For surface staining, cells were stained with various markers (supplemental table S1) at room
|
|
|
- Hubert Lambert
- 5 years ago
- Views:
Transcription
1 Supplemental materials and methods. Flow cytometry( details) : For surface staining, cells were stained with various markers (supplemental table S1) at room temperature for 15 minutes, washed with PBS and then re-suspended in PBS containing 1:200 dilution of LIVE/DEAD Fixable Near-IR stain. Cells were then incubate for 15 minutes under room temperature and were fixed with 1.5% formaldehyde for 20 minutes under room temperature, washed one time and resuspended in FACS buffer ( PBS with 5% fetal calf serum) before analysis on flow cytometer. For intracellular cytokine staining, PBMC were stimulated with PMA (25ng/ml, Sigma-Aldrich) and Ionomycin (500ng/ml, Sigma-Aldrich) in the presence of monensin (2mM; ebioscience) for 5 hours, or as described below under B10/B10 Pro conditions. Cells were then harvested and were stained with surface markers and then LIVE/DEAD Fixable Near-IR stain (Thermo Fisher Scientific) as described above, with the exception that monensin was added to all the staining buffers. Cells were then fixed with 1.5% formaldehyde for 20 minutes under room temperature, and were then washed twice with permeablization buffer (FACS buffer containing 0.25% Saponin, from Sigma-Aldrich), stained with appropriate cytokine antibodies( supplemental table 1), washed again with permeablization buffer, and were then analyzed by flow cytometer. For intracellular/intranuclear staining of Foxp3 and CTLA4, cells were first stained with surface maker and then labeled with LIVE/DEAD Fixable Near-IR stain (Thermo Fisher Scientific) as described above. Cells were then fixed/permeablized using the Foxp3 / Transcription Factor Staining Buffer Set (ebioscience) according to manufacturer s protocol and were stained with Foxp3 and Foxp3 antibodies (supplemental table S1). All the samples were analyzed using Beckman Coulter Gallios Flow Cytometer, which can detect up to 10 different fluorochrome conjugated antibodies simultaneously.
2 Analysis of IL-10 production by CLL cells. PBMC cells were resuspended ( cells/ml) in in Iscove's Modified Dulbecco's Media (IMDM) containing 10% fetal bovine serum (FBS), 200 μg/ml penicillin, 200 U/mL streptomycin, and 4mM L-glutamine (all from Gibco TM Thermo Fisher ) and stimulated with CpG (ODN 2006, 10 μg/ml; Invivogen), CD40L (1 μg/ml; R&D Systems), PMA (50 ng/ml; Sigma-Aldrich), Ionomycin (1 μg/ml; Sigma-Aldrich), monensin (2mM; ebioscience), as indicated in 48-well flat-bottom plates before staining and flow cytometry analysis. For B10 condition, cells were stimulated with CpG, PMA and Ionomycin in the presence of monensin for 5 hours. For B10 Pro condition, cells were stimulated with CpG/CD40L for 48 hours, with PMA/Ionomycin/ monensin added for the last 5 hours. After stimulation, cells were stained for surface markers, including CD19, CD5, CD3, CD4, and CD8. PECF-594 labeled CD14, CD11b, CD16, CD56 and CD123 were added as a dump channel to gate out corresponding cell types (supplemental table S1). After surface staining, cells were labeled with LIVE/DEAD Fixable Dead Cell Stains from ThermoFisher before being fixed with 1.5% Formaldehyde. Fixed cells were then permeablized with FACS buffer containing 0.25% Saponin and were stained with IL-10 antibody (supplemental table S1). Activation induced cell death in human T cells. : T cells were isolated from healthy human donors using EasySep Human T Cell Isolation Kit. Isolated T cells were stimulated in vitro with plate bound CD3/CD28 for 3 days. Cells were then rested in complete medium containing 50IU/ml IL-2 for additional 7-11 days before they were treated with vehicle, Ibrutinib or acalabrutinib for 30 minutes. Cells were then plated on to 48 well
3 plates coated with CD3; incubate for 6 hours (for flow cytometry based apoptosis assay) or 3 hours (to isolate mrna for qpcr to quantify FAS-L expression.) in the presence of IL2 to induce AICD. For AICD analysis, cells were stained with annexin-v fitc and Propidium Iodide (PI) using the BD biosciences 10X staining buffer according to the manufacturer s protocol before being analyzed on flow cytometer. For FAS Ligand mrna quantification, mrna were extracted from T cells after 3 hours of restimulation using QIAGEN RNeasy Mini RNA Isolation Kit. mrna was then reverse transcribed to cdna using the M-MLV Reverse Transcriptase from Thermo Fisher. Quantitative PCR for FAS- L were performed using the Taqman probe/primer mix (FAM labeled) from Thermo Fisher using GAPDH as internal control. Activation induced cell death in human NK cells Human CD56 + /CD3 - /14 - /20 - NK cells were isolated from peripheral blood leuko-paks from normal donors (American Red Cross) by incubation with an NK cell RosetteSep negative enrichment cocktail (Stem Cell Technology), followed by Ficoll-Hypaque density gradient centrifugation as previously described(96). NK cells were then sorted to greater than 99% purity with a FACSAria II cell sorter (BD Biosciences). Purified NK cells were plated at 5x10 4 cells/well in a 96-well round bottom plate and cultured for three days at 37 C. Medium consisted of RPMI 1640 supplemented with 10% fetal bovine serum (FBS), and 1% antibiotic/antimycotic (Life Technologies). The cytokines IL-2 (Peprotech) and IL-15 (National Cancer Institute) were supplemented as indicated for a final concentration of 10ng/mL. IL-12 (Miltenyi Biotec) was added where indicated at a concentration of 10ng/mL to induce activation induced cell death. Cell viability and apoptosis were assessed after three days in culture by annexin V (BD Biosciences) apoptotic and TO-PRO-3 (Molecular Probes) viability flow cytometric analysis(97). NK cells were harvested and stained with annexin V per manufacturer s instructions (BD
4 Biosciences). TO-PRO-3 was added immediately prior to acquisition, and all samples were analyzed with a LSRII cytometer (BD Biosciences) within one hour of annexin V staining. Analysis of dual staining of annexin V and TO-PRO-3 was analyzed using FlowJo (TreeStar).
5 Figure S1 A. Patients treated with Ibrutinib Total CD8+ T cells Naïve CD8+ T cells Central memory (CM) CD8+ T cells Effector memory (EM) CD8+ T cells CD45RA+ EM ( EMRA ) CD8+ T cells % total PBMC P=0.066 % total CD8 T cells P=0.046 P=0.006 P=0.017 % total PBMC Total CD4+ T cells P=0.051 % total CD4 T cells Naïve CD4+ T cells Central memory (CM) CD4+ T cells P=0.001 P=0.015 Effector memory (EM) CD4+ T cells P=0.102 CD45RA+ EM ( EMRA ) CD4+ T cells P=0.106 P=0.010 B. Patients treated with acalabrutinib total CD8+ T cells Naïve CD8+ T cells Central memory (CM) CD8+ T cells Effector memory (EM) CD8+ T cells CD45RA+ EM ( EMRA ) CD8+ T cells P=0.032 % total PBMC % total CD8 T cells % total PBMC total CD4+ T cells P=0.024 % total CD4 T cells Naïve CD4+ T cells Central memory (CM) CD4+ T cells Effector memory (EM) CD4+ T cells CD45RA+ EM ( EMRA ) CD4+ T cells Figure S1: The effect of ibrutinib or acalabrutinib treatment on the frequency of different subsets of peripheral T cells. (A). Percentage of different T cell subsets among total CD8 T cells( upper panel) and CD4 T cells (lower panel) before and after ibrutinib Treatment (n=18). (B). Percentage of different T cell subsets among total CD8 T cells( upper panel) and CD4 T cells (lower panel) before and after acalabrutinib treatment (n=12). T cells are differentiated into subsets based on their expression of CCR7 and CD45RA: naïve T cells (CCR7+CD45RA+), central memory T cells (CCR7+CD45RA-), effector memory T cells (CCR7-CD45RA-), and most differentiated effector memory T cells (T- EMRA, CCR7-CD45RA+).
6 Figure S2 PI A. B. C. vehicle Ibrutinib Dead Dead apoptotic apoptotic % Apoptotic+Necrotic cells P<0.001 P<0.001 Annexin V Fitc D. E. % Apoptotic+Necrotic cells P<0.05 % Apoptotic+Necrotic cells P<0.05 P<0.05 Figure S2: Ibrutinib treatment of human T cells or NK cells protects against activation induced cell death in a dose dependent manor. (A)- (C), T cells were isolated from healthy human donors blood samples, stimulated in vitro with CD3/CD28 for 3 days, rested in culture medium containing 50 IU IL-2 for 11 days, then were restimulated with plate bound CD3 for 6 hours (as in A. and B.) or 3 hours (as in C.) in the presence of IL2 to induce activation induced cell death in the presence of absence of ibrutinib. Each indicated condition. (A). representative FACS dot plots of Annexin V & PI staining. (B). Bar graphs that show the percentage of non-viable was done in triplicate. Data shown are representative of three independent experiments (apoptotic + necrotic, as defined by Annexin V positive and PI positive cells) cells after induction of AICD. (C). FAS-L mrna upregulation in activated T cells upon induction of AICD was impaired by ibrutinib treatment. mrna was isolated from the T cells after induction of AICD, cdna was synthesized and qpcr for FAS-L and GAPDH was done. Figure A, B and C represent 3 independent experiments. (D) & (E), Human CD56 + /CD3 - /14 - /20 - NK cells were isolated from peripheral blood from normal donors ( N=3) by negative enrichment, and were then sorted to greater than 99% purity by FACSAria II sorter. Purified NK cells were plated at 5x10 4 cells/well and were cultured for three days. IL-15 (D) and IL-2 (E) were added as indicated for a final concentration of 10ng/mL. IL-12 (Miltenyi Biotec) was added where indicated at a concentration of 10ng/mL to induce activation induced cell death. Bar graphs that show the percentage of non-viable (apoptotic + necrotic, as defined by Annexin V positive and TO-PRO-3 positive cells) cells after induction of AICD. (N=3)
7 Figure S3 A. Patients treated with Ibrutinib Total CD4 T cells Naïve CD4+ cells CM CD4+ cells EM CD4+ cells EMRA CD4+ cells Percentage of PD1+ cells P=0.001 P=0.009 P<0.001 P=0.001 P<0.001 P=0.042 Percentage of PD1+ cells EM CD4+ cells (CD27+) EM CD4+ cells (CD27-) EMRA CD4+ cells (CD27+) EMRA CD4+ cells (CD27-) P<0.001 P=0.014 P<0.001 B. Patients treated with acalabrutinib Percentage of PD1+ cells Total CD4 T cells Naïve CD4+ cells CM CD4+ cells EM CD4+ cells EMRA CD4+ cells P=0.099 P=0.002 P<0.001 P=0.023 P<0.001 P=0.018 P=0.007 P=0.002 P=0.097 Percentage of PD1+ cells EM CD4+ cells (CD27+) EM CD4+ cells (CD27-) EM RA CD4+ cells (CD27+) EMRA CD4+ cells (CD27-) P=0.005 P<0.001 P=0.026 P=0.078 Figure S3: Treatment with ibrutinib, as well as with acalabrutinib, leads to a significant reduction in the frequency of PD-1 positive cells in CD4 T cell populations. (A). Percentage of PD1 positive cells among different subsets of CD4 T cells from CLL patients before and after ibrutinib treatment. (n=17) (B). Percentage of PD1 positive cells among different subsets of CD4 T cells from CLL patients before and after acalabrutinib treatment. (n=10)
8 Figure S4. A. Patients treated with Ibrutinib % CTLA-4 positive Cells among total CD8 T cells % CTLA-4 positive Cells among CD45RA- CD8 T cells % CTLA-4 positive Cells among CD45RA+ CD8 T cells Percentage of CTLA-4+ cells P= P=0.004 P=0.006 P=0.007 P=0.038 P=0.027 B. Patients treated with acalabrutinib % CTLA-4 positive Cells among total CD8 T cells % CTLA-4 positive Cells among CD45RA- CD8 T cells % CTLA-4 positive Cells among CD45RA+ CD8 T cells Percentage of CTLA-4+ cells P=0.002 P=0.005 P=0.010 P=0.004 P=0.002 P=0.002 Figure S4: Treatment with ibrutinib, as well as with acalabrutinib, leads to a significant reduction in the frequency of intracellular CTLA4 positive cells in CD8 T cell populations. (A). Percentage of CTLA4 (intracellular) positive cells among total CD8 T cells, CD45RA- CD8 T cells and CD45RA+ CD8 T cells from CLL patients before and after ibrutinib treatment.(n=18). (B).Percentage of CTLA4 (intracellular) positive cells among total CD8 T cells, CD45RA- CD8 T cells and CD45RA+ CD8 T cells from CLL patients before and after acalabrutinib treatment.(n=9).
9 Figure S5 IL2/TNFa CD4+ IL4/IL-17 CD4+ IL2/IFNg CD4+ 6 month ibrutinib Tx Before ibrutinib Tx Figure S5: Representative FACS plot of intracellular cytokine staining( n=15). PBMC from CLL patient before and after ibrutinib treatment were stimulated with in vitro with PMA/Ionomycin in the presence of Monensin for 5 hours. Cytokine production (IFNγ, IL4, IL17, TNFα and IL2) were detected by intracellular cytokine staining.
10 Figure S6 All CD8+ T cells Naïve CD8+ T cells Central memory (CM) CD8+ T cells Effector memory (EM) CD8+ T cells CD45RA+ EM ( EMRA ) CD8+ T cells % PD1+ positive cells % PD1+ positive cells % PD1+ positive cells % PD1+ positive cells % PD1+ positive cells % PD1+ positive cells % PD1+ positive cells % PD1+ positive cells % PD1+ positive cells CD27+ Effector Memory CD8+ T cells CD27- Effector Memory CD8+ T cells CD27+ EMRA CD8+ T cells CD27- EMRA CD8+ T cells Healthy volunteer (HV) CLL patient, baseline Figure S6: PD-1 expression is increased in all of the T cell subsets in CLL patients comparing to healthy donors. The increase is most prominent in naïve and central memory T cell compartment. Frequencies of PD-1 positive cells among different CD8 T cell subsets were shown. n=11 for healthy donor, n=15 for CLL patients.
11 Figure S7: Short term Ibrutinib treatment (2 or 4 days) does not increase circulating T cell numbers. Wild type B6 mice were engrafted with CLL cells ( splenocytes from leukemic Eμ-TCL1 transgenic mice). seven weeks post leukemia engraftment, the mice were treated with ibrutinib and the circulating T cell numbers in peripheral blood were monitored before starting ibrutinib, 2 days and 4 days post starting ibrutinib( corresponding to the time when CLL cells numbers were transiently increased in peripheral blood in this model(1)). N=14.
12 Figure S8: Ibrutinib treatment increases the number of activated leukemia specific T cells. Mice were engrafted with AML cell line (C1498) expressing OVA (a model antigen). OT-1 transgenic T cells ( recognize OVA) were then adoptively transferred into AML engrafted mice. The mice were treated with Ibrutinib versus vehicle. Mice were sacrificed at day 6, spleens were harvested, the frequency and number of leukemia specific OT-1 T cells were counted and plotted. N=7 for each group.
13 Figure S9 Figure S9: Human CD56+/CD3-/14-/20- NK cells were isolated from peripheral blood from normal donors by negative enrichment, and were then sorted to greater than 99% purity by FACSAria II sorter. Purified NK cells were plated at 5x104 cells/well and were cultured for three days. IL-15 and IL-2 were added as indicated for a final concentration of 10ng/mL. IL-12 (Miltenyi Biotec) was added where indicated at a concentration of 10ng/mL to induce activation induced cell death. Vehicle control versus acalabrutinib 500nM, acalabrutinib 1000nM were added to rescue cytokine induced NK cell deah.. Bar graphs that show the percentage of non-viable (apoptotic + necrotic, as defined by Annexin V positive and TO-PRO-3 positive cells) cells after induction of AICD. ACP: acalabrutinib ( ACP-196)
14 Table S1: IC50 values for inhibition of enzymatic activity by ibrutinib versus acalabrutinib. Kinase IC50 (nm) of acalabrutinib IC50 (nm) of ibrutinib BTK 5.1 ± ± 0.2 BMX 46 ± ± 0.1 ITK > ± 1.2 TEC 93 ± 35 7 ± 2.5 TXK 368 ± ± 0.3 EGFR > ± 1.3 ERBB2 ~ ± 1.8 ERBB4 16 ± ± 1.3 JAK3 > ± 15 BLK > ± 0.0 FGR > ± 1.1 FYN > ± 0 HCK > ± 0 LCK > ± 1.3 LYN > ± 1 SRC > ± 1 YES1 > ± 0.2 CSK BRK FLT
15 Exp Table S2: detailed patient information gender time of treatment year of diagnosis Rai stage at the time of treatment 1 F 10/11/ line of prior Tx Del 13 Del 11q Del 17q Kipps Regimen x1 cycle March 2013 Cygoegenetics/FISH results Trisomy 12 6q21 2p12 14q32 8q24 complex karyotype? IgVH mutation status (%) % bone marrow invovlement baseline ALC cycle 3 ALC Cycle 6 ALC age at treatment positive positive no 7.6 > age at diagnosis 2 M 7/9/ BR x 6, 7/ /2012 5/16/13: Ofatumumab + Dinaciclib on protocol Aa by 2015: pancytopenia, platelet transfusion dependent no M 1/18/ FCR X6 finished by 8/ yes F 4/1/ FR X8 by 8/2012, Rituxan + steroid for AIHA no no no no 73.2 yes F 10/29/ /4 1st: chlorambucil from May 2005 through June 2006 with rituximab given in spring She had a partial response, lasted 2 years (progressive lymphocytosis and abdominal pain and GI symptoms with bulky adenopathy in the abdomen) 2nd: chlorambucil in October 2008 given with prednisone in February No response 3rd: PCR (pentostatin, cyclophosphamide, rituximab) for two cycles in March and April this was complicated by pneumonia. Treatment free for one year. 4th: rituximab weekly x 4 (March April 2010), progressive lymphocytosis, bulky abdominal nodes an splenomegaly, six months 5th: CVP for one cycle in October 2010, then R-CVP on 11/17/2010. complicated by jaw pain and constipation she had decreased adenopathy though. Treatment free for 3 months,, 6th bendamustine + rituximab x 2 cycles February & March th: High dose methylprednisone + rituximab June - July pretreatment - WBC 208.3, Hgb 8.4, Platelets 70,000, following therapy on 7/23/12, she had WBC 113.7, hemoglobin 10.5, and platelets 68. She had increasing adenopathy. 8th: Ofatumumab: 8 weekly doses from July to Sep Last dose was 09/24/ no F 6/11/ Ofatumumab 1-3/2012. per no 0.3 > M 7/24/ ofatumumab starting 10/2011 and received 8 weeks followed by 1 maintenance dose in February no 3.68 > F 7/18/ March 2011 weekly x 4 rituximab ending 3/30/ yes M 10/21/ no FCR + Campath X6 CR 2/ Rit + Revlamid 3 cycles - pneumonia #1: chlorambucil, prednisone, and fludarabine. #2: 9 cycles of CHOP which led to reduction in the number of leukemic cells in his bone marrow. #3: allo SCT in 03/1997. remission for about 9 years till 2006, 10 M 10/17/ no #4: single agent Rituxan for about 2 years, and subsequently with rituxan-bendamustine combination. #5: Revlimid, at first at 10mg every other day. #6: January 2011, briefly treated on study with ofatumumab,,pancytopenia and sepsis after 1st dose he had a good hematologic response 11 M 9/26/ M 11/5/ cycles of R-CVP, completing in April 2010 in Tampa Florida. He returned to Dayton, OH and had a bone marrow biopsy because of thrombocytopenia in July 2010 which showed persistent CLL involvement of somewhere between 30-50%. He was feeling well with normal counts and continued to be monitored. His WBC count 5/2011 had risen to the 30,000 range and his physician repeated his BM biopsy which showed 80% cellularity with 70% involvement by CLL cycles of BR from Aug 2011 to Jan 2012 at local facility 3. 4 weekly Rituxan and prednisone in June 2012 due to AIHA * Radiation: to cervical LAD in 2004 * Rituxan: 4 weekly treatments in 2004 * Rituxan: 4 weekly treatments in 2005 * Chlorambucil: * Rituxan: on/off along with Chlorambucil * FCR: x 6 cycles in CR * Rituxan/solumedrol: 08/2012 * Revlimid: on study in Jacksonville, FL at Mayo clinic 09/07/12-10/01/ yes no FCR x 6 cycles: 6/2003 through 11/2003; best response = CR 13 M 11/7/ yes R-lenalinomide: 6/2008 through 12/2008; discontinued secondary to severe thrombocytopenia 3. FCR x 4 cycles 2009/ Kipps regimen (solu-medrol, rituximab) x 2 cycle starting 8/20/ fludarabine x 6 in April fludarabine x 2 in October rituximab weekly x 4 in early M 12/4/ yes rituximab weekly in May rituixmab weekly in January rituximab + fludarabine on an abbreviated schedule x 3 in 7/ fludarabine + rituximab +Neulasta x 4 cycles in 2010, - rituximab weekly in October F 2/20/ FCR done in 2008, maintenance R X 6 months. BR starting 8/2011, only 1 cycle. ofatumumab locally from Jan to July yes F 1/28/ FC X 1, FCR x 1 (cytopenias, rituximab infusion reaction) 2. CVP x 6 --> rituximab weekly x 4 3. FCR x 4 (completed 3/2006) no BR + CAL101 (5/2011), discontinued secondary to rash 5. Ofatumumab (7-12/2011) 7/16/ /2001 Fludarabine x 6. Nearly achieved CR. 17 M 3/25/ yes /2004: FCR x 4 for progressive lymphocytosis to prepare for allosct - dose reduction of cyclophosphamide and fludarabine secondary to cytopenias. Achieved significant cytoreduction. 9/2004: RIC flu/bu/tbi allosct from matched sibling (brother), Some rash, but unclear if GVHD. 10/05: Bone marrow showed CR 18 F 7/2/ M 6/25/ > FCR x 6 cycles in 8/2004 1/2005: eight weekly doses of rituximab PR. 10/2006 to 4/2007 with eight treatments two times a month. fenretinide from July of 2007 to January of 2008 on a clinical trial with four doses of rituximab from 7-8/2007 and then continued on oral fenretinide. 6 cycles of FCR from March to August no 6.1 > no
16 Table S3, list of antibodies used. Antigen fluorochrome vendor Catalog # clone BTLA PE Biolegend MIH26 CD122 BV421 BD Biosciences Mik-β3 CD123 PE Efluor 610 ebiosciences H6 CD14 PE-CF594 BD Biosciences MφP9 CD16 FITC ebiosciences ebiocb16 (CB16) CD160 APC Biolegend BY55 CCR7 PE Cy7 Biolegend G043H7 CD11b PerCP- Eflour710 ebiosciences ICRF44 CD11c PE Efluor 610 ebiosciences CD160 PE Cy7 Biolegend BY55 CD180 MD1 PE ebiosciences MHR73-11 CD19 PerCP- Eflour710 ebiosciences SJ25C1 CD19 APC-R700 BD Biosciences HIB19, CD19 BB515 BD Biosciences HIB19 CD2 fitxc BD Biosciences S5.2 CD200 APC ebiosciences OX104 CD24 PE-CF594 BD Biosciences ML5 CD244 PE C1.7 CD244 PerCP-Cy5.5 biolegend C1.7 CD25 PE Cy7 ebiosciences BC96 CD25 PE cy7 ebiosciences BC96 CD26 PE-CF594 BD Biosciences M-A261 CD26 PerCp cy5.5 biolgend BA5b CD27 PE BD Biosciences M-T271 CD27 efluor 450 ebiosciences O323 CD27 BB515 BD Biosciences M-T271 CD28 PE ebiosciences CD28.2 CD290 TLR10 PE ebiosciences C10C5 CD3 APC-R700 BD Biosciences SK7 (Leu-4) CD31 APC ebiosciences WM59 CD33 APC ebiosciences WM-53 CD38 PE Cy7 ebiosciences HB7 CD4 Apc R700 BD Biosciences RPA-T4 CD4 BB515 BD Biosciences RPA-T4 CD45RA PerCP-Cy5.5 BD Biosciences HI100 CD5 BV510 BD Biosciences UCHT2
17 CD57 PE-CF594 BD Biosciences NK-1 CD62L BV510 BD Biosciences DREG-56 CD7 BB515 BD Biosciences M-T701 CD8 BV510 Biolegend SK1 CD8 PECF594 BD Biosciences RPA-T8 CD8 PE-Cy7 ebiosciences SK1 CD9 PerCp cy5.5 BD Biosciences M-L13 CD95 BB515 BD Biosciences DX2 CTLA4 BV421 BD Biosciences BNI3 CTLA4 PE-CF594 BD Biosciences BNI3 EOMES PE ebiosciences WD1928 FCRL3 BB515 BD Biosciences H5 Foxp3 PE BD Biosciences D/C7 HLA-A2 PECy7 ebiosciences BB7.2 HLADR BV421 BD Biosciences G46-6 IDO eflour 660 ebiosciences eyedio IFNg PerCP-Cy5.5 ebiosciences S.B3 IgD BV421 BD Biosciences IA6-2 IgG Alexa700 BD Biosciences G IgM APC BD Biosciences G IL-10 PE ebiosciences JES3-9D7 IL-17A APC ebiosciences ebio64dec17 IL2 BV421 BD Biosciences IL-4 PE BD Biosciences D4-8 KLRG1 BV421 biolgend F1 LAG3 PE Cy7 ebiosciences DS223H LAG3 APC ebiosciences DS223H PD1 BB515 BD Biosciences EH12.1 PD1 BV421 BD Biosciences MIH18 PTK7 PE miltenyibiotec B Tim3 APC ebiosciences F38-2E2 Tim3 BV421 Biolgend F38-2E2 TNFa PE Cy7 ebiosciences MAb11 TNFa efluor 450 ebiosciences MAb11
18 Table S4. P-values from initial and updated cohorts An initial analysis was performed using data from 17 patients treated with ibrutinib and 9 patients treated with acalabrutinib. Later, an additional 2 patients with ibrutinib and 4 patients with acalabrutinib were added to the original cohorts and the analysis was redone; this second analysis was not planned at the time of the first analysis. P-values from the initial and new analyses are shown below for comparison purposes. Note that in the original analysis, p-values within each figure were adjusted for multiple comparisons using Hochberg s procedure, while unadjusted p-values are presented in the new analysis. Not all experiments were performed on each patient s serial sample, therefore the actual N for each experiment was less than 19 and 13 for ibrutinib and acalabrutinib treated patients, respectively. Figure 1A (ibrutinib): CD8+ T cells 1A (ibrutinib): CD4+ T cells 1B (acalabrutinib): CD8+ T cells 1B (acalabrutinib): CD4+ T cells Comparison Initial Cohort Updated Cohort N P-value N P-value Total CD8+ T cells: cycle 3 vs baseline <.001 Total CD8+ T cells: cycle 6 vs baseline Naïve CD8+ T cells: cycle 3 vs baseline <.001 Naïve CD8+ T cells: cycle 6 vs baseline CM CD8+ T cells: cycle 3 vs baseline CM CD8+ T cells: cycle 6 vs baseline EM CD8+ T cells: cycle 3 vs baseline <.001 EM CD8+ T cells: cycle 6 vs baseline EMRA CD8+ T cells: cycle 3 vs baseline 16 < <.001 EMRA CD8+ T cells: cycle 6 vs baseline 16 < Total CD8+ T cells: cycle 3 vs baseline <.001 Total CD8+ T cells: cycle 6 vs baseline Naïve CD8+ T cells: cycle 3 vs baseline Naïve CD8+ T cells: cycle 6 vs baseline CM CD8+ T cells: cycle 3 vs baseline CM CD8+ T cells: cycle 6 vs baseline EM CD8+ T cells: cycle 3 vs baseline EM CD8+ T cells: cycle 6 vs baseline EMRA CD8+ T cells: cycle 3 vs baseline <.001 EMRA CD8+ T cells: cycle 6 vs baseline 16 < <.001 Total CD8+ T cells: cycle 3 vs baseline Total CD8+ T cells: cycle 6 vs baseline Naïve CD8+ T cells: cycle 3 vs baseline Naïve CD8+ T cells: cycle 6 vs baseline CM CD8+ T cells: cycle 3 vs baseline CM CD8+ T cells: cycle 6 vs baseline EM CD8+ T cells: cycle 3 vs baseline EM CD8+ T cells: cycle 6 vs baseline EMRA CD8+ T cells: cycle 3 vs baseline EMRA CD8+ T cells: cycle 6 vs baseline Total CD8+ T cells: cycle 3 vs baseline Total CD8+ T cells: cycle 6 vs baseline Naïve CD8+ T cells: cycle 3 vs baseline Naïve CD8+ T cells: cycle 6 vs baseline CM CD8+ T cells: cycle 3 vs baseline
19 Figure 2C (ibrutinib): Absolute cell # 2C (ibrutinib): Percentage 3A (ibrutinib): CD8 T cell subsets 3B (acalabrutinib): CD8 T cell subsets 4A (ibrutinib): CTLA4 Comparison Initial Cohort Updated Cohort N P-value N P-value CM CD8+ T cells: cycle 6 vs baseline EM CD8+ T cells: cycle 3 vs baseline EM CD8+ T cells: cycle 6 vs baseline EMRA CD8+ T cells: cycle 3 vs baseline EMRA CD8+ T cells: cycle 6 vs baseline Cycle 3 vs. baseline Cycle 6 vs. baseline Cycle 3 vs. baseline Cycle 6 vs. baseline Total CD8 T cells: cycle 3 vs baseline Total CD8 T cells: cycle 6 vs baseline <.001 Naïve CD8+ T cells: cycle 3 vs baseline Naïve CD8+ T cells: cycle 6 vs baseline T-CM CD8+ cells: cycle 3 vs baseline T-CM CD8+ cells: cycle 6 vs baseline <.001 T-EM CD8+ cells: cycle 3 vs baseline T-EM CD8+ cells: cycle 6 vs baseline <.001 T-EMRA CD8+ cells: cycle 3 vs baseline T-EMRA CD8+ cells: cycle 6 vs baseline T-EM CD8+ cells (CD27+): cycle 3 vs baseline T-EM CD8+ cells (CD27+): cycle 6 vs baseline T-EM CD8+ cells (CD27-): cycle 3 vs baseline T-EM CD8+ cells (CD27-): cycle 6 vs baseline <.001 T-EMRA CD8+ cells (CD27+): cycle 3 vs baseline T-EMRA CD8+ cells (CD27+): cycle 6 vs baseline T-EMRA CD8+ cells (CD27-): cycle 3 vs baseline T-EMRA CD8+ cells (CD27-): cycle 6 vs baseline Total CD8 T cells: cycle 3 vs baseline Total CD8 T cells: cycle 6 vs baseline <.001 Naïve CD8+ T cells: cycle 3 vs baseline Naïve CD8+ T cells: cycle 6 vs baseline T-CM CD8+ cells: cycle 3 vs baseline T-CM CD8+ cells: cycle 6 vs baseline <.001 T-EM CD8+ cells: cycle 3 vs baseline T-EM CD8+ cells: cycle 6 vs baseline T-EMRA CD8+ cells: cycle 3 vs baseline T-EMRA CD8+ cells: cycle 6 vs baseline T-EM CD8+ cells (CD27+): cycle 3 vs baseline T-EM CD8+ cells (CD27+): cycle 6 vs baseline T-EM CD8+ cells (CD27-): cycle 3 vs baseline T-EM CD8+ cells (CD27-): cycle 6 vs baseline T-EMRA CD8+ cells (CD27+): cycle 3 vs baseline T-EMRA CD8+ cells (CD27+): cycle 6 vs baseline <.001 T-EMRA CD8+ cells (CD27-): cycle 3 vs baseline T-EMRA CD8+ cells (CD27-): cycle 6 vs baseline Total CD4 T cells: cycle 3 vs baseline <.001 Total CD4 T cells: cycle 6 vs baseline 16 < <.001 CD45RA- CD4 T cells: cycle 3 vs baseline <.001 CD45RA- CD4 T cells: cycle 6 vs baseline 16 < <.001
20 Figure Comparison Initial Cohort Updated Cohort N P-value N P-value CD45RA+ CD4 T cells: cycle 3 vs baseline CD45RA+ CD4 T cells: cycle 6 vs baseline <.001 4B (acalabrutinib): Total CD4 T cells: cycle 3 vs baseline CTLA4 Total CD4 T cells: cycle 6 vs baseline CD45RA- CD4 T cells: cycle 3 vs baseline CD45RA- CD4 T cells: cycle 6 vs baseline CD45RA+ CD4 T cells: cycle 3 vs baseline CD45RA+ CD4 T cells: cycle 6 vs baseline A (ibrutinib): IFNγ: cycle 3 vs. baseline cytokines IFNγ: cycle 6 vs. baseline TNFα: cycle 3 vs. baseline TNFα: cycle 6 vs. baseline IL-2: cycle 3 vs. baseline IL-2: cycle 6 vs. baseline IL-4: cycle 3 vs. baseline IL-4: cycle 6 vs. baseline IL-17: cycle 3 vs. baseline IL-17: cycle 6 vs. baseline B (acalabrutinib): IFNγ: cycle 3 vs. baseline cytokines IFNγ: cycle 6 vs. baseline TNFα: cycle 3 vs. baseline TNFα: cycle 6 vs. baseline IL-2: cycle 3 vs. baseline IL-2: cycle 6 vs. baseline IL-4: cycle 3 vs. baseline IL-4: cycle 6 vs. baseline IL-17: cycle 3 vs. baseline IL-17: cycle 6 vs. baseline B (ibrutinib): Percentage: cycle 3 vs. baseline 16 < <.001 Foxp3+ cells Percentage: cycle 6 vs. baseline 16 < <.001 Absolute number: cycle 3 vs. baseline Absolute number: cycle 6 vs. baseline C (acalabrutinib): Percentage: cycle 3 vs. baseline Foxp3+ cells Percentage: cycle 6 vs. baseline Absolute number: cycle 3 vs. baseline Absolute number: cycle 6 vs. baseline A (ibrutinib): CD200, BTLA 7B (acalabrutinib): CD200, BTLA 8C (ibrutinib): IL10+ cells CD200: Cycle 3 vs. baseline 16 < <.001 CD200: Cycle 6 vs. baseline 16 < <.001 BTLA: Cycle 3 vs. baseline 14 < <.001 BTLA: Cycle 6 vs. baseline 14 < <.001 CD200: Cycle 3 vs. baseline CD200: Cycle 6 vs. baseline <.001 BTLA: Cycle 3 vs. baseline 8 < <.001 BTLA: Cycle 6 vs. baseline 8 < < hours (B10): Cycle 3 vs. baseline hours (B10): Cycle 6 vs. baseline hours (B10 Pro): Cycle 3 vs. baseline 16 < < hours (B10 Pro): Cycle 6 vs. baseline 16 < <.001
21 Figure 8C (acalabrutinib): IL10+ cells S1A (ibrutinib): CD8+ T cells S1A (ibrutinib): CD4+ T cells S1B (acalabrutinib): CD8+ T cells S1B (acalabrutinib): CD4+ T cells S3A (ibrutinib): CD4 T cell subsets Comparison Initial Cohort Updated Cohort N P-value N P-value 5 hours (B10): Cycle 3 vs. baseline hours (B10): Cycle 6 vs. baseline hours (B10 Pro): Cycle 3 vs. baseline < hours (B10 Pro): Cycle 6 vs. baseline <.001 Total CD8+ T cells: cycle 3 vs baseline Total CD8+ T cells: cycle 6 vs baseline Naïve CD8+ T cells: cycle 3 vs baseline Naïve CD8+ T cells: cycle 6 vs baseline CM CD8+ T cells: cycle 3 vs baseline CM CD8+ T cells: cycle 6 vs baseline EM CD8+ T cells: cycle 3 vs baseline EM CD8+ T cells: cycle 6 vs baseline EMRA CD8+ T cells: cycle 3 vs baseline EMRA CD8+ T cells: cycle 6 vs baseline Total CD8+ T cells: cycle 3 vs baseline Total CD8+ T cells: cycle 6 vs baseline Naïve CD8+ T cells: cycle 3 vs baseline Naïve CD8+ T cells: cycle 6 vs baseline CM CD8+ T cells: cycle 3 vs baseline CM CD8+ T cells: cycle 6 vs baseline EM CD8+ T cells: cycle 3 vs baseline EM CD8+ T cells: cycle 6 vs baseline EMRA CD8+ T cells: cycle 3 vs baseline EMRA CD8+ T cells: cycle 6 vs baseline Total CD8+ T cells: cycle 3 vs baseline Total CD8+ T cells: cycle 6 vs baseline Naïve CD8+ T cells: cycle 3 vs baseline Naïve CD8+ T cells: cycle 6 vs baseline CM CD8+ T cells: cycle 3 vs baseline CM CD8+ T cells: cycle 6 vs baseline EM CD8+ T cells: cycle 3 vs baseline EM CD8+ T cells: cycle 6 vs baseline EMRA CD8+ T cells: cycle 3 vs baseline EMRA CD8+ T cells: cycle 6 vs baseline Total CD8+ T cells: cycle 3 vs baseline Total CD8+ T cells: cycle 6 vs baseline Naïve CD8+ T cells: cycle 3 vs baseline Naïve CD8+ T cells: cycle 6 vs baseline CM CD8+ T cells: cycle 3 vs baseline CM CD8+ T cells: cycle 6 vs baseline EM CD8+ T cells: cycle 3 vs baseline EM CD8+ T cells: cycle 6 vs baseline EMRA CD8+ T cells: cycle 3 vs baseline EMRA CD8+ T cells: cycle 6 vs baseline Total CD4 T cells: cycle 3 vs baseline Total CD4 T cells: cycle 6 vs baseline Naïve CD4+ T cells: cycle 3 vs baseline Naïve CD4+ T cells: cycle 6 vs baseline T-CM CD4+ cells: cycle 3 vs baseline
22 Figure S3B (acalabrutinib): CD4 T cell subsets S4A (ibrutinib): CTLA4 S4B (acalabrutinib): CTLA4 Comparison Initial Cohort Updated Cohort N P-value N P-value T-CM CD4+ cells: cycle 6 vs baseline <.001 T-EM CD4+ cells: cycle 3 vs baseline T-EM CD4+ cells: cycle 6 vs baseline T-EMRA CD4+ cells: cycle 3 vs baseline T-EMRA CD4+ cells: cycle 6 vs baseline <.001 T-EM CD4+ cells (CD27+): cycle 3 vs baseline T-EM CD4+ cells (CD27+): cycle 6 vs baseline <.001 T-EM CD4+ cells (CD27-): cycle 3 vs baseline T-EM CD4+ cells (CD27-): cycle 6 vs baseline T-EMRA CD4+ cells (CD27+): cycle 3 vs baseline T-EMRA CD4+ cells (CD27+): cycle 6 vs baseline <.001 T-EMRA CD4+ cells (CD27-): cycle 3 vs baseline T-EMRA CD4+ cells (CD27-): cycle 6 vs baseline Total CD4 T cells: cycle 3 vs baseline Total CD4 T cells: cycle 6 vs baseline Naïve CD4+ T cells: cycle 3 vs baseline Naïve CD4+ T cells: cycle 6 vs baseline T-CM CD4+ cells: cycle 3 vs baseline T-CM CD4+ cells: cycle 6 vs baseline <.001 T-EM CD4+ cells: cycle 3 vs baseline T-EM CD4+ cells: cycle 6 vs baseline T-EMRA CD4+ cells: cycle 3 vs baseline T-EMRA CD4+ cells: cycle 6 vs baseline <.001 T-EM CD4+ cells (CD27+): cycle 3 vs baseline T-EM CD4+ cells (CD27+): cycle 6 vs baseline T-EM CD4+ cells (CD27-): cycle 3 vs baseline T-EM CD4+ cells (CD27-): cycle 6 vs baseline T-EMRA CD4+ cells (CD27+): cycle 3 vs baseline T-EMRA CD4+ cells (CD27+): cycle 6 vs baseline <.001 T-EMRA CD4+ cells (CD27-): cycle 3 vs baseline T-EMRA CD4+ cells (CD27-): cycle 6 vs baseline Total CD4 T cells: cycle 3 vs baseline Total CD4 T cells: cycle 6 vs baseline CD45RA- CD4 T cells: cycle 3 vs baseline CD45RA- CD4 T cells: cycle 6 vs baseline CD45RA+ CD4 T cells: cycle 3 vs baseline CD45RA+ CD4 T cells: cycle 6 vs baseline Total CD4 T cells: cycle 3 vs baseline Total CD4 T cells: cycle 6 vs baseline CD45RA- CD4 T cells: cycle 3 vs baseline CD45RA- CD4 T cells: cycle 6 vs baseline CD45RA+ CD4 T cells: cycle 3 vs baseline CD45RA+ CD4 T cells: cycle 6 vs baseline
Blocking antibodies and peptides. Rat anti-mouse PD-1 (29F.1A12, rat IgG2a, k), PD-
 Supplementary Methods Blocking antibodies and peptides. Rat anti-mouse PD-1 (29F.1A12, rat IgG2a, k), PD- L1 (10F.9G2, rat IgG2b, k), and PD-L2 (3.2, mouse IgG1) have been described (24). Anti-CTLA-4 (clone
Supplementary Methods Blocking antibodies and peptides. Rat anti-mouse PD-1 (29F.1A12, rat IgG2a, k), PD- L1 (10F.9G2, rat IgG2b, k), and PD-L2 (3.2, mouse IgG1) have been described (24). Anti-CTLA-4 (clone
Supplementary Table e-1. Flow cytometry reagents and staining combinations
 Supplementary data Supplementary Table e-1. Flow cytometry reagents and staining combinations Reagents Antibody Fluorochrome Clone Source conjugation CD3 FITC UCHT1 BD Biosciences CD3 PerCP-Cy5.5 SK7 Biolegend
Supplementary data Supplementary Table e-1. Flow cytometry reagents and staining combinations Reagents Antibody Fluorochrome Clone Source conjugation CD3 FITC UCHT1 BD Biosciences CD3 PerCP-Cy5.5 SK7 Biolegend
Fluorochrome Panel 1 Panel 2 Panel 3 Panel 4 Panel 5 CTLA-4 CTLA-4 CD15 CD3 FITC. Bio) PD-1 (MIH4, BD) ICOS (C398.4A, Biolegend) PD-L1 (MIH1, BD)
 Additional file : Table S. Antibodies used for panel stain to identify peripheral immune cell subsets. Panel : PD- signaling; Panel : CD + T cells, CD + T cells, B cells; Panel : Tregs; Panel :, -T, cdc,
Additional file : Table S. Antibodies used for panel stain to identify peripheral immune cell subsets. Panel : PD- signaling; Panel : CD + T cells, CD + T cells, B cells; Panel : Tregs; Panel :, -T, cdc,
MATERIALS AND METHODS. Neutralizing antibodies specific to mouse Dll1, Dll4, J1 and J2 were prepared as described. 1,2 All
 MATERIALS AND METHODS Antibodies (Abs), flow cytometry analysis and cell lines Neutralizing antibodies specific to mouse Dll1, Dll4, J1 and J2 were prepared as described. 1,2 All other antibodies used
MATERIALS AND METHODS Antibodies (Abs), flow cytometry analysis and cell lines Neutralizing antibodies specific to mouse Dll1, Dll4, J1 and J2 were prepared as described. 1,2 All other antibodies used
Cover Page. The handle holds various files of this Leiden University dissertation.
 Cover Page The handle http://hdl.handle.net/1887/23854 holds various files of this Leiden University dissertation. Author: Marel, Sander van der Title: Gene and cell therapy based treatment strategies
Cover Page The handle http://hdl.handle.net/1887/23854 holds various files of this Leiden University dissertation. Author: Marel, Sander van der Title: Gene and cell therapy based treatment strategies
Detailed step-by-step operating procedures for NK cell and CTL degranulation assays
 Supplemental methods Detailed step-by-step operating procedures for NK cell and CTL degranulation assays Materials PBMC isolated from patients, relatives and healthy donors as control K562 cells (ATCC,
Supplemental methods Detailed step-by-step operating procedures for NK cell and CTL degranulation assays Materials PBMC isolated from patients, relatives and healthy donors as control K562 cells (ATCC,
Primary Adult Naïve CD4+ CD45RA+ Cells. Prepared by: David Randolph at University of Alabama, Birmingham
 Primary Adult Naïve CD4+ CD45RA+ Cells Prepared by: David Randolph (drdrdr@uab.edu) at University of Alabama, Birmingham Goal: To obtain large numbers of highly pure primary CD4+ CD45RO- CD25- cells from
Primary Adult Naïve CD4+ CD45RA+ Cells Prepared by: David Randolph (drdrdr@uab.edu) at University of Alabama, Birmingham Goal: To obtain large numbers of highly pure primary CD4+ CD45RO- CD25- cells from
Supplementary Figure 1. Efficient DC depletion in CD11c.DOG transgenic mice
 Supplementary Figure 1. Efficient DC depletion in CD11c.DOG transgenic mice (a) CD11c.DOG transgenic mice (tg) were treated with 8 ng/g body weight (b.w.) diphtheria toxin (DT) i.p. on day -1 and every
Supplementary Figure 1. Efficient DC depletion in CD11c.DOG transgenic mice (a) CD11c.DOG transgenic mice (tg) were treated with 8 ng/g body weight (b.w.) diphtheria toxin (DT) i.p. on day -1 and every
Supplementary Materials for
 immunology.sciencemag.org/cgi/content/full/2/16/eaan6049/dc1 Supplementary Materials for Enzymatic synthesis of core 2 O-glycans governs the tissue-trafficking potential of memory CD8 + T cells Jossef
immunology.sciencemag.org/cgi/content/full/2/16/eaan6049/dc1 Supplementary Materials for Enzymatic synthesis of core 2 O-glycans governs the tissue-trafficking potential of memory CD8 + T cells Jossef
Supplementary Figure 1. Ex vivo IFNγ production by Tregs. Nature Medicine doi: /nm % CD127. Empty SSC 98.79% CD25 CD45RA.
 SSC CD25 1.8% CD127 Empty 98.79% FSC CD45RA CD45RA Foxp3 %IFNγ + cells 4 3 2 1 + IL-12 P =.3 IFNγ p=.9 %IL-4+ cells 3 2 1 IL-4 P =.4 c %IL-1 + cells IFNγ 4 3 2 1 Control Foxp3 IL-1 P =.41.64 4.76 MS 2.96
SSC CD25 1.8% CD127 Empty 98.79% FSC CD45RA CD45RA Foxp3 %IFNγ + cells 4 3 2 1 + IL-12 P =.3 IFNγ p=.9 %IL-4+ cells 3 2 1 IL-4 P =.4 c %IL-1 + cells IFNγ 4 3 2 1 Control Foxp3 IL-1 P =.41.64 4.76 MS 2.96
Supplementary Figure 1. Immune profiles of untreated and PD-1 blockade resistant EGFR and Kras mouse lung tumors (a) Total lung weight of untreated
 1 Supplementary Figure 1. Immune profiles of untreated and PD-1 blockade resistant EGFR and Kras mouse lung tumors (a) Total lung weight of untreated (U) EGFR TL mice (n=7), Kras mice (n=7), PD-1 blockade
1 Supplementary Figure 1. Immune profiles of untreated and PD-1 blockade resistant EGFR and Kras mouse lung tumors (a) Total lung weight of untreated (U) EGFR TL mice (n=7), Kras mice (n=7), PD-1 blockade
Ex vivo Human Antigen-specific T Cell Proliferation and Degranulation Willemijn Hobo 1, Wieger Norde 1 and Harry Dolstra 2*
 Ex vivo Human Antigen-specific T Cell Proliferation and Degranulation Willemijn Hobo 1, Wieger Norde 1 and Harry Dolstra 2* 1 Department of Laboratory Medicine - Laboratory of Hematology, Radboud University
Ex vivo Human Antigen-specific T Cell Proliferation and Degranulation Willemijn Hobo 1, Wieger Norde 1 and Harry Dolstra 2* 1 Department of Laboratory Medicine - Laboratory of Hematology, Radboud University
Supplementary Figure 1. IL-12 serum levels and frequency of subsets in FL patients. (A) IL-12
 1 Supplementary Data Figure legends Supplementary Figure 1. IL-12 serum levels and frequency of subsets in FL patients. (A) IL-12 serum levels measured by multiplex ELISA (Luminex) in FL patients before
1 Supplementary Data Figure legends Supplementary Figure 1. IL-12 serum levels and frequency of subsets in FL patients. (A) IL-12 serum levels measured by multiplex ELISA (Luminex) in FL patients before
Cell isolation. Spleen and lymph nodes (axillary, inguinal) were removed from mice
 Supplementary Methods: Cell isolation. Spleen and lymph nodes (axillary, inguinal) were removed from mice and gently meshed in DMEM containing 10% FBS to prepare for single cell suspensions. CD4 + CD25
Supplementary Methods: Cell isolation. Spleen and lymph nodes (axillary, inguinal) were removed from mice and gently meshed in DMEM containing 10% FBS to prepare for single cell suspensions. CD4 + CD25
sequences of a styx mutant reveals a T to A transversion in the donor splice site of intron 5
 sfigure 1 Styx mutant mice recapitulate the phenotype of SHIP -/- mice. (A) Analysis of the genomic sequences of a styx mutant reveals a T to A transversion in the donor splice site of intron 5 (GTAAC
sfigure 1 Styx mutant mice recapitulate the phenotype of SHIP -/- mice. (A) Analysis of the genomic sequences of a styx mutant reveals a T to A transversion in the donor splice site of intron 5 (GTAAC
York criteria, 6 RA patients and 10 age- and gender-matched healthy controls (HCs).
 MATERIALS AND METHODS Study population Blood samples were obtained from 15 patients with AS fulfilling the modified New York criteria, 6 RA patients and 10 age- and gender-matched healthy controls (HCs).
MATERIALS AND METHODS Study population Blood samples were obtained from 15 patients with AS fulfilling the modified New York criteria, 6 RA patients and 10 age- and gender-matched healthy controls (HCs).
NK cell flow cytometric assay In vivo DC viability and migration assay
 NK cell flow cytometric assay 6 NK cells were purified, by negative selection with the NK Cell Isolation Kit (Miltenyi iotec), from spleen and lymph nodes of 6 RAG1KO mice, injected the day before with
NK cell flow cytometric assay 6 NK cells were purified, by negative selection with the NK Cell Isolation Kit (Miltenyi iotec), from spleen and lymph nodes of 6 RAG1KO mice, injected the day before with
Supporting Information
 Supporting Information Desnues et al. 10.1073/pnas.1314121111 SI Materials and Methods Mice. Toll-like receptor (TLR)8 / and TLR9 / mice were generated as described previously (1, 2). TLR9 / mice were
Supporting Information Desnues et al. 10.1073/pnas.1314121111 SI Materials and Methods Mice. Toll-like receptor (TLR)8 / and TLR9 / mice were generated as described previously (1, 2). TLR9 / mice were
Interferon γ regulates idiopathic pneumonia syndrome, a. Th17 + CD4 + T-cell-mediated GvH disease
 Interferon γ regulates idiopathic pneumonia syndrome, a Th17 + CD4 + T-cell-mediated GvH disease Nora Mauermann, Julia Burian, Christophe von Garnier, Stefan Dirnhofer, Davide Germano, Christine Schuett,
Interferon γ regulates idiopathic pneumonia syndrome, a Th17 + CD4 + T-cell-mediated GvH disease Nora Mauermann, Julia Burian, Christophe von Garnier, Stefan Dirnhofer, Davide Germano, Christine Schuett,
SUPPLEMENTARY FIGURES
 SUPPLEMENTARY FIGURES Supplementary Figure 1: Chemokine receptor expression profiles of CCR6 + and CCR6 - CD4 + IL-17A +/ex and Treg cells. Quantitative PCR analysis of chemokine receptor transcript abundance
SUPPLEMENTARY FIGURES Supplementary Figure 1: Chemokine receptor expression profiles of CCR6 + and CCR6 - CD4 + IL-17A +/ex and Treg cells. Quantitative PCR analysis of chemokine receptor transcript abundance
Commercially available HLA Class II tetramers (Beckman Coulter) conjugated to
 Class II tetramer staining Commercially available HLA Class II tetramers (Beckman Coulter) conjugated to PE were combined with dominant HIV epitopes (DRB1*0101-DRFYKTLRAEQASQEV, DRB1*0301- PEKEVLVWKFDSRLAFHH,
Class II tetramer staining Commercially available HLA Class II tetramers (Beckman Coulter) conjugated to PE were combined with dominant HIV epitopes (DRB1*0101-DRFYKTLRAEQASQEV, DRB1*0301- PEKEVLVWKFDSRLAFHH,
Optimizing Intracellular Flow Cytometry:
 Optimizing Intracellular Flow Cytometry: Simultaneous Detection of Cytokines and Transcription Factors An encore presentation by Jurg Rohrer, PhD, BD Biosciences 10.26.10 Outline Introduction Cytokines
Optimizing Intracellular Flow Cytometry: Simultaneous Detection of Cytokines and Transcription Factors An encore presentation by Jurg Rohrer, PhD, BD Biosciences 10.26.10 Outline Introduction Cytokines
Recommended Protocols for Phospho Protein Detection in Human Cells
 Recommended Protocols for Phospho Protein Detection in Human Cells Depending on subcellular localization of the phospho protein of interest, as well as epitope susceptibility to cell fixing and permeabilizing
Recommended Protocols for Phospho Protein Detection in Human Cells Depending on subcellular localization of the phospho protein of interest, as well as epitope susceptibility to cell fixing and permeabilizing
MAIT cell function is modulated by PD-1 signaling in patients with active
 MAIT cell function is modulated by PD-1 signaling in patients with active tuberculosis Jing Jiang, M.D., Xinjing Wang, M.D., Hongjuan An, M.Sc., Bingfen Yang, Ph.D., Zhihong Cao, M.Sc., Yanhua Liu, Ph.D.,
MAIT cell function is modulated by PD-1 signaling in patients with active tuberculosis Jing Jiang, M.D., Xinjing Wang, M.D., Hongjuan An, M.Sc., Bingfen Yang, Ph.D., Zhihong Cao, M.Sc., Yanhua Liu, Ph.D.,
BD Flow Cytometry Reagents Multicolor Panels Designed for Optimal Resolution with the BD LSRFortessa X-20 Cell Analyzer
 Multicolor Panels Designed for Optimal Resolution with the BD LSRFortessa X-2 Cell Analyzer Proper multicolor panel design takes into account fluorochrome brightness, antigen density, co-expression, and
Multicolor Panels Designed for Optimal Resolution with the BD LSRFortessa X-2 Cell Analyzer Proper multicolor panel design takes into account fluorochrome brightness, antigen density, co-expression, and
PBMC from each patient were suspended in AIM V medium (Invitrogen) with 5% human
 Anti-CD19-CAR transduced T-cell preparation PBMC from each patient were suspended in AIM V medium (Invitrogen) with 5% human AB serum (Gemini) and 300 international units/ml IL-2 (Novartis). T cell proliferation
Anti-CD19-CAR transduced T-cell preparation PBMC from each patient were suspended in AIM V medium (Invitrogen) with 5% human AB serum (Gemini) and 300 international units/ml IL-2 (Novartis). T cell proliferation
Supplementary Materials
 Supplementary Materials 43 Figure S1. CD123 in acute lymphoblastic leukemia and leukemia-initiating cells. A. CD123 (histograms) is highly and homogenously expressed in B-ALL blasts (as defined by live,
Supplementary Materials 43 Figure S1. CD123 in acute lymphoblastic leukemia and leukemia-initiating cells. A. CD123 (histograms) is highly and homogenously expressed in B-ALL blasts (as defined by live,
Application Information Bulletin: Human NK Cells Phenotypic characterizing of human Natural Killer (NK) cell populations in peripheral blood
 Application Information Bulletin: Human NK Cells Phenotypic characterizing of human Natural Killer (NK) cell populations in peripheral blood Christopher A Fraker, Ph.D., University of Miami - Miami, Florida
Application Information Bulletin: Human NK Cells Phenotypic characterizing of human Natural Killer (NK) cell populations in peripheral blood Christopher A Fraker, Ph.D., University of Miami - Miami, Florida
L-selectin Is Essential for Delivery of Activated CD8 + T Cells to Virus-Infected Organs for Protective Immunity
 Cell Reports Supplemental Information L-selectin Is Essential for Delivery of Activated CD8 + T Cells to Virus-Infected Organs for Protective Immunity Rebar N. Mohammed, H. Angharad Watson, Miriam Vigar,
Cell Reports Supplemental Information L-selectin Is Essential for Delivery of Activated CD8 + T Cells to Virus-Infected Organs for Protective Immunity Rebar N. Mohammed, H. Angharad Watson, Miriam Vigar,
The encephalitogenicity of TH17 cells is dependent on IL-1- and IL-23- induced production of the cytokine GM-CSF
 CORRECTION NOTICE Nat.Immunol. 12, 568 575 (2011) The encephalitogenicity of TH17 cells is dependent on IL-1- and IL-23- induced production of the cytokine GM-CSF Mohamed El-Behi, Bogoljub Ciric, Hong
CORRECTION NOTICE Nat.Immunol. 12, 568 575 (2011) The encephalitogenicity of TH17 cells is dependent on IL-1- and IL-23- induced production of the cytokine GM-CSF Mohamed El-Behi, Bogoljub Ciric, Hong
PBMC from each patient were suspended in AIM V medium (Life Technologies) with 5%
 Allogeneic T-cells expressing an anti-cd19 chimeric antigen receptor induce remissions of B-cell malignancies that progress after allogeneic hematopoietic stem cell transplantation without causing graft-versus-host
Allogeneic T-cells expressing an anti-cd19 chimeric antigen receptor induce remissions of B-cell malignancies that progress after allogeneic hematopoietic stem cell transplantation without causing graft-versus-host
SUPPLEMENTARY INFORMATION. Involvement of IL-21 in the epidermal hyperplasia of psoriasis
 SUPPLEMENTARY INFORMATION Involvement of IL-21 in the epidermal hyperplasia of psoriasis Roberta Caruso 1, Elisabetta Botti 2, Massimiliano Sarra 1, Maria Esposito 2, Carmine Stolfi 1, Laura Diluvio 2,
SUPPLEMENTARY INFORMATION Involvement of IL-21 in the epidermal hyperplasia of psoriasis Roberta Caruso 1, Elisabetta Botti 2, Massimiliano Sarra 1, Maria Esposito 2, Carmine Stolfi 1, Laura Diluvio 2,
Rapid antigen-specific T cell enrichment (Rapid ARTE)
 Direct ex vivo characterization of human antigen-specific CD154+CD4+ T cell Rapid antigen-specific T cell enrichment (Rapid ARTE) Introduction Workflow Antigen (ag)-specific T cells play a central role
Direct ex vivo characterization of human antigen-specific CD154+CD4+ T cell Rapid antigen-specific T cell enrichment (Rapid ARTE) Introduction Workflow Antigen (ag)-specific T cells play a central role
Supplemental Table I.
 Supplemental Table I Male / Mean ± SEM n Mean ± SEM n Body weight, g 29.2±0.4 17 29.7±0.5 17 Total cholesterol, mg/dl 534.0±30.8 17 561.6±26.1 17 HDL-cholesterol, mg/dl 9.6±0.8 17 10.1±0.7 17 Triglycerides,
Supplemental Table I Male / Mean ± SEM n Mean ± SEM n Body weight, g 29.2±0.4 17 29.7±0.5 17 Total cholesterol, mg/dl 534.0±30.8 17 561.6±26.1 17 HDL-cholesterol, mg/dl 9.6±0.8 17 10.1±0.7 17 Triglycerides,
SUPPORTING INFORMATIONS
 SUPPORTING INFORMATIONS Mice MT/ret RetCD3ε KO α-cd25 treated MT/ret Age 1 month 3 mnths 6 months 1 month 3 months 6 months 1 month 3 months 6 months 2/87 Survival 87/87 incidence of 17/87 1 ary tumor
SUPPORTING INFORMATIONS Mice MT/ret RetCD3ε KO α-cd25 treated MT/ret Age 1 month 3 mnths 6 months 1 month 3 months 6 months 1 month 3 months 6 months 2/87 Survival 87/87 incidence of 17/87 1 ary tumor
Supplemental Information. T Cells Enhance Autoimmunity by Restraining Regulatory T Cell Responses via an Interleukin-23-Dependent Mechanism
 Immunity, Volume 33 Supplemental Information T Cells Enhance Autoimmunity by Restraining Regulatory T Cell Responses via an Interleukin-23-Dependent Mechanism Franziska Petermann, Veit Rothhammer, Malte
Immunity, Volume 33 Supplemental Information T Cells Enhance Autoimmunity by Restraining Regulatory T Cell Responses via an Interleukin-23-Dependent Mechanism Franziska Petermann, Veit Rothhammer, Malte
In vitro human regulatory T cell expansion
 - 1 - Human CD4 + CD25 + CD127 dim/- regulatory T cell Workflow isolation, in vitro expansion and analysis In vitro human regulatory T cell expansion Introduction Regulatory T (Treg) cells are a subpopulation
- 1 - Human CD4 + CD25 + CD127 dim/- regulatory T cell Workflow isolation, in vitro expansion and analysis In vitro human regulatory T cell expansion Introduction Regulatory T (Treg) cells are a subpopulation
T H 1, T H 2 and T H 17 polarization of naïve CD4 + mouse T cells
 A complete workflow for cell preparation, isolation, polarization and analysis T H 1, T H 2 and T H 17 polarization of naïve CD4 + mouse T cells Introduction Workflow CD4 + T helper (T H) cells play a
A complete workflow for cell preparation, isolation, polarization and analysis T H 1, T H 2 and T H 17 polarization of naïve CD4 + mouse T cells Introduction Workflow CD4 + T helper (T H) cells play a
Direct ex vivo characterization of human antigen-specific CD154 + CD4 + T cells Rapid antigen-reactive T cell enrichment (Rapid ARTE)
 Direct ex vivo characterization of human antigen-specific CD154 + CD4 + T cells Rapid antigen-reactive T cell enrichment (Rapid ARTE) Introduction Workflow Antigen (ag)-specific T cells play a central
Direct ex vivo characterization of human antigen-specific CD154 + CD4 + T cells Rapid antigen-reactive T cell enrichment (Rapid ARTE) Introduction Workflow Antigen (ag)-specific T cells play a central
Supplemental Methods In vitro T cell assays Inhibition of perforin- and FasL-mediated cytotoxicity Flow Cytometry
 Supplemental Methods In vitro T cell assays Cell lines Jurkat (ATCC #TIB-152), CCRF-CEM (ATCC #CCL-119), MOLT-4 (ATCC #CRL- 1582), Hut 78 (ATCC #TIB-161), SupT1 (ATCC #CRL-1942), Raji (ATCC #CCL-86) and
Supplemental Methods In vitro T cell assays Cell lines Jurkat (ATCC #TIB-152), CCRF-CEM (ATCC #CCL-119), MOLT-4 (ATCC #CRL- 1582), Hut 78 (ATCC #TIB-161), SupT1 (ATCC #CRL-1942), Raji (ATCC #CCL-86) and
In vitro human regulatory T cell expansion
 - 1 - Human CD4 + CD25 + regulatory T cell isolation, Workflow in vitro expansion and analysis In vitro human regulatory T cell expansion Introduction Regulatory T (Treg) cells are a subpopulation of T
- 1 - Human CD4 + CD25 + regulatory T cell isolation, Workflow in vitro expansion and analysis In vitro human regulatory T cell expansion Introduction Regulatory T (Treg) cells are a subpopulation of T
Supplementary Figure 1. Characterization of basophils after reconstitution of SCID mice
 Supplementary figure legends Supplementary Figure 1. Characterization of after reconstitution of SCID mice with CD4 + CD62L + T cells. (A-C) SCID mice (n = 6 / group) were reconstituted with 2 x 1 6 CD4
Supplementary figure legends Supplementary Figure 1. Characterization of after reconstitution of SCID mice with CD4 + CD62L + T cells. (A-C) SCID mice (n = 6 / group) were reconstituted with 2 x 1 6 CD4
Nature Medicine doi: /nm.3957
 Supplementary Fig. 1. p38 alternative activation, IL-21 expression, and T helper cell transcription factors in PDAC tissue. (a) Tissue microarrays of pancreatic tissue from 192 patients with pancreatic
Supplementary Fig. 1. p38 alternative activation, IL-21 expression, and T helper cell transcription factors in PDAC tissue. (a) Tissue microarrays of pancreatic tissue from 192 patients with pancreatic
Ibrutinib treatment improves T cell number and function in CLL patients
 Ibrutinib treatment improves T cell number and function in CLL patients Meixiao Long,, Natarajan Muthusamy, John C. Byrd J Clin Invest. 2017;127(8):3052-3064. https://doi.org/10.1172/jci89756. Clinical
Ibrutinib treatment improves T cell number and function in CLL patients Meixiao Long,, Natarajan Muthusamy, John C. Byrd J Clin Invest. 2017;127(8):3052-3064. https://doi.org/10.1172/jci89756. Clinical
Supplementary Data. Treg phenotype
 Supplementary Data Additional Experiment An additional experiment was performed using cryopreserved peripheral blood mononuclear cells (PBMC) derived from five renal cell carcinoma (RCC) patients [see
Supplementary Data Additional Experiment An additional experiment was performed using cryopreserved peripheral blood mononuclear cells (PBMC) derived from five renal cell carcinoma (RCC) patients [see
Supplementary Figure S1. Flow cytometric analysis of the expression of Thy1 in NH cells. Flow cytometric analysis of the expression of T1/ST2 and
 Supplementary Figure S1. Flow cytometric analysis of the expression of Thy1 in NH cells. Flow cytometric analysis of the expression of T1/ST2 and Thy1 in NH cells derived from the lungs of naïve mice.
Supplementary Figure S1. Flow cytometric analysis of the expression of Thy1 in NH cells. Flow cytometric analysis of the expression of T1/ST2 and Thy1 in NH cells derived from the lungs of naïve mice.
Supplemental Figure 1. Signature gene expression in in vitro differentiated Th0, Th1, Th2, Th17 and Treg cells. (A) Naïve CD4 + T cells were cultured
 Supplemental Figure 1. Signature gene expression in in vitro differentiated Th0, Th1, Th2, Th17 and Treg cells. (A) Naïve CD4 + T cells were cultured under Th0, Th1, Th2, Th17, and Treg conditions. mrna
Supplemental Figure 1. Signature gene expression in in vitro differentiated Th0, Th1, Th2, Th17 and Treg cells. (A) Naïve CD4 + T cells were cultured under Th0, Th1, Th2, Th17, and Treg conditions. mrna
B6/COLODR/SPL/11C/83/LAP/#2.006 B6/COLODR/SPL/11C/86/LAP/#2.016 CD11C B6/COLODR/SPL/11C/80/LAP/#2.011 CD11C
 CD3-specific antibody-induced immune tolerance and suppression of autoimmune encephalomyelitis involves TGF-β production through phagocytes digesting apoptotic T cells Sylvain Perruche 1,3, Pin Zhang 1,
CD3-specific antibody-induced immune tolerance and suppression of autoimmune encephalomyelitis involves TGF-β production through phagocytes digesting apoptotic T cells Sylvain Perruche 1,3, Pin Zhang 1,
Supplementary Figure 1
 Supplementary Figure 1 Identification of IFN-γ-producing CD8 + and CD4 + T cells with naive phenotype by alternative gating and sample-processing strategies. a. Contour 5% probability plots show definition
Supplementary Figure 1 Identification of IFN-γ-producing CD8 + and CD4 + T cells with naive phenotype by alternative gating and sample-processing strategies. a. Contour 5% probability plots show definition
well for 2 h at rt. Each dot represents an individual mouse and bar is the mean ±
 Supplementary data: Control DC Blimp-1 ko DC 8 6 4 2-2 IL-1β p=.5 medium 8 6 4 2 IL-2 Medium p=.16 8 6 4 2 IL-6 medium p=.3 5 4 3 2 1-1 medium IL-1 n.s. 25 2 15 1 5 IL-12(p7) p=.15 5 IFNγ p=.65 4 3 2 1
Supplementary data: Control DC Blimp-1 ko DC 8 6 4 2-2 IL-1β p=.5 medium 8 6 4 2 IL-2 Medium p=.16 8 6 4 2 IL-6 medium p=.3 5 4 3 2 1-1 medium IL-1 n.s. 25 2 15 1 5 IL-12(p7) p=.15 5 IFNγ p=.65 4 3 2 1
Comprehensive evaluation of human immune system reconstitution in NSG. and NSG -SGM3 mouse models toward the development of a novel ONCO-HU
 Comprehensive evaluation of human immune system reconstitution in NSG and NSG -SGM3 mouse models toward the development of a novel ONCO-HU xenograft model Aaron Middlebrook, 1 Eileen Snowden, 2 Warren
Comprehensive evaluation of human immune system reconstitution in NSG and NSG -SGM3 mouse models toward the development of a novel ONCO-HU xenograft model Aaron Middlebrook, 1 Eileen Snowden, 2 Warren
Nature Immunology: doi: /ni Supplementary Figure 1. Huwe1 has high expression in HSCs and is necessary for quiescence.
 Supplementary Figure 1 Huwe1 has high expression in HSCs and is necessary for quiescence. (a) Heat map visualizing expression of genes with a known function in ubiquitin-mediated proteolysis (KEGG: Ubiquitin
Supplementary Figure 1 Huwe1 has high expression in HSCs and is necessary for quiescence. (a) Heat map visualizing expression of genes with a known function in ubiquitin-mediated proteolysis (KEGG: Ubiquitin
January 25, 2017 Scientific Research Process Name of Journal: ESPS Manuscript NO: Manuscript Type: Title: Authors: Correspondence to
 January 25, 2017 Scientific Research Process Name of Journal: World Journal of Gastroenterology ESPS Manuscript NO: 31928 Manuscript Type: ORIGINAL ARTICLE Title: Thiopurine use associated with reduced
January 25, 2017 Scientific Research Process Name of Journal: World Journal of Gastroenterology ESPS Manuscript NO: 31928 Manuscript Type: ORIGINAL ARTICLE Title: Thiopurine use associated with reduced
Effect of Antiretroviral Therapy on HIV-mediated Impairment of the Neutrophil
 Online Data Supplement Effect of Antiretroviral Therapy on HIV-mediated Impairment of the Neutrophil Antimycobacterial Response David M Lowe, Nonzwakazi Bangani, Rene Goliath, Beate Kampmann, Katalin A
Online Data Supplement Effect of Antiretroviral Therapy on HIV-mediated Impairment of the Neutrophil Antimycobacterial Response David M Lowe, Nonzwakazi Bangani, Rene Goliath, Beate Kampmann, Katalin A
Peptide stimulation and Intracellular Cytokine Staining
 v Peptide stimulation and Intracellular Cytokine Staining Authors A. Cosma, S. Allgayer MATERIALS Date 01-02-2007 REAGENTS: Version 1.0 - PBMCs - Culture medium - Costimulating antibodies - Peptide pools
v Peptide stimulation and Intracellular Cytokine Staining Authors A. Cosma, S. Allgayer MATERIALS Date 01-02-2007 REAGENTS: Version 1.0 - PBMCs - Culture medium - Costimulating antibodies - Peptide pools
Optimizing Intracellular Flow Cytometry:
 Optimizing Intracellular Flow Cytometry: Simultaneous Detection of Cytokines and Transcription Factors Presented by Jurg Rohrer, PhD, BD Biosciences 23-10780-00 Outline Introduction Cytokines Transcription
Optimizing Intracellular Flow Cytometry: Simultaneous Detection of Cytokines and Transcription Factors Presented by Jurg Rohrer, PhD, BD Biosciences 23-10780-00 Outline Introduction Cytokines Transcription
Gladstone Institutes, University of California (UCSF), San Francisco, USA
 Fluorescence-linked Antigen Quantification (FLAQ) Assay for Fast Quantification of HIV-1 p24 Gag Marianne Gesner, Mekhala Maiti, Robert Grant and Marielle Cavrois * Gladstone Institutes, University of
Fluorescence-linked Antigen Quantification (FLAQ) Assay for Fast Quantification of HIV-1 p24 Gag Marianne Gesner, Mekhala Maiti, Robert Grant and Marielle Cavrois * Gladstone Institutes, University of
Supplementary Data Table of Contents:
 Supplementary Data Table of Contents: - Supplementary Methods - Supplementary Figures S1(A-B) - Supplementary Figures S2 (A-B) - Supplementary Figures S3 - Supplementary Figures S4(A-B) - Supplementary
Supplementary Data Table of Contents: - Supplementary Methods - Supplementary Figures S1(A-B) - Supplementary Figures S2 (A-B) - Supplementary Figures S3 - Supplementary Figures S4(A-B) - Supplementary
Supporting Information
 Supporting Information van der Windt et al. 10.1073/pnas.1221740110 SI Materials and Methods Mice and Reagents. C57BL/6 and major histocompatibility complex class I-restricted OVA-specific T-cell receptor
Supporting Information van der Windt et al. 10.1073/pnas.1221740110 SI Materials and Methods Mice and Reagents. C57BL/6 and major histocompatibility complex class I-restricted OVA-specific T-cell receptor
Supplementalgfigureg1gSchematicgdiagramgofgtumor1modellingg
 SChinjectionh F:LuchLCLsh IVhinjectionh T:cellsh Monitorhforhtumorh growthhandhxeno: reactivehgvhd GVLgexperimentg kcbgvsgpbgt1cellse Xeno1reactiveg experimentg kcbgvsgpbgt1cellse IVhinjectionh 5xh,N^6
SChinjectionh F:LuchLCLsh IVhinjectionh T:cellsh Monitorhforhtumorh growthhandhxeno: reactivehgvhd GVLgexperimentg kcbgvsgpbgt1cellse Xeno1reactiveg experimentg kcbgvsgpbgt1cellse IVhinjectionh 5xh,N^6
Eosinophils are required. for the maintenance of plasma cells in the bone marrow
 Eosinophils are required for the maintenance of plasma cells in the bone marrow Van Trung Chu, Anja Fröhlich, Gudrun Steinhauser, Tobias Scheel, Toralf Roch, Simon Fillatreau, James J. Lee, Max Löhning
Eosinophils are required for the maintenance of plasma cells in the bone marrow Van Trung Chu, Anja Fröhlich, Gudrun Steinhauser, Tobias Scheel, Toralf Roch, Simon Fillatreau, James J. Lee, Max Löhning
Supplementary Figures
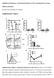 Inhibition of Pulmonary Anti Bacterial Defense by IFN γ During Recovery from Influenza Infection By Keer Sun and Dennis W. Metzger Supplementary Figures d a Ly6G Percentage survival f 1 75 5 1 25 1 5 1
Inhibition of Pulmonary Anti Bacterial Defense by IFN γ During Recovery from Influenza Infection By Keer Sun and Dennis W. Metzger Supplementary Figures d a Ly6G Percentage survival f 1 75 5 1 25 1 5 1
Supplemental Figure 1
 Supplemental Figure 1 1a 1c PD-1 MFI fold change 6 5 4 3 2 1 IL-1α IL-2 IL-4 IL-6 IL-1 IL-12 IL-13 IL-15 IL-17 IL-18 IL-21 IL-23 IFN-α Mut Human PD-1 promoter SBE-D 5 -GTCTG- -1.2kb SBE-P -CAGAC- -1.kb
Supplemental Figure 1 1a 1c PD-1 MFI fold change 6 5 4 3 2 1 IL-1α IL-2 IL-4 IL-6 IL-1 IL-12 IL-13 IL-15 IL-17 IL-18 IL-21 IL-23 IFN-α Mut Human PD-1 promoter SBE-D 5 -GTCTG- -1.2kb SBE-P -CAGAC- -1.kb
IL-6Rα IL-6RαT-KO KO. IL-6Rα f/f bp. f/f 628 bp deleted 368 bp. 500 bp
 STD H 2 O WT KO IL-6Rα f/f IL-6Rα IL-6RαT-KO KO 1000 bp 500 bp f/f 628 bp deleted 368 bp Supplementary Figure 1 Confirmation of T-cell IL-6Rα deficiency. (a) Representative histograms and (b) quantification
STD H 2 O WT KO IL-6Rα f/f IL-6Rα IL-6RαT-KO KO 1000 bp 500 bp f/f 628 bp deleted 368 bp Supplementary Figure 1 Confirmation of T-cell IL-6Rα deficiency. (a) Representative histograms and (b) quantification
Supplemental Materials for. Effects of sphingosine-1-phosphate receptor 1 phosphorylation in response to. FTY720 during neuroinflammation
 Supplemental Materials for Effects of sphingosine-1-phosphate receptor 1 phosphorylation in response to FTY7 during neuroinflammation This file includes: Supplemental Table 1. EAE clinical parameters of
Supplemental Materials for Effects of sphingosine-1-phosphate receptor 1 phosphorylation in response to FTY7 during neuroinflammation This file includes: Supplemental Table 1. EAE clinical parameters of
Supplemental Figure 1. IL-3 blockade with Fab CSL362 depletes plasmacytoid dendritic cells (pdcs), but not basophils, at higher doses.
 Supplemental Figure 1. IL-3 blockade with Fab CSL362 depletes plasmacytoid dendritic cells (pdcs), but not basophils, at higher doses. Percentage of viable (A) pdcs (Sytox Blue-, Lin1-, HLADR+, BDCA2++)
Supplemental Figure 1. IL-3 blockade with Fab CSL362 depletes plasmacytoid dendritic cells (pdcs), but not basophils, at higher doses. Percentage of viable (A) pdcs (Sytox Blue-, Lin1-, HLADR+, BDCA2++)
Supplementary Data 1. Alanine substitutions and position variants of APNCYGNIPL. Applied in
 Supplementary Data 1. Alanine substitutions and position variants of APNCYGNIPL. Applied in Supplementary Fig. 2 Substitution Sequence Position variant Sequence original APNCYGNIPL original APNCYGNIPL
Supplementary Data 1. Alanine substitutions and position variants of APNCYGNIPL. Applied in Supplementary Fig. 2 Substitution Sequence Position variant Sequence original APNCYGNIPL original APNCYGNIPL
Glow. Save. Let it. and. Get a Discount of 10% BD Biosciences Year End Promotion on a huge Selection of Reagent Solutions
 Let it Glow and Save BD Biosciences Year End Promotion 2014 Get a Discount of 10% on a huge Selection of Reagent Solutions All fluorochrome labeled antibodies The complete BD CBA portfolio Kits and reagents
Let it Glow and Save BD Biosciences Year End Promotion 2014 Get a Discount of 10% on a huge Selection of Reagent Solutions All fluorochrome labeled antibodies The complete BD CBA portfolio Kits and reagents
MATERIALS. Peptide stimulation and Intracellular Cytokine Staining- EFFECTOR 45RA PANEL
 v Peptide stimulation and Intracellular Cytokine Staining- EFFECTOR 45RA PANEL Authors S. Kutscher, A. Cosma, MATERIALS REAGENTS: - PBMCs - Culture medium - Costimulating antibodies - Peptide pools including
v Peptide stimulation and Intracellular Cytokine Staining- EFFECTOR 45RA PANEL Authors S. Kutscher, A. Cosma, MATERIALS REAGENTS: - PBMCs - Culture medium - Costimulating antibodies - Peptide pools including
Hua Tang, Weiping Cao, Sudhir Pai Kasturi, Rajesh Ravindran, Helder I Nakaya, Kousik
 SUPPLEMENTARY FIGURES 1-19 T H 2 response to cysteine-proteases requires dendritic cell-basophil cooperation via ROS mediated signaling Hua Tang, Weiping Cao, Sudhir Pai Kasturi, Rajesh Ravindran, Helder
SUPPLEMENTARY FIGURES 1-19 T H 2 response to cysteine-proteases requires dendritic cell-basophil cooperation via ROS mediated signaling Hua Tang, Weiping Cao, Sudhir Pai Kasturi, Rajesh Ravindran, Helder
SUPPLEMENT Supplementary Figure 1: (A) (B)
 SUPPLEMENT Supplementary Figure 1: CD4 + naïve effector T cells (CD4 effector) were labeled with CFSE, stimulated with α-cd2/cd3/cd28 coated beads (at 2 beads/cell) and cultured alone or cocultured with
SUPPLEMENT Supplementary Figure 1: CD4 + naïve effector T cells (CD4 effector) were labeled with CFSE, stimulated with α-cd2/cd3/cd28 coated beads (at 2 beads/cell) and cultured alone or cocultured with
and follicular helper T cells is Egr2-dependent. (a) Diagrammatic representation of the
 Supplementary Figure 1. LAG3 + Treg-mediated regulation of germinal center B cells and follicular helper T cells is Egr2-dependent. (a) Diagrammatic representation of the experimental protocol for the
Supplementary Figure 1. LAG3 + Treg-mediated regulation of germinal center B cells and follicular helper T cells is Egr2-dependent. (a) Diagrammatic representation of the experimental protocol for the
Supplementary Information
 Supplementary Information Methods Lymphocyte subsets analysis was performed on samples of 7 subjects by flow cytometry on blood samples collected in ethylenediaminetetraacetic acid (EDTA)-containing tubes
Supplementary Information Methods Lymphocyte subsets analysis was performed on samples of 7 subjects by flow cytometry on blood samples collected in ethylenediaminetetraacetic acid (EDTA)-containing tubes
CD14 + S100A9 + Monocytic Myeloid-Derived Suppressor Cells and Their Clinical Relevance in Non-Small Cell Lung Cancer
 CD14 + S1A9 + Monocytic Myeloid-Derived Suppressor Cells and Their Clinical Relevance in Non-Small Cell Lung Cancer Po-Hao, Feng M.D., Kang-Yun, Lee, M.D. Ph.D., Ya-Ling Chang, Yao-Fei Chan, Lu- Wei, Kuo,Ting-Yu
CD14 + S1A9 + Monocytic Myeloid-Derived Suppressor Cells and Their Clinical Relevance in Non-Small Cell Lung Cancer Po-Hao, Feng M.D., Kang-Yun, Lee, M.D. Ph.D., Ya-Ling Chang, Yao-Fei Chan, Lu- Wei, Kuo,Ting-Yu
D CD8 T cell number (x10 6 )
 IFNγ Supplemental Figure 1. CD T cell number (x1 6 ) 18 15 1 9 6 3 CD CD T cells CD6L C CD5 CD T cells CD6L D CD8 T cell number (x1 6 ) 1 8 6 E CD CD8 T cells CD6L F Log(1)CFU/g Feces 1 8 6 p
IFNγ Supplemental Figure 1. CD T cell number (x1 6 ) 18 15 1 9 6 3 CD CD T cells CD6L C CD5 CD T cells CD6L D CD8 T cell number (x1 6 ) 1 8 6 E CD CD8 T cells CD6L F Log(1)CFU/g Feces 1 8 6 p
Reviewers' comments: Reviewer #1 Expert in EAE and IL-17a (Remarks to the Author):
 Reviewers' comments: Reviewer #1 Expert in EAE and IL-17a (Remarks to the Author): This study shows that the inducible camp early repressor (ICER) is involved in development of Th17 cells that are pathogenic
Reviewers' comments: Reviewer #1 Expert in EAE and IL-17a (Remarks to the Author): This study shows that the inducible camp early repressor (ICER) is involved in development of Th17 cells that are pathogenic
Supplementary Information:
 Supplementary Information: Follicular regulatory T cells with Bcl6 expression suppress germinal center reactions by Yeonseok Chung, Shinya Tanaka, Fuliang Chu, Roza Nurieva, Gustavo J. Martinez, Seema
Supplementary Information: Follicular regulatory T cells with Bcl6 expression suppress germinal center reactions by Yeonseok Chung, Shinya Tanaka, Fuliang Chu, Roza Nurieva, Gustavo J. Martinez, Seema
Optimizing Intracellular Flow Cytometry
 Optimizing Intracellular Flow Cytometry Detection of Cytokines, Transcription Factors, and Phosphoprotein by Flow Cytometry Presented by Erika O Donnell, PhD, BD Biosciences 23-14876-00 Outline Basic principles
Optimizing Intracellular Flow Cytometry Detection of Cytokines, Transcription Factors, and Phosphoprotein by Flow Cytometry Presented by Erika O Donnell, PhD, BD Biosciences 23-14876-00 Outline Basic principles
Supplementary Figure S1: Alignment of CD28H. (a) Alignment of human CD28H with other known B7 receptors. (b) Alignment of CD28H orthologs.
 Supplementary Figure S1: Alignment of CD28H. (a) Alignment of human CD28H with other known B7 receptors. (b) Alignment of CD28H orthologs. Predicted CD28H protein sequences from human, chimpanzee (pan
Supplementary Figure S1: Alignment of CD28H. (a) Alignment of human CD28H with other known B7 receptors. (b) Alignment of CD28H orthologs. Predicted CD28H protein sequences from human, chimpanzee (pan
of whole cell cultures in U-bottomed wells of a 96-well plate are shown. 2
 Supplementary online material Supplementary figure legends Supplementary Figure 1 Exposure to T reg cells causes loss of T resp cells in co-cultures. T resp cells were stimulated with CD3+CD28 alone or
Supplementary online material Supplementary figure legends Supplementary Figure 1 Exposure to T reg cells causes loss of T resp cells in co-cultures. T resp cells were stimulated with CD3+CD28 alone or
In vitro bactericidal assay Fig. S8 Gentamicin protection assay Phagocytosis assay
 In vitro bactericidal assay Mouse bone marrow was isolated from the femur and the tibia. Cells were suspended in phosphate buffered saline containing.5% BSA and 2 mm EDTA and filtered through a cell strainer.
In vitro bactericidal assay Mouse bone marrow was isolated from the femur and the tibia. Cells were suspended in phosphate buffered saline containing.5% BSA and 2 mm EDTA and filtered through a cell strainer.
SUPPLEMENTARY INFORMATION
 Complete but curtailed T-cell response to very-low-affinity antigen Dietmar Zehn, Sarah Y. Lee & Michael J. Bevan Supp. Fig. 1: TCR chain usage among endogenous K b /Ova reactive T cells. C57BL/6 mice
Complete but curtailed T-cell response to very-low-affinity antigen Dietmar Zehn, Sarah Y. Lee & Michael J. Bevan Supp. Fig. 1: TCR chain usage among endogenous K b /Ova reactive T cells. C57BL/6 mice
Supplemental Information. Human Circulating PD-1 + CXCR3 CXCR5 + Memory. Tfh Cells Are Highly Functional and Correlate
 Immunity, Volume 39 Supplemental Information Human Circulating PD-1 + CXCR3 CXCR5 + Memory Tfh Cells Are Highly Functional and Correlate with Broadly Neutralizing HIV Antibody Responses Michela Locci,
Immunity, Volume 39 Supplemental Information Human Circulating PD-1 + CXCR3 CXCR5 + Memory Tfh Cells Are Highly Functional and Correlate with Broadly Neutralizing HIV Antibody Responses Michela Locci,
W/T Itgam -/- F4/80 CD115. F4/80 hi CD115 + F4/80 + CD115 +
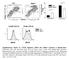 F4/8 % in the peritoneal lavage 6 4 2 p=.15 n.s p=.76 CD115 F4/8 hi CD115 + F4/8 + CD115 + F4/8 hi CD115 + F4/8 + CD115 + MHCII MHCII Supplementary Figure S1. CD11b deficiency affects the cellular responses
F4/8 % in the peritoneal lavage 6 4 2 p=.15 n.s p=.76 CD115 F4/8 hi CD115 + F4/8 + CD115 + F4/8 hi CD115 + F4/8 + CD115 + MHCII MHCII Supplementary Figure S1. CD11b deficiency affects the cellular responses
SUPPLEMENTARY INFORMATION. CXCR4 inhibitors could benefit to HER2 but not to Triple-Negative. breast cancer patients
 SUPPLEMENTARY INFORMATION CXCR4 inhibitors could benefit to HER2 but not to Triple-Negative breast cancer patients Lefort S. 1,2, Thuleau A. 3, Kieffer Y. 1,2, Sirven P. 1,2, Bieche I. 4, Marangoni E.
SUPPLEMENTARY INFORMATION CXCR4 inhibitors could benefit to HER2 but not to Triple-Negative breast cancer patients Lefort S. 1,2, Thuleau A. 3, Kieffer Y. 1,2, Sirven P. 1,2, Bieche I. 4, Marangoni E.
x Lymphocyte count /µl CD8+ count/µl 800 Calculated
 % Lymphocyte in CBC A. 50 40 30 20 10 Lymphocyte count /µl B. x10 3 2.5 1.5 C. 50 D. 1000 % CD3+CD8+ Cells 40 30 20 Calculated CD8+ count/µl 800 600 400 200 10 0 #61 #63 #64 #65 #68 #71 #72 #75 Figure
% Lymphocyte in CBC A. 50 40 30 20 10 Lymphocyte count /µl B. x10 3 2.5 1.5 C. 50 D. 1000 % CD3+CD8+ Cells 40 30 20 Calculated CD8+ count/µl 800 600 400 200 10 0 #61 #63 #64 #65 #68 #71 #72 #75 Figure
HIV-infected subjects were recruited from an ongoing clinic-based cohort (SCOPE)
 SUPPLEMENTAL METHODS Populations studied HIV-infected subjects were recruited from an ongoing clinic-based cohort (SCOPE) based at the University of California, San Francisco. From this cohort, we recruited
SUPPLEMENTAL METHODS Populations studied HIV-infected subjects were recruited from an ongoing clinic-based cohort (SCOPE) based at the University of California, San Francisco. From this cohort, we recruited
CD25-PE (BD Biosciences) and labeled with anti-pe-microbeads (Miltenyi Biotec) for depletion of CD25 +
 Supplements Supplemental Materials and Methods Depletion of CD25 + T-cells from PBMC. Fresh or HD precultured PBMC were stained with the conjugate CD25-PE (BD Biosciences) and labeled with anti-pe-microbeads
Supplements Supplemental Materials and Methods Depletion of CD25 + T-cells from PBMC. Fresh or HD precultured PBMC were stained with the conjugate CD25-PE (BD Biosciences) and labeled with anti-pe-microbeads
MED B Form CLL. Johannes Schetelig. London 09/April/
 www.ebmt.org MED B Form CLL Johannes Schetelig London 09/April/2013 Content Update on CLL (15 ) Experiment with mini MED B CLL Assessment of pre-treatment in CLL Cytogenetics in CLL What is the IGVH-Gene
www.ebmt.org MED B Form CLL Johannes Schetelig London 09/April/2013 Content Update on CLL (15 ) Experiment with mini MED B CLL Assessment of pre-treatment in CLL Cytogenetics in CLL What is the IGVH-Gene
ACALABRUTINIB IN MCL
 ACALABRUTINIB IN MCL Simon Rule Professor of Clinical Haematology Consultant Haematologist Derriford Hospital and Peninsula Medical School Plymouth UK BRUTON S TYROSINE KINASE (BTK): A CRITICAL KINASE
ACALABRUTINIB IN MCL Simon Rule Professor of Clinical Haematology Consultant Haematologist Derriford Hospital and Peninsula Medical School Plymouth UK BRUTON S TYROSINE KINASE (BTK): A CRITICAL KINASE
Islet viability assay and Glucose Stimulated Insulin Secretion assay RT-PCR and Western Blot
 Islet viability assay and Glucose Stimulated Insulin Secretion assay Islet cell viability was determined by colorimetric (3-(4,5-dimethylthiazol-2-yl)-2,5- diphenyltetrazolium bromide assay using CellTiter
Islet viability assay and Glucose Stimulated Insulin Secretion assay Islet cell viability was determined by colorimetric (3-(4,5-dimethylthiazol-2-yl)-2,5- diphenyltetrazolium bromide assay using CellTiter
Essential Medium, containing 10% fetal bovine serum, 100 U/ml penicillin and 100 µg/ml streptomycin. Huvec were cultured in
 Supplemental data Methods Cell culture media formulations A-431 and U-87 MG cells were maintained in Dulbecco s Modified Eagle s Medium. FaDu cells were cultured in Eagle's Minimum Essential Medium, containing
Supplemental data Methods Cell culture media formulations A-431 and U-87 MG cells were maintained in Dulbecco s Modified Eagle s Medium. FaDu cells were cultured in Eagle's Minimum Essential Medium, containing
DURACLONE IF BE CERTAIN ABOUT THE RESPONSE. l res. a il n c n. For Research Use Only - Not for use in Diagnostic procedures
 DURACLONE IF earch tria l res lc a om ic il n c n nio pa Yo ur BE CERTAIN ABOUT THE RESPONSE For Research Use Only - Not for use in Diagnostic procedures BE CERTAIN ABOUT THE RESPONSE The sensitive and
DURACLONE IF earch tria l res lc a om ic il n c n nio pa Yo ur BE CERTAIN ABOUT THE RESPONSE For Research Use Only - Not for use in Diagnostic procedures BE CERTAIN ABOUT THE RESPONSE The sensitive and
IMMUNOLOGY. Source, Isolate, Culture, And Analyze Immune Cells. Scientists Helping Scientists
 IMMUNOLOGY Source, Isolate, Culture, And Analyze Immune Cells Scientists Helping Scientists WWW.STEMCELL.COM TABLE OF CONTENTS Tools For Your Immunology Research 4 Primary Cells: It All Starts with The
IMMUNOLOGY Source, Isolate, Culture, And Analyze Immune Cells Scientists Helping Scientists WWW.STEMCELL.COM TABLE OF CONTENTS Tools For Your Immunology Research 4 Primary Cells: It All Starts with The
Canberra, Australia). CD11c-DTR-OVA-GFP (B6.CD11c-OVA), B6.luc + and. Cancer Research Center, Germany). B6 or BALB/c.FoxP3-DTR-GFP mice were
 Supplemental Materials and Methods Mice Female C57BL/6 (B6, I-E null, H-2 b ), BALB/c (H-2 d ) + ), FVB/N (H-2 q, I-E null, CD45.1 + ), and B6D2F1 (H-2 b/d ) mice were purchased from the Animal Resources
Supplemental Materials and Methods Mice Female C57BL/6 (B6, I-E null, H-2 b ), BALB/c (H-2 d ) + ), FVB/N (H-2 q, I-E null, CD45.1 + ), and B6D2F1 (H-2 b/d ) mice were purchased from the Animal Resources
Interleukin-2-Dependent Allergen-Specific Tissue-Resident Memory Cells Drive Asthma
 Immunity Supplemental Information Interleukin-2-Dependent Allergen-Specific Tissue-Resident Memory Cells Drive Asthma Brian D. Hondowicz, Dowon An, Jason M. Schenkel, Karen S. Kim, Holly R. Steach, Akshay
Immunity Supplemental Information Interleukin-2-Dependent Allergen-Specific Tissue-Resident Memory Cells Drive Asthma Brian D. Hondowicz, Dowon An, Jason M. Schenkel, Karen S. Kim, Holly R. Steach, Akshay
Supplementary Information
 Supplementary Information Supplementary Figure 1. CD4 + T cell activation and lack of apoptosis after crosslinking with anti-cd3 + anti-cd28 + anti-cd160. (a) Flow cytometry of anti-cd160 (5D.10A11) binding
Supplementary Information Supplementary Figure 1. CD4 + T cell activation and lack of apoptosis after crosslinking with anti-cd3 + anti-cd28 + anti-cd160. (a) Flow cytometry of anti-cd160 (5D.10A11) binding
Supplementary Figure 1: Expression of NFAT proteins in Nfat2-deleted B cells (a+b) Protein expression of NFAT2 (a) and NFAT1 (b) in isolated splenic
 Supplementary Figure 1: Expression of NFAT proteins in Nfat2-deleted B cells (a+b) Protein expression of NFAT2 (a) and NFAT1 (b) in isolated splenic B cells from WT Nfat2 +/+, TCL1 Nfat2 +/+ and TCL1 Nfat2
Supplementary Figure 1: Expression of NFAT proteins in Nfat2-deleted B cells (a+b) Protein expression of NFAT2 (a) and NFAT1 (b) in isolated splenic B cells from WT Nfat2 +/+, TCL1 Nfat2 +/+ and TCL1 Nfat2
X P. Supplementary Figure 1. Nature Medicine: doi: /nm Nilotinib LSK LT-HSC. Cytoplasm. Cytoplasm. Nucleus. Nucleus
 a b c Supplementary Figure 1 c-kit-apc-eflu780 Lin-FITC Flt3-Linc-Kit-APC-eflu780 LSK Sca-1-PE-Cy7 d e f CD48-APC LT-HSC CD150-PerCP-cy5.5 g h i j Cytoplasm RCC1 X Exp 5 mir 126 SPRED1 SPRED1 RAN P SPRED1
a b c Supplementary Figure 1 c-kit-apc-eflu780 Lin-FITC Flt3-Linc-Kit-APC-eflu780 LSK Sca-1-PE-Cy7 d e f CD48-APC LT-HSC CD150-PerCP-cy5.5 g h i j Cytoplasm RCC1 X Exp 5 mir 126 SPRED1 SPRED1 RAN P SPRED1
Chitin Activates Parallel Immune Modules that Direct Distinct Inflammatory Responses via Innate Lymphoid Type 2 and T Cells
 Immunity, Volume 4 Supplemental Information Chitin Activates Parallel Immune Modules that Direct Distinct Inflammatory Responses via Innate Lymphoid Type 2 and T Cells Steven J. Van Dyken, Alexander Mohapatra,
Immunity, Volume 4 Supplemental Information Chitin Activates Parallel Immune Modules that Direct Distinct Inflammatory Responses via Innate Lymphoid Type 2 and T Cells Steven J. Van Dyken, Alexander Mohapatra,
