Identification of suitable reference genes for the relative quantification of micrornas in pleural effusion
|
|
|
- Roy Flowers
- 5 years ago
- Views:
Transcription
1 ONCOLOGY LETTERS 8: , 2014 Identification of suitable reference genes for the relative quantification of micrornas in pleural effusion HYE SUK HAN 1, YEONG NANG JO 2, JIN YONG LEE 3, SONG YI CHOI 4, YUSOOK JEONG 1, JIEUN YUN 2 and OK JUN LEE 4 1 Department of Internal Medicine, College of Medicine, Chungbuk National University, Cheongju ; 2 Bioevaluation Center, Korea Research Institute of Bioscience and Biotechnology, Cheongwon ; 3 Public Health Medical Service, SMG SNU Boramae Medical Center, Seoul ; 4 Department of Pathology, College of Medicine, Chungbuk National University, Cheongju , Republic of Korea Received December 21, 2013; Accepted July 11, 2014 DOI: /ol Abstract. Circulating cell free micrornas (mirnas) are potential biomarkers of cancer. Reverse transcription quantitative polymerase chain reaction (RT qpcr) is widely used in mirna expression studies. The aim of this study was to identify suitable reference genes for RT qpcr analyses of mirna expression levels in pleural effusion. The expression levels of candidate reference mirnas were investigated in 10 benign pleural effusion (BPE) and 10 lung adenocarcinoma associated malignant pleural effusion (LA MPE) samples using mirna microarrays. The expression levels of candidate reference mirnas, together with those of U6 small nuclear RNA (snrna), RNU6B, RNU44 and RNU48 small RNAs, in 46 BPE and 45 LA MPE samples were validated by RT qpcr, and were analyzed using the NormFinder and BestKeeper algorithms. The impact of different normalization approaches on the detection of differential expression levels of mir 198 in BPE and LA MPE samples was also assessed. As determined by the mirna microarray data, five candidate reference mirnas were identified. Following RT qpcr validation, U6 snrna, mir 192, mir 20a, mir 221, mir 222 and mir 16 were evaluated using the NormFinder and BestKeeper software programs. U6 snrna and mir 192 were identified as single reference genes and the combination of these genes was preferred for the relative quantification of mirna expression Correspondence to: Dr Jieun Yun, Bioevaluation Center, Korea Research Institute of Bioscience and Biotechnology, Yangcheong ri, Ochang eup, Cheongwon , Republic of Korea E mail: jyun@kribb.re.kr Dr Ok Jun Lee, Department of Pathology, College of Medicine, Chungbuk National University, 52 Naesudong-ro, Seowon-gu, Cheongju , Republic of Korea E mail: ok5218@hanmail.net Key words: micrornas, pleural effusion, reference levels in pleural effusion. Normalization of mir 98 expression levels to those of U6 snrna, mir 192 or a combination of these genes enabled the detection of a significant difference between BPE and LA MPE samples. Therefore, U6 snrna and mir 192 are recommended as reference genes for the relative quantification of mirna expression levels in pleural effusion. Introduction MicroRNAs (mirnas) constitute a class of small non coding RNAs nucleotides in length that function as post transcriptional regulators of gene expression (1). mirnas exert important regulatory roles in the majority of cellular and developmental processes, and have been implicated in numerous human diseases, including cancer (2). Since their recent identification, the investigation of the potential for using extracellular and circulating mirnas as non invasive cancer biomarkers in body fluids, such as serum and plasma, has rapidly expanded (3,4). Circulating mirnas are stabilized and protected from RNase degradation by incorporation into various protein complexes or membranous particles, including exosomes and microvesicles (4). Pleural effusion is a common clinical manifestation of the various types of non small cell lung cancer, particularly adenocarcinoma, and the diagnosis of pleural effusion in these patients is of particular clinical importance. According to the Cancer Staging system, the existence of malignant pleural effusion (MPE) at stage IV indicates systemic disease; these patients cannot be treated with local therapeutic methods, such as surgical resection or radiotherapy (5). Currently the diagnosis of MPE relies on cytological analysis of pleural fluid; however, this method has limited sensitivity (6) and alternative methods are required to improve the diagnosis of pleural effusion. Recently, several studies have reported that the cell free mirnas present in pleural effusion may be useful diagnostic biomarkers for discriminating between benign pleural effusion (BPE) and MPE (7 9). mirna expression levels have been analyzed using traditional semi quantitative methods, including northern blotting, bead based flow cytometry and microarray technology;
2 1890 HAN et al: REFERENCE GENES IN PLEURAL EFFUSION however, reverse transcription quantitative polymerase chain reaction (RT qpcr) is the most sensitive, reproducible and widely used approach for the evaluation of mirnas (10). To prevent errors in datasets and to overcome the experimental variation associated with RT qpcr analysis procedures, such as RNA isolation, cdna synthesis and PCR kinetics, the relative quantification of mirnas through normalization to one or more stably expressed reference gene is the preferred approach (10). Suitable reference genes should be expressed constitutively and the expression levels should be unaffected by biological change, disease or treatment. The normalization of experimental gene RT qpcr data to those of unreliable reference genes may result in incorrect quantification of the mirnas of interest; the importance of validating suitable candidate endogenous control reference genes has been previously demonstrated (11 15). Although several mirna expression profiling studies in pleural effusion specimens have been performed (7 9,16), validation strategies for the selection of reference genes for normalization have, to the best of our knowledge, not been reported. Therefore, the aim of the present study was to identify suitable reference genes for normalizing the expression levels of circulating mirnas in pleural effusion identified by RT qpcr. Materials and methods Patients and pleural effusion samples. Pleural effusion samples were derived from 111 patients who visited Chungbuk National University Hospital (Cheongju, Korea) or Kangwon National University Hospital (Chuncheon, Korea) between February 2009 and September The pleural effusions were diagnosed as benign, as determined by the clinical context and the absence of malignant cells in at least two separate samples from the same patient. All lung adenocarcinoma associated MPE (LA MPE) samples were obtained from patients with tumors that were histologically and clinically diagnosed as primary adenocarcinoma of the lung, and that contained adenocarcinoma cells, as confirmed by pathologists. All samples were transported to the laboratory within 30 min of collection. The samples were then centrifuged at 11,300 x g for 5 min, and the supernatant and sediment fractions were aliquoted into separate microcentrifuge tubes and stored at 80 C prior to use. All patients provided written informed consent, and the study was reviewed and approved by the Institutional Review Board of Chungbuk National University Hospital. RNA extraction. RNA was extracted from 500 µl of each sample by the use of a Genolution urine mirna purification kit (Genolution Pharmaceuticals, Inc., Seoul, Korea), according to the manufacturer's instructions. The quantity of RNA extracted from each sample was examined using a NanoDrop ND 1000 spectrophotometer (Thermo Scientific, Wilmington, DE, USA). Microarray based mirna analysis. Candidate reference mirnas were selected from 10 BPE samples and 10 LA MPE samples using data from mirna microarray analyses. The BPE and LA MPE mirna expression profiles were generated by hybridizing small RNAs extracted from the pleural effusion samples to locked nucleic acid probes targeting 160 human mirnas. A PANArray TM mirna expression profiling kit (Panagene Inc., Daejeon, Korea) was used to identify the candidate reference mirnas. This kit is a peptide nucleic acid (PNA) based microarray, which uses a novel mirna labeling method to examine the expression profiles of cancer and stem cell related mirnas. RT qpcr. The expression levels of the candidate reference mirnas in 46 BPE samples and 45 LA MPE samples were validated using RT qpcr. The expression levels of two small nuclear RNAs (snrnas; U6 snrna and RNU6B) and two small nucleolar RNAs (snornas; RNU44 and RNU48) were also measured. Reverse transcription of 100 ng isolated mirna was conducted using a high capacity cdna reverse transcription kit (Applied Biosystems, Foster City, CA, USA), according to the manufacturer's instructions, and a specific mirna primer provided with a TaqMan MicroRNA Assay kit (Applied Biosystems). RT qpcr was performed using the Applied Biosystems 7,500 Fast Real Time PCR system along with a TaqMan MicroRNA Assay, TaqMan Universal PCR Master mix and No AmpErase UNG (Applied Biosystems). All reactions were performed in triplicate and Cq data were determined using the default threshold settings. The expression levels of the candidate reference mirnas and small RNAs included in the final analysis were calculated using the 2 ΔΔCt method. Data analysis. Non parametric tests [Mann Whitney U test and Kruskal Wallis one way analysis of variance (ANOVA) with Dunn's multiple comparisons correction] were used to determine any statistically significant differences between independent groups. Spearman's correlation coefficients were used to calculate the associations between the clinical variables and the expression levels of the candidate reference genes. P<0.05 was considered to indicate a statistically significant difference. All statistical analyses were performed using SPSS software for Windows, version 15.0 (SPSS, Inc., Chicago, IL, USA). The NormFinder ( and BestKeeper ( html) software programs were used to analyze the expression stabilities of the reference mirnas. NormFinder is a Microsoft Excel add in that uses an ANOVA based model to calculate stability values from a panel of candidate genes in different subgroups by combining the intra and inter group expression variation (17). BestKeeper determines the geometric mean and coefficient of variance (18). Genes with standard deviation >1 were considered to be inconstant. Inter gene associations were examined by pairwise correlation analysis. This calculation was used to determine whether the genes exhibited similar expression behavior. Candidate reference genes that exhibited high correlation with one another were included in the BestKeeper Index calculation. In the two programs, the lowest stability values indicate the most stably expressed genes. To determine the expression levels of the target mirnas relative to suitable reference genes, the comparative ΔCt method was used, with the relative quantities (Q rel ) calculated as follows: Q rel = 2 Δ(Cq test sample Cq normalizer). When normalized to the sample volume, the relative expression level of a target mirna was calculated as follows: Q rel = 2 Δ(Cq test sample Cq average of all samples).
3 ONCOLOGY LETTERS 8: , Results Selection of candidate mirna reference genes by microarray analysis. A total of 10 BPE samples and 10 LA MPE samples were analyzed using mirna microarrays. The following criteria were used to identify candidate reference mirnas: mirnas were detected in all 20 pleural effusion samples, the mean change in mirna expression levels was <1.1 fold, and no significant differences in mirna expression levels (P>0.05) between the BPE and LA MPE samples was detected. To avoid artifacts resulting from normalization of the microarray data, raw microarray expression data were also used. Of the 160 human mirnas included in the PANArray mirna expression profiling kit, 59 were identified in all pleural effusion samples. Five of these mirnas (mir 192, mir 20a, mir 221, mir 222 and mir 16) exhibited mean fold changes of <1.1 fold, with expression levels that did not differ significantly between the BPE and LA MPE samples. Validation of candidate reference genes by RT qpcr. The baseline characteristics of the patients in the validation cohort are listed in Table I. RT qpcr was performed to further evaluate the expression patterns of the five candidate reference mirnas identified by microarray analysis. A cohort of 91 pleural effusion samples, including 46 BPE samples and 45 LA MPE samples, was used for the RT qpcr experiments. Furthermore, the set of candidate reference mirnas was extended by inclusion of the U6 snrna, RNU6B, RNU44 and RNU48 small RNAs, which are commonly used for mirna expression level normalization. Details of the functions and PCR amplification efficiencies of each of the candidate reference mirnas and small RNAs are listed in Table II (19 25). To evaluate the range of detectable Cq values, the mir 192 expression levels in 10 fold serial dilutions of a pleural effusion sample were examined. A plot of Cq values versus the log concentration revealed a linear correlation for Cq values between 31 and 44 (Fig. 1). These results demonstrate that this method of measurement was appropriate for the present study. RNU6B, RNU44 and RNU48 were not detected in the majority of pleural effusion samples; therefore, these small RNAs were excluded from further analysis. The expression levels of the remaining six candidate reference genes, namely mir 192, mir 20a, mir 221, mir 222, mir 16 and U6 snrna, varied widely across the samples tested, with Cq values ranging between and (Fig. 2A). The expression levels of all mirnas did not differ significantly between BPE and LA MPE samples (Fig. 2B). The expression levels of the five mirnas and U6 snrna in BPE and LA MPE samples did not depend on age (Spearman's rank correlation ranged between and 0.128, P-value ranged from to 0.988), gender (Mann Whitney U test, P value ranged between and 0.808) or diagnosis (Kruskal Wallis test, P value ranged between and 0.874). Stability of the candidate reference gene expression levels. Variations in the expression stabilities of the reference genes were further assessed using the NormFinder and BestKeeper algorithms. The ranking of the genes by these software programs is summarized in Table III; lower stability values indicate higher gene stability. NormFinder identified U6 snrna as the most stably expressed gene, with a stability Table I. Baseline characteristics of the patients. Clinical characteristics BPE, n (%) LA MPE, n(%) (n=46) (n=45) Age (years) a 63 (22 92) 68 (33 92) Gender Male 31 (67.4) 23 (51.1) Female 15 (32.6) 22 (49.9) Diagnosis Tuberculosis 22 (47.8) - Pneumonia 19 (41.3) - Transudate 5 (10.9) - Metastatic sites M1a - 18 (40.0) M1b - 27 (60.0) a Median (range). BPE, benign pleural effusion; LA MPE, lung adenocarcinoma associated malignant pleural effusion; M1a, separate tumor nodule(s) in a contralateral lobe, a tumor with pleural nodules or malignant pleural (or pericardial) effusion; M1b, distant metastasis. value of Combining the U6 snrna and mir 192 gene data further reduced the NormFinder stability value to Using the BestKeeper algorithm, a similar ranking order of the candidate reference genes was observed; mir 192 was identified as the most stable gene (stability value of 0.160), followed by U6 snrna (stability value of 0.177). Influence of reference genes on the relative quantification of mir 198. Cell free mir 198 expression levels have been previously demonstrated to be significantly lower in LA MPE samples than in BPE samples (9). Therefore, in the present study, to assess the impact of reference gene selection on the validity of mirna expression data, the effect of normalization to the following variables on the relative quantification of mir 198 expression levels was evaluated: Sample volume; U6 snrna expression levels (the most stable candidate gene ranked by NormFinder); mir 192 expression levels (the most stable candidate gene ranked by BestKeeper); and the combination of U6 snrna and mir 192 expression levels. For normalization of the RT qpcr data to the sample volume, RNA was extracted from 500 µl of each pleural effusion sample and the Cq values were directly converted to the relative quantification values using the formula described above. Using this method, no significant differences in the levels of mir 198 expression were detected between the BPE and the LA MPE samples (P=0.178; Fig. 3A). However, when data were normalized to the expression levels of U6 snrna or mir 192, the mir 198 expression levels were significantly lower in the LA MPE samples than in the BPE samples (P<0.001 for U6 snrna and mir 192; Fig. 3B and C, respectively). Significant differences in the levels of mir 198 expression between BPE and LA MPE samples were also detected when the combination of U6 snrna and mir 192 expression levels served as a normalizer (P<0.001; Fig. 3D).
4 1892 HAN et al: REFERENCE GENES IN PLEURAL EFFUSION Table II. Details of candidate reference genes. Gene RNA species Accession number Function Reference PCR efficiency (%) mir 192 mirna MI Regulates dihydrofolate reductase and cellular proliferation mir 20a mirna MI Negatively regulates autophagy mir 221 mirna MI Regulates cell cycle mir 222 mirna MI Regulates cell cycle mir 16 mirna MI Induces apoptosis by targeting BCL U6 snrna snrna NR_ Guides post transcriptional modification of cellular RNAs RNU6B snrna NR_ Guides post transcriptional modification of cellular RNAs RNU44 snorna NR_ Guides site specific ribosomal 24 Undetected RNA modification RNU48 snorna NR_ Guides methylation of 28S 25 Undetected ribosomal RNA PCR, polymerase chain reaction; mirna, microrna; snrna, small nuclear RNA; snorna, small nucleolar RNA; BCL2, B cell lymphoma 2. Discussion To the best of our knowledge, the present study is the first systematic investigation of suitable reference genes for the relative quantification of mirnas in pleural effusion. In the present study, four strategies were used to identify suitable reference genes. mirna microarray analyses of BPE and MPE samples were performed to identify the invariant mirnas that are stably expressed. These candidate reference mirnas, as well as the U6 snrna, RNU6B, RNU44 and RNU48 small RNAs, which are the most frequently used reference genes in mirna expression studies (10,11,26), were validated by RT qpcr. The statistical algorithms NormFinder and BestKeeper were used to identify the most useful endogenous reference genes for relative quantification. The impact of different normalization approaches on the ability to detect reduced mir 198 expression levels in LA MPE samples was also examined. The results emphasize the importance of an appropriate normalization approach to identify U6 snrna and mir 192 as suitable reference genes for mirna analyses in pleural effusion samples. The selection of appropriate reference genes as normalizers for the relative quantification of mirna expression levels is required to avoid erroneous results and to improve the comparability of mirna expression level data amongst studies (10). The conventional approach to mirna RT qpcr data analysis is to normalize the values to the expression levels of snrnas, such as U6 snrna or RNU6B, or snornas, such as RNU44 or RNU48. Alternatively, synthetic mirnas, including ath mir 156a or ath mir 159a, are commonly added to the samples and used as normalizers. However, these mirnas may exhibit variable expression levels depending on the samples used and may introduce noise when used as internal controls (11 15). Due to differences in length, the physicochemical properties, isolation efficiencies and degradation rates of small RNAs differ from Figure 1. Linear correlation between Cq values and log concentration of mir 192 in a serially diluted pleural effusion sample. mir 192 expression levels were measured in 10 fold serial dilutions of the pleural effusion sample. The concentration of undiluted pleural effusion was arbitrarily designated as 1. Each point represents the mean of duplicate measurements. The regression curve of Cq values versus log concentration reveals a linear correlation (R 2 =0.9863) for Cq values between 31 and 44. those of mirnas (26). In the present study, RNU6B was not detected in the majority of pleural effusion samples; the poor quality of RNU6B as a reference gene has previously been reported in mirna expression studies of several other types of malignancy (11 14). Unlike mirnas, RNU6B is a high molecular weight RNA that lacks resistance to degradation by RNase in body fluids (27); these properties may account for the low abundance of RNU6B in the pleural effusion samples included in the present study. Recent studies have also reported that snornas exhibit highly variable levels of expression, and the use of these species as reference genes may introduce noise and result in inaccuracies in the data (15,28). Furthermore, the expression levels of snornas have been associated with cancer prognosis in multiple studies (15,29,30). This evidence demonstrates that these
5 ONCOLOGY LETTERS 8: , Table III. Ranking and best combination of candidate reference genes in pleural effusion samples as determined by the expression stability values calculated using NormFinder and BestKeeper. NormFinder BestKeeper Rank Gene Stability value a Gene Stability value a 1 U6 snrna mir mir U6 snrna mir-20a mir-20a mir mir mir mir mir mir Best combination U6 snrna/mir a High expression stability is indicated by a low stability value. snrna, small nuclear RNA; mirna, microrna. A B Figure 2. Expression levels of candidate reference genes in benign plural effusion (BPE) and lung adenocarcinoma associated malignant pleural effusion (LA MPE) samples. Reverse transcription quantitative polymerase chain reaction (RT qpcr) analyses of 46 BPE and 45 LA MPE samples were performed. (A) Box and whisker plot of the primary Cq values of U6 snrna and five candidate reference mirnas (mir 192, mir 20a, mir 221, mir 222 and mir 16) in the 91 pleural effusion samples. (B) Box and whisker plot of the Cq values of candidate reference genes in the BPE and LA MPE samples. The expression levels of all candidate reference genes did not differ significantly between BPE and LA MPE samples (P>0.05). The Cq values were corrected to PCR efficiency and two inter plate controls. The boxes (light gray, BPE and dark gray, LA MPE) signify the upper and lower quartiles, the median is indicated by a horizontal line, and the whiskers depict the 10th and 90th percentiles. The P values of the Mann Whiney U tests are indicated above each plot. molecules do not meet the requirement that reference genes be unaffected by biological changes, disease or treatments. Non human synthetic mirna also frequently serves as a spike in control to normalize for technical variability (31). In two recent studies that attempted to identify potential diagnostic or prognostic mirna biomarkers in pleural effusion, the relative mirna expression levels were directly normalized using ath mir156a as a spiked in reference gene (8,16). However, spike in controls may not correct for the variability arising from differences in template quality and/or the efficiency of the reverse transcription reaction. Therefore, candidate reference genes must be validated for each individual study design, as even frequently used reference genes are unstable under certain conditions. Thus, the identification of a reference gene that is most suited to the specific samples and experimental conditions of the study is recommended. In the current study, the candidate reference genes identified by the microarray and subsequent RT qpcr experiments were analyzed using the NormFinder and BestKeeper algorithms. NormFinder calculated the intra and inter group variations and identified U6 snrna as the single most stable reference gene with a stability value of The use of more than one reference gene is generally considered to increase the accuracy of mirna quantification, and the combination of U6 snrna and mir 192 expression exhibited an improved NormFinder stability value of NormFinder and BestKeeper did not produce the same reference gene ranking order for normalization; this difference may be attributable to the different calculation models used in the tools. However, NormFinder and BestKeeper ranked U6 snrna and mir 192 as the top two reference genes. The impact of different normalization strategies on the accuracy of RT qpcr results was examined by analyzing the expression levels of cell free mir 198 in pleural effusions. In a previous study, the levels of mir 198 expression were found to be significantly lower in LA MPE samples than in BPE samples, indicating that this mirna may have diagnostic potential for differentiating BPE and MPE (9). In the present study, the relative quantification of mir 198 was assessed using different normalization strategies. When
6 1894 HAN et al: REFERENCE GENES IN PLEURAL EFFUSION A B C D Figure 3. Effects of different normalization methods on the measurement of mir 198 expression levels. (A) No significant differences in the mir 198 expression levels in pleural effusion samples were detected when the data were normalized to serum volume. When the data were normalized to (B) U6 small nuclear RNA expression levels, (C) mir 192 expression levels and (D) a combination of U6 snrna and mir 192 expression levels, significant differences between benign pleural effusion (BPE) and lung adenocarcinoma associated malignant pleural effusion (LA MPE) samples were observed (P<0.001). The boxes signify the upper and lower quartiles, the median is indicated by a horizontal line, and the whiskers depict the 10th and 90th percentiles. The Mann Whiney U test P values are indicated in the upper right corner of each plot. the mir 198 data were normalized to the sample volume, no significant differences between BPE and LA MPE samples were detected. However, when the data were normalized to U6 snrna expression levels, mir 192 expression levels, and a combination of U6 snrna and mir 192 expression levels, significant differences between BPE and LA MPE samples were detected. These results indicate that normalization of mirna expression level data to those of systematically selected reference genes detects smaller differences than those identified by normalization of the data to the sample volume. The present study has certain limitations. A relatively small number of well known mirnas (160 of the >1,000 currently available) were screened using the PNA based microarray. Furthermore, the number of candidate reference genes examined was small; the use of multiple reference genes is critical to achieve more reliable expression data (32). Another limitation of the study was that the identification of U6 snrna and mir 192 as reference genes was only confirmed by normalization of the expression levels of a single mirna (mir 198) in the pleural effusion samples. If further studies investigating mirnas as potential diagnostic, prognostic and predictive biomarkers in pleural effusion are to be undertaken, the use of U6 snrna and mir 192 as reference genes must be further validated. Another issue is that all MPE samples used in the present study were obtained from lung adenocarcinoma specimens; whether U6 snrna and mir 192 may be used as reference genes for other types of MPE remains to be determined. In conclusion, the use of the expression levels of U6 snrna, mir 192 or a combination of these genes for the relative quantification of mirna expression levels in pleural effusion is recommended, as determined by the results from the present study. Acknowledgements This study was supported by a Basic Science Research Program through the National Research Foundation of Korea, funded by the Ministry of Education, Science and Technology (grant no ). All effusion samples examined in this study were provided by Chungbuk National University Hospital and Kangwon National University Hospital, members of the National Biobank of Korea, which is supported by the Ministry of Health, Welfare and Family Affairs. All samples derived from the National Biobank of Korea were obtained with informed consent under procedures approved by the institutional review board. The authors appreciate the assistance of the cancer research team at the Bioevaluation Center (Korea Research Institute of Bioscience and Biotechnology, Cheongju, Republic of Korea). References 1. Bartel DP: MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116: , Cho WC: MicroRNAs: potential biomarkers for cancer diagnosis, prognosis and targets for therapy. Int J Biochem Cell Biol 42: , Zhou L, Zhao YP, Liu WJ, et al: Circulating micrornas in cancer: diagnostic and prognostic significance. Expert Rev Anticancer Ther 12: , Cortez MA, Bueso Ramos C, Ferdin J, et al: MicroRNAs in body fluids the mix of hormones and biomarkers. Nat Rev Clin Oncol 8: , 2011.
7 ONCOLOGY LETTERS 8: , Tsim S, O'Dowd CA, Milroy R and Davidson S: Staging of non small cell lung cancer (NSCLC): a review. Respir Med 104: , Nance KV, Shermer RW and Askin FB: Diagnostic efficacy of pleural biopsy as compared with that of pleural fluid examination. Mod Pathol 4: , Xie L, Chen X, Wang L, et al: Cell free mirnas may indicate diagnosis and docetaxel sensitivity of tumor cells in malignant effusions. BMC Cancer 10: 591, Xie L, Wang T, Yu S, et al: Cell free mir 24 and mir 30d, potential diagnostic biomarkers in malignant effusions. Clin Biochem 44: , Han HS, Yun J, Lim SN, et al: Downregulation of cell free mir 198 as a diagnostic biomarker for lung adenocarcinoma associated malignant pleural effusion. Int J Cancer 133: , Mestdagh P, Van Vlierberghe P, De Weer A, et al: A novel and universal method for microrna RT qpcr data normalization. Genome Biol 10: R64, Ratert N, Meyer HA, Jung M, et al: Reference mirnas for mirnaome analysis of urothelial carcinomas. PLoS One 7: e39309, Song J, Bai Z, Han W, et al: Identification of suitable reference genes for qpcr analysis of serum microrna in gastric cancer patients. Dig Dis Sci 57: , Wotschofsky Z, Meyer HA, Jung M, et al: Reference genes for the relative quantification of micrornas in renal cell carcinomas and their metastases. Anal Biochem 417: , Chang KH, Mestdagh P, Vandesompele J, Kerin MJ and Miller N: MicroRNA expression profiling to identify and validate reference genes for relative quantification in colorectal cancer. BMC Cancer 10: 173, Gee HE, Buffa FM, Camps C, et al: The small nucleolar RNAs commonly used for microrna normalisation correlate with tumour pathology and prognosis. Br J Cancer 104: , Wang T, Lv M, Shen S, et al: Cell free microrna expression profiles in malignant effusion associated with patient survival in non small cell lung cancer. PLoS One 7: e43268, Andersen CL, Jensen JL and Ørntoft TF: Normalization of real time quantitative reverse transcription PCR data: a model based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Cancer Res 64: , Pfaffl MW, Tichopad A, Prgomet C and Neuvians TP: Determination of stable housekeeping genes, differentially regulated target genes and sample integrity: BestKeeper Excel based tool using pair wise correlations. Biotechnol Lett 26: , Song B, Wang Y, Kudo K, et al: mir 192 Regulates Dihydrofolate Reductase and Cellular Proliferation through the p53 microrna Circuit. Clin Cancer Res 14: , Wu H, Wang F, Hu S, et al: MiR 20a and mir 106b negatively regulate autophagy induced by leucine deprivation via suppression of ULK1 expression in C2C12 myoblasts. Cell Signal 24: , Visone R, Russo L, Pallante P, et al: MicroRNAs (mir) 221 and mir 222, both overexpressed in human thyroid papillary carcinomas, regulate p27kip1 protein levels and cell cycle. Endocr Relat Cancer 14: , Cimmino A, Calin GA, Fabbri M, et al: mir 15 and mir 16 induce apoptosis by targeting BCL2. Proc Natl Acad Sci USA 102: , Kiss T: Small nucleolar RNA guided post transcriptional modification of cellular RNAs. EMBO J 20: , Dieci G, Preti M and Montanini B: Eukaryotic snornas: a paradigm for gene expression flexibility. Genomics 94: 83 88, Galardi S, Fatica A, Bachi A, et al: Purified box C/D snornps are able to reproduce site specific 2' O methylation of target RNA in vitro. Mol Cell Biol 22: , Peltier HJ and Latham GJ: Normalization of microrna expression levels in quantitative RT PCR assays: identification of suitable reference RNA targets in normal and cancerous human solid tissues. RNA 14: , Chen X, Ba Y, Ma L, et al: Characterization of micrornas in serum: a novel class of biomarkers for diagnosis of cancer and other diseases. Cell Res 18: , Git A, Dvinge H, Salmon Divon M, et al: Systematic comparison of microarray profiling, real time PCR, and next generation sequencing technologies for measuring differential microrna expression. RNA 16: , Dong XY, Guo P, Boyd J, et al: Implication of snorna U50 in human breast cancer. J Genet Genomics 36: , Mourtada-Maarabouni M, Pickard MR, Hedge VL, Farzaneh F and Williams GT: GAS5, a non-protein-coding RNA, controls apoptosis and is downregulated in breast cancer. Oncogene 28: , Kroh EM, Parkin RK, Mitchell PS and Tewari M: Analysis of circulating microrna biomarkers in plasma and serum using quantitative reverse transcription PCR (qrt PCR). Methods 50: , Tricarico C, Pinzani P, Bianchi S, et al: Quantitative real time reverse transcription polymerase chain reaction: normalization to rrna or single housekeeping genes is inappropriate for human tissue biopsies. Anal Biochem 309: , 2002.
Downregulation of serum mir-17 and mir-106b levels in gastric cancer and benign gastric diseases
 Brief Communication Downregulation of serum mir-17 and mir-106b levels in gastric cancer and benign gastric diseases Qinghai Zeng 1 *, Cuihong Jin 2 *, Wenhang Chen 2, Fang Xia 3, Qi Wang 3, Fan Fan 4,
Brief Communication Downregulation of serum mir-17 and mir-106b levels in gastric cancer and benign gastric diseases Qinghai Zeng 1 *, Cuihong Jin 2 *, Wenhang Chen 2, Fang Xia 3, Qi Wang 3, Fan Fan 4,
Identification of Suitable Reference Genes for qpcr Analysis of Serum microrna in Gastric Cancer Patients
 DOI 10.1007/s10620-011-1981-7 ORIGINAL ARTICLE Identification of Suitable Reference Genes for qpcr Analysis of Serum microrna in Gastric Cancer Patients Jianning Song Zhigang Bai Wei Han Jun Zhang Hua
DOI 10.1007/s10620-011-1981-7 ORIGINAL ARTICLE Identification of Suitable Reference Genes for qpcr Analysis of Serum microrna in Gastric Cancer Patients Jianning Song Zhigang Bai Wei Han Jun Zhang Hua
From reference genes to global mean normalization
 From reference genes to global mean normalization Jo Vandesompele professor, Ghent University co-founder and CEO, Biogazelle qpcr Symposium USA November 9, 2009 Millbrae, CA outline what is normalization
From reference genes to global mean normalization Jo Vandesompele professor, Ghent University co-founder and CEO, Biogazelle qpcr Symposium USA November 9, 2009 Millbrae, CA outline what is normalization
A novel and universal method for microrna RT-qPCR data normalization
 A novel and universal method for microrna RT-qPCR data normalization Jo Vandesompele professor, Ghent University co-founder and CEO, Biogazelle 4 th International qpcr Symposium Weihenstephan, March 1,
A novel and universal method for microrna RT-qPCR data normalization Jo Vandesompele professor, Ghent University co-founder and CEO, Biogazelle 4 th International qpcr Symposium Weihenstephan, March 1,
microrna Presented for: Presented by: Date:
 microrna Presented for: Presented by: Date: 2 micrornas Non protein coding, endogenous RNAs of 21-22nt length Evolutionarily conserved Regulate gene expression by binding complementary regions at 3 regions
microrna Presented for: Presented by: Date: 2 micrornas Non protein coding, endogenous RNAs of 21-22nt length Evolutionarily conserved Regulate gene expression by binding complementary regions at 3 regions
Profiles of gene expression & diagnosis/prognosis of cancer. MCs in Advanced Genetics Ainoa Planas Riverola
 Profiles of gene expression & diagnosis/prognosis of cancer MCs in Advanced Genetics Ainoa Planas Riverola Gene expression profiles Gene expression profiling Used in molecular biology, it measures the
Profiles of gene expression & diagnosis/prognosis of cancer MCs in Advanced Genetics Ainoa Planas Riverola Gene expression profiles Gene expression profiling Used in molecular biology, it measures the
Title: Dysregulated mir-183 Inhibits Migration in Breast Cancer
 Author's response to reviews Title: Dysregulated mir-183 Inhibits Migration in Breast Cancer Authors: Aoife J Lowery (aoife.lowery@gmail.com) Nicola Miller (nicola.miller@nuigalway.ie) Roisin M Dwyer (roisin.dwyer@nuigalway.ie)
Author's response to reviews Title: Dysregulated mir-183 Inhibits Migration in Breast Cancer Authors: Aoife J Lowery (aoife.lowery@gmail.com) Nicola Miller (nicola.miller@nuigalway.ie) Roisin M Dwyer (roisin.dwyer@nuigalway.ie)
Supplementary webappendix
 Supplementary webappendix This webappendix formed part of the original submission and has been peer reviewed. We post it as supplied by the authors. Supplement to: Kratz JR, He J, Van Den Eeden SK, et
Supplementary webappendix This webappendix formed part of the original submission and has been peer reviewed. We post it as supplied by the authors. Supplement to: Kratz JR, He J, Van Den Eeden SK, et
Mir-595 is a significant indicator of poor patient prognosis in epithelial ovarian cancer
 European Review for Medical and Pharmacological Sciences 2017; 21: 4278-4282 Mir-595 is a significant indicator of poor patient prognosis in epithelial ovarian cancer Q.-H. ZHOU 1, Y.-M. ZHAO 2, L.-L.
European Review for Medical and Pharmacological Sciences 2017; 21: 4278-4282 Mir-595 is a significant indicator of poor patient prognosis in epithelial ovarian cancer Q.-H. ZHOU 1, Y.-M. ZHAO 2, L.-L.
Patnaik SK, et al. MicroRNAs to accurately histotype NSCLC biopsies
 Patnaik SK, et al. MicroRNAs to accurately histotype NSCLC biopsies. 2014. Supplemental Digital Content 1. Appendix 1. External data-sets used for associating microrna expression with lung squamous cell
Patnaik SK, et al. MicroRNAs to accurately histotype NSCLC biopsies. 2014. Supplemental Digital Content 1. Appendix 1. External data-sets used for associating microrna expression with lung squamous cell
RNA preparation from extracted paraffin cores:
 Supplementary methods, Nielsen et al., A comparison of PAM50 intrinsic subtyping with immunohistochemistry and clinical prognostic factors in tamoxifen-treated estrogen receptor positive breast cancer.
Supplementary methods, Nielsen et al., A comparison of PAM50 intrinsic subtyping with immunohistochemistry and clinical prognostic factors in tamoxifen-treated estrogen receptor positive breast cancer.
Serum mirna expression profile as a prognostic biomarker of stage II/III
 Serum mirna expression profile as a prognostic biomarker of stage II/III colorectal adenocarcinoma Jialu Li a,b,, Yang Liu c,, Cheng Wang d,, Ting Deng a,, Hongwei Liang b, Yifei Wang e, Dingzhi Huang
Serum mirna expression profile as a prognostic biomarker of stage II/III colorectal adenocarcinoma Jialu Li a,b,, Yang Liu c,, Cheng Wang d,, Ting Deng a,, Hongwei Liang b, Yifei Wang e, Dingzhi Huang
Low levels of serum mir-99a is a predictor of poor prognosis in breast cancer
 Low levels of serum mir-99a is a predictor of poor prognosis in breast cancer J. Li 1, Z.J. Song 2, Y.Y. Wang 1, Y. Yin 1, Y. Liu 1 and X. Nan 1 1 Tumor Research Department, Shaanxi Provincial Tumor Hospital,
Low levels of serum mir-99a is a predictor of poor prognosis in breast cancer J. Li 1, Z.J. Song 2, Y.Y. Wang 1, Y. Yin 1, Y. Liu 1 and X. Nan 1 1 Tumor Research Department, Shaanxi Provincial Tumor Hospital,
A two-microrna signature in urinary exosomes for diagnosis of prostate cancer
 Poster # B4 A two-microrna signature in urinary exosomes for diagnosis of prostate cancer Anne Karin Ildor Rasmussen 1, Peter Mouritzen 1, Karina Dalsgaard Sørensen 3, Thorarinn Blondal 1, Jörg Krummheuer
Poster # B4 A two-microrna signature in urinary exosomes for diagnosis of prostate cancer Anne Karin Ildor Rasmussen 1, Peter Mouritzen 1, Karina Dalsgaard Sørensen 3, Thorarinn Blondal 1, Jörg Krummheuer
Expression of mir-1294 is downregulated and predicts a poor prognosis in gastric cancer
 European Review for Medical and Pharmacological Sciences 2018; 22: 5525-5530 Expression of mir-1294 is downregulated and predicts a poor prognosis in gastric cancer Y.-X. SHI, B.-L. YE, B.-R. HU, X.-J.
European Review for Medical and Pharmacological Sciences 2018; 22: 5525-5530 Expression of mir-1294 is downregulated and predicts a poor prognosis in gastric cancer Y.-X. SHI, B.-L. YE, B.-R. HU, X.-J.
Reference mirnas for mirnaome Analysis of Urothelial Carcinomas
 Reference mirnas for mirnaome Analysis of Urothelial Carcinomas Nadine Ratert 1,2, Hellmuth-Alexander Meyer 1,3, Monika Jung 1, Hans-Joachim Mollenkopf 4, Ina Wagner 4, Kurt Miller 1, Ergin Kilic 5, Andreas
Reference mirnas for mirnaome Analysis of Urothelial Carcinomas Nadine Ratert 1,2, Hellmuth-Alexander Meyer 1,3, Monika Jung 1, Hans-Joachim Mollenkopf 4, Ina Wagner 4, Kurt Miller 1, Ergin Kilic 5, Andreas
RNA extraction, RT-PCR and real-time PCR. Total RNA were extracted using
 Supplementary Information Materials and Methods RNA extraction, RT-PCR and real-time PCR. Total RNA were extracted using Trizol reagent (Invitrogen,Carlsbad, CA) according to the manufacturer's instructions.
Supplementary Information Materials and Methods RNA extraction, RT-PCR and real-time PCR. Total RNA were extracted using Trizol reagent (Invitrogen,Carlsbad, CA) according to the manufacturer's instructions.
mir-218 tissue expression level is associated with aggressive progression of gastric cancer
 mir-218 tissue expression level is associated with aggressive progression of gastric cancer X.X. Wang 1, S.J. Ge 2, X.L. Wang 2, L.X. Jiang 1, M.F. Sheng 2 and J.J. Ma 2 1 Department of General Surgery,
mir-218 tissue expression level is associated with aggressive progression of gastric cancer X.X. Wang 1, S.J. Ge 2, X.L. Wang 2, L.X. Jiang 1, M.F. Sheng 2 and J.J. Ma 2 1 Department of General Surgery,
Cellecta Overview. Started Operations in 2007 Headquarters: Mountain View, CA
 Cellecta Overview Started Operations in 2007 Headquarters: Mountain View, CA Focus: Development of flexible, scalable, and broadly parallel genetic screening assays to expedite the discovery and characterization
Cellecta Overview Started Operations in 2007 Headquarters: Mountain View, CA Focus: Development of flexible, scalable, and broadly parallel genetic screening assays to expedite the discovery and characterization
MicroRNA-92gene expression in epithelial ovarian cancer using a novel Real-Time Polymerase change reaction
 www.muthjm.com Muthanna Medical Journal 2016; 3(1):23-33 MicroRNA-92gene expression in epithelial ovarian cancer using a novel Real-Time Polymerase change reaction Shoroq Mohammed AL-Temimi 1* Abstract
www.muthjm.com Muthanna Medical Journal 2016; 3(1):23-33 MicroRNA-92gene expression in epithelial ovarian cancer using a novel Real-Time Polymerase change reaction Shoroq Mohammed AL-Temimi 1* Abstract
Reduced mirna-218 expression in pancreatic cancer patients as a predictor of poor prognosis
 Reduced mirna-218 expression in pancreatic cancer patients as a predictor of poor prognosis B.-S. Li, H. Liu and W.-L. Yang Department of Gastrointestinal and Pancreatic Surgery, The Third Xiangya Hospital
Reduced mirna-218 expression in pancreatic cancer patients as a predictor of poor prognosis B.-S. Li, H. Liu and W.-L. Yang Department of Gastrointestinal and Pancreatic Surgery, The Third Xiangya Hospital
Evaluation of altered expression of mir-9 and mir-106a as an early diagnostic approach in gastric cancer
 Original Article Evaluation of altered expression of mir-9 and mir-106a as an early diagnostic approach in gastric cancer Khadije Shirmohammadi, Sareh Sohrabi, Zahra Jafarzadeh Samani, Hosein Effatpanah,
Original Article Evaluation of altered expression of mir-9 and mir-106a as an early diagnostic approach in gastric cancer Khadije Shirmohammadi, Sareh Sohrabi, Zahra Jafarzadeh Samani, Hosein Effatpanah,
Molecular in vitro diagnostic test for the quantitative detection of the mrna expression of ERBB2, ESR1, PGR and MKI67 in breast cancer tissue.
 Innovation for your breast cancer diagnostics PGR G ATA G C G A C G AT C G A A A G A A G T TA G ATA G C G A C G AT C G A A A G A A G T TA G ATA G C G A C G AT C G A A A G A A G T TA G ATA G C G A C ERBB2
Innovation for your breast cancer diagnostics PGR G ATA G C G A C G AT C G A A A G A A G T TA G ATA G C G A C G AT C G A A A G A A G T TA G ATA G C G A C G AT C G A A A G A A G T TA G ATA G C G A C ERBB2
Association of mir-21 with esophageal cancer prognosis: a meta-analysis
 Association of mir-21 with esophageal cancer prognosis: a meta-analysis S.-W. Wen 1, Y.-F. Zhang 1, Y. Li 1, Z.-X. Liu 2, H.-L. Lv 1, Z.-H. Li 1, Y.-Z. Xu 1, Y.-G. Zhu 1 and Z.-Q. Tian 1 1 Department of
Association of mir-21 with esophageal cancer prognosis: a meta-analysis S.-W. Wen 1, Y.-F. Zhang 1, Y. Li 1, Z.-X. Liu 2, H.-L. Lv 1, Z.-H. Li 1, Y.-Z. Xu 1, Y.-G. Zhu 1 and Z.-Q. Tian 1 1 Department of
microrna analysis for the detection of autologous blood transfusion in doping control
 microrna analysis for the detection of autologous blood transfusion in doping control Application note Doping Control Authors Francesco Donati*, Fiorella Boccia, Xavier de la Torre, Alessandra Stampella,
microrna analysis for the detection of autologous blood transfusion in doping control Application note Doping Control Authors Francesco Donati*, Fiorella Boccia, Xavier de la Torre, Alessandra Stampella,
MicroRNA expression profiling and functional analysis in prostate cancer. Marco Folini s.c. Ricerca Traslazionale DOSL
 MicroRNA expression profiling and functional analysis in prostate cancer Marco Folini s.c. Ricerca Traslazionale DOSL What are micrornas? For almost three decades, the alteration of protein-coding genes
MicroRNA expression profiling and functional analysis in prostate cancer Marco Folini s.c. Ricerca Traslazionale DOSL What are micrornas? For almost three decades, the alteration of protein-coding genes
developing new tools for diagnostics Join forces with IMGM Laboratories to make your mirna project a success
 micrornas developing new tools for diagnostics Join forces with IMGM Laboratories to make your mirna project a success Dr. Carola Wagner IMGM Laboratories GmbH Martinsried, Germany qpcr 2009 Symposium
micrornas developing new tools for diagnostics Join forces with IMGM Laboratories to make your mirna project a success Dr. Carola Wagner IMGM Laboratories GmbH Martinsried, Germany qpcr 2009 Symposium
MEDICAL POLICY Gene Expression Profiling for Cancers of Unknown Primary Site
 POLICY: PG0364 ORIGINAL EFFECTIVE: 04/22/16 LAST REVIEW: 07/26/18 MEDICAL POLICY Gene Expression Profiling for Cancers of Unknown Primary Site GUIDELINES This policy does not certify benefits or authorization
POLICY: PG0364 ORIGINAL EFFECTIVE: 04/22/16 LAST REVIEW: 07/26/18 MEDICAL POLICY Gene Expression Profiling for Cancers of Unknown Primary Site GUIDELINES This policy does not certify benefits or authorization
mir-125a-5p expression is associated with the age of breast cancer patients
 mir-125a-5p expression is associated with the age of breast cancer patients H. He 1 *, F. Xu 2 *, W. Huang 1, S.Y. Luo 1, Y.T. Lin 1, G.H. Zhang 1, Q. Du 1 and R.H. Duan 1 1 The State Key Laboratory of
mir-125a-5p expression is associated with the age of breast cancer patients H. He 1 *, F. Xu 2 *, W. Huang 1, S.Y. Luo 1, Y.T. Lin 1, G.H. Zhang 1, Q. Du 1 and R.H. Duan 1 1 The State Key Laboratory of
Research Article Malignancy Associated MicroRNA Expression Changes in Canine Mammary Cancer of Different Malignancies
 ISRN Veterinary Science, Article ID 148597, 5 pages http://dx.doi.org/10.1155/2014/148597 Research Article Malignancy Associated MicroRNA Expression Changes in Canine Mammary Cancer of Different Malignancies
ISRN Veterinary Science, Article ID 148597, 5 pages http://dx.doi.org/10.1155/2014/148597 Research Article Malignancy Associated MicroRNA Expression Changes in Canine Mammary Cancer of Different Malignancies
Decreased expression of mir-490-3p in osteosarcoma and its clinical significance
 European Review for Medical and Pharmacological Sciences Decreased expression of mir-490-3p in osteosarcoma and its clinical significance B. TANG, C. LIU, Q.-M. ZHANG, M. NI Department of Orthopedics,
European Review for Medical and Pharmacological Sciences Decreased expression of mir-490-3p in osteosarcoma and its clinical significance B. TANG, C. LIU, Q.-M. ZHANG, M. NI Department of Orthopedics,
Original Article Up-regulation of mir-10a and down-regulation of mir-148b serve as potential prognostic biomarkers for osteosarcoma
 Int J Clin Exp Pathol 2016;9(1):186-190 www.ijcep.com /ISSN:1936-2625/IJCEP0018032 Original Article Up-regulation of mir-10a and down-regulation of mir-148b serve as potential prognostic biomarkers for
Int J Clin Exp Pathol 2016;9(1):186-190 www.ijcep.com /ISSN:1936-2625/IJCEP0018032 Original Article Up-regulation of mir-10a and down-regulation of mir-148b serve as potential prognostic biomarkers for
Only Estrogen receptor positive is not enough to predict the prognosis of breast cancer
 Young Investigator Award, Global Breast Cancer Conference 2018 Only Estrogen receptor positive is not enough to predict the prognosis of breast cancer ㅑ Running head: Revisiting estrogen positive tumors
Young Investigator Award, Global Breast Cancer Conference 2018 Only Estrogen receptor positive is not enough to predict the prognosis of breast cancer ㅑ Running head: Revisiting estrogen positive tumors
microrna PCR System (Exiqon), following the manufacturer s instructions. In brief, 10ng of
 SUPPLEMENTAL MATERIALS AND METHODS Quantitative RT-PCR Quantitative RT-PCR analysis was performed using the Universal mircury LNA TM microrna PCR System (Exiqon), following the manufacturer s instructions.
SUPPLEMENTAL MATERIALS AND METHODS Quantitative RT-PCR Quantitative RT-PCR analysis was performed using the Universal mircury LNA TM microrna PCR System (Exiqon), following the manufacturer s instructions.
A circulating serum mirna panel as early detection biomarkers of cervical intraepithelial neoplasia
 European Review for Medical and Pharmacological Sciences 2016; 20: 4846-4851 A circulating serum mirna panel as early detection biomarkers of cervical intraepithelial neoplasia F. XIN 1, P. LIU 1, C-F.
European Review for Medical and Pharmacological Sciences 2016; 20: 4846-4851 A circulating serum mirna panel as early detection biomarkers of cervical intraepithelial neoplasia F. XIN 1, P. LIU 1, C-F.
Association between downexpression of mir-1301 and poor prognosis in patients with glioma
 European Review for Medical and Pharmacological Sciences 2017; 21: 4298-4303 Association between downexpression of mir-1301 and poor prognosis in patients with glioma Q.-L. BAI, C.-W. HU, X.-R. WANG, G.-F.
European Review for Medical and Pharmacological Sciences 2017; 21: 4298-4303 Association between downexpression of mir-1301 and poor prognosis in patients with glioma Q.-L. BAI, C.-W. HU, X.-R. WANG, G.-F.
Original Article High serum mir-203 predicts the poor prognosis in patients with pancreatic cancer
 Int J Clin Exp Pathol 2017;10(4):4688-4693 www.ijcep.com /ISSN:1936-2625/IJCEP0047128 Original Article High serum mir-203 predicts the poor prognosis in patients with pancreatic cancer Jun Ma, Xiaokun
Int J Clin Exp Pathol 2017;10(4):4688-4693 www.ijcep.com /ISSN:1936-2625/IJCEP0047128 Original Article High serum mir-203 predicts the poor prognosis in patients with pancreatic cancer Jun Ma, Xiaokun
Extracellular Vesicle RNA isolation Kits
 Extracellular Vesicle RNA isolation Kits Summary Section 4 Introduction 42 Exo-TotalRNA and TumorExo-TotalRNA isolation kits 43 Extracellular Vesicle RNA extraction kits Ordering information Products can
Extracellular Vesicle RNA isolation Kits Summary Section 4 Introduction 42 Exo-TotalRNA and TumorExo-TotalRNA isolation kits 43 Extracellular Vesicle RNA extraction kits Ordering information Products can
Diagnostic performance of microrna-29a for colorectal cancer: a meta-analysis
 Diagnostic performance of microrna-29a for colorectal cancer: a meta-analysis M.L. Zhi 1 *, Z.J. Liu 2 *, X.Y. Yi 1, L.J. Zhang 1 and Y.X. Bao 1 1 Department of Clinical Laboratory, The Second Affiliated
Diagnostic performance of microrna-29a for colorectal cancer: a meta-analysis M.L. Zhi 1 *, Z.J. Liu 2 *, X.Y. Yi 1, L.J. Zhang 1 and Y.X. Bao 1 1 Department of Clinical Laboratory, The Second Affiliated
Products for cfdna and mirna isolation. Subhead Circulating Cover nucleic acids from plasma
 MACHEREY-NAGEL Products for cfdna and mirna isolation Bioanalysis Subhead Circulating Cover nucleic acids from plasma n Flexible solutions for small and large blood plasma volumes n Highly efficient recovery
MACHEREY-NAGEL Products for cfdna and mirna isolation Bioanalysis Subhead Circulating Cover nucleic acids from plasma n Flexible solutions for small and large blood plasma volumes n Highly efficient recovery
ARTICLE IN PRESS. Critical Reviews in Oncology/Hematology xxx (2010) xxx xxx
 Critical Reviews in Oncology/Hematology xxx (2010) xxx xxx Circulating micrornas: Association with disease and potential use as biomarkers Glen Reid, Michaela B. Kirschner, Nico van Zandwijk Asbestos Diseases
Critical Reviews in Oncology/Hematology xxx (2010) xxx xxx Circulating micrornas: Association with disease and potential use as biomarkers Glen Reid, Michaela B. Kirschner, Nico van Zandwijk Asbestos Diseases
Clinical significance of serum mir-21 in breast cancer compared with CA153 and CEA
 Original Article Clinical significance of serum mir-21 in breast cancer compared with CA153 and CEA Jianjian Gao, Qingyun Zhang, Jianjun Xu, Lijuan Guo, Xuefeng Li Key Laboratory of Carcinogenesis and
Original Article Clinical significance of serum mir-21 in breast cancer compared with CA153 and CEA Jianjian Gao, Qingyun Zhang, Jianjun Xu, Lijuan Guo, Xuefeng Li Key Laboratory of Carcinogenesis and
Serum mir-200c expression level as a prognostic biomarker for gastric cancer
 Serum mir-200c expression level as a prognostic biomarker for gastric cancer H.-P. Zhang 1 *, F.-B. Sun 2 * and S.-J. Li 2 1 Department of Gastrointestinal Surgery, Yantai Yuhuangding Hospital, Yantai,
Serum mir-200c expression level as a prognostic biomarker for gastric cancer H.-P. Zhang 1 *, F.-B. Sun 2 * and S.-J. Li 2 1 Department of Gastrointestinal Surgery, Yantai Yuhuangding Hospital, Yantai,
Supplementary Information Titles Journal: Nature Medicine
 Supplementary Information Titles Journal: Nature Medicine Article Title: Corresponding Author: Supplementary Item & Number Supplementary Fig.1 Fig.2 Fig.3 Fig.4 Fig.5 Fig.6 Fig.7 Fig.8 Fig.9 Fig. Fig.11
Supplementary Information Titles Journal: Nature Medicine Article Title: Corresponding Author: Supplementary Item & Number Supplementary Fig.1 Fig.2 Fig.3 Fig.4 Fig.5 Fig.6 Fig.7 Fig.8 Fig.9 Fig. Fig.11
RECENT ADVANCES IN THE MOLECULAR DIAGNOSIS OF BREAST CANCER
 Technology Transfer in Diagnostic Pathology. 6th Central European Regional Meeting. Cytopathology. Balatonfüred, Hungary, April 7-9, 2011. RECENT ADVANCES IN THE MOLECULAR DIAGNOSIS OF BREAST CANCER Philippe
Technology Transfer in Diagnostic Pathology. 6th Central European Regional Meeting. Cytopathology. Balatonfüred, Hungary, April 7-9, 2011. RECENT ADVANCES IN THE MOLECULAR DIAGNOSIS OF BREAST CANCER Philippe
Molecular in vitro diagnostic test for the quantitative detection of the mrna expression of ERBB2, ESR1, PGR and MKI67 in breast cancer tissue.
 Innovation for your breast cancer diagnostics Now valid ated on: -Roche cob as z 480 A nalyzer -Roche L ightcycle r 480 II - ABI 750 0 Fast -Versant kpcr Cyc ler PGR ESR1 MKI67 Molecular in vitro diagnostic
Innovation for your breast cancer diagnostics Now valid ated on: -Roche cob as z 480 A nalyzer -Roche L ightcycle r 480 II - ABI 750 0 Fast -Versant kpcr Cyc ler PGR ESR1 MKI67 Molecular in vitro diagnostic
Phosphate buffered saline (PBS) for washing the cells TE buffer (nuclease-free) ph 7.5 for use with the PrimePCR Reverse Transcription Control Assay
 Catalog # Description 172-5080 SingleShot Cell Lysis Kit, 100 x 50 µl reactions 172-5081 SingleShot Cell Lysis Kit, 500 x 50 µl reactions For research purposes only. Introduction The SingleShot Cell Lysis
Catalog # Description 172-5080 SingleShot Cell Lysis Kit, 100 x 50 µl reactions 172-5081 SingleShot Cell Lysis Kit, 500 x 50 µl reactions For research purposes only. Introduction The SingleShot Cell Lysis
ncounter TM Analysis System
 ncounter TM Analysis System Molecules That Count TM www.nanostring.com Agenda NanoString Technologies History Introduction to the ncounter Analysis System CodeSet Design and Assay Principals System Performance
ncounter TM Analysis System Molecules That Count TM www.nanostring.com Agenda NanoString Technologies History Introduction to the ncounter Analysis System CodeSet Design and Assay Principals System Performance
Chapter 18. mirna Expression Profiling: From Reference Genes to Global Mean Normalization
 Chapter 18 mirna Expression Profiling: From Reference Genes to Global Mean Normalization Barbara D haene, Pieter Mestdagh, Jan Hellemans, and Jo Vandesompele Abstract MicroRNAs (mirnas) are an important
Chapter 18 mirna Expression Profiling: From Reference Genes to Global Mean Normalization Barbara D haene, Pieter Mestdagh, Jan Hellemans, and Jo Vandesompele Abstract MicroRNAs (mirnas) are an important
micrornas (mirna) and Biomarkers
 micrornas (mirna) and Biomarkers Small RNAs Make Big Splash mirnas & Genome Function Biomarkers in Cancer Future Prospects Javed Khan M.D. National Cancer Institute EORTC-NCI-ASCO November 2007 The Human
micrornas (mirna) and Biomarkers Small RNAs Make Big Splash mirnas & Genome Function Biomarkers in Cancer Future Prospects Javed Khan M.D. National Cancer Institute EORTC-NCI-ASCO November 2007 The Human
Expression of mir-148/152 Family as Potential Biomarkers in Non-Small-Cell Lung Cancer
 e-issn 1643-375 Med Sci Monit, 215; 21: 1155-1161 DOI: 1.12659/MSM.89294 Received: 214.11.3 Accepted: 214.11.13 Published: 215.4.23 Expression of mir-148/152 Family as Potential Biomarkers in Non-Small-Cell
e-issn 1643-375 Med Sci Monit, 215; 21: 1155-1161 DOI: 1.12659/MSM.89294 Received: 214.11.3 Accepted: 214.11.13 Published: 215.4.23 Expression of mir-148/152 Family as Potential Biomarkers in Non-Small-Cell
EXOSOMES & MICROVESICLES
 The Sample Preparation Experts EXOSOMES & MICROVESICLES The New Standard in Exosome Purification and RNA Isolation Best-in-Class, Pure & Simple Exosome Purification, Fractionation of Exosomal Free & Circulating
The Sample Preparation Experts EXOSOMES & MICROVESICLES The New Standard in Exosome Purification and RNA Isolation Best-in-Class, Pure & Simple Exosome Purification, Fractionation of Exosomal Free & Circulating
EXOTESTTM. ELISA assay for exosome capture, quantification and characterization from cell culture supernatants and biological fluids
 DATA SHEET EXOTESTTM ELISA assay for exosome capture, quantification and characterization from cell culture supernatants and biological fluids INTRODUCTION Exosomes are small endosome-derived lipid nanoparticles
DATA SHEET EXOTESTTM ELISA assay for exosome capture, quantification and characterization from cell culture supernatants and biological fluids INTRODUCTION Exosomes are small endosome-derived lipid nanoparticles
Research Article Plasma HULC as a Promising Novel Biomarker for the Detection of Hepatocellular Carcinoma
 BioMed Research International Volume 213, Article ID 13616, 5 pages http://dx.doi.org/1.1155/213/13616 Research Article Plasma HULC as a Promising Novel Biomarker for the Detection of Hepatocellular Carcinoma
BioMed Research International Volume 213, Article ID 13616, 5 pages http://dx.doi.org/1.1155/213/13616 Research Article Plasma HULC as a Promising Novel Biomarker for the Detection of Hepatocellular Carcinoma
CircHIPK3 is upregulated and predicts a poor prognosis in epithelial ovarian cancer
 European Review for Medical and Pharmacological Sciences 2018; 22: 3713-3718 CircHIPK3 is upregulated and predicts a poor prognosis in epithelial ovarian cancer N. LIU 1, J. ZHANG 1, L.-Y. ZHANG 1, L.
European Review for Medical and Pharmacological Sciences 2018; 22: 3713-3718 CircHIPK3 is upregulated and predicts a poor prognosis in epithelial ovarian cancer N. LIU 1, J. ZHANG 1, L.-Y. ZHANG 1, L.
95% CI: , P=0.037).
 Biomedical Research 2017; 28 (21): 9248-9252 Reduced expression level of mir-205 predicts poor prognosis in Qiang Yang 1*#, Peng Sun 2#, Haotian Feng 1, Xin Li 1, Jianmin Li 1 1 Department of Orthopedics,
Biomedical Research 2017; 28 (21): 9248-9252 Reduced expression level of mir-205 predicts poor prognosis in Qiang Yang 1*#, Peng Sun 2#, Haotian Feng 1, Xin Li 1, Jianmin Li 1 1 Department of Orthopedics,
Analysis of small RNAs from Drosophila Schneider cells using the Small RNA assay on the Agilent 2100 bioanalyzer. Application Note
 Analysis of small RNAs from Drosophila Schneider cells using the Small RNA assay on the Agilent 2100 bioanalyzer Application Note Odile Sismeiro, Jean-Yves Coppée, Christophe Antoniewski, and Hélène Thomassin
Analysis of small RNAs from Drosophila Schneider cells using the Small RNA assay on the Agilent 2100 bioanalyzer Application Note Odile Sismeiro, Jean-Yves Coppée, Christophe Antoniewski, and Hélène Thomassin
The detection of differentially expressed micrornas from the serum of ovarian cancer patients using a novel real-time PCR platform
 Available online at www.sciencedirect.com Gynecologic Oncology 112 (2009) 55 59 www.elsevier.com/locate/ygyno The detection of differentially expressed micrornas from the serum of ovarian cancer patients
Available online at www.sciencedirect.com Gynecologic Oncology 112 (2009) 55 59 www.elsevier.com/locate/ygyno The detection of differentially expressed micrornas from the serum of ovarian cancer patients
RNA-Seq profiling of circular RNAs in human colorectal Cancer liver metastasis and the potential biomarkers
 Xu et al. Molecular Cancer (2019) 18:8 https://doi.org/10.1186/s12943-018-0932-8 LETTER TO THE EDITOR RNA-Seq profiling of circular RNAs in human colorectal Cancer liver metastasis and the potential biomarkers
Xu et al. Molecular Cancer (2019) 18:8 https://doi.org/10.1186/s12943-018-0932-8 LETTER TO THE EDITOR RNA-Seq profiling of circular RNAs in human colorectal Cancer liver metastasis and the potential biomarkers
Circular RNA_LARP4 is lower expressed and serves as a potential biomarker of ovarian cancer prognosis
 European Review for Medical and Pharmacological Sciences 2018; 22: 7178-7182 Circular RNA_LARP4 is lower expressed and serves as a potential biomarker of ovarian cancer prognosis T. ZOU, P.-L. WANG, Y.
European Review for Medical and Pharmacological Sciences 2018; 22: 7178-7182 Circular RNA_LARP4 is lower expressed and serves as a potential biomarker of ovarian cancer prognosis T. ZOU, P.-L. WANG, Y.
Identification of micrornas in blood and urine as tumour markers for the detection of urinary bladder cancer
 ONCOLOGY REPORTS 30: 1949-1956, 2013 Identification of micrornas in blood and urine as tumour markers for the detection of urinary bladder cancer ANGELIKA TÖLLE 1, MONIKA JUNG 1, SILKE RABENHORST 1, ERGIN
ONCOLOGY REPORTS 30: 1949-1956, 2013 Identification of micrornas in blood and urine as tumour markers for the detection of urinary bladder cancer ANGELIKA TÖLLE 1, MONIKA JUNG 1, SILKE RABENHORST 1, ERGIN
(DNA) Real-time PCR. Exicycler 96 Rotor-Gene Q/6000 PCR
 Real-Time (DNA) Real-time Exicycler 96 Rotor-Gene Q/6000 IU Mix1 Mix2 IU/μl IU/μl IU/μl IU/μl IU/μl IPC NTC C 1 Lot# 2 Freeze & thawing 1 MSDS: Material Safety Data Sheets (TaqMan Real-time ' FAM ' BHQ1
Real-Time (DNA) Real-time Exicycler 96 Rotor-Gene Q/6000 IU Mix1 Mix2 IU/μl IU/μl IU/μl IU/μl IU/μl IPC NTC C 1 Lot# 2 Freeze & thawing 1 MSDS: Material Safety Data Sheets (TaqMan Real-time ' FAM ' BHQ1
mirna in Plasma Exosome is Stable under Different Storage Conditions
 Molecules 2014, 19, 1568-1575; doi:10.3390/molecules19021568 Article OPEN ACCESS molecules ISSN 1420-3049 www.mdpi.com/journal/molecules mirna in Plasma Exosome is Stable under Different Storage Conditions
Molecules 2014, 19, 1568-1575; doi:10.3390/molecules19021568 Article OPEN ACCESS molecules ISSN 1420-3049 www.mdpi.com/journal/molecules mirna in Plasma Exosome is Stable under Different Storage Conditions
QIAGEN Complete Solutions for Liquid Biopsy Molecular Testing
 QIAGEN Complete Solutions for Liquid Biopsy Molecular Testing Christopher Swagell, PhD Market Development Manager, Advanced Molecular Pathology QIAGEN 1 Agenda QIAGEN Solid Tumor Testing and Liquid Biopsy
QIAGEN Complete Solutions for Liquid Biopsy Molecular Testing Christopher Swagell, PhD Market Development Manager, Advanced Molecular Pathology QIAGEN 1 Agenda QIAGEN Solid Tumor Testing and Liquid Biopsy
Original Article Tissue expression level of lncrna UCA1 is a prognostic biomarker for colorectal cancer
 Int J Clin Exp Pathol 2016;9(4):4241-4246 www.ijcep.com /ISSN:1936-2625/IJCEP0012296 Original Article Tissue expression level of lncrna UCA1 is a prognostic biomarker for colorectal cancer Hong Jiang,
Int J Clin Exp Pathol 2016;9(4):4241-4246 www.ijcep.com /ISSN:1936-2625/IJCEP0012296 Original Article Tissue expression level of lncrna UCA1 is a prognostic biomarker for colorectal cancer Hong Jiang,
Validation of the T descriptor in the new 8th TNM classification for non-small cell lung cancer
 Original Article Validation of the T descriptor in the new 8th TNM classification for non-small cell lung cancer Hee Suk Jung 1, Jin Gu Lee 2, Chang Young Lee 2, Dae Joon Kim 2, Kyung Young Chung 2 1 Department
Original Article Validation of the T descriptor in the new 8th TNM classification for non-small cell lung cancer Hee Suk Jung 1, Jin Gu Lee 2, Chang Young Lee 2, Dae Joon Kim 2, Kyung Young Chung 2 1 Department
Digitizing the Proteomes From Big Tissue Biobanks
 Digitizing the Proteomes From Big Tissue Biobanks Analyzing 24 Proteomes Per Day by Microflow SWATH Acquisition and Spectronaut Pulsar Analysis Jan Muntel 1, Nick Morrice 2, Roland M. Bruderer 1, Lukas
Digitizing the Proteomes From Big Tissue Biobanks Analyzing 24 Proteomes Per Day by Microflow SWATH Acquisition and Spectronaut Pulsar Analysis Jan Muntel 1, Nick Morrice 2, Roland M. Bruderer 1, Lukas
A complete next-generation sequencing workfl ow for circulating cell-free DNA isolation and analysis
 APPLICATION NOTE Cell-Free DNA Isolation Kit A complete next-generation sequencing workfl ow for circulating cell-free DNA isolation and analysis Abstract Circulating cell-free DNA (cfdna) has been shown
APPLICATION NOTE Cell-Free DNA Isolation Kit A complete next-generation sequencing workfl ow for circulating cell-free DNA isolation and analysis Abstract Circulating cell-free DNA (cfdna) has been shown
Analysis of circulating long non-coding RNA UCA1 as potential biomarkers for diagnosis and prognosis of osteosarcoma
 European Review for Medical and Pharmacological Sciences 2017; 21: 498-503 Analysis of circulating long non-coding RNA UCA1 as potential biomarkers for diagnosis and prognosis of osteosarcoma J.-J. WEN
European Review for Medical and Pharmacological Sciences 2017; 21: 498-503 Analysis of circulating long non-coding RNA UCA1 as potential biomarkers for diagnosis and prognosis of osteosarcoma J.-J. WEN
Exosomal Del 1 as a potent diagnostic marker for breast cancer : A prospective cohort study
 GBCC 2017: ABS-0017 Exosomal Del 1 as a potent diagnostic marker for breast cancer : A prospective cohort study Soo Jung Lee 1, Jeeyeon Lee 2, Jin Hyang Jung 2, Ho Yong Park 2, Chan Hyeong Lee 3, Pyong
GBCC 2017: ABS-0017 Exosomal Del 1 as a potent diagnostic marker for breast cancer : A prospective cohort study Soo Jung Lee 1, Jeeyeon Lee 2, Jin Hyang Jung 2, Ho Yong Park 2, Chan Hyeong Lee 3, Pyong
Hepatitis B Antiviral Drug Development Multi-Marker Screening Assay
 Hepatitis B Antiviral Drug Development Multi-Marker Screening Assay Background ImQuest BioSciences has developed and qualified a single-plate method to expedite the screening of antiviral agents against
Hepatitis B Antiviral Drug Development Multi-Marker Screening Assay Background ImQuest BioSciences has developed and qualified a single-plate method to expedite the screening of antiviral agents against
A preliminary study on the relationship between circulating tumor cells count and clinical features in patients with non-small cell lung cancer
 Original Article Page 1 of 5 A preliminary study on the relationship between circulating tumor cells count and clinical features in patients with non-small cell lung cancer Jia-Wei Wan 1,2 *, Ming-Zhu
Original Article Page 1 of 5 A preliminary study on the relationship between circulating tumor cells count and clinical features in patients with non-small cell lung cancer Jia-Wei Wan 1,2 *, Ming-Zhu
MicroRNA and Male Infertility: A Potential for Diagnosis
 Review Article MicroRNA and Male Infertility: A Potential for Diagnosis * Abstract MicroRNAs (mirnas) are small non-coding single stranded RNA molecules that are physiologically produced in eukaryotic
Review Article MicroRNA and Male Infertility: A Potential for Diagnosis * Abstract MicroRNAs (mirnas) are small non-coding single stranded RNA molecules that are physiologically produced in eukaryotic
Validation of Housekeeping Genes for Gene Expression Analysis in Glioblastoma Using Quantitative Real-Time Polymerase Chain Reaction
 ORIGINAL ARTICLE Brain Tumor Res Treat 2015;3(1):24-29 / pissn 2288-2405 / eissn 2288-2413 http://dx.doi.org/10.14791/btrt.2015.3.1.24 Validation of Housekeeping Genes for Gene Expression Analysis in Glioblastoma
ORIGINAL ARTICLE Brain Tumor Res Treat 2015;3(1):24-29 / pissn 2288-2405 / eissn 2288-2413 http://dx.doi.org/10.14791/btrt.2015.3.1.24 Validation of Housekeeping Genes for Gene Expression Analysis in Glioblastoma
Total genomic DNA was extracted from either 6 DBS punches (3mm), or 0.1ml of peripheral or
 Material and methods Measurement of telomere length (TL) Total genomic DNA was extracted from either 6 DBS punches (3mm), or 0.1ml of peripheral or cord blood using QIAamp DNA Mini Kit and a Qiacube (Qiagen).
Material and methods Measurement of telomere length (TL) Total genomic DNA was extracted from either 6 DBS punches (3mm), or 0.1ml of peripheral or cord blood using QIAamp DNA Mini Kit and a Qiacube (Qiagen).
SUPPLEMENTAL MATERIAL
 SUPPLEMENTAL MATERIAL Clinical perspective It was recently discovered that small RNAs, called micrornas, circulate freely and stably in human plasma. This finding has sparked interest in the potential
SUPPLEMENTAL MATERIAL Clinical perspective It was recently discovered that small RNAs, called micrornas, circulate freely and stably in human plasma. This finding has sparked interest in the potential
Plasma Bmil mrna as a potential prognostic biomarker for distant metastasis in colorectal cancer patients
 Title Plasma Bmil mrna as a potential prognostic biomarker for distant metastasis in colorectal cancer patients Author(s) Pun, JC; Chan, JY; Chun, BK; Ng, KW; Tsui, SY; Wan, TMH; Lo, OSH; Poon, TCJ; Ng,
Title Plasma Bmil mrna as a potential prognostic biomarker for distant metastasis in colorectal cancer patients Author(s) Pun, JC; Chan, JY; Chun, BK; Ng, KW; Tsui, SY; Wan, TMH; Lo, OSH; Poon, TCJ; Ng,
Santosh Patnaik, MD, PhD! Assistant Member! Department of Thoracic Surgery! Roswell Park Cancer Institute!
 Santosh Patnaik, MD, PhD Assistant Member Department of Thoracic Surgery Roswell Park Cancer Institute MicroRNA biology, techniques and applications History Biogenesis Nomenclature Tissue specificity Mechanisms
Santosh Patnaik, MD, PhD Assistant Member Department of Thoracic Surgery Roswell Park Cancer Institute MicroRNA biology, techniques and applications History Biogenesis Nomenclature Tissue specificity Mechanisms
The epidermal growth factor receptor (EGFR) gene encodes
 ORIGINAL ARTICLE Detection of EGFR Mutation Status in Lung Adenocarcinoma Specimens with Different Proportions of Tumor Cells Using Two Methods of Differential Sensitivity Hye-Suk Han, MD,* Sung-nam Lim,
ORIGINAL ARTICLE Detection of EGFR Mutation Status in Lung Adenocarcinoma Specimens with Different Proportions of Tumor Cells Using Two Methods of Differential Sensitivity Hye-Suk Han, MD,* Sung-nam Lim,
Profiles of gene expression & diagnosis/prognosis of cancer Lorena Roa de la Cruz
 Genomics Profiles of gene expression & diagnosis/prognosis of cancer Lorena Roa de la Cruz Gene expression profiling Measurement of the activity of thousands of genes at once Techniques used for gene expression
Genomics Profiles of gene expression & diagnosis/prognosis of cancer Lorena Roa de la Cruz Gene expression profiling Measurement of the activity of thousands of genes at once Techniques used for gene expression
Corporate Medical Policy
 Corporate Medical Policy Microarray-based Gene Expression Testing for Cancers of Unknown File Name: Origination: Last CAP Review: Next CAP Review: Last Review: microarray-based_gene_expression_testing_for_cancers_of_unknown_primary
Corporate Medical Policy Microarray-based Gene Expression Testing for Cancers of Unknown File Name: Origination: Last CAP Review: Next CAP Review: Last Review: microarray-based_gene_expression_testing_for_cancers_of_unknown_primary
Identifying Relevant micrornas in Bladder Cancer using Multi-Task Learning
 Identifying Relevant micrornas in Bladder Cancer using Multi-Task Learning Adriana Birlutiu (1),(2), Paul Bulzu (1), Irina Iereminciuc (1), and Alexandru Floares (1) (1) SAIA & OncoPredict, Cluj-Napoca,
Identifying Relevant micrornas in Bladder Cancer using Multi-Task Learning Adriana Birlutiu (1),(2), Paul Bulzu (1), Irina Iereminciuc (1), and Alexandru Floares (1) (1) SAIA & OncoPredict, Cluj-Napoca,
Clinical value of combined detection of mir 1202 and mir 195 in early diagnosis of cervical cancer
 ONCOLOGY LETTERS 17: 3387-3391, 2019 Clinical value of combined detection of mir 1202 and mir 195 in early diagnosis of cervical cancer XIELAN YANG 1, ZHILING YAN 1, HONGYING YANG 1, HUIJING NI 1, LEI
ONCOLOGY LETTERS 17: 3387-3391, 2019 Clinical value of combined detection of mir 1202 and mir 195 in early diagnosis of cervical cancer XIELAN YANG 1, ZHILING YAN 1, HONGYING YANG 1, HUIJING NI 1, LEI
For in vitro Veterinary Diagnostics only. Kylt Rotavirus A. Real-Time RT-PCR Detection.
 For in vitro Veterinary Diagnostics only. Kylt Rotavirus A Real-Time RT-PCR Detection www.kylt.eu DIRECTION FOR USE Kylt Rotavirus A Real-Time RT-PCR Detection A. General Kylt Rotavirus A products are
For in vitro Veterinary Diagnostics only. Kylt Rotavirus A Real-Time RT-PCR Detection www.kylt.eu DIRECTION FOR USE Kylt Rotavirus A Real-Time RT-PCR Detection A. General Kylt Rotavirus A products are
Supplementary Figure 1: Experimental design. DISCOVERY PHASE VALIDATION PHASE (N = 88) (N = 20) Healthy = 20. Healthy = 6. Endometriosis = 33
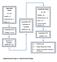 DISCOVERY PHASE (N = 20) Healthy = 6 Endometriosis = 7 EAOC = 7 Quantitative PCR (mirnas = 1113) a Quantitative PCR Verification of Candidate mirnas (N = 24) VALIDATION PHASE (N = 88) Healthy = 20 Endometriosis
DISCOVERY PHASE (N = 20) Healthy = 6 Endometriosis = 7 EAOC = 7 Quantitative PCR (mirnas = 1113) a Quantitative PCR Verification of Candidate mirnas (N = 24) VALIDATION PHASE (N = 88) Healthy = 20 Endometriosis
Assaying micrornas in biofluids for detection of drug induced cardiac injury. HESI Annual Meeting State of the Science Session June 8, 2011
 Assaying micrornas in biofluids for detection of drug induced cardiac injury HESI Annual Meeting State of the Science Session June 8, 2011 Karol Thompson, PhD Center for Drug Evaluation & Research US Food
Assaying micrornas in biofluids for detection of drug induced cardiac injury HESI Annual Meeting State of the Science Session June 8, 2011 Karol Thompson, PhD Center for Drug Evaluation & Research US Food
following radiotherapy
 British Journal of Cancer (1995) 72, 1536-154 r) 1995 Stockton Press All rights reserved 7-92/95 $12. Serum tumour markers in carcinoma of the uterine cervix and outcome following radiotherapy ARM Sproston',
British Journal of Cancer (1995) 72, 1536-154 r) 1995 Stockton Press All rights reserved 7-92/95 $12. Serum tumour markers in carcinoma of the uterine cervix and outcome following radiotherapy ARM Sproston',
m 6 A mrna methylation regulates AKT activity to promote the proliferation and tumorigenicity of endometrial cancer
 SUPPLEMENTARY INFORMATION Articles https://doi.org/10.1038/s41556-018-0174-4 In the format provided by the authors and unedited. m 6 A mrna methylation regulates AKT activity to promote the proliferation
SUPPLEMENTARY INFORMATION Articles https://doi.org/10.1038/s41556-018-0174-4 In the format provided by the authors and unedited. m 6 A mrna methylation regulates AKT activity to promote the proliferation
On the Reproducibility of TCGA Ovarian Cancer MicroRNA Profiles
 On the Reproducibility of TCGA Ovarian Cancer MicroRNA Profiles Ying-Wooi Wan 1,2,4, Claire M. Mach 2,3, Genevera I. Allen 1,7,8, Matthew L. Anderson 2,4,5 *, Zhandong Liu 1,5,6,7 * 1 Departments of Pediatrics
On the Reproducibility of TCGA Ovarian Cancer MicroRNA Profiles Ying-Wooi Wan 1,2,4, Claire M. Mach 2,3, Genevera I. Allen 1,7,8, Matthew L. Anderson 2,4,5 *, Zhandong Liu 1,5,6,7 * 1 Departments of Pediatrics
Value of serum galectin-3 and midkine level determination for assessing tumor severity in patients with thyroid cancer
 148 Journal of Hainan Medical University 2017; 23(3): 148-152 Journal of Hainan Medical University http://www.hnykdxxb.com Value of serum galectin-3 and midkine level determination for assessing tumor
148 Journal of Hainan Medical University 2017; 23(3): 148-152 Journal of Hainan Medical University http://www.hnykdxxb.com Value of serum galectin-3 and midkine level determination for assessing tumor
Implications of microrna-197 downregulated expression in esophageal cancer with poor prognosis
 Implications of microrna-197 downregulated expression in esophageal cancer with poor prognosis T.-Y. Wang 1, S.-G. Liu 2, B.-S. Zhao 2, B. Qi 2, X.-G. Qin 2 and W.-J. Yao 2 1 Department of Biochemistry
Implications of microrna-197 downregulated expression in esophageal cancer with poor prognosis T.-Y. Wang 1, S.-G. Liu 2, B.-S. Zhao 2, B. Qi 2, X.-G. Qin 2 and W.-J. Yao 2 1 Department of Biochemistry
ANALYTISCHE STRATEGIE Tissue Imaging. Bernd Bodenmiller Institute of Molecular Life Sciences University of Zurich
 ANALYTISCHE STRATEGIE Tissue Imaging Bernd Bodenmiller Institute of Molecular Life Sciences University of Zurich Quantitative Breast single cancer cell analysis Switzerland Brain Breast Lung Colon-rectum
ANALYTISCHE STRATEGIE Tissue Imaging Bernd Bodenmiller Institute of Molecular Life Sciences University of Zurich Quantitative Breast single cancer cell analysis Switzerland Brain Breast Lung Colon-rectum
RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays
 Supplementary Materials RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays Junhee Seok 1*, Weihong Xu 2, Ronald W. Davis 2, Wenzhong Xiao 2,3* 1 School of Electrical Engineering,
Supplementary Materials RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays Junhee Seok 1*, Weihong Xu 2, Ronald W. Davis 2, Wenzhong Xiao 2,3* 1 School of Electrical Engineering,
Identification of mirna expression profiles for diagnosis and prognosis of prostate cancer
 Identification of mirna expression profiles for diagnosis and prognosis of prostate cancer To my grandfather Curt "Cula" Carlsson "Research is to see what everybody else has seen, and to think what nobody
Identification of mirna expression profiles for diagnosis and prognosis of prostate cancer To my grandfather Curt "Cula" Carlsson "Research is to see what everybody else has seen, and to think what nobody
The clinical relevance of circulating, cell-free and exosomal micrornas as biomarkers for gynecological tumors
 Department of Tumor Biology The clinical relevance of circulating, cell-free and exosomal micrornas as biomarkers for gynecological tumors cfdna Copenhagen April 6-7, 2017 Heidi Schwarzenbach, PhD Tumor
Department of Tumor Biology The clinical relevance of circulating, cell-free and exosomal micrornas as biomarkers for gynecological tumors cfdna Copenhagen April 6-7, 2017 Heidi Schwarzenbach, PhD Tumor
Long non-coding RNA Loc is a potential prognostic biomarker in non-small cell lung cancer
 European Review for Medical and Pharmacological Sciences 2017; 21: 3808-3812 Long non-coding RNA Loc344887 is a potential prognostic biomarker in non-small cell lung cancer B. WU 1, X.-J. ZHANG 2, X.-G.
European Review for Medical and Pharmacological Sciences 2017; 21: 3808-3812 Long non-coding RNA Loc344887 is a potential prognostic biomarker in non-small cell lung cancer B. WU 1, X.-J. ZHANG 2, X.-G.
Supplementary Material
 Supplementary Material Identification of mir-187 and mir-182 as biomarkers for early diagnosis and prognosis in prostate cancer patients treated with radical prostatectomy Irene Casanova-Salas 1, José
Supplementary Material Identification of mir-187 and mir-182 as biomarkers for early diagnosis and prognosis in prostate cancer patients treated with radical prostatectomy Irene Casanova-Salas 1, José
High Expression of Forkhead Box Protein C2 is Related to Poor Prognosis in Human Gliomas
 DOI:http://dx.doi.org/10.7314/APJCP.2014.15.24.10621 RESEARCH ARTICLE High Expression of Forkhead Box Protein C2 is Related to Poor Prognosis in Human Gliomas Yao-Wu Wang 1, Chun-Li Yin 2, Hong-Yi Zhang
DOI:http://dx.doi.org/10.7314/APJCP.2014.15.24.10621 RESEARCH ARTICLE High Expression of Forkhead Box Protein C2 is Related to Poor Prognosis in Human Gliomas Yao-Wu Wang 1, Chun-Li Yin 2, Hong-Yi Zhang
(DNA) Real-time PCR. Exicycler 96 Rotor-Gene Q/6000 PCR
 Real-Time (DNA) Real-time Exicycler 96 Rotor-Gene Q/6000 IU Mix1 Mix2 IU/μl IU/μl IU/μl IU/μl IU/μl IPC NTC C 1 Lot# 2 Freeze & thawing 1 MSDS: Material Safety Data Sheets Real- (TaqMan time ' FAM ' BHQ1
Real-Time (DNA) Real-time Exicycler 96 Rotor-Gene Q/6000 IU Mix1 Mix2 IU/μl IU/μl IU/μl IU/μl IU/μl IPC NTC C 1 Lot# 2 Freeze & thawing 1 MSDS: Material Safety Data Sheets Real- (TaqMan time ' FAM ' BHQ1
Urine Biomarker Calibration for Ovarian Cancer Diagnosis
 , pp.218-222 http://dx.doi.org/10.14257/astl.2013.29.45 Urine Biomarker Calibration for Ovarian Cancer Diagnosis Eun-Suk Yang, Yu-Seop Kim, Chan-Young Park, Hye-Jeong Song, Jong-Dae Kim Dept. of Ubiquitous
, pp.218-222 http://dx.doi.org/10.14257/astl.2013.29.45 Urine Biomarker Calibration for Ovarian Cancer Diagnosis Eun-Suk Yang, Yu-Seop Kim, Chan-Young Park, Hye-Jeong Song, Jong-Dae Kim Dept. of Ubiquitous
