Supplementary Figures and Notes for Digestion and depletion of abundant
|
|
|
- Randall Hawkins
- 5 years ago
- Views:
Transcription
1 Supplementary Figures and Notes for Digestion and depletion of abundant proteins improves proteomic coverage Bryan R. Fonslow, Benjamin D. Stein, Kristofor J. Webb, Tao Xu, Jeong Choi, Sung Kyu Park, and John R. Yates III Supplementary Figure 1 Supplementary Figure 2 Supplementary Figure 3 Supplementary Figure 4 Supplementary Figure 5 Supplementary Figure 6 Supplementary Figure 7 Supplementary Figure 8 Supplementary Note 1 Supplementary Note 2 Supplementary Note 3 Supplementary Note 4 Supplementary Note 5 Supplementary Note 6 Log-log correlation plot of MW-corrected normalized spectral abundance factor (NSAF) for yeast proteins to their absolute cell copy number Analysis of digestion and depletion of proteins from a tryptic yeast lysate Analysis of digestion and depletion of tryptic peptides from a yeast lysate Comparison of proteins and peptides from triplicate Control and DigDeAPr runs of tryptic HEK lysates Statistical comparison of peptide quality scores from HEK cell lysates Peptide precursor intensity histogram comparison from HEK cell lysates Surface plots of HEK protein physicochemical properties based on relative spectral count changes Statistical comparison of proteins between two separate triplicate Control analyses of tryptic HEK lysates Derivation of Michaelis-Menten equation to describe protein digestion in a complex mixture Estimation and evaluation of initial DigDeAPr conditions Validation of DigDeAPr with known yeast protein copy numbers Improvements to peptide quantitation metrics with DigDeAPr Comparison of dynamic range compression mechanisms between DigDeAPr and hexapeptide-based ProteoMiner Analysis of potential protein losses from the MWCO spin-filter
2 Supplementary Figure 1: Log-log correlation plot of molecular weight-corrected normalized spectral abundance factor (NSAF) for yeast proteins to their absolute cell copy number. The data was fit using a nonlinear regression. The best fit line and 95% confidence interval are plotted.
3 Supplementary Figure 2: Analysis of digestion and depletion of proteins from a tryptic Yeast lysate. (a) Volcano plot of the log 2 average protein spectral count ratio between triplicate DigDeAPr and Control runs plotted against the p-value. Data points are plotted based on average spectral counts from triplicate Control runs: 1-9 spectral counts (black), (green), (magenta), and more than 1000 spectral counts (yellow with black outline). (b) Correlation of spectral count depletion to absolute protein abundance in Yeast. The data was fit using a linear regression. The fit line and 95% confidence interval are plotted. Box whisker plots of spectral count depletion for (c) all identified Yeast proteins and (d) low abundance Yeast proteins.
4 Supplementary Figure 3: Analysis of digestion and depletion of peptides from a yeast lysate. (a) Volcano plot of the log 2 average peptide spectral count ratio between triplicate DigDeAPr and Control runs plotted against the p-value. Data points are plotted based on average spectral counts from triplicate Control runs: 1-9 spectral counts (black), (green), and (magenta) spectral counts. (b) Correlation of peptide spectral count changes to absolute protein abundance in yeast. The data was fit using a linear regression. The best fit line and 95% confidence interval are plotted. Box whisker plots of peptide spectral count changes for (c) all identified yeast peptides and (d) low abundance yeast peptides plotted binned according to their protein abundance.
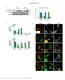
5 Supplementary Figure 4: Comparison of proteins and peptides from triplicate Control and DigDeAPr runs of HEK lysates. Venn diagrams for total (a) proteins and (b) peptides among triplicate Control and DigDeAPr runs. Venn diagrams for proteins among triplicate (c) Control and (d) DigDeAPr runs. Venn diagrams for peptides among triplicate (e) Control and (f) DigDeAPr runs.
6 Supplementary Figure 5: Statistical comparison of peptide quality scores from HEK cell lysates. Volcano plots of the log 2 ratio of the average (a) peptide XCorr and (b) peptide CN between triplicate DigDeAPr and Control runs plotted against the p-value. Data points are plotted based on average spectral counts from triplicate Control runs: 1-9 spectral counts (black), 1-99 spectral counts (green), and more than 100 spectral counts (magenta). XCorr is a measure of the spectral matching quality between theoretical and experimental spectra. CN is the difference between XCorr values of the 1 st and 2 nd candidate peptide sequences and is an indicator of peptide spectrum match correctness.
7 Supplementary Figure 6: HEK cell lysate peptide precursor intensity histogram comparison for triplicate Control (red) and DigDeAPr (blue) runs with error bars representing standard deviation. A systematic increase in peptide precursor intensity was found for all peptides identified in DigDeAPr runs relative to Control runs.
8 Supplementary Figure 7: Surface plots of HEK protein physicochemical properties based on relative spectral count abundance changes. Histograms for the relative frequency of protein (a) isoelectric point, (b) Kyte-Doolittle Score, (c) number of tryptic peptides, and (d) molecular weight were determined for each relative spectral count change bin. Only proteins with greater than 5 spectral counts in either Control or DigDeAPr runs were considered.
9 Supplementary Figure 8. Statistical comparison of proteins between two separate triplicate Control analyses of tryptic HEK lysates. Volcano plots of the log 2 ratio of the average from replicate triplicate Control runs for (a) protein spectral counts and (b) protein sequence coverage plotted against the p-value. Data points are plotted based on average spectral counts from triplicate Control runs: 1-9 spectral counts (black), (green), and more than 100 (magenta).
10 Supplementary Note 1. Derivation of Michaelis-Menten equation to describe abundance-based proteome digestion. The selective digestion of high abundance proteins and not low abundance proteins from the proteome requires specific conditions. Qualitatively, we knew that these conditions would be both diffusion-limited and trypsin-limited to ensure that trypsin would digest proteins at a controllable rate, based on the rate at which it forms a complex with abundant proteins. Since the complex formation rate is in part defined by K M, we also knew the protein concentration relative to K M would be an important parameter to consider. To quantitatively estimate the appropriate conditions for selective digestion of high abundance proteins we first derived an equation from the Michaelis-Menten equation to describe the digestion of low abundance proteins in a proteome from a complex cell lysate. For any enzyme reaction dependent on formation of a complex between the enzyme (E) and substrate (S) to form products (P) (1) the velocity ( ) or rate of product formation (d[p]/dt) can be defined by the rate constant k 2 and concentration of the Michaelis enzyme:substrate complex ([ES]), assuming the complex has reached steady-state equilibrium (d[es]/dt = 0): (2)
11 The enzyme:substrate complex is commonly defined by the total enzyme concentration ([E T ]), substrate concentration ([S]), and Michaelis constant (K M ) as: (3) Inserting equation 3 into equation 2 and defining the maximum reaction velocity ( max ) as [E T ]k 2 yields the Michaelis-Mention equation: (4) These derivations are well known. This form of the equation can be easily used to define the rate of digestion of one protein substrate at different concentrations of substrates relative to K M. In this equation, the relative relationship between the substrate concentration and the Michaelis constant are important. Most simply, when K M and the substrate concentration ([S]) are of the same order, equation 4 defines the velocity of the reaction ( ). When a reaction is performed where [S] is much greater than K M, such as with highly abundant proteins, the K M drops out of the equation and the [S] s cancel to yield a velocity that is the maximum velocity: (5) Conversely, when the substrate concentration is less than K M, such as when a protein is of low abundance in a lysate, the [S] in the denominator drops out and the velocity is equal to maximum velocity times the ratio of the substrate concentration and the Michaelis constant: (6)
12 As stated, during a protease digestion highly abundant proteins will be digested at a rate defined by equation 5 or 6, depending on whether they are above or below the K M. Note that under typical digestion condition for shotgun proteomics, where the protein:protease ratio is ~100:1, a majority of the proteins will be digested at V max. Unlike high abundance proteins, low abundance proteins will not be digested at a rate defined by equation 5 or 6. That is, low abundance protein digestion velocities should be defined by both their concentrations and the total protein concentration of the lysate. Thus, we derived an equation to describe the rate of digestion of individual low abundance proteins within a lysate when their concentrations are below the proteases K M. The K M of sequence grade trypsin is ~3 M, 1 well above the concentration of low abundance proteins (< 1 nm) in a common protein digestion concentration of 1 mg/ml (~10 M). We started with the equation generated from inserting equation 3 into equation 2: (5) Under the conditions described, the total enzyme concentration ([E T ]) is no longer available for formation of a complex with low abundance proteins since it is in a complex with high abundance proteins. Thus we can substitute [E T ] for the free protease concentration ([E]) and the individual low abundance protein concentration ([P i ]) for the substrate concentration ([S]). Further, since the protein concentration is much lower than K M, the [P i ] in the denominator drops out of the equation, yielding: (6)
13 The k 2 and K M can be separated out and the total enzyme concentration ([E T ]) can be substituted by the difference of the total enzyme concentration and the protease:protein complex concentration ([E T ] [EP T ]). Note the total protein (P T ) denotation to illustrate the complex concentration is defined by the total protein concentration and not the individual low abundance protein concentration (P i ). (7) The protease:protein complex concentration can be replaced by equation 3 yielding: (8) A common denominator is created for [E T ] by multiplying the numerator and denominator by K M + [P T ]: (9) The numerator is further expanded: (10) and subtracted using the common denominator: (11) The two [E T ][P T ] terms subtract out leaving : (12)
14 The K M s cancel leaving: (13) The numerator is the definition of V max, simplying the equation to: (14) This equation takes the same form of the general Michaelis-Menten equation. However, the substrate concentration for an individual protein ([P i ]) and the total protein substrate concentration ([P T ]) now together define the velocity of digestion for the individual protein (P i ). Further, the equation clearly defines that if the total protein concentration ([P T ]) is on the same order as K M, that the individual protein digestion velocity is a function of the sum of K M and the total protein concentration ([P T ]). Under conditions where the total protein concentration ([P T ]) is greater than K M and the individual protein concentrations [P i ] are still less than K M, then the velocity for digestion of individual low abundance proteins is: (15) Through inspection of equations 14 and 15 it becomes apparent that in order to achieve selective increases to the digestion velocity of abundant proteins over low abundant proteins, the total protein concentration ([P T ]) should not be less than K M. That is, the digestion velocity of abundant proteins will approach V max and be proportional to the mole percentage of the protein within the proteome. From equation 14, the digestion velocity for low abundance proteins will be proportional to their concentration ([P i ]) and
15 inversely proportional to the sum of the K M and the total protein concentration ([P T ]). Further, the extent to which [P T ] is greater than K M will define the number of proteins within a proteome that have a concentration also equal to or greater than K M. Equations 14 and 15 illustrate that the adequate conditions for digestion depletion are dependent on five parameters. Two of the parameters are biologically defined constants the K M of the protease and the protein abundance dynamic range of a proteome ([P T ] [P i ]), assumed to be greater than 10 6 for the HEK lysate. 2, 3 The other three parameters are biochemical variables - the protein and enzyme concentrations and the time of digestion. The next section (Supplementary Note 2) describes the estimation and evaluation of the three biochemical variables.
16 Supplementary Note 2. Estimation and evaluation of initial DigDeAPr conditions. As with most analytical work, we estimated that an order-of-magnitude adjustment in protein dynamic range was necessary to see a noticeable improvement in proteomic metrics. Thus we wanted to digest ~90% of the proteome away. This defined a reasonable starting protein mass of 1 mg since 100 g of protein lysate is regularly used for protein analysis with MudPIT. Our protein lysates were around 10 mg/ml, thus our digestions could be performed at a concentration of 10 mg/ml or lower. Somewhat arbitrarily assuming an average molecular weight of proteins in a lysate to be 67 kda, this would yield protein concentrations less than 150 M. Assuming we want high abundance proteins to be above the K M of trypsin and low abundance proteins to be below it described in the previous section, we chose 1 mg/ml, corresponding to 15 M (five times greater than the 3 M K M of trypsin). 1 Next we estimated a trypsin concentration based on a reasonable digestion time (8 hrs). In order to digest 90% of proteins at a concentration of 15 M in 8 hours, a V max of 500 pm s -1 would be required. Thus, an adequate trypsin concentration can be calculated by dividing the V max by the k cat of 0.44 s -1, 1 yielding a trypsin concentration of 1 nm or 25 ng/ml. These were the initial conditions we tested which also ultimately gave some of the best results. However, we also tested longer digestion times (12, 18, and 24 hr), different trypsin concentrations (2.5 ng/ml, 10 ng/ml, and 100 ng/ml), and a 10-fold lower 1 mg HEK protein lysate concentration (100 g/ml). Since the mass spectrometry-based proteomic analysis of all the initial digestion depletion optimization conditions would have been excessively time consuming, we
17 focused first on the mass balance of proteins during the course of digestion depletion (Online Methods). We would suggest using a similar mass balance strategy for implementing DigDeAPr on other sample types where the protein abundance dynamic range may be different. Time point analysis of the digestion depletion progress at fixed trypsin and lysate concentrations was the most informative optimization experiment we performed. It provided a confirmation of the kinetics of the digestion depletion and an adequate timeframe for which other concentration modifications could be performed. With varied digestion depletion time the progress of the depletion was established as either incomplete (< 75%), complete (75 95%), or over-depleted (> 95%), based on protein mass depletion measured by BCA mass balance. Appropriate concentration and/or digestion time changes were made. For instance, if the sample was overdepleted the trypsin concentration or digestion depletion time were reduced 50%. Incomplete depletion could be increased by doubling either of these parameters. The less straightforward, and potentially most important, optimization condition was selection of protein lysate concentration. An optimum lysate concentration is likely to be different even for human cell lines. For instance, from our previous analysis of cancer-derived HeLa cells with hexapeptide bead protein abundance adjustment we found 7 proteins with greater than 1000 spectral counts and another 8 proteins with greater than 500 spectral counts. 4 While with neuron-like HEK cells in this study, we only found one protein with close to 1000 spectral counts and 5 proteins around 500 spectral counts (which were the same as the HeLa proteins with greater than 1000 spectral counts). The protein lysate concentration defines which portion of the proteome is above (depleted), equal to (partially depleted), or below (non-depleted) the
18 K M of trypsin. Thus, in order to optimize the protein lysate concentration, proteomic analysis was performed and would be suggested for adopters of this method since it provides protein-specific abundance changes not evident with the mass balance approach. After proteomic analysis if all proteins appear over-depleted (including low abundance ones), decrease the lysate concentration. This shifts more individual protein concentrations below the K M of trypsin, focusing the depletion on the highest abundance proteins. If the sample is under-depleted (i.e. too few high abundance proteins are depleted), the lysate concentration can be increased. Conversely, this concentration change will shift more individual protein concentrations above the K M of trypsin, expanding the depletion range of abundant proteins. These concentration changes may also affect the rate of digestion depletion, so the appropriate changes to either protein mass, trypsin concentration, or digestion depletion time may be necessary. Fortunately, these adjustments can be monitored by mass balance without more time consuming mass spectrometry experiments.
19 Supplementary Note 3. Validation of digestion and depletion of abundant proteins with known yeast protein copy numbers To directly correlate protein spectral count changes to protein abundance changes we also performed triplicate Control and DigDeAPr runs on a yeast lysate. The analysis of log-phase yeast allows for direct comparison of spectral counts to absolute protein copy numbers per cell (Supplementary Fig. 1) determined by global western blot analysis. 5 In this case, we use protein copy numbers to correlate spectral count changes to protein abundance changes. We found similar changes in yeast protein abundance (Supplementary Fig. 2a) as HEK proteins (Fig. 2a) based on spectral counting using the same digestion depletion conditions for the yeast lysate as the HEK cell lysate. Further, similar trends were also observed at the peptide level for yeast proteins (Supplementary Fig. 3a) as with the HEK peptides (Fig. 2e). This highlights the robustness of DigDeAPr since mammalian and yeast cell lysates have significantly different numbers of proteins (~6,750 versus >10,000) and protein abundance profiles (10 6 versus 10 7 ). 2, 3, 5 Nevertheless, improvements in protein identifications and sequence coverages from DigDeAPr were more dramatic due to the higher complexity and dynamic range of the HEK proteome. For the yeast lysate data, when spectral count changes of yeast proteins are plotted versus their absolute abundance it is evident that the higher abundance proteins and peptides are selectively depleted as expected (Supplementary Figs. 2b and 3b). These results are further highlighted when the data is represented in box-whisker plots for all (Supplementary Figs. 2c and 3c) and lower abundance (Supplementary Figs. 2d and 3d) proteins and peptides.
20 Supplementary Note 4. Improvements to peptide quantitation metrics with DigDeAPr A comparison of the protein spectral count relative standard deviations (RSDs) between Control and DigDeAPr runs (Fig. 2f) can be used to assess the capabilities of DigDeAPr for spectral counting-based quantitation. High abundance protein spectral counting quantitation can be easily performed without DigDeAPr, thus the comparison of lower abundance protein spectral count RSDs are more relevant for applying DigDeAPr for quantitation. For spectral counts, median RSDs were 0.30 (Control) and 0.33 (DigDeAPr) and for 1-9 spectral counts, 0.43 and 0.46, respectively. The similarity of RSDs within these spectral count ranges indicates DigDeAPr should be just as precise as routinely used shotgun proteomic spectral counting methods. The higher RSDs for low spectral count proteins in both Control and DigDeAPr runs highlight the potential quantitation gains of DigDeAPr, as the methodology shifts low abundance proteins into more reproducible, higher spectral count ranges (i.e spectral counts). Additionally, these results indicate that our digestion depletions with MWCO spin-filters were within the spectral counting error of data-dependent shotgun proteomics. Despite the obvious expected changes to spectral counts of proteins with DigDeAPr, it should be a viable method for spectral counting-based protein quantitation. For peptides identified by both Control and DigDeAPr runs, we also investigated their changes in precursor intensity, an important factor for fragmentation spectra quality and quantitation. We found dramatic, statistically significant increases in precursor intensity from DigDeAPr for all peptide abundances in both Control and DigDeAPr runs (Fig. 3a) and for all peptides identified (Supplementary Fig. 6). Increased precursor intensities also directly led to greater chromatographic peak areas (Fig. 3b), as they are
21 theoretically proportional and their trends are quite similar. Precursor intensity and peak area are both relevant to label-free and metabolically-labeled protein quantitation methods. However, the potential quantitation gains may be best illustrated by improved peptide precursor S/N (Fig. 3c). Slightly more than half of the peptides (13,358 out of 23,932) with calculated ratio changes had low S/N (< 20) in Control runs. Notably, 79% of these low S/N peptides had higher S/N from employing DigDeAPr. Further, of the 6,442 peptides with S/N less than the limit of quantitation (LOQ = S/N > 10) in Control runs, 89% of these were raised above the LOQ threshold with DigDeAPr. Presumably these S/N gains would increase the number of total peptides that could be used for quantitation by ~25%, corresponding mostly to low abundance proteins. Conversely, peptides with high S/N in Control runs had minimal, acceptable decreases in S/N from digestion depletion. That is, of the 10,574 peptides with S/N greater than 20, only 5% (562) fell below the LOQ with an average S/N decrease of 18%. Thus, these precursor intensity, peak area, and S/N gains should improve accuracy, precision, and comprehensiveness of MS-based quantitation. Multiple existing and developing quantitative methods also employ MS/MS-based quantitation. Data-independent acquisition (DIA) methods for fragmenting, identifying, and quantifying peptides are more reproducible and sensitive than traditional datadependent acquisition (DDA) of peptide precursors 6 and with other similar methods emerging, 7, 8 is expected to become the norm. With DIA, peptide fragment ion intensities that contribute to the peptide identification can be summed within an MS/MS spectra and used to generate a reconstructed chromatogram from successive identifications with better S/N than from MS precursor ion spectra. Thus, we also
22 investigated the changes in summed MS/MS ion intensities (Fig. 3d) from the use of DigDeAPr. As with MS precursor intensities, all summed MS/MS intensities had similar gains to precursor intensity across all peptide abundances. Since we used DDA in this work, we could not reconstruct chromatograms from successive MS/MS spectra of the same peptide as in DIA. However, the MS/MS signal gains should translate very similarly to those in the MS scan where high precursor intensity yields both greater peak areas and higher S/N for quantitation. Additionally, although we haven t directly investigated how reporter-based MS/MS quantitation would be affected by DigDeAPr, the higher precursor intensities and S/N we found in the MS scans are also an indicator of potential gains in reporter ion intensities and S/N. Thus, DigDeAPr should ultimately also improve MS2-based reporter ion and data-independent quantitation methods.
23 Supplementary Note 5. Comparison of dynamic range compression mechanisms between DigDeAPr and hexapeptide-based ProteoMiner We previously showed that freeing of chromatographic and ion trap space using hexapeptide equalization of proteins leads to more new peptide identifications through sampling of higher precursor intensities for low abundance peptides. 4 This also appears to be the case for DigDeAPr as precursor intensities increased over all peptide abundances (Fig. 3a). These results were not surprising as the mechanisms for protein abundance adjustment using hexapeptide ligand libraries and DigDeAPr are similar, relying on either formation of protein:hexapeptide complexes with an affinity constant (K A ) or the formation of protein:protease complexes with a Michaelis constant (K M ), respectively. However, DigDeAPr appears to be less sensitive to protein isoelectric point (Supplementary Fig. 7a), with a nearly uniform relative distribution of isoelectric points over all spectral count changes, and is similarly insensitive to protein hydrophobicity (Supplementary Fig. 7b). Thus, digestion depletion may be more versatile than hexapeptide depletion and easier to implement than other common targeted depletion methods such as antibody depletion arrays. The advantages of relying on a single protease K M instead of many different protein-ligand K A s are (1) the selectivity differences are almost entirely based on protein abundance, not affinity and (2) the K M is less biased for a defined set of proteins as with a hexapeptide ligand library. Protein size may provide a slight bias with DigDeAPr since the number of tryptic sites generally scale with protein molecular weight (Supplementary Fig. 7c-d), but protein abundance remains the dominant depletion factor.
24 Supplementary Note 6. Analysis of potential protein loss from the MWCO spin-filter In addition to the expected abundance changes, some low spectral count proteins (< 10 spectral counts) also appeared depleted from DigDeAPr. Of these proteins, ~ 50 were consistently depleted (log 2 ratio < -1 and p < 0.05) in comparison to Control runs. However, when we examined this small number of proteins physicochemical properties, their isoelectric point, molecular weight, and hydrophobicity appeared equally distributed in comparison to all the proteins identified in Control runs. Overall, the low abundance protein spectral count decreases could be attributed to aspects of the DigDeAPr methodology, such as losses from the MWCO spin-filter, but without an obvious physicochemical trend it appears unlikely. Most of the low abundance protein depletion trend can likely be attributed to the pseudo-random sampling of low abundance peptide precursor ions in shotgun proteomics 9 and the inability to measure losses of low abundance proteins with data-dependent acquisition. This is illustrated by the comparison of two separate sets of triplicate Control analyses (Control versus Control). The comparison shows uniform ratio distributions for spectral counts and sequence coverages for all abundances (Supplementary Fig. 8), unlike DigDeAPr versus Control comparisons (Fig. 2c-d) where high abundance proteins are skewed towards decreasing ratios and low abundance proteins towards increasing ratios. This comparison not only further validates the spectral count changes observed with DigDeAPr, but also illustrates the expected variance in spectral counts based on abundance. That is, the low abundance proteins which appear depleted by DigDeAPr actually fall within the variance of spectral counting comparison. The spread of spectral count ratios is also represented as spectral count relative standard deviations (RSDs)
25 for Control and DigDeAPr runs (Fig. 2f). Comparison of these RSDs allows for the evaluation of the reproducibility of our digestion depletions. We found independently performed digestion depletions of HEK lysates, estimated by BCA mass balance analysis to be 85 ± 10% depleted, resulted in similar spectral count RSDs as Control runs for proteins with less than 100 spectral counts. Proteins with greater than 100 spectral counts had slightly higher median RSDs for DigDeAPr (0.32) versus Control (0.24). These RSDs directly illustrate the variation of the digestion depletion since high abundance proteins should be most affected by these steps. Thus, the 8% higher RSD than Control runs is consistent with our ability to measure the digestion depletion with BCA mass balance to ± 10%.
26 References 1. Finehout, E.J., Cantor, J.R. & Lee, K.H. Proteomics 5, (2005). 2. Mense, S.M. et al. Physiological genomics 25, (2006). 3. Schwanhausser, B. et al. Nature 473, (2011). 4. Fonslow, B.R. et al. Journal of proteome research 10, (2011). 5. Ghaemmaghami, S. et al. Nature 425, (2003). 6. Venable, J.D., Dong, M.Q., Wohlschlegel, J., Dillin, A. & Yates, J.R. Nature methods 1, (2004). 7. Gillet, L.C. et al. Mol Cell Proteomics 11, O (2012). 8. Panchaud, A., Jung, S., Shaffer, S.A., Aitchison, J.D. & Goodlett, D.R. Anal Chem 83, (2011). 9. Liu, H., Sadygov, R.G. & Yates, J.R., 3rd Anal Chem 76, (2004).
Digitizing the Proteomes From Big Tissue Biobanks
 Digitizing the Proteomes From Big Tissue Biobanks Analyzing 24 Proteomes Per Day by Microflow SWATH Acquisition and Spectronaut Pulsar Analysis Jan Muntel 1, Nick Morrice 2, Roland M. Bruderer 1, Lukas
Digitizing the Proteomes From Big Tissue Biobanks Analyzing 24 Proteomes Per Day by Microflow SWATH Acquisition and Spectronaut Pulsar Analysis Jan Muntel 1, Nick Morrice 2, Roland M. Bruderer 1, Lukas
Symposium on Protein Fractionation
 Symposium on Protein Fractionation Proteomic Analysis of Organ-Specific Breast Cancer Metastasis Speaker: Emily I. Chen, Ph.D The Scripps Research Institute Department of Cell Biology CANCER METASTASIS
Symposium on Protein Fractionation Proteomic Analysis of Organ-Specific Breast Cancer Metastasis Speaker: Emily I. Chen, Ph.D The Scripps Research Institute Department of Cell Biology CANCER METASTASIS
NIH Public Access Author Manuscript J Proteome Res. Author manuscript; available in PMC 2014 July 05.
 NIH Public Access Author Manuscript Published in final edited form as: J Proteome Res. 2013 July 5; 12(7): 3071 3086. doi:10.1021/pr3011588. Evaluation and Optimization of Mass Spectrometric Settings during
NIH Public Access Author Manuscript Published in final edited form as: J Proteome Res. 2013 July 5; 12(7): 3071 3086. doi:10.1021/pr3011588. Evaluation and Optimization of Mass Spectrometric Settings during
[ APPLICATION NOTE ] High Sensitivity Intact Monoclonal Antibody (mab) HRMS Quantification APPLICATION BENEFITS INTRODUCTION WATERS SOLUTIONS KEYWORDS
![[ APPLICATION NOTE ] High Sensitivity Intact Monoclonal Antibody (mab) HRMS Quantification APPLICATION BENEFITS INTRODUCTION WATERS SOLUTIONS KEYWORDS [ APPLICATION NOTE ] High Sensitivity Intact Monoclonal Antibody (mab) HRMS Quantification APPLICATION BENEFITS INTRODUCTION WATERS SOLUTIONS KEYWORDS](/thumbs/79/80328542.jpg) Yun Wang Alelyunas, Henry Shion, Mark Wrona Waters Corporation, Milford, MA, USA APPLICATION BENEFITS mab LC-MS method which enables users to achieve highly sensitive bioanalysis of intact trastuzumab
Yun Wang Alelyunas, Henry Shion, Mark Wrona Waters Corporation, Milford, MA, USA APPLICATION BENEFITS mab LC-MS method which enables users to achieve highly sensitive bioanalysis of intact trastuzumab
The distribution of log 2 ratio (H/L) for quantified peptides. cleavage sites in each bin of log 2 ratio of quantified. peptides
 Journal: Nature Methods Article Title: Corresponding Author: Protein digestion priority is independent of their abundances Mingliang Ye and Hanfa Zou Supplementary Figure 1 Supplementary Figure 2 The distribution
Journal: Nature Methods Article Title: Corresponding Author: Protein digestion priority is independent of their abundances Mingliang Ye and Hanfa Zou Supplementary Figure 1 Supplementary Figure 2 The distribution
Figure S6. A-J) Annotated UVPD mass spectra for top ten peptides found among the peptides identified by Byonic but not SEQUEST + Percolator.
 Extending Proteome Coverage by Combining MS/MS Methods and a Modified Bioinformatics Platform adapted for Database Searching of Positive and Negative Polarity 193 nm Ultraviolet Photodissociation Mass
Extending Proteome Coverage by Combining MS/MS Methods and a Modified Bioinformatics Platform adapted for Database Searching of Positive and Negative Polarity 193 nm Ultraviolet Photodissociation Mass
Nature Methods: doi: /nmeth.3177
 Supplementary Figure 1 Characterization of LysargiNase, trypsin and LysN missed cleavages. (a) Proportion of peptides identified in LysargiNase and trypsin digests of MDA-MB-231 cell lysates carrying 0,
Supplementary Figure 1 Characterization of LysargiNase, trypsin and LysN missed cleavages. (a) Proportion of peptides identified in LysargiNase and trypsin digests of MDA-MB-231 cell lysates carrying 0,
Application Note # LCMS-89 High quantification efficiency in plasma targeted proteomics with a full-capability discovery Q-TOF platform
 Application Note # LCMS-89 High quantification efficiency in plasma targeted proteomics with a full-capability discovery Q-TOF platform Abstract Targeted proteomics for biomarker verification/validation
Application Note # LCMS-89 High quantification efficiency in plasma targeted proteomics with a full-capability discovery Q-TOF platform Abstract Targeted proteomics for biomarker verification/validation
Supporting Information. Lysine Propionylation to Boost Proteome Sequence. Coverage and Enable a Silent SILAC Strategy for
 Supporting Information Lysine Propionylation to Boost Proteome Sequence Coverage and Enable a Silent SILAC Strategy for Relative Protein Quantification Christoph U. Schräder 1, Shaun Moore 1,2, Aaron A.
Supporting Information Lysine Propionylation to Boost Proteome Sequence Coverage and Enable a Silent SILAC Strategy for Relative Protein Quantification Christoph U. Schräder 1, Shaun Moore 1,2, Aaron A.
Nature Structural & Molecular Biology: doi: /nsmb.3218
 Supplementary Figure 1 Endogenous EGFR trafficking and responses depend on biased ligands. (a) Lysates from HeLa cells stimulated for 2 min. with increasing concentration of ligands were immunoblotted
Supplementary Figure 1 Endogenous EGFR trafficking and responses depend on biased ligands. (a) Lysates from HeLa cells stimulated for 2 min. with increasing concentration of ligands were immunoblotted
Unsupervised Identification of Isotope-Labeled Peptides
 Unsupervised Identification of Isotope-Labeled Peptides Joshua E Goldford 13 and Igor GL Libourel 124 1 Biotechnology institute, University of Minnesota, Saint Paul, MN 55108 2 Department of Plant Biology,
Unsupervised Identification of Isotope-Labeled Peptides Joshua E Goldford 13 and Igor GL Libourel 124 1 Biotechnology institute, University of Minnesota, Saint Paul, MN 55108 2 Department of Plant Biology,
PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System
 Application Note LC/MS PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System Purpose This application note describes an automated workflow
Application Note LC/MS PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System Purpose This application note describes an automated workflow
Bioanalytical Quantitation of Biotherapeutics Using Intact Protein vs. Proteolytic Peptides by LC-HR/AM on a Q Exactive MS
 Bioanalytical Quantitation of Biotherapeutics Using Intact Protein vs. Proteolytic Peptides by LC-HR/AM on a Q Exactive MS Jenny Chen, Hongxia Wang, Zhiqi Hao, Patrick Bennett, and Greg Kilby Thermo Fisher
Bioanalytical Quantitation of Biotherapeutics Using Intact Protein vs. Proteolytic Peptides by LC-HR/AM on a Q Exactive MS Jenny Chen, Hongxia Wang, Zhiqi Hao, Patrick Bennett, and Greg Kilby Thermo Fisher
Quantitation of Protein Phosphorylation Using Multiple Reaction Monitoring
 Quantitation of Protein Phosphorylation Using Multiple Reaction Monitoring Application Note Authors Ning Tang, Christine A. Miller and Keith Waddell Agilent Technologies, Inc. Santa Clara, CA USA This
Quantitation of Protein Phosphorylation Using Multiple Reaction Monitoring Application Note Authors Ning Tang, Christine A. Miller and Keith Waddell Agilent Technologies, Inc. Santa Clara, CA USA This
Electronic Supplementary Information NOVEL NON-TARGET ANALYSIS FOR FLUORINE COMPOUNDS USING ICPMS/MS AND HPLC-ICPMS/MS
 Electronic Supplementary Material (ESI) for Journal of Analytical Atomic Spectrometry. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information NOVEL NON-TARGET ANALYSIS
Electronic Supplementary Material (ESI) for Journal of Analytical Atomic Spectrometry. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information NOVEL NON-TARGET ANALYSIS
Comprehensive Forensic Toxicology Screening in Serum using On-Line SPE LC-MS/MS
 Comprehensive Forensic Toxicology Screening in Serum using On-Line SPE LC-MS/MS SCIEX QTRAP 4500 LC-MS/MS System and Spark Holland PICO Adrian M. Taylor 1, Peter Ringeling 2, Martin Sibum 2, Stefan Sturm
Comprehensive Forensic Toxicology Screening in Serum using On-Line SPE LC-MS/MS SCIEX QTRAP 4500 LC-MS/MS System and Spark Holland PICO Adrian M. Taylor 1, Peter Ringeling 2, Martin Sibum 2, Stefan Sturm
Lecture 3. Tandem MS & Protein Sequencing
 Lecture 3 Tandem MS & Protein Sequencing Nancy Allbritton, M.D., Ph.D. Department of Physiology & Biophysics 824-9137 (office) nlallbri@uci.edu Office- Rm D349 Medical Science D Bldg. Tandem MS Steps:
Lecture 3 Tandem MS & Protein Sequencing Nancy Allbritton, M.D., Ph.D. Department of Physiology & Biophysics 824-9137 (office) nlallbri@uci.edu Office- Rm D349 Medical Science D Bldg. Tandem MS Steps:
Improve Protein Analysis with the New, Mass Spectrometry- Compatible ProteasMAX Surfactant
 Improve Protein Analysis with the New, Mass Spectrometry- Compatible Surfactant ABSTRACT Incomplete solubilization and digestion and poor peptide recovery are frequent limitations in protein sample preparation
Improve Protein Analysis with the New, Mass Spectrometry- Compatible Surfactant ABSTRACT Incomplete solubilization and digestion and poor peptide recovery are frequent limitations in protein sample preparation
4th Multidimensional Chromatography Workshop Toronto (January, 2013) Herman C. Lam, Ph.D. Calibration & Validation Group
 4th Multidimensional Chromatography Workshop Toronto (January, 2013) Herman C. Lam, Ph.D. Calibration & Validation Group MDLC for Shotgun Proteomics Introduction General concepts Advantages Challenges
4th Multidimensional Chromatography Workshop Toronto (January, 2013) Herman C. Lam, Ph.D. Calibration & Validation Group MDLC for Shotgun Proteomics Introduction General concepts Advantages Challenges
Supplementary Figure S1. Appearacne of new acetyl groups in acetylated lysines using 2,3-13 C 6 pyruvate as a tracer instead of labeled glucose.
 Supplementary Figure S1. Appearacne of new acetyl groups in acetylated lysines using 2,3-13 C 6 pyruvate as a tracer instead of labeled glucose. (a) Relative levels of both new acetylation and all acetylation
Supplementary Figure S1. Appearacne of new acetyl groups in acetylated lysines using 2,3-13 C 6 pyruvate as a tracer instead of labeled glucose. (a) Relative levels of both new acetylation and all acetylation
MS1 and MS2 crosstalk in label free quantitation of mass spectrometry data independent acquisitions
 MS1 and MS2 crosstalk in label free quantitation of mass spectrometry data independent acquisitions MS1 528.18 +++ m/z 568.98 ++ m/z 678.34 ++ m/z MS2/SWATH June 9th, 2013 Matthew J. Rardin SIRT3 regulated
MS1 and MS2 crosstalk in label free quantitation of mass spectrometry data independent acquisitions MS1 528.18 +++ m/z 568.98 ++ m/z 678.34 ++ m/z MS2/SWATH June 9th, 2013 Matthew J. Rardin SIRT3 regulated
APPLICATION NOTE. Highly reproducible and Comprehensive Proteome Profiling of Formalin-Fixed Paraffin-Embedded (FFPE) Tissues Slices
 APPLICATION NOTE Highly reproducible and Comprehensive Proteome Profiling of Formalin-Fixed Paraffin-Embedded (FFPE) Tissues Slices INTRODUCTION Preservation of tissue biopsies is a critical step to This
APPLICATION NOTE Highly reproducible and Comprehensive Proteome Profiling of Formalin-Fixed Paraffin-Embedded (FFPE) Tissues Slices INTRODUCTION Preservation of tissue biopsies is a critical step to This
MS/MS to Targeted Proteomics (MRM)
 MS/MS to Targeted Proteomics (MRM) How it worked on the Human Lens Proteome Jayson Falkner PhD jay@singleorganism.com Genes Show Limited Value in Predicting Diseases With only a few exceptions, what the
MS/MS to Targeted Proteomics (MRM) How it worked on the Human Lens Proteome Jayson Falkner PhD jay@singleorganism.com Genes Show Limited Value in Predicting Diseases With only a few exceptions, what the
Extended Mass Range Triple Quadrupole for Routine Analysis of High Mass-to-charge Peptide Ions
 Extended Mass Range Triple Quadrupole for Routine Analysis of High Mass-to-charge Peptide Ions Application Note Targeted Proteomics Authors Linfeng Wu, Christine A. Miller, Jordy Hsiao, Te-wei Chu, Behrooz
Extended Mass Range Triple Quadrupole for Routine Analysis of High Mass-to-charge Peptide Ions Application Note Targeted Proteomics Authors Linfeng Wu, Christine A. Miller, Jordy Hsiao, Te-wei Chu, Behrooz
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/6/278/rs11/dc1 Supplementary Materials for In Vivo Phosphoproteomics Analysis Reveals the Cardiac Targets of β-adrenergic Receptor Signaling Alicia Lundby,* Martin
www.sciencesignaling.org/cgi/content/full/6/278/rs11/dc1 Supplementary Materials for In Vivo Phosphoproteomics Analysis Reveals the Cardiac Targets of β-adrenergic Receptor Signaling Alicia Lundby,* Martin
Nature Biotechnology: doi: /nbt Supplementary Figure 1
 Supplementary Figure 1 The timeline of the NGAG method for extraction of N-linked glycans and glycosite-containing peptides. The timeline can be changed based on the number of samples. Supplementary Figure
Supplementary Figure 1 The timeline of the NGAG method for extraction of N-linked glycans and glycosite-containing peptides. The timeline can be changed based on the number of samples. Supplementary Figure
Conditional spectrum-based ground motion selection. Part II: Intensity-based assessments and evaluation of alternative target spectra
 EARTHQUAKE ENGINEERING & STRUCTURAL DYNAMICS Published online 9 May 203 in Wiley Online Library (wileyonlinelibrary.com)..2303 Conditional spectrum-based ground motion selection. Part II: Intensity-based
EARTHQUAKE ENGINEERING & STRUCTURAL DYNAMICS Published online 9 May 203 in Wiley Online Library (wileyonlinelibrary.com)..2303 Conditional spectrum-based ground motion selection. Part II: Intensity-based
Shotgun metaproteomics of the human distal gut microbiota. Present by Lei Chen
 Shotgun metaproteomics of the human distal gut microbiota Present by Lei Chen (lc6@indana.edu) Outline Background What are the goals? Materials and Methods Results Discussion Background The human gastrointestinal
Shotgun metaproteomics of the human distal gut microbiota Present by Lei Chen (lc6@indana.edu) Outline Background What are the goals? Materials and Methods Results Discussion Background The human gastrointestinal
2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry
 Dr. Sanjeeva Srivastava 1. Fundamental of Mass Spectrometry Role of MS and basic concepts 2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry 2 1 MS basic concepts Mass spectrometry - technique
Dr. Sanjeeva Srivastava 1. Fundamental of Mass Spectrometry Role of MS and basic concepts 2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry 2 1 MS basic concepts Mass spectrometry - technique
Characterization of Disulfide Linkages in Proteins by 193 nm Ultraviolet Photodissociation (UVPD) Mass Spectrometry. Supporting Information
 Characterization of Disulfide Linkages in Proteins by 193 nm Ultraviolet Photodissociation (UVPD) Mass Spectrometry M. Montana Quick, Christopher M. Crittenden, Jake A. Rosenberg, and Jennifer S. Brodbelt
Characterization of Disulfide Linkages in Proteins by 193 nm Ultraviolet Photodissociation (UVPD) Mass Spectrometry M. Montana Quick, Christopher M. Crittenden, Jake A. Rosenberg, and Jennifer S. Brodbelt
 a RT:. - 1.1 1 95 9 85 8 27.64 27.87 28.1 27.42 28.24 38.98 Low ph RPLC NL: 2.29E9 TIC MS TrypticBSA_Dig est_replicate1_ 2Mar18_Preci ous_18-1-6 75 7 65 48.75 65.8 Relative Abundance 6 55 5 45 28.54 48.98
a RT:. - 1.1 1 95 9 85 8 27.64 27.87 28.1 27.42 28.24 38.98 Low ph RPLC NL: 2.29E9 TIC MS TrypticBSA_Dig est_replicate1_ 2Mar18_Preci ous_18-1-6 75 7 65 48.75 65.8 Relative Abundance 6 55 5 45 28.54 48.98
Structural Characterization of Prion-like Conformational Changes of the Neuronal Isoform of Aplysia CPEB
 Structural Characterization of Prion-like Conformational Changes of the Neuronal Isoform of Aplysia CPEB Bindu L. Raveendra, 1,5 Ansgar B. Siemer, 2,6 Sathyanarayanan V. Puthanveettil, 1,3,7 Wayne A. Hendrickson,
Structural Characterization of Prion-like Conformational Changes of the Neuronal Isoform of Aplysia CPEB Bindu L. Raveendra, 1,5 Ansgar B. Siemer, 2,6 Sathyanarayanan V. Puthanveettil, 1,3,7 Wayne A. Hendrickson,
Amadeo R. Fernández-Alba
 % of compounds % of compounds % of compounds % of compounds Amadeo R. Fernández-Alba LC-Orbitrap QExactive Focus Instrumental LOQ 1% 9% 8% 7% 6% 5% 4% 3% 2% 1% %.1 mg/g.2 mg/g Tomato.5 mg/g ddms2 Target
% of compounds % of compounds % of compounds % of compounds Amadeo R. Fernández-Alba LC-Orbitrap QExactive Focus Instrumental LOQ 1% 9% 8% 7% 6% 5% 4% 3% 2% 1% %.1 mg/g.2 mg/g Tomato.5 mg/g ddms2 Target
Quantification with Proteome Discoverer. Bernard Delanghe
 Quantification with Proteome Discoverer Bernard Delanghe Overview: Which approach to use? Proteome Discoverer Quantification Method What When to use Metabolic labeling SILAC Cell culture systems Small
Quantification with Proteome Discoverer Bernard Delanghe Overview: Which approach to use? Proteome Discoverer Quantification Method What When to use Metabolic labeling SILAC Cell culture systems Small
Protein Reports CPTAC Common Data Analysis Pipeline (CDAP)
 Protein Reports CPTAC Common Data Analysis Pipeline (CDAP) v. 05/03/2016 Summary The purpose of this document is to describe the protein reports generated as part of the CPTAC Common Data Analysis Pipeline
Protein Reports CPTAC Common Data Analysis Pipeline (CDAP) v. 05/03/2016 Summary The purpose of this document is to describe the protein reports generated as part of the CPTAC Common Data Analysis Pipeline
Vitamin D Metabolite Analysis in Biological Samples Using Agilent Captiva EMR Lipid
 Vitamin D Metabolite Analysis in Biological Samples Using Agilent Captiva EMR Lipid Application Note Clinical Research Authors Derick Lucas and Limian Zhao Agilent Technologies, Inc. Abstract Lipids from
Vitamin D Metabolite Analysis in Biological Samples Using Agilent Captiva EMR Lipid Application Note Clinical Research Authors Derick Lucas and Limian Zhao Agilent Technologies, Inc. Abstract Lipids from
Shotgun Proteomics MS/MS. Protein Mixture. proteolysis. Peptide Mixture. Time. Abundance. Abundance. m/z. Abundance. m/z 2. Abundance.
 Abundance Abundance Abundance Abundance Abundance Shotgun Proteomics Protein Mixture 1 2 3 MS/MS proteolysis m/z 2 3 Time µlc m/z MS 1 m/z Peptide Mixture m/z Block Diagram of a Mass Spectrometer Sample
Abundance Abundance Abundance Abundance Abundance Shotgun Proteomics Protein Mixture 1 2 3 MS/MS proteolysis m/z 2 3 Time µlc m/z MS 1 m/z Peptide Mixture m/z Block Diagram of a Mass Spectrometer Sample
Mass Spectrometry and Proteomics - Lecture 4 - Matthias Trost Newcastle University
 Mass Spectrometry and Proteomics - Lecture 4 - Matthias Trost Newcastle University matthias.trost@ncl.ac.uk previously Peptide fragmentation Hybrid instruments 117 The Building Blocks of Life DNA RNA Proteins
Mass Spectrometry and Proteomics - Lecture 4 - Matthias Trost Newcastle University matthias.trost@ncl.ac.uk previously Peptide fragmentation Hybrid instruments 117 The Building Blocks of Life DNA RNA Proteins
Quantitation by High Resolution Mass Spectrometry: Case Study of TOF MS for the Quantitation of Allopurinol from Human Plasma
 Quantitation by High Resolution Mass Spectrometry: Case Study of TOF MS for the Quantitation of Allopurinol from Human Plasma Shaokun Pang 1, Weixing Sun 2, Adrien Musuku 2, Xavier J. Misonne 1 1 SCIEX,
Quantitation by High Resolution Mass Spectrometry: Case Study of TOF MS for the Quantitation of Allopurinol from Human Plasma Shaokun Pang 1, Weixing Sun 2, Adrien Musuku 2, Xavier J. Misonne 1 1 SCIEX,
Nature Neuroscience: doi: /nn Supplementary Figure 1. Behavioral training.
 Supplementary Figure 1 Behavioral training. a, Mazes used for behavioral training. Asterisks indicate reward location. Only some example mazes are shown (for example, right choice and not left choice maze
Supplementary Figure 1 Behavioral training. a, Mazes used for behavioral training. Asterisks indicate reward location. Only some example mazes are shown (for example, right choice and not left choice maze
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature13418 Supplementary Results: USP30 opposes autophagic flux In HEK-293 cells, USP30 overexpression increased basal LC3-II levels, dependent on enzymatic activity,
SUPPLEMENTARY INFORMATION doi:10.1038/nature13418 Supplementary Results: USP30 opposes autophagic flux In HEK-293 cells, USP30 overexpression increased basal LC3-II levels, dependent on enzymatic activity,
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/8/398/rs12/dc1 Supplementary Materials for Quantitative phosphoproteomics reveals new roles for the protein phosphatase PP6 in mitotic cells Scott F. Rusin, Kate
www.sciencesignaling.org/cgi/content/full/8/398/rs12/dc1 Supplementary Materials for Quantitative phosphoproteomics reveals new roles for the protein phosphatase PP6 in mitotic cells Scott F. Rusin, Kate
The MOLECULES of LIFE
 The MOLECULES of LIFE Physical and Chemical Principles Solutions Manual Prepared by James Fraser and Samuel Leachman Chapter 16 Principles of Enzyme Catalysis Problems True/False and Multiple Choice 1.
The MOLECULES of LIFE Physical and Chemical Principles Solutions Manual Prepared by James Fraser and Samuel Leachman Chapter 16 Principles of Enzyme Catalysis Problems True/False and Multiple Choice 1.
SYNAPT G2-S High Definition MS (HDMS) System
 SYNAPT G2-S High Definition MS (HDMS) System High performance, versatility, and workflow efficiency of your MS system all play a crucial role in your ability to successfully reach your scientific and business
SYNAPT G2-S High Definition MS (HDMS) System High performance, versatility, and workflow efficiency of your MS system all play a crucial role in your ability to successfully reach your scientific and business
Considerations of the use of Triple Quadrupoles or Ion Traps in Quantitative Applications
 Considerations of the use of Triple Quadrupoles or Ion Traps in Quantitative Applications Triple Stage Quadrupole API MS / MS Full Scan Products / - IONS AND NEUTRALS FORMED IN API SOURCE Q0 LENS TRANSPORTS
Considerations of the use of Triple Quadrupoles or Ion Traps in Quantitative Applications Triple Stage Quadrupole API MS / MS Full Scan Products / - IONS AND NEUTRALS FORMED IN API SOURCE Q0 LENS TRANSPORTS
Probability-Based Protein Identification for Post-Translational Modifications and Amino Acid Variants Using Peptide Mass Fingerprint Data
 Probability-Based Protein Identification for Post-Translational Modifications and Amino Acid Variants Using Peptide Mass Fingerprint Data Tong WW, McComb ME, Perlman DH, Huang H, O Connor PB, Costello
Probability-Based Protein Identification for Post-Translational Modifications and Amino Acid Variants Using Peptide Mass Fingerprint Data Tong WW, McComb ME, Perlman DH, Huang H, O Connor PB, Costello
ACQUITY UPLC WITH PDA DETECTION: DETERMINING THE SENSITIVITY LIMITS OF OXYBUTYNIN AND RELATED COMPOUNDS
 ACQUITY UPLC WITH PDA DETECTION: DETERMINING THE SENSITIVITY LIMITS OF OXYBUTYNIN AND RELATED COMPOUNDS Tanya Jenkins Waters Corporation, Milford, MA, USA INTRODUCTION Some of the most challenging methods
ACQUITY UPLC WITH PDA DETECTION: DETERMINING THE SENSITIVITY LIMITS OF OXYBUTYNIN AND RELATED COMPOUNDS Tanya Jenkins Waters Corporation, Milford, MA, USA INTRODUCTION Some of the most challenging methods
Enhancing Sequence Coverage in Proteomics Studies by Using a Combination of Proteolytic Enzymes
 Enhancing Sequence Coverage in Proteomics Studies by Using a Combination of Proteolytic Enzymes Dominic Baeumlisberger 2, Christopher Kurz 3, Tabiwang N. Arrey, Marion Rohmer 2, Carola Schiller 3, Thomas
Enhancing Sequence Coverage in Proteomics Studies by Using a Combination of Proteolytic Enzymes Dominic Baeumlisberger 2, Christopher Kurz 3, Tabiwang N. Arrey, Marion Rohmer 2, Carola Schiller 3, Thomas
Double charge of 33kD peak A1 A2 B1 B2 M2+ M/z. ABRF Proteomics Research Group - Qualitative Proteomics Study Identifier Number 14146
 Abstract The 2008 ABRF Proteomics Research Group Study offers participants the chance to participate in an anonymous study to identify qualitative differences between two protein preparations. We used
Abstract The 2008 ABRF Proteomics Research Group Study offers participants the chance to participate in an anonymous study to identify qualitative differences between two protein preparations. We used
Moving from targeted towards non-targeted approaches
 Gesundheitsdirektion Moving from targeted towards non-targeted approaches Anton Kaufmann Official Food Control Authority of the Canton of Zurich () Switzerland 2 Overview I From single residue to multi
Gesundheitsdirektion Moving from targeted towards non-targeted approaches Anton Kaufmann Official Food Control Authority of the Canton of Zurich () Switzerland 2 Overview I From single residue to multi
Chapter 11: Enzyme Catalysis
 Chapter 11: Enzyme Catalysis Matching A) high B) deprotonated C) protonated D) least resistance E) motion F) rate-determining G) leaving group H) short peptides I) amino acid J) low K) coenzymes L) concerted
Chapter 11: Enzyme Catalysis Matching A) high B) deprotonated C) protonated D) least resistance E) motion F) rate-determining G) leaving group H) short peptides I) amino acid J) low K) coenzymes L) concerted
Using Software Tools to Improve the Detection of Impurities by LC/MS. Application Note. Christine Miller Agilent Technologies.
 Using Software Tools to Improve the Detection of Impurities Application Note Christine Miller Introduction The analysis of raw materials and finished products for or impurities presents a challenge in
Using Software Tools to Improve the Detection of Impurities Application Note Christine Miller Introduction The analysis of raw materials and finished products for or impurities presents a challenge in
MASS SPECTROMETRY BASED METABOLOMICS. Pavel Aronov. ABRF2010 Metabolomics Research Group March 21, 2010
 MASS SPECTROMETRY BASED METABOLOMICS Pavel Aronov ABRF2010 Metabolomics Research Group March 21, 2010 Types of Experiments in Metabolomics targeted non targeted Number of analyzed metabolites is limited
MASS SPECTROMETRY BASED METABOLOMICS Pavel Aronov ABRF2010 Metabolomics Research Group March 21, 2010 Types of Experiments in Metabolomics targeted non targeted Number of analyzed metabolites is limited
Supporting Information. Post translational Modifications of Serotonin Type 4 Receptor Heterologously Expressed in. Mouse Rod Cells
 Supporting Information Post translational Modifications of Serotonin Type 4 Receptor Heterologously Expressed in Mouse Rod Cells David Salom,, Benlian Wang,, Zhiqian Dong, Wenyu Sun, Pius Padayatti, Steven
Supporting Information Post translational Modifications of Serotonin Type 4 Receptor Heterologously Expressed in Mouse Rod Cells David Salom,, Benlian Wang,, Zhiqian Dong, Wenyu Sun, Pius Padayatti, Steven
LC/MS/MS SOLUTIONS FOR LIPIDOMICS. Biomarker and Omics Solutions FOR DISCOVERY AND TARGETED LIPIDOMICS
 LC/MS/MS SOLUTIONS FOR LIPIDOMICS Biomarker and Omics Solutions FOR DISCOVERY AND TARGETED LIPIDOMICS Lipids play a key role in many biological processes, such as the formation of cell membranes and signaling
LC/MS/MS SOLUTIONS FOR LIPIDOMICS Biomarker and Omics Solutions FOR DISCOVERY AND TARGETED LIPIDOMICS Lipids play a key role in many biological processes, such as the formation of cell membranes and signaling
Learning Objectives. Overview of topics to be discussed 10/25/2013 HIGH RESOLUTION MASS SPECTROMETRY (HRMS) IN DISCOVERY PROTEOMICS
 HIGH RESOLUTION MASS SPECTROMETRY (HRMS) IN DISCOVERY PROTEOMICS A clinical proteomics perspective Michael L. Merchant, PhD School of Medicine, University of Louisville Louisville, KY Learning Objectives
HIGH RESOLUTION MASS SPECTROMETRY (HRMS) IN DISCOVERY PROTEOMICS A clinical proteomics perspective Michael L. Merchant, PhD School of Medicine, University of Louisville Louisville, KY Learning Objectives
Chapter 1: Exploring Data
 Chapter 1: Exploring Data Key Vocabulary:! individual! variable! frequency table! relative frequency table! distribution! pie chart! bar graph! two-way table! marginal distributions! conditional distributions!
Chapter 1: Exploring Data Key Vocabulary:! individual! variable! frequency table! relative frequency table! distribution! pie chart! bar graph! two-way table! marginal distributions! conditional distributions!
Automating Mass Spectrometry-Based Quantitative Glycomics using Tandem Mass Tag (TMT) Reagents with SimGlycan
 PREMIER Biosoft Automating Mass Spectrometry-Based Quantitative Glycomics using Tandem Mass Tag (TMT) Reagents with SimGlycan Ne uaca2-3galb1-4glc NAcb1 6 Gal NAca -Thr 3 Ne uaca2-3galb1 Ningombam Sanjib
PREMIER Biosoft Automating Mass Spectrometry-Based Quantitative Glycomics using Tandem Mass Tag (TMT) Reagents with SimGlycan Ne uaca2-3galb1-4glc NAcb1 6 Gal NAca -Thr 3 Ne uaca2-3galb1 Ningombam Sanjib
UPLC/MS Monitoring of Water-Soluble Vitamin Bs in Cell Culture Media in Minutes
 UPLC/MS Monitoring of Water-Soluble Vitamin Bs in Cell Culture Media in Minutes Catalin E. Doneanu, Weibin Chen, and Jeffrey R. Mazzeo Waters Corporation, Milford, MA, U.S. A P P L I C AT ION B E N E F
UPLC/MS Monitoring of Water-Soluble Vitamin Bs in Cell Culture Media in Minutes Catalin E. Doneanu, Weibin Chen, and Jeffrey R. Mazzeo Waters Corporation, Milford, MA, U.S. A P P L I C AT ION B E N E F
Search settings MaxQuant
 Search settings MaxQuant Briefly, we used MaxQuant version 1.5.0.0 with the following settings. As variable modifications we allowed Acetyl (Protein N-terminus), methionine oxidation and glutamine to pyroglutamate
Search settings MaxQuant Briefly, we used MaxQuant version 1.5.0.0 with the following settings. As variable modifications we allowed Acetyl (Protein N-terminus), methionine oxidation and glutamine to pyroglutamate
Identification of Steroids in Water by Ion Trap LC/MS/MS Application
 Identification of Steroids in Water by Ion Trap LC/MS/MS Application Environmental Author Paul Zavitsanos and Christine Miller Agilent Technologies, Inc. Centerville Road Wilmington, DE 9- USA Abstract
Identification of Steroids in Water by Ion Trap LC/MS/MS Application Environmental Author Paul Zavitsanos and Christine Miller Agilent Technologies, Inc. Centerville Road Wilmington, DE 9- USA Abstract
Electronic Supporting Information
 Electronic Supplementary Material (ESI) for Green Chemistry. This journal is The Royal Society of Chemistry 2018 Electronic Supporting Information Two-phase systems developed with hydrophilic and hydrophobic
Electronic Supplementary Material (ESI) for Green Chemistry. This journal is The Royal Society of Chemistry 2018 Electronic Supporting Information Two-phase systems developed with hydrophilic and hydrophobic
Journal of Chemical and Pharmaceutical Research
 Available on line www.jocpr.com Journal of Chemical and Pharmaceutical Research ISSN No: 0975-7384 CODEN(USA): JCPRC5 J. Chem. Pharm. Res., 2011, 3(2):770-775 Validation of Rapid Liquid Chromatographic
Available on line www.jocpr.com Journal of Chemical and Pharmaceutical Research ISSN No: 0975-7384 CODEN(USA): JCPRC5 J. Chem. Pharm. Res., 2011, 3(2):770-775 Validation of Rapid Liquid Chromatographic
Comparison of Relative Quantification of Monoclonal Antibody N-glycans Using Fluorescence and MS Detection
 Comparison of Relative Quantification of Monoclonal ntibody N-glycans Using Fluorescence and MS Detection pplication Note iotherapeutics & iologics uthors scar Potter and Gregory Staples gilent Technologies,
Comparison of Relative Quantification of Monoclonal ntibody N-glycans Using Fluorescence and MS Detection pplication Note iotherapeutics & iologics uthors scar Potter and Gregory Staples gilent Technologies,
Comparison of mass spectrometers performances
 Comparison of mass spectrometers performances Instrument Mass Mass Sensitivity resolution accuracy Quadrupole 1 x 10 3 0.1 Da* 0.5-1.0 pmol DE-MALDI 2 x 10 4 20 ppm 1-10 fmol peptide 1-5 pmol protein Ion
Comparison of mass spectrometers performances Instrument Mass Mass Sensitivity resolution accuracy Quadrupole 1 x 10 3 0.1 Da* 0.5-1.0 pmol DE-MALDI 2 x 10 4 20 ppm 1-10 fmol peptide 1-5 pmol protein Ion
Don t miss a thing on your peptide mapping journey How to get full coverage peptide maps using high resolution accurate mass spectrometry
 Don t miss a thing on your peptide mapping journey How to get full coverage peptide maps using high resolution accurate mass spectrometry Kai Scheffler, PhD BioPharma Support Expert,LSMS Europe The world
Don t miss a thing on your peptide mapping journey How to get full coverage peptide maps using high resolution accurate mass spectrometry Kai Scheffler, PhD BioPharma Support Expert,LSMS Europe The world
Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random
 S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
Dr. Erin E. Chambers Waters Corporation. Presented by Dr. Diego Rodriguez Cabaleiro Waters Europe Waters Corporation 1
 Development of an SPE-LC/MS/MS Assay for the Simultaneous Quantification of Amyloid Beta Peptides in Cerebrospinal Fluid in Support of Alzheimer s Research Dr. Erin E. Chambers Waters Corporation Presented
Development of an SPE-LC/MS/MS Assay for the Simultaneous Quantification of Amyloid Beta Peptides in Cerebrospinal Fluid in Support of Alzheimer s Research Dr. Erin E. Chambers Waters Corporation Presented
Kidney. Heart. Lung. Sirt1. Gapdh. Mouse IgG DAPI. Rabbit IgG DAPI
 a e Na V 1.5 Ad-LacZ Ad- 110KD b Scn5a/ (relative to Ad-LacZ) f 150 100 50 0 p = 0.65 Ad-LacZ Ad- c Heart Lung Kidney Spleen 110KD d fl/fl c -/- DAPI 20 µm Na v 1.5 250KD fl/fl Rabbit IgG DAPI fl/fl Mouse
a e Na V 1.5 Ad-LacZ Ad- 110KD b Scn5a/ (relative to Ad-LacZ) f 150 100 50 0 p = 0.65 Ad-LacZ Ad- c Heart Lung Kidney Spleen 110KD d fl/fl c -/- DAPI 20 µm Na v 1.5 250KD fl/fl Rabbit IgG DAPI fl/fl Mouse
New Developments in LC-IMS-MS Proteomic Measurements and Informatic Analyses
 New Developments in LC-IMS-MS Proteomic Measurements and Informatic Analyses Erin Shammel Baker Kristin E. Burnum-Johnson, Xing Zhang, Cameron P. Casey, Yehia M. Ibrahim, Matthew E. Monroe, Tao Liu, Brendan
New Developments in LC-IMS-MS Proteomic Measurements and Informatic Analyses Erin Shammel Baker Kristin E. Burnum-Johnson, Xing Zhang, Cameron P. Casey, Yehia M. Ibrahim, Matthew E. Monroe, Tao Liu, Brendan
Glycosylation analysis of blood plasma proteins
 Glycosylation analysis of blood plasma proteins Thesis booklet Eszter Tóth Doctoral School of Pharmaceutical Sciences Semmelweis University Supervisor: Károly Vékey DSc Official reviewers: Borbála Dalmadiné
Glycosylation analysis of blood plasma proteins Thesis booklet Eszter Tóth Doctoral School of Pharmaceutical Sciences Semmelweis University Supervisor: Károly Vékey DSc Official reviewers: Borbála Dalmadiné
MS/MS Library Creation of Q-TOF LC/MS Data for MassHunter PCDL Manager
 MS/MS Library Creation of Q-TOF LC/MS Data for MassHunter PCDL Manager Quick Start Guide Step 1. Calibrate the Q-TOF LC/MS for low m/z ratios 2 Step 2. Set up a Flow Injection Analysis (FIA) method for
MS/MS Library Creation of Q-TOF LC/MS Data for MassHunter PCDL Manager Quick Start Guide Step 1. Calibrate the Q-TOF LC/MS for low m/z ratios 2 Step 2. Set up a Flow Injection Analysis (FIA) method for
Universal sample preparation method for proteome analysis
 nature methods Universal sample preparation method for proteome analysis Jacek R Wi niewski, Alexandre Zougman, Nagarjuna Nagaraj & Matthias Mann Supplementary figures and text: Supplementary Figure 1
nature methods Universal sample preparation method for proteome analysis Jacek R Wi niewski, Alexandre Zougman, Nagarjuna Nagaraj & Matthias Mann Supplementary figures and text: Supplementary Figure 1
The use of mass spectrometry in lipidomics. Outlines
 The use of mass spectrometry in lipidomics Jeevan Prasain jprasain@uab.edu 6-2612 utlines Brief introduction to lipidomics Analytical methodology: MS/MS structure elucidation of phospholipids Phospholipid
The use of mass spectrometry in lipidomics Jeevan Prasain jprasain@uab.edu 6-2612 utlines Brief introduction to lipidomics Analytical methodology: MS/MS structure elucidation of phospholipids Phospholipid
Profiling the Distribution of N-Glycosylation in Therapeutic Antibodies using the QTRAP 6500 System
 Profiling the Distribution of N-Glycosylation in Therapeutic Antibodies using the QTRAP 6500 System Scheduled MRM Pro Algorithm for Increased Efficiency of Targeted Detection Jenny Albanese 1, Christie
Profiling the Distribution of N-Glycosylation in Therapeutic Antibodies using the QTRAP 6500 System Scheduled MRM Pro Algorithm for Increased Efficiency of Targeted Detection Jenny Albanese 1, Christie
SUPPLEMENTAL MATERIAL
 1 SUPPLEMENTAL MATERIAL Response time and signal detection time distributions SM Fig. 1. Correct response time (thick solid green curve) and error response time densities (dashed red curve), averaged across
1 SUPPLEMENTAL MATERIAL Response time and signal detection time distributions SM Fig. 1. Correct response time (thick solid green curve) and error response time densities (dashed red curve), averaged across
New Instruments and Services
 New Instruments and Services Liwen Zhang Mass Spectrometry and Proteomics Facility The Ohio State University Summer Workshop 2016 Thermo Orbitrap Fusion http://planetorbitrap.com/orbitrap fusion Thermo
New Instruments and Services Liwen Zhang Mass Spectrometry and Proteomics Facility The Ohio State University Summer Workshop 2016 Thermo Orbitrap Fusion http://planetorbitrap.com/orbitrap fusion Thermo
New Mass Spectrometry Tools to Transform Metabolomics and Lipidomics
 New Mass Spectrometry Tools to Transform Metabolomics and Lipidomics July.3.13 Ken Miller Vice President of Marketing, Life Sciences Mass Spectrometry 1 The world leader in serving science Omics & the
New Mass Spectrometry Tools to Transform Metabolomics and Lipidomics July.3.13 Ken Miller Vice President of Marketing, Life Sciences Mass Spectrometry 1 The world leader in serving science Omics & the
For reprints and correspondence:
 APPLICATION REPORT SEPTEMBER 30, 2015 New Proteomic Workflows Combine Albumin Depletion and On- Bead Digestion, for Quantitative Cancer Serum Haiyan Zheng 1 ; Caifeng Zhao 1 ; Meiqian Qian 1 ; Swapan Roy
APPLICATION REPORT SEPTEMBER 30, 2015 New Proteomic Workflows Combine Albumin Depletion and On- Bead Digestion, for Quantitative Cancer Serum Haiyan Zheng 1 ; Caifeng Zhao 1 ; Meiqian Qian 1 ; Swapan Roy
Sequence Identification And Spatial Distribution of Rat Brain Tryptic Peptides Using MALDI Mass Spectrometric Imaging
 Sequence Identification And Spatial Distribution of Rat Brain Tryptic Peptides Using MALDI Mass Spectrometric Imaging AB SCIEX MALDI TOF/TOF* Systems Patrick Pribil AB SCIEX, Canada MALDI mass spectrometric
Sequence Identification And Spatial Distribution of Rat Brain Tryptic Peptides Using MALDI Mass Spectrometric Imaging AB SCIEX MALDI TOF/TOF* Systems Patrick Pribil AB SCIEX, Canada MALDI mass spectrometric
LECTURE-15. itraq Clinical Applications HANDOUT. Isobaric Tagging for Relative and Absolute quantitation (itraq) is a quantitative MS
 LECTURE-15 itraq Clinical Applications HANDOUT PREAMBLE Isobaric Tagging for Relative and Absolute quantitation (itraq) is a quantitative MS based method for quantifying proteins subject to various different
LECTURE-15 itraq Clinical Applications HANDOUT PREAMBLE Isobaric Tagging for Relative and Absolute quantitation (itraq) is a quantitative MS based method for quantifying proteins subject to various different
Mass Spectrometry. Mass spectrometer MALDI-TOF ESI/MS/MS. Basic components. Ionization source Mass analyzer Detector
 Mass Spectrometry MALDI-TOF ESI/MS/MS Mass spectrometer Basic components Ionization source Mass analyzer Detector 1 Principles of Mass Spectrometry Proteins are separated by mass to charge ratio (limit
Mass Spectrometry MALDI-TOF ESI/MS/MS Mass spectrometer Basic components Ionization source Mass analyzer Detector 1 Principles of Mass Spectrometry Proteins are separated by mass to charge ratio (limit
Journal of Chemical and Pharmaceutical Research
 Available on line www.jocpr.com Journal of Chemical and Pharmaceutical Research ISSN No: 0975-7384 CODEN(USA): JCPRC5 J. Chem. Pharm. Res., 2011, 3(1):138-144 Simultaneous RP HPLC determination of Latanoprost
Available on line www.jocpr.com Journal of Chemical and Pharmaceutical Research ISSN No: 0975-7384 CODEN(USA): JCPRC5 J. Chem. Pharm. Res., 2011, 3(1):138-144 Simultaneous RP HPLC determination of Latanoprost
Databehandling. 3. Mark e.g. the first fraction (1: 0-45 min, 2: min, 3; min, 4: min, 5: min, 6: min).
 Databehandling Data analysis 1. Choose Open in the Data analysis window. 2. Press the Open folder and choose the desired analysis. Click the + button, so that the Chromatograms line appears. Click the
Databehandling Data analysis 1. Choose Open in the Data analysis window. 2. Press the Open folder and choose the desired analysis. Click the + button, so that the Chromatograms line appears. Click the
A computational framework for discovery of glycoproteomic biomarkers
 A computational framework for discovery of glycoproteomic biomarkers Haixu Tang, Anoop Mayampurath, Chuan-Yih Yu Indiana University, Bloomington Yehia Mechref, Erwang Song Texas Tech University 1 Goal:
A computational framework for discovery of glycoproteomic biomarkers Haixu Tang, Anoop Mayampurath, Chuan-Yih Yu Indiana University, Bloomington Yehia Mechref, Erwang Song Texas Tech University 1 Goal:
SCIEX Vitamin D 200M Assay for the Topaz System
 The First FDA-Cleared LC-MS/MS Assay for Vitamin D SCIEX Vitamin D 200M Assay for the Topaz System The first FDA-cleared LC-MS/MS assay for Vitamin D Vitamin D is an important building block for human
The First FDA-Cleared LC-MS/MS Assay for Vitamin D SCIEX Vitamin D 200M Assay for the Topaz System The first FDA-cleared LC-MS/MS assay for Vitamin D Vitamin D is an important building block for human
CHAPTER 6 FUNCTIONAL PROPERTIES OF PROTEIN HYDROLYSATES
 68 CHAPTER 6 FUNCTIONAL PROPERTIES OF PROTEIN HYDROLYSATES 6.1 INTRODUCTION Functional properties can be defined as the overall physicochemical properties of proteins in food systems during processing,
68 CHAPTER 6 FUNCTIONAL PROPERTIES OF PROTEIN HYDROLYSATES 6.1 INTRODUCTION Functional properties can be defined as the overall physicochemical properties of proteins in food systems during processing,
Advances in Hybrid Mass Spectrometry
 The world leader in serving science Advances in Hybrid Mass Spectrometry ESAC 2008 Claire Dauly Field Marketing Specialist, Proteomics New hybrids instruments LTQ Orbitrap XL with ETD MALDI LTQ Orbitrap
The world leader in serving science Advances in Hybrid Mass Spectrometry ESAC 2008 Claire Dauly Field Marketing Specialist, Proteomics New hybrids instruments LTQ Orbitrap XL with ETD MALDI LTQ Orbitrap
VaTx1 VaTx2 VaTx3. VaTx min Retention Time (min) Retention Time (min)
 a Absorbance (mau) 5 2 5 3 4 5 6 7 8 9 6 2 3 4 5 6 VaTx2 High Ca 2+ Low Ca 2+ b 38.2 min Absorbance (mau) 3 2 3 4 5 3 2 VaTx2 39.3 min 3 4 5 3 2 4. min 3 4 5 Supplementary Figure. Toxin Purification For
a Absorbance (mau) 5 2 5 3 4 5 6 7 8 9 6 2 3 4 5 6 VaTx2 High Ca 2+ Low Ca 2+ b 38.2 min Absorbance (mau) 3 2 3 4 5 3 2 VaTx2 39.3 min 3 4 5 3 2 4. min 3 4 5 Supplementary Figure. Toxin Purification For
Integrated Targeted Quantitation Method for Insulin and its Therapeutic Analogs
 Integrated Targeted Quantitation Method for Insulin and its Therapeutic Analogs Eric Niederkofler, 1 Dobrin Nedelkov, 1 Urban Kiernan, 1 David Phillips, 1 Kemmons Tubbs, 1 Scott Peterman, 2 Bryan Krastins,
Integrated Targeted Quantitation Method for Insulin and its Therapeutic Analogs Eric Niederkofler, 1 Dobrin Nedelkov, 1 Urban Kiernan, 1 David Phillips, 1 Kemmons Tubbs, 1 Scott Peterman, 2 Bryan Krastins,
Latest Innovations in LC/MS/MS from Waters for Metabolism and Bioanalytical Applications
 Latest Innovations in LC/MS/MS from Waters for Metabolism and Bioanalytical Applications Ignatius J. Kass Senior Field Marketing Manager Pharmaceutical MS Challenges in Pharmaceutical Sample Analysis Quantitative
Latest Innovations in LC/MS/MS from Waters for Metabolism and Bioanalytical Applications Ignatius J. Kass Senior Field Marketing Manager Pharmaceutical MS Challenges in Pharmaceutical Sample Analysis Quantitative
Determination of red blood cell fatty acid profiles in clinical research
 Application Note Clinical Research Determination of red blood cell fatty acid profiles in clinical research Chemical ionization gas chromatography tandem mass spectrometry Authors Yvonne Schober 1, Hans
Application Note Clinical Research Determination of red blood cell fatty acid profiles in clinical research Chemical ionization gas chromatography tandem mass spectrometry Authors Yvonne Schober 1, Hans
Supporting Information
 Supporting Information Development of a High Coverage Pseudotargeted Lipidomics Method Based on Ultra-High Performance Liquid Chromatography-Mass Spectrometry Qiuhui Xuan 1,2#, Chunxiu Hu 1#, Di Yu 1,2,
Supporting Information Development of a High Coverage Pseudotargeted Lipidomics Method Based on Ultra-High Performance Liquid Chromatography-Mass Spectrometry Qiuhui Xuan 1,2#, Chunxiu Hu 1#, Di Yu 1,2,
BIOCHEMISTRY I HOMEWORK III DUE 10/15/03 66 points total + 2 bonus points = 68 points possible Swiss-PDB Viewer Exercise Attached
 BIOCHEMISTRY I HOMEWORK III DUE 10/15/03 66 points total + 2 bonus points = 68 points possible Swiss-PDB Viewer Exercise Attached 1). 20 points total T or F (2 points each; if false, briefly state why
BIOCHEMISTRY I HOMEWORK III DUE 10/15/03 66 points total + 2 bonus points = 68 points possible Swiss-PDB Viewer Exercise Attached 1). 20 points total T or F (2 points each; if false, briefly state why
Noninvasive Blood Glucose Analysis using Near Infrared Absorption Spectroscopy. Abstract
 Progress Report No. 2-3, March 31, 1999 The Home Automation and Healthcare Consortium Noninvasive Blood Glucose Analysis using Near Infrared Absorption Spectroscopy Prof. Kamal Youcef-Toumi Principal Investigator
Progress Report No. 2-3, March 31, 1999 The Home Automation and Healthcare Consortium Noninvasive Blood Glucose Analysis using Near Infrared Absorption Spectroscopy Prof. Kamal Youcef-Toumi Principal Investigator
RNA-Seq Preparation Comparision Summary: Lexogen, Standard, NEB
 RNA-Seq Preparation Comparision Summary: Lexogen, Standard, NEB CSF-NGS January 22, 214 Contents 1 Introduction 1 2 Experimental Details 1 3 Results And Discussion 1 3.1 ERCC spike ins............................................
RNA-Seq Preparation Comparision Summary: Lexogen, Standard, NEB CSF-NGS January 22, 214 Contents 1 Introduction 1 2 Experimental Details 1 3 Results And Discussion 1 3.1 ERCC spike ins............................................
Achieve Broad Lipid Quantitation using a High-Throughput Targeted Lipidomics Method
 Achieve Broad Lipid Quantitation using a High-Throughput Targeted Lipidomics Method LC-Based Approach for Lipid Class Separation and Quantitation on QTRAP 6500+ System Mackenzie Pearson, Santosh Kumar
Achieve Broad Lipid Quantitation using a High-Throughput Targeted Lipidomics Method LC-Based Approach for Lipid Class Separation and Quantitation on QTRAP 6500+ System Mackenzie Pearson, Santosh Kumar
Development of a Bioanalytical Method for Quantification of Amyloid Beta Peptides in Cerebrospinal Fluid
 Development of a Bioanalytical Method for Quantification of Amyloid Beta Peptides in Cerebrospinal Fluid Joanne ( 乔安妮 ) Mather Senior Scientist Waters Corporation Data courtesy of Erin Chambers and Mary
Development of a Bioanalytical Method for Quantification of Amyloid Beta Peptides in Cerebrospinal Fluid Joanne ( 乔安妮 ) Mather Senior Scientist Waters Corporation Data courtesy of Erin Chambers and Mary
Statistics is the science of collecting, organizing, presenting, analyzing, and interpreting data to assist in making effective decisions
 Readings: OpenStax Textbook - Chapters 1 5 (online) Appendix D & E (online) Plous - Chapters 1, 5, 6, 13 (online) Introductory comments Describe how familiarity with statistical methods can - be associated
Readings: OpenStax Textbook - Chapters 1 5 (online) Appendix D & E (online) Plous - Chapters 1, 5, 6, 13 (online) Introductory comments Describe how familiarity with statistical methods can - be associated
Improving Selectivity in Quantitative Analysis Using MS 3 on a Hybrid Quadrupole-Linear Ion Trap Mass Spectrometer
 Improving Selectivity in Quantitative Analysis Using MS 3 on a Hybrid Quadrupole-Linear Ion Trap Mass Spectrometer Overview Increased selectivity is achieved in quantitative analysis by using MS 3. The
Improving Selectivity in Quantitative Analysis Using MS 3 on a Hybrid Quadrupole-Linear Ion Trap Mass Spectrometer Overview Increased selectivity is achieved in quantitative analysis by using MS 3. The
