Direct Evidence of Lower Viral Replication Rates In Vivo in Human Immunodeficiency Virus Type 2 (HIV-2) Infection than in HIV-1 Infection
|
|
|
- Mervin Fisher
- 6 years ago
- Views:
Transcription
1 JOURNAL OF VIROLOGY, May 2007, p Vol. 81, No X/07/$ doi: /jvi Copyright 2007, American Society for Microbiology. All Rights Reserved. Direct Evidence of Lower Viral Replication Rates In Vivo in Human Immunodeficiency Virus Type 2 (HIV-2) Infection than in HIV-1 Infection Adam MacNeil, 1 Abdoulaye Dieng Sarr, 1 Jean-Louis Sankalé, 1 Seema Thakore Meloni, 1 Souleymane Mboup, 2 and Phyllis Kanki 1 * Department of Immunology and Infectious Diseases, Harvard School of Public Health, Boston, Massachusetts, 1 and Laboratoire de Bacteriologie et Virologie, Universite Cheikh Anta Diop, Dakar, Senegal 2 Received 28 November 2006/Accepted 20 February 2007 Studies have shown that human immunodeficiency virus type 2 (HIV-2) is less pathogenic than HIV-1, with a lower rate of disease progression. Similarly, plasma viral loads are lower in HIV-2 infection, suggesting that HIV-2 replication is restricted in vivo in comparison to that of HIV-1. However, to date, in vivo studies characterizing replication intermediates in the viral life cycle of HIV-2 have been limited. In order to test the hypothesis that HIV-2 has a lower replication rate in vivo than HIV-1 does, we quantified total viral DNA, integrated proviral DNA, cell-associated viral mrna, and plasma viral loads in peripheral blood samples from groups of therapy-naïve HIV-1-infected (n 21) and HIV-2-infected (n 18) individuals from Dakar, Senegal, with CD4 T-cell counts of >200/ l. Consistent with our previous findings, total viral DNA loads were similar between HIV-1 and HIV-2 and plasma viral loads were higher among HIV-1-infected individuals. Proportions of DNA in the integrated form were also similar between these viruses. In contrast, levels of viral mrna were lower in HIV-2 infection. Our study indicates that HIV-2 is able to establish a stable, integrated proviral infection in vivo, but that accumulation of viral mrna is attenuated in HIV-2 infection relative to that in HIV-1 infection. The differences in viral mrna are consistent with the differences in plasma viral loads between HIV-1 and HIV-2 and suggest that lower plasma viral loads, and possibly the attenuated pathogenesis of HIV-2, can be explained by lower rates of viral replication in vivo. Two closely related human lentiviruses, human immunodeficiency virus type 1 (HIV-1) and HIV-2, have been shown to cause AIDS. However, it is now well recognized that the in vivo pathogenicity of HIV-2 is attenuated relative to that of HIV-1, with significantly lower rates of disease progression and transmission (20, 27, 28, 43). Consistent with these observations, plasma viral loads are significantly lower in people infected with HIV-2 (1, 2, 36, 37, 42). Interestingly, studies have shown that quantities of total viral DNA in peripheral blood mononuclear cells (PBMCs) are similar in people infected with HIV-1 and people infected with HIV-2 (5, 32, 36). Within an infected cell, HIV DNA is found in a number of forms, including integrated proviral DNA, unintegrated linear viral DNA, and unintegrated one- and two-long-terminal-repeat (LTR) episomal viral DNA (31). The integration of proviral DNA is necessary for viral replication (10, 11, 21, 44). In addition, the integration of proviral DNA can result in a stable long-term infection. HIV-1 has been shown to establish latent infection in vivo within resting memory CD4 T cells, a reservoir in which the integrated proviral genome can persist for decades (reviewed in references 22 and 35). Although integration is necessary for viral replication, studies of HIV-1 have shown that the majority of in vivo viral DNA exists in the * Corresponding author. Mailing address: Department of Immunology and Infectious Diseases, Harvard School of Public Health, 651 Huntington Ave., Boston, MA Phone: (617) Fax: (617) pkanki@hsph.harvard.edu. Published ahead of print on 28 February linear unintegrated form (8), while integrated proviral DNA accounts for a minor fraction of the total DNA load in vivo (6, 8, 9, 15, 16). While higher plasma viral loads in HIV-1 infection suggest higher replication rates, no studies have clearly shown a difference in replication in vivo between HIV-1 and HIV-2. Additionally, previous studies comparing viral DNA in HIV-1 and HIV-2 infection have focused on total viral DNA and, to date, no studies have quantified integrated proviral DNA in people infected with HIV-2. In this study, we measured viral life cycle intermediates (integrated proviral DNA and viral mrna) in conjunction with total viral DNA loads and plasma viral loads in people infected with HIV-1 or HIV-2 to determine whether quantitative differences that can explain the lower plasma viral loads observed in HIV-2 infection exist at a specific point of the viral life cycle. MATERIALS AND METHODS Sample acquisition. PBMC samples were obtained from a cohort of female sex workers in Dakar, Senegal, that have been followed since Epidemiologic and clinical aspects of this cohort have previously been described elsewhere (19). All subjects signed informed consents and participated in protocols approved by the Counseil National de Lutte Contre le Sida Comite Ethique et Juridique and the Harvard School of Public Health Human Subjects Committee. CD4 T-cell counts were determined, and serum samples were diagnosed for HIV-1- and HIV-2-specific antibodies as previously described (19). All subjects enrolled in this study were antiretroviral therapy naïve and had CD4 T-cell counts above 200/ l at the time of sample acquisition. DNA and mrna extraction. Cryopreserved PBMC samples were thawed and rested overnight in RPMI 1640 medium supplemented with 10% (vol/vol) fetal bovine serum and 1% antibiotics to ensure sample viability and remove dead 5325
2 5326 MACNEIL ET AL. J. VIROL. cells. The following day, PBMC samples were divided equally, with half used for the extraction of DNA and half used for the extraction of mrna. DNA was extracted (blood and cell culture DNA mini kit; QIAGEN), and DNA concentrations were determined by an optical density reading at 260 nm. Polyadenylated mrna was extracted (Oligotex mrna direct mini kit; QIAGEN) and immediately converted to cdna (TaqMan reverse transcription reagents; Applied Biosystems) by using an equal-volume mixture of random hexamer and poly(t) primers. Quantification of total viral DNA and viral mrna. HIV-1 total viral DNA and mrna were quantified using a previously described real-time PCR assay (25) that amplifies a highly conserved region corresponding to the HIV-1 LTR-gag junction. For the quantification of HIV-2 total viral DNA and mrna, we developed a novel HIV-2 real-time PCR assay that amplifies a highly conserved region corresponding to the HIV-2 LTR-gag junction. To construct a quantification standard, a fragment of HIV-2 LTR-gag was amplified by PCR from a previously cloned HIV-2 DNA sample using primers AM2gag1f (AGACCCTG GTCTGTTAGGACCCTT) and AM2gag1r (CAATTCATTCGCTGCCCA CAC) cloned into an expression vector (pcr2.1; Invitrogen) and transformed into competent cells (TOP10; Invitrogen). Cloned plasmid was purified (SNAP MiniPrep kit; Invitrogen), and the concentration of purified plasmid was determined by an optical density reading at 260 nm. For the quantification of HIV-2 samples, real-time PCRs were performed by using primers HIV2gagF (CCAA CCACGACGGAGTGCTC) and HIV2gagR (CTCTCAAGACGGAGTTTCT CGC) and detected by using the probe HIV2gagP (AGGCCTCCGGGTGAAG GTAAG), which contained a 5 6-carboxyfluorescein fluorescent reporter and a 3 MGB nonfluorescent quencher. For HIV-1 and HIV-2 real-time PCR assays, real-time PCR reagents and reaction conditions were as previously described (25). Viral mrna was standardized to cellular beta-actin mrna. Beta-actin mrna was quantified using commercially available reagents (TaqMan B-actin detection reagents; Applied Biosystems) according to manufacturer recommendations. Quantification of integrated proviral DNA. Quantities of integrated proviral DNA were determined for HIV-1 and for HIV-2 by using an endpoint dilution Alu-PCR approach as previously described (9). A series of six threefold serial dilutions were performed in duplicate from a starting quantity of 1 g genomic DNA, and Alu-PCR was performed using primers Alu1 (GCCTCCCAAAGTG CTGGGATTACAG) (34) and AM1C1r (CTTAATACTGACGCTCTCGCA CCC) for HIV-1 samples and Alu1 and AM2C1r (AAGGGTCCTAACAGAC CAGGGTCT) for HIV-2 samples in a total volume of 100 l under the following reaction conditions: denaturation at 94 for 5 min; 35 PCR cycles of 94 C for 30 s, 60 C for 30 s, and 72 C for 3 min; and a final extension at 72 C for 7 min. The amplification of integrated proviral DNA was assayed by real-time PCR with 5 l of the first-round Alu-PCR product by using previously described HIV-1 and HIV-2 real-time PCR assays (25). Real-time PCR assays were performed for each sample in parallel with equivalent concentrations of nonamplified genomic DNA from that respective sample to control for nonamplified viral DNA carryover from the Alu-PCR. Integrated proviral DNA copy numbers were determined by using the QUALITY program (38). Quantification of plasma viral loads. HIV-1 plasma viral loads were determined by using a commercially available assay (Amplicor HIV-1 monitor test, version 1.5; Roche). To determine HIV-2 viral loads, we developed an in-house real-time PCR viral load assay. Briefly, we amplified a fragment of HIV-2 LTR-gag from a previously cloned HIV-2 DNA sample using primers HIV2gagF and AM2gag1r and synthesized a T7 promoter-driven in vitro transcription construct by using commercially available reagents (BLOCK-iT RNAi TOPO transcription kit; Invitrogen) and primer AM2gag1r. HIV-2 RNA was generated by in vitro transcription (MEGAshortscript; Ambion). In vitro-transcribed HIV-2 RNA was purified (MEGAclear; Ambion), treated with DNase (New England Biolabs), and repurified, and the concentration of in vitro-transcribed HIV-2 RNA was determined by an optical density reading at 260 nm. To create a full-length HIV-2 genomic RNA standard, full-length HIV-2 genomic RNA from a previously described in vitro infection (25) was extracted (QIAamp viral RNA mini kit; QIAGEN), treated with DNase (New England Biolabs), and repurified (MEGAclear; Ambion). Quantities of full-length HIV-2 genomic RNA were determined by using the in vitro-transcribed HIV-2 RNA as a quantitative standard in a two-step, real-time reverse transcription-pcr (TaqMan reverse transcription reagents and TaqMan universal PCR master mix; Applied Biosystems) using the primer HIV2gagR for the cdna synthesis reaction and the HIV-2 LTR-gag real-time PCR primers and probe described above. For the quantification of plasma viral loads, RNA standard curves were generated from the full-length HIV-2 genomic RNA standard, spiked to noninfected lysis buffer, and extracted in parallel with 280 l of plasma from HIV-2-infected subjects (QIAamp viral RNA mini kit; QIAGEN). HIV-2 plasma viral loads were quantified relative to the full-length HIV-2 genomic RNA standard curve in a twostep, real-time reverse transcription-pcr, as described above. Statistical analyses. Statistical analyses were performed using SAS 9.1. For analytical purposes, HIV-2-infected subjects with plasma viral loads of 400/ml were assigned values of 400/ml in accordance with the detection limit of the HIV-1 plasma viral load assay (Amplicor HIV-1 monitor test, version 1.5; Roche). The Wilcoxon rank sum test was used for an unadjusted comparison of CD4 T-cell counts, age, total viral DNA, integrated proviral DNA, viral mrna, plasma viral load, and percentage of total viral DNA integrated between HIV-1- and HIV-2-infected subjects; two-sided P values are reported. Because of the limited number of study subjects available, we were unable to directly match HIV-1-infected and HIV-2-infected individuals on the basis of CD4 T-cell count and age. Alternatively, to adjust for CD4 T-cell counts and age in the comparison of HIV-1-infected and HIV-2- infected individuals, regressions were performed using the ROBUSTREG procedure, which provides stable results in the presence of outlying values, and values of total viral DNA, integrated proviral DNA, viral mrna, and plasma viral loads were log transformed. Correlations of total viral DNA, integrated proviral DNA, percentage of total DNA integrated, and viral mrna with plasma viral loads were examined using the Pearson s correlation coefficient (for HIV-1) and Spearman s correlation coefficient (for HIV-2) by using log-transformed values of total viral DNA, integrated proviral DNA, viral mrna, and plasma viral loads. RESULTS For this study, we examined 21 subjects infected with HIV-1 and 18 subjects infected with HIV-2 from Dakar, Senegal. All had CD4 T-cell counts above 200/ l and were antiretroviral therapy naïve. Although all subjects were in the asymptomatic phase of infection, CD4 T-cell counts were lower (P ) in HIV-1-infected subjects (median and interquartile range values were 560 and 404 to 614 cells/ l, respectively [for HIV-2-infected subjects, median and interquartile range values were 946 and 534 to 1,503 cells/ l, respectively]). In addition, HIV-1-infected subjects (median and interquartile range values were 34.9 and 27.4 to 42.2 years, respectively) were younger (P ) than HIV-2-infected subjects (median and interquartile range values were 44.2 and 40.1 to 48.9 years, respectively). Previous studies have indicated that plasma viral loads are higher in people infected with HIV-1 than in people infected with HIV-2 (1, 2, 36, 37, 42), despite their having similar amounts of total viral DNA (5, 32, 36). For this study, we quantified total viral DNA by using real-time PCR assays for HIV-1 and HIV-2 that amplified highly conserved regions of the viral genomes that correspond to the LTR-gag junction. Twenty of 21 HIV-1-infected and 17 of 18 HIV-2-infected individuals had detectable total viral DNA. Among study subjects, total viral DNA loads were slightly higher in HIV-1- infected subjects than in HIV-2-infected subjects (Fig. 1). However, after adjusting for CD4 T-cell counts and age, there was no statistical difference in total viral DNA levels between HIV-1- and HIV-2-infected subjects. In contrast, in both unadjusted and adjusted analyses, plasma viral loads were significantly higher in HIV-1-infected subjects than those in HIV-2-infected subjects. In order to compare amounts of viral mrna between HIV-1 and HIV-2, viral transcripts isolated though the extraction of polyadenylated cellular mrna were quantified using the same real-time PCR assays that were used to quantify total viral DNA. Unlike for HIV-1-infected subjects, in which viral transcripts were detectable for the majority of subjects, amounts of viral mrna were below the limit of detection for
3 VOL. 81, 2007 LOW HIV-2 REPLICATION RATE IN VIVO 5327 FIG. 1. Total viral DNA, integrated proviral DNA, viral mrna, and plasma viral loads for HIV-1-infected and HIV-2-infected subjects. Units refer to copies per 10 6 PBMCs (total viral DNA and integrated proviral DNA), copies per 10 5 copies beta-actin (viral mrna), and viral load per milliliter of plasma (plasma viral load). Unadjusted P values refer to Wilcoxon rank sum two-sided P values. Adjusted P values refer to robust regression P values adjusted for CD4 T-cell count and age. Differences between HIV-1 and HIV-2 that reach statistical significance are in bold. most HIV-2-infected individuals. The difference in quantities of viral mrna between HIV-1 and HIV-2 were statistically significant in both unadjusted and adjusted analyses. In order to compare the amounts of integrated proviral DNA in PBMCs of subjects infected with HIV-1 with those of subjects infected with HIV-2, we used a previously described endpoint dilution Alu-PCR approach (9) that selectively amplifies integrated proviral DNA by using a virusspecific primer and a primer specific to conserved sequences in Alu elements. Alu elements are retrotransposons, and approximately 1.2 million copies of Alu elements are found in the human genome, accounting for 10 to 11% of the entire genome (18). To avoid unequal amplification between HIV-1 samples and HIV-2 samples, HIV-1 and HIV-2 primers were approximately equidistant from the 5 end of the respective viral genomes, and have previously been shown to amplify integrated proviral DNA (25). Twenty of 21 HIV- 1-infected subjects and 16 of 18 HIV-2-infected subjects had detectable integrated proviral DNA. Amounts of integrated proviral DNA were larger for HIV-1 than HIV-2, and values were slightly different after adjusting for CD4 T-cell counts and age. Previous studies of HIV-1 indicate that only a small proportion of total viral DNA is integrated in vivo (6, 8, 9, 15, 16). Among study subjects with detectable quantities of integrated proviral DNA, we observed similar results (Fig. 2) for both HIV-1 (the median percentage of total DNA integrated was 9.2%) and HIV-2 (the median percentage of total DNA integrated was 3.1%). There was no statistically significant difference in the percentage of DNA integrated between HIV-1- infected subjects and HIV-2-infected subjects. We examined the relationship of total viral DNA, integrated proviral DNA, percentage of total DNA integrated, and viral mrna to plasma viral loads for HIV-1-infected subjects and HIV-2-infected subjects (Table 1). As expected, for HIV-1, quantities of total viral DNA, integrated proviral DNA, and viral mrna all significantly correlated with plasma viral loads. In contrast, for HIV-1, there was no correlation between the percentage of total viral DNA integrated and plasma viral loads. Among HIV-2-infected subjects, we observed no significant correlation between plasma viral loads and total viral DNA or integrated proviral DNA. However, despite the small number of HIV-2-infected subjects having detectable mrna, quantities of viral mrna positively correlated with plasma viral loads among HIV-2-infected subjects. As with HIV-1, there was no relationship between the percentage of total viral DNA integrated and plasma viral loads. FIG. 2. Percentage of total DNA in the integrated form for HIV- 1-infected and HIV-2-infected subjects. Values are plotted for all subjects who had detectable integrated proviral DNA (n 20 for HIV-1; n 16 for HIV-2). Unadjusted P values refer to Wilcoxon rank sum two-sided P values. Adjusted P values refer to robust regression P values adjusted for CD4 T-cell count and age.
4 5328 MACNEIL ET AL. J. VIROL. TABLE 1. Correlation of total viral DNA, integrated proviral DNA, percentage of total DNA integrated, and viral mrna, with plasma viral loads for HIV-1-infected subjects and HIV-2-infected subjects Virus Total viral DNA Correlation statistics for viral load and a : Integrated proviral DNA DISCUSSION % of total DNA integrated Viral mrna r P r P r P r P HIV HIV a Correlations were examined using log-transformed total viral DNA, integrated proviral DNA, viral mrna, and plasma viral loads. Correlation coefficients (r) and P values refer to Pearson s correlation coefficient (for HIV-1) and Spearman s correlation coefficient (for HIV-2). Correlations that reach statistical significance are in bold. In HIV-1 infection, higher plasma viral loads are predictors of more rapid disease progression (3, 23, 29, 30, 33). Plasma viral loads have been clearly demonstrated to be lower in HIV-2 infection than in HIV-1 infection (1, 2, 36, 37, 42), possibly explaining the lower rate of disease progression of HIV-2 (12). This observation, in conjunction with similar total viral DNA loads in people infected with these viruses (5, 32, 36), suggests that HIV-2 maintains a lower rate of replication in vivo. However, host factors, such as differences in adaptive immune responses, could also account for the difference in plasma viral loads between these viruses. To address the hypothesis that HIV-2 has a lower replication rate in vivo than HIV-1 does, we quantified replication intermediates in the viral life cycles of HIV-1 and HIV-2 in vivo. Consistent with previous reports, we observed significantly higher plasma viral loads in people infected with HIV-1 than in people infected with HIV-2, and after an adjustment for CD4 T-cell counts and age, we observed no difference in total viral DNA. The interpretation of total viral DNA with regard to viral replication is difficult, as studies have shown that only a minority of HIV-1 DNA is actually integrated in vivo (6, 8, 9, 15, 16). Furthermore, this measurement includes one- and two-ltr circles, which are dead-end products. To better compare the nature of viral DNA between HIV-1 and HIV-2 in vivo, we quantified levels of integrated proviral DNA. We observed only modestly higher levels of integrated proviral DNA in HIV-1-infected individuals, and after comparing the percentage of viral DNA in the integrated form, we observed no difference between HIV-1 and HIV-2. While integrated proviral DNA clearly represents a minor proportion of the overall viral DNA load in HIV-2 infection, our data indicate that the integration of viral DNA is not overtly impaired in HIV-2 infection relative to that in HIV-1. The HIV-1 and HIV-2 real-time PCR assays used in this study to quantify viral mrna amplify a fragment of the LTRgag junction, which is encoded in processive unspliced viral mrna. Levels of viral mrna in HIV-2-infected individuals were significantly lower than those in HIV-1-infected individuals. This difference is consistent with the lower plasma viral loads among the HIV-2-infected individuals and implies that lower plasma viral loads can be accounted for by lower viral mrna levels in HIV-2 infection in vivo. Hermankova et al. (14) reported extremely little accumulation of processive viral mrna in latently infected resting CD4 T cells during HIV-1 infection in vivo. Given the observations that HIV-2 is able to establish integrated proviral infection, yet transcription remains limited, HIV-2 may have a much higher propensity for establishing latent infection in vivo than HIV-1 does. In support of this notion, the Nef protein of HIV-2 has been shown to downmodulate T-cell receptor-cd3 within infected cells and block responsiveness to T-cell activation, whereas the HIV-1 Nef protein fails to do so, contributing to increased cell death in HIV-1-infected cells and possibly increased persistence in HIV-2-infected cells (40). Among HIV-1-infected individuals, total viral DNA, integrated proviral DNA, and viral mrna all correlated with plasma viral loads. Of these measures, viral mrna had the strongest correlation with plasma viral load. Interestingly, despite the small number of individuals with detectable mrna, viral mrna also correlated with plasma viral load for HIV-2. These findings support the notion that viral mrna is a relevant marker of viral replication. For both HIV-1 and HIV-2, no correlation was observed between the percentage of total viral DNA integrated and plasma viral load, indicating that the proportion of integrated proviral DNA is not a pertinent measure for viral pathogenesis. Our data suggest that HIV-2 is able to establish a stable integrated proviral infection within PBMCs and that replication is attenuated at some point following integration, but prior to the accumulation of late transcripts. The reason for the block remains unknown. HIV-1 and HIV-2 have different regulatory elements within their promoter regions and have been shown to respond differently to cellular stimuli (13), suggesting possible differences in transcriptional initiation between these viruses. In addition, integration within heterochromatin has been shown to be associated with transcriptional inhibition and viral latency during in vitro HIV-1 infection (17, 24) and we have recently reported evidence of proviral integration within heterochromatin in a higher proportion of people infected with HIV-2 than with HIV-1 (25). Differences between HIV-1 and HIV-2 may also exist following transcriptional initiation. For instance, the HIV-2 LTR is significantly larger than the HIV-1 LTR, has been shown to undergo 5 LTR splicing, and has a complex secondary structure that may affect transcriptional trans-activation (4, 7). While the exact effect this has on mrna accumulation in vivo remains unknown, it is possible that differences in the LTR structure result in less-efficient transcriptional elongation and/or the switch from early to late transcripts in HIV-2 infection. To our knowledge, this is the first study to report direct evidence of a difference in the replication rates of HIV-1 and HIV-2 in vivo. In a recent report from our lab, we prospectively examined viral evolution over a decade of infection among eight HIV-2-infected individuals and observed significantly lower rates of viral evolution for HIV-2 than for HIV-1 (26), which also supports the notion that HIV-2 replication is attenuated in vivo in comparison to HIV-1 replication. It should be noted that the results of the current study do not exclude the possibility that differences in adaptive immune responses may also contribute to the lower plasma viral loads observed with HIV-2 infection. Recent data from our laboratory indicate a
5 VOL. 81, 2007 LOW HIV-2 REPLICATION RATE IN VIVO 5329 broad neutralizing response in people infected with HIV-2 (39). While it is well recognized that disease progression is significantly slower in HIV-2 infection, the virologic explanations for the attenuated pathogenicity of HIV-2 have remained unclear. Studies in vitro imply no difference in cytopathogenicity between HIV-1 and HIV-2 (41). However, cytopathogenicity is partially a function of viral replication. Through characterizing intermediates of the viral life cycles of HIV-1 and HIV-2 in people infected with these viruses, we have found direct evidence that replication rates are lower for HIV-2 than for HIV-1 in vivo. Additionally, we note that viral replication rates for HIV-1 and HIV-2 correspond with relative rates of disease progression for these viruses, implying that the difference in the pathogenicity of HIV-1 and HIV-2 may be explained by differences in viral replication. Since this difference appears to be manifested after the viral integration step, our data suggest that HIV-2 is capable of efficient cellular infection but has a propensity for viral latency. These observations are consistent with in vivo and epidemiologic population data, whereby HIV-2 persists despite a significant impairment in replicative capacity relative to HIV-1. ACKNOWLEDGMENTS We thank Shaun Rodriguez for helpful suggestions and Beth Chaplin and Christopher Mullins for technical assistance. This work was supported by grants from the National Institutes of Health (RO1AI46187, AI A1, and 5 T32 AI007638). REFERENCES 1. Alabi, A. S., S. Jaffar, K. Ariyoshi, T. Blanchard, M. Schim van der Loeff, A. A. Awasana, T. Corrah, S. Sabally, R. Sarge-Njie, F. Cham-Jallow, A. Jaye, N. Berry, and H. Whittle Plasma viral load, CD4 cell percentage, HLA and survival of HIV-1, HIV-2, and dually infected Gambian patients. AIDS 17: Andersson, S., H. Norrgren, Z. Da Silva, A. Biague, S. Bamba, S. Kwok, C. Christopherson, G. Biberfeld, and J. Albert Plasma viral load in HIV-1 and HIV-2 singly and dually infected individuals in Guinea-Bissau, West Africa: significantly lower plasma virus set point in HIV-2 infection than in HIV-1 infection. Arch. Intern. Med. 160: Arduino, J. M., M. A. Fischl, K. Stanley, A. C. Collier, and D. Spiegelman Do HIV type 1 RNA levels provide additional prognostic value to CD4 T lymphocyte counts in patients with advanced HIV type 1 infection? AIDS Res. Hum. Retrovir. 17: Berkhout, B Structural features in TAR RNA of human and simian immunodeficiency viruses: a phylogenetic analysis. Nucleic Acids Res. 20: Berry, N., K. Ariyoshi, O. Jobe, P. T. Ngum, T. Corrah, A. Wilkins, H. Whittle, and R. Tedder HIV type 2 proviral load measured by quantitative polymerase chain reaction correlates with CD4 lymphopenia in HIV type 2-infected individuals. AIDS Res. Hum. Retrovir. 10: Calcaterra, S., G. Cappiello, A. Di Caro, A. R. Garbuglia, and A. Benedetto Comparative analysis of total and integrated HIV-1 DNA in peripheral CD4 lymphocytes and monocytes after long treatment with HAART. J. Infect. 43: Chatterjee, P., A. Garzino-Demo, P. Swinney, and S. K. Arya Human immunodeficiency virus type 2 multiply spliced transcripts. AIDS Res. Hum. Retrovir. 9: Chun, T. W., L. Carruth, D. Finzi, X. Shen, J. A. DiGiuseppe, H. Taylor, M. Hermankova, K. Chadwick, J. Margolick, T. C. Quinn, Y. H. Kuo, R. Brookmeyer, M. A. Zeiger, P. Barditch-Crovo, and R. F. Siliciano Quantification of latent tissue reservoirs and total body viral load in HIV-1 infection. Nature 387: Chun, T. W., L. Stuyver, S. B. Mizell, L. A. Ehler, J. A. Mican, M. Baseler, A. L. Lloyd, M. A. Nowak, and A. S. Fauci Presence of an inducible HIV-1 latent reservoir during highly active antiretroviral therapy. Proc. Natl. Acad. Sci. USA 94: Engelman, A., G. Englund, J. M. Orenstein, M. A. Martin, and R. Craigie Multiple effects of mutations in human immunodeficiency virus type 1 integrase on viral replication. J. Virol. 69: Englund, G., T. S. Theodore, E. O. Freed, A. Engelman, and M. A. Martin Integration is required for productive infection of monocyte-derived macrophages by human immunodeficiency virus type 1. J. Virol. 69: Gottlieb, G. S., P. S. Sow, S. E. Hawes, I. Ndoye, M. Redman, A. M. Coll-Seck, M. A. Faye-Niang, A. Diop, J. M. Kuypers, C. W. Critchlow, R. Respess, J. I. Mullins, and N. B. Kiviat Equal plasma viral loads predict a similar rate of CD4 T-cell decline in human immunodeficiency virus (HIV) type 1- and HIV-2-infected individuals from Senegal, West Africa. J. Infect. Dis. 185: Hannibal, M. C., D. M. Markovitz, N. Clark, and G. J. Nabel Differential activation of human immunodeficiency virus type 1 and 2 transcription by specific T-cell activation signals. J. Virol. 67: Hermankova, M., J. D. Siliciano, Y. Zhou, D. Monie, K. Chadwick, J. B. Margolick, T. C. Quinn, and R. F. Siliciano Analysis of human immunodeficiency virus type 1 gene expression in latently infected resting CD4 T lymphocytes in vivo. J. Virol. 77: Ibanez, A., T. Puig, J. Elias, B. Clotet, L. Ruiz, and M. A. Martinez Quantification of integrated and total HIV-1 DNA after long-term highly active antiretroviral therapy in HIV-1-infected patients. AIDS 13: Izopet, J., M. Cazabat, C. Pasquier, K. Sandres-Saune, E. Bonnet, B. Marchou, P. Massip, and J. Puel Evolution of total and integrated HIV-1 DNA and change in DNA sequences in patients with sustained plasma virus suppression. Virology 302: Jordan, A., D. Bisgrove, and E. Verdin HIV reproducibly establishes a latent infection after acute infection of T cells in vitro. EMBO J. 22: Jurka, J Evolutionary impact of human Alu repetitive elements. Curr. Opin. Genet. Dev. 14: Kanki, P., S. M Boup, R. Marlink, K. Travers, C. C. Hsieh, A. Gueye, C. Boye, J. L. Sankale, C. Donnelly, W. Leisenring, et al Prevalence and risk determinants of human immunodeficiency virus type 2 (HIV-2) and human immunodeficiency virus type 1 (HIV-1) in West African female prostitutes. Am. J. Epidemiol. 136: Kanki, P. J Human immunodeficiency virus type 2 (HIV-2). AIDS Rev. 1: LaFemina, R. L., C. L. Schneider, H. L. Robbins, P. L. Callahan, K. LeGrow, E. Roth, W. A. Schleif, and E. A. Emini Requirement of active human immunodeficiency virus type 1 integrase enzyme for productive infection of human T-lymphoid cells. J. Virol. 66: Lassen, K., Y. Han, Y. Zhou, J. Siliciano, and R. F. Siliciano The multifactorial nature of HIV-1 latency. Trends Mol. Med. 10: Lefrère, J. J., F. Roudot-Thoraval, M. Mariotti, M. Thauvin, J. Lerable, J. Salpetrier, and L. Morand-Joubert The risk of disease progression is determined during the first year of human immunodeficiency virus type 1 infection. J. Infect. Dis. 177: Lewinski, M. K., D. Bisgrove, P. Shinn, H. Chen, C. Hoffmann, S. Hannenhalli, E. Verdin, C. C. Berry, J. R. Ecker, and F. D. Bushman Genome-wide analysis of chromosomal features repressing human immunodeficiency virus transcription. J. Virol. 79: MacNeil, A., J. L. Sankale, S. T. Meloni, A. D. Sarr, S. Mboup, and P. Kanki Genomic sites of human immunodeficiency virus type 2 (HIV-2) integration: similarities to HIV-1 in vitro and possible differences in vivo. J. Virol. 80: MacNeil, A., J. L. Sankale, S. T. Meloni, A. D. Sarr, S. Mboup, and P. Kanki Long-term intrapatient viral evolution during HIV-2 infection. J. Infect. Dis. 195: Marlink, R., P. Kanki, I. Thior, K. Travers, G. Eisen, T. Siby, I. Traore, C. C. Hsieh, M. C. Dia, E. H. Gueye, et al Reduced rate of disease development after HIV-2 infection as compared to HIV-1. Science 265: Marlink, R. G., D. Ricard, S. M Boup, P. J. Kanki, J. L. Romet-Lemonne, I. N Doye, K. Diop, M. A. Simpson, F. Greco, M. J. Chou, et al Clinical, hematologic, and immunologic cross-sectional evaluation of individuals exposed to human immunodeficiency virus type-2 (HIV-2). AIDS Res. Hum. Retrovir. 4: Mellors, J. W., A. Munoz, J. V. Giorgi, J. B. Margolick, C. J. Tassoni, P. Gupta, L. A. Kingsley, J. A. Todd, A. J. Saah, R. Detels, J. P. Phair, and C. R. Rinaldo, Jr Plasma viral load and CD4 lymphocytes as prognostic markers of HIV-1 infection. Ann. Intern. Med. 126: Mellors, J. W., C. R. Rinaldo, Jr., P. Gupta, R. M. White, J. A. Todd, and L. A. Kingsley Prognosis in HIV-1 infection predicted by the quantity of virus in plasma. Science 272: Meyerhans, A., T. Breinig, J. Vartanian, and S. Wain-Hobson Forms and function of intracellular HIV DNA, p In T. Leitner, B. Foley, B. Hahn, P. Marx, F. McCutchan, J. W. Mellors, S. Wolinsky, and B. Korber (ed.), HIV sequence compendium Theoretical Biology and Biophysics Group, Los Alamos, NM. 32. Norrgren, H., S. Marquina, T. Leitner, P. Aaby, M. Melbye, A. G. Poulsen, O. Larsen, F. Dias, D. Escanilla, S. Andersson, J. Albert, and A. Naucler HIV-2 genetic variation and DNA load in asymptomatic carriers and AIDS cases in Guinea-Bissau. J. Acquir. Immune Defic. Syndr. Hum. Retrovirol. 16: O Brien, T. R., W. A. Blattner, D. Waters, E. Eyster, M. W. Hilgartner, A. R.
6 5330 MACNEIL ET AL. J. VIROL. Cohen, N. Luban, A. Hatzakis, L. M. Aledort, P. S. Rosenberg, W. J. Miley, B. L. Kroner, and J. J. Goedert Serum HIV-1 RNA levels and time to development of AIDS in the Multicenter Hemophilia Cohort Study. JAMA 276: O Doherty, U., W. J. Swiggard, D. Jeyakumar, D. McGain, and M. H. Malim A sensitive, quantitative assay for human immunodeficiency virus type 1 integration. J. Virol. 76: Persaud, D., Y. Zhou, J. M. Siliciano, and R. F. Siliciano Latency in human immunodeficiency virus type 1 infection: no easy answers. J. Virol. 77: Popper, S. J., A. D. Sarr, A. Gueye-Ndiaye, S. Mboup, M. E. Essex, and P. J. Kanki Low plasma human immunodeficiency virus type 2 viral load is independent of proviral load: low virus production in vivo. J. Virol. 74: Popper, S. J., A. D. Sarr, K. U. Travers, A. Gueye-Ndiaye, S. Mboup, M. E. Essex, and P. J. Kanki Lower human immunodeficiency virus (HIV) type 2 viral load reflects the difference in pathogenicity of HIV-1 and HIV-2. J. Infect. Dis. 180: Rodrigo, A. G., P. C. Goracke, K. Rowhanian, and J. I. Mullins Quantitation of target molecules from polymerase chain reaction-based limiting dilution assays. AIDS Res. Hum. Retrovir. 13: Rodriguez, S., A. D. Sarr, A. MacNeil, S. Thakore-Meloni, A. Gueye-Ndiaye, I. Traoré, M. C. Dia, S. Mboup, and P. J. Kanki Comparison of heterologous neutralizing antibody responses of human immunodeficiency virus type 1 (HIV-1)- and HIV-2-infected Senegalese patients: distinct patterns of breadth and magnitude distinguish HIV-1 and HIV-2 infections. J. Virol. 81: Schindler, M., J. Munch, O. Kutsch, H. Li, M. L. Santiago, F. Bibollet- Ruche, M. C. Muller-Trutwin, F. J. Novembre, M. Peeters, V. Courgnaud, E. Bailes, P. Roques, D. L. Sodora, G. Silvestri, P. M. Sharp, B. H. Hahn, and F. Kirchhoff Nef-mediated suppression of T cell activation was lost in a lentiviral lineage that gave rise to HIV-1. Cell 125: Schramm, B., M. L. Penn, E. H. Palacios, R. M. Grant, F. Kirchhoff, and M. A. Goldsmith Cytopathicity of human immunodeficiency virus type 2 (HIV-2) in human lymphoid tissue is coreceptor dependent and comparable to that of HIV-1. J. Virol. 74: Simon, F., S. Matheron, C. Tamalet, I. Loussert-Ajaka, S. Bartczak, J. M. Pepin, C. Dhiver, E. Gamba, C. Elbim, J. A. Gastaut, et al Cellular and plasma viral load in patients infected with HIV-2. AIDS 7: Whittle, H., J. Morris, J. Todd, T. Corrah, S. Sabally, J. Bangali, P. T. Ngom, M. Rolfe, and A. Wilkins HIV-2-infected patients survive longer than HIV-1-infected patients. AIDS 8: Wiskerchen, M., and M. A. Muesing Human immunodeficiency virus type 1 integrase: effects of mutations on viral ability to integrate, direct viral gene expression from unintegrated viral DNA templates, and sustain viral propagation in primary cells. J. Virol. 69:
Genomic Sites of Human Immunodeficiency Virus Type 2 (HIV- 2) Integration: Similarities to HIV-1 In Vitro and Possible Differences In Vivo
 Genomic Sites of Human Immunodeficiency Virus Type 2 (HIV- 2) Integration: Similarities to HIV-1 In Vitro and Possible Differences In Vivo The Harvard community has made this article openly available.
Genomic Sites of Human Immunodeficiency Virus Type 2 (HIV- 2) Integration: Similarities to HIV-1 In Vitro and Possible Differences In Vivo The Harvard community has made this article openly available.
Supplemental Materials and Methods Plasmids and viruses Quantitative Reverse Transcription PCR Generation of molecular standard for quantitative PCR
 Supplemental Materials and Methods Plasmids and viruses To generate pseudotyped viruses, the previously described recombinant plasmids pnl4-3-δnef-gfp or pnl4-3-δ6-drgfp and a vector expressing HIV-1 X4
Supplemental Materials and Methods Plasmids and viruses To generate pseudotyped viruses, the previously described recombinant plasmids pnl4-3-δnef-gfp or pnl4-3-δ6-drgfp and a vector expressing HIV-1 X4
HIV-2 infected Senegalese patients: Distinct patterns of breadth and magnitude ACCEPTED
 JVI Accepts, published online ahead of print on 1 February 00 J. Virol. doi:./jvi.0-0 Copyright 00, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved. Title:
JVI Accepts, published online ahead of print on 1 February 00 J. Virol. doi:./jvi.0-0 Copyright 00, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved. Title:
Sixteen years of HIV surveillance in a West African research clinic reveals divergent epidemic trends of HIV-1 and HIV-2
 Published by Oxford University Press on behalf of the International Epidemiological Association International Journal of Epidemiology 2006;35:1322 1328 Ó The Author 2006; all rights reserved. Advance Access
Published by Oxford University Press on behalf of the International Epidemiological Association International Journal of Epidemiology 2006;35:1322 1328 Ó The Author 2006; all rights reserved. Advance Access
ORIGINAL INVESTIGATION. Plasma Viral Load in HIV-1 and HIV-2 Singly and Dually Infected Individuals in Guinea-Bissau, West Africa
 ORIGINAL INVESTIGATION Plasma Viral Load in HIV-1 and HIV-2 Singly and Dually Infected Individuals in Guinea-Bissau, West Africa Significantly Lower Plasma Virus Set Point in HIV-2 Infection Than in HIV-1
ORIGINAL INVESTIGATION Plasma Viral Load in HIV-1 and HIV-2 Singly and Dually Infected Individuals in Guinea-Bissau, West Africa Significantly Lower Plasma Virus Set Point in HIV-2 Infection Than in HIV-1
Comparison of Human Immunodeficiency Virus (HIV)-Specific T-Cell Responses in HIV-1- and HIV-2-Infected Individuals in Senegal
 JOURNAL OF VIROLOGY, Dec. 2004, p. 13934 13942 Vol. 78, No. 24 0022-538X/04/$08.00 0 DOI: 10.1128/JVI.78.24.13934 13942.2004 Copyright 2004, American Society for Microbiology. All Rights Reserved. Comparison
JOURNAL OF VIROLOGY, Dec. 2004, p. 13934 13942 Vol. 78, No. 24 0022-538X/04/$08.00 0 DOI: 10.1128/JVI.78.24.13934 13942.2004 Copyright 2004, American Society for Microbiology. All Rights Reserved. Comparison
Higher Homologous and Lower Cross-Reactive Gag-Specific T-Cell Responses in Human Immunodeficiency Virus Type 2 (HIV-2) Than in HIV-1 Infection
 JOURNAL OF VIROLOGY, Sept. 2008, p. 8619 8628 Vol. 82, No. 17 0022-538X/08/$08.00 0 doi:10.1128/jvi.00027-08 Copyright 2008, American Society for Microbiology. All Rights Reserved. Higher Homologous and
JOURNAL OF VIROLOGY, Sept. 2008, p. 8619 8628 Vol. 82, No. 17 0022-538X/08/$08.00 0 doi:10.1128/jvi.00027-08 Copyright 2008, American Society for Microbiology. All Rights Reserved. Higher Homologous and
HIV-DNA: nuovo marcatore virologico. Metodiche a confronto per la quantificazione di HIV-DNA
 HIV-DNA: nuovo marcatore virologico Metodiche a confronto per la quantificazione di HIV-DNA Maria Carla Re Laboratorio Retrovirus e Agenti infettivi HIV correlati UO di Microbiologia, Università di Bologna
HIV-DNA: nuovo marcatore virologico Metodiche a confronto per la quantificazione di HIV-DNA Maria Carla Re Laboratorio Retrovirus e Agenti infettivi HIV correlati UO di Microbiologia, Università di Bologna
Quantification of Proviral Load of Human Immunodeficiency Virus Type 2 Subtypes A and B Using Real-Time PCR
 JOURNAL OF CLINICAL MICROBIOLOGY, Dec. 2001, p. 4264 4268 Vol. 39, No. 12 0095-1137/01/$04.00 0 DOI: 10.1128/JCM.39.12.4264 4268.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved.
JOURNAL OF CLINICAL MICROBIOLOGY, Dec. 2001, p. 4264 4268 Vol. 39, No. 12 0095-1137/01/$04.00 0 DOI: 10.1128/JCM.39.12.4264 4268.2001 Copyright 2001, American Society for Microbiology. All Rights Reserved.
Fayth K. Yoshimura, Ph.D. September 7, of 7 HIV - BASIC PROPERTIES
 1 of 7 I. Viral Origin. A. Retrovirus - animal lentiviruses. HIV - BASIC PROPERTIES 1. HIV is a member of the Retrovirus family and more specifically it is a member of the Lentivirus genus of this family.
1 of 7 I. Viral Origin. A. Retrovirus - animal lentiviruses. HIV - BASIC PROPERTIES 1. HIV is a member of the Retrovirus family and more specifically it is a member of the Lentivirus genus of this family.
DATA SHEET. Provided: 500 µl of 5.6 mm Tris HCl, 4.4 mm Tris base, 0.05% sodium azide 0.1 mm EDTA, 5 mg/liter calf thymus DNA.
 Viral Load DNA >> Standard PCR standard 0 Copies Catalog Number: 1122 Lot Number: 150298 Release Category: A Provided: 500 µl of 5.6 mm Tris HCl, 4.4 mm Tris base, 0.05% sodium azide 0.1 mm EDTA, 5 mg/liter
Viral Load DNA >> Standard PCR standard 0 Copies Catalog Number: 1122 Lot Number: 150298 Release Category: A Provided: 500 µl of 5.6 mm Tris HCl, 4.4 mm Tris base, 0.05% sodium azide 0.1 mm EDTA, 5 mg/liter
HIV-1 Viral Load Real Time (RG)
 -1 Viral Load Real Time (RG) Real Time RT-PCR type 1 RNA quantification assay MSP Reg. pending Valdense 3616. 11700. Montevideo. Uruguay. phone (598) 2 336 83 01. Fax (598) 2 336 71 60. Info@atgen.com.uy
-1 Viral Load Real Time (RG) Real Time RT-PCR type 1 RNA quantification assay MSP Reg. pending Valdense 3616. 11700. Montevideo. Uruguay. phone (598) 2 336 83 01. Fax (598) 2 336 71 60. Info@atgen.com.uy
Hepatitis B Antiviral Drug Development Multi-Marker Screening Assay
 Hepatitis B Antiviral Drug Development Multi-Marker Screening Assay Background ImQuest BioSciences has developed and qualified a single-plate method to expedite the screening of antiviral agents against
Hepatitis B Antiviral Drug Development Multi-Marker Screening Assay Background ImQuest BioSciences has developed and qualified a single-plate method to expedite the screening of antiviral agents against
Recurrent HIV-1 Integration at the BACH2 Locus in Resting CD4 + T Cell Populations during Effective Highly Active Antiretroviral Therapy
 M A J O R A R T I C L E Recurrent HIV-1 Integration at the BACH2 Locus in Resting CD4 + T Cell Populations during Effective Highly Active Antiretroviral Therapy Terumasa Ikeda, Junji Shibata, Kazuhisa
M A J O R A R T I C L E Recurrent HIV-1 Integration at the BACH2 Locus in Resting CD4 + T Cell Populations during Effective Highly Active Antiretroviral Therapy Terumasa Ikeda, Junji Shibata, Kazuhisa
Maintenance of HIV-Specific CD4 + T Cell Help Distinguishes HIV-2 from HIV-1 Infection
 This information is current as of September 1, 2018. References Subscription Permissions Email Alerts Maintenance of HIV-Specific CD4 + T Cell Help Distinguishes HIV-2 from HIV-1 Infection Melody G. Duvall,
This information is current as of September 1, 2018. References Subscription Permissions Email Alerts Maintenance of HIV-Specific CD4 + T Cell Help Distinguishes HIV-2 from HIV-1 Infection Melody G. Duvall,
Inhibition of HIV-1 Integration in Ex Vivo-Infected CD4 T Cells from Elite Controllers
 JOURNAL OF VIROLOGY, Sept. 2011, p. 9646 9650 Vol. 85, No. 18 0022-538X/11/$12.00 doi:10.1128/jvi.05327-11 Copyright 2011, American Society for Microbiology. All Rights Reserved. Inhibition of HIV-1 Integration
JOURNAL OF VIROLOGY, Sept. 2011, p. 9646 9650 Vol. 85, No. 18 0022-538X/11/$12.00 doi:10.1128/jvi.05327-11 Copyright 2011, American Society for Microbiology. All Rights Reserved. Inhibition of HIV-1 Integration
Human Immunodeficiency Virus
 Human Immunodeficiency Virus Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Viruses and hosts Lentivirus from Latin lentis (slow), for slow progression of disease
Human Immunodeficiency Virus Virion Genome Genes and proteins Viruses and hosts Diseases Distinctive characteristics Viruses and hosts Lentivirus from Latin lentis (slow), for slow progression of disease
Potent Inhibition of Human Immunodeficiency Virus Type 1 (HIV-1) Gene Expression and Virus Production by an HIV-2 Tat Activation-Response RNA Decoy
 JOURNAL OF VIROLOGY, June 1999, p. 5191 5195 Vol. 73, No. 6 0022-538X/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. Potent Inhibition of Human Immunodeficiency Virus
JOURNAL OF VIROLOGY, June 1999, p. 5191 5195 Vol. 73, No. 6 0022-538X/99/$04.00 0 Copyright 1999, American Society for Microbiology. All Rights Reserved. Potent Inhibition of Human Immunodeficiency Virus
Fayth K. Yoshimura, Ph.D. September 7, of 7 RETROVIRUSES. 2. HTLV-II causes hairy T-cell leukemia
 1 of 7 I. Diseases Caused by Retroviruses RETROVIRUSES A. Human retroviruses that cause cancers 1. HTLV-I causes adult T-cell leukemia and tropical spastic paraparesis 2. HTLV-II causes hairy T-cell leukemia
1 of 7 I. Diseases Caused by Retroviruses RETROVIRUSES A. Human retroviruses that cause cancers 1. HTLV-I causes adult T-cell leukemia and tropical spastic paraparesis 2. HTLV-II causes hairy T-cell leukemia
Micropathology Ltd. University of Warwick Science Park, Venture Centre, Sir William Lyons Road, Coventry CV4 7EZ
 www.micropathology.com info@micropathology.com Micropathology Ltd Tel 24hrs: +44 (0) 24-76 323222 Fax / Ans: +44 (0) 24-76 - 323333 University of Warwick Science Park, Venture Centre, Sir William Lyons
www.micropathology.com info@micropathology.com Micropathology Ltd Tel 24hrs: +44 (0) 24-76 323222 Fax / Ans: +44 (0) 24-76 - 323333 University of Warwick Science Park, Venture Centre, Sir William Lyons
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Figure 1: Cryopreservation alters CD62L expression by CD4 T cells. Freshly isolated (left) or cryopreserved PBMCs (right) were stained with the mix of antibodies described
Supplementary Figure 1 Supplementary Figure 1: Cryopreservation alters CD62L expression by CD4 T cells. Freshly isolated (left) or cryopreserved PBMCs (right) were stained with the mix of antibodies described
altona RealStar Instructions for Use RealStar CMV PCR Kit /2017 EN DIAGNOSTICS
 altona DIAGNOSTICS Instructions for Use RealStar CMV PCR Kit 1.2 08/2017 EN RealStar RealStar CMV PCR Kit 1.2 For research use only! (RUO) 021202 INS-021200-EN-S01 48 08 2017 altona Diagnostics GmbH Mörkenstr.
altona DIAGNOSTICS Instructions for Use RealStar CMV PCR Kit 1.2 08/2017 EN RealStar RealStar CMV PCR Kit 1.2 For research use only! (RUO) 021202 INS-021200-EN-S01 48 08 2017 altona Diagnostics GmbH Mörkenstr.
VIRAL TITER COUNTS. The best methods of measuring infectious lentiviral titer
 VIRAL TITER COUNTS The best methods of measuring infectious lentiviral titer FLUORESCENCE CYCLES qpcr of Viral RNA SUMMARY Viral vectors are now routinely used for gene transduction in a wide variety of
VIRAL TITER COUNTS The best methods of measuring infectious lentiviral titer FLUORESCENCE CYCLES qpcr of Viral RNA SUMMARY Viral vectors are now routinely used for gene transduction in a wide variety of
Targeted Derepression of the Human Immunodeficiency Virus Type 1 Long Terminal Repeat by Pyrrole-Imidazole Polyamides
 JOURNAL OF VIROLOGY, Dec. 2002, p. 12349 12354 Vol. 76, No. 23 0022-538X/02/$04.00 0 DOI: 10.1128/JVI.76.23.12349 12354.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved. Targeted
JOURNAL OF VIROLOGY, Dec. 2002, p. 12349 12354 Vol. 76, No. 23 0022-538X/02/$04.00 0 DOI: 10.1128/JVI.76.23.12349 12354.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved. Targeted
NIH Public Access Author Manuscript J Acquir Immune Defic Syndr. Author manuscript; available in PMC 2013 September 01.
 NIH Public Access Author Manuscript Published in final edited form as: J Acquir Immune Defic Syndr. 2012 September 1; 61(1): 19 22. doi:10.1097/qai.0b013e318264460f. Evaluation of HIV-1 Ambiguous Nucleotide
NIH Public Access Author Manuscript Published in final edited form as: J Acquir Immune Defic Syndr. 2012 September 1; 61(1): 19 22. doi:10.1097/qai.0b013e318264460f. Evaluation of HIV-1 Ambiguous Nucleotide
The New England Journal of Medicine
 PERSISTENCE OF HIV-1 TRANSCRIPTION IN PERIPHERAL-BLOOD MONONUCLEAR CELLS IN PATIENTS RECEIVING POTENT ANTIRETROVIRAL THERAPY MANOHAR R. FURTADO, PH.D., DUNCAN S. CALLAWAY, B.S., JOHN P. PHAIR, M.D., KEVIN
PERSISTENCE OF HIV-1 TRANSCRIPTION IN PERIPHERAL-BLOOD MONONUCLEAR CELLS IN PATIENTS RECEIVING POTENT ANTIRETROVIRAL THERAPY MANOHAR R. FURTADO, PH.D., DUNCAN S. CALLAWAY, B.S., JOHN P. PHAIR, M.D., KEVIN
HIV INFECTION: An Overview
 HIV INFECTION: An Overview UNIVERSITY OF PAPUA NEW GUINEA SCHOOL OF MEDICINE AND HEALTH SCIENCES DIVISION OF BASIC MEDICAL SCIENCES DISCIPLINE OF BIOCHEMISTRY & MOLECULAR BIOLOGY PBL MBBS II SEMINAR VJ
HIV INFECTION: An Overview UNIVERSITY OF PAPUA NEW GUINEA SCHOOL OF MEDICINE AND HEALTH SCIENCES DIVISION OF BASIC MEDICAL SCIENCES DISCIPLINE OF BIOCHEMISTRY & MOLECULAR BIOLOGY PBL MBBS II SEMINAR VJ
Trends in molecular diagnostics
 Trends in molecular diagnostics Detection of target genes of interest Quantification Infectious diseases HIV Hepatitis C & B TB / MAC Cytomegalovirus Herpes simplex Varicella zoster CT/GC HPV Profiling
Trends in molecular diagnostics Detection of target genes of interest Quantification Infectious diseases HIV Hepatitis C & B TB / MAC Cytomegalovirus Herpes simplex Varicella zoster CT/GC HPV Profiling
Interaction with Human Immunodeficiency Virus (HIV) Type 2 Predicts HIV Type 1 Genotype
 Virology 268, 402 410 (2000) doi:10.1006/viro.2000.0192, available online at http://www.idealibrary.com on Interaction with Human Immunodeficiency Virus (HIV) Type 2 Predicts HIV Type 1 Genotype Abdoulaye
Virology 268, 402 410 (2000) doi:10.1006/viro.2000.0192, available online at http://www.idealibrary.com on Interaction with Human Immunodeficiency Virus (HIV) Type 2 Predicts HIV Type 1 Genotype Abdoulaye
Genetic Recombination between Human Immunodeficiency Virus Type 1 (HIV-1) and HIV-2, Two Distinct Human Lentiviruses
 JOURNAL OF VIROLOGY, Feb. 2008, p. 1923 1933 Vol. 82, No. 4 0022-538X/08/$08.00 0 doi:10.1128/jvi.01937-07 Genetic Recombination between Human Immunodeficiency Virus Type 1 (HIV-1) and HIV-2, Two Distinct
JOURNAL OF VIROLOGY, Feb. 2008, p. 1923 1933 Vol. 82, No. 4 0022-538X/08/$08.00 0 doi:10.1128/jvi.01937-07 Genetic Recombination between Human Immunodeficiency Virus Type 1 (HIV-1) and HIV-2, Two Distinct
Diversity and Tropism of HIV-1 Plasma Rebound Virus after Treatment Discontinuation
 Diversity and Tropism of HIV-1 Plasma Rebound Virus after Treatment Discontinuation By: Blake M. Hauser Senior Honors Thesis Department of Biology College of Arts and Sciences The University of North Carolina
Diversity and Tropism of HIV-1 Plasma Rebound Virus after Treatment Discontinuation By: Blake M. Hauser Senior Honors Thesis Department of Biology College of Arts and Sciences The University of North Carolina
Viral Vectors In The Research Laboratory: Just How Safe Are They? Dawn P. Wooley, Ph.D., SM(NRM), RBP, CBSP
 Viral Vectors In The Research Laboratory: Just How Safe Are They? Dawn P. Wooley, Ph.D., SM(NRM), RBP, CBSP 1 Learning Objectives Recognize hazards associated with viral vectors in research and animal
Viral Vectors In The Research Laboratory: Just How Safe Are They? Dawn P. Wooley, Ph.D., SM(NRM), RBP, CBSP 1 Learning Objectives Recognize hazards associated with viral vectors in research and animal
Lecture 2: Virology. I. Background
 Lecture 2: Virology I. Background A. Properties 1. Simple biological systems a. Aggregates of nucleic acids and protein 2. Non-living a. Cannot reproduce or carry out metabolic activities outside of a
Lecture 2: Virology I. Background A. Properties 1. Simple biological systems a. Aggregates of nucleic acids and protein 2. Non-living a. Cannot reproduce or carry out metabolic activities outside of a
It takes more than just a single target
 It takes more than just a single target As the challenges you face evolve... HIV mutates No HIV-1 mutation can be considered to be neutral 1 Growing evidence indicates all HIV subtypes may be prone to
It takes more than just a single target As the challenges you face evolve... HIV mutates No HIV-1 mutation can be considered to be neutral 1 Growing evidence indicates all HIV subtypes may be prone to
Laboratory for Clinical and Biological Studies, University of Miami Miller School of Medicine, Miami, FL, USA.
 000000 00000 0000 000 00 0 bdna () 00000 0000 000 00 0 Nuclisens () 000 00 0 000000 00000 0000 000 00 0 Amplicor () Comparison of Amplicor HIV- monitor Test, NucliSens HIV- QT and bdna Versant HIV RNA
000000 00000 0000 000 00 0 bdna () 00000 0000 000 00 0 Nuclisens () 000 00 0 000000 00000 0000 000 00 0 Amplicor () Comparison of Amplicor HIV- monitor Test, NucliSens HIV- QT and bdna Versant HIV RNA
Technical Bulletin No. 161
 CPAL Central Pennsylvania Alliance Laboratory Technical Bulletin No. 161 cobas 6800 HIV-1 Viral Load Assay - New Platform - June 1, 2017 Contact: Heather Habig, MLS (ASCP) CM, MB CM, 717-851-1422 Operations
CPAL Central Pennsylvania Alliance Laboratory Technical Bulletin No. 161 cobas 6800 HIV-1 Viral Load Assay - New Platform - June 1, 2017 Contact: Heather Habig, MLS (ASCP) CM, MB CM, 717-851-1422 Operations
Instructions for Use. RealStar Influenza Screen & Type RT-PCR Kit /2017 EN
 Instructions for Use RealStar Influenza Screen & Type RT-PCR Kit 4.0 05/2017 EN RealStar Influenza Screen & Type RT-PCR Kit 4.0 For research use only! (RUO) 164003 INS-164000-EN-S01 96 05 2017 altona
Instructions for Use RealStar Influenza Screen & Type RT-PCR Kit 4.0 05/2017 EN RealStar Influenza Screen & Type RT-PCR Kit 4.0 For research use only! (RUO) 164003 INS-164000-EN-S01 96 05 2017 altona
Chapter 19: The Genetics of Viruses and Bacteria
 Chapter 19: The Genetics of Viruses and Bacteria What is Microbiology? Microbiology is the science that studies microorganisms = living things that are too small to be seen with the naked eye Microorganisms
Chapter 19: The Genetics of Viruses and Bacteria What is Microbiology? Microbiology is the science that studies microorganisms = living things that are too small to be seen with the naked eye Microorganisms
AIDS - Knowledge and Dogma. Conditions for the Emergence and Decline of Scientific Theories Congress, July 16/ , Vienna, Austria
 AIDS - Knowledge and Dogma Conditions for the Emergence and Decline of Scientific Theories Congress, July 16/17 2010, Vienna, Austria Reliability of PCR to detect genetic sequences from HIV Juan Manuel
AIDS - Knowledge and Dogma Conditions for the Emergence and Decline of Scientific Theories Congress, July 16/17 2010, Vienna, Austria Reliability of PCR to detect genetic sequences from HIV Juan Manuel
Instructions for Use. RealStar Influenza S&T RT-PCR Kit /2017 EN
 Instructions for Use RealStar Influenza S&T RT-PCR Kit 3.0 01/2017 EN RealStar Influenza S&T RT-PCR Kit 3.0 For research use only! (RUO) 163003 INS-163000-EN-S02 96 01 2017 altona Diagnostics GmbH Mörkenstr.
Instructions for Use RealStar Influenza S&T RT-PCR Kit 3.0 01/2017 EN RealStar Influenza S&T RT-PCR Kit 3.0 For research use only! (RUO) 163003 INS-163000-EN-S02 96 01 2017 altona Diagnostics GmbH Mörkenstr.
HIV & AIDS: Overview
 HIV & AIDS: Overview UNIVERSITY OF PAPUA NEW GUINEA SCHOOL OF MEDICINE AND HEALTH SCIENCES DIVISION OF BASIC MEDICAL SCIENCES DISCIPLINE OF BIOCHEMISTRY & MOLECULAR BIOLOGY PBL SEMINAR VJ TEMPLE 1 What
HIV & AIDS: Overview UNIVERSITY OF PAPUA NEW GUINEA SCHOOL OF MEDICINE AND HEALTH SCIENCES DIVISION OF BASIC MEDICAL SCIENCES DISCIPLINE OF BIOCHEMISTRY & MOLECULAR BIOLOGY PBL SEMINAR VJ TEMPLE 1 What
Alexander O. Pasternak, Mirte Scherpenisse, Ben Berkhout
 Cell-associated HIV-1 unspliced to multiply spliced RNA ratio at 12 weeks ART correlates with markers of immune activation and apoptosis and predicts the CD4 + T-cell count at 96 weeks ART Alexander O.
Cell-associated HIV-1 unspliced to multiply spliced RNA ratio at 12 weeks ART correlates with markers of immune activation and apoptosis and predicts the CD4 + T-cell count at 96 weeks ART Alexander O.
Pelagia Research Library. European Journal of Experimental Biology, 2015, 5(10):1-5
 Available online at www.pelagiaresearchlibrary.com European Journal of Experimental Biology, 2015, 5(10):1-5 ISSN: 2248 9215 CODEN (USA): EJEBAU Molecular diagnosis of human immuno deficiency virus (HIV)
Available online at www.pelagiaresearchlibrary.com European Journal of Experimental Biology, 2015, 5(10):1-5 ISSN: 2248 9215 CODEN (USA): EJEBAU Molecular diagnosis of human immuno deficiency virus (HIV)
HIV-1 replication and immune dynamics are affected by raltegravir intensification of HAART-suppressed subjects
 HIV-1 replication and immune dynamics are affected by raltegravir intensification of HAART-suppressed subjects Maria J Buzón 1,9, Marta Massanella 1,9, Josep M Llibre 2, Anna Esteve 3, Viktor Dahl 4, Maria
HIV-1 replication and immune dynamics are affected by raltegravir intensification of HAART-suppressed subjects Maria J Buzón 1,9, Marta Massanella 1,9, Josep M Llibre 2, Anna Esteve 3, Viktor Dahl 4, Maria
Norgen s HIV proviral DNA PCR Kit was developed and validated to be used with the following PCR instruments: Qiagen Rotor-Gene Q BioRad icycler
 3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com HIV Proviral DNA PCR Kit Product # 33840 Product Insert Background Information
3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com HIV Proviral DNA PCR Kit Product # 33840 Product Insert Background Information
Transcriptional Regulation of Latent Feline Immunodeficiency Virus in Peripheral CD4+ T-lymphocytes
 Viruses 2012, 4, 878-888; doi:10.3390/v4050878 Brief Report OPEN ACCESS viruses ISSN 1999-4915 www.mdpi.com/journal/viruses Transcriptional Regulation of Latent Feline Immunodeficiency Virus in Peripheral
Viruses 2012, 4, 878-888; doi:10.3390/v4050878 Brief Report OPEN ACCESS viruses ISSN 1999-4915 www.mdpi.com/journal/viruses Transcriptional Regulation of Latent Feline Immunodeficiency Virus in Peripheral
Norgen s HIV Proviral DNA PCR Kit was developed and validated to be used with the following PCR instruments: Qiagen Rotor-Gene Q BioRad T1000 Cycler
 3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: 866-667-4362 (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com HIV Proviral DNA PCR Kit Product# 33840 Product Insert Intended
3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: 866-667-4362 (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com HIV Proviral DNA PCR Kit Product# 33840 Product Insert Intended
HIV-2 and the Immune Response
 AIDS Rev 2001; 3: 11-23 HIV-2 and the Immune Response Sören Andersson Department of Immunology, Swedish Institute for Infectious Disease Control, Stockholm, Sweden The Department of Immunology and Clinical
AIDS Rev 2001; 3: 11-23 HIV-2 and the Immune Response Sören Andersson Department of Immunology, Swedish Institute for Infectious Disease Control, Stockholm, Sweden The Department of Immunology and Clinical
Molecular Diagnosis Future Directions
 Molecular Diagnosis Future Directions Philip Cunningham NSW State Reference Laboratory for HIV/AIDS & Molecular Diagnostic Medicine Laboratory, SydPath St Vincent s Hospital Sydney Update on Molecular
Molecular Diagnosis Future Directions Philip Cunningham NSW State Reference Laboratory for HIV/AIDS & Molecular Diagnostic Medicine Laboratory, SydPath St Vincent s Hospital Sydney Update on Molecular
VQA Control SOP Version 4.0 Roche Amplicor HIV-1 DNA Test, v August 2007
 1. PRINCIPLE 1.1. The Virology Quality Assurance (VQA) Laboratory provides external cell pellet controls for use in the validation of assays that detect HIV proviral DNA. 1.2. HIV seronegative peripheral
1. PRINCIPLE 1.1. The Virology Quality Assurance (VQA) Laboratory provides external cell pellet controls for use in the validation of assays that detect HIV proviral DNA. 1.2. HIV seronegative peripheral
Biol115 The Thread of Life"
 Biol115 The Thread of Life" Lecture 9" Gene expression and the Central Dogma"... once (sequential) information has passed into protein it cannot get out again. " ~Francis Crick, 1958! Principles of Biology
Biol115 The Thread of Life" Lecture 9" Gene expression and the Central Dogma"... once (sequential) information has passed into protein it cannot get out again. " ~Francis Crick, 1958! Principles of Biology
Introduction retroposon
 17.1 - Introduction A retrovirus is an RNA virus able to convert its sequence into DNA by reverse transcription A retroposon (retrotransposon) is a transposon that mobilizes via an RNA form; the DNA element
17.1 - Introduction A retrovirus is an RNA virus able to convert its sequence into DNA by reverse transcription A retroposon (retrotransposon) is a transposon that mobilizes via an RNA form; the DNA element
HIV 101: Fundamentals of HIV Infection
 HIV 101: Fundamentals of HIV Infection David H. Spach, MD Professor of Medicine University of Washington Seattle, Washington Learning Objectives After attending this presentation, learners will be able
HIV 101: Fundamentals of HIV Infection David H. Spach, MD Professor of Medicine University of Washington Seattle, Washington Learning Objectives After attending this presentation, learners will be able
Low immune activation despite high levels of pathogenic HIV-1 results in long-term asymptomatic disease
 Low immune activation despite high levels of pathogenic HIV-1 results in long-term asymptomatic disease Shailesh K. Choudhary 1 *, Nienke Vrisekoop 2 *, Christine A. Jansen 2, Sigrid A. Otto 2, Hanneke
Low immune activation despite high levels of pathogenic HIV-1 results in long-term asymptomatic disease Shailesh K. Choudhary 1 *, Nienke Vrisekoop 2 *, Christine A. Jansen 2, Sigrid A. Otto 2, Hanneke
The HIV life cycle. integration. virus production. entry. transcription. reverse transcription. nuclear import
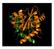 The HIV life cycle entry reverse transcription transcription integration virus production nuclear import Hazuda 2012 Integration Insertion of the viral DNA into host chromosomal DNA, essential step in
The HIV life cycle entry reverse transcription transcription integration virus production nuclear import Hazuda 2012 Integration Insertion of the viral DNA into host chromosomal DNA, essential step in
Towards a block and lock strategy: LEDGINs hamper the establishment of a reactivation competent HIV reservoir.
 Abstract no. MOLBPEA13 Towards a block and lock strategy: LEDGINs hamper the establishment of a reactivation competent HIV reservoir. G. Vansant,,A. Bruggemans, L. Vranckx, S. Saleh, I. Zurnic, F. Christ,
Abstract no. MOLBPEA13 Towards a block and lock strategy: LEDGINs hamper the establishment of a reactivation competent HIV reservoir. G. Vansant,,A. Bruggemans, L. Vranckx, S. Saleh, I. Zurnic, F. Christ,
Towards an HIV Cure. Steven G. Deeks Professor of Medicine University of California, San Francisco
 Towards an HIV Cure Steven G. Deeks Professor of Medicine University of California, San Francisco Why are we now talking about a cure? Emerging recognition that HAART does not fully restore health and/or
Towards an HIV Cure Steven G. Deeks Professor of Medicine University of California, San Francisco Why are we now talking about a cure? Emerging recognition that HAART does not fully restore health and/or
7.014 Problem Set 7 Solutions
 MIT Department of Biology 7.014 Introductory Biology, Spring 2005 7.014 Problem Set 7 Solutions Question 1 Part A Antigen binding site Antigen binding site Variable region Light chain Light chain Variable
MIT Department of Biology 7.014 Introductory Biology, Spring 2005 7.014 Problem Set 7 Solutions Question 1 Part A Antigen binding site Antigen binding site Variable region Light chain Light chain Variable
Immunodeficiency. (2 of 2)
 Immunodeficiency (2 of 2) Acquired (secondary) immunodeficiencies More common Many causes such as therapy, cancer, sarcoidosis, malnutrition, infection & renal disease The most common of which is therapy-related
Immunodeficiency (2 of 2) Acquired (secondary) immunodeficiencies More common Many causes such as therapy, cancer, sarcoidosis, malnutrition, infection & renal disease The most common of which is therapy-related
The monitoring of HIV-infected patients is based on
 AIDS RESEARCH AND HUMAN RETROVIRUSES Volume 32, Number 6, 2016 ª Mary Ann Liebert, Inc. DOI: 10.1089/aid.2015.0348 Prognostic Value of HIV-1 RNA on CD4 Trajectories and Disease Progression Among Antiretroviral-Naive
AIDS RESEARCH AND HUMAN RETROVIRUSES Volume 32, Number 6, 2016 ª Mary Ann Liebert, Inc. DOI: 10.1089/aid.2015.0348 Prognostic Value of HIV-1 RNA on CD4 Trajectories and Disease Progression Among Antiretroviral-Naive
~Lentivirus production~
 ~Lentivirus production~ May 30, 2008 RNAi core R&D group member Lentivirus Production Session Lentivirus!!! Is it health threatening to lab technician? What s so good about this RNAi library? How to produce
~Lentivirus production~ May 30, 2008 RNAi core R&D group member Lentivirus Production Session Lentivirus!!! Is it health threatening to lab technician? What s so good about this RNAi library? How to produce
Inhibition of HIV-1 Disease Progression by Contemporaneous HIV-2 Infection
 T h e n e w e ngl a nd j o u r na l o f m e dic i n e original article Inhibition of HIV-1 Disease Progression by Contemporaneous HIV-2 Infection Joakim Esbjörnsson, Ph.D., Fredrik Månsson, M.D., Ph.D.,
T h e n e w e ngl a nd j o u r na l o f m e dic i n e original article Inhibition of HIV-1 Disease Progression by Contemporaneous HIV-2 Infection Joakim Esbjörnsson, Ph.D., Fredrik Månsson, M.D., Ph.D.,
For in vitro Veterinary Diagnostics only. Kylt Rotavirus A. Real-Time RT-PCR Detection.
 For in vitro Veterinary Diagnostics only. Kylt Rotavirus A Real-Time RT-PCR Detection www.kylt.eu DIRECTION FOR USE Kylt Rotavirus A Real-Time RT-PCR Detection A. General Kylt Rotavirus A products are
For in vitro Veterinary Diagnostics only. Kylt Rotavirus A Real-Time RT-PCR Detection www.kylt.eu DIRECTION FOR USE Kylt Rotavirus A Real-Time RT-PCR Detection A. General Kylt Rotavirus A products are
Identification and Characterization of CD4 T cells actively transcribing HIV RNA in Peripheral Blood
 Dale and Betty Bumpers Vaccine Research Center National Institute of Allergy and Infectious Diseases National Institutes of Health Identification and Characterization of CD4 T cells actively transcribing
Dale and Betty Bumpers Vaccine Research Center National Institute of Allergy and Infectious Diseases National Institutes of Health Identification and Characterization of CD4 T cells actively transcribing
SUPPLEMENTARY INFORMATION. Divergent TLR7/9 signaling and type I interferon production distinguish
 SUPPLEMENTARY INFOATION Divergent TLR7/9 signaling and type I interferon production distinguish pathogenic and non-pathogenic AIDS-virus infections Judith N. Mandl, Ashley P. Barry, Thomas H. Vanderford,
SUPPLEMENTARY INFOATION Divergent TLR7/9 signaling and type I interferon production distinguish pathogenic and non-pathogenic AIDS-virus infections Judith N. Mandl, Ashley P. Barry, Thomas H. Vanderford,
Fig. 1: Schematic diagram of basic structure of HIV
 UNIVERSITY OF PAPUA NEW GUINEA SCHOOL OF MEDICINE AND HEALTH SCIENCES DIVISION OF BASIC MEDICAL SCIENCES DISCIPLINE OF BIOCHEMISTRY & MOLECULAR BIOLOGY PBL SEMINAR HIV & AIDS: An Overview What is HIV?
UNIVERSITY OF PAPUA NEW GUINEA SCHOOL OF MEDICINE AND HEALTH SCIENCES DIVISION OF BASIC MEDICAL SCIENCES DISCIPLINE OF BIOCHEMISTRY & MOLECULAR BIOLOGY PBL SEMINAR HIV & AIDS: An Overview What is HIV?
L I F E S C I E N C E S
 1a L I F E S C I E N C E S 5 -UUA AUA UUC GAA AGC UGC AUC GAA AAC UGU GAA UCA-3 5 -TTA ATA TTC GAA AGC TGC ATC GAA AAC TGT GAA TCA-3 3 -AAT TAT AAG CTT TCG ACG TAG CTT TTG ACA CTT AGT-5 OCTOBER 31, 2006
1a L I F E S C I E N C E S 5 -UUA AUA UUC GAA AGC UGC AUC GAA AAC UGU GAA UCA-3 5 -TTA ATA TTC GAA AGC TGC ATC GAA AAC TGT GAA TCA-3 3 -AAT TAT AAG CTT TCG ACG TAG CTT TTG ACA CTT AGT-5 OCTOBER 31, 2006
A VACCINE FOR HIV BIOE 301 LECTURE 10 MITALI BANERJEE HAART
 BIOE 301 LECTURE 10 MITALI BANERJEE A VACCINE FOR HIV HIV HAART Visit wikipedia.org and learn the mechanism of action of the five classes of antiretroviral drugs. (1) Reverse transcriptase inhibitors (RTIs)
BIOE 301 LECTURE 10 MITALI BANERJEE A VACCINE FOR HIV HIV HAART Visit wikipedia.org and learn the mechanism of action of the five classes of antiretroviral drugs. (1) Reverse transcriptase inhibitors (RTIs)
VQA HIV DNA Control SOP Version 5.0 HIV DNA Testing 13 March 2012
 1. PRINCIPLE 1.1. The Virology Quality Assurance (VQA) Laboratory provides external cell pellet controls for use in the validation of assays that detect HIV proviral DNA. 1.2. HIV seronegative peripheral
1. PRINCIPLE 1.1. The Virology Quality Assurance (VQA) Laboratory provides external cell pellet controls for use in the validation of assays that detect HIV proviral DNA. 1.2. HIV seronegative peripheral
Analysis of small RNAs from Drosophila Schneider cells using the Small RNA assay on the Agilent 2100 bioanalyzer. Application Note
 Analysis of small RNAs from Drosophila Schneider cells using the Small RNA assay on the Agilent 2100 bioanalyzer Application Note Odile Sismeiro, Jean-Yves Coppée, Christophe Antoniewski, and Hélène Thomassin
Analysis of small RNAs from Drosophila Schneider cells using the Small RNA assay on the Agilent 2100 bioanalyzer Application Note Odile Sismeiro, Jean-Yves Coppée, Christophe Antoniewski, and Hélène Thomassin
Julianne Edwards. Retroviruses. Spring 2010
 Retroviruses Spring 2010 A retrovirus can simply be referred to as an infectious particle which replicates backwards even though there are many different types of retroviruses. More specifically, a retrovirus
Retroviruses Spring 2010 A retrovirus can simply be referred to as an infectious particle which replicates backwards even though there are many different types of retroviruses. More specifically, a retrovirus
Early Transcription from Nonintegrated DNA in Human Immunodeficiency Virus Infection
 JOURNAL OF VIROLOGY, Oct. 2003, p. 10376 10382 Vol. 77, No. 19 0022-538X/03/$08.00 0 DOI: 10.1128/JVI.77.19.10376 10382.2003 Early Transcription from Nonintegrated DNA in Human Immunodeficiency Virus Infection
JOURNAL OF VIROLOGY, Oct. 2003, p. 10376 10382 Vol. 77, No. 19 0022-538X/03/$08.00 0 DOI: 10.1128/JVI.77.19.10376 10382.2003 Early Transcription from Nonintegrated DNA in Human Immunodeficiency Virus Infection
WHO Prequalification of In Vitro Diagnostics PUBLIC REPORT. Product: Alere q HIV-1/2 Detect WHO reference number: PQDx
 WHO Prequalification of In Vitro Diagnostics PUBLIC REPORT Product: Alere q HIV-1/2 Detect WHO reference number: PQDx 0226-032-00 Alere q HIV-1/2 Detect with product codes 270110050, 270110010 and 270300001,
WHO Prequalification of In Vitro Diagnostics PUBLIC REPORT Product: Alere q HIV-1/2 Detect WHO reference number: PQDx 0226-032-00 Alere q HIV-1/2 Detect with product codes 270110050, 270110010 and 270300001,
Commercially available HLA Class II tetramers (Beckman Coulter) conjugated to
 Class II tetramer staining Commercially available HLA Class II tetramers (Beckman Coulter) conjugated to PE were combined with dominant HIV epitopes (DRB1*0101-DRFYKTLRAEQASQEV, DRB1*0301- PEKEVLVWKFDSRLAFHH,
Class II tetramer staining Commercially available HLA Class II tetramers (Beckman Coulter) conjugated to PE were combined with dominant HIV epitopes (DRB1*0101-DRFYKTLRAEQASQEV, DRB1*0301- PEKEVLVWKFDSRLAFHH,
Chapter 19: Viruses. 1. Viral Structure & Reproduction. 2. Bacteriophages. 3. Animal Viruses. 4. Viroids & Prions
 Chapter 19: Viruses 1. Viral Structure & Reproduction 2. Bacteriophages 3. Animal Viruses 4. Viroids & Prions 1. Viral Structure & Reproduction Chapter Reading pp. 393-396 What exactly is a Virus? Viruses
Chapter 19: Viruses 1. Viral Structure & Reproduction 2. Bacteriophages 3. Animal Viruses 4. Viroids & Prions 1. Viral Structure & Reproduction Chapter Reading pp. 393-396 What exactly is a Virus? Viruses
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells
 MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
Gene transcript profiles to diagnose infectious diseases. ESCMID Conference on Diagnosing Infectious Diseases: Future and Innovation
 Gene transcript profiles to diagnose infectious diseases ESCMID Conference on Diagnosing Infectious Diseases: Future and Innovation 24-26 October 2011, Venice, Italy Matthew Berry Consultant Respiratory
Gene transcript profiles to diagnose infectious diseases ESCMID Conference on Diagnosing Infectious Diseases: Future and Innovation 24-26 October 2011, Venice, Italy Matthew Berry Consultant Respiratory
HIV-1 DNA levels after antiretroviral therapy in primary infection predict disease progression: the SPARTAC Trial
 HIV-1 DNA levels after antiretroviral therapy in primary infection predict disease progression: the SPARTAC Trial James Williams 1,2,3, Jacob Hurst 1,2,3, Nicola Robinson 1,2,3, Sarah Fidler 4, Jonathan
HIV-1 DNA levels after antiretroviral therapy in primary infection predict disease progression: the SPARTAC Trial James Williams 1,2,3, Jacob Hurst 1,2,3, Nicola Robinson 1,2,3, Sarah Fidler 4, Jonathan
Received 4 December 2001/Accepted 29 April 2002
 JOURNAL OF VIROLOGY, Aug. 2002, p. 8433 8445 Vol. 76, No. 16 0022-538X/02/$04.00 0 DOI: 10.1128/JVI.76.16.8433 8445.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved. The Relationship
JOURNAL OF VIROLOGY, Aug. 2002, p. 8433 8445 Vol. 76, No. 16 0022-538X/02/$04.00 0 DOI: 10.1128/JVI.76.16.8433 8445.2002 Copyright 2002, American Society for Microbiology. All Rights Reserved. The Relationship
Application of μmacs Streptavidin MicroBeads for the analysis of HIV-1 directly from patient plasma
 Excerpt from MACS&more Vol 8 1/2004 Application of μmacs Streptavidin MicroBeads for the analysis of HIV-1 directly from patient plasma L. Davis Lupo and Salvatore T. Butera HIV and Retrovirology Branch,
Excerpt from MACS&more Vol 8 1/2004 Application of μmacs Streptavidin MicroBeads for the analysis of HIV-1 directly from patient plasma L. Davis Lupo and Salvatore T. Butera HIV and Retrovirology Branch,
Sysmex Educational Enhancement and Development No
 SEED Haematology No 1 2015 Introduction to the basics of CD4 and HIV Viral Load Testing The purpose of this newsletter is to provide an introduction to the basics of the specific laboratory tests that
SEED Haematology No 1 2015 Introduction to the basics of CD4 and HIV Viral Load Testing The purpose of this newsletter is to provide an introduction to the basics of the specific laboratory tests that
Massive infection and loss of memory CD4 + T cells in multiple tissues during acute SIV infection
 Massive infection and loss of memory CD4 + T cells in multiple tissues during acute SIV infection Joseph J. Mattapallil 1, Daniel C. Douek 2, Brenna Hill 2, Yoshiaki Nishimura 3, Malcolm Martin 3 & Mario
Massive infection and loss of memory CD4 + T cells in multiple tissues during acute SIV infection Joseph J. Mattapallil 1, Daniel C. Douek 2, Brenna Hill 2, Yoshiaki Nishimura 3, Malcolm Martin 3 & Mario
Citation for published version (APA): Von Eije, K. J. (2009). RNAi based gene therapy for HIV-1, from bench to bedside
 UvA-DARE (Digital Academic Repository) RNAi based gene therapy for HIV-1, from bench to bedside Von Eije, K.J. Link to publication Citation for published version (APA): Von Eije, K. J. (2009). RNAi based
UvA-DARE (Digital Academic Repository) RNAi based gene therapy for HIV-1, from bench to bedside Von Eije, K.J. Link to publication Citation for published version (APA): Von Eije, K. J. (2009). RNAi based
Retroviruses. ---The name retrovirus comes from the enzyme, reverse transcriptase.
 Retroviruses ---The name retrovirus comes from the enzyme, reverse transcriptase. ---Reverse transcriptase (RT) converts the RNA genome present in the virus particle into DNA. ---RT discovered in 1970.
Retroviruses ---The name retrovirus comes from the enzyme, reverse transcriptase. ---Reverse transcriptase (RT) converts the RNA genome present in the virus particle into DNA. ---RT discovered in 1970.
Technical Bulletin No. 162
 CPAL Central Pennsylvania Alliance Laboratory Technical Bulletin No. 162 cobas 6800 HCV Viral Load Assay - New Platform - June 1, 2017 Contact: Heather Habig, MLS (ASCP) CM, MB CM, 717-851-1422 Operations
CPAL Central Pennsylvania Alliance Laboratory Technical Bulletin No. 162 cobas 6800 HCV Viral Load Assay - New Platform - June 1, 2017 Contact: Heather Habig, MLS (ASCP) CM, MB CM, 717-851-1422 Operations
New Insights on Mechanisms of Foamy Macrophage (FM) Induction and Persistence
 New Insights on Mechanisms of Foamy Macrophage (FM) Induction and Persistence Marian Laderoute, Ph.D. Medical Sciences -Immunology Lab Director Immune System Management Clinic & Lab 80 Aberdeen Street,
New Insights on Mechanisms of Foamy Macrophage (FM) Induction and Persistence Marian Laderoute, Ph.D. Medical Sciences -Immunology Lab Director Immune System Management Clinic & Lab 80 Aberdeen Street,
Figure S1. Schematic presentation of genomic replication of idsiv after transfection and infection. After transfection of idsiv plasmid DNA into 293T
 Figure S1. Schematic presentation of genomic replication of idsiv after transfection and infection. After transfection of idsiv plasmid DNA into 293T cells, the RNA genomes with all modifications are generated
Figure S1. Schematic presentation of genomic replication of idsiv after transfection and infection. After transfection of idsiv plasmid DNA into 293T cells, the RNA genomes with all modifications are generated
HIV viral load testing in the era of ART. Christian Noah Labor Lademannbogen, Hamburg
 HIV viral load testing in the era of ART Christian Noah Labor Lademannbogen, Hamburg 1 Life expectancy of patients on ART Data from the UK Collaborative HIV Cohort (UK CHIC) Requirements: Early diagnosis
HIV viral load testing in the era of ART Christian Noah Labor Lademannbogen, Hamburg 1 Life expectancy of patients on ART Data from the UK Collaborative HIV Cohort (UK CHIC) Requirements: Early diagnosis
Human Rotavirus A. genesig Standard Kit. Non structural protein 5 (NSP5) 150 tests. Primerdesign Ltd. For general laboratory and research use only
 TM Primerdesign Ltd Human Rotavirus A Non structural protein 5 (NSP5) genesig Standard Kit 150 tests For general laboratory and research use only 1 Introduction to Human Rotavirus A Rotavirus is a genus
TM Primerdesign Ltd Human Rotavirus A Non structural protein 5 (NSP5) genesig Standard Kit 150 tests For general laboratory and research use only 1 Introduction to Human Rotavirus A Rotavirus is a genus
Chapter 19: Viruses. 1. Viral Structure & Reproduction. What exactly is a Virus? 11/7/ Viral Structure & Reproduction. 2.
 Chapter 19: Viruses 1. Viral Structure & Reproduction 2. Bacteriophages 3. Animal Viruses 4. Viroids & Prions 1. Viral Structure & Reproduction Chapter Reading pp. 393-396 What exactly is a Virus? Viruses
Chapter 19: Viruses 1. Viral Structure & Reproduction 2. Bacteriophages 3. Animal Viruses 4. Viroids & Prions 1. Viral Structure & Reproduction Chapter Reading pp. 393-396 What exactly is a Virus? Viruses
Diagnostic Methods of HBV and HDV infections
 Diagnostic Methods of HBV and HDV infections Zohreh Sharifi,ph.D Blood Transfusion Research Center, High Institute for Research and Education in Transfusion Medicine Hepatitis B-laboratory diagnosis Detection
Diagnostic Methods of HBV and HDV infections Zohreh Sharifi,ph.D Blood Transfusion Research Center, High Institute for Research and Education in Transfusion Medicine Hepatitis B-laboratory diagnosis Detection
HIV/AIDS. Biology of HIV. Research Feature. Related Links. See Also
 6/1/2011 Biology of HIV Biology of HIV HIV belongs to a class of viruses known as retroviruses. Retroviruses are viruses that contain RNA (ribonucleic acid) as their genetic material. After infecting a
6/1/2011 Biology of HIV Biology of HIV HIV belongs to a class of viruses known as retroviruses. Retroviruses are viruses that contain RNA (ribonucleic acid) as their genetic material. After infecting a
Product # Kit Components
 3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com Pneumocystis jirovecii PCR Kit Product # 42820 Product Insert Background Information
3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com Pneumocystis jirovecii PCR Kit Product # 42820 Product Insert Background Information
Professor Jonathan Weber
 HIV/AIDS at 30: Back to the Future BHIVA / Wellcome Trust Multidisciplinary Event to mark World AIDS Day 2011 British HIV Association (BHIVA) 2011 HIV/AIDS at 30: Back to the Future BHIVA / Wellcome Trust
HIV/AIDS at 30: Back to the Future BHIVA / Wellcome Trust Multidisciplinary Event to mark World AIDS Day 2011 British HIV Association (BHIVA) 2011 HIV/AIDS at 30: Back to the Future BHIVA / Wellcome Trust
cure research HIV & AIDS
 Glossary of terms HIV & AIDS cure research Antiretroviral Therapy (ART) ART involves the use of several (usually a cocktail of three or more) antiretroviral drugs to halt HIV replication. ART drugs may
Glossary of terms HIV & AIDS cure research Antiretroviral Therapy (ART) ART involves the use of several (usually a cocktail of three or more) antiretroviral drugs to halt HIV replication. ART drugs may
Oligo Sequence* bp %GC Tm Hair Hm Ht Position Size Ref. HIVrt-F 5 -CTA-gAA-CTT-TRA-ATg-CAT-ggg-TAA-AAg-TA
 Human immunodeficiency virus (HIV) detection & quantitation by qrt-pcr (Taqman). Created on: Oct 26, 2010; Last modified by: Jul 17, 2017; Version: 3.0 This protocol describes the qrt-pcr taqman based
Human immunodeficiency virus (HIV) detection & quantitation by qrt-pcr (Taqman). Created on: Oct 26, 2010; Last modified by: Jul 17, 2017; Version: 3.0 This protocol describes the qrt-pcr taqman based
Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression.
 Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Supplementary Figure 1 Lentiviral Delivery of Combinatorial mirna Expression Constructs Provides Efficient Target Gene Repression. a, Design for lentiviral combinatorial mirna expression and sensor constructs.
Human diagnostics. Better be Sure: Quantify HDV & HBV viral load. RoboGene product family
 Human diagnostics Better be Sure: Quantify HDV & HBV viral load. RoboGene product family 2 RoboGene Product Family Improved patient management: Standardized monitoring of HBV DNA and HDV RNA viral load.
Human diagnostics Better be Sure: Quantify HDV & HBV viral load. RoboGene product family 2 RoboGene Product Family Improved patient management: Standardized monitoring of HBV DNA and HDV RNA viral load.
Restricted VZV transcription in human trigeminal ganglia
 JVI Accepts, published online ahead of print on 27 June 2012 J. Virol. doi:10.1128/jvi.01331-12 Copyright 2012, American Society for Microbiology. All Rights Reserved. 1 2 Restricted VZV transcription
JVI Accepts, published online ahead of print on 27 June 2012 J. Virol. doi:10.1128/jvi.01331-12 Copyright 2012, American Society for Microbiology. All Rights Reserved. 1 2 Restricted VZV transcription
Dealing with Post-market Issues: PCV Case Study
 Dealing with Post-market Issues: PCV Case Study CASE STUDY: Adventitious agent in raw material ISSUE: Presence of porcine circovirus (PCV-1) DNA detected in marketed rotavirus vaccine by an independent
Dealing with Post-market Issues: PCV Case Study CASE STUDY: Adventitious agent in raw material ISSUE: Presence of porcine circovirus (PCV-1) DNA detected in marketed rotavirus vaccine by an independent
