White Paper Estimating Genotype-Specific Incidence for One or Several Loci
|
|
|
- Molly Kelly
- 5 years ago
- Views:
Transcription
1 White Paper Estimating Genotype-Specific Incidence for One or Several Loci Authors: Mike Macpherson Brian Naughton Andro Hsu Joanna Mountain Created: September 5, 2007 Last Edited: November 18, 2007 Summary: We wish to estimate the incidence for a given trait based on an individual s genotype at one or more SNPs associated with the trait and any available phenotypic data for that individual. Here we describe the methods used to estimate these incidences, first for the case of a single SNP, then for the case of multiple SNPs.
2 Single Locus Calculation In practice, we provide an estimate of trait incidence conditional on the individual s genotype and available phenotypic information. The calculations below, however, hold whether incidence or prevalence estimates are computed. We henceforth use the generic term risk instead of incidence to indicate this fact. We assume a binary trait D and a single associated locus at which there are three possible genotypes G 1, G 2, and G 3, ordered so that G 1 is the lower-risk homozygote and G 3 is the higher-risk homozygote. The individual for whom we wish to estimate the incidence has genotype G m : m {1, 2, 3}. The quantity we wish to compute is Pr(D G m ), or the probability that the individual is affected given their genotype. The unconditional risk for the trait, denoted Pr(D), is assumed known for a subpopulation of which the individual is a member. For instance, this might be an estimate of Type 2 diabetes prevalence for Asian subjects between the ages of 20 and 40. We further assume that estimates of the three genotype frequencies Pr(G i ) are available for the same subpopulation. Lastly, we assume that estimates of the three genotype-specific odds ratios OR 1, OR 2, and OR 3 are available, where OR 1 =1by definition. Under these assumptions, we may compute the Pr(D G i ) by solving the following system of equations: Pr(D) = Pr(D G 1 )Pr(G 1 ) + Pr(D G 2 )Pr(G 2 ) + Pr(D G 3 )Pr(G 3 ) (1) OR 2 = Pr(D G 2)/(1 Pr(D G 2 )) Pr(D G 1 )/(1 Pr(D G 1 )) (2) OR 3 = Pr(D G 3)/(1 Pr(D G 3 )) Pr(D G 1 )/(1 Pr(D G 1 )) (3) Equation 1 follows from basic probability theory, and Equations 2 and 3 follow from the definition of an odds ratio. The estimated genotype-specific risk at the single locus is then Pr(D G m ). To convey the genotype-specific risk relative to the average risk in the population, we compute the quantity OR m = odds(d G m )/odds(d), where, for brevity, we have introduced the function odds(x) = Pr(X)/(1 Pr(X)). The inverse odds function will be used later on, and is odds 1 (X) = Pr(X)/(1 + Pr(X)). The superscript asterisk on OR m is a convention we adopt to distinguish an odds ratio computed relative to the average odds, rather than relative to the odds of lowerrisk homozygote, which is the standard definition. 1
3 Complications Odds Ratio Estimates: In the association study literature, odds ratios are commonly estimated via logistic regression assuming an additive model. This means that the logarithm of the odds ratio is assumed to relate linearly to the number of copies of the higher-risk allele (cf. Jewell, 2003). When the odds ratio is estimated in this way, the higher-risk homozygote s estimated log odds ratio is, by definition, exactly twice that of the heterozygote s log odds ratio, from which it follows that the higher-risk homozygote s odds ratio is the square of the heterozygote s odds ratio. Thus the odds ratio reported when the additive model is assumed is that associated with one copy of the higher-risk allele, which could be called an allelespecific odds ratio. In such cases we set the allele-specific odds ratio equal to the heterozygous genotype s odds ratio, OR 2, and set the higher-risk homozygote s odds ratio OR 3 =(OR 2 ) 2. In cases for which genotype-specific odds ratios are estimated separately, we use those estimates directly. Prevalence v. Incidence: We typically report genotype-specific risk in terms of incidence. However, the formulas in this white paper apply equally well to prevalence and incidence data. Specifically, the value Pr(D) may represent either a prevalence or an incidence datum. Mismatched Datasets: In practice, we rely on association studies that are most often based on individuals of European descent for our odds ratio estimates. We obtain genotype frequency estimates primarily from HapMap, which provides one population of European descent, one of Yoruban descent, and one of Asian (Chinese & Japanese) descent. We obtain prevalence/incidence estimates from a variety of sources, and these estimates often pertain to individuals of European descent alone. Ideally, we would have perfectly-matched odds ratio, genotype frequency, and prevalence estimates, meaning that, if the prevalence estimate pertains to Asian subjects between 20 and 40, we would also have genotype frequency estimates and odds ratio estimates for Asians between 20 and 40. It is most often the case that we do not have such matched estimates. At this writing, we assume that the overall HapMap genotype frequency estimates apply across all age ranges, i.e. that the genotype frequencies within any age range are identical to the overall frequencies. We also assume that odds ratios apply across age ranges. We record whether an odds ratio estimate derives from a European, African, or Asian population, and do the same for prevalence estimates. By derives we mean that we consider the population for which a given odds ratio or prevalence estimate, and (subjectively) decide whether that population is near enough to one of the three 2
4 HapMap populations for us to use the estimate. We only report genotype-specific risk estimates when the underlying estimates match at the population level. For example, when an odds ratio estimate exists for an Asian population, a genotype frequency estimate exists for an Asian population, and a prevalence estimate exists for an Asian population, we will report the genotype-specific risk estimates, even if the prevalence estimate is age-range specific. At this writing, we have not studied how this policy affects the accuracy of the estimates. Higher Moments: At this writing, we do not provide estimates of our certainty in the reported genotype-specific risk point estimate. It is standard practice to provide confidence intervals for odds ratios reported in the literature. We have sample sizes for the HapMap frequency data, and often have sample sizes for prevalence estimates, so it should be relatively straightforward to produce risk confidence intervals, and informative to our users. Multiple Locus Calculation For many traits, associations exist between the trait and several SNPs. We wish to combine the genotype-specific risk estimates from multiple loci into a single, genotype-specific composite risk estimate for an individual s genotype. Here we assume that the composite odds ratio, OR C, is given by the product of the individual s odds ratios at each locus. This is very similar to what is done in multiple logistic regression under an additive model, where each copy of a risk allele at a given locus is assumed to add one unit of the log odds ratio specific to that locus to the overall log odds ratio (cf. Jewell, 2003; Risch, 1990). It is not exactly the same because a multiple regression would be performed simultaneously on all the data, where we use individual odds ratios obtained from separate studies, which may, for instance, differ widely in sample size. We assume that there are K loci of interest, and denote the kth odds ratio of the ith genotype OR i,k. Similarly, the recentered odds ratios are denoted ORi,k. We denote the genotypes at the kth locus G 1,k,G 2,k,G 3,k, and denote the individual s genotype at the kth locus G mk,k. Thus OR C = K ORm k,k (4) k=1 The quantity OR C has the interpretation 3
5 OR C = odds(d K k=1 G mk,k)/odds(d), (5) where odds(d K k=1 G m k,k) is the odds of the individual s multilocus genotype. In computing the product ORC, we implicitly assume that the point where log(odds(d)) intersects the respective logistic regression line for each locus is the same point in each of the individual regression calculations, as would be true if a multiple logistic regression had been performed. Then Pr(D K k=1 G mk,k) =odds 1 [OR C odds(d)]. (6) Complications Higher Moments: As in the single-locus case, we do not calculate confidence intervals for the estimates we provide. Provided confidence intervals for individual risk at a single locus, it is straightforward to compute them for the composite risk. References Jewell, Nicholas P Statistics for Epidemiology. Chapman & Hall/CRC. Risch, N Linkage strategies for genetically complex traits. I. Multilocus models. Am J Hum Genet, 46(2),
(b) What is the allele frequency of the b allele in the new merged population on the island?
 2005 7.03 Problem Set 6 KEY Due before 5 PM on WEDNESDAY, November 23, 2005. Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. Two populations (Population One
2005 7.03 Problem Set 6 KEY Due before 5 PM on WEDNESDAY, November 23, 2005. Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. Two populations (Population One
GENETIC LINKAGE ANALYSIS
 Atlas of Genetics and Cytogenetics in Oncology and Haematology GENETIC LINKAGE ANALYSIS * I- Recombination fraction II- Definition of the "lod score" of a family III- Test for linkage IV- Estimation of
Atlas of Genetics and Cytogenetics in Oncology and Haematology GENETIC LINKAGE ANALYSIS * I- Recombination fraction II- Definition of the "lod score" of a family III- Test for linkage IV- Estimation of
Pedigree Analysis Why do Pedigrees? Goals of Pedigree Analysis Basic Symbols More Symbols Y-Linked Inheritance
 Pedigree Analysis Why do Pedigrees? Punnett squares and chi-square tests work well for organisms that have large numbers of offspring and controlled mating, but humans are quite different: Small families.
Pedigree Analysis Why do Pedigrees? Punnett squares and chi-square tests work well for organisms that have large numbers of offspring and controlled mating, but humans are quite different: Small families.
HARDY- WEINBERG PRACTICE PROBLEMS
 HARDY- WEINBERG PRACTICE PROBLEMS PROBLEMS TO SOLVE: 1. The proportion of homozygous recessives of a certain population is 0.09. If we assume that the gene pool is large and at equilibrium and all genotypes
HARDY- WEINBERG PRACTICE PROBLEMS PROBLEMS TO SOLVE: 1. The proportion of homozygous recessives of a certain population is 0.09. If we assume that the gene pool is large and at equilibrium and all genotypes
The Loss of Heterozygosity (LOH) Algorithm in Genotyping Console 2.0
 The Loss of Heterozygosity (LOH) Algorithm in Genotyping Console 2.0 Introduction Loss of erozygosity (LOH) represents the loss of allelic differences. The SNP markers on the SNP Array 6.0 can be used
The Loss of Heterozygosity (LOH) Algorithm in Genotyping Console 2.0 Introduction Loss of erozygosity (LOH) represents the loss of allelic differences. The SNP markers on the SNP Array 6.0 can be used
White Paper Estimating Complex Phenotype Prevalence Using Predictive Models
 White Paper 23-12 Estimating Complex Phenotype Prevalence Using Predictive Models Authors: Nicholas A. Furlotte Aaron Kleinman Robin Smith David Hinds Created: September 25 th, 2015 September 25th, 2015
White Paper 23-12 Estimating Complex Phenotype Prevalence Using Predictive Models Authors: Nicholas A. Furlotte Aaron Kleinman Robin Smith David Hinds Created: September 25 th, 2015 September 25th, 2015
DEFINITIONS OF HISTOCOMPATIBILITY TYPING TERMS
 DEFINITIONS OF HISTOCOMPATIBILITY TYPING TERMS The definitions below are intended as general concepts. There will be exceptions to these general definitions. These definitions do not imply any specific
DEFINITIONS OF HISTOCOMPATIBILITY TYPING TERMS The definitions below are intended as general concepts. There will be exceptions to these general definitions. These definitions do not imply any specific
Analysis of single gene effects 1. Quantitative analysis of single gene effects. Gregory Carey, Barbara J. Bowers, Jeanne M.
 Analysis of single gene effects 1 Quantitative analysis of single gene effects Gregory Carey, Barbara J. Bowers, Jeanne M. Wehner From the Department of Psychology (GC, JMW) and Institute for Behavioral
Analysis of single gene effects 1 Quantitative analysis of single gene effects Gregory Carey, Barbara J. Bowers, Jeanne M. Wehner From the Department of Psychology (GC, JMW) and Institute for Behavioral
GENOME-WIDE ASSOCIATION STUDIES
 GENOME-WIDE ASSOCIATION STUDIES SUCCESSES AND PITFALLS IBT 2012 Human Genetics & Molecular Medicine Zané Lombard IDENTIFYING DISEASE GENES??? Nature, 15 Feb 2001 Science, 16 Feb 2001 IDENTIFYING DISEASE
GENOME-WIDE ASSOCIATION STUDIES SUCCESSES AND PITFALLS IBT 2012 Human Genetics & Molecular Medicine Zané Lombard IDENTIFYING DISEASE GENES??? Nature, 15 Feb 2001 Science, 16 Feb 2001 IDENTIFYING DISEASE
Decomposition of the Genotypic Value
 Decomposition of the Genotypic Value 1 / 17 Partitioning of Phenotypic Values We introduced the general model of Y = G + E in the first lecture, where Y is the phenotypic value, G is the genotypic value,
Decomposition of the Genotypic Value 1 / 17 Partitioning of Phenotypic Values We introduced the general model of Y = G + E in the first lecture, where Y is the phenotypic value, G is the genotypic value,
Example HLA-B and abacavir. Roujeau 2014
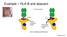 Example HLA-B and abacavir Roujeau 2014 FDA requires testing for abacavir Treatment with abacavir is generally well tolerated, but 5% of the patients experience hypersensitivity reactions that can be life
Example HLA-B and abacavir Roujeau 2014 FDA requires testing for abacavir Treatment with abacavir is generally well tolerated, but 5% of the patients experience hypersensitivity reactions that can be life
White Paper Guidelines on Vetting Genetic Associations
 White Paper 23-03 Guidelines on Vetting Genetic Associations Authors: Andro Hsu Brian Naughton Shirley Wu Created: November 14, 2007 Revised: February 14, 2008 Revised: June 10, 2010 (see end of document
White Paper 23-03 Guidelines on Vetting Genetic Associations Authors: Andro Hsu Brian Naughton Shirley Wu Created: November 14, 2007 Revised: February 14, 2008 Revised: June 10, 2010 (see end of document
Supplementary Figures
 Supplementary Figures Supplementary Fig 1. Comparison of sub-samples on the first two principal components of genetic variation. TheBritishsampleisplottedwithredpoints.The sub-samples of the diverse sample
Supplementary Figures Supplementary Fig 1. Comparison of sub-samples on the first two principal components of genetic variation. TheBritishsampleisplottedwithredpoints.The sub-samples of the diverse sample
Introduction to linkage and family based designs to study the genetic epidemiology of complex traits. Harold Snieder
 Introduction to linkage and family based designs to study the genetic epidemiology of complex traits Harold Snieder Overview of presentation Designs: population vs. family based Mendelian vs. complex diseases/traits
Introduction to linkage and family based designs to study the genetic epidemiology of complex traits Harold Snieder Overview of presentation Designs: population vs. family based Mendelian vs. complex diseases/traits
IN SILICO EVALUATION OF DNA-POOLED ALLELOTYPING VERSUS INDIVIDUAL GENOTYPING FOR GENOME-WIDE ASSOCIATION STUDIES OF COMPLEX DISEASE.
 IN SILICO EVALUATION OF DNA-POOLED ALLELOTYPING VERSUS INDIVIDUAL GENOTYPING FOR GENOME-WIDE ASSOCIATION STUDIES OF COMPLEX DISEASE By Siddharth Pratap Thesis Submitted to the Faculty of the Graduate School
IN SILICO EVALUATION OF DNA-POOLED ALLELOTYPING VERSUS INDIVIDUAL GENOTYPING FOR GENOME-WIDE ASSOCIATION STUDIES OF COMPLEX DISEASE By Siddharth Pratap Thesis Submitted to the Faculty of the Graduate School
Systems of Mating: Systems of Mating:
 8/29/2 Systems of Mating: the rules by which pairs of gametes are chosen from the local gene pool to be united in a zygote with respect to a particular locus or genetic system. Systems of Mating: A deme
8/29/2 Systems of Mating: the rules by which pairs of gametes are chosen from the local gene pool to be united in a zygote with respect to a particular locus or genetic system. Systems of Mating: A deme
An Introduction to Quantitative Genetics I. Heather A Lawson Advanced Genetics Spring2018
 An Introduction to Quantitative Genetics I Heather A Lawson Advanced Genetics Spring2018 Outline What is Quantitative Genetics? Genotypic Values and Genetic Effects Heritability Linkage Disequilibrium
An Introduction to Quantitative Genetics I Heather A Lawson Advanced Genetics Spring2018 Outline What is Quantitative Genetics? Genotypic Values and Genetic Effects Heritability Linkage Disequilibrium
IB BIO I Genetics Test Madden
 Name Date Multiple Choice 1. What does the genotype X H X h indicate? A. A co-dominant female B. A heterozygous male C. A heterozygous female D. A co-dominant male 2. A pure breeding tall plant with smooth
Name Date Multiple Choice 1. What does the genotype X H X h indicate? A. A co-dominant female B. A heterozygous male C. A heterozygous female D. A co-dominant male 2. A pure breeding tall plant with smooth
Association-heterogeneity mapping identifies an Asian-specific association of the GTF2I locus with rheumatoid arthritis
 Supplementary Material Association-heterogeneity mapping identifies an Asian-specific association of the GTF2I locus with rheumatoid arthritis Kwangwoo Kim 1,, So-Young Bang 1,, Katsunori Ikari 2,3, Dae
Supplementary Material Association-heterogeneity mapping identifies an Asian-specific association of the GTF2I locus with rheumatoid arthritis Kwangwoo Kim 1,, So-Young Bang 1,, Katsunori Ikari 2,3, Dae
MENDELIAN GENETICS. MENDEL RULE AND LAWS Please read and make sure you understand the following instructions and knowledge before you go on.
 MENDELIAN GENETICS Objectives Upon completion of this lab, students should: 1. Understand the principles and terms used in Mendelian genetics. 2. Know how to complete a Punnett square to estimate phenotypic
MENDELIAN GENETICS Objectives Upon completion of this lab, students should: 1. Understand the principles and terms used in Mendelian genetics. 2. Know how to complete a Punnett square to estimate phenotypic
Non-parametric methods for linkage analysis
 BIOSTT516 Statistical Methods in Genetic Epidemiology utumn 005 Non-parametric methods for linkage analysis To this point, we have discussed model-based linkage analyses. These require one to specify a
BIOSTT516 Statistical Methods in Genetic Epidemiology utumn 005 Non-parametric methods for linkage analysis To this point, we have discussed model-based linkage analyses. These require one to specify a
Supplementary Figure 1. Principal components analysis of European ancestry in the African American, Native Hawaiian and Latino populations.
 Supplementary Figure. Principal components analysis of European ancestry in the African American, Native Hawaiian and Latino populations. a Eigenvector 2.5..5.5. African Americans European Americans e
Supplementary Figure. Principal components analysis of European ancestry in the African American, Native Hawaiian and Latino populations. a Eigenvector 2.5..5.5. African Americans European Americans e
Inbreeding and Inbreeding Depression
 Inbreeding and Inbreeding Depression Inbreeding is mating among relatives which increases homozygosity Why is Inbreeding a Conservation Concern: Inbreeding may or may not lead to inbreeding depression,
Inbreeding and Inbreeding Depression Inbreeding is mating among relatives which increases homozygosity Why is Inbreeding a Conservation Concern: Inbreeding may or may not lead to inbreeding depression,
The Association Design and a Continuous Phenotype
 PSYC 5102: Association Design & Continuous Phenotypes (4/4/07) 1 The Association Design and a Continuous Phenotype The purpose of this note is to demonstrate how to perform a population-based association
PSYC 5102: Association Design & Continuous Phenotypes (4/4/07) 1 The Association Design and a Continuous Phenotype The purpose of this note is to demonstrate how to perform a population-based association
Mendel s Methods: Monohybrid Cross
 Mendel s Methods: Monohybrid Cross Mendel investigated whether the white-flowered form disappeared entirely by breeding the F1 purple flowers with each other. Crossing two purple F1 monohybrid plants is
Mendel s Methods: Monohybrid Cross Mendel investigated whether the white-flowered form disappeared entirely by breeding the F1 purple flowers with each other. Crossing two purple F1 monohybrid plants is
Supplementary Figure 1: Attenuation of association signals after conditioning for the lead SNP. a) attenuation of association signal at the 9p22.
 Supplementary Figure 1: Attenuation of association signals after conditioning for the lead SNP. a) attenuation of association signal at the 9p22.32 PCOS locus after conditioning for the lead SNP rs10993397;
Supplementary Figure 1: Attenuation of association signals after conditioning for the lead SNP. a) attenuation of association signal at the 9p22.32 PCOS locus after conditioning for the lead SNP rs10993397;
Statistical Tests for X Chromosome Association Study. with Simulations. Jian Wang July 10, 2012
 Statistical Tests for X Chromosome Association Study with Simulations Jian Wang July 10, 2012 Statistical Tests Zheng G, et al. 2007. Testing association for markers on the X chromosome. Genetic Epidemiology
Statistical Tests for X Chromosome Association Study with Simulations Jian Wang July 10, 2012 Statistical Tests Zheng G, et al. 2007. Testing association for markers on the X chromosome. Genetic Epidemiology
For more information about how to cite these materials visit
 Author(s): Kerby Shedden, Ph.D., 2010 License: Unless otherwise noted, this material is made available under the terms of the Creative Commons Attribution Share Alike 3.0 License: http://creativecommons.org/licenses/by-sa/3.0/
Author(s): Kerby Shedden, Ph.D., 2010 License: Unless otherwise noted, this material is made available under the terms of the Creative Commons Attribution Share Alike 3.0 License: http://creativecommons.org/licenses/by-sa/3.0/
Pedigree Construction Notes
 Name Date Pedigree Construction Notes GO TO à Mendelian Inheritance (http://www.uic.edu/classes/bms/bms655/lesson3.html) When human geneticists first began to publish family studies, they used a variety
Name Date Pedigree Construction Notes GO TO à Mendelian Inheritance (http://www.uic.edu/classes/bms/bms655/lesson3.html) When human geneticists first began to publish family studies, they used a variety
Replacing IBS with IBD: The MLS Method. Biostatistics 666 Lecture 15
 Replacing IBS with IBD: The MLS Method Biostatistics 666 Lecture 5 Previous Lecture Analysis of Affected Relative Pairs Test for Increased Sharing at Marker Expected Amount of IBS Sharing Previous Lecture:
Replacing IBS with IBD: The MLS Method Biostatistics 666 Lecture 5 Previous Lecture Analysis of Affected Relative Pairs Test for Increased Sharing at Marker Expected Amount of IBS Sharing Previous Lecture:
Bayesian approaches to handling missing data: Practical Exercises
 Bayesian approaches to handling missing data: Practical Exercises 1 Practical A Thanks to James Carpenter and Jonathan Bartlett who developed the exercise on which this practical is based (funded by ESRC).
Bayesian approaches to handling missing data: Practical Exercises 1 Practical A Thanks to James Carpenter and Jonathan Bartlett who developed the exercise on which this practical is based (funded by ESRC).
UNIVERSITY OF CALIFORNIA, LOS ANGELES
 UNIVERSITY OF CALIFORNIA, LOS ANGELES BERKELEY DAVIS IRVINE LOS ANGELES MERCED RIVERSIDE SAN DIEGO SAN FRANCISCO UCLA SANTA BARBARA SANTA CRUZ DEPARTMENT OF EPIDEMIOLOGY SCHOOL OF PUBLIC HEALTH CAMPUS
UNIVERSITY OF CALIFORNIA, LOS ANGELES BERKELEY DAVIS IRVINE LOS ANGELES MERCED RIVERSIDE SAN DIEGO SAN FRANCISCO UCLA SANTA BARBARA SANTA CRUZ DEPARTMENT OF EPIDEMIOLOGY SCHOOL OF PUBLIC HEALTH CAMPUS
A Unified Sampling Approach for Multipoint Analysis of Qualitative and Quantitative Traits in Sib Pairs
 Am. J. Hum. Genet. 66:1631 1641, 000 A Unified Sampling Approach for Multipoint Analysis of Qualitative and Quantitative Traits in Sib Pairs Kung-Yee Liang, 1 Chiung-Yu Huang, 1 and Terri H. Beaty Departments
Am. J. Hum. Genet. 66:1631 1641, 000 A Unified Sampling Approach for Multipoint Analysis of Qualitative and Quantitative Traits in Sib Pairs Kung-Yee Liang, 1 Chiung-Yu Huang, 1 and Terri H. Beaty Departments
Genetics Unit Exam. Number of progeny with following phenotype Experiment Red White #1: Fish 2 (red) with Fish 3 (red) 100 0
 Genetics Unit Exam Question You are working with an ornamental fish that shows two color phenotypes, red or white. The color is controlled by a single gene. These fish are hermaphrodites meaning they can
Genetics Unit Exam Question You are working with an ornamental fish that shows two color phenotypes, red or white. The color is controlled by a single gene. These fish are hermaphrodites meaning they can
Ascertainment Through Family History of Disease Often Decreases the Power of Family-based Association Studies
 Behav Genet (2007) 37:631 636 DOI 17/s10519-007-9149-0 ORIGINAL PAPER Ascertainment Through Family History of Disease Often Decreases the Power of Family-based Association Studies Manuel A. R. Ferreira
Behav Genet (2007) 37:631 636 DOI 17/s10519-007-9149-0 ORIGINAL PAPER Ascertainment Through Family History of Disease Often Decreases the Power of Family-based Association Studies Manuel A. R. Ferreira
Mendel explained how a dominant allele can mask the presence of a recessive allele.
 Section 2: Mendel explained how a dominant allele can mask the presence of a recessive allele. K What I Know W What I Want to Find Out L What I Learned Essential Questions What is the significance of Mendel
Section 2: Mendel explained how a dominant allele can mask the presence of a recessive allele. K What I Know W What I Want to Find Out L What I Learned Essential Questions What is the significance of Mendel
Assessing Gene-Environment Interactions in Genome-Wide Association Studies: Statistical Approaches
 Research Report Assessing Gene-Environment Interactions in Genome-Wide Association Studies: Statistical Approaches Philip C. Cooley, Robert F. Clark, and Ralph E. Folsom May 2014 RTI Press About the Authors
Research Report Assessing Gene-Environment Interactions in Genome-Wide Association Studies: Statistical Approaches Philip C. Cooley, Robert F. Clark, and Ralph E. Folsom May 2014 RTI Press About the Authors
Single Gene (Monogenic) Disorders. Mendelian Inheritance: Definitions. Mendelian Inheritance: Definitions
 Single Gene (Monogenic) Disorders Mendelian Inheritance: Definitions A genetic locus is a specific position or location on a chromosome. Frequently, locus is used to refer to a specific gene. Alleles are
Single Gene (Monogenic) Disorders Mendelian Inheritance: Definitions A genetic locus is a specific position or location on a chromosome. Frequently, locus is used to refer to a specific gene. Alleles are
Statistical Evaluation of Sibling Relationship
 The Korean Communications in Statistics Vol. 14 No. 3, 2007, pp. 541 549 Statistical Evaluation of Sibling Relationship Jae Won Lee 1), Hye-Seung Lee 2), Hyo Jung Lee 3) and Juck-Joon Hwang 4) Abstract
The Korean Communications in Statistics Vol. 14 No. 3, 2007, pp. 541 549 Statistical Evaluation of Sibling Relationship Jae Won Lee 1), Hye-Seung Lee 2), Hyo Jung Lee 3) and Juck-Joon Hwang 4) Abstract
Role of Genomics in Selection of Beef Cattle for Healthfulness Characteristics
 Role of Genomics in Selection of Beef Cattle for Healthfulness Characteristics Dorian Garrick dorian@iastate.edu Iowa State University & National Beef Cattle Evaluation Consortium Selection and Prediction
Role of Genomics in Selection of Beef Cattle for Healthfulness Characteristics Dorian Garrick dorian@iastate.edu Iowa State University & National Beef Cattle Evaluation Consortium Selection and Prediction
Introduction to Quantitative Genetics
 Introduction to Quantitative Genetics 1 / 17 Historical Background Quantitative genetics is the study of continuous or quantitative traits and their underlying mechanisms. The main principals of quantitative
Introduction to Quantitative Genetics 1 / 17 Historical Background Quantitative genetics is the study of continuous or quantitative traits and their underlying mechanisms. The main principals of quantitative
Chapter 4 PEDIGREE ANALYSIS IN HUMAN GENETICS
 Chapter 4 PEDIGREE ANALYSIS IN HUMAN GENETICS Chapter Summary In order to study the transmission of human genetic traits to the next generation, a different method of operation had to be adopted. Instead
Chapter 4 PEDIGREE ANALYSIS IN HUMAN GENETICS Chapter Summary In order to study the transmission of human genetic traits to the next generation, a different method of operation had to be adopted. Instead
Exam #2 BSC Fall. NAME_Key correct answers in BOLD FORM A
 Exam #2 BSC 2011 2004 Fall NAME_Key correct answers in BOLD FORM A Before you begin, please write your name and social security number on the computerized score sheet. Mark in the corresponding bubbles
Exam #2 BSC 2011 2004 Fall NAME_Key correct answers in BOLD FORM A Before you begin, please write your name and social security number on the computerized score sheet. Mark in the corresponding bubbles
Selection at one locus with many alleles, fertility selection, and sexual selection
 Selection at one locus with many alleles, fertility selection, and sexual selection Introduction It s easy to extend the Hardy-Weinberg principle to multiple alleles at a single locus. In fact, we already
Selection at one locus with many alleles, fertility selection, and sexual selection Introduction It s easy to extend the Hardy-Weinberg principle to multiple alleles at a single locus. In fact, we already
Heritability and genetic correlations explained by common SNPs for MetS traits. Shashaank Vattikuti, Juen Guo and Carson Chow LBM/NIDDK
 Heritability and genetic correlations explained by common SNPs for MetS traits Shashaank Vattikuti, Juen Guo and Carson Chow LBM/NIDDK The Genomewide Association Study. Manolio TA. N Engl J Med 2010;363:166-176.
Heritability and genetic correlations explained by common SNPs for MetS traits Shashaank Vattikuti, Juen Guo and Carson Chow LBM/NIDDK The Genomewide Association Study. Manolio TA. N Engl J Med 2010;363:166-176.
SNPrints: Defining SNP signatures for prediction of onset in complex diseases
 SNPrints: Defining SNP signatures for prediction of onset in complex diseases Linda Liu, Biomedical Informatics, Stanford University Daniel Newburger, Biomedical Informatics, Stanford University Grace
SNPrints: Defining SNP signatures for prediction of onset in complex diseases Linda Liu, Biomedical Informatics, Stanford University Daniel Newburger, Biomedical Informatics, Stanford University Grace
Roadmap. Inbreeding How inbred is a population? What are the consequences of inbreeding?
 1 Roadmap Quantitative traits What kinds of variation can selection work on? How much will a population respond to selection? Heritability How can response be restored? Inbreeding How inbred is a population?
1 Roadmap Quantitative traits What kinds of variation can selection work on? How much will a population respond to selection? Heritability How can response be restored? Inbreeding How inbred is a population?
CS2220 Introduction to Computational Biology
 CS2220 Introduction to Computational Biology WEEK 8: GENOME-WIDE ASSOCIATION STUDIES (GWAS) 1 Dr. Mengling FENG Institute for Infocomm Research Massachusetts Institute of Technology mfeng@mit.edu PLANS
CS2220 Introduction to Computational Biology WEEK 8: GENOME-WIDE ASSOCIATION STUDIES (GWAS) 1 Dr. Mengling FENG Institute for Infocomm Research Massachusetts Institute of Technology mfeng@mit.edu PLANS
Meiosis and Genetics
 Meiosis and Genetics Humans have chromosomes in each cell What pattern do you notice in the human karyotype (a technique that organizes chromosomes by type and size)? Humans are diploid 1 Gametes are produced
Meiosis and Genetics Humans have chromosomes in each cell What pattern do you notice in the human karyotype (a technique that organizes chromosomes by type and size)? Humans are diploid 1 Gametes are produced
Complex Traits Activity INSTRUCTION MANUAL. ANT 2110 Introduction to Physical Anthropology Professor Julie J. Lesnik
 Complex Traits Activity INSTRUCTION MANUAL ANT 2110 Introduction to Physical Anthropology Professor Julie J. Lesnik Introduction Human variation is complex. The simplest form of variation in a population
Complex Traits Activity INSTRUCTION MANUAL ANT 2110 Introduction to Physical Anthropology Professor Julie J. Lesnik Introduction Human variation is complex. The simplest form of variation in a population
Bio 312, Spring 2017 Exam 3 ( 1 ) Name:
 Bio 312, Spring 2017 Exam 3 ( 1 ) Name: Please write the first letter of your last name in the box; 5 points will be deducted if your name is hard to read or the box does not contain the correct letter.
Bio 312, Spring 2017 Exam 3 ( 1 ) Name: Please write the first letter of your last name in the box; 5 points will be deducted if your name is hard to read or the box does not contain the correct letter.
Solutions to Genetics Unit Exam
 Solutions to Genetics Unit Exam Question 1 You are working with an ornamental fish that shows two color phenotypes, red or white. The color is controlled by a single gene. These fish are hermaphrodites
Solutions to Genetics Unit Exam Question 1 You are working with an ornamental fish that shows two color phenotypes, red or white. The color is controlled by a single gene. These fish are hermaphrodites
A Comparison of Sample Size and Power in Case-Only Association Studies of Gene-Environment Interaction
 American Journal of Epidemiology ª The Author 2010. Published by Oxford University Press on behalf of the Johns Hopkins Bloomberg School of Public Health. This is an Open Access article distributed under
American Journal of Epidemiology ª The Author 2010. Published by Oxford University Press on behalf of the Johns Hopkins Bloomberg School of Public Health. This is an Open Access article distributed under
Tutorial on Genome-Wide Association Studies
 Tutorial on Genome-Wide Association Studies Assistant Professor Institute for Computational Biology Department of Epidemiology and Biostatistics Case Western Reserve University Acknowledgements Dana Crawford
Tutorial on Genome-Wide Association Studies Assistant Professor Institute for Computational Biology Department of Epidemiology and Biostatistics Case Western Reserve University Acknowledgements Dana Crawford
The Making of the Fittest: Natural Selection in Humans
 INTRODUCTION MENDELIAN GENETICS, PROBABILITY, PEDIGREES, AND CHI-SQUARE STATISTICS Hemoglobin is a protein found in red blood cells (RBCs) that transports oxygen throughout the body. The hemoglobin protein
INTRODUCTION MENDELIAN GENETICS, PROBABILITY, PEDIGREES, AND CHI-SQUARE STATISTICS Hemoglobin is a protein found in red blood cells (RBCs) that transports oxygen throughout the body. The hemoglobin protein
Lecture 17: Human Genetics. I. Types of Genetic Disorders. A. Single gene disorders
 Lecture 17: Human Genetics I. Types of Genetic Disorders A. Single gene disorders B. Multifactorial traits 1. Mutant alleles at several loci acting in concert C. Chromosomal abnormalities 1. Physical changes
Lecture 17: Human Genetics I. Types of Genetic Disorders A. Single gene disorders B. Multifactorial traits 1. Mutant alleles at several loci acting in concert C. Chromosomal abnormalities 1. Physical changes
CSE 258 Lecture 1.5. Web Mining and Recommender Systems. Supervised learning Regression
 CSE 258 Lecture 1.5 Web Mining and Recommender Systems Supervised learning Regression What is supervised learning? Supervised learning is the process of trying to infer from labeled data the underlying
CSE 258 Lecture 1.5 Web Mining and Recommender Systems Supervised learning Regression What is supervised learning? Supervised learning is the process of trying to infer from labeled data the underlying
A. Incorrect! Cells contain the units of genetic they are not the unit of heredity.
 MCAT Biology Problem Drill PS07: Mendelian Genetics Question No. 1 of 10 Question 1. The smallest unit of heredity is. Question #01 (A) Cell (B) Gene (C) Chromosome (D) Allele Cells contain the units of
MCAT Biology Problem Drill PS07: Mendelian Genetics Question No. 1 of 10 Question 1. The smallest unit of heredity is. Question #01 (A) Cell (B) Gene (C) Chromosome (D) Allele Cells contain the units of
DOES THE BRCAX GENE EXIST? FUTURE OUTLOOK
 CHAPTER 6 DOES THE BRCAX GENE EXIST? FUTURE OUTLOOK Genetic research aimed at the identification of new breast cancer susceptibility genes is at an interesting crossroad. On the one hand, the existence
CHAPTER 6 DOES THE BRCAX GENE EXIST? FUTURE OUTLOOK Genetic research aimed at the identification of new breast cancer susceptibility genes is at an interesting crossroad. On the one hand, the existence
BIOL 364 Population Biology Fairly testing the theory of evolution by natural selection with playing cards
 BIOL 364 Population Biology Fairly testing the theory of evolution by natural selection with playing cards Game I: The Basics Scenario: Our classroom is now a closed population (no immigration or emigration)
BIOL 364 Population Biology Fairly testing the theory of evolution by natural selection with playing cards Game I: The Basics Scenario: Our classroom is now a closed population (no immigration or emigration)
12 MENDEL, GENES, AND INHERITANCE
 12 MENDEL, GENES, AND INHERITANCE Chapter Outline 12.1 THE BEGINNINGS OF GENETICS: MENDEL S GARDEN PEAS Mendel chose true-breeding garden peas for his experiments Mendel first worked with single-character
12 MENDEL, GENES, AND INHERITANCE Chapter Outline 12.1 THE BEGINNINGS OF GENETICS: MENDEL S GARDEN PEAS Mendel chose true-breeding garden peas for his experiments Mendel first worked with single-character
Mendelism: the Basic principles of Inheritance
 Chapter 3. Mendelism: the Basic principles of Inheritance 1. Mendel s Study of Heredity 2. Applications of Mendel s Principles 3. Formulating and Testing Genetic Hypothesis 4. Mendelian Principles in Human
Chapter 3. Mendelism: the Basic principles of Inheritance 1. Mendel s Study of Heredity 2. Applications of Mendel s Principles 3. Formulating and Testing Genetic Hypothesis 4. Mendelian Principles in Human
Gregor Mendel and Genetics Worksheets
 Gregor Mendel and Genetics Worksheets Douglas Wilkin, Ph.D. (DWilkin) Say Thanks to the Authors Click http://www.ck12.org/saythanks (No sign in required) To access a customizable version of this book,
Gregor Mendel and Genetics Worksheets Douglas Wilkin, Ph.D. (DWilkin) Say Thanks to the Authors Click http://www.ck12.org/saythanks (No sign in required) To access a customizable version of this book,
UNIT 3 GENETICS LESSON #30: TRAITS, GENES, & ALLELES. Many things come in many forms. Give me an example of something that comes in many forms.
 UNIT 3 GENETICS LESSON #30: TRAITS, GENES, & ALLELES Many things come in many forms. Give me an example of something that comes in many forms. Genes, too, come in many forms. Main Idea #1 The same gene
UNIT 3 GENETICS LESSON #30: TRAITS, GENES, & ALLELES Many things come in many forms. Give me an example of something that comes in many forms. Genes, too, come in many forms. Main Idea #1 The same gene
Handling Immunogenetic Data Managing and Validating HLA Data
 Handling Immunogenetic Data Managing and Validating HLA Data Steven J. Mack PhD Children s Hospital Oakland Research Institute 16 th IHIW & Joint Conference Sunday 3 June, 2012 Overview 1. Master Analytical
Handling Immunogenetic Data Managing and Validating HLA Data Steven J. Mack PhD Children s Hospital Oakland Research Institute 16 th IHIW & Joint Conference Sunday 3 June, 2012 Overview 1. Master Analytical
Welcome Back! 2/6/18. A. GGSS B. ggss C. ggss D. GgSs E. Ggss. 1. A species of mice can have gray or black fur
 Welcome Back! 2/6/18 1. A species of mice can have gray or black fur and long or short tails. A cross between blackfurred, long-tailed mice and gray-furred, shorttailed mice produce all black-furred, long-tailed
Welcome Back! 2/6/18 1. A species of mice can have gray or black fur and long or short tails. A cross between blackfurred, long-tailed mice and gray-furred, shorttailed mice produce all black-furred, long-tailed
additive genetic component [d] = rded
![additive genetic component [d] = rded additive genetic component [d] = rded](/thumbs/92/108220661.jpg) Heredity (1976), 36 (1), 31-40 EFFECT OF GENE DISPERSION ON ESTIMATES OF COMPONENTS OF GENERATION MEANS AND VARIANCES N. E. M. JAYASEKARA* and J. L. JINKS Department of Genetics, University of Birmingham,
Heredity (1976), 36 (1), 31-40 EFFECT OF GENE DISPERSION ON ESTIMATES OF COMPONENTS OF GENERATION MEANS AND VARIANCES N. E. M. JAYASEKARA* and J. L. JINKS Department of Genetics, University of Birmingham,
New Enhancements: GWAS Workflows with SVS
 New Enhancements: GWAS Workflows with SVS August 9 th, 2017 Gabe Rudy VP Product & Engineering 20 most promising Biotech Technology Providers Top 10 Analytics Solution Providers Hype Cycle for Life sciences
New Enhancements: GWAS Workflows with SVS August 9 th, 2017 Gabe Rudy VP Product & Engineering 20 most promising Biotech Technology Providers Top 10 Analytics Solution Providers Hype Cycle for Life sciences
Lesson Overview 11.2 Applying Mendel s Principles
 THINK ABOUT IT Nothing in life is certain. Lesson Overview 11.2 Applying Mendel s Principles If a parent carries two different alleles for a certain gene, we can t be sure which of those alleles will be
THINK ABOUT IT Nothing in life is certain. Lesson Overview 11.2 Applying Mendel s Principles If a parent carries two different alleles for a certain gene, we can t be sure which of those alleles will be
HERITABILITY INTRODUCTION. Objectives
 36 HERITABILITY In collaboration with Mary Puterbaugh and Larry Lawson Objectives Understand the concept of heritability. Differentiate between broad-sense heritability and narrowsense heritability. Learn
36 HERITABILITY In collaboration with Mary Puterbaugh and Larry Lawson Objectives Understand the concept of heritability. Differentiate between broad-sense heritability and narrowsense heritability. Learn
Review Statistics review 11: Assessing risk Viv Bewick 1, Liz Cheek 1 and Jonathan Ball 2
 Available online http://ccforum.com/content/8/4/87 Review Statistics review 11: Assessing risk Viv Bewick 1, Liz Cheek 1 and Jonathan Ball 1 Senior Lecturer, School of Computing, Mathematical and Information
Available online http://ccforum.com/content/8/4/87 Review Statistics review 11: Assessing risk Viv Bewick 1, Liz Cheek 1 and Jonathan Ball 1 Senior Lecturer, School of Computing, Mathematical and Information
Title:Validation study of candidate single nucleotide polymorphisms associated with left ventricular hypertrophy in the Korean population
 Author's response to reviews Title:Validation study of candidate single nucleotide polymorphisms associated with left ventricular hypertrophy in the Korean population Authors: Jin-Kyu Park (cardiohy@gmail.com)
Author's response to reviews Title:Validation study of candidate single nucleotide polymorphisms associated with left ventricular hypertrophy in the Korean population Authors: Jin-Kyu Park (cardiohy@gmail.com)
META-ANALYSIS OF THE EFFECTS OF ALCOHOL DEHYDROGENASE GENOTYPE ON ALCOHOL DEPENDENCE AND ALCOHOLIC LIVER DISEASE
 Alcohol & Alcoholism Vol. 32, No. 5, pp. 613-619, 1997 META-ANALYSIS OF THE EFFECTS OF ALCOHOL DEHYDROGENASE GENOTYPE ON ALCOHOL DEPENDENCE AND ALCOHOLIC LIVER DISEASE J. B. WHITFIELD Department of Clinical
Alcohol & Alcoholism Vol. 32, No. 5, pp. 613-619, 1997 META-ANALYSIS OF THE EFFECTS OF ALCOHOL DEHYDROGENASE GENOTYPE ON ALCOHOL DEPENDENCE AND ALCOHOLIC LIVER DISEASE J. B. WHITFIELD Department of Clinical
DEFINITIONS: POPULATION: a localized group of individuals belonging to the same species
 DEFINITIONS: POPULATION: a localized group of individuals belonging to the same species SPECIES: a group of populations whose individuals have the potential to interbreed and produce fertile offspring
DEFINITIONS: POPULATION: a localized group of individuals belonging to the same species SPECIES: a group of populations whose individuals have the potential to interbreed and produce fertile offspring
The Making of the Fittest: Natural Selection in Humans
 MENDELIAN GENETICS, PROBABILITY, PEDIGREES, AND CHI-SQUARE STATISTICS INTRODUCTION Hemoglobin is a protein found in red blood cells that transports oxygen throughout the body. The hemoglobin protein consists
MENDELIAN GENETICS, PROBABILITY, PEDIGREES, AND CHI-SQUARE STATISTICS INTRODUCTION Hemoglobin is a protein found in red blood cells that transports oxygen throughout the body. The hemoglobin protein consists
Human population sub-structure and genetic association studies
 Human population sub-structure and genetic association studies Stephanie A. Santorico, Ph.D. Department of Mathematical & Statistical Sciences Stephanie.Santorico@ucdenver.edu Global Similarity Map from
Human population sub-structure and genetic association studies Stephanie A. Santorico, Ph.D. Department of Mathematical & Statistical Sciences Stephanie.Santorico@ucdenver.edu Global Similarity Map from
breast cancer; relative risk; risk factor; standard deviation; strength of association
 American Journal of Epidemiology The Author 2015. Published by Oxford University Press on behalf of the Johns Hopkins Bloomberg School of Public Health. All rights reserved. For permissions, please e-mail:
American Journal of Epidemiology The Author 2015. Published by Oxford University Press on behalf of the Johns Hopkins Bloomberg School of Public Health. All rights reserved. For permissions, please e-mail:
Quantitative Methods in Managment. An introduction to GLMs and measurement theory
 Quantitative Methods in Managment An Introduction to GLMs and measurement theory Graeme D. Hutcheson 1 Luiz Moutinho 2 1 School of Education Manchester university 2 Department of Management Glasgow University
Quantitative Methods in Managment An Introduction to GLMs and measurement theory Graeme D. Hutcheson 1 Luiz Moutinho 2 1 School of Education Manchester university 2 Department of Management Glasgow University
How Populations Evolve
 Chapter 16: pp. 283-298 BIOLOGY 10th Edition How Populations Evolve 10% of population Copyright The McGraw-Hill Companies, Inc. Permission required for reproduction or display. natural disaster kills five
Chapter 16: pp. 283-298 BIOLOGY 10th Edition How Populations Evolve 10% of population Copyright The McGraw-Hill Companies, Inc. Permission required for reproduction or display. natural disaster kills five
Genetic association analysis incorporating intermediate phenotypes information for complex diseases
 University of Iowa Iowa Research Online Theses and Dissertations Fall 2011 Genetic association analysis incorporating intermediate phenotypes information for complex diseases Yafang Li University of Iowa
University of Iowa Iowa Research Online Theses and Dissertations Fall 2011 Genetic association analysis incorporating intermediate phenotypes information for complex diseases Yafang Li University of Iowa
Essential Skills for Evidence-based Practice: Statistics for Therapy Questions
 Essential Skills for Evidence-based Practice: Statistics for Therapy Questions Jeanne Grace Corresponding author: J. Grace E-mail: Jeanne_Grace@urmc.rochester.edu Jeanne Grace RN PhD Emeritus Clinical
Essential Skills for Evidence-based Practice: Statistics for Therapy Questions Jeanne Grace Corresponding author: J. Grace E-mail: Jeanne_Grace@urmc.rochester.edu Jeanne Grace RN PhD Emeritus Clinical
Sexual Reproduction and Genetics. Section 1. Meiosis
 Chromosomes and Chromosome Number! Human body cells have 46 chromosomes! Each parent contributes 23 chromosomes! Homologous chromosomes one of two paired chromosomes, one from each parent Chromosomes and
Chromosomes and Chromosome Number! Human body cells have 46 chromosomes! Each parent contributes 23 chromosomes! Homologous chromosomes one of two paired chromosomes, one from each parent Chromosomes and
Modelling Research Productivity Using a Generalization of the Ordered Logistic Regression Model
 Modelling Research Productivity Using a Generalization of the Ordered Logistic Regression Model Delia North Temesgen Zewotir Michael Murray Abstract In South Africa, the Department of Education allocates
Modelling Research Productivity Using a Generalization of the Ordered Logistic Regression Model Delia North Temesgen Zewotir Michael Murray Abstract In South Africa, the Department of Education allocates
Comparison of Linkage-Disequilibrium Methods for Localization of Genes Influencing Quantitative Traits in Humans
 Am. J. Hum. Genet. 64:1194 105, 1999 Comparison of Linkage-Disequilibrium Methods for Localization of Genes Influencing Quantitative Traits in Humans Grier P. Page and Christopher I. Amos Department of
Am. J. Hum. Genet. 64:1194 105, 1999 Comparison of Linkage-Disequilibrium Methods for Localization of Genes Influencing Quantitative Traits in Humans Grier P. Page and Christopher I. Amos Department of
Punne% Square Quiz A AP Tes2ng this week 15-Week Grades due next week Note: media center is hos2ng tes2ng Turn in all make-up work
 Biology Monday, May 2, 2016 Do-Now: Punne% Square Quiz A 1. Write down today s FLT 2. What do we use Punne@ Squares for? 3. A purple flower (Pp) and a white flower are crossed. What % of the offspring
Biology Monday, May 2, 2016 Do-Now: Punne% Square Quiz A 1. Write down today s FLT 2. What do we use Punne@ Squares for? 3. A purple flower (Pp) and a white flower are crossed. What % of the offspring
INTERACTION BETWEEN NATURAL SELECTION FOR HETEROZYGOTES AND DIRECTIONAL SELECTION
 INTERACTION BETWEEN NATURAL SELECTION FOR HETEROZYGOTES AND DIRECTIONAL SELECTION MARGRITH WEHRLI VERGHESE 1228 Kingston Ridge Driue, Cary, N.C. 27511 Manuscript received May 5, 1973 Revised copy received
INTERACTION BETWEEN NATURAL SELECTION FOR HETEROZYGOTES AND DIRECTIONAL SELECTION MARGRITH WEHRLI VERGHESE 1228 Kingston Ridge Driue, Cary, N.C. 27511 Manuscript received May 5, 1973 Revised copy received
NOTES: Exceptions to Mendelian Genetics!
 NOTES: 11.3 Exceptions to Mendelian Genetics! Beyond Dominant and Recessive Alleles Some alleles are neither dominant nor recessive, and many traits are controlled by multiple alleles OR multiple genes.
NOTES: 11.3 Exceptions to Mendelian Genetics! Beyond Dominant and Recessive Alleles Some alleles are neither dominant nor recessive, and many traits are controlled by multiple alleles OR multiple genes.
Ct=28.4 WAT 92.6% Hepatic CE (mg/g) P=3.6x10-08 Plasma Cholesterol (mg/dl)
 a Control AAV mtm6sf-shrna8 Ct=4.3 Ct=8.4 Ct=8.8 Ct=8.9 Ct=.8 Ct=.5 Relative TM6SF mrna Level P=.5 X -5 b.5 Liver WAT Small intestine Relative TM6SF mrna Level..5 9.6% Control AAV mtm6sf-shrna mtm6sf-shrna6
a Control AAV mtm6sf-shrna8 Ct=4.3 Ct=8.4 Ct=8.8 Ct=8.9 Ct=.8 Ct=.5 Relative TM6SF mrna Level P=.5 X -5 b.5 Liver WAT Small intestine Relative TM6SF mrna Level..5 9.6% Control AAV mtm6sf-shrna mtm6sf-shrna6
When bad things happen to good genes: mutation vs. selection
 When bad things happen to good genes: mutation vs. selection Selection tends to increase the frequencies of alleles with higher marginal fitnesses. Does this mean that genes are perfect? No, mutation can
When bad things happen to good genes: mutation vs. selection Selection tends to increase the frequencies of alleles with higher marginal fitnesses. Does this mean that genes are perfect? No, mutation can
Dan Koller, Ph.D. Medical and Molecular Genetics
 Design of Genetic Studies Dan Koller, Ph.D. Research Assistant Professor Medical and Molecular Genetics Genetics and Medicine Over the past decade, advances from genetics have permeated medicine Identification
Design of Genetic Studies Dan Koller, Ph.D. Research Assistant Professor Medical and Molecular Genetics Genetics and Medicine Over the past decade, advances from genetics have permeated medicine Identification
Activities to Accompany the Genetics and Evolution App for ipad and iphone
 Activities to Accompany the Genetics and Evolution App for ipad and iphone All of the following questions can be answered using the ipad version of the Genetics and Evolution App. When using the iphone
Activities to Accompany the Genetics and Evolution App for ipad and iphone All of the following questions can be answered using the ipad version of the Genetics and Evolution App. When using the iphone
Bio 1M: Evolutionary processes
 Bio 1M: Evolutionary processes Evolution by natural selection Is something missing from the story I told last chapter? Heritable variation in traits Selection (i.e., differential reproductive success)
Bio 1M: Evolutionary processes Evolution by natural selection Is something missing from the story I told last chapter? Heritable variation in traits Selection (i.e., differential reproductive success)
Will now consider in detail the effects of relaxing the assumption of infinite-population size.
 FINITE POPULATION SIZE: GENETIC DRIFT READING: Nielsen & Slatkin pp. 21-27 Will now consider in detail the effects of relaxing the assumption of infinite-population size. Start with an extreme case: a
FINITE POPULATION SIZE: GENETIC DRIFT READING: Nielsen & Slatkin pp. 21-27 Will now consider in detail the effects of relaxing the assumption of infinite-population size. Start with an extreme case: a
Effects of age-at-diagnosis and duration of diabetes on GADA and IA-2A positivity
 Effects of age-at-diagnosis and duration of diabetes on GADA and IA-2A positivity Duration of diabetes was inversely correlated with age-at-diagnosis (ρ=-0.13). However, as backward stepwise regression
Effects of age-at-diagnosis and duration of diabetes on GADA and IA-2A positivity Duration of diabetes was inversely correlated with age-at-diagnosis (ρ=-0.13). However, as backward stepwise regression
MULTIPLE LINEAR REGRESSION 24.1 INTRODUCTION AND OBJECTIVES OBJECTIVES
 24 MULTIPLE LINEAR REGRESSION 24.1 INTRODUCTION AND OBJECTIVES In the previous chapter, simple linear regression was used when you have one independent variable and one dependent variable. This chapter
24 MULTIPLE LINEAR REGRESSION 24.1 INTRODUCTION AND OBJECTIVES In the previous chapter, simple linear regression was used when you have one independent variable and one dependent variable. This chapter
Pedigree Analysis. A = the trait (a genetic disease or abnormality, dominant) a = normal (recessive)
 Pedigree Analysis Introduction A pedigree is a diagram of family relationships that uses symbols to represent people and lines to represent genetic relationships. These diagrams make it easier to visualize
Pedigree Analysis Introduction A pedigree is a diagram of family relationships that uses symbols to represent people and lines to represent genetic relationships. These diagrams make it easier to visualize
Your Vocabulary words-- write into your journal:
 HUMAN INHERITANCE Your Vocabulary words-- write into your journal: 1. Multiple alleles: three or more forms of a gene that code for a single trait. 2. Sex chromosomes: these carry genes that determine
HUMAN INHERITANCE Your Vocabulary words-- write into your journal: 1. Multiple alleles: three or more forms of a gene that code for a single trait. 2. Sex chromosomes: these carry genes that determine
The laws of Heredity. Allele: is the copy (or a version) of the gene that control the same characteristics.
 The laws of Heredity 1. Definition: Heredity: The passing of traits from parents to their offspring by means of the genes from the parents. Gene: Part or portion of a chromosome that carries genetic information
The laws of Heredity 1. Definition: Heredity: The passing of traits from parents to their offspring by means of the genes from the parents. Gene: Part or portion of a chromosome that carries genetic information
4. A homozygous tall plant and a heterozygous tall plant are crossed. What is the percent probability of short offspring?
 LEVEL ZERO VOICE POP QUIZ (4 minutes, individual work): 1. Define gene: 2. Define phenotype: 3. A heterozygous white rabbit is crossed with a homozygous black rabbit. If they have 160 offspring, how many
LEVEL ZERO VOICE POP QUIZ (4 minutes, individual work): 1. Define gene: 2. Define phenotype: 3. A heterozygous white rabbit is crossed with a homozygous black rabbit. If they have 160 offspring, how many
Model of an F 1 and F 2 generation
 Mendelian Genetics Casual observation of a population of organisms (e.g. cats) will show variation in many visible characteristics (e.g. color of fur). While members of a species will have the same number
Mendelian Genetics Casual observation of a population of organisms (e.g. cats) will show variation in many visible characteristics (e.g. color of fur). While members of a species will have the same number
