Insilico Docking of Various Inhibitors of E.Faecalis Folate Pathway
|
|
|
- Molly Washington
- 6 years ago
- Views:
Transcription
1 International Journal of Scientific and Research Publications, Volume 7, Issue 3, March ISSN Insilico Docking of Various Inhibitors of E.Faecalis Folate Pathway ARCHANA MOON 1*, DEEBA KHAN 2*, PRANJALI GAJBHIYE 3* & MONALI JARIYA 4* 1,2,3&4 University Department of Biochemistry, Rashtrasant Tukadoji Maharaj Nagpur University, Nagpur *, Professor, University Department of Biochemistry, Rashtrasant Tukadoji Maharaj Nagpur University, Nagpur , Contact number: *, Project fellow, University Department of Biochemistry, Rashtrasant Tukadoji Maharaj Nagpur University, Nagpur , Contact number: Abstract Drug resistance to therapeutic antibiotics pose a challenge to the identification of novel targets and drugs for the treatment of infectious diseases. Infections caused by Enterococcus faecalis are a major health problem. Moreover, among UTI causing enterococci, multidrug resistant E. faecalis such as vancomycinresistant strains (VRE) have been reported increasingly in many countries. SMX & TMP are the commonly prescribed inhibitors of DHFR & DHPS, the enzymes of the folate biosynthetic pathway. In this study, insilico docking of various ligands/inhibitors to the Enterococcus faecalis DHFR & DHPS proteins has been performed by using Autodock Suite, version 1.5 6rC2. Index Terms Dihydrofolate reductase (DHFR), Dihydropteroate synthase (DHPS), Docking, Folate Pathway Inhibitors. M I. INTRODUCTION ost clinical isolates from urine samples of UTI patients show presence of Enterococcus faecalis and account for 80 90% of clinical strains. E. faecium accounts for the remaining 5 10% of such isolates (4). Enterococci currently ranks fourth in frequency among bacteria isolated from hospitalized patients (4). They are nosocomial pathogens and are associated with high mortality. The treatment of these infections pose a great challenge, due to the inherent resistance of Enterococci to many antibiotics (1). In addition, they have the capacity to easily acquire and express new resistance genes and can thus tolerate antibiotic selective pressure (2). Enterococci are Grampositive ubiquitous bacteria that are widely found in all types of animals and in the environment. They are typically harmless inhabitants of various body sites particularly the intestinal tract. However, enterococci are opportunistic pathogens (3). In addition, enterococci are inherently resistant to many antimicrobials, including penicillin, clindamycin, trimethoprim sulfamethoxazole, and low levels of aminoglycosides, and they are poorly responsive to cephalosporins and fluoroquinolones in vivo (3). They were traditionally regarded as low grade pathogens but have emerged as second leading cause of nosocomial infections and third most common cause of bacteremia. The most frequent infections caused by enterococci are UTI, endocarditis, bacteremia, intraabdominal and intrapelvic abscesses (5,6). This is amplified due to their acquired resistance to all currently available antibiotics that leaves the clinicians with limited treatment options and results in the selection and spreading of multidrugresistant (MDR) strains in hospitals (7). Trimethoprim (TMP) and sulfamethoxazole (SMX) are inhibitors of bacterial enzymes involved in the folate synthesis pathway. Folic acid is necessary to carry out a variety of important cellular functions, including synthesis of nucleic acids, particularly thymidine. Most bacteria are unable to take up exogenous folate from the environment and instead must synthesize it from the pamino benzoic acid precursor (Refer Fig:1). TMP and SMX inhibit successive enzymes in this pathway, limiting the production of dihydrofolate and its subsequent conversion to tetrahydrofolate (8). Use of computational methods is a cost effective strategy for speeding up the process of drug discovery and development process. Hence, understanding binding interactions between receptor and ligand is very essential for drug discovery scientists (9). Molecular docking, a computational method of studying binding interactions in terms of binding energies is immensely used in the process of drug discovery to save on cost and time. In this method, computer generated representation of a small molecule or ligand is placed into the active site of the target or protein s computational structure in a variety of positions, conformations and orientations. The position, orientation and conformation of the ligand in the active site of protein is called as a pose. In order to identify the energetically most favorable pose, each pose of the ligand is evaluated for binding energy computationally. The main objective of molecular docking method is to find a pose which has the lowest binding energy (9). AutoDock abbreviated as AD, is an automated suite of proteinligand docking tools. It is designed to predict the protein interactions with small molecules such as drug molecule and substrate. The application of this tool is immense, ranging from structure based drug design, lead molecule optimisation, proteinligand docking, proteinprotein docking, analysis and validation of mechanism of action of drug molecules, etc., AutoDock has two versions, namely, AutoDock4 and AutoDock Vina. The prior has been used in this study, AutoDock4 analyzes the interactions of ligand molecules at the specified target site of the protein. The users can define this specific target sites with the use of GridBox. AutoDock4 has two executable key programs, i.e., Autogrid4 and AutoDock4. Autogrid4 prepares a grid map of the amino acids presents within the GridBox defined by the user. AutoDock4
2 International Journal of Scientific and Research Publications, Volume 7, Issue 3, March ISSN then analyzes the interactions of those amino acids with the ligand molecule (10). Fig(1): FOLATE PATHWAY IN BACTERIA. II. MATERIALS & METHODS In silico studies In silico studies utilizing molecular docking is an important tool to study the interaction of ligands with active site residues of the receptor (11, 12). The docking involves the use of sampling algorithm and a scoring function to evaluate the proper orientation and pose of ligand molecule in relation to the binding energy. The correct identification of this binding pose of one or more related ligands is important in establishing a structureactivity relationship in lead optimization. The second use of scoring functions is to rank different ligands to predict their relative experimental activity (1214). In silico studies were performed using Autodock 4 suite (version 1.5 6rC2). The ligands viz., Chlorogenic acid, Ellagic acid, Gallic Acid, Hippuric acid, Quercetin and Standard antibiotics viz., Clavulanic acid, Cephalosporin, Cephalosporin C, Penicillin, Sulfamethoxazole and Trimethoprim were docked with DHFR & DHPS enzymes of E.faecalis. The ligands and the Standard antibiotics were selected on the basis of reported antibacterial activity and prescribed drugs. The DHFR (PDB Id 4M7U) protein structure of E.faecalis was downloaded from PDB ( X ray diffraction of 2.1A 0 ). The crystal structure of DHPS E.faecalis is unavailable in PDB hence, to obtain structural information of DHPS, the homology model was generated using Swiss model and PdbSum. DHPS E.faecalis protein was found to have 40.86% sequence identities with PDB ID: W767. This structure was used for the preparation of the model of DHPS of E. faecalis. The prepared model was further validated by Ramachandran plot with the help of PROCHECK. This plot verified the DHPS protein (W767) and hence was used for docking studies Fig (2). These models were further used to analyse and compare the effect of binding efficiency of DHPS towards commonly prescribed antibiotics as well as various inhibitors (15). Next, the PubSum database yielded the ligands with their Ligplots. Ligplots give interacting sites of the DHFR Fig (5) & DHPS Fig (4). Fig (5A& B) depicts the Ramachandran Plots of DHFR and DHPS E.faecalis respectively. The structure of ligands were downloaded from Pubchem (chemical structure data base) online portal and drawn in Marvin Sketch version Fig (6,7). After docking, the results were analyzed on the basis of their binding energy and their interactions (15).
3 International Journal of Scientific and Research Publications, Volume 7, Issue 3, March ISSN Fig(2): PDB STUCTURE OF DHFR AND DHPS PROTEIN OF E.FAECALIS. Fig(3): Ligplot of DHFR E.faecalis Ligand: Ligand Nap201(A) (Ser65, Thr64,, Glu105, Val101, Ser100, Thr46, ) interaction are shown by green dashed line.
4 International Journal of Scientific and Research Publications, Volume 7, Issue 3, March ISSN Fig(4): Ligplot of DHPS E.faecalis Ligand: Ligand HH2 1(_) ( Ser54, His244, Arg242, Lys207, Asp171, Asn108,, Ser20,, ) interaction are shown by green dashed line. Fig (5): The Ramachandran plot shows the phipsi torsion angles for all residues in the structure. Glycine residues are separately identified by triangles as these are not restricted to the regions of the plot appropriate to the other sidechain types. The colouring/shading on the plot represents the different regions: the darkest areas (here shown in red) correspond to the "core" regions representing the most favourable combinations of phipsi values.
5 International Journal of Scientific and Research Publications, Volume 7, Issue 3, March ISSN Fig(6): STRUCTURES OF INHIBITORS (A) Chlorogenic acid,(b) Ellagic acid, (C) Gallic acid, (D) Hippuric acid and (E) Quercetin. Fig(7) : STRUCTURES OF ANTIBIOTICS: (A)Clavulanic acid, (B) Cephalosporin,(C) CephalosporinC, (D) Penicillin, (E) Sulfamethoxazole and (F) Trimethoprim. Preparation of Proteins and Ligands: The ligands viz., Chlorogenic acid, Ellagic acid, Gallic Acid, Hippuric acid and Quercetin and Standard Antibiotics viz., Clavulanic acid, Cephalosporin, Cephalosporin C, Penicillin, Sulfamethoxazole and Trimethoprim that have exhibited prominent antibacterial activity towards isolated multidrugresistant bacteria and have been reported were selected for molecular docking analysis (15). The structures of DHFR & DHPS were opened in Biovia Discovery Studio 2016 version The structure of protein was cleared (i.e. the extra groups which includes water molecules, ligand groups were removed) by deleting the heteroatoms present in the protein (16). Only the protein and active site for docking is required, hence was saved in the PDB format. The structure of ligands were downloaded from Pubchem and drawn in Marvin Sketch view version and cleaned in 2D and 3D. This cleared the 2 dimensional and 3 dimensional structure of the ligand. For
6 International Journal of Scientific and Research Publications, Volume 7, Issue 3, March ISSN docking, the protein structure was obtained in PDB format and ligands in triposmol format or PDB format (16). Grid formation by Autodock Grid points generate the coordinates or interaction points where the ligand is docked. The grid box was generated at 60x60x60 A 0 to cover all the active site residues, and allowed the flexible rotation of ligands. The GA (genetic algorithm) and number of generation were set to 10 and for DHFR and DHPS respectively. The Lamarckian genetic algorithm was followed for ligand confirmation. All the above parameters decide the different confirmation of ligand in which the ligand will be docked. Other parameters for example, free energy (after docking is complete we get the value of free energy), rotatable bonds (number of rotatable bonds varies according to the ligand structure), number of torsions (16) etc were used as default (16). RESULT: Docking studies revealed the interaction of the protein with the ligands, w.r.t binding energy, type of interaction and amino acids involved in interactions. Binding energy should be ideally negative. More negative the binding energy, better the binding affinity of ligand and protein (16). Table 1 & 2 give the binding energy of ligands with DHFR & DHPS proteins respectively with inhibitors viz., Chlorogenic acid, Ellagic acid, Gallic Acid, Hippuric acid and Quercetin and Standard Antibiotics viz., Clavulanic acid, Cephalosporin, Cephalosporin C, Penicillin, Sulfamethoxazole and Trimethoprim. TABLE NO.I: BINDING ENERGY OF LIGANDS UPON DOCKING WITH DHFR PROTEIN OF E.FAECALIS. LIGANDS Binding Energy INHIBITORS Chlorogenic acid 6.68 Ellagic acid 7.47 Gallic acid 5.19 Hippuric acid 6.03 Quercetin 7.47 ANTIBIOTICS Clavulanic acid 5.43 Cephalosporin 8.26 Cephalosporin C 7.54 Penicillin 8.35 Sulfamethoxazole 7.67 Trimethoprim 6.38 Inhibitors (Chlorogenic acid, Ellagic acid, Gallic Acid, Hippuric acid and Quercetin) and Standard antibiotics (Clavulanic acid, Cephalosporin, Cephalosporin C, Penicillin, Sulfamethoxazole and Trimethoprim) were docked and the results obtained provide a comparative insight into the potency of inhibitors and standard antibiotics through analysis of their binding capacities. The binding energies of DHFR of E.faecalis with Penicillin, Cephalosporin, Ellagic acid & Quercetin show highest binding than other ligands. Table 1 clearly shows that the standard antibiotics viz., Penicillin and Cephalosporin are more effective than the inhibitors docked, but E.faecalis has emerged resistant to these antibiotics (15). TABLE NO.II: BINDING ENERGY OF LIGANDS UPON DOCKING WITH DHPS PROTEIN OF E.FAECALIS LIGANDS Binding Energy INHIBITORS Chlorogenic acid 7.93 Ellagic acid 6.84 Gallic acid 5.81 Hippuric acid 6.66 Quercetin 8.22 ANTIBIOTICS Clavulanic acid 6.77 Cephalosporin 9.31 Cephalosporin C 7.61 Penicillin 8.68 Sulfamethoxazole 7.77 Trimethoprim 7.58 Inhibitors (Chlorogenic acid, Ellagic acid, Gallic Acid, Hippuric acid and Quercetin) and standard antibiotics (Clavulanic acid, Cephalosporin, Cephalosporin C, Penicillin, Sulfamethoxazole and Trimethoprim) were docked and the results obtained provide a comparative insight into the potency of inhibitors and standard antibiotics through analysis of their binding capacities. The binding energies of DHPS protein of E.faecalis with Cephalosporin, Penicillin, Quercetin & Chlorogenic acid are showing highest binding than other ligands. From Table 2, it is clear that standard antibiotics viz., Cephalosporin and Penicillin are more effective than the inhibitors i.e Chlorogenic acid and Quercetin. Table 3 shows the interaction of various ligands with DHFR & DHPS i.e. hydrogen bond length, hydrogen bond name and amino acid involved in the interaction. As these ligands have proven antibacterial ( Sulfamethoxazole and Trimethoprim (20), Clavulanic Acid (21), Penicillin (22) Cephalosporin (25), Cephalosporin C (24) ) antimicrobial ( Gallic Acid (23) ) and anticancer activities ( Chlorogenic acid (19), Quercetin (19), Ellagic acid (18) and Gallic acid (23) ) these ligands can be further used as lead compounds in treatment of multidrug resistant urinary tract infection caused by E.faecalis. The interacting sites of DHFR & DHPS inhibitors and standard antibiotics matches with Ligplots of both DHFR and DHPS are shown in Fig 8, 9, 10 and 11 respectively.
7 International Journal of Scientific and Research Publications, Volume 7, Issue 3, March ISSN TABLE NO.III: INTERACTIONS OF LIGANDS WITH DHFR AND DHPS PROTEINS OF E.FAECALIS LIGANDS DHFR E.faecalis DHPS E.faecalis Hydrogen Bond Hydrogen Interacting Hydrogen Bond Length In A Bond Name Sites Length In A Chlorogenic acid Thr Gly18 Ser Ellagic acid Gallic acid Hippuric acid Quercetin Clavulanic acid Cephalosporin Cephalosporin C Ser65 Val101 Val102 Ala45 Thr46 Ala45 Ser100 Val101 Ala45 Ala45 Val80 Thr46 Ser65 Ser65 Glu105 Ser65 Thr46 Ser100 Thr64 Glu105 Ser100 Ser65 Glu105 Glu105 Thr64 Thr46 Glu105 Ser100 Penicillin Ser65 Thr64 Ser100 Sulfamethoxazole Trimethoprim Ser100 Val102 Val101 Thr64 Glu105 Hydrogen Bond Name Interacting Sites Ser54 Asp171 Arg Lys207 Arg242 Arg242 Arg208 Lys207 Ser20 Ser20 Arg208 Arg Lys Lys207 Arg242 Arg242 Lys207 Arg242
8 International Journal of Scientific and Research Publications, Volume 7, Issue 3, March ISSN Fig (8): Interacting Sites of DHFR Protein of E.faecalis with Inhibitors i.e. (A) Chlorogenic acid,(b) Ellagic acid, (C) Gallic acid, (D) Hippuric acid and(e) Quercetin.
9 International Journal of Scientific and Research Publications, Volume 7, Issue 3, March ISSN Fig(9): Interacting Sites of DHFR Protein of E.faecalis with Antibiotics i.e. (A)Clavulanic acid, (B) Cephalosporin,(C) CephalosporinC, (D) Penicillin, (E) Sulfamethoxazole and (F) Trimethoprim.
10 International Journal of Scientific and Research Publications, Volume 7, Issue 3, March ISSN Fig(10): Interacting Sites of DHPS Protein of E.faecalis with Inhibitors i.e. (A) Chlorogenic acid,(b) Ellagic acid, (C) Gallic acid, (D) Hippuric acid and (E) Quercetin.
11 International Journal of Scientific and Research Publications, Volume 7, Issue 3, March ISSN Fig(11): Interacting Sites of DHFR Protein of E.faecalis with Antibiotics i.e. (A)Clavulanic acid, (B) Cephalosporin,(C) CephalosporinC, (D) Penicillin, (E) Sulfamethoxazole and (F) Trimethoprim. III. DISCUSSION (UTIs) are the most common infections caused by Enterococcus faecalis. Little is known about the bacterial factors necessary for E. faecalis to cause infections in general, and even less has been reported related to the urinary tract. Many researchers have exposed the emergence of multidrug resistance in Enterococci faecalis to all clinically useful antibiotics (15, 17). Insilico studies with DHFR & DHPS showed a high binding affinity towards the ligands viz., Chlorogenic acid, Ellagic acid, Gallic Acid, Hippuric acid and Quercetin and as well as the prescribed standard antibiotics viz., Clavulanic acid, Cephalosporin, Cephalosporin C, Penicillin, Sulfamethoxazole and Trimethoprim. Molecular docking has been carried out to check the efficiency of these ligands and Standard antibiotics to bind to the active site of the DHFR protein of the folate pathway. On comparing the various inhibitors and standard antibiotics docked
12 International Journal of Scientific and Research Publications, Volume 7, Issue 3, March ISSN upon DHFR and DHPS of E.faecalis, Quercetin, Ellagic acid, Penicillin and Cephalosporin show higher binding affinity. Hence, it can be concluded that the compounds Quercetin and Ellagic acid have significant potential to bind to the active site of DHFR. Our study also shows that, inhibitors like Clavalunic acid and Gallic acid efficiently bind to the active site of the DHPS of E.faecalis. Quercetin and Chlorogenic acid have the highest binding affinity to the ligand binding pocket of DHPS of E.faecalis. When compared to inhibitors i.e Quercetin and Chlorogenic acid, antibiotics like Cephalosporin & Penicillin show higher interactions towards DHPS of E.faecalis. Even though, Standard antibiotics viz., Cephalosporin and Penicillin do show strong interactions with both the proteins of the folate synthesis pathway, the clinical isolates of E.faecalis have developed resistance to these. Hence, a better alternative, which binds to DHFR is Quercetin and Ellagic acid while inhibitors that binds to DHPS are Quercetin and Chlorogenic acid. Protein binding with various ligands, indicate that various inhibitors viz., Chlorogenic acid, Ellagic acid, Gallic Acid, Hippuric acid and Quercetin of DHFR and DHPS can be utilized for the treatment of MDR UTI after due invivo, invitro and ADMET testing, since they have been proved to posses potential antibacterial activities. IV. CONCLUSION Antibiotics have been a high success till date for curbing bacterial infections. But, the widespread and uncontrolled use of antibiotics has led to the emergence of multidrugresistant (MDR) bacteria (15). Along with limited treatment options and increased mortality the MDR seems grave. Hence, there is an urgent need to search for a new antibacterial agent. The molecular docking programs aid to establish new ligands/inhibitors for the selected target receptor proteins from the different available databases, based on their efficiency to bind the active sites on the receptor (15). Our study shows that inhibitors viz., Quercetin and Ellagic acid show best interactions and binding energy with DHFR while Quercetin and Chlorogenic acid show best interactions and binding energy with DHPS of the folate synthesis pathway of E.faecalis. These in silico studies supported with invivo, invitro and ADMET testing will certainly help towards developing candidates for treatment of MDRUTI in the future. More studies on mutations are needed for corroborating the role of quercetin, chlorogenic acid and ellagic acid as antibacterial agents to treat MDR E. faecalis mediated uropathological infections. ACKNOWLEDGMENTS We acknowledge the grant received from R & I, Technology Transfer Project funded by RUSA, Maharashtra Government, India for Rs. 35 lacs, June 2016, (Sanction No. RUSA/ order/ R&I/ / 273) Dt.18 /6/ REFERENCES [1] Purva Mathur, Arti Kapil, Rachna Chandra, Pratibha Sharma & Bimal Das, Antimicrobial resistance in Enterococcus faecalis at a tertiary care centre of northern India, Indian J Med Res 118, July 2003, pp 2528 [2] CindyLove Tremblay 1 Ann Letellier 1 Sylvain Quessy 1 Danielle Daignault, 2 and Marie Archambaulti*AntibioticResistant Enterococcus faecalis in Abattoir Pigs and Plasmid Colocalization and Cotransfer of tet(m) and erm(b) Genes, Journal of Food Protection, Vol. 75, No. 9, 2012, Pages [3] J. Scott Weese, DVM, DVSc, DACVIM MultidrugResistant Enterococcal Infections, University of Guelph. [4] Cecilia Pozzi,a Stefania Ferrari,b Debora Cortesi,b Rosaria Luciani,b Robert M. Stroud,c Alessia Catalano,d Maria Paola Costib* and Stefano Mangania*, The structure of Enterococcus faecalis thymidylate synthase provides clues about folate bacterial metabolism, ISSN [5] Saraswathy MP, Multidrug resistant Enterococci isolated from urine samples at a tertiary care hospital. [6] *Suddhanshu Bhardwaj, Kalyani Bhamre Jayashri Dhawale, Mahendra Patil and Sunil Divase, Enterococcus faecium and Enterococcus faecalis, the nosocomial pathogens with special reference to multidrug resistance and phenotypic characterization, International Journal of Pharmaceutical Science and Practice. Volume 2, Number 1 (2013) pp 110. [7] Maj Puneet Bhatt a,*, Anubha Patel b, Brig A.K. Sahni c, Surg Cmde A.K. Praharaj, (Retd)d, Col Naveen Grover e, Surg Cdr C.N. Chaudhari f, Nikunja Kumar Das b, Mayuri Kulkarni b Emergence of multidrug resistant enterococci at a tertiary care centre Medical Journal armed forces India 71 (2015) [8] William R Miller 1, Jose M Munita 1,2, and Cesar A Arias*,1,3 Mechanisms of antibiotic resistance in enterococci Expert Rev Anti Infect Ther October ; 12(10): doi: / [9] Sudha Ramachandra 1, Vinay Chavan 2 A Genetic Algorithm for Conformation Search Optimization in Molecular Docking (IJCSIT) International Journal of Computer Science and Information Technologies, Vol. 6 (6), 2015, [10] Lokesh Ravi, Kannabiran K*, A Handbook on ProteinLigand Docking tool, ISSN Innovare Journal of medical sciences. [11] Brooijmans N, Kuntz I. Molecular recognition and docking algorithms. Annu Rev Biophys Biomol Struct 2003;32: [12] Kitchen D, Decornez H, Furr J, Bajorath J. Docking and scoring in virtual screening for drug discovery: methods and applications. Nat Rev Drug Discovery 2004;3: [13] Hartshorn M, Murray C, Cleasby A, Frederickson M, Tickle I, Jhoti H. Fragmentbased lead discovery using Xray crystallography. J Med Chem 2005;48: [14] David E, Stephen N. Virtual screening of DNA minor groove binders. J Med Chem 2006;49: [15] Pallavi Sahare 11, Archana Moon. In silico modelling of βlactam resistant Enterococcus faecalis PBP4 and its interactions with various phytoligands.. International Journal of Pharmacy and Pharmaceutical Sciences, Vol 8, Issue 7, [16] P. Sahare and A. Moon * Insilico docking studies of phytoligands against e. Coli PBP3: approachtowards novel antibacterial therapeutic agent IJPSR, 2016; Vol. 7(9): International Journal of Pharmaceutical Sciences and Research. [17] Brian LH, Louis BR. Intrinsic and acquired resistance mechanisms in Enterococcus. Virulence 2012;3: [18] Maryam Zahin, 1,2 Iqbal Ahmad, 1 Ramesh C. Gupta, 2,3 and Farrukh Aqil 2,4, Punicalagin and Ellagic Acid Demonstrate Antimutagenic Activity and Inhibition of Benzo[a]pyrene Induced DNA Adducts, Hindawi Publishing Corporation BioMed Research International Volume 2014, Article ID , 10 pages. [19] Marzena Matejczyk, Biological and Anticancer activity of selected Natural Poducts, Bialystok University of Technology, Faculty of Civil Engineering and Environmental Engineering, Department of Sanitary Biology and Biotechnology, Medical and Biological Sciences, 2015, 29/3, [20] Dragana D. Božić, Marina Milenković, Branka Ivković* & Ivana Ćirković** Antibacterial activity of three newlysynthesized chalcones & synergism with antibiotics against clinical isolates of methicillinresistant Staphylococcus aureus, Indian J Med Res 140, July 2014, pp
13 International Journal of Scientific and Research Publications, Volume 7, Issue 3, March ISSN [21] M. Matsuura,' H. Nakazawa, 1 T. Hashimoto, 2 and S. MitsuhashiI', Combined Antibacterial Activity of Amoxicillin with Clavulanic Acid Against AmpicillinResistant Strains, Antibacterial Agents and Chemotherapy, June 1980, p [22] R. Knox and J. T. Smith, Antibacterial Activity, Penicillinase Stability and Inducing Ability of Different Penicillins, J. gen. Microbiol. (1962), 28, [23] Jureerut Daduang 1*, Adisak Palasap 1, Sakda Daduang 2, Patcharee Boonsiri 3, Prasit Suwannalert 4, Temduang Limpaiboon 1 Gallic Acid Enhancement of Gold Nanoparticle Anticancer Activity in Cervical Cancer Cells, Asian Pac J Cancer Prev, 16 (1), [24] Venkata Ratna Ravi Kumar Dasari, Sri Rami Reddy Donthireddy, Murali Yugandhar Nikku and Hanumantha Rao Garapati*, Optimization of medium constituents for Cephalosporin C production using response surface methodology and artificial neural networks, J Biochem Tech (2009) 1(3):6974 ISSN: [25] Kumar Gaurav a*, Subir Kundu a and Richa Srivastav b, Synthesis and In Vitro antibacterial activity of some Novel Cephem Antibiotics, International Journal of Pharmacy and Pharmaceutical Sciences. AUTHORS First Author Archana Moon, Professor, University Department of Biochemistry, Rashtrasant Tukadoji Maharaj Nagpur University, Nagpur , moon.archana@gmail.com, Contact number: Second Author Deeba Khan, Project fellow, University Department of Biochemistry, Rashtrasant Tukadoji Maharaj Nagpur University, Nagpur , deebabiochemistry@gmail.com, Contact number: Third Author Pranjali Gajbhiye, University Department of Biochemistry, Rashtrasant Tukadoji Maharaj Nagpur University, Nagpur Fourth Author Monali Jariya, University Department of Biochemistry, Rashtrasant Tukadoji Maharaj Nagpur University, Nagpur
INSILICO DOCKING AND INTERACTION ANALYSIS OF ELLAGIC ACID AND CURCUMIN DERIVATIVES AGAINST HUMAN CANCER
 ISSN: 0976-2876 (Print) ISSN: 2250-0138(Online) INSILICO DOCKING AND INTERACTION ANALYSIS OF ELLAGIC ACID AND CURCUMIN DERIVATIVES AGAINST HUMAN CANCER NAYANA MOHAN a AND M.S. LATHA b1 ab Department of
ISSN: 0976-2876 (Print) ISSN: 2250-0138(Online) INSILICO DOCKING AND INTERACTION ANALYSIS OF ELLAGIC ACID AND CURCUMIN DERIVATIVES AGAINST HUMAN CANCER NAYANA MOHAN a AND M.S. LATHA b1 ab Department of
Section 5. Enzymes, Equilibrium, Energy and the Sulfonamides
 Section 5 Enzymes, Equilibrium, Energy and the Sulfonamides Monday: ESKAPE handout describing them (Tiffany will provide). M-W Tie the metabolism back to the nutritional requirements and media choice,
Section 5 Enzymes, Equilibrium, Energy and the Sulfonamides Monday: ESKAPE handout describing them (Tiffany will provide). M-W Tie the metabolism back to the nutritional requirements and media choice,
Available Online through
 ISS: 0975-766X CDE: IJPTFI Available nline through Research Article www.ijptonline.com RELATISHIP BETWEE SCRIG FUCTI AD THE IC 50 IHIBITRY F PI3K IHIBITRS Awantika Shrivastava*, Dr.K.Durga Prasad 1, Dr.
ISS: 0975-766X CDE: IJPTFI Available nline through Research Article www.ijptonline.com RELATISHIP BETWEE SCRIG FUCTI AD THE IC 50 IHIBITRY F PI3K IHIBITRS Awantika Shrivastava*, Dr.K.Durga Prasad 1, Dr.
CS612 - Algorithms in Bioinformatics
 Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
Identification of polyphenolic compound inhibitors for 1, 3 β D- glucan synthase in the treatment of dermatophytoses by molecular docking
 International Journal of Advanced Research in Medical Sciences 2017 Volume 1 Issue 1 Pages: 1-5 Original Research Paper Identification of polyphenolic compound inhibitors for 1, 3 β D- glucan synthase
International Journal of Advanced Research in Medical Sciences 2017 Volume 1 Issue 1 Pages: 1-5 Original Research Paper Identification of polyphenolic compound inhibitors for 1, 3 β D- glucan synthase
Molecular Docking Studies of Flavonoids of Noni Fruit (Morinda citrifolia L.) to Peroxisome Proliferator-Activated Receptor-Gamma (PPARγ)
 3rd International Conference on Computation for Science and Technology (ICCST-3) Molecular Docking Studies of Flavonoids of Noni Fruit (Morinda citrifolia L.) to Peroxisome Proliferator-Activated Receptor-Gamma
3rd International Conference on Computation for Science and Technology (ICCST-3) Molecular Docking Studies of Flavonoids of Noni Fruit (Morinda citrifolia L.) to Peroxisome Proliferator-Activated Receptor-Gamma
Molecular docking analysis of UniProtKB nitrate reductase enzyme with known natural flavonoids
 www.bioinformation.net Volume 12(12) Hypothesis Molecular docking analysis of UniProtKB nitrate reductase enzyme with known natural flavonoids Ayub Shaik 1, Vishnu Thumma 2, Aruna Kumari Kotha 2, Sandhya
www.bioinformation.net Volume 12(12) Hypothesis Molecular docking analysis of UniProtKB nitrate reductase enzyme with known natural flavonoids Ayub Shaik 1, Vishnu Thumma 2, Aruna Kumari Kotha 2, Sandhya
A Novel Lantibiotic Acting on Bacterial Cell Wall Synthesis Produced by the Uncommon FLAVIA MARINELLI. DBSM, University of Insubria, Varese Italy
 A Novel Lantibiotic Acting on Bacterial Cell Wall Synthesis Produced by the Uncommon Actinomycete Planomonospora sp. FLAVIA MARINELLI DBSM, University of Insubria, Varese Italy Vicuron Pharmaceuticals,
A Novel Lantibiotic Acting on Bacterial Cell Wall Synthesis Produced by the Uncommon Actinomycete Planomonospora sp. FLAVIA MARINELLI DBSM, University of Insubria, Varese Italy Vicuron Pharmaceuticals,
Journal of Chemical and Pharmaceutical Research, 2015, 7(12): Research Article
 Available online www.jocpr.com Journal of Chemical and Pharmaceutical Research, 2015, 7(12):1082-1086 Research Article ISSN : 0975-7384 CODEN(USA) : JCPRC5 Determination of three dimensional structure
Available online www.jocpr.com Journal of Chemical and Pharmaceutical Research, 2015, 7(12):1082-1086 Research Article ISSN : 0975-7384 CODEN(USA) : JCPRC5 Determination of three dimensional structure
Molecular Docking Studies on Pyrazolopyrimidine and their Derivatives as Human Phosphoinositide 3-Kinase Inhibitors
 Cloud Publications International Journal of Advanced Bioinformatics and Computational Biology 2012, Volume 1, Issue 1, pp. 1-11, Article ID Med-30 Research Article Open Access Molecular Docking Studies
Cloud Publications International Journal of Advanced Bioinformatics and Computational Biology 2012, Volume 1, Issue 1, pp. 1-11, Article ID Med-30 Research Article Open Access Molecular Docking Studies
Exploring in silico affinity of flavonoids and tannins to human fibroblast growth factorinducible14 (Fn14), a member of TNF receptor super family
 www.bioinformation.net Hypothesis Volume 9(12) Exploring in silico affinity of flavonoids and tannins to human fibroblast growth factorinducible14 (Fn14), a member of TNF receptor super family CVS Siva
www.bioinformation.net Hypothesis Volume 9(12) Exploring in silico affinity of flavonoids and tannins to human fibroblast growth factorinducible14 (Fn14), a member of TNF receptor super family CVS Siva
Identification of Targetable Virulence Factor and Drug Screening For Bacterial Pneumonia
 IOSR Journal of Pharmacy and Biological Sciences (IOSR-JPBS) e-issn: 2278-3008, p-issn:2319-7676. Volume 10, Issue 2 Ver. 1 (Mar -Apr. 2015), PP 20-24 www.iosrjournals.org Identification of Targetable
IOSR Journal of Pharmacy and Biological Sciences (IOSR-JPBS) e-issn: 2278-3008, p-issn:2319-7676. Volume 10, Issue 2 Ver. 1 (Mar -Apr. 2015), PP 20-24 www.iosrjournals.org Identification of Targetable
In silico approach of compounds in Cissus quadrangularis targeting multiproteins as anticancer agents
 Research Article In silico approach of compounds in Cissus quadrangularis targeting multiproteins as anticancer agents C. N. Hemalatha 1, Elancheziyan K 1, D. Pavithra 2, M. Vijey Aanandhi 3 * ABSTRACT
Research Article In silico approach of compounds in Cissus quadrangularis targeting multiproteins as anticancer agents C. N. Hemalatha 1, Elancheziyan K 1, D. Pavithra 2, M. Vijey Aanandhi 3 * ABSTRACT
New Insights into Peptidoglycan Biosynthesis. Louis B. Rice Warren Alpert Medical School of Brown University Providence, RI
 New Insights into Peptidoglycan Biosynthesis Louis B. Rice Warren Alpert Medical School of Brown University Providence, RI Outline of Presentation Review role of penicillin-binding proteins in cell wall
New Insights into Peptidoglycan Biosynthesis Louis B. Rice Warren Alpert Medical School of Brown University Providence, RI Outline of Presentation Review role of penicillin-binding proteins in cell wall
Natural Flavonoids as CDK5 inhibitors: A Homology modeling and Molecular Docking Study
 International Journal of ChemTech Research CDEN (USA): IJCRGG, ISSN: 0974-4290, ISSN(nline):2455-9555 Vol.9, No.09 pp 210-215, 2016 Natural Flavonoids as CDK5 inhibitors: A Homology modeling and Molecular
International Journal of ChemTech Research CDEN (USA): IJCRGG, ISSN: 0974-4290, ISSN(nline):2455-9555 Vol.9, No.09 pp 210-215, 2016 Natural Flavonoids as CDK5 inhibitors: A Homology modeling and Molecular
University Department of Biochemistry (Established in 1946) July 2016; Vol. 4 Issue 2
 ...An e Newsletter of University Department of Biochemistry July 2016; Vol. 4 Issue 2 University Department of Biochemistry (Established in 1946) e-mail: rtmnubiochem@gmail.com Editorial Board Chief Editor
...An e Newsletter of University Department of Biochemistry July 2016; Vol. 4 Issue 2 University Department of Biochemistry (Established in 1946) e-mail: rtmnubiochem@gmail.com Editorial Board Chief Editor
Research Article. Screening of Annona muricata for E. coli enoyl acyl carrier protein reductase inhibitors by molecular docking
 Available online www.jocpr.com Journal of Chemical and Pharmaceutical Research, 2015, 7(4):452-457 Research Article ISSN : 0975-7384 CODEN(USA) : JCPRC5 Screening of Annona muricata for E. coli enoyl acyl
Available online www.jocpr.com Journal of Chemical and Pharmaceutical Research, 2015, 7(4):452-457 Research Article ISSN : 0975-7384 CODEN(USA) : JCPRC5 Screening of Annona muricata for E. coli enoyl acyl
A molecular docking study of anticancer drug paclitaxel and its analogues
 Indian Journal of Biochemistry & Biophysics Vol. 48, April 2011, pp. 101-105 A molecular docking study of anticancer drug paclitaxel and its analogues Ruma Sinha, Ambarish Sharan Vidyarthi and Shankaracharya*
Indian Journal of Biochemistry & Biophysics Vol. 48, April 2011, pp. 101-105 A molecular docking study of anticancer drug paclitaxel and its analogues Ruma Sinha, Ambarish Sharan Vidyarthi and Shankaracharya*
Amino Acids. Review I: Protein Structure. Amino Acids: Structures. Amino Acids (contd.) Rajan Munshi
 Review I: Protein Structure Rajan Munshi BBSI @ Pitt 2005 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2005 Amino Acids Building blocks of proteins 20 amino acids
Review I: Protein Structure Rajan Munshi BBSI @ Pitt 2005 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2005 Amino Acids Building blocks of proteins 20 amino acids
IN-SILICO APPROACH FOR THE ASSESSMENT OF ORAL CANCER PROPERTY ON LIMONIA ACIDISSIMA
 IJPSR (2016), Vol. 7, Issue 3 (Research Article) Received on 16 September, 2015; received in revised form, 29 October, 2015; accepted, 12 December, 2015; published 01 March, 2016 IN-SILICO APPROACH FOR
IJPSR (2016), Vol. 7, Issue 3 (Research Article) Received on 16 September, 2015; received in revised form, 29 October, 2015; accepted, 12 December, 2015; published 01 March, 2016 IN-SILICO APPROACH FOR
This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is worth 2 points.
 MBB 407/511 Molecular Biology and Biochemistry First Examination - October 1, 2002 Name Social Security Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is
MBB 407/511 Molecular Biology and Biochemistry First Examination - October 1, 2002 Name Social Security Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is
Chemical Mechanism of Enzymes
 Chemical Mechanism of Enzymes Enzyme Engineering 5.2 Definition of the mechanism 1. The sequence from substrate(s) to product(s) : Reaction steps 2. The rates at which the complex are interconverted 3.
Chemical Mechanism of Enzymes Enzyme Engineering 5.2 Definition of the mechanism 1. The sequence from substrate(s) to product(s) : Reaction steps 2. The rates at which the complex are interconverted 3.
open access Hypothesis Volume 10(7) Rajesh Kumar Pathak 1, 2, Mamta Baunthiyal 1 *, Gohar Taj 2 & Anil Kumar 2
 www.bioinformation.net Hypothesis Volume 10(7) Virtual screening of natural inhibitors to the predicted HBx protein structure of Hepatitis B Virus using molecular docking for identification of potential
www.bioinformation.net Hypothesis Volume 10(7) Virtual screening of natural inhibitors to the predicted HBx protein structure of Hepatitis B Virus using molecular docking for identification of potential
Comparative Modeling of Gingival Protein and Docking Studies with Natural Flavonoid Inhibitors
 DOI:10.7598/cst2016.1226 Chemical Science Transactions ISSN:2278-3458 2016, 5(2), 387-392 RESEARCH ARTICLE Comparative Modeling of Gingival Protein and Docking Studies with Natural Flavonoid Inhibitors
DOI:10.7598/cst2016.1226 Chemical Science Transactions ISSN:2278-3458 2016, 5(2), 387-392 RESEARCH ARTICLE Comparative Modeling of Gingival Protein and Docking Studies with Natural Flavonoid Inhibitors
Research Article. Insilico predictions of inhibitors of novel statin structural analogues with HMG-CoA reductase
 Available online www.jocpr.com Journal of Chemical and Pharmaceutical Research, 2015, 7(3):942-950 Research Article ISSN : 0975-7384 CODEN(USA) : JCPRC5 Insilico predictions of inhibitors of novel statin
Available online www.jocpr.com Journal of Chemical and Pharmaceutical Research, 2015, 7(3):942-950 Research Article ISSN : 0975-7384 CODEN(USA) : JCPRC5 Insilico predictions of inhibitors of novel statin
Copyright 2008 Pearson Education, Inc., publishing as Pearson Benjamin Cummings
 Concept 5.4: Proteins have many structures, resulting in a wide range of functions Proteins account for more than 50% of the dry mass of most cells Protein functions include structural support, storage,
Concept 5.4: Proteins have many structures, resulting in a wide range of functions Proteins account for more than 50% of the dry mass of most cells Protein functions include structural support, storage,
LAB#23: Biochemical Evidence of Evolution Name: Period Date :
 LAB#23: Biochemical Evidence of Name: Period Date : Laboratory Experience #23 Bridge Worth 80 Lab Minutes If two organisms have similar portions of DNA (genes), these organisms will probably make similar
LAB#23: Biochemical Evidence of Name: Period Date : Laboratory Experience #23 Bridge Worth 80 Lab Minutes If two organisms have similar portions of DNA (genes), these organisms will probably make similar
Molecular Docking Studies of Shc1 Protein: A Drug Target to Treat Obesity
 IOSR Journal of Pharmacy and Biological Sciences (IOSR-JPBS) e-issn: 2278-3008, p-issn:2319-7676. Volume 9, Issue 6 Ver. III (Nov -Dec. 2014), PP 63-68 Molecular Docking Studies of Shc1 Protein: A Drug
IOSR Journal of Pharmacy and Biological Sciences (IOSR-JPBS) e-issn: 2278-3008, p-issn:2319-7676. Volume 9, Issue 6 Ver. III (Nov -Dec. 2014), PP 63-68 Molecular Docking Studies of Shc1 Protein: A Drug
Introduction to Protein Structure Collection
 Introduction to Protein Structure Collection Teaching Points This collection is designed to introduce students to the concepts of protein structure and biochemistry. Different activities guide students
Introduction to Protein Structure Collection Teaching Points This collection is designed to introduce students to the concepts of protein structure and biochemistry. Different activities guide students
Trans-10,cis-12 Conjugated Linoleic Acid a Novel Inhibitor of Inflammatory Mediators
 American Journal of Bioinformatics Research 2016, 6(2): 27-31 DOI: 10.5923/j.bioinformatics.20160602.01 Trans-10,cis-12 Conjugated Linoleic Acid a Novel Inhibitor of Inflammatory Mediators Rashmi H. K.,
American Journal of Bioinformatics Research 2016, 6(2): 27-31 DOI: 10.5923/j.bioinformatics.20160602.01 Trans-10,cis-12 Conjugated Linoleic Acid a Novel Inhibitor of Inflammatory Mediators Rashmi H. K.,
Structure of the measles virus hemagglutinin bound to the CD46 receptor. César Santiago, María L. Celma, Thilo Stehle and José M.
 Supporting Figures and Table for Structure of the measles virus hemagglutinin bound to the CD46 receptor César Santiago, María L. Celma, Thilo Stehle and José M. Casasnovas This PDF file includes: Supplementary
Supporting Figures and Table for Structure of the measles virus hemagglutinin bound to the CD46 receptor César Santiago, María L. Celma, Thilo Stehle and José M. Casasnovas This PDF file includes: Supplementary
Ch. Charan Singh University, Meerut (250004), India
 EFFICIENT IDENTIFICATION OF NOVEL ANTIFUNGAL COMPOUNDS TARGETING CYTOCHROME-B OF FUSARIUM OXYSPORUM F. SP. LYCOPERSICI BY VIRTUAL SCREENING AND STATISTICAL ANALYSIS 1 Reetu Gour, 2 A.P. Garg and 3 Ajay
EFFICIENT IDENTIFICATION OF NOVEL ANTIFUNGAL COMPOUNDS TARGETING CYTOCHROME-B OF FUSARIUM OXYSPORUM F. SP. LYCOPERSICI BY VIRTUAL SCREENING AND STATISTICAL ANALYSIS 1 Reetu Gour, 2 A.P. Garg and 3 Ajay
II. METHODS AND MATERIALS A.
 IN-SIlico Docking Analysis of Sterculia Lychnophora Compounds against Proteins Causing Alzheimer s Disease Pooja Shetty 1, Anuradha Palve 1, Mukesh Pimpliskar 2, R. N. Jadhav 3* and Pramod Shinde 1 1 Department
IN-SIlico Docking Analysis of Sterculia Lychnophora Compounds against Proteins Causing Alzheimer s Disease Pooja Shetty 1, Anuradha Palve 1, Mukesh Pimpliskar 2, R. N. Jadhav 3* and Pramod Shinde 1 1 Department
PHARMACOLOGY II. Dr Shariq Syed Associate Professor AIKTC, SoP
 PHARMACOLOGY II Dr Shariq Syed Associate Professor AIKTC, SoP INTRODUCTION TO BACTERIA! INTRODUCTION TO BACTERIA! THEY COME IN DIFFERENT SHAPES ANTIMICROBIAL SITES OF ACTION SULPHONAMIDES 1930, Physician/researcher
PHARMACOLOGY II Dr Shariq Syed Associate Professor AIKTC, SoP INTRODUCTION TO BACTERIA! INTRODUCTION TO BACTERIA! THEY COME IN DIFFERENT SHAPES ANTIMICROBIAL SITES OF ACTION SULPHONAMIDES 1930, Physician/researcher
Spectrum of vancomycin and susceptibility testing
 Spectrum of vancomycin and susceptibility testing Olivier Denis Service de Microbiologie Laboratoire de bactériologie Service de Microbiologie Hôpital Erasme Glycopeptides Vancomycin 1958 < Amycolatopsis
Spectrum of vancomycin and susceptibility testing Olivier Denis Service de Microbiologie Laboratoire de bactériologie Service de Microbiologie Hôpital Erasme Glycopeptides Vancomycin 1958 < Amycolatopsis
Computer Aided Docking Studies on Anticancer Drugs for Breast Cancer
 International Journal of Information & Computation Technology. ISSN 0974-2239 Volume 2, Number 1 (2012), pp. 49-55 International Research Publications House http://www. ripublication.com Computer Aided
International Journal of Information & Computation Technology. ISSN 0974-2239 Volume 2, Number 1 (2012), pp. 49-55 International Research Publications House http://www. ripublication.com Computer Aided
SUPPORTING INFORMATION FOR. A Computational Approach to Enzyme Design: Using Docking and MM- GBSA Scoring
 SUPPRTING INFRMATIN FR A Computational Approach to Enzyme Design: Predicting ω- Aminotransferase Catalytic Activity Using Docking and MM- GBSA Scoring Sarah Sirin, 1 Rajesh Kumar, 2 Carlos Martinez, 2
SUPPRTING INFRMATIN FR A Computational Approach to Enzyme Design: Predicting ω- Aminotransferase Catalytic Activity Using Docking and MM- GBSA Scoring Sarah Sirin, 1 Rajesh Kumar, 2 Carlos Martinez, 2
Computational Analysis of Oral Disease (Periodontitis) for Drug Targets
 Year: 2014; Volume: 1; Issue: 1 Article ID: BCI14 01; Pages: 1-6 Research Article Computational Analysis of Oral Disease (Periodontitis) for Drug Targets V. Madhumathi and *S. Vijaya kumar P.G. and Research
Year: 2014; Volume: 1; Issue: 1 Article ID: BCI14 01; Pages: 1-6 Research Article Computational Analysis of Oral Disease (Periodontitis) for Drug Targets V. Madhumathi and *S. Vijaya kumar P.G. and Research
Introduction to proteins and protein structure
 Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
3.2 Ligand-Binding at Nicotinic Acid Receptor Subtypes GPR109A/B
 3.2 Ligand-Binding at Nicotinic Acid Receptor Subtypes GPR109A/B 3.2.1 Characterization of the Ligand Binding Site at GPR109A Receptor Ligands of GPR109A Receptor are Carboxylic Acids Nicotinic acid (pyridine-3-carboxylic
3.2 Ligand-Binding at Nicotinic Acid Receptor Subtypes GPR109A/B 3.2.1 Characterization of the Ligand Binding Site at GPR109A Receptor Ligands of GPR109A Receptor are Carboxylic Acids Nicotinic acid (pyridine-3-carboxylic
Design and Modeling Studies on Liriodenine derivatives as novel topoisomerase II inhibitors
 International Journal of ChemTech Research CODEN( USA): IJCRGG ISSN : 0974-4290 Vol. 3, No.3,pp 1622-1627, July-Sept 2011 Design and Modeling Studies on Liriodenine derivatives as novel topoisomerase II
International Journal of ChemTech Research CODEN( USA): IJCRGG ISSN : 0974-4290 Vol. 3, No.3,pp 1622-1627, July-Sept 2011 Design and Modeling Studies on Liriodenine derivatives as novel topoisomerase II
ANTICANCER DRUGS AS PROSPECTIVE EFFLUX PUMP INHIBITORS FOR SALMONELLA TYPHI PRODUCE CONFLICTING RESULTS IN IN SILICO AND IN VITRO STUDIES
 International Journal of Pharmacy and Pharmaceutical Sciences ISSN- 0975-1491 Vol 8, Issue 12, 2016 Original Article ANTICANCER DRUGS AS PROSPECTIVE EFFLUX PUMP INHIBITORS FOR SALMONELLA TYPHI PRODUCE
International Journal of Pharmacy and Pharmaceutical Sciences ISSN- 0975-1491 Vol 8, Issue 12, 2016 Original Article ANTICANCER DRUGS AS PROSPECTIVE EFFLUX PUMP INHIBITORS FOR SALMONELLA TYPHI PRODUCE
Investigators Meeting
 Outcomes of Urinary Tract Infection Management by Pharmacists (R x OUTMAP) Investigators Meeting June 11, 2017 Overview 1. Introductions and Opening Remarks 2. Epidemiology and Definitions 3. UTI Assessment
Outcomes of Urinary Tract Infection Management by Pharmacists (R x OUTMAP) Investigators Meeting June 11, 2017 Overview 1. Introductions and Opening Remarks 2. Epidemiology and Definitions 3. UTI Assessment
Molecular modeling of Ruellia tuberosa compounds as -amylase inhibitor: an in silico comparation between human and rat enzyme model
 www.bioinformation.net Volume 10(4) Hypothesis Molecular modeling of Ruellia tuberosa L compounds as amylase inhibitor: an in silico comparation between human and rat enzyme model Dyah Ratna Wulan 1, 2,
www.bioinformation.net Volume 10(4) Hypothesis Molecular modeling of Ruellia tuberosa L compounds as amylase inhibitor: an in silico comparation between human and rat enzyme model Dyah Ratna Wulan 1, 2,
fibromyalgia and early genetic studies revealed that the gene SLC6A4 showed an increased frequency among patients
 ISSN: 0975-766X CODEN: IJPTFI Available Online through Research Article www.ijptonline.com IN-SILICO SCREENING FOR THE IDENTIFICATION OF SLC6A4 DRUG TARGETS AND ANTI- RHEUMATIC ACTIVITY OF PHYTOLIGANDS
ISSN: 0975-766X CODEN: IJPTFI Available Online through Research Article www.ijptonline.com IN-SILICO SCREENING FOR THE IDENTIFICATION OF SLC6A4 DRUG TARGETS AND ANTI- RHEUMATIC ACTIVITY OF PHYTOLIGANDS
Molecular modelling and insilico drug docking studies on breast cancer target protein (TNRC9) using cheminformatics software and tools
 Available online at www.ijntps.org ISSN: 2277 2782 INTERNATIONAL JOURNAL OF NOVEL TRENDS IN PHARMACEUTICAL SCIENCES Molecular modelling and insilico drug docking studies on breast cancer target protein
Available online at www.ijntps.org ISSN: 2277 2782 INTERNATIONAL JOURNAL OF NOVEL TRENDS IN PHARMACEUTICAL SCIENCES Molecular modelling and insilico drug docking studies on breast cancer target protein
Protein Refinement in Discovery Studio 1.7. Presented by: Francisco Hernandez-Guzman, PhD
 Protein Refinement in Discovery Studio 1.7 Presented by: Francisco Hernandez-Guzman, PhD February 2007 Agenda Part I: Protein Refinement in Discovery Studio 1.7 (20mins) The problem Novel Algorithms at
Protein Refinement in Discovery Studio 1.7 Presented by: Francisco Hernandez-Guzman, PhD February 2007 Agenda Part I: Protein Refinement in Discovery Studio 1.7 (20mins) The problem Novel Algorithms at
BUGS and DRUGS Part 2 March 13, 2013 Marieke Kruidering- Hall
 BUGS and DRUGS Part March, 0 Marieke Kruidering- Hall BIOGRAPHY: Marieke Kruidering- Hall is Associate Professor in the Department of Cellular & Molecular Pharmacology. She was born in the Netherlands.
BUGS and DRUGS Part March, 0 Marieke Kruidering- Hall BIOGRAPHY: Marieke Kruidering- Hall is Associate Professor in the Department of Cellular & Molecular Pharmacology. She was born in the Netherlands.
Urinary tract infection. Mohamed Ahmed Fouad Lecturer of pediatrics Jazan faculty of medicine
 Urinary tract infection Mohamed Ahmed Fouad Lecturer of pediatrics Jazan faculty of medicine Objectives To differentiate between types of urinary tract infections To recognize the epidemiology of UTI in
Urinary tract infection Mohamed Ahmed Fouad Lecturer of pediatrics Jazan faculty of medicine Objectives To differentiate between types of urinary tract infections To recognize the epidemiology of UTI in
AutoLigand: A Tool for Finding Ligand Binding Sites. CUHK workshop Dec, 2011
 AutoLigand: A Tool for Finding Ligand Binding Sites CUHK workshop Dec, 2011 Stefano Forli, Ph.D., TSRI Rodney M. Harris, Ph.D., 1060 Discovery Engineering Overview Description and uses of AutoLigand Algorithms
AutoLigand: A Tool for Finding Ligand Binding Sites CUHK workshop Dec, 2011 Stefano Forli, Ph.D., TSRI Rodney M. Harris, Ph.D., 1060 Discovery Engineering Overview Description and uses of AutoLigand Algorithms
Cases in employees. Cases. Day of onset (July)
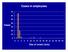 Cases in employees 70 60 50 Cases 40 30 20 10 0 0 2 4 6 8 10 12 14 16 18 20 22 24 26 28 30 Day of onset (July) 70 60 50 40 Cases 30 Cases in employees and visitors Employees Visitors 20 10 0 0 2 4 6 8
Cases in employees 70 60 50 Cases 40 30 20 10 0 0 2 4 6 8 10 12 14 16 18 20 22 24 26 28 30 Day of onset (July) 70 60 50 40 Cases 30 Cases in employees and visitors Employees Visitors 20 10 0 0 2 4 6 8
Review II: The Molecules of Life
 Review II: The Molecules of Life Judy Wieber BBSI @ Pitt 2007 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2007 Outline Introduction Proteins Carbohydrates Lipids
Review II: The Molecules of Life Judy Wieber BBSI @ Pitt 2007 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2007 Outline Introduction Proteins Carbohydrates Lipids
Final Report. Identification of Residue Mutations that Increase the Binding Affinity of LSTc to HA RBD. Wendy Fong ( 方美珍 ) CNIC; Beijing, China
 Final Report Identification of Residue Mutations that Increase the Binding Affinity of LSTc to HA RBD Wendy Fong ( 方美珍 ) CNIC; Beijing, China Influenza s Viral Life Cycle 1. HA on virus binds to sialic
Final Report Identification of Residue Mutations that Increase the Binding Affinity of LSTc to HA RBD Wendy Fong ( 方美珍 ) CNIC; Beijing, China Influenza s Viral Life Cycle 1. HA on virus binds to sialic
International Journal of Pharma and Bio Sciences V1(2)2010 IN SILICO PHARMACOGENOMIC ANALYSIS OF ALCOHOL DEHYDROGENASE INVOLVED IN ALCOHOLISM
 SINGH SATENDRA, MECARTY S. D., JAIN P.A., GAUTAM B., FARMER R., YADAV P.K. AND RAM G.D. 1 Department of Computational Biology & Bioinformatics, JSBBE, SHIATS, Allahabad-211007,India 1 Department of Tissue
SINGH SATENDRA, MECARTY S. D., JAIN P.A., GAUTAM B., FARMER R., YADAV P.K. AND RAM G.D. 1 Department of Computational Biology & Bioinformatics, JSBBE, SHIATS, Allahabad-211007,India 1 Department of Tissue
STRUCTURE OF COMMONLY USED PENICILLINS
 PENICILLINS Alice Prince I. CHEMISTRY A basic structure of penicillins consists of a nucleus with three components: a thiazolidine ring, a β-lactam ring and a side chain. The side chain determines, in
PENICILLINS Alice Prince I. CHEMISTRY A basic structure of penicillins consists of a nucleus with three components: a thiazolidine ring, a β-lactam ring and a side chain. The side chain determines, in
The Structure and Function of Macromolecules
 The Structure and Function of Macromolecules Macromolecules are polymers Polymer long molecule consisting of many similar building blocks. Monomer the small building block molecules. Carbohydrates, proteins
The Structure and Function of Macromolecules Macromolecules are polymers Polymer long molecule consisting of many similar building blocks. Monomer the small building block molecules. Carbohydrates, proteins
From the Structure and Function of the Ribosome to new Antibiotics
 From the Structure and Function of the Ribosome to new Antibiotics Crick s central dogma of molecular biology: DNA makes DNA makes RNA makes protein Jim Watson, 1964 J.A. Lake, 1976 (J.M.B. 105, 131)
From the Structure and Function of the Ribosome to new Antibiotics Crick s central dogma of molecular biology: DNA makes DNA makes RNA makes protein Jim Watson, 1964 J.A. Lake, 1976 (J.M.B. 105, 131)
Adenium Biotech. Management: - Peter Nordkild, MD, CEO, ex Novo Nordisk, Ferring, Egalet - Søren Neve, PhD, project director, ex Lundbeck, Novozymes
 Adenium Biotech Management: - Peter Nordkild, MD, CEO, ex Novo Nordisk, Ferring, Egalet - Søren Neve, PhD, project director, ex Lundbeck, Novozymes Board of Directors: - Stephan Christgau, PhD, chairman,
Adenium Biotech Management: - Peter Nordkild, MD, CEO, ex Novo Nordisk, Ferring, Egalet - Søren Neve, PhD, project director, ex Lundbeck, Novozymes Board of Directors: - Stephan Christgau, PhD, chairman,
Page 8/6: The cell. Where to start: Proteins (control a cell) (start/end products)
 Page 8/6: The cell Where to start: Proteins (control a cell) (start/end products) Page 11/10: Structural hierarchy Proteins Phenotype of organism 3 Dimensional structure Function by interaction THE PROTEIN
Page 8/6: The cell Where to start: Proteins (control a cell) (start/end products) Page 11/10: Structural hierarchy Proteins Phenotype of organism 3 Dimensional structure Function by interaction THE PROTEIN
Targeting PPIs in Oncology using a Fragment-Based Drug Discovery Approach Justin F. Bower CRUK Beatson Institute
 Targeting PPIs in Oncology using a Fragment-Based Drug Discovery Approach Justin F. Bower CRUK Beatson Institute Cancer Research UK Beatson Institute Establish an outstanding basic research programme into
Targeting PPIs in Oncology using a Fragment-Based Drug Discovery Approach Justin F. Bower CRUK Beatson Institute Cancer Research UK Beatson Institute Establish an outstanding basic research programme into
Antimicrobial resistance Fact sheet N 194 Updated April 2014
 Antimicrobial resistance Fact sheet N 194 Updated April 2014 Key facts Antimicrobial resistance (AMR) threatens the effective prevention and treatment of an ever-increasing range of infections caused by
Antimicrobial resistance Fact sheet N 194 Updated April 2014 Key facts Antimicrobial resistance (AMR) threatens the effective prevention and treatment of an ever-increasing range of infections caused by
Treatment of febrile neutropenia in patients with neoplasia
 Treatment of febrile neutropenia in patients with neoplasia George Samonis MD, PhD Medical Oncologist Infectious Diseases Specialist Professor of Medicine The University of Crete, Heraklion,, Crete, Greece
Treatment of febrile neutropenia in patients with neoplasia George Samonis MD, PhD Medical Oncologist Infectious Diseases Specialist Professor of Medicine The University of Crete, Heraklion,, Crete, Greece
Methicillin-Resistant Staphylococcus aureus (MRSA) S urveillance Report 2008 Background Methods
 Methicillin-Resistant Staphylococcus aureus (MRSA) Surveillance Report 2008 Oregon Active Bacterial Core Surveillance (ABCs) Office of Disease Prevention & Epidemiology Oregon Department of Human Services
Methicillin-Resistant Staphylococcus aureus (MRSA) Surveillance Report 2008 Oregon Active Bacterial Core Surveillance (ABCs) Office of Disease Prevention & Epidemiology Oregon Department of Human Services
BIOCHEMISTRY REVIEW. Overview of Biomolecules. Chapter 4 Protein Sequence
 BIOCHEMISTRY REVIEW Overview of Biomolecules Chapter 4 Protein Sequence 2 3 4 Are You Getting It?? A molecule of hemoglobin is compared with a molecule of lysozyme. Which characteristics do they share?
BIOCHEMISTRY REVIEW Overview of Biomolecules Chapter 4 Protein Sequence 2 3 4 Are You Getting It?? A molecule of hemoglobin is compared with a molecule of lysozyme. Which characteristics do they share?
Practice Problems 3. a. What is the name of the bond formed between two amino acids? Are these bonds free to rotate?
 Life Sciences 1a Practice Problems 3 1. Draw the oligopeptide for Ala-Phe-Gly-Thr-Asp. You do not need to indicate the stereochemistry of the sidechains. Denote with arrows the bonds formed between the
Life Sciences 1a Practice Problems 3 1. Draw the oligopeptide for Ala-Phe-Gly-Thr-Asp. You do not need to indicate the stereochemistry of the sidechains. Denote with arrows the bonds formed between the
INTERACTIONS OF GANODERIOL-F WITH ASPARTIC PROTEASES OF HIV AND PLASMEPSIN FOR ANTI-HIV AND ANTI-MALARIA DISCOVERY
 Innovare Academic Sciences International Journal of Pharmacy and Pharmaceutical Sciences ISSN- 0975-1491 Vol 6, Issue 5, 2014 Original Article INTERACTIONS OF GANODERIOL-F WITH ASPARTIC PROTEASES OF HIV
Innovare Academic Sciences International Journal of Pharmacy and Pharmaceutical Sciences ISSN- 0975-1491 Vol 6, Issue 5, 2014 Original Article INTERACTIONS OF GANODERIOL-F WITH ASPARTIC PROTEASES OF HIV
Microbiology - Problem Drill 16: Antibiotics. Question No. 1 of 10. Question. Feedback. Question
 Microbiology - Problem Drill 16: Antibiotics No. 1 of 10 1. An effective chemotherapeutic drug should have. (A) Low therapeutic index (B) More toxicity (C) Selective toxicity (D) Mutation inducing properties
Microbiology - Problem Drill 16: Antibiotics No. 1 of 10 1. An effective chemotherapeutic drug should have. (A) Low therapeutic index (B) More toxicity (C) Selective toxicity (D) Mutation inducing properties
Hydrogen Bonds and Biological Activities
 ugo Kubinyi, www.kubinyi.de ydrogen Bonds and Biological Activities ugo Kubinyi Germany EMail kubinyi@tonline.de omepage www.kubinyi.de ugo Kubinyi, www.kubinyi.de ature of the ydrogen Bond C=... a hydrogen
ugo Kubinyi, www.kubinyi.de ydrogen Bonds and Biological Activities ugo Kubinyi Germany EMail kubinyi@tonline.de omepage www.kubinyi.de ugo Kubinyi, www.kubinyi.de ature of the ydrogen Bond C=... a hydrogen
Bioinformation Volume 5
 Identification of sequence mutations affecting hemagglutinin specificity to sialic acid receptor in influenza A virus subtypes Usman Sumo Friend Tambunan*, Ramdhan Department of Chemistry, Faculty of Mathematics
Identification of sequence mutations affecting hemagglutinin specificity to sialic acid receptor in influenza A virus subtypes Usman Sumo Friend Tambunan*, Ramdhan Department of Chemistry, Faculty of Mathematics
Potential antibacterial drug targets for Quercetin and Rutin: An in silico study using AutoDock
 Available online at www.scholarsresearchlibrary.com Scholars Research Library Der Pharmacia Lettre, 2015, 7 (11):68-72 (http://scholarsresearchlibrary.com/archive.html) ISSN 0975-5071 USA CODEN: DPLEB4
Available online at www.scholarsresearchlibrary.com Scholars Research Library Der Pharmacia Lettre, 2015, 7 (11):68-72 (http://scholarsresearchlibrary.com/archive.html) ISSN 0975-5071 USA CODEN: DPLEB4
Supplementary Information
 Supplementary Information Two common structural motifs for TCR recognition by staphylococcal enterotoxins Karin Erica Johanna Rödström 1, Paulina Regenthal 1, Christopher Bahl 2, Alex Ford 2, David Baker
Supplementary Information Two common structural motifs for TCR recognition by staphylococcal enterotoxins Karin Erica Johanna Rödström 1, Paulina Regenthal 1, Christopher Bahl 2, Alex Ford 2, David Baker
Bacterial Infections of the Urinary System *
 OpenStax-CNX module: m64804 1 Bacterial Infections of the Urinary System * Douglas Risser This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 4.0 1 Learning
OpenStax-CNX module: m64804 1 Bacterial Infections of the Urinary System * Douglas Risser This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 4.0 1 Learning
News. Volume 8: November 2, 2009
 FightAIDS@Home News Volume 8: November 2, 2009 The FightAIDS@Home Project uses the volunteered computing power of the World Community Grid to test candidate compounds against the variations (or mutants
FightAIDS@Home News Volume 8: November 2, 2009 The FightAIDS@Home Project uses the volunteered computing power of the World Community Grid to test candidate compounds against the variations (or mutants
Prevalence of Extended Spectrum -Lactamases In E.coli and Klebsiella spp. in a Tertiary Care Hospital
 ISSN: 2319-7706 Volume 3 Number 10 (2014) pp. 474-478 http://www.ijcmas.com Original Research Article Prevalence of Extended Spectrum -Lactamases In E.coli and Klebsiella spp. in a Tertiary Care Hospital
ISSN: 2319-7706 Volume 3 Number 10 (2014) pp. 474-478 http://www.ijcmas.com Original Research Article Prevalence of Extended Spectrum -Lactamases In E.coli and Klebsiella spp. in a Tertiary Care Hospital
Multiple-Choice Questions Answer ALL 20 multiple-choice questions on the Scantron Card in PENCIL
 Multiple-Choice Questions Answer ALL 20 multiple-choice questions on the Scantron Card in PENCIL For Questions 1-10 choose ONE INCORRECT answer. 1. Which ONE of the following statements concerning the
Multiple-Choice Questions Answer ALL 20 multiple-choice questions on the Scantron Card in PENCIL For Questions 1-10 choose ONE INCORRECT answer. 1. Which ONE of the following statements concerning the
*Corresponding Author KEY WORDS Swine Flu, Hemagglutinin, Neuraminidase, Influenza A virus, Molecular Docking.
 Page33 IJPBS Volume 3 Issue 3 JUL-SEP 2013 33-45 Research Article Biological Sciences INHIBITORY ACTIVITY OF HEMAGGLUTININ AND NEURAMINIDASE PROTEIN CASEPASE ACTIVITY IN SWINE FLU [H1N1] Sanjay.V*, Tanusree
Page33 IJPBS Volume 3 Issue 3 JUL-SEP 2013 33-45 Research Article Biological Sciences INHIBITORY ACTIVITY OF HEMAGGLUTININ AND NEURAMINIDASE PROTEIN CASEPASE ACTIVITY IN SWINE FLU [H1N1] Sanjay.V*, Tanusree
1. Name: Dr Purva Mathur 2. Designation and Qualification: Assistant Professor M.D. Microbiology
 1. Name: Dr Purva Mathur 2. Designation and Qualification: Assistant Professor M.D. Microbiology Present employment: Faculty in Department of Laboratory Medicine JPNA Trauma Centre, AIIMS, since 26.09.2005
1. Name: Dr Purva Mathur 2. Designation and Qualification: Assistant Professor M.D. Microbiology Present employment: Faculty in Department of Laboratory Medicine JPNA Trauma Centre, AIIMS, since 26.09.2005
Available online Research Article
 Available online www.jocpr.com Journal of Chemical and Pharmaceutical Research, 2014, 6(12):860-865 Research Article ISSN : 0975-7384 CODEN(USA) : JCPRC5 In silico binding and interaction studies of aldose
Available online www.jocpr.com Journal of Chemical and Pharmaceutical Research, 2014, 6(12):860-865 Research Article ISSN : 0975-7384 CODEN(USA) : JCPRC5 In silico binding and interaction studies of aldose
320 MBIO Microbial Diagnosis. Aljawharah F. Alabbad Noorah A. Alkubaisi 2017
 320 MBIO Microbial Diagnosis Aljawharah F. Alabbad Noorah A. Alkubaisi 2017 Pathogens of the Urinary tract The urinary system is composed of organs that regulate the chemical composition and volume of
320 MBIO Microbial Diagnosis Aljawharah F. Alabbad Noorah A. Alkubaisi 2017 Pathogens of the Urinary tract The urinary system is composed of organs that regulate the chemical composition and volume of
Virtual screening using the ligand ZINC database for novel lipoxygenase-3 inhibitors
 www.bioinformation.net Hypothesis Volume 9(11) Virtual screening using the ligand ZINC database for novel lipoxygenase-3 inhibitors Monika 1 *, Janmeet Kour 1 & Kulwinder Singh 2 1Department of Biotechnology,
www.bioinformation.net Hypothesis Volume 9(11) Virtual screening using the ligand ZINC database for novel lipoxygenase-3 inhibitors Monika 1 *, Janmeet Kour 1 & Kulwinder Singh 2 1Department of Biotechnology,
Properties of amino acids in proteins
 Properties of amino acids in proteins one of the primary roles of DNA (but far from the only one!!!) is to code for proteins A typical bacterium builds thousands types of proteins, all from ~20 amino acids
Properties of amino acids in proteins one of the primary roles of DNA (but far from the only one!!!) is to code for proteins A typical bacterium builds thousands types of proteins, all from ~20 amino acids
Molecular Docking Study of Beta-Glucosidase with Cellobiose, Cellotetraose and Cellotetriose
 www.bioinformation.net Hypothesis Volume 9(16) Molecular Docking Study of Beta-Glucosidase with Cellobiose, Cellotetraose and Cellotetriose Nurul Bahiyah Ahmad Khairudin* & Nur Shima Fadhilah Mazlan Malaysia
www.bioinformation.net Hypothesis Volume 9(16) Molecular Docking Study of Beta-Glucosidase with Cellobiose, Cellotetraose and Cellotetriose Nurul Bahiyah Ahmad Khairudin* & Nur Shima Fadhilah Mazlan Malaysia
Statin inhibition of HMG-CoA reductase: a 3-dimensional view
 Atherosclerosis Supplements 4 (2003) 3/8 www.elsevier.com/locate/atherosclerosis Statin inhibition of HMG-CoA reductase: a 3-dimensional view Eva Istvan * Department of Molecular Microbiology, Howard Hughes
Atherosclerosis Supplements 4 (2003) 3/8 www.elsevier.com/locate/atherosclerosis Statin inhibition of HMG-CoA reductase: a 3-dimensional view Eva Istvan * Department of Molecular Microbiology, Howard Hughes
The Basics: A general review of molecular biology:
 The Basics: A general review of molecular biology: DNA Transcription RNA Translation Proteins DNA (deoxy-ribonucleic acid) is the genetic material It is an informational super polymer -think of it as the
The Basics: A general review of molecular biology: DNA Transcription RNA Translation Proteins DNA (deoxy-ribonucleic acid) is the genetic material It is an informational super polymer -think of it as the
Mutational studies of novel screened molecules against wild and mutated HIV-1 integrase using molecular docking studies
 Research Article Mutational studies of novel screened molecules against wild and mutated HIV-1 integrase using molecular docking studies Pawan Gupta 1,2 *, Prabha Garg 2 ABSTRACT Background and Aim: The
Research Article Mutational studies of novel screened molecules against wild and mutated HIV-1 integrase using molecular docking studies Pawan Gupta 1,2 *, Prabha Garg 2 ABSTRACT Background and Aim: The
Enhanced EARS-Net Surveillance 2017 First Half
 1 Enhanced EARS-Net Surveillance 2017 First Half In this report Main results for 2017, first half Breakdown of factors by organism and resistance subtype Device-association Data quality assessment Key
1 Enhanced EARS-Net Surveillance 2017 First Half In this report Main results for 2017, first half Breakdown of factors by organism and resistance subtype Device-association Data quality assessment Key
Cells N5 Homework book
 1 Cells N5 Homework book 2 Homework 1 3 4 5 Homework2 Cell Ultrastructure and Membrane 1. Name and give the function of the numbered organelles in the cell below: A E B D C 2. Name 3 structures you might
1 Cells N5 Homework book 2 Homework 1 3 4 5 Homework2 Cell Ultrastructure and Membrane 1. Name and give the function of the numbered organelles in the cell below: A E B D C 2. Name 3 structures you might
Synopsis of the thesis on
 Synopsis of the thesis on COMPUTATIONAL INVESTIGATION OF PROTEIN-LIGAND INTERACTIONS IN ANTI-DIABETIC AGENTS A dissertation submitted in the partial fulfillment for the award of the degree of DOCTOR OF
Synopsis of the thesis on COMPUTATIONAL INVESTIGATION OF PROTEIN-LIGAND INTERACTIONS IN ANTI-DIABETIC AGENTS A dissertation submitted in the partial fulfillment for the award of the degree of DOCTOR OF
Objective: You will be able to explain how the subcomponents of
 Objective: You will be able to explain how the subcomponents of nucleic acids determine the properties of that polymer. Do Now: Read the first two paragraphs from enduring understanding 4.A Essential knowledge:
Objective: You will be able to explain how the subcomponents of nucleic acids determine the properties of that polymer. Do Now: Read the first two paragraphs from enduring understanding 4.A Essential knowledge:
Proteins are sometimes only produced in one cell type or cell compartment (brain has 15,000 expressed proteins, gut has 2,000).
 Lecture 2: Principles of Protein Structure: Amino Acids Why study proteins? Proteins underpin every aspect of biological activity and therefore are targets for drug design and medicinal therapy, and in
Lecture 2: Principles of Protein Structure: Amino Acids Why study proteins? Proteins underpin every aspect of biological activity and therefore are targets for drug design and medicinal therapy, and in
VIRTUAL SCREENING and RAPID SCREENING ASSAYS in the discovery of NEW ANTIPARASITIC AGENTS
 VITUAL SCEEIG and APID SCEEIG ASSAYS in the discovery of EW ATIPAASITIC AGETS Stefania Ferrari DUG DISCVEY PCESS target identification and validation high-throughput technologies hit compounds identification
VITUAL SCEEIG and APID SCEEIG ASSAYS in the discovery of EW ATIPAASITIC AGETS Stefania Ferrari DUG DISCVEY PCESS target identification and validation high-throughput technologies hit compounds identification
Atypical Natural Killer T-cell receptor recognition of CD1d-lipid antigens supplementary Information.
 Atypical Natural Killer T-cell receptor recognition of CD1d-lipid antigens supplementary Information. Supplementary Figure 1. Phenotypic analysis of TRBV25-1 + and TRBV25-1 - CD1d-α-GalCerreactive cells.
Atypical Natural Killer T-cell receptor recognition of CD1d-lipid antigens supplementary Information. Supplementary Figure 1. Phenotypic analysis of TRBV25-1 + and TRBV25-1 - CD1d-α-GalCerreactive cells.
Development of C sporins. Beta-lactam antibiotics - Cephalosporins. Second generation C sporins. Targets - PBP s
 Beta-lactam antibiotics - Cephalosporins Development of C sporins Targets - PBP s Activity - Cidal - growing organisms (like the penicillins) Principles of action - Affinity for PBP s Permeability properties
Beta-lactam antibiotics - Cephalosporins Development of C sporins Targets - PBP s Activity - Cidal - growing organisms (like the penicillins) Principles of action - Affinity for PBP s Permeability properties
2. Which of the following amino acids is most likely to be found on the outer surface of a properly folded protein?
 Name: WHITE Student Number: Answer the following questions on the computer scoring sheet. 1 mark each 1. Which of the following amino acids would have the highest relative mobility R f in normal thin layer
Name: WHITE Student Number: Answer the following questions on the computer scoring sheet. 1 mark each 1. Which of the following amino acids would have the highest relative mobility R f in normal thin layer
CHAPTER 21: Amino Acids, Proteins, & Enzymes. General, Organic, & Biological Chemistry Janice Gorzynski Smith
 CHAPTER 21: Amino Acids, Proteins, & Enzymes General, Organic, & Biological Chemistry Janice Gorzynski Smith CHAPTER 21: Amino Acids, Proteins, Enzymes Learning Objectives: q The 20 common, naturally occurring
CHAPTER 21: Amino Acids, Proteins, & Enzymes General, Organic, & Biological Chemistry Janice Gorzynski Smith CHAPTER 21: Amino Acids, Proteins, Enzymes Learning Objectives: q The 20 common, naturally occurring
Workshop on Analysis and prediction of contacts in proteins
 Workshop on Analysis and prediction of contacts in proteins 1.09.09 Eran Eyal 1, Vladimir Potapov 2, Ronen Levy 3, Vladimir Sobolev 3 and Marvin Edelman 3 1 Sheba Medical Center, Ramat Gan, Israel; Departments
Workshop on Analysis and prediction of contacts in proteins 1.09.09 Eran Eyal 1, Vladimir Potapov 2, Ronen Levy 3, Vladimir Sobolev 3 and Marvin Edelman 3 1 Sheba Medical Center, Ramat Gan, Israel; Departments
Structural Analysis of Tyrosine SpecificProtein Kinase (TPK) and Interaction Studies with Inhibitors for Lung Cancer.
 Structural Analysis of Tyrosine SpecificProtein Kinase (TPK) and Interaction Studies with Inhibitors for Lung Cancer. Aishwarya Hipparagi 1, Shambhu M G 1, Kusum Paul 1 1 Department of Biotechnology, The
Structural Analysis of Tyrosine SpecificProtein Kinase (TPK) and Interaction Studies with Inhibitors for Lung Cancer. Aishwarya Hipparagi 1, Shambhu M G 1, Kusum Paul 1 1 Department of Biotechnology, The
Chapter 3: Amino Acids and Peptides
 Chapter 3: Amino Acids and Peptides BINF 6101/8101, Spring 2018 Outline 1. Overall amino acid structure 2. Amino acid stereochemistry 3. Amino acid sidechain structure & classification 4. Non-standard
Chapter 3: Amino Acids and Peptides BINF 6101/8101, Spring 2018 Outline 1. Overall amino acid structure 2. Amino acid stereochemistry 3. Amino acid sidechain structure & classification 4. Non-standard
THE EFFECT OF DIABETES MELLITUS ON THE CLINICAL AND MICRO-BIOLOGICAL OUTCOMES IN PATIENTS WITH ACUTE PYELONEPHRITIS
 American Journal of Infectious Diseases 10 (2): 71-76, 2014 ISSN: 1553-6203 2014 Science Publication doi:10.3844/ajidsp.2014.71.76 Published Online 10 (2) 2014 (http://www.thescipub.com/ajid.toc) THE EFFECT
American Journal of Infectious Diseases 10 (2): 71-76, 2014 ISSN: 1553-6203 2014 Science Publication doi:10.3844/ajidsp.2014.71.76 Published Online 10 (2) 2014 (http://www.thescipub.com/ajid.toc) THE EFFECT
Bioinformatics for molecular biology
 Bioinformatics for molecular biology Structural bioinformatics tools, predictors, and 3D modeling Structural Biology Review Dr Research Scientist Department of Microbiology, Oslo University Hospital -
Bioinformatics for molecular biology Structural bioinformatics tools, predictors, and 3D modeling Structural Biology Review Dr Research Scientist Department of Microbiology, Oslo University Hospital -
