Quantitative proteomics reveals the kinetics of trypsin-catalyzed protein digestion
|
|
|
- Mildred James
- 5 years ago
- Views:
Transcription
1 Anal Bioanal Chem (2014) 406: DOI /s RESEARCH PAPER Quantitative proteomics reveals the kinetics of trypsin-catalyzed protein digestion Yanbo Pan & Kai Cheng & Jiawei Mao & Fangjie Liu & Jing Liu & Mingliang Ye & Hanfa Zou Received: 9 June 2014 /Revised: 14 July 2014 /Accepted: 25 July 2014 /Published online: 19 August 2014 # Springer-Verlag Berlin Heidelberg 2014 Abstract Trypsin is the popular protease to digest proteins into peptides in shotgun proteomics, but few studies have attempted to systematically investigate the kinetics of trypsin-catalyzed protein digestion in proteome samples. In this study, we applied quantitative proteomics via triplex stable isotope dimethyl labeling to investigate the kinetics of trypsin-catalyzed cleavage. It was found that trypsin cleaves the C-terminal to lysine (K) and arginine (R) residues with higher rates for R. And the cleavage sites surrounded by neutral residues could be quickly cut, while those with neighboring charged residues (D/E/K/R) or proline residue (P) could be slowly cut. In a proteome sample, a huge number of proteins with different physical chemical properties coexists. If any type of protein could be preferably digested, then limited digestion could be applied to reduce the sample complexity. However, we found that protein abundance and other physicochemical properties, such as molecular weight (Mw), grand average of hydropathicity (GRAVY), aliphatic index, and isoelectric point (pi) have no notable correlation with digestion priority of proteins. Keywords Trypsin. Protein digestion. Kinetics. Stable isotope dimethyl labeling. Mass spectrometry Electronic supplementary material The online version of this article (doi: /s ) contains supplementary material, which is available to authorized users. Y. Pan: K. Cheng : J. Mao : F. Liu : J. Liu : M. Ye (*) : H. Zou (*) Key Lab of Separation Sciences for Analytical Chemistry, National Chromatographic Research and Analysis Center, Dalian Institute of Chemical Physics, Chinese Academy of Sciences, Dalian , China mingliang@dicp.ac.cn hanfazou@dicp.ac.cn Y. Pan: K. Cheng : J. Mao : F. Liu : J. Liu University of Chinese Academy of Sciences, Beijing , China Introduction Shotgun proteomics, i.e., bottom-up proteomics, is a powerful strategy for identifying proteins from complex protein mixture [1]. It relies on enzymatic digestion of proteins into peptides prior to liquid chromatography-coupled tandem mass spectrometry (LC-MS/MS) analysis [2]. A variety of proteases including trypsin, Lys-C, Glu-C, etc. are applied to digest proteins in proteome research. Among these proteases, trypsin is the most commonly used enzyme. This is mainly attributed to the facts that tryptic peptides have highly basic residues at the C-termini of peptides, and they are in the preferred mass range for effective fragmentation by MS/MS. Thus, the fragmentation of tryptic peptides generally leads to a series of y- ion series and makes tandem mass spectra more easily interpretable [3]. Trypsin catalyzes the hydrolysis of peptide bonds immediately after lysine/arginine (K/R) residues in proteins. There are many potential trypsin cleavage sites in proteins, while not all these sites could be cut during a trypsin digestion. These sites are called missed cleavage sites [4 6]. Clearly, the missed cleavage sites have very slow kinetics to be hydrolyzed by trypsin. Though the trypsin digestion plays an important role in proteome analysis, systematic studying on the kinetics of trypsin-catalyzed reactions in proteome samples was not reported. In a proteome sample, huge number of proteins with different physical chemical properties coexists. If any type of proteins could be preferably digested, then limited digestion could be applied to reduce the sample complexity. However, no one has attempted to investigate the digestion priority of different proteins in a typical proteome study. To better understand the trypsin-catalyzed digestion process, kinetics study of this important process is required. There are numerous studies on kinetics analysis of trypsin-catalyzed hydrolysis of lysine or arginine derivatives [7, 8]. In these studies, a single substrate with only one cleavage site was
2 6248 Y. Pan et al. incubated with trypsin for determination of kinetics constants. The kinetics constants for peptides can also be determined in the same way when mass spectrometry was applied to monitor the reactions [9]. For these studies, one enzymatic reaction can only determine the kinetics constants for one substrate. In theory, the cleavage priority of cleavage sites in all proteins in proteome samples could be determined by synthesizing peptides centered with these sites. However, this approach is expensive and time-consuming. Though a single protein was also reported to study trypsin digestion kinetics, these studies cannot reflect the kinetics of the cleavage sites [10, 11]. Walmsley et al. [12] compared peptide abundances in 2- and 18-h human serum albumin (HSA) digests using label-free quantification and principal components analysis (PCA) and found that many cleavage sites showed variable digestion kinetics patterns. However, systematic investigation of the kinetics of the cleavage sites on proteins in complex proteome samples is still needed. Quantitative proteomics, especially stable isotope labeling, is a powerful tool for determining the different peptide amount between different proteome samples in high throughput. Recently, we have demonstrated in a preliminary study that quantitative proteomics could be a powerful tool to study the kinetics and digestion priority of trypsin digestion [13]. Triplex stable isotope dimethyl-labeling approach enable the simultaneous analysis of three samples in one time, and it is a reliable, a cost-effective, and an undemanding procedure that can be easily automated and applied in high-throughput proteomics experiments [14]. So, we employed the triplex stable isotope dimethyl-labeling approach to investigate the kinetics behavior of trypsin digestion in detail in this study. Proteome samples were digested with three different times, and then the resultant digests were labeled with stable isotope dimethyl labels, respectively. After quantitative proteomics analysis, 10,483 unique peptides from 2,270 proteins were quantified. Based on the abundance changes of the generated peptides during this time course study, four types of cleavage sites, i.e., very fast, fast, slow, and very slow, were determined. This enabled the investigation of cut priority of cleavage sites surrounded with different residues. It was found that the cleavage sites surrounded by neutral residues could be quickly cut, while those with neighboring charged residues (D/E/K/R) or proline residue (P) could be slowly cut. Because the quantified peptides could be classified into early and later generated peptides, the digestion priority of proteins with different physicochemical properties can also be investigated. In general, the results show that protein abundance and other physicochemical properties, such as molecular weight (Mw), grand average of hydropathicity (GRAVY), aliphatic index, and isoelectric point (pi) has no notable influence on the digestion priority of proteins. Experimental section Reagents and chemicals All the water used in this experiment was prepared using a Milli-Q system (Millipore, Bedford, MA). Formic acid (FA) was provided by Fluka (Buchs, Germany). Acetonitrile (ACN, HPLC grade) was purchased from Merck (Darmstadt, Germany). All the other chemicals and reagents were purchased from Sigma (St. Louis, MO). Fused silica capillaries with 75 μm i.d. were obtained from Polymicro Technologies (Phoenix, AZ). Cell growth and lysis The HeLa cells were grown according to Bian et al. [15]. The cell pellets were softly homogenized in a cold lysis buffer containing 8 M urea, 50 mm triethyl ammonium bicarbonate (TEAB; ph=8.0), 2 % protease cocktail (v/v), 1 % Triton X-100 (v/v), 65 mm dithiothreitol (DTT), 1 mm EDTA, 1 mm EDGA, 1 mm PMSF, 1 mm NaF, and 1 mm Na 3 VO 4, sonicated for 400 W 120 s, and centrifuged at 23,000g for 1 h. The supernatant containing the total cell proteins was precipitated with five volumes of cold acetone/ ethanol/acetic acid (v/v/v=50/50/0.1) at 20 C. Protein precipitant was centrifuged at 15,000g for 30 min. The pellet was washed separately with acetone and 75 % ethanol, then lyophilized to dryness, and stored at 80 C. Protein digestion and stable isotope dimethyl labeling Proteins (1 mg, the protein concentration was determined by Bradford assay) were denatured and reduced in 1 ml of 8 M urea, 100 mm TEAB, ph 8.0, and 10 mm DTT at 56 C for 40 min and then alkylated by 20 mm IAA in the darkness at room temperature for 30 min. The sample was diluted to 8 ml (1 M urea) with 100 mm TEAB, ph 8.0. Trypsin (Sigma) was added at a 40:1 protein/protease mass ratio along with CaCl 2 to 1 mm for digestion at 37 C; after digested for 1, 4, and 18 h, three same aliquots (100 μg) of samples were removed from the tube. To prevent further digestion, the trypsin was inhibited by addition of aprotinin (final concentration, 2.0 μg/ ml). Then, the three aliquots above were desalted by solidphase extraction (SPE) column, lyophilized, and labeled with light, intermediate, and heavy dimethyl, respectively. For the triplex stable isotope dimethyl labeling [14, 16], 50 μlofch 2 O(4%,v/v)CD 2 O(4%,v/v)and 13 CD 2 O(4%, v/v) were added into the sample solutions, respectively, and then 50 μl of freshly prepared NaBH 3 CN (0.6 M), NaBH 3 CN (0.6 M), and NaBD 3 CN (0.6 M) were added subsequently. The resultant mixture was incubated for 1 h at room temperature. Then, 10 μl of ammonia (25 %) and 25 μloffawere added to consume the excess labeling reagents and to acidify
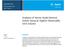
3 Quantitative proteomics reveals the kinetics of protein digestion 6249 the sample. After mixing in a ratio of 1:1:1 on the basis of the total peptide amount, the labeled peptide mixture was desalted by the SPE column. Dry the samples by vacuum centrifugation and stored at 80 C until used. Nano LC-MS/MS analysis For 2D strong cation exchange (SCX)-RP LC-MS/MS analysis with the LTQ-Orbitrap mass spectrometer (Velos, Thermo Fisher Scientific), a capillary monolithic column (5 cm 200 μm ID) with phosphate functional groups was applied as an SCX trap column in the first dimension. The sample loading and analysis procedures were as follows: the peptide samples were first dissolved in 0.1 % (v/v) formicacidin water, and then loaded onto the monolith SCX trap column; the trap column was equilibrated with 0.1 % (v/v)formicacid in water for 10 min [17]. After that, it was directly connected to an RP analytical column in tandem by a union. Then, six gradient elution steps were applied to gradually elute peptides from the SCX trap column to the RP analytical column with ammonium acetate solution concentrations of 50, 150, 250, 350, 500, and 1,000 mm, respectively. After each elution step, a subsequent RP LC-MS/MS was executed in 150-min gradient time with 0.1 % (v/v)formicacidinacetonitrilefrom5to 35 % (v/v); a capillary column was first manually pulled to a fine point as spray tip, and then packed with C18 AQ beads (3 μm, 120 Å, Michrom Bio Resources). All MS and MS/MS spectra were acquired in the data-dependent mode with the 20 most intense ions fragmented by CID. multiplied by 1,000 and divided by its amino acid frequency of occurrence in the database to form the normalized peptide sequences [20]. The pi, GRAVY, and aliphatic index of proteins and peptides were calculated according to ExPASy ( Results and discussion Time course study of trypsin digestion of proteome sample Time course investigation of trypsin digestion of proteome samples was performed with quantitative proteomics at three time points. As shown in Fig. 1, the lysate of HeLa cells (1 mg) was firstly subjected to trypsin digestion, and the same aliquots (100 μg) of the digestion were removed at time points of 0.5, 2, and 18 h, respectively. The trypsin inhibitor aprotinin were added to the three-time course digestion immediately to avoid further digestion. Then, quantitative proteomics using triplex stable isotope dimethyl labeling was applied to monitor the abundance variation of generated peptides at different time points. The digests from the three time points were labeled with light (0.5-h digestion), intermediate (2-h digestion), and heavy dimethyl (18-h digestion), respectively. The labeled Data analysis Protein quantification was performed using MaxQuant (version , [18]. The raw files were searched against UniProt database of human downloaded from (released on 12/11/ 2013); carbamidomethylation on cysteine was set as a fixed modification, and oxidation on methionine was set as variable modifications. Peptides were searched using fully tryptic cleavage constraints and up to four missed cleavage sites were allowed; the mass tolerances for the precursor ions and fragment ions were set to 6 ppm and 0.5 Da, respectively. For quantification, stable isotope dimethyl labeling, different quantification modes integrated into MaxQuant were selected, respectively. The other settings were the same to the conventional search. Sequence logos were automatically generated by the WebLogo ( logo.cgi) [19]. The raw sequences for WebLogo analysis were centered at the cleavage site and extended 13 residues (±6 residues). The N- or C-terminal sequences that could not be extended were excluded. In order to eliminate the influence of the relative occurrence of different amino acids in the proteome, the raw sequences for WebLogo analysis were Fig. 1 Experimental scheme for investigating the kinetics of trypsincatalyzed protein digestion by quantitative proteomics
4 6250 Y. Pan et al. peptides from the above three aliquots (10 μg each) were combined and analyzed with 2D RP LC-MS/MS. The acquired raw files from two technical replication runs were processed using the MaxQuant platform. To keep only the highly reliable quantified results, more strict criteria were applied to filter the data, the peptide should be quantified with RSD <50 % in both runs [16, 17, 21], which led to the quantification of 10,483 unique peptides from 2,270 proteins. It was found that 42.6 % (11,128/23,346) peptides have at least one missed cleavage sites. In a control experiment where the digestion was performed as in the general proteomics experiment, it was found that 25.8 % (814/3,157) peptides have more than one missed cleavage sites. The high frequency of missed cleavage sites observed for the peptides quantified in this time course study indicated that the digestion in the initial digestion stage is not complete. The distributions of log2 ratios M/L, H/M, and H/L were given in Fig. S1 in the Electronic Supplementary Material; the percentages of peptides with log2 ratios M/L, H/M, and H/L out of [ 1, 1] were 61.6, 34.0, and 47.5 %, respectively. In quantitative proteomics experiments, the percentages are usually less than 2 % if the same amount of sample was used [22]. Clearly, the concentration of many peptides changed significantly during the different digestion time which indicated that these peptides were generated at different speed. To directly reflect the change of peptide abundance during the time course, the peak areas of the three isotopic peaks were normalized by that of the medium one. Therefore, the abundances of the generated peptides were represented as L/M, M/M, and H/M. The log2 ratios out of [ 1, 1] were considered as significant change and between [ 1, 1] were considered as unchanged. Except the log2 ratios L/M, M/M, and H/M, log2 ratios H/L were employed to remove the uncorrected ones. Because there are three time points, in theory, the dynamic change of the peptide concentration during the digestion can be clustered into nine types (Fig. 2 and Electronic Supplementary Material (ESM) Table S1). Cluster 1: The concentration of these peptides does not change significantly for all the three time points. Cluster 2: the peptide concentrations do not change much from 0.5 to 2 h, but decreased in 18 h. Cluster 3: the peptide concentration decreased from 0.5 to 2 h, but do not change much in 18 h. Cluster 4: the peptide concentrations decreased during all the digestion steps. The peptides from above four clusters have their peak concentrations at the time point of 1 h, indicating that these peptides were mainly generated in the first digestion step, and were classified as the early generated peptides (Table 1). Except cluster 1, the concentrations of other peptides decreased with further digestion, indicating that these peptides were gradually degraded. We compared the percentage of peptides with missed cleavage sites for these four clusters. It was found that 32.7%ofpeptidesincluster1havemissedcleavagesites, while 64.8, 90.0, and 91.3 % peptides in other three clusters have at least one missed cleavage sites (Fig. 3a). These data illustrated that the peptides in clusters 2, 3, and 4 were degraded with further digestion probably because the missed cleavage sites were slow ones which were cut afterwards. The rest of the peptides have their peak concentrations at other time points (Fig. 2). Cluster 5: the concentration of these peptides increased from 0.5 to 2 h, but remains unchanged afterwards. Cluster 6: the peptide concentration increased from 0.5 to 2 h, and then decreased. The peptides in above two clusters have their peak concentrations at 2 h, and they were mainly generated at the second digestion stage. Cluster 7: the peptides with concentration increased all the time. Cluster 8: the abundance of these peptides does not change from 0.5 to 2 h, but increased in 18 h. These peptides have their peak concentrations at 18 h, and they were mainly generated at the last digestion stage. The last cluster 9: the abundance of these peptides decreased from 0.5 to 2 h, and then increased from 2 to 18 h. It is hard to explain the abundance change for these peptides. Because only 0.25 % peptides (21/8,334) belong to this cluster, these peptides were not considered seriously in this study. Because the peptides in the last four clusters were mainly generated in the last two digestion step, they were classified as the late generated peptides (Table 1). It can be seen from Fig. 3a that the peptides in clusters 2, 3, 4, 6, and 9 have higher frequency of missed cleavage sites. Interestingly, the peptides in these clusters decreased in abundance at least in one interval of the three time points as evidenced in Fig. 2. And if the peptides in the cluster did not decrease in abundance in any of the intervals (clusters 1, 5, 7, 8), their frequencies of having missed cleavage were much lower. This is because the peptides with missed cleavage sites tend to be further cut during the digestion, which led to the decrease of their abundance. The high consistent of the frequency of missed cleavage sites with the abundance change during the time course indicated that these peptides were accurately quantified. Investigation of the cut priority of cleavage sites surrounded with different residues Each protein has many trypsin cleavage sites. After trypsin digestion, proteins will be digested into many peptides. The above quantitative proteomics study indicated that these peptides are not generated at the same speed. The reason that some peptides were generated earlier and some peptide generated later is that the trypsin-catalyzed hydrolysis rates are different for different cleavage sites. Though this quantitative proteomics cannot directly determine the kinetics constants for this enzymatic reaction, it can reveal the cut priority of the cleavage sites. The sequence surrounding the cleavage site can be described as P 4 -P 3 -P 2 -P 1 -P 1 -P 2 -P 3 -P 4 [23], where cleavage occurs between P 1 and P 1. For trypsin-catalyzed cleavage, all P 1 positions are either K or R. It is of interest to
5 Quantitative proteomics reveals the kinetics of protein digestion 6251 Fig. 2 Clusters of the peptides according to their abundance change during the time course investigate which types of residues surrounding the cleavage sites affect their digestion kinetics. Except for the peptides generated from protein N-/C-termini, the majority of peptides are generated by two trypsin cleavages. Thus, for each identified peptide, it typically has two terminal cleavage sites. Take a quantified peptide, FIDTTSKFGHGR, as the example. The N-terminal residue (F) on this peptide is the P 1 residue for the N-terminal cleavage site, while the C-terminal residue (R) is the P 1 residue. To extract the residues surrounding the cleavage sites, the identified peptides should be mapped to their parent proteins. For above the peptide, the sequences surrounding the two cleavage sites were determined to be DLK.FID and HGR.FQT after mapping to their parent protein Table 1 The numbers of the early and late generated peptides Early generated peptides Late generated peptides Cluster 1 (2,156) Cluster 5 (2,932) Cluster 2 (1,179) Cluster 6 (1,692) Cluster 3 (189) Cluster 7 (6) Cluster 4 (126) Cluster 8 (12) sequence, respectively. In this way, the residues around the peptide terminal cleavage sites could be determined. It is well known that K/R with a neighboring P residue on the C-terminal side (P 1 position) and K/R with an aspartic acid (D) or glutamic acid (E) residue on either the N- or C-terminal side (P 2 or P 1 position) were difficult to cut [3 6, 24]. We first investigate if there were notable differences in distribution of P on P 1 position and D/E on (P 2 and P 1 position) for the terminal cleavage sites on above nine clusters of peptides. As shown in Fig. 3b, the percentages of P residue on this position were all <0.15 %, which is far less than the nature P residue frequency of the human proteome (about 6.3 % as shown in ESM Fig. S2) [25, 26] and the frequency of P followed by K/R in database (5.7 %). It indicated that K/R followed with P was not likely cut by trypsin. This is consistent to Keil rules [27], the commonly accepted rule for a trypsin cut site is K/R.P. The percentages of negative charged amino acid residues D/E (P 2 or P 1 position) were less than the native frequency of the human proteome ( 13 %) for clusters 1 4, while the percentages for clusters 5 8 were higher than the native frequency of the human proteome. Based on the quantified ratios, the peptides in clusters 1 4 were generated earlier than those in clusters 5 8 during the digestion.
6 6252 Y. Pan et al. Fig. 3 Percentages of peptides (a) with missed cleavage sites; (b) with D/E (on P 2 and P 1 position), K/R (on P 1 position), and P (on P 1 position) for terminal cleavage sites; and (c) with D/E, K/R, and P (their positions were as in (b)) for missed cleavage sites in nine clusters
7 Quantitative proteomics reveals the kinetics of protein digestion 6253 Fig. 4 Sequence logos of four cleavage site types with different kinetics (very fast, fast, slow, and very slow sites). a All sequences, b the sequences only consider K as the cleavage site, and c the sequences only consider R as the cleavage site. The frequencies of amino acids in the peptides were normalized by their occurrence frequency in the proteome database The high frequency of acidic residues in late generated peptides means that the trypsin-catalyzed reaction is slow when K/R with neighboring D/E. This is also consistent with the fact that K/R with neighboring D/E is likely to be missed cleavage. In addition to the cleavage sites revealed from the peptide termini, there were missed cleavage sites on some of the identified peptides. Still, take the identified peptide FIDTTSKFGHGR as an example. It has one missed cleavage site. The sequence centered with the cleavage site was TSK.FGH. We then compared the distributions of P on P 1 position and D/E on P 2 or P 1 position for the missed cleavage sites on above nine clusters of peptides. As shown in Fig. 3c,the percentages for P were far higher than the N- and C-terminal cleavage sites, and similar with or slightly higher than the nature P amino acid composition of the human proteome. This confirmed that K/R followed by P was not likely cut by trypsin. The percentages of negative charged amino acids D/E (>18.4 %) were all higher than the native composition of the human proteome (D/E about 12 %) for all clusters. These percentages are higher than those of cleavage sites revealed by the terminal sites of the early generated peptides (clusters 1 4 in Fig.3b) while are quite similar with those of the terminal sites for the late generated peptides (clusters 5 8inFig.3b). This is not surprising since the missed cleavage sites were relatively slow. As shown above, the cleavage sites could be revealed by the quantified peptides with either terminal sites or missed cleavage sites. To investigate the cut priority of cleavage sites, they must be sorted in the order of their kinetics. Depending on when the sites got cut, the cleavage sites are classified into four types, i.e., very fast, fast, slow, and very slow sites (for details, see the Supplementary Note and Table S2 in the ESM). The fast cleavage sites (5,942) were the sites got cut in the first digestion step. Both C- and N-terminal sites for the early generated peptides (clusters 1 4) are fast cleavage sites because they were generated in the first digestion step. The slow cleavage sites (105) were the sites got cut in the second digestion step. The missed cleavage sites in peptides of cluster 3 containing one missed cleavage site belongs to this class because the concentrations of peptides in this cluster do not change much from 0.5 to 2 h, but decreased in 18 h, indicating that these peptides got cut in the second digestion step. The slower cleavage sites (1,161) were the sites got cut in the third digestion step. Based on the change of peptide concentration, the missed cleavage sites on peptides of cluster 2 and cluster 6 with one missed cleavage site belong to this type. The slowest cleavage sites (1,682) were the sites cannot be cut at any digestion steps. They are the missed cleavage sites on peptides from cluster 1, cluster 5, and cluster 8.
8 6254 Y. Pan et al. The sequence logos for the normalized peptide sequences centered with above four types of cleavage sites were generated by the WebLogo and are shown in Fig. 4. It is obvious that the cleavage sites K/R surrounded by neutral residues could be quickly cut, while those with neighboring charged residues (D/E/K/R) or P could be slowly cut (Fig. 4a). For the two types of fast cleavage sites, i.e., very fast and fast sites, R residues on P 1 position account for 55.3 and 44.2 % of all sites, respectively. While for the two types of slow cleavage sites, the R sites account for less than 25 % (ESM Fig. S3). This indicated that trypsin cleaves the C-terminal to K and R residues with higher rates for R. We are curious if there is any difference in the effects of surrounding residues on the kinetics of cleavage sites K and R. For this purpose, we generated the sequence logos for cleavage sites K and R separately (Fig. 4b, c). In general, they are quite similar but there are some differences. To examine the effects of surrounding residues on the kinetics of cleavage sites in detail, we compared the distribution of the neighboring residues (ESM Fig. S4). An interesting phenomenon is that the K/R following P (P 2 position) tends to be very fast cleavage sites, the special configuration of proline and the small side chain probably allow trypsin access more easily the cleavage sites. The presence of D/E on P 2,P 1,andP 2 positions of cleavage sites K make the kinetics slow, and the P 2 position is more sensitive to D/E; the similar situation was observed in cleavage sites R. Cleavage sites R with alkaline amino acid R on P 2 position were more difficult to be cut than acidic amino acids, the opposite situation occurred at the cleavage sites K. Investigation of the digestion priority of proteins with different physicochemical properties It is of interest to investigate if the peptides generated at different time points have correlation with different types of proteins. The quantified peptides were classified into two types: (1) the early generated peptides, these peptides are mainly generated at the first digestion stage (clusters 1 4) and (2) the late generated peptides, these peptides are mainly generated at the second and third digestion stages (clusters 5 8). There were 3,650 and 4,642 unique peptides that correspond to 1,397 and 1,346 proteins for the above two types of peptides, respectively. If a specific type of proteins is digested earlier during a digestion, then more peptides should present in the pool of the early generated peptides. Therefore, the protein digestion priority could be judged according to the distribution of early and late generated peptides across different physicochemical properties of their parent proteins. We have investigated the digestion priority of proteins relative to their abundances with a much smaller dataset [13]. In this study, we further investigated this correlation with much bigger dataset. The quantified proteins were classified into 16 bins according to their spectra counts, which approximately represented their abundances. Then, the distributions of early and late generated peptides across the spectra counts were investigated (Fig. 5). The overall distributions between the two types of peptides are quite similar, suggesting that the digestion priority of individual proteins is almost independent of their abundances. However, there is a subtle difference between these two distributions. The percentages for early generated peptides are slightly higher than those for late generated peptides in the low spectra count range, while the difference is opposite in the high spectra count range but the difference can be ignored when the spectra counts were larger than 32. If the protein spectra count accurately reflects the protein abundance [28 30], then the low abundance proteins are digested slightly earlier than the high abundance ones in general. We then investigated if the early/late generated peptides preferably derived from proteins with some types of physicochemical properties. We first investigated the digestion priority for proteins with different sizes. The Mw for the majority of the identified proteins were in the range of 30,000 to 100,000 Da. If the proteins are not denatured, then trypsin may have difficulty to access the cleavage sites buried inside big proteins, and so, the more late generated peptides should be derived from the big proteins. However, it was found that the distributions of the two types of peptides across the Mw of proteins are also very similar (ESM Fig. S5), indicating the digestion priority of proteins does not depend on their sizes. This is not surprising since the proteins are denatured, and so, Fig. 5 The distribution of the early generated peptides and the late generated peptides across the protein spectra counts. The percentages on the Y-axis are the percentages of the early or late generated peptides within each bin of log2 (spectra counts) of the proteins they derived from
9 Quantitative proteomics reveals the kinetics of protein digestion 6255 the accessibility of trypsin to the cleavage sites on the proteins with different sizes is similar. As trypsin cut the sites surrounded with neutral residues with high rate, it is of interest to investigate if the digestion priority depends on their hydrophobicity. The GRAVY value for a protein is calculated as the sum of hydropathicity values of all the amino acid residues divided by the number of residues in the sequence [31]. The aliphatic index of a protein is defined as the relative volume occupied by aliphatic side chains (alanine, valine, isoleucine, and leucine). An increase in the aliphatic index increases the thermostability of globular proteins [32]. Both GRAVY and aliphatic index reflect the hydrophobicity of proteins. It can be found that the distributions of early and late generated peptides across either GRAVY value or aliphatic index have no notable differences (ESM Fig. S6 and S7), indicating that there are no much differences in the digestion priority of proteins with different hydrophobicity. The GRAVY values of most proteins were <0, showing that the digested proteins were relatively hydrophilic because of the lysis buffer solution used here. Finally, we investigated the dependence of digestion priority on protein s pi values and no dependence was observed either (ESM Fig. S8). Above data indicated that digestion priority does not depend on the physicochemical properties of proteins investigated in general. Conclusions To study the kinetics of trypsin-catalyzed protein digestion, quantitative proteomics was applied to monitor the dynamics of the generated peptides from trypsin digestion of a proteome sample in a time course study. According to the dynamic change of peptide abundance, the peptides were divided into two types: the early generated peptides and the late generated peptides. In general, the trypsin-catalyzed digestion priority of individual proteins in a proteome sample is independent of their abundances and other physicochemical properties, such as Mw, GRAVY, aliphatic index, and pi. Thus, selective enrichment or depletion of specific type of proteins via limited digestion is likely impossible. The data also indicate that the priority order of cleavage depends largely on the kinetics properties of the cleavage sites, i.e., the residues surrounding the cleavage sites in proteins. The high consistency of the slow cleavage sites with the reported missed cleavage sites indicated that the quantitative proteomics approach is a good approach to compare the kinetics of trypsin-catalyzed cleavage. Acknowledgments This work was supported by the China State Key Basic Research Program Grant (2013CB911202, 2012CB910101, and 2012CB910604), the Creative Research Group Project of NSFC ( ), the National Natural Science Foundation of China ( , , , and ), National Key Special Program on Infection diseases (2012ZX ), and Analytical Method Innovation Program of MOST (2012IM030900). References 1. Hunt DF, Yates JR, Shabanowitz J, Winston S, Hauer CR (1986) Protein sequencing by tandem mass spectrometry. Proc Natl Acad Sci U S A 83(17): Wu C, Tran JC, Zamdborg L, Durbin KR, Li M, Ahlf DR, Early BP, Thomas PM, Sweedler JV, Kelleher NL (2012) A protease for middle-down proteomics. Nat Methods 9(8): Olsen JV, Ong S-E, Mann M (2004) Trypsin cleaves exclusively C- terminal to arginine and lysine residues. Mol Cell Proteomics 3(6): Siepen JA, Keevil E-J, Knight D, Hubbard SJ (2007) Prediction of missed cleavage sites in tryptic peptides aids protein identification in proteomics. J Proteome Res 6(1): Lawless C, Hubbard SJ (2012) Prediction of missed proteolytic cleavages for the selection of surrogate peptides for quantitative proteomics. Omics 16(9): Gershon PD (2013) Cleaved and missed sites for trypsin, Lys-C and Lys-N can be predicted with high confidence on the basis of sequence context. J Proteome Res 13(2): Wang S-S, Carpenter FH (1968) Kinetic studies at high ph of the trypsin-catalyzed hydrolysis of N α -benzoyl derivatives of L- arginamide, L-lysinamide, and S-2-aminoethyl-L-cysteinamide and related compounds. J Biol Chem 243(13): Simpson B, Haard N (1984) Purification and characterization of trypsin from the Greenland cod (Gadus ogac). 1. Kinetic and thermodynamic characteristics. Can J Biochem Cell Biol 62(9): Caprioli RM, Smith L (1986) Determination of K m and V max for tryptic peptide hydrolysis using fast atom bombardment mass spectrometry. Anal Chem 58(6): Fraser D, Powell RE (1950) The kinetics of trypsin digestion. J Biol Chem 187: Halsey JF, Harrington WF (1973) Substructure of paramyosin. Correlation of helix stability, trypsin digestion kinetics, and amino acid composition. Biochemistry 12(4): Walmsley SJ, Rudnick PA, Liang Y, Dong Q, Stein SE, Nesvizhskii AI (2013) Comprehensive analysis of protein digestion using six trypsins reveals the origin of trypsin as a significant source of variability in proteomics. J Proteome Res 12(12): Ye M, Pan Y, Cheng K, Zou H (2014) Protein digestion priority is independent of protein abundances. Nat Methods 11(3): Boersema PJ, Raijmakers R, Lemeer S, Mohammed S, Heck AJR (2009) Multiplex peptide stable isotope dimethyl labeling for quantitative proteomics. Nat Protoc 4(4): Bian Y, Ye M, Song C, Cheng K, Wang C, Wei X, Zhu J, Chen R, Wang F, Zou H (2012) Improve the coverage for the analysis of phosphoproteome of HeLa cells by a tandem digestion approach. J Proteome Res 11(5): Song C, Wang F, Ye M, Cheng K, Chen R, Zhu J, Tan Y, Wang H, Figeys D, Zou H (2011) Improvement of the quantification accuracy and throughput for phosphoproteome analysis by a pseudo triplex stable isotope dimethyl labeling approach. Anal Chem 83(20): WangF,ChenR,ZhuJ,SunD,SongC,WuY,YeM,WangL,ZouH (2010) A fully automated system with online sample loading, isotope dimethyl labeling and multidimensional separation for highthroughput quantitative proteome analysis. Anal Chem 82(7): Cox J, Mann M (2008) MaxQuant enables high peptide identification rates, individualized ppb-range mass accuracies and proteome-wide protein quantification. Nat Biotechnol 26(12): Crooks GE, Hon G, Chandonia J-M, Brenner SE (2004) WebLogo: a sequence logo generator. Genome Res 14(6): Rodriguez J, Gupta N, Smith RD, Pevzner PA (2007) Does trypsin cut before proline? J Proteome Res 7(1):
10 6256 Y. Pan et al. 21. Yang S, Nie A, Zhang L, Yan G, Yao J, Xie L, Lu H, Yang P (2012) A novel quantitative proteomics workflow by isobaric terminal labeling. J Proteome 75(18): Li Z, Adams RM, Chourey K, Hurst GB, Hettich RL, Pan C (2012) Systematic comparison of label-free, metabolic labeling, and isobaric chemical labeling for quantitative proteomics on LTQ Orbitrap velos. J Proteome Res 11(3): Schechter I, Berger A (1967) On the size of the active site in proteases. I Papain. Biochem Biophys Res Commun 27(2): Thiede B, Lamer S, Mattow J, Siejak F, Dimmler C, Rudel T, Jungblut PR (2000) Analysis of missed cleavage sites, tryptophan oxidation and N-terminal pyroglutamylation after in-gel tryptic digestion. Rapid Commun Mass Spectrom 14(6): Switzar L, Giera M, Niessen WM (2013) Protein digestion: an overview of the available techniques and recent developments. J Proteome Res 12(3): Apweiler R, Biswas M, Fleischmann W, Kanapin A, Karavidopoulou Y, Kersey P, Kriventseva EV, Mittard V, Mulder N, Phan I (2001) Proteome analysis database: online application of InterPro and CluSTr for the functional classification of proteins in whole genomes. Nucleic Acids Res 29(1): Keil B (1992) Specificity of proteolysis. Springer, Berlin 28. Liu H, Sadygov RG, Yates JR (2004) A model for random sampling and estimation of relative protein abundance in shotgun proteomics. Anal Chem 76(14): Old WM, Meyer-Arendt K, Aveline-Wolf L, Pierce KG, Mendoza A, Sevinsky JR, Resing KA, Ahn NG (2005) Comparison of label-free methods for quantifying human proteins by shotgun proteomics. Mol Cell Proteomics 4(10): Ning K, Fermin D, Nesvizhskii AI (2012) Comparative analysis of different label-free mass spectrometry based protein abundance estimates and their correlation with RNA-Seq gene expression data. J Proteome Res 11(4): Kyte J, Doolittle RF (1982) A simple method for displaying the hydropathic character of a protein. J Mol Biol 157(1): Atsushi I (1980) Thermostability and aliphatic index of globular proteins. J Biochem 88(6):
The distribution of log 2 ratio (H/L) for quantified peptides. cleavage sites in each bin of log 2 ratio of quantified. peptides
 Journal: Nature Methods Article Title: Corresponding Author: Protein digestion priority is independent of their abundances Mingliang Ye and Hanfa Zou Supplementary Figure 1 Supplementary Figure 2 The distribution
Journal: Nature Methods Article Title: Corresponding Author: Protein digestion priority is independent of their abundances Mingliang Ye and Hanfa Zou Supplementary Figure 1 Supplementary Figure 2 The distribution
Application of a new capillary HPLC- ICP-MS interface to the identification of selenium-containing proteins in selenized yeast
 Application of a new capillary HPLC- ICP-MS interface to the identification of selenium-containing proteins in selenized yeast Application note Food supplements Authors Juliusz Bianga and Joanna Szpunar
Application of a new capillary HPLC- ICP-MS interface to the identification of selenium-containing proteins in selenized yeast Application note Food supplements Authors Juliusz Bianga and Joanna Szpunar
Characterization of Disulfide Linkages in Proteins by 193 nm Ultraviolet Photodissociation (UVPD) Mass Spectrometry. Supporting Information
 Characterization of Disulfide Linkages in Proteins by 193 nm Ultraviolet Photodissociation (UVPD) Mass Spectrometry M. Montana Quick, Christopher M. Crittenden, Jake A. Rosenberg, and Jennifer S. Brodbelt
Characterization of Disulfide Linkages in Proteins by 193 nm Ultraviolet Photodissociation (UVPD) Mass Spectrometry M. Montana Quick, Christopher M. Crittenden, Jake A. Rosenberg, and Jennifer S. Brodbelt
4th Multidimensional Chromatography Workshop Toronto (January, 2013) Herman C. Lam, Ph.D. Calibration & Validation Group
 4th Multidimensional Chromatography Workshop Toronto (January, 2013) Herman C. Lam, Ph.D. Calibration & Validation Group MDLC for Shotgun Proteomics Introduction General concepts Advantages Challenges
4th Multidimensional Chromatography Workshop Toronto (January, 2013) Herman C. Lam, Ph.D. Calibration & Validation Group MDLC for Shotgun Proteomics Introduction General concepts Advantages Challenges
Mass Spectrometry and Proteomics - Lecture 4 - Matthias Trost Newcastle University
 Mass Spectrometry and Proteomics - Lecture 4 - Matthias Trost Newcastle University matthias.trost@ncl.ac.uk previously Peptide fragmentation Hybrid instruments 117 The Building Blocks of Life DNA RNA Proteins
Mass Spectrometry and Proteomics - Lecture 4 - Matthias Trost Newcastle University matthias.trost@ncl.ac.uk previously Peptide fragmentation Hybrid instruments 117 The Building Blocks of Life DNA RNA Proteins
In-Solution Digestion for proteomics
 In-Solution Digestion for proteomics Guidelines for sample preparation (How to protect your samples from contamination with keratin) 1. Try to avoid any contact of samples and solutions with dust, skin
In-Solution Digestion for proteomics Guidelines for sample preparation (How to protect your samples from contamination with keratin) 1. Try to avoid any contact of samples and solutions with dust, skin
Nature Methods: doi: /nmeth.3177
 Supplementary Figure 1 Characterization of LysargiNase, trypsin and LysN missed cleavages. (a) Proportion of peptides identified in LysargiNase and trypsin digests of MDA-MB-231 cell lysates carrying 0,
Supplementary Figure 1 Characterization of LysargiNase, trypsin and LysN missed cleavages. (a) Proportion of peptides identified in LysargiNase and trypsin digests of MDA-MB-231 cell lysates carrying 0,
2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry
 Dr. Sanjeeva Srivastava 1. Fundamental of Mass Spectrometry Role of MS and basic concepts 2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry 2 1 MS basic concepts Mass spectrometry - technique
Dr. Sanjeeva Srivastava 1. Fundamental of Mass Spectrometry Role of MS and basic concepts 2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry 2 1 MS basic concepts Mass spectrometry - technique
Trypsin Mass Spectrometry Grade
 058PR-03 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Trypsin Mass Spectrometry Grade A Chemically Modified, TPCK treated, Affinity Purified
058PR-03 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Trypsin Mass Spectrometry Grade A Chemically Modified, TPCK treated, Affinity Purified
Supporting information
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2014 Supporting information Glycan Reductive Isotope-coded Amino Acid Labeling (GRIAL) for Mass Spectrometry-based
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2014 Supporting information Glycan Reductive Isotope-coded Amino Acid Labeling (GRIAL) for Mass Spectrometry-based
Multiplex Protein Quantitation using itraq Reagents in a Gel-Based Workflow
 Multiplex Protein Quantitation using itraq Reagents in a Gel-Based Workflow Purpose Described herein is a workflow that combines the isobaric tagging reagents, itraq Reagents, with the separation power
Multiplex Protein Quantitation using itraq Reagents in a Gel-Based Workflow Purpose Described herein is a workflow that combines the isobaric tagging reagents, itraq Reagents, with the separation power
Shotgun Proteomics MS/MS. Protein Mixture. proteolysis. Peptide Mixture. Time. Abundance. Abundance. m/z. Abundance. m/z 2. Abundance.
 Abundance Abundance Abundance Abundance Abundance Shotgun Proteomics Protein Mixture 1 2 3 MS/MS proteolysis m/z 2 3 Time µlc m/z MS 1 m/z Peptide Mixture m/z Block Diagram of a Mass Spectrometer Sample
Abundance Abundance Abundance Abundance Abundance Shotgun Proteomics Protein Mixture 1 2 3 MS/MS proteolysis m/z 2 3 Time µlc m/z MS 1 m/z Peptide Mixture m/z Block Diagram of a Mass Spectrometer Sample
REDOX PROTEOMICS. Roman Zubarev.
 REDOX PROTEOMICS Roman Zubarev Roman.Zubarev@ki.se Physiological Chemistry I, Department for Medical Biochemistry & Biophysics, Karolinska Institutet, Stockholm What is (RedOx) Proteomics? Proteomics -
REDOX PROTEOMICS Roman Zubarev Roman.Zubarev@ki.se Physiological Chemistry I, Department for Medical Biochemistry & Biophysics, Karolinska Institutet, Stockholm What is (RedOx) Proteomics? Proteomics -
NIH Public Access Author Manuscript J Proteome Res. Author manuscript; available in PMC 2014 July 05.
 NIH Public Access Author Manuscript Published in final edited form as: J Proteome Res. 2013 July 5; 12(7): 3071 3086. doi:10.1021/pr3011588. Evaluation and Optimization of Mass Spectrometric Settings during
NIH Public Access Author Manuscript Published in final edited form as: J Proteome Res. 2013 July 5; 12(7): 3071 3086. doi:10.1021/pr3011588. Evaluation and Optimization of Mass Spectrometric Settings during
Quantitation of Protein Phosphorylation Using Multiple Reaction Monitoring
 Quantitation of Protein Phosphorylation Using Multiple Reaction Monitoring Application Note Authors Ning Tang, Christine A. Miller and Keith Waddell Agilent Technologies, Inc. Santa Clara, CA USA This
Quantitation of Protein Phosphorylation Using Multiple Reaction Monitoring Application Note Authors Ning Tang, Christine A. Miller and Keith Waddell Agilent Technologies, Inc. Santa Clara, CA USA This
Molecular Cell, Volume 46. Supplemental Information
 Molecular Cell, Volume 46 Supplemental Information Mapping N-Glycosylation Sites across Seven Evolutionary Distant Species Reveals a Divergent Substrate Proteome Despite a Common Core Machinery Dorota
Molecular Cell, Volume 46 Supplemental Information Mapping N-Glycosylation Sites across Seven Evolutionary Distant Species Reveals a Divergent Substrate Proteome Despite a Common Core Machinery Dorota
User Guide. Protein Clpper. Statistical scoring of protease cleavage sites. 1. Introduction Protein Clpper Analysis Procedure...
 User Guide Protein Clpper Statistical scoring of protease cleavage sites Content 1. Introduction... 2 2. Protein Clpper Analysis Procedure... 3 3. Input and Output Files... 9 4. Contact Information...
User Guide Protein Clpper Statistical scoring of protease cleavage sites Content 1. Introduction... 2 2. Protein Clpper Analysis Procedure... 3 3. Input and Output Files... 9 4. Contact Information...
Lecture 3. Tandem MS & Protein Sequencing
 Lecture 3 Tandem MS & Protein Sequencing Nancy Allbritton, M.D., Ph.D. Department of Physiology & Biophysics 824-9137 (office) nlallbri@uci.edu Office- Rm D349 Medical Science D Bldg. Tandem MS Steps:
Lecture 3 Tandem MS & Protein Sequencing Nancy Allbritton, M.D., Ph.D. Department of Physiology & Biophysics 824-9137 (office) nlallbri@uci.edu Office- Rm D349 Medical Science D Bldg. Tandem MS Steps:
Supplementary Figures and Notes for Digestion and depletion of abundant
 Supplementary Figures and Notes for Digestion and depletion of abundant proteins improves proteomic coverage Bryan R. Fonslow, Benjamin D. Stein, Kristofor J. Webb, Tao Xu, Jeong Choi, Sung Kyu Park, and
Supplementary Figures and Notes for Digestion and depletion of abundant proteins improves proteomic coverage Bryan R. Fonslow, Benjamin D. Stein, Kristofor J. Webb, Tao Xu, Jeong Choi, Sung Kyu Park, and
Lys-C/Arg-C, a More Specific and Efficient Digestion Approach for
 Lys-C/Arg-C, a More Specific and Efficient Digestion Approach for Proteomics Studies Zhen Wu, Jichang Huang, Jingnan Huang, Qingqing Li, Xumin Zhang *, State Key Laboratory of Genetic Engineering, Department
Lys-C/Arg-C, a More Specific and Efficient Digestion Approach for Proteomics Studies Zhen Wu, Jichang Huang, Jingnan Huang, Qingqing Li, Xumin Zhang *, State Key Laboratory of Genetic Engineering, Department
Improve Protein Analysis with the New, Mass Spectrometry- Compatible ProteasMAX Surfactant
 Improve Protein Analysis with the New, Mass Spectrometry- Compatible Surfactant ABSTRACT Incomplete solubilization and digestion and poor peptide recovery are frequent limitations in protein sample preparation
Improve Protein Analysis with the New, Mass Spectrometry- Compatible Surfactant ABSTRACT Incomplete solubilization and digestion and poor peptide recovery are frequent limitations in protein sample preparation
Agilent Protein In-Gel Tryptic Digestion Kit
 Agilent 5188-2749 Protein In-Gel Tryptic Digestion Kit Agilent Protein In-Gel Tryptic Digestion Kit Instructions Kit Contents The Protein In-Gel Tryptic Digestion Kit includes sufficient reagents for approximately
Agilent 5188-2749 Protein In-Gel Tryptic Digestion Kit Agilent Protein In-Gel Tryptic Digestion Kit Instructions Kit Contents The Protein In-Gel Tryptic Digestion Kit includes sufficient reagents for approximately
PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System
 Application Note LC/MS PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System Purpose This application note describes an automated workflow
Application Note LC/MS PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System Purpose This application note describes an automated workflow
Quantitative chromatin proteomics reveals a dynamic histone. post-translational modification landscape that defines asexual
 Quantitative chromatin proteomics reveals a dynamic histone post-translational modification landscape that defines asexual and sexual Plasmodium falciparum parasites Nanika Coetzee 1, Simone Sidoli 2,
Quantitative chromatin proteomics reveals a dynamic histone post-translational modification landscape that defines asexual and sexual Plasmodium falciparum parasites Nanika Coetzee 1, Simone Sidoli 2,
Integration of Cell Lysis, Protein Extraction, and Digestion into One Step for Ultrafast Sample Preparation for Phosphoproteome Analysis
 pubs.acs.org/ac Integration of Cell Lysis, Protein Extraction, and Digestion into One Step for Ultrafast Sample Preparation for Phosphoproteome Analysis Fangjie Liu,, Mingliang Ye,*, Yanbo Pan,, Yi Zhang,,
pubs.acs.org/ac Integration of Cell Lysis, Protein Extraction, and Digestion into One Step for Ultrafast Sample Preparation for Phosphoproteome Analysis Fangjie Liu,, Mingliang Ye,*, Yanbo Pan,, Yi Zhang,,
Quantitative Analysis of Underivatized Amino Acids in Plant Matrix by Hydrophilic Interaction Chromatography (HILIC) with LC/MS Detection
 Application Note Food Testing, Metabolomics, Agricultural Chemistry, Environmental Quantitative Analysis of Underivatized Amino Acids in Plant Matrix by Hydrophilic Interaction Chromatography (HILIC) with
Application Note Food Testing, Metabolomics, Agricultural Chemistry, Environmental Quantitative Analysis of Underivatized Amino Acids in Plant Matrix by Hydrophilic Interaction Chromatography (HILIC) with
Universal sample preparation method for proteome analysis
 nature methods Universal sample preparation method for proteome analysis Jacek R Wi niewski, Alexandre Zougman, Nagarjuna Nagaraj & Matthias Mann Supplementary figures and text: Supplementary Figure 1
nature methods Universal sample preparation method for proteome analysis Jacek R Wi niewski, Alexandre Zougman, Nagarjuna Nagaraj & Matthias Mann Supplementary figures and text: Supplementary Figure 1
Quantification with Proteome Discoverer. Bernard Delanghe
 Quantification with Proteome Discoverer Bernard Delanghe Overview: Which approach to use? Proteome Discoverer Quantification Method What When to use Metabolic labeling SILAC Cell culture systems Small
Quantification with Proteome Discoverer Bernard Delanghe Overview: Which approach to use? Proteome Discoverer Quantification Method What When to use Metabolic labeling SILAC Cell culture systems Small
Supporting Information
 Supporting Information Mechanism of inactivation of -aminobutyric acid aminotransferase by (1S,3S)-3-amino-4-difluoromethylenyl-1- cyclopentanoic acid (CPP-115) Hyunbeom Lee, 1, Emma H. Doud, 1,2 Rui Wu,
Supporting Information Mechanism of inactivation of -aminobutyric acid aminotransferase by (1S,3S)-3-amino-4-difluoromethylenyl-1- cyclopentanoic acid (CPP-115) Hyunbeom Lee, 1, Emma H. Doud, 1,2 Rui Wu,
Supplementary Figure 1. Method development.
 Supplementary Figure 1 Method development. Titration experiments to determine standard antibody:lysate concentration. Lysates (~2 mg of total proteins) were prepared from cells expressing FLAG- tagged
Supplementary Figure 1 Method development. Titration experiments to determine standard antibody:lysate concentration. Lysates (~2 mg of total proteins) were prepared from cells expressing FLAG- tagged
Biomolecular Mass Spectrometry
 Lipids ot different than other organic small molecules Carbohydrates Polymers of monosaccharides linked via glycosidic bonds (acetals/ ketals) many different combinationsvery interesting no time ucleic
Lipids ot different than other organic small molecules Carbohydrates Polymers of monosaccharides linked via glycosidic bonds (acetals/ ketals) many different combinationsvery interesting no time ucleic
Supporting Information. Lysine Propionylation to Boost Proteome Sequence. Coverage and Enable a Silent SILAC Strategy for
 Supporting Information Lysine Propionylation to Boost Proteome Sequence Coverage and Enable a Silent SILAC Strategy for Relative Protein Quantification Christoph U. Schräder 1, Shaun Moore 1,2, Aaron A.
Supporting Information Lysine Propionylation to Boost Proteome Sequence Coverage and Enable a Silent SILAC Strategy for Relative Protein Quantification Christoph U. Schräder 1, Shaun Moore 1,2, Aaron A.
Enhancing Sequence Coverage in Proteomics Studies by Using a Combination of Proteolytic Enzymes
 Enhancing Sequence Coverage in Proteomics Studies by Using a Combination of Proteolytic Enzymes Dominic Baeumlisberger 2, Christopher Kurz 3, Tabiwang N. Arrey, Marion Rohmer 2, Carola Schiller 3, Thomas
Enhancing Sequence Coverage in Proteomics Studies by Using a Combination of Proteolytic Enzymes Dominic Baeumlisberger 2, Christopher Kurz 3, Tabiwang N. Arrey, Marion Rohmer 2, Carola Schiller 3, Thomas
Learning Objectives. Overview of topics to be discussed 10/25/2013 HIGH RESOLUTION MASS SPECTROMETRY (HRMS) IN DISCOVERY PROTEOMICS
 HIGH RESOLUTION MASS SPECTROMETRY (HRMS) IN DISCOVERY PROTEOMICS A clinical proteomics perspective Michael L. Merchant, PhD School of Medicine, University of Louisville Louisville, KY Learning Objectives
HIGH RESOLUTION MASS SPECTROMETRY (HRMS) IN DISCOVERY PROTEOMICS A clinical proteomics perspective Michael L. Merchant, PhD School of Medicine, University of Louisville Louisville, KY Learning Objectives
Supplementary material: Materials and suppliers
 Supplementary material: Materials and suppliers Electrophoresis consumables including tris-glycine, acrylamide, SDS buffer and Coomassie Brilliant Blue G-2 dye (CBB) were purchased from Ameresco (Solon,
Supplementary material: Materials and suppliers Electrophoresis consumables including tris-glycine, acrylamide, SDS buffer and Coomassie Brilliant Blue G-2 dye (CBB) were purchased from Ameresco (Solon,
In-Gel Tryptic Digestion Kit
 INSTRUCTIONS In-Gel Tryptic Digestion Kit 3747 N. Meridian Road P.O. Box 117 Rockford, IL 61105 89871 1468.2 Number Description 89871 In-Gel Tryptic Digestion Kit, sufficient reagents for approximately
INSTRUCTIONS In-Gel Tryptic Digestion Kit 3747 N. Meridian Road P.O. Box 117 Rockford, IL 61105 89871 1468.2 Number Description 89871 In-Gel Tryptic Digestion Kit, sufficient reagents for approximately
Trypsin Digestion Mix
 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name 239PR Trypsin Digestion Mix Provides optimal buffered conditions for in gel trypsin digestion
G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name 239PR Trypsin Digestion Mix Provides optimal buffered conditions for in gel trypsin digestion
Supporting Information
 Supporting Information Development of a High Coverage Pseudotargeted Lipidomics Method Based on Ultra-High Performance Liquid Chromatography-Mass Spectrometry Qiuhui Xuan 1,2#, Chunxiu Hu 1#, Di Yu 1,2,
Supporting Information Development of a High Coverage Pseudotargeted Lipidomics Method Based on Ultra-High Performance Liquid Chromatography-Mass Spectrometry Qiuhui Xuan 1,2#, Chunxiu Hu 1#, Di Yu 1,2,
Determination of β2-agonists in Pork Using Agilent SampliQ SCX Solid-Phase Extraction Cartridges and Liquid Chromatography-Tandem Mass Spectrometry
 Determination of β2-agonists in Pork Using Agilent SampliQ SCX Solid-Phase Extraction Cartridges and Liquid Chromatography-Tandem Mass Spectrometry Application Note Food Safety Authors Chenhao Zhai Agilent
Determination of β2-agonists in Pork Using Agilent SampliQ SCX Solid-Phase Extraction Cartridges and Liquid Chromatography-Tandem Mass Spectrometry Application Note Food Safety Authors Chenhao Zhai Agilent
AccuMAP Low ph Protein Digestion Kits
 TECHNICAL MANUAL AccuMAP Low ph Protein Digestion Kits Instruc ons for Use of Products VA1040 and VA1050 5/17 TM504 AccuMAP Low ph Protein Digestion Kits All technical literature is available at: www.promega.com/protocols/
TECHNICAL MANUAL AccuMAP Low ph Protein Digestion Kits Instruc ons for Use of Products VA1040 and VA1050 5/17 TM504 AccuMAP Low ph Protein Digestion Kits All technical literature is available at: www.promega.com/protocols/
Analysis of Amino Acids Derived Online Using an Agilent AdvanceBio AAA Column
 Application Note Pharmaceutical and Food Testing Analysis of Amino Acids Derived Online Using an Agilent AdvanceBio AAA Column Author Lu Yufei Agilent Technologies, Inc. Abstract A liquid chromatographic
Application Note Pharmaceutical and Food Testing Analysis of Amino Acids Derived Online Using an Agilent AdvanceBio AAA Column Author Lu Yufei Agilent Technologies, Inc. Abstract A liquid chromatographic
TECHNICAL BULLETIN. R 2 GlcNAcβ1 4GlcNAcβ1 Asn
 GlycoProfile II Enzymatic In-Solution N-Deglycosylation Kit Product Code PP0201 Storage Temperature 2 8 C TECHNICAL BULLETIN Product Description Glycosylation is one of the most common posttranslational
GlycoProfile II Enzymatic In-Solution N-Deglycosylation Kit Product Code PP0201 Storage Temperature 2 8 C TECHNICAL BULLETIN Product Description Glycosylation is one of the most common posttranslational
Bioanalytical Quantitation of Biotherapeutics Using Intact Protein vs. Proteolytic Peptides by LC-HR/AM on a Q Exactive MS
 Bioanalytical Quantitation of Biotherapeutics Using Intact Protein vs. Proteolytic Peptides by LC-HR/AM on a Q Exactive MS Jenny Chen, Hongxia Wang, Zhiqi Hao, Patrick Bennett, and Greg Kilby Thermo Fisher
Bioanalytical Quantitation of Biotherapeutics Using Intact Protein vs. Proteolytic Peptides by LC-HR/AM on a Q Exactive MS Jenny Chen, Hongxia Wang, Zhiqi Hao, Patrick Bennett, and Greg Kilby Thermo Fisher
Up to now, gel electrophoresis remains the most powerful
 pubs.acs.org/ac Polyacrylamide Gel with Switchable Trypsin Activity for Analysis of Proteins Fangjie Liu,, Mingliang Ye,*, Chunli Wang,, Zhengyan Hu,, Yi Zhang,, Hongqiang Qin,, Kai Cheng,, and Hanfa Zou*,
pubs.acs.org/ac Polyacrylamide Gel with Switchable Trypsin Activity for Analysis of Proteins Fangjie Liu,, Mingliang Ye,*, Chunli Wang,, Zhengyan Hu,, Yi Zhang,, Hongqiang Qin,, Kai Cheng,, and Hanfa Zou*,
Automating Mass Spectrometry-Based Quantitative Glycomics using Tandem Mass Tag (TMT) Reagents with SimGlycan
 PREMIER Biosoft Automating Mass Spectrometry-Based Quantitative Glycomics using Tandem Mass Tag (TMT) Reagents with SimGlycan Ne uaca2-3galb1-4glc NAcb1 6 Gal NAca -Thr 3 Ne uaca2-3galb1 Ningombam Sanjib
PREMIER Biosoft Automating Mass Spectrometry-Based Quantitative Glycomics using Tandem Mass Tag (TMT) Reagents with SimGlycan Ne uaca2-3galb1-4glc NAcb1 6 Gal NAca -Thr 3 Ne uaca2-3galb1 Ningombam Sanjib
ProteaseMAX Surfactant, Trypsin Enhancer
 Technical Bulletin ProteaseMAX Surfactant, Trypsin Enhancer INSTRUCTIONS FOR USE OF PRODUCTS V2071 AND V2072. PRINTED IN USA. Revised 1/10 ProteaseMAX Surfactant, Trypsin Enhancer All technical literature
Technical Bulletin ProteaseMAX Surfactant, Trypsin Enhancer INSTRUCTIONS FOR USE OF PRODUCTS V2071 AND V2072. PRINTED IN USA. Revised 1/10 ProteaseMAX Surfactant, Trypsin Enhancer All technical literature
Analysis of Testosterone, Androstenedione, and Dehydroepiandrosterone Sulfate in Serum for Clinical Research
 Analysis of Testosterone, Androstenedione, and Dehydroepiandrosterone Sulfate in Serum for Clinical Research Dominic Foley, Michelle Wills, and Lisa Calton Waters Corporation, Wilmslow, UK APPLICATION
Analysis of Testosterone, Androstenedione, and Dehydroepiandrosterone Sulfate in Serum for Clinical Research Dominic Foley, Michelle Wills, and Lisa Calton Waters Corporation, Wilmslow, UK APPLICATION
Figure S6. A-J) Annotated UVPD mass spectra for top ten peptides found among the peptides identified by Byonic but not SEQUEST + Percolator.
 Extending Proteome Coverage by Combining MS/MS Methods and a Modified Bioinformatics Platform adapted for Database Searching of Positive and Negative Polarity 193 nm Ultraviolet Photodissociation Mass
Extending Proteome Coverage by Combining MS/MS Methods and a Modified Bioinformatics Platform adapted for Database Searching of Positive and Negative Polarity 193 nm Ultraviolet Photodissociation Mass
LC-MS/MS Method for the Determination of Tenofovir from Plasma
 LC-MS/MS Method for the Determination of Tenofovir from Plasma Kimberly Phipps, Thermo Fisher Scientific, Runcorn, Cheshire, UK Application Note 687 Key Words SPE, SOLA CX, Hypersil GOLD, tenofovir Abstract
LC-MS/MS Method for the Determination of Tenofovir from Plasma Kimberly Phipps, Thermo Fisher Scientific, Runcorn, Cheshire, UK Application Note 687 Key Words SPE, SOLA CX, Hypersil GOLD, tenofovir Abstract
Mass Spectrometry. Mass spectrometer MALDI-TOF ESI/MS/MS. Basic components. Ionization source Mass analyzer Detector
 Mass Spectrometry MALDI-TOF ESI/MS/MS Mass spectrometer Basic components Ionization source Mass analyzer Detector 1 Principles of Mass Spectrometry Proteins are separated by mass to charge ratio (limit
Mass Spectrometry MALDI-TOF ESI/MS/MS Mass spectrometer Basic components Ionization source Mass analyzer Detector 1 Principles of Mass Spectrometry Proteins are separated by mass to charge ratio (limit
Nature Methods: doi: /nmeth Supplementary Figure 1
 Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
Biological Mass spectrometry in Protein Chemistry
 Biological Mass spectrometry in Protein Chemistry Tuula Nyman Institute of Biotechnology tuula.nyman@helsinki.fi MASS SPECTROMETRY is an analytical technique that identifies the chemical composition of
Biological Mass spectrometry in Protein Chemistry Tuula Nyman Institute of Biotechnology tuula.nyman@helsinki.fi MASS SPECTROMETRY is an analytical technique that identifies the chemical composition of
Development of a Human Cell-Free Expression System to Generate Stable-Isotope-Labeled Protein Standards for Quantitative Mass Spectrometry
 Development of a Human Cell-Free Expression System to Generate Stable-Isotope-Labeled Protein Standards for Quantitative Mass Spectrometry Ryan D. omgarden 1, Derek aerenwald 2, Eric Hommema 1, Scott Peterman
Development of a Human Cell-Free Expression System to Generate Stable-Isotope-Labeled Protein Standards for Quantitative Mass Spectrometry Ryan D. omgarden 1, Derek aerenwald 2, Eric Hommema 1, Scott Peterman
Section 1 Proteins and Proteomics
 Section 1 Proteins and Proteomics Learning Objectives At the end of this assignment, you should be able to: 1. Draw the chemical structure of an amino acid and small peptide. 2. Describe the difference
Section 1 Proteins and Proteomics Learning Objectives At the end of this assignment, you should be able to: 1. Draw the chemical structure of an amino acid and small peptide. 2. Describe the difference
Mass Spectrometry and Proteomics. Professor Xudong Yao Bioanalytical Chemistry Spring 2007
 Mass Spectrometry and Proteomics Professor Xudong Yao Bioanalytical Chemistry Spring 2007 Proteomics and -omics Roles of mass spectrometry Comparative proteomics Chemical proteomics Protein, Proteome and
Mass Spectrometry and Proteomics Professor Xudong Yao Bioanalytical Chemistry Spring 2007 Proteomics and -omics Roles of mass spectrometry Comparative proteomics Chemical proteomics Protein, Proteome and
INLIGHT Glycan Tagging Kit Protocol
 Cambridge Isotope Laboratories, Inc. isotope.com INLIGHT Glycan Tagging Kit Protocol A Glycan Tagging Kit for Comparative Quantification of N-linked Glycans INLIGHT Glycan Tagging Kit Catalog No. GTK-1000
Cambridge Isotope Laboratories, Inc. isotope.com INLIGHT Glycan Tagging Kit Protocol A Glycan Tagging Kit for Comparative Quantification of N-linked Glycans INLIGHT Glycan Tagging Kit Catalog No. GTK-1000
SPE-LC-MS/MS Method for the Determination of Nicotine, Cotinine, and Trans-3-hydroxycotinine in Urine
 SPE-LC-MS/MS Method for the Determination of Nicotine, Cotinine, and Trans-3-hydroxycotinine in Urine J. Jones, Thermo Fisher Scientific, Runcorn, Cheshire, UK Application Note 709 Key Words SPE, SOLA
SPE-LC-MS/MS Method for the Determination of Nicotine, Cotinine, and Trans-3-hydroxycotinine in Urine J. Jones, Thermo Fisher Scientific, Runcorn, Cheshire, UK Application Note 709 Key Words SPE, SOLA
Identification of Ginsenosides Using the SCIEX X500R QTOF System
 Identification of Ginsenosides Using the SCIEX X500R QTOF System Wang Sha, Cheng Haiyan, Liu Ting, Li Lijun, Jin Wenhai[Author] SCIEX, Pacific Applications Support Center (Beijing). China Background Ginseng
Identification of Ginsenosides Using the SCIEX X500R QTOF System Wang Sha, Cheng Haiyan, Liu Ting, Li Lijun, Jin Wenhai[Author] SCIEX, Pacific Applications Support Center (Beijing). China Background Ginseng
Screening and Speciation of Raw and Processed Meat Products
 vmethod Application for Food Testing Screening and Speciation of Raw and Processed Meat Products A Selective and Robust LC-MS/MS Method for Multiple Meat Speciation and Authentication on the QTRAP 4500
vmethod Application for Food Testing Screening and Speciation of Raw and Processed Meat Products A Selective and Robust LC-MS/MS Method for Multiple Meat Speciation and Authentication on the QTRAP 4500
Phosphotyrosine biased enrichment of tryptic peptides from cancer cells by combining py-mip and TiO 2 affinity resins
 Phosphotyrosine biased enrichment of tryptic peptides from cancer cells by combining py-mip and TiO 2 affinity resins Loreta Bllaci, Silje B. Torsetnes,, Celina Wierzbicka, Sudhirkumar Shinde, Börje Sellergren,
Phosphotyrosine biased enrichment of tryptic peptides from cancer cells by combining py-mip and TiO 2 affinity resins Loreta Bllaci, Silje B. Torsetnes,, Celina Wierzbicka, Sudhirkumar Shinde, Börje Sellergren,
Broad Spectrum Protease Inhibitor Cocktail
 Broad Spectrum Protease Inhibitor Cocktail Catalog number: AR1182 Boster s Broad Spectrum Protease Inhibitor Cocktail is a complex of various protease inhibitors, which has been tested for inhibiting proteases
Broad Spectrum Protease Inhibitor Cocktail Catalog number: AR1182 Boster s Broad Spectrum Protease Inhibitor Cocktail is a complex of various protease inhibitors, which has been tested for inhibiting proteases
Protein Precipitation for Biological Fluid Samples Using Agilent Captiva EMR Lipid 96-Well Plates
 Application Note Clinical Research Protein Precipitation for Biological Fluid Samples Using Agilent Captiva EMR Lipid 96-Well Plates Authors Limian Zhao and Megan Juck Agilent Technologies, Inc. Abstract
Application Note Clinical Research Protein Precipitation for Biological Fluid Samples Using Agilent Captiva EMR Lipid 96-Well Plates Authors Limian Zhao and Megan Juck Agilent Technologies, Inc. Abstract
2D-LC as an Automated Desalting Tool for MSD Analysis
 2D-LC as an Automated Desalting Tool for MSD Analysis Direct Mass Selective Detection of a Pharmaceutical Peptide from an MS-Incompatible USP Method Application Note Biologics and Biosimilars Author Sonja
2D-LC as an Automated Desalting Tool for MSD Analysis Direct Mass Selective Detection of a Pharmaceutical Peptide from an MS-Incompatible USP Method Application Note Biologics and Biosimilars Author Sonja
Babu Antharavally, Ryan Bomgarden, and John Rogers Thermo Fisher Scientific, Rockford, IL
 A Versatile High-Recovery Method for Removing Detergents from Low-Concentration Protein or Peptide Samples for Mass Spectrometry Sample Preparation and Analysis Babu Antharavally, Ryan Bomgarden, and John
A Versatile High-Recovery Method for Removing Detergents from Low-Concentration Protein or Peptide Samples for Mass Spectrometry Sample Preparation and Analysis Babu Antharavally, Ryan Bomgarden, and John
Supplementary Information. Effects of Perfluorooctanoic Acid on Metabolic Profiles in Brain and Liver of Mouse by a
 Supplementary Information Effects of Perfluorooctanoic Acid on Metabolic Profiles in Brain and Liver of Mouse by a High-throughput Targeted Metabolomics Approach Nanyang Yu, Si Wei, *, Meiying Li, Jingping
Supplementary Information Effects of Perfluorooctanoic Acid on Metabolic Profiles in Brain and Liver of Mouse by a High-throughput Targeted Metabolomics Approach Nanyang Yu, Si Wei, *, Meiying Li, Jingping
Fig.S1 ESI-MS spectrum of reaction of ApA and THPTb after 16 h.
 Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2014 Experiment Cleavage of dinucleotides Dinucleotides (ApA, CpC, GpG, UpU) were purchased from
Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2014 Experiment Cleavage of dinucleotides Dinucleotides (ApA, CpC, GpG, UpU) were purchased from
LC/MS Method for Comprehensive Analysis of Plasma Lipids
 Application Note omics LC/MS Method for Comprehensive Analysis of Plasma s Authors Tomas Cajka and Oliver Fiehn West Coast Metabolomics Center, University of California Davis, 451 Health Sciences Drive,
Application Note omics LC/MS Method for Comprehensive Analysis of Plasma s Authors Tomas Cajka and Oliver Fiehn West Coast Metabolomics Center, University of California Davis, 451 Health Sciences Drive,
AMINO ACIDS STRUCTURE, CLASSIFICATION, PROPERTIES. PRIMARY STRUCTURE OF PROTEINS
 AMINO ACIDS STRUCTURE, CLASSIFICATION, PROPERTIES. PRIMARY STRUCTURE OF PROTEINS Elena Rivneac PhD, Associate Professor Department of Biochemistry and Clinical Biochemistry State University of Medicine
AMINO ACIDS STRUCTURE, CLASSIFICATION, PROPERTIES. PRIMARY STRUCTURE OF PROTEINS Elena Rivneac PhD, Associate Professor Department of Biochemistry and Clinical Biochemistry State University of Medicine
For reprints and correspondence:
 APPLICATION REPORT SEPTEMBER 30, 2015 New Proteomic Workflows Combine Albumin Depletion and On- Bead Digestion, for Quantitative Cancer Serum Haiyan Zheng 1 ; Caifeng Zhao 1 ; Meiqian Qian 1 ; Swapan Roy
APPLICATION REPORT SEPTEMBER 30, 2015 New Proteomic Workflows Combine Albumin Depletion and On- Bead Digestion, for Quantitative Cancer Serum Haiyan Zheng 1 ; Caifeng Zhao 1 ; Meiqian Qian 1 ; Swapan Roy
Faster Mass Spec: Same-Day Sample Prep Now a Reality. Michael M. Rosenblatt, Ph.D.
 Faster Mass Spec: Same-Day Sample Prep Now a Reality Michael M. Rosenblatt, Ph.D. Applications of Mass Spec in Biology Biomarker Discovery Protein Interactions Protein Expression Drug Discovery Subcellular
Faster Mass Spec: Same-Day Sample Prep Now a Reality Michael M. Rosenblatt, Ph.D. Applications of Mass Spec in Biology Biomarker Discovery Protein Interactions Protein Expression Drug Discovery Subcellular
SUPPLEMENTARY MATERIAL
 SUPPLEMENTARY MATERIAL Purification and biochemical properties of SDS-stable low molecular weight alkaline serine protease from Citrullus Colocynthis Muhammad Bashir Khan, 1,3 Hidayatullah khan, 2 Muhammad
SUPPLEMENTARY MATERIAL Purification and biochemical properties of SDS-stable low molecular weight alkaline serine protease from Citrullus Colocynthis Muhammad Bashir Khan, 1,3 Hidayatullah khan, 2 Muhammad
Robust extraction, separation, and quantitation of structural isomer steroids from human plasma by SPE-UHPLC-MS/MS
 TECHNICAL NOTE 21882 Robust extraction, separation, and quantitation of structural isomer steroids human plasma by SPE-UHPLC-MS/MS Authors Jon Bardsley 1, Kean Woodmansey 1, and Stacy Tremintin 2 1 Thermo
TECHNICAL NOTE 21882 Robust extraction, separation, and quantitation of structural isomer steroids human plasma by SPE-UHPLC-MS/MS Authors Jon Bardsley 1, Kean Woodmansey 1, and Stacy Tremintin 2 1 Thermo
Quantitative Analysis of Vit D Metabolites in Human Plasma using Exactive System
 Quantitative Analysis of Vit D Metabolites in Human Plasma using Exactive System Marta Kozak Clinical Research Applications Group Thermo Fisher Scientific San Jose CA Clinical Research use only, Not for
Quantitative Analysis of Vit D Metabolites in Human Plasma using Exactive System Marta Kozak Clinical Research Applications Group Thermo Fisher Scientific San Jose CA Clinical Research use only, Not for
Determination of 6-Chloropicolinic Acid (6-CPA) in Crops by Liquid Chromatography with Tandem Mass Spectrometry Detection. EPL-BAS Method No.
 Page 1 of 10 Determination of 6-Chloropicolinic Acid (6-CPA) in Crops by Liquid Chromatography with Tandem Mass Spectrometry Detection EPL-BAS Method No. 205G881B Method Summary: Residues of 6-CPA are
Page 1 of 10 Determination of 6-Chloropicolinic Acid (6-CPA) in Crops by Liquid Chromatography with Tandem Mass Spectrometry Detection EPL-BAS Method No. 205G881B Method Summary: Residues of 6-CPA are
A Robustness Study for the Agilent 6470 LC-MS/MS Mass Spectrometer
 A Robustness Study for the Agilent 7 LC-MS/MS Mass Spectrometer Application Note Clinical Research Authors Linda Côté, Siji Joseph, Sreelakshmy Menon, and Kevin McCann Agilent Technologies, Inc. Abstract
A Robustness Study for the Agilent 7 LC-MS/MS Mass Spectrometer Application Note Clinical Research Authors Linda Côté, Siji Joseph, Sreelakshmy Menon, and Kevin McCann Agilent Technologies, Inc. Abstract
Introduction to Peptide Sequencing
 Introduction to Peptide equencing Quadrupole Ion Traps tructural Biophysics Course December 3, 2014 12/8/14 Introduction to Peptide equencing - athan Yates 1 Why are ion traps used to sequence peptides?
Introduction to Peptide equencing Quadrupole Ion Traps tructural Biophysics Course December 3, 2014 12/8/14 Introduction to Peptide equencing - athan Yates 1 Why are ion traps used to sequence peptides?
Work-flow: protein sample preparation Precipitation methods Removal of interfering substances Specific examples:
 Dr. Sanjeeva Srivastava IIT Bombay Work-flow: protein sample preparation Precipitation methods Removal of interfering substances Specific examples: Sample preparation for serum proteome analysis Sample
Dr. Sanjeeva Srivastava IIT Bombay Work-flow: protein sample preparation Precipitation methods Removal of interfering substances Specific examples: Sample preparation for serum proteome analysis Sample
Relative Quantitation of Human Polymorphonuclear Leukocyte Cell Membrane GPEtn Lipids
 Relative Quantitation of Human Polymorphonuclear Leukocyte Cell Membrane GPEtn Lipids Using the QTRAP System with mtraq Reagents Karin A. Zemski-Berry 1, John M. Hevko 2, and Robert C. Murphy 1 1 Department
Relative Quantitation of Human Polymorphonuclear Leukocyte Cell Membrane GPEtn Lipids Using the QTRAP System with mtraq Reagents Karin A. Zemski-Berry 1, John M. Hevko 2, and Robert C. Murphy 1 1 Department
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/8/398/rs12/dc1 Supplementary Materials for Quantitative phosphoproteomics reveals new roles for the protein phosphatase PP6 in mitotic cells Scott F. Rusin, Kate
www.sciencesignaling.org/cgi/content/full/8/398/rs12/dc1 Supplementary Materials for Quantitative phosphoproteomics reveals new roles for the protein phosphatase PP6 in mitotic cells Scott F. Rusin, Kate
Don t miss a thing on your peptide mapping journey How to get full coverage peptide maps using high resolution accurate mass spectrometry
 Don t miss a thing on your peptide mapping journey How to get full coverage peptide maps using high resolution accurate mass spectrometry Kai Scheffler, PhD BioPharma Support Expert,LSMS Europe The world
Don t miss a thing on your peptide mapping journey How to get full coverage peptide maps using high resolution accurate mass spectrometry Kai Scheffler, PhD BioPharma Support Expert,LSMS Europe The world
Development of a Bioanalytical Method for Quantification of Amyloid Beta Peptides in Cerebrospinal Fluid
 Development of a Bioanalytical Method for Quantification of Amyloid Beta Peptides in Cerebrospinal Fluid Joanne ( 乔安妮 ) Mather Senior Scientist Waters Corporation Data courtesy of Erin Chambers and Mary
Development of a Bioanalytical Method for Quantification of Amyloid Beta Peptides in Cerebrospinal Fluid Joanne ( 乔安妮 ) Mather Senior Scientist Waters Corporation Data courtesy of Erin Chambers and Mary
Supporting information
 Supporting information Figure legends Supplementary Table 1. Specific product ions obtained from fragmentation of lithium adducts in the positive ion mode comparing the different positional isomers of
Supporting information Figure legends Supplementary Table 1. Specific product ions obtained from fragmentation of lithium adducts in the positive ion mode comparing the different positional isomers of
Unsupervised Identification of Isotope-Labeled Peptides
 Unsupervised Identification of Isotope-Labeled Peptides Joshua E Goldford 13 and Igor GL Libourel 124 1 Biotechnology institute, University of Minnesota, Saint Paul, MN 55108 2 Department of Plant Biology,
Unsupervised Identification of Isotope-Labeled Peptides Joshua E Goldford 13 and Igor GL Libourel 124 1 Biotechnology institute, University of Minnesota, Saint Paul, MN 55108 2 Department of Plant Biology,
Symposium on Protein Fractionation
 Symposium on Protein Fractionation Proteomic Analysis of Organ-Specific Breast Cancer Metastasis Speaker: Emily I. Chen, Ph.D The Scripps Research Institute Department of Cell Biology CANCER METASTASIS
Symposium on Protein Fractionation Proteomic Analysis of Organ-Specific Breast Cancer Metastasis Speaker: Emily I. Chen, Ph.D The Scripps Research Institute Department of Cell Biology CANCER METASTASIS
MALDI-TOF. Introduction. Schematic and Theory of MALDI
 MALDI-TOF Proteins and peptides have been characterized by high pressure liquid chromatography (HPLC) or SDS PAGE by generating peptide maps. These peptide maps have been used as fingerprints of protein
MALDI-TOF Proteins and peptides have been characterized by high pressure liquid chromatography (HPLC) or SDS PAGE by generating peptide maps. These peptide maps have been used as fingerprints of protein
Amino acids-incorporated nanoflowers with an
 Amino acids-incorporated nanoflowers with an intrinsic peroxidase-like activity Zhuo-Fu Wu 1,2,+, Zhi Wang 1,+, Ye Zhang 3, Ya-Li Ma 3, Cheng-Yan He 4, Heng Li 1, Lei Chen 1, Qi-Sheng Huo 3, Lei Wang 1,*
Amino acids-incorporated nanoflowers with an intrinsic peroxidase-like activity Zhuo-Fu Wu 1,2,+, Zhi Wang 1,+, Ye Zhang 3, Ya-Li Ma 3, Cheng-Yan He 4, Heng Li 1, Lei Chen 1, Qi-Sheng Huo 3, Lei Wang 1,*
Proteomics Grade. Protocol. Catalog # Agilent Technologies. Research Use Only. Not for use in Diagnostic Procedures. Version A, January 2010
 Proteomics Grade Trypsin Catalog #204310 Protocol Version A, January 2010 Research Use Only. Not for use in Diagnostic Procedures. Agilent Technologies Notices Agilent Technologies, Inc. 2010 No part of
Proteomics Grade Trypsin Catalog #204310 Protocol Version A, January 2010 Research Use Only. Not for use in Diagnostic Procedures. Agilent Technologies Notices Agilent Technologies, Inc. 2010 No part of
Proteomics of body liquids as a source for potential methods for medical diagnostics Prof. Dr. Evgeny Nikolaev
 Proteomics of body liquids as a source for potential methods for medical diagnostics Prof. Dr. Evgeny Nikolaev Institute for Biochemical Physics, Rus. Acad. Sci., Moscow, Russia. Institute for Energy Problems
Proteomics of body liquids as a source for potential methods for medical diagnostics Prof. Dr. Evgeny Nikolaev Institute for Biochemical Physics, Rus. Acad. Sci., Moscow, Russia. Institute for Energy Problems
Mass Spectrometry based metabolomics
 Mass Spectrometry based metabolomics Metabolomics- A realm of small molecules (
Mass Spectrometry based metabolomics Metabolomics- A realm of small molecules (
Kit for assay of thioredoxin
 FkTRX-02-V2 Kit for assay of thioredoxin The thioredoxin system is the major protein disulfide reductase in cells and comprises thioredoxin, thioredoxin reductase and NADPH (1). Thioredoxin systems are
FkTRX-02-V2 Kit for assay of thioredoxin The thioredoxin system is the major protein disulfide reductase in cells and comprises thioredoxin, thioredoxin reductase and NADPH (1). Thioredoxin systems are
Jose Castro-Perez, Henry Shion, Kate Yu, John Shockcor, Emma Marsden-Edwards, Jeff Goshawk Waters Corporation, Milford, MA, U.S. and Manchester, UK
 HIGH-THRUGHPUT REACTIVE METABLITE SCREEIG FR DICLFEAC BY UPLC AD XEV TQ MS WITH SCAWAVE Jose Castro-Perez, Henry Shion, Kate Yu, John Shockcor, Emma Marsden-Edwards, Jeff Goshawk Waters Corporation, Milford,
HIGH-THRUGHPUT REACTIVE METABLITE SCREEIG FR DICLFEAC BY UPLC AD XEV TQ MS WITH SCAWAVE Jose Castro-Perez, Henry Shion, Kate Yu, John Shockcor, Emma Marsden-Edwards, Jeff Goshawk Waters Corporation, Milford,
Ultra Performance Liquid Chromatography Coupled to Orthogonal Quadrupole TOF MS(MS) for Metabolite Identification
 22 SEPARATION SCIENCE REDEFINED MAY 2005 Ultra Performance Liquid Chromatography Coupled to Orthogonal Quadrupole TOF MS(MS) for Metabolite Identification In the drug discovery process the detection and
22 SEPARATION SCIENCE REDEFINED MAY 2005 Ultra Performance Liquid Chromatography Coupled to Orthogonal Quadrupole TOF MS(MS) for Metabolite Identification In the drug discovery process the detection and
(III) MALDI instrumentation
 Dr. Sanjeeva Srivastava (I) Basics of MALDI-TF (II) Sample preparation In-gel digestion Zip-tip sample clean-up Matrix and sample plating (III) MALDI instrumentation 2 1 (I) Basics of MALDI-TF Analyte
Dr. Sanjeeva Srivastava (I) Basics of MALDI-TF (II) Sample preparation In-gel digestion Zip-tip sample clean-up Matrix and sample plating (III) MALDI instrumentation 2 1 (I) Basics of MALDI-TF Analyte
Supporting Information
 Notes Bull. Korean Chem. Soc. 2013, Vol. 34, No. 1 1 http://dx.doi.org/10.5012/bkcs.2013.34.1.xxx Supporting Information Chemical Constituents of Ficus drupacea Leaves and their α-glucosidase Inhibitory
Notes Bull. Korean Chem. Soc. 2013, Vol. 34, No. 1 1 http://dx.doi.org/10.5012/bkcs.2013.34.1.xxx Supporting Information Chemical Constituents of Ficus drupacea Leaves and their α-glucosidase Inhibitory
Supplementary Material (ESI) for Chemical Communications This journal is (c) The Royal Society of Chemistry 2008
 Experimental Details Unless otherwise noted, all chemicals were purchased from Sigma-Aldrich Chemical Company and were used as received. 2-DOS and neamine were kindly provided by Dr. F. Huang. Paromamine
Experimental Details Unless otherwise noted, all chemicals were purchased from Sigma-Aldrich Chemical Company and were used as received. 2-DOS and neamine were kindly provided by Dr. F. Huang. Paromamine
Core E Analysis of Neutral Lipids from Human Plasma June 4, 2010 Thomas J. Leiker and Robert M. Barkley
 Core E Analysis of Neutral Lipids from Human Plasma June 4, 2010 Thomas J. Leiker and Robert M. Barkley This protocol describes the extraction and direct measurement of cholesterol esters (CEs) and triacylglycerols
Core E Analysis of Neutral Lipids from Human Plasma June 4, 2010 Thomas J. Leiker and Robert M. Barkley This protocol describes the extraction and direct measurement of cholesterol esters (CEs) and triacylglycerols
LECTURE-15. itraq Clinical Applications HANDOUT. Isobaric Tagging for Relative and Absolute quantitation (itraq) is a quantitative MS
 LECTURE-15 itraq Clinical Applications HANDOUT PREAMBLE Isobaric Tagging for Relative and Absolute quantitation (itraq) is a quantitative MS based method for quantifying proteins subject to various different
LECTURE-15 itraq Clinical Applications HANDOUT PREAMBLE Isobaric Tagging for Relative and Absolute quantitation (itraq) is a quantitative MS based method for quantifying proteins subject to various different
Protein and peptide separations,
 A Quarterly Technical Newsletter Spring Grace Vydac: Leading the Way in Separations for Proteomics Protein and peptide separations, always important to protein chemists and enzymologists, have recently
A Quarterly Technical Newsletter Spring Grace Vydac: Leading the Way in Separations for Proteomics Protein and peptide separations, always important to protein chemists and enzymologists, have recently
A Definitive Lipidomics Workflow for Human Plasma Utilizing Off-line Enrichment and Class Specific Separation of Phospholipids
 A Definitive Lipidomics Workflow for Human Plasma Utilizing Off-line Enrichment and Class Specific Separation of Phospholipids Jeremy Netto, 1 Stephen Wong, 1 Federico Torta, 2 Pradeep Narayanaswamy, 2
A Definitive Lipidomics Workflow for Human Plasma Utilizing Off-line Enrichment and Class Specific Separation of Phospholipids Jeremy Netto, 1 Stephen Wong, 1 Federico Torta, 2 Pradeep Narayanaswamy, 2
