Application of the Mole-8.5 supercomputer: Probing the whole influenza virion at the atomic level
|
|
|
- Lewis Mitchell
- 6 years ago
- Views:
Transcription
1 Article Molecular Biology July 2011 Vol.56 No.20: doi: /s SPECIAL TOPICS: Application of the Mole-8.5 supercomputer: Probing the whole influenza virion at the atomic level XU Ji 1,2, WANG XiaoWei 1, HE XianFeng 1, REN Ying 1*, GE Wei 1* & LI JingHai 1 1 State Key Laboratory of Multiphase Complex Systems, Institute of Process Engineering, Chinese Academy of Sciences, Beijing , China; 2 Graduate University of the Chinese Academy of Sciences, Beijing , China Received March 18, 2011; accepted April 7, 2011 Discrete computer simulations are quite helpful in understanding dynamic structures in complex systems. Recently, using the Mole-8.5 supercomputer and molecular dynamics simulations as a computational microscope, we simulated the dynamic structure of a whole H1N1 influenza virion in solution for the first time at the atomic level. In total, 300 million atoms in a periodic cube with an edge length of nm were simulated. Using 288 low level hybrids with 1728 C2050 GPUs and a software package developed specifically for the hardware, the simulation executed 770 ps/d with an integration time step of 1 fs, and analyzed the dynamic structure. With the tremendous computational power of GPUs, efficient software packages for various hardware designs, and consistent physical models, more challenging applications will be carried out in the near future. influenza virion, Mole-8.5, supercomputer, GPU, molecular dynamics simulation Citation: Xu J, Wang X W, He X F, et al. Application of the Mole-8.5 supercomputer: Probing the whole influenza virion at the atomic level. Chinese Sci Bull, 2011, 56: , doi: /s During the past few decades, discrete simulation has been used as a computational microscope to probe the dynamic structures of complex systems, especially when inspection instruments are insufficient in resolving capability, or operational conditions are difficult to realize in experiments. In chemical engineering, detailed information of individual particles is necessary to understand the heterogeneity structure in both gas-solid and liquid-solid systems. In life science, an understanding of the dynamic structure of the influenza virion in vivo is quite useful in preventing influenza epidemics and designing anti-influenza drugs. However, these processes are too fast and too delicate to capture using current experimental facilities. On the other hand, computer simulations of these systems usually involve billions of particles, which is beyond the reach of existing supercomputers. In a wide range of scientific fields where such challenges occur, urgent demands for large-scale simulations are calling for the provision of *Corresponding authors ( yren@home.ipe.ac.cn; wge@home.ipe.ac.cn) more powerful computational capability. 1 Computational facility A supercomputer called Mole-8.5 with about 2.0 Petaflops peak performance in single precision and 1.0 Petaflops in double precision was formally established in April 2010 [1]. It was designed and implemented by the Institute of Process Engineering (IPE), Chinese Academy of Sciences (CAS), one of the NVIDIA CUDA Centers of Excellence (CCOE) (CUDA, Computed Unified Device Architecture, a prevailing programming infrastructure for general purpose graphic processing units, or GPUs). The Mole-8.5 embodies a design philosophy utilizing the structural similarity between hardware, software, and the problems to be solved [2], based on the multi-scale method and discrete simulation approaches developed at IPE [3 5]. This supercomputer, which includes 362 nodes, is the first GPGPU supercomputer using NVIDIA Tesla C2050 boards in the world. It The Author(s) This article is published with open access at Springerlink.com csb.scichina.com
2 Xu J, et al. Chinese Sci Bull July (2011) Vol.56 No contains a multi-level structure, with high level CPU computing nodes, mid-level visualization-computing nodes connected to a 3 6 DID LCD array for online visualization, and low level hybrid computing nodes. The simulation work is mainly carried out by the low level nodes, which use 2 Intel(R) Xeon(R) E GHz CPUs, 48 GB DDR MHz memory, and 6 C2050 GPU boards to accelerate the computation. All the nodes are connected via a Gigabit Ethernet and QDR Infiniband network. If computation of the application running on the Mole-8.5 is fast enough, the results can be displayed online on the display wall of 3 6 DID LCD panels at the same time as the computation, thereby realizing realtime simulation. The general structure of the whole system is given in Figure 1 [1]. The Mole-8.5 has some unique advantages over a HPC system with the same performance based on CPUs; that is, the high performance/price ratio, and the fact that it occupies only about 150 m 2. Though the system is mainly designed for multiscale discrete simulation, the Linpack performance of its 320 low level nodes still reaches 207 TFlops, with a power efficiency of 431 Mflop/s/Watt, debuting at No. 8 on the Green500 list in June 2010 ( lists/2010/11/top/list.php?from=1&to=100), No. 19 on the 35th TOP500 list in June 2010 ( 2010/06/100), and No. 3 on the China TOP100 list in October 2010 [6]. With the specifically developed software package for the hardware structure described above, the Mole-8.5 has exhibited significant acceleration compared with traditional CPU-based clusters and has been used to perform large- scale simulations in chemical engineering, such as a continuum-scale direct numerical simulation of a gas-solid flow system with up to 672 GPUs 1) and a quasi-realtime simulation of an industrial scale rotating drum using the discrete element method with more than 200 GPUs 2). Recently, using a molecular dynamics (MD) simulation software package developed from our previous work [7], it has become feasible to probe the dynamic structure of the whole H1N1 virion at the atomic level. 2 Stationary structure The influenza virion, an enveloped RNA virus belonging to the Orthomyxoviridae family, is a major cause of global infection and mortality. The H1N1 virus, with the numbers denoting the subtypes of hemagglutinin (HA) and neuraminidase (NA), respectively, is a subtype of the influenza A virus and was responsible for the flu pandemic in Influenza virions are highly pleomorphic, with sizes ranging from spherical particles with a diameter of approximately nm to filamentous particles with a length of several mm [8]. As shown in Figure 2, a rough model of the general structure of influenza virions and crystal structures of single macromolecules comprising the virions have been provided by electron microscopes, X-ray diffraction, etc. The influenza A virion is studded with glycoprotein spikes of a large number of HA and NA, at a ratio of approximately four to one, projecting from a lipid membrane [9]. HA is evenly spaced, while NA is located in clusters. The Figure 1 Global architecture of the entire Mole-8.5 system [1]. 1) The EMMS group (Principal authors: Wei Ge, Wei Wang, Ning Yang, Jinghai Li, Mooson Kwauk). Meso-scale oriented simulation towards virtual process engineering (VPE) the EMMS Paradigm. Chem Eng Sci, 2011, doi: /j.ces ) Xu J, Qi H, Fang X, et al. Quasi-realtime simulation of rotating drum using discrete element method with parallel GPU computing. Accepted by Particuology, 2011
3 2116 Xu J, et al. Chinese Sci Bull July (2011) Vol.56 No.20 Figure 2 The main procedure for probing H1N1 dynamic structure at the atomic level. Figures of the electron microscopy and the model were taken from ref. [9]. A quarter of the outer sphere is not shown, and components are shown in different colors for better visualization. membrane also contains the matrix protein M 2, although in much lower quantities (1: M 2 :HA) [10]. Matrix protein M 1, the most abundant component of the virion, lies just beneath the lipid envelope and its three integral membrane proteins, resulting in an enclosure of the virion core [11]. The viral core comprises eight helical viral ribonucleoproteins (vrnps), each containing a segment of single-stranded negative-sense RNA, multiple copies of nucleoprotein (NP), and three polymerase proteins (PB1, PB2, PA). The information above has already provided a static picture of the 3D structure of influenza virions, though the model is rough and the structures of some componential molecules are still unknown or not completely known. Computer simulations can be utilized as an important tool to complement the current theoretical and experimental understanding. First, the missing details should be reconstructed to provide an atomic structure of the virion, and then, an explicit solvent MD study should be performed to detect further dynamics of the virion in vivo. The globular lipid membrane is made up of a double layer of dipalmitoylphosphatidylcholine (DPPC), resulting in a globular membrane with an external diameter of 106 nm. Three transmembrane proteins, NA, HA, and M 2 are constructed following similar procedures, that is, reconstruction of single macromolecular structure at atomic-scale followed by preparation of a matrix of the macromolecules on a spherical surface. A total of 374 HA and 98 NA molecules are thus located on a spherical sphere with their tails embedded in the lipid membrane as appropriate. Forty-seven proton channels penetrate the membrane, with each proton channel formed by 4 parallel monomers of M 2. The spherical protein layer of the M 1 matrix is constructed in the same way as the membrane. A vrnp is constructed by placing NP monomers in a helical structure and locating a packed single-stranded negative-sense RNA strand on the surface of the protein scaffold. The simulated H1N1 virion is finally constructed with 2363 proteins, DPPC molecules and 8 RNA strands, with details of the components given in Table 1. Each component has been equilibrated before being assembled into the virion. The complex is then solvated into SPC [12] water with an appropriate concentration of ions, resulting in 300 million atoms in a periodic cube with each side nm long. The entire system is equilibrated again, and then simulated at 300 K with a slightly modified GROMOS 53a6 force field [13] against RNA strands. Berendsen s temperature coupling method [14] is adopted in this work. Table 1 Components of the H1N1 virion Component PDB ID No. of molecules HA 3LZG 374 NA 2HTY 98 M1 1AA M2 2RLF 164 NP 2IQH 147 Lipids RNA 8
4 Xu J, et al. Chinese Sci Bull July (2011) Vol.56 No Software The software package used here is based on GPU_MD [7], which is the single GPU version software for simulating macromolecules with high efficiency. Parallel simulation on multiple GPUs via three-dimensional domain decomposition [15] is implemented to probe the whole virus structure at the atomic level. One domain also needs the particle data of its surrounding space, usually called the virtual boundary, besides its own to perform the calculations in CPU_MD The particle data of the virtual boundary are, in turn, constructed along the direction of decomposition [5], i.e., X, Y, Z. All particles reside in the same process in a single GPU computation, while they transfer among the processes in the parallel implementation. After the particles have moved in or out, the original arrays of particles will have empty spaces in them. To make full use of the memory bandwidth and avoid vacant memory access, the particles should be rearranged in memory after a given number of simulation steps as some particles move out to other processes, executed by the CPU, and others move in, that is, transfer back to the GPU. The bonded interaction relations are changed after the rearrangement of particles, so the bonded interaction lists should be reconstructed correspondingly. To maximize GPU efficiency, the amount and times of data transfer between GPU and CPU should be minimized. Only the positions of the particles in the boundary space are transferred at each step. Regarding this point, all the particle data needed by one process is constructed and stored locally. Then the computational task in each GPU can be performed in the same way as those of the GPU_MD The computations are carried out on the low level hybrid computing nodes, the main computational resources of the Mole-8.5, where GPUs deal with simple computational tasks and CPUs deal with the more complex and high level decision-making tasks. With 288 nodes comprising 1,728 C2050 GPUs, the simulation reached a speed of 770 ps/d with an integration time step of 1 fs. Starting from the predefined structure, the virus experiences a significant change to obtain a stable structure. As shown in Figure 3, the 3 components of the radius of gyration (R gx, R gy and R gz ), calculated for all the atoms in the lipid membrane and transmembrane proteins, show similar behaviors during the 0.5 ns simulation. The profiles decrease sharply during the first 0.1 ns, suggesting a quick adjustment of the unfavorable structures; then the profiles slowly increase in the next 0.05 ns and remain comparatively stable thereafter. In accordance with the structural changes, the potential energy (Figure 4(a)), including both intra- and inter-molecular Van der Waals interactions and columbic interactions, together with the solvation energy of the virion (Figure 4(b)), show a significant decrease in the first 0.1 ns, but later become relatively stable. The plots of the structural changes and energetic properties suggest the virion slowly evolves to a stable structure in the aqueous solution. The hardware and software used in this work serve as nutrients to keep the virion alive. This endeavor may help us to investigate the biological behaviors of the influenza virus, find its weakness, and thus provide valuable knowledge to prevent future influenza epidemics. 5 Conclusions For a long time, molecular simulations have focused on Figure 3 R gx, R gy and R gz of the lipid membrane and transmembrane proteins of a virus as a function of simulation time. 4 Computational results and discussion Figure 4 Potential energy (a) and solvation energy (b) of the virion as a function of simulation time.
5 2118 Xu J, et al. Chinese Sci Bull July (2011) Vol.56 No.20 individual biological molecules. In recent years, the capacity of computational resources has increased to macrobiomolecular complexes. Now, it is encouraging to see that the new era of simulating biological agents has arrived. With the tremendous computational power of GPUs, efficient software packages for various hardware designs, and consistent physical models, more challenging applications will be carried out in the near future. This work was supported by Ministry of Finance (ZDYZ2008-2), the National Key Science and Technology Project (2008ZX HZ), the Knowledge Innovation Project of Chinese Academy of Sciences (KGCX2-YW-124), and the National Natural Science Foundation of China ( ). 1 Wang X, Ge W, He X, et al. Development and application of a HPC system for multi-scale discrete simulation-mole-8.5. International Supercomputing Conference, Germany, Chen F, Ge W, Guo L, et al. Multi-scale HPC system for multi-scale discrete simulation Development and application of a supercomputer with 1 Petaflops peak performance in single precision. Particuology, 2009, 7: Ge W, Li J. General approach for discrete simulation of complex systems. Chinese Sci Bull, 2002, 47: Tang D. A general method of parallel computation of particle method and its preliminary applications. Doctor Dissertation. Beijing: Chinese Academy of Sciences, Wang X. A framework for parallel simulation of particle systems with pairadditive local interactions-toward a general approach. Doctor Dissertation. Beijing: Chinese Academy of Sciences, Zhang Y, Sun J, Yuan G, et al. Perspectives of China s HPC system development: A view from the 2009 China HPC TOP100 list. Front Comput Sci China, 2010, 4: Xu J, Ren Y, Ge W, et al. Molecular dynamics simulation of macromolecules using graphics processing unit. Mol Simulat, 2010, 36: Roberts P C, Compans R W. Host cell dependence of viral morphology. Proc Natl Acad Sci USA, 1998, 95: Lamb R A, Krug R M. Orthomyxoviridae: The viruses and their replication. In: Knipe D M, Howley P M, eds. Field s Virology. Philadelphia: Lippincott-Raven Publishers, Zebedee S L, Lamb R A. Influenza A virus M2 protein: Monoclonal antibody restriction of virus growth and detection of M 2 in virions. J Virol, 1988, 62: Yasuda J, Bucher D J, Ishihama A. Growth control of influenza A virus by M 1 protein: Analysis of transfectant viruses carrying the chimeric M gene. J Virol, 1994, 68: Berendsen H J C, Postma J P M, Van Gunsteren W F, et al. Interaction models for water in relation to protein hydration. Intermol Forces, 1981, 11(Suppl 1): Oostenbrink C, Villa A, Mark A E, et al. A biomolecular force field based on the free enthalpy of hydration and solvation: The GROMOS force-field parameter sets 53A5 and 53A6. J Comput Chem, 2004, 25: Berendsen H J C, Postma J P M, Van Gunsteren W F, et al. Molecular dynamics with coupling to an external bath. J Chem Phys, 1984, 81: Shaw D E. A fast, scalable method for the parallel evaluation of distance-limited pairwise particle interactions. J Comput Chem, 2005, 26: 1803 Open Access This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Influenza viruses. Virion. Genome. Genes and proteins. Viruses and hosts. Diseases. Distinctive characteristics
 Influenza viruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Enveloped particles, quasi-spherical or filamentous Diameter 80-120 nm Envelope is derived
Influenza viruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Enveloped particles, quasi-spherical or filamentous Diameter 80-120 nm Envelope is derived
Supplementary Information
 Coarse-Grained Molecular Dynamics Simulations of Photoswitchable Assembly and Disassembly Xiaoyan Zheng, Dong Wang,* Zhigang Shuai,* MOE Key Laboratory of Organic Optoelectronics and Molecular Engineering,
Coarse-Grained Molecular Dynamics Simulations of Photoswitchable Assembly and Disassembly Xiaoyan Zheng, Dong Wang,* Zhigang Shuai,* MOE Key Laboratory of Organic Optoelectronics and Molecular Engineering,
Project PRACE 1IP, WP7.4
 Project PRACE 1IP, WP7.4 Plamenka Borovska, Veska Gancheva Computer Systems Department Technical University of Sofia The Team is consists of 5 members: 2 Professors; 1 Assist. Professor; 2 Researchers;
Project PRACE 1IP, WP7.4 Plamenka Borovska, Veska Gancheva Computer Systems Department Technical University of Sofia The Team is consists of 5 members: 2 Professors; 1 Assist. Professor; 2 Researchers;
University of Groningen. Computational studies of influenza hemagglutinin Boonstra, Sander
 University of Groningen Computational studies of influenza hemagglutinin Boonstra, Sander IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's PDF) if you wish to cite from it.
University of Groningen Computational studies of influenza hemagglutinin Boonstra, Sander IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's PDF) if you wish to cite from it.
Viral structure م.م رنا مشعل
 Viral structure م.م رنا مشعل Viruses must reproduce (replicate) within cells, because they cannot generate energy or synthesize proteins. Because they can reproduce only within cells, viruses are obligate
Viral structure م.م رنا مشعل Viruses must reproduce (replicate) within cells, because they cannot generate energy or synthesize proteins. Because they can reproduce only within cells, viruses are obligate
Supplementary information for Effects of Stretching Speed on. Mechanical Rupture of Phospholipid/Cholesterol Bilayers: Molecular
 Supplementary information for Effects of Stretching Speed on Mechanical Rupture of Phospholipid/Cholesterol Bilayers: Molecular Dynamics Simulation Taiki Shigematsu, Kenichiro Koshiyama*, and Shigeo Wada
Supplementary information for Effects of Stretching Speed on Mechanical Rupture of Phospholipid/Cholesterol Bilayers: Molecular Dynamics Simulation Taiki Shigematsu, Kenichiro Koshiyama*, and Shigeo Wada
Interaction of Functionalized C 60 Nanoparticles with Lipid Membranes
 Interaction of Functionalized C 60 Nanoparticles with Lipid Membranes Kevin Gasperich Advisor: Dmitry Kopelevich ABSTRACT The recent rapid development of applications for fullerenes has led to an increase
Interaction of Functionalized C 60 Nanoparticles with Lipid Membranes Kevin Gasperich Advisor: Dmitry Kopelevich ABSTRACT The recent rapid development of applications for fullerenes has led to an increase
ERA: Architectures for Inference
 ERA: Architectures for Inference Dan Hammerstrom Electrical And Computer Engineering 7/28/09 1 Intelligent Computing In spite of the transistor bounty of Moore s law, there is a large class of problems
ERA: Architectures for Inference Dan Hammerstrom Electrical And Computer Engineering 7/28/09 1 Intelligent Computing In spite of the transistor bounty of Moore s law, there is a large class of problems
Influenza Virus Genotypes Circulating In Central Greece During And Vaccine Strain Match
 ISPUB.COM The Internet Journal of Microbiology Volume 13 Number 1 Influenza Virus Genotypes Circulating In Central Greece During 2012-2014 And Vaccine Strain Match E Plakokefalos, A Vontas, Z Florou, G
ISPUB.COM The Internet Journal of Microbiology Volume 13 Number 1 Influenza Virus Genotypes Circulating In Central Greece During 2012-2014 And Vaccine Strain Match E Plakokefalos, A Vontas, Z Florou, G
Virus Structure. Characteristics of capsids. Virus envelopes. Virion assembly John Wiley & Sons, Inc. All rights reserved.
 Virus Structure Characteristics of capsids Virus envelopes Virion assembly Capsids package viral genomes and transmit them to a new host cell Capsid rigid, symmetrical container composed of viral protein
Virus Structure Characteristics of capsids Virus envelopes Virion assembly Capsids package viral genomes and transmit them to a new host cell Capsid rigid, symmetrical container composed of viral protein
 http://nmhm.washingtondc.museum/collections/archives/agalleries/1918flu/ncp1603.jpg 1 https://assets-production-webvanta-com.s3-us-west- 2 2.amazonaws.com/000000/47/62/original/images/img_109_influenza/Spanish_flu_death_chart.jpg
http://nmhm.washingtondc.museum/collections/archives/agalleries/1918flu/ncp1603.jpg 1 https://assets-production-webvanta-com.s3-us-west- 2 2.amazonaws.com/000000/47/62/original/images/img_109_influenza/Spanish_flu_death_chart.jpg
Virology. *Viruses can be only observed by electron microscope never by light microscope. The size of the virus: nm in diameter.
 Virology We are going to start with general introduction about viruses, they are everywhere around us; in food; within the environment; in direct contact to etc.. They may cause viral infection by itself
Virology We are going to start with general introduction about viruses, they are everywhere around us; in food; within the environment; in direct contact to etc.. They may cause viral infection by itself
We parameterized a coarse-grained fullerene consistent with the MARTINI coarse-grained force field
 Parameterization of the fullerene coarse-grained model We parameterized a coarse-grained fullerene consistent with the MARTINI coarse-grained force field for lipids 1 and proteins 2. In the MARTINI force
Parameterization of the fullerene coarse-grained model We parameterized a coarse-grained fullerene consistent with the MARTINI coarse-grained force field for lipids 1 and proteins 2. In the MARTINI force
Evolution of influenza
 Evolution of influenza Today: 1. Global health impact of flu - why should we care? 2. - what are the components of the virus and how do they change? 3. Where does influenza come from? - are there animal
Evolution of influenza Today: 1. Global health impact of flu - why should we care? 2. - what are the components of the virus and how do they change? 3. Where does influenza come from? - are there animal
Supporting Information
 Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 018 Supporting Information Modulating Interactions between Ligand-Coated Nanoparticles and Phase-Separated
Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 018 Supporting Information Modulating Interactions between Ligand-Coated Nanoparticles and Phase-Separated
Medical Virology. Herpesviruses, Orthomyxoviruses, and Retro virus. - Herpesviruses Structure & Composition: Herpesviruses
 Medical Virology Lecture 2 Asst. Prof. Dr. Dalya Basil Herpesviruses, Orthomyxoviruses, and Retro virus - Herpesviruses Structure & Composition: Herpesviruses Enveloped DNA viruses. All herpesviruses have
Medical Virology Lecture 2 Asst. Prof. Dr. Dalya Basil Herpesviruses, Orthomyxoviruses, and Retro virus - Herpesviruses Structure & Composition: Herpesviruses Enveloped DNA viruses. All herpesviruses have
BIOL*1090 Introduction To Molecular and Cellular Biology Fall 2014
 Last time... BIOL*1090 Introduction To Molecular and Cellular Biology Fall 2014 Lecture 3 - Sept. 15, 2014 Viruses Biological Membranes Karp 7th ed: Chpt. 4; sections 4-1, 4-3 to 4-7 1 2 VIRUS Non-cellular
Last time... BIOL*1090 Introduction To Molecular and Cellular Biology Fall 2014 Lecture 3 - Sept. 15, 2014 Viruses Biological Membranes Karp 7th ed: Chpt. 4; sections 4-1, 4-3 to 4-7 1 2 VIRUS Non-cellular
Grade Level: Grades 9-12 Estimated Time Allotment Part 1: One 50- minute class period Part 2: One 50- minute class period
 The History of Vaccines Lesson Plan: Viruses and Evolution Overview and Purpose: The purpose of this lesson is to prepare students for exploring the biological basis of vaccines. Students will explore
The History of Vaccines Lesson Plan: Viruses and Evolution Overview and Purpose: The purpose of this lesson is to prepare students for exploring the biological basis of vaccines. Students will explore
AFM SURFACE ANALYSIS OF FUNGAL MODIFIED CTMP FIBERS
 CELLULOSE CHEMISTRY AND TECHNOLOGY AFM SURFACE ANALYSIS OF FUNGAL MODIFIED CTMP FIBERS CHONG-XING HUANG, QI-FENG YANG and SHUANG-FEI WANG College of Light Industry and Food Engineering, Guangxi University,
CELLULOSE CHEMISTRY AND TECHNOLOGY AFM SURFACE ANALYSIS OF FUNGAL MODIFIED CTMP FIBERS CHONG-XING HUANG, QI-FENG YANG and SHUANG-FEI WANG College of Light Industry and Food Engineering, Guangxi University,
Overview of virus life cycle
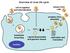 Overview of virus life cycle cell recognition and internalization release from cells progeny virus assembly membrane breaching nucleus capsid disassembly and genome release replication and translation
Overview of virus life cycle cell recognition and internalization release from cells progeny virus assembly membrane breaching nucleus capsid disassembly and genome release replication and translation
Research on Digital Testing System of Evaluating Characteristics for Ultrasonic Transducer
 Sensors & Transducers 2014 by IFSA Publishing, S. L. http://www.sensorsportal.com Research on Digital Testing System of Evaluating Characteristics for Ultrasonic Transducer Qin Yin, * Liang Heng, Peng-Fei
Sensors & Transducers 2014 by IFSA Publishing, S. L. http://www.sensorsportal.com Research on Digital Testing System of Evaluating Characteristics for Ultrasonic Transducer Qin Yin, * Liang Heng, Peng-Fei
Plasmid-Driven Formation of Influenza Virus-Like Particles
 JOURNAL OF VIROLOGY, Jan. 2000, p. 547 551 Vol. 74, No. 1 0022-538X/00/$04.00 0 Copyright 2000, American Society for Microbiology. All Rights Reserved. Plasmid-Driven Formation of Influenza Virus-Like
JOURNAL OF VIROLOGY, Jan. 2000, p. 547 551 Vol. 74, No. 1 0022-538X/00/$04.00 0 Copyright 2000, American Society for Microbiology. All Rights Reserved. Plasmid-Driven Formation of Influenza Virus-Like
Overview: Chapter 19 Viruses: A Borrowed Life
 Overview: Chapter 19 Viruses: A Borrowed Life Viruses called bacteriophages can infect and set in motion a genetic takeover of bacteria, such as Escherichia coli Viruses lead a kind of borrowed life between
Overview: Chapter 19 Viruses: A Borrowed Life Viruses called bacteriophages can infect and set in motion a genetic takeover of bacteria, such as Escherichia coli Viruses lead a kind of borrowed life between
19 2 Viruses Slide 1 of 34
 1 of 34 What Is a Virus? What Is a Virus? Viruses are particles of nucleic acid, protein, and in some cases, lipids. Viruses can reproduce only by infecting living cells. 2 of 34 What Is a Virus? Viruses
1 of 34 What Is a Virus? What Is a Virus? Viruses are particles of nucleic acid, protein, and in some cases, lipids. Viruses can reproduce only by infecting living cells. 2 of 34 What Is a Virus? Viruses
DIGITIZING HUMAN BRAIN: BLUE BRAIN PROJECT
 DIGITIZING HUMAN BRAIN: BLUE BRAIN PROJECT Diwijesh 1, Ms. Pooja Khanna 2 1 M.tech CSE, Amity University 2 Associate Professor, ASET ABSTRACT Human brain is most complex and unique creation of nature which
DIGITIZING HUMAN BRAIN: BLUE BRAIN PROJECT Diwijesh 1, Ms. Pooja Khanna 2 1 M.tech CSE, Amity University 2 Associate Professor, ASET ABSTRACT Human brain is most complex and unique creation of nature which
Coronaviruses cause acute, mild upper respiratory infection (common cold).
 Coronaviruses David A. J. Tyrrell Steven H. Myint GENERAL CONCEPTS Clinical Presentation Coronaviruses cause acute, mild upper respiratory infection (common cold). Structure Spherical or pleomorphic enveloped
Coronaviruses David A. J. Tyrrell Steven H. Myint GENERAL CONCEPTS Clinical Presentation Coronaviruses cause acute, mild upper respiratory infection (common cold). Structure Spherical or pleomorphic enveloped
Chapter 12: Membranes. Voet & Voet: Pages
 Chapter 12: Membranes Voet & Voet: Pages 390-415 Slide 1 Membranes Essential components of all living cells (define boundry of cells) exclude toxic ions and compounds; accumulation of nutrients energy
Chapter 12: Membranes Voet & Voet: Pages 390-415 Slide 1 Membranes Essential components of all living cells (define boundry of cells) exclude toxic ions and compounds; accumulation of nutrients energy
Supporting Information for Origin of 1/f noise in hydration dynamics on lipid membrane surfaces
 Supporting Informati for Origin of /f noise in hydrati dynamics lipid membrane surfaces Eiji Yamamoto, Takuma kimoto, Masato Yasui, 2 and Kenji Yasuoka Department of Mechanical Engineering, Keio University,
Supporting Informati for Origin of /f noise in hydrati dynamics lipid membrane surfaces Eiji Yamamoto, Takuma kimoto, Masato Yasui, 2 and Kenji Yasuoka Department of Mechanical Engineering, Keio University,
Modeling the Antigenic Evolution of Influenza Viruses from Sequences
 Modeling the Antigenic Evolution of Influenza Viruses from Sequences Taijiao Jiang Center of Systems Medicine, Chinese Academy of Medical Sciences Suzhou Institute of Systems Medicine October 8-10, 2015.
Modeling the Antigenic Evolution of Influenza Viruses from Sequences Taijiao Jiang Center of Systems Medicine, Chinese Academy of Medical Sciences Suzhou Institute of Systems Medicine October 8-10, 2015.
Palindromes drive the re-assortment in Influenza A
 www.bioinformation.net Hypothesis Volume 7(3) Palindromes drive the re-assortment in Influenza A Abdullah Zubaer 1, 2 * & Simrika Thapa 1, 2 1Swapnojaatra Bioresearch Laboratory, DataSoft Systems, Dhaka-15,
www.bioinformation.net Hypothesis Volume 7(3) Palindromes drive the re-assortment in Influenza A Abdullah Zubaer 1, 2 * & Simrika Thapa 1, 2 1Swapnojaatra Bioresearch Laboratory, DataSoft Systems, Dhaka-15,
19 Viruses BIOLOGY. Outline. Structural Features and Characteristics. The Good the Bad and the Ugly. Structural Features and Characteristics
 9 Viruses CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson Outline I. Viruses A. Structure of viruses B. Common Characteristics of Viruses C. Viral replication D. HIV Lecture Presentation
9 Viruses CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson Outline I. Viruses A. Structure of viruses B. Common Characteristics of Viruses C. Viral replication D. HIV Lecture Presentation
Chapter 19: Viruses. 1. Viral Structure & Reproduction. What exactly is a Virus? 11/7/ Viral Structure & Reproduction. 2.
 Chapter 19: Viruses 1. Viral Structure & Reproduction 2. Bacteriophages 3. Animal Viruses 4. Viroids & Prions 1. Viral Structure & Reproduction Chapter Reading pp. 393-396 What exactly is a Virus? Viruses
Chapter 19: Viruses 1. Viral Structure & Reproduction 2. Bacteriophages 3. Animal Viruses 4. Viroids & Prions 1. Viral Structure & Reproduction Chapter Reading pp. 393-396 What exactly is a Virus? Viruses
Coarse-Grained Molecular Dynamics for Copolymer- Vesicle Self-Assembly. Case Study: Sterically Stabilized Liposomes.
 Coarse-Grained Molecular Dynamics for Copolymer- Vesicle Self-Assembly. Case Study: Sterically Stabilized Liposomes. Alexander Kantardjiev 1 and Pavletta Shestakova 1 1 Institute of Organic Chemistry with
Coarse-Grained Molecular Dynamics for Copolymer- Vesicle Self-Assembly. Case Study: Sterically Stabilized Liposomes. Alexander Kantardjiev 1 and Pavletta Shestakova 1 1 Institute of Organic Chemistry with
Bioinformation Volume 5
 Identification of sequence mutations affecting hemagglutinin specificity to sialic acid receptor in influenza A virus subtypes Usman Sumo Friend Tambunan*, Ramdhan Department of Chemistry, Faculty of Mathematics
Identification of sequence mutations affecting hemagglutinin specificity to sialic acid receptor in influenza A virus subtypes Usman Sumo Friend Tambunan*, Ramdhan Department of Chemistry, Faculty of Mathematics
Supporting material. Membrane permeation induced by aggregates of human islet amyloid polypeptides
 Supporting material Membrane permeation induced by aggregates of human islet amyloid polypeptides Chetan Poojari Forschungszentrum Jülich GmbH, Institute of Complex Systems: Structural Biochemistry (ICS-6),
Supporting material Membrane permeation induced by aggregates of human islet amyloid polypeptides Chetan Poojari Forschungszentrum Jülich GmbH, Institute of Complex Systems: Structural Biochemistry (ICS-6),
PROCEEDINGS OF THE YEREVAN STATE UNIVERSITY
 PROCEEDINGS OF THE YEREVAN STATE UNIVERSITY Physical and Mathematical Sciences 2018, 52(3), p. 217 221 P h y s i c s STUDY OF THE SWELLING OF THE PHOSPHOLIPID BILAYER, DEPENDING ON THE ANGLE BETWEEN THE
PROCEEDINGS OF THE YEREVAN STATE UNIVERSITY Physical and Mathematical Sciences 2018, 52(3), p. 217 221 P h y s i c s STUDY OF THE SWELLING OF THE PHOSPHOLIPID BILAYER, DEPENDING ON THE ANGLE BETWEEN THE
Modeling of epidemic spreading with white Gaussian noise
 Article Statistical Physics and Mathematics for Complex Systems December 20 Vol.56 No.34: 3683 3688 doi: 0.007/s434-0-4753-z SPECIAL TOPICS: Modeling of epidemic spreading with white Gaussian noise GU
Article Statistical Physics and Mathematics for Complex Systems December 20 Vol.56 No.34: 3683 3688 doi: 0.007/s434-0-4753-z SPECIAL TOPICS: Modeling of epidemic spreading with white Gaussian noise GU
Avian Influenza Virus H7N9. Dr. Di Liu Network Information Center Institute of Microbiology Chinese Academy of Sciences
 Avian Influenza Virus H7N9 Dr. Di Liu Network Information Center Institute of Microbiology Chinese Academy of Sciences Avian Influenza Virus RNA virus, Orthomyxoviruses Influenza A virus Eight Gene segments
Avian Influenza Virus H7N9 Dr. Di Liu Network Information Center Institute of Microbiology Chinese Academy of Sciences Avian Influenza Virus RNA virus, Orthomyxoviruses Influenza A virus Eight Gene segments
Growing knowledge of cell biology at the fundamental molecular level opened the door to new ways of inventing drugs
 E-tutorial 5 Rational drug design - The way we find new medicines was modernised as the 20 th and 21 st centuries progressed - Modernisation was driven by many factors, but above all by an improved understanding
E-tutorial 5 Rational drug design - The way we find new medicines was modernised as the 20 th and 21 st centuries progressed - Modernisation was driven by many factors, but above all by an improved understanding
Chapter 19: Viruses. 1. Viral Structure & Reproduction. 2. Bacteriophages. 3. Animal Viruses. 4. Viroids & Prions
 Chapter 19: Viruses 1. Viral Structure & Reproduction 2. Bacteriophages 3. Animal Viruses 4. Viroids & Prions 1. Viral Structure & Reproduction Chapter Reading pp. 393-396 What exactly is a Virus? Viruses
Chapter 19: Viruses 1. Viral Structure & Reproduction 2. Bacteriophages 3. Animal Viruses 4. Viroids & Prions 1. Viral Structure & Reproduction Chapter Reading pp. 393-396 What exactly is a Virus? Viruses
Reverse Genetics of RNA Viruses
 Reverse Genetics of RNA Viruses Reverse Genetics (RG) he creation of a virus with a fulllength copy of the viral genome he most powerful tool in modern virology RG of RNA viruses Generation or recovery
Reverse Genetics of RNA Viruses Reverse Genetics (RG) he creation of a virus with a fulllength copy of the viral genome he most powerful tool in modern virology RG of RNA viruses Generation or recovery
INFLUENZA VIRUS. INFLUENZA VIRUS CDC WEBSITE
 INFLUENZA VIRUS INFLUENZA VIRUS CDC WEBSITE http://www.cdc.gov/ncidod/diseases/flu/fluinfo.htm 1 THE IMPACT OF INFLUENZA Deaths: PANDEMICS 1918-19 S p a n is h flu 5 0 0,0 0 0 U S 2 0,0 0 0,0 0 0 w o rld
INFLUENZA VIRUS INFLUENZA VIRUS CDC WEBSITE http://www.cdc.gov/ncidod/diseases/flu/fluinfo.htm 1 THE IMPACT OF INFLUENZA Deaths: PANDEMICS 1918-19 S p a n is h flu 5 0 0,0 0 0 U S 2 0,0 0 0,0 0 0 w o rld
Final Report. Identification of Residue Mutations that Increase the Binding Affinity of LSTc to HA RBD. Wendy Fong ( 方美珍 ) CNIC; Beijing, China
 Final Report Identification of Residue Mutations that Increase the Binding Affinity of LSTc to HA RBD Wendy Fong ( 方美珍 ) CNIC; Beijing, China Influenza s Viral Life Cycle 1. HA on virus binds to sialic
Final Report Identification of Residue Mutations that Increase the Binding Affinity of LSTc to HA RBD Wendy Fong ( 方美珍 ) CNIC; Beijing, China Influenza s Viral Life Cycle 1. HA on virus binds to sialic
BIOPHYSICS II. By Prof. Xiang Yang Liu Department of Physics,
 BIOPHYSICS II By Prof. Xiang Yang Liu Department of Physics, NUS 1 Hydrogen bond and the stability of macromolecular structure Membrane Model Amphiphilic molecule self-assembly at the surface and din the
BIOPHYSICS II By Prof. Xiang Yang Liu Department of Physics, NUS 1 Hydrogen bond and the stability of macromolecular structure Membrane Model Amphiphilic molecule self-assembly at the surface and din the
Lecture 15. Membrane Proteins I
 Lecture 15 Membrane Proteins I Introduction What are membrane proteins and where do they exist? Proteins consist of three main classes which are classified as globular, fibrous and membrane proteins. A
Lecture 15 Membrane Proteins I Introduction What are membrane proteins and where do they exist? Proteins consist of three main classes which are classified as globular, fibrous and membrane proteins. A
Origins and evolutionary genomics of the novel avian-origin H7N9 influenza A virus in China: Early findings
 Origins and evolutionary genomics of the novel 2013 avian-origin H7N9 influenza A virus in : Early findings Jiankui He*, Luwen Ning, Yin Tong Department of Biology, South University of Science and Technology
Origins and evolutionary genomics of the novel 2013 avian-origin H7N9 influenza A virus in : Early findings Jiankui He*, Luwen Ning, Yin Tong Department of Biology, South University of Science and Technology
Molecular Dynamics of HIV-1 Reverse Transcriptase
 Molecular Dynamics of HIV-1 Reverse Transcriptase Abderrahmane Benghanem Rensselaer Polytechnic Institute,Troy, NY Mentor: Dr. Maria Kurnikova Carnegie Mellon, Pittsburgh PA Outline HIV-1 Reverse Transcriptase
Molecular Dynamics of HIV-1 Reverse Transcriptase Abderrahmane Benghanem Rensselaer Polytechnic Institute,Troy, NY Mentor: Dr. Maria Kurnikova Carnegie Mellon, Pittsburgh PA Outline HIV-1 Reverse Transcriptase
Viral reproductive cycle
 Lecture 29: Viruses Lecture outline 11/11/05 Types of viruses Bacteriophage Lytic and lysogenic life cycles viruses viruses Influenza Prions Mad cow disease 0.5 µm Figure 18.4 Viral structure of capsid
Lecture 29: Viruses Lecture outline 11/11/05 Types of viruses Bacteriophage Lytic and lysogenic life cycles viruses viruses Influenza Prions Mad cow disease 0.5 µm Figure 18.4 Viral structure of capsid
Inter-Species Cross-Seeding: Stability and Assembly of Rat - Human Amylin Aggregates. Workalemahu M. Berhanu
 Inter-Species Cross-Seeding: Stability and Assembly of Rat - Human Amylin Aggregates Workalemahu M. Berhanu University of Oklahoma Department of Chemistry and Biochemistry STRUCTURE OF AMYLOIDS & ROLE
Inter-Species Cross-Seeding: Stability and Assembly of Rat - Human Amylin Aggregates Workalemahu M. Berhanu University of Oklahoma Department of Chemistry and Biochemistry STRUCTURE OF AMYLOIDS & ROLE
PENETRANT TESTING: ABILITY EXTENSION AND SENSITIVITY EVALUATION
 PENETRANT TESTING: ABILITY EXTENSION AND SENSITIVITY EVALUATION Migoun N. P. Institute of Applied Physics of National Academy of Sciences of Belarus, Minsk, Belarus Introduction. Several ways of the increasing
PENETRANT TESTING: ABILITY EXTENSION AND SENSITIVITY EVALUATION Migoun N. P. Institute of Applied Physics of National Academy of Sciences of Belarus, Minsk, Belarus Introduction. Several ways of the increasing
Biology 350: Microbial Diversity
 Biology 350: Microbial Diversity Strange Invaders: Viruses, viroids, and prions. Lecture #27 7 November 2007-1- Notice handouts and announcements for today: Outline and study questions A 1999 paper discussing
Biology 350: Microbial Diversity Strange Invaders: Viruses, viroids, and prions. Lecture #27 7 November 2007-1- Notice handouts and announcements for today: Outline and study questions A 1999 paper discussing
High brightness electron beam for radiation therapy A new approach
 High brightness electron beam for radiation therapy A new approach Wei Gai ( 盖炜 ) Engineering Physics Department, Tsinghua University, Beijing, China Argonne National Laboratory, Argonne, IL 60439, USA
High brightness electron beam for radiation therapy A new approach Wei Gai ( 盖炜 ) Engineering Physics Department, Tsinghua University, Beijing, China Argonne National Laboratory, Argonne, IL 60439, USA
Cellular membranes are fluid mosaics of lipids and proteins.
 Study Guide e Plasma Membrane You should be able to write out the definitions to each of the following terms in your own words: plasma membrane fluid mosaic integral proteins peripheral proteins receptor
Study Guide e Plasma Membrane You should be able to write out the definitions to each of the following terms in your own words: plasma membrane fluid mosaic integral proteins peripheral proteins receptor
Structure of viruses
 Structure of viruses Lecture 4 Biology 3310/4310 Virology Spring 2018 In order to create something that functions properly - a container, a chair, a house - its essence has to be explored, for it should
Structure of viruses Lecture 4 Biology 3310/4310 Virology Spring 2018 In order to create something that functions properly - a container, a chair, a house - its essence has to be explored, for it should
Structure of viruses
 Structure of viruses Lecture 4 Biology 3310/4310 Virology Spring 2017 In order to create something that functions properly - a container, a chair, a house - its essence has to be explored, for it should
Structure of viruses Lecture 4 Biology 3310/4310 Virology Spring 2017 In order to create something that functions properly - a container, a chair, a house - its essence has to be explored, for it should
Plasma membrane structure and dynamics explored via a combined AFM/FCS approach
 Plasma membrane structure and dynamics explored via a combined AFM/FCS approach Salvatore Chiantia Molekulare Biophysik, Dept. Of Biology Humboldt-Universität zu Berlin Dresden nanoseminar, May 2013 Outline
Plasma membrane structure and dynamics explored via a combined AFM/FCS approach Salvatore Chiantia Molekulare Biophysik, Dept. Of Biology Humboldt-Universität zu Berlin Dresden nanoseminar, May 2013 Outline
GPU Accelerated Apps Momentum
 GPU Accelerated Apps Momentum Key codes are GPU Accelerated! Molecular Dynamics Abalone GPU only code ACEMD GPU only code AMBER CHARMM DL_POLY GROMACS HOOMD-Blue GPU only code LAMMPS NAMD Quantum Chemistry
GPU Accelerated Apps Momentum Key codes are GPU Accelerated! Molecular Dynamics Abalone GPU only code ACEMD GPU only code AMBER CHARMM DL_POLY GROMACS HOOMD-Blue GPU only code LAMMPS NAMD Quantum Chemistry
Patricia Fitzgerald-Bocarsly
 FLU Patricia Fitzgerald-Bocarsly October 23, 2008 Orthomyxoviruses Orthomyxo virus (ortho = true or correct ) Negative-sense RNA virus (complementary to mrna) Five different genera Influenza A, B, C Thogotovirus
FLU Patricia Fitzgerald-Bocarsly October 23, 2008 Orthomyxoviruses Orthomyxo virus (ortho = true or correct ) Negative-sense RNA virus (complementary to mrna) Five different genera Influenza A, B, C Thogotovirus
H 2 O. Liquid, solid, and vapor coexist in the same environment
 Water H 2 O Liquid, solid, and vapor coexist in the same environment WATER MOLECULES FORM HYDROGEN BONDS Water is a fundamental requirement for life, so it is important to understand the structural and
Water H 2 O Liquid, solid, and vapor coexist in the same environment WATER MOLECULES FORM HYDROGEN BONDS Water is a fundamental requirement for life, so it is important to understand the structural and
CALCIUM CASEINATE. What Is Casein?
 CALCIUM CASEINATE A high quality milk protein that is Calcium Rich, manufactured from fresh pasteurized skimmed milk through precipitation of casein followed by neutralization and natural drying, which
CALCIUM CASEINATE A high quality milk protein that is Calcium Rich, manufactured from fresh pasteurized skimmed milk through precipitation of casein followed by neutralization and natural drying, which
brought to you by and REFERENCES
 brought to you by www.thebacteriophages.org and www.phage.org REFERENCES 1. Butcher, S. J., T. Dokland, P. M. Ojala, D. H. Bamford, and S. D. Fuller. 1997. Intermediates in the assembly pathway of the
brought to you by www.thebacteriophages.org and www.phage.org REFERENCES 1. Butcher, S. J., T. Dokland, P. M. Ojala, D. H. Bamford, and S. D. Fuller. 1997. Intermediates in the assembly pathway of the
Week 5 Section. Junaid Malek, M.D.
 Week 5 Section Junaid Malek, M.D. HIV: Anatomy Membrane (partiallystolen from host cell) 2 Glycoproteins (proteins modified by added sugar) 2 copies of RNA Capsid HIV Genome Encodes: Structural Proteins
Week 5 Section Junaid Malek, M.D. HIV: Anatomy Membrane (partiallystolen from host cell) 2 Glycoproteins (proteins modified by added sugar) 2 copies of RNA Capsid HIV Genome Encodes: Structural Proteins
Viral Capsids and Envelopes: Structure and Function
 Viral Capsids and Envelopes: Structure and Function William Lucas, Harvard Medical School, Boston, Massachusetts, USA David M Knipe, Harvard Medical School, Boston, Massachusetts, USA Virus particles contain
Viral Capsids and Envelopes: Structure and Function William Lucas, Harvard Medical School, Boston, Massachusetts, USA David M Knipe, Harvard Medical School, Boston, Massachusetts, USA Virus particles contain
Phylogenetic Methods
 Phylogenetic Methods Multiple Sequence lignment Pairwise distance matrix lustering algorithms: NJ, UPM - guide trees Phylogenetic trees Nucleotide vs. amino acid sequences for phylogenies ) Nucleotides:
Phylogenetic Methods Multiple Sequence lignment Pairwise distance matrix lustering algorithms: NJ, UPM - guide trees Phylogenetic trees Nucleotide vs. amino acid sequences for phylogenies ) Nucleotides:
STRUCTURE, GENERAL CHARACTERISTICS AND REPRODUCTION OF VIRUSES
 STRUCTURE, GENERAL CHARACTERISTICS AND REPRODUCTION OF VIRUSES Introduction Viruses are noncellular genetic elements that use a living cell for their replication and have an extracellular state. Viruses
STRUCTURE, GENERAL CHARACTERISTICS AND REPRODUCTION OF VIRUSES Introduction Viruses are noncellular genetic elements that use a living cell for their replication and have an extracellular state. Viruses
Lectures 11 12: Fibrous & Membrane Proteins. Lecturer: Prof. Brigita Urbanc
 Lectures 11 12: Fibrous & Membrane Proteins Lecturer: Prof. Brigita Urbanc (brigita@drexel.edu) 1 FIBROUS PROTEINS: function: structural microfilaments & microtubules fibrils, hair, silk reinforce membranes
Lectures 11 12: Fibrous & Membrane Proteins Lecturer: Prof. Brigita Urbanc (brigita@drexel.edu) 1 FIBROUS PROTEINS: function: structural microfilaments & microtubules fibrils, hair, silk reinforce membranes
Supplementary Information A Hydrophobic Barrier Deep Within the Inner Pore of the TWIK-1 K2P Potassium Channel Aryal et al.
 Supplementary Information A Hydrophobic Barrier Deep Within the Inner Pore of the TWIK-1 K2P Potassium Channel Aryal et al. Supplementary Figure 1 TWIK-1 stability during MD simulations in a phospholipid
Supplementary Information A Hydrophobic Barrier Deep Within the Inner Pore of the TWIK-1 K2P Potassium Channel Aryal et al. Supplementary Figure 1 TWIK-1 stability during MD simulations in a phospholipid
Virology Introduction. Definitions. Introduction. Structure of virus. Virus transmission. Classification of virus. DNA Virus. RNA Virus. Treatment.
 DEVH Virology Introduction Definitions. Introduction. Structure of virus. Virus transmission. Classification of virus. DNA Virus. RNA Virus. Treatment. Definitions Virology: The science which study the
DEVH Virology Introduction Definitions. Introduction. Structure of virus. Virus transmission. Classification of virus. DNA Virus. RNA Virus. Treatment. Definitions Virology: The science which study the
Some living things are made of ONE cell, and are called. Other organisms are composed of many cells, and are called. (SEE PAGE 6)
 Section: 1.1 Question of the Day: Name: Review of Old Information: N/A New Information: We tend to only think of animals as living. However, there is a great diversity of organisms that we consider living
Section: 1.1 Question of the Day: Name: Review of Old Information: N/A New Information: We tend to only think of animals as living. However, there is a great diversity of organisms that we consider living
Influenza or flu is a
 Clinical and Research Area Infectious Diseases Influenza Virus Types A and B Influenza or flu is a respiratory illness that is caused by influenza viruses. Influenza viruses type A and type B cause seasonal
Clinical and Research Area Infectious Diseases Influenza Virus Types A and B Influenza or flu is a respiratory illness that is caused by influenza viruses. Influenza viruses type A and type B cause seasonal
Human Immunodeficiency Virus. Acquired Immune Deficiency Syndrome AIDS
 Human Immunodeficiency Virus Acquired Immune Deficiency Syndrome AIDS Sudden outbreak in USA of opportunistic infections and cancers in young men in 1981 Pneumocystis carinii pneumonia (PCP), Kaposi s
Human Immunodeficiency Virus Acquired Immune Deficiency Syndrome AIDS Sudden outbreak in USA of opportunistic infections and cancers in young men in 1981 Pneumocystis carinii pneumonia (PCP), Kaposi s
Ralf Wagner Paul-Ehrlich-Institut
 www.pei.de Other Assays for the Detection of Neuraminidase (NA)-Specific Antibodies Ralf Wagner Paul-Ehrlich-Institut Overview to presented assays Assay principle based on: Chemical substrates: Protein
www.pei.de Other Assays for the Detection of Neuraminidase (NA)-Specific Antibodies Ralf Wagner Paul-Ehrlich-Institut Overview to presented assays Assay principle based on: Chemical substrates: Protein
Petascale Bimolecular Simulation with NAMD on Titan, Blue Waters, and Summit
 Petascale Bimolecular Simulation with NAMD on Titan, Blue Waters, and Summit James Phillips Beckman Institute, University of Illinois http://www.ks.uiuc.edu/research/namd/ SC15 NVIDIA Biomedical Technology
Petascale Bimolecular Simulation with NAMD on Titan, Blue Waters, and Summit James Phillips Beckman Institute, University of Illinois http://www.ks.uiuc.edu/research/namd/ SC15 NVIDIA Biomedical Technology
Coarse grained simulations of Lipid Bilayer Membranes
 Coarse grained simulations of Lipid Bilayer Membranes P. B. Sunil Kumar Department of Physics IIT Madras, Chennai 600036 sunil@iitm.ac.in Atomistic MD: time scales ~ 10 ns length scales ~100 nm 2 To study
Coarse grained simulations of Lipid Bilayer Membranes P. B. Sunil Kumar Department of Physics IIT Madras, Chennai 600036 sunil@iitm.ac.in Atomistic MD: time scales ~ 10 ns length scales ~100 nm 2 To study
Research on Microscopic Pore Structure and Permeability of Air-Entrained Fly Ash Concrete Subjected to Freezing and Thawing Action
 4 th International Conference on the Durability of Concrete Structures 26 July 14 Purdue University, West Lafayette, IN, USA Research on Microscopic Pore Structure and Permeability of Air-Entrained Fly
4 th International Conference on the Durability of Concrete Structures 26 July 14 Purdue University, West Lafayette, IN, USA Research on Microscopic Pore Structure and Permeability of Air-Entrained Fly
Research on Extraction Process of Gallic Acid from Penthorum chinense Pursh by Aqueous Ethanol
 Green and Sustainable Chemistry, 2015, 5, 63-69 Published Online May 2015 in SciRes. http://www.scirp.org/journal/gsc http://dx.doi.org/10.4236/gsc.2015.52009 Research on Extraction Process of Gallic Acid
Green and Sustainable Chemistry, 2015, 5, 63-69 Published Online May 2015 in SciRes. http://www.scirp.org/journal/gsc http://dx.doi.org/10.4236/gsc.2015.52009 Research on Extraction Process of Gallic Acid
Structural biology of viruses
 Structural biology of viruses Biophysical Chemistry 1, Fall 2010 Coat proteins DNA/RNA packaging Reading assignment: Chap. 15 Virus particles self-assemble from coat monomers Virus Structure and Function
Structural biology of viruses Biophysical Chemistry 1, Fall 2010 Coat proteins DNA/RNA packaging Reading assignment: Chap. 15 Virus particles self-assemble from coat monomers Virus Structure and Function
Regulation of cell signaling cascades by influenza A virus
 The Second International Symposium on Optimization and Systems Biology (OSB 08) Lijiang, China, October 31 November 3, 2008 Copyright 2008 ORSC & APORC, pp. 389 394 Regulation of cell signaling cascades
The Second International Symposium on Optimization and Systems Biology (OSB 08) Lijiang, China, October 31 November 3, 2008 Copyright 2008 ORSC & APORC, pp. 389 394 Regulation of cell signaling cascades
Severe Acute Respiratory Syndrome (SARS) Coronavirus
 Severe Acute Respiratory Syndrome (SARS) Coronavirus Coronaviruses Coronaviruses are single stranded enveloped RNA viruses that have a helical geometry. Coronaviruses are the largest of RNA viruses with
Severe Acute Respiratory Syndrome (SARS) Coronavirus Coronaviruses Coronaviruses are single stranded enveloped RNA viruses that have a helical geometry. Coronaviruses are the largest of RNA viruses with
Lipid raft-a gateway for passing through the cell membrane for pathogens
 16 3 2004 6 Chinese Bulletin of Life Sciences Vol. 16, No. 3 Jun., 2004 10040374(2004)03014404 200031 / GPI (GPI) Q241 Q257 R37 A Lipid rafta gateway for passing through the cell membrane for pathogens
16 3 2004 6 Chinese Bulletin of Life Sciences Vol. 16, No. 3 Jun., 2004 10040374(2004)03014404 200031 / GPI (GPI) Q241 Q257 R37 A Lipid rafta gateway for passing through the cell membrane for pathogens
2.1 VIRUSES. 2.1 Learning Goals
 2.1 VIRUSES 2.1 Learning Goals To understand the structure, function, and how Viruses replicate To understand the difference between Viruses to Prokaryotes and Eukaryotes; namely that viruses are not classified
2.1 VIRUSES 2.1 Learning Goals To understand the structure, function, and how Viruses replicate To understand the difference between Viruses to Prokaryotes and Eukaryotes; namely that viruses are not classified
Coronaviruses. Virion. Genome. Genes and proteins. Viruses and hosts. Diseases. Distinctive characteristics
 Coronaviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Spherical enveloped particles studded with clubbed spikes Diameter 120-160 nm Coiled helical
Coronaviruses Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Virion Spherical enveloped particles studded with clubbed spikes Diameter 120-160 nm Coiled helical
Dr. Ahmed K. Ali Attachment and entry of viruses into cells
 Lec. 6 Dr. Ahmed K. Ali Attachment and entry of viruses into cells The aim of a virus is to replicate itself, and in order to achieve this aim it needs to enter a host cell, make copies of itself and
Lec. 6 Dr. Ahmed K. Ali Attachment and entry of viruses into cells The aim of a virus is to replicate itself, and in order to achieve this aim it needs to enter a host cell, make copies of itself and
Strategies for containing an emerging influenza pandemic in South East Asia 1
 Strategies for containing an emerging influenza pandemic in South East Asia 1 Modeling pandemic spread and possible control plans of avian flu H5N1 BBSI, Nicole Kennerly, Shlomo Ta asan 1 Nature. 2005
Strategies for containing an emerging influenza pandemic in South East Asia 1 Modeling pandemic spread and possible control plans of avian flu H5N1 BBSI, Nicole Kennerly, Shlomo Ta asan 1 Nature. 2005
Where Health Care Meets Policy. with Dr. Mike Magee
 Where Health Care Meets Policy with Dr. Mike Magee The Threat of Bird Flu Understanding Bird Flu and the Influenza Virus 3 types of the influenza virus: A, B and C reflect differences in the M protein
Where Health Care Meets Policy with Dr. Mike Magee The Threat of Bird Flu Understanding Bird Flu and the Influenza Virus 3 types of the influenza virus: A, B and C reflect differences in the M protein
Human Genome Complexity, Viruses & Genetic Variability
 Human Genome Complexity, Viruses & Genetic Variability (Learning Objectives) Learn the types of DNA sequences present in the Human Genome other than genes coding for functional proteins. Review what you
Human Genome Complexity, Viruses & Genetic Variability (Learning Objectives) Learn the types of DNA sequences present in the Human Genome other than genes coding for functional proteins. Review what you
LESSON 1.4 WORKBOOK. Viral sizes and structures. Workbook Lesson 1.4
 Eukaryotes organisms that contain a membrane bound nucleus and organelles. Prokaryotes organisms that lack a nucleus or other membrane-bound organelles. Viruses small, non-cellular (lacking a cell), infectious
Eukaryotes organisms that contain a membrane bound nucleus and organelles. Prokaryotes organisms that lack a nucleus or other membrane-bound organelles. Viruses small, non-cellular (lacking a cell), infectious
How to Build the Management Mode for the Gymnasiums in Ordinary Universities in China
 Journal of Sports Science 4 (2016) 226-231 doi: 10.17265/2332-7839/2016.04.006 D DAVID PUBLISHING How to Build the Management Mode for the Gymnasiums in in China Fengquan Yu Sports Sociology and Humanities,
Journal of Sports Science 4 (2016) 226-231 doi: 10.17265/2332-7839/2016.04.006 D DAVID PUBLISHING How to Build the Management Mode for the Gymnasiums in in China Fengquan Yu Sports Sociology and Humanities,
LESSON 1.4 WORKBOOK. Viral structures. Just how small are viruses? Workbook Lesson 1.4 1
 Eukaryotes- organisms that contain a membrane bound nucleus and organelles Prokaryotes- organisms that lack a nucleus or other membrane-bound organelles Viruses-small acellular (lacking a cell) infectious
Eukaryotes- organisms that contain a membrane bound nucleus and organelles Prokaryotes- organisms that lack a nucleus or other membrane-bound organelles Viruses-small acellular (lacking a cell) infectious
Flu, Avian Flu and emerging aspects (H1N1 resistance)
 EU-CIS Seminar New trends in Infectious Diseases 26 28 November 2008 / Lyon, France Flu, Avian Flu and emerging aspects (H1N1 resistance) Pr. Florence MORFIN FRE 3011 Université Lyon 1 - CNRS Laboratory
EU-CIS Seminar New trends in Infectious Diseases 26 28 November 2008 / Lyon, France Flu, Avian Flu and emerging aspects (H1N1 resistance) Pr. Florence MORFIN FRE 3011 Université Lyon 1 - CNRS Laboratory
Improved Processing Research on Arc Tooth Cylindrical Gear
 International Journal of Environmental Monitoring and Analysis 2017; 5(3): 91-95 http://www.sciencepublishinggroup.com/j/ijema doi: 10.11648/j.ijema.20170503.14 ISSN: 2328-7659 (Print); ISSN: 2328-7667
International Journal of Environmental Monitoring and Analysis 2017; 5(3): 91-95 http://www.sciencepublishinggroup.com/j/ijema doi: 10.11648/j.ijema.20170503.14 ISSN: 2328-7659 (Print); ISSN: 2328-7667
Supplementary materials
 Supplementary materials Chemical library from ChemBridge 50,240 structurally diverse small molecule compounds dissolved in DMSO Hits Controls: No virus added μ Primary screening at 20 g/ml of compounds
Supplementary materials Chemical library from ChemBridge 50,240 structurally diverse small molecule compounds dissolved in DMSO Hits Controls: No virus added μ Primary screening at 20 g/ml of compounds
Penetration of Gold Nanoparticle through Human Skin: Unraveling Its Mechanisms at the Molecular Scale
 Penetration of Gold Nanoparticle through Human Skin: Unraveling Its Mechanisms at the Molecular Scale Rakesh Gupta and Beena Rai* TATA Research Development & Design Centre, TCS Innovation labs, Pune India,
Penetration of Gold Nanoparticle through Human Skin: Unraveling Its Mechanisms at the Molecular Scale Rakesh Gupta and Beena Rai* TATA Research Development & Design Centre, TCS Innovation labs, Pune India,
Inhibition of Fibril Formation of Beta-Amyloid Peptides
 John von Neumann Institute for Computing Inhibition of Fibril Formation of Beta-Amyloid Peptides N. S. Lam, M. Kouza, H. Zung, M. S. Li published in From Computational Biophysics to Systems Biology (CBSB08),
John von Neumann Institute for Computing Inhibition of Fibril Formation of Beta-Amyloid Peptides N. S. Lam, M. Kouza, H. Zung, M. S. Li published in From Computational Biophysics to Systems Biology (CBSB08),
Influenza Viruses A Review
 Influenza Viruses A Review AVIAN INFLUENZA: INTERSECTORAL COLLABORATION Larnaca, Cyprus 20 22 July 2009 Kate Glynn Scientific and Technical Department, OIE Influenza Viruses C. Goldsmith,1981 Influenza
Influenza Viruses A Review AVIAN INFLUENZA: INTERSECTORAL COLLABORATION Larnaca, Cyprus 20 22 July 2009 Kate Glynn Scientific and Technical Department, OIE Influenza Viruses C. Goldsmith,1981 Influenza
Quantifying Lipid Contents in Enveloped Virus Particles with Plasmonic Nanoparticles
 Quantifying Lipid Contents in Enveloped Virus Particles with Plasmonic Nanoparticles Amin Feizpour Reinhard Lab Department of Chemistry and the Photonics Center, Boston University, Boston, MA May 2014
Quantifying Lipid Contents in Enveloped Virus Particles with Plasmonic Nanoparticles Amin Feizpour Reinhard Lab Department of Chemistry and the Photonics Center, Boston University, Boston, MA May 2014
11/15/2011. Outline. Structural Features and Characteristics. The Good the Bad and the Ugly. Viral Genomes. Structural Features and Characteristics
 Chapter 19 - Viruses Outline I. Viruses A. Structure of viruses B. Common Characteristics of Viruses C. Viral replication D. HIV II. Prions The Good the Bad and the Ugly Viruses fit into the bad category
Chapter 19 - Viruses Outline I. Viruses A. Structure of viruses B. Common Characteristics of Viruses C. Viral replication D. HIV II. Prions The Good the Bad and the Ugly Viruses fit into the bad category
hemagglutinin and the neuraminidase genes (RNA/recombinant viruses/polyacrylamide gel electrophoresis/genetics)
 Proc. Natl. Acad. Sci. USA Vol. 73, No. 6, pp. 242-246, June 976 Microbiology Mapping of the influenza virus genome: Identification of the hemagglutinin and the neuraminidase genes (RNA/recombinant viruses/polyacrylamide
Proc. Natl. Acad. Sci. USA Vol. 73, No. 6, pp. 242-246, June 976 Microbiology Mapping of the influenza virus genome: Identification of the hemagglutinin and the neuraminidase genes (RNA/recombinant viruses/polyacrylamide
Xianren Zhang ( 张现仁 )
 The interaction between nanoparticles and membranes: from cytotoxicity to drug delivery Xianren Zhang ( 张现仁 ) zhangxr@mail.buct.edu.cn State Key Laboratory of Organic-Inorganic Composites, Beijing University
The interaction between nanoparticles and membranes: from cytotoxicity to drug delivery Xianren Zhang ( 张现仁 ) zhangxr@mail.buct.edu.cn State Key Laboratory of Organic-Inorganic Composites, Beijing University
Envelope e_ :------, Envelope glycoproteins e_ ~ Single-stranded RNA ----, Nucleocapsid
 Virus Antib(tdies Your Expertise, Our Antibodies, Accelerated Discovery. Envelope e_---------:------, Envelope glycoproteins e_-------1~ Single-stranded RNA ----, Nucleocapsid First identified in 1989,
Virus Antib(tdies Your Expertise, Our Antibodies, Accelerated Discovery. Envelope e_---------:------, Envelope glycoproteins e_-------1~ Single-stranded RNA ----, Nucleocapsid First identified in 1989,
