C R I T F C T E C H N I C A L R E P O R T 11-19
|
|
|
- Johnathan Bond
- 6 years ago
- Views:
Transcription
1 Columbia River Inter-Tribal Fish Commission 729 NE Oregon, Suite Portland, OR C R I T F C T E C H N I C A L R E P O R T Genetic Stock Structure, Relative Productivity and Migration (Gene Flow) of White Sturgeon Among Bonneville, The Dalles, John Day and McNary Reservoirs in the Lower Mid-Columbia River Region Andrew Matala and Blaine Parker October 31, 2011
2 Document ID #P Annual Progress Report Genetic stock structure, relative productivity and migration (gene flow) of white sturgeon among Bonneville, The Dalles, John Day and McNary reservoirs in the lower mid-columbia River region. Prepared by: Andrew Matala 1 and Blaine Parker 2 Columbia River Inter-Tribal Fish Commission 1 Hagerman Fish Culture Experiment Station 3059-F National Fish Hatchery Rd. Hagerman, ID N. E. Oregon, Suite 200 Portland, OR Prepared for: U.S. Department of Energy Bonneville Power Administration Division of Fish and Wildlife P.O. Box 3621 Portland, OR Project Number: Contract Number: Reporting Period: 11/1/10-10/31/11 Submitted: October 31,
3 Abstract This report presents the results of genetic monitoring of white sturgeon (Acipenser transmontanus) in the second of a ten year implementation period for the project. These results will be used to address long-term objectives for monitoring and conservation of populations among four upstream impoundments in the mid Columbia River Basin (CRB): Bonneville, The Dalles, John Day, and McNary reservoirs. Our project objectives include evaluation of population differentiation and migration (gene flow) among reservoirs, and relatedness and effective population size within each reservoir. Further, long-term goals also include exploring parentage assignment between sampled young-of-the-year and mature adult individuals, and the eventual genetic characterization of individuals captured for broodstock in a proposed Yakama Nation sturgeon hatchery program. Table of Contents Acknowledgements...2 Introduction..3 Methods...3 Results...4 Discussion 5 References 6 List of Figures Figure 1.) Numbers of observed alleles...11 Figure 2.) Principal coordinates analysis plot..12 Figure 3.) Scatter plot of allele frequency...13 Figure 4.) Bar plots of proportional population membership...14 Figure 5.) Scatter plot of allele frequency.17 Figure 5.) Principal coordinates analysis plot..18 List of Tables Table 1: Description of nine white sturgeon collections evaluated in Table 2: Population pairwise matrix of among-group variation ( PT) Table 3: Allele frequency comparisons among collections Acknowledgements The authors of this report thank the Bonneville Power Administration for funding this research project, and the BPA staff for their assistance. We appreciate the contributions by all of the project participants. Tissue samples were collected by technicians and staff from the Yakama Nation, Oregon Department of Fish and Game, and the Washington Department of Fish and Wildlife. Linda Lemmon of the Blind Canyon Aqua Ranch provided Snake River broodstock samples for population comparisons. Thanks to Donella Miller for continued support and for serving as a resource for sturgeon management. We appreciate the support of Andrea Drauch- Schreier and UC Davis colleagues in conference to discuss sturgeon issues in the Columbia River, and for data analysis contributions. We are grateful for data collection and quality control contributions in the genetics laboratory from CRITFC technician Lori Maxwell. 2
4 Introduction Our long-term objectives are to characterize the genetic population structure of white sturgeon among four impoundments, to assess migration and gene flow, and effective population size, and relative productivity. In 2011 white sturgeon collections were comprised of 1032 individuals described in the previous (2010) report, and an additional 599 sturgeon sampled in field efforts. Fish were sampled during young of the year (YOY) gill net surveys, during tribal broodstock collection efforts, and during Yakama Nation cooperative tagging efforts. Some individuals were pared from the data set due to incomplete genotypes and duplicate sampling (see Table 1 for sample sizes) as determined by identical genotypes for a pair of sampled fish. In 2011, the sampling surveys were focused on The Dalles reservoir and therefore a disproportionate number of individuals were collected and analyzed from among the four described reservoirs sampled in Between 2010 and 2011, the largest number of samples overall have been collected from John Day Reservoir (Table 1). Additional collections evaluated this report year include a sample from the lower Columbia River below Bonneville dam, and a Snake River reference group for comparison (Table 1). Sampled individuals were young of the year (YOY) and/or juvenile fish. In addition we also analyzed a limited number of fish considered to be potentially mature adults based on fork length measurements. However given the disproportionate numbers of adults across impoundments, the current data set is not yet appropriate for evaluating effective population size and productivity (e.g. through parentage analysis), which will require genotypes from a greater number of sexually mature fish that better represent inclusive populations. Based on the preliminary results presented here, we offer some reasonable speculation on likely genetic structure and the prevalence of gene flow, along with project efforts in the following years to benefit increased population structure resolution and clarification of results. Projected efforts (projected n=1000 per year) in subsequent years will continue to address project needs; the current baseline of genotypes, as additional data accumulates annually, will be valuable over the course of the next few years in evaluating all relevant and feasible population parameter estimates. Methods Genetic evaluation of population structure was largely conducted using the program GenAlEx version 6.41 (Peakall and Smouse 2006). To understand relative levels of population diversity, we evaluated genotypes across a suite of 13 microsatellite loci ( SAT). Observed amplified DNA fragments (here after referred to as alleles) may number up to eight for octoploid white sturgeon. Therefore, alleles are treated as individual loci, and were scored by presence (1) or absence (0) for each individual fish. We calculated sample allele frequencies, defined here as the proportion of individuals within a given population for which a specific allele was observed. In addition, we calculated numbers of observed alleles per collection and locus, and private alleles (occurring in one population exclusively) among collections. Multivariate principle coordinates analysis (PCA; Orloci 1978) was performed to describe population similarities and distinctions. The method reduces redundant variables into a smaller subset of the most informative, where each successive PCA axis explains proportionately less of the total variation. Generally the first 2-3 axes will reveal most of the separation among distinct groups. We used the standardized covariance matrix option based on Nei s unbiased genetic distance (Nei 1978); this measure is one of the most widely used for estimating genetic distance among populations. Partitioning of within- and among- components of total genetic variation were evaluated with analysis of molecular variance (AMOVA) and pairwise genetic distance ( PT) in GenAlEx. The PT statistic is a Euclidean metric analogous to Wrights Fst, and appropriate for binary data. Bayesian cluster analysis was performed using the program STRUCTURE version 2 (Pritchard and Donnelly 2000) to estimate the membership coefficients (Q), or fractional membership in k 3
5 inferred populations for each individual sample in each population. The STRUCTURE results infer levels of gene flow or proportional migration between reservoirs. Results In the 2011 report year we included additional temporal collections from the four impoundments as well as several additional collections (Table 1) including a group of juvenile sturgeon that are the progeny of broodstock captured in the Snake River between Brownlee Dam and Bliss Dam (Linda Lemmon pers. comm.; Kruse-MaIIe 1993; Patterson et al. 1992), and a collection from the Willamette River in the lower Columbia. A second temporal collection from the lower Columbia River was included, and although McNary Reservoir has seen limited numbers of sampled fish to date, we included a collection of adult sturgeon (many believed to be mature) sampled from below Priest Rapids Dam (PRD) downstream to McNary Reservoir (Table 1). In our 2010 report, results of spatial ordination of data using PCA (Figure 1) indicated a distinct sub-cluster of individuals within the John Day impoundment sample, and a second from the Yakama Hatchery broodstock (reference) group. The previously described distinctions in the John Day Reservoir ( type-1 in this report) and in the Yakama captive broodstock both persist in the current analysis. Further, in the updated PCA analysis, we have identified the Snake River collection as a third similarly distinct but unique collection. The remainder of samples from the John Day, Bonneville, McNary, and The Dalles (TDA) impoundments, as well as the Lower Columbia River samples all tended to overlap substantially in PCA (Figure 2). The cumulative percent of total variation among groups that is explained by the first three axes is 82.4%, and is partitioned as follows: x-axis (45.9%), y-axis (21.5%), z-axis (15.0%). In AMOVA analysis, a significant proportion of the total variation across the data set (5%) was attributable to variation among collections, and it was significant (P<0.001). The AMOVA results also revealed significant among-group variation between collections from below Bonneville Dam and those above, but this was largely driven by a small number of pairwise PT comparisons (Table 2). In fact, Willamette River was only significantly different from PRD MCN 2011 in pairwise comparisons with temporal collections from the four impoundments (Table 2). Based on AMOVA and PCA results, it was deemed appropriate to combine temporal samples by impoundment-of-origin from 2010 and Numbers of observed alleles and allele frequency (indicating degree of heterozygosity) was highly variable among collections (Figure 1; Table 3). Numbers of alleles was greatest in the four impoundments but varied across years, likely an artifact of sample sizes. Fewest number of observed alleles occurred in the John Day type-1 collection, followed by the two domesticated stocks in the Snake River and Yakama hatchery. When collections within each impoundment were pooled across years the numbers of observed alleles was relatively stable (Figure 1). The John Day Reservoir collections had the greatest number of private alleles, followed by Bonneville impoundment (Figure 1; Table 3). For eight of the 13 loci involved, John Day reservoir collections accounted for the greatest number of observed alleles, and at three loci John Day shared that distinction with at least one other group (Table 3). This is not surprising given that the John Day sample sizes are significantly larger than the other collections. This may indicate that actual genetic diversity in the other impoundments and collections has not yet been well represented or accurately estimated. Results of Bayesian cluster analysis revealed five likely or inferred populations (k=5). Assigning group membership of individual fish across collections (Table 1; Figure 4) to inferred clusters gave results that substantiated the PCA analysis. In addition, the Bayesian results indicated high mean membership fidelity (76%) of Yakama broodstock to cluster #2 (Q2 in Table 1), Snake River (96.5%) to cluster #4 (Q4 in Table 1), and John Day ( type-1) to cluster #3 4
6 (98%). In fact, in all three outlier groups nearly all individuals could be characterized by greater than 9% membership to respective group (Figure 4; first three histograms). In the Yakama broodstock there are 12 individuals that appear to be of an entirely different origin than the remainder of that collection. The LCR collections were predominately highest for membership coefficient Q1 (75%), and to a lesser degree (64%) the Willamette River collection. By comparison, the McNary, John Day, and Bonneville impoundment collections were generally evenly split between Q1 and Q5. Note that in a plot of allele frequencies, the Yakama, Snake River and John Day type-1 collections have relatively high frequencies or complete absence of the allele, while all other collections track very closely with one another (Figure 5; example alleles are depicted from among those with the greatest variation across collections). This indicates occurance of relatively rare alleles, but also less diversity in these populations. Of special interest among clustering analysis results was the observation of three individuals in the PRD to McNary collection that appear to share a high degree of similarity to the John Day type-1 group from 2010 (Figure 4d). Similarly, there are several individuals from throughout the remaining collections that belong to the Snake River population with membership coefficients in excess of 50%. For a separate analysis, those individuals were pared from their respective collections (impoundment) of origin. A group named like Snake River and identified by greater than 50% membership in the Snake River group included: 1 Bonneville individual, 3 PRD-McNary individuals, 3 The Dalles individuals, and 19 John Day individuals. A group named PRD (like JD type-1) included three individuals collected from near Priest Rapids Dam with greater than 94% membership in the John Day type-1 group. The two additional populations were treated uniquely in PCA analysis and results confirmed their population similarities (Figure 6). By portioning these individuals from their original putative populations of origin, the overall PCA results more accurately resolved the total variation in the data set. The cumulative percent of total variation among groups that is explained by the first three axes was revised to 92.2%, and is partitioned as follows: x-axis (59.7%), y-axis (19.7%), z-axis (12.9%). Discussion Interpretation of the body of results from this second year of analysis in the proposed ongoing ten year project, may represent unique population distinctions, but should be approached with caution. There are obvious biases in the data largely related to skewed and deficient sample sizes for some collections. Moreover, in the absence of more definitive relatedness data and results, it is difficult to evaluate relative productivity or even population structure among the CRB impoundments. With that caveat, it appears that collections among lower Columbia River, Willamette River, Bonneville, The Dalles, John Day and McNary impoundments are largely similar with little discernible genetic differentiation. However, the similarity appears to occur on somewhat of a cline. Recall that mid Columbia collections are more similar to each other in the STRUCTURE analysis, split between two inferred population clusters. The proportional similarity of downstream (below Bonneville Dam) collections is more homogeneous. Further, those collections located geographically intermediate (i.e., the mid Columbia from McNary reservoir to Bonneville Dam) exhibit an influence by a possible distinct upstream population/s in the Snake River (a second potentially distinct influence may also originate from the upper Columbia River above Priest Rapids Dam). One may speculate that this is indicative of a high rate of gene flow that is one-directional, as juveniles may be more likely to migrate downstream through the reservoirs while tendencies to travel upstream via ladders is less likely. The Snake River collection may confirm to this suggestion, yet the barriers involved in the Snake River region may not provide comparable juvenile passage capability, and adults are essentially excluded from upstream movement. Given that the lower Columbia River sturgeon populations is large (total abundance) in comparison, influences from smaller upstream populations may be more difficult to detect. 5
7 Lastly, there is reasonable evidence to suggest that impoundment specific productivity varies, where a likely cohort of individuals that is markedly different in genetic signature was identified in the John Day reservoir in 2010 (John Day type-1 ). However, in 2011 we observed large, potentially mature individuals far upstream that appear to be of the same origin. This may suggest the influence of an as yet unidentified population (e.g., Kootenai or other upstream population) contributing to the populations sampled in the data set. The degree of distinction of the John Day type-1 collection in comparison to the other (natural-origin) sampled individuals is on par with the degree of distinction recognized among a long established broodstock of unknown origin at the Yakama Hatchery (YAK) and a newly added aquaculture collection originating from the Snake River. Lower observed allelic diversity may suggest a founder effect and small effective population sizes. Alternatively, rather than representing unique populations it could be argued that those broodstock and/or artificially reared individuals (and natural origin groups) share either a high level of relatedness (i.e., full siblings) or are exhibiting a high degree of inbreeding among individuals sampled from each group. With the continuation of this project and a more balanced sampling approach that represents all reservoirs equally, we expect that the current two-year accumulation of results will be greatly clarified and/or validated with improved estimates and conclusions regarding the genetic relationship (e.g. gene flow) between the four impoundments. Of particular concern or specific interest is a private sturgeon hatchery operated by Henry Pelfry that has contributed to both mitigation releases in the John Day Reservoir and in the Willamette River. Moreover, it is believed that much of the current domesticated stock at the Yakama artificial propagation facility originated from Henry Pelfry s hatchery (Donella Miller, pers. comm.). Lastly, a hatchery stock reared at Abernathy Fish Technology Center was recently transferred to the Yakama artificial propagation facility (Smith and Godfrey 2009), and this stock also originated from Henry Pelfry s hatchery. It will therefore be beneficial to genetically characterize this stock in order to discern what influences it may have imparted on the current population structure among the mid- Columbia River impoundments. We will continue to target both YOY and older (mature) fish in subsequent years to allow for implementation of additional kinds of proposed analyses defined in our overall objectives (e.g. parentage), which will help to better understand the relative sizes and productivity potentials of populations among the four impoundments References Evanno, G., S. Regnaut, and J. Goudet Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Molecular Ecology 14: Kruse-MaIIe, G. O White sturgeon evaluations in the Snake River. Job Performance Report, Project F-73-R-15, Subproject IV, Study IV. Volume 93, Article 07. Nei, M Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics, 89: Orloci, L Multivariate analysis in vegetation research. The Hague: Dr W. Junk B. V. Peakall, R. and P. E. Smouse GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Molecular Ecology Notes 6: Patterson, T., K. A. Apperson and J. T. Siple Snake River White Sturgeon Culture Volume 100, Article 01. 6
8 Peakall, R., and P. E. Smouse GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Molecular Ecology Notes. 6, Pritchard, J. K., M. Stephens and P. Donnelly Inference of population structure Using multilocus genotype data. Genetics, 155: Smith, C. and L. Godfrey Relatedness among white sturgeon sent from Abernathy Fish Technology Center to Yakama Nation Prosser Facility in United States Fish and Wildlife Service Report. Abernathy Fish Technology Center, Applied Program in Conservation Genetics, Longview, WA 98632, 7
9 Table 1.) Description of nine white sturgeon collections analyzed between 2010 and Some collections are described in our 2010 report. Temporal collections were combined for analysis. Note the limited availability of samples from the Willamette River and McNary impoundment. Values in parenthesis indicate number of excluded individuals due to duplicate sampling. The symbol (*) indicates a sample size less than the total due to incomplete data for all individuals. The (---) entry represents no available data. The mean Q-values from the program STRUCTURE (Pritchard et al. 2007) are proportional membership in k=5 inferred groups (Q1-5). Bolded values indicate the highest proportional membership assignment. fork length (cm) membership coefficient population total (n) (n) min max mean life history stage Q1 Q2 Q3 Q4 Q5 Lower Columbia 81 34* YOY/juvenile 2* subadult adult Willamette YOY/juvenile subadult adult Bonneville YOY/juvenile subadult adult The Dalles 498; (6) 267* YOY/juvenile 218* subadult * adult John Day 42; (1) YOY/juvenile ("type-1") subadult adult John Day 578: (10) YOY/juvenile subadult adult PRD - McNary 44; (1) YOY/juvenile 8
10 subadult adult Snake River 102; (1) broodstock YN broodstock broodstock
11 Table 2: Population pairwise matrix of among-group variation ( PT). The PT values reside below the diagonal, and the corresponding P-values are above the diagonal. Significant pairwise comparisons (P<0.01) are indicated by bold italics. PT PRD - MCN (2010) PRD - MCN (2011) Snake River (2011) Bonneville (2010) Bonneville (2011) John Day ("type-1") John Day (2010) John Day (2011) Lower Columbia (2010) Lower Columbia (2011) The Dalles (2010) The Dalles (2011) Willamette (2011) YN broostock (2009) PRD - MCN (2010) PRD - MCN (2011) Snake River (2011) Bonneville (2010) Bonneville (2011) John Day ("type-1") John Day (2010) John Day (2011) Lower Columbia (2010) Lower Columbia (2011) The Dalles (2010) The Dalles (2011) Willamette (2011) YN broostock (2009)
12 Figure 1. Numbers of observed alleles (fragments) per temporal collection from 2010 to Number of observed private alleles (secondary axis) are indicated by filled circles. Dashed lines represent numbers of observed alleles for each location when temporal collections are pooled; note similar numbers for four mid-columbia impoundments total # observed alleles # private alleles
13 Figure 2. Principle coordinates analysis plots. The two perspectives (A and B) show the same ordination of the data points in space but with different rotations on the x and y axes to show depth and separation in three dimensions. A. y 1.) Lower Columbia River ) Lower Columbia River ) Willamette River ) Bonneville Impoundment ) Bonneville Impoundment ) John Day Impoundment ) The Dalles Impoundment ) The Dalles Impoundment ) John Day Impoundment ) PRD - McNary Impoundment ) PRD - McNary Impoundment ) Yakama Broodstock ) Snake River 14.) John Day TYPE-1 (2010) z x B y 14 x z 13 12
14 Figure 3. Scatter plots showing allele frequencies for all 13 microsatellite loci employed. Plots represent alleles with representatively high variation among all alleles observed. A list of all observed alleles is presented in Table 2. Willamette Lower Columbia Bonneville The Dalles JohnDay John Day "type-1" PDR/ McNary Snake River YN broodstock Population proportion Population proportion 13
15 Figure 4. Bayesian Cluster Analysis histograms generated using the program STRUCTURE. The x-axis is individual sample and the y-axis is proportional membership. Note the first three panels ( A-C) represent highly homogenous collections: Snake River is Q4, John Day type-1 is Q3, and Yakama broodstock is Q2 (Table 1). The remaining panels (D-M) depict temporal collections in the Lower Columbia River where Q1 is represented by dark grey and Q5 is represented by light grey. A. 100% Snake River : Blind Canyon Aqua-Ranch 90% 80% 70% 60% 50% 40% 30% 20% 10% 0% B. 100% 90% 80% 70% 60% 50% 40% 30% 20% 10% 0% John Day Impoundment TYPE-1 (2010) C. 100% Yakama Nation Broodstock (2009) 90% 80% 70% 60% 50% 40% 30% 20% 10% 0% 14
16 D. 100% Priest Rapids Dam to McNary Reservoir ( ) G. 100% Lower Columbia River % 90% 80% 80% 70% 70% 60% 60% 50% 50% 40% 40% 30% 30% 20% 20% 10% 10% 0% 0% E. 100% Bonneville Impoundment 2010 H. 100% Lower Columbia River % 90% 80% 80% 70% 70% 60% 60% 50% 50% 40% 40% 30% 30% 20% 20% 10% 10% 0% 0% F. 100% Bonneville Impoundment 2011 I. 100% Willamette River % 90% 80% 80% 70% 70% 60% 60% 50% 50% 40% 40% 30% 30% 20% 20% 10% 10% 0% 0% 15
17 J. K. 100% 90% 80% 70% 60% 50% 40% 30% 20% 10% 0% 100% 90% 80% 70% 60% 50% 40% 30% 20% 10% 0% John Day Impoundment 2010 John Day Impoundment 2011 L. 100% 90% 80% 70% 60% 50% 40% 30% 20% 10% 0% M. 100% 90% 80% 70% 60% 50% 40% 30% 20% 10% 0% The Dalles Impoundment 2010 The Dalles Impoundment
18 Figure 5. Scatter plots showing allele frequencies for all 13 microsatellite loci employed (see Figure 3). All collections except the three outlier collections are represented by black dashes, and the connecting dashed lines indicate similarity in trend of allele frequencies across collections. John Day "type-1" Snake River YN broodstock Population proportion Population proportion 17
19 Figure 6. Principle coordinates analysis plots. The two perspectives (A and B) show the same ordination of the data points in space but with different rotations on the x and y axes to show depth and separation in three dimensions. Plots show association or similarity in clustering between collections (Table 1) and outlier individuals identified as members of groups different from collection of origin (Figure 4). A. 1.) Lower Columbia River 2.) Willamette River 3.) Bonneville Impoundment 4.) John Day Impoundment 5.) The Dalles Impoundment 6.) PRD - McNary Impoundment ) Yakama Broodstock ) Snake River 9.) John Day TYPE-1 (2010) y x PRD (Like JD type-1 ) 9 7 Like Snake River z 8 B. y z x PRD (Like JD type-1 ) 9 Like Snake River 8 18
20 Table 3.) Allele (fragment) frequency comparisons at 13 SAT loci, among 14 collections from the Columbia River Basin sampled between 2010 and 2011 (see Figure 1). Collections are defined in Table 1. Alleles are designated by size in nucleotide base pairs (bp). Total number of observed alleles appears in parentheses for each locus overall, and within each collection. Private alleles (P) are those that were observed in only a single collection (underlined and bolded). Locus Allele Lower Columbia (2010) Lower Columbia (2011) Willamette Bonneville (2010) Bonneville (2011) The Dalles (2010) The Dalles (2011) John Day ("type-1) John Day (2010) John Day (2011) PRD-McNary (2010) PRD-McNary (2011) Snake River YN broodstock AciG2 (7) (6) (7) (6) (6) (6) (6) (6) (3) (6) (6) (5) (6) (4) (6) P AciG53 (10) (8) (8) (6) (9) (8) (8) (8) (4) (9) (5) (3) (6) (5) (6)
21 Actm177 (7) (6) (5) (5) (6) (5) (6) (6) (4) (7) (5) (4) (6) (4) (5) Actm43 (25) (15) (16) (14) (20) (13) (18) (19) (6) (22) (14) (9) (16) (7) (7) P
22 P P P P Actm52 (27) (23) (19) (15) (23) (18) (22) (24) (10) (24) (16) (15) (19) (15) (16) P P P
23 Atr105 (10) (6) (7) (5) (7) (6) (7) (9) (4) (10) (7) (5) (8) (6) (5) P AciG35 (21) (18) (18) (15) (20) (18) (19) (20) (9) (21) (16) (12) (18) (13) (11)
24 Atr1101 (9) (8) (7) (6) (9) (7) (9) (9) (5) (9) (5) (4) (7) (5) (5) Atr117 (31) (20) (19) (17) (25) (14) (23) (26) (6) (28) (15) (8) (16) (12) (11) P
25 P P AciG110 (25) (21) (20) (17) (23) (14) (18) (22) (10) (25) (12) (10) (16) (10) (10)
26 Atr107 (29) (22) (24) (16) (26) (18) (27) (26) (8) (27) (20) (11) (20) (15) (16)
27 P P Atr109 (34) (23) (20) (19) (28) (20) (24) (28) (9) (30) (20) (13) (23) (12) (12) P
Reliability of Ordination Analyses
 Reliability of Ordination Analyses Objectives: Discuss Reliability Define Consistency and Accuracy Discuss Validation Methods Opening Thoughts Inference Space: What is it? Inference space can be defined
Reliability of Ordination Analyses Objectives: Discuss Reliability Define Consistency and Accuracy Discuss Validation Methods Opening Thoughts Inference Space: What is it? Inference space can be defined
Preliminary results of parentage analyses of 2010 Tobique River pre-smolt
 Preliminary results of parentage analyses of 2010 Tobique River pre-smolt Patrick O Reilly, Ross Jones, Trevor Goff, Stephanie Ratelle, and Lorraine Hamilton Presented by: Sherisse McWilliam-Hughes The
Preliminary results of parentage analyses of 2010 Tobique River pre-smolt Patrick O Reilly, Ross Jones, Trevor Goff, Stephanie Ratelle, and Lorraine Hamilton Presented by: Sherisse McWilliam-Hughes The
Relative reproductive success of hatchery and wild Steelhead in the Hood River
 Relative reproductive success of hatchery and wild Steelhead in the Hood River Final report on work conducted under BPA Intergovernmental Contract 9245, Project # 1988-053-12 and ODFW Interagency agreement
Relative reproductive success of hatchery and wild Steelhead in the Hood River Final report on work conducted under BPA Intergovernmental Contract 9245, Project # 1988-053-12 and ODFW Interagency agreement
July 2017 Report EVALUATING SPRING CHINOOK SALMON REINTRODUCTIONS ABOVE DETROIT DAM, ON THE NORTH SANTIAM RIVER, USING GENETIC PARENTAGE ANALYSIS
 July 2017 Report EVALUATING SPRING CHINOOK SALMON REINTRODUCTIONS ABOVE DETROIT DAM, ON THE NORTH SANTIAM RIVER, USING GENETIC PARENTAGE ANALYSIS Prepared for: U. S. ARMY CORPS OF ENGINEERS PORTLAND DISTRICT
July 2017 Report EVALUATING SPRING CHINOOK SALMON REINTRODUCTIONS ABOVE DETROIT DAM, ON THE NORTH SANTIAM RIVER, USING GENETIC PARENTAGE ANALYSIS Prepared for: U. S. ARMY CORPS OF ENGINEERS PORTLAND DISTRICT
Nature Neuroscience: doi: /nn Supplementary Figure 1. Behavioral training.
 Supplementary Figure 1 Behavioral training. a, Mazes used for behavioral training. Asterisks indicate reward location. Only some example mazes are shown (for example, right choice and not left choice maze
Supplementary Figure 1 Behavioral training. a, Mazes used for behavioral training. Asterisks indicate reward location. Only some example mazes are shown (for example, right choice and not left choice maze
STATUS REPORT - PINNIPED PREDATION AND DETERRENT ACTIVITIES AT BONNEVILLE DAM, 2009
 STATUS REPORT - PINNIPED PREDATION AND DETERRENT ACTIVITIES AT BONNEVILLE DAM, 29 Robert Stansell, Sean Tackley, and Karrie Gibbons - (541) 374-881 Fisheries Field Unit U.S. Army Corps of Engineers Bonneville
STATUS REPORT - PINNIPED PREDATION AND DETERRENT ACTIVITIES AT BONNEVILLE DAM, 29 Robert Stansell, Sean Tackley, and Karrie Gibbons - (541) 374-881 Fisheries Field Unit U.S. Army Corps of Engineers Bonneville
Operational Guidelines for Pacific Salmon Hatcheries Production Planning, Broodstock Collection and Spawning Scope of Guidelines
 Operational Guidelines for Pacific Salmon Hatcheries Production Planning, Broodstock Collection and Spawning Scope of Guidelines These guidelines have been developed to guide production planning, broodstock
Operational Guidelines for Pacific Salmon Hatcheries Production Planning, Broodstock Collection and Spawning Scope of Guidelines These guidelines have been developed to guide production planning, broodstock
STATUS REPORT - PINNIPED PREDATION AND DETERRENT ACTIVITIES AT BONNEVILLE DAM, 2009
 STATUS REPORT - PINNIPED PREDATION AND DETERRENT ACTIVITIES AT BONNEVILLE DAM, 29 Robert Stansell, Sean Tackley, and Karrie Gibbons - (541) 374-881 Fisheries Field Unit U.S. Army Corps of Engineers Bonneville
STATUS REPORT - PINNIPED PREDATION AND DETERRENT ACTIVITIES AT BONNEVILLE DAM, 29 Robert Stansell, Sean Tackley, and Karrie Gibbons - (541) 374-881 Fisheries Field Unit U.S. Army Corps of Engineers Bonneville
Oregon Pinnipeds: Status, Trends, & Management. Robin Brown Oregon Department of Fish and Wildlife Marine Mammal Program
 Oregon Pinnipeds: Status, Trends, & Management Robin Brown Oregon Department of Fish and Wildlife Marine Mammal Program Acknowledgments NOAA Fisheries National Marine Mammal Laboratory Washington Department
Oregon Pinnipeds: Status, Trends, & Management Robin Brown Oregon Department of Fish and Wildlife Marine Mammal Program Acknowledgments NOAA Fisheries National Marine Mammal Laboratory Washington Department
Parentage and relatedness determination in farmed Atlantic salmon. Ashie T. Norris*, Daniel G. Bradley and Edward P. Cunningham.
 Parentage and relatedness determination in farmed Atlantic salmon (Salmo salar) using microsatellite markers Ashie T. Norris*, Daniel G. Bradley and Edward P. Cunningham. Department of Genetics, Trinity
Parentage and relatedness determination in farmed Atlantic salmon (Salmo salar) using microsatellite markers Ashie T. Norris*, Daniel G. Bradley and Edward P. Cunningham. Department of Genetics, Trinity
STATUS REPORT - PINNIPED PREDATION AND DETERRENT ACTIVITIES AT BONNEVILLE DAM, 2009
 STATUS REPORT - PINNIPED PREDATION AND DETERRENT ACTIVITIES AT BONNEVILLE DAM, 29 Robert Stansell, Sean Tackley, and Karrie Gibbons - (541) 374-881 Fisheries Field Unit U.S. Army Corps of Engineers Bonneville
STATUS REPORT - PINNIPED PREDATION AND DETERRENT ACTIVITIES AT BONNEVILLE DAM, 29 Robert Stansell, Sean Tackley, and Karrie Gibbons - (541) 374-881 Fisheries Field Unit U.S. Army Corps of Engineers Bonneville
Supportive breeding boosts natural population abundance with minimal negative impacts on fitness of a wild population of Chinook salmon
 Molecular Ecology (2012) 21, 5236 5250 doi: 10.1111/mec.12046 Supportive breeding boosts natural population abundance with minimal negative impacts on fitness of a wild population of Chinook salmon MAUREEN
Molecular Ecology (2012) 21, 5236 5250 doi: 10.1111/mec.12046 Supportive breeding boosts natural population abundance with minimal negative impacts on fitness of a wild population of Chinook salmon MAUREEN
STATUS REPORT PINNIPED PREDATION AND DETERRENT ACTIVITIES AT BONNEVILLE LOCK AND DAM. May 3, 2017
 STATUS REPORT PINNIPED PREDATION AND DETERRENT ACTIVITIES AT BONNEVILLE LOCK AND DAM May 3, 217 Prepared by: Kyle Tidwell, Bjorn van der Leeuw, Thomas Van Hevelingen Fisheries Field Unit U.S. Army Corps
STATUS REPORT PINNIPED PREDATION AND DETERRENT ACTIVITIES AT BONNEVILLE LOCK AND DAM May 3, 217 Prepared by: Kyle Tidwell, Bjorn van der Leeuw, Thomas Van Hevelingen Fisheries Field Unit U.S. Army Corps
Supplemental Figures. Figure S1: 2-component Gaussian mixture model of Bourdeau et al. s fold-change distribution
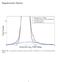 Supplemental Figures Figure S1: 2-component Gaussian mixture model of Bourdeau et al. s fold-change distribution 1 All CTCF Unreponsive RefSeq Frequency 0 2000 4000 6000 0 500 1000 1500 2000 Block Size
Supplemental Figures Figure S1: 2-component Gaussian mixture model of Bourdeau et al. s fold-change distribution 1 All CTCF Unreponsive RefSeq Frequency 0 2000 4000 6000 0 500 1000 1500 2000 Block Size
Supplementary Figure 1: Features of IGLL5 Mutations in CLL: a) Representative IGV screenshot of first
 Supplementary Figure 1: Features of IGLL5 Mutations in CLL: a) Representative IGV screenshot of first intron IGLL5 mutation depicting biallelic mutations. Red arrows highlight the presence of out of phase
Supplementary Figure 1: Features of IGLL5 Mutations in CLL: a) Representative IGV screenshot of first intron IGLL5 mutation depicting biallelic mutations. Red arrows highlight the presence of out of phase
Evaluating minijack rates in spring Chinook Salmon CONFIRMING THAT SPRING PLASMA 11-KETOTESTOSTERONE LEVEL ACCURATELY PREDICTS FALL MATURATION STATUS
 Evaluating minijack rates in spring Chinook Salmon CONFIRMING THAT SPRING PLASMA 11-KETOTESTOSTERONE LEVEL ACCURATELY PREDICTS FALL MATURATION STATUS Background information What are minijacks and why do
Evaluating minijack rates in spring Chinook Salmon CONFIRMING THAT SPRING PLASMA 11-KETOTESTOSTERONE LEVEL ACCURATELY PREDICTS FALL MATURATION STATUS Background information What are minijacks and why do
period. The distribution of PREs and TCs, stratified by synoptic category
 3. Climatology of PREs during 1988 2008 3.1 Overview A total of 56 PREs associated with 38 Atlantic basin TCs were identified for the 1988 2008 period. The distribution of PREs and TCs, stratified by synoptic
3. Climatology of PREs during 1988 2008 3.1 Overview A total of 56 PREs associated with 38 Atlantic basin TCs were identified for the 1988 2008 period. The distribution of PREs and TCs, stratified by synoptic
TECHNICAL ANNEX with country details
 WG ECOSTAT report on common understanding of using mitigation s for reaching Good Ecological Potential for heavily modified water bodies Part 1: Impacted by water storage TECHNICAL ANNEX with country details
WG ECOSTAT report on common understanding of using mitigation s for reaching Good Ecological Potential for heavily modified water bodies Part 1: Impacted by water storage TECHNICAL ANNEX with country details
Monitoring for precocious male maturation (minijacks) in hatchery programs: A case study of upper Columbia River summer Chinook salmon
 Monitoring for precocious male maturation (minijacks) in hatchery programs: A case study of upper Columbia River summer Chinook salmon Don Larsen, Brian Beckman, Paul Parkins, Kathy Cooper, Deb Harstad,
Monitoring for precocious male maturation (minijacks) in hatchery programs: A case study of upper Columbia River summer Chinook salmon Don Larsen, Brian Beckman, Paul Parkins, Kathy Cooper, Deb Harstad,
Relationship between Project Operations and Spring Chinook Adult Passage at Little Goose Dam
 FISH PASSAGE CENTER 847 N.E. 19 th Avenue, #250, Portland, Oregon 97232 Phone: (503) 833-3900 Fax: (503) 232-1259 www.fpc.org e-mail us at fpcstaff@fpc.org MEMORANDUM TO: Michele DeHart FROM: Bobby Hsu,
FISH PASSAGE CENTER 847 N.E. 19 th Avenue, #250, Portland, Oregon 97232 Phone: (503) 833-3900 Fax: (503) 232-1259 www.fpc.org e-mail us at fpcstaff@fpc.org MEMORANDUM TO: Michele DeHart FROM: Bobby Hsu,
SUPPLEMENTARY INFORMATION In format provided by Javier DeFelipe et al. (MARCH 2013)
 Supplementary Online Information S2 Analysis of raw data Forty-two out of the 48 experts finished the experiment, and only data from these 42 experts are considered in the remainder of the analysis. We
Supplementary Online Information S2 Analysis of raw data Forty-two out of the 48 experts finished the experiment, and only data from these 42 experts are considered in the remainder of the analysis. We
Feeding A Natural Marine Based Diet. To Improve Health and Survival of Reconditioned Kelt Steelhead
 Feeding A Natural Marine Based Diet To Improve Health and Survival of Reconditioned Kelt Steelhead Project Background In 2003, the Colville Confederated Tribes began participation in a reproductive success
Feeding A Natural Marine Based Diet To Improve Health and Survival of Reconditioned Kelt Steelhead Project Background In 2003, the Colville Confederated Tribes began participation in a reproductive success
Clarification of December 9, 2011 memo regarding Little Goose spill operations and adult Chinook conversion rates and travel times
 FISH PASSAGE CENTER 1827 NE 44 th Ave., Suite 240, Portland, OR 97213 Phone: (503) 230-4099 Fax: (503) 230-7559 http://www.fpc.org/ e-mail us at fpcstaff@fpc.org MEMORANDUM TO: Fish Passage Advisory Committee
FISH PASSAGE CENTER 1827 NE 44 th Ave., Suite 240, Portland, OR 97213 Phone: (503) 230-4099 Fax: (503) 230-7559 http://www.fpc.org/ e-mail us at fpcstaff@fpc.org MEMORANDUM TO: Fish Passage Advisory Committee
Ecological Statistics
 A Primer of Ecological Statistics Second Edition Nicholas J. Gotelli University of Vermont Aaron M. Ellison Harvard Forest Sinauer Associates, Inc. Publishers Sunderland, Massachusetts U.S.A. Brief Contents
A Primer of Ecological Statistics Second Edition Nicholas J. Gotelli University of Vermont Aaron M. Ellison Harvard Forest Sinauer Associates, Inc. Publishers Sunderland, Massachusetts U.S.A. Brief Contents
Michael Jepson, Ken Tolotti, Christopher Peery, and Brian Burke University of Idaho and NOAA Fisheries
 To: From: David Clugston, USACE Portland District Michael Jepson, Ken Tolotti, Christopher Peery, and Brian Burke University of Idaho and NOAA Fisheries RE: Radio-telemetry data for Chinook salmon at Bonneville
To: From: David Clugston, USACE Portland District Michael Jepson, Ken Tolotti, Christopher Peery, and Brian Burke University of Idaho and NOAA Fisheries RE: Radio-telemetry data for Chinook salmon at Bonneville
Mike Jepson, Ken Tolotti, Chris Peery, and Brian Burke University of Idaho and NOAA Fisheries
 To: From: David Clugston, USACE Portland District Mike Jepson, Ken Tolotti, Chris Peery, and Brian Burke University of Idaho and NOAA Fisheries RE: Radio-telemetry data for Chinook salmon at Bonneville
To: From: David Clugston, USACE Portland District Mike Jepson, Ken Tolotti, Chris Peery, and Brian Burke University of Idaho and NOAA Fisheries RE: Radio-telemetry data for Chinook salmon at Bonneville
Biology of Breeding: Considerations for maximizing genetic diversity of breeding groups
 Dillon Damuth 03/01/2015 Biology of Breeding: Considerations for maximizing genetic diversity of breeding groups When a person joins the hobby of reptile keeping and make the decision to breed animals
Dillon Damuth 03/01/2015 Biology of Breeding: Considerations for maximizing genetic diversity of breeding groups When a person joins the hobby of reptile keeping and make the decision to breed animals
How is genetic taken into account in captive breeding program? Asan Hilal Dubois Anne-Cécile
 How is genetic taken into account in captive breeding program? Asan Hilal Dubois Anne-Cécile Content What is captive breeding? Disadvantages of captive breeding Genetic as a solution? How genetic is used?
How is genetic taken into account in captive breeding program? Asan Hilal Dubois Anne-Cécile Content What is captive breeding? Disadvantages of captive breeding Genetic as a solution? How genetic is used?
CRYOPRESERVATION OF KOOTENAI RIVER WHITE STURGEON
 CRYOPRESERVATION OF KOOTENAI RIVER WHITE STURGEON (Acipenser transmontanus) GAMETES 2009 ANNUAL REPORT A report to the Kootenai Tribe of Idaho September 2009 Prepared by Steven J. Patton and Joseph G.
CRYOPRESERVATION OF KOOTENAI RIVER WHITE STURGEON (Acipenser transmontanus) GAMETES 2009 ANNUAL REPORT A report to the Kootenai Tribe of Idaho September 2009 Prepared by Steven J. Patton and Joseph G.
Non-Invasive Sampling of Schistosomes from Humans Requires Correcting for Family Structure
 Non-Invasive Sampling of Schistosomes from Humans Requires Correcting for Family Structure Michelle L. Steinauer 1 *, Mark R. Christie 2, Michael S. Blouin 2, Lelo E. Agola 3, Ibrahim N. Mwangi 3, Geoffrey
Non-Invasive Sampling of Schistosomes from Humans Requires Correcting for Family Structure Michelle L. Steinauer 1 *, Mark R. Christie 2, Michael S. Blouin 2, Lelo E. Agola 3, Ibrahim N. Mwangi 3, Geoffrey
WEST BRANCH MONTREAL RIVER INTERNET FLOW STUDY OCTOBER 2007
 AMERICAN WHITEWATER PO BOX 1540 CULLOWHEE, NC 28723 WEST BRANCH MONTREAL RIVER INTERNET FLOW STUDY OCTOBER 2007 EVAN STANFORD and THOMAS O KEEFE AMERICAN WHITEWATER www.americanwhitewater.org ABSTRACT
AMERICAN WHITEWATER PO BOX 1540 CULLOWHEE, NC 28723 WEST BRANCH MONTREAL RIVER INTERNET FLOW STUDY OCTOBER 2007 EVAN STANFORD and THOMAS O KEEFE AMERICAN WHITEWATER www.americanwhitewater.org ABSTRACT
Analysis of Environmental Data Conceptual Foundations: En viro n m e n tal Data
 Analysis of Environmental Data Conceptual Foundations: En viro n m e n tal Data 1. Purpose of data collection...................................................... 2 2. Samples and populations.......................................................
Analysis of Environmental Data Conceptual Foundations: En viro n m e n tal Data 1. Purpose of data collection...................................................... 2 2. Samples and populations.......................................................
Supporting Information
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 Supporting Information Variances and biases of absolute distributions were larger in the 2-line
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 Supporting Information Variances and biases of absolute distributions were larger in the 2-line
Operation Compliance Monitoring Plan (License Article 407) Annual Report for Water Year July 2014 June 2015
 Henry M. Jackson Hydroelectric Project (FERC No. 2157) Operation Compliance Monitoring Plan (License Article 407) Annual Report for Water Year July 2014 June 2015 Prepared By: Everett, WA August 2015 Final
Henry M. Jackson Hydroelectric Project (FERC No. 2157) Operation Compliance Monitoring Plan (License Article 407) Annual Report for Water Year July 2014 June 2015 Prepared By: Everett, WA August 2015 Final
Summary. Introduction. Atypical and Duplicated Samples. Atypical Samples. Noah A. Rosenberg
 doi: 10.1111/j.1469-1809.2006.00285.x Standardized Subsets of the HGDP-CEPH Human Genome Diversity Cell Line Panel, Accounting for Atypical and Duplicated Samples and Pairs of Close Relatives Noah A. Rosenberg
doi: 10.1111/j.1469-1809.2006.00285.x Standardized Subsets of the HGDP-CEPH Human Genome Diversity Cell Line Panel, Accounting for Atypical and Duplicated Samples and Pairs of Close Relatives Noah A. Rosenberg
Managing Precocious Maturation in Chinook Salmon Captive Broodstock
 Managing Precocious Maturation in Chinook Salmon Captive Broodstock Paul Adelizi, Jamie McGrath-Castro and Brian Erlandsen California Department of Fish and Wildlife 2017 Northwest Fish Culture Concepts
Managing Precocious Maturation in Chinook Salmon Captive Broodstock Paul Adelizi, Jamie McGrath-Castro and Brian Erlandsen California Department of Fish and Wildlife 2017 Northwest Fish Culture Concepts
WDHS Curriculum Map Probability and Statistics. What is Statistics and how does it relate to you?
 WDHS Curriculum Map Probability and Statistics Time Interval/ Unit 1: Introduction to Statistics 1.1-1.3 2 weeks S-IC-1: Understand statistics as a process for making inferences about population parameters
WDHS Curriculum Map Probability and Statistics Time Interval/ Unit 1: Introduction to Statistics 1.1-1.3 2 weeks S-IC-1: Understand statistics as a process for making inferences about population parameters
Elms, Hayes, Shelburne 1
 Elms, Hayes, Shelburne 1 Potential Influential Factors of Cognitive Decline and Engagement in Participants of Adult Day Services Lauren Elms, Cat Hayes, Will Shelburne Acknowledgements The authors would
Elms, Hayes, Shelburne 1 Potential Influential Factors of Cognitive Decline and Engagement in Participants of Adult Day Services Lauren Elms, Cat Hayes, Will Shelburne Acknowledgements The authors would
MEMORANDUM. Michele DeHart. David A. Benner. DATE: December 9, NWPPC Mainstem Amendment Analysis Review
 FISH PASSAGE CENTER 2501 SW First Avenue, Suite 230, Portland, OR 97201-4752 Phone: (503) 230-4099 Fax: (503) 230-7559 http://www.fpc.org e-mail us at fpcstaff@fpc.org MEMORANDUM TO: FROM: Michele DeHart
FISH PASSAGE CENTER 2501 SW First Avenue, Suite 230, Portland, OR 97201-4752 Phone: (503) 230-4099 Fax: (503) 230-7559 http://www.fpc.org e-mail us at fpcstaff@fpc.org MEMORANDUM TO: FROM: Michele DeHart
Quantitative Methods in Computing Education Research (A brief overview tips and techniques)
 Quantitative Methods in Computing Education Research (A brief overview tips and techniques) Dr Judy Sheard Senior Lecturer Co-Director, Computing Education Research Group Monash University judy.sheard@monash.edu
Quantitative Methods in Computing Education Research (A brief overview tips and techniques) Dr Judy Sheard Senior Lecturer Co-Director, Computing Education Research Group Monash University judy.sheard@monash.edu
Any inbreeding will have similar effect, but slower. Overall, inbreeding modifies H-W by a factor F, the inbreeding coefficient.
 Effect of finite population. Two major effects 1) inbreeding 2) genetic drift Inbreeding Does not change gene frequency; however, increases homozygotes. Consider a population where selfing is the only
Effect of finite population. Two major effects 1) inbreeding 2) genetic drift Inbreeding Does not change gene frequency; however, increases homozygotes. Consider a population where selfing is the only
CS2220 Introduction to Computational Biology
 CS2220 Introduction to Computational Biology WEEK 8: GENOME-WIDE ASSOCIATION STUDIES (GWAS) 1 Dr. Mengling FENG Institute for Infocomm Research Massachusetts Institute of Technology mfeng@mit.edu PLANS
CS2220 Introduction to Computational Biology WEEK 8: GENOME-WIDE ASSOCIATION STUDIES (GWAS) 1 Dr. Mengling FENG Institute for Infocomm Research Massachusetts Institute of Technology mfeng@mit.edu PLANS
Tutorial 3: MANOVA. Pekka Malo 30E00500 Quantitative Empirical Research Spring 2016
 Tutorial 3: Pekka Malo 30E00500 Quantitative Empirical Research Spring 2016 Step 1: Research design Adequacy of sample size Choice of dependent variables Choice of independent variables (treatment effects)
Tutorial 3: Pekka Malo 30E00500 Quantitative Empirical Research Spring 2016 Step 1: Research design Adequacy of sample size Choice of dependent variables Choice of independent variables (treatment effects)
Understandable Statistics
 Understandable Statistics correlated to the Advanced Placement Program Course Description for Statistics Prepared for Alabama CC2 6/2003 2003 Understandable Statistics 2003 correlated to the Advanced Placement
Understandable Statistics correlated to the Advanced Placement Program Course Description for Statistics Prepared for Alabama CC2 6/2003 2003 Understandable Statistics 2003 correlated to the Advanced Placement
Outlier Analysis. Lijun Zhang
 Outlier Analysis Lijun Zhang zlj@nju.edu.cn http://cs.nju.edu.cn/zlj Outline Introduction Extreme Value Analysis Probabilistic Models Clustering for Outlier Detection Distance-Based Outlier Detection Density-Based
Outlier Analysis Lijun Zhang zlj@nju.edu.cn http://cs.nju.edu.cn/zlj Outline Introduction Extreme Value Analysis Probabilistic Models Clustering for Outlier Detection Distance-Based Outlier Detection Density-Based
Oregon Hatchery Research Center. Research Plan 2 December 2015 NWFCC
 Oregon Hatchery Research Center Research Plan 2 December 2015 NWFCC OHRC David Noakes Professor Fisheries & Wildlife Department, OSU Director, OHRC david.noakes@oregonstate.edu OHRC GOAL 1: Understand
Oregon Hatchery Research Center Research Plan 2 December 2015 NWFCC OHRC David Noakes Professor Fisheries & Wildlife Department, OSU Director, OHRC david.noakes@oregonstate.edu OHRC GOAL 1: Understand
Evaluation of Full-Sib Families of Douglas-fir in a Nelder Design. ABSTRACT
 Evaluation of Full-Sib Families of Douglas-fir in a Nelder Design. Roy Stonecypher and Rex McCullough ABSTRACT Thirty Douglas-fir full-sib families and an unselected check were evaluated in a Nelder's
Evaluation of Full-Sib Families of Douglas-fir in a Nelder Design. Roy Stonecypher and Rex McCullough ABSTRACT Thirty Douglas-fir full-sib families and an unselected check were evaluated in a Nelder's
HERITABILITY AND ITS GENETIC WORTH FOR PLANT BREEDING
 HERITABILITY AND ITS GENETIC WORTH FOR PLANT BREEDING Author: Prasanta Kumar Majhi M. Sc. (Agri.), Junior Research Scholar, Department of Genetics and Plant Breeding, College of Agriculture, UAS, Dharwad,
HERITABILITY AND ITS GENETIC WORTH FOR PLANT BREEDING Author: Prasanta Kumar Majhi M. Sc. (Agri.), Junior Research Scholar, Department of Genetics and Plant Breeding, College of Agriculture, UAS, Dharwad,
SELECTED OBSERVATIONS OF CORALS AND SPONGES
 APPENDIX D SELECTED OBSERVATIONS OF CORALS AND SPONGES Appendix D maps depict the spatial distribution of selected observations of corals and sponges from visual surveys conducted by a number of agencies
APPENDIX D SELECTED OBSERVATIONS OF CORALS AND SPONGES Appendix D maps depict the spatial distribution of selected observations of corals and sponges from visual surveys conducted by a number of agencies
The SAGE Encyclopedia of Educational Research, Measurement, and Evaluation Multivariate Analysis of Variance
 The SAGE Encyclopedia of Educational Research, Measurement, Multivariate Analysis of Variance Contributors: David W. Stockburger Edited by: Bruce B. Frey Book Title: Chapter Title: "Multivariate Analysis
The SAGE Encyclopedia of Educational Research, Measurement, Multivariate Analysis of Variance Contributors: David W. Stockburger Edited by: Bruce B. Frey Book Title: Chapter Title: "Multivariate Analysis
Seventy percent of people who are candidates for allogeneic hematopoietic
 274 7th Biennial Symposium on Minorities, the Medically Underserved and Cancer Supplement to Cancer The National Marrow Donor Program Meeting the Needs of the Medically Underserved Dennis L. Confer, M.D.
274 7th Biennial Symposium on Minorities, the Medically Underserved and Cancer Supplement to Cancer The National Marrow Donor Program Meeting the Needs of the Medically Underserved Dennis L. Confer, M.D.
STATUS REPORT PINNIPED PREDATION AND HAZING AT BONNEVILLE DAM IN Robert Stansell, Sean Tackley, and Karrie Gibbons 4/6/07
 STATUS REPORT PINNIPED PREDATION AND HAZING AT BONNEVILLE DAM IN 27 Robert Stansell, Sean Tackley, and Karrie Gibbons 4/6/7 This report is the fifth of regular status reports on the pinniped predation
STATUS REPORT PINNIPED PREDATION AND HAZING AT BONNEVILLE DAM IN 27 Robert Stansell, Sean Tackley, and Karrie Gibbons 4/6/7 This report is the fifth of regular status reports on the pinniped predation
Nature Genetics: doi: /ng Supplementary Figure 1. Country distribution of GME samples and designation of geographical subregions.
 Supplementary Figure 1 Country distribution of GME samples and designation of geographical subregions. GME samples collected across 20 countries and territories from the GME. Pie size corresponds to the
Supplementary Figure 1 Country distribution of GME samples and designation of geographical subregions. GME samples collected across 20 countries and territories from the GME. Pie size corresponds to the
Applications. DSC 410/510 Multivariate Statistical Methods. Discriminating Two Groups. What is Discriminant Analysis
 DSC 4/5 Multivariate Statistical Methods Applications DSC 4/5 Multivariate Statistical Methods Discriminant Analysis Identify the group to which an object or case (e.g. person, firm, product) belongs:
DSC 4/5 Multivariate Statistical Methods Applications DSC 4/5 Multivariate Statistical Methods Discriminant Analysis Identify the group to which an object or case (e.g. person, firm, product) belongs:
Nature Neuroscience: doi: /nn Supplementary Figure 1
 Supplementary Figure 1 Illustration of the working of network-based SVM to confidently predict a new (and now confirmed) ASD gene. Gene CTNND2 s brain network neighborhood that enabled its prediction by
Supplementary Figure 1 Illustration of the working of network-based SVM to confidently predict a new (and now confirmed) ASD gene. Gene CTNND2 s brain network neighborhood that enabled its prediction by
Supplementary Figure 1. Nature Neuroscience: doi: /nn.4547
 Supplementary Figure 1 Characterization of the Microfetti mouse model. (a) Gating strategy for 8-color flow analysis of peripheral Ly-6C + monocytes from Microfetti mice 5-7 days after TAM treatment. Living
Supplementary Figure 1 Characterization of the Microfetti mouse model. (a) Gating strategy for 8-color flow analysis of peripheral Ly-6C + monocytes from Microfetti mice 5-7 days after TAM treatment. Living
HOST-PATHOGEN CO-EVOLUTION THROUGH HIV-1 WHOLE GENOME ANALYSIS
 HOST-PATHOGEN CO-EVOLUTION THROUGH HIV-1 WHOLE GENOME ANALYSIS Somda&a Sinha Indian Institute of Science, Education & Research Mohali, INDIA International Visiting Research Fellow, Peter Wall Institute
HOST-PATHOGEN CO-EVOLUTION THROUGH HIV-1 WHOLE GENOME ANALYSIS Somda&a Sinha Indian Institute of Science, Education & Research Mohali, INDIA International Visiting Research Fellow, Peter Wall Institute
AD (Leave blank) TITLE: Genomic Characterization of Brain Metastasis in Non-Small Cell Lung Cancer Patients
 AD (Leave blank) Award Number: W81XWH-12-1-0444 TITLE: Genomic Characterization of Brain Metastasis in Non-Small Cell Lung Cancer Patients PRINCIPAL INVESTIGATOR: Mark A. Watson, MD PhD CONTRACTING ORGANIZATION:
AD (Leave blank) Award Number: W81XWH-12-1-0444 TITLE: Genomic Characterization of Brain Metastasis in Non-Small Cell Lung Cancer Patients PRINCIPAL INVESTIGATOR: Mark A. Watson, MD PhD CONTRACTING ORGANIZATION:
Statistics is a broad mathematical discipline dealing with
 Statistical Primer for Cardiovascular Research Descriptive Statistics and Graphical Displays Martin G. Larson, SD Statistics is a broad mathematical discipline dealing with techniques for the collection,
Statistical Primer for Cardiovascular Research Descriptive Statistics and Graphical Displays Martin G. Larson, SD Statistics is a broad mathematical discipline dealing with techniques for the collection,
Identification of Tissue Independent Cancer Driver Genes
 Identification of Tissue Independent Cancer Driver Genes Alexandros Manolakos, Idoia Ochoa, Kartik Venkat Supervisor: Olivier Gevaert Abstract Identification of genomic patterns in tumors is an important
Identification of Tissue Independent Cancer Driver Genes Alexandros Manolakos, Idoia Ochoa, Kartik Venkat Supervisor: Olivier Gevaert Abstract Identification of genomic patterns in tumors is an important
Ch. 23 The Evolution of Populations
 Ch. 23 The Evolution of Populations 1 Essential question: Do populations evolve? 2 Mutation and Sexual reproduction produce genetic variation that makes evolution possible What is the smallest unit of
Ch. 23 The Evolution of Populations 1 Essential question: Do populations evolve? 2 Mutation and Sexual reproduction produce genetic variation that makes evolution possible What is the smallest unit of
DNA and morphometric diagnostics for apple and snowberry maggot flies
 FINAL REPORT Project Title: DURATION: 1 YEAR DNA and morphometric diagnostics for apple and snowberry maggot flies PI: Wee Yee Co-PI(2): Jeff Feder Organization: USDA-ARS Organization: University of Notre
FINAL REPORT Project Title: DURATION: 1 YEAR DNA and morphometric diagnostics for apple and snowberry maggot flies PI: Wee Yee Co-PI(2): Jeff Feder Organization: USDA-ARS Organization: University of Notre
Human population sub-structure and genetic association studies
 Human population sub-structure and genetic association studies Stephanie A. Santorico, Ph.D. Department of Mathematical & Statistical Sciences Stephanie.Santorico@ucdenver.edu Global Similarity Map from
Human population sub-structure and genetic association studies Stephanie A. Santorico, Ph.D. Department of Mathematical & Statistical Sciences Stephanie.Santorico@ucdenver.edu Global Similarity Map from
Sampling Problems in Estimating Small Mammal Population Size1
 Sampling Problems in Estimating Small Mammal Population Size1 George E. Menkens, Jr.2 and Stanley H. Anderson3 Abstract. -Estimates of population size are influenced by four sources of error: measurement,
Sampling Problems in Estimating Small Mammal Population Size1 George E. Menkens, Jr.2 and Stanley H. Anderson3 Abstract. -Estimates of population size are influenced by four sources of error: measurement,
Chapter 1: Exploring Data
 Chapter 1: Exploring Data Key Vocabulary:! individual! variable! frequency table! relative frequency table! distribution! pie chart! bar graph! two-way table! marginal distributions! conditional distributions!
Chapter 1: Exploring Data Key Vocabulary:! individual! variable! frequency table! relative frequency table! distribution! pie chart! bar graph! two-way table! marginal distributions! conditional distributions!
Unit 1 Exploring and Understanding Data
 Unit 1 Exploring and Understanding Data Area Principle Bar Chart Boxplot Conditional Distribution Dotplot Empirical Rule Five Number Summary Frequency Distribution Frequency Polygon Histogram Interquartile
Unit 1 Exploring and Understanding Data Area Principle Bar Chart Boxplot Conditional Distribution Dotplot Empirical Rule Five Number Summary Frequency Distribution Frequency Polygon Histogram Interquartile
White Paper Estimating Complex Phenotype Prevalence Using Predictive Models
 White Paper 23-12 Estimating Complex Phenotype Prevalence Using Predictive Models Authors: Nicholas A. Furlotte Aaron Kleinman Robin Smith David Hinds Created: September 25 th, 2015 September 25th, 2015
White Paper 23-12 Estimating Complex Phenotype Prevalence Using Predictive Models Authors: Nicholas A. Furlotte Aaron Kleinman Robin Smith David Hinds Created: September 25 th, 2015 September 25th, 2015
CareerCruising. MatchMaker Reliability and Validity Analysis
 CareerCruising MatchMaker Reliability and Validity Analysis 1 Career Cruising MatchMaker Analysis Reliability and Validity Analysis CECS contracted with Career Cruising to conduct psychometric analyses
CareerCruising MatchMaker Reliability and Validity Analysis 1 Career Cruising MatchMaker Analysis Reliability and Validity Analysis CECS contracted with Career Cruising to conduct psychometric analyses
GIZZARD SHAD FROM CAESAR CREEK LAKE, OHIO 1
 Copyright 985 Ohio Acad. Sci. TANAORHAMPHUS LONGIROSTRIS (ACANTHOCEPHALA) IN GIZZARD SHAD FROM CAESAR CREEK LAKE, OHIO JERRY H. HUBSCHMAN, Department of Biological Sciences, Wright State University, Dayton,
Copyright 985 Ohio Acad. Sci. TANAORHAMPHUS LONGIROSTRIS (ACANTHOCEPHALA) IN GIZZARD SHAD FROM CAESAR CREEK LAKE, OHIO JERRY H. HUBSCHMAN, Department of Biological Sciences, Wright State University, Dayton,
Introduction to Statistical Data Analysis I
 Introduction to Statistical Data Analysis I JULY 2011 Afsaneh Yazdani Preface What is Statistics? Preface What is Statistics? Science of: designing studies or experiments, collecting data Summarizing/modeling/analyzing
Introduction to Statistical Data Analysis I JULY 2011 Afsaneh Yazdani Preface What is Statistics? Preface What is Statistics? Science of: designing studies or experiments, collecting data Summarizing/modeling/analyzing
STATUS REPORT PINNIPED PREDATION AND HAZING AT BONNEVILLE DAM IN Robert Stansell, Sean Tackley, and Karrie Gibbons 3/9/07
 STATUS REPORT PINNIPED PREDATION AND HAZING AT BONNEVILLE DAM IN 27 Robert Stansell, Sean Tackley, and Karrie Gibbons 3/9/7 This report is the first of regular status reports on the pinniped predation
STATUS REPORT PINNIPED PREDATION AND HAZING AT BONNEVILLE DAM IN 27 Robert Stansell, Sean Tackley, and Karrie Gibbons 3/9/7 This report is the first of regular status reports on the pinniped predation
English summary Modus Via. Fine-tuning geographical offender profiling.
 English summary Modus Via. Fine-tuning geographical offender profiling. When an offender goes out to commit a crime, say a burglary, he needs a target location to commit that crime. The offender will most
English summary Modus Via. Fine-tuning geographical offender profiling. When an offender goes out to commit a crime, say a burglary, he needs a target location to commit that crime. The offender will most
Genetic diversity in the fungal pathogen Dothistroma septosporum. Angie Dale Masters of Science, NRES Supervisor: Kathy Lewis
 Genetic diversity in the fungal pathogen Dothistroma septosporum Angie Dale Masters of Science, NRES Supervisor: Kathy Lewis 1 Outline Introduction Research Questions Methods Results to date Implications
Genetic diversity in the fungal pathogen Dothistroma septosporum Angie Dale Masters of Science, NRES Supervisor: Kathy Lewis 1 Outline Introduction Research Questions Methods Results to date Implications
Small Group Presentations
 Admin Assignment 1 due next Tuesday at 3pm in the Psychology course centre. Matrix Quiz during the first hour of next lecture. Assignment 2 due 13 May at 10am. I will upload and distribute these at the
Admin Assignment 1 due next Tuesday at 3pm in the Psychology course centre. Matrix Quiz during the first hour of next lecture. Assignment 2 due 13 May at 10am. I will upload and distribute these at the
STATUS REPORT PINNIPED PREDATION AND HAZING AT BONNEVILLE DAM IN Robert Stansell, Sean Tackley, and Karrie Gibbons 3/23/07
 STATUS REPORT PINNIPED PREDATION AND HAZING AT BONNEVILLE DAM IN 7 Robert Stansell, Sean Tackley, and Karrie Gibbons 3/23/7 This report is the third of regular status reports on the pinniped predation
STATUS REPORT PINNIPED PREDATION AND HAZING AT BONNEVILLE DAM IN 7 Robert Stansell, Sean Tackley, and Karrie Gibbons 3/23/7 This report is the third of regular status reports on the pinniped predation
Dr Rick Tearle Senior Applications Specialist, EMEA Complete Genomics Complete Genomics, Inc.
 Dr Rick Tearle Senior Applications Specialist, EMEA Complete Genomics Topics Overview of Data Processing Pipeline Overview of Data Files 2 DNA Nano-Ball (DNB) Read Structure Genome : acgtacatgcattcacacatgcttagctatctctcgccag
Dr Rick Tearle Senior Applications Specialist, EMEA Complete Genomics Topics Overview of Data Processing Pipeline Overview of Data Files 2 DNA Nano-Ball (DNB) Read Structure Genome : acgtacatgcattcacacatgcttagctatctctcgccag
OREGON SAGE GROUSE FRAMEWORK
 OREGON SAGE GROUSE FRAMEWORK Protecting sage grouse habitat from human disturbance 1. Existing Conditions Oregon has effectively protected Greater Sage Grouse core habitat from most human disturbance.
OREGON SAGE GROUSE FRAMEWORK Protecting sage grouse habitat from human disturbance 1. Existing Conditions Oregon has effectively protected Greater Sage Grouse core habitat from most human disturbance.
LTA Analysis of HapMap Genotype Data
 LTA Analysis of HapMap Genotype Data Introduction. This supplement to Global variation in copy number in the human genome, by Redon et al., describes the details of the LTA analysis used to screen HapMap
LTA Analysis of HapMap Genotype Data Introduction. This supplement to Global variation in copy number in the human genome, by Redon et al., describes the details of the LTA analysis used to screen HapMap
Abstract. Optimization strategy of Copy Number Variant calling using Multiplicom solutions APPLICATION NOTE. Introduction
 Optimization strategy of Copy Number Variant calling using Multiplicom solutions Michael Vyverman, PhD; Laura Standaert, PhD and Wouter Bossuyt, PhD Abstract Copy number variations (CNVs) represent a significant
Optimization strategy of Copy Number Variant calling using Multiplicom solutions Michael Vyverman, PhD; Laura Standaert, PhD and Wouter Bossuyt, PhD Abstract Copy number variations (CNVs) represent a significant
Chapter 7: Descriptive Statistics
 Chapter Overview Chapter 7 provides an introduction to basic strategies for describing groups statistically. Statistical concepts around normal distributions are discussed. The statistical procedures of
Chapter Overview Chapter 7 provides an introduction to basic strategies for describing groups statistically. Statistical concepts around normal distributions are discussed. The statistical procedures of
Lecture 1 An introduction to statistics in Ichthyology and Fisheries Science
 Lecture 1 An introduction to statistics in Ichthyology and Fisheries Science What is statistics and why do we need it? Statistics attempts to make inferences about unknown values that are common to a population
Lecture 1 An introduction to statistics in Ichthyology and Fisheries Science What is statistics and why do we need it? Statistics attempts to make inferences about unknown values that are common to a population
19th AWCBR (Australian Winter Conference on Brain Research), 2001, Queenstown, AU
 19th AWCBR (Australian Winter Conference on Brain Research), 21, Queenstown, AU https://www.otago.ac.nz/awcbr/proceedings/otago394614.pdf Do local modification rules allow efficient learning about distributed
19th AWCBR (Australian Winter Conference on Brain Research), 21, Queenstown, AU https://www.otago.ac.nz/awcbr/proceedings/otago394614.pdf Do local modification rules allow efficient learning about distributed
Nature Structural & Molecular Biology: doi: /nsmb.2419
 Supplementary Figure 1 Mapped sequence reads and nucleosome occupancies. (a) Distribution of sequencing reads on the mouse reference genome for chromosome 14 as an example. The number of reads in a 1 Mb
Supplementary Figure 1 Mapped sequence reads and nucleosome occupancies. (a) Distribution of sequencing reads on the mouse reference genome for chromosome 14 as an example. The number of reads in a 1 Mb
GENETIC DRIFT & EFFECTIVE POPULATION SIZE
 Instructor: Dr. Martha B. Reiskind AEC 450/550: Conservation Genetics Spring 2018 Lecture Notes for Lectures 3a & b: In the past students have expressed concern about the inbreeding coefficient, so please
Instructor: Dr. Martha B. Reiskind AEC 450/550: Conservation Genetics Spring 2018 Lecture Notes for Lectures 3a & b: In the past students have expressed concern about the inbreeding coefficient, so please
EVALUATION OF PINNIPED PREDATION ON ADULT SALMONIDS AND OTHER FISH IN THE BONNEVILLE DAM TAILRACE,
 EVALUATION OF PINNIPED PREDATION ON ADULT SALMONIDS AND OTHER FISH IN THE BONNEVILLE DAM TAILRACE, 2008-2010 Robert J. Stansell, Karrie M. Gibbons, and William T. Nagy U.S. Army Corps of Engineers Portland
EVALUATION OF PINNIPED PREDATION ON ADULT SALMONIDS AND OTHER FISH IN THE BONNEVILLE DAM TAILRACE, 2008-2010 Robert J. Stansell, Karrie M. Gibbons, and William T. Nagy U.S. Army Corps of Engineers Portland
The Inheritance of Complex Traits
 The Inheritance of Complex Traits Differences Among Siblings Is due to both Genetic and Environmental Factors VIDEO: Designer Babies Traits Controlled by Two or More Genes Many phenotypes are influenced
The Inheritance of Complex Traits Differences Among Siblings Is due to both Genetic and Environmental Factors VIDEO: Designer Babies Traits Controlled by Two or More Genes Many phenotypes are influenced
Population activity structure of excitatory and inhibitory neurons
 RESEARCH ARTICLE Population activity structure of excitatory and inhibitory neurons Sean R. Bittner 1,3, Ryan C. Williamson 3,5, Adam C. Snyder 1,3,7, Ashok Litwin-Kumar 8, Brent Doiron 3,6, Steven M.
RESEARCH ARTICLE Population activity structure of excitatory and inhibitory neurons Sean R. Bittner 1,3, Ryan C. Williamson 3,5, Adam C. Snyder 1,3,7, Ashok Litwin-Kumar 8, Brent Doiron 3,6, Steven M.
A Comparison of Collaborative Filtering Methods for Medication Reconciliation
 A Comparison of Collaborative Filtering Methods for Medication Reconciliation Huanian Zheng, Rema Padman, Daniel B. Neill The H. John Heinz III College, Carnegie Mellon University, Pittsburgh, PA, 15213,
A Comparison of Collaborative Filtering Methods for Medication Reconciliation Huanian Zheng, Rema Padman, Daniel B. Neill The H. John Heinz III College, Carnegie Mellon University, Pittsburgh, PA, 15213,
Effects of Disturbance on Populations of Marine Mammals
 DISTRIBUTION STATEMENT A. Approved for public release; distribution is unlimited. Effects of Disturbance on Populations of Marine Mammals Erica Fleishman, Ph.D., Principal Investigator John Muir Institute
DISTRIBUTION STATEMENT A. Approved for public release; distribution is unlimited. Effects of Disturbance on Populations of Marine Mammals Erica Fleishman, Ph.D., Principal Investigator John Muir Institute
Principal Components Factor Analysis in the Literature. Stage 1: Define the Research Problem
 Principal Components Factor Analysis in the Literature This problem is taken from the research article: Charles P. Flynn and Suzanne R. Kunkel, "Deprivation, Compensation, and Conceptions of an Afterlife."
Principal Components Factor Analysis in the Literature This problem is taken from the research article: Charles P. Flynn and Suzanne R. Kunkel, "Deprivation, Compensation, and Conceptions of an Afterlife."
Block-upscaling of transport in heterogeneous aquifers
 158 Calibration and Reliability in Groundwater Modelling: From Uncertainty to Decision Making (Proceedings of ModelCARE 2005, The Hague, The Netherlands, June 2005). IAHS Publ. 304, 2006. Block-upscaling
158 Calibration and Reliability in Groundwater Modelling: From Uncertainty to Decision Making (Proceedings of ModelCARE 2005, The Hague, The Netherlands, June 2005). IAHS Publ. 304, 2006. Block-upscaling
Inter-country mixing in HIV transmission clusters: A pan-european phylodynamic study
 Inter-country mixing in HIV transmission clusters: A pan-european phylodynamic study Prabhav Kalaghatgi Max Planck Institute for Informatics March 20th 2013 HIV epidemic (2009) Prabhav Kalaghatgi 2/18
Inter-country mixing in HIV transmission clusters: A pan-european phylodynamic study Prabhav Kalaghatgi Max Planck Institute for Informatics March 20th 2013 HIV epidemic (2009) Prabhav Kalaghatgi 2/18
STAT 503X Case Study 1: Restaurant Tipping
 STAT 503X Case Study 1: Restaurant Tipping 1 Description Food server s tips in restaurants may be influenced by many factors including the nature of the restaurant, size of the party, table locations in
STAT 503X Case Study 1: Restaurant Tipping 1 Description Food server s tips in restaurants may be influenced by many factors including the nature of the restaurant, size of the party, table locations in
How to interpret scientific & statistical graphs
 How to interpret scientific & statistical graphs Theresa A Scott, MS Department of Biostatistics theresa.scott@vanderbilt.edu http://biostat.mc.vanderbilt.edu/theresascott 1 A brief introduction Graphics:
How to interpret scientific & statistical graphs Theresa A Scott, MS Department of Biostatistics theresa.scott@vanderbilt.edu http://biostat.mc.vanderbilt.edu/theresascott 1 A brief introduction Graphics:
Principles of Experimental Design
 Principles of Experimental Design Bret Hanlon and Bret Larget Department of Statistics University of Wisconsin Madison November 15, 2011 Designing Experiments 1 / 31 Experimental Design Many interesting
Principles of Experimental Design Bret Hanlon and Bret Larget Department of Statistics University of Wisconsin Madison November 15, 2011 Designing Experiments 1 / 31 Experimental Design Many interesting
Survey research (Lecture 1) Summary & Conclusion. Lecture 10 Survey Research & Design in Psychology James Neill, 2015 Creative Commons Attribution 4.
 Summary & Conclusion Lecture 10 Survey Research & Design in Psychology James Neill, 2015 Creative Commons Attribution 4.0 Overview 1. Survey research 2. Survey design 3. Descriptives & graphing 4. Correlation
Summary & Conclusion Lecture 10 Survey Research & Design in Psychology James Neill, 2015 Creative Commons Attribution 4.0 Overview 1. Survey research 2. Survey design 3. Descriptives & graphing 4. Correlation
Survey research (Lecture 1)
 Summary & Conclusion Lecture 10 Survey Research & Design in Psychology James Neill, 2015 Creative Commons Attribution 4.0 Overview 1. Survey research 2. Survey design 3. Descriptives & graphing 4. Correlation
Summary & Conclusion Lecture 10 Survey Research & Design in Psychology James Neill, 2015 Creative Commons Attribution 4.0 Overview 1. Survey research 2. Survey design 3. Descriptives & graphing 4. Correlation
SCATTER PLOTS AND TREND LINES
 1 SCATTER PLOTS AND TREND LINES LEARNING MAP INFORMATION STANDARDS 8.SP.1 Construct and interpret scatter s for measurement to investigate patterns of between two quantities. Describe patterns such as
1 SCATTER PLOTS AND TREND LINES LEARNING MAP INFORMATION STANDARDS 8.SP.1 Construct and interpret scatter s for measurement to investigate patterns of between two quantities. Describe patterns such as
Principles of Experimental Design
 Principles of Experimental Design Bret Hanlon and Bret Larget Department of Statistics University of Wisconsin Madison November 15, 2011 Designing Experiments 1 / 31 Experimental Design Many interesting
Principles of Experimental Design Bret Hanlon and Bret Larget Department of Statistics University of Wisconsin Madison November 15, 2011 Designing Experiments 1 / 31 Experimental Design Many interesting
