing for CNV detection in hemato/onco routine diagnostics? Groupe Francophone de Cytogénétique Hématologique du 1 er février 2018 Paris
2 Low pass sequencing (LPS) or shallow sequencing Depth of coverage method: infers copy number from the observed sequence depth across the genome and does not require both ends of the molecule to be sequenced Divide the reference genome into bins and count the number of reads into each bin: copy number is inferred from the observed read counts across the genome. QDNA seq pipeline No normal reference necessary (reduces cost by half, samples for which normal reference is not available can be analyzed, avoids measurement noise from the reference sample) Use of fixed sized bins (=key feature of QDNAseq), allows relatively rapid analysis BAM files as input (produced by BWA algorithm) Read depth Reference Genome bin Mapped reads loss gain Bin position
3 Method: QDNA Seq
4 HiSeq run yields Gb of data for 1x50bp 8 lanes per run ( bp per lane) One lane contained 14 to 18 samples Sequence 35 bp per read On average: 46,9 million total reads per sample (on average: human genome has 3,2 billion nucleotides)
5 Output of QDNAseq Read counts per bin Corrected for GC content and sequence mappability Filtered (of problematic genomic regions) Median normalized Log 2 transformed Picking up aberrations: calls starting from 500 à 600kb Mean coverage +/ 0,17 [(PF_READS_ALIGNED * Read length ) / genome length; with genome X +Y hg19: and read length 35 nt]
6 Noise in copy number profiles comes from the statistics of counting as well as from the many steps during the analysis of the sample Noise sources: Sample acquisition and storage/transportation DNA extraction Array: DNA labeling, hybridization NGS: library prep, sequencing Computational processing The variances of all these independent sources of noise are additive Look at the variances of the profiles to investigate their noise characteristics Noise in LPS is dominated by counting statistics for most of the samples, so can be reduced by read depth
7 Compare signal to noise ratio of array versus shallow sequencing
8 Array CGH Leuven Oxford Gene Technology s CytoSure Haematological Cancer + SNP array (8x60k format) spots Targets regions important for CLL, MM, MPN and MDS Backbone: 1 probe every 117kb Average gene resolution: 1 probe every 68 kb LOH regions > 30 Mb Segmentation based normalisation => Dye Swap: every sample is analysed twice
9 CLL: current routine diagnostic methods Karyotype at diagnosis after 3 day culture stimulated with CpG/IL2 FISH after culture LSI ATM (SO) / CEP12 (SG), 11q22/centromere 12 (Vysis) LSI13 (RB1) (SG) / D13S319 (SO), 13q14 (Vysis) LSI TP53 (SO) / CEP17 (SG), 17p13.1/centromere 17 (Vysis) CLL FU: FISH for TP53 if the patient needs (new) treatment
10 CLL: DNA for array CGH and LPS DNA extracted from uncultured whole blood sample (<3ml; EDTA) Chemagen B 4K method Magnetic polyvinyl alcohol beads (M PVA beads) which will bind the DNA after lysis of the blood DNA concentration and purity: measured by DropSense96 (UV VIS absorption analysis) (Beer Lambert law)
11 CLL case 8: karyotype and FISH Caryotype: 48,XY,+12,+19[1]/48,sl,t(8;9)(q24;p12)[9] Anomalies clonales complexes (3). Pronostic moins favorable. Loss of 13q14 not detected in karyotype; but only using karyotyping the t(8;9) is observed!
12 CLL case 8: karyotype and FISH FISH results: Probe Result LSI ATM (SO) / CEP12 (SG), 11q22/centromere 12 (Vysis) gain 12 (51%) LSI13 (RB1) (SG) / D13S319 (SO), 13q14 (Vysis) loss 13q14 (51%) 17p13.1/centromere 17 (Vysis) normal 17p13
13 CLL case 8: array CGH profile, whole genome view Control of the quality parameters: OK DLR (derivative log ratio): < 0,20 Red signal intensity: >300 Green signal intensity: >200 Red background noise: <15 Green background noise: <15 S/N (red/green signial to noise): > 30 FISH, array CGH or LPS: same conclusion: 13q loss (containing DLEU1, DLEU2, DLEU7, MIR15A and MIR16 1 and NOT RB1) and trisomy 12 are recurrent abnormalities in CLL. Trisomy 12 is associated with an intermediate prognosis. Note: chromosome 19 not analysed by FISH
14 CLL case 8: array CGH profile, whole genome view bins of 100 kbp
15 CLL case 8: more noise with array than LPS Dye Swap (2 experiments visualized)
16 CLL case 8: more noise with array than LPS Visualisation without DyeSwap
17 Chromosome 12: array vs LPS Outliers are frequently observed, all over the genome Log2 ratio: 0,31 The trisomy 12 is very clear Clean profile
18 Chromosome 13: array vs LPS error in OGT array design Log2 ratio: 0,37
19 Chromosome 19: array vs LPS Fluctuations like this are common using these arrays, all over the genome Log2 ratio: 0,30
20 Output LPS data in Excel chr start end Size sample AVG_score regcount DupDel ISCN_notation ,6 Mb GC042604_100 0, GAIN arr[hg19]12p13.33q24,33(400, ,000,000)x ,5 Mb GC042604_100 0,49875 LOSS arr[hg19]13q14.2q14.3(50,600,000 51,100,000)x ,4 Mb GC042604_100 0, GAIN arr[hg19]19p13.2q13,43(7,500,000 56,900,000)x3 Visualisation software? Output Array data from CytoSure in pdf
21 With CytoSure analysis/visualisation software for arrays: you can quickly SEE that RB1 is not in the deletion
22 CLL case 5 Probe Result LSI ATM (SO) / CEP12 (SG), 11q22/centromere 12 (Vysis) normal 12, normal 11q22 LSI13 (RB1) (SG) / D13S319 (SO), 13q14 loss 13q14 (11,5%), without (Vysis) loss of RB1 17p13.1/centromere 17 (Vysis) normal 17p13 FISH on peripheral blood after culture (on 10 metaphases and 200 interphase cells): Loss of 13q14 (D13S319), without loss of RB1 locus in 4/10 metaphases [loss of D13S319 on the der(13)t(9;13)] and in 11,5% of the interphase cells. Good prognosis in CLL.
23 CLL case 5 Karyotype: 46,XX,t(9;13)(q13;q12)[4]/46,XX[11]
24 CLL case 5: 13q14 loss not found by array nor LPS Whole genome view Chromosome 13 overview array LPS
25 CLL case 6: aberrations in +/ 20 30% of cells are detected by FISH and karyotyping FISH results: Probe Result normal 12, loss of 11q22 in 7/10 LSI ATM (SO) / CEP12 (SG), 11q22/centromere 12 (Vysis) metaphases and 23% of interphase cells LSI13 (RB1) (SG) / D13S319 (SO), 13q14 (Vysis) loss 13q14 (28%), without loss of RB1 17p13.1/centromere 17 (Vysis) normal 17p13 FISH on peripheral blood after culture (on 10 metaphases and 200 interphase cells): Loss of 11q22 (ATM) in 7/10 metaphases and in 23% of the interphase cells. Loss of 13q14 in 28% of the interphase cells. Worse prognosis in CLL.
26 CLL case 6: aberrations in +/ 20 30% of cells are detected by FISH and karyotyping Karyotype: 46,XY,del(11)(q22)[9]/46,XY[1]
27 CLL case 6: aberrations in +/ 20 30% of cells are NOT always detected by array (unclear, below cut off)??
28 Chromosome 11 CLL case 6: aberrations in +/ 20 30% of cells are NOT always detected by array (unclear, below cut off)
29 CLL case 6: aberrations in +/ 20 30% of cells are NOT always detected by array (unclear, below cut off) Chromosome 11 => zoomed in on 11q14.1q26 33,58 Mb Deletion?? Unclear and below cut off, log2= 0,1222
30 Chromosome 13 CLL case 6: aberrations in +/ 20 30% of cells are NOT always detected by array (unclear, below cut off)
31 CLL case 6: aberrations in +/ 20 30% of cells are NOT always detected by array (unclear, below cut off) Chromosome 13 => zoomed in on 13q13.3q31,1 33,07 Mb Deletion?? Unclear and below cut off ( 0,36); log2 ratio= 0,2001
32 CLL case 6: aberrations in +/ 20 30% of cells are detected by low pass sequencing
33 CLL case 6: aberrations in +/ 20 30% of cells are detected by low pass sequencing
34 CLL case 6: aberrations in +/ 20 30% of cells are detected by low pass sequencing
35 Multiple Myeloma (MM): current routine diagnostic method Direct FISH on cytospin: LSI TP53 (SO) / CEP17 (SG), 17p13.1/centromere 17 (Vysis) LSI IGH DC BA, 14q32 (Vysis) CEP9 (SO), centromere 9 (Vysis) CKS1B (TR) / CDKN2C (SG) [1q21.3/1p32.3, Cytocell] / Chromosome 1 Control Probe [1p11.1 1q12, Empire Genomics] further work up of IGH if split in IGH (IGH/CCND1 IGH/FGFR3 IGH/MAF MYC BA IGH/MAFB IGH/MYC/CEP8) FU MM: LSI TP53 (SO) / CEP17 (SG), 17p13.1/centromere 17 (Vysis) LSI MYC DC BA, 8q24.2 (Vysis) CKS1B (TR) / CDKN2C (SG) [1q21.3/1p32.3, Cytocell] / Chromosome 1 Control Probe [1p11.1 1q12, Empire Genomics] probes most important aberration at time of diagnosis
36 Low pass sequencing (cfdna and DNA CD138+ cells) for the detection of copy number aberrations in Multiple Myeloma 31 MM cases peripheral blood (strecktube) bone marrow aspirates extraction of ccfdna cells purified by CD138 selection (EasySep) plasma cells Sequencing 18 cases Work in progress: last cases using new libraryprepmethod Truseqnano Cytospin DNA extraction FISH 31 cases array CGH 15 cases More cases can be done if necessary low pass sequencing 20 cases 36
37 Analysis of low pass sequencing data of cfdna: GR and z score Genomic Representation (GR) Profiling 1. filtering and aligning of reads constitutional pregnancy/malignancy homozygous deletion heterozygous deletion gain homozygous deletion heterozygous deletion gain 2. GR and z score number of reads in a chromosome/partition/bin GR total of autosomal reads z score GR measured GR SD normal chr20 chr21
38 Analysis of low pass sequencing data of cfdna: GR and z score
39 DNA concentration: 500 ng standard, but 50 ng or lower is possible LPS op Hiseq4000 Paired end DNA input: 500 ng Run 2 GC LPS op Hiseq4000 Single end DNA input: 500 ng Run 3 GC LPS op Hiseq4000 Single end DNA input: 100 ng Run 3 GC LPS op Hiseq4000 Single end DNA input: 50 ng Run 3 GC Result FISH: Trisomy 1q (zonder trisomie van centromeer) in 12% of the interphase cells ~ close to cut off. Balanced t(4;14)(p16;q32) in 98% of the interphase cells
40 FISH Matrix: bone marrow plasma cells Per sonde werden 48 tot 100 interfasekernen onderzocht. Dit onderzoek toont: trisomie 4p16 in 53% van de kernen. trisomie 6p21 in 31 43% van de kernen. tri/tetrasomie 9 in 29% van de kernen. trisomie 11q13 in 45% van de kernen. trisomie 20q12 in 32 43% van de kernen. een afwijking van de regio 14q32: partieel verlies van één VH signaal in 37% van de kernen. Bijkomend onderzoek met specifieke probes (IgH/CCND1, FGFR3, CMAF, CCND3, MAFB) toont noch t(11;14), t(4;14), t(14;16), t(6;14), t(14;20). de andere regio's lijken normaal. (chromosoom 8 is normaal) => Klonale afwijkingen. Hyperdiploïdie, prognostisch gunstig (standard risico) bij MM. Low pass sequencing Matrix: cfdna C1D1 (22/03/2016) MM case 2 Plasma cells: 13,3 % Hyperdiploid case: both array and LPS are NOT correct! Karyotype bone marrow after 5d culture: Array CGH Matrix: bone marrow plasma cells Low pass sequencing Matrix: bone marrow plasma cells 46,XY[10] 40
41 FISH (Matrix: bone marrow plasma cells) Per sonde werden 100 interfasekernen onderzocht. Dit onderzoek toont: tri/tetrasomie 9 in 89% van de kernen. trisomie 11q13 in 88% van de kernen. een afwijking van de regio 14q32: één gesplitst CH/VH signaal in 94% van de kernen. Bijkomend onderzoek met specifieke probes (IgH/CCND1, FGFR3, CMAF, CCND3, MAFB, CMYC) toont een niet gebalanceerde t(8;14)(q24.3;q32) in 100% van de kernen. de andere regio's lijken normaal. => Klonale afwijkingen, o.a. t(8;14) (Burkitt translocatie). CMYC translocaties zijn laattijdige of "secundaire" afwijkingen in MM (prognostisch eerder ongunstig). Low pass sequencing Matrix: cfdna C1D1(29/07/2016) MM17 Plasma cell percentage : 67 % (28/07/2016) Array CGH Matrix: bone marrow plasma cells (28/07/2016) Low pass sequencing Matrix: bone marrow plasma cells (28/07/2016) 41
42 MM17 Plasma cell percentage : 67 % (28/07/2016) Profile cfdna normalises rapidly during treatment Low pass sequencing Matrix: cfdna C1D1 (29/07/2016) cfdna C2D1 (9/09/2016) M piek in gammafractie (bloed): 28/06/2016: 47,4 g/ L M piek in gammafractie (bloed): 9/09/2016: 11,8 g/ L Laagste Z score C1D1: 7,7 hoogste Z score C2D1: 2,9 C3D1 en C4D1 werden ook afgenomen, maar profiel al genormaliseerd daarom niet meer gesequenced. 42
43 FISH Matrix: bone marrow plasma cells Per sonde werden 100 interfasekernen onderzocht. Dit onderzoek toont: een afwijking van de regio 14q32: één gesplitst CH/VH signaal in 82% van de kernen. Bijkomend onderzoek met specifieke probes (IgH/CCND1) toont een gebalanceerde t(11;14)(q13;q32) in 74% van de kernen. de andere regio's lijken normaal. Klonale afwijking. t(11;14), prognostisch gunstig (standard risico) bij MM Low pass sequencing Matrix: cfdna voor therapie (24/03/2017) MM4 Plasma cell percentage (24/03/2016): 43 % Karyotype Matrix: bone marrow 48,XY,add(6)(q14),+add(6),,add(9)(p13),t(11;14)(q13;q32),+18,+mar[6] Array CGH Matrix: bone marrow (niet opgezuiverd maar geprikt in plasmacytoom) Cfr karyotype Low pass sequencing Matrix: bone marrow (niet opgezuiverd maar geprikt in plasmacytoom) 43
44 MM4 Plasma cell percentage (24/03/2016): 43 % Extra aberration? Voor C1D1 (24/03/2017) C1D1 (22/04/2016) C2D1 (20/05/2016) C3D1 (17/06/2016) C5D1 (9/09/2016) C9D1 (30/12/2016) M piek in gammafractie (bloed): 24/03/2016: 39,2 g/ L M piek in gammafractie (bloed): 20/05/2016: 30,1 g/ L M piek in gammafractie (bloed): 17/06/2016: 26,3 g/ L M piek in gammafractie (bloed): 9/09/2016: 20,3 g/ L M piek in gammafractie (bloed): 9/09/2016: 18,2 g/ L CcfDNA sequencing profiles before and after treatment of MM patient: no respons to treatment. 44
45 MDS: several examples with +8: trisomie 8 is easily detected, even if present in low percentage of cells MDS17 GC clonal B cell population, pancytopenia progressive: MDS, NHL. MDS TYPE RCMD MDS23 GC MDS, refractory anaemia MDS31 GC RAEB 1 follow up MDS42 GC Pancytopenia, macrocytosis : MDS, MDS RAEB(2)? LPS: +8, Y IPSS: Intermediate outcome IPSS R: 2 intermediate prognosis LPS: +8 IPSS: Intermediate outcome IPSS R: 2 intermediate prognosis LPS: +8 IPSS: Intermediate outcome IPSS R: 2 intermediate prognosis LPS: +8 IPSS: Intermediate outcome IPSS R: 2 intermediate prognosis Karyotype: IPSS: IPSS R: 47,XY,+8[4]/45,X, Y[6] Intermediate outcome 2 intermediate prognosis Karyotype: 47,XX,+8[8]/46,XX [2] IPSS: Intermediate outcome IPSS R: 2 intermediate prognosis Karyotype: IPSS: IPSS R: 47,XY,+8[10] Intermediate outcome 2 intermediate prognosis Karyotype: IPSS: IPSS R: 47,XY,+8[2]/46,XY[8] Intermediate outcome 2 intermediate prognosis
46 MDS: several examples with 7 in karyotype MDS21 Hypocellular marrow, dysplasia in the red lineage; MDS? Karyotype: 46,XY,del(7)(q21q35)[3]/47,XY,+14[2]/46,XY[5] IPSS R: 3 poor prognosis? All other chromosomes have a flat profile trisomy 14? LPS: IPSS R: del(7)(q21q35),+14? 3 poor prognosis
47 MDS: several examples with 7 in karyotype MDS44 MDS RCMD, Female 60y, status chimerism after transplantation with male donor A quoi pensez vous??
48 MDS: several examples with 7 in karyotype MDS44; MDS RCMD, Female 60y, status chimerism after transplantation with male donor Karyotype: IPSS R: 46,XX, 7,+r[4]/46,XX[6]//46,XY[1] 3 poorprognosis Conclusion karyotype: 90% receptor hematopoiesis. Pseudodiploïd karyotype with (partial?) loss of chromosome 7. Classic aberration in MDS with a bad prognosis (IPSS R cytogenetic score 3).
49 MDS: several examples with 7 in karyotype MDS44; MDS RCMD, Female 60y, status chimerism after transplantation with male donor Karyotype: IPSS R: 46,XX, 7,+r[4]/46,XX[6]//46,XY[1] 3 poorprognosis Conclusion karyotype: 90% receptorhematopoiesis. Pseudodiploïd karyotype with (partial?) loss of chromosome 7. Classic aberration in MDS with a bad prognosis (IPSS R cytogenetic score 3). Ring of chromosome 7 Other chromsomes: rather flat profile.
50 Serial dilution: samples +8/ 7 in karyotype Karyotype MDS31: 47,XY,+8[10] Karyotype MDS32: 45,XY, 7[10] 100% trisomy 8 GC % trisomy 8/ 5% monosomy 7 GC % trisomy 8/ 10% monosomy 7 GC % trisomy 8/ 20% monosomy 7 GC % trisomy 8/ 80% monosomy 7 GC % trisomy 8/ 90% monosomy 7 GC % trisomy/ 95% monosomy 7 GC % monosomy 7 GC Low pass sequencing is very sensitive and can detect aberrations in low % of cells: 10% in normal background is reliably detected
51 Cost Plasmocyte purification (EasySep II): 34,25 euro/sample DNA extraction: 25 euro/sample Library Prep (KAPA HTP; automated): 115 euro (this cost can be reduced, KAPA Hyper prep) Sequencing cost: 1345/lane (75 à 96 euro/sample) Cost array (x2: Dye Swap): 285 euro Array is more labor intensive
52 Conclusions Low pass sequencing costs less, shows less outliers and yields a better signal to noise ratio than array Detection of CN LOH with LPS: ongoing
Haematology Probes for Multiple Myeloma
 Haematology Probes for Multiple Myeloma MULTIPLE MYELOMA Multiple myeloma (MM) is a plasma cell neoplasm, characterised by the accumulation of clonal plasma cells in the bone marrow and by very complex
Haematology Probes for Multiple Myeloma MULTIPLE MYELOMA Multiple myeloma (MM) is a plasma cell neoplasm, characterised by the accumulation of clonal plasma cells in the bone marrow and by very complex
Outline. Chromosomal analysis FISH. Chromosomal abnormalities in cancer. Clinical application of cytogenetics. Procedure Nomenclature
 Outline Chromosomal analysis Procedure Nomenclature FISH Procedure Probes Multicolor-FISH CGH Chromosomal abnormalities in cancer CML, MPD, MDS, AML, ALL, CLL, myeloma, lymphoma Clinical application of
Outline Chromosomal analysis Procedure Nomenclature FISH Procedure Probes Multicolor-FISH CGH Chromosomal abnormalities in cancer CML, MPD, MDS, AML, ALL, CLL, myeloma, lymphoma Clinical application of
The role of cytogenomics in the diagnostic work-up of Chronic Lymphocytic Leukaemia
 The role of cytogenomics in the diagnostic work-up of Chronic Lymphocytic Leukaemia Adrian Zordan, Meaghan Wall, Ruth MacKinnon, Pina D Achille & Lynda Campbell Victorian Cancer Cytogenetics Service (VCCS)
The role of cytogenomics in the diagnostic work-up of Chronic Lymphocytic Leukaemia Adrian Zordan, Meaghan Wall, Ruth MacKinnon, Pina D Achille & Lynda Campbell Victorian Cancer Cytogenetics Service (VCCS)
Role of FISH in Hematological Cancers
 Role of FISH in Hematological Cancers Thomas S.K. Wan PhD,FRCPath,FFSc(RCPA) Honorary Professor, Department of Pathology & Clinical Biochemistry, Queen Mary Hospital, University of Hong Kong. e-mail: wantsk@hku.hk
Role of FISH in Hematological Cancers Thomas S.K. Wan PhD,FRCPath,FFSc(RCPA) Honorary Professor, Department of Pathology & Clinical Biochemistry, Queen Mary Hospital, University of Hong Kong. e-mail: wantsk@hku.hk
Identification of Prognostically Relevant Chromosomal Abnormalities in Routine Diagnostics of Multiple Myeloma Using Genomic Profiling
 Identification of Prognostically Relevant Chromosomal Abnormalities in Routine Diagnostics of Multiple Myeloma Using Genomic Profiling EIGIL KJELDSEN Haemodiagnostic Laboratory, Cancer Cytogenetics Section,
Identification of Prognostically Relevant Chromosomal Abnormalities in Routine Diagnostics of Multiple Myeloma Using Genomic Profiling EIGIL KJELDSEN Haemodiagnostic Laboratory, Cancer Cytogenetics Section,
DNA-seq Bioinformatics Analysis: Copy Number Variation
 DNA-seq Bioinformatics Analysis: Copy Number Variation Elodie Girard elodie.girard@curie.fr U900 institut Curie, INSERM, Mines ParisTech, PSL Research University Paris, France NGS Applications 5C HiC DNA-seq
DNA-seq Bioinformatics Analysis: Copy Number Variation Elodie Girard elodie.girard@curie.fr U900 institut Curie, INSERM, Mines ParisTech, PSL Research University Paris, France NGS Applications 5C HiC DNA-seq
Cytogenetics 101: Clinical Research and Molecular Genetic Technologies
 Cytogenetics 101: Clinical Research and Molecular Genetic Technologies Topics for Today s Presentation 1 Classical vs Molecular Cytogenetics 2 What acgh? 3 What is FISH? 4 What is NGS? 5 How can these
Cytogenetics 101: Clinical Research and Molecular Genetic Technologies Topics for Today s Presentation 1 Classical vs Molecular Cytogenetics 2 What acgh? 3 What is FISH? 4 What is NGS? 5 How can these
Integration of microarray analysis into the clinical diagnosis of hematological malignancies: How much can we improve cytogenetic testing?
 /, Vol. 6, No. 22 Integration of microarray analysis into the clinical diagnosis of hematological malignancies: How much can we improve cytogenetic testing? Jess F. Peterson 1,2,6, Nidhi Aggarwal 3, Clayton
/, Vol. 6, No. 22 Integration of microarray analysis into the clinical diagnosis of hematological malignancies: How much can we improve cytogenetic testing? Jess F. Peterson 1,2,6, Nidhi Aggarwal 3, Clayton
CLL Complete SM Report
 Reported: 02/01/2012 Σ CGI ID No:5 Client:r Client Address: CLINICAL DATA: Lymphoma No CBC results provided. CLL Complete SM Report FINAL DIAGNOSIS: CD19+ B cell lymphoma, ZAP-70 + (44%), with borderline
Reported: 02/01/2012 Σ CGI ID No:5 Client:r Client Address: CLINICAL DATA: Lymphoma No CBC results provided. CLL Complete SM Report FINAL DIAGNOSIS: CD19+ B cell lymphoma, ZAP-70 + (44%), with borderline
Addressing the challenges of genomic characterization of hematologic malignancies using microarrays
 Addressing the challenges of genomic characterization of hematologic malignancies using microarrays Sarah South, PhD, FACMG Medical Director, ARUP Laboratories Department of Pediatrics and Pathology University
Addressing the challenges of genomic characterization of hematologic malignancies using microarrays Sarah South, PhD, FACMG Medical Director, ARUP Laboratories Department of Pediatrics and Pathology University
Volume 7, Issue 1 January 2012
 The Hong Kong College of Pathologists, Incorporated in Hong Kong with Limited Liability Volume 7, Issue 1 January 2012 Editorial note: Chronic lymphocytic leukaemia (CLL) is the commonest chronic lymphoproliferative
The Hong Kong College of Pathologists, Incorporated in Hong Kong with Limited Liability Volume 7, Issue 1 January 2012 Editorial note: Chronic lymphocytic leukaemia (CLL) is the commonest chronic lymphoproliferative
Chromosomal Aberrations
 Chromosomal Aberrations Chromosomal Aberrations Abnormalities of chromosomes may be either numerical or structural and may involve one or more autosomes, sex chromosomes, or both simultaneously. Numerical
Chromosomal Aberrations Chromosomal Aberrations Abnormalities of chromosomes may be either numerical or structural and may involve one or more autosomes, sex chromosomes, or both simultaneously. Numerical
Approach to Mental Retardation and Developmental Delay. SR Ghaffari MSc MD PhD
 Approach to Mental Retardation and Developmental Delay SR Ghaffari MSc MD PhD Introduction Objectives Definition of MR and DD Classification Epidemiology (prevalence, recurrence risk, ) Etiology Importance
Approach to Mental Retardation and Developmental Delay SR Ghaffari MSc MD PhD Introduction Objectives Definition of MR and DD Classification Epidemiology (prevalence, recurrence risk, ) Etiology Importance
Molecular Markers. Marcie Riches, MD, MS Associate Professor University of North Carolina Scientific Director, Infection and Immune Reconstitution WC
 Molecular Markers Marcie Riches, MD, MS Associate Professor University of North Carolina Scientific Director, Infection and Immune Reconstitution WC Overview Testing methods Rationale for molecular testing
Molecular Markers Marcie Riches, MD, MS Associate Professor University of North Carolina Scientific Director, Infection and Immune Reconstitution WC Overview Testing methods Rationale for molecular testing
Reporting cytogenetics Can it make sense? Daniel Weisdorf MD University of Minnesota
 Reporting cytogenetics Can it make sense? Daniel Weisdorf MD University of Minnesota Reporting cytogenetics What is it? Terminology Clinical value What details are important Diagnostic Tools for Leukemia
Reporting cytogenetics Can it make sense? Daniel Weisdorf MD University of Minnesota Reporting cytogenetics What is it? Terminology Clinical value What details are important Diagnostic Tools for Leukemia
Introduction to Cytogenetics
 Introduction to Cytogenetics Catherine McCarthy Pathology Qld Cytogenetic abnormalities constitutional acquired: clonal: related or unrelated non-clonal Investigating constitutional abnormalities peripheral
Introduction to Cytogenetics Catherine McCarthy Pathology Qld Cytogenetic abnormalities constitutional acquired: clonal: related or unrelated non-clonal Investigating constitutional abnormalities peripheral
Monoallelic and biallelic deletions of 13q14 in a group of CLL/SLL patients investigated by CGH Haematological Cancer and SNP array (8x60K)
 Grygalewicz et al. Molecular Cytogenetics (2016) 9:1 DOI 10.1186/s13039-015-0212-x RESEARCH Open Access Monoallelic and biallelic deletions of 13q14 in a group of CLL/SLL patients investigated by CGH Haematological
Grygalewicz et al. Molecular Cytogenetics (2016) 9:1 DOI 10.1186/s13039-015-0212-x RESEARCH Open Access Monoallelic and biallelic deletions of 13q14 in a group of CLL/SLL patients investigated by CGH Haematological
Detection of aneuploidy in a single cell using the Ion ReproSeq PGS View Kit
 APPLICATION NOTE Ion PGM System Detection of aneuploidy in a single cell using the Ion ReproSeq PGS View Kit Key findings The Ion PGM System, in concert with the Ion ReproSeq PGS View Kit and Ion Reporter
APPLICATION NOTE Ion PGM System Detection of aneuploidy in a single cell using the Ion ReproSeq PGS View Kit Key findings The Ion PGM System, in concert with the Ion ReproSeq PGS View Kit and Ion Reporter
Implementation of the DDD/ClinGen OGT (CytoSure v3) Microarray
 Implementation of the DDD/ClinGen OGT (CytoSure v3) Microarray OGT UGM Birmingham 08/09/2016 Dom McMullan Birmingham Women's NHS Trust WM chromosomal microarray (CMA) testing Population of ~6 million (10%)
Implementation of the DDD/ClinGen OGT (CytoSure v3) Microarray OGT UGM Birmingham 08/09/2016 Dom McMullan Birmingham Women's NHS Trust WM chromosomal microarray (CMA) testing Population of ~6 million (10%)
CYTOGENETICS INTRODUCTION SPECIAL INSTRUCTIONS ON SAMPLE COLLECTION AND HANDLING
 INTRODUCTION The Cytogenetics Laboratory offers a comprehensive array of chromosome investigations for cancers, constitutional abnormalities, and prenatal and postnatal diagnosis. Analyses are performed
INTRODUCTION The Cytogenetics Laboratory offers a comprehensive array of chromosome investigations for cancers, constitutional abnormalities, and prenatal and postnatal diagnosis. Analyses are performed
Clinical Interpretation of Cancer Genomes
 IGENZ Ltd, Auckland, New Zealand Clinical Interpretation of Cancer Genomes Dr Amanda Dixon-McIver www.igenz.co.nz 1992 Slovenia and Croatia gain independence USA and Russia declare the Cold War over Steffi
IGENZ Ltd, Auckland, New Zealand Clinical Interpretation of Cancer Genomes Dr Amanda Dixon-McIver www.igenz.co.nz 1992 Slovenia and Croatia gain independence USA and Russia declare the Cold War over Steffi
Chromosome Abnormalities
 Chromosome Abnormalities Chromosomal abnormalities vs. molecular mutations Simply a matter of size Chromosomal abnormalities are big errors Two types of abnormalities 1. Constitutional problem present
Chromosome Abnormalities Chromosomal abnormalities vs. molecular mutations Simply a matter of size Chromosomal abnormalities are big errors Two types of abnormalities 1. Constitutional problem present
CYTOGENETICS Dr. Mary Ann Perle
 CYTOGENETICS Dr. Mary Ann Perle I) Mitosis and metaphase chromosomes A) Chromosomes are most fully condensed and clearly distinguishable during mitosis. B) Mitosis (M phase) takes 1 to 2 hrs and is divided
CYTOGENETICS Dr. Mary Ann Perle I) Mitosis and metaphase chromosomes A) Chromosomes are most fully condensed and clearly distinguishable during mitosis. B) Mitosis (M phase) takes 1 to 2 hrs and is divided
Genomic complexity and arrays in CLL. Gian Matteo Rigolin, MD, PhD St. Anna University Hospital Ferrara, Italy
 Genomic complexity and arrays in CLL Gian Matteo Rigolin, MD, PhD St. Anna University Hospital Ferrara, Italy Clinical relevance of genomic complexity (GC) in CLL GC has been identified as a critical negative
Genomic complexity and arrays in CLL Gian Matteo Rigolin, MD, PhD St. Anna University Hospital Ferrara, Italy Clinical relevance of genomic complexity (GC) in CLL GC has been identified as a critical negative
CNV Detection and Interpretation in Genomic Data
 CNV Detection and Interpretation in Genomic Data Benjamin W. Darbro, M.D., Ph.D. Assistant Professor of Pediatrics Director of the Shivanand R. Patil Cytogenetics and Molecular Laboratory Overview What
CNV Detection and Interpretation in Genomic Data Benjamin W. Darbro, M.D., Ph.D. Assistant Professor of Pediatrics Director of the Shivanand R. Patil Cytogenetics and Molecular Laboratory Overview What
Your single-source laboratory solution. FISH Probe Library
 Your single-source laboratory solution. FISH Probe Library Alphabetical List by Probe with Clinical Indication Probe Clinical Indication 1p36 (TP73)/19q13 (GLTSCR) 1q21 (CKS1B) 5q (CSF1R/RPS14) 7q (MDFIC)
Your single-source laboratory solution. FISH Probe Library Alphabetical List by Probe with Clinical Indication Probe Clinical Indication 1p36 (TP73)/19q13 (GLTSCR) 1q21 (CKS1B) 5q (CSF1R/RPS14) 7q (MDFIC)
Challenges of CGH array testing in children with developmental delay. Dr Sally Davies 17 th September 2014
 Challenges of CGH array testing in children with developmental delay Dr Sally Davies 17 th September 2014 CGH array What is CGH array? Understanding the test Benefits Results to expect Consent issues Ethical
Challenges of CGH array testing in children with developmental delay Dr Sally Davies 17 th September 2014 CGH array What is CGH array? Understanding the test Benefits Results to expect Consent issues Ethical
UTILIZATION OF A SNP MICROARRAY FOR CHRONIC LYMPHOCYTIC LEUKEMIA: EFFICACY, INFORMATIVE FINDINGS AND PROGNOSTIC CAPABILITIES
 UTILIZATION OF A SNP MICROARRAY FOR CHRONIC LYMPHOCYTIC LEUKEMIA: EFFICACY, INFORMATIVE FINDINGS AND PROGNOSTIC CAPABILITIES S Schwartz, Z Hosseini, S Schepis, P Papenhausen Laboratory Corporation of America
UTILIZATION OF A SNP MICROARRAY FOR CHRONIC LYMPHOCYTIC LEUKEMIA: EFFICACY, INFORMATIVE FINDINGS AND PROGNOSTIC CAPABILITIES S Schwartz, Z Hosseini, S Schepis, P Papenhausen Laboratory Corporation of America
AVENIO ctdna Analysis Kits The complete NGS liquid biopsy solution EMPOWER YOUR LAB
 Analysis Kits The complete NGS liquid biopsy solution EMPOWER YOUR LAB Analysis Kits Next-generation performance in liquid biopsies 2 Accelerating clinical research From liquid biopsy to next-generation
Analysis Kits The complete NGS liquid biopsy solution EMPOWER YOUR LAB Analysis Kits Next-generation performance in liquid biopsies 2 Accelerating clinical research From liquid biopsy to next-generation
Clinical utility of FISH analysis in addition to G-banded karyotype in hematologic malignancies and proposal of a practical approach
 VOLUME 45 ㆍ NUMBER 3 ㆍ September 2010 THE KOREAN JOURNAL OF HEMATOLOGY ORIGINAL ARTICLE Clinical utility of FISH analysis in addition to G-banded karyotype in hematologic malignancies and proposal of a
VOLUME 45 ㆍ NUMBER 3 ㆍ September 2010 THE KOREAN JOURNAL OF HEMATOLOGY ORIGINAL ARTICLE Clinical utility of FISH analysis in addition to G-banded karyotype in hematologic malignancies and proposal of a
AVENIO family of NGS oncology assays ctdna and Tumor Tissue Analysis Kits
 AVENIO family of NGS oncology assays ctdna and Tumor Tissue Analysis Kits Accelerating clinical research Next-generation sequencing (NGS) has the ability to interrogate many different genes and detect
AVENIO family of NGS oncology assays ctdna and Tumor Tissue Analysis Kits Accelerating clinical research Next-generation sequencing (NGS) has the ability to interrogate many different genes and detect
Understanding the Human Karyotype Colleen Jackson Cook, Ph.D.
 Understanding the Human Karyotype Colleen Jackson Cook, Ph.D. SUPPLEMENTAL READING Nussbaum, RL, McInnes, RR, and Willard HF (2007) Thompson and Thompson Genetics in Medicine, 7th edition. Saunders: Philadelphia.
Understanding the Human Karyotype Colleen Jackson Cook, Ph.D. SUPPLEMENTAL READING Nussbaum, RL, McInnes, RR, and Willard HF (2007) Thompson and Thompson Genetics in Medicine, 7th edition. Saunders: Philadelphia.
Using the Bravo Liquid-Handling System for Next Generation Sequencing Sample Prep
 Using the Bravo Liquid-Handling System for Next Generation Sequencing Sample Prep Tom Walsh, PhD Division of Medical Genetics University of Washington Next generation sequencing Sanger sequencing gold
Using the Bravo Liquid-Handling System for Next Generation Sequencing Sample Prep Tom Walsh, PhD Division of Medical Genetics University of Washington Next generation sequencing Sanger sequencing gold
Global variation in copy number in the human genome
 Global variation in copy number in the human genome Redon et. al. Nature 444:444-454 (2006) 12.03.2007 Tarmo Puurand Study 270 individuals (HapMap collection) Affymetrix 500K Whole Genome TilePath (WGTP)
Global variation in copy number in the human genome Redon et. al. Nature 444:444-454 (2006) 12.03.2007 Tarmo Puurand Study 270 individuals (HapMap collection) Affymetrix 500K Whole Genome TilePath (WGTP)
P53 Gene Deletion Detected By Fluorescence In Situ Hybridization is an Adverse
 Blood First Edition Paper, prepublished online August 31, 2004; DOI 10.1182/blood-2004-04-1363 P53 Gene Deletion Detected By Fluorescence In Situ Hybridization is an Adverse Prognostic Factor for Patients
Blood First Edition Paper, prepublished online August 31, 2004; DOI 10.1182/blood-2004-04-1363 P53 Gene Deletion Detected By Fluorescence In Situ Hybridization is an Adverse Prognostic Factor for Patients
Clinical Genomics. Ina E. Amarillo, PhD FACMGG
 Clinical Genomics Ina E. Amarillo, PhD FACMGG Associate Medical Director, Cytogenetics Lab (CaTG), Lab and Genomic Medicine Assistant Professor, Pathology and Immunology Outline Clinical Genomics Testing
Clinical Genomics Ina E. Amarillo, PhD FACMGG Associate Medical Director, Cytogenetics Lab (CaTG), Lab and Genomic Medicine Assistant Professor, Pathology and Immunology Outline Clinical Genomics Testing
Applications of Chromosomal Microarray Analysis (CMA) in pre- and postnatal Diagnostic: advantages, limitations and concerns
 Applications of Chromosomal Microarray Analysis (CMA) in pre- and postnatal Diagnostic: advantages, limitations and concerns جواد کریمزاد حق PhD of Medical Genetics آزمايشگاه پاتوبيولوژي و ژنتيك پارسه
Applications of Chromosomal Microarray Analysis (CMA) in pre- and postnatal Diagnostic: advantages, limitations and concerns جواد کریمزاد حق PhD of Medical Genetics آزمايشگاه پاتوبيولوژي و ژنتيك پارسه
Chapter 1 : Genetics 101
 Chapter 1 : Genetics 101 Understanding the underlying concepts of human genetics and the role of genes, behavior, and the environment will be important to appropriately collecting and applying genetic
Chapter 1 : Genetics 101 Understanding the underlying concepts of human genetics and the role of genes, behavior, and the environment will be important to appropriately collecting and applying genetic
Application of Whole Genome Microarrays in Cancer: You should be doing this test!!
 Application of Whole Genome Microarrays in Cancer: You should be doing this test!! Daynna Wolff, Ph.D. Director, Cytogenetics and Genomics Disclosures Clinical Laboratory Director and Employee, Medical
Application of Whole Genome Microarrays in Cancer: You should be doing this test!! Daynna Wolff, Ph.D. Director, Cytogenetics and Genomics Disclosures Clinical Laboratory Director and Employee, Medical
Fluorescent in situ hybridization studies in multiple myeloma
 Fluorescent in situ hybridization studies in multiple myeloma Ozge Ozalp Yuregir 1, Feride Iffet Sahin 1, Zerrin Yilmaz 1, Ebru Kizilkilic 2, Sema Karakus 2 and Hakan Ozdogu 2 1 Department of Medical Genetics
Fluorescent in situ hybridization studies in multiple myeloma Ozge Ozalp Yuregir 1, Feride Iffet Sahin 1, Zerrin Yilmaz 1, Ebru Kizilkilic 2, Sema Karakus 2 and Hakan Ozdogu 2 1 Department of Medical Genetics
Molecular Genetics of Paediatric Tumours. Gino Somers MBBS, BMedSci, PhD, FRCPA Pathologist-in-Chief Hospital for Sick Children, Toronto, ON, CANADA
 Molecular Genetics of Paediatric Tumours Gino Somers MBBS, BMedSci, PhD, FRCPA Pathologist-in-Chief Hospital for Sick Children, Toronto, ON, CANADA Financial Disclosure NanoString - conference costs for
Molecular Genetics of Paediatric Tumours Gino Somers MBBS, BMedSci, PhD, FRCPA Pathologist-in-Chief Hospital for Sick Children, Toronto, ON, CANADA Financial Disclosure NanoString - conference costs for
Oncology Genetics: Cytogenetics and FISH 17/09/2014
 Oncology Genetics: Cytogenetics and FISH 17/09/2014 Chris Wragg Head of Oncology Genomics, BGL BGL Bristol Genetics Laboratory (BGL) CPA accredited Genetics laboratory serving a core population of 4-5million
Oncology Genetics: Cytogenetics and FISH 17/09/2014 Chris Wragg Head of Oncology Genomics, BGL BGL Bristol Genetics Laboratory (BGL) CPA accredited Genetics laboratory serving a core population of 4-5million
MYELODYSPLASTIC SYNDROMES: A diagnosis often missed
 MYELODYSPLASTIC SYNDROMES: A diagnosis often missed D R. EMMA W YPKEMA C O N S U LTA N T H A E M AT O L O G I S T L A N C E T L A B O R AT O R I E S THE MYELODYSPLASTIC SYNDROMES DEFINITION The Myelodysplastic
MYELODYSPLASTIC SYNDROMES: A diagnosis often missed D R. EMMA W YPKEMA C O N S U LTA N T H A E M AT O L O G I S T L A N C E T L A B O R AT O R I E S THE MYELODYSPLASTIC SYNDROMES DEFINITION The Myelodysplastic
Myeloma Genetics what do we know and where are we going?
 in partnership with Myeloma Genetics what do we know and where are we going? Dr Brian Walker Thames Valley Cancer Network 14 th September 2015 Making the discoveries that defeat cancer Myeloma Genome:
in partnership with Myeloma Genetics what do we know and where are we going? Dr Brian Walker Thames Valley Cancer Network 14 th September 2015 Making the discoveries that defeat cancer Myeloma Genome:
Molecular Pathogenesis of Multiple Myeloma:
 Molecular Pathogenesis of Multiple Myeloma: Ig translocations hyperdiploid vs non-hyperdiploid CYCLIN D dysregulation other oncogenic events Michael Kuehl MM: post-germinal center tumor of long-lived BM
Molecular Pathogenesis of Multiple Myeloma: Ig translocations hyperdiploid vs non-hyperdiploid CYCLIN D dysregulation other oncogenic events Michael Kuehl MM: post-germinal center tumor of long-lived BM
Prenatal Diagnosis: Are There Microarrays in Your Future?
 Financial Disclosure UCSF Antepartum Intrapartum Management Course June 8 I have no financial relationship with any aspect of private industry Prenatal Diagnosis: Are There Microarrays in Your Future?
Financial Disclosure UCSF Antepartum Intrapartum Management Course June 8 I have no financial relationship with any aspect of private industry Prenatal Diagnosis: Are There Microarrays in Your Future?
Determination of Genomic Imbalances by Genome-wide Screening Approaches
 Overview Determination of Genomic Imbalances by Genome-wide Screening Approaches Károly Szuhai Introduction/Methodologies Applications/Results Conclusion Approaches Introduction/Methodologies Chromosome
Overview Determination of Genomic Imbalances by Genome-wide Screening Approaches Károly Szuhai Introduction/Methodologies Applications/Results Conclusion Approaches Introduction/Methodologies Chromosome
BHS training course. Laboratory Hematology Cytogenetics. Lucienne Michaux. Centrum voor Menselijke Erfelijkheid, UZLeuven
 BHS training course Laboratory Hematology Cytogenetics Lucienne Michaux Centrum voor Menselijke Erfelijkheid, UZLeuven 18/11/2017 Organization of the Lecture Definition and principles Tools Applications
BHS training course Laboratory Hematology Cytogenetics Lucienne Michaux Centrum voor Menselijke Erfelijkheid, UZLeuven 18/11/2017 Organization of the Lecture Definition and principles Tools Applications
Cytogenetic analyses in malignant hematological disorders
 Cytogenetic analyses in malignant hematological disorders general concepts Lucienne Michaux Lessenreeks 21/11/2017 Plan Definition History Pathophysiology of malignant hematological disorders Techniques
Cytogenetic analyses in malignant hematological disorders general concepts Lucienne Michaux Lessenreeks 21/11/2017 Plan Definition History Pathophysiology of malignant hematological disorders Techniques
Comprehensive Chromosome Screening Is NextGen Likely to be the Final Best Platform and What are its Advantages and Quirks?
 Comprehensive Chromosome Screening Is NextGen Likely to be the Final Best Platform and What are its Advantages and Quirks? Embryo 1 Embryo 2 combine samples for a single sequencing chip Barcode 1 CTAAGGTAAC
Comprehensive Chromosome Screening Is NextGen Likely to be the Final Best Platform and What are its Advantages and Quirks? Embryo 1 Embryo 2 combine samples for a single sequencing chip Barcode 1 CTAAGGTAAC
Generating Spontaneous Copy Number Variants (CNVs) Jennifer Freeman Assistant Professor of Toxicology School of Health Sciences Purdue University
 Role of Chemical lexposure in Generating Spontaneous Copy Number Variants (CNVs) Jennifer Freeman Assistant Professor of Toxicology School of Health Sciences Purdue University CNV Discovery Reference Genetic
Role of Chemical lexposure in Generating Spontaneous Copy Number Variants (CNVs) Jennifer Freeman Assistant Professor of Toxicology School of Health Sciences Purdue University CNV Discovery Reference Genetic
Chromosome aberrations in a series of 120 multiple myeloma cases with abnormal karyotypes
 Chromosome aberrations in a series of 120 multiple myeloma cases with abnormal karyotypes Anwar N. Mohamed, 1,2 * Gail Bentley, 1 Michelle L. Bonnett, 1 Jeff Zonder, 3 and Ayad Al-Katib 3 1 Department
Chromosome aberrations in a series of 120 multiple myeloma cases with abnormal karyotypes Anwar N. Mohamed, 1,2 * Gail Bentley, 1 Michelle L. Bonnett, 1 Jeff Zonder, 3 and Ayad Al-Katib 3 1 Department
MYC Translocations In Multiple Myeloma Involve Recruitment Of Enhancer Elements Resulting In Over- Expression and Decreased Overall Survival
 in partnership with MYC Translocations In Multiple Myeloma Involve Recruitment Of Enhancer Elements Resulting In Over- Expression and Decreased Overall Survival Brian A Walker Centre for Myeloma Research,
in partnership with MYC Translocations In Multiple Myeloma Involve Recruitment Of Enhancer Elements Resulting In Over- Expression and Decreased Overall Survival Brian A Walker Centre for Myeloma Research,
Multiplex target enrichment using DNA indexing for ultra-high throughput variant detection
 Multiplex target enrichment using DNA indexing for ultra-high throughput variant detection Dr Elaine Kenny Neuropsychiatric Genetics Research Group Institute of Molecular Medicine Trinity College Dublin
Multiplex target enrichment using DNA indexing for ultra-high throughput variant detection Dr Elaine Kenny Neuropsychiatric Genetics Research Group Institute of Molecular Medicine Trinity College Dublin
Myeloma cytogenetics: a personal and historical perspective. Fiona Ross UKMF 19/03/2014
 Myeloma cytogenetics: a personal and historical perspective Fiona Ross UKMF 19/03/2014 Chromosome abnormalities in myeloma Abnormal myeloma karyotypes started to be published in 1970s but many cases throughout
Myeloma cytogenetics: a personal and historical perspective Fiona Ross UKMF 19/03/2014 Chromosome abnormalities in myeloma Abnormal myeloma karyotypes started to be published in 1970s but many cases throughout
Significance of Chromosome Changes in Hematological Disorders and Solid Tumors
 Significance of Chromosome Changes in Hematological Disorders and Solid Tumors Size of Components of Human Genome Size of haploid genome! Estimated genetic constitution! Size of average chromosome
Significance of Chromosome Changes in Hematological Disorders and Solid Tumors Size of Components of Human Genome Size of haploid genome! Estimated genetic constitution! Size of average chromosome
Significance of Chromosome Changes in Hematological Disorders and Solid Tumors
 Significance of Chromosome Changes in Hematological Disorders and Solid Tumors Size of Components of Human Genome Size of haploid genome 3.3 X 10 9 DNA basepairs Estimated genetic constitution 30,000
Significance of Chromosome Changes in Hematological Disorders and Solid Tumors Size of Components of Human Genome Size of haploid genome 3.3 X 10 9 DNA basepairs Estimated genetic constitution 30,000
Advance Your Genomic Research Using Targeted Resequencing with SeqCap EZ Library
 Advance Your Genomic Research Using Targeted Resequencing with SeqCap EZ Library Marilou Wijdicks International Product Manager Research For Life Science Research Only. Not for Use in Diagnostic Procedures.
Advance Your Genomic Research Using Targeted Resequencing with SeqCap EZ Library Marilou Wijdicks International Product Manager Research For Life Science Research Only. Not for Use in Diagnostic Procedures.
MPS for translocations
 MPS for translocations Filip Van Nieuwerburgh, Ph.D. Lab of Pharmaceutical Biotechnology NXTGNT massively parallel sequencing facility, Ghent University In collaboration with: Center for Medical Genetics,
MPS for translocations Filip Van Nieuwerburgh, Ph.D. Lab of Pharmaceutical Biotechnology NXTGNT massively parallel sequencing facility, Ghent University In collaboration with: Center for Medical Genetics,
Cytogenetics and FISH Studies in Multiple Myeloma A Retrospective Study from Western India
 American Journal of Current Biology Gadhia P et al. American Journal of Current Biology 2014, 2:1-7 American Journal Page 1 of of Current 7 Biology http://www.ivyunion.org/index.php/ajcurrb Vol. 2, Article
American Journal of Current Biology Gadhia P et al. American Journal of Current Biology 2014, 2:1-7 American Journal Page 1 of of Current 7 Biology http://www.ivyunion.org/index.php/ajcurrb Vol. 2, Article
TEST MENU TEST CPT CODES TAT. Chromosome Analysis Bone Marrow x 2, 88264, x 3, Days
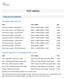 TEST MENU CANCER/LEUKEMIA CHROMOSOME ANALYSIS Chromosome Analysis Bone Marrow 88237 x 2, 88264, 88280 x 3, 88291 4 Days Chromosome Analysis Bone Marrow Core 88237 x 2, 88264, 88280 x 3, 88291 4 Days Chromosome
TEST MENU CANCER/LEUKEMIA CHROMOSOME ANALYSIS Chromosome Analysis Bone Marrow 88237 x 2, 88264, 88280 x 3, 88291 4 Days Chromosome Analysis Bone Marrow Core 88237 x 2, 88264, 88280 x 3, 88291 4 Days Chromosome
Investigating rare diseases with Agilent NGS solutions
 Investigating rare diseases with Agilent NGS solutions Chitra Kotwaliwale, Ph.D. 1 Rare diseases affect 350 million people worldwide 7,000 rare diseases 80% are genetic 60 million affected in the US, Europe
Investigating rare diseases with Agilent NGS solutions Chitra Kotwaliwale, Ph.D. 1 Rare diseases affect 350 million people worldwide 7,000 rare diseases 80% are genetic 60 million affected in the US, Europe
Clinical Validation of Cytocell Pathology Probes
 Clinical Validation of Cytocell Pathology Probes Shivanand Richardson Specialized Technician Molecular Pathology Department of Pathology, University Medical Center Utrecht The Netherlands 5th annual Cytocell
Clinical Validation of Cytocell Pathology Probes Shivanand Richardson Specialized Technician Molecular Pathology Department of Pathology, University Medical Center Utrecht The Netherlands 5th annual Cytocell
Whole-Genome SNP Array Analysis Genomic Complexity and Clinical Relevance in Prenatal, Postnatal, and Oncology Testing
 The Evolution of Cytogenetics Testing Whole-Genome SNP Array Analysis Genomic Complexity and Clinical Relevance in Prenatal, Postnatal, and Oncology Testing Mark Micale, Ph.D., F.A.C.M.G. Medical Director,
The Evolution of Cytogenetics Testing Whole-Genome SNP Array Analysis Genomic Complexity and Clinical Relevance in Prenatal, Postnatal, and Oncology Testing Mark Micale, Ph.D., F.A.C.M.G. Medical Director,
Agilent s Copy Number Variation (CNV) Portfolio
 Technical Overview Agilent s Copy Number Variation (CNV) Portfolio Abstract Copy Number Variation (CNV) is now recognized as a prevalent form of structural variation in the genome contributing to human
Technical Overview Agilent s Copy Number Variation (CNV) Portfolio Abstract Copy Number Variation (CNV) is now recognized as a prevalent form of structural variation in the genome contributing to human
Comparison of FISH, CpG-stimulation, chromosomal microarray and mate pair sequencing in 20 patients with CLL or lymphoma
 Comparison of FISH, CpG-stimulation, chromosomal microarray and mate pair sequencing in 20 patients with CLL or lymphoma Stephanie Smoley Cancer Genomics Consortium August 9 th, 2017 2016 MFMER slide-1
Comparison of FISH, CpG-stimulation, chromosomal microarray and mate pair sequencing in 20 patients with CLL or lymphoma Stephanie Smoley Cancer Genomics Consortium August 9 th, 2017 2016 MFMER slide-1
PG-Seq NGS Kit for Preimplantation Genetic Screening
 Application Note: PG-Seq Validation Study PG-Seq NGS Kit for Preimplantation Genetic Screening Validation using Multi (5-10) Cells and Single Cells from euploid and aneuploid cell lines Introduction Advances
Application Note: PG-Seq Validation Study PG-Seq NGS Kit for Preimplantation Genetic Screening Validation using Multi (5-10) Cells and Single Cells from euploid and aneuploid cell lines Introduction Advances
(Fully automated FISH Analysis for B-cell Chronic Lymphocytic Leukemia: a faster alternative to manual double scoring)
 Routine integration of fully automated image analysis of Kreatech CLL FISH Probes, using the CytoVision GSL Scanning System (Fully automated FISH Analysis for B-cell Chronic Lymphocytic Leukemia: a faster
Routine integration of fully automated image analysis of Kreatech CLL FISH Probes, using the CytoVision GSL Scanning System (Fully automated FISH Analysis for B-cell Chronic Lymphocytic Leukemia: a faster
Nature Biotechnology: doi: /nbt.1904
 Supplementary Information Comparison between assembly-based SV calls and array CGH results Genome-wide array assessment of copy number changes, such as array comparative genomic hybridization (acgh), is
Supplementary Information Comparison between assembly-based SV calls and array CGH results Genome-wide array assessment of copy number changes, such as array comparative genomic hybridization (acgh), is
What s the Human Genome Project Got to Do with Developmental Disabilities?
 What s the Human Genome Project Got to Do with Developmental Disabilities? Disclosures Neither speaker has anything to disclose. Phase Two: Interpretation Officially started in October 1990 Goals of the
What s the Human Genome Project Got to Do with Developmental Disabilities? Disclosures Neither speaker has anything to disclose. Phase Two: Interpretation Officially started in October 1990 Goals of the
Genomic structural variation
 Genomic structural variation Mario Cáceres The new genomic variation DNA sequence differs across individuals much more than researchers had suspected through structural changes A huge amount of structural
Genomic structural variation Mario Cáceres The new genomic variation DNA sequence differs across individuals much more than researchers had suspected through structural changes A huge amount of structural
The Utilization of Karyotyping, ifish, and MLPA for the Detection of Recurrence Genetic Aberrations in Multiple Myeloma
 DOI:10.22034/APJCP.2017.18.11.3135 The Utilization of Karyotyping, ifish and MLPA for the Detection of Recurrence Genetic Aberrations in Multiple Myeloma RESEARCH ARTICLE The Utilization of Karyotyping,
DOI:10.22034/APJCP.2017.18.11.3135 The Utilization of Karyotyping, ifish and MLPA for the Detection of Recurrence Genetic Aberrations in Multiple Myeloma RESEARCH ARTICLE The Utilization of Karyotyping,
Abstract. Hematopathology / Improved Cytogenetics in Lymphoma. Key Words: Cytogenetics; B-cell lymphoma; CpG-oligonucleotide DSP30
 Hematopathology / Improved Cytogenetics in Lymphoma Improved Detection Rate of Cytogenetic Abnormalities in Chronic Lymphocytic Leukemia and Other Mature B-Cell Neoplasms With Use of CpG-Oligonucleotide
Hematopathology / Improved Cytogenetics in Lymphoma Improved Detection Rate of Cytogenetic Abnormalities in Chronic Lymphocytic Leukemia and Other Mature B-Cell Neoplasms With Use of CpG-Oligonucleotide
Oncology Cytogenetics Diagnostic Service - User Guide 2014
 Oncology Cytogenetics Diagnostic Service - User Guide 2014 Contact details Address: Cytogenetics Department 5 th Floor Tower Wing Guy s Hospital Great Maze Pond London SE1 9RT General enquiries 0207 188
Oncology Cytogenetics Diagnostic Service - User Guide 2014 Contact details Address: Cytogenetics Department 5 th Floor Tower Wing Guy s Hospital Great Maze Pond London SE1 9RT General enquiries 0207 188
Abstract. Optimization strategy of Copy Number Variant calling using Multiplicom solutions APPLICATION NOTE. Introduction
 Optimization strategy of Copy Number Variant calling using Multiplicom solutions Michael Vyverman, PhD; Laura Standaert, PhD and Wouter Bossuyt, PhD Abstract Copy number variations (CNVs) represent a significant
Optimization strategy of Copy Number Variant calling using Multiplicom solutions Michael Vyverman, PhD; Laura Standaert, PhD and Wouter Bossuyt, PhD Abstract Copy number variations (CNVs) represent a significant
June 11, Ella Noel, D.O., FACOI 1717 West Broadway Madison, WI
 June 11, 2018 Ella Noel, D.O., FACOI 1717 West Broadway Madison, WI 53713 policycomments@wpsic.com RE: Draft Local Coverage Determination: MolDX: MDS FISH (DL37772) Dear Dr. Noel Thank you for the opportunity
June 11, 2018 Ella Noel, D.O., FACOI 1717 West Broadway Madison, WI 53713 policycomments@wpsic.com RE: Draft Local Coverage Determination: MolDX: MDS FISH (DL37772) Dear Dr. Noel Thank you for the opportunity
Multiple Copy Number Variations in a Patient with Developmental Delay ASCLS- March 31, 2016
 Multiple Copy Number Variations in a Patient with Developmental Delay ASCLS- March 31, 2016 Marwan Tayeh, PhD, FACMG Director, MMGL Molecular Genetics Assistant Professor of Pediatrics Department of Pediatrics
Multiple Copy Number Variations in a Patient with Developmental Delay ASCLS- March 31, 2016 Marwan Tayeh, PhD, FACMG Director, MMGL Molecular Genetics Assistant Professor of Pediatrics Department of Pediatrics
Cytogenetic and molecular abnormalities in AML. Dr Elizabeth Tegg Director of haematology Pathology West
 Cytogenetic and molecular abnormalities in AML Dr Elizabeth Tegg Director of haematology Pathology West Outline Classification of AML Types of genetic changes Next generation sequencing in HM Outline Classification
Cytogenetic and molecular abnormalities in AML Dr Elizabeth Tegg Director of haematology Pathology West Outline Classification of AML Types of genetic changes Next generation sequencing in HM Outline Classification
Karyology. Preparation and study of karyotypes is part of Cytogenetics.
 Chromosomal Karyotyping Karyology Karyotyping - process of pairing and ordering all chromosomes of an organism, thus providing a genome-wide snapshot of an individual's chromosomes. Karyotypes describe
Chromosomal Karyotyping Karyology Karyotyping - process of pairing and ordering all chromosomes of an organism, thus providing a genome-wide snapshot of an individual's chromosomes. Karyotypes describe
Statistical Analysis of Single Nucleotide Polymorphism Microarrays in Cancer Studies
 Statistical Analysis of Single Nucleotide Polymorphism Microarrays in Cancer Studies Stanford Biostatistics Workshop Pierre Neuvial with Henrik Bengtsson and Terry Speed Department of Statistics, UC Berkeley
Statistical Analysis of Single Nucleotide Polymorphism Microarrays in Cancer Studies Stanford Biostatistics Workshop Pierre Neuvial with Henrik Bengtsson and Terry Speed Department of Statistics, UC Berkeley
Objectives. Morphology and IHC. Flow and Cyto FISH. Testing for Heme Malignancies 3/20/2013
 Molecular Markers in Hematologic Malignancy: Ways to locate the needle in the haystack. Objectives Review the types of testing for hematologic malignancies Understand rationale for molecular testing Marcie
Molecular Markers in Hematologic Malignancy: Ways to locate the needle in the haystack. Objectives Review the types of testing for hematologic malignancies Understand rationale for molecular testing Marcie
at least 5 probes standard 8 probes (13, 15, 16, 18, 21, 22, 15, X, Y) at least 5 probes standard 8 probes (13, 15, 16, 18, 21, 22, X, Y)
 Management of FISH probe testing Petra Musilová et al. Repromeda, Brno, Czech Rep. Veterinary Research Institute, Brno Genprogress, Brno, Czech Rep. Aneuploidy screening at least 5 probes standard 8 probes
Management of FISH probe testing Petra Musilová et al. Repromeda, Brno, Czech Rep. Veterinary Research Institute, Brno Genprogress, Brno, Czech Rep. Aneuploidy screening at least 5 probes standard 8 probes
Supplementary note: Comparison of deletion variants identified in this study and four earlier studies
 Supplementary note: Comparison of deletion variants identified in this study and four earlier studies Here we compare the results of this study to potentially overlapping results from four earlier studies
Supplementary note: Comparison of deletion variants identified in this study and four earlier studies Here we compare the results of this study to potentially overlapping results from four earlier studies
Kim et al. Journal of Hematology & Oncology 2013, 6:76
 Kim et al. Journal of Hematology & Oncology 2, 6:76 JOURNAL OF HEMATOLOGY & ONCOLOGY RESEARCH Open Access Clinical significance of cytogenetic aberrations in bone marrow of patients with diffuse large
Kim et al. Journal of Hematology & Oncology 2, 6:76 JOURNAL OF HEMATOLOGY & ONCOLOGY RESEARCH Open Access Clinical significance of cytogenetic aberrations in bone marrow of patients with diffuse large
Structural Variation and Medical Genomics
 Structural Variation and Medical Genomics Andrew King Department of Biomedical Informatics July 8, 2014 You already know about small scale genetic mutations Single nucleotide polymorphism (SNPs) Deletions,
Structural Variation and Medical Genomics Andrew King Department of Biomedical Informatics July 8, 2014 You already know about small scale genetic mutations Single nucleotide polymorphism (SNPs) Deletions,
Chapter 10 Chromosomes and Cell Reproduction
 Chapter 10 Chromosomes and Cell Reproduction Chromosomes Organisms grow by dividing of cells Binary Fission form of asexual reproduction that produces identical offspring (Bacteria) Eukaryotes have two
Chapter 10 Chromosomes and Cell Reproduction Chromosomes Organisms grow by dividing of cells Binary Fission form of asexual reproduction that produces identical offspring (Bacteria) Eukaryotes have two
Fluorescence in-situ Hybridization (FISH) ETO(RUNX1T1)/AML1(RUNX1) or t(8;21)(q21.3;q22)
 PML/RARA t(15;17) Translocation Assay Result : nuc ish(pml 2)(RARA 2)[200] : 200/200(100%) interphase nuclei show normal 2O 2G signals for PML/RARA : is Negative for t(15;17)(q22;q21.1) 2 Orange 2 Green
PML/RARA t(15;17) Translocation Assay Result : nuc ish(pml 2)(RARA 2)[200] : 200/200(100%) interphase nuclei show normal 2O 2G signals for PML/RARA : is Negative for t(15;17)(q22;q21.1) 2 Orange 2 Green
Chapter 3 Chromosomal Aberrations
 MEDICAL GENETICS Chapter 3 Chromosomal Aberrations Abnormalities of chromosomes may be either numerical or structural and may involve one or more autosomes, sex chromosomes, or both simultaneously. Numerical
MEDICAL GENETICS Chapter 3 Chromosomal Aberrations Abnormalities of chromosomes may be either numerical or structural and may involve one or more autosomes, sex chromosomes, or both simultaneously. Numerical
Table 1: biological tests in SMD
 Table 1: biological tests in SMD Tests Mandatory Recommended Under validation Morphology Marrow aspirate Marrow biopsy 1 Iron staining Quantification of dysplasia WHO 2008 Classification Cytogenetics Conventional
Table 1: biological tests in SMD Tests Mandatory Recommended Under validation Morphology Marrow aspirate Marrow biopsy 1 Iron staining Quantification of dysplasia WHO 2008 Classification Cytogenetics Conventional
Do acgh analysis have a place in routine cytogenetic workup in leukemia/cancer? - A single institution experience. Cambridge, April 9 th 2013
 Do acgh analysis have a place in routine cytogenetic workup in leukemia/cancer? - A single institution experience. Cambridge, April 9 th 2013 Aarhus University Hospital Eigil Kjeldsen, Cancercytogenetic
Do acgh analysis have a place in routine cytogenetic workup in leukemia/cancer? - A single institution experience. Cambridge, April 9 th 2013 Aarhus University Hospital Eigil Kjeldsen, Cancercytogenetic
XCyting DNA FISH Probes
 XCyting DNA FISH Catalog 2017/18 MetaSystems 1. Industriestr. 7 D-68804 Altlussheim Germany Phone: +49 6205 2927610 Fax: +49 6205 2927629 E-Mail info: info@metasystems-probes.com E-Mail orders: orders@metasystems-probes.com
XCyting DNA FISH Catalog 2017/18 MetaSystems 1. Industriestr. 7 D-68804 Altlussheim Germany Phone: +49 6205 2927610 Fax: +49 6205 2927629 E-Mail info: info@metasystems-probes.com E-Mail orders: orders@metasystems-probes.com
Genomic Medicine: What every pathologist needs to know
 Genomic Medicine: What every pathologist needs to know Stephen P. Ethier, Ph.D. Professor, Department of Pathology and Laboratory Medicine, MUSC Director, MUSC Center for Genomic Medicine Genomics and
Genomic Medicine: What every pathologist needs to know Stephen P. Ethier, Ph.D. Professor, Department of Pathology and Laboratory Medicine, MUSC Director, MUSC Center for Genomic Medicine Genomics and
September 04, 2008
 27027 Tourney Road Valencia, CA 91355 800 421 7110 www.specialtylabs.com Test Updates September 04, 2008 Dear Valued Client: As you may be aware, in recent years there has been a tremendous challenge in
27027 Tourney Road Valencia, CA 91355 800 421 7110 www.specialtylabs.com Test Updates September 04, 2008 Dear Valued Client: As you may be aware, in recent years there has been a tremendous challenge in
Allogeneic Hematopoietic Stem-Cell Transplantation for Myelodysplastic Syndromes and Myeloproliferative Neoplasms. Policy Specific Section:
 Medical Policy Allogeneic Hematopoietic Stem-Cell Transplantation for Myelodysplastic Syndromes and Myeloproliferative Type: Medical Necessity and Investigational / Experimental Policy Specific Section:
Medical Policy Allogeneic Hematopoietic Stem-Cell Transplantation for Myelodysplastic Syndromes and Myeloproliferative Type: Medical Necessity and Investigational / Experimental Policy Specific Section:
Initial Diagnosis and Treatment 81 Male
 Case SH2017-0359 Shiraz Fidai 1, Sandeep Gurbuxani 1, Girish Venkataraman 1, Gordana Raca 2, Madina Sukhanova 3, Michelle M Le Beau 3, Y. Lynn Wang 4, Mir Alikhan 4, Megan M.McNerney 4, Yuri Kobzev 4,
Case SH2017-0359 Shiraz Fidai 1, Sandeep Gurbuxani 1, Girish Venkataraman 1, Gordana Raca 2, Madina Sukhanova 3, Michelle M Le Beau 3, Y. Lynn Wang 4, Mir Alikhan 4, Megan M.McNerney 4, Yuri Kobzev 4,
Mutation Detection and CNV Analysis for Illumina Sequencing data from HaloPlex Target Enrichment Panels using NextGENe Software for Clinical Research
 Mutation Detection and CNV Analysis for Illumina Sequencing data from HaloPlex Target Enrichment Panels using NextGENe Software for Clinical Research Application Note Authors John McGuigan, Megan Manion,
Mutation Detection and CNV Analysis for Illumina Sequencing data from HaloPlex Target Enrichment Panels using NextGENe Software for Clinical Research Application Note Authors John McGuigan, Megan Manion,
Collected: , PM Sent: , PM Received: , PM Preliminary: , PM. Notification Status: COMPREHENSIVE DIAGNOSIS
 PATIENT Name:HIPAA, Compliant DOB: 03-25-1945 (60 yr) ID#: xxx-xx-0000 Sex: M Tel: xxx-xxx-xxxx SPECIMEN Your No:WS05-xxxx Case No:C05-00xxx Req. No:Txxxxx Collected: 06-08-05, PM Sent: 06-09-05, PM Received:
PATIENT Name:HIPAA, Compliant DOB: 03-25-1945 (60 yr) ID#: xxx-xx-0000 Sex: M Tel: xxx-xxx-xxxx SPECIMEN Your No:WS05-xxxx Case No:C05-00xxx Req. No:Txxxxx Collected: 06-08-05, PM Sent: 06-09-05, PM Received:
CHAPTER-VII : SUMMARY AND CONCLUSIONS
 CHAPTER-VII : SUMMARY AND CONCLUSIONS 199 SUMMARY AND CONCLUSIONS t The rapid development of human genetics during the past couple of decades and the discovery of numerous cytogenetic abnormalities have
CHAPTER-VII : SUMMARY AND CONCLUSIONS 199 SUMMARY AND CONCLUSIONS t The rapid development of human genetics during the past couple of decades and the discovery of numerous cytogenetic abnormalities have
SEAMLESS CGH DIAGNOSTIC TESTING
 SEAMLESS CGH DIAGNOSTIC TESTING GENETISURE DX POSTNATAL ASSAY Informed decisions start with a complete microarray platform for postnatal analysis For In Vitro Diagnostic Use INTENDED USE: GenetiSure Dx
SEAMLESS CGH DIAGNOSTIC TESTING GENETISURE DX POSTNATAL ASSAY Informed decisions start with a complete microarray platform for postnatal analysis For In Vitro Diagnostic Use INTENDED USE: GenetiSure Dx
