Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
|
|
|
- Iris Ryan
- 5 years ago
- Views:
Transcription
1 Supplementary Figure 1 Design of isolated protein and RNC constructs, and homogeneity of purified RNCs. (a) Schematic depicting the design and nomenclature used for all the isolated proteins and RNCs used in this study. Sequence boundaries are indicated. The isolated FLN constructs comprise: a range of FLN5 C-terminal truncations in which between 2 and 21 amino acids have been removed, FLNΔX, in which X refers to the extent of the truncation, the folding incompetent variant FLN5 Y719E harboring a destabilizing Glu mutation at the site indicated in orange, and FLN5-6Δ18 in which FLN6 has the removal of 18 amino acids from its C-terminus (Hsu, S.T. et al. Proc Natl Acad Sci U S A 104, , 2007). In SecM-stalled FLN5 RNCs, the FLN5 sequence is tethered to the PTC via increasing lengths of the FLN6 sequence and the SecM translational arrest motif; for simplicity this will be referred to here as the linker. The linker, ranging from 21 to 110 amino acids, therefore corresponds to the distance (in residues) between its most C-terminal residue, G750, and the PTC. Folding-incompetent RNCs have the Y719E mutation in FLN5 and are referred to as FLN5+L Y719E RNCs. For the poly(gs) RNCs, FLN6 is replaced with a repeating poly(gs) motif, GGGG/S, to generate equivalent linkers of between 21 to 67 residues in length, as present in the FLN5 RNCs. (b) Representative PAGE gel of a purified RNC (FLN5+42) and of 70S ribosomes visualized with Coomassie blue stain (left) and silver stain (right). The gels show a banding pattern characteristic of ribosomal proteins and show that the RNCs are essentially free of extraneous proteins. The bands corresponding to ribosomal proteins L1, L2, L6, S5, L29, L31, L35 are highlighted. (c) An anti-his western blot of FLN5+42 RNC and of 70S ribosomes, showing the trna-bound form of the RNC and the absence of any non-specific detection of ribosomal proteins in untranslating 70S ribosomes. The extent of nascent chain attachment to ribosomes was determined to be > 90% across all samples. (d) The amount of residual trigger factor (TF) and DnaK present within the purified RNC samples was assessed by densitometry analysis of anti-tf and anti-dnak western blots, respectively. The purified samples are essentially free of the effects of TF and DnaK ( 1.2%). Highlighted in asterisks are the RNCs for which TF or DnaK levels were below levels of detection. (e) Representative western blots of RNC samples which were presented in a modified form in Fig. 2c are shown here for clarity, where the RNCs shown in Fig. 2c are marked by asterisks.
2 Supplementary Figure 2 Monitoring of the integrity of RNCs. An integrated approach using NMR and biochemical analyses was used to determine the integrity of the RNCs over time. (a) For each of FLN5+21, 45, 47, 67 and 110 RNCs, the 1 H- 13 C correlation spectrum is shown (identical to Fig. 3a), overlaid with that of isolated FLN5 (cyan). Resonances arising from labelled 70S proteins can be observed in spectra at ca. H =0.8 p.p.m. (b) The integrity of these RNCs was monitored over time and the timeframe during which the nascent chain was determined to be attached in each RNC as derived by NMR methods is indicated by the shaded region: 13 C-edited diffusion experiments ( 1 H STE- 1 H, 13 C-HMQC) were used to determine the diffusion coefficient associated with the nascent chain in FLN5+67 and 110 RNCs (cyan). The changes in intensities of the cross-peaks observed in the correlation spectra were used to monitor the FLN5+45 and 47 RNCs (green) and the diffusion coefficients (from 13 C-edited experiments) of the combined ribosome-derived resonances were monitored for FLN5+21 RNC (yellow). (c) Western blots against either the N-terminal His-tag or C-terminal SecM sequence report on both the trna-bound and released forms of the nascent chain. (d) Densitometry analysis of the trna-bound form over time as assessed by the anti-his western blot. In the analyses shown in panels b, c and d, the shaded region represents the timeframe corresponding to an exclusively ribosome-bound nascent chain and for which the 2D correlation spectra are summed and presented in panel a. (e) As a representative example for 15 N- labelled RNCs, a 1 H- 15 N SOFAST-HMQC spectrum (identical to that in Fig. 3b) of FLN5+21 RNC (black) is overlaid with a 2D correlation spectrum of isolated FLN5 12 (blue) and a selection of unambiguously assigned resonances is indicated. (f) Diffusion coefficient for FLN5+21 RNC resonances assessed over time (calculated from 15 N XSTE spectra; single spectrum shown on left). A signal attenuation (I 95 /I 5 ) by a factor of > 0.62 corresponds to a diffusion coefficient of an intact ribosomal particle (D = 2.0 ± m 2 s -1 ) at 25 C. The timeframe during which the nascent chain is assessed as being intact is shaded in grey. (g) 15 N XSTE spectra of FLN5+21 RNC recorded at gradient strengths 5% and 95% of the maximum gradient strength G max, T m -1 for the timeframe during which the nascent chain was deemed to be intact. (h) Anti-His and anti-secm western blot analyses of FLN5+21 RNC.
3
4 Supplementary Figure 3 U- 15 N labeled spectra: assignments, chemical-shift analyses and linewidth measurements on FLN5 RNCs and isolated FLN5 variants. 1 H- 15 N correlation spectra and resonance assignments of: (a) FLN5 12 (b) FLN (c) the 70S ribosome for which the ribosomal protein L7/L12 gives rise to most resonances that are observed (Christodoulou, J. et al. Proc Natl Acad Sci U S A 101, , 2004). (d) 1 H- 15 N correlation spectrum of FLN+21 RNC. (e) Overlay of the 1 H- 15 N correlation spectra of FLN5+21 RNC (black) and isolated FLN5 12 (blue), demonstrating that the disordered region of FLN5+21 RNC spectrum ( ppm in 1 H dimension) overlays closely with that of disordered FLN5. (f) Overlay of FLN5+21 RNC (black) and FLN (green); the resonances in the disordered region of the RNC spectrum do not correspond to those of unfolded FLN6. (g) Overlay of the spectra of FLN5+21 RNC (black) and of the 70S ribosome particle (magenta), the limited overlap with ribosomal protein L7/L12 resonances indicating an essentially negligible level of background signal of ribosomal proteins in the RNC. (h) 1 H- 15 N correlation spectrum of FLN5+110 RNC. (i) Overlay of 1 H- 15 N correlation spectra of FLN5+110 RNC (black) and FLN5 12 (blue); the resonances in the disordered region of the RNC spectrum clearly do not correspond to those of unfolded isolated FLN5. (j) Overlay of 1 H- 15 N correlation spectra of FLN5+110 RNC (black) and FLN (green); the unfolded region of FLN5+110 RNC overlays closely with that of disordered FLN6. (k) Overlay of 1 H- 15 N correlation spectra of FLN5+110 RNC (black) and 70S ribosomes (magenta); there is a negligible contribution of resonances arising from the presence of labelled 70S ribosomal proteins. (l) 1 H- 15 N correlation spectrum of FLN5 Y719E (orange), overlaid with that of a C- terminally truncated variant, FLN5 12 (black). The close overlay demonstrates that the Y719E mutation (generated based upon predictions using the PoPMuSiC algorithm (Dehouck, Y., Kwasigroch, J.M., Gilis, D. & Rooman, M. BMC Bioinformatics 12, 151, 2011) results in an unfolded conformation that is highly comparable to that observed in FLN N R 2 relaxation rates (at 277 K, data not shown) averaged 2.8 ± 0.6 s -1 for the 81 residues analysed (error taken as the standard deviation), indicating a highly disordered protein. The 1 H- 15 N spectra of both proteins were completely assigned and the key chemical shift changes between the two are indicated with dotted lines. (m) Chemical shift differences between FLN5 Y719E and FLN5 12 are mapped against the amino acid sequence, as calculated by the formula. This shows that the chemical shifts are closely similar between the two constructs apart from at the mutation site (marked by an asterisk) and at the C-terminus of FLN5 12. (n) Comparison of linewidths between isolated FLN5 16 and FLN5+21, +31, +37 and +42 RNCs as measured for resonances Ala683, Ala694 and Val682. (o) 1 H cross-sections of FLN5+21, 31, 37 and 42 RNC resonances used for lineshape fitting (as described in the Online Methods): Ala683 and Ala694 are fitted simultaneously to Lorentzian lineshapes. RNC linewidths (blue) are greater than those observed for isolated FLN5 16 (grey).
5 Supplementary Figure 4 PEGylation of FLN5 Y719E RNCs. (a) Autoradiography of SDS-PAGE gels as presented in a cropped form in Fig. 4b is provided here for clarity. The arrows indicate the different forms of FLN5 nascent chains observed: trna-bound nascent chain (green), PEGylated trna-bound nascent chain (red) and released nascent chain (cyan), PEGylated released nascent chain (blue). Approximate migration of molecular weight standards is indicated on the left. (b) RNase A treated FLN5 Y719E RNCs, showing the PEGylation achievable on the released nascent chains (color coding as in a). Approximate migration of molecular weight standards is indicated on the left.
6 Supplementary Figure 5 1 H- 15 N correlation spectra of FLN5 C-terminal truncations. (a) Natively folded full-length FLN5 domain. (b) FLN5 8. The removal of eight C-terminal residues results in FLN5 sampling both unfolded and native-like conformational states. (c) FLN5 16, shows a characteristic spectrum of an unfolded polypeptide. Below, the regions removed by the C-terminal truncation are highlighted on ribbon diagrams of FLN5 (PDB: 1QFH), in magenta for FLN5 8 and blue for FLN5 16.
7 Supplementary Figure 6 FLN5 RNCs in which FLN6 is substituted with a poly(gs) linker. (a) Schematic of the poly(gs)-linker RNCs, in which the FLN6 domain is substituted with a poly(gs) sequence. The FLN6 residues that have been substituted in two RNCs analysed by NMR, FLN5+31 and +42 to generate FLN5+31 GS and +42 GS, respectively, are shown in the highlighted box. (b) Anti-His western blot of FLN5+31 GS and 42 GS RNCs in their released and trna-bound forms, respectively. (c) The emergence of FLN5 from the exit tunnel in a series of poly(gs)-linker RNCs was monitored by PEGylation of G751C variants of the FLN5 RNCs in a manner similar to that shown in Fig. 4 for the FLN6 domain. (d) Overlay of 1 H- 15 N correlation spectra of FLN5+42 GS RNC and FLN5+42 RNC. The latter was recorded at 950 MHz and with enhanced resolution, however FLN5+42 and +42 GS RNCs have similar intrinsic linewidths. The close overlay suggests shows the similar unfolded conformational preferences between the two nascent chains i.e. FLN5 is unfolded in the FLN5+42 GS RNC. (e) Overlay of 1 H- 15 N correlation spectra FLN5+42 GS RNC when it is intact (dark green) and released (after ~20 h of NMR acquisition, light green), which shows the appearance of additional glycine resonances arising from the poly(gs) linker in the released nascent chain. (f) Relative intensities of FLN5+42 (upper panel) and FLN5+31 (lower panel) GS RNCs compared to isolated, unfolded FLN5 (the green trace represents a 5- point moving average). 5-point moving average plots of the relative intensities of FLN5+42 (upper panel) and FLN5+31 (lower panel) RNCs compared to isolated unfolded FLN5 are also shown for comparison (pink). The close overlay of the two indicates that altering the linker does not seem to impart significantly on the conformation and/or ribosome interactions of unfolded FLN5. (g) Overlay of 1 H- 15 N correlation spectra of FLN5+42 GS RNC after ~20 h of NMR acquisition and of natively folded FLN5: upon release from the ribosome, resonances corresponding to natively folded FLN5 are observed.
8 Supplementary Figure 7 Interactions between folded and unfolded isolated FLN5 variants with 70S ribosomes. Relative peak intensities of: (a) FLN5 in the presence of 1 molar equivalent of 70S ribosomes relative to FLN5 alone, at 5 µm (as presented in Fig. 6b) (b) FLN5 12 in the presence of 1 molar equivalent of 70S ribosomes relative to FLN5 12 alone, at 5 µm. (c) FLN5 Y719E in the presence of 1 molar equivalent of 70S ribosomes relative to FLN5 Y719E alone, at 8 µm (as presented in Fig. 6b). Shaded areas highlight the peak broadenings observed.
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION TITLE: Structural Basis of Signal Sequence Surveillance and Selection by the SRP-SR Complex AUTHORS and AFFILIATIONS Ottilie von Loeffelholz 1,2, Kèvin Knoops 1,2,6, Aileen Ariosa
SUPPLEMENTARY INFORMATION TITLE: Structural Basis of Signal Sequence Surveillance and Selection by the SRP-SR Complex AUTHORS and AFFILIATIONS Ottilie von Loeffelholz 1,2, Kèvin Knoops 1,2,6, Aileen Ariosa
Transient Ribosomal Attenuation Coordinates Protein Synthesis and Co-translational Folding
 SUPPLEMENTARY INFORMATION: Transient Ribosomal Attenuation Coordinates Protein Synthesis and Co-translational Folding Gong Zhang 1,2, Magdalena Hubalewska 1 & Zoya Ignatova 1,2 1 Department of Cellular
SUPPLEMENTARY INFORMATION: Transient Ribosomal Attenuation Coordinates Protein Synthesis and Co-translational Folding Gong Zhang 1,2, Magdalena Hubalewska 1 & Zoya Ignatova 1,2 1 Department of Cellular
(B D) Three views of the final refined 2Fo-Fc electron density map of the Vpr (red)-ung2 (green) interacting region, contoured at 1.4σ.
 Supplementary Figure 1 Overall structure of the DDB1 DCAF1 Vpr UNG2 complex. (A) The final refined 2Fo-Fc electron density map, contoured at 1.4σ of Vpr, illustrating well-defined side chains. (B D) Three
Supplementary Figure 1 Overall structure of the DDB1 DCAF1 Vpr UNG2 complex. (A) The final refined 2Fo-Fc electron density map, contoured at 1.4σ of Vpr, illustrating well-defined side chains. (B D) Three
SUPPORTING INFORMATION. Lysine Carbonylation is a Previously Unrecognized Contributor. to Peroxidase Activation of Cytochrome c by Chloramine-T
 Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2019 SUPPORTING INFORMATION Lysine Carbonylation is a Previously Unrecognized Contributor to
Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2019 SUPPORTING INFORMATION Lysine Carbonylation is a Previously Unrecognized Contributor to
Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site.
 Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
Structural Characterization of Prion-like Conformational Changes of the Neuronal Isoform of Aplysia CPEB
 Structural Characterization of Prion-like Conformational Changes of the Neuronal Isoform of Aplysia CPEB Bindu L. Raveendra, 1,5 Ansgar B. Siemer, 2,6 Sathyanarayanan V. Puthanveettil, 1,3,7 Wayne A. Hendrickson,
Structural Characterization of Prion-like Conformational Changes of the Neuronal Isoform of Aplysia CPEB Bindu L. Raveendra, 1,5 Ansgar B. Siemer, 2,6 Sathyanarayanan V. Puthanveettil, 1,3,7 Wayne A. Hendrickson,
Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation
 Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation Hang Yu, 1, 2, a) Wei Han, 1, 3, b) Wen Ma, 1, 2 1, 2, 3, c) and Klaus Schulten 1) Beckman
Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation Hang Yu, 1, 2, a) Wei Han, 1, 3, b) Wen Ma, 1, 2 1, 2, 3, c) and Klaus Schulten 1) Beckman
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 The UBL and RING1 interface remains associated in the complex structures of Parkin and pub. a) Asymmetric Unit of crystal structure of UBLR0RBR and pub complex showing UBL (green),
Supplementary Figure 1 The UBL and RING1 interface remains associated in the complex structures of Parkin and pub. a) Asymmetric Unit of crystal structure of UBLR0RBR and pub complex showing UBL (green),
Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.
 Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.5 % SDS-PAGE gel under reducing and non-reducing conditions.
Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.5 % SDS-PAGE gel under reducing and non-reducing conditions.
Introduction 1/3. The protagonist of our story: the prokaryotic ribosome
 Introduction 1/3 The protagonist of our story: the prokaryotic ribosome Introduction 2/3 The core functional domains of the ribosome (the peptidyl trasferase center and the decoding center) are composed
Introduction 1/3 The protagonist of our story: the prokaryotic ribosome Introduction 2/3 The core functional domains of the ribosome (the peptidyl trasferase center and the decoding center) are composed
Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random
 S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
Unveiling transient protein-protein interactions that modulate inhibition of alpha-synuclein aggregation
 Supplementary information Unveiling transient protein-protein interactions that modulate inhibition of alpha-synuclein aggregation by beta-synuclein, a pre-synaptic protein that co-localizes with alpha-synuclein.
Supplementary information Unveiling transient protein-protein interactions that modulate inhibition of alpha-synuclein aggregation by beta-synuclein, a pre-synaptic protein that co-localizes with alpha-synuclein.
Figure S1. (A) SDS-PAGE separation of GST-fusion proteins purified from E.coli BL21 strain is shown. An equal amount of GST-tag control, LRRK2 LRR
 Figure S1. (A) SDS-PAGE separation of GST-fusion proteins purified from E.coli BL21 strain is shown. An equal amount of GST-tag control, LRRK2 LRR and LRRK2 WD40 GST fusion proteins (5 µg) were loaded
Figure S1. (A) SDS-PAGE separation of GST-fusion proteins purified from E.coli BL21 strain is shown. An equal amount of GST-tag control, LRRK2 LRR and LRRK2 WD40 GST fusion proteins (5 µg) were loaded
Chapter 31. Completing the Protein Life Cycle: Folding, Processing and Degradation. Biochemistry by Reginald Garrett and Charles Grisham
 Chapter 31 Completing the Protein Life Cycle: Folding, Processing and Degradation Biochemistry by Reginald Garrett and Charles Grisham Essential Question How are newly synthesized polypeptide chains transformed
Chapter 31 Completing the Protein Life Cycle: Folding, Processing and Degradation Biochemistry by Reginald Garrett and Charles Grisham Essential Question How are newly synthesized polypeptide chains transformed
Supplementary Figure 1. Structure models of the c4 variant proteins based on the structures of maltose binding protein (MBP) in red and TEM-1
 Supplementary Figure 1. Structure models of the c4 variant proteins based on the structures of maltose binding protein (MBP) in red and TEM-1 β-lactamase (BLA) in blue. (a) c4 model and the close-up view
Supplementary Figure 1. Structure models of the c4 variant proteins based on the structures of maltose binding protein (MBP) in red and TEM-1 β-lactamase (BLA) in blue. (a) c4 model and the close-up view
Supplementary Figure 1. Properties of various IZUMO1 monoclonal antibodies and behavior of SPACA6. (a) (b) (c) (d) (e) (f) (g) .
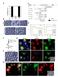 Supplementary Figure 1. Properties of various IZUMO1 monoclonal antibodies and behavior of SPACA6. (a) The inhibitory effects of new antibodies (Mab17 and Mab18). They were investigated in in vitro fertilization
Supplementary Figure 1. Properties of various IZUMO1 monoclonal antibodies and behavior of SPACA6. (a) The inhibitory effects of new antibodies (Mab17 and Mab18). They were investigated in in vitro fertilization
Supplementary Materials for
 www.sciencemag.org/cgi/content/full/science.aal4326/dc1 Supplementary Materials for Structure of a eukaryotic voltage-gated sodium channel at near-atomic resolution Huaizong Shen, Qiang Zhou, Xiaojing
www.sciencemag.org/cgi/content/full/science.aal4326/dc1 Supplementary Materials for Structure of a eukaryotic voltage-gated sodium channel at near-atomic resolution Huaizong Shen, Qiang Zhou, Xiaojing
KDM2A. Reactions. containing. Reactions
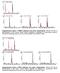 Supplementary Figure 1 KDM2A catalyses only lysine demethylation.. MALDI-TOF MS of demethylation of the shown variant histone peptides as catalysed by recombinant KDM2A. Reactions containing enzyme are
Supplementary Figure 1 KDM2A catalyses only lysine demethylation.. MALDI-TOF MS of demethylation of the shown variant histone peptides as catalysed by recombinant KDM2A. Reactions containing enzyme are
SUPPLEMENTARY INFORMATION. Supplementary Figures
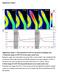 SUPPLEMENTARY INFORMATION Supplementary Figures Supplementary Figure 1: Characterization of CerTN-L15 expressed in Arabidopsis roots. a. Ratiometric images of CerTN-L15 in roots under osmotic stress Ratiometric
SUPPLEMENTARY INFORMATION Supplementary Figures Supplementary Figure 1: Characterization of CerTN-L15 expressed in Arabidopsis roots. a. Ratiometric images of CerTN-L15 in roots under osmotic stress Ratiometric
Supplementary Information
 Supplementary Information Archaeal Elp3 catalyzes trna wobble uridine modification at C5 via a radical mechanism Kiruthika Selvadurai, Pei Wang, Joseph Seimetz & Raven H Huang* Department of Biochemistry,
Supplementary Information Archaeal Elp3 catalyzes trna wobble uridine modification at C5 via a radical mechanism Kiruthika Selvadurai, Pei Wang, Joseph Seimetz & Raven H Huang* Department of Biochemistry,
crossmark Ca V subunits interact with the voltage-gated calcium channel
 crossmark THE JOURNAL OF BIOLOGICAL CHEMISTRY VOL. 291, NO. 39, pp. 20402 20416, September 23, 2016 Author s Choice 2016 by The American Society for Biochemistry and Molecular Biology, Inc. Published in
crossmark THE JOURNAL OF BIOLOGICAL CHEMISTRY VOL. 291, NO. 39, pp. 20402 20416, September 23, 2016 Author s Choice 2016 by The American Society for Biochemistry and Molecular Biology, Inc. Published in
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10913 Supplementary Figure 1 2F o -F c electron density maps of cognate and near-cognate trna Leu 2 in the A site of the 70S ribosome. The maps are contoured at 1.2 sigma and some of
doi:10.1038/nature10913 Supplementary Figure 1 2F o -F c electron density maps of cognate and near-cognate trna Leu 2 in the A site of the 70S ribosome. The maps are contoured at 1.2 sigma and some of
Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular
 Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular weight markers (M). Supplementary Figure-2. Overlay of
Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular weight markers (M). Supplementary Figure-2. Overlay of
Thermal shift binding experiments were carried out using Thermofluor 384 ELS system. Protein
 Supplementary Methods Thermal shift assays Thermal shift binding experiments were carried out using Thermofluor 384 ELS system. Protein unfolding was examined by monitoring the fluorescence of ANS (1-anilinonaphthalene-8-
Supplementary Methods Thermal shift assays Thermal shift binding experiments were carried out using Thermofluor 384 ELS system. Protein unfolding was examined by monitoring the fluorescence of ANS (1-anilinonaphthalene-8-
Nature Structural & Molecular Biology: doi: /nsmb.2419
 Supplementary Figure 1 Mapped sequence reads and nucleosome occupancies. (a) Distribution of sequencing reads on the mouse reference genome for chromosome 14 as an example. The number of reads in a 1 Mb
Supplementary Figure 1 Mapped sequence reads and nucleosome occupancies. (a) Distribution of sequencing reads on the mouse reference genome for chromosome 14 as an example. The number of reads in a 1 Mb
SDS-Assisted Protein Transport Through Solid-State Nanopores
 Supplementary Information for: SDS-Assisted Protein Transport Through Solid-State Nanopores Laura Restrepo-Pérez 1, Shalini John 2, Aleksei Aksimentiev 2 *, Chirlmin Joo 1 *, Cees Dekker 1 * 1 Department
Supplementary Information for: SDS-Assisted Protein Transport Through Solid-State Nanopores Laura Restrepo-Pérez 1, Shalini John 2, Aleksei Aksimentiev 2 *, Chirlmin Joo 1 *, Cees Dekker 1 * 1 Department
Chapter 8. Interaction between the phosphatidylinositol 3- kinase SH3 domain and a photocleavable cyclic peptide
 Interaction between the phosphatidylinositol 3- kinase SH3 domain and a photocleavable cyclic peptide 129 Abstract The interaction of the PI3K SH3 domain with a cyclic photocleavable peptide and the linear
Interaction between the phosphatidylinositol 3- kinase SH3 domain and a photocleavable cyclic peptide 129 Abstract The interaction of the PI3K SH3 domain with a cyclic photocleavable peptide and the linear
Table S1: Kinetic parameters of drug and substrate binding to wild type and HIV-1 protease variants. Data adapted from Ref. 6 in main text.
 Dynamical Network of HIV-1 Protease Mutants Reveals the Mechanism of Drug Resistance and Unhindered Activity Rajeswari Appadurai and Sanjib Senapati* BJM School of Biosciences and Department of Biotechnology,
Dynamical Network of HIV-1 Protease Mutants Reveals the Mechanism of Drug Resistance and Unhindered Activity Rajeswari Appadurai and Sanjib Senapati* BJM School of Biosciences and Department of Biotechnology,
Supplementary Figure 1. Amino acid sequences of GodA and GodA*. Inserted. residues are colored red. Numbers indicate the position of each residue.
 Supplementary Figure 1. Amino acid sequences of GodA and GodA*. Inserted residues are colored red. Numbers indicate the position of each residue. Supplementary Figure 2. SDS-PAGE analysis of purified recombinant
Supplementary Figure 1. Amino acid sequences of GodA and GodA*. Inserted residues are colored red. Numbers indicate the position of each residue. Supplementary Figure 2. SDS-PAGE analysis of purified recombinant
Problem-solving Test: The Mechanism of Protein Synthesis
 Q 2009 by The International Union of Biochemistry and Molecular Biology BIOCHEMISTRY AND MOLECULAR BIOLOGY EDUCATION Vol. 37, No. 1, pp. 58 62, 2009 Problem-based Learning Problem-solving Test: The Mechanism
Q 2009 by The International Union of Biochemistry and Molecular Biology BIOCHEMISTRY AND MOLECULAR BIOLOGY EDUCATION Vol. 37, No. 1, pp. 58 62, 2009 Problem-based Learning Problem-solving Test: The Mechanism
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature22394 Supplementary Table 1 Observed intermolecular interactions within the GLP-1:GLP-1R TMD interface. Superscripts refer to the Wootten residue numbering system
SUPPLEMENTARY INFORMATION doi:10.1038/nature22394 Supplementary Table 1 Observed intermolecular interactions within the GLP-1:GLP-1R TMD interface. Superscripts refer to the Wootten residue numbering system
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/8/398/rs12/dc1 Supplementary Materials for Quantitative phosphoproteomics reveals new roles for the protein phosphatase PP6 in mitotic cells Scott F. Rusin, Kate
www.sciencesignaling.org/cgi/content/full/8/398/rs12/dc1 Supplementary Materials for Quantitative phosphoproteomics reveals new roles for the protein phosphatase PP6 in mitotic cells Scott F. Rusin, Kate
Alpha thalassemia mental retardation X-linked. Acquired alpha-thalassemia myelodysplastic syndrome
 Alpha thalassemia mental retardation X-linked Acquired alpha-thalassemia myelodysplastic syndrome (Alpha thalassemia mental retardation X-linked) Acquired alpha-thalassemia myelodysplastic syndrome Schematic
Alpha thalassemia mental retardation X-linked Acquired alpha-thalassemia myelodysplastic syndrome (Alpha thalassemia mental retardation X-linked) Acquired alpha-thalassemia myelodysplastic syndrome Schematic
reads observed in trnas from the analysis of RNAs carrying a 5 -OH ends isolated from cells induced to express
 Supplementary Figure 1. VapC-mt4 cleaves trna Ala2 in E. coli. Histograms representing the fold change in reads observed in trnas from the analysis of RNAs carrying a 5 -OH ends isolated from cells induced
Supplementary Figure 1. VapC-mt4 cleaves trna Ala2 in E. coli. Histograms representing the fold change in reads observed in trnas from the analysis of RNAs carrying a 5 -OH ends isolated from cells induced
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Figure 1 SNARE Probes for FRET/2pFLIM Analysis Used in the Present Study. mturquoise (mtq) and Venus (Ven) are in blue and yellow, respectively. The soluble N-ethylmaleimide-sensitive
Supplementary Figure 1 Supplementary Figure 1 SNARE Probes for FRET/2pFLIM Analysis Used in the Present Study. mturquoise (mtq) and Venus (Ven) are in blue and yellow, respectively. The soluble N-ethylmaleimide-sensitive
Analysis of Uncomplexed and Copper-complexed Methanobactin with UV/Visible Spectrophotometry, Mass Spectrometry and NMR Spectrometry
 Analysis of Uncomplexed and Copper-complexed Methanobactin with UV/Visible Spectrophotometry, Mass Spectrometry and NMR Spectrometry Lee Behling, Alan DiSpirito, Scott Hartsel, Larry Masterson, Gianluigi
Analysis of Uncomplexed and Copper-complexed Methanobactin with UV/Visible Spectrophotometry, Mass Spectrometry and NMR Spectrometry Lee Behling, Alan DiSpirito, Scott Hartsel, Larry Masterson, Gianluigi
Supplementary Figure 1. Using DNA barcode-labeled MHC multimers to generate TCR fingerprints
 Supplementary Figure 1 Using DNA barcode-labeled MHC multimers to generate TCR fingerprints (a) Schematic overview of the workflow behind a TCR fingerprint. Each peptide position of the original peptide
Supplementary Figure 1 Using DNA barcode-labeled MHC multimers to generate TCR fingerprints (a) Schematic overview of the workflow behind a TCR fingerprint. Each peptide position of the original peptide
Characterizing fatty acids with advanced multinuclear NMR methods
 Characterizing fatty acids with advanced multinuclear NMR methods Fatty acids consist of long carbon chains ending with a carboxylic acid on one side and a methyl group on the other. Most naturally occurring
Characterizing fatty acids with advanced multinuclear NMR methods Fatty acids consist of long carbon chains ending with a carboxylic acid on one side and a methyl group on the other. Most naturally occurring
SUPPLEMENTARY INFORMATION
 Figure S1. Cleavage of uniquitin AAA -CPP TAT in vitro and in cells. a, b. In vitro two-dimensional 1 H- 15 N correlation spectrum of ubiquitin AAA -CPP TAT before (a) and after (b) Yeast Ubiquitin Hydrolase
Figure S1. Cleavage of uniquitin AAA -CPP TAT in vitro and in cells. a, b. In vitro two-dimensional 1 H- 15 N correlation spectrum of ubiquitin AAA -CPP TAT before (a) and after (b) Yeast Ubiquitin Hydrolase
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10962 Supplementary Figure 1. Expression of AvrAC-FLAG in protoplasts. Total protein extracted from protoplasts described in Fig. 1a was subjected to anti-flag immunoblot to detect AvrAC-FLAG
doi:10.1038/nature10962 Supplementary Figure 1. Expression of AvrAC-FLAG in protoplasts. Total protein extracted from protoplasts described in Fig. 1a was subjected to anti-flag immunoblot to detect AvrAC-FLAG
SUPPLEMENTARY INFORMATION. An orthogonal ribosome-trnas pair by the engineering of
 SUPPLEMENTARY INFORMATION An orthogonal ribosome-trnas pair by the engineering of peptidyl transferase center Naohiro Terasaka 1 *, Gosuke Hayashi 1 *, Takayuki Katoh 1, and Hiroaki Suga 1,2 1 Department
SUPPLEMENTARY INFORMATION An orthogonal ribosome-trnas pair by the engineering of peptidyl transferase center Naohiro Terasaka 1 *, Gosuke Hayashi 1 *, Takayuki Katoh 1, and Hiroaki Suga 1,2 1 Department
Cbl ubiquitin ligase: Lord of the RINGs
 Cbl ubiquitin ligase: Lord of the RINGs Not just quite interesting - really interesting! A cell must be able to degrade proteins when their activity is no longer required. Many eukaryotic proteins are
Cbl ubiquitin ligase: Lord of the RINGs Not just quite interesting - really interesting! A cell must be able to degrade proteins when their activity is no longer required. Many eukaryotic proteins are
STRUCTURE OF A UBIQUITIN-LOADED HECT LIGASE REVEALS THE MOLECULAR BASIS FOR CATALYTIC PRIMING SUPPLEMENTARY INFORMATION
 STRUCTURE OF A UBIQUITINLOADED ECT LIGASE REVEALS TE MOLECULAR BASIS FOR CATALYTIC PRIMING Elena Maspero 1, Eleonora Valentini 1, Sara Mari 1, Valentina Cecatiello 2, Paolo Soffientini 1, Sebastiano Pasqualato
STRUCTURE OF A UBIQUITINLOADED ECT LIGASE REVEALS TE MOLECULAR BASIS FOR CATALYTIC PRIMING Elena Maspero 1, Eleonora Valentini 1, Sara Mari 1, Valentina Cecatiello 2, Paolo Soffientini 1, Sebastiano Pasqualato
Double charge of 33kD peak A1 A2 B1 B2 M2+ M/z. ABRF Proteomics Research Group - Qualitative Proteomics Study Identifier Number 14146
 Abstract The 2008 ABRF Proteomics Research Group Study offers participants the chance to participate in an anonymous study to identify qualitative differences between two protein preparations. We used
Abstract The 2008 ABRF Proteomics Research Group Study offers participants the chance to participate in an anonymous study to identify qualitative differences between two protein preparations. We used
Nature Neuroscience: doi: /nn Supplementary Figure 1. Neuron class-specific arrangements of Khc::nod::lacZ label in dendrites.
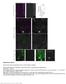 Supplementary Figure 1 Neuron class-specific arrangements of Khc::nod::lacZ label in dendrites. Staining with fluorescence antibodies to detect GFP (Green), β-galactosidase (magenta/white). (a, b) Class
Supplementary Figure 1 Neuron class-specific arrangements of Khc::nod::lacZ label in dendrites. Staining with fluorescence antibodies to detect GFP (Green), β-galactosidase (magenta/white). (a, b) Class
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2988 Supplementary Figure 1 Kif7 L130P encodes a stable protein that does not localize to cilia tips. (a) Immunoblot with KIF7 antibody in cell lysates of wild-type, Kif7 L130P and Kif7
DOI: 10.1038/ncb2988 Supplementary Figure 1 Kif7 L130P encodes a stable protein that does not localize to cilia tips. (a) Immunoblot with KIF7 antibody in cell lysates of wild-type, Kif7 L130P and Kif7
Data are contained in multiple tabs in Excel spreadsheets and in CSV files.
 Contents Overview Curves Methods Measuring enzymatic activity (figure 2) Enzyme characterisation (Figure S1, S2) Enzyme kinetics (Table 3) Effect of ph on activity (figure 3B) Effect of metals and inhibitors
Contents Overview Curves Methods Measuring enzymatic activity (figure 2) Enzyme characterisation (Figure S1, S2) Enzyme kinetics (Table 3) Effect of ph on activity (figure 3B) Effect of metals and inhibitors
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/4/3/eaaq0762/dc1 Supplementary Materials for Structures of monomeric and oligomeric forms of the Toxoplasma gondii perforin-like protein 1 Tao Ni, Sophie I. Williams,
advances.sciencemag.org/cgi/content/full/4/3/eaaq0762/dc1 Supplementary Materials for Structures of monomeric and oligomeric forms of the Toxoplasma gondii perforin-like protein 1 Tao Ni, Sophie I. Williams,
a b G75 G60 Sw-2 Sw-1 Supplementary Figure 1. Structure predictions by I-TASSER Server.
 a b G75 2 2 G60 Sw-2 Sw-1 Supplementary Figure 1. Structure predictions by I-TASSER Server. a. Overlay of top 10 models generated by I-TASSER illustrates the potential effect of 7 amino acid insertion
a b G75 2 2 G60 Sw-2 Sw-1 Supplementary Figure 1. Structure predictions by I-TASSER Server. a. Overlay of top 10 models generated by I-TASSER illustrates the potential effect of 7 amino acid insertion
Imaging energy status in live cells with a fluorescent biosensor of the intracellular ATP-to-ADP. ratio
 Imaging energy status in live cells with a fluorescent biosensor of the intracellular ATP-to-ADP ratio Mathew Tantama, Juan Ramón Martínez-François, Rebecca Mongeon, Gary Yellen* Department of Neurobiology,
Imaging energy status in live cells with a fluorescent biosensor of the intracellular ATP-to-ADP ratio Mathew Tantama, Juan Ramón Martínez-François, Rebecca Mongeon, Gary Yellen* Department of Neurobiology,
BIO 311C Spring Lecture 15 Friday 26 Feb. 1
 BIO 311C Spring 2010 Lecture 15 Friday 26 Feb. 1 Illustration of a Polypeptide amino acids peptide bonds Review Polypeptide (chain) See textbook, Fig 5.21, p. 82 for a more clear illustration Folding and
BIO 311C Spring 2010 Lecture 15 Friday 26 Feb. 1 Illustration of a Polypeptide amino acids peptide bonds Review Polypeptide (chain) See textbook, Fig 5.21, p. 82 for a more clear illustration Folding and
Nature Structural and Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Mutational analysis of the SA2-Scc1 interaction in vitro and in human cells. (a) Autoradiograph (top) and Coomassie stained gel (bottom) of 35 S-labeled Myc-SA2 proteins (input)
Supplementary Figure 1 Mutational analysis of the SA2-Scc1 interaction in vitro and in human cells. (a) Autoradiograph (top) and Coomassie stained gel (bottom) of 35 S-labeled Myc-SA2 proteins (input)
Atypical Natural Killer T-cell receptor recognition of CD1d-lipid antigens supplementary Information.
 Atypical Natural Killer T-cell receptor recognition of CD1d-lipid antigens supplementary Information. Supplementary Figure 1. Phenotypic analysis of TRBV25-1 + and TRBV25-1 - CD1d-α-GalCerreactive cells.
Atypical Natural Killer T-cell receptor recognition of CD1d-lipid antigens supplementary Information. Supplementary Figure 1. Phenotypic analysis of TRBV25-1 + and TRBV25-1 - CD1d-α-GalCerreactive cells.
Supplemental Information. Melanopsin-Encoded Response Properties. of Intrinsically Photosensitive. Retinal Ganglion Cells
 Neuron, Volume 90 Supplemental Information Melanopsin-Encoded Response Properties of Intrinsically Photosensitive Retinal Ganglion Cells Ludovic S. Mure, Megumi Hatori, Quansheng Zhu, James Demas, Irene
Neuron, Volume 90 Supplemental Information Melanopsin-Encoded Response Properties of Intrinsically Photosensitive Retinal Ganglion Cells Ludovic S. Mure, Megumi Hatori, Quansheng Zhu, James Demas, Irene
This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is worth 2 points.
 MBB 407/511 Molecular Biology and Biochemistry First Examination - October 1, 2002 Name Social Security Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is
MBB 407/511 Molecular Biology and Biochemistry First Examination - October 1, 2002 Name Social Security Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is
Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase. Biol 405 Molecular Medicine
 Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase Biol 405 Molecular Medicine 1998 Crystal structure of phenylalanine hydroxylase solved. The polypeptide consists of three regions: Regulatory
Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase Biol 405 Molecular Medicine 1998 Crystal structure of phenylalanine hydroxylase solved. The polypeptide consists of three regions: Regulatory
CS612 - Algorithms in Bioinformatics
 Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/2/4/e1500980/dc1 Supplementary Materials for The crystal structure of human dopamine -hydroxylase at 2.9 Å resolution Trine V. Vendelboe, Pernille Harris, Yuguang
advances.sciencemag.org/cgi/content/full/2/4/e1500980/dc1 Supplementary Materials for The crystal structure of human dopamine -hydroxylase at 2.9 Å resolution Trine V. Vendelboe, Pernille Harris, Yuguang
Nature Biotechnology: doi: /nbt.3828
 Supplementary Figure 1 Development of a FRET-based MCS. (a) Linker and MA2 modification are indicated by single letter amino acid code. indicates deletion of amino acids and N or C indicate the terminus
Supplementary Figure 1 Development of a FRET-based MCS. (a) Linker and MA2 modification are indicated by single letter amino acid code. indicates deletion of amino acids and N or C indicate the terminus
Genetic information flows from mrna to protein through the process of translation
 Genetic information flows from mrn to protein through the process of translation TYPES OF RN (RIBONUCLEIC CID) RN s job - protein synthesis (assembly of amino acids into proteins) Three main types: 1.
Genetic information flows from mrn to protein through the process of translation TYPES OF RN (RIBONUCLEIC CID) RN s job - protein synthesis (assembly of amino acids into proteins) Three main types: 1.
Chapter VI. Increased affinity in the complex of yeast cytochrome c and cytochrome c peroxidase
 Chapter VI Increased affinity in the complex of yeast cytochrome c and cytochrome c peroxidase Chapter VI Abstract We report the study of T12A Cyt c CcP complex using isothermal titration calorimetry (ITC),
Chapter VI Increased affinity in the complex of yeast cytochrome c and cytochrome c peroxidase Chapter VI Abstract We report the study of T12A Cyt c CcP complex using isothermal titration calorimetry (ITC),
Heparin Sodium ヘパリンナトリウム
 Heparin Sodium ヘパリンナトリウム Add the following next to Description: Identification Dissolve 1 mg each of Heparin Sodium and Heparin Sodium Reference Standard for physicochemical test in 1 ml of water, and
Heparin Sodium ヘパリンナトリウム Add the following next to Description: Identification Dissolve 1 mg each of Heparin Sodium and Heparin Sodium Reference Standard for physicochemical test in 1 ml of water, and
T H E J O U R N A L O F C E L L B I O L O G Y
 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Krenn et al., http://www.jcb.org/cgi/content/full/jcb.201110013/dc1 Figure S1. Levels of expressed proteins and demonstration that C-terminal
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Krenn et al., http://www.jcb.org/cgi/content/full/jcb.201110013/dc1 Figure S1. Levels of expressed proteins and demonstration that C-terminal
Introduction to proteins and protein structure
 Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
Crystallization-grade After D After V3 cocktail. Time (s) Time (s) Time (s) Time (s) Time (s) Time (s)
 Ligand Type Name 6 Crystallization-grade After 447-52D After V3 cocktail Receptor CD4 Resonance Units 5 1 5 1 5 1 Broadly neutralizing antibodies 2G12 VRC26.9 Resonance Units Resonance Units 3 1 15 1 5
Ligand Type Name 6 Crystallization-grade After 447-52D After V3 cocktail Receptor CD4 Resonance Units 5 1 5 1 5 1 Broadly neutralizing antibodies 2G12 VRC26.9 Resonance Units Resonance Units 3 1 15 1 5
Nature Neuroscience doi: /nn Supplementary Figure 1. Characterization of viral injections.
 Supplementary Figure 1 Characterization of viral injections. (a) Dorsal view of a mouse brain (dashed white outline) after receiving a large, unilateral thalamic injection (~100 nl); demonstrating that
Supplementary Figure 1 Characterization of viral injections. (a) Dorsal view of a mouse brain (dashed white outline) after receiving a large, unilateral thalamic injection (~100 nl); demonstrating that
Main Functions maintain homeostasis
 The Cell Membrane Main Functions The main goal is to maintain homeostasis. Regulates materials moving in and out of the cell. Provides a large surface area on which specific chemical reactions can occur.
The Cell Membrane Main Functions The main goal is to maintain homeostasis. Regulates materials moving in and out of the cell. Provides a large surface area on which specific chemical reactions can occur.
Supplementary Information Janssen et al.
 Supplementary Information Janssen et al. Insights into complement convertase formation based on the structure of the factor B CVF complex Bert J.C. Janssen 1, Lucio Gomes 1, Roman I. Koning 2, Dmitri I.
Supplementary Information Janssen et al. Insights into complement convertase formation based on the structure of the factor B CVF complex Bert J.C. Janssen 1, Lucio Gomes 1, Roman I. Koning 2, Dmitri I.
Supplementary Materials: An NMR Guided Screening Method for Selective Fragment Docking and Synthesis of a Warhead Inhibitor
 Molecules 216, 21, 846; doi:1.339/molecules217846 S1 of S14 Supplementary Materials: An NMR Guided Screening Method for Selective Fragment Docking and Synthesis of a Warhead Inhibitor Ram B. Khattri, Daniel
Molecules 216, 21, 846; doi:1.339/molecules217846 S1 of S14 Supplementary Materials: An NMR Guided Screening Method for Selective Fragment Docking and Synthesis of a Warhead Inhibitor Ram B. Khattri, Daniel
High-sensitivity Orbitrap mass analysis of intact macromolecular assemblies. R. J. Rose, E. Damoc, E. Denisov, A. Makarov, A. J. R.
 High-sensitivity Orbitrap mass analysis of intact macromolecular assemblies R. J. Rose, E. Damoc, E. Denisov, A. Makarov, A. J. R. Heck SUPPLEMENTARY INFORMATION HCD multipole C -trap Transport octapole
High-sensitivity Orbitrap mass analysis of intact macromolecular assemblies R. J. Rose, E. Damoc, E. Denisov, A. Makarov, A. J. R. Heck SUPPLEMENTARY INFORMATION HCD multipole C -trap Transport octapole
Structural analysis of fungus-derived FAD glucose dehydrogenase
 Structural analysis of fungus-derived FAD glucose dehydrogenase Hiromi Yoshida 1, Genki Sakai 2, Kazushige Mori 3, Katsuhiro Kojima 3, Shigehiro Kamitori 1, and Koji Sode 2,3,* 1 Life Science Research
Structural analysis of fungus-derived FAD glucose dehydrogenase Hiromi Yoshida 1, Genki Sakai 2, Kazushige Mori 3, Katsuhiro Kojima 3, Shigehiro Kamitori 1, and Koji Sode 2,3,* 1 Life Science Research
Nature Methods: doi: /nmeth Supplementary Figure 1
 Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
Multiple-Choice Questions Answer ALL 20 multiple-choice questions on the Scantron Card in PENCIL
 Multiple-Choice Questions Answer ALL 20 multiple-choice questions on the Scantron Card in PENCIL For Questions 1-10 choose ONE INCORRECT answer. 1. Which ONE of the following statements concerning the
Multiple-Choice Questions Answer ALL 20 multiple-choice questions on the Scantron Card in PENCIL For Questions 1-10 choose ONE INCORRECT answer. 1. Which ONE of the following statements concerning the
Nature Methods: doi: /nmeth Supplementary Figure 1. Salipro lipid particles.
 Supplementary Figure 1 Salipro lipid particles. (a) Gel filtration analysis of Saposin A after incubation with the indicated detergent solubilised lipid solutions. The generation of Saposin A-lipid complexes
Supplementary Figure 1 Salipro lipid particles. (a) Gel filtration analysis of Saposin A after incubation with the indicated detergent solubilised lipid solutions. The generation of Saposin A-lipid complexes
Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels. Supplementary Information
 Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels Craig T. Armstrong, Philip E. Mason, J. L. Ross Anderson and Christopher E. Dempsey
Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels Craig T. Armstrong, Philip E. Mason, J. L. Ross Anderson and Christopher E. Dempsey
Characterization of Disulfide Linkages in Proteins by 193 nm Ultraviolet Photodissociation (UVPD) Mass Spectrometry. Supporting Information
 Characterization of Disulfide Linkages in Proteins by 193 nm Ultraviolet Photodissociation (UVPD) Mass Spectrometry M. Montana Quick, Christopher M. Crittenden, Jake A. Rosenberg, and Jennifer S. Brodbelt
Characterization of Disulfide Linkages in Proteins by 193 nm Ultraviolet Photodissociation (UVPD) Mass Spectrometry M. Montana Quick, Christopher M. Crittenden, Jake A. Rosenberg, and Jennifer S. Brodbelt
Biology Open (2014) 000, 1 10 doi: /bio
 (2014) 000, 1 10 doi:10.1242/bio.201410041 Supplementary Material Michael Brauchle et al. doi: 10.1242/bio.201410041 Fig. S1. Alignment of GFP, sfgfp, egfp, eyfp, mcherry and mruby2. Sequence-based alignment
(2014) 000, 1 10 doi:10.1242/bio.201410041 Supplementary Material Michael Brauchle et al. doi: 10.1242/bio.201410041 Fig. S1. Alignment of GFP, sfgfp, egfp, eyfp, mcherry and mruby2. Sequence-based alignment
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 Relationship of the Icelandic psychosis family to five individuals (X1 X5) who carry the haplotype harboring RBM12 c.2377g>t but not RBM12 c.2377g>t. An asterisk marks whole-genome-sequenced
Supplementary Figure 1 Relationship of the Icelandic psychosis family to five individuals (X1 X5) who carry the haplotype harboring RBM12 c.2377g>t but not RBM12 c.2377g>t. An asterisk marks whole-genome-sequenced
d e f Spatiotemporal quantification of subcellular ATP levels in a single HeLa cell during changes in morphology Supplementary Information
 Ca 2+ level (a. u.) Area (a. u.) Normalized distance Normalized distance Center Edge Center Edge Relative ATP level Relative ATP level Supplementary Information Spatiotemporal quantification of subcellular
Ca 2+ level (a. u.) Area (a. u.) Normalized distance Normalized distance Center Edge Center Edge Relative ATP level Relative ATP level Supplementary Information Spatiotemporal quantification of subcellular
Cahn - Ingold - Prelog system. Proteins: Evolution, and Analysis Lecture 7 9/15/2009. The Fischer Convention (1) G (2) (3)
 Chapter 4 (1) G Proteins: Evolution, and Analysis Lecture 7 9/15/2009 A V L I M P F W Chapter 4 (2) S (3) T N Q Y C K R H D E The Fischer Convention Absolute configuration about an asymmetric carbon related
Chapter 4 (1) G Proteins: Evolution, and Analysis Lecture 7 9/15/2009 A V L I M P F W Chapter 4 (2) S (3) T N Q Y C K R H D E The Fischer Convention Absolute configuration about an asymmetric carbon related
Title. Issue Date Doc URL. Type. Additional There Information. File Information. Author(s) Yoshikawa, Shinya; Ishimori, Koichiro
 Title NMR basis for interprotein electron transfer gating Sakamoto, Koichi; Kamiya, Masakatsu; Imai, Mizue; Sh Author(s) Yoshikawa, Shinya; Ishimori, Koichiro CitationProceedings of the National Academy
Title NMR basis for interprotein electron transfer gating Sakamoto, Koichi; Kamiya, Masakatsu; Imai, Mizue; Sh Author(s) Yoshikawa, Shinya; Ishimori, Koichiro CitationProceedings of the National Academy
BCH Graduate Survey of Biochemistry
 BCH 5045 Graduate Survey of Biochemistry Instructor: Charles Guy Producer: Ron Thomas Director: Glen Graham Lecture 10 Slide sets available at: http://hort.ifas.ufl.edu/teach/guyweb/bch5045/index.html
BCH 5045 Graduate Survey of Biochemistry Instructor: Charles Guy Producer: Ron Thomas Director: Glen Graham Lecture 10 Slide sets available at: http://hort.ifas.ufl.edu/teach/guyweb/bch5045/index.html
Supplementary Figure 1. PD-L1 is glycosylated in cancer cells. (a) Western blot analysis of PD-L1 in breast cancer cells. (b) Western blot analysis
 Supplementary Figure 1. PD-L1 is glycosylated in cancer cells. (a) Western blot analysis of PD-L1 in breast cancer cells. (b) Western blot analysis of PD-L1 in ovarian cancer cells. (c) Western blot analysis
Supplementary Figure 1. PD-L1 is glycosylated in cancer cells. (a) Western blot analysis of PD-L1 in breast cancer cells. (b) Western blot analysis of PD-L1 in ovarian cancer cells. (c) Western blot analysis
Nature Structural & Molecular Biology: doi: /nsmb.1933
 The structural basis of open channel block in a prokaryotic pentameric ligand-gated ion channel Ricarda J. C. Hilf, Carlo Bertozzi, Iwan Zimmermann, Alwin Reiter, Dirk Trauner and Raimund Dutzler a GLIC
The structural basis of open channel block in a prokaryotic pentameric ligand-gated ion channel Ricarda J. C. Hilf, Carlo Bertozzi, Iwan Zimmermann, Alwin Reiter, Dirk Trauner and Raimund Dutzler a GLIC
HPLC '88. Poster Presentation. Isolation of Thymosin B4 from Thymosin Fraction 5 by Reverse Phase HPLC
 Essentials in HPLC '88 Poster Presentation Isolation of Thymosin B4 from Thymosin Fraction 5 by Reverse Phase HPLC M. Badamchian, M.P. Strickler, M.J. Stone, A.L. Goldstein for Waters.bioresearchThe absolute,
Essentials in HPLC '88 Poster Presentation Isolation of Thymosin B4 from Thymosin Fraction 5 by Reverse Phase HPLC M. Badamchian, M.P. Strickler, M.J. Stone, A.L. Goldstein for Waters.bioresearchThe absolute,
ATP-independent reversal of a membrane protein aggregate by a chloroplast SRP
 ATP-independent reversal of a membrane protein aggregate by a chloroplast SRP Peera Jaru-Ampornpan 1, Kuang Shen 1,3, Vinh Q. Lam 1,3, Mona Ali 2, Sebastian Doniach 2, Tony Z. Jia 1, Shu-ou Shan 1 Supplementary
ATP-independent reversal of a membrane protein aggregate by a chloroplast SRP Peera Jaru-Ampornpan 1, Kuang Shen 1,3, Vinh Q. Lam 1,3, Mona Ali 2, Sebastian Doniach 2, Tony Z. Jia 1, Shu-ou Shan 1 Supplementary
SUPPLEMENTARY INFORMATION
 doi: 10.1038/nature07422 SUPPLEMENTARY INFRMATIN K S(P) R S I M Q(L4) R M 7 6 Sp Q I K R 5 4 3 2 1 L(L0) L L S E 0 +1 +2 +3 Figure S1a Difference electron density (mfo DFc) for the peptide (Qpeptide),
doi: 10.1038/nature07422 SUPPLEMENTARY INFRMATIN K S(P) R S I M Q(L4) R M 7 6 Sp Q I K R 5 4 3 2 1 L(L0) L L S E 0 +1 +2 +3 Figure S1a Difference electron density (mfo DFc) for the peptide (Qpeptide),
1. to understand how proteins find their destination in prokaryotic and eukaryotic cells 2. to know how proteins are bio-recycled
 Protein Targeting Objectives 1. to understand how proteins find their destination in prokaryotic and eukaryotic cells 2. to know how proteins are bio-recycled As a protein is being synthesized, decisions
Protein Targeting Objectives 1. to understand how proteins find their destination in prokaryotic and eukaryotic cells 2. to know how proteins are bio-recycled As a protein is being synthesized, decisions
Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid.
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 Supplementary Figure 1. SC35M polymerase activity in the presence of Bat or SC35M NP encoded from the phw2000 rescue plasmid. HEK293T
Tivadar Orban, Beata Jastrzebska, Sayan Gupta, Benlian Wang, Masaru Miyagi, Mark R. Chance, and Krzysztof Palczewski
 Structure, Volume Supplemental Information Conformational Dynamics of Activation for the Pentameric Complex of Dimeric G Protein-Coupled Receptor and Heterotrimeric G Protein Tivadar Orban, Beata Jastrzebska,
Structure, Volume Supplemental Information Conformational Dynamics of Activation for the Pentameric Complex of Dimeric G Protein-Coupled Receptor and Heterotrimeric G Protein Tivadar Orban, Beata Jastrzebska,
underlying metastasis and recurrence in HNSCC, we analyzed two groups of patients. The
 Supplementary Figures Figure S1. Patient cohorts and study design. To define and interrogate the genetic alterations underlying metastasis and recurrence in HNSCC, we analyzed two groups of patients. The
Supplementary Figures Figure S1. Patient cohorts and study design. To define and interrogate the genetic alterations underlying metastasis and recurrence in HNSCC, we analyzed two groups of patients. The
Supplementary Figure 1. Normal T lymphocyte populations in Dapk -/- mice. (a) Normal thymic development in Dapk -/- mice. Thymocytes from WT and Dapk
 Supplementary Figure 1. Normal T lymphocyte populations in Dapk -/- mice. (a) Normal thymic development in Dapk -/- mice. Thymocytes from WT and Dapk -/- mice were stained for expression of CD4 and CD8.
Supplementary Figure 1. Normal T lymphocyte populations in Dapk -/- mice. (a) Normal thymic development in Dapk -/- mice. Thymocytes from WT and Dapk -/- mice were stained for expression of CD4 and CD8.
chapter 1 - fig. 2 Mechanism of transcriptional control by ppar agonists.
 chapter 1 - fig. 1 The -omics subdisciplines. chapter 1 - fig. 2 Mechanism of transcriptional control by ppar agonists. 201 figures chapter 1 chapter 2 - fig. 1 Schematic overview of the different steps
chapter 1 - fig. 1 The -omics subdisciplines. chapter 1 - fig. 2 Mechanism of transcriptional control by ppar agonists. 201 figures chapter 1 chapter 2 - fig. 1 Schematic overview of the different steps
Behavioral generalization
 Supplementary Figure 1 Behavioral generalization. a. Behavioral generalization curves in four Individual sessions. Shown is the conditioned response (CR, mean ± SEM), as a function of absolute (main) or
Supplementary Figure 1 Behavioral generalization. a. Behavioral generalization curves in four Individual sessions. Shown is the conditioned response (CR, mean ± SEM), as a function of absolute (main) or
Supplementary Figure 1. Nature Neuroscience: doi: /nn.4547
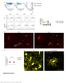 Supplementary Figure 1 Characterization of the Microfetti mouse model. (a) Gating strategy for 8-color flow analysis of peripheral Ly-6C + monocytes from Microfetti mice 5-7 days after TAM treatment. Living
Supplementary Figure 1 Characterization of the Microfetti mouse model. (a) Gating strategy for 8-color flow analysis of peripheral Ly-6C + monocytes from Microfetti mice 5-7 days after TAM treatment. Living
Supplementary Table 1. Properties of lysates of E. coli strains expressing CcLpxI point mutants
 Supplementary Table 1. Properties of lysates of E. coli strains expressing CcLpxI point mutants Species UDP-2,3- diacylglucosamine hydrolase specific activity (nmol min -1 mg -1 ) Fold vectorcontrol specific
Supplementary Table 1. Properties of lysates of E. coli strains expressing CcLpxI point mutants Species UDP-2,3- diacylglucosamine hydrolase specific activity (nmol min -1 mg -1 ) Fold vectorcontrol specific
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3076 Supplementary Figure 1 btrcp targets Cep68 for degradation during mitosis. a) Cep68 immunofluorescence in interphase and metaphase. U-2OS cells were transfected with control sirna
DOI: 10.1038/ncb3076 Supplementary Figure 1 btrcp targets Cep68 for degradation during mitosis. a) Cep68 immunofluorescence in interphase and metaphase. U-2OS cells were transfected with control sirna
Sum of Neurally Distinct Stimulus- and Task-Related Components.
 SUPPLEMENTARY MATERIAL for Cardoso et al. 22 The Neuroimaging Signal is a Linear Sum of Neurally Distinct Stimulus- and Task-Related Components. : Appendix: Homogeneous Linear ( Null ) and Modified Linear
SUPPLEMENTARY MATERIAL for Cardoso et al. 22 The Neuroimaging Signal is a Linear Sum of Neurally Distinct Stimulus- and Task-Related Components. : Appendix: Homogeneous Linear ( Null ) and Modified Linear
SUPPLEMENTARY INFORMATION
 sirna pool: Control Tetherin -HA-GFP HA-Tetherin -Tubulin Supplementary Figure S1. Knockdown of HA-tagged tetherin expression by tetherin specific sirnas. HeLa cells were cotransfected with plasmids expressing
sirna pool: Control Tetherin -HA-GFP HA-Tetherin -Tubulin Supplementary Figure S1. Knockdown of HA-tagged tetherin expression by tetherin specific sirnas. HeLa cells were cotransfected with plasmids expressing
File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description:
 File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description: Supplementary Figure 1. Schematic of Ras biochemical coupled assay.
File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description: Supplementary Figure 1. Schematic of Ras biochemical coupled assay.
