RapidFire Selected Therapeutic Areas and Screening Techniques
|
|
|
- Madison Norton
- 5 years ago
- Views:
Transcription
1 RapidFire Selected Therapeutic Areas and Screening Techniques RapidFire: SPE/MS/MS Life Sciences Group William A. LaMarr, Ph.D. Senior R&D Manager, RapidFire
2 RapidFire Mass Spectrometry Ultra-fast autosampler & online SPE system Replaces LC in LC/MS Reusable SPE cartridge Integrates with standard ESI MS instruments Cycle time = 7-13 s/sample Compatible with various matrices Sub-cellular fractions Cell culture supernatants Cell / tissue extracts Biological fluids
3 Advantages of mass spectrometry True label-free detection Direct, quantitative measurements P 4 Native reaction substrates & products (no radioactivity, no surrogate analytes, no indirect or secondary components) Functional biochemical assays instead of : P 4 (rather than target binding assays)
4 Limitations of MS Molecules must be charged Desalting step required Sample purification is Serial slow Instrumentation is expensive, not easily scalable to meet demand
5 Applications of the RapidFire Platform 1) Native Analyte Detection - surrogate substrates can introduce confounding factors, effect enzyme kinetics, and produce data artifacts 2) Replace Intractable Assays - assays may present challenges in workflow, may be resource intensive, may be cost prohibitive, may present regulatory issues (radioactivity) 3) Enable Target Classes - multiple modification events on the same substrate are impossible to track by many common optical and radioactive methodlogies
6 ADS Customer Base Biotech Large Pharma Multiple Target Customers ne Target Customers Biopharma High Profile Targets Unique Targets Primary Secondary Screening Screening (27x) Primary Screening Secondary Screening (6x) & Support
7 Targeted Therapeutic Areas Metabolic Disorder Anti-Infectives ncology ther Inflammation Epigenetics Inflammation Cardiovascular Disease Metabolic Disorder
8 Metabolic Disorder
9 Metabolic Disorders Class of genetic disease that encompasses varied conditions Inborn errors of metabolism Congenital metabolic disease Most are due to a single enzyme mutation effecting conversion of substrate to product and often result in: accumulation of toxic substances or reduced ability to synthesize essential compounds Because of the diverse nature of the diseases in the group, accurate numbers for incidence are difficult to determine. ne Canadian study* found that approximately 15% of single gene disorders in the population are considered metabolic disorders. * Applegarth DA, Toone JR, Lowry RB (January 2000). "Incidence of inborn errors of metabolism in British Columbia, ". Pediatrics 105 (1): e10
10 Enzymes and associated disorder(s) An example of enzymes discussed in today s presentation: Enzyme Role Associated Metabolic Disorder(s) Serine palmitoyltransferase Involved in sphingolipid biosynthesis Hereditary sensory neuropathy type 1 Acetyl CoA Carboxylase Involved in fatty acid synthesis Type 2 Diabetes ATP citrate lyase GM3 Synthase Stearoyl CoA Desaturase Diacylglycerol Acyltransferase Crucial in many biosynthetic pathways including lipogenesis and cholesterolgenesis Involved in the biosynthesis of complex gangliosides Important for the desaturation of fatty acids Catalyzes the synthesis of triglycerides from digylcerides Fatty liver disease, type 2 diabetes Infantile seizure/epilepsy disorder besity, liver disease obesity
11 Acetyl-Coenzyme A Carboxylase + KH 14 C 3 AT P Acetyl Coenzyme A 2 14 C H Malonyl Coenzyme A
12 Acetyl-Coenzyme A AssayConditions 50 mm HEPES, ph mm MgCl 2 2 mm tripotassium citrate 2 mm DTT 0.75 mg/ml BSA 4 mm ATP 12.5 mm KHC 3 20 mm Acetyl-Coenzyme A *ACC assay conditions based on previously published 14C-incorporation assay protocol: Harwood, H.J., Jr. et al. J. Biol. Chem. (2003), 278 (39):
13 Acetyl-Coenzyme A (m.w. 809) Malonyl-Coenzyme A (m.w. 853)
14 * Sumper, M. Eur. J. Biochem. (1974), 49 (2): * Harwood, H.J., Jr. et al. Curr. pin. Investig. Drugs (2004), 5 (3):
15 Stearoyl-Coenzyme A Desaturase NH 2 3 H 3 H S H N H N H H P - P - NH 4 + NH 4 + N N N N H P H - NH 4 + NH 2 S + H N H N H H N N N N P P - NH NH 4 H P H - - NH H 3 H
16 Stearoyl-Coenzyme A leoyl-coenzyme A * Soulard, P., et al. Analytica Chimica Acta (2008), 627 (1):
17 Z' Score > 200,000 wells screened Plate Number
18
19 Diacylglycerol Acyltransferase Diolein 14 C 14 C 14 C H N NH 2 N H N N P H P H P H H N H H N S leoyl-coenzyme A H Triolein 14 C 14 C 14 C
20
21 IC 50 = 49.1 nm Z Score = 0.77
22
23 Epigenetics
24 Protein Methyltransferase Assay Strategies ThioGlo Assay SAH detection AlphaScreen Masoud Vedadi Pan- methylation or selective antibodies are needed for multiple states Amy Quinn F. Liu et.al., Journal of Medicinal Chemistry. 2009:
25 Protein Methyltransferase Assay Strategies Fluorescence Polarization Transcreener Epigen methyltransferase assay principle. Klink T A et al. J Biomol Screen 2011;17:59-70
26 Methylation-Sensitive Proteolysis Assay on EZ Reader II Wigle et al, Chemistry & Biology,
27 Co-Product Measurements -Histone Methyltransferases (HMTs) S-adenosyl-methionine (SAM) to S-adenosyl-homocysteine (SAH) -Histone Acetyltransferases (HATs) Acetyl-coenzyme A (ACoA) to Coenzyme A (CoA) -Sirtuin Deacetylases (SIRTs) Nicotinamide adenine dinucleotide (NAD + ) to nicotinamide acetyl-adp-ribose (AADPr)
28 Direct measurement of the AADPr product of sirtuin reactions SIRT1, SIRT2, SIRT3 & SIRT5 & SIRT6? Acetylated Protein De-Acetylated Protein NH 2 N N NH 2 N N P H P - N + NH 2 N N N N P H P - H + N NH 2 H H H H H H H NAD acetyl-adp-ribose Nicotinamide
29 SIRT1 - Enzyme Titration Timecourse Reaction Conditions: 50 mm Tris ph mm NaCl 2.7 mm KCl 1 mm MgCl % BSA 5 mm DTT 100 mm NAD + 10 mm p53 peptide (Anaspec cat # 62121) 2 --acetyl-adp-ribose Internal Standard ~12.5 minute analysis time
30 Initial Velocity (V 0 ) SIRT1 - Enzymatic Parameters (Linearity, K m & IC 50 ) 2 --acetyl-adp-ribose Based Analysis 1.2E E E E E E E+00 R² = Enzyme Dilution K m = 28 ± 5 mm K m = 54 ± 1 mm IC 50 = 77 ± 1 mm
31 Comparison of Peptide Based and 2 --acetyl- ADP-ribose Based Assay Parameters Peptide Read K m of NAD + K m of peptide IC 50 of nicotinamide 2AADPr Read Peptide Read 2AADPr Read Peptide Read 2AADPr Read SIRT1 38 ± 4 mm 54 ± 1 mm 25 ± 6 mm 28 ± 5 mm 62 ± 1 mm 77 ± 1 mm SIRT2 46 ± 11 mm 50 ± 8 mm 8 ± 3 mm 12 ± 1 mm 10 ± 1 mm 11 ± 1 mm SIRT3 118 ± 44 mm 144 ± 21 mm 4 ± 1 mm 6 ± 1 mm 31 ± 1 mm 39 ± 1 mm
32 Direct Peptide Measurements Histone Demethylases Lysine Demethylase 1 (LSD-1) uses Flavin Adenine Dinucleotide (FAD) Jumanji Domain 2a (JMJD2a) uses Fe+2 mediated oxidative chemistry Histone Deacetylases Histone Deacetylase 1 (HDAC-1) uses a metal dependent hydrolysis
33 SIRT1 - Enzyme Titration Timecourse Reaction Conditions: Deacetylated Peptide 180 minutes 120 minutes 240 minutes 50 mm Tris ph mm NaCl 2.7 mm KCl 1 mm MgCl % BSA 5 mm DTT 100 mm NAD + 10 mm p53 peptide (Anaspec cat # 62121) 30 minutes Enzyme 15 minutes 0 minutes Acetylated Peptide 45 minutes 60 minutes ~12.5 minute analysis time
34 Percent Conversion Initial Velocity (V 0 ) SIRT1 - Enzyme Linearity Enzyme Titration / Timecourse Enzyme Linearity x enzyme x enzyme x enzyme R² = Time (minutes) Enzyme Dilution
35 SIRT1 - Enzymatic Parameters (K m, IC 50, etc ) K m = 25 ± 6 mm K m = 38 ± 4 mm Deacetylated peptide IC 50 = 62 ± 1 mm Acetylated peptide Literature IC 50 value ~ 50 mm Bitterman et. al., J. Biol. Chem. (2002) 277: Marcotte et. al., Anal. Biochem. (2005) 332:90-99 Nicotinamide ~ 3 minute analysis time (8-point log-dilution IC 50 curve, n=3)
36 Initial Velocity SIRT2 - Enzymatic Parameters (Linearity, K m & IC 50 ) y = x R² = Enzyme Concentration (x stock) K m = 8 ± 2 mm K m = 46 ± 11 mm IC 50 = 9.6 ± 0.1 mm
37 Initial Velocity SIRT3 - Enzymatic Parameters (Linearity, K m & IC 50 ) y = x R² = Enzyme Concentration (x K m = 4.3 ± 1.0 mm K m = 118 ± 44 mm IC 50 = 31 mm
38 Labeled vs. Un-labeled Sirtuin Assay Free fluorophore Protein Labeled Acetylated Peptide Protein Unlabeled Acetylated Peptide H N CH C CH 2 CH 2 CH 2 CH 2 HN H N CH C CH 2 CH 2 CH 2 CH 2 HN Protein Protein SIRT1/Chymotrypsin Resveratrol Howitz et. al., Nature (2003) 425: SIRT1/Chymotrypsin Resveratrol Kaeberlein et. al., J. Biol. Chem. (2005) 280: Beher et. al., Chem. Biol. Rug Des. (2009) 74: Pacholec et. al., J. Biol. Chem. (2010) 285: Protein H 2 N CH C CH 2 CH 2 CH 2 CH 2 NH 2 Protein De-Acetylated Peptide H N CH C CH 2 CH 2 CH 2 CH 2 NH 2 Protein De-Acetylated Peptide Reaction Activation X No Reaction Activation
39 SIRT1 - Substrate Dependant Activation by Resveratrol Labeled Peptide Unlabeled Peptide Rye et. al., J. Biomol. Screen. (2011) 16: Milne et. al., Nature (2007) 450:
40 Multiple Modification Events on a Single Peptide potential acetylation site Preferred acetylation sites potential acetylation site -Human p53 ( ) -19-mer peptide - Six potential acetylation sites Rye et. al., J. Biomol. Screen. (2011) 16: Prives et. al., Nature (2007) 450:
41 Percent of Control Direct Measurement of Modification Events on Whole Histone Proteins Protein (Q-TF) AcCoA/CoA (QqQ) [Garcinol] (mm) Incorporation of a high-resolution (Q-TF) mass spectrometer into the RapidFire workflow allows the substitution of whole proteins for representative peptide based sequences in a high-throughput screening compatible mode. Rye et. al., J. Biomol. Screen. (2011) 16:
42 Screening Histone Demethylases in a Pharmaceutical Drug Discovery Setting Kruidenier et. al., Nature (2012) 488: Plant et. al., J. Anal. Biochem. (2011) 419: Hutchinson et. al., J. Biomol. Screen. (2012) 17: Melanie Leveridge, MipTec 2011, oral presentation
43 Conclusions Mass spectrometry is an excellent tool for epigenetic target based drug discovery because if it s ability to: - Directly measure native, label-free peptides and generic reaction co-products - Directly and independently measure multiple modification events on single substrates - Directly measure modifications to whole protein substrates
44 Fragment Based Drug Discovery
45 Fragment Based Drug Discovery
46 BACE-1 Assay: Fragment Based Screen Cary Eclipse Fluorescence Spectrophotometer
47 Assay Setup Fluorescent Peptide Fluorescence Plate Reader X Unlabeled Peptide Fluorescence Plate Reader Fluorescent Peptide Mass Spectrometer Unlabeled Peptide Mass Spectrometer
48 Assay System Characterization Unlabeled Substrate By MS (UMS) Fluorescently Labeled Substrate by FS (FS) Fluorescently Labeled Substrate by MS (LMS)
49 UMS LMS FS all 3 Initial Screening Results Hits by Assay Format Hits by Assay Format UMS UMS LMS FS all LMS FS
50 Hits bserved by MS nly Follow-up of selected hits confirmed that compound autofluorescence (AF) obscured several hits in the FS data, including the most potent analyte. Titration of that compound revealed a concentration-dependent increase in signal in the FS assay, suggesting AF, while the MS data were consistent with a traditional inhibition curve.
51 Hits bserved with the Labeled Peptide nly A second class of compounds was uncovered consisting of those molecules that appear as hits when the labeled peptide is employed (as in the FS and LMS assays), but do not show inhibition when the more native substrate is used (UMS). These results suggest that perhaps compounds exist that interfere with the enzyme s ability to bind the peptide carrying the bulky label but not with the tighter binding exhibited by the enzyme for the unlabeled substrate, raising the possibility of misleading data being produced when modified substrates are employed.
52 Hits bserved with the Unlabeled Peptide nly Yet another class of inhibitors was detected in the unlabeled assay (UMS) but was not found with the fluorescent peptide (FS or LMS). Because MS eliminates the need for unnatural modification of substrates, it allows the study of more biologically relevant molecules. These more realistic substrates could reveal activities that are lost with modified peptides, possibly due to altered binding, which in this case was clearly revealed by the K m experiments. Unlabeled Substrate By MS (UMS) Fluorescently Labeled Substrate by MS (LMS)
53 Conclusions Robust assays were developed for both a labeled and an unlabeled substrate of the BACE-1 enzyme. Screening a fragment library against both substrates using both detection methods produced three disparate hit sets. FS and MS produced different hit sets when used as complementary detection methods on the same samples. While some MS hits (including the most potent) were obscured by autofluorescence in the FS assay, this did not account for all of the differences between the methods. MS generated different hit sets for the labeled and the unlabeled peptide, underscoring the importance of substrate selection. Label-free screening by high-throughput MS has proven to be a valid method for conducting activity-based screens of fragment libraries that enables the study of more native molecules and is less susceptible to confounding factors, such as AF.
54 Target Binding Assays
55 Mass Spectrometry Approaches to Binding Assays
56 Questions? Agilent Technologies William A. LaMarr, Ph.D. Senior R&D Manager, RapidFire
Data Sheet. Fluorogenic HDAC 8 Assay Kit Catalog #: 50068
 Data Sheet Fluorogenic HDAC 8 Assay Kit Catalog #: 50068 DESCRIPTION: The Fluorogenic HDAC 8 Assay Kit is a complete assay system designed to measure histone deacetylase (HDAC) 8 activity for screening
Data Sheet Fluorogenic HDAC 8 Assay Kit Catalog #: 50068 DESCRIPTION: The Fluorogenic HDAC 8 Assay Kit is a complete assay system designed to measure histone deacetylase (HDAC) 8 activity for screening
Introduction. Figure 1 Experimental Strategy
 Beyond s: Interrogating the Purinome Tony A. Klink, Karen Kleman-Leyer, Matt Staeben, Robert G. Lowery BellBrook Labs, Madison, Wisconsin, USA 53711 Introduction The development of ATP-site ligands as
Beyond s: Interrogating the Purinome Tony A. Klink, Karen Kleman-Leyer, Matt Staeben, Robert G. Lowery BellBrook Labs, Madison, Wisconsin, USA 53711 Introduction The development of ATP-site ligands as
SensoLyte 520 HDAC Activity Assay Kit *Fluorimetric*
 SensoLyte 520 HDAC Activity Assay Kit *Fluorimetric* Catalog # 72084 Kit Size 100 Assays (96-well plate) Optimized Performance: This kit is optimized to detect HDAC activity. Enhanced Value: It provides
SensoLyte 520 HDAC Activity Assay Kit *Fluorimetric* Catalog # 72084 Kit Size 100 Assays (96-well plate) Optimized Performance: This kit is optimized to detect HDAC activity. Enhanced Value: It provides
Coenzymes. Coenzymes 9/11/2018. BCMB 3100 Introduction to Coenzymes & Vitamins
 BCMB 3100 Introduction to Coenzymes & Vitamins Cofactors Essential ions Coenzymes Cosubstrates Prosthetic groups Coenzymes structure/function/active group Vitamins 1 Coenzymes Some enzymes require for
BCMB 3100 Introduction to Coenzymes & Vitamins Cofactors Essential ions Coenzymes Cosubstrates Prosthetic groups Coenzymes structure/function/active group Vitamins 1 Coenzymes Some enzymes require for
BCMB 3100 Introduction to Coenzymes & Vitamins
 BCMB 3100 Introduction to Coenzymes & Vitamins Cofactors Essential ions Coenzymes Cosubstrates Prosthetic groups Coenzymes structure/function/active group Vitamins 1 Coenzymes Some enzymes require for
BCMB 3100 Introduction to Coenzymes & Vitamins Cofactors Essential ions Coenzymes Cosubstrates Prosthetic groups Coenzymes structure/function/active group Vitamins 1 Coenzymes Some enzymes require for
ab Histone Deacetylase (HDAC) Activity Assay Kit (Fluorometric)
 ab156064 Histone Deacetylase (HDAC) Activity Assay Kit (Fluorometric) Instructions for Use For the quantitative measurement of Histone Deacetylase activity in cell lysates This product is for research
ab156064 Histone Deacetylase (HDAC) Activity Assay Kit (Fluorometric) Instructions for Use For the quantitative measurement of Histone Deacetylase activity in cell lysates This product is for research
Chapter 12 Nutrition
 Chapter 12 Nutrition Nutrients macronutrients: large required daily quantities carbohydrates, lipids, proteins micronutrients: small required daily quantities vitamins, minerals Also required: water and
Chapter 12 Nutrition Nutrients macronutrients: large required daily quantities carbohydrates, lipids, proteins micronutrients: small required daily quantities vitamins, minerals Also required: water and
Use of RapidFire/MS for Metabolomics. - from untargeted to targeted and Pathway driven analysis. RapidFire/MS. Moritz Wagner Agilent Technologies
 Use of RapidFire/MS for Metabolomics RapidFire/MS - from untargeted to targeted and Pathway driven analysis Moritz Wagner Agilent Technologies 1 Agilent RapidFire TM High-throughput Mass Spectrometry System
Use of RapidFire/MS for Metabolomics RapidFire/MS - from untargeted to targeted and Pathway driven analysis Moritz Wagner Agilent Technologies 1 Agilent RapidFire TM High-throughput Mass Spectrometry System
Epigenase HDAC Activity/Inhibition Direct Assay Kit (Fluorometric)
 Epigenase HDAC Activity/Inhibition Direct Assay Kit (Fluorometric) Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE Uses: The Epigenase HDAC Activity/Inhibition Direct Assay Kit (Fluorometric)
Epigenase HDAC Activity/Inhibition Direct Assay Kit (Fluorometric) Base Catalog # PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE Uses: The Epigenase HDAC Activity/Inhibition Direct Assay Kit (Fluorometric)
Coenzymes. Coenzymes 9/15/2014. BCMB 3100 Introduction to Coenzymes & Vitamins
 BCMB 3100 Introduction to Coenzymes & Vitamins Cofactors Essential ions Coenzymes Cosubstrates Prosthetic groups Coenzymes structure/function/active group Vitamins 1 Coenzymes Some enzymes require for
BCMB 3100 Introduction to Coenzymes & Vitamins Cofactors Essential ions Coenzymes Cosubstrates Prosthetic groups Coenzymes structure/function/active group Vitamins 1 Coenzymes Some enzymes require for
9/16/2015. Coenzymes. Coenzymes. BCMB 3100 Introduction to Coenzymes & Vitamins. Types of cofactors
 BCMB 3100 Introduction to Coenzymes & Vitamins Cofactors Essential ions Coenzymes Cosubstrates Prosthetic groups Coenzymes structure/function/active group Vitamins 1 Coenzymes Some enzymes require for
BCMB 3100 Introduction to Coenzymes & Vitamins Cofactors Essential ions Coenzymes Cosubstrates Prosthetic groups Coenzymes structure/function/active group Vitamins 1 Coenzymes Some enzymes require for
The Role of Protein Domains in Cell Signaling
 The Role of Protein Domains in Cell Signaling Growth Factors RTK Ras Shc PLCPLC-γ P P P P P P GDP Ras GTP PTB CH P Sos Raf SH2 SH2 PI(3)K SH3 SH3 MEK Grb2 MAPK Phosphotyrosine binding domains: PTB and
The Role of Protein Domains in Cell Signaling Growth Factors RTK Ras Shc PLCPLC-γ P P P P P P GDP Ras GTP PTB CH P Sos Raf SH2 SH2 PI(3)K SH3 SH3 MEK Grb2 MAPK Phosphotyrosine binding domains: PTB and
Streamlining ATPase and Kinase Assay Development Using Direct ADP Detection. Biology at Work
 Streamlining ATPase and Kinase Assay Development Using Direct ADP Detection Summary Background on Transcreener ADP Assay, differentiators, competitive advantages Optimizing assays for kinases, a rule based
Streamlining ATPase and Kinase Assay Development Using Direct ADP Detection Summary Background on Transcreener ADP Assay, differentiators, competitive advantages Optimizing assays for kinases, a rule based
HDAC1 Inhibitor Screening Assay Kit
 HDAC1 Inhibitor Screening Assay Kit Catalog Number KA1320 96 assays Version: 03 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Background... 3 Principle of the Assay...
HDAC1 Inhibitor Screening Assay Kit Catalog Number KA1320 96 assays Version: 03 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Background... 3 Principle of the Assay...
Energy Production In A Cell (Chapter 25 Metabolism)
 Energy Production In A Cell (Chapter 25 Metabolism) Large food molecules contain a lot of potential energy in the form of chemical bonds but it requires a lot of work to liberate the energy. Cells need
Energy Production In A Cell (Chapter 25 Metabolism) Large food molecules contain a lot of potential energy in the form of chemical bonds but it requires a lot of work to liberate the energy. Cells need
Fatty acid breakdown
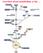 Fatty acids contain a long hydrocarbon chain and a terminal carboxylate group. Most contain between 14 and 24 carbon atoms. The chains may be saturated or contain double bonds. The complete oxidation of
Fatty acids contain a long hydrocarbon chain and a terminal carboxylate group. Most contain between 14 and 24 carbon atoms. The chains may be saturated or contain double bonds. The complete oxidation of
Fatty acids synthesis
 Fatty acids synthesis The synthesis start from Acetyl COA the first step requires ATP + reducing power NADPH! even though the oxidation and synthesis are different pathways but from chemical part of view
Fatty acids synthesis The synthesis start from Acetyl COA the first step requires ATP + reducing power NADPH! even though the oxidation and synthesis are different pathways but from chemical part of view
Introduction to Metabolism Cell Structure and Function
 Introduction to Metabolism Cell Structure and Function Cells can be divided into two primary types prokaryotes - Almost all prokaryotes are bacteria eukaryotes - Eukaryotes include all cells of multicellular
Introduction to Metabolism Cell Structure and Function Cells can be divided into two primary types prokaryotes - Almost all prokaryotes are bacteria eukaryotes - Eukaryotes include all cells of multicellular
Chapter 9 Overview. Aerobic Metabolism I: The Citric Acid Cycle. Live processes - series of oxidation-reduction reactions. Aerobic metabolism I
 n n Chapter 9 Overview Aerobic Metabolism I: The Citric Acid Cycle Live processes - series of oxidation-reduction reactions Ingestion of proteins, carbohydrates, lipids Provide basic building blocks for
n n Chapter 9 Overview Aerobic Metabolism I: The Citric Acid Cycle Live processes - series of oxidation-reduction reactions Ingestion of proteins, carbohydrates, lipids Provide basic building blocks for
Assay Report. Histone Deacetylase (HDAC) Inhibitor Assays Enzymatic Study of Compounds from Client
 Assay Report Histone Deacetylase (HDAC) Inhibitor Assays Enzymatic Study of Compounds from Client Page 1 of 27 Client_HDAC _Year Month Day 1 Client_HDAC_Year Month Year HDAC Inhibitor Assays Study Sponsor:
Assay Report Histone Deacetylase (HDAC) Inhibitor Assays Enzymatic Study of Compounds from Client Page 1 of 27 Client_HDAC _Year Month Day 1 Client_HDAC_Year Month Year HDAC Inhibitor Assays Study Sponsor:
Epigenase HDAC Activity/Inhibition Direct Assay Kit (Colorimetric)
 Epigenase HDAC Activity/Inhibition Direct Assay Kit (Colorimetric) Base Catalog # P-4034 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE Uses: The Epigenase HDAC Activity/Inhibition Direct Assay Kit (Colorimetric)
Epigenase HDAC Activity/Inhibition Direct Assay Kit (Colorimetric) Base Catalog # P-4034 PLEASE READ THIS ENTIRE USER GUIDE BEFORE USE Uses: The Epigenase HDAC Activity/Inhibition Direct Assay Kit (Colorimetric)
How Cells Harvest Energy. Chapter 7. Respiration
 How Cells Harvest Energy Chapter 7 Respiration Organisms classified on how they obtain energy: autotrophs: produce their own organic molecules through photosynthesis heterotrophs: live on organic compounds
How Cells Harvest Energy Chapter 7 Respiration Organisms classified on how they obtain energy: autotrophs: produce their own organic molecules through photosynthesis heterotrophs: live on organic compounds
Lecture 1- Metabolism: Basic Concepts and Design. Introduction. Introduction. Introduction. Questions we will focus on this semester:
 Lecture 1- Metabolism: Basic Concepts and Design Chem 454: Regulatory Mechanisms in Biochemistry University of Wisconsin-Eau Claire 1 Introduction Questions we will focus on this semester: How does a cell
Lecture 1- Metabolism: Basic Concepts and Design Chem 454: Regulatory Mechanisms in Biochemistry University of Wisconsin-Eau Claire 1 Introduction Questions we will focus on this semester: How does a cell
Synthesis of Fatty Acids and Triacylglycerol
 Fatty Acid Synthesis Synthesis of Fatty Acids and Triacylglycerol Requires Carbon Source: Reducing Power: NADPH Energy Input: ATP Why Energy? Why Energy? Fatty Acid Fatty Acid + n(atp) ΔG o : -ve Fatty
Fatty Acid Synthesis Synthesis of Fatty Acids and Triacylglycerol Requires Carbon Source: Reducing Power: NADPH Energy Input: ATP Why Energy? Why Energy? Fatty Acid Fatty Acid + n(atp) ΔG o : -ve Fatty
PhosFree TM Phosphate Assay Biochem Kit
 PhosFree TM Phosphate Assay Biochem Kit (Cat. # BK050) ORDERING INFORMATION To order by phone: (303) - 322-2254 To order by Fax: (303) - 322-2257 To order by e-mail: cservice@cytoskeleton.com Technical
PhosFree TM Phosphate Assay Biochem Kit (Cat. # BK050) ORDERING INFORMATION To order by phone: (303) - 322-2254 To order by Fax: (303) - 322-2257 To order by e-mail: cservice@cytoskeleton.com Technical
Fatty Acid Oxidation Assay on the XF24 Analyzer
 Fatty Acid Oxidation Assay on the XF24 Analyzer Mitochondria oxidize a variety of fuels to generate ATP through oxidative phosphorylation. Cells can utilize fatty acid, glucose and amino acids as their
Fatty Acid Oxidation Assay on the XF24 Analyzer Mitochondria oxidize a variety of fuels to generate ATP through oxidative phosphorylation. Cells can utilize fatty acid, glucose and amino acids as their
Supplementary Information
 Supplementary Information Archaeal Elp3 catalyzes trna wobble uridine modification at C5 via a radical mechanism Kiruthika Selvadurai, Pei Wang, Joseph Seimetz & Raven H Huang* Department of Biochemistry,
Supplementary Information Archaeal Elp3 catalyzes trna wobble uridine modification at C5 via a radical mechanism Kiruthika Selvadurai, Pei Wang, Joseph Seimetz & Raven H Huang* Department of Biochemistry,
Cells extract energy from their environment and use the energy for a host of biological activities including biosynthesis.
 ATP=cellular energy Cells extract energy from their environment and use the energy for a host of biological activities including biosynthesis. The reactions of energy extraction and energy use are called
ATP=cellular energy Cells extract energy from their environment and use the energy for a host of biological activities including biosynthesis. The reactions of energy extraction and energy use are called
Minjia Tan, Ph.D Shanghai Institute of Materia Medica, Chinese Academy of Sciences
 Lysine Glutarylation Is a Protein Posttranslational Modification Regulated by SIRT5 Minjia Tan, Ph.D Shanghai Institute of Materia Medica, Chinese Academy of Sciences Lysine: the most frequently modified
Lysine Glutarylation Is a Protein Posttranslational Modification Regulated by SIRT5 Minjia Tan, Ph.D Shanghai Institute of Materia Medica, Chinese Academy of Sciences Lysine: the most frequently modified
Synthesis of Fatty Acids and Triacylglycerol
 Synthesis of Fatty Acids and Triacylglycerol Lippincott s Chapter 16 Fatty Acid Synthesis Mainly in the Liver Requires Carbon Source: Acetyl CoA Reducing Power: NADPH 8 CH 3 COO C 15 H 33 COO Energy Input:
Synthesis of Fatty Acids and Triacylglycerol Lippincott s Chapter 16 Fatty Acid Synthesis Mainly in the Liver Requires Carbon Source: Acetyl CoA Reducing Power: NADPH 8 CH 3 COO C 15 H 33 COO Energy Input:
ANSC/NUTR 618 LIPIDS & LIPID METABOLISM. Fatty Acid Elongation and Desaturation
 ANSC/NUTR 618 LIPIDS & LIPID METABOLISM I. Fatty acid elongation A. General 1. At least 60% of fatty acids in triacylglycerols are C18. 2. Free palmitic acid (16:0) synthesized in cytoplasm is elongated
ANSC/NUTR 618 LIPIDS & LIPID METABOLISM I. Fatty acid elongation A. General 1. At least 60% of fatty acids in triacylglycerols are C18. 2. Free palmitic acid (16:0) synthesized in cytoplasm is elongated
Chemistry 1506: Allied Health Chemistry 2. Section 13: Biosynthetic Pathways. Making Complex Biomolecules. Outline
 Chemistry 1506 Dr. Hunter s Class Section 13 Notes - Page 1/9 Chemistry 1506: Allied Health Chemistry 2 Section 13: Biosynthetic Pathways Making Complex Biomolecules utline SECTIN 13.1 INTRDUCTIN...2 SECTIN
Chemistry 1506 Dr. Hunter s Class Section 13 Notes - Page 1/9 Chemistry 1506: Allied Health Chemistry 2 Section 13: Biosynthetic Pathways Making Complex Biomolecules utline SECTIN 13.1 INTRDUCTIN...2 SECTIN
Epigenetics q&more 01.11
 Laurie. Knight, istockphoto.com Epigenetics 6 Bookmarks About the reading of genes in the Book of Life Prof. Dr. Manfred Jung, Julia M. Wagner, Institute for Pharmaceutical Sciences, Albert-Ludwig-University
Laurie. Knight, istockphoto.com Epigenetics 6 Bookmarks About the reading of genes in the Book of Life Prof. Dr. Manfred Jung, Julia M. Wagner, Institute for Pharmaceutical Sciences, Albert-Ludwig-University
Improved Stability of the LANCE Ultra Signal in Kinase Assays
 Improved Stability of the LANCE Ultra Signal in Kinase Assays LANCE Ultra is a high throughput screening (HTS) technology platform optimized for homogeneous time-resolved fluorescence resonance energy
Improved Stability of the LANCE Ultra Signal in Kinase Assays LANCE Ultra is a high throughput screening (HTS) technology platform optimized for homogeneous time-resolved fluorescence resonance energy
Post-translational modifications of proteins in gene regulation under hypoxic conditions
 203 Review Article Post-translational modifications of proteins in gene regulation under hypoxic conditions 1, 2) Olga S. Safronova 1) Department of Cellular Physiological Chemistry, Tokyo Medical and
203 Review Article Post-translational modifications of proteins in gene regulation under hypoxic conditions 1, 2) Olga S. Safronova 1) Department of Cellular Physiological Chemistry, Tokyo Medical and
Chemical Energy. Valencia College
 9 Pathways that Harvest Chemical Energy Valencia College 9 Pathways that Harvest Chemical Energy Chapter objectives: How Does Glucose Oxidation Release Chemical Energy? What Are the Aerobic Pathways of
9 Pathways that Harvest Chemical Energy Valencia College 9 Pathways that Harvest Chemical Energy Chapter objectives: How Does Glucose Oxidation Release Chemical Energy? What Are the Aerobic Pathways of
HDAC Fluorometric Activity Assay Kit
 HDAC Fluorometric Activity Assay Kit Item. 10011563 www.caymanchem.com Customer Service 800.364.9897 Technical Support 888.526.5351 1180 E. Ellsworth Rd Ann Arbor, MI USA TABLE OF CONTENTS GENERAL INFORMATION
HDAC Fluorometric Activity Assay Kit Item. 10011563 www.caymanchem.com Customer Service 800.364.9897 Technical Support 888.526.5351 1180 E. Ellsworth Rd Ann Arbor, MI USA TABLE OF CONTENTS GENERAL INFORMATION
ANSC/NUTR 618 Lipids & Lipid Metabolism
 I. Overall concepts A. Definitions ANC/NUTR 618 Lipids & Lipid Metabolism 1. De novo synthesis = synthesis from non-fatty acid precursors a. Carbohydrate precursors (glucose, lactate, and pyruvate) b.
I. Overall concepts A. Definitions ANC/NUTR 618 Lipids & Lipid Metabolism 1. De novo synthesis = synthesis from non-fatty acid precursors a. Carbohydrate precursors (glucose, lactate, and pyruvate) b.
HDAC Fluorometric Activity Assay Kit
 HDAC Fluorometric Activity Assay Kit Item. 10011563 Customer Service 800.364.9897 * Technical Support 888.526.5351 www.caymanchem.com TABLE OF CONTENTS GENERAL INFORMATION 3 Materials Supplied 4 Precautions
HDAC Fluorometric Activity Assay Kit Item. 10011563 Customer Service 800.364.9897 * Technical Support 888.526.5351 www.caymanchem.com TABLE OF CONTENTS GENERAL INFORMATION 3 Materials Supplied 4 Precautions
ANSC/NUTR 618 Lipids & Lipid Metabolism
 I. verall concepts A. Definitions ANSC/NUTR 618 Lipids & Lipid Metabolism 1. De novo synthesis = synthesis from non-fatty acid precursors a. Carbohydrate precursors (glucose and lactate) 1) Uses glucose
I. verall concepts A. Definitions ANSC/NUTR 618 Lipids & Lipid Metabolism 1. De novo synthesis = synthesis from non-fatty acid precursors a. Carbohydrate precursors (glucose and lactate) 1) Uses glucose
Energy Transformation: Cellular Respiration Outline 1. Sources of cellular ATP 2. Turning chemical energy of covalent bonds between C-C into energy
 Energy Transformation: Cellular Respiration Outline 1. Sources of cellular ATP 2. Turning chemical energy of covalent bonds between C-C into energy for cellular work (ATP) 3. Importance of electrons and
Energy Transformation: Cellular Respiration Outline 1. Sources of cellular ATP 2. Turning chemical energy of covalent bonds between C-C into energy for cellular work (ATP) 3. Importance of electrons and
Chemistry 1506: Allied Health Chemistry 2. Section 11: Bioenergetics. Energy Generation in the Cell. Outline
 Chemistry 1506 Dr. unter s Class Section 11 Notes - Page 1/17 Chemistry 1506: Allied ealth Chemistry 2 Section 11: Bioenergetics Energy Generation in the Cell utline SECTIN 11.1 INTRDUCTIN & MITCNDRIA...2
Chemistry 1506 Dr. unter s Class Section 11 Notes - Page 1/17 Chemistry 1506: Allied ealth Chemistry 2 Section 11: Bioenergetics Energy Generation in the Cell utline SECTIN 11.1 INTRDUCTIN & MITCNDRIA...2
HDAC Assay Kit. (Fluorescent) (version B1) Catalog No
 HDAC Assay Kit (Fluorescent) (version B1) Catalog No. 56200 Active Motif North America 1914 Palomar Oaks Way, Suite 150 Carlsbad, California 92008, USA Toll free: 877 222 9543 Telephone: 760 431 1263 Fax:
HDAC Assay Kit (Fluorescent) (version B1) Catalog No. 56200 Active Motif North America 1914 Palomar Oaks Way, Suite 150 Carlsbad, California 92008, USA Toll free: 877 222 9543 Telephone: 760 431 1263 Fax:
A cell has enough ATP to last for about three seconds.
 Energy Transformation: Cellular Respiration Outline 1. Energy and carbon sources in living cells 2. Sources of cellular ATP 3. Turning chemical energy of covalent bonds between C-C into energy for cellular
Energy Transformation: Cellular Respiration Outline 1. Energy and carbon sources in living cells 2. Sources of cellular ATP 3. Turning chemical energy of covalent bonds between C-C into energy for cellular
Lecture Sixteen: METABOLIC ENERGY: [Based on GENERATION Chapter 15
 Lecture Sixteen: METABOLIC ENERGY: [Based on GENERATION Chapter 15 AND STORAGE Berg, (Figures in red are for the 7th Edition) Tymoczko (Figures in Blue are for the 8th Edition) & Stryer] Two major questions
Lecture Sixteen: METABOLIC ENERGY: [Based on GENERATION Chapter 15 AND STORAGE Berg, (Figures in red are for the 7th Edition) Tymoczko (Figures in Blue are for the 8th Edition) & Stryer] Two major questions
ADP, ATP and Cellular Respiration
 ADP, ATP and Cellular Respiration What Is ATP? Energy used by all Cells Adenosine Triphosphate Organic molecule containing highenergy Phosphate bonds Chemical Structure of ATP Adenine Base 3 Phosphates
ADP, ATP and Cellular Respiration What Is ATP? Energy used by all Cells Adenosine Triphosphate Organic molecule containing highenergy Phosphate bonds Chemical Structure of ATP Adenine Base 3 Phosphates
BIOSYNTHESIS OF FATTY ACIDS. doc. Ing. Zenóbia Chavková, CSc.
 BIOSYNTHESIS OF FATTY ACIDS doc. Ing. Zenóbia Chavková, CSc. The pathway for the of FAs is not the reversal of the oxidation pathway Both pathways are separated within different cellular compartments In
BIOSYNTHESIS OF FATTY ACIDS doc. Ing. Zenóbia Chavková, CSc. The pathway for the of FAs is not the reversal of the oxidation pathway Both pathways are separated within different cellular compartments In
SensoLyte Rh110 Cathepsin K Assay Kit *Fluorimetric* Revision#1.2 Last Updated: May 2017 Catalog # Kit Size
 SensoLyte Rh110 Cathepsin K Assay Kit *Fluorimetric* Revision#1.2 Last Updated: May 2017 Catalog # 72152 Kit Size 100 Assays (96-well plate) Optimized Performance: This kit detects Cathepsin K activity.
SensoLyte Rh110 Cathepsin K Assay Kit *Fluorimetric* Revision#1.2 Last Updated: May 2017 Catalog # 72152 Kit Size 100 Assays (96-well plate) Optimized Performance: This kit detects Cathepsin K activity.
PFK Activity Assay Kit (Colorimetric)
 PFK Activity Assay Kit (Colorimetric) Catalog Number KA3761 100 assays Version: 02 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Background... 3 General Information...
PFK Activity Assay Kit (Colorimetric) Catalog Number KA3761 100 assays Version: 02 Intended for research use only www.abnova.com Table of Contents Introduction... 3 Background... 3 General Information...
Chemistry 506: Allied Health Chemistry 2. Chapter 22: Biosynthetic Pathways. Making Complex Biomolecules
 Chemistry 506: Allied Health Chemistry 2 1 Chapter 22: Biosynthetic Pathways Making Complex Biomolecules Introduction to General, rganic & Biochemistry, 5 th Edition by Bettelheim and March: Chapter 22,
Chemistry 506: Allied Health Chemistry 2 1 Chapter 22: Biosynthetic Pathways Making Complex Biomolecules Introduction to General, rganic & Biochemistry, 5 th Edition by Bettelheim and March: Chapter 22,
Kit for assay of thioredoxin
 FkTRX-02-V2 Kit for assay of thioredoxin The thioredoxin system is the major protein disulfide reductase in cells and comprises thioredoxin, thioredoxin reductase and NADPH (1). Thioredoxin systems are
FkTRX-02-V2 Kit for assay of thioredoxin The thioredoxin system is the major protein disulfide reductase in cells and comprises thioredoxin, thioredoxin reductase and NADPH (1). Thioredoxin systems are
ANSC/NUTR 618 Lipids & Lipid Metabolism
 Fatty Acid ynthesis I. verall concepts A. Definitions ANC/NUTR 618 Lipids & Lipid Metabolism Fatty Acid ynthesis 1. De novo synthesis = synthesis from non-fatty acid precursors a. Carbohydrate precursors
Fatty Acid ynthesis I. verall concepts A. Definitions ANC/NUTR 618 Lipids & Lipid Metabolism Fatty Acid ynthesis 1. De novo synthesis = synthesis from non-fatty acid precursors a. Carbohydrate precursors
Syllabus for BASIC METABOLIC PRINCIPLES
 Syllabus for BASIC METABOLIC PRINCIPLES The video lecture covers basic principles you will need to know for the lectures covering enzymes and metabolism in Principles of Metabolism and elsewhere in the
Syllabus for BASIC METABOLIC PRINCIPLES The video lecture covers basic principles you will need to know for the lectures covering enzymes and metabolism in Principles of Metabolism and elsewhere in the
ASSAY THEORY 2 ADAPTA ASSAY CONDITIONS 3 ADAPTA ASSAY CONTROLS 4 ADAPTA DATA ANALYSIS 5
 Page 1 of 11 ASSAY THEORY 2 ADAPTA ASSAY CONDITIONS 3 ADAPTA ASSAY CONTROLS 4 ADAPTA DATA ANALYSIS 5 KINASE-SPECIFIC ASSAY CONDITIONS 6 CAMK1 (CaMK1) 6 CDK4/cyclin D1 6 CDK4/cyclin D3 6 CDK6/cyclin D1
Page 1 of 11 ASSAY THEORY 2 ADAPTA ASSAY CONDITIONS 3 ADAPTA ASSAY CONTROLS 4 ADAPTA DATA ANALYSIS 5 KINASE-SPECIFIC ASSAY CONDITIONS 6 CAMK1 (CaMK1) 6 CDK4/cyclin D1 6 CDK4/cyclin D3 6 CDK6/cyclin D1
WHY IS THIS IMPORTANT?
 CHAPTER 3 ESSENTIALS OF METABOLISM WHY IS THIS IMPORTANT? It is important to have a basic understanding of metabolism because it governs the survival and growth of microorganisms The growth of microorganisms
CHAPTER 3 ESSENTIALS OF METABOLISM WHY IS THIS IMPORTANT? It is important to have a basic understanding of metabolism because it governs the survival and growth of microorganisms The growth of microorganisms
III. 6. Test. Respiració cel lular
 III. 6. Test. Respiració cel lular Chapter Questions 1) What is the term for metabolic pathways that release stored energy by breaking down complex molecules? A) anabolic pathways B) catabolic pathways
III. 6. Test. Respiració cel lular Chapter Questions 1) What is the term for metabolic pathways that release stored energy by breaking down complex molecules? A) anabolic pathways B) catabolic pathways
CELLULAR RESPIRATION SUMMARY EQUATION. C 6 H 12 O 6 + O 2 6CO2 + 6H 2 O + energy (ATP) STEPWISE REDOX REACTION
 CELLULAR RESPIRATION SUMMARY EQUATION C 6 H 12 O 6 + O 2 6CO2 + 6H 2 O + energy (ATP) STEPWISE REDOX REACTION Oxidation: partial or complete loss of electrons Reduction: partial or complete gain of electrons
CELLULAR RESPIRATION SUMMARY EQUATION C 6 H 12 O 6 + O 2 6CO2 + 6H 2 O + energy (ATP) STEPWISE REDOX REACTION Oxidation: partial or complete loss of electrons Reduction: partial or complete gain of electrons
Leen Alsahele. Razan Al-zoubi ... Faisal
 25 Leen Alsahele Razan Al-zoubi... Faisal last time we started talking about regulation of fatty acid synthesis and degradation *regulation of fatty acid synthesis by: 1- regulation of acetyl CoA carboxylase
25 Leen Alsahele Razan Al-zoubi... Faisal last time we started talking about regulation of fatty acid synthesis and degradation *regulation of fatty acid synthesis by: 1- regulation of acetyl CoA carboxylase
Enzymes and Metabolism
 PowerPoint Lecture Slides prepared by Vince Austin, University of Kentucky Enzymes and Metabolism Human Anatomy & Physiology, Sixth Edition Elaine N. Marieb 1 Protein Macromolecules composed of combinations
PowerPoint Lecture Slides prepared by Vince Austin, University of Kentucky Enzymes and Metabolism Human Anatomy & Physiology, Sixth Edition Elaine N. Marieb 1 Protein Macromolecules composed of combinations
Ch. 9 Cell Respiration. Title: Oct 15 3:24 PM (1 of 53)
 Ch. 9 Cell Respiration Title: Oct 15 3:24 PM (1 of 53) Essential question: How do cells use stored chemical energy in organic molecules and to generate ATP? Title: Oct 15 3:28 PM (2 of 53) Title: Oct 19
Ch. 9 Cell Respiration Title: Oct 15 3:24 PM (1 of 53) Essential question: How do cells use stored chemical energy in organic molecules and to generate ATP? Title: Oct 15 3:28 PM (2 of 53) Title: Oct 19
CELLULAR METABOLISM. Metabolic pathways can be linear, branched, cyclic or spiral
 CHM333 LECTURE 24 & 25: 3/27 29/13 SPRING 2013 Professor Christine Hrycyna CELLULAR METABOLISM What is metabolism? - How cells acquire, transform, store and use energy - Study reactions in a cell and how
CHM333 LECTURE 24 & 25: 3/27 29/13 SPRING 2013 Professor Christine Hrycyna CELLULAR METABOLISM What is metabolism? - How cells acquire, transform, store and use energy - Study reactions in a cell and how
SelectScreen Biochemical Kinase Profiling Service
 Page 1 of 11 ASSAY THEORY 2 ADAPTA ASSAY CONDITIONS 3 ADAPTA ASSAY CONTROLS 4 ADAPTA DATA ANALYSIS 5 KINASE-SPECIFIC ASSAY CONDITIONS 6 CAMK1 (CaMK1) 6 CDK7/cyclin H/MNAT1 6 CDK9/cyclin T1 6 CHUK (IKK
Page 1 of 11 ASSAY THEORY 2 ADAPTA ASSAY CONDITIONS 3 ADAPTA ASSAY CONTROLS 4 ADAPTA DATA ANALYSIS 5 KINASE-SPECIFIC ASSAY CONDITIONS 6 CAMK1 (CaMK1) 6 CDK7/cyclin H/MNAT1 6 CDK9/cyclin T1 6 CHUK (IKK
Chemistry 506: Allied Health Chemistry 2. Chapter 20: Bioenergetics. Energy Generation in the Cell
 Chemistry 506 Dr. unter s Class Chapter 20. Chemistry 506: Allied ealth Chemistry 2 1 Chapter 20: Bioenergetics Energy Generation in the Cell Introduction to General, rganic & Biochemistry, 5 th Edition
Chemistry 506 Dr. unter s Class Chapter 20. Chemistry 506: Allied ealth Chemistry 2 1 Chapter 20: Bioenergetics Energy Generation in the Cell Introduction to General, rganic & Biochemistry, 5 th Edition
LAST LECTURE. 1. Things the course missed 2. Things you might be tested on
 LAST LECTURE 1. Things the course missed 2. Things you might be tested on Things I didn t tell you that might make more sense now that you have had BIS105 From Regulation of Cellular Metabolism by Protein
LAST LECTURE 1. Things the course missed 2. Things you might be tested on Things I didn t tell you that might make more sense now that you have had BIS105 From Regulation of Cellular Metabolism by Protein
number Done by Corrected by Doctor Faisal Al-Khatibe
 number 24 Done by Mohammed tarabieh Corrected by Doctor Faisal Al-Khatibe 1 P a g e *Please look over the previous sheet about fatty acid synthesis **Oxidation(degradation) of fatty acids, occurs in the
number 24 Done by Mohammed tarabieh Corrected by Doctor Faisal Al-Khatibe 1 P a g e *Please look over the previous sheet about fatty acid synthesis **Oxidation(degradation) of fatty acids, occurs in the
Plate-Based Assay Methods for the Assessment of Cellular Health
 Plate-Based Assay Methods for the Assessment of Cellular Health Andrew L. Niles, Senior Research Scientist 2012, Promega Corporation. Biological Outcomes in Cell Culture Treatment -Small molecule -Bio-molecule
Plate-Based Assay Methods for the Assessment of Cellular Health Andrew L. Niles, Senior Research Scientist 2012, Promega Corporation. Biological Outcomes in Cell Culture Treatment -Small molecule -Bio-molecule
HDAC1 Inhibitor Screening Assay Kit
 HDAC1 Inhibitor Screening Assay Kit Item No. 10011564 www.caymanchem.com Customer Service 800.364.9897 Technical Support 888.526.5351 1180 E. Ellsworth Rd Ann Arbor, MI USA TABLE OF CONTENTS GENERAL INFORMATION
HDAC1 Inhibitor Screening Assay Kit Item No. 10011564 www.caymanchem.com Customer Service 800.364.9897 Technical Support 888.526.5351 1180 E. Ellsworth Rd Ann Arbor, MI USA TABLE OF CONTENTS GENERAL INFORMATION
CHE 242 Exam 3 Practice Questions
 CHE 242 Exam 3 Practice Questions Glucose metabolism 1. Below is depicted glucose catabolism. Indicate on the pathways the following: A) which reaction(s) of glycolysis are irreversible B) where energy
CHE 242 Exam 3 Practice Questions Glucose metabolism 1. Below is depicted glucose catabolism. Indicate on the pathways the following: A) which reaction(s) of glycolysis are irreversible B) where energy
Ahmad Ulnar. Faisal Nimri ... Dr.Faisal
 24 Ahmad Ulnar Faisal Nimri... Dr.Faisal Fatty Acid Synthesis - Occurs mainly in the Liver (to store excess carbohydrates as triacylglycerols(fat)) and in lactating mammary glands (for the production of
24 Ahmad Ulnar Faisal Nimri... Dr.Faisal Fatty Acid Synthesis - Occurs mainly in the Liver (to store excess carbohydrates as triacylglycerols(fat)) and in lactating mammary glands (for the production of
Notes CELLULAR RESPIRATION SUMMARY EQUATION C 6 H 12 O 6 + O 2. 6CO 2 + 6H 2 O + energy (ATP) STEPWISE REDOX REACTION
 AP BIOLOGY CELLULAR ENERGETICS ACTIVITY #2 Notes NAME DATE HOUR SUMMARY EQUATION CELLULAR RESPIRATION C 6 H 12 O 6 + O 2 6CO 2 + 6H 2 O + energy (ATP) STEPWISE REDOX REACTION Oxidation: partial or complete
AP BIOLOGY CELLULAR ENERGETICS ACTIVITY #2 Notes NAME DATE HOUR SUMMARY EQUATION CELLULAR RESPIRATION C 6 H 12 O 6 + O 2 6CO 2 + 6H 2 O + energy (ATP) STEPWISE REDOX REACTION Oxidation: partial or complete
Respiration. Respiration. Respiration. How Cells Harvest Energy. Chapter 7
 How Cells Harvest Energy Chapter 7 Organisms can be classified based on how they obtain energy: autotrophs: are able to produce their own organic molecules through photosynthesis heterotrophs: live on
How Cells Harvest Energy Chapter 7 Organisms can be classified based on how they obtain energy: autotrophs: are able to produce their own organic molecules through photosynthesis heterotrophs: live on
Expert Intelligence for Better Decisions Epigenetics:
 Expert Intelligence for Better Decisions Epigenetics: Emerging Targets, Available Technologies, Expert Interviews, and an Epigenetic Community Perspective Using This Document Insight Pharma Reports are
Expert Intelligence for Better Decisions Epigenetics: Emerging Targets, Available Technologies, Expert Interviews, and an Epigenetic Community Perspective Using This Document Insight Pharma Reports are
TCA CYCLE (Citric Acid Cycle)
 TCA CYCLE (Citric Acid Cycle) TCA CYCLE The Citric Acid Cycle is also known as: Kreb s cycle Sir Hans Krebs Nobel prize, 1953 TCA (tricarboxylic acid) cycle The citric acid cycle requires aerobic conditions!!!!
TCA CYCLE (Citric Acid Cycle) TCA CYCLE The Citric Acid Cycle is also known as: Kreb s cycle Sir Hans Krebs Nobel prize, 1953 TCA (tricarboxylic acid) cycle The citric acid cycle requires aerobic conditions!!!!
ENZYMES: CLASSIFICATION, STRUCTURE
 ENZYMES: CLASSIFICATION, STRUCTURE Enzymes - catalysts of biological reactions Accelerate reactions by a millions fold Common features for enzymes and inorganic catalysts: 1. Catalyze only thermodynamically
ENZYMES: CLASSIFICATION, STRUCTURE Enzymes - catalysts of biological reactions Accelerate reactions by a millions fold Common features for enzymes and inorganic catalysts: 1. Catalyze only thermodynamically
Gene Expression DNA RNA. Protein. Metabolites, stress, environment
 Gene Expression DNA RNA Protein Metabolites, stress, environment 1 EPIGENETICS The study of alterations in gene function that cannot be explained by changes in DNA sequence. Epigenetic gene regulatory
Gene Expression DNA RNA Protein Metabolites, stress, environment 1 EPIGENETICS The study of alterations in gene function that cannot be explained by changes in DNA sequence. Epigenetic gene regulatory
Ch 9: Cellular Respiration
 Ch 9: Cellular Respiration Cellular Respiration An overview Exergonic reactions and catabolic pathway Energy stored in bonds of food molecules is transferred to ATP Cellular respiration provides the energy
Ch 9: Cellular Respiration Cellular Respiration An overview Exergonic reactions and catabolic pathway Energy stored in bonds of food molecules is transferred to ATP Cellular respiration provides the energy
Releasing Chemical Energy
 Releasing Chemical Energy Ø Energy From Carbohydrates Ø Aerobic Respiration/ Stages Ø Fermentation Ø Food as a Source of Energy How Do Cells Access the Chemical Energy in Carbohydrayes? Aerobic Respiration
Releasing Chemical Energy Ø Energy From Carbohydrates Ø Aerobic Respiration/ Stages Ø Fermentation Ø Food as a Source of Energy How Do Cells Access the Chemical Energy in Carbohydrayes? Aerobic Respiration
Glycolysis. Cellular Respiration
 Glucose is the preferred carbohydrate of cells. In solution, it can change from a linear chain to a ring. Energy is stored in the bonds of the carbohydrates. Breaking these bonds releases that energy.
Glucose is the preferred carbohydrate of cells. In solution, it can change from a linear chain to a ring. Energy is stored in the bonds of the carbohydrates. Breaking these bonds releases that energy.
Roles of Lipids. principal form of stored energy major constituents of cell membranes vitamins messengers intra and extracellular
 Roles of Lipids principal form of stored energy major constituents of cell membranes vitamins messengers intra and extracellular = Oxidation of fatty acids Central energy-yielding pathway in animals. O
Roles of Lipids principal form of stored energy major constituents of cell membranes vitamins messengers intra and extracellular = Oxidation of fatty acids Central energy-yielding pathway in animals. O
Energy storage in cells
 Energy storage in cells Josef Fontana EC - 58 Overview of the lecture Introduction to the storage substances of human body Overview of storage compounds in the body Glycogen metabolism Structure of glycogen
Energy storage in cells Josef Fontana EC - 58 Overview of the lecture Introduction to the storage substances of human body Overview of storage compounds in the body Glycogen metabolism Structure of glycogen
Anabolism of Fatty acids (Anabolic Lynen spiral) Glycerol and Triglycerides
 Anabolism of Fatty acids (Anabolic Lynen spiral) Glycerol and Triglycerides Anabolism of fatty acids Fatty acids are not stored in the body free. They are a source of energy in the form of triglycerides
Anabolism of Fatty acids (Anabolic Lynen spiral) Glycerol and Triglycerides Anabolism of fatty acids Fatty acids are not stored in the body free. They are a source of energy in the form of triglycerides
Biology 30 Structure & Function of Cells (Part 2) Bioenergetics: Energy: Potential energy: Examples: Kinetic energy. Examples:
 Biology 30 Structure & Function of Cells (Part 2) Bioenergetics: Energy: Potential energy: Examples: Kinetic energy Examples: Energy can be transformed: Thermodynamics: First law of Thermodynamics: Second
Biology 30 Structure & Function of Cells (Part 2) Bioenergetics: Energy: Potential energy: Examples: Kinetic energy Examples: Energy can be transformed: Thermodynamics: First law of Thermodynamics: Second
Chapter 14. Energy conversion: Energy & Behavior
 Chapter 14 Energy conversion: Energy & Behavior Why do you Eat and Breath? To generate ATP Foods, Oxygen, and Mitochodria Cells Obtain Energy by the Oxidation of Organic Molecules Food making ATP making
Chapter 14 Energy conversion: Energy & Behavior Why do you Eat and Breath? To generate ATP Foods, Oxygen, and Mitochodria Cells Obtain Energy by the Oxidation of Organic Molecules Food making ATP making
ANSC 689 PHYSIOLOGICAL CHEMISTRY OF LIVESTOCK SPECIDS. Enzyme Kinetics and Control Reactions
 Handout Enzyme Kinetics and Control Reactions ANSC 689 PHYSIOLOGICAL CHEMISTRY OF LIVESTOCK SPECIDS Enzyme Kinetics and Control Reactions I. Kinetics A. Reaction rates 1. First order (reaction rate is
Handout Enzyme Kinetics and Control Reactions ANSC 689 PHYSIOLOGICAL CHEMISTRY OF LIVESTOCK SPECIDS Enzyme Kinetics and Control Reactions I. Kinetics A. Reaction rates 1. First order (reaction rate is
SensoLyte 520 HIV-1 Protease Assay Kit *Fluorimetric*
 SensoLyte 520 HIV-1 Protease Assay Kit *Fluorimetric* Catalog # 71147 Kit Size 100 assays (96-well) or 500 assays (384-well) Convenient Format: Complete kit including all the assay components. Optimized
SensoLyte 520 HIV-1 Protease Assay Kit *Fluorimetric* Catalog # 71147 Kit Size 100 assays (96-well) or 500 assays (384-well) Convenient Format: Complete kit including all the assay components. Optimized
Mangifera indica activates key metabolic master switch enzymes The innovation for well-aging
 Mangifera indica activates key metabolic master switch enzymes The innovation for well-aging Dr. S. Buchwald-Werner 6. May 2015 Vitafoods Europe Conference Healthy Aging Session Consumer demand for healthy-aging
Mangifera indica activates key metabolic master switch enzymes The innovation for well-aging Dr. S. Buchwald-Werner 6. May 2015 Vitafoods Europe Conference Healthy Aging Session Consumer demand for healthy-aging
MRP2 TR ATPase Assay Protocol CAT. NO. SBAT03
 MRP2 TR ATPase CAT. NO. SBAT03 Page 1 of 18 Determination of the interaction of drugs with the human MRP2 (ABCC2) transporter using the ATPase Assay For the following membrane products: SB-MRP2-Sf9-ATPase
MRP2 TR ATPase CAT. NO. SBAT03 Page 1 of 18 Determination of the interaction of drugs with the human MRP2 (ABCC2) transporter using the ATPase Assay For the following membrane products: SB-MRP2-Sf9-ATPase
Coenzymes, vitamins and trace elements 209. Petr Tůma Eva Samcová
 Coenzymes, vitamins and trace elements 209 Petr Tůma Eva Samcová History and nomenclature of enzymes 1810, Gay-Lussac made an experiment with yeats alter saccharide to ethanol and CO 2 Fermentation From
Coenzymes, vitamins and trace elements 209 Petr Tůma Eva Samcová History and nomenclature of enzymes 1810, Gay-Lussac made an experiment with yeats alter saccharide to ethanol and CO 2 Fermentation From
Metabolism Energy Pathways Biosynthesis. Catabolism Anabolism Enzymes
 Topics Microbial Metabolism Metabolism Energy Pathways Biosynthesis 2 Metabolism Catabolism Catabolism Anabolism Enzymes Breakdown of complex organic molecules in order to extract energy and dform simpler
Topics Microbial Metabolism Metabolism Energy Pathways Biosynthesis 2 Metabolism Catabolism Catabolism Anabolism Enzymes Breakdown of complex organic molecules in order to extract energy and dform simpler
Lecture 11 - Biosynthesis of Amino Acids
 Lecture 11 - Biosynthesis of Amino Acids Chem 454: Regulatory Mechanisms in Biochemistry University of Wisconsin-Eau Claire 1 Introduction Biosynthetic pathways for amino acids, nucleotides and lipids
Lecture 11 - Biosynthesis of Amino Acids Chem 454: Regulatory Mechanisms in Biochemistry University of Wisconsin-Eau Claire 1 Introduction Biosynthetic pathways for amino acids, nucleotides and lipids
Chapter 5. Microbial Metabolism
 Chapter 5 Microbial Metabolism Metabolism Collection of controlled biochemical reactions that take place within a microbe Ultimate function of metabolism is to reproduce the organism Metabolic Processes
Chapter 5 Microbial Metabolism Metabolism Collection of controlled biochemical reactions that take place within a microbe Ultimate function of metabolism is to reproduce the organism Metabolic Processes
Lecture 16. Finish lipid metabolism (Triglycerides, Isoprenoids/Steroids, Glyoxylate cycle) Amino acid metabolism (Urea cycle) Google Man III
 Lecture 16 Finish lipid metabolism (Triglycerides, Isoprenoids/Steroids, Glyoxylate cycle) Amino acid metabolism (Urea cycle) Google Man III The Powertrain of Human Metabolism (verview) CARBHYDRATES PRTEINS
Lecture 16 Finish lipid metabolism (Triglycerides, Isoprenoids/Steroids, Glyoxylate cycle) Amino acid metabolism (Urea cycle) Google Man III The Powertrain of Human Metabolism (verview) CARBHYDRATES PRTEINS
Regulation of Enzyme Activity
 Regulation of Enzyme Activity Enzyme activity must be regulated so that the proper levels of products are produced at all times and places This control occurs in several ways: - biosynthesis at the genetic
Regulation of Enzyme Activity Enzyme activity must be regulated so that the proper levels of products are produced at all times and places This control occurs in several ways: - biosynthesis at the genetic
Jayanti Tokas 1, Puneet Tokas 2, Shailini Jain 3 and Hariom Yadav 3
 Jayanti Tokas 1, Puneet Tokas 2, Shailini Jain 3 and Hariom Yadav 3 1 Department of Biotechnology, JMIT, Radaur, Haryana, India 2 KITM, Kurukshetra, Haryana, India 3 NIDDK, National Institute of Health,
Jayanti Tokas 1, Puneet Tokas 2, Shailini Jain 3 and Hariom Yadav 3 1 Department of Biotechnology, JMIT, Radaur, Haryana, India 2 KITM, Kurukshetra, Haryana, India 3 NIDDK, National Institute of Health,
Notes CELLULAR RESPIRATION SUMMARY EQUATION C 6 H 12 O 6 + O 2. 6CO 2 + 6H 2 O + energy (ATP) STEPWISE REDOX REACTION
 AP BIOLOGY CELLULAR ENERGETICS ACTIVITY #2 Notes NAME DATE HOUR SUMMARY EQUATION CELLULAR RESPIRATION C 6 H 12 O 6 + O 2 6CO 2 + 6H 2 O + energy (ATP) STEPWISE REDOX REACTION Oxidation: partial or complete
AP BIOLOGY CELLULAR ENERGETICS ACTIVITY #2 Notes NAME DATE HOUR SUMMARY EQUATION CELLULAR RESPIRATION C 6 H 12 O 6 + O 2 6CO 2 + 6H 2 O + energy (ATP) STEPWISE REDOX REACTION Oxidation: partial or complete
SensoLyte Generic MMP Assay Kit *Colorimetric*
 SensoLyte Generic MMP Assay Kit *Colorimetric* Revision#1.2 Catalog # Kit Size Last updated: May2017 AS-72095 100 Assays (96-well plate) Optimized Performance: This kit is optimized to detect MMP activity
SensoLyte Generic MMP Assay Kit *Colorimetric* Revision#1.2 Catalog # Kit Size Last updated: May2017 AS-72095 100 Assays (96-well plate) Optimized Performance: This kit is optimized to detect MMP activity
