Nature Neuroscience: doi: /nn Supplementary Figure 1
|
|
|
- Griffin Leo Lamb
- 5 years ago
- Views:
Transcription
1 Supplementary Figure 1 Illustration of the working of network-based SVM to confidently predict a new (and now confirmed) ASD gene. Gene CTNND2 s brain network neighborhood that enabled its prediction by the SVM. E1-E4 denote genes with various levels of evidence (high to low) for known association with ASD. CTNND2 gene ranks 16th in our prediction and has been recently discovered as a high-confidence gene associated with ASD in female-enriched multiplex (Turner et al. (2015) Nature 520(7545), 51 6), although our classifier did not see this gene during training (at any level of confidence). In our brain network, CTNND2 is tightly linked to highevidence genes SHANK2 and NRXN1 (E1), and several lower-evidence genes such as ATP2B2, DPP6 and SNAP25. In addition, it also shares common neighbors with E1 genes SHANK2, GRIN2B and NRXN1. The combination of local connectivity and global interaction pattern together enable the SVM to accurately predict CTNND2 as an ASD-related gene. Both DAWN (Liu et al. (2015) The Annals of Applied Statistics 9(3), ) (DAWN-2015 rank 6079/8488) and NETBAG+ (Chang et al. (2014) Nature Neuroscience 18(2), ), recent network-based methods focused on autism, fail to predict this gene s relevance for autism, mainly because previous genetic studies have not strongly linked this gene to ASD. This is evident from CTNND2 s high P-value (0.90) using TADA (De Rubeis et al. (2014) Nature 515(7526), ), a method to summarize prior ASD genetic data (all types of mutations) into gene P-values for ASD-risk. Our method, on the other hand, without any previous genetic evidence about this gene (this gene is also not in our gold standard), ranks it 16 th out of more than 25,000 genes in the genome.
2 Supplementary Figure 2 Robustness of ASD-gene predictions to changes in our gold-standard gene sets. To test the robustness of our predictions, we made predictions by subsampling the gold standard, each time using 4/5 of the negatives and positives. The rank based correlation coefficient between our original prediction and the 100 sets of predictions made on the resampled gold standard is 0.993, indicating that our genome-wide predictions are highly robust to noise in the gold standard.
3 Supplementary Figure 3 Permutation-based P-values for our genome-wide predictions. In order to improve the interpretability of our genome-wide ranking, we calculated a permutation-based P-value and a corresponding Q- value for each gene (see Methods). The plot shows the distribution of these P-values, with the red dashed line indicating mean frequency.
4 Supplementary Figure 4 Extended evaluation of autism-gene predictions on empirical data from an independent sequencing study. All evaluations below were performed and presented similar to those in Figure 2b and 2c. The resulting trends significant enrichment of proband LGD mutations and non-enrichment of sibling LGD mutations are consistent with those observed in Figure 2. (a) Rankbased enrichment test (without top-decile cutoffs) on data from all and unpublished families, for All and Novel ASD genes (Fig. 2, and Fig S4 b, c and d), showing trends similar to those presented in Figure 2. All families All genes is in Figure 2b, and the rest are here. All plots present the z-score quantifying the enrichment of the gene-set of interest towards the top of our genome-wide ranking of genes (see Methods). The three mutation gene-sets in each case are colored differently (labeled in the legend below) with the number of genes in parenthesis below. The P-values recorded at top of each bar were calculated using a permutation test described in Methods in detail. (b) Novel de novo LGDs from all families: This data set is derived from mutations recorded from all families published in 2014, but restricted to only genes that were not part of our training gold-standard (completely novel ASD genes). The total number of genes in the gene-set and the P-value from the binomial test are given in parenthesis just below. (c) All de novo LGDs from unpublished families: This data set is derived from mutations recorded only from SSC families that were unpublished in the 2012 studies and subsequently published in 2014 (all genes from completely unseen families). (d) Novel de novo LGDs from unpublished families: This data set is derived from mutations recorded from families that were unpublished in the 2012 studies and subsequently published in 2014, further restricted to only genes that were not part of our training gold-standard (completely novel ASD genes from unseen families).
5 Supplementary Figure 5 Histograms of prior genetic evidence scores and the distributions of top genes as predicted by DAWN-2015 and our method. (a) Distribution of 2014 TADA (De Rubeis et al. (2014) Nature 515(7526), ) P-values of 8488 genes (white), which is the input genetic evidence for DAWN-2015 (Liu et al. (2015) The Annals of Applied Statistics 9(3), ). Overlaid on top (blue) is the distribution of TADA P-values of DAWN s top 333 predicted genes. (b) Similar to (a), here overlaid on top (red) with the distribution of 2014 TADA P-values of a comparable set of our top 333 predicted genes.
6 Supplementary Figure 6 Comparison of our method to a previous method (DAWN). We compare our method to DAWN-2015 (Liu et al. (2015) The Annals of Applied Statistics 9(3), ) by evaluating the ASD gene ranking produced by the two methods in their ability to prioritize (a) novel LGD-targets, (b) novel protein-protein interaction (PPI) partners, and (c) ASD-associated genes identified by genome-wide association studies (GWAS). Precision is presented as fold-overrandom, measured as observed precision over expected baseline-precision (calculated as Precision/[P/(P+N)], where P is the number of positives and N is the number of negatives). For (a), gene targets of novel de novo LGDs observed in SSC probands and siblings were, respectively, used as positive and negative examples. For (b), novel PPI partners of potential ASD genes identified in a genomewide assay (Corominas et al. (2014) Nature Communications 5) were used as positives, and all other proteins tested in that assay were used as negatives. For (c), genes associated with autism as documented in the GWAS catalog were used as positives, while all other genes in the catalogue were used as negatives. P-values were calculated by Wilcoxon rank-sum test.
7 Supplementary Figure 7 Specificity of our genome-wide ranking to ASD. (a) Enrichment of ASD-gene ranking on genes associated with various neurological/brain diseases. To test the specificity of our genome-wide ranking to ASD, we tested the ranking of genes associated with five neurological diseases annotated in the OMIM database. We found no significant enrichment for unrelated diseases, indicating that our top-ranked genes are indeed specific for ASD, and distinct from genes associated with other neuronal disease. (b) Evaluation of ASD-gene ranking on genes linked to disorders closely related to ASD. We obtained genes associated with intellectual disability (ID; left; (Parikshak et al. (2013) Cell 155(5), )), schizophrenia (middle; (Fromer et al. (2014) Nature 506(7487), )), and developmental disorders (DDD; right; (TDDD Study (2015) Nature 519(7542), 223 8)) identified in large sequencing studies, removed the genes in our ASD positive gold standards, and test their distribution in our genome-wide ASD-gene ranking. We observe the expected significant overlap with ID and schizophrenia, while noting that our ranking prioritizes hundreds of additional genes not implicated in these disorders. The enrichment of genes in the DDD dataset is not surprising because the underlying cohort includes cases with ASD as well as several other disorders that have comorbidity with ASD (e.g. ID, heart development disorders).
8 Supplementary Figure 8 Top decile genes tend to be more constrained (have less common variation). (a) Boxplot of distribution of RVIS score (Petrovski et al. (2013) PLoS Genetics 9(8), e ) as a function of our autism gene ranking. (b) Histogram of fraction of constrained genes (out of 1,003) (Samocha et al. (2014) Nature Genetics 46(9), ), along autism gene ranking. Testing the two constrained sets against our predictions (see Methods) showed similar results both within the topdecile (using Fisher s exact test) or without any cutoff (using a rank-based permutation test): top-ranked genes do tend to be more constrained relative to all the other genes (RVIS, top-decile Wilcoxon test P < 2.2e-16, rank-based permutation test P = 1e-6; Constrained set, Fisher s exact test P = 1.36e-86, rank-based permutation test P = 1e-6).
9 Supplementary Figure 9 Spatiotemporal analysis is not biased by windows with dramatic changes. Spatiotemporal signature for each window was derived by controlling for both brain region and developmental stage. The permutation test used to identify association of each signature with ASD also controls for the number of genes in that signature. The plot shows that ASD-association of all signatures (indicated using the negative logarithm of the enrichment Q-value) as a function of the number of genes in each signature, demonstrating that there is no correlation between signature-size and ASD-association.
10 Supplementary Figure 10 Statistical analysis of ASD-gene clustering in the brain network. Results of a permutation test to evaluate the clustering of the ASD genes in a randomized brain network. Shared k-nearest-neighborbased analysis was repeated a 100 times, each time randomizing the k-nearest neighbors of each node in the network of top ASD genes. For a random set of nearly 30,000 gene pairs, the bulk of the clustering scores ranged between 0 and 0.3 based on random k neighbors, significantly lower than the cutoff of 0.9 used in our analysis of the real network. Our ASD functional modules thus are significantly and substantially more cohesive than random.
11 Supplementary Figure 11 Comparison of predicted ASD ranks of genes within autism-associated CNVs that have prior genetic and functional evidence. Boxplots show the distribution of ASD ranks of genes within the 8 ASD-associated CNVs that have different types of prior evidence: genetic (strong; red; n = 13), functional (weak; blue; n = 8), or none (grey; n = 166). *: P 0.05; ***: P
12 Supplementary Figure 12 Illustration of CNV diagram. Top decile genes within ASD-associated CNVs (blue circles), and known ASD genes (red circles) are linked in the brain network via intermediate genes (green circles). Statistically significant intermediate genes are identified using a permutation test against random genomic intervals. The biological processes enriched among significant intermediate genes (green bubble) that are shared with multiple ASD-associated CNVs are shown in Figure 6.
13 Supplementary Figure 13 Illustration of network visualization in the ASD web server. The interactive ASD web-interface enables biologists to explore their genes of interest (left) in the human brain-specific network (right). Users can easily explore the contribution of each of the different data types to any predicted brain-specific interaction (window at bottom right), including a summary of which data types contributed the most for example, co-expression and GSEA mirna targets in this case as well as evidence weights for each individual dataset. Users can click on any dataset to be redirected to the underlying data.
14 Supplementary Figure 14 New predictions based on training with updated gold standards correspond closely to the predictions used in this study. As an example of how we can regularly update the results in our web-server based on newly identified genes, we have made available a new set of predictions at by training on an updated version of the SFARI gene database that includes all the results from the 2014 study (Iossifov et al. (2014) Nature 515(7526), ). The scatter plot shows the original (v1) and new ranks (v2) of each gene in the genome. The dotted lines correspond to the top-decile of the predicted genes. The new predictions are overall quite consistent with the original one used throughout this study, having a correlation coefficient of 0.93 between the genomewide rankings with 83.3% of top decile genes consistent between v1 and v2. In addition, as our web server demonstrates, analyses results (including GO enrichments of predictions and neurodevelopmental analysis) also remain nearly identical.
15 Supplementary Figure 15 A potential use case: estimating the relevance of a high-resolution spatiotemporal window in the brain to autism. A biomedical researcher could use our predictions as a framework for analyzing their data from high- or low-throughput assays, allowing high-resolution study of autism genetics in the functional and physiological contexts of their interest. For example, a researcher who has generated gene expression or proteomics data from a new high-resolution spatiotemporal window in the brain (either human or in a model organism) can use our approach to assess the molecular relevance of that window to ASD. This approach identifying a characteristic gene signature from the samples and estimating its enrichment in our genome-wide ASD-gene ranking (similar to results presented in Figure 3) can provide results of increasing specificity with increasing spatiotemporal resolution in the available data. (S)he could then focus on the highly-ranked genes in her/his expression signature in combination with the specific functional contexts identified for these genes ( ASD-associated functional modules as presented in Figure 4) to generate hypotheses and design experiments to further characterize the genes expressed in the window in relation to autism.
38 Int'l Conf. Bioinformatics and Computational Biology BIOCOMP'16
 38 Int'l Conf. Bioinformatics and Computational Biology BIOCOMP'16 PGAR: ASD Candidate Gene Prioritization System Using Expression Patterns Steven Cogill and Liangjiang Wang Department of Genetics and
38 Int'l Conf. Bioinformatics and Computational Biology BIOCOMP'16 PGAR: ASD Candidate Gene Prioritization System Using Expression Patterns Steven Cogill and Liangjiang Wang Department of Genetics and
Nature Genetics: doi: /ng Supplementary Figure 1. PCA for ancestry in SNV data.
 Supplementary Figure 1 PCA for ancestry in SNV data. (a) EIGENSTRAT principal-component analysis (PCA) of SNV genotype data on all samples. (b) PCA of only proband SNV genotype data. (c) PCA of SNV genotype
Supplementary Figure 1 PCA for ancestry in SNV data. (a) EIGENSTRAT principal-component analysis (PCA) of SNV genotype data on all samples. (b) PCA of only proband SNV genotype data. (c) PCA of SNV genotype
Nature Methods: doi: /nmeth.3115
 Supplementary Figure 1 Analysis of DNA methylation in a cancer cohort based on Infinium 450K data. RnBeads was used to rediscover a clinically distinct subgroup of glioblastoma patients characterized by
Supplementary Figure 1 Analysis of DNA methylation in a cancer cohort based on Infinium 450K data. RnBeads was used to rediscover a clinically distinct subgroup of glioblastoma patients characterized by
Supplementary. properties of. network types. randomly sampled. subsets (75%
 Supplementary Information Gene co-expression network analysis reveals common system-level prognostic genes across cancer types properties of Supplementary Figure 1 The robustness and overlap of prognostic
Supplementary Information Gene co-expression network analysis reveals common system-level prognostic genes across cancer types properties of Supplementary Figure 1 The robustness and overlap of prognostic
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 Illustrative example of ptdt using height The expected value of a child s polygenic risk score (PRS) for a trait is the average of maternal and paternal PRS values. For example,
Supplementary Figure 1 Illustrative example of ptdt using height The expected value of a child s polygenic risk score (PRS) for a trait is the average of maternal and paternal PRS values. For example,
Strength of functional signature correlates with effect size in autism
 Ballouz and Gillis Genome Medicine (217) 9:64 DOI 1.1186/s1373-17-455-8 RESEARCH Open Access Strength of functional signature correlates with effect size in autism Sara Ballouz and Jesse Gillis * Abstract
Ballouz and Gillis Genome Medicine (217) 9:64 DOI 1.1186/s1373-17-455-8 RESEARCH Open Access Strength of functional signature correlates with effect size in autism Sara Ballouz and Jesse Gillis * Abstract
Nature Genetics: doi: /ng Supplementary Figure 1. SEER data for male and female cancer incidence from
 Supplementary Figure 1 SEER data for male and female cancer incidence from 1975 2013. (a,b) Incidence rates of oral cavity and pharynx cancer (a) and leukemia (b) are plotted, grouped by males (blue),
Supplementary Figure 1 SEER data for male and female cancer incidence from 1975 2013. (a,b) Incidence rates of oral cavity and pharynx cancer (a) and leukemia (b) are plotted, grouped by males (blue),
OncoPPi Portal A Cancer Protein Interaction Network to Inform Therapeutic Strategies
 OncoPPi Portal A Cancer Protein Interaction Network to Inform Therapeutic Strategies 2017 Contents Datasets... 2 Protein-protein interaction dataset... 2 Set of known PPIs... 3 Domain-domain interactions...
OncoPPi Portal A Cancer Protein Interaction Network to Inform Therapeutic Strategies 2017 Contents Datasets... 2 Protein-protein interaction dataset... 2 Set of known PPIs... 3 Domain-domain interactions...
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Experimental design and workflow utilized to generate the WMG Protein Atlas.
 Supplementary Figure 1 Experimental design and workflow utilized to generate the WMG Protein Atlas. (a) Illustration of the plant organs and nodule infection time points analyzed. (b) Proteomic workflow
Supplementary Figure 1 Experimental design and workflow utilized to generate the WMG Protein Atlas. (a) Illustration of the plant organs and nodule infection time points analyzed. (b) Proteomic workflow
Supplementary Figure 1. Metabolic landscape of cancer discovery pipeline. RNAseq raw counts data of cancer and healthy tissue samples were downloaded
 Supplementary Figure 1. Metabolic landscape of cancer discovery pipeline. RNAseq raw counts data of cancer and healthy tissue samples were downloaded from TCGA and differentially expressed metabolic genes
Supplementary Figure 1. Metabolic landscape of cancer discovery pipeline. RNAseq raw counts data of cancer and healthy tissue samples were downloaded from TCGA and differentially expressed metabolic genes
Expanded View Figures
 EMO Molecular Medicine Proteomic map of squamous cell carcinomas Hanibal ohnenberger et al Expanded View Figures Figure EV1. Technical reproducibility. Pearson s correlation analysis of normalised SILC
EMO Molecular Medicine Proteomic map of squamous cell carcinomas Hanibal ohnenberger et al Expanded View Figures Figure EV1. Technical reproducibility. Pearson s correlation analysis of normalised SILC
Nature Immunology: doi: /ni Supplementary Figure 1. Transcriptional program of the TE and MP CD8 + T cell subsets.
 Supplementary Figure 1 Transcriptional program of the TE and MP CD8 + T cell subsets. (a) Comparison of gene expression of TE and MP CD8 + T cell subsets by microarray. Genes that are 1.5-fold upregulated
Supplementary Figure 1 Transcriptional program of the TE and MP CD8 + T cell subsets. (a) Comparison of gene expression of TE and MP CD8 + T cell subsets by microarray. Genes that are 1.5-fold upregulated
Gene Ontology and Functional Enrichment. Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein
 Gene Ontology and Functional Enrichment Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein The parsimony principle: A quick review Find the tree that requires the fewest
Gene Ontology and Functional Enrichment Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein The parsimony principle: A quick review Find the tree that requires the fewest
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature13908 Supplementary Tables Supplementary Table 1: Families in this study (.xlsx) All families included in the study are listed. For each family, we show: the genders of the probands and
doi:10.1038/nature13908 Supplementary Tables Supplementary Table 1: Families in this study (.xlsx) All families included in the study are listed. For each family, we show: the genders of the probands and
Supplementary Materials for
 www.sciencetranslationalmedicine.org/cgi/content/full/7/283/283ra54/dc1 Supplementary Materials for Clonal status of actionable driver events and the timing of mutational processes in cancer evolution
www.sciencetranslationalmedicine.org/cgi/content/full/7/283/283ra54/dc1 Supplementary Materials for Clonal status of actionable driver events and the timing of mutational processes in cancer evolution
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10866 a b 1 2 3 4 5 6 7 Match No Match 1 2 3 4 5 6 7 Turcan et al. Supplementary Fig.1 Concepts mapping H3K27 targets in EF CBX8 targets in EF H3K27 targets in ES SUZ12 targets in ES
doi:10.1038/nature10866 a b 1 2 3 4 5 6 7 Match No Match 1 2 3 4 5 6 7 Turcan et al. Supplementary Fig.1 Concepts mapping H3K27 targets in EF CBX8 targets in EF H3K27 targets in ES SUZ12 targets in ES
SAPLING: A Tool for Gene Network Analysis focusing on Psychiatric Genetics
 SAPLING: A Tool for Gene Network Analysis focusing on Psychiatric Genetics sapling.cshl.edu Wim Verleyen, Ph.D. Gillis Lab Outline Motivation Disease-gene analysis Enrichment analysis Gene network analysis:
SAPLING: A Tool for Gene Network Analysis focusing on Psychiatric Genetics sapling.cshl.edu Wim Verleyen, Ph.D. Gillis Lab Outline Motivation Disease-gene analysis Enrichment analysis Gene network analysis:
Using large-scale human genetic variation to inform variant prioritization in neuropsychiatric disorders
 Using large-scale human genetic variation to inform variant prioritization in neuropsychiatric disorders Kaitlin E. Samocha Hurles lab, Wellcome Trust Sanger Institute ACGS Summer Scientific Meeting 27
Using large-scale human genetic variation to inform variant prioritization in neuropsychiatric disorders Kaitlin E. Samocha Hurles lab, Wellcome Trust Sanger Institute ACGS Summer Scientific Meeting 27
Autism Pathways Analysis: A Functional Framework and Clues for Further Investigation. Martha Herbert PhD MD Ya Wen PhD July 2016
 Autism Pathways Analysis: A Functional Framework and Clues for Further Investigation Martha Herbert PhD MD Ya Wen PhD July 2016 1 Report on pathway network analyses in autism, based on open-access paper
Autism Pathways Analysis: A Functional Framework and Clues for Further Investigation Martha Herbert PhD MD Ya Wen PhD July 2016 1 Report on pathway network analyses in autism, based on open-access paper
Supplementary Figure 1
 Supplementary Figure 1 An example of the gene-term-disease network automatically generated by Phenolyzer web server for 'autism'. The largest word represents the user s input term, Autism. The pink round
Supplementary Figure 1 An example of the gene-term-disease network automatically generated by Phenolyzer web server for 'autism'. The largest word represents the user s input term, Autism. The pink round
Figure S2. Distribution of acgh probes on all ten chromosomes of the RIL M0022
 96 APPENDIX B. Supporting Information for chapter 4 "changes in genome content generated via segregation of non-allelic homologs" Figure S1. Potential de novo CNV probes and sizes of apparently de novo
96 APPENDIX B. Supporting Information for chapter 4 "changes in genome content generated via segregation of non-allelic homologs" Figure S1. Potential de novo CNV probes and sizes of apparently de novo
Supplementary Figure 1: Features of IGLL5 Mutations in CLL: a) Representative IGV screenshot of first
 Supplementary Figure 1: Features of IGLL5 Mutations in CLL: a) Representative IGV screenshot of first intron IGLL5 mutation depicting biallelic mutations. Red arrows highlight the presence of out of phase
Supplementary Figure 1: Features of IGLL5 Mutations in CLL: a) Representative IGV screenshot of first intron IGLL5 mutation depicting biallelic mutations. Red arrows highlight the presence of out of phase
SFARI Gene 2.0 User Guide
 1 SFARI Gene 2.0 User Guide This document is designed to acquaint the new user with SFARI Gene release 4.0, an integrated resource for autism research, and to provide enough information to allow the user
1 SFARI Gene 2.0 User Guide This document is designed to acquaint the new user with SFARI Gene release 4.0, an integrated resource for autism research, and to provide enough information to allow the user
SUPPLEMENTAL INFORMATION
 SUPPLEMENTAL INFORMATION GO term analysis of differentially methylated SUMIs. GO term analysis of the 458 SUMIs with the largest differential methylation between human and chimp shows that they are more
SUPPLEMENTAL INFORMATION GO term analysis of differentially methylated SUMIs. GO term analysis of the 458 SUMIs with the largest differential methylation between human and chimp shows that they are more
Gene Ontology 2 Function/Pathway Enrichment. Biol4559 Thurs, April 12, 2018 Bill Pearson Pinn 6-057
 Gene Ontology 2 Function/Pathway Enrichment Biol4559 Thurs, April 12, 2018 Bill Pearson wrp@virginia.edu 4-2818 Pinn 6-057 Function/Pathway enrichment analysis do sets (subsets) of differentially expressed
Gene Ontology 2 Function/Pathway Enrichment Biol4559 Thurs, April 12, 2018 Bill Pearson wrp@virginia.edu 4-2818 Pinn 6-057 Function/Pathway enrichment analysis do sets (subsets) of differentially expressed
A quick review. The clustering problem: Hierarchical clustering algorithm: Many possible distance metrics K-mean clustering algorithm:
 The clustering problem: partition genes into distinct sets with high homogeneity and high separation Hierarchical clustering algorithm: 1. Assign each object to a separate cluster. 2. Regroup the pair
The clustering problem: partition genes into distinct sets with high homogeneity and high separation Hierarchical clustering algorithm: 1. Assign each object to a separate cluster. 2. Regroup the pair
Nature Neuroscience: doi: /nn Supplementary Figure 1. Behavioral training.
 Supplementary Figure 1 Behavioral training. a, Mazes used for behavioral training. Asterisks indicate reward location. Only some example mazes are shown (for example, right choice and not left choice maze
Supplementary Figure 1 Behavioral training. a, Mazes used for behavioral training. Asterisks indicate reward location. Only some example mazes are shown (for example, right choice and not left choice maze
Supplementary Materials Extracting a Cellular Hierarchy from High-dimensional Cytometry Data with SPADE
 Supplementary Materials Extracting a Cellular Hierarchy from High-dimensional Cytometry Data with SPADE Peng Qiu1,4, Erin F. Simonds2, Sean C. Bendall2, Kenneth D. Gibbs Jr.2, Robert V. Bruggner2, Michael
Supplementary Materials Extracting a Cellular Hierarchy from High-dimensional Cytometry Data with SPADE Peng Qiu1,4, Erin F. Simonds2, Sean C. Bendall2, Kenneth D. Gibbs Jr.2, Robert V. Bruggner2, Michael
Expanded View Figures
 Solip Park & Ben Lehner Epistasis is cancer type specific Molecular Systems Biology Expanded View Figures A B G C D E F H Figure EV1. Epistatic interactions detected in a pan-cancer analysis and saturation
Solip Park & Ben Lehner Epistasis is cancer type specific Molecular Systems Biology Expanded View Figures A B G C D E F H Figure EV1. Epistatic interactions detected in a pan-cancer analysis and saturation
ARTICLE RESEARCH. Macmillan Publishers Limited. All rights reserved
 Extended Data Figure 6 Annotation of drivers based on clinical characteristics and co-occurrence patterns. a, Putative drivers affecting greater than 10 patients were assessed for enrichment in IGHV mutated
Extended Data Figure 6 Annotation of drivers based on clinical characteristics and co-occurrence patterns. a, Putative drivers affecting greater than 10 patients were assessed for enrichment in IGHV mutated
Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63.
 Supplementary Figure Legends Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63. A. Screenshot of the UCSC genome browser from normalized RNAPII and RNA-seq ChIP-seq data
Supplementary Figure Legends Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63. A. Screenshot of the UCSC genome browser from normalized RNAPII and RNA-seq ChIP-seq data
Supporting Information
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 Supporting Information Variances and biases of absolute distributions were larger in the 2-line
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 Supporting Information Variances and biases of absolute distributions were larger in the 2-line
Journal: Nature Methods
 Journal: Nature Methods Article Title: Network-based stratification of tumor mutations Corresponding Author: Trey Ideker Supplementary Item Supplementary Figure 1 Supplementary Figure 2 Supplementary Figure
Journal: Nature Methods Article Title: Network-based stratification of tumor mutations Corresponding Author: Trey Ideker Supplementary Item Supplementary Figure 1 Supplementary Figure 2 Supplementary Figure
Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes.
 Supplementary Figure 1 Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes. (a,b) Values of coefficients associated with genomic features, separately
Supplementary Figure 1 Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes. (a,b) Values of coefficients associated with genomic features, separately
Case Studies on High Throughput Gene Expression Data Kun Huang, PhD Raghu Machiraju, PhD
 Case Studies on High Throughput Gene Expression Data Kun Huang, PhD Raghu Machiraju, PhD Department of Biomedical Informatics Department of Computer Science and Engineering The Ohio State University Review
Case Studies on High Throughput Gene Expression Data Kun Huang, PhD Raghu Machiraju, PhD Department of Biomedical Informatics Department of Computer Science and Engineering The Ohio State University Review
Visualization of Human Disorders and Genes Network
 Visualization of Human Disorders and Genes Network Bryan (Dung) Ta Dept. of Computer Science University of Maryland, College Park bryanta@cs.umd.edu I. INTRODUCTION Goh et al. [4] has provided an useful
Visualization of Human Disorders and Genes Network Bryan (Dung) Ta Dept. of Computer Science University of Maryland, College Park bryanta@cs.umd.edu I. INTRODUCTION Goh et al. [4] has provided an useful
Probability-Based Protein Identification for Post-Translational Modifications and Amino Acid Variants Using Peptide Mass Fingerprint Data
 Probability-Based Protein Identification for Post-Translational Modifications and Amino Acid Variants Using Peptide Mass Fingerprint Data Tong WW, McComb ME, Perlman DH, Huang H, O Connor PB, Costello
Probability-Based Protein Identification for Post-Translational Modifications and Amino Acid Variants Using Peptide Mass Fingerprint Data Tong WW, McComb ME, Perlman DH, Huang H, O Connor PB, Costello
Supplementary information for: Human micrornas co-silence in well-separated groups and have different essentialities
 Supplementary information for: Human micrornas co-silence in well-separated groups and have different essentialities Gábor Boross,2, Katalin Orosz,2 and Illés J. Farkas 2, Department of Biological Physics,
Supplementary information for: Human micrornas co-silence in well-separated groups and have different essentialities Gábor Boross,2, Katalin Orosz,2 and Illés J. Farkas 2, Department of Biological Physics,
SUPPLEMENTARY FIGURES
 SUPPLEMENTARY FIGURES Figure S1. Clinical significance of ZNF322A overexpression in Caucasian lung cancer patients. (A) Representative immunohistochemistry images of ZNF322A protein expression in tissue
SUPPLEMENTARY FIGURES Figure S1. Clinical significance of ZNF322A overexpression in Caucasian lung cancer patients. (A) Representative immunohistochemistry images of ZNF322A protein expression in tissue
What you should know before you collect data. BAE 815 (Fall 2017) Dr. Zifei Liu
 What you should know before you collect data BAE 815 (Fall 2017) Dr. Zifei Liu Zifeiliu@ksu.edu Types and levels of study Descriptive statistics Inferential statistics How to choose a statistical test
What you should know before you collect data BAE 815 (Fall 2017) Dr. Zifei Liu Zifeiliu@ksu.edu Types and levels of study Descriptive statistics Inferential statistics How to choose a statistical test
Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types.
 Supplementary Figure 1 Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types. (a) Pearson correlation heatmap among open chromatin profiles of different
Supplementary Figure 1 Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types. (a) Pearson correlation heatmap among open chromatin profiles of different
Supplement to SCnorm: robust normalization of single-cell RNA-seq data
 Supplement to SCnorm: robust normalization of single-cell RNA-seq data Supplementary Note 1: SCnorm does not require spike-ins, since we find that the performance of spike-ins in scrna-seq is often compromised,
Supplement to SCnorm: robust normalization of single-cell RNA-seq data Supplementary Note 1: SCnorm does not require spike-ins, since we find that the performance of spike-ins in scrna-seq is often compromised,
Detection of aneuploidy in a single cell using the Ion ReproSeq PGS View Kit
 APPLICATION NOTE Ion PGM System Detection of aneuploidy in a single cell using the Ion ReproSeq PGS View Kit Key findings The Ion PGM System, in concert with the Ion ReproSeq PGS View Kit and Ion Reporter
APPLICATION NOTE Ion PGM System Detection of aneuploidy in a single cell using the Ion ReproSeq PGS View Kit Key findings The Ion PGM System, in concert with the Ion ReproSeq PGS View Kit and Ion Reporter
Supplementary Figure 1 Information on transgenic mouse models and their recording and optogenetic equipment. (a) 108 (b-c) (d) (e) (f) (g)
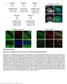 Supplementary Figure 1 Information on transgenic mouse models and their recording and optogenetic equipment. (a) In four mice, cre-dependent expression of the hyperpolarizing opsin Arch in pyramidal cells
Supplementary Figure 1 Information on transgenic mouse models and their recording and optogenetic equipment. (a) In four mice, cre-dependent expression of the hyperpolarizing opsin Arch in pyramidal cells
Supplementary Information. Supplementary Figures
 Supplementary Information Supplementary Figures.8 57 essential gene density 2 1.5 LTR insert frequency diversity DEL.5 DUP.5 INV.5 TRA 1 2 3 4 5 1 2 3 4 1 2 Supplementary Figure 1. Locations and minor
Supplementary Information Supplementary Figures.8 57 essential gene density 2 1.5 LTR insert frequency diversity DEL.5 DUP.5 INV.5 TRA 1 2 3 4 5 1 2 3 4 1 2 Supplementary Figure 1. Locations and minor
Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor suppressor genes
 Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor suppressor genes Kaifu Chen 1,2,3,4,5,10, Zhong Chen 6,10, Dayong Wu 6, Lili Zhang 7, Xueqiu Lin 1,2,8,
Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor suppressor genes Kaifu Chen 1,2,3,4,5,10, Zhong Chen 6,10, Dayong Wu 6, Lili Zhang 7, Xueqiu Lin 1,2,8,
Cancer Informatics Lecture
 Cancer Informatics Lecture Mayo-UIUC Computational Genomics Course June 22, 2018 Krishna Rani Kalari Ph.D. Associate Professor 2017 MFMER 3702274-1 Outline The Cancer Genome Atlas (TCGA) Genomic Data Commons
Cancer Informatics Lecture Mayo-UIUC Computational Genomics Course June 22, 2018 Krishna Rani Kalari Ph.D. Associate Professor 2017 MFMER 3702274-1 Outline The Cancer Genome Atlas (TCGA) Genomic Data Commons
Sum of Neurally Distinct Stimulus- and Task-Related Components.
 SUPPLEMENTARY MATERIAL for Cardoso et al. 22 The Neuroimaging Signal is a Linear Sum of Neurally Distinct Stimulus- and Task-Related Components. : Appendix: Homogeneous Linear ( Null ) and Modified Linear
SUPPLEMENTARY MATERIAL for Cardoso et al. 22 The Neuroimaging Signal is a Linear Sum of Neurally Distinct Stimulus- and Task-Related Components. : Appendix: Homogeneous Linear ( Null ) and Modified Linear
Understandable Statistics
 Understandable Statistics correlated to the Advanced Placement Program Course Description for Statistics Prepared for Alabama CC2 6/2003 2003 Understandable Statistics 2003 correlated to the Advanced Placement
Understandable Statistics correlated to the Advanced Placement Program Course Description for Statistics Prepared for Alabama CC2 6/2003 2003 Understandable Statistics 2003 correlated to the Advanced Placement
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Heatmap of GO terms for differentially expressed genes. The terms were hierarchically clustered using the GO term enrichment beta. Darker red, higher positive
Supplementary Figures Supplementary Figure 1. Heatmap of GO terms for differentially expressed genes. The terms were hierarchically clustered using the GO term enrichment beta. Darker red, higher positive
The Amazing Brain Webinar Series: Select Topics in Neuroscience and Child Development for the Clinician
 The Amazing Brain Webinar Series: Select Topics in Neuroscience and Child Development for the Clinician Part VII Recent Advances in the Genetics of Autism Spectrum Disorders Abha R. Gupta, MD, PhD Jointly
The Amazing Brain Webinar Series: Select Topics in Neuroscience and Child Development for the Clinician Part VII Recent Advances in the Genetics of Autism Spectrum Disorders Abha R. Gupta, MD, PhD Jointly
Theta sequences are essential for internally generated hippocampal firing fields.
 Theta sequences are essential for internally generated hippocampal firing fields. Yingxue Wang, Sandro Romani, Brian Lustig, Anthony Leonardo, Eva Pastalkova Supplementary Materials Supplementary Modeling
Theta sequences are essential for internally generated hippocampal firing fields. Yingxue Wang, Sandro Romani, Brian Lustig, Anthony Leonardo, Eva Pastalkova Supplementary Materials Supplementary Modeling
User Guide. Association analysis. Input
 User Guide TFEA.ChIP is a tool to estimate transcription factor enrichment in a set of differentially expressed genes using data from ChIP-Seq experiments performed in different tissues and conditions.
User Guide TFEA.ChIP is a tool to estimate transcription factor enrichment in a set of differentially expressed genes using data from ChIP-Seq experiments performed in different tissues and conditions.
Digitizing the Proteomes From Big Tissue Biobanks
 Digitizing the Proteomes From Big Tissue Biobanks Analyzing 24 Proteomes Per Day by Microflow SWATH Acquisition and Spectronaut Pulsar Analysis Jan Muntel 1, Nick Morrice 2, Roland M. Bruderer 1, Lukas
Digitizing the Proteomes From Big Tissue Biobanks Analyzing 24 Proteomes Per Day by Microflow SWATH Acquisition and Spectronaut Pulsar Analysis Jan Muntel 1, Nick Morrice 2, Roland M. Bruderer 1, Lukas
Lecture 20. Disease Genetics
 Lecture 20. Disease Genetics Michael Schatz April 12 2018 JHU 600.749: Applied Comparative Genomics Part 1: Pre-genome Era Sickle Cell Anaemia Sickle-cell anaemia (SCA) is an abnormality in the oxygen-carrying
Lecture 20. Disease Genetics Michael Schatz April 12 2018 JHU 600.749: Applied Comparative Genomics Part 1: Pre-genome Era Sickle Cell Anaemia Sickle-cell anaemia (SCA) is an abnormality in the oxygen-carrying
Comparison of Gene Set Analysis with Various Score Transformations to Test the Significance of Sets of Genes
 Comparison of Gene Set Analysis with Various Score Transformations to Test the Significance of Sets of Genes Ivan Arreola and Dr. David Han Department of Management of Science and Statistics, University
Comparison of Gene Set Analysis with Various Score Transformations to Test the Significance of Sets of Genes Ivan Arreola and Dr. David Han Department of Management of Science and Statistics, University
Two distinct mechanisms for experiencedependent
 SUPPLEMENTARY INFORMATION Articles https://doi.org/10.1038/s41593-018-0150-0 In the format provided by the authors and unedited. Two distinct mechanisms for experiencedependent homeostasis Michelle C.
SUPPLEMENTARY INFORMATION Articles https://doi.org/10.1038/s41593-018-0150-0 In the format provided by the authors and unedited. Two distinct mechanisms for experiencedependent homeostasis Michelle C.
STATISTICS AND RESEARCH DESIGN
 Statistics 1 STATISTICS AND RESEARCH DESIGN These are subjects that are frequently confused. Both subjects often evoke student anxiety and avoidance. To further complicate matters, both areas appear have
Statistics 1 STATISTICS AND RESEARCH DESIGN These are subjects that are frequently confused. Both subjects often evoke student anxiety and avoidance. To further complicate matters, both areas appear have
A Practical Guide to Integrative Genomics by RNA-seq and ChIP-seq Analysis
 A Practical Guide to Integrative Genomics by RNA-seq and ChIP-seq Analysis Jian Xu, Ph.D. Children s Research Institute, UTSW Introduction Outline Overview of genomic and next-gen sequencing technologies
A Practical Guide to Integrative Genomics by RNA-seq and ChIP-seq Analysis Jian Xu, Ph.D. Children s Research Institute, UTSW Introduction Outline Overview of genomic and next-gen sequencing technologies
SUPPLEMENTARY APPENDIX
 SUPPLEMENTARY APPENDIX 1) Supplemental Figure 1. Histopathologic Characteristics of the Tumors in the Discovery Cohort 2) Supplemental Figure 2. Incorporation of Normal Epidermal Melanocytic Signature
SUPPLEMENTARY APPENDIX 1) Supplemental Figure 1. Histopathologic Characteristics of the Tumors in the Discovery Cohort 2) Supplemental Figure 2. Incorporation of Normal Epidermal Melanocytic Signature
Recurrence rates provide evidence for sex-differential, familial genetic liability for autism spectrum disorders in multiplex families and twins
 Werling and Geschwind Molecular Autism (2015) 6:27 DOI 10.1186/s13229-015-0004-5 RESEARCH Open Access Recurrence rates provide evidence for sex-differential, familial genetic liability for autism spectrum
Werling and Geschwind Molecular Autism (2015) 6:27 DOI 10.1186/s13229-015-0004-5 RESEARCH Open Access Recurrence rates provide evidence for sex-differential, familial genetic liability for autism spectrum
IMPaLA tutorial.
 IMPaLA tutorial http://impala.molgen.mpg.de/ 1. Introduction IMPaLA is a web tool, developed for integrated pathway analysis of metabolomics data alongside gene expression or protein abundance data. It
IMPaLA tutorial http://impala.molgen.mpg.de/ 1. Introduction IMPaLA is a web tool, developed for integrated pathway analysis of metabolomics data alongside gene expression or protein abundance data. It
Nature Neuroscience: doi: /nn Supplementary Figure 1. Trial structure for go/no-go behavior
 Supplementary Figure 1 Trial structure for go/no-go behavior a, Overall timeline of experiments. Day 1: A1 mapping, injection of AAV1-SYN-GCAMP6s, cranial window and headpost implantation. Water restriction
Supplementary Figure 1 Trial structure for go/no-go behavior a, Overall timeline of experiments. Day 1: A1 mapping, injection of AAV1-SYN-GCAMP6s, cranial window and headpost implantation. Water restriction
Sequencing studies implicate inherited mutations in autism
 NEWS Sequencing studies implicate inherited mutations in autism BY EMILY SINGER 23 JANUARY 2013 1 / 5 Unusual inheritance: Researchers have found a relatively mild mutation in a gene linked to Cohen syndrome,
NEWS Sequencing studies implicate inherited mutations in autism BY EMILY SINGER 23 JANUARY 2013 1 / 5 Unusual inheritance: Researchers have found a relatively mild mutation in a gene linked to Cohen syndrome,
RNA-Seq Preparation Comparision Summary: Lexogen, Standard, NEB
 RNA-Seq Preparation Comparision Summary: Lexogen, Standard, NEB CSF-NGS January 22, 214 Contents 1 Introduction 1 2 Experimental Details 1 3 Results And Discussion 1 3.1 ERCC spike ins............................................
RNA-Seq Preparation Comparision Summary: Lexogen, Standard, NEB CSF-NGS January 22, 214 Contents 1 Introduction 1 2 Experimental Details 1 3 Results And Discussion 1 3.1 ERCC spike ins............................................
CoINcIDE: A framework for discovery of patient subtypes across multiple datasets
 Planey and Gevaert Genome Medicine (2016) 8:27 DOI 10.1186/s13073-016-0281-4 METHOD CoINcIDE: A framework for discovery of patient subtypes across multiple datasets Catherine R. Planey and Olivier Gevaert
Planey and Gevaert Genome Medicine (2016) 8:27 DOI 10.1186/s13073-016-0281-4 METHOD CoINcIDE: A framework for discovery of patient subtypes across multiple datasets Catherine R. Planey and Olivier Gevaert
Supplementary Figure 1. ALVAC-protein vaccines and macaque immunization. (A) Maximum likelihood
 Supplementary Figure 1. ALVAC-protein vaccines and macaque immunization. (A) Maximum likelihood tree illustrating CRF01_AE gp120 protein sequence relationships between 107 Envs sampled in the RV144 trial
Supplementary Figure 1. ALVAC-protein vaccines and macaque immunization. (A) Maximum likelihood tree illustrating CRF01_AE gp120 protein sequence relationships between 107 Envs sampled in the RV144 trial
PBZ FT01_PBZ FT01_TZ FT01_NZ. interface zone (I) tumor zone (TZ) necrotic zone (NZ)
 Oncotarget, Supplementary Materials www.impactjournals.com/oncotarget/ SUPPLEMENTRY FLES ndividuals factor map (P) FT_ FT_ FT_ Dim (.%) Dim (.%) >% peripheral brain zone () around % interface zone () FT
Oncotarget, Supplementary Materials www.impactjournals.com/oncotarget/ SUPPLEMENTRY FLES ndividuals factor map (P) FT_ FT_ FT_ Dim (.%) Dim (.%) >% peripheral brain zone () around % interface zone () FT
SUPPLEMENTARY INFORMATION
 Supplementary text Collectively, we were able to detect ~14,000 expressed genes with RPKM (reads per kilobase per million) > 1 or ~16,000 with RPKM > 0.1 in at least one cell type from oocyte to the morula
Supplementary text Collectively, we were able to detect ~14,000 expressed genes with RPKM (reads per kilobase per million) > 1 or ~16,000 with RPKM > 0.1 in at least one cell type from oocyte to the morula
User s Manual Version 1.0
 User s Manual Version 1.0 #639 Longmian Avenue, Jiangning District, Nanjing,211198,P.R.China. http://tcoa.cpu.edu.cn/ Contact us at xiaosheng.wang@cpu.edu.cn for technical issue and questions Catalogue
User s Manual Version 1.0 #639 Longmian Avenue, Jiangning District, Nanjing,211198,P.R.China. http://tcoa.cpu.edu.cn/ Contact us at xiaosheng.wang@cpu.edu.cn for technical issue and questions Catalogue
Nature Neuroscience: doi: /nn Supplementary Figure 1. Missense damaging predictions as a function of allele frequency
 Supplementary Figure 1 Missense damaging predictions as a function of allele frequency Percentage of missense variants classified as damaging by eight different classifiers and a classifier consisting
Supplementary Figure 1 Missense damaging predictions as a function of allele frequency Percentage of missense variants classified as damaging by eight different classifiers and a classifier consisting
Application of Artificial Neural Networks in Classification of Autism Diagnosis Based on Gene Expression Signatures
 Application of Artificial Neural Networks in Classification of Autism Diagnosis Based on Gene Expression Signatures 1 2 3 4 5 Kathleen T Quach Department of Neuroscience University of California, San Diego
Application of Artificial Neural Networks in Classification of Autism Diagnosis Based on Gene Expression Signatures 1 2 3 4 5 Kathleen T Quach Department of Neuroscience University of California, San Diego
TADA: Analyzing De Novo, Transmission and Case-Control Sequencing Data
 TADA: Analyzing De Novo, Transmission and Case-Control Sequencing Data Each person inherits mutations from parents, some of which may predispose the person to certain diseases. Meanwhile, new mutations
TADA: Analyzing De Novo, Transmission and Case-Control Sequencing Data Each person inherits mutations from parents, some of which may predispose the person to certain diseases. Meanwhile, new mutations
EXPression ANalyzer and DisplayER
 EXPression ANalyzer and DisplayER Tom Hait Aviv Steiner Igor Ulitsky Chaim Linhart Amos Tanay Seagull Shavit Rani Elkon Adi Maron-Katz Dorit Sagir Eyal David Roded Sharan Israel Steinfeld Yossi Shiloh
EXPression ANalyzer and DisplayER Tom Hait Aviv Steiner Igor Ulitsky Chaim Linhart Amos Tanay Seagull Shavit Rani Elkon Adi Maron-Katz Dorit Sagir Eyal David Roded Sharan Israel Steinfeld Yossi Shiloh
Stepwise Knowledge Acquisition in a Fuzzy Knowledge Representation Framework
 Stepwise Knowledge Acquisition in a Fuzzy Knowledge Representation Framework Thomas E. Rothenfluh 1, Karl Bögl 2, and Klaus-Peter Adlassnig 2 1 Department of Psychology University of Zurich, Zürichbergstraße
Stepwise Knowledge Acquisition in a Fuzzy Knowledge Representation Framework Thomas E. Rothenfluh 1, Karl Bögl 2, and Klaus-Peter Adlassnig 2 1 Department of Psychology University of Zurich, Zürichbergstraße
Supplementary Figure 1. Nature Neuroscience: doi: /nn.4547
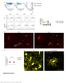 Supplementary Figure 1 Characterization of the Microfetti mouse model. (a) Gating strategy for 8-color flow analysis of peripheral Ly-6C + monocytes from Microfetti mice 5-7 days after TAM treatment. Living
Supplementary Figure 1 Characterization of the Microfetti mouse model. (a) Gating strategy for 8-color flow analysis of peripheral Ly-6C + monocytes from Microfetti mice 5-7 days after TAM treatment. Living
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Frequency of alternative-cassette-exon engagement with the ribosome is consistent across data from multiple human cell types and from mouse stem cells. Box plots showing AS frequency
Supplementary Figure 1 Frequency of alternative-cassette-exon engagement with the ribosome is consistent across data from multiple human cell types and from mouse stem cells. Box plots showing AS frequency
Integrated systems analysis reveals a molecular network underlying autism spectrum disorders
 Article Integrated systems analysis reveals a molecular network underlying autism spectrum disorders Jingjing Li 1,, Minyi Shi 1,, Zhihai Ma 1,, Shuchun Zhao 2, Ghia Euskirchen 1, Jennifer Ziskin 2, Alexander
Article Integrated systems analysis reveals a molecular network underlying autism spectrum disorders Jingjing Li 1,, Minyi Shi 1,, Zhihai Ma 1,, Shuchun Zhao 2, Ghia Euskirchen 1, Jennifer Ziskin 2, Alexander
SSM signature genes are highly expressed in residual scar tissues after preoperative radiotherapy of rectal cancer.
 Supplementary Figure 1 SSM signature genes are highly expressed in residual scar tissues after preoperative radiotherapy of rectal cancer. Scatter plots comparing expression profiles of matched pretreatment
Supplementary Figure 1 SSM signature genes are highly expressed in residual scar tissues after preoperative radiotherapy of rectal cancer. Scatter plots comparing expression profiles of matched pretreatment
Autism shares brain signature with schizophrenia, bipolar disorder
 NEWS Autism shares brain signature with schizophrenia, bipolar disorder BY NICHOLETTE ZELIADT 8 FEBRUARY 2018 1 / 5 Gene expression patterns in the brains of people with autism are similar to those of
NEWS Autism shares brain signature with schizophrenia, bipolar disorder BY NICHOLETTE ZELIADT 8 FEBRUARY 2018 1 / 5 Gene expression patterns in the brains of people with autism are similar to those of
Single SNP/Gene Analysis. Typical Results of GWAS Analysis (Single SNP Approach) Typical Results of GWAS Analysis (Single SNP Approach)
 High-Throughput Sequencing Course Gene-Set Analysis Biostatistics and Bioinformatics Summer 28 Section Introduction What is Gene Set Analysis? Many names for gene set analysis: Pathway analysis Gene set
High-Throughput Sequencing Course Gene-Set Analysis Biostatistics and Bioinformatics Summer 28 Section Introduction What is Gene Set Analysis? Many names for gene set analysis: Pathway analysis Gene set
R2: web-based genomics analysis and visualization platform
 R2: web-based genomics analysis and visualization platform Overview Jan Koster Department of Oncogenomics Academic Medical Center (AMC) UvA, the Netherlands jankoster@amc.uva.nl jankoster@amc.uva.nl 1
R2: web-based genomics analysis and visualization platform Overview Jan Koster Department of Oncogenomics Academic Medical Center (AMC) UvA, the Netherlands jankoster@amc.uva.nl jankoster@amc.uva.nl 1
a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation,
 Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays
 Supplementary Materials RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays Junhee Seok 1*, Weihong Xu 2, Ronald W. Davis 2, Wenzhong Xiao 2,3* 1 School of Electrical Engineering,
Supplementary Materials RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays Junhee Seok 1*, Weihong Xu 2, Ronald W. Davis 2, Wenzhong Xiao 2,3* 1 School of Electrical Engineering,
Sbarrato_Supplementary_Fig1
 Sbarrato_Supplementary_Fig1 Supplementary Figure 1. Translatome analysis of CLL patients based on IGVH mutational status ) Profile for the translatome of unmutated IGVH CLL versus mutated IGVH CLL pa9ents.
Sbarrato_Supplementary_Fig1 Supplementary Figure 1. Translatome analysis of CLL patients based on IGVH mutational status ) Profile for the translatome of unmutated IGVH CLL versus mutated IGVH CLL pa9ents.
Gene Expression Analysis Web Forum. Jonathan Gerstenhaber Field Application Specialist
 Gene Expression Analysis Web Forum Jonathan Gerstenhaber Field Application Specialist Our plan today: Import Preliminary Analysis Statistical Analysis Additional Analysis Downstream Analysis 2 Copyright
Gene Expression Analysis Web Forum Jonathan Gerstenhaber Field Application Specialist Our plan today: Import Preliminary Analysis Statistical Analysis Additional Analysis Downstream Analysis 2 Copyright
Variant Detection & Interpretation in a diagnostic context. Christian Gilissen
 Variant Detection & Interpretation in a diagnostic context Christian Gilissen c.gilissen@gen.umcn.nl 28-05-2013 So far Sequencing Johan den Dunnen Marja Jakobs Ewart de Bruijn Mapping Victor Guryev Variant
Variant Detection & Interpretation in a diagnostic context Christian Gilissen c.gilissen@gen.umcn.nl 28-05-2013 So far Sequencing Johan den Dunnen Marja Jakobs Ewart de Bruijn Mapping Victor Guryev Variant
TITLE: A Data-Driven Approach to Patient Risk Stratification for Acute Respiratory Distress Syndrome (ARDS)
 TITLE: A Data-Driven Approach to Patient Risk Stratification for Acute Respiratory Distress Syndrome (ARDS) AUTHORS: Tejas Prahlad INTRODUCTION Acute Respiratory Distress Syndrome (ARDS) is a condition
TITLE: A Data-Driven Approach to Patient Risk Stratification for Acute Respiratory Distress Syndrome (ARDS) AUTHORS: Tejas Prahlad INTRODUCTION Acute Respiratory Distress Syndrome (ARDS) is a condition
The Pretest! Pretest! Pretest! Assignment (Example 2)
 The Pretest! Pretest! Pretest! Assignment (Example 2) May 19, 2003 1 Statement of Purpose and Description of Pretest Procedure When one designs a Math 10 exam one hopes to measure whether a student s ability
The Pretest! Pretest! Pretest! Assignment (Example 2) May 19, 2003 1 Statement of Purpose and Description of Pretest Procedure When one designs a Math 10 exam one hopes to measure whether a student s ability
Computational Identification and Prediction of Tissue-Specific Alternative Splicing in H. Sapiens. Eric Van Nostrand CS229 Final Project
 Computational Identification and Prediction of Tissue-Specific Alternative Splicing in H. Sapiens. Eric Van Nostrand CS229 Final Project Introduction RNA splicing is a critical step in eukaryotic gene
Computational Identification and Prediction of Tissue-Specific Alternative Splicing in H. Sapiens. Eric Van Nostrand CS229 Final Project Introduction RNA splicing is a critical step in eukaryotic gene
Burning debate: What s the best way to nab real autism genes?
 OPINION, VIEWPOINT Burning debate: What s the best way to nab real autism genes? BY BRIAN O'ROAK 27 JUNE 2017 Over the past 10 years researchers have made tremendous progress in understanding the genetic
OPINION, VIEWPOINT Burning debate: What s the best way to nab real autism genes? BY BRIAN O'ROAK 27 JUNE 2017 Over the past 10 years researchers have made tremendous progress in understanding the genetic
DNA methylation signatures for 2016 WHO classification subtypes of diffuse gliomas
 Paul et al. Clinical Epigenetics (2017) 9:32 DOI 10.1186/s13148-017-0331-9 RESEARCH Open Access DNA methylation signatures for 2016 WHO classification subtypes of diffuse gliomas Yashna Paul, Baisakhi
Paul et al. Clinical Epigenetics (2017) 9:32 DOI 10.1186/s13148-017-0331-9 RESEARCH Open Access DNA methylation signatures for 2016 WHO classification subtypes of diffuse gliomas Yashna Paul, Baisakhi
Nature Neuroscience doi: /nn Supplementary Figure 1. Characterization of viral injections.
 Supplementary Figure 1 Characterization of viral injections. (a) Dorsal view of a mouse brain (dashed white outline) after receiving a large, unilateral thalamic injection (~100 nl); demonstrating that
Supplementary Figure 1 Characterization of viral injections. (a) Dorsal view of a mouse brain (dashed white outline) after receiving a large, unilateral thalamic injection (~100 nl); demonstrating that
Supplementary Materials
 1 Supplementary Materials Rotger et al. Table S1A: Demographic characteristics of study participants. VNP RP EC CP (n=6) (n=66) (n=9) (n=5) Male gender, n(%) 5 (83) 54 (82) 5 (56) 3 (60) White ethnicity,
1 Supplementary Materials Rotger et al. Table S1A: Demographic characteristics of study participants. VNP RP EC CP (n=6) (n=66) (n=9) (n=5) Male gender, n(%) 5 (83) 54 (82) 5 (56) 3 (60) White ethnicity,
Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation
 Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation Hang Yu, 1, 2, a) Wei Han, 1, 3, b) Wen Ma, 1, 2 1, 2, 3, c) and Klaus Schulten 1) Beckman
Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation Hang Yu, 1, 2, a) Wei Han, 1, 3, b) Wen Ma, 1, 2 1, 2, 3, c) and Klaus Schulten 1) Beckman
MATS: a Bayesian framework for flexible detection of differential alternative splicing from RNA-Seq data
 Nucleic Acids Research Advance Access published February 1, 2012 Nucleic Acids Research, 2012, 1 13 doi:10.1093/nar/gkr1291 MATS: a Bayesian framework for flexible detection of differential alternative
Nucleic Acids Research Advance Access published February 1, 2012 Nucleic Acids Research, 2012, 1 13 doi:10.1093/nar/gkr1291 MATS: a Bayesian framework for flexible detection of differential alternative
Supplementary Online Content
 Supplementary Online Content Viktorin A, Uher R, Kolevzon A, Reichenberg A, Levine SZ, Sandin S. Association of antidepressant medication use during pregnancy with intellectual disability in offspring.
Supplementary Online Content Viktorin A, Uher R, Kolevzon A, Reichenberg A, Levine SZ, Sandin S. Association of antidepressant medication use during pregnancy with intellectual disability in offspring.
7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans.
 Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
Supplementary Materials for
 www.sciencemag.org/content/355/6332/eaai8478/suppl/dc1 Supplementary Materials for Decoupling genetics, lineages, and microenvironment in IDH-mutant gliomas by single-cell RNA-seq Andrew S. Venteicher,
www.sciencemag.org/content/355/6332/eaai8478/suppl/dc1 Supplementary Materials for Decoupling genetics, lineages, and microenvironment in IDH-mutant gliomas by single-cell RNA-seq Andrew S. Venteicher,
Supplementary material
 Supplementary material S1. Event-related potentials Event-related potentials (ERPs) were calculated for stimuli for each visual field (mean of low, medium and high spatial frequency stimuli). For each
Supplementary material S1. Event-related potentials Event-related potentials (ERPs) were calculated for stimuli for each visual field (mean of low, medium and high spatial frequency stimuli). For each
