Cbl ubiquitin ligase: Lord of the RINGs
|
|
|
- Shawn Elliott
- 5 years ago
- Views:
Transcription
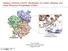
1 Cbl ubiquitin ligase: Lord of the RINGs Not just quite interesting - really interesting! A cell must be able to degrade proteins when their activity is no longer required. Many eukaryotic proteins are labelled for destruction by addition of a flag called ubiquitin. This ubiquitination directs proteins along a pathway for degradation. Defects in such pathways are associated with diseases such as cancer, neurodegenerative disorders and viral infections. Fig1. RING E3 ligase transfers ubiquitin from the E2 active site to the substrate. Flagging a protein with ubiquitin is done by enzymes called ubiquitin ligases. There are many such enzymes in the cell which flag different sorts of proteins, and one which is quite interesting is Cbl (also called c-cbl) [view-1]. Cbl is a member of the RING E3 class of ubiquitin ligases, so called because it contains a RING (Really Interesting New Gene) domain. Members of the RING E3 family are the final protein in a cascade of three enzymes and act to transfer ubiquitin from a donor protein, E2 ubiquitinconjugating enzyme (E2), to the protein that is to be flagged for destruction. (graphic 1). Cbl specifically transfers ubiquitin to proteins in the receptor tyrosine kinase (RTK) family which is important in cell signalling. One RING to bind E2 Cbl is a multi-domain protein (view-1). A tyrosine kinase-binding domain (TKBD) which binds RTK substrates is found at the N-terminus. It is connected to the RING domain via a linker-helix region. The RING domain is followed by a proline-rich region and, finally, a ubiquitin-associated (UBA) domain. Only the first three domains, TKBD, linker helix and RING, are needed for activity. The structure of this portion of Cbl has been solved (PDB 1fbv) in complex with its two substrates, E2 and a peptide from the tyrosine kinase.
2 The structure reveals the domain architecture of Cbl and the location of the substratebinding sites. It turns out that the RING domain binds the ubiquitin-donor protein E2 and the TKBD binds the RTK substrate. However, these two substrates, ubiquitin donor and acceptor, are located on opposite sides of the Cbl molecule, nearly 70 Å apart! (view- 2). It would be impossible to transfer the ubiquitin between the two substrates in this conformation. Cbl acts not only as a E3 ubiquitin ligase but also as an adaptor in RTK signal transduction. In this role, the enzyme must bind its target protein, but crucially should not modify it by addition of ubiquitin, i.e., its enzymatic function should be suppressed. Which of the two functions Cbl performs is controlled by phosphorylation of a single tyrosine residue at position 371, located in the linker helix. Phosphorylation enhances the ligase activity and removing the phosphate drastically reduces it. Defects or mutations in Cbl, particularly those involving Tyr371, are related to human myeloid disorders. In fact, this residue is frequently mutated in people with certain forms of leukemia. Obviously, it would be really interesting to find out how phosphorylation of Tyr371 controls the enzyme s activity. The Huang laboratory in Glasgow has recently determined the structure of Cbl with and without substrates. They have also determined the structures of Cbl in both the dephosphorylated and phosphorylated forms, shedding light on how the phosphorylation switch activates Cbl and influences the substrate-binding sites (Reference 1). Moving the RING In de-phosphorylated Cbl (PDB 2y1m), the RING domain is packed against the TKBD, with its E2-binding surface buried, thus preventing E2 from binding. This state (view-3) is therefore an auto-inhibited conformation. Binding of a substrate peptide to the opposite face of the TKBD induces a slight movement of the linker helix (PDB 2y1n), establishing a new hydrophobic interaction between the linker helix and the TKBD that shifts the RING domain into a half-open conformation (view-4). In this conformation, E2 is able to bind to the RING, but it competes with the TKBD for binding. Tyr371 and Tyr368, residues located in the linker helix, are buried in the TKBD, thereby locking the linker helix to the TKBD (view-5). Therefore, when Cbl is de-phosphorylated and substrate is bound, the RING domain fluctuates between a closed, auto-inhibited conformation and an open conformation. Though this open state may be catalytically competent, the E2 and the substrate are still on opposite sides of Cbl and the enzyme has a very low catalytic rate. This makes the de-phosphorylated Cbl a poor enzyme, but allows it to act as an ideal adaptor. RING flipping Upon phosphorylation, Tyr371 can no longer be accommodated by the hydrophobic pocket on the TKBD, and therefore the linker helix is unlocked from the TKBD. This movement fully exposes the E2-binding face of the RING domain, enabling it to bind E2 without competition. Thus, phosphorylation of Tyr371 enhances ligase activity by abolishing the auto-inhibition of Cbl.
3 But phosphorylation doesn t just free up the RING domain, it also induces huge conformational changes that flip the linker helix 180 compared with the dephosphorylated state (PDB 4a4c) (view-6). As a result, the RING domain and its bound E2 move to the same side of Cbl as the substrate peptide. This shortens the distance between the active site of the E2, where ubiquitin is attached, and the TKBD-substratebinding site to 28 Å. This is still a large distance, but in this structure only a small fragment of the substrate is bound; the real substrate is a protein of 619 amino acids. The structures show that phophorylation of Tyr371 enhances Cbl s enzymatic activity both by preventing the RING domain from adopting its auto-inhibited state and by moving it into a much more favourable position for ubiquitin transfer. Mutations or deletions in the linker helix and RING domains would affect the activity of Cbl and potentially provide a starting point for the design of new anti-cancer therapeutics. This Quips article was developed in collaboration with Hao Dou and Danny Huang at the Beatson Institute for Cancer Research. References Dou H. et al. Nat.Struct.Mol.Biol Jan 22;19(2): PubMed PMID: VIEWS
4 View 1 Domain structure of Cbl E3 Ubiquitin Ligase. The Tyrosine Kinase Binding Domain (TKBD) is shown in blue and the RING domain in orange. The linker-helix region which connects these two domains is yellow. View 2 Substrate binding to Cbl. E2 Ubiquitin-conjugating enzyme (which donates ubiquitin) is shown in cyan. A peptide from the substrate molecule to which ubiquitin is transferred is shown as a spacefilling model. These two molecules are positioned more than 70Å apart on opposite sides of Cbl.
5 View 3 Without substrates, the RING domain and linker helix are packed against the TKBD. The RING domain (orange) is bound to the TKBD, occluding its E2 binding surface and making E2 recruitment impossible. Residues involved in the interface are highlighted. View 4 Binding a substrate peptide causes movement of the distant RING domain. On binding a substrate peptide (space-filling model), the RING domain (orange) moves to expose the E2 binding face.
6 View 5 Key tyrosine residues in the linker helix (yellow) are buried in the TKBD (blue). Tyrosines 368 (cyan) and 371 (pink) are buried in hydrophobic pockets on the TKBD surface, anchoring the linker helix to the surface of the TKBD and locking its position. Tyr 371 is unable to fit in this position once phosphorylated. View 6 Phosphorylation of Tyr 371 dislodges the linker helix from the TKBD triggers a large-scale conformational change. Following phosphorylation, the linker helix and RING domain, along with the bound E2 are able to swing 180 to bring E2 (cyan) within range of the substrate (space-filling model).
Effects of Second Messengers
 Effects of Second Messengers Inositol trisphosphate Diacylglycerol Opens Calcium Channels Binding to IP 3 -gated Channel Cooperative binding Activates Protein Kinase C is required Phosphorylation of many
Effects of Second Messengers Inositol trisphosphate Diacylglycerol Opens Calcium Channels Binding to IP 3 -gated Channel Cooperative binding Activates Protein Kinase C is required Phosphorylation of many
Signal-Transduction Cascades - 2. The Phosphoinositide Cascade
 Signal-Transduction Cascades - 2 The Phosphoinositide Cascade Calcium ion as a second messenger Tyrosine kinase and receptor dimerization scribd.com Faisal Khatib JU The Phosphoinositide Cascade Used by
Signal-Transduction Cascades - 2 The Phosphoinositide Cascade Calcium ion as a second messenger Tyrosine kinase and receptor dimerization scribd.com Faisal Khatib JU The Phosphoinositide Cascade Used by
(B D) Three views of the final refined 2Fo-Fc electron density map of the Vpr (red)-ung2 (green) interacting region, contoured at 1.4σ.
 Supplementary Figure 1 Overall structure of the DDB1 DCAF1 Vpr UNG2 complex. (A) The final refined 2Fo-Fc electron density map, contoured at 1.4σ of Vpr, illustrating well-defined side chains. (B D) Three
Supplementary Figure 1 Overall structure of the DDB1 DCAF1 Vpr UNG2 complex. (A) The final refined 2Fo-Fc electron density map, contoured at 1.4σ of Vpr, illustrating well-defined side chains. (B D) Three
Molecular Medicine: Gleevec and Chronic Myelogenous Leukemia. Dec 14 & 19, 2006 Prof. Erin O Shea Prof. Dan Kahne
 Molecular Medicine: Gleevec and Chronic Myelogenous Leukemia Dec 14 & 19, 2006 Prof. Erin Shea Prof. Dan Kahne 1 Cancer, Kinases and Gleevec: 1. What is CML? a. Blood cell maturation b. Philadelphia Chromosome
Molecular Medicine: Gleevec and Chronic Myelogenous Leukemia Dec 14 & 19, 2006 Prof. Erin Shea Prof. Dan Kahne 1 Cancer, Kinases and Gleevec: 1. What is CML? a. Blood cell maturation b. Philadelphia Chromosome
Tala Saleh. Ahmad Attari. Mamoun Ahram
 23 Tala Saleh Ahmad Attari Minna Mushtaha Mamoun Ahram In the previous lecture, we discussed the mechanisms of regulating enzymes through inhibitors. Now, we will start this lecture by discussing regulation
23 Tala Saleh Ahmad Attari Minna Mushtaha Mamoun Ahram In the previous lecture, we discussed the mechanisms of regulating enzymes through inhibitors. Now, we will start this lecture by discussing regulation
This PDF file includes: Supplementary Figures 1 to 6 Supplementary Tables 1 to 2 Supplementary Methods Supplementary References
 Structure of the catalytic core of p300 and implications for chromatin targeting and HAT regulation Manuela Delvecchio, Jonathan Gaucher, Carmen Aguilar-Gurrieri, Esther Ortega, Daniel Panne This PDF file
Structure of the catalytic core of p300 and implications for chromatin targeting and HAT regulation Manuela Delvecchio, Jonathan Gaucher, Carmen Aguilar-Gurrieri, Esther Ortega, Daniel Panne This PDF file
Patrick, An Introduction to Medicinal Chemistry 4e Chapter 5 Receptors and signal transduction
 atrick, An Introduction to Medicinal Chemistry 4e Answers to end-of-chapter questions 1) The diagram in the question shows two important hydrogen bonding interactions where AT acts both as a hydrogen bond
atrick, An Introduction to Medicinal Chemistry 4e Answers to end-of-chapter questions 1) The diagram in the question shows two important hydrogen bonding interactions where AT acts both as a hydrogen bond
G-Protein Signaling. Introduction to intracellular signaling. Dr. SARRAY Sameh, Ph.D
 G-Protein Signaling Introduction to intracellular signaling Dr. SARRAY Sameh, Ph.D Cell signaling Cells communicate via extracellular signaling molecules (Hormones, growth factors and neurotransmitters
G-Protein Signaling Introduction to intracellular signaling Dr. SARRAY Sameh, Ph.D Cell signaling Cells communicate via extracellular signaling molecules (Hormones, growth factors and neurotransmitters
Previous Class. Today. Detection of enzymatic intermediates: Protein tyrosine phosphatase mechanism. Protein Kinase Catalytic Properties
 Previous Class Detection of enzymatic intermediates: Protein tyrosine phosphatase mechanism Today Protein Kinase Catalytic Properties Protein Phosphorylation Phosphorylation: key protein modification
Previous Class Detection of enzymatic intermediates: Protein tyrosine phosphatase mechanism Today Protein Kinase Catalytic Properties Protein Phosphorylation Phosphorylation: key protein modification
Enzymes Part III: regulation II. Dr. Mamoun Ahram Summer, 2017
 Enzymes Part III: regulation II Dr. Mamoun Ahram Summer, 2017 Advantage This is a major mechanism for rapid and transient regulation of enzyme activity. A most common mechanism is enzyme phosphorylation
Enzymes Part III: regulation II Dr. Mamoun Ahram Summer, 2017 Advantage This is a major mechanism for rapid and transient regulation of enzyme activity. A most common mechanism is enzyme phosphorylation
Excerpt from J. Mol. Biol. (2002) 320, :
 Excerpt from J. Mol. Biol. (2002) 320, 1095 1108: Crystal Structure of the Ternary Complex of the Catalytic Domain of Human Phenylalanine Hydroxylase with Tetrahydrobiopterin and 3-(2-Thienyl)-L-alanine,
Excerpt from J. Mol. Biol. (2002) 320, 1095 1108: Crystal Structure of the Ternary Complex of the Catalytic Domain of Human Phenylalanine Hydroxylase with Tetrahydrobiopterin and 3-(2-Thienyl)-L-alanine,
Enzyme-coupled Receptors. Cell-surface receptors 1. Ion-channel-coupled receptors 2. G-protein-coupled receptors 3. Enzyme-coupled receptors
 Enzyme-coupled Receptors Cell-surface receptors 1. Ion-channel-coupled receptors 2. G-protein-coupled receptors 3. Enzyme-coupled receptors Cell-surface receptors allow a flow of ions across the plasma
Enzyme-coupled Receptors Cell-surface receptors 1. Ion-channel-coupled receptors 2. G-protein-coupled receptors 3. Enzyme-coupled receptors Cell-surface receptors allow a flow of ions across the plasma
The elements of G protein-coupled receptor systems
 The elements of G protein-coupled receptor systems Prostaglandines Sphingosine 1-phosphate a receptor that contains 7 membrane-spanning domains a coupled trimeric G protein which functions as a switch
The elements of G protein-coupled receptor systems Prostaglandines Sphingosine 1-phosphate a receptor that contains 7 membrane-spanning domains a coupled trimeric G protein which functions as a switch
Signaling. Dr. Sujata Persad Katz Group Centre for Pharmacy & Health research
 Signaling Dr. Sujata Persad 3-020 Katz Group Centre for Pharmacy & Health research E-mail:sujata.persad@ualberta.ca 1 Growth Factor Receptors and Other Signaling Pathways What we will cover today: How
Signaling Dr. Sujata Persad 3-020 Katz Group Centre for Pharmacy & Health research E-mail:sujata.persad@ualberta.ca 1 Growth Factor Receptors and Other Signaling Pathways What we will cover today: How
Amino Acids. Review I: Protein Structure. Amino Acids: Structures. Amino Acids (contd.) Rajan Munshi
 Review I: Protein Structure Rajan Munshi BBSI @ Pitt 2005 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2005 Amino Acids Building blocks of proteins 20 amino acids
Review I: Protein Structure Rajan Munshi BBSI @ Pitt 2005 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2005 Amino Acids Building blocks of proteins 20 amino acids
Chemistry 135, First Exam. September 23, Chem 135, Exam 1 SID:
 Chemistry 135, First Exam September 23, 2015 This exam will be worth 15% of your overall grade. Please read all instructions/questions carefully and provide answers in the space provided. There should
Chemistry 135, First Exam September 23, 2015 This exam will be worth 15% of your overall grade. Please read all instructions/questions carefully and provide answers in the space provided. There should
The ubiquitin-proteasome system. in bone biology. University of Nottingham. ECTS PhD Training July 5 th Dr Rob Layfield
 The ubiquitin-proteasome system in bone biology Dr Rob Layfield University of Nottingham ECTS PhD Training July 5 th 2009 Images reproduced from: http://www.bostonbiochem.com/overview.php?prod=ubchains
The ubiquitin-proteasome system in bone biology Dr Rob Layfield University of Nottingham ECTS PhD Training July 5 th 2009 Images reproduced from: http://www.bostonbiochem.com/overview.php?prod=ubchains
Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site.
 Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
Chapter 11. Cell Communication. Signal Transduction Pathways
 Chapter 11 Cell Communication Signal Transduction Pathways Signal-Transduction Pathway Signal on a cell s surface is converted into a specific cellular response Local signaling (short distance) - Paracrine
Chapter 11 Cell Communication Signal Transduction Pathways Signal-Transduction Pathway Signal on a cell s surface is converted into a specific cellular response Local signaling (short distance) - Paracrine
3.2 Ligand-Binding at Nicotinic Acid Receptor Subtypes GPR109A/B
 3.2 Ligand-Binding at Nicotinic Acid Receptor Subtypes GPR109A/B 3.2.1 Characterization of the Ligand Binding Site at GPR109A Receptor Ligands of GPR109A Receptor are Carboxylic Acids Nicotinic acid (pyridine-3-carboxylic
3.2 Ligand-Binding at Nicotinic Acid Receptor Subtypes GPR109A/B 3.2.1 Characterization of the Ligand Binding Site at GPR109A Receptor Ligands of GPR109A Receptor are Carboxylic Acids Nicotinic acid (pyridine-3-carboxylic
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 The UBL and RING1 interface remains associated in the complex structures of Parkin and pub. a) Asymmetric Unit of crystal structure of UBLR0RBR and pub complex showing UBL (green),
Supplementary Figure 1 The UBL and RING1 interface remains associated in the complex structures of Parkin and pub. a) Asymmetric Unit of crystal structure of UBLR0RBR and pub complex showing UBL (green),
Chapter 31. Completing the Protein Life Cycle: Folding, Processing and Degradation. Biochemistry by Reginald Garrett and Charles Grisham
 Chapter 31 Completing the Protein Life Cycle: Folding, Processing and Degradation Biochemistry by Reginald Garrett and Charles Grisham Essential Question How are newly synthesized polypeptide chains transformed
Chapter 31 Completing the Protein Life Cycle: Folding, Processing and Degradation Biochemistry by Reginald Garrett and Charles Grisham Essential Question How are newly synthesized polypeptide chains transformed
Introduction to proteins and protein structure
 Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
Structural biology of viruses
 Structural biology of viruses Biophysical Chemistry 1, Fall 2010 Coat proteins DNA/RNA packaging Reading assignment: Chap. 15 Virus particles self-assemble from coat monomers Virus Structure and Function
Structural biology of viruses Biophysical Chemistry 1, Fall 2010 Coat proteins DNA/RNA packaging Reading assignment: Chap. 15 Virus particles self-assemble from coat monomers Virus Structure and Function
Lecture 10 More about proteins
 Lecture 10 More about proteins Today we're going to extend our discussion of protein structure. This may seem far-removed from gene cloning, but it is the path to understanding the genes that we are cloning.
Lecture 10 More about proteins Today we're going to extend our discussion of protein structure. This may seem far-removed from gene cloning, but it is the path to understanding the genes that we are cloning.
Lecture 5: Drug targets (continued)
 Lecture 5: Drug targets (continued) IIa. Enzymes as drug targets (HMG-CoA example) Many drugs are inhibitors of enzymes that catalyze biologically important reactions. The conversion of HMG-CoA to mevalonic
Lecture 5: Drug targets (continued) IIa. Enzymes as drug targets (HMG-CoA example) Many drugs are inhibitors of enzymes that catalyze biologically important reactions. The conversion of HMG-CoA to mevalonic
Cell Biology Lecture 9 Notes Basic Principles of cell signaling and GPCR system
 Cell Biology Lecture 9 Notes Basic Principles of cell signaling and GPCR system Basic Elements of cell signaling: Signal or signaling molecule (ligand, first messenger) o Small molecules (epinephrine,
Cell Biology Lecture 9 Notes Basic Principles of cell signaling and GPCR system Basic Elements of cell signaling: Signal or signaling molecule (ligand, first messenger) o Small molecules (epinephrine,
Cellular Signaling Pathways. Signaling Overview
 Cellular Signaling Pathways Signaling Overview Signaling steps Synthesis and release of signaling molecules (ligands) by the signaling cell. Transport of the signal to the target cell Detection of the
Cellular Signaling Pathways Signaling Overview Signaling steps Synthesis and release of signaling molecules (ligands) by the signaling cell. Transport of the signal to the target cell Detection of the
Molecular Medicine: Gleevec and Chronic Myelogenous Leukemia
 Molecular Medicine: Gleevec and Chronic Myelogenous Leukemia Dec 14 & 19, 2006 Prof. Erin Shea Prof. Dan Kahne Cancer, Kinases and Gleevec: 1. What is CML? a. Blood cell maturation b. Philadelphia Chromosome
Molecular Medicine: Gleevec and Chronic Myelogenous Leukemia Dec 14 & 19, 2006 Prof. Erin Shea Prof. Dan Kahne Cancer, Kinases and Gleevec: 1. What is CML? a. Blood cell maturation b. Philadelphia Chromosome
Proteins consist of joined amino acids They are joined by a Also called an Amide Bond
 Lecture Two: Peptide Bond & Protein Structure [Chapter 2 Berg, Tymoczko & Stryer] (Figures in Red are for the 7th Edition) (Figures in Blue are for the 8th Edition) Proteins consist of joined amino acids
Lecture Two: Peptide Bond & Protein Structure [Chapter 2 Berg, Tymoczko & Stryer] (Figures in Red are for the 7th Edition) (Figures in Blue are for the 8th Edition) Proteins consist of joined amino acids
Bio 111 Study Guide Chapter 11 Cell Communication
 Bio 111 Study Guide Chapter 11 Cell Communication BEFORE CLASS: Reading: Read the introduction on p. 210, and for Concept 11.1, read from the first full paragraph on p. 212. Read all of Concept 11.2. Pay
Bio 111 Study Guide Chapter 11 Cell Communication BEFORE CLASS: Reading: Read the introduction on p. 210, and for Concept 11.1, read from the first full paragraph on p. 212. Read all of Concept 11.2. Pay
Signal Transduction: G-Protein Coupled Receptors
 Signal Transduction: G-Protein Coupled Receptors Federle, M. (2017). Lectures 4-5: Signal Transduction parts 1&2: nuclear receptors and GPCRs. Lecture presented at PHAR 423 Lecture in UIC College of Pharmacy,
Signal Transduction: G-Protein Coupled Receptors Federle, M. (2017). Lectures 4-5: Signal Transduction parts 1&2: nuclear receptors and GPCRs. Lecture presented at PHAR 423 Lecture in UIC College of Pharmacy,
Chapter 9. Cellular Signaling
 Chapter 9 Cellular Signaling Cellular Messaging Page 215 Cells can signal to each other and interpret the signals they receive from other cells and the environment Signals are most often chemicals The
Chapter 9 Cellular Signaling Cellular Messaging Page 215 Cells can signal to each other and interpret the signals they receive from other cells and the environment Signals are most often chemicals The
Simulate enzymatic actions. Explain enzymatic specificity. Investigate two types of enzyme inhibitors used in regulating enzymatic activity.
 Name: Enzymes in Action Objectives: You will use the model pieces in the kit to: Simulate enzymatic actions. Explain enzymatic specificity. Investigate two types of enzyme inhibitors used in regulating
Name: Enzymes in Action Objectives: You will use the model pieces in the kit to: Simulate enzymatic actions. Explain enzymatic specificity. Investigate two types of enzyme inhibitors used in regulating
Regulation of cell function by intracellular signaling
 Regulation of cell function by intracellular signaling Objectives: Regulation principle Allosteric and covalent mechanisms, Popular second messengers, Protein kinases, Kinase cascade and interaction. regulation
Regulation of cell function by intracellular signaling Objectives: Regulation principle Allosteric and covalent mechanisms, Popular second messengers, Protein kinases, Kinase cascade and interaction. regulation
Chapter 23 Enzymes 1
 Chapter 23 Enzymes 1 Enzymes Ribbon diagram of cytochrome c oxidase, the enzyme that directly uses oxygen during respiration. 2 Enzyme Catalysis Enzyme: A biological catalyst. With the exception of some
Chapter 23 Enzymes 1 Enzymes Ribbon diagram of cytochrome c oxidase, the enzyme that directly uses oxygen during respiration. 2 Enzyme Catalysis Enzyme: A biological catalyst. With the exception of some
CHAPTER 10: REGULATORY STRATEGIES. Traffic signals control the flow of traffic
 CHAPTER 10: REGULATORY STRATEGIES Traffic signals control the flow of traffic INTRODUCTION CHAPTER 10 The activity of enzymes must often be regulated so that they function at the proper time and place.
CHAPTER 10: REGULATORY STRATEGIES Traffic signals control the flow of traffic INTRODUCTION CHAPTER 10 The activity of enzymes must often be regulated so that they function at the proper time and place.
Enzymes: Regulation 2-3
 Enzymes: Regulation 2-3 Reversible covalent modification Association with regulatory proteins Irreversible covalent modification/proteolytic cleavage Reading: Berg, Tymoczko & Stryer, 6th ed., Chapter
Enzymes: Regulation 2-3 Reversible covalent modification Association with regulatory proteins Irreversible covalent modification/proteolytic cleavage Reading: Berg, Tymoczko & Stryer, 6th ed., Chapter
Receptor mediated Signal Transduction
 Receptor mediated Signal Transduction G-protein-linked receptors adenylyl cyclase camp PKA Organization of receptor protein-tyrosine kinases From G.M. Cooper, The Cell. A molecular approach, 2004, third
Receptor mediated Signal Transduction G-protein-linked receptors adenylyl cyclase camp PKA Organization of receptor protein-tyrosine kinases From G.M. Cooper, The Cell. A molecular approach, 2004, third
Main Functions maintain homeostasis
 The Cell Membrane Main Functions The main goal is to maintain homeostasis. Regulates materials moving in and out of the cell. Provides a large surface area on which specific chemical reactions can occur.
The Cell Membrane Main Functions The main goal is to maintain homeostasis. Regulates materials moving in and out of the cell. Provides a large surface area on which specific chemical reactions can occur.
Statin inhibition of HMG-CoA reductase: a 3-dimensional view
 Atherosclerosis Supplements 4 (2003) 3/8 www.elsevier.com/locate/atherosclerosis Statin inhibition of HMG-CoA reductase: a 3-dimensional view Eva Istvan * Department of Molecular Microbiology, Howard Hughes
Atherosclerosis Supplements 4 (2003) 3/8 www.elsevier.com/locate/atherosclerosis Statin inhibition of HMG-CoA reductase: a 3-dimensional view Eva Istvan * Department of Molecular Microbiology, Howard Hughes
Tyrosine kinases. Cell surface receptors ligand binding. Producer cell RNA. Target cell
 Tyrosine kinases http://msbl.helsinki.fi/tkseminar roducer cell Signaling molecules Receptor binding Signal transduction Target cell Activation of Gene expression RNA Biological responses proliferation,
Tyrosine kinases http://msbl.helsinki.fi/tkseminar roducer cell Signaling molecules Receptor binding Signal transduction Target cell Activation of Gene expression RNA Biological responses proliferation,
Cancer and Tyrosine Kinase Inhibition
 Cancer and Tyrosine Kinase Inhibition 1 Motivation (1) In first world countries cancer is the second most common cause of death after cardiovascular diseases. For most tumors, treatment is limited to surgery,
Cancer and Tyrosine Kinase Inhibition 1 Motivation (1) In first world countries cancer is the second most common cause of death after cardiovascular diseases. For most tumors, treatment is limited to surgery,
Signal Transduction Cascades
 Signal Transduction Cascades Contents of this page: Kinases & phosphatases Protein Kinase A (camp-dependent protein kinase) G-protein signal cascade Structure of G-proteins Small GTP-binding proteins,
Signal Transduction Cascades Contents of this page: Kinases & phosphatases Protein Kinase A (camp-dependent protein kinase) G-protein signal cascade Structure of G-proteins Small GTP-binding proteins,
Chapter 15: Signal transduction
 Chapter 15: Signal transduction Know the terminology: Enzyme-linked receptor, G-protein linked receptor, nuclear hormone receptor, G-protein, adaptor protein, scaffolding protein, SH2 domain, MAPK, Ras,
Chapter 15: Signal transduction Know the terminology: Enzyme-linked receptor, G-protein linked receptor, nuclear hormone receptor, G-protein, adaptor protein, scaffolding protein, SH2 domain, MAPK, Ras,
The three important structural features of proteins:
 The three important structural features of proteins: a. Primary (1 o ) The amino acid sequence (coded by genes) b. Secondary (2 o ) The interaction of amino acids that are close together or far apart in
The three important structural features of proteins: a. Primary (1 o ) The amino acid sequence (coded by genes) b. Secondary (2 o ) The interaction of amino acids that are close together or far apart in
Protein Secondary Structure
 Protein Secondary Structure Reading: Berg, Tymoczko & Stryer, 6th ed., Chapter 2, pp. 37-45 Problems in textbook: chapter 2, pp. 63-64, #1,5,9 Directory of Jmol structures of proteins: http://www.biochem.arizona.edu/classes/bioc462/462a/jmol/routines/routines.html
Protein Secondary Structure Reading: Berg, Tymoczko & Stryer, 6th ed., Chapter 2, pp. 37-45 Problems in textbook: chapter 2, pp. 63-64, #1,5,9 Directory of Jmol structures of proteins: http://www.biochem.arizona.edu/classes/bioc462/462a/jmol/routines/routines.html
Molecular Cell Biology - Problem Drill 19: Cell Signaling Pathways and Gene Expression
 Molecular Cell Biology - Problem Drill 19: Cell Signaling Pathways and Gene Expression Question No. 1 of 10 1. Which statement about cell signaling is correct? Question #1 (A) Cell signaling involves receiving
Molecular Cell Biology - Problem Drill 19: Cell Signaling Pathways and Gene Expression Question No. 1 of 10 1. Which statement about cell signaling is correct? Question #1 (A) Cell signaling involves receiving
ARV Mode of Action. Mode of Action. Mode of Action NRTI. Immunopaedia.org.za
 ARV Mode of Action Mode of Action Mode of Action - NRTI Mode of Action - NNRTI Mode of Action - Protease Inhibitors Mode of Action - Integrase inhibitor Mode of Action - Entry Inhibitors Mode of Action
ARV Mode of Action Mode of Action Mode of Action - NRTI Mode of Action - NNRTI Mode of Action - Protease Inhibitors Mode of Action - Integrase inhibitor Mode of Action - Entry Inhibitors Mode of Action
Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase. Biol 405 Molecular Medicine
 Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase Biol 405 Molecular Medicine 1998 Crystal structure of phenylalanine hydroxylase solved. The polypeptide consists of three regions: Regulatory
Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase Biol 405 Molecular Medicine 1998 Crystal structure of phenylalanine hydroxylase solved. The polypeptide consists of three regions: Regulatory
REGULATION OF ENZYME ACTIVITY. Medical Biochemistry, Lecture 25
 REGULATION OF ENZYME ACTIVITY Medical Biochemistry, Lecture 25 Lecture 25, Outline General properties of enzyme regulation Regulation of enzyme concentrations Allosteric enzymes and feedback inhibition
REGULATION OF ENZYME ACTIVITY Medical Biochemistry, Lecture 25 Lecture 25, Outline General properties of enzyme regulation Regulation of enzyme concentrations Allosteric enzymes and feedback inhibition
Cell Signaling part 2
 15 Cell Signaling part 2 Functions of Cell Surface Receptors Other cell surface receptors are directly linked to intracellular enzymes. The largest family of these is the receptor protein tyrosine kinases,
15 Cell Signaling part 2 Functions of Cell Surface Receptors Other cell surface receptors are directly linked to intracellular enzymes. The largest family of these is the receptor protein tyrosine kinases,
This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is worth 2 points.
 MBB 407/511 Molecular Biology and Biochemistry First Examination - October 1, 2002 Name Social Security Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is
MBB 407/511 Molecular Biology and Biochemistry First Examination - October 1, 2002 Name Social Security Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Design of isolated protein and RNC constructs, and homogeneity of purified RNCs. (a) Schematic depicting the design and nomenclature used for all the isolated proteins and RNCs used
Supplementary Figure 1 Design of isolated protein and RNC constructs, and homogeneity of purified RNCs. (a) Schematic depicting the design and nomenclature used for all the isolated proteins and RNCs used
Chapter 10. Regulatory Strategy
 Chapter 10 Regulatory Strategy Regulation of enzymatic activity: 1. Allosteric Control. Allosteric proteins have a regulatory site(s) and multiple functional sites Activity of proteins is regulated by
Chapter 10 Regulatory Strategy Regulation of enzymatic activity: 1. Allosteric Control. Allosteric proteins have a regulatory site(s) and multiple functional sites Activity of proteins is regulated by
Biochemistry 2 Recita0on Amino Acid Metabolism
 Biochemistry 2 Recita0on Amino Acid Metabolism 04-20- 2015 Glutamine and Glutamate as key entry points for NH 4 + Amino acid catabolism Glutamine synthetase enables toxic NH 4 + to combine with glutamate
Biochemistry 2 Recita0on Amino Acid Metabolism 04-20- 2015 Glutamine and Glutamate as key entry points for NH 4 + Amino acid catabolism Glutamine synthetase enables toxic NH 4 + to combine with glutamate
An Introduction to Enzyme Structure and Function
 An Introduction to Enzyme Structure and Function Enzymes Many reactions in living systems are similar to laboratory reactions. 1. Reactions in living systems often occur with the aid of enzymes. 2. Enzymes
An Introduction to Enzyme Structure and Function Enzymes Many reactions in living systems are similar to laboratory reactions. 1. Reactions in living systems often occur with the aid of enzymes. 2. Enzymes
Enzymes: The Catalysts of Life
 Chapter 6 Enzymes: The Catalysts of Life Lectures by Kathleen Fitzpatrick Simon Fraser University Activation Energy and the Metastable State Many thermodynamically feasible reactions in a cell that could
Chapter 6 Enzymes: The Catalysts of Life Lectures by Kathleen Fitzpatrick Simon Fraser University Activation Energy and the Metastable State Many thermodynamically feasible reactions in a cell that could
Protein tyrosine kinase signaling
 rotein tyrosine kinase signaling Serge ROCHE CRBM CNRS/Montpellier University serge.roche@crbm.cnrs.fr rotein phosphorylation on Tyr A central mechanism to control cell communication in a multicellular
rotein tyrosine kinase signaling Serge ROCHE CRBM CNRS/Montpellier University serge.roche@crbm.cnrs.fr rotein phosphorylation on Tyr A central mechanism to control cell communication in a multicellular
Biochem 503 Fall Protein Tyr Phosphatases
 Biochem 503 Fall 2005 Protein Tyr Phosphatases David Brautigan assigned reading: Stoker (2005) J. Endocrin. 185:19-33 History 1981-1982 First description of P-Tyr specific phosphohydrolyase activity in
Biochem 503 Fall 2005 Protein Tyr Phosphatases David Brautigan assigned reading: Stoker (2005) J. Endocrin. 185:19-33 History 1981-1982 First description of P-Tyr specific phosphohydrolyase activity in
Complexity DNA. Genome RNA. Transcriptome. Protein. Proteome. Metabolites. Metabolome
 DNA Genome Complexity RNA Transcriptome Systems Biology Linking all the components of a cell in a quantitative and temporal manner Protein Proteome Metabolites Metabolome Where are the functional elements?
DNA Genome Complexity RNA Transcriptome Systems Biology Linking all the components of a cell in a quantitative and temporal manner Protein Proteome Metabolites Metabolome Where are the functional elements?
Plasma membranes. Plasmodesmata between plant cells. Gap junctions between animal cells Cell junctions. Cell-cell recognition
 Cell Communication Cell Signaling Cell-to-cell communication is essential for multicellular organisms Communicate by chemical messengers Animal and plant cells have cell junctions that directly connect
Cell Communication Cell Signaling Cell-to-cell communication is essential for multicellular organisms Communicate by chemical messengers Animal and plant cells have cell junctions that directly connect
MCB*4010 Midterm Exam / Winter 2008
 MCB*4010 Midterm Exam / Winter 2008 Name: ID: Instructions: Answer all 4 questions. The number of marks for each question indicates how many points you need to provide. Write your answers in point form,
MCB*4010 Midterm Exam / Winter 2008 Name: ID: Instructions: Answer all 4 questions. The number of marks for each question indicates how many points you need to provide. Write your answers in point form,
Cell Communication. Cell Communication. Cell Communication. Cell Communication. Cell Communication. Chapter 9. Communication between cells requires:
 Chapter 9 Communication between cells requires: ligand: the signaling molecule receptor protein: the molecule to which the receptor binds -may be on the plasma membrane or within the cell 2 There are four
Chapter 9 Communication between cells requires: ligand: the signaling molecule receptor protein: the molecule to which the receptor binds -may be on the plasma membrane or within the cell 2 There are four
RAS Genes. The ras superfamily of genes encodes small GTP binding proteins that are responsible for the regulation of many cellular processes.
 ۱ RAS Genes The ras superfamily of genes encodes small GTP binding proteins that are responsible for the regulation of many cellular processes. Oncogenic ras genes in human cells include H ras, N ras,
۱ RAS Genes The ras superfamily of genes encodes small GTP binding proteins that are responsible for the regulation of many cellular processes. Oncogenic ras genes in human cells include H ras, N ras,
biochem480 [Autumn2014] Enzyme bio-informatics project
![biochem480 [Autumn2014] Enzyme bio-informatics project biochem480 [Autumn2014] Enzyme bio-informatics project](/thumbs/91/104514013.jpg) biochem480 [Autumn2014] Enzyme bio-informatics project Edit out any stuff in italics for the final version Student Name: IUBSystematic Name: 3-hydroxy-3-methylglutaryl-CoA reductase Other protein Names:
biochem480 [Autumn2014] Enzyme bio-informatics project Edit out any stuff in italics for the final version Student Name: IUBSystematic Name: 3-hydroxy-3-methylglutaryl-CoA reductase Other protein Names:
Comprehensive and Easy Course Notes for BIOL1040 Exams and Assessment
 Comprehensive and Easy Course Notes for BIOL1040 Exams and Assessment MODULE 1: PRINCIPLES OF CELL FUNCTION Membrane Structure & Function Cellular membranes are fluid mosaics of lipids and proteins Phospholipids
Comprehensive and Easy Course Notes for BIOL1040 Exams and Assessment MODULE 1: PRINCIPLES OF CELL FUNCTION Membrane Structure & Function Cellular membranes are fluid mosaics of lipids and proteins Phospholipids
Supplementary Material
 Supplementary Material Materials and methods Enzyme assay The enzymatic activity of -glucosidase toward salicin was measured with the Miller method (Miller, 1959) using glucose as the standard. A total
Supplementary Material Materials and methods Enzyme assay The enzymatic activity of -glucosidase toward salicin was measured with the Miller method (Miller, 1959) using glucose as the standard. A total
Biochemistry - I. Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture 1 Amino Acids I
 Biochemistry - I Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture 1 Amino Acids I Hello, welcome to the course Biochemistry 1 conducted by me Dr. S Dasgupta,
Biochemistry - I Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture 1 Amino Acids I Hello, welcome to the course Biochemistry 1 conducted by me Dr. S Dasgupta,
SUPPLEMENTARY INFORMATION. Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity
 SUPPLEMENTARY INFORMATION Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity Christopher Woods 1, Maturos Malaisree 1, Ben Long 2, Simon McIntosh-Smith
SUPPLEMENTARY INFORMATION Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity Christopher Woods 1, Maturos Malaisree 1, Ben Long 2, Simon McIntosh-Smith
P450 CYCLE. All P450s follow the same catalytic cycle of;
 P450 CYCLE All P450s follow the same catalytic cycle of; 1. Initial substrate binding 2. First electron reduction 3. Oxygen binding 4. Second electron transfer 5 and 6. Proton transfer/dioxygen cleavage
P450 CYCLE All P450s follow the same catalytic cycle of; 1. Initial substrate binding 2. First electron reduction 3. Oxygen binding 4. Second electron transfer 5 and 6. Proton transfer/dioxygen cleavage
Src-INACTIVE / Src-INACTIVE
 Biology 169 -- Exam 1 February 2003 Answer each question, noting carefully the instructions for each. Repeat- Read the instructions for each question before answering!!! Be as specific as possible in each
Biology 169 -- Exam 1 February 2003 Answer each question, noting carefully the instructions for each. Repeat- Read the instructions for each question before answering!!! Be as specific as possible in each
Copyright Mark Brandt, Ph.D. 46
 Examples of tein Structures tein types teins fall into three general classes, based on their overall three-dimensional structure and on their functional role: fibrous, membrane, and globular. Fibrous proteins
Examples of tein Structures tein types teins fall into three general classes, based on their overall three-dimensional structure and on their functional role: fibrous, membrane, and globular. Fibrous proteins
Molecular Graphics Perspective of Protein Structure and Function
 Molecular Graphics Perspective of Protein Structure and Function VMD Highlights > 20,000 registered Users Platforms: Unix (16 builds) Windows MacOS X Display of large biomolecules and simulation trajectories
Molecular Graphics Perspective of Protein Structure and Function VMD Highlights > 20,000 registered Users Platforms: Unix (16 builds) Windows MacOS X Display of large biomolecules and simulation trajectories
Signal Transduction Pathways. Part 2
 Signal Transduction Pathways Part 2 GPCRs G-protein coupled receptors > 700 GPCRs in humans Mediate responses to senses taste, smell, sight ~ 1000 GPCRs mediate sense of smell in mouse Half of all known
Signal Transduction Pathways Part 2 GPCRs G-protein coupled receptors > 700 GPCRs in humans Mediate responses to senses taste, smell, sight ~ 1000 GPCRs mediate sense of smell in mouse Half of all known
Review II: The Molecules of Life
 Review II: The Molecules of Life Judy Wieber BBSI @ Pitt 2007 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2007 Outline Introduction Proteins Carbohydrates Lipids
Review II: The Molecules of Life Judy Wieber BBSI @ Pitt 2007 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2007 Outline Introduction Proteins Carbohydrates Lipids
2013 W. H. Freeman and Company. 12 Signal Transduction
 2013 W. H. Freeman and Company 12 Signal Transduction CHAPTER 12 Signal Transduction Key topics: General features of signal transduction Structure and function of G protein coupled receptors Structure
2013 W. H. Freeman and Company 12 Signal Transduction CHAPTER 12 Signal Transduction Key topics: General features of signal transduction Structure and function of G protein coupled receptors Structure
Enzyme Catalytic Mechanisms. Dr. Kevin Ahern
 Enzyme Catalytic Mechanisms Dr. Kevin Ahern Cleave Peptide Bonds Specificity of Cutting Common Active Site Composition/Structure Mechanistically Well Studied Chymotrypsin Chymotrypsin Catalysis H2O Chymotrypsin
Enzyme Catalytic Mechanisms Dr. Kevin Ahern Cleave Peptide Bonds Specificity of Cutting Common Active Site Composition/Structure Mechanistically Well Studied Chymotrypsin Chymotrypsin Catalysis H2O Chymotrypsin
Eukaryotic transcription (III)
 Eukaryotic transcription (III) 1. Chromosome and chromatin structure Chromatin, chromatid, and chromosome chromatin Genomes exist as chromatins before or after cell division (interphase) but as chromatids
Eukaryotic transcription (III) 1. Chromosome and chromatin structure Chromatin, chromatid, and chromosome chromatin Genomes exist as chromatins before or after cell division (interphase) but as chromatids
Propagation of the Signal
 OpenStax-CNX module: m44452 1 Propagation of the Signal OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section,
OpenStax-CNX module: m44452 1 Propagation of the Signal OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section,
Review of Biochemistry
 Review of Biochemistry Chemical bond Functional Groups Amino Acid Protein Structure and Function Proteins are polymers of amino acids. Each amino acids in a protein contains a amino group, - NH 2,
Review of Biochemistry Chemical bond Functional Groups Amino Acid Protein Structure and Function Proteins are polymers of amino acids. Each amino acids in a protein contains a amino group, - NH 2,
Biol220 Cellular Signalling. Non-receptor tyrosine kinases
 Biol220 Cellular Signalling Non-receptor tyrosine kinases The 7TM receptors initiate signal transducton pathways through changes in tertiary structure that are induced by ligand binding. A fundamentally
Biol220 Cellular Signalling Non-receptor tyrosine kinases The 7TM receptors initiate signal transducton pathways through changes in tertiary structure that are induced by ligand binding. A fundamentally
FCC2 5CY7 FCC1 5CY8. Actinonin 5CVQ
 Table S1. Data collection and refinement statistics Data collection Apo 5E5D MA 5CPD MAS 5CP0 Actinonin 5CVQ FCC1 5CY8 FCC2 5CY7 FCC 5CWY FCC4 5CVK FCC5 5CWX FCC6 5CVP X-ray source 17A-KEK PAL4A PAL4A
Table S1. Data collection and refinement statistics Data collection Apo 5E5D MA 5CPD MAS 5CP0 Actinonin 5CVQ FCC1 5CY8 FCC2 5CY7 FCC 5CWY FCC4 5CVK FCC5 5CWX FCC6 5CVP X-ray source 17A-KEK PAL4A PAL4A
Function. BRAF (gene) From Wikipedia, the free encyclopedia
 BRAF (gene) From Wikipedia, the free encyclopedia BRAF is a human gene that makes a protein called B-Raf. The gene is also referred to as proto-oncogene B-Raf and v-raf murine sarcoma viral oncogene homolog
BRAF (gene) From Wikipedia, the free encyclopedia BRAF is a human gene that makes a protein called B-Raf. The gene is also referred to as proto-oncogene B-Raf and v-raf murine sarcoma viral oncogene homolog
SUPPORTING INFORMATION FOR. A Computational Approach to Enzyme Design: Using Docking and MM- GBSA Scoring
 SUPPRTING INFRMATIN FR A Computational Approach to Enzyme Design: Predicting ω- Aminotransferase Catalytic Activity Using Docking and MM- GBSA Scoring Sarah Sirin, 1 Rajesh Kumar, 2 Carlos Martinez, 2
SUPPRTING INFRMATIN FR A Computational Approach to Enzyme Design: Predicting ω- Aminotransferase Catalytic Activity Using Docking and MM- GBSA Scoring Sarah Sirin, 1 Rajesh Kumar, 2 Carlos Martinez, 2
Reading Assignments: Chapter 16: Cell Communication Pgs ; 545 & Figure 16-15; ; work Q-1, 3, 4, 10, 12, 15, 16, 17, 20 & 23
 Biol 205 Signal Transduction, the Social Contract and Rogue Cancer Cells Inside cancer web site http://www.insidecancer.org/ National Cancer Institute http://www.cancer.gov/cancerinfo/ Reading Assignments:
Biol 205 Signal Transduction, the Social Contract and Rogue Cancer Cells Inside cancer web site http://www.insidecancer.org/ National Cancer Institute http://www.cancer.gov/cancerinfo/ Reading Assignments:
Lecture: CHAPTER 13 Signal Transduction Pathways
 Lecture: 10 17 2016 CHAPTER 13 Signal Transduction Pathways Chapter 13 Outline Signal transduction cascades have many components in common: 1. Release of a primary message as a response to a physiological
Lecture: 10 17 2016 CHAPTER 13 Signal Transduction Pathways Chapter 13 Outline Signal transduction cascades have many components in common: 1. Release of a primary message as a response to a physiological
SUPPORTING INFORMATION. The Architecture of the TIR Domain Signalosome in the Toll-like Receptor-4. Signaling Pathway
 SUPPORTING INFORMATION The Architecture of the TIR Domain Signalosome in the Toll-like Receptor-4 Signaling Pathway Emine Guven-Maiorov 1, Ozlem Keskin 1, Attila Gursoy 2*, Carter VanWaes 3, Zhong Chen
SUPPORTING INFORMATION The Architecture of the TIR Domain Signalosome in the Toll-like Receptor-4 Signaling Pathway Emine Guven-Maiorov 1, Ozlem Keskin 1, Attila Gursoy 2*, Carter VanWaes 3, Zhong Chen
Structural analysis of fungus-derived FAD glucose dehydrogenase
 Structural analysis of fungus-derived FAD glucose dehydrogenase Hiromi Yoshida 1, Genki Sakai 2, Kazushige Mori 3, Katsuhiro Kojima 3, Shigehiro Kamitori 1, and Koji Sode 2,3,* 1 Life Science Research
Structural analysis of fungus-derived FAD glucose dehydrogenase Hiromi Yoshida 1, Genki Sakai 2, Kazushige Mori 3, Katsuhiro Kojima 3, Shigehiro Kamitori 1, and Koji Sode 2,3,* 1 Life Science Research
Chapter 20. Cell - Cell Signaling: Hormones and Receptors. Three general types of extracellular signaling. endocrine signaling. paracrine signaling
 Chapter 20 Cell - Cell Signaling: Hormones and Receptors Three general types of extracellular signaling endocrine signaling paracrine signaling autocrine signaling Endocrine Signaling - signaling molecules
Chapter 20 Cell - Cell Signaling: Hormones and Receptors Three general types of extracellular signaling endocrine signaling paracrine signaling autocrine signaling Endocrine Signaling - signaling molecules
The Tissue Engineer s Toolkit
 The Tissue Engineer s Toolkit Stimuli Detection and Response Ken Webb, Ph. D. Assistant Professor Dept. of Bioengineering Clemson University Environmental Stimulus-Cellular Response Environmental Stimuli
The Tissue Engineer s Toolkit Stimuli Detection and Response Ken Webb, Ph. D. Assistant Professor Dept. of Bioengineering Clemson University Environmental Stimulus-Cellular Response Environmental Stimuli
CYTOKINE RECEPTORS AND SIGNAL TRANSDUCTION
 CYTOKINE RECEPTORS AND SIGNAL TRANSDUCTION What is Cytokine? Secreted popypeptide (protein) involved in cell-to-cell signaling. Acts in paracrine or autocrine fashion through specific cellular receptors.
CYTOKINE RECEPTORS AND SIGNAL TRANSDUCTION What is Cytokine? Secreted popypeptide (protein) involved in cell-to-cell signaling. Acts in paracrine or autocrine fashion through specific cellular receptors.
NIH Public Access Author Manuscript J Am Chem Soc. Author manuscript; available in PMC 2008 September 29.
 NIH Public Access Author Manuscript Published in final edited form as: J Am Chem Soc. 2006 March 8; 128(9): 2812 2813. doi:10.1021/ja058211x. HIV-1 protease flaps spontaneously close to the correct structure
NIH Public Access Author Manuscript Published in final edited form as: J Am Chem Soc. 2006 March 8; 128(9): 2812 2813. doi:10.1021/ja058211x. HIV-1 protease flaps spontaneously close to the correct structure
Allosteric Inhibition of SHP2: Identification of a Potent, Selective, and Orally Efficacious Phosphatase Inhibitor!
 Allosteric Inhibition of SHP2: Identification of a Potent, Selective, and Orally Efficacious Phosphatase Inhibitor Allosteric pocket SHP2 Phosphatase ovel allosteric Phosphatase inhibitor Evan Carder Wipf
Allosteric Inhibition of SHP2: Identification of a Potent, Selective, and Orally Efficacious Phosphatase Inhibitor Allosteric pocket SHP2 Phosphatase ovel allosteric Phosphatase inhibitor Evan Carder Wipf
Chapter 3. Protein Structure and Function
 Chapter 3 Protein Structure and Function Broad functional classes So Proteins have structure and function... Fine! -Why do we care to know more???? Understanding functional architechture gives us POWER
Chapter 3 Protein Structure and Function Broad functional classes So Proteins have structure and function... Fine! -Why do we care to know more???? Understanding functional architechture gives us POWER
Cell Communication. Cell Communication. Communication between cells requires: ligand: the signaling molecule
 Cell Communication Cell Communication Communication between cells requires: ligand: the signaling molecule receptor protein: the molecule to which the ligand binds (may be on the plasma membrane or within
Cell Communication Cell Communication Communication between cells requires: ligand: the signaling molecule receptor protein: the molecule to which the ligand binds (may be on the plasma membrane or within
KEY CONCEPT QUESTIONS IN SIGNAL TRANSDUCTION
 Signal Transduction - Part 2 Key Concepts - Receptor tyrosine kinases control cell metabolism and proliferation Growth factor signaling through Ras Mutated cell signaling genes in cancer cells are called
Signal Transduction - Part 2 Key Concepts - Receptor tyrosine kinases control cell metabolism and proliferation Growth factor signaling through Ras Mutated cell signaling genes in cancer cells are called
Receptors. Dr. Sanaa Bardaweel
 Receptors Types and Theories Dr. Sanaa Bardaweel Some terms in receptor-drug interactions Agonists: drugs that mimic the natural messengers and activate receptors. Antagonist: drugs that block receptors.
Receptors Types and Theories Dr. Sanaa Bardaweel Some terms in receptor-drug interactions Agonists: drugs that mimic the natural messengers and activate receptors. Antagonist: drugs that block receptors.
ab E3 Ligase Auto- Ubiquitilylation Assay Kit
 ab139469 E3 Ligase Auto- Ubiquitilylation Assay Kit Instructions for Use For testing ubiquitin E3 ligase activity through assessment of their ability to undergo auto-ubiquitinylation This product is for
ab139469 E3 Ligase Auto- Ubiquitilylation Assay Kit Instructions for Use For testing ubiquitin E3 ligase activity through assessment of their ability to undergo auto-ubiquitinylation This product is for
Regulating Transmembrane Signaling Through Plexin- Neuropilin Oligomerization
 Regulating Transmembrane Signaling Through Plexin- Neuropilin Oligomerization Bryan W. Berger Department of Chemical Engineering Program in Bioengineering Components of Cell Membranes Approximately 30-40%
Regulating Transmembrane Signaling Through Plexin- Neuropilin Oligomerization Bryan W. Berger Department of Chemical Engineering Program in Bioengineering Components of Cell Membranes Approximately 30-40%
