Effect of oxidation on the properties of apolipoproteins A4 and A 4
|
|
|
- Dominick Williamson
- 5 years ago
- Views:
Transcription
1 Effect of oxidation on the properties of apolipoproteins A4 and A 4 G. M. Anantharamaiah, * Thomas A. Hughes, M. Iqbal, * Ali Gawish, * Peter J. Neame, * Mark F. Medley,* and Jere P. Segrestt Departments of Medicine* and Biochemistry, t and the Atherosclerosis Research Unit, University of Alabama at Birmingham, Birmingham, AL Abstract Purified apolipoprotein A-I has been separated by reversed-phase high performance liquid chromatography (HPLC) into multiple peaks and these peaks have been characterized. One peak, apoa-ib had a relatively longer retention time on HPLC but its retention time could be shortened by treatment by hydrogen peroxide. CNBr cleavage studies indicated that the differences in apoa-ib and in its oxidation product, apoa-la, were due to the different oxidation states of methionine. This phenomenon was also observed in apoa-11, where methionine oxidation produced two more forms of this apolipoprotein in addition to the native form. These isomers were found to have different secondary structures and affinities for lipid. Model peptide analogs of the amphipathic helix with the same sequence but with methionine and methionine sulfoxide at the nonpolar face of the amphipathic helix were synthesized and studied. It was found that the lipid affinities of these synthetic peptide isomers were very different. They also differed in their secondary structures as studied by circular dichroism (CD). a We propose that methionine oxidation introduces hydrophilic residues at the nonpolar face of the amphipathic helical domains of these apolipoproteins and, therefore, alters their secondary structure and lipid affinity. -Anantharamaiah, G. M., T. A. Hughes, M. Iqbal, A. Gawish, P. J. Neame, M. F. Medley, and J. P. Segrest. Effect of oxidation on the properties of apolipoproteins A-I and A-11. J. Lipid Res : Supplementary key words apoa-i - apoa-i1 methionine oxidation synthetic peptides amphipathic helix apolipoprotein isomers HPLC High density lipoproteins (HDL) are protein-lipid cornplexes found in plasma with a density range of 1.63 to 1.21 g/ml. There is now substantial evidence that HDL tholesterol concentration is inversely related to the incidence of coronary artery disease (1, 2). HDL is composed of approximately 5% protein of which apolipoproteins A-I and A-I1 make up about 9% of the total protein. The primary structures of these apolipoproteins are known (3). Although extensive investigations have been carried out on the behaviors of these apolipoproteins, particularly apoa-i, conflitting reports exist on their molecular properties (4). It is known that apoa-i and A-I1 contain repetitive helical domains with opposing nonpolar/polar faces (amphipathic helices). This structure is believed to contribute to the ability of these apolipoproteins to solubilize lipid in an aqueous environment (5). Recently, apoa-i has been reported to exist in different polymorphic forms (6-9). Some investigators have suggested that these forms arise due to a sequential deamidation process which gives rise to successively acidic isomers (8). It has also been suggested that differences in the polymorphic forms of apoa-i are due to oxidation of the protein (1). There are no reports in the literature on the characterization of apoa-i1 isoforms. This laboratory has recently developed a reversedphase HPLC methodo~ow for the apolipoprotein composition of plasma lipoproteins (11). Using this technique, we have two forms of apoa-i (apoa-la and apoa-ib) and three forms of apoa-i1 (apoa-iia, apoa-iib, and apoa-iic). In this report, we will present data that indicates that these forms of apo~-~ and A-II (as identified by HPLC) are due to the oxidation of methionine in these Proteins. TWO model Peptide analogs of the amphipathic helix (with the same amino acid sequence but with methionine or methionine sulfoxide on the nonpolar face of the amphipathic helix) have been synthesized and studied. Results from these studies support our contention that methionine oxidation of peptides with identical primary structures leads to differences in HPLC retention times. Furthermore, these peptides have very different lipid binding characteristics and secondary structural features indicating that methionine oxidation of apolipoproteins is likely to significantly alter their interactions with lipids. Abbreviations are in accordance with IUPAC-IUB nomenclature. Others used are: Met(()), methionine sulfoxide; HDL, high density lipoproteins; DMPC, dimyristoylphosphatidylcholine; SDS-PAGE, sodium dodecylsulfate-polyacrylamide gel electrophoresis; CD, circular dichroism; HPLC, high performance liquid chromatography. Present address: Department of Medicine, University of Tennessee, Memphis, TN Downloaded from by guest, on September 22, 218 Journal of Lipid Research Volume 29,
2 Lipoprotein isolation METHODS HDL (d g/ml) was isolated by gradient ultracentrifugation. Ten ml of plasma was raised to a density of 1.35 g/ml with 5.94 of KBr. A second layer of buffer (1 ml) (NaC1.9%, EDTA 1. mm, Tris 1 mm, sodium azide.1%, ph 8.5, containing protease inhibitors 6- aminohexanoic acid (1 mm), benzamidine-hcl(5 mm), and phenylmethylsulfonylfluoride (1. mm)) raised to a density of 1.2 g/ml with KBr, was added to the tube and, finally, the tube was filled with the above buffer (d 1.6 g/ml) to a final volume of approximately 4 ml. The tubes were sealed and spun to 8. x 1" rad'. sec-' at 7, rpm at 15OC in a 7 Ti rotor (approximately 4 hr and 15 min). HDL was dialyzed at 4OC against.1 M ammonium bicarbonate buffer,.1% NaN3, ph 8.. The lipoproteins were delipidated with five volumes of hexane isopropanol 3:2. The aqueous layer was dried and the proteins were solubilized in 3 M guanidine-hc1, and then analyzed by HPLC. Analytical HPLC procedure A Perkin Elmer Series 4 pump was used with an LC-55 or LC-75 UV detector at 214 nm. Peak areas were quanti- 2 HPLC of HDL P i tated by a Shimadzu data station. Analytical reversedphase HPLC was done on a Vydac C-18 column (4.6 x 25 mm) as a gradient of acetonitrile in water (containing.1% TFA) from 25 to 58% at l.o%/min, column temperature, 5OoC and a flow rate of 1.2 ml/min. Fig. 1 shows typical chromatograms of apohdl and apoa-i. These peaks were isolated and identified by amino acid analysis (Table l), N-terminal analysis, and SDS polyacrylamide gel electrophoresis. Apolipoprotein isomers with the following retention times on the analytical HPLC were named as follows: apoa-ia, 26.4 min; apoa-ib, 28.2 min; apoa-iic, 29.4 min; apoa-iia, 3.7 min; apoa-iib, 31.8 min. Purification of apolipoprotein A-I and A-I1 isomers The preparative HPLC method used to isolate these proteins in large amounts (5-1 mg) was similar to the analytical method described above, except that a larger column (1 mm diameter) and a slower gradient (.5%/ min) were used. Peaks were identified by analytical HPLC analysis. Circular dichroism The CD spectra were taken using a JASCO spectropolarimeter coupled to a DPN-5 integrator. ApoA-Ia s B HPLC Of AvA-1 Downloaded from by guest, on September 22, 218 Time(m in) Fig. 1. HPLC profile of apohdl and apoa-i. Panel A: human apohdl was analyzed by HPLC as described in Methods. The isomers of apoa-i and apoa-i1 are indicated. Panel B: apoa-i, purified by S-2 gel chromatography, was analyzed by HPLC. Isomers of apoa-i are identified. 31 Journal of Lipid Research Volume 29, 1988
3 ~ ~ ~ ~~ ~~ TABLE 1. Amino acid composition ratios and values for amino acids/molecule used to identify apoa-i and apoa-i1 Obtained Obtained Expected Expected ApoA-I A-la A-Ih ApoA-I1 A-iIa A-IIh A-llc His Met lie Asu/Glx Thr/Ala Leu/Phe Values for His, Met, and Ile are residues per molecule of apoa-i and per molecule of apoa-i1 dimer. Hydrolyses were performed at 1 iooc for 2 hr in 6 M HCI in the presence of dithiothreitol to reduce methionine sulfoxide to methionine. Amino acids were analyzed on a Durrum amino acid analyzer using post column derivatization with -phthalaldehyde. and apoa-ib (5 pglml) were dissolved in 1 mm phosphate buffer, 15 mm NaC1, ph 8.. Synthetic peptide analogs of apolipoproteins were also studied under the same conditions. Electron microscopy The apolipoprotein or synthetic peptides complexed with unilamellar vesicles of DMPC at 1:l or 12 weight ratio were stained with 2% potassium phosphotungstate, ph 5.9, and examined with a Philips EM4 microscope on carbon-coated Formvar grids. Gradient gel electrophoresis A 1-25% linear acrylamide gradient gel was formed. Electrophoresis was performed at 4OC with 3 V162.5 ma in the stacking gel and 5 VI8 ma in the running gel. The chamber buffer was Tris-glycine with.1% SDS at ph 8.5. The gel was stained in.25% Coomassie Blue. Oxidation with hydrogen peroxide Purified apolipoprotein isomers (7 pg of apoa-ib or 24 pg of apoa-iib) were treated with hydrogen peroxide solution (1 pl of 3% in the case of apoa-ib and 1, 3, and 2 ~1 of.3% in the case of apoa-iib) at room temperature for 5 min. The oxidation products were studied by analytical HPLC. Treatment with mercaptoethanol Apolipoproteins A-Ia, A-IIa, A-IIb, and A-IIc (5 pg) were treated with mercaptoethanol (3 pl) and heated in boiling water for 2 sec. The reduction products were examined by analytical HPLC. CNBr treatment One mg of each protein was dissolved in 8% HCOOH (.5 ml) and treated with the same amount of CNBr at room temperature for 24 hr. The reaction mixtures were diluted with 5 ml of H2 and lyophilized (12). Peptide synthesis Two peptide analogs of 18A ([Met3]18A and [Met()']18A) were synthesized and purified following our procedure described elsewhere (13). These peptide analogs differ in having Met or Met() at the third position from the N-terminus of the 18A peptide with the following sequence: Asp-Trp-Leu-Lys-Ala-Pherljrr-Asp-Lys-Val-~a-Glu-Lys-Leu- Lys-Glu-Ala-Phe, which has been well characterized by us (13). RESULTS Apolipoprotein isolation and characterization ApoA-I has three methionines at positions 86, 112, and 148 and apoa-i1 has one methionine at position 26 from the N-terminus. Based on the amino acid analysis and N-terminal analysis of individual peaks, the peaks seen on the HPLC chromatograph of apohdl were designated as shown in Fig. 1A. Fig. 1B shows that apoa-i, isolated by conventional gel filtration (on S-2 sephacryl in 3 M guanidine- HCl) is a mixture of two isomers (apoa-ia and apoa-ib). These isomers were subjected to SDS-PAGE, amino acid analyses, and N-terminal sequencing to three residues (Table 1). These studies formally identified the peaks observed by reversed-phase HPLC. Similar studies of the apoa-i1 isomers (Table 1) again identified all peaks as being apoa-11. No N-terminal residue for peaks due to apoa-i1 was found as they contain pyroglutamic acid which does not react with P IX used in the Edman degradation procedure. Trypsin digestion of the apolipoprotein isomers also produced HPLC peptide maps with similar fragments (data not shown). Oxidation Of apolipoproteins ApoA-Ib, apoa-ia, apohdl, and apoa-iib were treated with hydrogen peroxide, and subjected to HPLC analysis. Downloaded from by guest, on September 22, 218 Anantharamaiah et al. Oxidized forms of apolipoproteins A-I and A-I1 311
4 9 A-lb Control Y OD 1! t c -(II i v n n I I I Time( m in) I 1 I Fig. 2. HPLC chromatogram of apoa-ib and apoa-ib treated with hydrogen peroxide. Panel A: HPLC of untreated apoa-ib. Panel B: HPLC of aga-ib after treatment. The following results were observed. ApoA-Ib completely disappeared, and a new peak (with the same retention time as A-Ia in Fig. 1B) appeared (Fig. 2). The new peak is a chemically prepared apoa-ia and is designated as apoa-ia*. There was no change in the retention time of apoa-ia following treatment with hydrogen peroxide L,- 3 (results not shown). ApoA-1Ib was progressively converted to apoa-iia and then to apoa-iic with increasing amounts of hydrogen peroxide (Fig. 3). Similar changes were observed when whole HDL was treated with hydrogen peroxide. These results indicated that apoa-ia, apoa-iia, and apoa-iic are the oxidized forms of these apolipoproteins. C I Downloaded from by guest, on September 22, '-r- Timelmlnl I rimelmin) rim~lmln) Fig. 3. HPLC chromatogram of apoa-iib and its conversion products after treatment with increasing amounts of hydrogen peroxide. Panel A: apoa-iib before treatment. Panel B: apoa-iib after treatment with 1 pl of.3% hydrogen peroxide. Panel C: apoa-iib after treatment with 3 pl of.3% hydrogen peroxide. Panel D: apoa-iib after treatment with 2 pl of.3% hydrogen peroxide. 312 Journal of Lipid Research Volume 29, 1988
5 CNBr studies The results of CNBr digestion of apoa-ia and apoa-ib are shown in Fig. 4. CNBr specifically cleaves peptides at the methionine residues (12) but when methionine is oxidized, the protein becomes resistant to CNBr digestion. As can be seen in Fig. 4, apoa-ia* was almost completely resistant to CNBr, while apoa-ib underwent substantial fragmentation. These results indicate that apoa-ia* contains almost entirely oxidized methionine residues while apoa-ib contains reduced methionine residues. Since the CNBr/amino acid analysis methodology is unlikely to give complete results in either direction, the data are relative rather than absolute. That is, A-Ia is more oxidized than A-Ib. ApoA-I and A-I1 isomers were treated with CNBr for 1 hr under acidic conditions in order to convert the methionine residues to homoserine. This reaction does not alter oxidized methionine residues. Acid hydrolysis of each of the isomers was then performed under reducing conditions (which convert oxidized methionine to reduced methionine) and the hydrolyzate was analyzed for its amino acid composition. The amount of methionine mea TABLE 2. Apolipoprotein Determination of the degree of methionine oxidation in apoa-i and A-I1 Average Number of Methionine ResidueslMolecule A-Ib 1.47 f.12 A-la 1.95 f.16b A-IIb.4 f.5 A-IIa 1.14 f.2 A-IIc 1.64 f.5 Apolipoproteins (in triplicate) were incubated with CNBr for 1 hr under acidic conditions in order to convert methionine residues to homoserine. Methionine sulfoxide is not altered during this reaction. The protein was then hydrolyzed under reducing conditions to convert methionine sulfoxide to methionine. An amino acid analysis was then performed. The amount of methionine measured gives an estimate of the amount of methionine sulfoxide in the original protein. The values for apoa-i1 reflect for the dimer. bp <.1 versus value immediately above. sured, therefore, reflects the amount of oxidized methionine in the original sample. Table 2 shows that there were highly significant differences in the amount of oxidized methionine in the apolipoprotein isomers and that the shorter the HPLC retention time the greater the degree of methionine oxidation. One sample of apoa-ia* was analyzed and it was found to contain 95% oxidized methionine. This is consistent with the high degree of resistance to CNBr digestion noted above. Treatment of apolipoprotein isomers with mercaptoethanol ApoA-I1 is a dimer with two identical peptide monomers connected by a disulfide bond. Each monomer contains a single methionine residue at position 26 from the N- terminus. Fig. 5 shows that reduction of apoa-iia produced two peaks of almost equal size while reduction of apoa-iib resulted in a single peak with a retention time identical to the second peak from apoa-iia (Fig. 5). ApoA-IIc produced a single peak with the retention time identical to the first peak produced from apoa-iia. These results indicate that apoa-iib and apoa-iic are made up of identical monomers and apoa-iia is a mixed dimer. Since apoa-i does not contain disulfide bonds and it is unlikely that the mild reducing conditions used in these experiments would be able to reduce methionine residues, we would not expect mercaptoethanol to have any effect on apoa-ia or A-Ib retention times. This is what was observed. Downloaded from by guest, on September 22, 218 Fig. 4. SDS-polyacrylamide g el electrophoresis (1-25% gradient) of apoa-la, apoa-lb, and their CNRr fragments. Lancs 1 and 3: apoa-la and apoa-ib, respectively, prior to CNRr digestion. Lanes 2 and 4 apoa-la and apoa-lb, respectively, after CNBr digestion. Lane 5: molecular weight standards. Synthetic peptide analogs of the amphipathic helix In our previous studies, we have shown that our model synthetic amphipathic peptide analog (18A) with a sequence AspTip-Leu-Lys-Ala-PheTyr-Asp-Lys-Val-Ala-Glu-Lys-Leu- Lys-Glu-Ala-Phe, mimics the properties of apoa-i (13). In order to investigate the effects of methionine oxidation on Anantharamaiah et al. Oxidized forms of apolipoproteins A-I and A-I1 313
6 AA 3 A-lla Control A-lla+ Mercaptoethanol z -. t - I- O C,z A - I I I A-llc Control 2 lb A-llc+ Mercaptoethanol f N Downloaded from by guest, on September 22, 218 b Ib 2 Time( m in) Ib 2' $ Fig. 5. A: HPLC chromatogram of apoa-iia and its conversion products after treatment with mercaptoethanol. Panel A: apoa-iia before treatment. Panel B: apoa-iia after treatment with mercaptoethanol. B: HPLC chromatogram of apoa-iib and its conversion products after treatment with mercaptoethanol. Panel A: apaa-iib before treatment. Panel B: apoa-iib after treatment with mercaptoethanol. C: HPLC chromatogram of apaa-iic and its conversion products after treatment with mercaptoethanol. Panel A: apoa-iic before treatment. Panel B: apoa-iic after treatment with mercaptoethanol. 314 Journal of Lipid Research Volume 29, 1988
7 \ A-llb control h. F m 3 C I." cl" I the properties of the lipid binding domains of apolipoproteins, we synthesized two analogs of the 18A amphipathic peptide ([Met3]18A and [Met()3]18A). Position three of these analogs is in the center of the nonpolar face of the amphipathic helix (13). Peptide [Met()3]18A was found to have a shorter retention time on reversed-phase HPLC than [Met3]18A (Fig. 6). This was similar to the oxidized apolipoprotein isomers which also had a shorter retention time than the reduced isomers. Circular dichroism CD of the [Met()3]18A and [Met3]18A peptides in solution and in the presence of DMPC demonstrated that the oxidized peptide analog had less helicity than the reduced analog. Similar results were obtained with the apoa-i and A-I1 isomers (Table 1) except that apoa-iia (with one oxidized methionine) had slightly greater helicity than apoa-iib (no oxidized methionines). The fully oxidized apoa-i1 (A-IIc) had significantly less helicity than either of the other isomers. These studies indicate that an alteration in the hydrophobicity of a single amino acid residue on the nonpolar face of the amphipathic helix can cause changes in the secondary structural features. Apolipoprotein-lipid complexes Peptide analogs and apolipoprotein isomers were mixed with DMPC at 1:l weight ratio. While [Met3]18A clarified I I I unilamellar vesicles of DMPC instantly, [Met()']18A had little effect. Even at a 2:l peptide-lipid ratio, the analog with oxidized methionine only slowly clarified DMPC vesicles. Electron microscopy of the complexes (Fig. 7) showed that even at a higher peptide-lipid ratio, particles formed with [Met()']18A were larger (diameter 16.8 * 1.7 nm) than those formed by [Met3]18A at a 1:l weight ratio (diameter nm, P <.1). DISCUSSION That oxidation of a single methionine changes the properties of peptides and proteins is well documented in the literature (14-16). No such studies have been reported to date in the case of apolipoproteins. Apolipoproteins A-I, A-IV, and E have been shown to contain multiple amphipathic helical domains, which are lipid binding domains (5). In the case of human apoa-i, there are eight 22mer amphipathic helical domains (3). Three methionine residues are present at positions 86, 112, and 148 (helices 2, 3, and 5, respectively) (3). Examination of the helical wheel representations of these regions indicates that all three methionines are present in the nonpolar face of amphipathic helical domains. It has been shown by studies of synthetic peptide analogs of the amphipathic helix that an alteration in the hydrophobicity of the non- Downloaded from by guest, on September 22, 218 Anantharamaiah et al. Oxidized forms of apolipoproteins A-I and A-I1 315
8 A B f c Fig. 6. HPLC chromatogram of synthetic amphipathic peptide analogs of MA containing methionine and methionine sulfoxide. Panel A: [Met3]18A analog. Panel B: [Met()3]18A analog. polar face directly affects lipid-associating properties (17). Oxidation of methionine to methionine sulfoxide reduces the hydrophobicity of this amino acid and, therefore, should reduce the hydrophobicity of the protein. It would be expected that a reduction in the hydrophobicity of a protein would result in a lower affinity for the reversedphase HPLC column (shorter retention time) while a reduction in the hydrophobicity of the nonpolar face of the protein is expected to reduce the affinity for DMPC. Indeed, this was what was observed in the case of the synthetic analogs and apolipoprotein isomers. Our CD studies showed that the oxidized peptide analog, apoa-ia, and apoa-iic have less alpha-helicity than the corresponding reduced proteins, indicating that oxidation of methionine also alters the secondary structure of these proteins. Since there are three methionine residues in apoa-i at positions 86, 112, and 148, which belong to three different 22mer domains of apoa-i, we could expect more than two isomeric forms on HPLC. As shown in Fig. 1, apoa-ia and apoa-ib are doublets which may indicate that each of these peaks contain a mixture of apoa-i isomers. It is possible that the peak designated as apoa-ia is not completely oxidized and the peak apoa-ib is not completely reduced. However, it is clear from the amino acid analysis and CNBr studies that A-Ia is more oxidized than A-Ib. The HPLC may not be separating pure isomeric proteins. The reason for such a difference in HPLC retention time between the oxidized and reduced forms is not clear. It is possible that certain methionine residues are more important than others for determining the HPLC retention time. This is not surprising since each of the three methionine residues is situated at the nonpolar face of three different 22mer amphipathic helical domains which are in different environments in apoa-i. The amount of oxidized and reduced apoa-i in the conventionally isolated apoa-i may vary from preparation to preparation. It may, therefore, be difficult to get consistent results in CD studies and lipid-association properties from two different batches of apoa-i. Based on the CNBr studies (Table 3), we suggest that apoa-iic is more oxidized than apoa-iia which is more oxidized than apoa-iib. That is, apoa-iib has both methionines reduced, and apoa-iic has both methionines oxidized. If this is true, then when the dimer is cleaved, apoa-iib and apoa-iic would be reduced to monomers with a single peak on HPLC while apoa-iia would pro- Downloaded from by guest, on September 22, Journal of Lipid Research Volume 29, 1988
9 I Fig. 7. Negative stain electron micrographs of complexes formed by incubation of multilarnellar DMPC vesicles with peptide analogs. Panel A: IMetJ18A:DMPC at a 1:lweight ratio. Panel B: [Mct(O)]l8A:DMPC at a 1:l wight ratio. duce two peaks of equal size. This was what was observed reduced isomers to the oxidized forms with hydrogen (Fig. 5). Moreover, the first peak produced by apoa-iia reduction with mercaptoethanol corresponds to the peroxide. Since oxidized methionines are difficult to reduce with reducing agents, attempts to reduce the oximonomer peak produced by apoa-iic. Similarly, the second peak produced by apoa-iia corresponds to the monomer peak produced by apoa-iib. Thus, our studies indicate dized form have yielded mixtures that may be due to TABLE 3. Percent helicity as determined by CD studies of the that apoa-i1 exists as three isomers caused by differences apolipoprotein isomers and peptide analogs (12) in methionine oxidation. We have shown that hydrogen peroxide treatment of apoa-iib produces the other apoa-i1 Apolipoprotein ButTer DMPC I:I isomers (a and c). Since this protein has only a single A-Ib methionine in each of its monomers, which would thus A-Ia have identical environments, we would expect each of A-Ia' these residues to have an 35.6 equal influence on this protein's A-IIb 46.9 physical properties. This 38.5 appears to be the case as far as A-IIa 48.6 HPLC retention time is concerned. Each additional A-IIc methionine oxidation reduces apoa-i1 retention time to a [Met3]18A [Met(O)']l8A 11.2 similar degree. In summary, our data strongly suggest that methionine Protein concentrations were determined by the Coomassic Blue (Pierce oxidation is a major contributor to the alteration in the Chemical Company, Rockford, IL) method and by quantitative HPLC. HPLC retention time and lipid binding characteristics of was used in these studies. apoa-i and apoa-11. We have successfully converted the "CD could not be obtained because the solution was not clear. Fifty pglml of protein in.1 M ammonium bicarbonate buffer, ph 8., Downloaded from by guest, on September 22, 218 Ananthammaiah et al. Oxidized forms of apolipoproteins A-I and A-I1 317
10 degradation of protein. We have also synthesized analogs of the amphipathic helix containing methionine or methionine sulfoxide which behave similarly to the apolipoprotein isomers on HPLC. There are many reports in the literature that give conflicting results of apoa-i functional properties, including apoa-i self-association (4) and LCAT activation (6). We have shown that differences in the degree of methionine oxidation can alter the ability of apolipoproteins and synthetic peptides to associate with lipid. The reported differences in the properties of apoa-i may be partly due to differing amounts of apoa-ia and A-Ib. I We would like to thank Dr. Christie G. Brouillette for the excellent electron microscopy studies which we report in this manuscript. We would also like to thank Margaret A. Moore for her skilled technical assistance and Mr. Ken Beaudrie for his help in preparing this manuscript. This work was supported by National Institutes of Health Grant HL-34343, the Alabama Affiliate of the American Heart Association, and the Diabetes Trust Fund, Inc., located in the Diabetes Hospital at the University of Alabama at Birmingham. Manuscript received 23 June 1987, in revised form 26 August 1987, and in re-reuised form 16 September REFERENCES Gordon, T., W. P. Castelli, M. C. Hjortland, W. P. Kannel, and T. R. Dawber High density lipoprotein as a protective factor against coronary heart disease. Am. J. Med. 62: Glomset, J. A High density lipoproteins in human health and B disease. Adu. Internal Med. 25: Lou, C. C., W-H. Li, M. N. Moore, and L. Chan Structure and evolution of the apolipoprotein multigene family. J. Mol. Biol. 187: Osborne, J. C., Jr., and H. B. Brewer, Jr The plasma lipoproteins. Adu. Protein Chem. 31: Segrest, J. P., R. L. Jackson, D. L. Morrisett, and A. M. Gotto, J A molecular theory of lipid-protein interactions in the plasma lipoproteins. FEBS Lett. 38: Nestruck, A. C., G. Suzue, and Y. L. Marcel Studies on the polymorphism of human apolipoprotein A-I. Biochim. Biophys. Acta. 617: Miltrop, P., P. K. Weech, R. W. Milne, and Y. L. Marcel Immunochemical characterisation of apolipoprotein A-I from normal human plasma. Atherosclerosis. 6: Ghiselli, G., M. E Rohde, S. Tanenbaum, S. Krishnan, and A. M. Gotto, J Origin of apolipoprotein A-I polymorphism in plasma. J. Bioi. Chem. 26: Weech, P. K., R. W. Milne, P. Miltrop, and Y. L. Marcell Apolipoprotein A-I from normal human plasma: definition of three distinct antigenic determinants. Biochim. Biophys. Acta. 835: Ghisellini, M., M. Pecorari, and S. Calandra Changes in the main isoform of human apolipoprotein A-I following incubation of plasma. Atherosciemsis. 59: Hughes, T. A., M. A. Moore, P. Neame, M. E Medley, and B. H. Chung Rapid quantitative apolipoprotein analysis by gradient ultracentrifugation and reversed phase high performance liquid chromatography. J. Lipid Res. 29: Gross, E., and B. Witkop Nonenzymatic cleavage of peptide bonds: the methionine residues in bovine pancreatic ribonuclease. J. Biol. Chem Anantharamaiah, G. M., J. L. Jones, C. G. Brouillette, C. F. Schmidt, B. H. Chung, T. A. Hughes, A. S. Bhown, and J. P. Segrest Studies of synthetic peptide analogs of the amphipathic helix. J Bioi. Chem. 26: Brot, N., L. Weissbech, J. Werth, and H. Weissbech Enzymatic reduction of protein-bond methionine sulfoxide. Pmc. Natl. Acad. Sci. USA. 78: Brot, N., and H. Weissbach Biochemistry and physiological role of methionine sulfoxide residues in proteins. Arch. Biochem. Biophys. 223: Shechter, Y Selective oxidation and reduction of methionine residues in peptides and proteins by oxygen exchange between sulfoxide and sulfide. J Biol. Chem. 261: Anantharamaiah, G. M., A. Gawish, M. Iqbal, S. A. Khan, C. G. Brouillette, and J. P. Segrest Structurefunction of synthetic analogs of the amphipathic helix. Pmtides Biol. Fluidr. 34: Downloaded from by guest, on September 22, Journal of Lipid Research Volume 29, 1988
HPLC '88. Poster Presentation. Isolation of Thymosin B4 from Thymosin Fraction 5 by Reverse Phase HPLC
 Essentials in HPLC '88 Poster Presentation Isolation of Thymosin B4 from Thymosin Fraction 5 by Reverse Phase HPLC M. Badamchian, M.P. Strickler, M.J. Stone, A.L. Goldstein for Waters.bioresearchThe absolute,
Essentials in HPLC '88 Poster Presentation Isolation of Thymosin B4 from Thymosin Fraction 5 by Reverse Phase HPLC M. Badamchian, M.P. Strickler, M.J. Stone, A.L. Goldstein for Waters.bioresearchThe absolute,
This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is worth 2 points.
 MBB 407/511 Molecular Biology and Biochemistry First Examination - October 1, 2002 Name Social Security Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is
MBB 407/511 Molecular Biology and Biochemistry First Examination - October 1, 2002 Name Social Security Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is
Purification of Glucagon3 Interleukin-2 Fusion Protein Derived from E. coli
 Purification of Glucagon3 Interleukin-2 Fusion Protein Derived from E. coli Hye Soon Won Dept. of Chem. Eng. Chungnam National University INTRODUCTION Human interleukin-2(hil-2) - known as T Cell Growth
Purification of Glucagon3 Interleukin-2 Fusion Protein Derived from E. coli Hye Soon Won Dept. of Chem. Eng. Chungnam National University INTRODUCTION Human interleukin-2(hil-2) - known as T Cell Growth
Electronic Supplementary Information. Table of Contents
 Electronic Supplementary Information Examination of native chemical ligation using peptidyl prolyl thioester Takahiro Nakamura, Akira Shigenaga, Kohei Sato, Yusuke Tsuda, Ken Sakamoto, and Akira Otaka*
Electronic Supplementary Information Examination of native chemical ligation using peptidyl prolyl thioester Takahiro Nakamura, Akira Shigenaga, Kohei Sato, Yusuke Tsuda, Ken Sakamoto, and Akira Otaka*
Nature Methods: doi: /nmeth Supplementary Figure 1
 Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
BIOCHEMISTRY REVIEW. Overview of Biomolecules. Chapter 4 Protein Sequence
 BIOCHEMISTRY REVIEW Overview of Biomolecules Chapter 4 Protein Sequence 2 3 4 Are You Getting It?? A molecule of hemoglobin is compared with a molecule of lysozyme. Which characteristics do they share?
BIOCHEMISTRY REVIEW Overview of Biomolecules Chapter 4 Protein Sequence 2 3 4 Are You Getting It?? A molecule of hemoglobin is compared with a molecule of lysozyme. Which characteristics do they share?
Multiple-Choice Questions Answer ALL 20 multiple-choice questions on the Scantron Card in PENCIL
 Multiple-Choice Questions Answer ALL 20 multiple-choice questions on the Scantron Card in PENCIL For Questions 1-10 choose ONE INCORRECT answer. 1. Which ONE of the following statements concerning the
Multiple-Choice Questions Answer ALL 20 multiple-choice questions on the Scantron Card in PENCIL For Questions 1-10 choose ONE INCORRECT answer. 1. Which ONE of the following statements concerning the
Levels of Protein Structure:
 Levels of Protein Structure: PRIMARY STRUCTURE (1 ) - Defined, non-random sequence of amino acids along the peptide backbone o Described in two ways: Amino acid composition Amino acid sequence M-L-D-G-C-G
Levels of Protein Structure: PRIMARY STRUCTURE (1 ) - Defined, non-random sequence of amino acids along the peptide backbone o Described in two ways: Amino acid composition Amino acid sequence M-L-D-G-C-G
Mercaptoethanesulfonic acid as the reductive thiol-containing reagent employed for the derivatization of amino acids with o-phthaldialdehyde analysis
 Acta Univ. Sapientiae, Alimentaria, 1 (2008) 49 60 Mercaptoethanesulfonic acid as the reductive thiol-containing reagent employed for the derivatization of amino acids with o-phthaldialdehyde analysis
Acta Univ. Sapientiae, Alimentaria, 1 (2008) 49 60 Mercaptoethanesulfonic acid as the reductive thiol-containing reagent employed for the derivatization of amino acids with o-phthaldialdehyde analysis
Quantitative LC-MS/MS Analysis of Glucagon. Veniamin Lapko, Ph.D June 21, 2011
 Quantitative LC-MS/MS Analysis of Glucagon Veniamin Lapko, Ph.D June 21, 2011 Contents Comparison with small molecule LC-MS/MS LC-MS/MS sensitivity of peptides detection Stability: neat vs. matrix solutions
Quantitative LC-MS/MS Analysis of Glucagon Veniamin Lapko, Ph.D June 21, 2011 Contents Comparison with small molecule LC-MS/MS LC-MS/MS sensitivity of peptides detection Stability: neat vs. matrix solutions
130327SCH4U_biochem April 09, 2013
 Option B: B1.1 ENERGY Human Biochemistry If more energy is taken in from food than is used up, weight gain will follow. Similarly if more energy is used than we supply our body with, weight loss will occur.
Option B: B1.1 ENERGY Human Biochemistry If more energy is taken in from food than is used up, weight gain will follow. Similarly if more energy is used than we supply our body with, weight loss will occur.
Previous Class. Today. Detection of enzymatic intermediates: Protein tyrosine phosphatase mechanism. Protein Kinase Catalytic Properties
 Previous Class Detection of enzymatic intermediates: Protein tyrosine phosphatase mechanism Today Protein Kinase Catalytic Properties Protein Phosphorylation Phosphorylation: key protein modification
Previous Class Detection of enzymatic intermediates: Protein tyrosine phosphatase mechanism Today Protein Kinase Catalytic Properties Protein Phosphorylation Phosphorylation: key protein modification
2. Which of the following amino acids is most likely to be found on the outer surface of a properly folded protein?
 Name: WHITE Student Number: Answer the following questions on the computer scoring sheet. 1 mark each 1. Which of the following amino acids would have the highest relative mobility R f in normal thin layer
Name: WHITE Student Number: Answer the following questions on the computer scoring sheet. 1 mark each 1. Which of the following amino acids would have the highest relative mobility R f in normal thin layer
VaTx1 VaTx2 VaTx3. VaTx min Retention Time (min) Retention Time (min)
 a Absorbance (mau) 5 2 5 3 4 5 6 7 8 9 6 2 3 4 5 6 VaTx2 High Ca 2+ Low Ca 2+ b 38.2 min Absorbance (mau) 3 2 3 4 5 3 2 VaTx2 39.3 min 3 4 5 3 2 4. min 3 4 5 Supplementary Figure. Toxin Purification For
a Absorbance (mau) 5 2 5 3 4 5 6 7 8 9 6 2 3 4 5 6 VaTx2 High Ca 2+ Low Ca 2+ b 38.2 min Absorbance (mau) 3 2 3 4 5 3 2 VaTx2 39.3 min 3 4 5 3 2 4. min 3 4 5 Supplementary Figure. Toxin Purification For
SUPPLEMENTARY DATA. Materials and Methods
 SUPPLEMENTARY DATA Materials and Methods HPLC-UV of phospholipid classes and HETE isomer determination. Fractionation of platelet lipid classes was undertaken on a Spherisorb S5W 150 x 4.6 mm column (Waters
SUPPLEMENTARY DATA Materials and Methods HPLC-UV of phospholipid classes and HETE isomer determination. Fractionation of platelet lipid classes was undertaken on a Spherisorb S5W 150 x 4.6 mm column (Waters
Depleting Lipoproteins from Serum
 Depleting Lipoproteins from Serum Kathy K. Foxx Kalen Biomedical LLC President For decades, fetal bovine serum (FBS) has been used as a supplement for cell-culture media, providing the growth factors that
Depleting Lipoproteins from Serum Kathy K. Foxx Kalen Biomedical LLC President For decades, fetal bovine serum (FBS) has been used as a supplement for cell-culture media, providing the growth factors that
Antoine Bouchoux, Pierre-Emerson Cayemitte, Julien Jardin, Geneviève Gésan-Guiziou, and Bernard Cabane
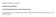 Biophysical Journal, Volume 96 Supplementary Material Casein Micelle Dispersions under Osmotic Stress Antoine Bouchoux, Pierre-Emerson Cayemitte, Julien Jardin, Geneviève Gésan-Guiziou, and Bernard Cabane
Biophysical Journal, Volume 96 Supplementary Material Casein Micelle Dispersions under Osmotic Stress Antoine Bouchoux, Pierre-Emerson Cayemitte, Julien Jardin, Geneviève Gésan-Guiziou, and Bernard Cabane
Molecular Biology. general transfer: occurs normally in cells. special transfer: occurs only in the laboratory in specific conditions.
 Chapter 9: Proteins Molecular Biology replication general transfer: occurs normally in cells transcription special transfer: occurs only in the laboratory in specific conditions translation unknown transfer:
Chapter 9: Proteins Molecular Biology replication general transfer: occurs normally in cells transcription special transfer: occurs only in the laboratory in specific conditions translation unknown transfer:
Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular
 Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular weight markers (M). Supplementary Figure-2. Overlay of
Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular weight markers (M). Supplementary Figure-2. Overlay of
Biomolecules: amino acids
 Biomolecules: amino acids Amino acids Amino acids are the building blocks of proteins They are also part of hormones, neurotransmitters and metabolic intermediates There are 20 different amino acids in
Biomolecules: amino acids Amino acids Amino acids are the building blocks of proteins They are also part of hormones, neurotransmitters and metabolic intermediates There are 20 different amino acids in
AA s are the building blocks of proteins
 Chamras Chemistry 106 Lecture otes Chapter 24: Amino Acids, Peptides, and Proteins General Formula: () n (') α-amino Acids: (n = 1) Example: Amino Acids and Proteins: Glycine Alanine Valine AA s are the
Chamras Chemistry 106 Lecture otes Chapter 24: Amino Acids, Peptides, and Proteins General Formula: () n (') α-amino Acids: (n = 1) Example: Amino Acids and Proteins: Glycine Alanine Valine AA s are the
Supporting Information
 Supporting Information Cyclic Peptidyl Inhibitors against Human Peptidyl-Prolyl Isomerase Pin1 Tao Liu, Yu Liu, Hung-Ying Kao, *,, and Dehua Pei Table of Contents: Table S1. Structures of amino acid building
Supporting Information Cyclic Peptidyl Inhibitors against Human Peptidyl-Prolyl Isomerase Pin1 Tao Liu, Yu Liu, Hung-Ying Kao, *,, and Dehua Pei Table of Contents: Table S1. Structures of amino acid building
BCHS 3304/ Exam I. September 26,
 Name: 5.5.# BCHS 3304/ Exam I September 26, 2002............... Instructions: 1. There are 13 pages to this exam. Count pages prior to beginning exam. You may use the back pages of the exam as scratch
Name: 5.5.# BCHS 3304/ Exam I September 26, 2002............... Instructions: 1. There are 13 pages to this exam. Count pages prior to beginning exam. You may use the back pages of the exam as scratch
Chapter 3. Structure of Enzymes. Enzyme Engineering
 Chapter 3. Structure of Enzymes Enzyme Engineering 3.1 Introduction With purified protein, Determining M r of the protein Determining composition of amino acids and the primary structure Determining the
Chapter 3. Structure of Enzymes Enzyme Engineering 3.1 Introduction With purified protein, Determining M r of the protein Determining composition of amino acids and the primary structure Determining the
Chapter 20 and GHW#10 Questions. Proteins
 Chapter 20 and GHW#10 Questions Proteins Proteins Naturally occurring bioorganic polyamide polymers containing a sequence of various combinations of 20 amino acids. Amino acids contain the elements carbon,
Chapter 20 and GHW#10 Questions Proteins Proteins Naturally occurring bioorganic polyamide polymers containing a sequence of various combinations of 20 amino acids. Amino acids contain the elements carbon,
BabyBio IMAC columns DATA SHEET DS
 BabyBio IMAC columns DATA SHEET DS 45 655 010 BabyBio columns for Immobilized Metal Ion Affinity Chromatography (IMAC) are ready-to-use for quick and easy purification of polyhistidine-tagged (His-tagged)
BabyBio IMAC columns DATA SHEET DS 45 655 010 BabyBio columns for Immobilized Metal Ion Affinity Chromatography (IMAC) are ready-to-use for quick and easy purification of polyhistidine-tagged (His-tagged)
Objective: You will be able to explain how the subcomponents of
 Objective: You will be able to explain how the subcomponents of nucleic acids determine the properties of that polymer. Do Now: Read the first two paragraphs from enduring understanding 4.A Essential knowledge:
Objective: You will be able to explain how the subcomponents of nucleic acids determine the properties of that polymer. Do Now: Read the first two paragraphs from enduring understanding 4.A Essential knowledge:
BIO 311C Spring Lecture 15 Friday 26 Feb. 1
 BIO 311C Spring 2010 Lecture 15 Friday 26 Feb. 1 Illustration of a Polypeptide amino acids peptide bonds Review Polypeptide (chain) See textbook, Fig 5.21, p. 82 for a more clear illustration Folding and
BIO 311C Spring 2010 Lecture 15 Friday 26 Feb. 1 Illustration of a Polypeptide amino acids peptide bonds Review Polypeptide (chain) See textbook, Fig 5.21, p. 82 for a more clear illustration Folding and
CS612 - Algorithms in Bioinformatics
 Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
BCH Graduate Survey of Biochemistry
 BCH 5045 Graduate Survey of Biochemistry Instructor: Charles Guy Producer: Ron Thomas Director: Glen Graham Lecture 7 Slide sets available at: http://hort.ifas.ufl.edu/teach/guyweb/bch5045/index.html David
BCH 5045 Graduate Survey of Biochemistry Instructor: Charles Guy Producer: Ron Thomas Director: Glen Graham Lecture 7 Slide sets available at: http://hort.ifas.ufl.edu/teach/guyweb/bch5045/index.html David
note on methodology I
 note on methodology I isolated per tube, and the preparation is very dilute and needs to be concentrated. We present here some modifications to this method in order to prepare large volumes of concentrated
note on methodology I isolated per tube, and the preparation is very dilute and needs to be concentrated. We present here some modifications to this method in order to prepare large volumes of concentrated
Trypsin Mass Spectrometry Grade
 058PR-03 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Trypsin Mass Spectrometry Grade A Chemically Modified, TPCK treated, Affinity Purified
058PR-03 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Trypsin Mass Spectrometry Grade A Chemically Modified, TPCK treated, Affinity Purified
CHAPTER 21: Amino Acids, Proteins, & Enzymes. General, Organic, & Biological Chemistry Janice Gorzynski Smith
 CHAPTER 21: Amino Acids, Proteins, & Enzymes General, Organic, & Biological Chemistry Janice Gorzynski Smith CHAPTER 21: Amino Acids, Proteins, Enzymes Learning Objectives: q The 20 common, naturally occurring
CHAPTER 21: Amino Acids, Proteins, & Enzymes General, Organic, & Biological Chemistry Janice Gorzynski Smith CHAPTER 21: Amino Acids, Proteins, Enzymes Learning Objectives: q The 20 common, naturally occurring
PAPER No. : 16, Bioorganic and biophysical chemistry MODULE No. : 22, Mechanism of enzyme catalyst reaction (I) Chymotrypsin
 Subject Paper No and Title 16 Bio-organic and Biophysical Module No and Title 22 Mechanism of Enzyme Catalyzed reactions I Module Tag CHE_P16_M22 Chymotrypsin TABLE OF CONTENTS 1. Learning outcomes 2.
Subject Paper No and Title 16 Bio-organic and Biophysical Module No and Title 22 Mechanism of Enzyme Catalyzed reactions I Module Tag CHE_P16_M22 Chymotrypsin TABLE OF CONTENTS 1. Learning outcomes 2.
Tenofovir disoproxil fumarate (Tenofoviri disoproxili fumaras)
 C 19 H 30 N 5 O 10 P. C 4 H 4 O 4 Relative molecular mass. 635.5. Chemical names. bis(1-methylethyl) 5-{[(1R)-2-(6-amino-9H-purin-9-yl)-1-methylethoxy]methyl}-5-oxo-2,4,6,8-tetraoxa-5-λ 5 - phosphanonanedioate
C 19 H 30 N 5 O 10 P. C 4 H 4 O 4 Relative molecular mass. 635.5. Chemical names. bis(1-methylethyl) 5-{[(1R)-2-(6-amino-9H-purin-9-yl)-1-methylethoxy]methyl}-5-oxo-2,4,6,8-tetraoxa-5-λ 5 - phosphanonanedioate
Toxins 2016, 8, 222; doi: /toxins
 S1 of S8 Supplementary Materials: Mapping Protein Protein Interactions of the Resistance Related Bacterial Zeta Toxin Epsilon Antitoxin Complex (ε2ζ2) with High Affinity Peptide Ligands Using Fluorescence
S1 of S8 Supplementary Materials: Mapping Protein Protein Interactions of the Resistance Related Bacterial Zeta Toxin Epsilon Antitoxin Complex (ε2ζ2) with High Affinity Peptide Ligands Using Fluorescence
Analysis of L- and D-Amino Acids Using UPLC Yuta Mutaguchi 1 and Toshihisa Ohshima 2*
 Analysis of L- and D-Amino Acids Using UPLC Yuta Mutaguchi 1 and Toshihisa Ohshima 2* 1 Department of Biotechnology, Akita Prefectural University, Akita City, Japan; 2 Department of Biomedical Engineering,
Analysis of L- and D-Amino Acids Using UPLC Yuta Mutaguchi 1 and Toshihisa Ohshima 2* 1 Department of Biotechnology, Akita Prefectural University, Akita City, Japan; 2 Department of Biomedical Engineering,
Computer programs to identify and classify amphipathic a helical domains
 Computer programs to identify and classify amphipathic a helical domains Martin K Jones, G M Anantharamaiah, and Jere P Segrest Atherosclerosis Research Unit, Departments of Medicine and Biochemisq, 630
Computer programs to identify and classify amphipathic a helical domains Martin K Jones, G M Anantharamaiah, and Jere P Segrest Atherosclerosis Research Unit, Departments of Medicine and Biochemisq, 630
The Different Folding Behavior of Insulin and Insulin-like Growth Factor 1 Is Mainly Controlled by Their B-Chain/Domain
 1556 Biochemistry 2002, 41, 1556-1567 The Different Folding Behavior of Insulin and Insulin-like Growth Factor 1 Is Mainly Controlled by Their B-Chain/Domain Zhan-Yun Guo, Lu Shen, and You-Min Feng* State
1556 Biochemistry 2002, 41, 1556-1567 The Different Folding Behavior of Insulin and Insulin-like Growth Factor 1 Is Mainly Controlled by Their B-Chain/Domain Zhan-Yun Guo, Lu Shen, and You-Min Feng* State
Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels. Supplementary Information
 Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels Craig T. Armstrong, Philip E. Mason, J. L. Ross Anderson and Christopher E. Dempsey
Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels Craig T. Armstrong, Philip E. Mason, J. L. Ross Anderson and Christopher E. Dempsey
Edman Sample Preparation Part 2. Bill Henzel Genentech Inc.
 Edman Sample Preparation Part 2 Bill Henzel Genentech Inc. Determining a Protein is Blocked Estimate the amount of protein applied to the protein sequencer. Compare the staining intensity of the band of
Edman Sample Preparation Part 2 Bill Henzel Genentech Inc. Determining a Protein is Blocked Estimate the amount of protein applied to the protein sequencer. Compare the staining intensity of the band of
Isolation of five carotenoid compounds from tangerine tomatoes
 Isolation of five carotenoid compounds from tangerine tomatoes Thesis Thomas Haufe Advisor: Steven J. Schwartz, Ph.D Department of Food Science and Technology Presented in fulfillment of the requirements
Isolation of five carotenoid compounds from tangerine tomatoes Thesis Thomas Haufe Advisor: Steven J. Schwartz, Ph.D Department of Food Science and Technology Presented in fulfillment of the requirements
PROTEINS. Building blocks, structure and function. Aim: You will have a clear picture of protein construction and their general properties
 PROTEINS Building blocks, structure and function Aim: You will have a clear picture of protein construction and their general properties Reading materials: Compendium in Biochemistry, page 13-49. Microbiology,
PROTEINS Building blocks, structure and function Aim: You will have a clear picture of protein construction and their general properties Reading materials: Compendium in Biochemistry, page 13-49. Microbiology,
Biological systems interact, and these systems and their interactions possess complex properties. STOP at enduring understanding 4A
 Biological systems interact, and these systems and their interactions possess complex properties. STOP at enduring understanding 4A Homework Watch the Bozeman video called, Biological Molecules Objective:
Biological systems interact, and these systems and their interactions possess complex properties. STOP at enduring understanding 4A Homework Watch the Bozeman video called, Biological Molecules Objective:
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/2/4/e1500980/dc1 Supplementary Materials for The crystal structure of human dopamine -hydroxylase at 2.9 Å resolution Trine V. Vendelboe, Pernille Harris, Yuguang
advances.sciencemag.org/cgi/content/full/2/4/e1500980/dc1 Supplementary Materials for The crystal structure of human dopamine -hydroxylase at 2.9 Å resolution Trine V. Vendelboe, Pernille Harris, Yuguang
Properties of amino acids in proteins
 Properties of amino acids in proteins one of the primary roles of DNA (but far from the only one!!!) is to code for proteins A typical bacterium builds thousands types of proteins, all from ~20 amino acids
Properties of amino acids in proteins one of the primary roles of DNA (but far from the only one!!!) is to code for proteins A typical bacterium builds thousands types of proteins, all from ~20 amino acids
Structural Characterization of Prion-like Conformational Changes of the Neuronal Isoform of Aplysia CPEB
 Structural Characterization of Prion-like Conformational Changes of the Neuronal Isoform of Aplysia CPEB Bindu L. Raveendra, 1,5 Ansgar B. Siemer, 2,6 Sathyanarayanan V. Puthanveettil, 1,3,7 Wayne A. Hendrickson,
Structural Characterization of Prion-like Conformational Changes of the Neuronal Isoform of Aplysia CPEB Bindu L. Raveendra, 1,5 Ansgar B. Siemer, 2,6 Sathyanarayanan V. Puthanveettil, 1,3,7 Wayne A. Hendrickson,
Chemistry 121 Winter 17
 Chemistry 121 Winter 17 Introduction to Organic Chemistry and Biochemistry Instructor Dr. Upali Siriwardane (Ph.D. Ohio State) E-mail: upali@latech.edu Office: 311 Carson Taylor Hall ; Phone: 318-257-4941;
Chemistry 121 Winter 17 Introduction to Organic Chemistry and Biochemistry Instructor Dr. Upali Siriwardane (Ph.D. Ohio State) E-mail: upali@latech.edu Office: 311 Carson Taylor Hall ; Phone: 318-257-4941;
Page 1 of 5 Biochemistry I Fall 2017 Practice for Exam2 Dr. Stone Name
 Page 1 of 5 Biochemistry I Fall 2017 Practice for Exam2 Dr. Stone ame o answers will be provided. ere are some constants and equations that may be useful: K a = [+][A-]/[A] p = pka + log [A-]/[A] K a for
Page 1 of 5 Biochemistry I Fall 2017 Practice for Exam2 Dr. Stone ame o answers will be provided. ere are some constants and equations that may be useful: K a = [+][A-]/[A] p = pka + log [A-]/[A] K a for
1-To know what is protein 2-To identify Types of protein 3- To Know amino acids 4- To be differentiate between essential and nonessential amino acids
 Amino acids 1-To know what is protein 2-To identify Types of protein 3- To Know amino acids 4- To be differentiate between essential and nonessential amino acids 5-To understand amino acids synthesis Amino
Amino acids 1-To know what is protein 2-To identify Types of protein 3- To Know amino acids 4- To be differentiate between essential and nonessential amino acids 5-To understand amino acids synthesis Amino
The Structure and Function of Macromolecules
 The Structure and Function of Macromolecules Macromolecules are polymers Polymer long molecule consisting of many similar building blocks. Monomer the small building block molecules. Carbohydrates, proteins
The Structure and Function of Macromolecules Macromolecules are polymers Polymer long molecule consisting of many similar building blocks. Monomer the small building block molecules. Carbohydrates, proteins
Moorpark College Chemistry 11 Fall Instructor: Professor Gopal. Examination # 5: Section Five May 7, Name: (print)
 Moorpark College Chemistry 11 Fall 2013 Instructor: Professor Gopal Examination # 5: Section Five May 7, 2013 Name: (print) Directions: Make sure your examination contains TEN total pages (including this
Moorpark College Chemistry 11 Fall 2013 Instructor: Professor Gopal Examination # 5: Section Five May 7, 2013 Name: (print) Directions: Make sure your examination contains TEN total pages (including this
RITONAVIRI COMPRESSI RITONAVIR TABLETS. Final text for addition to The International Pharmacopoeia (July 2012)
 July 2012 RITONAVIRI COMPRESSI RITONAVIR TABLETS Final text for addition to The International Pharmacopoeia (July 2012) This monograph was adopted at the Forty-sixth WHO Expert Committee on Specifications
July 2012 RITONAVIRI COMPRESSI RITONAVIR TABLETS Final text for addition to The International Pharmacopoeia (July 2012) This monograph was adopted at the Forty-sixth WHO Expert Committee on Specifications
Chapter 3. Protein Structure and Function
 Chapter 3 Protein Structure and Function Broad functional classes So Proteins have structure and function... Fine! -Why do we care to know more???? Understanding functional architechture gives us POWER
Chapter 3 Protein Structure and Function Broad functional classes So Proteins have structure and function... Fine! -Why do we care to know more???? Understanding functional architechture gives us POWER
AccuMAP Low ph Protein Digestion Kits
 TECHNICAL MANUAL AccuMAP Low ph Protein Digestion Kits Instruc ons for Use of Products VA1040 and VA1050 5/17 TM504 AccuMAP Low ph Protein Digestion Kits All technical literature is available at: www.promega.com/protocols/
TECHNICAL MANUAL AccuMAP Low ph Protein Digestion Kits Instruc ons for Use of Products VA1040 and VA1050 5/17 TM504 AccuMAP Low ph Protein Digestion Kits All technical literature is available at: www.promega.com/protocols/
Relative Measurement of Zeaxanthin Stereoisomers by Chiral HPLC
 Relative Measurement of Zeaxanthin Stereoisomers by Chiral HPLC Principle To measure the relative percentages of the (3R,3 R), (3R,3 S) and (3S,3 S) stereoisomers of zeaxanthin in dietary ingredient and
Relative Measurement of Zeaxanthin Stereoisomers by Chiral HPLC Principle To measure the relative percentages of the (3R,3 R), (3R,3 S) and (3S,3 S) stereoisomers of zeaxanthin in dietary ingredient and
paper and beads don t fall off. Then, place the beads in the following order on the pipe cleaner:
 Beady Pipe Cleaner Proteins Background: Proteins are the molecules that carry out most of the cell s dayto-day functions. While the DNA in the nucleus is "the boss" and controls the activities of the cell,
Beady Pipe Cleaner Proteins Background: Proteins are the molecules that carry out most of the cell s dayto-day functions. While the DNA in the nucleus is "the boss" and controls the activities of the cell,
TENOFOVIR TABLETS: Final text for addition to The International Pharmacopoeia (June 2010)
 June 2010 TENOFOVIR TABLETS: Final text for addition to The International Pharmacopoeia (June 2010) This monograph was adopted at the Forty-fourth WHO Expert Committee on Specifications for Pharmaceutical
June 2010 TENOFOVIR TABLETS: Final text for addition to The International Pharmacopoeia (June 2010) This monograph was adopted at the Forty-fourth WHO Expert Committee on Specifications for Pharmaceutical
Gentilucci, Amino Acids, Peptides, and Proteins. Peptides and proteins are polymers of amino acids linked together by amide bonds CH 3
 Amino Acids Peptides and proteins are polymers of amino acids linked together by amide bonds Aliphatic Side-Chain Amino Acids - - H CH glycine alanine 3 proline valine CH CH 3 - leucine - isoleucine CH
Amino Acids Peptides and proteins are polymers of amino acids linked together by amide bonds Aliphatic Side-Chain Amino Acids - - H CH glycine alanine 3 proline valine CH CH 3 - leucine - isoleucine CH
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/2/1/e1500678/dc1 Supplementary Materials for Chemical synthesis of erythropoietin glycoforms for insights into the relationship between glycosylation pattern and
advances.sciencemag.org/cgi/content/full/2/1/e1500678/dc1 Supplementary Materials for Chemical synthesis of erythropoietin glycoforms for insights into the relationship between glycosylation pattern and
CHM333 LECTURE 9: 9/16/09 FALL 2009 Professor Christine Hrycyna
 PROTEIN STRUCTURE I Each protein has a characteristic shape, size and function: Classification of Proteins on the Basis of Biological Role: 1. Structural Proteins a. Provide mechanical support to cells
PROTEIN STRUCTURE I Each protein has a characteristic shape, size and function: Classification of Proteins on the Basis of Biological Role: 1. Structural Proteins a. Provide mechanical support to cells
Supporting Information
 Electronic Supplementary Material (ESI) for Organic & Biomolecular Chemistry. This journal is The Royal Society of Chemistry 2014 Supporting Information 1. General Information...S2 a. Materials b. HPLC
Electronic Supplementary Material (ESI) for Organic & Biomolecular Chemistry. This journal is The Royal Society of Chemistry 2014 Supporting Information 1. General Information...S2 a. Materials b. HPLC
Development of a Cell-penetrating Peptide that Exhibits Responsive. Changes in its Secondary Structure in the Cellular Environment
 Development of a Cell-penetrating Peptide that Exhibits Responsive Changes in its Secondary Structure in the Cellular Environment iroko Yamashita, 1 Takuma Kato, 2 Makoto ba, 2 Takashi Misawa, 1 Takayuki
Development of a Cell-penetrating Peptide that Exhibits Responsive Changes in its Secondary Structure in the Cellular Environment iroko Yamashita, 1 Takuma Kato, 2 Makoto ba, 2 Takashi Misawa, 1 Takayuki
Turnover of Synthetic Class A Amphipathic Peptide Analogues of Exchangeable Apolipoproteins in Rats. Correlation With Physical Properties
 886 Turnover of Synthetic Class A Amphipathic Peptide Analogues of Exchangeable Apolipoproteins in Rats Correlation With Physical Properties David W. Garber, Y.V. Venkatachalapathi, Kiran B. Gupta, Jamal
886 Turnover of Synthetic Class A Amphipathic Peptide Analogues of Exchangeable Apolipoproteins in Rats Correlation With Physical Properties David W. Garber, Y.V. Venkatachalapathi, Kiran B. Gupta, Jamal
Human Biochemistry Option B
 Human Biochemistry Option B A look ahead... Your body has many functions to perform every day: Structural support, genetic information, communication, energy supply, metabolism Right now, thousands of
Human Biochemistry Option B A look ahead... Your body has many functions to perform every day: Structural support, genetic information, communication, energy supply, metabolism Right now, thousands of
v o = V max [S] rate = kt[s] e V max = k cat E t ΔG = -RT lnk eq K m + [S]
![v o = V max [S] rate = kt[s] e V max = k cat E t ΔG = -RT lnk eq K m + [S] v o = V max [S] rate = kt[s] e V max = k cat E t ΔG = -RT lnk eq K m + [S]](/thumbs/81/84404059.jpg) Exam 3 Spring 2017 Dr. Stone 8:00 Name There are 100 possible points on this exam. -ΔG / RT v o = V max [S] rate = kt[s] e V max = k cat E t ΔG = -RT lnk eq K m + [S] h rate forward = k forward [reactants]
Exam 3 Spring 2017 Dr. Stone 8:00 Name There are 100 possible points on this exam. -ΔG / RT v o = V max [S] rate = kt[s] e V max = k cat E t ΔG = -RT lnk eq K m + [S] h rate forward = k forward [reactants]
Proteins. Amino acids, structure and function. The Nobel Prize in Chemistry 2012 Robert J. Lefkowitz Brian K. Kobilka
 Proteins Amino acids, structure and function The Nobel Prize in Chemistry 2012 Robert J. Lefkowitz Brian K. Kobilka O O HO N N HN OH Ser65-Tyr66-Gly67 The Nobel prize in chemistry 2008 Osamu Shimomura,
Proteins Amino acids, structure and function The Nobel Prize in Chemistry 2012 Robert J. Lefkowitz Brian K. Kobilka O O HO N N HN OH Ser65-Tyr66-Gly67 The Nobel prize in chemistry 2008 Osamu Shimomura,
Isolation, Purification and Molecular Weight Determination of Antihypertensive Peptides Derived from
 2011, Vol. 32, No. 02 213 1,2 1 1, * 1 1 (1. 361012 2. 350002) AS.1398 (Porphyra haitanesis)ace Sephadex G-15 (RP-HPLC) (MALDI-TOF-MS) 2000D ACE IC50 0.73mg/mL Sephadex G-15 6 E IC50 0.67mg/mL E RP-HPLC
2011, Vol. 32, No. 02 213 1,2 1 1, * 1 1 (1. 361012 2. 350002) AS.1398 (Porphyra haitanesis)ace Sephadex G-15 (RP-HPLC) (MALDI-TOF-MS) 2000D ACE IC50 0.73mg/mL Sephadex G-15 6 E IC50 0.67mg/mL E RP-HPLC
Tivadar Orban, Beata Jastrzebska, Sayan Gupta, Benlian Wang, Masaru Miyagi, Mark R. Chance, and Krzysztof Palczewski
 Structure, Volume Supplemental Information Conformational Dynamics of Activation for the Pentameric Complex of Dimeric G Protein-Coupled Receptor and Heterotrimeric G Protein Tivadar Orban, Beata Jastrzebska,
Structure, Volume Supplemental Information Conformational Dynamics of Activation for the Pentameric Complex of Dimeric G Protein-Coupled Receptor and Heterotrimeric G Protein Tivadar Orban, Beata Jastrzebska,
Chemical Biology of Tea Catechins
 Workshop Argentina-Japan Bioscience and Biotechnology for the Promotion of Agriculture and Food Production August 4 th 2009 Chemical Biology of Tea Catechins Tsutomu NAKAYAMA Laboratory of Molecular Food
Workshop Argentina-Japan Bioscience and Biotechnology for the Promotion of Agriculture and Food Production August 4 th 2009 Chemical Biology of Tea Catechins Tsutomu NAKAYAMA Laboratory of Molecular Food
SYNOPSIS STUDIES ON THE PREPARATION AND CHARACTERISATION OF PROTEIN HYDROLYSATES FROM GROUNDNUT AND SOYBEAN ISOLATES
 1 SYNOPSIS STUDIES ON THE PREPARATION AND CHARACTERISATION OF PROTEIN HYDROLYSATES FROM GROUNDNUT AND SOYBEAN ISOLATES Proteins are important in food processing and food product development, as they are
1 SYNOPSIS STUDIES ON THE PREPARATION AND CHARACTERISATION OF PROTEIN HYDROLYSATES FROM GROUNDNUT AND SOYBEAN ISOLATES Proteins are important in food processing and food product development, as they are
Cahn - Ingold - Prelog system. Proteins: Evolution, and Analysis Lecture 7 9/15/2009. The Fischer Convention (1) G (2) (3)
 Chapter 4 (1) G Proteins: Evolution, and Analysis Lecture 7 9/15/2009 A V L I M P F W Chapter 4 (2) S (3) T N Q Y C K R H D E The Fischer Convention Absolute configuration about an asymmetric carbon related
Chapter 4 (1) G Proteins: Evolution, and Analysis Lecture 7 9/15/2009 A V L I M P F W Chapter 4 (2) S (3) T N Q Y C K R H D E The Fischer Convention Absolute configuration about an asymmetric carbon related
Wheat Amino acids & Peptides for Hair Care. INCI Name EU/USA CAS # EINECS # Hydrolyzed wheat protein
 KELYAMIN Wheat Amino acids & Peptides for Hair Care Identification INCI Name EU/USA CAS # EINECS # Hydrolyzed wheat protein 70084-87-6 305-225-0 Composition % Liquid Powder Aqua Hydrolyzed wheat protein
KELYAMIN Wheat Amino acids & Peptides for Hair Care Identification INCI Name EU/USA CAS # EINECS # Hydrolyzed wheat protein 70084-87-6 305-225-0 Composition % Liquid Powder Aqua Hydrolyzed wheat protein
Separation of Macrocyclic Lactones (Avermectins) on FLARE C18 MM & FLARE C18+ Columns
 Separation of Macrocyclic Lactones (Avermectins) on FLARE C8 MM & FLARE C8+ Columns Introduction Diamond Analytics Technical Note: T05- Avermectins are a series of 6-membered macrocyclic lactone derivatives
Separation of Macrocyclic Lactones (Avermectins) on FLARE C8 MM & FLARE C8+ Columns Introduction Diamond Analytics Technical Note: T05- Avermectins are a series of 6-membered macrocyclic lactone derivatives
Four melanocyte-stimulating hormones have the following amino acid sequences:
 Assignment 14: Melanocyte-stimulating hormone belongs to a group called the melanocortins. This group includes ACTH, alpha-msh, beta-msh and gamma-msh; these peptides are all cleavage products of a large
Assignment 14: Melanocyte-stimulating hormone belongs to a group called the melanocortins. This group includes ACTH, alpha-msh, beta-msh and gamma-msh; these peptides are all cleavage products of a large
Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random
 S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
Amino Acids. Review I: Protein Structure. Amino Acids: Structures. Amino Acids (contd.) Rajan Munshi
 Review I: Protein Structure Rajan Munshi BBSI @ Pitt 2005 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2005 Amino Acids Building blocks of proteins 20 amino acids
Review I: Protein Structure Rajan Munshi BBSI @ Pitt 2005 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2005 Amino Acids Building blocks of proteins 20 amino acids
DETERMINATION OF D-AMINO ACIDS. I. HYDROLYSIS OF DNP-L- AMINO ACID METHYL ESTERS WITH CARBOXYPEPTIDASE-Y
 Carlsberg Res. Commun. Vol. 49, p. 585-590, 1984 DETERMINATION OF D-AMINO ACIDS. I. HYDROLYSIS OF DNP-L- AMINO ACID METHYL ESTERS WITH CARBOXYPEPTIDASE-Y by PETER MARFEY" and MARTIN OTTESEN Department
Carlsberg Res. Commun. Vol. 49, p. 585-590, 1984 DETERMINATION OF D-AMINO ACIDS. I. HYDROLYSIS OF DNP-L- AMINO ACID METHYL ESTERS WITH CARBOXYPEPTIDASE-Y by PETER MARFEY" and MARTIN OTTESEN Department
Lipids: diverse group of hydrophobic molecules
 Lipids: diverse group of hydrophobic molecules Lipids only macromolecules that do not form polymers li3le or no affinity for water hydrophobic consist mostly of hydrocarbons nonpolar covalent bonds fats
Lipids: diverse group of hydrophobic molecules Lipids only macromolecules that do not form polymers li3le or no affinity for water hydrophobic consist mostly of hydrocarbons nonpolar covalent bonds fats
Thiol-Activated gem-dithiols: A New Class of Controllable. Hydrogen Sulfide (H 2 S) Donors
 Thiol-Activated gem-dithiols: A New Class of Controllable Hydrogen Sulfide (H 2 S) Donors Yu Zhao, Jianming Kang, Chung-Min Park, Powell E. Bagdon, Bo Peng, and Ming Xian * Department of Chemistry, Washington
Thiol-Activated gem-dithiols: A New Class of Controllable Hydrogen Sulfide (H 2 S) Donors Yu Zhao, Jianming Kang, Chung-Min Park, Powell E. Bagdon, Bo Peng, and Ming Xian * Department of Chemistry, Washington
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/NCHEM.2419 Diversification of Self-Replicating Molecules Jan W. Sadownik, Elio Mattia, Piotr Nowak, Sijbren Otto* University of Groningen, Center for Systems Chemistry, Stratingh Institute
DOI: 10.1038/NCHEM.2419 Diversification of Self-Replicating Molecules Jan W. Sadownik, Elio Mattia, Piotr Nowak, Sijbren Otto* University of Groningen, Center for Systems Chemistry, Stratingh Institute
Complete Amino Acid Sequence of the Protease Inhibitor BWI-4a from Buckwheat Seeds
 Biochemistry (Moscow), Vol. 65, No. 10, 2000, pp. 1140-1144. Translated from Biokhimiya, Vol. 65, No. 10, 2000, pp. 1347-1352. Original Russian Text Copyright 2000 by Belozersky, Dunaevsky, Musolyamov,
Biochemistry (Moscow), Vol. 65, No. 10, 2000, pp. 1140-1144. Translated from Biokhimiya, Vol. 65, No. 10, 2000, pp. 1347-1352. Original Russian Text Copyright 2000 by Belozersky, Dunaevsky, Musolyamov,
PNGase F Instruction Manual
 PNGase F Instruction Manual Catalog Number 170-6883 Bio-Rad Laboratories, 2000 Alfred Nobel Dr., Hercules, CA 94547 4006094 Rev A Table of Contents Section 1 Introduction...1 Section 2 Kit Components and
PNGase F Instruction Manual Catalog Number 170-6883 Bio-Rad Laboratories, 2000 Alfred Nobel Dr., Hercules, CA 94547 4006094 Rev A Table of Contents Section 1 Introduction...1 Section 2 Kit Components and
A Molecular Model of High Density Lipoproteins
 Proc. Nat. Acad. Sci. USA Vol. 71, No. 4, pp. 1534-1538, April 1974 A Molecular Model of High Density Lipoproteins (assembly of proteins and lipids/amphipathic a-helical regions/hydrophobicity of apoproteins)
Proc. Nat. Acad. Sci. USA Vol. 71, No. 4, pp. 1534-1538, April 1974 A Molecular Model of High Density Lipoproteins (assembly of proteins and lipids/amphipathic a-helical regions/hydrophobicity of apoproteins)
Hydrolysis of porcine b-casein by bovine plasmin and bovine chymosin
 Z Lebensm Unters Forsch A (1999) 208 : 83 89 Q Springer-Verlag 1999 ORIGINAL PAPER Daniel P. Gallagher 7 Tanoj K. Singh Daniel M. Mulvihill Hydrolysis of porcine b-casein by bovine plasmin and bovine chymosin
Z Lebensm Unters Forsch A (1999) 208 : 83 89 Q Springer-Verlag 1999 ORIGINAL PAPER Daniel P. Gallagher 7 Tanoj K. Singh Daniel M. Mulvihill Hydrolysis of porcine b-casein by bovine plasmin and bovine chymosin
Midterm 1 Last, First
 Midterm 1 BIS 105 Prof. T. Murphy April 23, 2014 There should be 6 pages in this exam. Exam instructions (1) Please write your name on the top of every page of the exam (2) Show all work for full credit
Midterm 1 BIS 105 Prof. T. Murphy April 23, 2014 There should be 6 pages in this exam. Exam instructions (1) Please write your name on the top of every page of the exam (2) Show all work for full credit
BIO th September, 1997
 BIO 451 26th September, 1997 EXAM I This exam will be taken apart for grading. Please PRINT your name on each page. If you do not have sufficient room for your answer in the space provided, please continue
BIO 451 26th September, 1997 EXAM I This exam will be taken apart for grading. Please PRINT your name on each page. If you do not have sufficient room for your answer in the space provided, please continue
Lipoproteins Metabolism
 Lipoproteins Metabolism LEARNING OBJECTIVES By the end of this Lecture, the student should be able to describe: What are Lipoproteins? Describe Lipoprotein Particles. Composition of Lipoproteins. The chemical
Lipoproteins Metabolism LEARNING OBJECTIVES By the end of this Lecture, the student should be able to describe: What are Lipoproteins? Describe Lipoprotein Particles. Composition of Lipoproteins. The chemical
(65 pts.) 27. (10 pts.) 28. (15 pts.) 29. (10 pts.) TOTAL (100 points) Moorpark College Chemistry 11 Spring Instructor: Professor Gopal
 Moorpark College Chemistry 11 Spring 2012 Instructor: Professor Gopal Examination # 5: Section Five May 1, 2012 Name: (print) GOOD LUCK! Directions: Make sure your examination contains TWELVE total pages
Moorpark College Chemistry 11 Spring 2012 Instructor: Professor Gopal Examination # 5: Section Five May 1, 2012 Name: (print) GOOD LUCK! Directions: Make sure your examination contains TWELVE total pages
Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.
 Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.5 % SDS-PAGE gel under reducing and non-reducing conditions.
Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.5 % SDS-PAGE gel under reducing and non-reducing conditions.
CHAPTER-6 IDENTIFICATION, AND CHARACTERISATION OF DEGRADATION IMPURITY IN VALSARTAN TABLETS
 129 CHAPTER-6 IDENTIFICATION, AND CHARACTERISATION OF DEGRADATION IMPURITY IN VALSARTAN TABLETS 130 6.1. Introduction Valsartan is an orally active specific angiotensin II blocker effective in lowering
129 CHAPTER-6 IDENTIFICATION, AND CHARACTERISATION OF DEGRADATION IMPURITY IN VALSARTAN TABLETS 130 6.1. Introduction Valsartan is an orally active specific angiotensin II blocker effective in lowering
For questions 1-4, match the carbohydrate with its size/functional group name:
 Chemistry 11 Fall 2013 Examination #5 PRACTICE 1 ANSWERS For the first portion of this exam, select the best answer choice for the questions below and mark the answers on your scantron. Then answer the
Chemistry 11 Fall 2013 Examination #5 PRACTICE 1 ANSWERS For the first portion of this exam, select the best answer choice for the questions below and mark the answers on your scantron. Then answer the
The Structure and Function of Large Biological Molecules Part 4: Proteins Chapter 5
 Key Concepts: The Structure and Function of Large Biological Molecules Part 4: Proteins Chapter 5 Proteins include a diversity of structures, resulting in a wide range of functions Proteins Enzymatic s
Key Concepts: The Structure and Function of Large Biological Molecules Part 4: Proteins Chapter 5 Proteins include a diversity of structures, resulting in a wide range of functions Proteins Enzymatic s
UV Tracer TM Maleimide NHS ester
 UV Tracer TM Maleimide HS ester Product o.: 1020 Product ame: UV-Tracer TM Maleimide-HS ester Chemical Structure: Chemical Composition: C 41 H 67 5 18 Molecular Weight: 1014.08 Appearance: Storage: Yellow
UV Tracer TM Maleimide HS ester Product o.: 1020 Product ame: UV-Tracer TM Maleimide-HS ester Chemical Structure: Chemical Composition: C 41 H 67 5 18 Molecular Weight: 1014.08 Appearance: Storage: Yellow
Lecture 3. Tandem MS & Protein Sequencing
 Lecture 3 Tandem MS & Protein Sequencing Nancy Allbritton, M.D., Ph.D. Department of Physiology & Biophysics 824-9137 (office) nlallbri@uci.edu Office- Rm D349 Medical Science D Bldg. Tandem MS Steps:
Lecture 3 Tandem MS & Protein Sequencing Nancy Allbritton, M.D., Ph.D. Department of Physiology & Biophysics 824-9137 (office) nlallbri@uci.edu Office- Rm D349 Medical Science D Bldg. Tandem MS Steps:
Student Biochemistry I Homework III Due 10/13/04 64 points total (48 points based on text; 16 points for Swiss-PDB viewer exercise)
 Biochemistry I Homework III Due 10/13/04 64 points total (48 points based on text; 16 points for Swiss-PDB viewer exercise) 1). 20 points total T or F; if false, provide a brief rationale as to why. Only
Biochemistry I Homework III Due 10/13/04 64 points total (48 points based on text; 16 points for Swiss-PDB viewer exercise) 1). 20 points total T or F; if false, provide a brief rationale as to why. Only
AMINO ACIDS STRUCTURE, CLASSIFICATION, PROPERTIES. PRIMARY STRUCTURE OF PROTEINS
 AMINO ACIDS STRUCTURE, CLASSIFICATION, PROPERTIES. PRIMARY STRUCTURE OF PROTEINS Elena Rivneac PhD, Associate Professor Department of Biochemistry and Clinical Biochemistry State University of Medicine
AMINO ACIDS STRUCTURE, CLASSIFICATION, PROPERTIES. PRIMARY STRUCTURE OF PROTEINS Elena Rivneac PhD, Associate Professor Department of Biochemistry and Clinical Biochemistry State University of Medicine
For questions 1-4, match the carbohydrate with its size/functional group name:
 Chemistry 11 Fall 2013 Examination #5 PRACTICE 1 For the first portion of this exam, select the best answer choice for the questions below and mark the answers on your scantron. Then answer the free response
Chemistry 11 Fall 2013 Examination #5 PRACTICE 1 For the first portion of this exam, select the best answer choice for the questions below and mark the answers on your scantron. Then answer the free response
Chemical Nature of the Amino Acids. Table of a-amino Acids Found in Proteins
 Chemical Nature of the Amino Acids All peptides and polypeptides are polymers of alpha-amino acids. There are 20 a- amino acids that are relevant to the make-up of mammalian proteins (see below). Several
Chemical Nature of the Amino Acids All peptides and polypeptides are polymers of alpha-amino acids. There are 20 a- amino acids that are relevant to the make-up of mammalian proteins (see below). Several
DELFIA Tb-N1 DTA Chelate & Terbium Standard
 AD0029P-1 (en) 1 DELFIA Tb-N1 DTA Chelate & AD0012 Terbium Standard For Research Use Only INTRODUCTION DELFIA Tb-N1 DTA Chelate is optimized for the terbium labeling of proteins and peptides for use in
AD0029P-1 (en) 1 DELFIA Tb-N1 DTA Chelate & AD0012 Terbium Standard For Research Use Only INTRODUCTION DELFIA Tb-N1 DTA Chelate is optimized for the terbium labeling of proteins and peptides for use in
