SUPPLEMENTARY INFORMATION
|
|
|
- Noah Gregory
- 5 years ago
- Views:
Transcription
1 SPPLEMENTY INFOMTION 2 =.75 SHPE + 1 (eplicate 2) SHPE + 1 (eplicate 1) Figure S1. eproducibility of SHPE measurements between biological replicates. eactivities corresponding to extension reactions performed with primer 9 are shown plotted on a logarithmic scale. eplicates were performed on independent HIV-1 N genome preparations, by different individuals (J.M.W. and.w.l.), roughly one year apart. 1
2 SPPLEMENTY INFOMTION ount (rel. units) a SHPE reactivity b 2.69 ±.61 (48).6 ±.9 (41) SHPE reactivity SHPE reactivity 1 unpaired internal pairs Sovent accessibility (Å 2 ) Figure S2. SHPE reactivities are strongly sensitive to N secondary structure, but not to solvent accessibility. These SHPE data are from the Nase P specificity domain N 1,11, which has a compact structure with significant, tightly packed, tertiary interactions 12. Solvent accessibility was calculated using a 1.4 Å radius probe for the ribose 2'-oxygen atom. (a) SHPE reactivities do not correlate with solvent accessibility ( 2 =.4). Small panels at top give SHPE reactivity distributions for nucleotides whose solvent accessibility are low, med or high (in red, orange and green, respectively). Distributions are similar and all regions contain nucleotides with both high and low reactivities. (b) SHPE reactivities strongly discriminate between unpaired and base paired nucleotides. Box plot representations are shown. Numerical values give the mean ± standard deviation; the number of measurements is in parentheses. Internal base pairs are defined as positions that are paired as visualized crystallographically and are adjacent to other canonically paired nucleotides 11. The strong predictive relationship between SHPE reactivity and secondary structure is consistent with benchmarks showing SHPE measures local disorder in N
3 SPPLEMENTY INFOMTION ccessibility of splice acceptor sites consensus SHPE reactivity N Splice acceptor S2 S3a S4 S4c S4a S4b S5 S7a S7b S7 S8 Position in genome Mean SHPE reactivity arely used Figure S3. SHPE reactivities at splice acceptor sites in the NL4-3 genome. Histograms show SHPE reactivities at each splice acceptor site across the five nucleotide consensus motif. The mean SHPE reactivity at each acceptor is colored according to the scale used throughout the manuscript. The mean SHPE reactivity across all five nucleotide windows in the HIV genome is.34; thus, all splice acceptors have high SHPE reactivities, except the three (underlined) characterized as rarely used sites
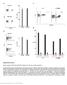
4 SPPLEMENTY INFOMTION a inter-protein linkers mean reactivity Number of bootstrapped data sets (out of 1,) b 2 p = protein domain junctions mean reactivity p = Mean SHPE eactivity Figure S4. Histograms comparing mean SHPE reactivities for inter-protein linker regions and protein domain junctions with distributions of equivalent-length sequences obtained from random regions in the HIV genome by a bootstrap statistical analysis. p-values give the probability that the low SHPE reactivities in the collection of genome elements occurred by chance. 4
5 SPPLEMENTY INFOMTION.6 M- yp loop -N N-p6 SHPE reactivity NL4-3 position Toeprint intnesity (au) ~ ~ ~ Figure S5. Distribution of ribosome pause sites in ag at the M- and -N junctions. Pause sites were identified by inhibition of reverse transcriptase-mediated primer extension, after first inhibiting ribosome processivity with cycloheximide. (Top panel) Median SHPE reactivities in ag over a 75 nt widow, reproduced from Fig. 1 in the main text, are shown in blue. ed line shows median SHPE reactivity over entire genome. Domain junctions and the cyclophilin loop are indicated explicitly. ray bars indicate regions scanned in the toeprinting experiment; the strongest pause sites are indicated with vertical bars. (Lower panels) Toeprinting intensity as a function of genome position. 5
6 SPPLEMENTY INFOMTION a P3 slippery 1625 sequence P2 159 frameshift stimulatory stem 158 P NL4-3 b M Y P3 M Y Y N Y Y Y Y P2 M Y Y M Y M Y P1 Y Y Y M K N group M consensus or or or or ny Base pair conservation 1% >9% <7% Figure S6. Structure of the HIV-1 gag-pol frameshift element. (a) SHPE-constrained secondary structure. Nucleotides are colored according to their SHPE reactivities using the scale shown in Fig. 3b. (b) Sequence and structural conservation for the 3-helix junction model across 37 HIV-1 group M reference sequences. 6
7 ' T 5' 5' poly signal PBS DIS SL2 PSI gag-pol frameshift PPT B D E F H I J K L M N O P Q S T V W X Structure of the NL4-3 HIV-1 N enome (5' Half) pol (IN) end vpr start gag () start gag (M) end gag (M) start gag (p2) start B D gag () end E gag (p2) end gag (N) start F gag (N) end gag (p1) start TF peptide start H I J gag (p1) end gag (p6) start K L TF peptide end pol (P) start M N gag (p6) end O pol (P) end pol (T) start P Q pol (T) end pol (Nase) start S pol (Nase) end pol (IN) start T vif start V W X Protein oding egions to 3' half m1 5' 3' tn 3 Lys 5' 3' SHPE eactivity not analyzed (53 nts) Figure S7. Detailed secondary structure for the NL4-3 HIV-1 genome, including nucleotide identities. Divided into two panels. doi: 1.138/nature8237 SPPLEMENTY INFOMTION 7
8 Structure of the NL4-3 HIV-1 N enome (3' Half) () n E I V IV III IIc IIb IIa ev B.S. 3' T 3' poly signal -3' PPT Ee Ff g Hh Ii Jj Kk Ll Mm Nn Oo Y Z a Bb c Dd 76 vif end tat exon1 start vpr end nef end Y Z a tat/rev exon 1 end rev exon 1 start vpu start Bb c Dd env signal peptide start env signal peptide end env (gp12) start env (gp41) start env (gp12) end Ee Ff g Hh Ii tat/rev exon 2 start rev end env end tat end nef start Jj Kk Ll Mm Nn Oo Protein oding egions to 5' half doi: 1.138/nature8237 SPPLEMENTY INFOMTION doi: 1.138/nature8237 SPPLEMENTY INFOMTION 8
9 SPPLEMENTY INFOMTION Table S1: NL4-3 SHPE primer sequences Number Primer Sequence Binding Site in the NL4-3 enome 1 TTTTT TTTTTTT TTTTTTTTTT TTTTTTTT TTTTTT TTTTTTTTT TTTTTTT TTTT TTTTTTTTTTTT TTTTTTTTT TTTT TTT TTTTTTTTTT TTTTTT TTTTTTT TTTT TTTTTTT TTTTTTTT TTTTT TTTTTTTT T TTTTTTTTTT TTTTTTT TTTTTT TTTTTT TTTTT TTTTTTT TTTTT TTTTTTTT TT TTTTTTTTTTTTTTTTTTTTTT / poly() 9
10 SPPLEMENTY INFOMTION Table S2: Protein Domains in Figure 1d Protein PDB (ref.) Protein esidues olor M/ 2OL Blue reen ed Yellow Pink Yellow (-terminal) ray P 3PHV Blue T/Nase H (p66) 1HMV ed Yellow Pink Yellow ray reen Magenta IN 1WJ ed Yellow 1BIS Pink Yellow 1QM Magenta gp Blue 18-2 Yellow Blue gp41 1SZT , ed 1
11 SPPLEMENTY INFOMTION Supplementary eferences 1. Kelly, B.N. et al. Implications for viral capsid assembly from crystal structures of HIV-1 ag(1-278) and (N)( ). Biochemistry 45, (26). 2. Worthylake, D.K., Wang, H., Yoo, S., Sundquist, W.I. & Hill,.P. Structures of the HIV-1 capsid protein dimerization domain at 2.6 Å resolution. cta rystallogr. D Biol. rystallogr. 55, (1999). 3. Lapatto,., Blundell, T., Hemmings,., Overington, J., Wilderspin,., Wood, S., Merson, J.., Whittle, P.J., Danley, D.E., eoghegan, K.F., et al. X-ray analysis of HIV- 1 proteinase at 2.7 Å resolution confirms structural homology among retroviral enzymes. Nature 342, (1989). 4. odgers, D.W. et al. The structure of unliganded reverse transcriptase from the human immunodeficiency virus type 1. Proc. Natl. cad. Sci. S 92, (1995). 5. ai, M. et al. Solution structure of the N-terminal zinc binding domain of HIV-1 integrase. Nat. Struct. Biol. 4, (1997). 6. oldgur, Y. et al. Three new structures of the core domain of HIV-1 integrase: an active site that binds magnesium. Proc. Natl. cad. Sci. S 95, (1998). 7. Eijkelenboom,.P. et al. The DN-binding domain of HIV-1 integrase has an SH3-like fold. Nat. Struct. Biol. 2, (1995). 8. Kwong, P.D. et al. Structure of an HIV gp12 envelope glycoprotein in complex with the D4 receptor and a neutralizing human antibody. Nature 393, (1998). 9. Tan, K., Liu, J., Wang, J., Shen, S. & Lu, M. tomic structure of a thermostable subdomain of HIV-1 gp41. Proc. Natl. cad. Sci. S 94, (1997). 1. Mortimer, S.. & Weeks, K.M. fast-acting reagent for accurate analysis of N secondary and tertiary structure by SHPE chemistry. J. m. hem. Soc. 129, (27). 11. Wilkinson, K.. et al. Influence of nucleotide identity on ribose 2'-hydroxyl reactivity in N. N 15, in press (29). 12. Krasilnikov,.S., Yang, X., Pan, T. & Mondragon,. rystal structure of the specificity domain of ribonuclease P. Nature 421, (23). 13. herghe,.m., Shajani, Z., Wilkinson, K.., Varani,. & Weeks, K.M. Strong correlation between SHPE chemistry and the generalized NM order parameter (S 2 ) in N. J. m. hem. Soc. 13, (28). 14. Purcell, D.F. & Martin, M.. lternative splicing of human immunodeficiency virus type 1 mn modulates viral protein expression, replication, and infectivity. J. Virol. 67, (1993). 11
Architecture and secondary structure of an entire HIV-1 RNA genome
 Vol 46 6 ugust 29 doi:1.138/nature8237 RTILES rchitecture and secondary structure of an entire HIV-1 RN genome Joseph M. Watts 1, Kristen K. Dang 2, Robert J. orelick 5, hristopher W. Leonard 1, Julian
Vol 46 6 ugust 29 doi:1.138/nature8237 RTILES rchitecture and secondary structure of an entire HIV-1 RN genome Joseph M. Watts 1, Kristen K. Dang 2, Robert J. orelick 5, hristopher W. Leonard 1, Julian
Retroviral RNA Processing and stability
 Retroviral RN Processing and stability m 7 gag pol env src Karen Beemon Johns Hopkins niversity m 7 env src m 7 src Retroviruses hijack host cell gene expression machinery to generate progeny virions Simple
Retroviral RN Processing and stability m 7 gag pol env src Karen Beemon Johns Hopkins niversity m 7 env src m 7 src Retroviruses hijack host cell gene expression machinery to generate progeny virions Simple
L I F E S C I E N C E S
 1a L I F E S C I E N C E S 5 -UUA AUA UUC GAA AGC UGC AUC GAA AAC UGU GAA UCA-3 5 -TTA ATA TTC GAA AGC TGC ATC GAA AAC TGT GAA TCA-3 3 -AAT TAT AAG CTT TCG ACG TAG CTT TTG ACA CTT AGT-5 OCTOBER 31, 2006
1a L I F E S C I E N C E S 5 -UUA AUA UUC GAA AGC UGC AUC GAA AAC UGU GAA UCA-3 5 -TTA ATA TTC GAA AGC TGC ATC GAA AAC TGT GAA TCA-3 3 -AAT TAT AAG CTT TCG ACG TAG CTT TTG ACA CTT AGT-5 OCTOBER 31, 2006
HOST-PATHOGEN CO-EVOLUTION THROUGH HIV-1 WHOLE GENOME ANALYSIS
 HOST-PATHOGEN CO-EVOLUTION THROUGH HIV-1 WHOLE GENOME ANALYSIS Somda&a Sinha Indian Institute of Science, Education & Research Mohali, INDIA International Visiting Research Fellow, Peter Wall Institute
HOST-PATHOGEN CO-EVOLUTION THROUGH HIV-1 WHOLE GENOME ANALYSIS Somda&a Sinha Indian Institute of Science, Education & Research Mohali, INDIA International Visiting Research Fellow, Peter Wall Institute
Human immunodeficiency virus type 1 splicing at the major splice donor site is controlled by highly conserved RNA sequence and structural elements
 Journal of eneral Virology (2015), 96, 3389 3395 DOI 10.1099/jgv.0.000288 Short ommunication Human immunodeficiency virus type 1 splicing at the major splice donor site is controlled by highly conserved
Journal of eneral Virology (2015), 96, 3389 3395 DOI 10.1099/jgv.0.000288 Short ommunication Human immunodeficiency virus type 1 splicing at the major splice donor site is controlled by highly conserved
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10913 Supplementary Figure 1 2F o -F c electron density maps of cognate and near-cognate trna Leu 2 in the A site of the 70S ribosome. The maps are contoured at 1.2 sigma and some of
doi:10.1038/nature10913 Supplementary Figure 1 2F o -F c electron density maps of cognate and near-cognate trna Leu 2 in the A site of the 70S ribosome. The maps are contoured at 1.2 sigma and some of
HIV INFECTION: An Overview
 HIV INFECTION: An Overview UNIVERSITY OF PAPUA NEW GUINEA SCHOOL OF MEDICINE AND HEALTH SCIENCES DIVISION OF BASIC MEDICAL SCIENCES DISCIPLINE OF BIOCHEMISTRY & MOLECULAR BIOLOGY PBL MBBS II SEMINAR VJ
HIV INFECTION: An Overview UNIVERSITY OF PAPUA NEW GUINEA SCHOOL OF MEDICINE AND HEALTH SCIENCES DIVISION OF BASIC MEDICAL SCIENCES DISCIPLINE OF BIOCHEMISTRY & MOLECULAR BIOLOGY PBL MBBS II SEMINAR VJ
Prokaryotic Biology. VIRAL STDs, HIV-1 AND AIDS
 Prokaryotic Biology VIRAL STDs, HIV-1 AND AIDS Prokaryotic Biology FROM THE CDC VIRAL STDs, HIV-1 AND AIDS VIRAL STDs & CONTACT VIRAL DISEASES A. GENITAL HERPES & COLD SORES 1. HERPES SIMPLEX VIRUS-2 (HHV-2)
Prokaryotic Biology VIRAL STDs, HIV-1 AND AIDS Prokaryotic Biology FROM THE CDC VIRAL STDs, HIV-1 AND AIDS VIRAL STDs & CONTACT VIRAL DISEASES A. GENITAL HERPES & COLD SORES 1. HERPES SIMPLEX VIRUS-2 (HHV-2)
HIV & AIDS: Overview
 HIV & AIDS: Overview UNIVERSITY OF PAPUA NEW GUINEA SCHOOL OF MEDICINE AND HEALTH SCIENCES DIVISION OF BASIC MEDICAL SCIENCES DISCIPLINE OF BIOCHEMISTRY & MOLECULAR BIOLOGY PBL SEMINAR VJ TEMPLE 1 What
HIV & AIDS: Overview UNIVERSITY OF PAPUA NEW GUINEA SCHOOL OF MEDICINE AND HEALTH SCIENCES DIVISION OF BASIC MEDICAL SCIENCES DISCIPLINE OF BIOCHEMISTRY & MOLECULAR BIOLOGY PBL SEMINAR VJ TEMPLE 1 What
Panther has new prey
 Raising the Bar for Performance Testing Panther has new prey The Aptima HIV-1 Quant Dx assay leads the hunt for HIV-1 diagnosis and viral load monitoring. Freedom to work the way you choose Run what assays
Raising the Bar for Performance Testing Panther has new prey The Aptima HIV-1 Quant Dx assay leads the hunt for HIV-1 diagnosis and viral load monitoring. Freedom to work the way you choose Run what assays
Sequence Analysis of Human Immunodeficiency Virus Type 1
 Sequence Analysis of Human Immunodeficiency Virus Type 1 Stephanie Lucas 1,2 Mentor: Panayiotis V. Benos 1,3 With help from: David L. Corcoran 4 1 Bioengineering and Bioinformatics Summer Institute, Department
Sequence Analysis of Human Immunodeficiency Virus Type 1 Stephanie Lucas 1,2 Mentor: Panayiotis V. Benos 1,3 With help from: David L. Corcoran 4 1 Bioengineering and Bioinformatics Summer Institute, Department
Citation for published version (APA): Von Eije, K. J. (2009). RNAi based gene therapy for HIV-1, from bench to bedside
 UvA-DARE (Digital Academic Repository) RNAi based gene therapy for HIV-1, from bench to bedside Von Eije, K.J. Link to publication Citation for published version (APA): Von Eije, K. J. (2009). RNAi based
UvA-DARE (Digital Academic Repository) RNAi based gene therapy for HIV-1, from bench to bedside Von Eije, K.J. Link to publication Citation for published version (APA): Von Eije, K. J. (2009). RNAi based
Retroviruses. ---The name retrovirus comes from the enzyme, reverse transcriptase.
 Retroviruses ---The name retrovirus comes from the enzyme, reverse transcriptase. ---Reverse transcriptase (RT) converts the RNA genome present in the virus particle into DNA. ---RT discovered in 1970.
Retroviruses ---The name retrovirus comes from the enzyme, reverse transcriptase. ---Reverse transcriptase (RT) converts the RNA genome present in the virus particle into DNA. ---RT discovered in 1970.
Mina John Institute for Immunology and Infectious Diseases Royal Perth Hospital & Murdoch University Perth, Australia
 Mina John Institute for Immunology and Infectious Diseases Royal Perth Hospital & Murdoch University Perth, Australia AIDSvaccine conference, 14 th September 2011 IMGT HLA database July 2011 >5000 class
Mina John Institute for Immunology and Infectious Diseases Royal Perth Hospital & Murdoch University Perth, Australia AIDSvaccine conference, 14 th September 2011 IMGT HLA database July 2011 >5000 class
Host Double Strand Break Repair Generates HIV-1 Strains Resistant to CRISPR/Cas9
 Host Double Strand Break Repair Generates HIV-1 Strains Resistant to CRISPR/Cas9 Kristine E. Yoder, a * and Ralf Bundschuh b a Department of Molecular Virology, Immunology and Medical Genetics, Center
Host Double Strand Break Repair Generates HIV-1 Strains Resistant to CRISPR/Cas9 Kristine E. Yoder, a * and Ralf Bundschuh b a Department of Molecular Virology, Immunology and Medical Genetics, Center
Going Nowhere Fast: Lentivirus genetic sequence evolution does not correlate with phenotypic evolution.
 Going Nowhere Fast: Lentivirus genetic sequence evolution does not correlate with phenotypic evolution. Brian T. Foley, PhD btf@lanl.gov HIV Genetic Sequences, Immunology, Drug Resistance and Vaccine Trials
Going Nowhere Fast: Lentivirus genetic sequence evolution does not correlate with phenotypic evolution. Brian T. Foley, PhD btf@lanl.gov HIV Genetic Sequences, Immunology, Drug Resistance and Vaccine Trials
How HIV Causes Disease Prof. Bruce D. Walker
 How HIV Causes Disease Howard Hughes Medical Institute Massachusetts General Hospital Harvard Medical School 1 The global AIDS crisis 60 million infections 20 million deaths 2 3 The screen versions of
How HIV Causes Disease Howard Hughes Medical Institute Massachusetts General Hospital Harvard Medical School 1 The global AIDS crisis 60 million infections 20 million deaths 2 3 The screen versions of
Molecular Dynamics of HIV-1 Reverse Transcriptase
 Molecular Dynamics of HIV-1 Reverse Transcriptase Abderrahmane Benghanem Rensselaer Polytechnic Institute,Troy, NY Mentor: Dr. Maria Kurnikova Carnegie Mellon, Pittsburgh PA Outline HIV-1 Reverse Transcriptase
Molecular Dynamics of HIV-1 Reverse Transcriptase Abderrahmane Benghanem Rensselaer Polytechnic Institute,Troy, NY Mentor: Dr. Maria Kurnikova Carnegie Mellon, Pittsburgh PA Outline HIV-1 Reverse Transcriptase
Under the Radar Screen: How Bugs Trick Our Immune Defenses
 Under the Radar Screen: How Bugs Trick Our Immune Defenses Session 7: Cytokines Marie-Eve Paquet and Gijsbert Grotenbreg Whitehead Institute for Biomedical Research HHV-8 Discovered in the 1980 s at the
Under the Radar Screen: How Bugs Trick Our Immune Defenses Session 7: Cytokines Marie-Eve Paquet and Gijsbert Grotenbreg Whitehead Institute for Biomedical Research HHV-8 Discovered in the 1980 s at the
Fayth K. Yoshimura, Ph.D. September 7, of 7 RETROVIRUSES. 2. HTLV-II causes hairy T-cell leukemia
 1 of 7 I. Diseases Caused by Retroviruses RETROVIRUSES A. Human retroviruses that cause cancers 1. HTLV-I causes adult T-cell leukemia and tropical spastic paraparesis 2. HTLV-II causes hairy T-cell leukemia
1 of 7 I. Diseases Caused by Retroviruses RETROVIRUSES A. Human retroviruses that cause cancers 1. HTLV-I causes adult T-cell leukemia and tropical spastic paraparesis 2. HTLV-II causes hairy T-cell leukemia
HIV 101: Fundamentals of HIV Infection
 HIV 101: Fundamentals of HIV Infection David H. Spach, MD Professor of Medicine University of Washington Seattle, Washington Learning Objectives After attending this presentation, learners will be able
HIV 101: Fundamentals of HIV Infection David H. Spach, MD Professor of Medicine University of Washington Seattle, Washington Learning Objectives After attending this presentation, learners will be able
Fig. 1: Schematic diagram of basic structure of HIV
 UNIVERSITY OF PAPUA NEW GUINEA SCHOOL OF MEDICINE AND HEALTH SCIENCES DIVISION OF BASIC MEDICAL SCIENCES DISCIPLINE OF BIOCHEMISTRY & MOLECULAR BIOLOGY PBL SEMINAR HIV & AIDS: An Overview What is HIV?
UNIVERSITY OF PAPUA NEW GUINEA SCHOOL OF MEDICINE AND HEALTH SCIENCES DIVISION OF BASIC MEDICAL SCIENCES DISCIPLINE OF BIOCHEMISTRY & MOLECULAR BIOLOGY PBL SEMINAR HIV & AIDS: An Overview What is HIV?
Some living things are made of ONE cell, and are called. Other organisms are composed of many cells, and are called. (SEE PAGE 6)
 Section: 1.1 Question of the Day: Name: Review of Old Information: N/A New Information: We tend to only think of animals as living. However, there is a great diversity of organisms that we consider living
Section: 1.1 Question of the Day: Name: Review of Old Information: N/A New Information: We tend to only think of animals as living. However, there is a great diversity of organisms that we consider living
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Design of isolated protein and RNC constructs, and homogeneity of purified RNCs. (a) Schematic depicting the design and nomenclature used for all the isolated proteins and RNCs used
Supplementary Figure 1 Design of isolated protein and RNC constructs, and homogeneity of purified RNCs. (a) Schematic depicting the design and nomenclature used for all the isolated proteins and RNCs used
List of Figures. List of Tables
 Supporting Information for: Signaling Domain of Sonic Hedgehog as Cannibalistic Calcium-Regulated Zinc-Peptidase Rocio Rebollido-Rios 1, Shyam Bandari 3, Christoph Wilms 1, Stanislav Jakuschev 1, Andrea
Supporting Information for: Signaling Domain of Sonic Hedgehog as Cannibalistic Calcium-Regulated Zinc-Peptidase Rocio Rebollido-Rios 1, Shyam Bandari 3, Christoph Wilms 1, Stanislav Jakuschev 1, Andrea
Human Immunodeficiency Virus
 Human Immunodeficiency Virus Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Viruses and hosts Lentivirus from Latin lentis (slow), for slow progression of disease
Human Immunodeficiency Virus Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Viruses and hosts Lentivirus from Latin lentis (slow), for slow progression of disease
CURRENT DEVELOMENTS AND FUTURE PROSPECTS FOR HIV GENE THERAPY USING INTERFERING RNA-BASED STRATEGIES
 [Frontiers in Bioscience 5, d527-555, May 1, 2000] CURRENT DEVELOMENTS AND FUTURE PROSPECTS FOR HIV GENE THERAPY USING INTERFERING RNA-BASED STRATEGIES Betty Lamothe, Sadhna Joshi Department of Medical
[Frontiers in Bioscience 5, d527-555, May 1, 2000] CURRENT DEVELOMENTS AND FUTURE PROSPECTS FOR HIV GENE THERAPY USING INTERFERING RNA-BASED STRATEGIES Betty Lamothe, Sadhna Joshi Department of Medical
HIV-1: Fifteen Proteins and an RNA
 Annu. Rev. Biochem. 1998. 67:1 25 Copyright c 1998 by Annual Reviews. All rights reserved HIV-1: Fifteen Proteins and an RNA Alan D. Frankel Department of Biochemistry and Biophysics, University of California,
Annu. Rev. Biochem. 1998. 67:1 25 Copyright c 1998 by Annual Reviews. All rights reserved HIV-1: Fifteen Proteins and an RNA Alan D. Frankel Department of Biochemistry and Biophysics, University of California,
HIV and drug resistance Simon Collins UK-CAB 1 May 2009
 HIV and drug resistance Simon Collins UK-CAB 1 May 2009 slides: thanks to Prof Clive Loveday, Intl. Clinical Virology Centre www.icvc.org.uk Tip of the iceberg = HIV result, CD4, VL Introduction: resistance
HIV and drug resistance Simon Collins UK-CAB 1 May 2009 slides: thanks to Prof Clive Loveday, Intl. Clinical Virology Centre www.icvc.org.uk Tip of the iceberg = HIV result, CD4, VL Introduction: resistance
Modeling the HIV-1 Intasome: A Prototype View of the Target of Integrase Inhibitors
 Viruses 2010, 2, 2777-2781; doi:10.3390/v2122777 OPEN ACCESS viruses ISSN 1999-4915 www.mdpi.com/journal/viruses Commentary Modeling the HIV-1 Intasome: A Prototype View of the Target of Integrase Inhibitors
Viruses 2010, 2, 2777-2781; doi:10.3390/v2122777 OPEN ACCESS viruses ISSN 1999-4915 www.mdpi.com/journal/viruses Commentary Modeling the HIV-1 Intasome: A Prototype View of the Target of Integrase Inhibitors
Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random
 S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
3. on T helper {cells / lymphocytes} ; 3. ACCEPT macrophages / dendritic cells / CD4 cells
 1(a) 1. (structure G is {glycoprotein / gp120} ; 2. used for {attachment / eq} to CD4 (molecules / receptors /antigens) ; 1. IGNORE gp 41 and gp 160 and other wrong numbers 3. on T helper {cells / lymphocytes}
1(a) 1. (structure G is {glycoprotein / gp120} ; 2. used for {attachment / eq} to CD4 (molecules / receptors /antigens) ; 1. IGNORE gp 41 and gp 160 and other wrong numbers 3. on T helper {cells / lymphocytes}
Fayth K. Yoshimura, Ph.D. September 7, of 7 HIV - BASIC PROPERTIES
 1 of 7 I. Viral Origin. A. Retrovirus - animal lentiviruses. HIV - BASIC PROPERTIES 1. HIV is a member of the Retrovirus family and more specifically it is a member of the Lentivirus genus of this family.
1 of 7 I. Viral Origin. A. Retrovirus - animal lentiviruses. HIV - BASIC PROPERTIES 1. HIV is a member of the Retrovirus family and more specifically it is a member of the Lentivirus genus of this family.
Biochemical and Biophysical Research Communications 305 (2003)
 Biochemical and Biophysical Research Communications 305 (2003) 322 326 BBRC www.elsevier.com/locate/ybbrc Comprehensive mutagenesis of HIV-1 protease: a computational geometry approach q Majid Masso and
Biochemical and Biophysical Research Communications 305 (2003) 322 326 BBRC www.elsevier.com/locate/ybbrc Comprehensive mutagenesis of HIV-1 protease: a computational geometry approach q Majid Masso and
Received 25 April 2002/Accepted 21 May 2002
 JOURNAL OF VIROLOGY, Sept. 2002, p. 8757 8768 Vol. 76, No. 17 0022-538X/02/$04.00 0 DOI: 10.1128/JVI.76.17.8757 8768.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved. Clustering
JOURNAL OF VIROLOGY, Sept. 2002, p. 8757 8768 Vol. 76, No. 17 0022-538X/02/$04.00 0 DOI: 10.1128/JVI.76.17.8757 8768.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved. Clustering
Section 6. Junaid Malek, M.D.
 Section 6 Junaid Malek, M.D. The Golgi and gp160 gp160 transported from ER to the Golgi in coated vesicles These coated vesicles fuse to the cis portion of the Golgi and deposit their cargo in the cisternae
Section 6 Junaid Malek, M.D. The Golgi and gp160 gp160 transported from ER to the Golgi in coated vesicles These coated vesicles fuse to the cis portion of the Golgi and deposit their cargo in the cisternae
HIV/AIDS. Biology of HIV. Research Feature. Related Links. See Also
 6/1/2011 Biology of HIV Biology of HIV HIV belongs to a class of viruses known as retroviruses. Retroviruses are viruses that contain RNA (ribonucleic acid) as their genetic material. After infecting a
6/1/2011 Biology of HIV Biology of HIV HIV belongs to a class of viruses known as retroviruses. Retroviruses are viruses that contain RNA (ribonucleic acid) as their genetic material. After infecting a
HIV Immunopathogenesis. Modeling the Immune System May 2, 2007
 HIV Immunopathogenesis Modeling the Immune System May 2, 2007 Question 1 : Explain how HIV infects the host Zafer Iscan Yuanjian Wang Zufferey Abhishek Garg How does HIV infect the host? HIV infection
HIV Immunopathogenesis Modeling the Immune System May 2, 2007 Question 1 : Explain how HIV infects the host Zafer Iscan Yuanjian Wang Zufferey Abhishek Garg How does HIV infect the host? HIV infection
HIV/AIDS: Molecular Biology and pathogenesis. George N. Pavlakis National Cancer Institute, USA
 HIV/AIDS: Molecular Biology and pathogenesis George N. Pavlakis National Cancer Institute, USA Infection RNA Export, packaging Virion formation RNA Reverse transcription DNA RNA Integration Transcription
HIV/AIDS: Molecular Biology and pathogenesis George N. Pavlakis National Cancer Institute, USA Infection RNA Export, packaging Virion formation RNA Reverse transcription DNA RNA Integration Transcription
Reliable Prediction of Viral RNA Structures
 Reliable Prediction of Viral RNA Structures Winterseminar 2016, Bled Roman Ochsenreiter TBI Wien Bled, 15.2.2016 Roman Ochsenreiter TBI Wien Reliable Prediction of Viral RNA Structures Bled, 15.2.2016
Reliable Prediction of Viral RNA Structures Winterseminar 2016, Bled Roman Ochsenreiter TBI Wien Bled, 15.2.2016 Roman Ochsenreiter TBI Wien Reliable Prediction of Viral RNA Structures Bled, 15.2.2016
Centers for Disease Control August 9, 2004
 HIV CDC site UNAIDS Aids Knowledge Base http://www.cdc.gov/hiv/dhap.htm http://hivinsite.ucsf.edu/insite.jsp?page=kb National Institute of Allergy and Infectious Diseases http://www.niaid.nih.gov/default.htm
HIV CDC site UNAIDS Aids Knowledge Base http://www.cdc.gov/hiv/dhap.htm http://hivinsite.ucsf.edu/insite.jsp?page=kb National Institute of Allergy and Infectious Diseases http://www.niaid.nih.gov/default.htm
Lentiviruses: HIV-1 Pathogenesis
 Lentiviruses: HIV-1 Pathogenesis Human Immunodeficiency Virus, HIV, computer graphic by Russell Kightley Tsafi Pe ery, Ph.D. Departments of Medicine and Biochemistry & Molecular Biology NJMS, UMDNJ. e-mail:
Lentiviruses: HIV-1 Pathogenesis Human Immunodeficiency Virus, HIV, computer graphic by Russell Kightley Tsafi Pe ery, Ph.D. Departments of Medicine and Biochemistry & Molecular Biology NJMS, UMDNJ. e-mail:
Introduction retroposon
 17.1 - Introduction A retrovirus is an RNA virus able to convert its sequence into DNA by reverse transcription A retroposon (retrotransposon) is a transposon that mobilizes via an RNA form; the DNA element
17.1 - Introduction A retrovirus is an RNA virus able to convert its sequence into DNA by reverse transcription A retroposon (retrotransposon) is a transposon that mobilizes via an RNA form; the DNA element
HIV-1 Dual Infection and Neurocognitive Impairment
 HIV-1 Dual Infection and Neurocognitive Impairment Gabriel Wagner, MD Assistant Professor of Medicine Infectious Diseases & Global Public Health UC San Diego HIV-Associated End Organ Damage Antiretroviral
HIV-1 Dual Infection and Neurocognitive Impairment Gabriel Wagner, MD Assistant Professor of Medicine Infectious Diseases & Global Public Health UC San Diego HIV-Associated End Organ Damage Antiretroviral
Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels. Supplementary Information
 Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels Craig T. Armstrong, Philip E. Mason, J. L. Ross Anderson and Christopher E. Dempsey
Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels Craig T. Armstrong, Philip E. Mason, J. L. Ross Anderson and Christopher E. Dempsey
Supplementary information. MARCH8 inhibits HIV-1 infection by reducing virion incorporation of envelope glycoproteins
 Supplementary information inhibits HIV-1 infection by reducing virion incorporation of envelope glycoproteins Takuya Tada, Yanzhao Zhang, Takayoshi Koyama, Minoru Tobiume, Yasuko Tsunetsugu-Yokota, Shoji
Supplementary information inhibits HIV-1 infection by reducing virion incorporation of envelope glycoproteins Takuya Tada, Yanzhao Zhang, Takayoshi Koyama, Minoru Tobiume, Yasuko Tsunetsugu-Yokota, Shoji
Contribution of PDZD8 to Stabilization of the Human Immunodeficiency Virus (HIV-1) Capsid
 Contribution of PDZD8 to Stabilization of the Human Immunodeficiency Virus (HIV-1) Capsid The Harvard community has made this article openly available. Please share how this access benefits you. Your story
Contribution of PDZD8 to Stabilization of the Human Immunodeficiency Virus (HIV-1) Capsid The Harvard community has made this article openly available. Please share how this access benefits you. Your story
ARV Mode of Action. Mode of Action. Mode of Action NRTI. Immunopaedia.org.za
 ARV Mode of Action Mode of Action Mode of Action - NRTI Mode of Action - NNRTI Mode of Action - Protease Inhibitors Mode of Action - Integrase inhibitor Mode of Action - Entry Inhibitors Mode of Action
ARV Mode of Action Mode of Action Mode of Action - NRTI Mode of Action - NNRTI Mode of Action - Protease Inhibitors Mode of Action - Integrase inhibitor Mode of Action - Entry Inhibitors Mode of Action
HUMAN IMMUNODEFICIENCY VIRUS
 Futuro promisorio de la terapia antirretroviral: Nuevos blancos terapéuticos. María José Míguez, M.D., PhD., Universidad de Miami, EE.UU. HUMAN IMMUNODEFICIENCY VIRUS REVERSE TRANSCRIPTASA REPLICATION
Futuro promisorio de la terapia antirretroviral: Nuevos blancos terapéuticos. María José Míguez, M.D., PhD., Universidad de Miami, EE.UU. HUMAN IMMUNODEFICIENCY VIRUS REVERSE TRANSCRIPTASA REPLICATION
Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site.
 Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
Rajesh Kannangai Phone: ; Fax: ; *Corresponding author
 Amino acid sequence divergence of Tat protein (exon1) of subtype B and C HIV-1 strains: Does it have implications for vaccine development? Abraham Joseph Kandathil 1, Rajesh Kannangai 1, *, Oriapadickal
Amino acid sequence divergence of Tat protein (exon1) of subtype B and C HIV-1 strains: Does it have implications for vaccine development? Abraham Joseph Kandathil 1, Rajesh Kannangai 1, *, Oriapadickal
Crystallization-grade After D After V3 cocktail. Time (s) Time (s) Time (s) Time (s) Time (s) Time (s)
 Ligand Type Name 6 Crystallization-grade After 447-52D After V3 cocktail Receptor CD4 Resonance Units 5 1 5 1 5 1 Broadly neutralizing antibodies 2G12 VRC26.9 Resonance Units Resonance Units 3 1 15 1 5
Ligand Type Name 6 Crystallization-grade After 447-52D After V3 cocktail Receptor CD4 Resonance Units 5 1 5 1 5 1 Broadly neutralizing antibodies 2G12 VRC26.9 Resonance Units Resonance Units 3 1 15 1 5
An integrated map of HIV genome-wide variation from a population perspective. Li et al.
 An integrated map of HIV genome-wide variation from a population perspective Li et al. Li et al. Retrovirology (2015) 12:18 DOI 10.1186/s12977-015-0148-6 Li et al. Retrovirology (2015) 12:18 DOI 10.1186/s12977-015-0148-6
An integrated map of HIV genome-wide variation from a population perspective Li et al. Li et al. Retrovirology (2015) 12:18 DOI 10.1186/s12977-015-0148-6 Li et al. Retrovirology (2015) 12:18 DOI 10.1186/s12977-015-0148-6
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Binding capacity of DNA-barcoded MHC multimers and recovery of antigen specificity
 Supplementary Figure 1 Binding capacity of DNA-barcoded MHC multimers and recovery of antigen specificity (a, b) Fluorescent-based determination of the binding capacity of DNA-barcoded MHC multimers (+barcode)
Supplementary Figure 1 Binding capacity of DNA-barcoded MHC multimers and recovery of antigen specificity (a, b) Fluorescent-based determination of the binding capacity of DNA-barcoded MHC multimers (+barcode)
Molecular Mechanisms by Which Human Immunodeficiency Virus Type 1 Integrase Stimulates the Early Steps of Reverse Transcription
 JOURNAL OF VIROLOGY, Sept. 2007, p. 10037 10046 Vol. 81, No. 18 0022-538X/07/$08.00 0 doi:10.1128/jvi.00519-07 Copyright 2007, American Society for Microbiology. All Rights Reserved. Molecular Mechanisms
JOURNAL OF VIROLOGY, Sept. 2007, p. 10037 10046 Vol. 81, No. 18 0022-538X/07/$08.00 0 doi:10.1128/jvi.00519-07 Copyright 2007, American Society for Microbiology. All Rights Reserved. Molecular Mechanisms
Structural biology of viruses
 Structural biology of viruses Biophysical Chemistry 1, Fall 2010 Coat proteins DNA/RNA packaging Reading assignment: Chap. 15 Virus particles self-assemble from coat monomers Virus Structure and Function
Structural biology of viruses Biophysical Chemistry 1, Fall 2010 Coat proteins DNA/RNA packaging Reading assignment: Chap. 15 Virus particles self-assemble from coat monomers Virus Structure and Function
Supporting Information
 Supporting Information Sui et al..7/pnas.997 Pre-CLP CM9 LA9 SL Tat# Pol Vif % Tetramer + CD + CD + Vac+IL- +IL- Vac Fig. S. Frequencies of six different CD + CD + Mamu-A*-tetramer + cells were measured
Supporting Information Sui et al..7/pnas.997 Pre-CLP CM9 LA9 SL Tat# Pol Vif % Tetramer + CD + CD + Vac+IL- +IL- Vac Fig. S. Frequencies of six different CD + CD + Mamu-A*-tetramer + cells were measured
Supplemental Information. An Atlas of b-glucuronidases. in the Human Intestinal Microbiome
 Structure, Volume 2 Supplemental Information An Atlas of b-glucuronidases in the Human Intestinal Microbiome Rebecca M. Pollet, Emma H. D'Agostino, William G. Walton, Yongmei Xu, Michael S. Little, Kristen
Structure, Volume 2 Supplemental Information An Atlas of b-glucuronidases in the Human Intestinal Microbiome Rebecca M. Pollet, Emma H. D'Agostino, William G. Walton, Yongmei Xu, Michael S. Little, Kristen
Supplementary Material
 Supplementary Material Supplementary Text Text S1. Time distributions of the high FRET efficiency at different concentrations of EF-G.GTP From Fig. 1, the model of ribosomal translocation at non-saturating
Supplementary Material Supplementary Text Text S1. Time distributions of the high FRET efficiency at different concentrations of EF-G.GTP From Fig. 1, the model of ribosomal translocation at non-saturating
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Frequency of alternative-cassette-exon engagement with the ribosome is consistent across data from multiple human cell types and from mouse stem cells. Box plots showing AS frequency
Supplementary Figure 1 Frequency of alternative-cassette-exon engagement with the ribosome is consistent across data from multiple human cell types and from mouse stem cells. Box plots showing AS frequency
NK mediated Antibody Dependent Cellular Cytotoxicity in HIV infections
 NK mediated Antibody Dependent Cellular Cytotoxicity in HIV infections Amy Chung Dr. Ivan Stratov Prof. Stephen Kent ADCC process consists of Target cell QuickTime and a TIFF (Uncompressed) FcγR decompressor
NK mediated Antibody Dependent Cellular Cytotoxicity in HIV infections Amy Chung Dr. Ivan Stratov Prof. Stephen Kent ADCC process consists of Target cell QuickTime and a TIFF (Uncompressed) FcγR decompressor
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/4/3/eaaq0762/dc1 Supplementary Materials for Structures of monomeric and oligomeric forms of the Toxoplasma gondii perforin-like protein 1 Tao Ni, Sophie I. Williams,
advances.sciencemag.org/cgi/content/full/4/3/eaaq0762/dc1 Supplementary Materials for Structures of monomeric and oligomeric forms of the Toxoplasma gondii perforin-like protein 1 Tao Ni, Sophie I. Williams,
Supplementary Figure 1. ALVAC-protein vaccines and macaque immunization. (A) Maximum likelihood
 Supplementary Figure 1. ALVAC-protein vaccines and macaque immunization. (A) Maximum likelihood tree illustrating CRF01_AE gp120 protein sequence relationships between 107 Envs sampled in the RV144 trial
Supplementary Figure 1. ALVAC-protein vaccines and macaque immunization. (A) Maximum likelihood tree illustrating CRF01_AE gp120 protein sequence relationships between 107 Envs sampled in the RV144 trial
Table S1: Kinetic parameters of drug and substrate binding to wild type and HIV-1 protease variants. Data adapted from Ref. 6 in main text.
 Dynamical Network of HIV-1 Protease Mutants Reveals the Mechanism of Drug Resistance and Unhindered Activity Rajeswari Appadurai and Sanjib Senapati* BJM School of Biosciences and Department of Biotechnology,
Dynamical Network of HIV-1 Protease Mutants Reveals the Mechanism of Drug Resistance and Unhindered Activity Rajeswari Appadurai and Sanjib Senapati* BJM School of Biosciences and Department of Biotechnology,
Analysis of protein modeling for envelope glycoprotein GP120 for HIV via bioinformatics approaches
 International Research Journal of Virology Vol. 1(1), pp. 002-006, March, 2014. www.premierpublishers.org ISSN: 2326-7193x IRJV Research Article Analysis of protein modeling for envelope glycoprotein GP120
International Research Journal of Virology Vol. 1(1), pp. 002-006, March, 2014. www.premierpublishers.org ISSN: 2326-7193x IRJV Research Article Analysis of protein modeling for envelope glycoprotein GP120
MODULE 4: SPLICING. Removal of introns from messenger RNA by splicing
 Last update: 05/10/2017 MODULE 4: SPLICING Lesson Plan: Title MEG LAAKSO Removal of introns from messenger RNA by splicing Objectives Identify splice donor and acceptor sites that are best supported by
Last update: 05/10/2017 MODULE 4: SPLICING Lesson Plan: Title MEG LAAKSO Removal of introns from messenger RNA by splicing Objectives Identify splice donor and acceptor sites that are best supported by
HIV Life Cycle & Genetics
 HIV Life Cycle & enetics! etroviruses (and transposable elements) appear to be part of every cell's genome! From bacteria to yeast, flies, fish, and humans! ome endogenous retroviruses (most notably in
HIV Life Cycle & enetics! etroviruses (and transposable elements) appear to be part of every cell's genome! From bacteria to yeast, flies, fish, and humans! ome endogenous retroviruses (most notably in
NC Pol RNA RNA DNA D C T RNA. retroviruses/ Gag Pol Env. E/psi. E/psi RNA. MA CA NC MA Env CA 2-4
 55 pp.153 160 2005 RN 1. 100nm RN D T http://www.ncbi.nlm.nih.gov/ retroviruses/ ag Pol Env ag 4 6 M N M Env 565-0871 3-1 TEL 06-6879-8348 FX 06-6879-8347 E-mail sakuragi@biken.osaka-u.ac.jp N Pol PR RT
55 pp.153 160 2005 RN 1. 100nm RN D T http://www.ncbi.nlm.nih.gov/ retroviruses/ ag Pol Env ag 4 6 M N M Env 565-0871 3-1 TEL 06-6879-8348 FX 06-6879-8347 E-mail sakuragi@biken.osaka-u.ac.jp N Pol PR RT
Peptide hydrolysis uncatalyzed half-life = ~450 years HIV protease-catalyzed half-life = ~3 seconds
 Uncatalyzed half-life Peptide hydrolysis uncatalyzed half-life = ~450 years IV protease-catalyzed half-life = ~3 seconds Life Sciences 1a Lecture Slides Set 9 Fall 2006-2007 Prof. David R. Liu In the absence
Uncatalyzed half-life Peptide hydrolysis uncatalyzed half-life = ~450 years IV protease-catalyzed half-life = ~3 seconds Life Sciences 1a Lecture Slides Set 9 Fall 2006-2007 Prof. David R. Liu In the absence
Diagnostic Tests for HIV
 Mountain West AIDS Education and Training Center Diagnostic Tests for HIV David Spach, MD Principal Investigator, Mountain West AETC Professor of Medicine, University of Washington Last Updated: June 22,
Mountain West AIDS Education and Training Center Diagnostic Tests for HIV David Spach, MD Principal Investigator, Mountain West AETC Professor of Medicine, University of Washington Last Updated: June 22,
Translation. Host Cell Shutoff 1) Initiation of eukaryotic translation involves many initiation factors
 Translation Questions? 1) How does poliovirus shutoff eukaryotic translation? 2) If eukaryotic messages are not translated how can poliovirus get its message translated? Host Cell Shutoff 1) Initiation
Translation Questions? 1) How does poliovirus shutoff eukaryotic translation? 2) If eukaryotic messages are not translated how can poliovirus get its message translated? Host Cell Shutoff 1) Initiation
SDS-Assisted Protein Transport Through Solid-State Nanopores
 Supplementary Information for: SDS-Assisted Protein Transport Through Solid-State Nanopores Laura Restrepo-Pérez 1, Shalini John 2, Aleksei Aksimentiev 2 *, Chirlmin Joo 1 *, Cees Dekker 1 * 1 Department
Supplementary Information for: SDS-Assisted Protein Transport Through Solid-State Nanopores Laura Restrepo-Pérez 1, Shalini John 2, Aleksei Aksimentiev 2 *, Chirlmin Joo 1 *, Cees Dekker 1 * 1 Department
To test the possible source of the HBV infection outside the study family, we searched the Genbank
 Supplementary Discussion The source of hepatitis B virus infection To test the possible source of the HBV infection outside the study family, we searched the Genbank and HBV Database (http://hbvdb.ibcp.fr),
Supplementary Discussion The source of hepatitis B virus infection To test the possible source of the HBV infection outside the study family, we searched the Genbank and HBV Database (http://hbvdb.ibcp.fr),
7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans.
 Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
Immunologic Methods in Diagnosis of HIV Infection. Tehran Medical Sciences Branch, Islamic Azad
 Immunologic Methods in Diagnosis of HIV Infection M Parsania, Ph.D. Tehran Medical Sciences Branch, Islamic Azad University Retroviridae Retroviruses (family Retroviridae) id ) are enveloped, single stranded
Immunologic Methods in Diagnosis of HIV Infection M Parsania, Ph.D. Tehran Medical Sciences Branch, Islamic Azad University Retroviridae Retroviruses (family Retroviridae) id ) are enveloped, single stranded
VIROLOGY. Engineering Viral Genomes: Retrovirus Vectors
 VIROLOGY Engineering Viral Genomes: Retrovirus Vectors Viral vectors Retrovirus replicative cycle Most mammalian retroviruses use trna PRO, trna Lys3, trna Lys1,2 The partially unfolded trna is annealed
VIROLOGY Engineering Viral Genomes: Retrovirus Vectors Viral vectors Retrovirus replicative cycle Most mammalian retroviruses use trna PRO, trna Lys3, trna Lys1,2 The partially unfolded trna is annealed
Running Head: AN UNDERSTANDING OF HIV- 1, SYMPTOMS, AND TREATMENTS. An Understanding of HIV- 1, Symptoms, and Treatments.
 Running Head: AN UNDERSTANDING OF HIV- 1, SYMPTOMS, AND TREATMENTS An Understanding of HIV- 1, Symptoms, and Treatments Benjamin Mills Abstract HIV- 1 is a virus that has had major impacts worldwide. Numerous
Running Head: AN UNDERSTANDING OF HIV- 1, SYMPTOMS, AND TREATMENTS An Understanding of HIV- 1, Symptoms, and Treatments Benjamin Mills Abstract HIV- 1 is a virus that has had major impacts worldwide. Numerous
SUPPLEMENTARY FIG. S1. MVC inhibition curves in NP2-CD4/CCR5 cells. Luciferase reporter viruses pseudotyped with baseline (black solid lines) and MVC
 Supplementary Data SUPPLEMENTARY FIG. S1. MVC inhibition curves in NP2-CD4/CCR5 cells. Luciferase reporter viruses pseudotyped with baseline (black solid lines) and MVC failure Envs (black dotted lines)
Supplementary Data SUPPLEMENTARY FIG. S1. MVC inhibition curves in NP2-CD4/CCR5 cells. Luciferase reporter viruses pseudotyped with baseline (black solid lines) and MVC failure Envs (black dotted lines)
Amino Acids. Review I: Protein Structure. Amino Acids: Structures. Amino Acids (contd.) Rajan Munshi
 Review I: Protein Structure Rajan Munshi BBSI @ Pitt 2005 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2005 Amino Acids Building blocks of proteins 20 amino acids
Review I: Protein Structure Rajan Munshi BBSI @ Pitt 2005 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2005 Amino Acids Building blocks of proteins 20 amino acids
 CDC site UNAIDS Aids Knowledge Base http://www.cdc.gov/hiv/dhap.htm http://hivinsite.ucsf.edu/insite.jsp?page=kb National Institute of Allergy and Infectious Diseases http://www.niaid.nih.gov/default.htm
CDC site UNAIDS Aids Knowledge Base http://www.cdc.gov/hiv/dhap.htm http://hivinsite.ucsf.edu/insite.jsp?page=kb National Institute of Allergy and Infectious Diseases http://www.niaid.nih.gov/default.htm
David S. Goodsell From the Department of Molecular Biology, The Scripps Research Institute, La Jolla, California 92037
 Q 2012 by The International Union of Biochemistry and Molecular Biology BIOCHEMISTRY AND MOLECULAR BIOLOGY EDUCATION Vol. 40, No. 5, pp. 291 296, 2012 Miniseries Illustrating the Machinery of Life: Viruses*
Q 2012 by The International Union of Biochemistry and Molecular Biology BIOCHEMISTRY AND MOLECULAR BIOLOGY EDUCATION Vol. 40, No. 5, pp. 291 296, 2012 Miniseries Illustrating the Machinery of Life: Viruses*
Human Immunodeficiency Virus
 Human Immunodeficiency Virus 1. Identification of the AIDS Virus a) opportunistic infections observed in homosexual men (all had T4 helper cell depletion) -> termed Acquired Immune Deficiency Syndrome;
Human Immunodeficiency Virus 1. Identification of the AIDS Virus a) opportunistic infections observed in homosexual men (all had T4 helper cell depletion) -> termed Acquired Immune Deficiency Syndrome;
Identification of Mutation(s) in. Associated with Neutralization Resistance. Miah Blomquist
 Identification of Mutation(s) in the HIV 1 gp41 Subunit Associated with Neutralization Resistance Miah Blomquist What is HIV 1? HIV-1 is an epidemic that affects over 34 million people worldwide. HIV-1
Identification of Mutation(s) in the HIV 1 gp41 Subunit Associated with Neutralization Resistance Miah Blomquist What is HIV 1? HIV-1 is an epidemic that affects over 34 million people worldwide. HIV-1
There are approximately 30,000 proteasomes in a typical human cell Each proteasome is approximately 700 kda in size The proteasome is made up of 3
 Proteasomes Proteasomes Proteasomes are responsible for degrading proteins that have been damaged, assembled improperly, or that are of no profitable use to the cell. The unwanted protein is literally
Proteasomes Proteasomes Proteasomes are responsible for degrading proteins that have been damaged, assembled improperly, or that are of no profitable use to the cell. The unwanted protein is literally
Defining kinetic properties of HIV-specific CD8 + T-cell responses in acute infection
 Defining kinetic properties of HIV-specific CD8 + T-cell responses in acute infection Yiding Yang 1 and Vitaly V. Ganusov 1,2,3 1 Department of Microbiology, University of Tennessee, Knoxville, TN 37996,
Defining kinetic properties of HIV-specific CD8 + T-cell responses in acute infection Yiding Yang 1 and Vitaly V. Ganusov 1,2,3 1 Department of Microbiology, University of Tennessee, Knoxville, TN 37996,
Impact of TLR7/8 Triggering on HIV Pathogenesis
 University Hospital Zurich u Division of Infectious Diseases and Hospital Epidemiology Impact of TLR7/8 Triggering on HIV Pathogenesis Roberto F. Speck, MD Hallmark of HIV-Infection is the Progressive
University Hospital Zurich u Division of Infectious Diseases and Hospital Epidemiology Impact of TLR7/8 Triggering on HIV Pathogenesis Roberto F. Speck, MD Hallmark of HIV-Infection is the Progressive
Aminoglycoside activity observed on single pre-translocation ribosome complexes
 correction notice Nat. Chem. Biol. 6, 54 62 (2010) Aminoglycoside activity observed on single pre-translocation ribosome complexes Michael B Feldman, Daniel S Terry, Roger B Altman & Scott C Blanchard
correction notice Nat. Chem. Biol. 6, 54 62 (2010) Aminoglycoside activity observed on single pre-translocation ribosome complexes Michael B Feldman, Daniel S Terry, Roger B Altman & Scott C Blanchard
Lecture Readings. Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell
 October 26, 2006 1 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
October 26, 2006 1 Vesicular Trafficking, Secretory Pathway, HIV Assembly and Exit from Cell 1. Secretory pathway a. Formation of coated vesicles b. SNAREs and vesicle targeting 2. Membrane fusion a. SNAREs
Lecture 10 More about proteins
 Lecture 10 More about proteins Today we're going to extend our discussion of protein structure. This may seem far-removed from gene cloning, but it is the path to understanding the genes that we are cloning.
Lecture 10 More about proteins Today we're going to extend our discussion of protein structure. This may seem far-removed from gene cloning, but it is the path to understanding the genes that we are cloning.
Supplementary Figure 1
 Supplementary Figure 1 Asymmetrical function of 5p and 3p arms of mir-181 and mir-30 families and mir-142 and mir-154. (a) Control experiments using mirna sensor vector and empty pri-mirna overexpression
Supplementary Figure 1 Asymmetrical function of 5p and 3p arms of mir-181 and mir-30 families and mir-142 and mir-154. (a) Control experiments using mirna sensor vector and empty pri-mirna overexpression
RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays
 Supplementary Materials RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays Junhee Seok 1*, Weihong Xu 2, Ronald W. Davis 2, Wenzhong Xiao 2,3* 1 School of Electrical Engineering,
Supplementary Materials RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays Junhee Seok 1*, Weihong Xu 2, Ronald W. Davis 2, Wenzhong Xiao 2,3* 1 School of Electrical Engineering,
Structure of the measles virus hemagglutinin bound to the CD46 receptor. César Santiago, María L. Celma, Thilo Stehle and José M.
 Supporting Figures and Table for Structure of the measles virus hemagglutinin bound to the CD46 receptor César Santiago, María L. Celma, Thilo Stehle and José M. Casasnovas This PDF file includes: Supplementary
Supporting Figures and Table for Structure of the measles virus hemagglutinin bound to the CD46 receptor César Santiago, María L. Celma, Thilo Stehle and José M. Casasnovas This PDF file includes: Supplementary
Abstract. Title of dissertation: HUMAN IMMUNODEFICIENCY VIRUS NUCLEOCAPSID NUCLEIC ACID CHAPERONE ACTIVITY.
 Abstract Title of dissertation: HUMAN IMMUNODEFICIENCY VIRUS NUCLEOCAPSID PROTEIN: ANALYSIS OF THE MECHANISM OF STRAND EXCHANGE AND THE ROLE OF THE ZINC FINGERS IN NUCLEIC ACID CHAPERONE ACTIVITY. Megan
Abstract Title of dissertation: HUMAN IMMUNODEFICIENCY VIRUS NUCLEOCAPSID PROTEIN: ANALYSIS OF THE MECHANISM OF STRAND EXCHANGE AND THE ROLE OF THE ZINC FINGERS IN NUCLEIC ACID CHAPERONE ACTIVITY. Megan
Molecular Cell Biology - Problem Drill 10: Gene Expression in Eukaryotes
 Molecular Cell Biology - Problem Drill 10: Gene Expression in Eukaryotes Question No. 1 of 10 1. Which of the following statements about gene expression control in eukaryotes is correct? Question #1 (A)
Molecular Cell Biology - Problem Drill 10: Gene Expression in Eukaryotes Question No. 1 of 10 1. Which of the following statements about gene expression control in eukaryotes is correct? Question #1 (A)
Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid.
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression.
 Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
COMPUTATIONAL ANALYSIS OF CONSERVED AND MUTATED AMINO ACIDS IN GP160 PROTEIN OF HIV TYPE-1
 Journal of Cell and Tissue Research Vol. 10(3) 2359-2364 (2010) ISSN: 0973-0028 (Available online at www.tcrjournals.com) Original Article COMPUTATIONAL ANALYSIS OF CONSERVED AND MUTATED AMINO ACIDS IN
Journal of Cell and Tissue Research Vol. 10(3) 2359-2364 (2010) ISSN: 0973-0028 (Available online at www.tcrjournals.com) Original Article COMPUTATIONAL ANALYSIS OF CONSERVED AND MUTATED AMINO ACIDS IN
Last time we talked about the few steps in viral replication cycle and the un-coating stage:
 Zeina Al-Momani Last time we talked about the few steps in viral replication cycle and the un-coating stage: Un-coating: is a general term for the events which occur after penetration, we talked about
Zeina Al-Momani Last time we talked about the few steps in viral replication cycle and the un-coating stage: Un-coating: is a general term for the events which occur after penetration, we talked about
HIV Anti-HIV Neutralizing Antibodies
 ,**/ The Japanese Society for AIDS Research The Journal of AIDS Research : HIV HIV Anti-HIV Neutralizing Antibodies * Junji SHIBATA and Shuzo MATSUSHITA * Division of Clinical Retrovirology and Infectious
,**/ The Japanese Society for AIDS Research The Journal of AIDS Research : HIV HIV Anti-HIV Neutralizing Antibodies * Junji SHIBATA and Shuzo MATSUSHITA * Division of Clinical Retrovirology and Infectious
Rice in vivo RNA structurome reveals RNA secondary structure conservation and divergence in plants
 Rice in vivo RN structurome reveals RN secondary structure conservation and divergence in plants Hongjing Deng 1,2,,5, Jitender heema 3, Hang Zhang 2, Hugh Woolfenden 2, Matthew Norris 2, Zhenshan Liu
Rice in vivo RN structurome reveals RN secondary structure conservation and divergence in plants Hongjing Deng 1,2,,5, Jitender heema 3, Hang Zhang 2, Hugh Woolfenden 2, Matthew Norris 2, Zhenshan Liu
