ncounter Data Analysis Guidelines for Copy Number Variation (CNV) Molecules That Count NanoString Technologies, Inc.
|
|
|
- Grant Watson
- 5 years ago
- Views:
Transcription
1 ncounter Data Analysis Guidelines for Copy Number Variation (CNV) NanoString Technologies, Inc. 530 Fairview Ave N Suite 2000 Seattle, Washington Tel: info@nanostring.com Molecules That Count Translational Research Gene Expression mirna Expression Copy Number Variation MAN-C
2 ncounter Data Analysis Guidelines for CNV FOR RESEARCH USE ONLY. Not for use in diagnostic procedures. Intellectual Property Rights This ncounter Analysis System manual and its contents are the property of NanoString Technologies, Inc. ( NanoString ), and is intended solely for the use of NanoString customers, for the purpose of operating the ncounter Analysis System. The ncounter Analysis System (including both its software and hardware components) and this User Guide and any other documentation provided to you by NanoString in connection therewith, are subject to patents, copyright, trade secret rights and other intellectual property rights owned by, or licensed to, NanoString. No part of the software or hardware, may be reproduced, transmitted, transcribed, stored in a retrieval system, or translated into other languages without the prior written consent of NanoString. Limited License Subject to the terms and conditions of the ncounter Analysis System contained in the product quotation, NanoString grants you a limited, nonexclusive, non-transferable, non-sublicensable, research use only license to use the proprietary ncounter Analysis System only in accordance with the manual and other written instructions provided by NanoString. Except as expressly set forth in the terms and conditions, no right or license, whether express, implied or statutory, is granted by NanoString under any intellectual property right owned by, or licensed to, NanoString by virtue of the supply of the proprietary ncounter Analysis System. Without limiting the foregoing, no right or license, whether express, implied or statutory, is granted by NanoString, to use the ncounter Analysis System with any third party product not supplied or licensed to you by NanoString, or recommended for use by NanoString in a manual or other written instruction provided by NanoString. Trademarks NanoString Technologies, NanoString, ncounter and Molecules That Count are registered trademarks or trademarks of NanoString Technologies, Inc., in the United States and/or other countries. All other trademarks and/or service marks not owned by NanoString that appear in this manual are the property of their respective owners. Copyright 2011 NanoString Technologies, Inc. All rights reserved. 2
3 NanoString Technologies PRODUCT MANUAL Preface... 4 Conventions Used... 4 Contact Information... 4 CHAPTER 1: Introduction ncounter Custom CNV Data Output... 5 Raw Data... 5 Code Classes... 6 Basic CNV Data Analysis Workflow... 6 Assay Quality Control Check... 6 Normalization to Invariant Probes... 8 Calculating Copy Number Estimates X and Y Chromosome Copy Number Calculation...11 Averaging Copy Number Estimates by Genomic Region...12 Generating Integer Copy Number Calls...12 Reference Sample Adjustments Adjusting the Copy Number Estimate Calculation Reference Documents and Support...15 Molecules That Count 3
4 ncounter Data Analysis Guidelines for CNV The following conventions are used throughout this manual and are described below for your reference: Special font formatting is used in this manual. Such formatting conventions are used in specific instances as described below: NanoString Technologies, Inc. 530 Fairview Ave N Suite 2000 Seattle, Washington USA Tel: NANO (6266) Fax: support@nanostring.com 4
5 1 Introduction The ncounter Custom Copy Number Variation (CNV) Assay utilizes NanoString s unique direct and multiplexed detection of nucleic acids in solution to generate estimates of copy number variation for hundreds of loci in a single reaction. Each NanoString Reporter and Capture Probe pair is complementary to ~100 nt of contiguous genomic DNA sequence at a user-specified locus. Genomic DNA is fragmented into small pieces ( bp) and denatured to produce single strands. The Custom CNV CodeSet is then hybridized to the fragmented, denatured DNA sample in a single multiplexed reaction (up to 800 genomic loci per CodeSet). Hybridized DNA-CodeSet complexes are purified by the fully automated ncounter Prep Station, and Reporters are counted by the ncounter Digital Analyzer. The results of an ncounter Custom CNV Assay experiment are compiled and displayed using the CNV Collector Tool software included with the assay kit. Detailed instructions for using this tool are in the CNV Collector Tool User Manual provided with the software. The following Data Analysis Guidelines for CNV are intended as a supplement to the ncounter CNV Collector Tool User Manual. They provide instructions and additional information for those who wish to do further QC and/or data manipulations with the data output from the CNV Collector Tool, such as: Modify the normalization method Apply additional assay quality control metrics Change reference samples within a single data set Use reference samples with known copy numbers that differ from 2 The basic output of the Custom CNV Assay is a spreadsheet containing the CodeSet probe identifiers, sample identifiers, and the digital counts recorded for each probe in each sample. This file, referred to as a Reporter Code Count (RCC) file, can be uploaded to the ncounter CNV Collector Tool for automated normalization and analysis. FIGURE 1.1: Report Code Count (RCC) file Molecules That Count Translational Research Gene Expression mirna Expression Copy Number Variation 5
6 ncounter Data Analysis Guidelines for CNV 1. POSITIVE ncounter Custom CNV CodeSets contains 6 positive dsdna control probes, each targeting a unique DNA sequence present in every assay. The concentrations of DNA target range from fm to 128 fm in the hybridization reaction. 2. NEGATIVE ncounter Custom CNV CodeSets contain 8 negative control probes, for which there is no DNA target present in the hybridization reaction. These probes monitor the nonspecific, or background, counts for every assay. 3. INVARIANT Each CodeSet contains a set of 10 probes (INV) designed to autosomal genomic regions predicted not to contain common CNVs. 4. RESTRICTION SITE Custom CNV CodeSets contain four control probes to monitor the efficiency of the DNA fragmentation and denaturation steps of the CNV Assay Protocol. Probes A and B are designed to a DNA sequence containing an AluI restriction site, and will return low count when the AluI fragmentation is working correctly. Probes C and D are designed to sequences that lack an AluI restriction site, and serve as controls for the presence of target DNA in the sample preparation step. When used according to the assay manual, these controls help identify problems in restriction enzyme fragmentation and denaturation steps of the assay. 5. ENDOGENOUS The Custom CNV Assay probes specified by the user, designed to specific regions of the genome. FIGURE 1.2: CNV Data Analysis Workflow Step 1 Assay QC È Step 2 Data Normalization È Step 3 Copy Number Estimation Relative to Reference Sample È Step 4 Averaging Probes per Region (if applicable) È Step 5 Integer Copy Number Prediction 6
7 NanoString Technologies PRODUCT MANUAL When used according to the assay manual, each Custom CNV Assay contains controls which monitor hybridization efficiency, sample DNA fragmentation denaturation, and sample DNA input amount. Before continuing with CNV analysis, first look at the Raw Data output file from the CNV Collector to gauge the performance of the assay. The positive control (POS) DNA targets are added in a linear titration to each codeset to generate a standard curve. The final concentration in the hybridization of each target (in fm) is indicated in parentheses next to the POS probe identifier. To check the linearity of the POS control standard curve, insert a new column in the Raw Data spreadsheet to the left of the first assay data column. Add the POS target concentrations to the first 6 rows of this new column. Then create a scatter plot of the target concentration vs. raw data for each target (Figure 1.3). FIGURE 1.3: Positive Controls In the scatter plot shown in Figure 1.3, the axes are represented in logarithmic scale and the linear regression (R 2 ) is shown in the inset. The R 2 value for the POS control probes should be > The Custom CNV Assay Kit comes with a set of four DNA controls that, when added to your genomic DNA sample prior to fragmentation, will monitor the efficiency of enzymatic digestion and heat denaturation. The DNA targets for probes labeled RESTRICTIONSITE+A and RESTRICTIONSITE+B contain an AluI restriction site such that, after complete digestion, the target site will be cleaved by the enzyme and low probe count will be observed. The DNA targets for probes labeled RESTRICTIONSITE-C and RESTRICTIONSITE-D do not contain AluI sites, and will generate probe counts even in the absence of fragmentation. These targets will serve as controls for proper addition of the control DNA to the sample and proper heat denaturation. If the DNA sample is not denatured prior to hybridization, you will observe low counts (generally < 200) for RESTRICTIONSITE-C and RESTRICTIONSITE-D probes. When the genomic DNA sample is completely digested with AluI enzyme and denatured, you should observe at least a 10-fold difference in counts between RESTRICTIONSITE+ (Probes A and B) and RESTRICTIONSITE- probes (Probes C and D): FIGURE 1.4: Restriction Fragmentation Controls Molecules That Count 7
8 ncounter Data Analysis Guidelines for CNV Each ncounter Custom CNV CodeSet contains probes designed to invariant regions of 10 autosomes. It is assumed that these 10 regions will represent 2 chromosomal copies in a vast majority of samples analyzed. Therefore, normalizing data to the counts obtained from these 10 invariant probes should correct for any differences in sample to sample genomic DNA input arising from pipetting error or inaccuracies in DNA quantitation. Normalization is performed automatically by the ncounter CNV Collector Tool, but can be done manually as follows: 1. To normalize data to the Invariant probes, calculate the average count value for the 10 INV probes (AVE INV) in the first lane (sample) as shown in Figure 1.5: FIGURE 1.5: Calculating the average count value. 8
9 NanoString Technologies PRODUCT MANUAL 2. Calculate a normalization factor for each assay by first calculating the mean of the average INV count values (mean AVE INV) across all lanes you wish to analyze (Figure 1.6): FIGURE 1.6: Calculating the mean of the average INV count values. To generate a normalization factor, divide the mean average value (mean AVE INV from Step 2) by the average count value (AVE INV from Step 1) for each lane: FIGURE 1.7: Generating a Normalization Factor Molecules That Count 9
10 ncounter Data Analysis Guidelines for CNV 3. Calculate INV-normalized counts for each probe. On a new sheet, generate normalized counts for each probe in the CodeSet by multiplying the RAW counts for each probe by the normalization factor for the lane as follows: FIGURE 1.8 For copy number analysis, it is not necessary to normalize the data for POS, NEG or Restriction Site controls. For simplicity, these probes can be left out of the INV normalization spreadsheet. The INV-normalization procedure should be carried out on the Invariant control and Endogenous code classes. CAUTION: Copy number data generated by the Custom CNV Assay can be negatively affected when DNA input amounts are too low, at which point sampling error can introduce unacceptably high levels of variation in the data. NanoString recommends a minimum of 100 counts for the average of the 10 Invariant control probes in the INV-normalized data set to ensure reliable copy number estimation. This is particularly important for the reference sample(s), since accurate copy number calculations depend upon high quality reference sample data. A poor quality reference sample will adversely affect copy number calls for all samples. A poor quality test sample will result in unreliable copy number call for that sample alone. In general for purified genomic DNA, free of RNA contamination, 100 INV-normalized counts will correspond to 100 ng genomic DNA. The basic data analysis strategy for determining copy numbers with the ncounter Custom CNV Assay is to calculate a copy number estimate for each probe relative to a reference sample (or samples). Each probe in the Custom CNV CodeSet is a unique sequence and bar code, and as a result small variations in probe efficiency can result in count variation between probes even when targeting genomic regions of equal copy number. However, this difference in counting efficiency will be constant for a given probe over all samples analyzed. Therefore, highly accurate copy number estimates can be generated by simply taking the ratio of counts from test samples to the counts of a fixed reference sample(s), and calculating copy numbers relative to that reference sample. The example analysis that follows will assume a copy number of 2 for all genomic loci being assayed in the reference sample. In a later section (Reference Sample Adjustments on page 10), we will consider alternative analysis strategies for genomic loci that differ from 2 copies in the reference sample(s). 10
11 NanoString Technologies PRODUCT MANUAL NOTE: If the copy number of your reference sample is not known, the analysis method presented here will only estimate the copy number relative to the unknown reference; an absolute integer copy number prediction will not be possible. First, start a new sheet for copy number calculations and select your reference sample. In the example below, the reference sample will be Sample 1 (Column D). Next, divide each test samples probe value (INV-normalized counts) by the corresponding probes in the reference samples INV-normalized counts. For autosomal chromosomes (1-22), multiply this quotient by 2 to account for the presence of two chromosomal copies in the diploid reference sample. (Hint: Use the $ shortcut command to hold the reference sample column constant in the formula.) FIGURE 1.9 The formula for determining copy number of the X and Y chromosomes must be adjusted depending on the gender of the reference sample. If the reference sample is female, the formulas should reflect two copies of the X chromosome and 0 copies of the Y chromosome as shown on Figure FIGURE 1.10 Molecules That Count 11
12 ncounter Data Analysis Guidelines for CNV In order to generate meaningful copy number estimates for the Y chromosome probe, it is best to use a reference sample that contains a Y chromosome (male). The formula for Y chromosome probes can then be adjusted to use this male reference to calculate copy numbers for only the Y chromosome probe(s). In the example below, the reference for the Y chromosome probe has been switched to Sample 2 (Column E): FIGURE 1.11 If your ncounter Custom CNV CodeSet contains multiple probes for a single genomic locus, it may be desirable to generate an average copy number estimate value based on all probes for that particular locus. This average value can then be used to generate an integer copy number assignment. If you are using a single probe per region, proceed to Generating Integer Copy Number Calls, below, for instructions on how to round the estimated copy number. To average probes for each locus, create a new spreadsheet. For each genomic region (locus), calculate the average of copy number estimate values using the =AVERAGE function as described in Normalization to Invariant Probes on page 5. To convert copy number estimates to integer copy number predictions, the copy number estimate values must be rounded. The simplest method is to round each estimated value to the nearest integer, using an IF function in Excel. In the following example we demonstrate an integer copy number +/- 0.4, although each investigator should determine the appropriate rounding criteria based on their own specific data analysis requirements. NOTE: Copy number estimates that are half-integer values (e.g., 1.5, 2.5, 3.5) will require further interpretation by the investigator to determine the integer copy number value. In the following example, such values are returned as the decimal copy number estimate value and are not rounded to the nearest integer. 12
13 NanoString Technologies PRODUCT MANUAL On a new sheet, enter an IF formula in the cell corresponding to the first probe and first sample following the example shown below. The values entered in the formula can be adjusted to alter the rounding criteria. For example, values between 0.8 and 1.2 (rather than 0.6 and 1.4) can be rounded to 1. Once the formula has been entered, click on the lower-right corner of the cell and drag the formula down (to apply to all Invariant and Endogenous probes), then across to apply to all samples. FIGURE 1.12 Here we have presented a simple three-step method for generating integer copy number calls from the ncounter CNV Assay raw data: normalization to INV probe counts, copy number estimation relative to a reference sample, and conversion to integer copy number calls. In the next section (Reference Sample Adjustments) we will consider situations when the copy number of the reference sample differs from 2, and when it is desirable to use multiple reference samples. Molecules That Count 13
14 ncounter Data Analysis Guidelines for CNV In some cases, the reference sample selected may contain genomic regions with a copy number of 0 (deletion), 1 (single-copy), or greater than 2 copies. Since it is not possible to calculate copy number estimates using a reference sample with 0 copies at a particular locus, it is necessary to change the reference to a sample that has counts registered for that specific region, and that has a known copy number. After the appropriate reference sample has been identified, it may be necessary to adjust the formula for calculating the copy number estimate. If the reference sample copy number is 1 for a particular genomic region, the formula for the corresponding probes can be altered by deleting the multiplication factor of 2. In the example below, the reference sample for probe1 (Chr1) is changed to Sample2 (Column E) and the copy number formula is adjusted for a copy number of 1 : NOTE: It may not be necessary to change the reference sample, only to alter the known copy number of the original reference sample following the same method outlined here. FIGURE
15 NanoString Technologies PRODUCT MANUAL If the copy number of the new reference is 3, the formula can be adjusted by changing the multiplication factor to 3 to generate the correct copy number. In the following example, the reference sample is Sample 1, but the formula is adjusted by adding a multiplication factor of 3 to reflect 3 copies of probe region1 in the reference sample: FIGURE 1.14 The adjusted formula can then be copied to the appropriate cells for each probe by clicking on the lower right corner of the highlighted cell and dragging over the cells you would like to change. For additional information on the ncounter CNV Assay and data analysis, please refer to these documents available for download at ncounter CNV Collector Tool User Manual ncounter Custom CNV Assay Manual Contacting Support For questions about the ncounter CNV Assay, ncounter CNV Collector Tool or data analysis, please contact: support@nanostring.com Phone: NANO or contact your NanoString Field Applications Scientist. Molecules That Count 15
16 ncounter Data Analysis Guidelines for CNV NanoString Technologies, Inc. CONTACT US SALES CONTACTS 530 Fairview Ave N Suite 2000 Seattle, Washington info@nanostring.com United States: us.sales@nanostring.com Tel: (888) Europe: europe.sales@nanostring.com Fax: (206) Japan: japan.sales@nanostring.com Other Regions: info@nanostring.com 2011 NanoString Technologies, Inc. All rights reserved. NanoString, NanoString Technologies, ncounter, and Molecules That Count are registered trademarks of NanoString Technologies, Inc., ( NanoString ) in the United States and/or other countries. All other trademarks and or service marks not owned by NanoString that appear in this document are the property of their respective owners. The manufacture, use and or sale of NanoString product(s) may be subject to one or more patents or pending patent applications owned by NanoString or licensed to NanoString from Life Technologies Corporation and other third parties. FOR RESEARCH USE ONLY. Not for use in diagnostic procedures. MAN-C
ncounter TM Analysis System
 ncounter TM Analysis System Molecules That Count TM www.nanostring.com Agenda NanoString Technologies History Introduction to the ncounter Analysis System CodeSet Design and Assay Principals System Performance
ncounter TM Analysis System Molecules That Count TM www.nanostring.com Agenda NanoString Technologies History Introduction to the ncounter Analysis System CodeSet Design and Assay Principals System Performance
Agile Product Lifecycle Management for Process
 Nutrition Surveillance Management User Guide Release 5.2.1 Part No. E13901-01 September 2008 Copyrights and Trademarks Copyright 1995, 2008, Oracle Corporation and/or its affiliates. All rights reserved.
Nutrition Surveillance Management User Guide Release 5.2.1 Part No. E13901-01 September 2008 Copyrights and Trademarks Copyright 1995, 2008, Oracle Corporation and/or its affiliates. All rights reserved.
Importance of Cell Stock Concentration for Accurate Target Cell Recovery
 TECHNICAL NOTE Importance of Cell Stock Concentration for Accurate Target Cell Recovery INTRODUCTION 10x Genomics Single Cell Protocols require suspensions of viable, single cells (Single Cell Protocols
TECHNICAL NOTE Importance of Cell Stock Concentration for Accurate Target Cell Recovery INTRODUCTION 10x Genomics Single Cell Protocols require suspensions of viable, single cells (Single Cell Protocols
Mutation Detection and CNV Analysis for Illumina Sequencing data from HaloPlex Target Enrichment Panels using NextGENe Software for Clinical Research
 Mutation Detection and CNV Analysis for Illumina Sequencing data from HaloPlex Target Enrichment Panels using NextGENe Software for Clinical Research Application Note Authors John McGuigan, Megan Manion,
Mutation Detection and CNV Analysis for Illumina Sequencing data from HaloPlex Target Enrichment Panels using NextGENe Software for Clinical Research Application Note Authors John McGuigan, Megan Manion,
Abstract. Optimization strategy of Copy Number Variant calling using Multiplicom solutions APPLICATION NOTE. Introduction
 Optimization strategy of Copy Number Variant calling using Multiplicom solutions Michael Vyverman, PhD; Laura Standaert, PhD and Wouter Bossuyt, PhD Abstract Copy number variations (CNVs) represent a significant
Optimization strategy of Copy Number Variant calling using Multiplicom solutions Michael Vyverman, PhD; Laura Standaert, PhD and Wouter Bossuyt, PhD Abstract Copy number variations (CNVs) represent a significant
AVENIO family of NGS oncology assays ctdna and Tumor Tissue Analysis Kits
 AVENIO family of NGS oncology assays ctdna and Tumor Tissue Analysis Kits Accelerating clinical research Next-generation sequencing (NGS) has the ability to interrogate many different genes and detect
AVENIO family of NGS oncology assays ctdna and Tumor Tissue Analysis Kits Accelerating clinical research Next-generation sequencing (NGS) has the ability to interrogate many different genes and detect
Advance Your Genomic Research Using Targeted Resequencing with SeqCap EZ Library
 Advance Your Genomic Research Using Targeted Resequencing with SeqCap EZ Library Marilou Wijdicks International Product Manager Research For Life Science Research Only. Not for Use in Diagnostic Procedures.
Advance Your Genomic Research Using Targeted Resequencing with SeqCap EZ Library Marilou Wijdicks International Product Manager Research For Life Science Research Only. Not for Use in Diagnostic Procedures.
A Penny for Your Thoughts: Scientific Measurements and Introduction to Excel
 A Penny for Your Thoughts: Scientific Measurements and Introduction to Excel Pre-lab Assignment: Introduction Reading: 1. Chapter sections 1.4 through 1.6 in your course text. 2. This lab handout. Questions:
A Penny for Your Thoughts: Scientific Measurements and Introduction to Excel Pre-lab Assignment: Introduction Reading: 1. Chapter sections 1.4 through 1.6 in your course text. 2. This lab handout. Questions:
Model 380B DNA Synthesizer Version 1.1
 Model 380B DNA Synthesizer Version 1.1 User s Manual Copyright 2001, Applied Biosystems. All rights reserved. For Research Use Only. Not for use in diagnostic procedures. APPLIED BIOSYSTEMS LIMITED WARRANTY
Model 380B DNA Synthesizer Version 1.1 User s Manual Copyright 2001, Applied Biosystems. All rights reserved. For Research Use Only. Not for use in diagnostic procedures. APPLIED BIOSYSTEMS LIMITED WARRANTY
To learn how to use the molar extinction coefficient in a real experiment, consider the following example.
 Week 3 - Phosphatase Data Analysis with Microsoft Excel Week 3 Learning Goals: To understand the effect of enzyme and substrate concentration on reaction rate To understand the concepts of Vmax, Km and
Week 3 - Phosphatase Data Analysis with Microsoft Excel Week 3 Learning Goals: To understand the effect of enzyme and substrate concentration on reaction rate To understand the concepts of Vmax, Km and
Integrated Analysis of Copy Number and Gene Expression
 Integrated Analysis of Copy Number and Gene Expression Nexus Copy Number provides user-friendly interface and functionalities to integrate copy number analysis with gene expression results for the purpose
Integrated Analysis of Copy Number and Gene Expression Nexus Copy Number provides user-friendly interface and functionalities to integrate copy number analysis with gene expression results for the purpose
Detection of aneuploidy in a single cell using the Ion ReproSeq PGS View Kit
 APPLICATION NOTE Ion PGM System Detection of aneuploidy in a single cell using the Ion ReproSeq PGS View Kit Key findings The Ion PGM System, in concert with the Ion ReproSeq PGS View Kit and Ion Reporter
APPLICATION NOTE Ion PGM System Detection of aneuploidy in a single cell using the Ion ReproSeq PGS View Kit Key findings The Ion PGM System, in concert with the Ion ReproSeq PGS View Kit and Ion Reporter
AND9020/D. Adaptive Feedback Cancellation 3 from ON Semiconductor APPLICATION NOTE INTRODUCTION
 Adaptive Feedback Cancellation 3 from ON Semiconductor APPLICATION NOTE INTRODUCTION This information note describes the feedback cancellation feature provided in ON Semiconductor s latest digital hearing
Adaptive Feedback Cancellation 3 from ON Semiconductor APPLICATION NOTE INTRODUCTION This information note describes the feedback cancellation feature provided in ON Semiconductor s latest digital hearing
WHO Prequalification of In Vitro Diagnostics PUBLIC REPORT. Product: Alere q HIV-1/2 Detect WHO reference number: PQDx
 WHO Prequalification of In Vitro Diagnostics PUBLIC REPORT Product: Alere q HIV-1/2 Detect WHO reference number: PQDx 0226-032-00 Alere q HIV-1/2 Detect with product codes 270110050, 270110010 and 270300001,
WHO Prequalification of In Vitro Diagnostics PUBLIC REPORT Product: Alere q HIV-1/2 Detect WHO reference number: PQDx 0226-032-00 Alere q HIV-1/2 Detect with product codes 270110050, 270110010 and 270300001,
Experiment 1: Scientific Measurements and Introduction to Excel
 Experiment 1: Scientific Measurements and Introduction to Excel Reading: Chapter 1 of your textbook and this lab handout. Learning Goals for Experiment 1: To use a scientific notebook as a primary record
Experiment 1: Scientific Measurements and Introduction to Excel Reading: Chapter 1 of your textbook and this lab handout. Learning Goals for Experiment 1: To use a scientific notebook as a primary record
Experiment 1: Scientific Measurements and Introduction to Excel
 Experiment 1: Scientific Measurements and Introduction to Excel Reading: Chapter 1 of your textbook and this lab handout. Learning Goals for Experiment 1: To use a scientific notebook as a primary record
Experiment 1: Scientific Measurements and Introduction to Excel Reading: Chapter 1 of your textbook and this lab handout. Learning Goals for Experiment 1: To use a scientific notebook as a primary record
Chapter 1: Managing workbooks
 Chapter 1: Managing workbooks Module A: Managing worksheets You use the Insert tab on the ribbon to insert new worksheets. True or False? Which of the following are options for moving or copying a worksheet?
Chapter 1: Managing workbooks Module A: Managing worksheets You use the Insert tab on the ribbon to insert new worksheets. True or False? Which of the following are options for moving or copying a worksheet?
Product Contents. 1 Specifications 1 Product Description. 2 Buffer Preparation... 3 Protocol. 3 Ordering Information 4
 INSTRUCTION MANUAL Quick-RNA Midiprep Kit Catalog No. R1056 Highlights 10 minute method for isolating RNA (up to 1 mg) from a wide range of cell types and tissue samples. Clean-Spin column technology allows
INSTRUCTION MANUAL Quick-RNA Midiprep Kit Catalog No. R1056 Highlights 10 minute method for isolating RNA (up to 1 mg) from a wide range of cell types and tissue samples. Clean-Spin column technology allows
Bioruptor DNA QC kit. Track the efficiency of your Bioruptor Pico. Cat. No. C
 Bioruptor DNA QC kit Track the efficiency of your Bioruptor Pico Cat. No. C40010002 DIAGENODE S.A. BELGIUM EUROPE LIEGE SCIENCE PARK Rue Bois Saint-Jean, 3 4102 Seraing - Belgium Tel: +32 4 364 20 50 Fax:
Bioruptor DNA QC kit Track the efficiency of your Bioruptor Pico Cat. No. C40010002 DIAGENODE S.A. BELGIUM EUROPE LIEGE SCIENCE PARK Rue Bois Saint-Jean, 3 4102 Seraing - Belgium Tel: +32 4 364 20 50 Fax:
SleepImage Website Instructions for Use
 SleepImage Website Instructions for Use Wellness Clinician Account Version 1 MyCardio SleepImage Website Copyright 2017 MyCardio. All rights reserved. Distributed by MyCardio LLC Issued Sept, 2017 Printed
SleepImage Website Instructions for Use Wellness Clinician Account Version 1 MyCardio SleepImage Website Copyright 2017 MyCardio. All rights reserved. Distributed by MyCardio LLC Issued Sept, 2017 Printed
A complete next-generation sequencing workfl ow for circulating cell-free DNA isolation and analysis
 APPLICATION NOTE Cell-Free DNA Isolation Kit A complete next-generation sequencing workfl ow for circulating cell-free DNA isolation and analysis Abstract Circulating cell-free DNA (cfdna) has been shown
APPLICATION NOTE Cell-Free DNA Isolation Kit A complete next-generation sequencing workfl ow for circulating cell-free DNA isolation and analysis Abstract Circulating cell-free DNA (cfdna) has been shown
Cloud Condensation Nuclei Counter (CCN) Module
 Particle Analysis and Display System (PADS): Cloud Condensation Nuclei Counter (CCN) Module Operator Manual DOC-0190 A-1 PADS 2.5.6, CCN Module 2.5.1 5710 Flatiron Parkway, Unit B Boulder, CO 80301 USA
Particle Analysis and Display System (PADS): Cloud Condensation Nuclei Counter (CCN) Module Operator Manual DOC-0190 A-1 PADS 2.5.6, CCN Module 2.5.1 5710 Flatiron Parkway, Unit B Boulder, CO 80301 USA
XCF TM COMPLETE Exosome and cfdna Isolation Kit (for Serum & Plasma)
 XCF TM COMPLETE Exosome and cfdna Isolation Kit (for Serum & Plasma) Cat# XCF100A-1 User Manual Store kit components at +4ºC and +25ºC Version 1 2/2/2017 A limited-use label license covers this product.
XCF TM COMPLETE Exosome and cfdna Isolation Kit (for Serum & Plasma) Cat# XCF100A-1 User Manual Store kit components at +4ºC and +25ºC Version 1 2/2/2017 A limited-use label license covers this product.
User Guide V: 3.0, August 2017
 User Guide V: 3.0, August 2017 a product of FAQ 3 General Information 1.1 System Overview 5 1.2 User Permissions 6 1.3 Points of Contact 7 1.4 Acronyms and Definitions 8 System Summary 2.1 System Configuration
User Guide V: 3.0, August 2017 a product of FAQ 3 General Information 1.1 System Overview 5 1.2 User Permissions 6 1.3 Points of Contact 7 1.4 Acronyms and Definitions 8 System Summary 2.1 System Configuration
LC/MS/MS SOLUTIONS FOR LIPIDOMICS. Biomarker and Omics Solutions FOR DISCOVERY AND TARGETED LIPIDOMICS
 LC/MS/MS SOLUTIONS FOR LIPIDOMICS Biomarker and Omics Solutions FOR DISCOVERY AND TARGETED LIPIDOMICS Lipids play a key role in many biological processes, such as the formation of cell membranes and signaling
LC/MS/MS SOLUTIONS FOR LIPIDOMICS Biomarker and Omics Solutions FOR DISCOVERY AND TARGETED LIPIDOMICS Lipids play a key role in many biological processes, such as the formation of cell membranes and signaling
Review Questions in Introductory Knowledge... 37
 Table of Contents Preface..... 17 About the Authors... 19 How This Book is Organized... 20 Who Should Buy This Book?... 20 Where to Find Answers to Review Questions and Exercises... 20 How to Report Errata...
Table of Contents Preface..... 17 About the Authors... 19 How This Book is Organized... 20 Who Should Buy This Book?... 20 Where to Find Answers to Review Questions and Exercises... 20 How to Report Errata...
To open a CMA file > Download and Save file Start CMA Open file from within CMA
 Example name Effect size Analysis type Level Tamiflu Symptom relief Mean difference (Hours to relief) Basic Basic Reference Cochrane Figure 4 Synopsis We have a series of studies that evaluated the effect
Example name Effect size Analysis type Level Tamiflu Symptom relief Mean difference (Hours to relief) Basic Basic Reference Cochrane Figure 4 Synopsis We have a series of studies that evaluated the effect
For in vitro Veterinary Diagnostics only. Kylt Rotavirus A. Real-Time RT-PCR Detection.
 For in vitro Veterinary Diagnostics only. Kylt Rotavirus A Real-Time RT-PCR Detection www.kylt.eu DIRECTION FOR USE Kylt Rotavirus A Real-Time RT-PCR Detection A. General Kylt Rotavirus A products are
For in vitro Veterinary Diagnostics only. Kylt Rotavirus A Real-Time RT-PCR Detection www.kylt.eu DIRECTION FOR USE Kylt Rotavirus A Real-Time RT-PCR Detection A. General Kylt Rotavirus A products are
Chapter 3 Software Packages to Install How to Set Up Python Eclipse How to Set Up Eclipse... 42
 Table of Contents Preface..... 21 About the Authors... 23 Acknowledgments... 24 How This Book is Organized... 24 Who Should Buy This Book?... 24 Where to Find Answers to Review Questions and Exercises...
Table of Contents Preface..... 21 About the Authors... 23 Acknowledgments... 24 How This Book is Organized... 24 Who Should Buy This Book?... 24 Where to Find Answers to Review Questions and Exercises...
Dr FuelCell Load Measurement Box
 1 (855) 251-0016 sales@fuelcellstore.com Dr FuelCell Load Measurement Box Instruction Manual www.fuelcellstore.com Instruction Manual for Dr FuelCell TM Load Measurement Box Version 1.0.1 November 2008
1 (855) 251-0016 sales@fuelcellstore.com Dr FuelCell Load Measurement Box Instruction Manual www.fuelcellstore.com Instruction Manual for Dr FuelCell TM Load Measurement Box Version 1.0.1 November 2008
E SERIES. Contents CALIBRATION PROCEDURE. Version 2.0
 CALIBRATION PROCEDURE E SERIES Version 2.0 Contents Introduction Document Scope... 2 Calibration Overview... 3 What Is Calibration?... 3 Why Calibrate?... 3 How Often Should You Calibrate?... 3 What Can
CALIBRATION PROCEDURE E SERIES Version 2.0 Contents Introduction Document Scope... 2 Calibration Overview... 3 What Is Calibration?... 3 Why Calibrate?... 3 How Often Should You Calibrate?... 3 What Can
EXOCET Exosome Quantitation Assay
 EXOCET Exosome Quantitation Assay EXOCET96A-1 User Manual Check Package Contents for Storage Temperatures Version 3 5/30/2017 A limited-use label license covers this product. By use of this product, you
EXOCET Exosome Quantitation Assay EXOCET96A-1 User Manual Check Package Contents for Storage Temperatures Version 3 5/30/2017 A limited-use label license covers this product. By use of this product, you
15.053x. OpenSolver (http://opensolver.org/)
 15.053x OpenSolver (http://opensolver.org/) 1 Table of Contents Introduction to OpenSolver slides 3-4 Example 1: Diet Problem, Set-Up slides 5-11 Example 1: Diet Problem, Dialog Box slides 12-17 Example
15.053x OpenSolver (http://opensolver.org/) 1 Table of Contents Introduction to OpenSolver slides 3-4 Example 1: Diet Problem, Set-Up slides 5-11 Example 1: Diet Problem, Dialog Box slides 12-17 Example
AUTISM SPECTRUM RATING SCALES (ASRS )
 AUTISM SPECTRUM RATING SCALES ( ) Scoring the for Individuals Who Do Not Speak or Speak Infrequently Sam Goldstein, Ph.D. & Jack A. Naglieri, Ph.D. Technical Report #1 This technical report was edited
AUTISM SPECTRUM RATING SCALES ( ) Scoring the for Individuals Who Do Not Speak or Speak Infrequently Sam Goldstein, Ph.D. & Jack A. Naglieri, Ph.D. Technical Report #1 This technical report was edited
Phosphate buffered saline (PBS) for washing the cells TE buffer (nuclease-free) ph 7.5 for use with the PrimePCR Reverse Transcription Control Assay
 Catalog # Description 172-5080 SingleShot Cell Lysis Kit, 100 x 50 µl reactions 172-5081 SingleShot Cell Lysis Kit, 500 x 50 µl reactions For research purposes only. Introduction The SingleShot Cell Lysis
Catalog # Description 172-5080 SingleShot Cell Lysis Kit, 100 x 50 µl reactions 172-5081 SingleShot Cell Lysis Kit, 500 x 50 µl reactions For research purposes only. Introduction The SingleShot Cell Lysis
QIAsymphony DSP Circulating DNA Kit
 QIAsymphony DSP Circulating DNA Kit February 2017 Performance Characteristics 937556 Sample to Insight Contents Performance Characteristics... 4 Basic performance... 4 Run precision... 6 Equivalent performance
QIAsymphony DSP Circulating DNA Kit February 2017 Performance Characteristics 937556 Sample to Insight Contents Performance Characteristics... 4 Basic performance... 4 Run precision... 6 Equivalent performance
Step-by-Step RECD Guide
 Precision Audiometric Instruments www.medrx-usa.com Step-by-Step RECD Guide The RECD task involves 4 steps: 1 - Complete Calibration of the Speakers and Probe Tube 2 - Measure an Ear Response 3 - Perform
Precision Audiometric Instruments www.medrx-usa.com Step-by-Step RECD Guide The RECD task involves 4 steps: 1 - Complete Calibration of the Speakers and Probe Tube 2 - Measure an Ear Response 3 - Perform
Unit 1: Introduction to the Operating System, Computer Systems, and Networks 1.1 Define terminology Prepare a list of terms with definitions
 AR Computer Applications I Correlated to Benchmark Microsoft Office 2010 (492490) Unit 1: Introduction to the Operating System, Computer Systems, and Networks 1.1 Define terminology 1.1.1 Prepare a list
AR Computer Applications I Correlated to Benchmark Microsoft Office 2010 (492490) Unit 1: Introduction to the Operating System, Computer Systems, and Networks 1.1 Define terminology 1.1.1 Prepare a list
Exo-Check Exosome Antibody Array (Neuro) Standard Kit
 Exo-Check Exosome Antibody Array (Neuro) Standard Kit Cat # EXORAY500A-4, EXORAY500A-8 User Manual Please see individual components for storage conditions Version 1 8/21/2018 A limited-use label license
Exo-Check Exosome Antibody Array (Neuro) Standard Kit Cat # EXORAY500A-4, EXORAY500A-8 User Manual Please see individual components for storage conditions Version 1 8/21/2018 A limited-use label license
Proteome Discoverer Version 1.3
 Xcalibur Proteome Discoverer Version 1.3 Installation Guide XCALI-97359 Revision A May 2011 2011 Thermo Fisher Scientific Inc. All rights reserved. Xcalibur is a registered trademark of Thermo Fisher Scientific
Xcalibur Proteome Discoverer Version 1.3 Installation Guide XCALI-97359 Revision A May 2011 2011 Thermo Fisher Scientific Inc. All rights reserved. Xcalibur is a registered trademark of Thermo Fisher Scientific
Urinary 8-Isoprostane ELISA kit
 Urinary 8-Isoprostane ELISA kit Catalog Number: 8isoU Store at -20 C. FOR RESEARCH USE ONLY V.020420 Introduction This competitive ELISA kit is for determination of free and glucronidated 8-isoprostane
Urinary 8-Isoprostane ELISA kit Catalog Number: 8isoU Store at -20 C. FOR RESEARCH USE ONLY V.020420 Introduction This competitive ELISA kit is for determination of free and glucronidated 8-isoprostane
Sleep Apnea Therapy Software Clinician Manual
 Sleep Apnea Therapy Software Clinician Manual Page ii Sleep Apnea Therapy Software Clinician Manual Notices Revised Notice Trademark Copyright Sleep Apnea Therapy Software Clinician Manual 103391 Rev A
Sleep Apnea Therapy Software Clinician Manual Page ii Sleep Apnea Therapy Software Clinician Manual Notices Revised Notice Trademark Copyright Sleep Apnea Therapy Software Clinician Manual 103391 Rev A
EnzChek Ultra Phytase Assay Kit
 EnzChek Ultra Phytase Assay Kit Catalog no. E33701 Table 1. Contents and storage information. Material Amount Storage Stability Amplex UltraRed reagent, MW = ~300 (Component A) Dimethylsulfoxide (DMSO),
EnzChek Ultra Phytase Assay Kit Catalog no. E33701 Table 1. Contents and storage information. Material Amount Storage Stability Amplex UltraRed reagent, MW = ~300 (Component A) Dimethylsulfoxide (DMSO),
Charts Worksheet using Excel Obesity Can a New Drug Help?
 Worksheet using Excel 2000 Obesity Can a New Drug Help? Introduction Obesity is known to be a major health risk. The data here arise from a study which aimed to investigate whether or not a new drug, used
Worksheet using Excel 2000 Obesity Can a New Drug Help? Introduction Obesity is known to be a major health risk. The data here arise from a study which aimed to investigate whether or not a new drug, used
Q: How do I get the protein concentration in mg/ml from the standard curve if the X-axis is in units of µg.
 Photometry Frequently Asked Questions Q: How do I get the protein concentration in mg/ml from the standard curve if the X-axis is in units of µg. Protein standard curves are traditionally presented as
Photometry Frequently Asked Questions Q: How do I get the protein concentration in mg/ml from the standard curve if the X-axis is in units of µg. Protein standard curves are traditionally presented as
Kofax VRS. Installation Guide
 Kofax VRS Installation Guide 2013-06-27 1999-2013 Kofax, Inc., 15211 Laguna Canyon Road, Irvine, California 92618, U.S.A. All rights reserved. Use is subject to license terms. Third-party software is copyrighted
Kofax VRS Installation Guide 2013-06-27 1999-2013 Kofax, Inc., 15211 Laguna Canyon Road, Irvine, California 92618, U.S.A. All rights reserved. Use is subject to license terms. Third-party software is copyrighted
Norgen s HIV Proviral DNA PCR Kit was developed and validated to be used with the following PCR instruments: Qiagen Rotor-Gene Q BioRad T1000 Cycler
 3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: 866-667-4362 (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com HIV Proviral DNA PCR Kit Product# 33840 Product Insert Intended
3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: 866-667-4362 (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com HIV Proviral DNA PCR Kit Product# 33840 Product Insert Intended
Small RNA Sequencing. Project Workflow. Service Description. Sequencing Service Specification BGISEQ-500 SERVICE OVERVIEW SAMPLE PREPARATION
 BGISEQ-500 SERVICE OVERVIEW Small RNA Sequencing Service Description Small RNAs are a type of non-coding RNA (ncrna) molecules that are less than 200nt in length. They are often involved in gene silencing
BGISEQ-500 SERVICE OVERVIEW Small RNA Sequencing Service Description Small RNAs are a type of non-coding RNA (ncrna) molecules that are less than 200nt in length. They are often involved in gene silencing
Clay Tablet Connector for hybris. User Guide. Version 1.5.0
 Clay Tablet Connector for hybris User Guide Version 1.5.0 August 4, 2016 Copyright Copyright 2005-2016 Clay Tablet Technologies Inc. All rights reserved. All rights reserved. This document and its content
Clay Tablet Connector for hybris User Guide Version 1.5.0 August 4, 2016 Copyright Copyright 2005-2016 Clay Tablet Technologies Inc. All rights reserved. All rights reserved. This document and its content
OneTouch Reveal Web Application. User Manual for Healthcare Professionals Instructions for Use
 OneTouch Reveal Web Application User Manual for Healthcare Professionals Instructions for Use Contents 2 Contents Chapter 1: Introduction...4 Product Overview...4 Intended Use...4 System Requirements...
OneTouch Reveal Web Application User Manual for Healthcare Professionals Instructions for Use Contents 2 Contents Chapter 1: Introduction...4 Product Overview...4 Intended Use...4 System Requirements...
MolecularMD. One-Step qrt-pcr BCR-ABL Kit. Product Description and User Manual. For Quantitative RT-PCR Analysis of BCR-ABL. Contact Us.
 Contact Us If you have any questions for or comments about MolecularMD, please feel free to contact us. Email Customer Service CustomerService@MolecularMD.com Technical Support TechSupport@MolecularMD.com
Contact Us If you have any questions for or comments about MolecularMD, please feel free to contact us. Email Customer Service CustomerService@MolecularMD.com Technical Support TechSupport@MolecularMD.com
User Bulletin 7500 Fast Precision Plate Holder for MicroAmp Tube Strips
 User Bulletin 7500 Fast Precision Plate Holder for MicroAmp Tube Strips December 19, 2007 SUBJECT: Using MicroAmp Fast 8-Tube Strips (0.1-mL) on the 7500 Fast Precision Plate Holder for MicroAmp Tube Strips
User Bulletin 7500 Fast Precision Plate Holder for MicroAmp Tube Strips December 19, 2007 SUBJECT: Using MicroAmp Fast 8-Tube Strips (0.1-mL) on the 7500 Fast Precision Plate Holder for MicroAmp Tube Strips
For purification of viral DNA and RNA from a wide range of sample materials
 QIAamp virus kits For purification of viral DNA and RNA from a wide range of sample materials Automatable on QIAGEN s proven QIAamp Kits set the standard for purification of viral DNA and RNA. QIAamp virus
QIAamp virus kits For purification of viral DNA and RNA from a wide range of sample materials Automatable on QIAGEN s proven QIAamp Kits set the standard for purification of viral DNA and RNA. QIAamp virus
Prosigna BREAST CANCER PROGNOSTIC GENE SIGNATURE ASSAY
 Prosigna BREAST CANCER PROGNOSTIC GENE SIGNATURE ASSAY Methodology The test is based on the reported 50-gene classifier algorithm originally named PAM50 and is performed on the ncounter Dx Analysis System
Prosigna BREAST CANCER PROGNOSTIC GENE SIGNATURE ASSAY Methodology The test is based on the reported 50-gene classifier algorithm originally named PAM50 and is performed on the ncounter Dx Analysis System
Prosigna BREAST CANCER PROGNOSTIC GENE SIGNATURE ASSAY
 Prosigna BREAST CANCER PROGNOSTIC GENE SIGNATURE ASSAY GENE EXPRESSION PROFILING WITH PROSIGNA What is Prosigna? Prosigna Breast Cancer Prognostic Gene Signature Assay is an FDA-approved assay which provides
Prosigna BREAST CANCER PROGNOSTIC GENE SIGNATURE ASSAY GENE EXPRESSION PROFILING WITH PROSIGNA What is Prosigna? Prosigna Breast Cancer Prognostic Gene Signature Assay is an FDA-approved assay which provides
Thrive Hearing Control Application
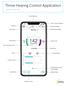 Thrive Hearing Control Application Apple Advanced Current Memory Thrive Virtual Assistant Settings User Guide Connection Status Edit Memory/Geotag Body Score Brain Score Thrive Wellness Score Heart Rate
Thrive Hearing Control Application Apple Advanced Current Memory Thrive Virtual Assistant Settings User Guide Connection Status Edit Memory/Geotag Body Score Brain Score Thrive Wellness Score Heart Rate
CNV PCA Search Tutorial
 CNV PCA Search Tutorial Release 8.1 Golden Helix, Inc. March 18, 2014 Contents 1. Data Preparation 2 A. Join Log Ratio Data with Phenotype Information.............................. 2 B. Activate only
CNV PCA Search Tutorial Release 8.1 Golden Helix, Inc. March 18, 2014 Contents 1. Data Preparation 2 A. Join Log Ratio Data with Phenotype Information.............................. 2 B. Activate only
Product Contents. 1 Specifications 1 Product Description. 2 Buffer Preparation... 3 Protocol. 3 Ordering Information 4 Related Products..
 INSTRUCTION MANUAL Quick-RNA MidiPrep Catalog No. R1056 Highlights 10 minute method for isolating RNA (up to 1 mg) from a wide range of cell types and tissue samples. Clean-Spin column technology allows
INSTRUCTION MANUAL Quick-RNA MidiPrep Catalog No. R1056 Highlights 10 minute method for isolating RNA (up to 1 mg) from a wide range of cell types and tissue samples. Clean-Spin column technology allows
AVENIO ctdna Analysis Kits The complete NGS liquid biopsy solution EMPOWER YOUR LAB
 Analysis Kits The complete NGS liquid biopsy solution EMPOWER YOUR LAB Analysis Kits Next-generation performance in liquid biopsies 2 Accelerating clinical research From liquid biopsy to next-generation
Analysis Kits The complete NGS liquid biopsy solution EMPOWER YOUR LAB Analysis Kits Next-generation performance in liquid biopsies 2 Accelerating clinical research From liquid biopsy to next-generation
SANAKO Lab 100 STS USER GUIDE
 SANAKO Lab 100 STS USER GUIDE Copyright 2008 SANAKO Corporation. All rights reserved. Microsoft is a registered trademark. Microsoft Windows 2000 and Windows XP are trademarks of Microsoft Corporation.
SANAKO Lab 100 STS USER GUIDE Copyright 2008 SANAKO Corporation. All rights reserved. Microsoft is a registered trademark. Microsoft Windows 2000 and Windows XP are trademarks of Microsoft Corporation.
The Essen Climate Evaluation Schema EssenCES
 The Essen Climate Evaluation Schema EssenCES Norbert Schalast Matthew Tonkin (Eds.) A Manual and More EssenCES The Essen Climate Evaluation Schema EssenCES A manual and more Norbert Schalast & Matthew
The Essen Climate Evaluation Schema EssenCES Norbert Schalast Matthew Tonkin (Eds.) A Manual and More EssenCES The Essen Climate Evaluation Schema EssenCES A manual and more Norbert Schalast & Matthew
Dementia Direct Enhanced Service
 Vision 3 Dementia Direct Enhanced Service England Outcomes Manager Copyright INPS Ltd 2015 The Bread Factory, 1A Broughton Street, Battersea, London, SW8 3QJ T: +44 (0) 207 501700 F:+44 (0) 207 5017100
Vision 3 Dementia Direct Enhanced Service England Outcomes Manager Copyright INPS Ltd 2015 The Bread Factory, 1A Broughton Street, Battersea, London, SW8 3QJ T: +44 (0) 207 501700 F:+44 (0) 207 5017100
THE OPTICAL SOCIETY ONLINE JOURNALS SINGLE SITE LICENSE AGREEMENT
 THE OPTICAL SOCIETY ONLINE JOURNALS SINGLE SITE LICENSE AGREEMENT BY THIS AGREEMENT between The Optical Society ( OSA ) and the named below ( Licensee ), OSA grants to Licensee access to the OSA online
THE OPTICAL SOCIETY ONLINE JOURNALS SINGLE SITE LICENSE AGREEMENT BY THIS AGREEMENT between The Optical Society ( OSA ) and the named below ( Licensee ), OSA grants to Licensee access to the OSA online
ExoQuick Exosome Isolation and RNA Purification Kits
 ExoQuick Exosome Isolation and RNA Purification Kits Cat # EQ806A-1, EQ806TC-1, EQ808A-1 User Manual Storage: Please see individual components Version 2 8/14/2018 A limited-use label license covers this
ExoQuick Exosome Isolation and RNA Purification Kits Cat # EQ806A-1, EQ806TC-1, EQ808A-1 User Manual Storage: Please see individual components Version 2 8/14/2018 A limited-use label license covers this
Norgen s HIV proviral DNA PCR Kit was developed and validated to be used with the following PCR instruments: Qiagen Rotor-Gene Q BioRad icycler
 3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com HIV Proviral DNA PCR Kit Product # 33840 Product Insert Background Information
3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: (905) 227-8848 Fax: (905) 227-1061 Email: techsupport@norgenbiotek.com HIV Proviral DNA PCR Kit Product # 33840 Product Insert Background Information
ExpressArt FFPE Clear RNAready kit
 Features and Example Results General problems with FFPE samples Formalin-fixation of tissues results in severe RNA fragmentation, as well as in RNA RNA, RNA-DNA and RNA protein cross-linking, which impairs
Features and Example Results General problems with FFPE samples Formalin-fixation of tissues results in severe RNA fragmentation, as well as in RNA RNA, RNA-DNA and RNA protein cross-linking, which impairs
Blood Urea Nitrogen Enzymatic Kit Manual Catalog #:
 BIOO LIFE SCIENCE PRODUCTS Blood Urea Nitrogen Enzymatic Kit Manual Catalog #: 5602-01 BIOO Scientific Corp. 2010 TABLE OF CONTENTS GENERAL INFORMATION... 1 Product Description... 1 Procedure Overview...
BIOO LIFE SCIENCE PRODUCTS Blood Urea Nitrogen Enzymatic Kit Manual Catalog #: 5602-01 BIOO Scientific Corp. 2010 TABLE OF CONTENTS GENERAL INFORMATION... 1 Product Description... 1 Procedure Overview...
ICD-10 Readiness (*9/14/15) By PracticeHwy.com, Inc.
 ICD-10 Readiness (*9/14/15) By PracticeHwy.com, Inc. Notice Information in this document is subject to change without notice and does not represent a commitment on the part of PracticeHwy.com, Inc. Companies,
ICD-10 Readiness (*9/14/15) By PracticeHwy.com, Inc. Notice Information in this document is subject to change without notice and does not represent a commitment on the part of PracticeHwy.com, Inc. Companies,
Cortex Gateway 2.0. Administrator Guide. September Document Version C
 Cortex Gateway 2.0 Administrator Guide September 2015 Document Version C Version C of the Cortex Gateway 2.0 Administrator Guide had been updated with editing changes. Contents Preface... 1 About Cortex
Cortex Gateway 2.0 Administrator Guide September 2015 Document Version C Version C of the Cortex Gateway 2.0 Administrator Guide had been updated with editing changes. Contents Preface... 1 About Cortex
IPA Advanced Training Course
 IPA Advanced Training Course October 2013 Academia sinica Gene (Kuan Wen Chen) IPA Certified Analyst Agenda I. Data Upload and How to Run a Core Analysis II. Functional Interpretation in IPA Hands-on Exercises
IPA Advanced Training Course October 2013 Academia sinica Gene (Kuan Wen Chen) IPA Certified Analyst Agenda I. Data Upload and How to Run a Core Analysis II. Functional Interpretation in IPA Hands-on Exercises
CaseBuilder - Quick Reference Guide
 ADP UNEMPLOYMENT COMPENSATION MANAGEMENT CaseBuilder - Quick Reference Guide After signing into CaseBuilder, the first screen the user will see is called the Dashboard. The user can then navigate to any
ADP UNEMPLOYMENT COMPENSATION MANAGEMENT CaseBuilder - Quick Reference Guide After signing into CaseBuilder, the first screen the user will see is called the Dashboard. The user can then navigate to any
Sanako Lab 100 STS USER GUIDE
 Sanako Lab 100 STS USER GUIDE Copyright 2002-2015 SANAKO Corporation. All rights reserved. Microsoft is a registered trademark. Microsoft Windows XP, Windows Vista and Windows 7 are trademarks of Microsoft
Sanako Lab 100 STS USER GUIDE Copyright 2002-2015 SANAKO Corporation. All rights reserved. Microsoft is a registered trademark. Microsoft Windows XP, Windows Vista and Windows 7 are trademarks of Microsoft
INSTRUCTION MANUAL. RNA Clean & Concentrator -5 Catalog Nos. R1015 & R1016. Highlights. Contents
 INSTRUCTION MANUAL Catalog Nos. R1015 & R1016 Highlights Quick (5 minute) method for cleaning and concentrating RNA. Ideal for purification of RNA from aqueous phase following an acid phenol extraction.
INSTRUCTION MANUAL Catalog Nos. R1015 & R1016 Highlights Quick (5 minute) method for cleaning and concentrating RNA. Ideal for purification of RNA from aqueous phase following an acid phenol extraction.
PENNDOT e-notification
 PENNDOT e-notification Bureau of Design Engineering Computing Management Division BRADD No. 027 August 30, 2010 Release of Version 3.1.5.0 PennDOT's Bridge Automated Design and Drafting Software (BRADD)
PENNDOT e-notification Bureau of Design Engineering Computing Management Division BRADD No. 027 August 30, 2010 Release of Version 3.1.5.0 PennDOT's Bridge Automated Design and Drafting Software (BRADD)
Appendix B. Nodulus Observer XT Instructional Guide. 1. Setting up your project p. 2. a. Observation p. 2. b. Subjects, behaviors and coding p.
 1 Appendix B Nodulus Observer XT Instructional Guide Sections: 1. Setting up your project p. 2 a. Observation p. 2 b. Subjects, behaviors and coding p. 3 c. Independent variables p. 4 2. Carry out an observation
1 Appendix B Nodulus Observer XT Instructional Guide Sections: 1. Setting up your project p. 2 a. Observation p. 2 b. Subjects, behaviors and coding p. 3 c. Independent variables p. 4 2. Carry out an observation
Diabetes Management Software V1.3 USER S MANUAL
 Diabetes Management Software V1.3 Manufacturer: BIONIME CORPORATION No. 100, Sec. 2, Daqing St., South Dist., Taichung City 40242, Taiwan http: //www.bionime.com E-mail: info@bionime.com Made in Taiwan
Diabetes Management Software V1.3 Manufacturer: BIONIME CORPORATION No. 100, Sec. 2, Daqing St., South Dist., Taichung City 40242, Taiwan http: //www.bionime.com E-mail: info@bionime.com Made in Taiwan
DNA-seq Bioinformatics Analysis: Copy Number Variation
 DNA-seq Bioinformatics Analysis: Copy Number Variation Elodie Girard elodie.girard@curie.fr U900 institut Curie, INSERM, Mines ParisTech, PSL Research University Paris, France NGS Applications 5C HiC DNA-seq
DNA-seq Bioinformatics Analysis: Copy Number Variation Elodie Girard elodie.girard@curie.fr U900 institut Curie, INSERM, Mines ParisTech, PSL Research University Paris, France NGS Applications 5C HiC DNA-seq
Thrive Hearing Control Application
 Thrive Hearing Control Application Android Advanced Current Memory Thrive Assistant Settings User Guide Connection Status Edit Memory/Geotag Body Score Brain Score Thrive Wellness Score Heart Rate Mute
Thrive Hearing Control Application Android Advanced Current Memory Thrive Assistant Settings User Guide Connection Status Edit Memory/Geotag Body Score Brain Score Thrive Wellness Score Heart Rate Mute
Content Part 2 Users manual... 4
 Content Part 2 Users manual... 4 Introduction. What is Kleos... 4 Case management... 5 Identity management... 9 Document management... 11 Document generation... 15 e-mail management... 15 Installation
Content Part 2 Users manual... 4 Introduction. What is Kleos... 4 Case management... 5 Identity management... 9 Document management... 11 Document generation... 15 e-mail management... 15 Installation
Workflow. Connecting the Pieces For Total Patient Care
 Workflow Connecting the Pieces For Total Patient Care Biocare provides a full line of IHC and molecular pathology products for cancer and infectious disease diagnosis. From a full range of equipment: including
Workflow Connecting the Pieces For Total Patient Care Biocare provides a full line of IHC and molecular pathology products for cancer and infectious disease diagnosis. From a full range of equipment: including
Cytogenetics 101: Clinical Research and Molecular Genetic Technologies
 Cytogenetics 101: Clinical Research and Molecular Genetic Technologies Topics for Today s Presentation 1 Classical vs Molecular Cytogenetics 2 What acgh? 3 What is FISH? 4 What is NGS? 5 How can these
Cytogenetics 101: Clinical Research and Molecular Genetic Technologies Topics for Today s Presentation 1 Classical vs Molecular Cytogenetics 2 What acgh? 3 What is FISH? 4 What is NGS? 5 How can these
Screening for novel oncology biomarker panels using both DNA and protein microarrays. John Anson, PhD VP Biomarker Discovery
 Screening for novel oncology biomarker panels using both DNA and protein microarrays John Anson, PhD VP Biomarker Discovery Outline of presentation Introduction to OGT and our approach to biomarker studies
Screening for novel oncology biomarker panels using both DNA and protein microarrays John Anson, PhD VP Biomarker Discovery Outline of presentation Introduction to OGT and our approach to biomarker studies
User Guide. Association analysis. Input
 User Guide TFEA.ChIP is a tool to estimate transcription factor enrichment in a set of differentially expressed genes using data from ChIP-Seq experiments performed in different tissues and conditions.
User Guide TFEA.ChIP is a tool to estimate transcription factor enrichment in a set of differentially expressed genes using data from ChIP-Seq experiments performed in different tissues and conditions.
ECDC HIV Modelling Tool User Manual version 1.0.0
 ECDC HIV Modelling Tool User Manual version 1 Copyright European Centre for Disease Prevention and Control, 2015 All rights reserved. No part of the contents of this document may be reproduced or transmitted
ECDC HIV Modelling Tool User Manual version 1 Copyright European Centre for Disease Prevention and Control, 2015 All rights reserved. No part of the contents of this document may be reproduced or transmitted
ExoQuick PLUS Exosome Purification Kit for Serum & Plasma
 ExoQuick PLUS Exosome Purification Kit for Serum & Plasma Cat# EQPL10A-1 User Manual Store kit at +4 0 C Version 2 2/22/2017 A limited-use label license covers this product. By use of this product, you
ExoQuick PLUS Exosome Purification Kit for Serum & Plasma Cat# EQPL10A-1 User Manual Store kit at +4 0 C Version 2 2/22/2017 A limited-use label license covers this product. By use of this product, you
Internal Validation Guide of Y-STR Systems for Forensic Laboratories Printed in USA. 11/12 Part# GE713
 REFERENCE MANUAL Internal Validation Guide of Y-STR Systems for Forensic Laboratories 11/12 Internal Validation Guide of Y-STR Systems for Forensic Laboratories PLEASE DISCARD PREVIOUS VERSIONS. All technical
REFERENCE MANUAL Internal Validation Guide of Y-STR Systems for Forensic Laboratories 11/12 Internal Validation Guide of Y-STR Systems for Forensic Laboratories PLEASE DISCARD PREVIOUS VERSIONS. All technical
ab Factor Xa Activity Assay Kit (Fluorometric)
 ab204711 Factor Xa Activity Assay Kit (Fluorometric) Instructions for Use For rapid, sensitive and accurate detection of Factor Xa activity. This product is for research use only and is not intended for
ab204711 Factor Xa Activity Assay Kit (Fluorometric) Instructions for Use For rapid, sensitive and accurate detection of Factor Xa activity. This product is for research use only and is not intended for
Implementation Guide for the DoseControl Dosimetry System
 1.0 SCOPE The GEX DoseControl System is intended for use as a single dosimetry system solution, specifically developed to satisfy the generic dosimetry needs of radiation sterilization application users
1.0 SCOPE The GEX DoseControl System is intended for use as a single dosimetry system solution, specifically developed to satisfy the generic dosimetry needs of radiation sterilization application users
Use Case 9: Coordinated Changes of Epigenomic Marks Across Tissue Types. Epigenome Informatics Workshop Bioinformatics Research Laboratory
 Use Case 9: Coordinated Changes of Epigenomic Marks Across Tissue Types Epigenome Informatics Workshop Bioinformatics Research Laboratory 1 Introduction Active or inactive states of transcription factor
Use Case 9: Coordinated Changes of Epigenomic Marks Across Tissue Types Epigenome Informatics Workshop Bioinformatics Research Laboratory 1 Introduction Active or inactive states of transcription factor
User Guide. Protein Clpper. Statistical scoring of protease cleavage sites. 1. Introduction Protein Clpper Analysis Procedure...
 User Guide Protein Clpper Statistical scoring of protease cleavage sites Content 1. Introduction... 2 2. Protein Clpper Analysis Procedure... 3 3. Input and Output Files... 9 4. Contact Information...
User Guide Protein Clpper Statistical scoring of protease cleavage sites Content 1. Introduction... 2 2. Protein Clpper Analysis Procedure... 3 3. Input and Output Files... 9 4. Contact Information...
User Instruction Guide
 User Instruction Guide Table of Contents Logging In and Logging Out of MMSx 1 Creating a TPN (Terminal Profile Number) 2 Single Merchant 2 From Navigation Bar 2 From Home Page Link 4 Multiple Merchants
User Instruction Guide Table of Contents Logging In and Logging Out of MMSx 1 Creating a TPN (Terminal Profile Number) 2 Single Merchant 2 From Navigation Bar 2 From Home Page Link 4 Multiple Merchants
For Research Use Only Ver
 INSTRUCTION MANUAL Quick-RNA Miniprep Kit Catalog Nos. R1054 & R1055 Highlights High-quality total RNA (including small RNAs) from a wide range of samples. You can opt to isolate small and large RNAs in
INSTRUCTION MANUAL Quick-RNA Miniprep Kit Catalog Nos. R1054 & R1055 Highlights High-quality total RNA (including small RNAs) from a wide range of samples. You can opt to isolate small and large RNAs in
For Research Use Only Ver
 INSTRUCTION MANUAL Quick-cfDNA/cfRNA Serum & Plasma Kit Catalog No. R1072 Highlights High-quality cell-free DNA and RNA are easily and robustly purified from up to 3 ml of plasma, serum, or other biological
INSTRUCTION MANUAL Quick-cfDNA/cfRNA Serum & Plasma Kit Catalog No. R1072 Highlights High-quality cell-free DNA and RNA are easily and robustly purified from up to 3 ml of plasma, serum, or other biological
Excel Solver. Table of Contents. Introduction to Excel Solver slides 3-4. Example 1: Diet Problem, Set-Up slides 5-11
 15.053 Excel Solver 1 Table of Contents Introduction to Excel Solver slides 3- : Diet Problem, Set-Up slides 5-11 : Diet Problem, Dialog Box slides 12-17 Example 2: Food Start-Up Problem slides 18-19 Note
15.053 Excel Solver 1 Table of Contents Introduction to Excel Solver slides 3- : Diet Problem, Set-Up slides 5-11 : Diet Problem, Dialog Box slides 12-17 Example 2: Food Start-Up Problem slides 18-19 Note
Software Version 1.0. User s Manual
 Software Version 1.0 User s Manual Table of Contents Contents 0 Important Information about Your FreeStyle Libre software...1 Intended Use...1 System Requirements...1 Customer Service...1 Getting to Know
Software Version 1.0 User s Manual Table of Contents Contents 0 Important Information about Your FreeStyle Libre software...1 Intended Use...1 System Requirements...1 Customer Service...1 Getting to Know
ab HRV 3C Protease Inhibitor Screening Kit (Colorimetric)
 Version 2 Last updated 2 March 2017 ab211089 HRV 3C Protease Inhibitor Screening Kit (Colorimetric) For the rapid, sensitive and accurate screening of potential HRV 3C Protease inhibitors. This product
Version 2 Last updated 2 March 2017 ab211089 HRV 3C Protease Inhibitor Screening Kit (Colorimetric) For the rapid, sensitive and accurate screening of potential HRV 3C Protease inhibitors. This product
ADVANCED VBA FOR PROJECT FINANCE Near Future Ltd. Registration no
 ADVANCED VBA FOR PROJECT FINANCE f i n a n c i a l f o r e c a s t i n G 2017 Near Future Ltd. Registration no. 10321258 www.nearfuturefinance.com info@nearfuturefinance.com COURSE OVERVIEW This course
ADVANCED VBA FOR PROJECT FINANCE f i n a n c i a l f o r e c a s t i n G 2017 Near Future Ltd. Registration no. 10321258 www.nearfuturefinance.com info@nearfuturefinance.com COURSE OVERVIEW This course
38 Int'l Conf. Bioinformatics and Computational Biology BIOCOMP'16
 38 Int'l Conf. Bioinformatics and Computational Biology BIOCOMP'16 PGAR: ASD Candidate Gene Prioritization System Using Expression Patterns Steven Cogill and Liangjiang Wang Department of Genetics and
38 Int'l Conf. Bioinformatics and Computational Biology BIOCOMP'16 PGAR: ASD Candidate Gene Prioritization System Using Expression Patterns Steven Cogill and Liangjiang Wang Department of Genetics and
User Manual second language
 User Manual second language GlucoTel Blood Glucose Monitoring and Diabetes Management System must be used with cell phones that have: Table of contents 2 3 Introduction 4 Bluetooth Wireless Technology
User Manual second language GlucoTel Blood Glucose Monitoring and Diabetes Management System must be used with cell phones that have: Table of contents 2 3 Introduction 4 Bluetooth Wireless Technology
