Identification of bacteria directly from positive blood culture samples by DNA pyrosequencing of the 16S rrna gene
|
|
|
- Deborah Preston
- 5 years ago
- Views:
Transcription
1 Journal of Medical Microbiology (2012), 61, DOI /jmm Identification of bacteria directly from positive blood culture samples by DNA pyrosequencing of the 16S rrna gene Maiko Motoshima, 1 Katsunori Yanagihara, 1 Yoshitomo Morinaga, 1 Junichi Matsuda, 1 Hiroo Hasegawa, 1 Shigeru Kohno 2,3 and Shimeru Kamihira 1 Correspondence Katsunori Yanagihara k-yanagi@nagasaki-u.ac.jp 1 Department of Laboratory Medicine, Nagasaki University Graduate School of Biomedical Sciences, Nagasaki, Japan 2 Second Department of Internal Medicine, Nagasaki University Graduate School of Biomedical Sciences, Nagasaki, Japan 3 Global COE Program, Nagasaki University, Nagasaki, Japan Received 26 June 2012 Accepted 14 August 2012 Rapid identification of the causative bacteria of sepsis in patients can contribute to the selection of appropriate antibiotics and improvement of patients prognosis. Genotypic identification is an emerging technology that may provide an alternative method to, or complement, established phenotypic identification procedures. We evaluated a rapid protocol for bacterial identification based on PCR and pyrosequencing of the V1 and V3 regions of the 16S rrna gene using DNA extracted directly from positive blood culture samples. One hundred and two positive blood culture bottles from 68 patients were randomly selected and the bacteria were identified by phenotyping and pyrosequencing. The results of pyrosequencing identification displayed 84.3 and 64.7 % concordance with the results of phenotypic identification at the genus and species levels, respectively. In the monomicrobial samples, the concordance between the results of pyrosequencing and phenotypic identification at the genus level was 87.0 %. Pyrosequencing identified one isolate in 60 % of polymicrobial samples, which were confirmed by culture analysis. Of the samples identified by pyrosequencing, 55.7 % showed consistent results in V1 and V3 targeted sequencing; other samples were identified based on the results of V1 (12.5 %) or V3 (31.8 %) sequencing alone. One isolate was erroneously identified by pyrosequencing due to high sequence similarity with another isolate. Pyrosequencing identified one isolate that was not detected by phenotypic identification. The process of pyrosequencing identification can be completed within ~4 h. The information provided by DNA-pyrosequencing for the identification of micro-organisms in positive blood culture bottles is accurate and could prove to be a rapid and useful tool in standard laboratory practice. INTRODUCTION Bloodstream infections causing severe sepsis and septic shock result in high mortality. Detection and identification of the causative micro-organisms of sepsis are crucial for selection of the appropriate antimicrobial agents. Blood culture is an important method for the growth and subsequent identification of causative micro-organisms and diagnostic laboratories are required to detect such microorganisms as rapidly as possible. Accurate identification of bacterial isolates is also an essential task of the clinical microbiology laboratory. While traditional phenotypic Abbreviation: MALDI-TOF MS, matrix-assisted laser desorption/ionization time-of-flight mass spectrometry. identification is universally used in clinical laboratories, this method has some disadvantages, for example, it is time consuming, is sometimes difficult and does not always accurately identify target micro-organisms. In addition, interpretation of the results obtained using phenotypic methods can be subjective (Stager & Davis, 1992). Genotypic identification of micro-organisms is an emerging technology that may provide an alternative or complementary method to established phenotypic identification procedures. Sequence analysis of the 16S rrna gene is a widely accepted tool for molecular identification of bacteria (Kolbert & Persing, 1999; Patel, 2001; Woese, 1987). Bacterial 16S rrna genes consist of eight highly conserved regions and nine variable regions (Woese, 1987) G 2012 SGM Printed in Great Britain
2 Pyrosequencing of 16S rrna gene from blood cultures V1 and V3 are two distinct variable regions of the 16S rrna gene that have been used as targets for sequencingbased identification assays (Luna et al., 2007). This method capitalizes on the highly conserved nature of 16S rrna genes by positioning amplification and sequencing primers in the conserved regions that flank the variable regions (specifically V1 and V3), thereby allowing primers to theoretically amplify most bacterial pathogens. Public databases such as GenBank, the Nucleotide Sequence Database at the European Molecular Biology Laboratory (EMBL-Bank), the DNA Data Bank of Japan (DDBJ) and the Ribosomal Database Project II (RDP II) contain a vast number of bacterial 16S rrna gene sequences, allowing rapid analysis and providing phylogenetically meaningful information (Bosshard et al., 2006). Pyrosequencing is a DNA sequencing technique that is based on the detection of pyrophosphate that is released during DNA synthesis and was introduced as a rapid alternative to traditional Sanger DNA sequencing (Ronaghi et al., 1996). The DNA base sequence is determined by measuring the strength of visible light that is generated in proportion to the number of incorporated nucleotides in a cascade of enzymic reactions (Ronaghi, 2001). The main advantage of pyrosequencing is its rapidity and lower price compared to conventional sequencing. Although the length of the sequence that can be obtained by pyrosequencing is fairly short and limited to about bases, carefully designed applications can provide information that is sufficient for the differentiation of gene sequences. Pyrosequencing has already been applied to the identification of bacteria (Jonasson et al., 2002; Luna et al., 2007; Ronaghi & Elahi, 2002). It has also been predicted that pyrosequencing of the 16S rrna gene may function as a molecular Gram stain for the identification of bacteria (Jordan et al., 2005). In order to effectively treat patients with sepsis, the rapid identification of causative bacteria is important; however, conventional phenotyping-based identification requires an extra day after blood culture tests become positive. Therefore, the rapidity of pyrosequencing-based identification is an attractive advantage for diagnosis. In fact, Jordan et al. (2009) reported that the combination of real-time PCR and pyrosequencing methods rapidly identified bacteria from positive blood culture samples and provided highly concordant results with those obtained using phenotypic identification. To rapidly identify clinical isolates from positive blood culture samples, we evaluated a rapid protocol for the identification of micro-organisms using PCR and pyrosequencing of 16S rrna genes. Using clinical samples from our hospital, we compared this molecular bacterial identification protocol with conventional culture identification techniques. METHODS Sample collection. This study was performed at the Nagasaki University Hospital, a tertiary hospital with about 850 beds, and was approved by the ethics committee of the Nagasaki University Hospital. Positive blood culture samples from both paediatrics and adults were randomly selected from blood culture bottles that were submitted during 2010 for routine microbiological testing. Blood sampling was performed according to the recommended methods in the hospital and 5 10 ml blood was collected into the each bottle. Bottles containing samples from the same patient but at different time points were excluded. Blood culture and phenotypic identification. Blood samples collected in BacT/ALERT FA or BacT/ALERT FN bottles (biomérieux) at the Nagasaki University Hospital were cultured using BacT/ALERT 3D (biomérieux), which is an automated microbial detection system that displays a positive result if microbial growth is detected by a fluorescent sensor. Each bottle was removed from the blood culture instrument within 12 h following a positive result and.1 ml samples were immediately extracted from the bottle. The sample was Gram stained and subcultured on the appropriate agar-based culture plates. All samples were identified according to standard biochemical identification methods using the VITEK 2 system (biomérieux) or the Phoenix100 system (Becton Dickinson). DNA extraction and amplification. Bacterial DNA was extracted directly from 1 ml blood culture fluid using the BiOstic bacteraemia DNA isolation kit (MoBio Laboratories) according to the manufacturer s instructions. Chromosomal DNA was eluted in a final volume of 50 ml elution buffer. The V1 (amplicon size 115 bp) and the V3 (amplicon size 81 bp) regions of 16S rrna genes were amplified according to a previously published method (Luna et al., 2007). Nucleotide positions refer to positions in the 16S rrna gene of Escherichia coli. Primers Bio-pBR5 (59-biotin-GAAGAGTTTGATCAT- GGCTCAG-39) and pbr-v1 (59-TTACTCACCCGTCCGCCACT-39) were used for amplification of the V1 region and Bio-B-V3 (59-biotin- ACGACAGCCATGCAGCACCT-39) and pjbs.v3 (59-GCAACGCGA- AGAACCTTACC-39) were used for amplification of the V3 region. Each 50 ml reaction mixture contained 25 ml Ampdirect (Shimadzu), 0.2 M each primer, 1.25 U AmpliTaq Gold DNA polymerase LD (Life Technologies) and 5 ml DNA template. PCRs were performed using the GeneAmp PCR system 9700 (Life Technologies) with the following cycling parameters: 95 uc for 10 min, followed by 35 cycles of 95 uc for 40 s, 55 uc for 40 s and 72 uc for 60 s, with a final elongation step at 72 uc for 60 s. DNA extracted from clinical isolates, including Staphylococcus aureus, Bacillus cereus and Escherichia coli, were used as positive controls for PCR and sterile water was used as a negative control. Each PCR product was verified by agarose gel electrophoresis. Samples without separate bands were amplified after 10- or 100-fold dilution and reconfirmed by agarose gel electrophoresis. DNA pyrosequencing. The amplified products of the 16S rrna V1 and V3 regions were prepared for pyrosequencing by using the vacuum prep tool (Qiagen) following the recommended protocol. For the preparation of each reaction, 40 ml of the biotinylated PCR product was used. To prepare the sequencing plate, purified PCR products were resuspended in 43 ml binding buffer and 3 ml streptavidin beads. Double-stranded DNA was then denatured to single-stranded DNA using 0.2 M NaOH. Subsequently, singlestranded DNA was resuspended in 40 ml annealing buffer with 0.3 mm sequencing primer and then annealed to the sequencing primer at 80 uc for 2 min. The primers pbr-v1 and pjbs.v3 were used as described above as DNA sequencing primers for the V1 and V3 regions, respectively. Pyrosequencing was performed on a PyroMark ID instrument (Qiagen) with eight cycles of a repetitive ACTG dispensation. Sequence similarities of the PCR products were determined using the DDBJ search program ( and a strain with.99 % sequence similarity was considered as an isolated strain
3 M. Motoshima and others RESULTS Culture results In the present study, 102 samples collected from 68 patients were cultured and 112 bacteria and one fungus (Candida albicans) were isolated. Two different microorganisms were isolated from each of 10 (9.8 %) samples collected from a total of seven cases. The cultured bacteria included a total of 15 genera and 28 species. One isolate was not identified and is referred to as an anaerobic Grampositive rod. Detection and identification of micro-organisms by DNA-pyrosequencing All 102 samples were successfully amplified by PCR, targeting the V1 or V3 region of the 16S rrna gene. Four samples required dilution for amplification of the products because of inhibition. DNA pyrosequencing-based identification was then performed using these PCR products. From the 102 samples, 88 (86.3 %) strains were detected to the genus level and 68 (66.7 %) strains were detected to the species level by DNA pyrosequencing, comprising 16 genera and 19 species, including 41 Gram-positive cocci, nine Gram-positive bacilli, 34 Gram-negative bacilli and four anaerobic organisms. Of the 68 patient cases, isolates from 61 (89.7 %) cases were detected to the genus level and isolates from 49 (72.1 %) cases were detected to the species level. Identification of isolates based on culture and pyrosequencing of V1 and V3 regions Pyrosequencing results corresponding to each culture-based organism were analysed. Of the 92 monomicrobial samples, 21 strains identified by pyrosequencing showed complete agreement with the results of culture-based identification with both V1 and V3 targeted pyrosequencing methods (Fig. 1) providing identification to the species level (Table 1), identifying strains of Staphylococcus aureus, Staphylococcus epidermidis, Corynebacterium striatum, Escherichia coli and Pseudomonas aeruginosa. The other concordant isolates that were identified to the species level were dependent on the results of either V1 (20 isolates) or V3 (22 isolates) region sequencing. In two isolates, pyrosequencing identified different strains from the culture-based identification method. One of these isolates was identified by both V1 and V3 region targeted sequencing and the other was dependent on a result from V3 region sequencing alone. In the polymicrobial samples, pyrosequencing failed to completely identify all of the bacteria that were identified by the culture method. However, one organism was identified in each of six (60 %) of these samples, and three of these six strains were identified to species level by pyrosequencing (Table 2). One strain was identified based on the results of both V1 and V3 region sequencing and the others were identified based on V1 sequencing alone. However, in some samples, the 16S rrna gene targets could not be used to successfully identify the bacterium at both the species and genus levels. Strains of the genus Enterobacter as well as strains of Bacteroides fragilis, Fusobacterium nucleatum and Bifidobacterium scardovii were not detected by the sequencing of the V1 region. In contrast, pyrosequencing of the V3 region failed to detect Citrobacter freundii and strains of the genus Clostridium. Concordance rate of sequence-based identification The percentage of concordance between the results of culturebased and pyrosequence-based identification was calculated (Table 3). Of the 92 monomicrobial samples identified by culture, 80 (87.0 %) samples identified at genus level and 63 (68.5 %) samples identified at species level were concordant with pyrosequence-based identifications. Two (2.2 %) samples showed discordant results and 10 (10.9 %) samples were unidentified by pyrosequencing. Ten samples were confirmed as polymicrobial by culturebased identification and pyrosequencing identified one micro-organism at the genus level in six of these samples and one micro-organism at species level in three of these samples, all of which were concordant with the results of culture based identification. Pyrosequencing did not detect two or more micro-organisms in any of the samples. The overall agreement between the results of culture- and pyrosequence-based identification was 84.3 % (86/102) at the genus level and 64.7 % (66/102) at the species level. Analysis of discordant results Two samples displayed discordant results following identification by DNA pyrosequencing and phenotyping. In one sample, the isolate was determined as Staphylococcus epidermidis by pyrosequencing; however, the characteristics of this isolate were inconsistent with biochemical data of Staphylococcus epidermidis and the Phoenix100 and VITEK2 system analyses identified this isolate as Staphylococcus capitis with 99 % probability. 16S rrna gene sequence similarity between Staphylococcus epidermidis and Staphylococcus capitis is 99 %. The pyrosequencing result was therefore interpreted as a false-positive result. In the other sample, the isolate was identified as Bifidobacterium scardovii by pyrosequencing but the conventional culture method identified it simply as an anaerobic Grampositive rod that could not be further classified because of poor data regarding morphological and biochemical characteristics. This isolate was ultimately determined as B. scardovii after confirmation following reproducibility testing. DISCUSSION Rapid identification of the causative bacteria of sepsis in patients allows the appropriate antibiotics to be selected 1558 Journal of Medical Microbiology 61
4 Pyrosequencing of 16S rrna gene from blood cultures (a) (b) E S G A C T G A C T G A C T G A C T G A C T G A C T G A C T G A C T G A C T G A C T G A C T G A C T G A C T G A C T G A C T GAATCCAGGAGCAAGCCCCTTCCTATCGCCTCGACTGCTGACT E S G A C T G A C T G A C T G A C T G A C T G A C T G A C T G A C T G A C T G A C T G A C T G A C T G A C T G A C T G A C T TGGCCTTGACATGCTGAGAACTTTCCAGAGATGGATTGGTGCCTTC Fig. 1. A representative pyrogram of targeted 16S rrna gene V1 (a) and V3 (b) regions. The sequence results are shown at the bottom of each pyrogram. This sample was identified as P. aeruginosa after a sequence similarity search. (Barenfanger et al., 1999) and improves prognosis (Barenfanger et al., 2001). Bacterial identification based on genetic methods can provide information that is useful for the selection of targeted antibiotics. The overall results for the identification of isolates by pyrosequencing and culture-based methods agreed in 84.3 % and 64.7 % of samples for identification at the genus and species levels, respectively. A previous study of DNA pyrosequencing identification using pure-cultured isolates reported ~90 % agreement between the isolates identified by the two methods (Luna et al., 2007). Considering that DNA was extracted directly from blood culture bottle fluids, we believe that the concordance between the results of the two methods used in the present study is reasonable. In addition, pyrosequencing resulted in only one error in sample identification, which was due to very high sequence similarity between two bacteria, implying that DNApyrosequencing is a very accurate method for the identification of bacteria. These results suggest that the identification of bacteria in positive blood culture samples by DNApyrosequencing will be useful for the evaluation of clinical samples and may influence the choice appropriate antimicrobial treatment, benefitting patient outcome. Highly accurate pyrosequencing identification of bacteria from positive blood culture bottles has been reported previously (Jordan et al., 2009). In this report, the 23S rrna gene was used as the target to improve the accuracy of identification of some specific bacteria, such as members of the family Enterobacteriaceae and the genus Streptococcus,andthe agreement between the results of pyrosequencing- and culture-based identification reached 97.8 %. In the present study, the rate of concordance between the two techniques was lower. This was partly because a larger number of polymicrobial samples were included in the present study than were present in the study by Jordan et al. (2009). Furthermore, the relatively large number of specific strains that went undetected, such as members of the genus Enterobacter, could also decrease the rate of concordance of this study. In the present study, pyrosequencing of the V1 and V3 regions produced similar results in many samples but highlighted different characteristics in some specific groups of bacteria. The V1 region can effectively be used to identify members of the genus Enterococcus and distinguish between E. faecalis and E. faecium; conversely, using the V3 region can have advantages in detecting S. epidermidis and E. coli. This suggests that sequencing both the V1 and V3 regions improves the accuracy of diagnosis. However, the optimal combination of variable regions of the 16S rrna gene for use in diagnosis has been a controversial issue (Sundquist et al., 2007; Wang et al., 2007). Conventional biochemical testing, especially for difficultto-identify pathogens, may result in incorrect pathogen identification, resulting in inconsistent information for the physician (Downes et al., 1998; Stager & Davis, 1992). Molecular methods provide novel strategies for bacterial pathogen identification (Tang et al., 1998). 16S rrna gene sequencing has previously been reported to detect relevant isolates of non-fermenting Gram-negative bacilli at high rates compared to phenotypic identification techniques. The reason for the low rate of phenotypic identification was that nearly half of the isolates that corresponded to species based on sequencing data were not included in the databases of conventional phenotypic identification systems (Bosshard et al., 2006). Molecular methods are considered useful for identification of Gram-positive bacteria or anaerobes as well as Gram-negative bacteria. In the present study, Bifidobacterium scardovii was identified by the genetic method but not by the usual laboratory procedures. Therefore, the genetic method described in this study may complement current methods of phenotypic identification. Although the method described in this study is considered to be a useful and convenient procedure for the rapid identification of micro-organisms, it also has some limitations in terms of efficacy of identification. First, pyrosequencing can fail to separate distinct bacteria that have similar sequences because it provides only short sequence lengths. Members of the genera Aeromonas, Bacillus and Staphylococcus typically have similar sequences in the target gene to other members within their respective genus. Therefore, organisms that belong to these genera were not
5 M. Motoshima and others Table 1. Distribution of pyrosequencing identification results in monomicrobial samples Values represent numbers of isolates identified by pyrosequencing, targeting either V1+V3, V1 or V3 regions, to the species level or to the genus level. Isolates identified only to the genus level by pyrosequencing and those results that were discordant or undetected by pyrosequencing were assigned to a species based on the results of culture-based identification. Taxa No. of isolates Number of concordant samples Species level Genus level Number of discordant samples Undetected V1+V3 V1 V3 Gram-positive cocci Enterococcus faecalis Enterococcus faecium Staphylococcus aureus Staphylococcus capitis * 1 Staphylococcus epidermidis Staphylococcus haemolyticus Staphylococcus hominis Staphylococcus schleifreri Staphylococcus simulans Staphylococcus agalactiae Gram-positive bacilli Bacillus cereus Bacillus thuringiensis Corynebacterium striatum Gram-negative bacilli Aeromonas hydrophila Aeromonas sobria Citrobacter freundii Enterobacter aerogenes Enterobacter cloacae Escherichia coli Haemophilus influenzae Klebsiella oxytoca Klebsiella pneumoniae Pseudomonas aeruginosa Pseudomonas putida Others (anaerobes) Bacteroides fragilis Fusobacterium nucleatum Genus Veillonella Anaerobic Gram-positive rod D 0 Total *This isolate was misidentified as Staphylococcus epidermidis by both V1 and V3 sequencing. DThis isolate was identified as Bifidobacterium scardovii by V3 sequencing. effectively identified at the species level by pyrosequencing but showed good agreement with the results of culturebased identification at the genus level. Other specific sequencing targets are required to identify the correct species of these bacteria. However, with the exception of the genus Staphylococcus, members of these genera are rarely isolated and their antibiotic resistance has not become problematic. Therefore, genetic methods to identify these bacteria at the species level may not be considered necessary. Second, in polymicrobial infections, pyrosequencing may not identify all of the different bacteria. Thus, when a sample for pyrosequencing contains polymicrobial genes, the result obtained from sequencing can consist of a mix of sequences from those micro-organisms. Therefore, pyrosequencing may not effectively detect organisms in patients with polymicrobial infections. Among the bacteria that were undetectable by pyrosequencing, some species, such as Enterobacter cloacae, Enterococcus faecalis, Bacteroides fragilis 1560 Journal of Medical Microbiology 61
6 Pyrosequencing of 16S rrna gene from blood cultures Table 2. Identification of micro-organisms in the 10 polymicrobial samples Phenotypic identification Pyrosequencing identification Result No. of samples Result Number of concordant samples* Undetected Species level Genus level Bacillus thuringiensis/ Staphylococcus saprophyticus Bacteroides fragilis/ Clostridium clostridiofome Enterobacter cloacae/ Enterococcus feacalis Enterobacter cloacae/ Genus Bacteroides Enterobacter cloacae/ Staphylococcus aureus Enterococcus faecium/ Staphylococcus heamolyticus Staphylococcus aureus/ Candida albicans 2 Genus Bacillus 0 2D 0 1 Genus Clostridium 0 1d 0 2 Undetected Undetected Staphylococcus aureus Enterococcus faecium 2d Undetected *When at least one of the micro-organisms was identified by both identification methods, to either the genus or the species level, the sample was considered concordant. DIdentification based on V3 sequence. didentification based on V1 sequence. Identification based on both V1 and V3 sequence. and Bacillus thuringiensis, were commonly observed and the samples that included these isolates were often polymicrobial. Most of the bacteria that were not detected by pyrosequencing in this study are commonly known as causative pathogens of intra-abdominal and urinary tract infections, in which polymicrobial infections are often observed (Reuben et al., 1989). Matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS) has been used as an accurate identification tool with fast and cost-effective benefits. Identification of bacteria directly from positive blood cultures by using MALDI-TOF MS has also been attempted and % of bacteria were correctly identified to the species level (Christner et al., 2010; Wimmer et al., 2012; Wüppenhorst et al., 2012). However, some bacteria including E. coli and species of the genus Shigella are known to be indistinguishable by MALDI-TOF MS. Members of the genus Streptococcus are also not reliably identifiable to the species level by using MALDI-TOF MS or 16S rrna pyrosequencing. Therefore, these rapid protocols require additional procedures to identify these bacteria correctly. The process of pyrosequencing identification of bacteria that was used in the present study, including sample preparation, the sequencing reaction and analysis of the Table 3. Summary of DNA pyrosequencing results and the concordance with results of phenotypic identification Results of pyrosequencing identification Number of isolates (%) Monomicrobial (n592) Polymicrobial (n510)* Total (n5102) Genus level Concordant 80 (87.0) 6 (60.0) 86 (84.3) Discordant 2 (2.2) 0 (0.0) 2 (2.0) Undetected 10 (10.9) 4 (40.0) 14 (13.7) Species level Concordant 63 (68.5) 3 (30.0) 66 (64.7) Discordant 2 (2.2) 0 (0.0) 2 (2.0) Undetected 27 (29.3) 7 (70.0) 34 (33.3) *When at least one isolate gave the same result, the sample was considered concordant
7 M. Motoshima and others results, can be completed within ~4 h. Repeated sequencing from the same sample bottle provided consistent results. This method would therefore be relatively easy to fit into a standard laboratory routine and obtaining information regarding the isolate within a day would be of great help in improving the outcome of sepsis. ACKNOWLEDGEMENTS This research was partially supported by a Grant-in-Aid for Scientific Research from the Japanese Ministry of Education, Culture, Sports, Science and Technology (no ) to K Y, and a grant from the Global Centers of Excellence Program, Nagasaki University. REFERRENCES Barenfanger, J., Drake, C. & Kacich, G. (1999). Clinical and financial benefits of rapid bacterial identification and antimicrobial susceptibility testing. J Clin Microbiol 37, Barenfanger, J., Short, M. A. & Groesch, A. A. (2001). Improved antimicrobial interventions have benefits. J Clin Microbiol 39, Bosshard, P. P., Zbinden, R., Abels, S., Böddinghaus, B., Altwegg, M. &Böttger, E. C. (2006). 16S rrna gene sequencing versus the API 20 NE system and the VITEK 2 ID-GNB card for identification of nonfermenting Gram-negative bacteria in the clinical laboratory. J Clin Microbiol 44, Christner,M.,Rohde,H.,Wolters,M.,Sobottka,I.,Wegscheider,K.& Aepfelbacher, M. (2010). Rapid identification of bacteria from positive blood culture bottles by use of matrix-assisted laser desorptionionization time of flight mass spectrometry fingerprinting. J Clin Microbiol 48, Downes, A., Thornhill, C. & Mallard, R. H. (1998). An evaluation of Vitek an automated system for bacterial identification and susceptibility testing. Commun Dis Public Health 1, Jonasson, J., Olofsson, M. & Monstein, H. J. (2002). Classification, identification and subtyping of bacteria based on pyrosequencing and signature matching of 16S rdna fragments. APMIS 110, Jordan, J. A., Butchko, A. R. & Durso, M. B. (2005). Use of pyrosequencing of 16S rrna fragments to differentiate between bacteria responsible for neonatal sepsis. J Mol Diagn 7, Jordan, J. A., Jones-Laughner, J. & Durso, M. B. (2009). Utility of pyrosequencing in identifying bacteria directly from positive blood culture bottles. J Clin Microbiol 47, Kolbert, C. P. & Persing, D. H. (1999). Ribosomal DNA sequencing as a tool for identification of bacterial pathogens. Curr Opin Microbiol 2, Luna,R.A.,Fasciano,L.R.,Jones,S.C.,Boyanton,B.L.,Jr,Ton,T.T.& Versalovic, J. (2007). DNA pyrosequencing-based bacterial pathogen identification in a pediatric hospital setting. JClinMicrobiol45, Patel, J. B. (2001). 16S rrna gene sequencing for bacterial pathogen identification in the clinical laboratory. Mol Diagn 6, Reuben, A. G., Musher, D. M., Hamill, R. J. & Broucke, I. (1989). Polymicrobial bacteremia: clinical and microbiologic patterns. Rev Infect Dis 11, Ronaghi, M. (2001). Pyrosequencing sheds light on DNA sequencing. Genome Res 11, Ronaghi, M. & Elahi, E. (2002). Pyrosequencing for microbial typing. J Chromatogr B Analyt Technol Biomed Life Sci 782, Ronaghi, M., Karamohamed, S., Pettersson, B., Uhlén, M. & Nyrén, P. (1996). Real-time DNA sequencing using detection of pyrophosphate release. Anal Biochem 242, Stager, C. E. & Davis, J. R. (1992). Automated systems for identification of microorganisms. Clin Microbiol Rev 5, Sundquist, A., Bigdeli, S., Jalili, R., Druzin, M. L., Waller, S., Pullen, K. M., El-Sayed, Y. Y., Taslimi, M. M., Batzoglou, S. & Ronaghi, M. (2007). Bacterial flora-typing with targeted, chip-based pyrosequencing. BMC Microbiol 7, 108. Tang, Y. W., Ellis, N. M., Hopkins, M. K., Smith, D. H., Dodge, D. E. & Persing, D. H. (1998). Comparison of phenotypic and genotypic techniques for identification of unusual aerobic pathogenic Gramnegative bacilli. J Clin Microbiol 36, Wang, Q., Garrity, G. M., Tiedje, J. M. & Cole, J. R. (2007). Naive Bayesian classifier for rapid assignment of rrna sequences into the new bacterial taxonomy. Appl Environ Microbiol 73, Wimmer, J. L., Long, S. W., Cernoch, P., Land, G. A., Davis, J. R., Musser, J. M. & Olsen, R. J. (2012). Strategy for rapid identification and antibiotic susceptibility testing of Gram-negative bacteria directly recovered from positive blood cultures using the Bruker MALDI Biotyper and the BD Phoenix system. J Clin Microbiol 50, Woese, C. R. (1987). Bacterial evolution. Microbiol Rev 51, Wüppenhorst, N., Consoir, C., Lörch, D. & Schneider, C. (2012). Direct identification of bacteria from charcoal-containing blood culture bottles using matrix-assisted laser desorption/ionisation time-of-flight mass spectrometry. Eur J Clin Microbiol Infect Dis (in Press) Journal of Medical Microbiology 61
DNA pyrosequencing of the 16S rrna
 NAOSITE: Nagasaki University's Ac Title Author(s) Identification of bacteria directly DNA pyrosequencing of the 16S rrna Motoshima, Maiko; Yanagihara, Katsu Matsuda, Junichi; Hasegawa, Hiroo; Citation
NAOSITE: Nagasaki University's Ac Title Author(s) Identification of bacteria directly DNA pyrosequencing of the 16S rrna Motoshima, Maiko; Yanagihara, Katsu Matsuda, Junichi; Hasegawa, Hiroo; Citation
OVERVIEW OF CURRENT IDENTIFICATION SYSTEMS AND DATABASES
 OVERVIEW OF CURRENT IDENTIFICATION SYSTEMS AND DATABASES EVERY STEP OF THE WAY 1 EVERY STEP OF THE WAY MICROBIAL IDENTIFICATION METHODS DNA RNA Genotypic Sequencing of ribosomal RNA regions of bacteria
OVERVIEW OF CURRENT IDENTIFICATION SYSTEMS AND DATABASES EVERY STEP OF THE WAY 1 EVERY STEP OF THE WAY MICROBIAL IDENTIFICATION METHODS DNA RNA Genotypic Sequencing of ribosomal RNA regions of bacteria
MALDI-TOF mass spectrometry for direct bacterial identification
 JCM Accepts, published online ahead of print on 17 February 2010 J. Clin. Microbiol. doi:10.1128/jcm.01780-09 Copyright 2010, American Society for Microbiology and/or the Listed Authors/Institutions. All
JCM Accepts, published online ahead of print on 17 February 2010 J. Clin. Microbiol. doi:10.1128/jcm.01780-09 Copyright 2010, American Society for Microbiology and/or the Listed Authors/Institutions. All
Optimizing MALDI-TOF Use. Clinical Impact Laboratory Impact
 Optimizing MALDI-TOF Use Clinical Impact Laboratory Impact Christine C. Ginocchio, PhD, MT (ASCP) Clinical Professor of Medicine Hofstra North Shore-LIJ School of Medicine, NY VP, Global Microbiology Affairs,
Optimizing MALDI-TOF Use Clinical Impact Laboratory Impact Christine C. Ginocchio, PhD, MT (ASCP) Clinical Professor of Medicine Hofstra North Shore-LIJ School of Medicine, NY VP, Global Microbiology Affairs,
Medical Microbiology, University Hospital of Liège, Liège, Belgium
 Journal of Medical Microbiology (2012), 61, 1511 1516 DOI 10.1099/jmm.0.044750-0 Direct identification of bacteria from BacT/ALERT anaerobic positive blood cultures by MALDI-TOF MS: MALDI Sepsityper kit
Journal of Medical Microbiology (2012), 61, 1511 1516 DOI 10.1099/jmm.0.044750-0 Direct identification of bacteria from BacT/ALERT anaerobic positive blood cultures by MALDI-TOF MS: MALDI Sepsityper kit
Comparative study of MALDI-TOF MS and VITEK 2 in bacteria identification
 MALDI-TOF MS in Clinical Microbiology Comparative study of MALDI-TOF MS and VITEK 2 in bacteria identification Ling Guo, Liyan Ye, Qiang Zhao, Yanning Ma, Jiyong Yang, Yanping Luo Department of Microbiology,
MALDI-TOF MS in Clinical Microbiology Comparative study of MALDI-TOF MS and VITEK 2 in bacteria identification Ling Guo, Liyan Ye, Qiang Zhao, Yanning Ma, Jiyong Yang, Yanping Luo Department of Microbiology,
Direct identification of bacteria from positive anaerobic BacT/Alert blood cultures by. Contents Category: Diagnostics, typing and identification
 1 2 3 Direct identification of bacteria from positive anaerobic BacT/Alert blood cultures by MALDI-TOF MS: MALDI Sepsityper kit (Bruker) versus in-house saponin method for bacterial extraction 4 5 Running
1 2 3 Direct identification of bacteria from positive anaerobic BacT/Alert blood cultures by MALDI-TOF MS: MALDI Sepsityper kit (Bruker) versus in-house saponin method for bacterial extraction 4 5 Running
Brief Communication Clinical Microbiology
 Brief Communication Clinical Microbiology Ann Lab Med 2017;37:531-535 https://doi.org/10.3343/alm.2017.37.6.531 ISSN 2234-3806 eissn 2234-3814 Comparison of a New Matrix-Assisted Laser Desorption/Ionization
Brief Communication Clinical Microbiology Ann Lab Med 2017;37:531-535 https://doi.org/10.3343/alm.2017.37.6.531 ISSN 2234-3806 eissn 2234-3814 Comparison of a New Matrix-Assisted Laser Desorption/Ionization
Cell counting (EtBr) Before cell-lysis. Cell-lysis by 3% SDS beads beating. After cell-lysis
 Key words Sample Cell counting (EtBr) DNA extraction Cell-lysis by 3% SDS beads beating Before cell-lysis After cell-lysis Cell-lysis efficiency (Maintained70%) PCR PCR amplification of partial fragments
Key words Sample Cell counting (EtBr) DNA extraction Cell-lysis by 3% SDS beads beating Before cell-lysis After cell-lysis Cell-lysis efficiency (Maintained70%) PCR PCR amplification of partial fragments
VITEK MS Implementation of Mass Spectrometry
 VITEK MS Implementation of Mass Spectrometry Willson Jang, Technical Leader wljang@providencehealth.bc.ca Microbiology & Virology Laboratory Providence Health Care St. Paul s Hospital October 23, 2013
VITEK MS Implementation of Mass Spectrometry Willson Jang, Technical Leader wljang@providencehealth.bc.ca Microbiology & Virology Laboratory Providence Health Care St. Paul s Hospital October 23, 2013
Tandem MS in Microbiology. Chris Doern, PhD D(ABMM) UT Southwestern Medical Center Dallas, TX
 Tandem MS in Microbiology Chris Doern, PhD D(ABMM) UT Southwestern Medical Center Dallas, TX Learning Objectives After the presentation you should be able to: 1. Understand the mass spectrometry methodology
Tandem MS in Microbiology Chris Doern, PhD D(ABMM) UT Southwestern Medical Center Dallas, TX Learning Objectives After the presentation you should be able to: 1. Understand the mass spectrometry methodology
Normal Human Flora. (Human Microbiome) Dr.Sarmad M.H. Zeiny Baghdad College of Medicine
 Normal Human Flora (Human Microbiome) Dr.Sarmad M.H. Zeiny Baghdad College of Medicine 2014-2015 Objectives Describe important human normal flora. Demonstrate the epidemiology of human normal flora. Determine
Normal Human Flora (Human Microbiome) Dr.Sarmad M.H. Zeiny Baghdad College of Medicine 2014-2015 Objectives Describe important human normal flora. Demonstrate the epidemiology of human normal flora. Determine
RESEARCH ARTICLE Torres et al., Journal of Medical Microbiology 2017;66: DOI /jmm
 RESEARCH ARTICLE Torres et al., Journal of Medical Microbiology 2017;66:1752 1758 DOI 10.1099/jmm.0.000643 Short-term incubation of positive blood cultures in brain-heart infusion broth accelerates identification
RESEARCH ARTICLE Torres et al., Journal of Medical Microbiology 2017;66:1752 1758 DOI 10.1099/jmm.0.000643 Short-term incubation of positive blood cultures in brain-heart infusion broth accelerates identification
Matrix-assisted laser desorption/ionization time-of-flight mass spectrometry for the identification of beta-hemolytic streptococci
 Original Article Matrix-assisted laser desorption/ionization time-of-flight mass spectrometry for the identification of beta-hemolytic streptococci Chunmei Zhou 1 *, Lili Tao 2 *, Bijie Hu 3, Jian Ma 4,
Original Article Matrix-assisted laser desorption/ionization time-of-flight mass spectrometry for the identification of beta-hemolytic streptococci Chunmei Zhou 1 *, Lili Tao 2 *, Bijie Hu 3, Jian Ma 4,
Rapid identification and resistance assessment: The future is mass spectrometry
 Rapid identification and resistance assessment: The future is mass spectrometry Dr Sanmarié Schlebusch Director of Microbiology Mater Pathology Brisbane Outline Introduction Plug and play Pre-prep and
Rapid identification and resistance assessment: The future is mass spectrometry Dr Sanmarié Schlebusch Director of Microbiology Mater Pathology Brisbane Outline Introduction Plug and play Pre-prep and
Rapid identification of bacteria from positive blood culture bottles by MALDI-TOF MS following short-term incubation on solid media
 Journal of Medical Microbiology (2015), 64, 1346 1352 DOI 10.1099/jmm.0.000168 Rapid identification of bacteria from positive blood culture bottles by MALDI-TOF MS following short-term incubation on solid
Journal of Medical Microbiology (2015), 64, 1346 1352 DOI 10.1099/jmm.0.000168 Rapid identification of bacteria from positive blood culture bottles by MALDI-TOF MS following short-term incubation on solid
ESCMID Online Lecture Library. by author
 Microbiological evaluation: how to report the results Alvaro Pascual MD, PhD Infectious Diseases and Clinical Microbiology Unit. University Hospital Virgen Macarena University of Sevilla BSI management
Microbiological evaluation: how to report the results Alvaro Pascual MD, PhD Infectious Diseases and Clinical Microbiology Unit. University Hospital Virgen Macarena University of Sevilla BSI management
MALDI-TOF Mass Spectrometry: A New Rapid ID Method in Clinical Microbiology
 MALDI-TOF Mass Spectrometry: A New Rapid ID Method in Clinical Microbiology Patrick R. Murray, PhD WW Director, Scientific Affairs BD Diagnostic Systems Outline MALDI-TOF is the most important innovation
MALDI-TOF Mass Spectrometry: A New Rapid ID Method in Clinical Microbiology Patrick R. Murray, PhD WW Director, Scientific Affairs BD Diagnostic Systems Outline MALDI-TOF is the most important innovation
Original Article Clinical Microbiology INTRODUCTION
 Original Article Clinical Microbiology Ann Lab Med 2015;35:416-422 http://dx.doi.org/10.3343/alm.2015.35.4.416 ISSN 2234-3806 eissn 2234-3814 Direct Identification of Urinary Tract Pathogens From Urine
Original Article Clinical Microbiology Ann Lab Med 2015;35:416-422 http://dx.doi.org/10.3343/alm.2015.35.4.416 ISSN 2234-3806 eissn 2234-3814 Direct Identification of Urinary Tract Pathogens From Urine
Received 1 March 2011/Returned for modification 19 April 2011/Accepted 20 May 2011
 JOURNAL OF CLINICAL MICROBIOLOGY, Aug. 2011, p. 2980 2984 Vol. 49, No. 8 0095-1137/11/$12.00 doi:10.1128/jcm.00431-11 Copyright 2011, American Society for Microbiology. All Rights Reserved. Utility of
JOURNAL OF CLINICAL MICROBIOLOGY, Aug. 2011, p. 2980 2984 Vol. 49, No. 8 0095-1137/11/$12.00 doi:10.1128/jcm.00431-11 Copyright 2011, American Society for Microbiology. All Rights Reserved. Utility of
A Strategy for Rapid Identification and Antibiotic Susceptibility. Testing of Gram-Negative Bacteria Directly Recovered from Positive
 JCM Accepts, published online ahead of print on 18 April 2012 J. Clin. Microbiol. doi:10.1128/jcm.00409-12 Copyright 2012, American Society for Microbiology. All Rights Reserved. 1 1 2 3 4 5 6 7 8 9 10
JCM Accepts, published online ahead of print on 18 April 2012 J. Clin. Microbiol. doi:10.1128/jcm.00409-12 Copyright 2012, American Society for Microbiology. All Rights Reserved. 1 1 2 3 4 5 6 7 8 9 10
Torres, Brian Carlmichael L., Ma. Luisa G. Daroy, Juan s. Lopez, Vanessa Oh, Marie Joan Loy, Prospero ma. Tuaño, and Ronald R. Matias Research and
 Torres, Brian Carlmichael L., Ma. Luisa G. Daroy, Juan s. Lopez, Vanessa Oh, Marie Joan Loy, Prospero ma. Tuaño, and Ronald R. Matias Research and Biotechnology Division and International Eye Institute
Torres, Brian Carlmichael L., Ma. Luisa G. Daroy, Juan s. Lopez, Vanessa Oh, Marie Joan Loy, Prospero ma. Tuaño, and Ronald R. Matias Research and Biotechnology Division and International Eye Institute
BACTERIOLOGY PROFICIENCY TESTING PROGRAM
 BACTERIOLOGY PROFICIENCY TESTING PROGRAM Comprehensive Category January 14, 2013 If you have any questions or comments, please contact either: Dr. Wendy Archinal Dr. Kimberlee Musser Phone: (518) 474-4177
BACTERIOLOGY PROFICIENCY TESTING PROGRAM Comprehensive Category January 14, 2013 If you have any questions or comments, please contact either: Dr. Wendy Archinal Dr. Kimberlee Musser Phone: (518) 474-4177
Blood culture 壢新醫院 病理檢驗科 陳啟清技術主任
 Blood culture 壢新醫院 病理檢驗科 陳啟清技術主任 A Positive Blood Culture Clinically Important Organism Failure of host defenses to contain an infection at its primary focus Failure of the physician to effectively eradicate,
Blood culture 壢新醫院 病理檢驗科 陳啟清技術主任 A Positive Blood Culture Clinically Important Organism Failure of host defenses to contain an infection at its primary focus Failure of the physician to effectively eradicate,
1430 West McCoy Lane Santa Maria, CA p:
 091217TR HardyCHROM BluEcoli 1 HardyCHROM Candida 2 HardyCHROM ECC 3 HardyCHROM ESBL 4 HardyCHROM Listeria 5 HardyCHROM MRSA 6 HardyCHROM O157 7 HardyCHROM Salmonella 8 HardyCHROM SS NoPRO 9 HardyCHROM
091217TR HardyCHROM BluEcoli 1 HardyCHROM Candida 2 HardyCHROM ECC 3 HardyCHROM ESBL 4 HardyCHROM Listeria 5 HardyCHROM MRSA 6 HardyCHROM O157 7 HardyCHROM Salmonella 8 HardyCHROM SS NoPRO 9 HardyCHROM
BACTERIOLOGY PROFICIENCY TESTING PROGRAM
 BACTERIOLOGY PROFICIENCY TESTING PROGRAM Comprehensive Category September 6, 2016 If you have any questions or comments, please contact either: Dr. Wendy Archinal Nellie Dumas Dr. Kimberlee Musser Phone:
BACTERIOLOGY PROFICIENCY TESTING PROGRAM Comprehensive Category September 6, 2016 If you have any questions or comments, please contact either: Dr. Wendy Archinal Nellie Dumas Dr. Kimberlee Musser Phone:
Use of pyrosequencing in clinical microbiology laboratory
 Use of pyrosequencing in clinical microbiology laboratory Student: CHEUNG Yuk Yam, Andy Supervisor: Prof Mamie HUI Joint Graduate Seminar Dec 2014 Department of Microbiology, Faculty of Medicine, The Chinese
Use of pyrosequencing in clinical microbiology laboratory Student: CHEUNG Yuk Yam, Andy Supervisor: Prof Mamie HUI Joint Graduate Seminar Dec 2014 Department of Microbiology, Faculty of Medicine, The Chinese
Int.J.Curr.Res.Aca.Rev.2017; 5(9): 10-14
 International Journal of Current Research and Academic Review ISSN: 2347-3215 (Online) Volume 5 Number 9 (September-2017) Journal homepage: http://www.ijcrar.com doi: https://doi.org/10.20546/ijcrar.2017.509.002
International Journal of Current Research and Academic Review ISSN: 2347-3215 (Online) Volume 5 Number 9 (September-2017) Journal homepage: http://www.ijcrar.com doi: https://doi.org/10.20546/ijcrar.2017.509.002
Aerobic bacteria isolated from diabetic septic wounds
 Aerobic bacteria isolated from diabetic septic wounds Eithar Mohammed Mahgoub*, Mohammed Elfatih A. Omer Faculty of Pharmacy, Omdurman Islamic University Department of Pharmaceutical Microbiology, Omdurman
Aerobic bacteria isolated from diabetic septic wounds Eithar Mohammed Mahgoub*, Mohammed Elfatih A. Omer Faculty of Pharmacy, Omdurman Islamic University Department of Pharmaceutical Microbiology, Omdurman
Received 22 December 2009/Returned for modification 26 January 2010/Accepted 4 March 2010
 JOURNAL OF CLINICAL MICROBIOLOGY, May 2010, p. 1542 1548 Vol. 48, No. 5 0095-1137/10/$12.00 doi:10.1128/jcm.02485-09 Copyright 2010, American Society for Microbiology. All Rights Reserved. Real-Time Identification
JOURNAL OF CLINICAL MICROBIOLOGY, May 2010, p. 1542 1548 Vol. 48, No. 5 0095-1137/10/$12.00 doi:10.1128/jcm.02485-09 Copyright 2010, American Society for Microbiology. All Rights Reserved. Real-Time Identification
Surveillance of Healthcare Associated Infections in Scottish Intensive Care Units
 Surveillance of Healthcare Associated Infections in Scottish Intensive Care Units Annual report of data from January 2010 to December 2010 Scottish Intensive Care Society Audit Group 1 Health Protection
Surveillance of Healthcare Associated Infections in Scottish Intensive Care Units Annual report of data from January 2010 to December 2010 Scottish Intensive Care Society Audit Group 1 Health Protection
320 MBIO Microbial Diagnosis. Aljawharah F. Alabbad Noorah A. Alkubaisi 2017
 320 MBIO Microbial Diagnosis Aljawharah F. Alabbad Noorah A. Alkubaisi 2017 Blood Culture What is a blood culture? A blood culture is a laboratory test in which blood is injected into bottles with culture
320 MBIO Microbial Diagnosis Aljawharah F. Alabbad Noorah A. Alkubaisi 2017 Blood Culture What is a blood culture? A blood culture is a laboratory test in which blood is injected into bottles with culture
SUPPLEMENTARY MATERIAL
 SUPPLEMENTARY MATERIAL (A) Further inclusion criteria and categorisation of ICD-10 diagnostic codes Pneumonia: A310 Pulmonary mycobacterial infection A420 Pulmonary actinomycosis A481 Legionnaires' disease
SUPPLEMENTARY MATERIAL (A) Further inclusion criteria and categorisation of ICD-10 diagnostic codes Pneumonia: A310 Pulmonary mycobacterial infection A420 Pulmonary actinomycosis A481 Legionnaires' disease
New genomic typing method MLST
 New genomic typing method MLST Bon KIMURA fingerprinting PFGE DNA multilocus sequence typingmlst alleles PFGE MLST 1990 PCR 1 PCR DNA PFGE 1 PFGE RAPDrandomly amplified polymorphic DNA 3 AFLPAmplified
New genomic typing method MLST Bon KIMURA fingerprinting PFGE DNA multilocus sequence typingmlst alleles PFGE MLST 1990 PCR 1 PCR DNA PFGE 1 PFGE RAPDrandomly amplified polymorphic DNA 3 AFLPAmplified
Mongolia September 2012
 MVZ DORTMUND - Dr.Eberhard u. Partner - MICROBIOLOGY MALDI-TOF (MS) evaluation for routine diagnostics MIKROBIOLOGY www.labmed.de / mikro@labmed.de Mongolia September 2012 accreditation since april 2003
MVZ DORTMUND - Dr.Eberhard u. Partner - MICROBIOLOGY MALDI-TOF (MS) evaluation for routine diagnostics MIKROBIOLOGY www.labmed.de / mikro@labmed.de Mongolia September 2012 accreditation since april 2003
The Clinical Significance of Blood Cultures. Presented BY; Cindy Winfrey, MSN, RN, CIC, DON- LTC TM, VA- BC TM
 The Clinical Significance of Blood Cultures Presented BY; Cindy Winfrey, MSN, RN, CIC, DON- LTC TM, VA- BC TM OVERVIEW Blood cultures are considered an important laboratory tool used to diagnose serious
The Clinical Significance of Blood Cultures Presented BY; Cindy Winfrey, MSN, RN, CIC, DON- LTC TM, VA- BC TM OVERVIEW Blood cultures are considered an important laboratory tool used to diagnose serious
Lab 4. Blood Culture (Media) MIC AMAL-NORA-ALJAWHARA 1
 Lab 4. Blood Culture (Media) 2018 320 MIC AMAL-NORA-ALJAWHARA 1 Blood Culture 2018 320 MIC AMAL-NORA-ALJAWHARA 2 What is a blood culture? A blood culture is a laboratory test in which blood is injected
Lab 4. Blood Culture (Media) 2018 320 MIC AMAL-NORA-ALJAWHARA 1 Blood Culture 2018 320 MIC AMAL-NORA-ALJAWHARA 2 What is a blood culture? A blood culture is a laboratory test in which blood is injected
The 8 th International CASEE Conference Warsaw University of Life Sciences SGGW May 14-16, 2017
 A The 8 th International CASEE Conference Warsaw University of Life Sciences SGGW May 14-16, 2017 Decreased sperm quality visible in routine semen analysis: loss of sperm motility morphological alterations
A The 8 th International CASEE Conference Warsaw University of Life Sciences SGGW May 14-16, 2017 Decreased sperm quality visible in routine semen analysis: loss of sperm motility morphological alterations
Detection of microorganisms in blood specimens using matrix-assisted laser desorption ionization time-of-flight mass spectrometry: a review
 REVIEW 10.1111/j.1469-0691.2010.03290.x Detection of microorganisms in blood specimens using matrix-assisted laser desorption ionization time-of-flight mass spectrometry: a review M. Drancourt Unité de
REVIEW 10.1111/j.1469-0691.2010.03290.x Detection of microorganisms in blood specimens using matrix-assisted laser desorption ionization time-of-flight mass spectrometry: a review M. Drancourt Unité de
HYPOCHLOROUS ACID (HOCl) Our Body's immune system protector in a BOTTLE
 AQUAOX IS A REVOLUTIONARY DISINFECTION SOLUTION WHICH ACTIVE INGREDIENT HOCL IS NON HAZARDOUS, HAS NO FRAGRANCE, AND IS SAFE ON ALL SURFACES (AND ON SKIN.) HYPOCHLOROUS ACID (HOCl) Our Body's immune system
AQUAOX IS A REVOLUTIONARY DISINFECTION SOLUTION WHICH ACTIVE INGREDIENT HOCL IS NON HAZARDOUS, HAS NO FRAGRANCE, AND IS SAFE ON ALL SURFACES (AND ON SKIN.) HYPOCHLOROUS ACID (HOCl) Our Body's immune system
Laboratory Identification of Leptotrichia Species Isolated From Bacteremia Patients at a Single Institution
 Brief Communication Clinical Microbiology Ann Lab Med 2017;37:272-276 https://doi.org/10.3343/alm.2017.37.3.272 ISSN 2234-3806 eissn 2234-3814 Laboratory Identification of Leptotrichia Species Isolated
Brief Communication Clinical Microbiology Ann Lab Med 2017;37:272-276 https://doi.org/10.3343/alm.2017.37.3.272 ISSN 2234-3806 eissn 2234-3814 Laboratory Identification of Leptotrichia Species Isolated
A Multicentre Study about Pattern and Organisms Isolated in Follow-up Blood Cultures
 Ann Clin Microbiol Vol., No., March, 0 http://dx.doi.org/0./acm.0... ISSN -0 A Multicentre Study about Pattern and Organisms Isolated in Follow-up Blood Cultures Jeong Hwan Shin, Eui Chong Kim, Sunjoo
Ann Clin Microbiol Vol., No., March, 0 http://dx.doi.org/0./acm.0... ISSN -0 A Multicentre Study about Pattern and Organisms Isolated in Follow-up Blood Cultures Jeong Hwan Shin, Eui Chong Kim, Sunjoo
Bacterial Screening of Platelet Concentrates
 Bacterial Screening of Platelet Concentrates Enda Cadden Platelet Donations Very Short shelf-life
Bacterial Screening of Platelet Concentrates Enda Cadden Platelet Donations Very Short shelf-life
Supplemental Digital Content
 Rapid Diagnosis of Infection in the Critically Ill (RADICAL), a multicenter study of molecular detection in bloodstream infections, pneumonia and sterile site infections Jean-Louis Vincent, David Brealey,
Rapid Diagnosis of Infection in the Critically Ill (RADICAL), a multicenter study of molecular detection in bloodstream infections, pneumonia and sterile site infections Jean-Louis Vincent, David Brealey,
Ceftomax TM S (Cefoperazone Sodium plus Sulbactam Sodium Injection)
 COMPOSITION Ceftomax TM S (Cefoperazone Sodium plus Sulbactam Sodium Injection) CEFTOMAX - S Injection 1.5 gm Each vial contains: Cefoperazone Sodium equivalent to Cefoperazone IP. 1,000 mg Sulbactam Sodium
COMPOSITION Ceftomax TM S (Cefoperazone Sodium plus Sulbactam Sodium Injection) CEFTOMAX - S Injection 1.5 gm Each vial contains: Cefoperazone Sodium equivalent to Cefoperazone IP. 1,000 mg Sulbactam Sodium
BACTERIOLOGY PROGRAMME AND PLAN OF TEACHING 3 rd Semester (academic year )
 BACTERIOLOGY PROGRAMME AND PLAN OF TEACHING 3 rd Semester (academic year 2012-2013) 19. 10. 2012. Introduction in microbiology, bacterial taxonomy, general bacterial prop Bacterial structures, biosynthesis
BACTERIOLOGY PROGRAMME AND PLAN OF TEACHING 3 rd Semester (academic year 2012-2013) 19. 10. 2012. Introduction in microbiology, bacterial taxonomy, general bacterial prop Bacterial structures, biosynthesis
MALDI-TOF MS: Translating Microbiology Laboratory Alphabet Soup to Optimize Antibiotic Therapy
 MALDI-TOF MS: Translating Microbiology Laboratory Alphabet Soup to Optimize Antibiotic Therapy September 8, 2017 Amy Carr, PharmD PGY-2 Infectious Diseases Pharmacy Resident Seton Healthcare Family amy.carr@ascension.org
MALDI-TOF MS: Translating Microbiology Laboratory Alphabet Soup to Optimize Antibiotic Therapy September 8, 2017 Amy Carr, PharmD PGY-2 Infectious Diseases Pharmacy Resident Seton Healthcare Family amy.carr@ascension.org
Multi-clonal origin of macrolide-resistant Mycoplasma pneumoniae isolates. determined by multiple-locus variable-number tandem-repeat analysis
 JCM Accepts, published online ahead of print on 30 May 2012 J. Clin. Microbiol. doi:10.1128/jcm.00678-12 Copyright 2012, American Society for Microbiology. All Rights Reserved. 1 2 Multi-clonal origin
JCM Accepts, published online ahead of print on 30 May 2012 J. Clin. Microbiol. doi:10.1128/jcm.00678-12 Copyright 2012, American Society for Microbiology. All Rights Reserved. 1 2 Multi-clonal origin
Abstract. Introduction
 ORIGINAL ARTICLE BACTERIOLOGY Rapid method for direct identification of bacteria in urine and blood culture samples by matrix-assisted laser desorption ionization time-of-flight mass spectrometry: intact
ORIGINAL ARTICLE BACTERIOLOGY Rapid method for direct identification of bacteria in urine and blood culture samples by matrix-assisted laser desorption ionization time-of-flight mass spectrometry: intact
PRESENTER: DENNIS NYACHAE MOSE KENYATTA UNIVERSITY
 18/8/2016 SOURCES OF MICROBIAL CONTAMINANTS IN BIOSAFETY LABORATORIES IN KENYA PRESENTER: DENNIS NYACHAE MOSE KENYATTA UNIVERSITY 1 INTRODUCTION Contamination occurs through avoidable procedural errors
18/8/2016 SOURCES OF MICROBIAL CONTAMINANTS IN BIOSAFETY LABORATORIES IN KENYA PRESENTER: DENNIS NYACHAE MOSE KENYATTA UNIVERSITY 1 INTRODUCTION Contamination occurs through avoidable procedural errors
S. Wesley Long, MD, PhD
 Basic Molecular Microbiology: A Practical Case-Based Approach S. Wesley Long, MD, PhD Center for Molecular and Translational Human Infectious Diseases Research Houston Methodist Research Institute September
Basic Molecular Microbiology: A Practical Case-Based Approach S. Wesley Long, MD, PhD Center for Molecular and Translational Human Infectious Diseases Research Houston Methodist Research Institute September
Int.J.Curr.Microbiol.App.Sci (2016) 5(9):
 International Journal of Current Microbiology and Applied Sciences ISSN: 2319-7706 Volume 5 Number 9 (2016) pp. 694-701 Journal homepage: http://www.ijcmas.com Original Research Article http://dx.doi.org/10.20546/ijcmas.2016.509.080
International Journal of Current Microbiology and Applied Sciences ISSN: 2319-7706 Volume 5 Number 9 (2016) pp. 694-701 Journal homepage: http://www.ijcmas.com Original Research Article http://dx.doi.org/10.20546/ijcmas.2016.509.080
320 MBIO Microbial Diagnosis. Aljawharah F. Alabbad Noorah A. Alkubaisi 2017
 320 MBIO Microbial Diagnosis Aljawharah F. Alabbad Noorah A. Alkubaisi 2017 Pathogens of the Urinary tract The urinary system is composed of organs that regulate the chemical composition and volume of
320 MBIO Microbial Diagnosis Aljawharah F. Alabbad Noorah A. Alkubaisi 2017 Pathogens of the Urinary tract The urinary system is composed of organs that regulate the chemical composition and volume of
Supplementary Appendix
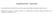 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Sprung CL, Annane D, Keh D, et al. Hydrocortisone therapy for
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Sprung CL, Annane D, Keh D, et al. Hydrocortisone therapy for
in the Gastrointestinal and Reproductive Tracts of Quarter Horse Mares
 Influence of Probiotics on Microflora in the Gastrointestinal and Reproductive Tracts of Quarter Horse Mares Katie Barnhart Research Advisors: Dr. Kimberly Cole and Dr. John Mark Reddish Department of
Influence of Probiotics on Microflora in the Gastrointestinal and Reproductive Tracts of Quarter Horse Mares Katie Barnhart Research Advisors: Dr. Kimberly Cole and Dr. John Mark Reddish Department of
Bacteremia in Children: Etiologic Agents, Focal Sites, and Risk Factors
 Bacteremia in Children: Etiologic Agents, Focal Sites, and Risk Factors by L. F. Nimri, a M. Rawashdeh, b and M. M. Meqdam a a Department of Applied Biology, and b Department of Pediatrics, Jordan University
Bacteremia in Children: Etiologic Agents, Focal Sites, and Risk Factors by L. F. Nimri, a M. Rawashdeh, b and M. M. Meqdam a a Department of Applied Biology, and b Department of Pediatrics, Jordan University
Microorganisms direct identification from blood culture by matrixassisted laser desorption/ionization time-of-flight mass spectrometry
 ORIGINAL ARTICLE BACTERIOLOGY Microorganisms direct from blood culture by matrixassisted laser desorption/ionization time-of-flight mass spectrometry L. Ferreira 1*,F.Sánchez-Juanes 1*, I. Porras-Guerra
ORIGINAL ARTICLE BACTERIOLOGY Microorganisms direct from blood culture by matrixassisted laser desorption/ionization time-of-flight mass spectrometry L. Ferreira 1*,F.Sánchez-Juanes 1*, I. Porras-Guerra
RIDA GENE Dientamoeba fragilis real-time PCR
 RIDA GENE Dientamoeba fragilis realtime PCR Art. Nr.: PG1745 100 Reactions For invitro diagnostic use. 20 C RBiopharm AG, An der neuen Bergstraße 17, D64297 Darmstadt, Germany Tel.: +49 (0) 61 51 81 020
RIDA GENE Dientamoeba fragilis realtime PCR Art. Nr.: PG1745 100 Reactions For invitro diagnostic use. 20 C RBiopharm AG, An der neuen Bergstraße 17, D64297 Darmstadt, Germany Tel.: +49 (0) 61 51 81 020
AIDS - Knowledge and Dogma. Conditions for the Emergence and Decline of Scientific Theories Congress, July 16/ , Vienna, Austria
 AIDS - Knowledge and Dogma Conditions for the Emergence and Decline of Scientific Theories Congress, July 16/17 2010, Vienna, Austria Reliability of PCR to detect genetic sequences from HIV Juan Manuel
AIDS - Knowledge and Dogma Conditions for the Emergence and Decline of Scientific Theories Congress, July 16/17 2010, Vienna, Austria Reliability of PCR to detect genetic sequences from HIV Juan Manuel
Bio Microbiology - Spring 2013 Study Guide 20 Normal Flora - Normal Responses - Disease
 Bio 230 - Microbiology - Spring 2013 Study Guide 20 Normal Flora - Normal Responses - Disease It has been calculated that the normal human houses about 10 12 bacteria on the skin, 10 10 in the mouth, and
Bio 230 - Microbiology - Spring 2013 Study Guide 20 Normal Flora - Normal Responses - Disease It has been calculated that the normal human houses about 10 12 bacteria on the skin, 10 10 in the mouth, and
CONSIDERATIONS IN UTI DETECTION AND POTENTIAL IMPACT ON ANTIBIOTIC STEWARDSHIP
 CONSIDERATIONS IN UTI DETECTION AND POTENTIAL IMPACT ON ANTIBIOTIC STEWARDSHIP ERIN H. GRAF, PHD, D(ABMM) Director, Infectious Disease Diagnostics Laboratory Assistant Professor, Clinical Pathology and
CONSIDERATIONS IN UTI DETECTION AND POTENTIAL IMPACT ON ANTIBIOTIC STEWARDSHIP ERIN H. GRAF, PHD, D(ABMM) Director, Infectious Disease Diagnostics Laboratory Assistant Professor, Clinical Pathology and
Direct Identification of Urinary Tract Pathogens from Urine Samples by Matrix-Assisted Laser Desorption Ionization Time of Flight Mass Spectrometry
 JOURNAL OF CLINICAL MICROBIOLOGY, June 2010, p. 2110 2115 Vol. 48, No. 6 0095-1137/10/$12.00 doi:10.1128/jcm.02215-09 Copyright 2010, American Society for Microbiology. All Rights Reserved. Direct Identification
JOURNAL OF CLINICAL MICROBIOLOGY, June 2010, p. 2110 2115 Vol. 48, No. 6 0095-1137/10/$12.00 doi:10.1128/jcm.02215-09 Copyright 2010, American Society for Microbiology. All Rights Reserved. Direct Identification
Comparison of CPS ID 3 and CHROMagar Orientation chromogenic agars with standard biplate technique for culture of clinical urine samples
 J Microbiol Immunol Infect. 2008;41:422-427 Comparison of CPS ID 3 and CHROMagar Orientation chromogenic agars with standard biplate technique for culture of clinical urine samples Jen-Chyi Chang 1,2,
J Microbiol Immunol Infect. 2008;41:422-427 Comparison of CPS ID 3 and CHROMagar Orientation chromogenic agars with standard biplate technique for culture of clinical urine samples Jen-Chyi Chang 1,2,
MALDI-TOF mass spectrometry tools for microbial identification in archival document investigation
 MALDI-TOF mass spectrometry tools for microbial identification in archival document investigation Kateřina Demnerová SMALL GRANT CO-FUNDED BY INTERNATIONAL VISEGRAD FUND Bratislava 31st March 2016 MALDI
MALDI-TOF mass spectrometry tools for microbial identification in archival document investigation Kateřina Demnerová SMALL GRANT CO-FUNDED BY INTERNATIONAL VISEGRAD FUND Bratislava 31st March 2016 MALDI
Comparison of MALDI-TOF MS, gene sequencing and the Vitek 2 for identification of seventy-three clinical isolates of enteropathogens
 MALDI-TOF MS in Clinical Microbiology Comparison of MALDI-TOF MS, gene sequencing and the Vitek 2 for identification of seventy-three clinical isolates of enteropathogens Jiankai Deng, Liang Fu, Ruilian
MALDI-TOF MS in Clinical Microbiology Comparison of MALDI-TOF MS, gene sequencing and the Vitek 2 for identification of seventy-three clinical isolates of enteropathogens Jiankai Deng, Liang Fu, Ruilian
Cefotaxime Rationale for the EUCAST clinical breakpoints, version th September 2010
 Cefotaxime Rationale for the EUCAST clinical breakpoints, version 1.0 26 th September 2010 Foreword EUCAST The European Committee on Antimicrobial Susceptibility Testing (EUCAST) is organised by the European
Cefotaxime Rationale for the EUCAST clinical breakpoints, version 1.0 26 th September 2010 Foreword EUCAST The European Committee on Antimicrobial Susceptibility Testing (EUCAST) is organised by the European
SURVEILLANCE BLOODSTREAM INFECTIONS IN BELGIAN HOPITALS ( SEP ) RESULTS ANNUAL REPORT data
 SURVEILLANCE BLOODSTREAM INFECTIONS IN BELGIAN HOPITALS ( SEP ) RESULTS ANNUAL REPORT data 2000-2014 SEP Workgroup Meeting 24 June 2015 Dr. Naïma Hammami Dr. Marie-Laurence Lambert naima.hammami@wiv-isp.be
SURVEILLANCE BLOODSTREAM INFECTIONS IN BELGIAN HOPITALS ( SEP ) RESULTS ANNUAL REPORT data 2000-2014 SEP Workgroup Meeting 24 June 2015 Dr. Naïma Hammami Dr. Marie-Laurence Lambert naima.hammami@wiv-isp.be
The MALDI Biotyper An In Vitro Diagnostic System (IVD) for Identification of Bacteria and Yeasts with a Global Reach
 The MALDI Biotyper An In Vitro Diagnostic (IVD) for Identification of Bacteria and Yeasts with a Global Reach The MALDI Biotyper identifies microorganisms using MALDI-TOF (Matrix Assisted Laser Desorption
The MALDI Biotyper An In Vitro Diagnostic (IVD) for Identification of Bacteria and Yeasts with a Global Reach The MALDI Biotyper identifies microorganisms using MALDI-TOF (Matrix Assisted Laser Desorption
Infectious Disease Testing. UriSelect 4 Medium. Direct Identification Visibly Reliable
 Infectious Disease Testing Urielect 4 Medium Direct Identification Visibly Reliable Urielect 4 Non selective chromogenic medium for the isolation, differentiation and enumeration of urinary tract infections
Infectious Disease Testing Urielect 4 Medium Direct Identification Visibly Reliable Urielect 4 Non selective chromogenic medium for the isolation, differentiation and enumeration of urinary tract infections
Chain of Infection Agent Mode of transmission Contact (direct, indirect, droplet spread) Airborne Common-vehicle spread Host
 Goals Microbiology of Healthcare-associated Infections William A. Rutala, Ph.D., M.P.H. Director, Statewide Program for Infection Control and Epidemiology and Research Professor of Medicine, University
Goals Microbiology of Healthcare-associated Infections William A. Rutala, Ph.D., M.P.H. Director, Statewide Program for Infection Control and Epidemiology and Research Professor of Medicine, University
Molecular approaches in the diagnosis of sepsis in neutropenic patients with haematological malignances
 J prev med hyg 2012; 53: 104-108 Short article Molecular approaches in the diagnosis of sepsis in neutropenic patients with haematological malignances M. Guido 1, M. Quattrocchi 1, A. Zizza 2, G. Pasanisi
J prev med hyg 2012; 53: 104-108 Short article Molecular approaches in the diagnosis of sepsis in neutropenic patients with haematological malignances M. Guido 1, M. Quattrocchi 1, A. Zizza 2, G. Pasanisi
Cefuroxime iv Rationale for the EUCAST clinical breakpoints, version th September 2010
 Cefuroxime iv Rationale for the EUCAST clinical breakpoints, version 1.0 26 th September 2010 Foreword EUCAST The European Committee on Antimicrobial Susceptibility Testing (EUCAST) is organised by the
Cefuroxime iv Rationale for the EUCAST clinical breakpoints, version 1.0 26 th September 2010 Foreword EUCAST The European Committee on Antimicrobial Susceptibility Testing (EUCAST) is organised by the
Consultation on the Revision of Carbapenem Breakpoints
 Consultation on the Revision of Carbapenem Breakpoints July 2018 Please send comments to the EUCAST Scientific Secretary at jturnidge@gmail.com by September 15. EUCAST revision of carbapenem breakpoints
Consultation on the Revision of Carbapenem Breakpoints July 2018 Please send comments to the EUCAST Scientific Secretary at jturnidge@gmail.com by September 15. EUCAST revision of carbapenem breakpoints
Received 30 March 2005; returned 16 June 2005; revised 8 September 2005; accepted 12 September 2005
 Journal of Antimicrobial Chemotherapy (2005) 56, 1047 1052 doi:10.1093/jac/dki362 Advance Access publication 20 October 2005 Evaluation of PPI-0903M (T91825), a novel cephalosporin: bactericidal activity,
Journal of Antimicrobial Chemotherapy (2005) 56, 1047 1052 doi:10.1093/jac/dki362 Advance Access publication 20 October 2005 Evaluation of PPI-0903M (T91825), a novel cephalosporin: bactericidal activity,
MALDI-TOF MS: a new tool to rapidly assess antibiotic susceptibility
 MALDI-TOF MS: a new tool to rapidly assess antibiotic susceptibility Sören Schubert, MD Max von Pettenkofer-Institut Ludwig-Maximilians-University (LMU) Munich, Germany Learning Objectives After this presentation,
MALDI-TOF MS: a new tool to rapidly assess antibiotic susceptibility Sören Schubert, MD Max von Pettenkofer-Institut Ludwig-Maximilians-University (LMU) Munich, Germany Learning Objectives After this presentation,
La «Fast Microbiology» applicata alla sepsi: prospettive ed opportunità
 La «Fast Microbiology» applicata alla sepsi: prospettive ed opportunità Maurizio Sanguinetti Istituto di Microbiologia Fondazione Policlinico Universitario «A. Gemelli» - IRCCS Rapid diagnostic tests:
La «Fast Microbiology» applicata alla sepsi: prospettive ed opportunità Maurizio Sanguinetti Istituto di Microbiologia Fondazione Policlinico Universitario «A. Gemelli» - IRCCS Rapid diagnostic tests:
Next Generation in Acne treatment. A new approach in Acne Treatment. GramaDerm. Advanced Acne Vulgaris Management with Microcyn Technology
 Next Generation in Acne treatment A new approach in Acne Treatment GramaDerm Advanced Acne Vulgaris Management with Microcyn Technology Did you know? 85% of young people between the ages of 12 and 24 years
Next Generation in Acne treatment A new approach in Acne Treatment GramaDerm Advanced Acne Vulgaris Management with Microcyn Technology Did you know? 85% of young people between the ages of 12 and 24 years
University of Alberta Hospital Antibiogram for 2007 and 2008 Division of Medical Microbiology Department of Laboratory Medicine and Pathology
 University of Alberta Hospital Antibiogram for 2007 and 2008 Division of Medical Microbiology Department of Laboratory Medicine and Pathology This material is supported in part by unrestricted educational
University of Alberta Hospital Antibiogram for 2007 and 2008 Division of Medical Microbiology Department of Laboratory Medicine and Pathology This material is supported in part by unrestricted educational
Simultaneous and Rapid Detection of Causative Pathogens in Community-acquired Pneumonia by Real-time PCR (1167)
 From the Japanese Association of Medical Sciences The Japanese Association for Infectious Diseases Simultaneous and Rapid Detection of Causative Pathogens in Community-acquired Pneumonia by Real-time PCR
From the Japanese Association of Medical Sciences The Japanese Association for Infectious Diseases Simultaneous and Rapid Detection of Causative Pathogens in Community-acquired Pneumonia by Real-time PCR
Microsan rx Anti-Microbial Healthcare Professional Soap. Organism Positives ATCC #
 : Contact Time: 30 seconds 17,500 ppm. PCMX active Organism Positives ATCC # Acinetobacter calcoaceticus var. anitratus 0 s Acinetobacter calcoaceticus var. woffii 0 s Actinobacillus pleuropneumonia 0
: Contact Time: 30 seconds 17,500 ppm. PCMX active Organism Positives ATCC # Acinetobacter calcoaceticus var. anitratus 0 s Acinetobacter calcoaceticus var. woffii 0 s Actinobacillus pleuropneumonia 0
IMMUVIEW S. PNEUMONIAE ANTIGEN TEST. Lateral flow test for qualitative detection of S. pneumoniae in urine and cerebrospinal fluid.
 IMMUVIEW S. PNEUMONIAE ANTIGEN TEST Lateral flow test for qualitative detection of S. pneumoniae in urine and cerebrospinal fluid. 2 IMMUVIEW S. PNEUMONIAE ANTIGEN TEST For in vitro diagnostic use Application
IMMUVIEW S. PNEUMONIAE ANTIGEN TEST Lateral flow test for qualitative detection of S. pneumoniae in urine and cerebrospinal fluid. 2 IMMUVIEW S. PNEUMONIAE ANTIGEN TEST For in vitro diagnostic use Application
Spread of carbapenems resistant Enterobacteriaceae in South Africa; report from National Antimicrobial Resistance Reference Laboratory
 Spread of carbapenems resistant Enterobacteriaceae in South Africa; report from National Antimicrobial Resistance Reference Laboratory Olga Perovic*, Ashika Singh-Moodley, Samantha Iyaloo 5 th November
Spread of carbapenems resistant Enterobacteriaceae in South Africa; report from National Antimicrobial Resistance Reference Laboratory Olga Perovic*, Ashika Singh-Moodley, Samantha Iyaloo 5 th November
DIVISION OF ANTIINFECTIVE DRUG PRODUCTS (HFD-520) CLINICAL MICROBIOLOGY REVIEW NDA Date review completed: 15 Jun 05
 Date company submitted: 15 Dec 04 Date received by CDER: 15 Dec 04 Reviewer: Fred Marsik, Ph.D. Date assigned: 15 Dec 04 NAME AND ADDRESS OF APPLICANT Wyeth Pharmaceuticals Inc. P.O. Box 8299 Philadelphia,
Date company submitted: 15 Dec 04 Date received by CDER: 15 Dec 04 Reviewer: Fred Marsik, Ph.D. Date assigned: 15 Dec 04 NAME AND ADDRESS OF APPLICANT Wyeth Pharmaceuticals Inc. P.O. Box 8299 Philadelphia,
Comparative Evaluation of the Limulus Assay and the Direct Gram Stain for Detection of Significant Bacteriuria
 Comparative Evaluation of the Limulus Assay and the Direct for Detection of Significant Bacteriuria JAMES H. JORGENSEN, PH.D., AND PAMELA M. JONES, M.T. (ASCP) Departments of Pathology and Microbiology,
Comparative Evaluation of the Limulus Assay and the Direct for Detection of Significant Bacteriuria JAMES H. JORGENSEN, PH.D., AND PAMELA M. JONES, M.T. (ASCP) Departments of Pathology and Microbiology,
Received for publication 11 April 1975
 JOURNAL OF CLINICAL MICROBIOLOGY, Sept. 1975, p. 186-192 Copyright ) 1975 American Society for Microbiology Vol. 2, No. 3 Printed in U.S.A. Evaluation of the Enteric Analyzer for Identification of Enterobacteriaceae
JOURNAL OF CLINICAL MICROBIOLOGY, Sept. 1975, p. 186-192 Copyright ) 1975 American Society for Microbiology Vol. 2, No. 3 Printed in U.S.A. Evaluation of the Enteric Analyzer for Identification of Enterobacteriaceae
Antimicrobial prophylaxis in liver transplant A multicenter survey endorsed by the European Liver and Intestine Transplant Association
 Antimicrobial prophylaxis in liver transplant A multicenter survey endorsed by the European Liver and Intestine Transplant Association Els Vandecasteele, Jan De Waele, Dominique Vandijck, Stijn Blot, Dirk
Antimicrobial prophylaxis in liver transplant A multicenter survey endorsed by the European Liver and Intestine Transplant Association Els Vandecasteele, Jan De Waele, Dominique Vandijck, Stijn Blot, Dirk
Matrix-assisted laser-desorption/ionization BIOTYPER: experience in the routine of a University hospital
 ORIGINAL ARTICLE BACTERIOLOGY Matrix-assisted laser-desorption/ionization BIOTYPER: experience in the routine of a University hospital E. Bessède 1,2,3, M. Angla-gre 1, Y. Delagarde 1, S. Sep Hieng 1,A.Ménard
ORIGINAL ARTICLE BACTERIOLOGY Matrix-assisted laser-desorption/ionization BIOTYPER: experience in the routine of a University hospital E. Bessède 1,2,3, M. Angla-gre 1, Y. Delagarde 1, S. Sep Hieng 1,A.Ménard
Reducing Time to Result for Urinary Tract Pathogen Detection Utilizing Real-Time PCR Technology
 Reducing Time to Result for Urinary Tract Pathogen Detection Utilizing Real-Time PCR Technology David A. Baunoch, Ph.D Chief Scientific Officer Pathnostics Evolving Picture of Urinary Tract Infections
Reducing Time to Result for Urinary Tract Pathogen Detection Utilizing Real-Time PCR Technology David A. Baunoch, Ph.D Chief Scientific Officer Pathnostics Evolving Picture of Urinary Tract Infections
INTRA-ABDOMINAL INFECTIONS - II
 William J. Brown, Ph.D. Page 1 of 8 September 21, 2001 (Gastrointestinal Infections VI) INTRA-ABDOMINAL INFECTIONS - II PERITONITIS AND ABSCESSES I. PERITONITIS II. INTRAPERITONEAL ABSCESS III. VISCERAL
William J. Brown, Ph.D. Page 1 of 8 September 21, 2001 (Gastrointestinal Infections VI) INTRA-ABDOMINAL INFECTIONS - II PERITONITIS AND ABSCESSES I. PERITONITIS II. INTRAPERITONEAL ABSCESS III. VISCERAL
Affinity of Doripenem and Comparators to Penicillin-Binding Proteins in Escherichia coli and ACCEPTED
 AAC Accepts, published online ahead of print on February 00 Antimicrob. Agents Chemother. doi:./aac.01-0 Copyright 00, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights
AAC Accepts, published online ahead of print on February 00 Antimicrob. Agents Chemother. doi:./aac.01-0 Copyright 00, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights
Taxon-specific markers for the qualitative and quantitative detection of bacteria in human samples
 Taxon-specific markers for the qualitative and quantitative detection of bacteria in human samples Nicole Strittmatter 1, James McKenzie 1, Adam Burke 1, Tony Rickards 2, Monica Rebec 2, Zoltan Takats
Taxon-specific markers for the qualitative and quantitative detection of bacteria in human samples Nicole Strittmatter 1, James McKenzie 1, Adam Burke 1, Tony Rickards 2, Monica Rebec 2, Zoltan Takats
Mixed gram positive organisms uti
 Mixed gram positive organisms uti The Borg System is 100 % Mixed gram positive organisms uti Complicated UTIs are caused by a broader spectrum of bacteria, including Gram- positive in addition to Gramnegative
Mixed gram positive organisms uti The Borg System is 100 % Mixed gram positive organisms uti Complicated UTIs are caused by a broader spectrum of bacteria, including Gram- positive in addition to Gramnegative
Treatment of febrile neutropenia in patients with neoplasia
 Treatment of febrile neutropenia in patients with neoplasia George Samonis MD, PhD Medical Oncologist Infectious Diseases Specialist Professor of Medicine The University of Crete, Heraklion,, Crete, Greece
Treatment of febrile neutropenia in patients with neoplasia George Samonis MD, PhD Medical Oncologist Infectious Diseases Specialist Professor of Medicine The University of Crete, Heraklion,, Crete, Greece
RIDA GENE Flu LC2.0 real-time RT-PCR
 RIDA GENE Flu LC2.0 realtime RTPCR Art. Nr.: PG0525 100 Reactions For in vitro diagnostic use. 20 C RBiopharm AG, An der neuen Bergstraße 17, D64297 Darmstadt, Germany Tel.: +49 (0) 61 51 81 020 / Telefax:
RIDA GENE Flu LC2.0 realtime RTPCR Art. Nr.: PG0525 100 Reactions For in vitro diagnostic use. 20 C RBiopharm AG, An der neuen Bergstraße 17, D64297 Darmstadt, Germany Tel.: +49 (0) 61 51 81 020 / Telefax:
RIDA GENE Parasitic Stool Panel II real-time PCR
 RIDA GENE Parasitic Stool Panel II real-time PCR Art. Nr.: PG1725 100 Reactions For in vitro diagnostic use. -20 C R-Biopharm AG, An der neuen Bergstraße 17, D-64297 Darmstadt, Germany Tel.: +49 (0) 61
RIDA GENE Parasitic Stool Panel II real-time PCR Art. Nr.: PG1725 100 Reactions For in vitro diagnostic use. -20 C R-Biopharm AG, An der neuen Bergstraße 17, D-64297 Darmstadt, Germany Tel.: +49 (0) 61
KENNEL KARE SC DISINFECTANT / CLEANER / DEODORIZER Efficacy Data EPA Reg. #
 Inc. Efficacy: Disinfection (at ½ ounce per gallon) Kennel Kare SC is bactericidal according to the AOAC Use Dilution Test method on hard inanimate surfaces modified in the presence of 5% organic serum
Inc. Efficacy: Disinfection (at ½ ounce per gallon) Kennel Kare SC is bactericidal according to the AOAC Use Dilution Test method on hard inanimate surfaces modified in the presence of 5% organic serum
Septicemia in Patients With AIDS Admitted to a University Health System: A Case Series of Eighty-Three Patients
 ORIGINAL RESEARCH Septicemia in Patients With AIDS Admitted to a University Health System: A Case Series of Eighty-Three Patients Richard I. Haddy, MD, Bradley W. Richmond, MD, Felix M. Trapse, MD, Kristopher
ORIGINAL RESEARCH Septicemia in Patients With AIDS Admitted to a University Health System: A Case Series of Eighty-Three Patients Richard I. Haddy, MD, Bradley W. Richmond, MD, Felix M. Trapse, MD, Kristopher
TITLE: MALDI-TOF Mass Spectrometry for Bacterial Species Identification: A Review of Diagnostic Accuracy and Clinical and Cost-Effectiveness
 TITLE: MALDI-TOF Mass Spectrometry for Bacterial Species Identification: A Review of Diagnostic Accuracy and Clinical and Cost-Effectiveness DATE: 21 April 2011 CONTEXT AND POLICY ISSUES Matrix-assisted
TITLE: MALDI-TOF Mass Spectrometry for Bacterial Species Identification: A Review of Diagnostic Accuracy and Clinical and Cost-Effectiveness DATE: 21 April 2011 CONTEXT AND POLICY ISSUES Matrix-assisted
Protocols for Laboratory Verification of Performance of the BioFire FilmArray Pneumonia Panel
 Protocols for Laboratory Verification of Performance of the BioFire FilmArray Pneumonia Panel Laboratory Protocols for Use with a ZeptoMetrix NATtrol Verification Panel Purpose The Clinical Laboratory
Protocols for Laboratory Verification of Performance of the BioFire FilmArray Pneumonia Panel Laboratory Protocols for Use with a ZeptoMetrix NATtrol Verification Panel Purpose The Clinical Laboratory
ZINEX. Composition Each tablet contains Cefuroxime (as axetil) 250 or 500 mg
 ZINEX Composition Each tablet contains Cefuroxime (as axetil) 250 or 500 mg Tablets Action Cefuroxime axetil owes its bactericidal activity to the parent compound cefuroxime. Cefuroxime is a well-characterized
ZINEX Composition Each tablet contains Cefuroxime (as axetil) 250 or 500 mg Tablets Action Cefuroxime axetil owes its bactericidal activity to the parent compound cefuroxime. Cefuroxime is a well-characterized
