CIC EDIZIONI INTERNAZIONALI
|
|
|
- Ethelbert Williams
- 5 years ago
- Views:
Transcription
1 Prenatal diagnosis of genomic disorders and chromosome abnormalities using array-based comparative genomic hybridization Francesca Gullotta 1 Michela Biancolella 1 Elena Costa 1 Isabella Colapietro 1 Anna Maria Nardone 1 Paolo Molinaro 1 Adalgisa Pietropolli 2 Marianovella Narcisi 2 Cristiana Di Rosa 1 Giuseppe Novelli 1 1 Dipartimento di Biopatologia e Diagnostica per Immagini, Università di Roma Tor Vergata and Azienda Ospedaliera Universitaria Policlinico Tor Vergata, Roma, Italy 2 Sezione di Clinica Ginecologica e Ostetrica, A.O.U. Policlinico Tor Vergata, Roma, Italy Reprint requests to: Prof. Giuseppe Novelli Laboratorio di Genetica Medica Azienda Ospedaliera Universitaria Policlinico Tor Vergata, Roma Viale Oxford, Roma Fax: +39/06/ E.mail: novelli@med.uniroma2.it Summary Cytogenetic analysis is a crucial tool of prenatal diagnosis. The ability to rapidly detect aneuploidy and identify small structural abnormalities of foetal chromosomes has been greatly improved by the use of molecular cytogenetic technologies. Microarray-based Comparative Genomic Hybridization (acgh) has been recently employed in postnatal diagnosis of cryptic chromosomal aberrations, but use in prenatal diagnosis is still limited. We set-up a diagnostic protocol which uses acgh technology on genomic DNA isolated from uncultured chorionic villus sampled at week s gestation. We used a commercially targeted microarray (MDTelArray, Technogenetics Srl Bouty Group, Sesto S. Giovanni, Milan, Italy) constituted by 167 genomic clones corresponding to 34 critical regions frequently involved in microdeletions and microduplications and 126 subtelomeric clones. Array validation has been carried-out via retrospective analysis of DNA isolated from a series of cytogenetically normal chorionic villus samples (CVS) and of DNA isolated from cytogenetically abnormal cultured amniocytes, CVS or peripheral blood. A pilot prospective study was undertaken analyzing 25 CVS obtained from foetuses at risk for chromosomal aberrations. acgh results both for retrospective and prospective studies were in agreement with data obtained using classical cytogenetic analysis, and/or FISH analysis or DNA testing. Although these preliminary data support the usefulness of acgh in prenatal diagnosis, further prospective studies are required to verify the feasibility of introducing this technique as part of the diagnostic armamentarium for identify affected foetuses. KEY WORDS: prenatal diagnosis, array-based comparative genomic hybridization (acgh), aneuploidy, cryptic chromosomal aberrations, chorionic villus samples (CVS). Introduction A potentially lethal or handicapping major defect occurs in 2-3% of liveborn infants (1). Congenital malformations have become the main cause of infant mortality during the first years of life (2, 3). Approximately 10-15% of stillborn and liveborn infants with malformations have chromosomal imbalances (4, 5). Since the development of chromosome banding techniques in the late 1960 s (6), microscopic karyotype analysis has been applied to prenatal testing and is still today considered the gold standard for prenatal diagnosis. This procedure results to be highly reliable for identifying chromosome copy number abnormalities (aneuploidy) and large structural rearrangements in foetal cells obtained invasively by either amniocentesis or chorionic villus sampling (CVS). However, even if this procedure results highly reliable, a number of limitations frequently occur. The resolution appear to be inadequate to detect deletions or duplications <10 Mb. In addition, the technique requires cells culture and a long time for definitive results generating frequently anxiety for parents during a pregnancy. Studies have demonstrated the ability of molecular techniques to detect aneuploidy and submicroscopic chromosomal anomalies within 24 hrs. These include, fluorescence in situ hybridization (FISH), quantitative fluorescence polymerase chain reaction (QF-PCR), and multiplex ligation-dependent probe amplification (MLPA) (7-9). However, all these techniques seems to be inadequate to perform a genomewide screening. Recent studies have demonstrated the ability of acgh to detect submicroscopic chromosomal anomalies in individuals with learning and developmental disability providing evidence for a genomewide screening strategy in detecting DNA copy number imbalances in a rapid and less labour-intensive manner (10, 11). This technique is similar in principle to conventional metaphase CGH (12, 13), but uses arrayed DNA sequences instead of metaphase 16 Journal of Prenatal Medicine 2007; 1 (1): 16-22
2 Prenatal diagnosis of genomic disorders and chromosome abnormalities using array-based comparative genomic hybridization chromosomes as targets for hybridization, thus providing a direct link between detected aberrations and the physical and genetic maps of the human genome. Patient and reference genomic DNAs labelled with two different fluorochromes are co-hybridized to an array of mapped DNA fragments immobilized on slides (12, 14). The genomic resolution depends on the physical distance between two clones and the sizes of individual clones. This technique is able to detect, in a single experiment, any dosage imbalances including aneuploidies, deletions or duplications, but it cannot detect balanced rearrangements such as reciprocal and Robertsonian translocations or inversions. acgh shows a number of advantages compared to conventional techniques in terms of clinical practice and cost implications. Here we present our experience in validation of an innovative acgh (MDTelArray, Technogenetics Srl Bouty Group, Sesto S. Giovanni, Milan, Italy) for prenatal diagnosis. For this purpose, we established an original protocol based on genomic DNA extracted from CVS during the first trimester of gestation. MDTelArray was arranged by 167 genomic clones corresponding to 34 critical genomic regions frequently involved in microdeletions and microduplications and 126 subtelomeric clones. The acgh was tested in a series of six retrospective unbalanced cases, and 16 normal DNA samples. In addition, 25 prospective samples of chorionic villi obtained from foetuses at high-risk for chromosomal aberrations were studied. Methods Retrospective series A series of 6 pathological DNA obtained from CVS or cultured amniotic cells of samples affected foetuses (n. 3 respectively, trisomy 21, Klinefelter syndrome, and Duchenne Type Muscular Dystrophy, DMD) or from affected children (n. 3 respectively, Turner syndrome, Prader-Willi syndrome and Smith-Magenis syndrome) and 16 normal DNA samples (CVS) obtained from unaffected foetuses, were studied. All retrospective series of samples were investigated by routine cytogenetics analysis and/or FISH, or DNA testing (DMD). Prospective series Twenty-five CVS obtained from foetuses at risk for chromosomal aberrations due to advanced maternal age, were obtained by the subjects obstetricians using their standard clinical procedures. All CVS were prepared for standard G-band karyotype for clinical laboratory protocols and, in some cases, for the molecular genetic analysis as well. The remainder of the foetal sample was collected for acgh analysis. For each experiment, we used at least 2 µg of DNA extracted both from fresh chorionic villus samples and corionic villus samples preserved at -20 C even for as long as one month. Test DNA was extracted using a Wizard Genomic DNA Purification Kit (Promega, Madison, WI), according to the manufacturer s protocols. Reference genomic DNA was derived from peripheral blood of phenotypically normal male and female control individuals (Promega, Madison, WI). Microarray constitution, hybridization and data analysis acgh analysis was performed using a targeted microarray (MDTelArray, Technogenetics Srl Bouty Group, Sesto S. Giovanni, Milan, Italy) constituted by 167 genomic clones corresponding to 34 critical regions frequently involved in microdeletions and microduplications (Table I) and 126 subtelomeric clones. Each BAC clone was spotted in triplicate and positive (pool of human BAC clone) and negative (rice DNA) control clones were also printed. This array enables the simultaneous analysis of all the regions above via a single hybridization, therefore saving both time and sample material. The array s high resolution, allows for more reliable and precise data. Moreover, two different hybridization areas were spotted on the array, consenting to perform the experiment in dye-reversal (Fig. 1). This permits to achieve high quality data and provides additional confirmation of true copy number alterations (15, 16). In dye-reversal technique, for each patient sample, two experiments were performed with reversal of the dye labels for the control and the test samples (17), followed by integration of the data from both dyereversed hybridizations to determine interferences for each case (Fig. 2). After purification (Zymo Research s Clean and Concentrator -25, Orange, CA), an equal amount (1 µg) of both test and reference DNA was labelled with Cy3-dCTP and Cy5-dCTP (Amersham Biosciences, Little Chalfont, UK), respectively, by random priming (Bioprime Array CGH Genomic Labelling Module, Invitrogen, Carlsbad, CA), purified with CyScribe GFX Purification Kit (Amersham Biosciences, Little Chalfont, UK), precipitated and hybridized following the microarray manufacturer s protocols (Technogenetics Srl Bouty Group, Sesto S. Giovanni, Milan, Italy). A dye-reversal experiment was performed for each patient sample. Images were acquired using GenePix 4000B dual-laser scanner and GenePix Pro 6.0 software (Axon Instruments, Sunnyvale, CA). The average ratios of the six spots for each patient were analyzed and plotted using BlueFuse for Microarrays 3.3 (5164) software (BlueGenome Ltd, Cambridge, UK). Increases (gains) and decreases (losses) in DNA sequence copy number were defined by test/reference ratios above 1.2 and below 0.8, respectively, based upon previous reports in which 1.2 (log =0.3) and 0.8 (log =-0.3) were selected as cut-off values based on experimental results of normal variation observed when two normal reference DNAs were co-hybridized to genomic microarrays (18, 10). To ensure detection of any sex chromosomes abnormalities if the sex of the foetus was different from the sex of the DNA reference, during array validation chromosomes mismatch experiments, using male vs. female DNA of cytogenetically normal individuals, were performed. Journal of Prenatal Medicine 2007; 1 (1):
3 F. Gullotta et al. Table I - Microdeletion syndromes represented in the MDTelArray. Monosomy 1p36 Syndrome [(OMIM #607872) (1p36)] Van der Woude Syndrome [(OMIM #119300) (1q32-q41)] Nephronophthisis 1 [(OMIM #256100) (2q13)] Brachydactyly-Mental Retardation Syndrome [(OMIM %600430) (2q37)] Wolf-Hirschhorn Syndrome [(OMIM #194190) (4p16.3)] Cri-du-Chat Syndrome [(OMIM #123450) (5p15.2)] Adenomatous Polyposis of the Colon [(OMIM ) (5q21-q22)] Sotos Syndrome [(OMIM #117550) (5q35)] Saethre-Chotzen Syndrome [(OMIM #101400) (7p21.1)] Williams-Beuren Syndrome [(OMIM #194050) (7q11.2)] Kallmann Syndrome 2 [(OMIM #147950) (8p11.2-p11.1)] Langer-Giedion Syndrome [(OMIM #150230) (8q24.11-q24.13)] Monosomy 9p Syndrome [(OMIM #158170)] HDR Syndrome [(OMIM #146255) (10p15)] DiGeorge Syndrome/Velocardiofacial Syndrome Spectrum of Malformation 2 [(OMIM %601362) (10p14-10p13)] Wagr Syndrome [(OMIM #194072) (11p13)] Potocki-Shaffer Syndrome [(OMIM #601224) (11p11.2)] Prader-Willi Syndrome / Angelman Syndrome [(OMIM #176270/#105830) (15q11-q13)] ATR-16 Syndrome [(OMIM #141750) (16pter-p13.3)] Miller-Dieker Lissencephaly Syndrome [(OMIM #247200) (17p13.3)] Charcot-Marie-Tooth Disease Neuropathy, Type 1A [(OMIM #118220) (17p11.2)] Neuropathy, Hereditary, with Liability to Pressure Palsies [(OMIM #162500) (17p11.2)] Smith-Magenis Syndrome [(OMIM #182290) (17p11.2)] Neurofibromatosis Familial Spinal [(OMIM #162210) (17q11.2)] Alagille Syndrome 1 [(OMIM #118450) (20p12)] DiGeorge Syndrome [(OMIM #188400) (22q11.2)] Neurofibromatosis, Type II [(OMIM #101000) (22q12.2)] Short Stature, Idiopathic, Autosomal [(OMIM #604271) (Xpter-p22.32, Ypter-p11.2)] Steroid Sulfatase Deficiency [(OMIM ) (Xp22.32)] Kallmann Syndrome 1 [(OMIM ) (Xp22.3)] Muscular Dystrophy, Duchenne Type [(OMIM #310200) (Xp21.2)] ATR-X Syndrome [(OMIM #301040) (Xq13)] Pelizaeus-Merzbacher Disease [(OMIM #312080) (Xq22)] XX Male Syndrome [(OMIM )] Figure 1 - MDTelArray imaging showing two different hybridization areas to perform the experiment in dye-reversal. 18 Journal of Prenatal Medicine 2007; 1 (1): 16-22
4 Prenatal diagnosis of genomic disorders and chromosome abnormalities using array-based comparative genomic hybridization A Figure 2 - An illustrative example of a microdeletion detected by MDTelArray using a dye-reversal experiment. For each patient sample, two experiments were performed with reversal of the dye labelling for the test and the reference samples (A and B). In A test (patient) DNA was labelled in Cy3 (green) and reference (control) DNA in Cy5 (red). In B, the same test DNA was labelled in Cy5 (red) and the reference DNA in Cy3 (green). Results and discussion Initially, we evaluated if the DNA extraction method influenced the acgh results. For this purpose, we compared hybridization results obtained using aliquots of the same sample, immediately treated or stored at -20 C for up to one month. No significant differences were scored between samples processed immediately respect to those stored. The acgh analytical validity was carried-out via retrospective analysis of DNA isolated from a series of cytogenetically normal CVS and cytogenetically abnormal DNA obtained from cultured amniocytes, CVS or peripheral blood. Chromosomal abnormalities included aneuploidies (trisomy 21, 47,XXY, 45,X) and microdeletions [del (X)(p21.2), del(15)(q11.2-q13) and del(17)(p11.2)] respectively associated to DMD, Prader-Willi syndrome, and Smith-Magenis syndrome. achg analysis of the sample with trisomy 21 had a single gain in copy number of clones on chromosome 21 that was represented by 3 BAC clones with a ratio = 1.38 ± DNA sample of kariotype 47,XXY also exhibited single copy gain of clones on chromosome Y as represented by 7 BAC clones with a ratio = 1.29 ± 0.04 (using a reference female DNA). acgh analysis of 45,X DNA sample showed a single copy number loss of all BAC clones on chromosome X (ratio = 0.77 ± 0.012) (using a reference female DNA). DMD DNA sample had a copy number loss on chromosome X as showed by 3 BAC clones with a ratio = 0.58 ± In Prader-Willi sample was evident the loss of copy number of 10 BAC clones (ratio = 0.71 ± 0.02) on chromosome 15. acgh analysis of Smith- Magenis DNA showed a single copy loss of 5 BAC clones on chromosome 17 with a ratio = 0.78 ± The acgh results were in agreement with those obtained by the classical cytogenetic analysis, FISH analysis and DNA testing achieving an analytical sensitivity and specificity of 100% in the examined samples. After array validation, 25 uncultured chorionic villus B samples obtained from foetuses at risk for chromosomal aberrations were analyzed. 24 out of 25 examined samples did not show any chromosomal abnormalities in the analyzed regions (Fig. 3). In one case, with abnormal ultrasound findings, a trisomy 18 was detected (Fig. 4). achg analysis of the sample had a single gain in copy number of clones on chromosome 18 that was represented by 7 BAC clones with a ratio = 1.35 ± The presence of this aneuploidy was confirmed by karyotype analysis. Cytogenetic analysis is an important component of prenatal diagnosis allowing the identification of aneuploidy and unbalanced structural rearrangements in foetuses including and/or one of the following: advanced maternal age, abnormal serum screening results, a high-risk for cytogenetic abnormalaties based on family history, abnormal ultrasound findings. Conventional karyotype analysis of banded chromosomes is still considered the gold standard method in prenatal diagnosis. Although highly reliable for identifying aneuploidies as well as large chromosomal rearrangements, its resolution size (5-10 Mb) significantly limits this technique. (reviewed in 19). In addition, the chromosomal origin of extra structurally abnormal chromosomes (ESACs) could not be revealed and microscopic rearrangements of subtelomeric regions cannot be detected (20, 21). Furthermore, the average time to receive the analysis results (approximately 14 days) represents another significant limitation. Molecular cytogenetic techniques were developed to overcome the resolution limitations of karyotype analysis, as well as reducing the average reporting time for a rapid screening of common aneuploidies (i.e. trisomy 13, 18 and 21, triploidy and aneuploidy of the sex chromosomes) (FISH and QF- PCR analyses) and MLPA (7-9). Although these techniques do not provide a genome wide screen, they are not likely to replace conventional karyotyping. In order to investigate structural chromosomal rearrangements associated with copy number variation in a genome wide scale, comparative genomic hybridization (CGH) has Journal of Prenatal Medicine 2007; 1 (1):
5 F. Gullotta et al. Figure 3 - MDTelArray linear plot showing normal results for all chromosomes. The thin green line represents the fluorescence intensity ratios between unaffected foetus (labbelled with Cy3) and control (labelled with Cy5) (green arrow). The thin red line represents the fluorescence intensity ratios obtained from a second hybridization in which the dyes have been reversed (control Cy3: unaffected foetus Cy5). A B C D Figure 4 - (A) MDTelArray linear plot showing the trisomy of chromosome 18 (affected foetus DNA Cy3: control Cy5). Duplication is evident as a deviation from a log 2 ratio of 0. (B) Chromosome 18 analysis showing fluorescence intensity ratios between affected foetus DNA (labbelled with Cy3) and control DNA (labelled with Cy5). (C) Partial of linear plot of the reverse experiment showing fluorescence intensity ratios between control DNA (labbelled with Cy3) and affected foetus DNA (labelled with Cy5). (D) Chromosome 18 analysis showing fluorescence intensity ratios between control DNA (labbelled with Cy3) and affected foetus DNA (labelled with Cy5). 20 Journal of Prenatal Medicine 2007; 1 (1): 16-22
6 Prenatal diagnosis of genomic disorders and chromosome abnormalities using array-based comparative genomic hybridization been introduced to assist prenatal diagnosis (22-24). However, CGH still requires metaphase chromosomes as targets for hybridization, limiting its resolution to approximately 5 Mb, like microscopic karyotype analysis (25). Microarray-based comparative genomic hybridization (acgh) is a recently developed technology that evolved from standard CGH on metaphase spreads. acgh uses arrayed DNA sequences instead of metaphase chromosomes as target of hybridization, thus providing a direct link between detected aberrations and the physical and genetic maps of the human genome. Its resolution is limited only by the size of the target and the density of these clones. This technique is able to detect any dosage imbalances including aneuploidies, deletions or duplications, but it can not detect balanced rearrangements such as reciprocal and Robertsonian translocations or inversions. The primary advantage of acgh over conventional cytogenetics and FISH analysis is its ability to detect DNA copy number changes simultaneously at multiple discrete loci in a genome. Moreover, acgh offers rapid, high throughput analysis on minimal amounts of DNA, two prerequisites for any platforms applied to prenatal diagnosis. In fact, its utility to identify chromosomal imbalances in prenatal samples has been recently reported (26-29). However, whole-genome arrays could generate data that are difficult to interpret and that are subject to multiple FISH verifications per patient. Furthermore, the array content must be well considered before its application in prenatal testing. Copy number variations (CNVs) of genome regions not previously associated to chromosomal phenotype may be difficult to interpret. In addition, recent studies have shown that familiar DNA gains and losses across the genome are numerous and common (30, 31). CNVs may have no clinical significance if equally detected in phenotipically normal and abnormal individuals or have clinical importance, especially if identified as de novo in a person with abnormal phenotype. The remaining CN- Vs may have some clinical implications, but it will be necessary to observe the same alterations in more abnormal cases to understand their clinical relevance (31-33). Depending on the resolution and representation of the genome of the microarray, CNVs may be frequently found, thus complicating the interpretation of results. Thus, microarray design is very important to obtain high quality results. In contrast to whole-genome acgh, targeted arrays constructed with clones mapping within chromosome regions frequently involved in genomic disorders or subchromosomal anomalies have been developed (34, 15, 35). These arrays have been used in the clinical diagnosis of chromosome abnormalities in children with birth defects, mental retardation, or developmental disabilities (16, 36, 37), however prenatal uses are limited at present (38). We present a preliminary study on validation and utilization in prenatal diagnosis, of a targeted array able to reveal specifically 34 critical chromosomal regions involved in microdeletions and/or microduplications that cause well known syndromes. The array contained also 41 subtelomeric regions target of mental retardation and learning and developmental disability. All acgh results agreed with those obtained by karyotyping or FISH analysis or DNA testing, detecting both aneuploidies and chromosomal microdeletions. Even if we analyzed only a restricted number of samples, our results confirmed the possibility to use this kind of acgh in prenatal diagnosis, minimalizing the difficulties of good interpretation of results. Furthermore, applying acgh technology on genomic DNA isolated from uncultured chorionic villus samples we obtained the same results as obtained by DNA isolated from cultured cells. Thus, the time taken to report results back to patients could be significantly reduced. We are confident that this targeted array improves the analysis of segmental aneuploidy in uterus and suggests that it may prove feasible to introduce acgh as part of the diagnostic armamentarium for detecting chromosomal rearrangements which would not be otherwise investigated in routine prenatal cytogenetic analysis. Acknowledgements We thank Dr. G. Galluzzi for the gift of DMD DNA sample, Dr. Antonio Novelli for his help and suggestions, the Technogenetics Srl (Bouty Group, Sesto S. Giovanni, Milan, Italy) for the opportunity to use the MDTelArray. This work was supported in part by a grant from the Italian Ministry of Health and Italian Ministry of University and Research. References 11. Kalter H, Warkany J. Medical progress. Congenital malformations: etiologic factors and their role in prevention (first of two parts). N Engl J Med 1983;308: Kalter H, Warkany J. Congenital malformations (second of two parts). N Engl J Med 1983;308: De Galan-Roosen AE, Kuijpers JC, Meershoek AP, van Velzen D. Contribution of congenital malformations to perinatal mortality. A 10 years prospective regional study in The Netherlands. Eur J Obstet Gynecol Reprod Biol 1998; 80: Nelson K, Holmes LB. Malformations due to presumed spontaneous mutations in newborn infants. N Engl J Med 1989;320: Stevenson RE. The genetic basis of human anomalies. In: Stevenson RE, Hall JG, Goodman RM, eds. Human malformations and related anomalies, Vol 1. New York: Oxford University Press 1993: Caspersson T, Farber S, Foley G.E et al. Chemical differentiation along metaphase chromosomes. Exp Cell Res 1968;49(1): Klinger K, Landes G, Shook D et al. Rapid detection of chromosome aneuploidies in uncultured amniocytes by using fluorescence in situ hybridization (FISH). Am J Hum Genet 1992;51(1): Adinolfi M, Sherlock J. Prenatal detection of chromosome disorders by QF-PCR. Lancet. 2001;358(9287): Slater HR, Bruno DL, Ren H, Pertile M, Schouten JP, Choo KH. Rapid, high throughput prenatal detection of aneuploidy using a novel quantitative method (MLPA). J Med Genet 2003;40: Vissers LE, de Vries BB, Osoegawa K et al. Array-based comparative genomic hybridization for the genomewide detection of submicroscopic chromosomal abnormalities. Am J Hum Genet 2003;73(6): Shaw-Smith C, Redon R, Rickman L et al. Microarray Journal of Prenatal Medicine 2007; 1 (1):
7 F. Gullotta et al. based comparative genomic hybridisation (array-cgh) detects submicroscopic chromosomal deletions and duplications in patients with learning disability/mental retardation and dysmorphic features. Am J Hum Genet 2003;73(6): Pinkel D, Segraves R, Sudar D et al. High resolution analysis of DNA copy number variation using comparative genomic hybridization to microarrays. Nat Genet 1998;20: Ishkanian AS, Malloff CA, Watson SK et al. A tiling resolution DNA microarray with complete coverage of the human genome. Nat Genet 2004;36: Snijders AM, Nowak N, Segraves R et al. Assembly of microarrays for genome-wide measurement of DNA copy number. Nat Genet 2001;29: Yu W, Ballif BC, Kashork CD et al. Development of a comparative genomic hybridization microarray and demonstration of its utility with 25 well-characterized 1p36 deletions. Hum Mol Genet 2003;12(17): Bejjani BA, Theisen AP, Ballif BC, Shaffer LG. Array-based comparative genomic hybridization in clinical diagnosis. Expert Rev Mol Diagn 2005;5(3): Wessendorf S, Fritz B, Wrobel G et al. Automated screening for genomic imbalances using matrix-based comparative genomic hybridization. Lab Invest 2002;82(1): Hui AB, Lo KW, Yin XL, Poon WS, Ng HK. Detection of multiple gene amplifications in glioblastoma multiforme using array-based comparative genomic hybridization. Lab Invest 2001;81(5): Shaffer LG, Bejjani BA. A cytogeneticist's perspective on genomic microarrays. Hum Reprod Update 2004;10(3): Flint J, Knight S. The use of telomere probes to investigate submicroscopic rearrangements associated with mental retardation. Curr Opin Genet Dev 2003;13(3): Knight, Flint J. The use of subtelomeric probes to study mental retardation. Methods Cell Biol 2004;75: Yu LC, Moore DH 2nd, Magrane G et al. Objective aneuploidy detection for fetal and neonatal screening using comparative genomic hybridization (CGH). Cytometry 1997; 28(3): Lapierre JM, Cacheux V, Collot N et al. Comparison of comparative genomic hybridization with conventional karyotype and classical fluorescence in situ hybridization for prenatal and postnatal diagnosis of unbalanced chromosome abnormalities. Ann Genet 1998;41(3): Daniely M, Barkai G, Goldman B, Aviram-Goldring A. Detection of numerical chromosome aberrations by comparative genomic hybridization. Prenat Diagn 1999;19(2): Kallioniemi A, Kallioniemi OP, Sudar D et al. Comparative genomic hybridization for molecular cytogenetic analysis of solid tumors. Science 1992;258(5083): Rickman L, Fiegler H, Shaw-Smith C et al. Prenatal detection of unbalanced chromosomal rearrangements by array CGH. J Med Genet 2006;43(4): Rickman L, Fiegler H, Carter NP, Bobrow M. Prenatal diagnosis by array-cgh. Eur J Med Genet 2005;48(3): Le Caignec C, Boceno M, Saugier-Veber P et al. Detection of genomic imbalances by array based comparative genomic hybridisation in fetuses with multiple malformations. J Med Genet 2005;42(2): Miura S, Miura K, Masuzaki H et al. Microarray comparative genomic hybridization (CGH)-based prenatal diagnosis for chromosome abnormalities using cell-free fetal DNA in amniotic fluid. J Hum Genet 2006;51(5): Cheung J, Estivill X, Khaja R et al. Genome-wide detection of segmental duplications and potential assembly errors in the human genome sequence. Genome Biol 2003;4(4): R Iafrate AJ, Feuk L, Rivera MN et al. Detection of largescale variation in the human genome. Nat Genet 2004; 36(9): Sebat J, Lakshmi B, Troge J et al. Large-scale copy number polymorphism in the human genome. Science 2004; 23;305(5683): Friedman JM, Baross A, Delaney AD et al. Oligonucleotide microarray analysis of genomic imbalance in children with mental retardation. Am J Hum Genet 2006;79(3): Veltman JA, Schoenmakers EF, Eussen BH et al. Highthroughput analysis of subtelomeric chromosome rearrangements by use of array-based comparative genomic hybridization. Am J Hum Genet 2002;70(5): Locke DP, Segraves R, Nicholls RD et al. BAC microarray analysis of 15q11-q13 rearrangements and the impact of segmental duplications. J Med Genet 2004;41(3): Cheung SW, Shaw CA, Yu W et al. Development and validation of a CGH microarray for clinical cytogenetic diagnosis. Genet Med 2005;7(6): Shaffer LG, Bejjani BA. Medical applications of array CGH and the transformation of clinical cytogenetics.cytogenet Genome Res 2006;115(3-4): Sahoo T, Cheung SW, Ward P et al. Prenatal diagnosis of chromosomal abnormalities using array-based comparative genomic hybridization. Genet Med 2006;8(11): Journal of Prenatal Medicine 2007; 1 (1): 16-22
Prenatal Diagnosis: Are There Microarrays in Your Future?
 Financial Disclosure UCSF Antepartum Intrapartum Management Course June 8 I have no financial relationship with any aspect of private industry Prenatal Diagnosis: Are There Microarrays in Your Future?
Financial Disclosure UCSF Antepartum Intrapartum Management Course June 8 I have no financial relationship with any aspect of private industry Prenatal Diagnosis: Are There Microarrays in Your Future?
New and Developing Technologies for Genetic Diagnostics National Genetics Reference Laboratory (Wessex) Salisbury, UK - July 2010 BACs on Beads
 New and Developing Technologies for Genetic Diagnostics National Genetics Reference Laboratory (Wessex) Salisbury, UK - July 2010 BACs on Beads Susan Gross, MD Division of Reproductive Genetics Professor
New and Developing Technologies for Genetic Diagnostics National Genetics Reference Laboratory (Wessex) Salisbury, UK - July 2010 BACs on Beads Susan Gross, MD Division of Reproductive Genetics Professor
CYTOGENETICS Dr. Mary Ann Perle
 CYTOGENETICS Dr. Mary Ann Perle I) Mitosis and metaphase chromosomes A) Chromosomes are most fully condensed and clearly distinguishable during mitosis. B) Mitosis (M phase) takes 1 to 2 hrs and is divided
CYTOGENETICS Dr. Mary Ann Perle I) Mitosis and metaphase chromosomes A) Chromosomes are most fully condensed and clearly distinguishable during mitosis. B) Mitosis (M phase) takes 1 to 2 hrs and is divided
Chromosome microarray analysis in routine prenatal diagnosis practice: a prospective study on 3000 consecutive clinical cases
 Chromosome microarray analysis in routine prenatal diagnosis practice: a prospective study on 3000 consecutive clinical cases Fiorentino F, Napoletano S, Caiazzo F, Sessa M, Bono S, Spizzichino L, Gordon
Chromosome microarray analysis in routine prenatal diagnosis practice: a prospective study on 3000 consecutive clinical cases Fiorentino F, Napoletano S, Caiazzo F, Sessa M, Bono S, Spizzichino L, Gordon
Application of Array-based Comparative Genome Hybridization in Children with Developmental Delay or Mental Retardation
 Pediatr Neonatol 2008;49(6):213 217 REVIEW ARTICLE Application of Array-based Comparative Genome Hybridization in Children with Developmental Delay or Mental Retardation Jao-Shwann Liang 1,2 *, Keiko Shimojima
Pediatr Neonatol 2008;49(6):213 217 REVIEW ARTICLE Application of Array-based Comparative Genome Hybridization in Children with Developmental Delay or Mental Retardation Jao-Shwann Liang 1,2 *, Keiko Shimojima
Understanding the Human Karyotype Colleen Jackson Cook, Ph.D.
 Understanding the Human Karyotype Colleen Jackson Cook, Ph.D. SUPPLEMENTAL READING Nussbaum, RL, McInnes, RR, and Willard HF (2007) Thompson and Thompson Genetics in Medicine, 7th edition. Saunders: Philadelphia.
Understanding the Human Karyotype Colleen Jackson Cook, Ph.D. SUPPLEMENTAL READING Nussbaum, RL, McInnes, RR, and Willard HF (2007) Thompson and Thompson Genetics in Medicine, 7th edition. Saunders: Philadelphia.
CHROMOSOMAL MICROARRAY (CGH+SNP)
 Chromosome imbalances are a significant cause of developmental delay, mental retardation, autism spectrum disorders, dysmorphic features and/or birth defects. The imbalance of genetic material may be due
Chromosome imbalances are a significant cause of developmental delay, mental retardation, autism spectrum disorders, dysmorphic features and/or birth defects. The imbalance of genetic material may be due
review Genetics IN Medicine 13
 January 2008 Vol. 10 No. 1 review Pre- and postnatal genetic testing by array-comparative genomic hybridization: genetic counseling perspectives Sandra Darilek, MS, Patricia Ward, MS, Amber Pursley, MS,
January 2008 Vol. 10 No. 1 review Pre- and postnatal genetic testing by array-comparative genomic hybridization: genetic counseling perspectives Sandra Darilek, MS, Patricia Ward, MS, Amber Pursley, MS,
CURRENT GENETIC TESTING TOOLS IN NEONATAL MEDICINE. Dr. Bahar Naghavi
 2 CURRENT GENETIC TESTING TOOLS IN NEONATAL MEDICINE Dr. Bahar Naghavi Assistant professor of Basic Science Department, Shahid Beheshti University of Medical Sciences, Tehran,Iran 3 Introduction Over 4000
2 CURRENT GENETIC TESTING TOOLS IN NEONATAL MEDICINE Dr. Bahar Naghavi Assistant professor of Basic Science Department, Shahid Beheshti University of Medical Sciences, Tehran,Iran 3 Introduction Over 4000
Challenges of CGH array testing in children with developmental delay. Dr Sally Davies 17 th September 2014
 Challenges of CGH array testing in children with developmental delay Dr Sally Davies 17 th September 2014 CGH array What is CGH array? Understanding the test Benefits Results to expect Consent issues Ethical
Challenges of CGH array testing in children with developmental delay Dr Sally Davies 17 th September 2014 CGH array What is CGH array? Understanding the test Benefits Results to expect Consent issues Ethical
Human Genetic Variation Copy Number Variation
 The Evolution of Laboratory Prenatal Diagnosis Lawrence D. Platt, MD Professor of Obstetrics and Gynecology David Geffen School of Medicine at UCLA Center for Fetal Medicine & Women s Ultrasound Los Angeles,
The Evolution of Laboratory Prenatal Diagnosis Lawrence D. Platt, MD Professor of Obstetrics and Gynecology David Geffen School of Medicine at UCLA Center for Fetal Medicine & Women s Ultrasound Los Angeles,
Cytogenetics 101: Clinical Research and Molecular Genetic Technologies
 Cytogenetics 101: Clinical Research and Molecular Genetic Technologies Topics for Today s Presentation 1 Classical vs Molecular Cytogenetics 2 What acgh? 3 What is FISH? 4 What is NGS? 5 How can these
Cytogenetics 101: Clinical Research and Molecular Genetic Technologies Topics for Today s Presentation 1 Classical vs Molecular Cytogenetics 2 What acgh? 3 What is FISH? 4 What is NGS? 5 How can these
Approach to Mental Retardation and Developmental Delay. SR Ghaffari MSc MD PhD
 Approach to Mental Retardation and Developmental Delay SR Ghaffari MSc MD PhD Introduction Objectives Definition of MR and DD Classification Epidemiology (prevalence, recurrence risk, ) Etiology Importance
Approach to Mental Retardation and Developmental Delay SR Ghaffari MSc MD PhD Introduction Objectives Definition of MR and DD Classification Epidemiology (prevalence, recurrence risk, ) Etiology Importance
Spontaneous abortions are common, with 10% to 15% of all
 ARTICLE Diagnosis of miscarriages by molecular karyotyping: Benefits and pitfalls Caroline Robberecht, MSc, Vicky Schuddinck, BSc, Jean-Pierre Fryns, MD, PhD, and Joris Robert Vermeesch, PhD Purpose: About
ARTICLE Diagnosis of miscarriages by molecular karyotyping: Benefits and pitfalls Caroline Robberecht, MSc, Vicky Schuddinck, BSc, Jean-Pierre Fryns, MD, PhD, and Joris Robert Vermeesch, PhD Purpose: About
New Molecular Techniques for the Prenatal Detection of Chromosomal Aneuploidy
 SOGC TECHNICAL UPDATE No. 210, July 2008 New Molecular Techniques for the Prenatal Detection of Chromosomal Aneuploidy This technical update has been prepared by the Genetics Committee and approved by
SOGC TECHNICAL UPDATE No. 210, July 2008 New Molecular Techniques for the Prenatal Detection of Chromosomal Aneuploidy This technical update has been prepared by the Genetics Committee and approved by
Canadian College of Medical Geneticists (CCMG) Cytogenetics Examination. May 4, 2010
 Canadian College of Medical Geneticists (CCMG) Cytogenetics Examination May 4, 2010 Examination Length = 3 hours Total Marks = 100 (7 questions) Total Pages = 8 (including cover sheet and 2 pages of prints)
Canadian College of Medical Geneticists (CCMG) Cytogenetics Examination May 4, 2010 Examination Length = 3 hours Total Marks = 100 (7 questions) Total Pages = 8 (including cover sheet and 2 pages of prints)
Copy number imbalances detected with a BAC-based array comparative genomic hybridization platform in congenital diaphragmatic hernia fetuses
 Copy number imbalances detected with a BAC-based array comparative genomic hybridization platform in congenital diaphragmatic hernia fetuses I.N. Machado 1,2, J.K. Heinrich 2, R. Barini 1 and C.F.A. Peralta
Copy number imbalances detected with a BAC-based array comparative genomic hybridization platform in congenital diaphragmatic hernia fetuses I.N. Machado 1,2, J.K. Heinrich 2, R. Barini 1 and C.F.A. Peralta
Association for Molecular Pathology Promoting Clinical Practice, Basic Research, and Education in Molecular Pathology
 Association for Molecular Pathology Promoting Clinical Practice, Basic Research, and Education in Molecular Pathology 9650 Rockville Pike, Bethesda, Maryland 20814 Tel: 301-634-7939 Fax: 301-634-7990 Email:
Association for Molecular Pathology Promoting Clinical Practice, Basic Research, and Education in Molecular Pathology 9650 Rockville Pike, Bethesda, Maryland 20814 Tel: 301-634-7939 Fax: 301-634-7990 Email:
Corporate Medical Policy
 Corporate Medical Policy Invasive Prenatal (Fetal) Diagnostic Testing File Name: Origination: Last CAP Review: Next CAP Review: Last Review: invasive_prenatal_(fetal)_diagnostic_testing 12/2014 3/2018
Corporate Medical Policy Invasive Prenatal (Fetal) Diagnostic Testing File Name: Origination: Last CAP Review: Next CAP Review: Last Review: invasive_prenatal_(fetal)_diagnostic_testing 12/2014 3/2018
SALSA MLPA probemix P372-B1 Microdeletion Syndromes 6 Lot B1-1016, B
 SALSA MLPA probemix P372-B1 Microdeletion Syndromes 6 Lot B1-1016, B1-0613. The purpose of the P372 probemix is to further investigate results found with the P245 Microdeletion Syndromes-1A probemix. The
SALSA MLPA probemix P372-B1 Microdeletion Syndromes 6 Lot B1-1016, B1-0613. The purpose of the P372 probemix is to further investigate results found with the P245 Microdeletion Syndromes-1A probemix. The
MLPA SAMPLES USER GUIDE
 MLPA SAMPLES USER GUIDE MLPA microdeletion studies Requirements: 1-2 mls (minimum) of peripheral blood in a lithium heparin tube (green or orange top) AND 1-2 mls of peripheral blood in a EDTA tube (purple
MLPA SAMPLES USER GUIDE MLPA microdeletion studies Requirements: 1-2 mls (minimum) of peripheral blood in a lithium heparin tube (green or orange top) AND 1-2 mls of peripheral blood in a EDTA tube (purple
SALSA MLPA probemix P371-A1 Microdeletion Syndromes 5 Lot A1-0509
 mix P371-A1 Microdeletion Syndromes 5 Lot A1-0509 The purpose of the P371 mix is to further investigate results found with the P245 Microdeletion mix. The P245 mix provides a possibility to screen samples
mix P371-A1 Microdeletion Syndromes 5 Lot A1-0509 The purpose of the P371 mix is to further investigate results found with the P245 Microdeletion mix. The P245 mix provides a possibility to screen samples
Detection of aneuploidy in a single cell using the Ion ReproSeq PGS View Kit
 APPLICATION NOTE Ion PGM System Detection of aneuploidy in a single cell using the Ion ReproSeq PGS View Kit Key findings The Ion PGM System, in concert with the Ion ReproSeq PGS View Kit and Ion Reporter
APPLICATION NOTE Ion PGM System Detection of aneuploidy in a single cell using the Ion ReproSeq PGS View Kit Key findings The Ion PGM System, in concert with the Ion ReproSeq PGS View Kit and Ion Reporter
Case 1B. 46,XY,-14,+t(14;21)
 Case 1B 46,XY,-14,+t(14;21) G-banded Chromosome telomere centromere G-dark bands AT-rich few genes G-pale bands GC-rich many genes telomere ideograms ideograms Conventional (light microscopy) p = short
Case 1B 46,XY,-14,+t(14;21) G-banded Chromosome telomere centromere G-dark bands AT-rich few genes G-pale bands GC-rich many genes telomere ideograms ideograms Conventional (light microscopy) p = short
Sharan Goobie, MD, MSc, FRCPC
 Sharan Goobie, MD, MSc, FRCPC Chromosome testing in 2014 Presenter Disclosure: Sharan Goobie has no potential for conflict of interest with this presentation Objectives Review of standard genetic investigations
Sharan Goobie, MD, MSc, FRCPC Chromosome testing in 2014 Presenter Disclosure: Sharan Goobie has no potential for conflict of interest with this presentation Objectives Review of standard genetic investigations
Microdeletion and Prenatal FISH Probes
 Microdeletion and Prenatal FISH Probes Rapid Prenatal Screening The Cytocell prenatal FISH assays are designed for the rapid and accurate detection of the most common foetal chromosomal disorders: Down
Microdeletion and Prenatal FISH Probes Rapid Prenatal Screening The Cytocell prenatal FISH assays are designed for the rapid and accurate detection of the most common foetal chromosomal disorders: Down
Applications of Chromosomal Microarray Analysis (CMA) in pre- and postnatal Diagnostic: advantages, limitations and concerns
 Applications of Chromosomal Microarray Analysis (CMA) in pre- and postnatal Diagnostic: advantages, limitations and concerns جواد کریمزاد حق PhD of Medical Genetics آزمايشگاه پاتوبيولوژي و ژنتيك پارسه
Applications of Chromosomal Microarray Analysis (CMA) in pre- and postnatal Diagnostic: advantages, limitations and concerns جواد کریمزاد حق PhD of Medical Genetics آزمايشگاه پاتوبيولوژي و ژنتيك پارسه
Determination of Genomic Imbalances by Genome-wide Screening Approaches
 Overview Determination of Genomic Imbalances by Genome-wide Screening Approaches Károly Szuhai Introduction/Methodologies Applications/Results Conclusion Approaches Introduction/Methodologies Chromosome
Overview Determination of Genomic Imbalances by Genome-wide Screening Approaches Károly Szuhai Introduction/Methodologies Applications/Results Conclusion Approaches Introduction/Methodologies Chromosome
Karyology. Preparation and study of karyotypes is part of Cytogenetics.
 Chromosomal Karyotyping Karyology Karyotyping - process of pairing and ordering all chromosomes of an organism, thus providing a genome-wide snapshot of an individual's chromosomes. Karyotypes describe
Chromosomal Karyotyping Karyology Karyotyping - process of pairing and ordering all chromosomes of an organism, thus providing a genome-wide snapshot of an individual's chromosomes. Karyotypes describe
scfish: A Primer Peter K. Rogan, Ph.D. Joan H. M. Knoll, Ph.D., FACMG, FCCMG Children s Mercy Hospitals and Clinics Tuesday, July 29, 2003
 scfish: A Primer Peter K. Rogan, Ph.D. Joan H. M. Knoll, Ph.D., FACMG, FCCMG Children s Mercy Hospitals and Clinics Tuesday, July 29, 2003 FISH: A Molecular Cytogenetic Test Complementary nucleic acid
scfish: A Primer Peter K. Rogan, Ph.D. Joan H. M. Knoll, Ph.D., FACMG, FCCMG Children s Mercy Hospitals and Clinics Tuesday, July 29, 2003 FISH: A Molecular Cytogenetic Test Complementary nucleic acid
MOLECULAR MECHANISMS FOR CONSTITUTIONAL CHROMOSOMAL REARRANGEMENTS IN HUMANS
 Annu. Rev. Genet. 2000. 34:297 329 Copyright c 2000 by Annual Reviews. All rights reserved MOLECULAR MECHANISMS FOR CONSTITUTIONAL CHROMOSOMAL REARRANGEMENTS IN HUMANS Lisa G. Shaffer 1 and James R. Lupski
Annu. Rev. Genet. 2000. 34:297 329 Copyright c 2000 by Annual Reviews. All rights reserved MOLECULAR MECHANISMS FOR CONSTITUTIONAL CHROMOSOMAL REARRANGEMENTS IN HUMANS Lisa G. Shaffer 1 and James R. Lupski
Genetic Testing 101: Interpreting the Chromosomes
 Genetic Testing 101: Interpreting the Chromosomes Kristin Lindstrom, MD Division of Genetics and Metabolism Phoenix Children s Hospital AzAAP Pediatrics in the Red Rocks I have no disclosures for this
Genetic Testing 101: Interpreting the Chromosomes Kristin Lindstrom, MD Division of Genetics and Metabolism Phoenix Children s Hospital AzAAP Pediatrics in the Red Rocks I have no disclosures for this
Clinical evaluation of microarray data
 Clinical evaluation of microarray data David Amor 19 th June 2011 Single base change Microarrays 3-4Mb What is a microarray? Up to 10 6 bits of Information!! Highly multiplexed FISH hybridisations. Microarray
Clinical evaluation of microarray data David Amor 19 th June 2011 Single base change Microarrays 3-4Mb What is a microarray? Up to 10 6 bits of Information!! Highly multiplexed FISH hybridisations. Microarray
C linical geneticists are notorious jargon
 1264 REVIEW Chromosome analysis: what and when to request F H Sharkey, E Maher, D R FitzPatrick... Chromosome abnormalities have long been recognised as an important cause of learning disability and multiple
1264 REVIEW Chromosome analysis: what and when to request F H Sharkey, E Maher, D R FitzPatrick... Chromosome abnormalities have long been recognised as an important cause of learning disability and multiple
Schedule of Accreditation issued by United Kingdom Accreditation Service 2 Pine Trees, Chertsey Lane, Staines-upon-Thames, TW18 3HR, UK
 SW Thames Regional Genetics Laboratory, Jenner Wing, SGUL Cranmer Terrace London SW17 0RE Contact: Mr John Short Tel: +44 (0) 208 725 5332 Fax: +44 (0) 208 725 3440 E-Mail: swtrgl@stgeorges.nhs.uk Website:
SW Thames Regional Genetics Laboratory, Jenner Wing, SGUL Cranmer Terrace London SW17 0RE Contact: Mr John Short Tel: +44 (0) 208 725 5332 Fax: +44 (0) 208 725 3440 E-Mail: swtrgl@stgeorges.nhs.uk Website:
Current controversies in prenatal diagnosis 3: is conventional chromosome analysis necessary in the post-array CGH era?
 PRENATAL DIAGNOSIS Prenat Diagn 2011; 31: 235 243. Published online 10 February 2011 in Wiley Online Library (wileyonlinelibrary.com).2722 DEBATE Current controversies in prenatal diagnosis 3: is conventional
PRENATAL DIAGNOSIS Prenat Diagn 2011; 31: 235 243. Published online 10 February 2011 in Wiley Online Library (wileyonlinelibrary.com).2722 DEBATE Current controversies in prenatal diagnosis 3: is conventional
National Medical Policy
 National Medical Policy Subject: Policy Number: Single Nucleotide Polymorphism (SNP) Chromosomal Microarray Analysis for Prenatal Testing and Intellectual Disability, Developmental Delay, and Multiple
National Medical Policy Subject: Policy Number: Single Nucleotide Polymorphism (SNP) Chromosomal Microarray Analysis for Prenatal Testing and Intellectual Disability, Developmental Delay, and Multiple
CHROMOSOME MICROARRAY TESTING
 CHROMOSOME MICROARRAY TESTING UnitedHealthcare Commercial Medical Policy Policy Number: 2017T0559J Effective Date: August 1, 2017 Table of Contents Page INSTRUCTIONS FOR USE... 1 BENEFIT CONSIDERATIONS...
CHROMOSOME MICROARRAY TESTING UnitedHealthcare Commercial Medical Policy Policy Number: 2017T0559J Effective Date: August 1, 2017 Table of Contents Page INSTRUCTIONS FOR USE... 1 BENEFIT CONSIDERATIONS...
Children s Hospital Zagreb, University of Zagreb Medical School, Zagreb, Croatia.
 Multiplex ligation-dependent probe amplification (MLPA) genetic testing in the diagnostics of children with developmental delay/intellectual disabilities Leona Morožin Pohovski, Ingeborg Barišid Children
Multiplex ligation-dependent probe amplification (MLPA) genetic testing in the diagnostics of children with developmental delay/intellectual disabilities Leona Morožin Pohovski, Ingeborg Barišid Children
chromosomal anomalies and mental pdf Chapter 8: Chromosomes and Chromosomal Anomalies (PDF) Chromosomal abnormalities -A review - ResearchGate
 DOWNLOAD OR READ : CHROMOSOMAL ANOMALIES AND MENTAL RETARDATION FROM GENOTYPES TO NEUROPSYCHOLOGICAL PHENOTYPES OF GENETIC SYNDROMES AT HIGH INCIDENCEGENOTYPE TO PHENOTYPE PDF EBOOK EPUB MOBI Page 1 Page
DOWNLOAD OR READ : CHROMOSOMAL ANOMALIES AND MENTAL RETARDATION FROM GENOTYPES TO NEUROPSYCHOLOGICAL PHENOTYPES OF GENETIC SYNDROMES AT HIGH INCIDENCEGENOTYPE TO PHENOTYPE PDF EBOOK EPUB MOBI Page 1 Page
Genomics and Genetics in Healthcare. By Mary Knutson, RN, MSN
 Genomics and Genetics in Healthcare By Mary Knutson, RN, MSN Clinical Objectives Understand the importance of genomics to provide effective nursing care Integrate genetic knowledge and skills into nursing
Genomics and Genetics in Healthcare By Mary Knutson, RN, MSN Clinical Objectives Understand the importance of genomics to provide effective nursing care Integrate genetic knowledge and skills into nursing
Chromosome Abnormalities
 Chromosome Abnormalities Chromosomal abnormalities vs. molecular mutations Simply a matter of size Chromosomal abnormalities are big errors Two types of abnormalities 1. Constitutional problem present
Chromosome Abnormalities Chromosomal abnormalities vs. molecular mutations Simply a matter of size Chromosomal abnormalities are big errors Two types of abnormalities 1. Constitutional problem present
Prior Authorization Review Panel MCO Policy Submission
 Prior Authorization Review Panel MCO Policy Submission A separate copy of this form must accompany each policy submitted for review. Policies submitted without this form will not be considered for review.
Prior Authorization Review Panel MCO Policy Submission A separate copy of this form must accompany each policy submitted for review. Policies submitted without this form will not be considered for review.
Clinical Interpretation of Cancer Genomes
 IGENZ Ltd, Auckland, New Zealand Clinical Interpretation of Cancer Genomes Dr Amanda Dixon-McIver www.igenz.co.nz 1992 Slovenia and Croatia gain independence USA and Russia declare the Cold War over Steffi
IGENZ Ltd, Auckland, New Zealand Clinical Interpretation of Cancer Genomes Dr Amanda Dixon-McIver www.igenz.co.nz 1992 Slovenia and Croatia gain independence USA and Russia declare the Cold War over Steffi
An Overview of Cytogenetics. Bridget Herschap, M.D. 9/23/2013
 An Overview of Cytogenetics Bridget Herschap, M.D. 9/23/2013 Objectives } History and Introduction of Cytogenetics } Overview of Current Techniques } Common cytogenetic tests and their clinical application
An Overview of Cytogenetics Bridget Herschap, M.D. 9/23/2013 Objectives } History and Introduction of Cytogenetics } Overview of Current Techniques } Common cytogenetic tests and their clinical application
%(6/31) [DOI] /j.issn
![%(6/31) [DOI] /j.issn %(6/31) [DOI] /j.issn](/thumbs/92/110077812.jpg) 750 2016 9 1 41 9 [ ] 2012 2 2015 5 31 20~37 21~27 31 G 4 1 26 23 3 11.54%(3/26) 4 / 21 6 19.35%(6/31) [ ] [ ] R714.53 [ ] A [ ] 0577-7402(2016)09-0750-08 [DOI] 10.11855/j.issn.0577-7402.2016.09.10 Application
750 2016 9 1 41 9 [ ] 2012 2 2015 5 31 20~37 21~27 31 G 4 1 26 23 3 11.54%(3/26) 4 / 21 6 19.35%(6/31) [ ] [ ] R714.53 [ ] A [ ] 0577-7402(2016)09-0750-08 [DOI] 10.11855/j.issn.0577-7402.2016.09.10 Application
Genome-wide analysis by SNP Array
 Genome-wide analysis by SNP Array SNP Array in the diagnosis of Mental Retardation and Congenital Abnormalities Contents Introduction...3 Conventional and molecular cytogenetics (FISH)...4 1. Karyotyping...4
Genome-wide analysis by SNP Array SNP Array in the diagnosis of Mental Retardation and Congenital Abnormalities Contents Introduction...3 Conventional and molecular cytogenetics (FISH)...4 1. Karyotyping...4
22q11.2 DELETION SYNDROME. Anna Mª Cueto González Clinical Geneticist Programa de Medicina Molecular y Genética Hospital Vall d Hebrón (Barcelona)
 22q11.2 DELETION SYNDROME Anna Mª Cueto González Clinical Geneticist Programa de Medicina Molecular y Genética Hospital Vall d Hebrón (Barcelona) Genomic disorders GENOMICS DISORDERS refers to those diseases
22q11.2 DELETION SYNDROME Anna Mª Cueto González Clinical Geneticist Programa de Medicina Molecular y Genética Hospital Vall d Hebrón (Barcelona) Genomic disorders GENOMICS DISORDERS refers to those diseases
Structural Chromosome Aberrations
 Structural Chromosome Aberrations 2 Structural chromosome aberrations or chromosome mutations represent apart from aneuploidies the most frequent pathologic findings in applied chromosome diagnostics.
Structural Chromosome Aberrations 2 Structural chromosome aberrations or chromosome mutations represent apart from aneuploidies the most frequent pathologic findings in applied chromosome diagnostics.
Most severely affected will be the probe for exon 15. Please keep an eye on the D-fragments (especially the 96 nt fragment).
 SALSA MLPA probemix P343-C3 Autism-1 Lot C3-1016. As compared to version C2 (lot C2-0312) five reference probes have been replaced, one reference probe added and several lengths have been adjusted. Warning:
SALSA MLPA probemix P343-C3 Autism-1 Lot C3-1016. As compared to version C2 (lot C2-0312) five reference probes have been replaced, one reference probe added and several lengths have been adjusted. Warning:
Faravareh Khordadpoor (PhD in molecular genetics) 1- Tehran Medical Genetics Laboratory 2- Science and research branch, Islamic Azad University
 Faravareh Khordadpoor (PhD in molecular genetics) 1- Tehran Medical Genetics Laboratory 2- Science and research branch, Islamic Azad University 1395 21 مشاوره ژنتیک و نقش آن در پیش گیری از معلولیت ها 20
Faravareh Khordadpoor (PhD in molecular genetics) 1- Tehran Medical Genetics Laboratory 2- Science and research branch, Islamic Azad University 1395 21 مشاوره ژنتیک و نقش آن در پیش گیری از معلولیت ها 20
Chapter 15 Notes 15.1: Mendelian inheritance chromosome theory of inheritance wild type 15.2: Sex-linked genes
 Chapter 15 Notes The Chromosomal Basis of Inheritance Mendel s hereditary factors were genes, though this wasn t known at the time Now we know that genes are located on The location of a particular gene
Chapter 15 Notes The Chromosomal Basis of Inheritance Mendel s hereditary factors were genes, though this wasn t known at the time Now we know that genes are located on The location of a particular gene
Mental retardation is one of the most common forms of THE USE OF GENOMIC MICROARRAYS TO STUDY CHROMOSOMAL ABNORMALITIES IN MENTAL RETARDATION
 MENTAL RETARDATION AND DEVELOPMENTAL DISABILITIES RESEARCH REVIEWS 11: 279 285 (2005) THE USE OF GENOMIC MICROARRAYS TO STUDY CHROMOSOMAL ABNORMALITIES IN MENTAL RETARDATION Rong Mao 1,2 and Jonathan Pevsner
MENTAL RETARDATION AND DEVELOPMENTAL DISABILITIES RESEARCH REVIEWS 11: 279 285 (2005) THE USE OF GENOMIC MICROARRAYS TO STUDY CHROMOSOMAL ABNORMALITIES IN MENTAL RETARDATION Rong Mao 1,2 and Jonathan Pevsner
POLICY PRODUCT VARIATIONS DESCRIPTION/BACKGROUND RATIONALE DEFINITIONS BENEFIT VARIATIONS DISCLAIMER CODING INFORMATION REFERENCES POLICY HISTORY
 Original Issue Date (Created): April 26, 2011 Most Recent Review Date (Revised): January 28, 2014 Effective Date: June 1, 2014 POLICY PRODUCT VARIATIONS DESCRIPTION/BACKGROUND RATIONALE DEFINITIONS BENEFIT
Original Issue Date (Created): April 26, 2011 Most Recent Review Date (Revised): January 28, 2014 Effective Date: June 1, 2014 POLICY PRODUCT VARIATIONS DESCRIPTION/BACKGROUND RATIONALE DEFINITIONS BENEFIT
Clinical Cytogenomics Laboratory. Laboratory. Leading diagnostics for better medicine. Beaumont
 Clinical Cytogenomics Laboratory Leading diagnostics for better medicine Beaumont Laboratory Beaumont Laboratory s Clinical Cytogenomics Lab Genome decoding is essential Knowledge of variations in genome
Clinical Cytogenomics Laboratory Leading diagnostics for better medicine Beaumont Laboratory Beaumont Laboratory s Clinical Cytogenomics Lab Genome decoding is essential Knowledge of variations in genome
Product Description SALSA MLPA Probemix P064-C2 Microdeletion Syndromes-1B To be used with the MLPA General Protocol.
 Product Description SALSA Probemix P064-C2 Microdeletion Syndromes-1B To be used with the MLPA General Protocol. Version C2. Small change in sequence of two probes. For complete product history see page
Product Description SALSA Probemix P064-C2 Microdeletion Syndromes-1B To be used with the MLPA General Protocol. Version C2. Small change in sequence of two probes. For complete product history see page
What s the Human Genome Project Got to Do with Developmental Disabilities?
 What s the Human Genome Project Got to Do with Developmental Disabilities? Disclosures Neither speaker has anything to disclose. Phase Two: Interpretation Officially started in October 1990 Goals of the
What s the Human Genome Project Got to Do with Developmental Disabilities? Disclosures Neither speaker has anything to disclose. Phase Two: Interpretation Officially started in October 1990 Goals of the
Medical Policy. MP Invasive Prenatal (Fetal) Diagnostic Testing
 Medical Policy MP 2.04.116 BCBSA Ref. Policy: 2.04.116 Last Review: 04/30/2018 Effective Date: 04/30/2018 Section: Medicine Related Policies 2.04.59 Genetic Testing for Developmental Delay/Intellectual
Medical Policy MP 2.04.116 BCBSA Ref. Policy: 2.04.116 Last Review: 04/30/2018 Effective Date: 04/30/2018 Section: Medicine Related Policies 2.04.59 Genetic Testing for Developmental Delay/Intellectual
SNP Array NOTE: THIS IS A SAMPLE REPORT AND MAY NOT REFLECT ACTUAL PATIENT DATA. FORMAT AND/OR CONTENT MAY BE UPDATED PERIODICALLY.
 SAMPLE REPORT SNP Array NOTE: THIS IS A SAMPLE REPORT AND MAY NOT REFLECT ACTUAL PATIENT DATA. FORMAT AND/OR CONTENT MAY BE UPDATED PERIODICALLY. RESULTS SNP Array Copy Number Variations Result: LOSS,
SAMPLE REPORT SNP Array NOTE: THIS IS A SAMPLE REPORT AND MAY NOT REFLECT ACTUAL PATIENT DATA. FORMAT AND/OR CONTENT MAY BE UPDATED PERIODICALLY. RESULTS SNP Array Copy Number Variations Result: LOSS,
Role of FISH in Hematological Cancers
 Role of FISH in Hematological Cancers Thomas S.K. Wan PhD,FRCPath,FFSc(RCPA) Honorary Professor, Department of Pathology & Clinical Biochemistry, Queen Mary Hospital, University of Hong Kong. e-mail: wantsk@hku.hk
Role of FISH in Hematological Cancers Thomas S.K. Wan PhD,FRCPath,FFSc(RCPA) Honorary Professor, Department of Pathology & Clinical Biochemistry, Queen Mary Hospital, University of Hong Kong. e-mail: wantsk@hku.hk
Multiple Copy Number Variations in a Patient with Developmental Delay ASCLS- March 31, 2016
 Multiple Copy Number Variations in a Patient with Developmental Delay ASCLS- March 31, 2016 Marwan Tayeh, PhD, FACMG Director, MMGL Molecular Genetics Assistant Professor of Pediatrics Department of Pediatrics
Multiple Copy Number Variations in a Patient with Developmental Delay ASCLS- March 31, 2016 Marwan Tayeh, PhD, FACMG Director, MMGL Molecular Genetics Assistant Professor of Pediatrics Department of Pediatrics
MULTIPLE CHOICE QUESTIONS
 SHORT ANSWER QUESTIONS-Please type your awesome answers on a separate sheet of paper. 1. What is an X-linked inheritance pattern? Use a specific example to explain the role of the father and mother in
SHORT ANSWER QUESTIONS-Please type your awesome answers on a separate sheet of paper. 1. What is an X-linked inheritance pattern? Use a specific example to explain the role of the father and mother in
GENOME-WIDE ANALYSIS BY SNP ARRAY
 GENOME-WIDE ANALYSIS BY SNP ARRAY SNP ARRAY IN THE DIAGNOSIS OF INTELLECTUAL DISABILITY AND C O N G E N I T A L ABNORMALITIES Contents Introduction... 3 Conventional and molecular cytogenetics (FISH)...
GENOME-WIDE ANALYSIS BY SNP ARRAY SNP ARRAY IN THE DIAGNOSIS OF INTELLECTUAL DISABILITY AND C O N G E N I T A L ABNORMALITIES Contents Introduction... 3 Conventional and molecular cytogenetics (FISH)...
SNP Array NOTE: THIS IS A SAMPLE REPORT AND MAY NOT REFLECT ACTUAL PATIENT DATA. FORMAT AND/OR CONTENT MAY BE UPDATED PERIODICALLY.
 SAMPLE REPORT SNP Array NOTE: THIS IS A SAMPLE REPORT AND MAY NOT REFLECT ACTUAL PATIENT DATA. FORMAT AND/OR CONTENT MAY BE UPDATED PERIODICALLY. RESULTS SNP Array Copy Number Variations Result: GAIN,
SAMPLE REPORT SNP Array NOTE: THIS IS A SAMPLE REPORT AND MAY NOT REFLECT ACTUAL PATIENT DATA. FORMAT AND/OR CONTENT MAY BE UPDATED PERIODICALLY. RESULTS SNP Array Copy Number Variations Result: GAIN,
TEST MENU TEST CPT CODES TAT. Chromosome Analysis Bone Marrow x 2, 88264, x 3, Days
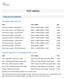 TEST MENU CANCER/LEUKEMIA CHROMOSOME ANALYSIS Chromosome Analysis Bone Marrow 88237 x 2, 88264, 88280 x 3, 88291 4 Days Chromosome Analysis Bone Marrow Core 88237 x 2, 88264, 88280 x 3, 88291 4 Days Chromosome
TEST MENU CANCER/LEUKEMIA CHROMOSOME ANALYSIS Chromosome Analysis Bone Marrow 88237 x 2, 88264, 88280 x 3, 88291 4 Days Chromosome Analysis Bone Marrow Core 88237 x 2, 88264, 88280 x 3, 88291 4 Days Chromosome
Structural Variation and Medical Genomics
 Structural Variation and Medical Genomics Andrew King Department of Biomedical Informatics July 8, 2014 You already know about small scale genetic mutations Single nucleotide polymorphism (SNPs) Deletions,
Structural Variation and Medical Genomics Andrew King Department of Biomedical Informatics July 8, 2014 You already know about small scale genetic mutations Single nucleotide polymorphism (SNPs) Deletions,
Certificate of Analysis
 COA Version 01; Issued on 30 June 2017 (v02) Certificate of Analysis SALSA probemix P064 Microdeletion Syndromes-1B Catalogue # Product name P064-025R, P064-50R, P064-100R Probemix P064 Microdeletion Syndromes-1B
COA Version 01; Issued on 30 June 2017 (v02) Certificate of Analysis SALSA probemix P064 Microdeletion Syndromes-1B Catalogue # Product name P064-025R, P064-50R, P064-100R Probemix P064 Microdeletion Syndromes-1B
One of the major causes of developmental delay, mental
 ARTICLE Detection of low-level mosaicism and placental mosaicism by oligonucleotide array comparative genomic hybridization Stuart A. Scott, PhD, FACMG 1, Ninette Cohen, PhD, FACMG 1, Tracy Brandt, PhD
ARTICLE Detection of low-level mosaicism and placental mosaicism by oligonucleotide array comparative genomic hybridization Stuart A. Scott, PhD, FACMG 1, Ninette Cohen, PhD, FACMG 1, Tracy Brandt, PhD
Short Report. B Lakhal a, R Braham b, R Berguigua a, N Bouali a, M Zaouali c, M Chaieb b, RA Veitia d,e,f, A Saad a,g and H Elghezal a,g
 Clin Genet 2010: 78: 181 185 Printed in Singapore. All rights reserved Short Report 2010 John Wiley & Sons A/S CLINICAL GENETICS doi: 10.1111/j.1399-0004.2009.01359.x Cytogenetic analyses of premature
Clin Genet 2010: 78: 181 185 Printed in Singapore. All rights reserved Short Report 2010 John Wiley & Sons A/S CLINICAL GENETICS doi: 10.1111/j.1399-0004.2009.01359.x Cytogenetic analyses of premature
7.1 Molecular Characterization of Fragile X Syndrome
 7 GENETIC DISORDERS Advances in knowledge of molecular genetics, cytogenetics and biochemical genetics have led to availability of diagnostic tests for various genetic disorders. The most important application
7 GENETIC DISORDERS Advances in knowledge of molecular genetics, cytogenetics and biochemical genetics have led to availability of diagnostic tests for various genetic disorders. The most important application
Genetic Testing for Single-Gene and Multifactorial Conditions
 Clinical Appropriateness Guidelines Genetic Testing for Single-Gene and Multifactorial Conditions EFFECTIVE DECEMBER 1, 2017 Appropriate.Safe.Affordable 2017 AIM Specialty Health 2069-1217 Table of Contents
Clinical Appropriateness Guidelines Genetic Testing for Single-Gene and Multifactorial Conditions EFFECTIVE DECEMBER 1, 2017 Appropriate.Safe.Affordable 2017 AIM Specialty Health 2069-1217 Table of Contents
Case Report Prenatal Diagnosis of Cystic Hygroma related to a Deletion of 16q24.1 with Haploinsufficiency of FOXF1 and FOXC2 Genes
 Case Reports in Genetics Volume 2012, Article ID 490408, 4 pages doi:10.1155/2012/490408 Case Report Prenatal Diagnosis of Cystic Hygroma related to a Deletion of 16q24.1 with Haploinsufficiency of FOXF1
Case Reports in Genetics Volume 2012, Article ID 490408, 4 pages doi:10.1155/2012/490408 Case Report Prenatal Diagnosis of Cystic Hygroma related to a Deletion of 16q24.1 with Haploinsufficiency of FOXF1
Clinical Genomics. Ina E. Amarillo, PhD FACMGG
 Clinical Genomics Ina E. Amarillo, PhD FACMGG Associate Medical Director, Cytogenetics Lab (CaTG), Lab and Genomic Medicine Assistant Professor, Pathology and Immunology Outline Clinical Genomics Testing
Clinical Genomics Ina E. Amarillo, PhD FACMGG Associate Medical Director, Cytogenetics Lab (CaTG), Lab and Genomic Medicine Assistant Professor, Pathology and Immunology Outline Clinical Genomics Testing
Copy Number Variants of Uncertain Significance in Prenatal diagnosis Are the Goalposts Moving? Lisa Burvill-Holmes Bristol Genetics Laboratory
 Copy Number Variants of Uncertain Significance in Prenatal diagnosis Are the Goalposts Moving? Lisa Burvill-Holmes Bristol Genetics Laboratory http://www.nbt.nhs.uk/genetics Microarray CGH in Prenatal
Copy Number Variants of Uncertain Significance in Prenatal diagnosis Are the Goalposts Moving? Lisa Burvill-Holmes Bristol Genetics Laboratory http://www.nbt.nhs.uk/genetics Microarray CGH in Prenatal
PROVIDER POLICIES & PROCEDURES
 PROVIDER POLICIES & PROCEDURES COMPARATIVE GENOMIC HYBRIDIZATION (CGH) MICROARRAY TESTING FOR DEVELOPMENTAL DELAY, AUTISM SPECTRUM DISORDER AND INTELLECTUAL DISABILITY The purpose of this policy is to
PROVIDER POLICIES & PROCEDURES COMPARATIVE GENOMIC HYBRIDIZATION (CGH) MICROARRAY TESTING FOR DEVELOPMENTAL DELAY, AUTISM SPECTRUM DISORDER AND INTELLECTUAL DISABILITY The purpose of this policy is to
Comprehensive Chromosome Screening Is NextGen Likely to be the Final Best Platform and What are its Advantages and Quirks?
 Comprehensive Chromosome Screening Is NextGen Likely to be the Final Best Platform and What are its Advantages and Quirks? Embryo 1 Embryo 2 combine samples for a single sequencing chip Barcode 1 CTAAGGTAAC
Comprehensive Chromosome Screening Is NextGen Likely to be the Final Best Platform and What are its Advantages and Quirks? Embryo 1 Embryo 2 combine samples for a single sequencing chip Barcode 1 CTAAGGTAAC
Genetic Disorders. PART ONE: Detecting Genetic Disorders. Amniocentesis Chorionic villus sampling Karyotype Triple Screen Blood Test
 Genetic Disorders PART ONE: Detecting Genetic Disorders Amniocentesis Chorionic villus sampling Karyotype Triple Screen Blood Test Amniocentesis A technique for determining genetic abnormalities in a fetus
Genetic Disorders PART ONE: Detecting Genetic Disorders Amniocentesis Chorionic villus sampling Karyotype Triple Screen Blood Test Amniocentesis A technique for determining genetic abnormalities in a fetus
CHROMOSOME MICROARRAY TESTING (NON-ONCOLOGY CONDITIONS)
 CHROMOSOME MICROARRAY TESTING (NON-ONCOLOGY CONDITIONS) UnitedHealthcare Oxford Clinical Policy Policy Number: LABORATORY 016.12 T2 Effective Date: June 1, 2018 Table of Contents Page INSTRUCTIONS FOR
CHROMOSOME MICROARRAY TESTING (NON-ONCOLOGY CONDITIONS) UnitedHealthcare Oxford Clinical Policy Policy Number: LABORATORY 016.12 T2 Effective Date: June 1, 2018 Table of Contents Page INSTRUCTIONS FOR
THE CHROMOSOMAL BASIS OF INHERITANCE CHAPTER 15
 THE CHROMOSOMAL BASIS OF INHERITANCE CHAPTER 15 What you must know: Inheritance in sex-linked genes. Inheritance of linked genes and chromosomal mapping. How alteration of chromosome number or structurally
THE CHROMOSOMAL BASIS OF INHERITANCE CHAPTER 15 What you must know: Inheritance in sex-linked genes. Inheritance of linked genes and chromosomal mapping. How alteration of chromosome number or structurally
Dr Dipti Deshmukh. Mayu Uemura. 11th September Introduction
 Genetic Findings in Children with Unexplained Developmental Delay Dr Dipti Deshmukh Neurodevelopmental Consultant, St. George's Hospital, London Mayu Uemura 4th year MBBS4, St George s University of London
Genetic Findings in Children with Unexplained Developmental Delay Dr Dipti Deshmukh Neurodevelopmental Consultant, St. George's Hospital, London Mayu Uemura 4th year MBBS4, St George s University of London
Genetics in Primary Care Curriculum Statement 6. Dr Dave Harniess PCME Stockport
 Genetics in Primary Care Curriculum Statement 6 Dr Dave Harniess PCME Stockport Learning Objectives Understanding of genetic component of disease Screening for genetic conditions and risk assessment in
Genetics in Primary Care Curriculum Statement 6 Dr Dave Harniess PCME Stockport Learning Objectives Understanding of genetic component of disease Screening for genetic conditions and risk assessment in
September 04, 2008
 27027 Tourney Road Valencia, CA 91355 800 421 7110 www.specialtylabs.com Test Updates September 04, 2008 Dear Valued Client: As you may be aware, in recent years there has been a tremendous challenge in
27027 Tourney Road Valencia, CA 91355 800 421 7110 www.specialtylabs.com Test Updates September 04, 2008 Dear Valued Client: As you may be aware, in recent years there has been a tremendous challenge in
Schedule of Accreditation issued by United Kingdom Accreditation Service 2 Pine Trees, Chertsey Lane, Staines-upon-Thames, TW18 3HR, UK
 2 Pine Trees, Chertsey Lane, Staines-upon-Thames, TW18 3HR, UK University Hospitals NHS Foundation Trust, Level 1, The Women s Centre John Radcliffe Hospital University Hospitals NHS Foundation Trust OX3
2 Pine Trees, Chertsey Lane, Staines-upon-Thames, TW18 3HR, UK University Hospitals NHS Foundation Trust, Level 1, The Women s Centre John Radcliffe Hospital University Hospitals NHS Foundation Trust OX3
Molecular cytogenetic evaluation of chromosomal
 http://dx.doi.org/10.4322/2357-9730.50295 Original Article Molecular cytogenetic evaluation of chromosomal microdeletions: the experience of a public hospital in Southern Brazil Mariluce Riegel 1,2, Natália
http://dx.doi.org/10.4322/2357-9730.50295 Original Article Molecular cytogenetic evaluation of chromosomal microdeletions: the experience of a public hospital in Southern Brazil Mariluce Riegel 1,2, Natália
SALSA MLPA KIT P036-E1 HUMAN TELOMERE-3 Lot 0808: As compared to the previous version (P036-D2), the probes for 1p and 4q have been replaced.
 SALSA MLPA KIT P036-E1 HUMAN TELOMERE-3 Lot 0808: As compared to the previous version (P036-D2), the probes for 1p and 4q have been replaced. MENTAL RETARDATION is caused by aberrant copy numbers of subtelomeric
SALSA MLPA KIT P036-E1 HUMAN TELOMERE-3 Lot 0808: As compared to the previous version (P036-D2), the probes for 1p and 4q have been replaced. MENTAL RETARDATION is caused by aberrant copy numbers of subtelomeric
Chromosome Mutations
 Chromosome Mutations Variation in Chromosome Number Euploidy: having full sets of chromosomes Haploid Diploid Triploid Aneuploidy: having anything other than full sets of chromosomes Monosomy Trisomy Variation
Chromosome Mutations Variation in Chromosome Number Euploidy: having full sets of chromosomes Haploid Diploid Triploid Aneuploidy: having anything other than full sets of chromosomes Monosomy Trisomy Variation
Genetic Testing for the Developmental Delay/Intellectual Disability, Autism Spectrum Disorder, and Congenital Anomalies
 Genetic Testing for the Developmental Delay/Intellectual Disability, Autism Spectrum Disorder, and Congenital Anomalies Policy Number: 2.04.59 Last Review: 5/2018 Origination: 5/2015 Next Review: 5/2019
Genetic Testing for the Developmental Delay/Intellectual Disability, Autism Spectrum Disorder, and Congenital Anomalies Policy Number: 2.04.59 Last Review: 5/2018 Origination: 5/2015 Next Review: 5/2019
The Chromosomal Basis of Inheritance
 Chapter 15 The Chromosomal Basis of Inheritance PowerPoint Lectures for Biology, Seventh Edition Neil Campbell and Jane Reece Lectures by Chris Romero Overview: Locating Genes on Chromosomes A century
Chapter 15 The Chromosomal Basis of Inheritance PowerPoint Lectures for Biology, Seventh Edition Neil Campbell and Jane Reece Lectures by Chris Romero Overview: Locating Genes on Chromosomes A century
Clinical Cytogenetics September 14 th 20 th, 2014 Goldrain, South Tyrol, Italy
 Thanks for their support to: E.C.A. European Cytogeneticists Association Clinical Cytogenetics September 14 th 20 th, 2014 Goldrain, South Tyrol, Italy DIRECTOR: A. Schinzel (Zurich, Switzerland) FACULTY:
Thanks for their support to: E.C.A. European Cytogeneticists Association Clinical Cytogenetics September 14 th 20 th, 2014 Goldrain, South Tyrol, Italy DIRECTOR: A. Schinzel (Zurich, Switzerland) FACULTY:
Information for you about Panorama s Microdeletion Screening BROUGHT TO YOU BY:
 Information for you about Panorama s Microdeletion Screening BROUGHT TO YOU BY: Panorama TM is a Non-Invasive Prenatal Test (NIPT) that screens for Down syndrome and other genetic abnormalities caused
Information for you about Panorama s Microdeletion Screening BROUGHT TO YOU BY: Panorama TM is a Non-Invasive Prenatal Test (NIPT) that screens for Down syndrome and other genetic abnormalities caused
Supplementary note: Comparison of deletion variants identified in this study and four earlier studies
 Supplementary note: Comparison of deletion variants identified in this study and four earlier studies Here we compare the results of this study to potentially overlapping results from four earlier studies
Supplementary note: Comparison of deletion variants identified in this study and four earlier studies Here we compare the results of this study to potentially overlapping results from four earlier studies
Schedule of Accreditation issued by United Kingdom Accreditation Service 2 Pine Trees, Chertsey Lane, Staines-upon-Thames, TW18 3HR, UK
 2 Pine Trees, Chertsey Lane, Staines-upon-Thames, TW18 3HR, UK Accredited to ISO 15189:2012 Laboratory Genetics Level 2B, Laboratory Medicine Queen Elizabeth University Hospital Govan Road Glasgow G51
2 Pine Trees, Chertsey Lane, Staines-upon-Thames, TW18 3HR, UK Accredited to ISO 15189:2012 Laboratory Genetics Level 2B, Laboratory Medicine Queen Elizabeth University Hospital Govan Road Glasgow G51
Human Genetic Mutations
 Human Genetic Mutations 2 Main Types of Mutations 1.) Chromosomal Mutations 2.) Gene Mutations What are chromosomes? Humans have 23 pairs of chromosomes, with one chromosome from each parent. The chromosomes
Human Genetic Mutations 2 Main Types of Mutations 1.) Chromosomal Mutations 2.) Gene Mutations What are chromosomes? Humans have 23 pairs of chromosomes, with one chromosome from each parent. The chromosomes
Pedigree Analysis. Genetic disorders. Dominant inheritance. Recessive inheritance. Autosomal vs. sex-linked traits. X-linked recessive inheritance
 Genetic disorders 4.2 Errors During Meiosis 5.3 Following Patterns of Human nheritance Pedigree Analysis 2005 Lee Bardwell Autosomal vs. sex-linked traits Autosomal traits are caused by genes on autosomes
Genetic disorders 4.2 Errors During Meiosis 5.3 Following Patterns of Human nheritance Pedigree Analysis 2005 Lee Bardwell Autosomal vs. sex-linked traits Autosomal traits are caused by genes on autosomes
LIST OF INVESTIGATIONS
 Karyotyping: K001 K002 LIST OF INVESTIGATIONS SAMPLE CONTAINER TYPE cells For Karyotyping [Single] cells For Karyotyping [Couple] Vacutainer Vacutainer 7-8 7-8 K003 Fetal Blood Sample For Karyotyping Vacutainer
Karyotyping: K001 K002 LIST OF INVESTIGATIONS SAMPLE CONTAINER TYPE cells For Karyotyping [Single] cells For Karyotyping [Couple] Vacutainer Vacutainer 7-8 7-8 K003 Fetal Blood Sample For Karyotyping Vacutainer
Invasive Prenatal (Fetal) Diagnostic Testing
 Protocol Invasive Prenatal (Fetal) Diagnostic Testing (204116) Medical Benefit Effective Date: 04/01/18 Next Review Date: 01/19 Preauthorization Yes Review Dates: 01/15, 01/16, 01/17, 01/18 Preauthorization
Protocol Invasive Prenatal (Fetal) Diagnostic Testing (204116) Medical Benefit Effective Date: 04/01/18 Next Review Date: 01/19 Preauthorization Yes Review Dates: 01/15, 01/16, 01/17, 01/18 Preauthorization
Generating Spontaneous Copy Number Variants (CNVs) Jennifer Freeman Assistant Professor of Toxicology School of Health Sciences Purdue University
 Role of Chemical lexposure in Generating Spontaneous Copy Number Variants (CNVs) Jennifer Freeman Assistant Professor of Toxicology School of Health Sciences Purdue University CNV Discovery Reference Genetic
Role of Chemical lexposure in Generating Spontaneous Copy Number Variants (CNVs) Jennifer Freeman Assistant Professor of Toxicology School of Health Sciences Purdue University CNV Discovery Reference Genetic
Original Policy Date
 MP 2.04.76 Genetic Counseling Medical Policy Section Medicine Issue 12:2013 Original Policy Date 12:2013 Last Review Status/Date Created Local Policy/ 12:2013 Return to Medical Policy Index Disclaimer
MP 2.04.76 Genetic Counseling Medical Policy Section Medicine Issue 12:2013 Original Policy Date 12:2013 Last Review Status/Date Created Local Policy/ 12:2013 Return to Medical Policy Index Disclaimer
Section: Medicine Effective Date: January15, 2016 Subsection: Pathology/Laboratory Original Policy Date: December 7, 2011 Subject:
 Section: Medicine Effective Date: January15, 2016 Last Review Status/Date: December 2015 Page: 1 of 22 Microarray Analysis and Next-Generation Autism Spectrum Disorder and/or Congenital Description Chromosomal
Section: Medicine Effective Date: January15, 2016 Last Review Status/Date: December 2015 Page: 1 of 22 Microarray Analysis and Next-Generation Autism Spectrum Disorder and/or Congenital Description Chromosomal
