W.M. Keck Foundation Biotechnology Resource Laboratories. Erol Gulcicek Christopher Colangelo Terence Wu
|
|
|
- May Powell
- 5 years ago
- Views:
Transcription
1 W.M. Keck Foundation Biotechnology Resource Laboratories Erol Gulcicek Christopher Colangelo Terence Wu
2 - Site is currently up with basic information - Overview of profiling technologies has recently been added since the url was first distributed - Ideas for future features are very welcome
3 Yale/NIDA Neuroproteomics Center Fact Sheet Title of Center Grant: Theme of Center: Principal Investigator: Co-Principal Investigator: #Grants Awarded: Total Funding Awarded to Yale: Yale/NIDA Neuroproteomics Research Center "Proteomics of Altered Signaling in Addiction" Kenneth R. Williams, Ph.D. Director, W.M. Keck Foundation Biotechnology Resource Laboratory Professor (Adj,) Research Dept. of Molecular Biophysics & Biochemistry Yale University School of Medicine Angus C. Nairn, Ph.D. Professor, Department of Psychiatry Yale University School of Medicine 2 Nationally $7.7 million Total Budget Period: 9/23/2004 5/31/2009 Center Cores: Center Investigators: Administrative, Bioinformatics and Biostatistics, Protein Identification, Protein Microarray; and Protein and Lipid Separation and Profiling 14 from Yale University, The Veterans Administration Medical Center (West Haven) Rockefeller University
4 Overall Goal: To substantially increase understanding of the biochemical mechanisms that underlie substance abuse and its treatment. Specific Aims: Bring together Yale faculty working at the forefront of such key neuroscience areas as Signal transduction, Plasticity, Neuronal morphogenesis, Lipid metabolism in neuronal signaling and Synaptic function Synaptic response to psychotropic drugs with experts in Proteomics, Biostatistics, Bioinformatics.
5 Proteomics of Altered Signaling in Addiction Receptor-Mediated Signaling Transcriptional Regulation Endocytosis and Exocytosis Regulation of the Cytoskeleton DIGE Neuronal Development Phosphoproteome Profiling ICAT Neuronal Plasticity 2D-LC Beckman Coulter System Cell Death Lipid Profiling MudPIT Translational Research Identification and Characterization of Signaling Proteins Whose Expression is Altered by Drug Addiction Serum Biomarkers Targeted Proteomics Clinical Studies of Drug Addiction
6 Yale/NIDA Neuroproteomics Program Initial Stage: - Meet individual PI s to: - Better understand the PI s research area - Jointly determine best to implement proteomics technologies available - Timelines from the Cores. - Review of current technologies - Development of new technologies - Some already know Core personnel and have already initiated studies in which case please continue working with these staff and additionally, I would be very glad to talk with you also if you have questions about other technologies. Erol E. Gulcicek erol.gulcicek@yale.edu
7 Commonly Used Protein Profiling Workflows in Mass Spectrometry Primary Biological Samples Organ, Tissue, cell extracts, biological fluids, etc. Proteins Peptides LC, MS and Tandem-MS Protein Sample Preparation Sub cellular components, fractionation, affinity enrichment, Protein complexes, etc. Protein Labeling Chemistries ICAT, PhIAT, SILAC, Protein Sample Simplification (or Purification): 18 O proteolysis, etc. 1D-SDS 2D-SDS Protein LC DIGE IEF PAGE PAGE (Q, S, RP, SEX, etc. FFE Digestion Digestion Digestion Digestion Enrichment step and/or Peptide Labeling Chemistries (SCX, IMAC, itraq TM etc.) (Non LC) MS/MS Top Down MS or MudPit MS/MS RP LC MS/MS Peptide Mass Fingerprint Protein MS/MS HT Protein ID & Protein sequence ID, (PMF) sequence ID, Protein Sequencing, Quantitation PTM and Quantitation Protein ID PTM and PTM SELDI or Predominantly Predominantly MALDI Predominantly Predominantly ESI QTOF& IT ESI QTOF & IT (3D and 2D), MALDI TOF/TOF MS. TOF MS MALDI TOF MS ESI FTMS (3D and 2D) Also FTMS and Triple-Q (incl. Quad. Trap) Biomarker patterns
8 Protein Profiling Overview Christopher M. Colangelo, PhD Director of Protein Profiling 300 George Street Room G006 (203) (W)
9 Brief Overview of the Keck Laboratory Founded in 1980: 40 Full time staff 100 Major instrument systems purchased at a cost of >$11 million dollars 25,000 ft 2 of space 12 individual Resources that provide a wide range of proteomic and genomic biotechnologies Completes 270,000 protein and nucleic acid syntheses and analyses annually for 260 Yale and 640 non-yale investigators at 300 institutions & companies in 27 countries NHLBI Proteomics Center (2002) NIH High End Instrumentation Grant (2002): FTICR-MS National Biodefense Center of Excellence: Proteomics Core (2003) NIH High End Instrumentation Grant: Center for High Performance Computation In Biomedicine (2004) NIDA Neuroproteomics Center (2004) (
10 Disease Based Proteomics Receptor-Mediated Signaling DIGE Phosphoproteome Profiling Transcriptional Regulation ICAT Endocytosis and Exocytosis 2D-LC Beckman Coulter System Lipid Profiling Regulation of the Cytoskeleton MudPIT Translational Research Identification and Characterization of Signaling Proteins Whose Expression is Up/Down Regulated Serum Biomarkers Targeted Proteomics Clinical Studies of Biomarker Efficacy
11 Keck MALDI-MS Based Disease Biomarker Discovery Sera from >50 normal patients Sera from >50 disease patients Automatically desalt each sample (submitted in 96 well plate format) using a Micromass MassPrep robot and C-18 Zip-Tips. Automated MALDI-MS on Micromass MALDI-L/R instrument: 130,000 m/z vs intensity datapoints acquired from 700 to 28,000 Da. Linearly normalize baseline-corrected intensities in each spectrum to the overall median for all normal + disease spectra in the training set. Filter out noise and all datapoints which fail to pass peak and unique peptde ion criteria. Use a Random Forest Algorithm customized by Keck Biostatistics Resource (Wu et, al 2003) to identify peptide markers whose intensities can be used to maximally discriminate all normal from disease samples. Optimize marker selection by increasing the size of the training set until the ability to correctly classify a testing set is maximized
12 Median MALDI-MS Spectra for Sera from 47 Ovarian Cancer (top) vs 42 Control (middle) Patients and the Resulting Difference Spectrum (bottom)
13 m/z Versus Intensity Distribution for 129 Samples Around the 2084 Ovarian Cancer Marker (relative importance = 2.9) Marker Position Median Cancer Median Control
14 Impact of Training Set Size & Number of Biomarkers on Classification of 34 "Unknown" Ovarian Cancer Sera #Biomarkers = 25 Training Set Size: 102 Sera 80% 80% Correctly Classified Test Sera 75% 70% 65% 60% 55% %Correctly Classified Test Sera 75% 70% 65% 50% % #Samples in Training Set Number of Biomarkers
15 Genomics The blueprint of life DNA Central Dogma PCR Mullis (1985) Human Genome Sequence Watson and Crick (1953) Nature 171, Crick (1970) Nature 227, Crick (1958) Symp. Soc. Exp. Biol. 138 The International Human Genome Mapping Consortium (2001) Nature 409, Venter et. al. (2001) Science Feb :
16 What is proteomics? Genomics provides us with the words, but doesn t provide us with any meaning Proteomics - provides us with the definitions for the genome The identification, characterization and quantification of all proteins involved in a particular pathway, organelle, cell, tissue, organ or organism that can be studied in concert to provide accurate and comprehensive data about that system. Protein Profiling provide us with the grammar and syntax to form a meaningful language Understanding how proteins interact with each other, their environment, and function with outside molecules Challenges - proteomics and protein profiling are dynamic languages
17 How do we bridge the gap from genomics to proteomics? Genomics Proteomics
18 Gene Microarrays First step towards bridging the gap between human genome and function Keck Human 4.6K cdna Array Cluster Analysis
19 mrna vs. Protein correlation (R=0.66) Comparing protein abundance and mrna expression levels on a genomic scale Greenbaum D, Colangelo C, Williams K, Gerstein M Genome Biol. 2003; 4(9): 117. Epub 2003 Aug 29.
20 Reality check for proteomics Avogadro s number - 6 x molecules/mole For MS proteomics we want 100 fmoles (10-14 moles) = 6.02 x molecules cell copy number cells needed for 100 fmoles 10 6 billion (6.02 x 10 9 ) million (6.02 x 10 8 ) million (6.02 x 10 7 ) 10,000 6 million (6.02 x 10 6 ) 100, million (6.02 x 10 5 ) 10cm culture dish = 10 5 cells 100,000 copy number limit 1g of tissue = 10 8 cells 1000 copy number limit
21 Levels of proteins in plasma Molecular & Cellular Proteomics 2003, Anderson and Anderson 2 (1): 50
22 Protein Profiling
23 Disease Based Proteomics Receptor-Mediated Signaling DIGE Phosphoproteome Profiling Transcriptional Regulation ICAT Endocytosis and Exocytosis 2D-LC Beckman Coulter System Lipid Profiling Regulation of the Cytoskeleton MudPIT Translational Research Identification and Characterization of Signaling Proteins Whose Expression is Up/Down Regulated Serum Biomarkers Targeted Proteomics Clinical Studies of Biomarker Efficacy
24 Cleavable ICAT Reagents Allows selection and concentration of cysteinecontaining peptides Removes biotin which improves MS/MS fragmentation Provide 9 Da which provides ratio of heavy/light peptide conc. Reacts with free cysteines Applied Biosystems
25 Differential expression analysis on human leukemic precursor cells vs. differentiated monocytes/neutrophils 16 SCX fractions collected 100ug of protein is labeled with acid cleavable ICAT TM reagent/trypsinized Cys-containing peptides affinity purified/ cleaved with TFA Online LC-MS/MS analysis on QStar XL System
26 ICAT-Determined Expression Ratio of Zinc Finger Protein 9 in Two Human Cell Lines Overall Average Expression Ratio of Protein 9 in Two Human Cell Lines: 3.4 fold decrease (average H/L = )
27 Web-based query results from Yale Protein Expression Database (YPED)
28 Disease Based Proteomics Receptor-Mediated Signaling DIGE Phosphoproteome Profiling Transcriptional Regulation ICAT Endocytosis and Exocytosis 2D-LC Beckman Coulter System Lipid Profiling Regulation of the Cytoskeleton MudPIT Translational Research Identification and Characterization of Signaling Proteins Whose Expression is Up/Down Regulated Serum Biomarkers Targeted Proteomics Clinical Studies of Biomarker Efficacy
29 Automated 2D Comparative LC Protein Profiling (Beckman-Coulter PF2D Platform) Partial pi/uv map of a colon cancer cell line before and after drug treatment. The RP-HPLC profiles illustrate differences in the pi fraction which are shown also by the color coded band depictions of these RP-HPLC profiles (from Beckman-Coulter).
30 Two-dimensional Comparison Plot of NB4 (Green) and NB4 + RA (Red) (Neutrophil) from Beckman PF2D System 5 mg of total protein Retention Time ph The 2D overlay plot shows the relative differential expression for each of the 21 ph fractions based on overlay comparisons of 214 nm reversed phase chromatograms Fraction
31 Overlay of NB4 and NB4+RA Reversed Phase Chromatograms from Fraction 7 The differential peaks in the UV chromatogram represent intact proteins that are expressed or modified between control and treated samples. Absorbance (211 nm) Retention Time
32 APEX FTMS APEX-Q Industrial Design Life Science Tools is our Business
33 Typical Project Milestones Weeks DIGE AAA results Differential Gel Preparative gel Mass Spectrometry Data Analysis ICAT AAA results ICAT Labeling Cationexchange Mass Spectrometry Data Analysis PF2D AAA results 2D Chromatography ICAT Labeling Mass Spectrometry Data Analysis
34 Disease Based Proteomics Receptor-Mediated Signaling DIGE Phosphoproteome Profiling Transcriptional Regulation ICAT Endocytosis and Exocytosis 2D-LC Beckman Coulter System Lipid Profiling Regulation of the Cytoskeleton MudPIT Translational Research Identification and Characterization of Signaling Proteins Whose Expression is Up/Down Regulated Serum Biomarkers Targeted Proteomics Clinical Studies of Biomarker Efficacy
35 2D Electrophoresis 2D gel electrophoresis a Gateway proteomic tool for the separation of complex protein mixtures from many different biological sample types Pros: -High resolution separation by pi and MW -Possible to resolve several thousand proteins on a single gel -IPG strips feature high reproducibility and resolution (single ph unit) -PTMs, such as phosphorylation and glycosylation are clearly visualized Cons: - Membrane/ hydrophobic proteins have solubility problems a) can be addressed by detergents, e.g., ASB-14, Triton X-100 b) some proteins are prone to precipitation at their IE point c) try an alternative technology, such as ICAT -Gel-to-gel reproducibility and therefore Quantitation is a problem
36 DIFFERENTIAL FLUORESCENCE GEL ELECTROPHORESIS PROTEOME PROFILING (DIGE)
37 DIGE Detection Limits # of cells protein copies/cell total # of proteins moles of protein ng of 50kd 1 x pmole 8,304 (~1 g of tissue) pmole 830 MS DIGE pmole fmole fmole fmol 0.08
38 DIGE method is highly reproducible but only with good sample preparation! -use of label compatible buffers 7M Urea, 2M Thiourea, 4% CHAPS, in 30 mm TRIS adjusted to ph 8.5 -ensure that the sample preparation is homogenous - clean -up of sample via precipitation, dialysis, nuclease -low abundance issues are best addressed at the sample preparation step a)subcellular fractionation b)directed biochemical enrichments/pre-fractionation c)microdissection/punches of specific tissue regions d)removal of high abundance species -quantitate samples prior to analysis to normalize, AAA is best -ensure there is enough sample for downstream studies!
39 Albumin and IgG depletion from serum
40 cy 3 image
41 cy 5 image
42 spots with a greater or = to 3X increase (Blue) or Decrease (Red) In cy5/cy3 spot volume ratio
43
Introduction to Proteomics 1.0
 Introduction to Proteomics 1.0 CMSP Workshop Pratik Jagtap Managing Director, CMSP Objectives Why are we here? For participants: Learn basics of MS-based proteomics Learn what s necessary for success using
Introduction to Proteomics 1.0 CMSP Workshop Pratik Jagtap Managing Director, CMSP Objectives Why are we here? For participants: Learn basics of MS-based proteomics Learn what s necessary for success using
PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System
 Application Note LC/MS PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System Purpose This application note describes an automated workflow
Application Note LC/MS PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System Purpose This application note describes an automated workflow
Mass Spectrometry. Mass spectrometer MALDI-TOF ESI/MS/MS. Basic components. Ionization source Mass analyzer Detector
 Mass Spectrometry MALDI-TOF ESI/MS/MS Mass spectrometer Basic components Ionization source Mass analyzer Detector 1 Principles of Mass Spectrometry Proteins are separated by mass to charge ratio (limit
Mass Spectrometry MALDI-TOF ESI/MS/MS Mass spectrometer Basic components Ionization source Mass analyzer Detector 1 Principles of Mass Spectrometry Proteins are separated by mass to charge ratio (limit
Proteomics/Peptidomics
 Proteomics/Peptidomics System biology tools and preclinical models for translational research in endometriosis, ESHRE Campus workshop, 4-5 September 2009 E. Waelkens Proteomics: What? Proteins Proteomics
Proteomics/Peptidomics System biology tools and preclinical models for translational research in endometriosis, ESHRE Campus workshop, 4-5 September 2009 E. Waelkens Proteomics: What? Proteins Proteomics
One Gene, Many Proteins. Applications of Mass Spectrometry to Proteomics. Why Proteomics? Raghothama Chaerkady, Ph.D.
 Applications of Mass Spectrometry to Proteomics Raghothama Chaerkady, Ph.D. McKusick-Nathans Institute of Genetic Medicine and the Department of Biological Chemistry Why Proteomics? One Gene, Many Proteins
Applications of Mass Spectrometry to Proteomics Raghothama Chaerkady, Ph.D. McKusick-Nathans Institute of Genetic Medicine and the Department of Biological Chemistry Why Proteomics? One Gene, Many Proteins
Improve Protein Analysis with the New, Mass Spectrometry- Compatible ProteasMAX Surfactant
 Improve Protein Analysis with the New, Mass Spectrometry- Compatible Surfactant ABSTRACT Incomplete solubilization and digestion and poor peptide recovery are frequent limitations in protein sample preparation
Improve Protein Analysis with the New, Mass Spectrometry- Compatible Surfactant ABSTRACT Incomplete solubilization and digestion and poor peptide recovery are frequent limitations in protein sample preparation
Lecture 3. Tandem MS & Protein Sequencing
 Lecture 3 Tandem MS & Protein Sequencing Nancy Allbritton, M.D., Ph.D. Department of Physiology & Biophysics 824-9137 (office) nlallbri@uci.edu Office- Rm D349 Medical Science D Bldg. Tandem MS Steps:
Lecture 3 Tandem MS & Protein Sequencing Nancy Allbritton, M.D., Ph.D. Department of Physiology & Biophysics 824-9137 (office) nlallbri@uci.edu Office- Rm D349 Medical Science D Bldg. Tandem MS Steps:
SUNY UPSTATE MEDICAL UNIVERSITY PROTEOMICS CORE
 SUNY UPSTATE MEDICAL UNIVERSITY PROTEOMICS CORE Location: SUNY Upstate Medical University Weiskotten Hall Addition, Room 4303 750 East Adams Street Syracuse, NY 13210 Contact Information: David Kakhniashvili,
SUNY UPSTATE MEDICAL UNIVERSITY PROTEOMICS CORE Location: SUNY Upstate Medical University Weiskotten Hall Addition, Room 4303 750 East Adams Street Syracuse, NY 13210 Contact Information: David Kakhniashvili,
Protein Post Translational Modification (PTM) & Profiling Core Erol E. Gulcicek
 Protein Post Translational Modification (PTM) & Profiling Core Erol E. Gulcicek Determine sites of Post Translational Modifications and changes in their Levels - Phosphorylation*** - Ubiquitination - Palmitoylation
Protein Post Translational Modification (PTM) & Profiling Core Erol E. Gulcicek Determine sites of Post Translational Modifications and changes in their Levels - Phosphorylation*** - Ubiquitination - Palmitoylation
Learning Objectives. Overview of topics to be discussed 10/25/2013 HIGH RESOLUTION MASS SPECTROMETRY (HRMS) IN DISCOVERY PROTEOMICS
 HIGH RESOLUTION MASS SPECTROMETRY (HRMS) IN DISCOVERY PROTEOMICS A clinical proteomics perspective Michael L. Merchant, PhD School of Medicine, University of Louisville Louisville, KY Learning Objectives
HIGH RESOLUTION MASS SPECTROMETRY (HRMS) IN DISCOVERY PROTEOMICS A clinical proteomics perspective Michael L. Merchant, PhD School of Medicine, University of Louisville Louisville, KY Learning Objectives
Shotgun Proteomics MS/MS. Protein Mixture. proteolysis. Peptide Mixture. Time. Abundance. Abundance. m/z. Abundance. m/z 2. Abundance.
 Abundance Abundance Abundance Abundance Abundance Shotgun Proteomics Protein Mixture 1 2 3 MS/MS proteolysis m/z 2 3 Time µlc m/z MS 1 m/z Peptide Mixture m/z Block Diagram of a Mass Spectrometer Sample
Abundance Abundance Abundance Abundance Abundance Shotgun Proteomics Protein Mixture 1 2 3 MS/MS proteolysis m/z 2 3 Time µlc m/z MS 1 m/z Peptide Mixture m/z Block Diagram of a Mass Spectrometer Sample
New Instruments and Services
 New Instruments and Services Liwen Zhang Mass Spectrometry and Proteomics Facility The Ohio State University Summer Workshop 2016 Thermo Orbitrap Fusion http://planetorbitrap.com/orbitrap fusion Thermo
New Instruments and Services Liwen Zhang Mass Spectrometry and Proteomics Facility The Ohio State University Summer Workshop 2016 Thermo Orbitrap Fusion http://planetorbitrap.com/orbitrap fusion Thermo
Double charge of 33kD peak A1 A2 B1 B2 M2+ M/z. ABRF Proteomics Research Group - Qualitative Proteomics Study Identifier Number 14146
 Abstract The 2008 ABRF Proteomics Research Group Study offers participants the chance to participate in an anonymous study to identify qualitative differences between two protein preparations. We used
Abstract The 2008 ABRF Proteomics Research Group Study offers participants the chance to participate in an anonymous study to identify qualitative differences between two protein preparations. We used
MS/MS to Targeted Proteomics (MRM)
 MS/MS to Targeted Proteomics (MRM) How it worked on the Human Lens Proteome Jayson Falkner PhD jay@singleorganism.com Genes Show Limited Value in Predicting Diseases With only a few exceptions, what the
MS/MS to Targeted Proteomics (MRM) How it worked on the Human Lens Proteome Jayson Falkner PhD jay@singleorganism.com Genes Show Limited Value in Predicting Diseases With only a few exceptions, what the
LC/MS/MS SOLUTIONS FOR LIPIDOMICS. Biomarker and Omics Solutions FOR DISCOVERY AND TARGETED LIPIDOMICS
 LC/MS/MS SOLUTIONS FOR LIPIDOMICS Biomarker and Omics Solutions FOR DISCOVERY AND TARGETED LIPIDOMICS Lipids play a key role in many biological processes, such as the formation of cell membranes and signaling
LC/MS/MS SOLUTIONS FOR LIPIDOMICS Biomarker and Omics Solutions FOR DISCOVERY AND TARGETED LIPIDOMICS Lipids play a key role in many biological processes, such as the formation of cell membranes and signaling
Mass Spectrometry. - Introduction - Ion sources & sample introduction - Mass analyzers - Basics of biomolecule MS - Applications
 - Introduction - Ion sources & sample introduction - Mass analyzers - Basics of biomolecule MS - Applications Adapted from Mass Spectrometry in Biotechnology Gary Siuzdak,, Academic Press 1996 1 Introduction
- Introduction - Ion sources & sample introduction - Mass analyzers - Basics of biomolecule MS - Applications Adapted from Mass Spectrometry in Biotechnology Gary Siuzdak,, Academic Press 1996 1 Introduction
Advances in Hybrid Mass Spectrometry
 The world leader in serving science Advances in Hybrid Mass Spectrometry ESAC 2008 Claire Dauly Field Marketing Specialist, Proteomics New hybrids instruments LTQ Orbitrap XL with ETD MALDI LTQ Orbitrap
The world leader in serving science Advances in Hybrid Mass Spectrometry ESAC 2008 Claire Dauly Field Marketing Specialist, Proteomics New hybrids instruments LTQ Orbitrap XL with ETD MALDI LTQ Orbitrap
New Instruments and Services
 New Instruments and Services http://planetorbitrap.com/orbitrap fusion Combining the best of quadrupole, Orbitrap, and ion trap mass analysis in a revolutionary Tribrid architecture, the Orbitrap Fusion
New Instruments and Services http://planetorbitrap.com/orbitrap fusion Combining the best of quadrupole, Orbitrap, and ion trap mass analysis in a revolutionary Tribrid architecture, the Orbitrap Fusion
Mass Spectrometry and Proteomics - Lecture 4 - Matthias Trost Newcastle University
 Mass Spectrometry and Proteomics - Lecture 4 - Matthias Trost Newcastle University matthias.trost@ncl.ac.uk previously Peptide fragmentation Hybrid instruments 117 The Building Blocks of Life DNA RNA Proteins
Mass Spectrometry and Proteomics - Lecture 4 - Matthias Trost Newcastle University matthias.trost@ncl.ac.uk previously Peptide fragmentation Hybrid instruments 117 The Building Blocks of Life DNA RNA Proteins
2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry
 Dr. Sanjeeva Srivastava 1. Fundamental of Mass Spectrometry Role of MS and basic concepts 2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry 2 1 MS basic concepts Mass spectrometry - technique
Dr. Sanjeeva Srivastava 1. Fundamental of Mass Spectrometry Role of MS and basic concepts 2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry 2 1 MS basic concepts Mass spectrometry - technique
Glycosylation analysis of blood plasma proteins
 Glycosylation analysis of blood plasma proteins Thesis booklet Eszter Tóth Doctoral School of Pharmaceutical Sciences Semmelweis University Supervisor: Károly Vékey DSc Official reviewers: Borbála Dalmadiné
Glycosylation analysis of blood plasma proteins Thesis booklet Eszter Tóth Doctoral School of Pharmaceutical Sciences Semmelweis University Supervisor: Károly Vékey DSc Official reviewers: Borbála Dalmadiné
1. Sample Introduction to MS Systems:
 MS Overview: 9.10.08 1. Sample Introduction to MS Systems:...2 1.1. Chromatography Interfaces:...3 1.2. Electron impact: Used mainly in Protein MS hard ionization source...4 1.3. Electrospray Ioniztion:
MS Overview: 9.10.08 1. Sample Introduction to MS Systems:...2 1.1. Chromatography Interfaces:...3 1.2. Electron impact: Used mainly in Protein MS hard ionization source...4 1.3. Electrospray Ioniztion:
SCS Mass Spectrometry Laboratory
 SCS Mass Spectrometry Laboratory Contact Information Staff 31 Noyes Laboratory (8:00-5:00 M-F) 217-333-2545 http://scs.illinois.edu/massspec/ Furong Sun (frs@illinois.edu) Furong Sun Director Training
SCS Mass Spectrometry Laboratory Contact Information Staff 31 Noyes Laboratory (8:00-5:00 M-F) 217-333-2545 http://scs.illinois.edu/massspec/ Furong Sun (frs@illinois.edu) Furong Sun Director Training
Comparison of mass spectrometers performances
 Comparison of mass spectrometers performances Instrument Mass Mass Sensitivity resolution accuracy Quadrupole 1 x 10 3 0.1 Da* 0.5-1.0 pmol DE-MALDI 2 x 10 4 20 ppm 1-10 fmol peptide 1-5 pmol protein Ion
Comparison of mass spectrometers performances Instrument Mass Mass Sensitivity resolution accuracy Quadrupole 1 x 10 3 0.1 Da* 0.5-1.0 pmol DE-MALDI 2 x 10 4 20 ppm 1-10 fmol peptide 1-5 pmol protein Ion
Characterization of Disulfide Linkages in Proteins by 193 nm Ultraviolet Photodissociation (UVPD) Mass Spectrometry. Supporting Information
 Characterization of Disulfide Linkages in Proteins by 193 nm Ultraviolet Photodissociation (UVPD) Mass Spectrometry M. Montana Quick, Christopher M. Crittenden, Jake A. Rosenberg, and Jennifer S. Brodbelt
Characterization of Disulfide Linkages in Proteins by 193 nm Ultraviolet Photodissociation (UVPD) Mass Spectrometry M. Montana Quick, Christopher M. Crittenden, Jake A. Rosenberg, and Jennifer S. Brodbelt
Multiplex Protein Quantitation using itraq Reagents in a Gel-Based Workflow
 Multiplex Protein Quantitation using itraq Reagents in a Gel-Based Workflow Purpose Described herein is a workflow that combines the isobaric tagging reagents, itraq Reagents, with the separation power
Multiplex Protein Quantitation using itraq Reagents in a Gel-Based Workflow Purpose Described herein is a workflow that combines the isobaric tagging reagents, itraq Reagents, with the separation power
The Detection of Allergens in Food Products with LC-MS
 The Detection of Allergens in Food Products with LC-MS Something for the future? Jacqueline van der Wielen Scope of Organisation Dutch Food and Consumer Product Safety Authority: Law enforcement Control
The Detection of Allergens in Food Products with LC-MS Something for the future? Jacqueline van der Wielen Scope of Organisation Dutch Food and Consumer Product Safety Authority: Law enforcement Control
Technical Note # TN-31 Redefining MALDI-TOF/TOF Performance
 Bruker Daltonics Technical Note # TN-31 Redefining MALDI-TOF/TOF Performance The new ultraflextreme exceeds all current expectations of MALDI-TOF/TOF technology: A proprietary khz smartbeam-ii TM MALDI
Bruker Daltonics Technical Note # TN-31 Redefining MALDI-TOF/TOF Performance The new ultraflextreme exceeds all current expectations of MALDI-TOF/TOF technology: A proprietary khz smartbeam-ii TM MALDI
PosterREPRINT A NOVEL APPROACH TO MALDI-TOF-MS SAMPLE PREPARATION. Presented at ABRF 2002, Austin, Texas, USA, 9th - 12th March 2002.
 Introduction A NOVEL APPROACH TO MALDI-TOF-MS SAMPLE PREPARATION Ed Bouvier 2, Jeff Brown 1, Emmanuelle Claude 1, John L. Gebler 2, Weibin Chen 2, *Dominic Gostick 1, Kevin Howes 1, James Langridge 1,
Introduction A NOVEL APPROACH TO MALDI-TOF-MS SAMPLE PREPARATION Ed Bouvier 2, Jeff Brown 1, Emmanuelle Claude 1, John L. Gebler 2, Weibin Chen 2, *Dominic Gostick 1, Kevin Howes 1, James Langridge 1,
Quantification with Proteome Discoverer. Bernard Delanghe
 Quantification with Proteome Discoverer Bernard Delanghe Overview: Which approach to use? Proteome Discoverer Quantification Method What When to use Metabolic labeling SILAC Cell culture systems Small
Quantification with Proteome Discoverer Bernard Delanghe Overview: Which approach to use? Proteome Discoverer Quantification Method What When to use Metabolic labeling SILAC Cell culture systems Small
Welcome! Mass Spectrometry meets Cheminformatics WCMC Metabolomics Course 2014 Tobias Kind. Course: Search of MS/MS files with the NIST MS Search GUI
 Biology Informatics Chemistry Welcome! Mass Spectrometry meets Cheminformatics WCMC Metabolomics Course 2014 Tobias Kind Course: Search of MS/MS files with the NIST MS Search GUI http://fiehnlab.ucdavis.edu/staff/kind
Biology Informatics Chemistry Welcome! Mass Spectrometry meets Cheminformatics WCMC Metabolomics Course 2014 Tobias Kind Course: Search of MS/MS files with the NIST MS Search GUI http://fiehnlab.ucdavis.edu/staff/kind
DetergentOUT GBS10 Spin Plates
 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name DetergentOUT GBS10 Spin Plates 96-Well Plates for the Removal of Detergents from Peptide
G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name DetergentOUT GBS10 Spin Plates 96-Well Plates for the Removal of Detergents from Peptide
Chapter 10apter 9. Chapter 10. Summary
 Chapter 10apter 9 Chapter 10 The field of proteomics has developed rapidly in recent years. The essence of proteomics is to characterize the behavior of a group of proteins, the system rather than the
Chapter 10apter 9 Chapter 10 The field of proteomics has developed rapidly in recent years. The essence of proteomics is to characterize the behavior of a group of proteins, the system rather than the
Metabolomic and Proteomics Solutions for Integrated Biology. Christine Miller Omics Market Manager ASMS 2015
 Metabolomic and Proteomics Solutions for Integrated Biology Christine Miller Omics Market Manager ASMS 2015 Integrating Biological Analysis Using Pathways Protein A R HO R Protein B Protein X Identifies
Metabolomic and Proteomics Solutions for Integrated Biology Christine Miller Omics Market Manager ASMS 2015 Integrating Biological Analysis Using Pathways Protein A R HO R Protein B Protein X Identifies
Stress-Related Biomarkers of Relapse Vulnerability
 Stress-Related Biomarkers of Relapse Vulnerability in Addictive Disorders. December 3, 2008 Robert D. Beech, MD, PhD, Assistant Professor Rajita Sinha, PhD, Professor Yale University Department of Psychiatry
Stress-Related Biomarkers of Relapse Vulnerability in Addictive Disorders. December 3, 2008 Robert D. Beech, MD, PhD, Assistant Professor Rajita Sinha, PhD, Professor Yale University Department of Psychiatry
Applications of HPLC-MALDI-TOF MS/MS Phosphoproteomic Analysis in Oncological Clinical Diagnostics
 Current Proteomics, 2011, 8, 153-167 153 Applications of HPLC-MALDI-TOF MS/MS Phosphoproteomic Analysis in Oncological Clinical Diagnostics Courtney L. Haddock*,1, Barry Holtz 1, Neil Senzer 1,2,3,4 and
Current Proteomics, 2011, 8, 153-167 153 Applications of HPLC-MALDI-TOF MS/MS Phosphoproteomic Analysis in Oncological Clinical Diagnostics Courtney L. Haddock*,1, Barry Holtz 1, Neil Senzer 1,2,3,4 and
MASS SPECTROMETRY BASED METABOLOMICS. Pavel Aronov. ABRF2010 Metabolomics Research Group March 21, 2010
 MASS SPECTROMETRY BASED METABOLOMICS Pavel Aronov ABRF2010 Metabolomics Research Group March 21, 2010 Types of Experiments in Metabolomics targeted non targeted Number of analyzed metabolites is limited
MASS SPECTROMETRY BASED METABOLOMICS Pavel Aronov ABRF2010 Metabolomics Research Group March 21, 2010 Types of Experiments in Metabolomics targeted non targeted Number of analyzed metabolites is limited
MALDI-TOF. Introduction. Schematic and Theory of MALDI
 MALDI-TOF Proteins and peptides have been characterized by high pressure liquid chromatography (HPLC) or SDS PAGE by generating peptide maps. These peptide maps have been used as fingerprints of protein
MALDI-TOF Proteins and peptides have been characterized by high pressure liquid chromatography (HPLC) or SDS PAGE by generating peptide maps. These peptide maps have been used as fingerprints of protein
5 Identification of Binding Partners of the Annexin A2 / P11 Complex by Chemical Cross-Linking
 5 Identification of Binding Partners of the Annexin A2 / P11 Complex by Chemical Cross-Linking In the quest of the omics sciences for holistic schemes, the identification of binding partners of proteins
5 Identification of Binding Partners of the Annexin A2 / P11 Complex by Chemical Cross-Linking In the quest of the omics sciences for holistic schemes, the identification of binding partners of proteins
REDOX PROTEOMICS. Roman Zubarev.
 REDOX PROTEOMICS Roman Zubarev Roman.Zubarev@ki.se Physiological Chemistry I, Department for Medical Biochemistry & Biophysics, Karolinska Institutet, Stockholm What is (RedOx) Proteomics? Proteomics -
REDOX PROTEOMICS Roman Zubarev Roman.Zubarev@ki.se Physiological Chemistry I, Department for Medical Biochemistry & Biophysics, Karolinska Institutet, Stockholm What is (RedOx) Proteomics? Proteomics -
MALDI Imaging Mass Spectrometry
 MALDI Imaging Mass Spectrometry Nan Kleinholz Mass Spectrometry and Proteomics Facility The Ohio State University Mass Spectrometry and Proteomics Workshop What is MALDI Imaging? MALDI: Matrix Assisted
MALDI Imaging Mass Spectrometry Nan Kleinholz Mass Spectrometry and Proteomics Facility The Ohio State University Mass Spectrometry and Proteomics Workshop What is MALDI Imaging? MALDI: Matrix Assisted
Proteome analysis Cellular proteomics Plasma proteomics
 Proteome analysis Cellular proteomics Plasma proteomics Janne Lehtiö Maria Pernemalm Karolinska Biomics Center Karolinska Institutet / Karolinska University Hospital Janne Lehtio 2009-02-09 1 KBC Proteomics
Proteome analysis Cellular proteomics Plasma proteomics Janne Lehtiö Maria Pernemalm Karolinska Biomics Center Karolinska Institutet / Karolinska University Hospital Janne Lehtio 2009-02-09 1 KBC Proteomics
Proteomics of body liquids as a source for potential methods for medical diagnostics Prof. Dr. Evgeny Nikolaev
 Proteomics of body liquids as a source for potential methods for medical diagnostics Prof. Dr. Evgeny Nikolaev Institute for Biochemical Physics, Rus. Acad. Sci., Moscow, Russia. Institute for Energy Problems
Proteomics of body liquids as a source for potential methods for medical diagnostics Prof. Dr. Evgeny Nikolaev Institute for Biochemical Physics, Rus. Acad. Sci., Moscow, Russia. Institute for Energy Problems
Digitizing the Proteomes From Big Tissue Biobanks
 Digitizing the Proteomes From Big Tissue Biobanks Analyzing 24 Proteomes Per Day by Microflow SWATH Acquisition and Spectronaut Pulsar Analysis Jan Muntel 1, Nick Morrice 2, Roland M. Bruderer 1, Lukas
Digitizing the Proteomes From Big Tissue Biobanks Analyzing 24 Proteomes Per Day by Microflow SWATH Acquisition and Spectronaut Pulsar Analysis Jan Muntel 1, Nick Morrice 2, Roland M. Bruderer 1, Lukas
MASS SPECTROMETRY IN METABOLOMICS
 For personal use only. Please do not reuse or reproduce without the author s permission MASS SPECTRMETRY IN METABLMICS Pavel Aronov Stanford Mass Spectrometry Users Meeting August 21, 2008 rigin of Metabolomics
For personal use only. Please do not reuse or reproduce without the author s permission MASS SPECTRMETRY IN METABLMICS Pavel Aronov Stanford Mass Spectrometry Users Meeting August 21, 2008 rigin of Metabolomics
The Analysis of Proteomic Spectra from Serum Samples. Keith Baggerly Biostatistics & Applied Mathematics MD Anderson Cancer Center
 The Analysis of Proteomic Spectra from Serum Samples Keith Baggerly Biostatistics & Applied Mathematics MD Anderson Cancer Center PROTEOMICS 1 What Are Proteomic Spectra? DNA makes RNA makes Protein Microarrays
The Analysis of Proteomic Spectra from Serum Samples Keith Baggerly Biostatistics & Applied Mathematics MD Anderson Cancer Center PROTEOMICS 1 What Are Proteomic Spectra? DNA makes RNA makes Protein Microarrays
LECTURE-15. itraq Clinical Applications HANDOUT. Isobaric Tagging for Relative and Absolute quantitation (itraq) is a quantitative MS
 LECTURE-15 itraq Clinical Applications HANDOUT PREAMBLE Isobaric Tagging for Relative and Absolute quantitation (itraq) is a quantitative MS based method for quantifying proteins subject to various different
LECTURE-15 itraq Clinical Applications HANDOUT PREAMBLE Isobaric Tagging for Relative and Absolute quantitation (itraq) is a quantitative MS based method for quantifying proteins subject to various different
Quantification by Mass Spectrometry
 Quantification by Mass Spectrometry PC219 Lecture 5 30 April, 2010 sguan@cgl.ucsf.edu 1 Applications of Quantitative Proteomics Qualitative protein identification is NOT sufficient to describe a biological
Quantification by Mass Spectrometry PC219 Lecture 5 30 April, 2010 sguan@cgl.ucsf.edu 1 Applications of Quantitative Proteomics Qualitative protein identification is NOT sufficient to describe a biological
Nature Methods: doi: /nmeth.3177
 Supplementary Figure 1 Characterization of LysargiNase, trypsin and LysN missed cleavages. (a) Proportion of peptides identified in LysargiNase and trypsin digests of MDA-MB-231 cell lysates carrying 0,
Supplementary Figure 1 Characterization of LysargiNase, trypsin and LysN missed cleavages. (a) Proportion of peptides identified in LysargiNase and trypsin digests of MDA-MB-231 cell lysates carrying 0,
DetergentOUT Tween. DetergentOUT GBS10. OrgoSol DetergentOUT
 252PR 01 G-Biosciences, St Louis, MO. USA 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name DetergentOUT Detergent Removal Systems For the Removal of Detergents
252PR 01 G-Biosciences, St Louis, MO. USA 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name DetergentOUT Detergent Removal Systems For the Removal of Detergents
Automating Mass Spectrometry-Based Quantitative Glycomics using Tandem Mass Tag (TMT) Reagents with SimGlycan
 PREMIER Biosoft Automating Mass Spectrometry-Based Quantitative Glycomics using Tandem Mass Tag (TMT) Reagents with SimGlycan Ne uaca2-3galb1-4glc NAcb1 6 Gal NAca -Thr 3 Ne uaca2-3galb1 Ningombam Sanjib
PREMIER Biosoft Automating Mass Spectrometry-Based Quantitative Glycomics using Tandem Mass Tag (TMT) Reagents with SimGlycan Ne uaca2-3galb1-4glc NAcb1 6 Gal NAca -Thr 3 Ne uaca2-3galb1 Ningombam Sanjib
Mass Spectrometry Infrastructure
 Mass Spectrometry Infrastructure Todd Williams, Ph.D. Director KU Mass Spectrometry and Analytical Proteomics Laboratory Mass Spectrometry Lab B025 Malott Hall Mission The Mass Spectrometry and analytical
Mass Spectrometry Infrastructure Todd Williams, Ph.D. Director KU Mass Spectrometry and Analytical Proteomics Laboratory Mass Spectrometry Lab B025 Malott Hall Mission The Mass Spectrometry and analytical
Thermo Scientific LipidSearch Software for Lipidomics Workflows. Automated Identification and Relative. Quantitation of Lipids by LC/MS
 Thermo Scientific LipidSearch Software for Lipidomics Workflows Automated Identification and Relative of Lipids by LC/MS The promise of lipidomics Lipidomics is a new field of study crucial for understanding
Thermo Scientific LipidSearch Software for Lipidomics Workflows Automated Identification and Relative of Lipids by LC/MS The promise of lipidomics Lipidomics is a new field of study crucial for understanding
DetergentOUT Detergent Removal Systems
 252PR-04 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name DetergentOUT Detergent Removal Systems For the Removal of Detergents from Peptide
252PR-04 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name DetergentOUT Detergent Removal Systems For the Removal of Detergents from Peptide
Mass Spectrometry and Proteomics. Professor Xudong Yao Bioanalytical Chemistry Spring 2007
 Mass Spectrometry and Proteomics Professor Xudong Yao Bioanalytical Chemistry Spring 2007 Proteomics and -omics Roles of mass spectrometry Comparative proteomics Chemical proteomics Protein, Proteome and
Mass Spectrometry and Proteomics Professor Xudong Yao Bioanalytical Chemistry Spring 2007 Proteomics and -omics Roles of mass spectrometry Comparative proteomics Chemical proteomics Protein, Proteome and
What Impact will Proteomics, Including CSF Analysis, have on the Near-term Development of Biomarkers for Nervous System Diseases?
 What Impact will Proteomics, Including CSF Analysis, have on the Near-term Development of Biomarkers for Nervous System Diseases? Daumier 1 Howard Schulman (howard.schulman@menlo.ppdi.com) Biomarker Discovery,
What Impact will Proteomics, Including CSF Analysis, have on the Near-term Development of Biomarkers for Nervous System Diseases? Daumier 1 Howard Schulman (howard.schulman@menlo.ppdi.com) Biomarker Discovery,
Biological Mass spectrometry in Protein Chemistry
 Biological Mass spectrometry in Protein Chemistry Tuula Nyman Institute of Biotechnology tuula.nyman@helsinki.fi MASS SPECTROMETRY is an analytical technique that identifies the chemical composition of
Biological Mass spectrometry in Protein Chemistry Tuula Nyman Institute of Biotechnology tuula.nyman@helsinki.fi MASS SPECTROMETRY is an analytical technique that identifies the chemical composition of
Agilent Protein In-Gel Tryptic Digestion Kit
 Agilent 5188-2749 Protein In-Gel Tryptic Digestion Kit Agilent Protein In-Gel Tryptic Digestion Kit Instructions Kit Contents The Protein In-Gel Tryptic Digestion Kit includes sufficient reagents for approximately
Agilent 5188-2749 Protein In-Gel Tryptic Digestion Kit Agilent Protein In-Gel Tryptic Digestion Kit Instructions Kit Contents The Protein In-Gel Tryptic Digestion Kit includes sufficient reagents for approximately
Nature Biotechnology: doi: /nbt Supplementary Figure 1
 Supplementary Figure 1 The timeline of the NGAG method for extraction of N-linked glycans and glycosite-containing peptides. The timeline can be changed based on the number of samples. Supplementary Figure
Supplementary Figure 1 The timeline of the NGAG method for extraction of N-linked glycans and glycosite-containing peptides. The timeline can be changed based on the number of samples. Supplementary Figure
Using CART to Mine SELDI ProteinChip Data for Biomarkers and Disease Stratification
 Using CART to Mine SELDI ProteinChip Data for Biomarkers and Disease Stratification Kenna Mawk, D.V.M. Informatics Product Manager Ciphergen Biosystems, Inc. Outline Introduction to ProteinChip Technology
Using CART to Mine SELDI ProteinChip Data for Biomarkers and Disease Stratification Kenna Mawk, D.V.M. Informatics Product Manager Ciphergen Biosystems, Inc. Outline Introduction to ProteinChip Technology
Suppression of the Malignant Phenotype of Bladder Cancer Cells Investigations of the Proteome in Relation to Phenotype and Gene Expression
 Suppression of the Malignant Phenotype of Bladder Cancer Cells Investigations of the Proteome in Relation to Phenotype and Gene Expression Topics Modulation of Phenotype by Extracellular Matrix. Bioinformatics
Suppression of the Malignant Phenotype of Bladder Cancer Cells Investigations of the Proteome in Relation to Phenotype and Gene Expression Topics Modulation of Phenotype by Extracellular Matrix. Bioinformatics
MSSimulator. Simulation of Mass Spectrometry Data. Chris Bielow, Stephan Aiche, Sandro Andreotti, Knut Reinert FU Berlin, Germany
 Chris Bielow Algorithmic Bioinformatics, Institute for Computer Science MSSimulator Chris Bielow, Stephan Aiche, Sandro Andreotti, Knut Reinert FU Berlin, Germany Simulation of Mass Spectrometry Data Motivation
Chris Bielow Algorithmic Bioinformatics, Institute for Computer Science MSSimulator Chris Bielow, Stephan Aiche, Sandro Andreotti, Knut Reinert FU Berlin, Germany Simulation of Mass Spectrometry Data Motivation
RDP Cores Highlights: the CF Analytics Core. Facundo M. Fernández School of Chemistry and Biochemistry Georgia Institute of Technology
 CF@LANTA RDP Cores Highlights: the CF Analytics Core Facundo M. Fernández School of Chemistry and Biochemistry Georgia Institute of Technology CF@LANTA RDP Center The CF@LANTA RDP Center at Emory University
CF@LANTA RDP Cores Highlights: the CF Analytics Core Facundo M. Fernández School of Chemistry and Biochemistry Georgia Institute of Technology CF@LANTA RDP Center The CF@LANTA RDP Center at Emory University
New Developments in LC-IMS-MS Proteomic Measurements and Informatic Analyses
 New Developments in LC-IMS-MS Proteomic Measurements and Informatic Analyses Erin Shammel Baker Kristin E. Burnum-Johnson, Xing Zhang, Cameron P. Casey, Yehia M. Ibrahim, Matthew E. Monroe, Tao Liu, Brendan
New Developments in LC-IMS-MS Proteomic Measurements and Informatic Analyses Erin Shammel Baker Kristin E. Burnum-Johnson, Xing Zhang, Cameron P. Casey, Yehia M. Ibrahim, Matthew E. Monroe, Tao Liu, Brendan
Chemical Biology, Option II Mechanism Based Proteomic Tagging Case History CH1
 Proteome Wide Screening of Serine Protease Activity Proc Natl Acad Sci 1999, 97, 14694; Proteomics 2001, 1, 1067; Proc Natl Acad Sci 2002, 99, 10335; Biochemistry 2001, 40, 4005; J. Am. Chem. Soc., 2005,
Proteome Wide Screening of Serine Protease Activity Proc Natl Acad Sci 1999, 97, 14694; Proteomics 2001, 1, 1067; Proc Natl Acad Sci 2002, 99, 10335; Biochemistry 2001, 40, 4005; J. Am. Chem. Soc., 2005,
Protein and peptide separations,
 A Quarterly Technical Newsletter Spring Grace Vydac: Leading the Way in Separations for Proteomics Protein and peptide separations, always important to protein chemists and enzymologists, have recently
A Quarterly Technical Newsletter Spring Grace Vydac: Leading the Way in Separations for Proteomics Protein and peptide separations, always important to protein chemists and enzymologists, have recently
Probability-Based Protein Identification for Post-Translational Modifications and Amino Acid Variants Using Peptide Mass Fingerprint Data
 Probability-Based Protein Identification for Post-Translational Modifications and Amino Acid Variants Using Peptide Mass Fingerprint Data Tong WW, McComb ME, Perlman DH, Huang H, O Connor PB, Costello
Probability-Based Protein Identification for Post-Translational Modifications and Amino Acid Variants Using Peptide Mass Fingerprint Data Tong WW, McComb ME, Perlman DH, Huang H, O Connor PB, Costello
4th Multidimensional Chromatography Workshop Toronto (January, 2013) Herman C. Lam, Ph.D. Calibration & Validation Group
 4th Multidimensional Chromatography Workshop Toronto (January, 2013) Herman C. Lam, Ph.D. Calibration & Validation Group MDLC for Shotgun Proteomics Introduction General concepts Advantages Challenges
4th Multidimensional Chromatography Workshop Toronto (January, 2013) Herman C. Lam, Ph.D. Calibration & Validation Group MDLC for Shotgun Proteomics Introduction General concepts Advantages Challenges
LOCALISATION, IDENTIFICATION AND SEPARATION OF MOLECULES. Gilles Frache Materials Characterization Day October 14 th 2016
 LOCALISATION, IDENTIFICATION AND SEPARATION OF MOLECULES Gilles Frache Materials Characterization Day October 14 th 2016 1 MOLECULAR ANALYSES Which focus? LOCALIZATION of molecules by Mass Spectrometry
LOCALISATION, IDENTIFICATION AND SEPARATION OF MOLECULES Gilles Frache Materials Characterization Day October 14 th 2016 1 MOLECULAR ANALYSES Which focus? LOCALIZATION of molecules by Mass Spectrometry
High-Throughput Analysis of Oligonucleotides using Automated Electrospray Ionization Mass Spectrometry
 High-Throughput Analysis of Oligonucleotides using Automated Electrospray Ionization Mass Spectrometry Mark E. Hail 1, Brian Elliott 2, Kerry Nugent 3, Jeffrey L. Whitney 1, and David J. Detlefsen 1 1
High-Throughput Analysis of Oligonucleotides using Automated Electrospray Ionization Mass Spectrometry Mark E. Hail 1, Brian Elliott 2, Kerry Nugent 3, Jeffrey L. Whitney 1, and David J. Detlefsen 1 1
PosterREPRINT AN AUTOMATED METHOD TO SELF-CALIBRATE AND REJECT NOISE FROM MALDI PEPTIDE MASS FINGERPRINT SPECTRA
 Overview AN AUTOMATED METHOD TO SELF-CALIBRATE AND REJECT NOISE FROM MALDI PEPTIDE MASS FINGERPRINT SPECTRA Jeffery M Brown, Neil Swainston, Dominic O. Gostick, Keith Richardson, Richard Denny, Steven
Overview AN AUTOMATED METHOD TO SELF-CALIBRATE AND REJECT NOISE FROM MALDI PEPTIDE MASS FINGERPRINT SPECTRA Jeffery M Brown, Neil Swainston, Dominic O. Gostick, Keith Richardson, Richard Denny, Steven
New Mass Spectrometry Tools to Transform Metabolomics and Lipidomics
 New Mass Spectrometry Tools to Transform Metabolomics and Lipidomics July.3.13 Ken Miller Vice President of Marketing, Life Sciences Mass Spectrometry 1 The world leader in serving science Omics & the
New Mass Spectrometry Tools to Transform Metabolomics and Lipidomics July.3.13 Ken Miller Vice President of Marketing, Life Sciences Mass Spectrometry 1 The world leader in serving science Omics & the
Mass spectra of peptides and proteins - and LC analysis of proteomes Stephen Barnes, PhD
 Mass spectra of peptides and proteins - and LC analysis of proteomes Stephen Barnes, PhD 4-7117 sbarnes@uab.edu Overview A mass spectrum Electrospray MS Analysis of intact proteins Molecular weight calculations
Mass spectra of peptides and proteins - and LC analysis of proteomes Stephen Barnes, PhD 4-7117 sbarnes@uab.edu Overview A mass spectrum Electrospray MS Analysis of intact proteins Molecular weight calculations
The 1997 ABRF Mass Spectrometry Committee Collaborative Study: Identification of Phosphopeptides in a Tryptic Digest of Apomyoglobin
 The 1997 ABRF Mass Spectrometry Committee Collaborative Study: Identification of Phosphopeptides in a Tryptic Digest of Apomyoglobin Kristine Swiderek, Farzin Gharahdaghi, Lowell Ericsson, Murray Hackett,
The 1997 ABRF Mass Spectrometry Committee Collaborative Study: Identification of Phosphopeptides in a Tryptic Digest of Apomyoglobin Kristine Swiderek, Farzin Gharahdaghi, Lowell Ericsson, Murray Hackett,
FPO. Label-Free Molecular Imaging. Innovation with Integrity. Discover, localize and quantify biochemical changes and molecular markers
 FPO Label-Free Molecular Imaging Discover, localize and quantify biochemical changes and molecular markers Innovation with Integrity Mass Spectrometry Bruker, the global leader in MALDI-MS technology,
FPO Label-Free Molecular Imaging Discover, localize and quantify biochemical changes and molecular markers Innovation with Integrity Mass Spectrometry Bruker, the global leader in MALDI-MS technology,
Trypsin Mass Spectrometry Grade
 058PR-03 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Trypsin Mass Spectrometry Grade A Chemically Modified, TPCK treated, Affinity Purified
058PR-03 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Trypsin Mass Spectrometry Grade A Chemically Modified, TPCK treated, Affinity Purified
Bruker Daltonics. autoflex III smartbeam. The Standard in MALDI-TOF Performance MALDI-TOF/TOF. think forward
 Bruker Daltonics autoflex III smartbeam The Standard in MALDI-TOF Performance think forward MALDI-TOF/TOF Designed for a Routine High Level of Performance The autoflex III smartbeam brings the power of
Bruker Daltonics autoflex III smartbeam The Standard in MALDI-TOF Performance think forward MALDI-TOF/TOF Designed for a Routine High Level of Performance The autoflex III smartbeam brings the power of
Primary Structure Analysis. Automated Evaluation. LC-MS Data Sets
 Primary Structure Analysis by Automated Evaluation of LC-MS Data Sets Mass Spec 29 Dr. Wozny, MassMap GmbH & Co. KG 1 LC-MS Peptide Mapping Origin of Signals Peptides with and without post-translational
Primary Structure Analysis by Automated Evaluation of LC-MS Data Sets Mass Spec 29 Dr. Wozny, MassMap GmbH & Co. KG 1 LC-MS Peptide Mapping Origin of Signals Peptides with and without post-translational
Don t miss a thing on your peptide mapping journey How to get full coverage peptide maps using high resolution accurate mass spectrometry
 Don t miss a thing on your peptide mapping journey How to get full coverage peptide maps using high resolution accurate mass spectrometry Kai Scheffler, PhD BioPharma Support Expert,LSMS Europe The world
Don t miss a thing on your peptide mapping journey How to get full coverage peptide maps using high resolution accurate mass spectrometry Kai Scheffler, PhD BioPharma Support Expert,LSMS Europe The world
Ion Source. Mass Analyzer. Detector. intensity. mass/charge
 Proteomics Informatics Overview of spectrometry (Week 2) Ion Source Analyzer Detector Peptide Fragmentation Ion Source Analyzer 1 Fragmentation Analyzer 2 Detector b y Liquid Chromatography (LC)-MS/MS
Proteomics Informatics Overview of spectrometry (Week 2) Ion Source Analyzer Detector Peptide Fragmentation Ion Source Analyzer 1 Fragmentation Analyzer 2 Detector b y Liquid Chromatography (LC)-MS/MS
SYNAPT G2-S High Definition MS (HDMS) System
 SYNAPT G2-S High Definition MS (HDMS) System High performance, versatility, and workflow efficiency of your MS system all play a crucial role in your ability to successfully reach your scientific and business
SYNAPT G2-S High Definition MS (HDMS) System High performance, versatility, and workflow efficiency of your MS system all play a crucial role in your ability to successfully reach your scientific and business
SUPPORTING INFORMATION. Lysine Carbonylation is a Previously Unrecognized Contributor. to Peroxidase Activation of Cytochrome c by Chloramine-T
 Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2019 SUPPORTING INFORMATION Lysine Carbonylation is a Previously Unrecognized Contributor to
Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2019 SUPPORTING INFORMATION Lysine Carbonylation is a Previously Unrecognized Contributor to
AB Sciex QStar XL. AIMS Instrumentation & Sample Report Documentation. chemistry
 Mass Spectrometry Laboratory AIMS Instrumentation & Sample Report Documentation AB Sciex QStar XL chemistry UNIVERSITY OF TORONTO AIMS Mass Spectrometry Laboratory Department of Chemistry, University of
Mass Spectrometry Laboratory AIMS Instrumentation & Sample Report Documentation AB Sciex QStar XL chemistry UNIVERSITY OF TORONTO AIMS Mass Spectrometry Laboratory Department of Chemistry, University of
N α -Acetylation of yeast ribosomal proteins and its effect on protein synthesis
 JOURNAL OF PROTEOMICS 74 (2011) 431 441 available at www.sciencedirect.com www.elsevier.com/locate/jprot N α -Acetylation of yeast ribosomal proteins and its effect on protein synthesis Masahiro Kamita
JOURNAL OF PROTEOMICS 74 (2011) 431 441 available at www.sciencedirect.com www.elsevier.com/locate/jprot N α -Acetylation of yeast ribosomal proteins and its effect on protein synthesis Masahiro Kamita
MALDI Imaging Drug Imaging Detlev Suckau Head of R&D MALDI Bruker Daltonik GmbH. December 19,
 MALDI Imaging Drug Imaging Detlev Suckau Head of R&D MALDI Bruker Daltonik GmbH December 19, 2014 1 The principle of MALDI imaging Spatially resolved mass spectra are recorded Each mass signal represents
MALDI Imaging Drug Imaging Detlev Suckau Head of R&D MALDI Bruker Daltonik GmbH December 19, 2014 1 The principle of MALDI imaging Spatially resolved mass spectra are recorded Each mass signal represents
An Overview of Recent Developments for Mass Spectrometry in the Field of Proteomics
 An Overview of Recent Developments for Mass Spectrometry in the Field of Proteomics Jeffrey W. Finch, Ph.D. 44th Rocky Mountain Conference July 30th, 2002 Outline 1. Background a. Proteomics b. MS Tools
An Overview of Recent Developments for Mass Spectrometry in the Field of Proteomics Jeffrey W. Finch, Ph.D. 44th Rocky Mountain Conference July 30th, 2002 Outline 1. Background a. Proteomics b. MS Tools
Nature Structural & Molecular Biology: doi: /nsmb.3218
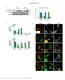 Supplementary Figure 1 Endogenous EGFR trafficking and responses depend on biased ligands. (a) Lysates from HeLa cells stimulated for 2 min. with increasing concentration of ligands were immunoblotted
Supplementary Figure 1 Endogenous EGFR trafficking and responses depend on biased ligands. (a) Lysates from HeLa cells stimulated for 2 min. with increasing concentration of ligands were immunoblotted
Babu Antharavally, Ryan Bomgarden, and John Rogers Thermo Fisher Scientific, Rockford, IL
 A Versatile High-Recovery Method for Removing Detergents from Low-Concentration Protein or Peptide Samples for Mass Spectrometry Sample Preparation and Analysis Babu Antharavally, Ryan Bomgarden, and John
A Versatile High-Recovery Method for Removing Detergents from Low-Concentration Protein or Peptide Samples for Mass Spectrometry Sample Preparation and Analysis Babu Antharavally, Ryan Bomgarden, and John
NIH Public Access Author Manuscript J Proteome Res. Author manuscript; available in PMC 2014 July 05.
 NIH Public Access Author Manuscript Published in final edited form as: J Proteome Res. 2013 July 5; 12(7): 3071 3086. doi:10.1021/pr3011588. Evaluation and Optimization of Mass Spectrometric Settings during
NIH Public Access Author Manuscript Published in final edited form as: J Proteome Res. 2013 July 5; 12(7): 3071 3086. doi:10.1021/pr3011588. Evaluation and Optimization of Mass Spectrometric Settings during
PosterREPRINT INTRODUCTION. 2-D PAGE of Mouse Liver Samples. 2-D PAGE of E.coli Samples. Digestion / Cleanup. EXPERIMENTAL 1-D PAGE of BSA Samples
 INTRODUCTION Identification and characterization of low abundance proteins separated by 2D gel electrophoresis is complicated by two important factors; the use of suitable staining techniques for the visualization
INTRODUCTION Identification and characterization of low abundance proteins separated by 2D gel electrophoresis is complicated by two important factors; the use of suitable staining techniques for the visualization
Chapter 3. Protein Structure and Function
 Chapter 3 Protein Structure and Function Broad functional classes So Proteins have structure and function... Fine! -Why do we care to know more???? Understanding functional architechture gives us POWER
Chapter 3 Protein Structure and Function Broad functional classes So Proteins have structure and function... Fine! -Why do we care to know more???? Understanding functional architechture gives us POWER
Mass Spectrometry and Proteomics Xudong Yao
 Mass Spectrometry and Proteomics Xudong Yao Dept of Chemistry University of Connecticut Storrs, CT April 19, 2005 Proteomics and -omics Roles of mass spectrometry Comparative proteomics Gel or non-gel
Mass Spectrometry and Proteomics Xudong Yao Dept of Chemistry University of Connecticut Storrs, CT April 19, 2005 Proteomics and -omics Roles of mass spectrometry Comparative proteomics Gel or non-gel
AccuMAP Low ph Protein Digestion Kits
 TECHNICAL MANUAL AccuMAP Low ph Protein Digestion Kits Instruc ons for Use of Products VA1040 and VA1050 5/17 TM504 AccuMAP Low ph Protein Digestion Kits All technical literature is available at: www.promega.com/protocols/
TECHNICAL MANUAL AccuMAP Low ph Protein Digestion Kits Instruc ons for Use of Products VA1040 and VA1050 5/17 TM504 AccuMAP Low ph Protein Digestion Kits All technical literature is available at: www.promega.com/protocols/
Mass Spectrometry at the Laboratory of Food Chemistry. Edwin Bakx Laboratory of Food Chemistry Wageningen University
 Mass Spectrometry at the Wageningen University Mass Spectrometry at the 3 UPLC/CE - ESI - Ion trap MS systems UPLC Thermo Acella with a Velos or VelosPro CE Beckman PA800 with a Thermo VelosPro 1 UPLC-
Mass Spectrometry at the Wageningen University Mass Spectrometry at the 3 UPLC/CE - ESI - Ion trap MS systems UPLC Thermo Acella with a Velos or VelosPro CE Beckman PA800 with a Thermo VelosPro 1 UPLC-
Case Center for Proteomics and Bioinformatics Workshop Series
 Case Center for Proteomics and Bioinformatics Workshop Series March 5: Masaru Miyagi and Chao Yuan Will Present: Quantitative proteomic analysis using stable isotopic labeling April 2: Elizabeth Yohannes
Case Center for Proteomics and Bioinformatics Workshop Series March 5: Masaru Miyagi and Chao Yuan Will Present: Quantitative proteomic analysis using stable isotopic labeling April 2: Elizabeth Yohannes
Metabolomics: quantifying the phenotype
 Metabolomics: quantifying the phenotype Metabolomics Promises Quantitative Phenotyping What can happen GENOME What appears to be happening Bioinformatics TRANSCRIPTOME What makes it happen PROTEOME Systems
Metabolomics: quantifying the phenotype Metabolomics Promises Quantitative Phenotyping What can happen GENOME What appears to be happening Bioinformatics TRANSCRIPTOME What makes it happen PROTEOME Systems
Mass Spectrometry Course Árpád Somogyi Chemistry and Biochemistry MassSpectrometry Facility) University of Debrecen, April 12-23, 2010
 Mass Spectrometry Course Árpád Somogyi Chemistry and Biochemistry MassSpectrometry Facility) University of Debrecen, April 12-23, 2010 Introduction, Ionization Methods Mass Analyzers, Ion Activation Methods
Mass Spectrometry Course Árpád Somogyi Chemistry and Biochemistry MassSpectrometry Facility) University of Debrecen, April 12-23, 2010 Introduction, Ionization Methods Mass Analyzers, Ion Activation Methods
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/8/398/rs12/dc1 Supplementary Materials for Quantitative phosphoproteomics reveals new roles for the protein phosphatase PP6 in mitotic cells Scott F. Rusin, Kate
www.sciencesignaling.org/cgi/content/full/8/398/rs12/dc1 Supplementary Materials for Quantitative phosphoproteomics reveals new roles for the protein phosphatase PP6 in mitotic cells Scott F. Rusin, Kate
SIEVE 2.1 Proteomics Example
 SIEVE 2.1 Proteomics Example Software Overview What is SIEVE? SIEVE is Thermo Scientific s differential software solution. SIEVE will continue to enhance our current product for label-free differential
SIEVE 2.1 Proteomics Example Software Overview What is SIEVE? SIEVE is Thermo Scientific s differential software solution. SIEVE will continue to enhance our current product for label-free differential
SELYMATRA: Web Application for the analysis. of mass spectra
 SELYMATRA: Web Application for the analysis of mass spectra arxiv:1710.05914v1 [q-bio.qm] 15 Oct 2017 Davide Nardone, Angelo Ciaramella, Mariangela Cerreta, Salvatore Pulcrano, Gian Carlo Bellenchi, Giuseppe
SELYMATRA: Web Application for the analysis of mass spectra arxiv:1710.05914v1 [q-bio.qm] 15 Oct 2017 Davide Nardone, Angelo Ciaramella, Mariangela Cerreta, Salvatore Pulcrano, Gian Carlo Bellenchi, Giuseppe
