MCMDA: Matrix Completion for MiRNA-Disease Association prediction
|
|
|
- Shauna Simpson
- 5 years ago
- Views:
Transcription
1 /, Advance Publications 2017 MCMDA: Matrix Completion for MiRNA-Disease Association prediction Jian-Qiang Li 1,*, Zhi-Hao Rong 2,*, Xing Chen 3, Gui-Ying Yan 4, Zhu-Hong You 5 1 College of Computer Science and Software Engineering, Shenzhen University, Shenzhen, , China 2 School of Software, Beihang University, Beijing, , China 3 School of Information and Control Engineering, China University of Mining and Technology, Xuzhou, , China 4 Academy of Mathematics and Systems Science, Chinese Academy of Sciences, Beijing, , China 5 Xinjiang Technical Institute of Physics and Chemistry, Chinese Academy of Science, ürümqi, , China * The authors wish it to be known that, in their opinion, The first two authors should be regarded as joint First Authors Correspondence to: Xing Chen, xingchen@amss.ac.cn Keywords: mirna, disease, mirna-disease association, matrix completion Received: October 17, 2016 Accepted: January 09, 2017 Published: February 03, 2017 ABSTRACT Nowadays, researchers have realized that micrornas (mirnas) are playing a significant role in many important biological processes and they are closely connected with various complex human diseases. However, since there are too many possible mirna-disease associations to analyze, it remains difficult to predict the potential mirnas related to human diseases without a systematic and effective method. In this study, we developed a Matrix Completion for MiRNA-Disease Association prediction model (MCMDA) based on the known mirna-disease associations in HMDD database. MCMDA model utilized the matrix completion algorithm to update the adjacency matrix of known mirna-disease associations and furthermore predict the potential associations. To evaluate the performance of MCMDA, we performed leave-oneout cross validation (LOOCV) and 5-fold cross validation to compare MCMDA with three previous classical computational models (RLSMDA, HDMP, and WBSMDA). As a result, MCMDA achieved AUCs of in global LOOCV, in local LOOCV and average AUC of / in 5-fold cross validation. Moreover, the prediction results associated with colon neoplasms, kidney neoplasms, lymphoma and prostate neoplasms were verified. As a consequence, 84%, 86%, 78% and 90% of the top 50 potential mirnas for these four diseases were respectively confirmed by recent experimental discoveries. Therefore, MCMDA model is superior to the previous models in that it improves the prediction performance although it only depends on the known mirna-disease associations. INTRODUCTION MicroRNA (mirna) is a kind of short noncoding single-stranded RNA (~22nt) which can regulate the gene expression by binding to the 3 untranslated regions (UTRs) of its target messenger RNA (mrna) through base pairing [1, 2]. There are significant differences between the mirnas in different tissues and different growth stages, which means that mirnas have differential spatial and temporal expression patterns [3]. Based on plenty of biological experiments, researchers now believe that these small molecules have a wide range of regulation effects on eukaryotic gene expression, not only in human genes but also in genes of many other species [4]. Up to now, researchers have discovered that mirnas are involved in a series of critical life processes, including early cell growth, proliferation, differentiation [5, 6], apoptosis, death [7], fat metabolism and so on. Therefore, it is no wonder that mirnas are closely related to many complex human diseases [8, 9]. For example, studies have implicated that mirna-143 and mirna-145 are constantly down-regulated in colorectal 1
2 tumors [10] and recently Croce et al. also have shown that the downregulation of these mirnas is a common occurrence in breast carcinomas [11]. Besides, studies by Takamizawa et al. [12] and Yanaihara et al. [13] have presented evidence that transcripts of certain let-7 homologs are significantly downregulated in human lung cancer. Based on real-time polymerase chain reaction (PCR), the analysis of mirna arrays using pooled RNA samples from five gastric cancer patients indicates that the expression of mirna-107, mirna-21, mirna-196a, mirna-26b, mirna-9, mirna-142-3p, mirna-30b, mirna-150, mirna-191, and mirna-17 was found to be upregulated [14]. However, it is expensive and timeconsuming to identify the associations between mirnas and diseases using experimental methods. Considering that large numbers of mirna-associated datasets are available, computational methods are efficient in predicting mirnadisease associations in that they can select the most promising associated mirnas for further experimental studies [15 17]. Therefore, it is necessary for us to make further efforts and develop efficient computational models to predict the potential mirna-disease associations [16, 18 31]. Many computational methods have been established to predict the potential associations between mirnas and diseases depending on the assumption that mirnas with similar functions are more likely to have connections with diseases which share similar phenotypes [32, 33]. Jiang et al. [34] proposed a hypergeometric distribution-based model to predict mirna-disease associations based on disease phenotype similarity network, mirna functional similarity network, and known human disease-mirna association network. However, this method strongly depends on the mirna-target interactions with a high rate of false positive and false negative samples. Moreover, Shi et al. [35] presented a new model by implementing random walk algorithm on protein-protein interaction (PPI) network based on the idea that mirnas whose target genes are related to certain diseases are more likely to be associated with these diseases. They made use of the mirna target interactions, disease gene associations, and PPIs to acquire potential associations between the mirnas and diseases. Mork et al. [36] proposed a mirpd method with the help of protein-disease interactions as well as protein-mirna interactions, where not only disease-related mirnas but also potential disease-related proteins were analyzed. By integrating known disease gene associations and mirna-target interactions, Xu et al. [37] introduced a mirna prioritization method which need not rely on the known mirna-disease associations. Instead, what they needed to do was to evaluate the similarity between the targets of mirnas and disease genes. Nevertheless, all the methods mentioned above suffered from the mirna-target interactions with high false positive and false negative samples, which could significantly reduce the accuracy of the aforementioned models. Researchers also proposed some other computational models without relying on mirna-target interactions. Based on mirna functional similarity, disease semantic similarity, disease phenotype similarity, and mirnadisease associations, Xuan et al. [38] presented an HDMP model which analyzed the mirnas related to the diseases by considering the functional similarities of the mirna s k most similar neighbors. Compared with the previous methods, HDMP assigned higher weight to the mirnas in the cluster and family since they are more likely to be associated with similar diseases. When applied to new diseases without some known related mirnas, however, HDMP is unable to work since it strongly depends on the neighbors of the mirnas. Besides, HDMP is based on a local similarity measure rather than a global measure which can notably promote the prediction performance. Xuan et al. [39] introduced another model called MIDP based on random walk, which exploited the characteristics of the nodes and the various ranges of topologies. The labeled nodes in MIDP were assigned higher transition weight than the unlabeled nodes, which efficiently exploited the prior information of nodes and various ranges of topologies. What is worth mentioning is that MIDP effectively relieved the negative effect of noisy data. MIDP also extended the walk on a mirna-disease bilayer network to predict candidate specially for the diseases without any known mirnas. Recently, Zeng et al. [40] utilized matrix completion to predict the mirna-disease associations based on mirna-mirna network and disease-disease network. The method contributed multiple feature sets to address problems related to insufficient mirna-disease association data. The method could be applied to predict unknown mirna-disease associations and new pathogenic mirnas for well-characterized diseases. Chen et al. [41] proposed RWRMDA model which integrated mirna-mirna functional similarity and known mirna-disease associations information to predict mirna-disease associations. RWRMDA was motivated based on the investigation that global similarity measures are better in predicting the associations between mirnas and diseases than the previous local network similarity measures. Still, this method fails to predict mirnas associated with new diseases without any known related mirnas. Chen et al. [16] presented another model called WBSMDA based on mirna functional similarity, disease semantic similarity, mirna-disease associations, and Gaussian interaction profile kernel similarity for mirnas and diseases. WBSMDA makes a breakthrough in that it succeeds in predicting related mirnas for new diseases without known related mirnas and new mirnas without known related diseases. Recently, Chen et al. [42] presented a model of HGIMDA using mirna functional similarity, disease semantic similarity, 2
3 mirna-disease associations, and Gaussian interaction profile kernel similarities. In HGIMDA, the new mirna functional similarity network was obtained by combining mirna functional similarity network with Gaussian interaction profile kernel similarities for mirnas. The process of calculating new disease similarity network was quite similar. Then, a heterogeneous graph was obtained by combining new mirna functional similarity network, new disease similarity network and known mirnadisease associations. Moreover, the potential association between a disease and a mirna could be inferred based on an iterative equation if they didn t have known association. It has been verified that HGIMDA obtained a high prediction performance. In addition, several computational models have considered machine learning methods. For instance, Xu et al. [43] developed a mirna target-dysregulated network (MTDN) based on mirna-target interactions as well as mirna and mrna expression profiles. Besides, MTDN implemented support vector machine (SVM) classifier to distinguish positive mirna-disease associations from negative ones. Nevertheless, it is still fairly difficult to obtain the negative mirna-disease associations today, which seriously decreases the prediction performance of this computational model. Chen et al. [15] presented a RLSMDA model based on semi-supervised learning which calculated the semantic similarity between different diseases. It is worth mentioning that RLSMDA could identify related mirnas for diseases without any known associated mirnas, meanwhile avoiding the problem of using negative associations between mirnas and diseases. The trouble of RLSMDA is how to find the appropriate parameters and how to combine the classifiers from mirna space and disease space together. Chen et al. [19] developed another computational model called RBMMMDA based on mirna-disease associations which presented restricted Boltzmann machine (RBM) which is a two-layer undirected graphical model consisting of layers of visible and hidden units. Compared to the previous models, RBMMMDA could obtain not only new mirnadisease associations but also corresponding association types. However, it is still too difficult to learn the complex parameters. In this study, we developed an effective computational model of Matrix Completion for MiRNA- Disease Association prediction model (MCMDA) using matrix completion algorithm based on the known mirna-disease associations to predict the potential mirna-disease associations. Compared to the previous computational models, MCMDA predicts the mirnadisease associations by using the matrix completion algorithm, which is of high efficiency to update the low-rank mirna-disease matrix. Besides, negative associations which are required in some previous computational models are not needed in MCMDA. To evaluate the effectiveness of MCMDA, global and local LOOCV as well as 5-fold cross validation were introduced. The AUCs of global and local LOOCV were respectively and , and the model obtained the average AUC of / on 5-fold cross validation. Besides, the top 10 and top 50 mirnas related to colon neoplasms, kidney neoplasms, lymphoma and prostate neoplasms obtained by MCMDA were examined in dbdemc [44] and mir2disease [45] database. As a result, 84%, 86%, 78% and 90% of the top 50 potential mirnas for these four complex diseases were respectively confirmed by recent experimental discoveries. Thus, it proves that MCMDA is effective in predicting potential mirna-disease associations and it has significant advantages over the previous methods although MCMDA only depends on known mirna-disease associations. RESULTS Performance evaluation We used global and local LOOCV as well as 5-fold cross validation based on the known mirnadisease associations in HMDD database to evaluate the performance of MCMDA. Meanwhile, MCMDA were compared with three previous classical computational methods: WBSMDA [16], RLSMDA [15] and HDMP [38]. In LOOCV evaluation, each known association in the database was regarded as the test sample in turn while the other known associations were regarded as training samples. The mirna-diseases without known association evidences were considered as candidate samples. The scores of all mirna-disease pairs could be obtained after MCMDA was implemented. In global LOOCV, the score of the test sample was compared with the scores of all the candidate samples while in local LOOCV, the test sample was merely compared with the scores of the candidate samples which included the particular disease in the test sample. In 5-fold cross validation, the known mirna-disease associations were randomly divided into five disjoint parts. Each time, one part was picked out as test samples and the other four parts were treated as training samples. Still, the mirna-disease pairs without known association evidences were regarded as candidate samples. Then, the score of each test sample were compared with the scores of all the candidate samples, respectively. This procedure was repeated five times until each known association was used as test sample and its score was compared with the scores of the candidate samples. Those test samples whose ranks exceeded the given threshold were considered to predict the mirnadisease associations correctly. Finally, we drew a receiver operating characteristics curve (ROC) to compare MCMDA with all the previous methods. In this curve, the true positive rate (TPR, sensitivity) and false positive rate (FPR, 1-specificity) were plotted [46]. Sensitivity represents the percentage 3
4 of mirna-disease test samples whose ranks exceeded the given threshold while specificity represents the percentage of negative mirna-disease associations whose ranks were lower than the threshold [47]. The area under the ROC curve (AUC) was calculated to evaluate the accuracy of MCMDA. If AUC=1, MCMDA proves to be a prefect performance. AUC of 0.5 means that the method merely has a random prediction performance. As a result, the AUCs of MCMDA, WBSMDA, RLSMDA and HDMP were , , , and , respectively in global LOOCV. For local LOOCV, MCMDA, WBSMDA, RLSMDA and HDMP acquired AUCs of , , and , respectively. The average AUCs of MCMDA, WBSMDA, RLSMDA, HDMP were / , / , / and / , respectively in 5-fold cross validation (See Figure 1). All in all, MCMDA turns out to be more effective in predicting potential mirnadisease associations compared with the previous methods, especially considering that MCMDA merely depends on the known mirna-disease associations in the database. Case studies Furthermore, case studies of four significant diseases related to human health were implemented to practically evaluate the prediction accuracy of MCMDA. The top 10 and top 50 predicted mirnas related with these diseases were examined by another two mirna-disease databases, dbdemc [44] and mir2disease [45]. Colon Neoplasms is a malignant cancer which is commonly found in the boundary of rectum and sigmoid colon [48]. It is the third most common cancer and the third leading cause of cancer death for both men and women in the United States [49]. However, early patients of colon neoplasms only suffer from subtle symptoms [50], making the disease difficult to be detected. To make things worse, it is reported that its occurrence rate has an increasing trend these years [51]. Thus, it is urgent to predict the potential mirnas related to colon neoplasms. With the help of the modern iatrology, many mirnas have been confirmed to be correlated with colon neoplasms. For instance, mirna-145 targets the insulin receptor substrate-1 and thus inhibits the growth of colon cancer cells [52]. Besides, mirna-126, which is frequently lost in colon neoplasms cells, has the function of suppressing the growth of neoplastic cells by targeting phosphatidylinositol 3-kinase signaling [53]. MCMDA was implemented to predict the top 50 mirnas associated with colon neoplasms. Therefore, 9 of the top 10 and 42 of the top 50 predicted mirnas associated with colon neoplasms were verified by dbdemc and mir2disease database (See Table 1). Kidney neoplasms, also known as renal cancer, is a cancer starting in the cells of kidney that includes many different types [54]. The two most common types of kidney Figure 1: Performance evaluation comparison between MCMDA and three previous prediction models (RLSMDA, HDMP, WBSMDA) in terms of ROC curve and AUC based on global LOOCV and local LOOCV tested by known mirna-disease associations in the HMDD database. MCMDA achieved AUC of in global LOOCV and in local LOOCV. Thus, the performance of MCMDA is almost better than all the previous models in some degree and it proves to be effective in predicting the potential mirna-disease associations. 4
5 Table 1: Prediction of the top 50 predicted mirnas associated with colon neoplasms based on known associations in HMDD database mirna Evidence mirna Evidence hsa-mir-146a dbdemc hsa-mir-196a dbdemc;mir2disease hsa-mir-155 dbdemc;mir2disease hsa-mir-29c dbdemc hsa-mir-122 unconfirmed hsa-mir-223 dbdemc;mir2disease hsa-mir-21 dbdemc;mir2disease hsa-mir-143 dbdemc;mir2disease hsa-mir-34a dbdemc;mir2disease hsa-let-7a unconfirmed hsa-mir-221 dbdemc;mir2disease hsa-mir-195 dbdemc;mir2disease hsa-mir-16 dbdemc hsa-mir-200b dbdemc hsa-mir-125b dbdemc hsa-mir-214 dbdemc hsa-mir-29a dbdemc;mir2disease hsa-mir-106b dbdemc;mir2disease hsa-mir-29b dbdemc;mir2disease hsa-mir-23a mir2disease hsa-mir-15a dbdemc hsa-mir-142 unconfirmed hsa-mir-133a dbdemc;mir2disease hsa-mir-31 dbdemc;mir2disease hsa-mir-222 dbdemc hsa-mir-34c mir2disease hsa-mir-20a dbdemc;mir2disease hsa-mir-141 dbdemc;mir2disease hsa-mir-199a unconfirmed hsa-mir-148a dbdemc hsa-mir-26a dbdemc;mir2disease hsa-mir-182 dbdemc;mir2disease hsa-mir-1 dbdemc;mir2disease hsa-mir-200a unconfirmed hsa-mir-19b dbdemc;mir2disease hsa-let-7c dbdemc hsa-mir-19a dbdemc;mir2disease hsa-mir-101 unconfirmed hsa-mir-15b mir2disease hsa-mir-192 dbdemc;mir2disease hsa-mir-18a mir2disease hsa-mir-181a dbdemc;mir2disease hsa-mir-92a dbdemc hsa-mir-9 dbdemc;mir2disease hsa-mir-206 unconfirmed hsa-mir-133b dbdemc;mir2disease hsa-mir-30b dbdemc;mir2disease hsa-mir-34b dbdemc;mir2disease hsa-mir-150 dbdemc;mir2disease hsa-mir-183 dbdemc;mir2disease The first column records top 1-25 related mirnas. The second column records the top related mirnas. cancer are renal cell carcinoma (RCC) and transitional cell carcinoma (TCC, also known as urothelial cell carcinoma) of the renal pelvis [55]. The most common symptoms of kidney neoplasms patients are pains in the lumbar and hematuria [56]. Many existing kidney neoplasm-related mirnas have been reported based on recent biological experiments. For example, the common target ACVR2B of five mirnas (mirna-192, mirna-194, mirna-215, mirna-200c and mirna-141) is strongly expressed in renal childhood neoplasms [57]. In addition, mirna-23b, by targeting proline oxidase, a novel tumor suppressor protein, could function as an oncogene in renal cancer [58]. Thus, the decreasing mirna-23b expression may prove to be an effective way of inhibiting kidney tumor growth [58]. Based on MCMDA, 7 of the top 10 potential mirnas associated with kidney neoplasms were confirmed by dedemc and mir2disease database while 43 were verified of the top 50 (See Table 2). Lymphoma is a malignant tumor originating in the lymphatic hematopoietic system [59] which consists of two categories: non-hodgkinlymphoma (NHL) and Hodgkin'slymphoma (HL) [60]. Lymphoma is thought 5
6 Table 2: Prediction of the top 50 predicted mirnas associated with kidney neoplasms based on known associations in HMDD database mirna Evidence mirna Evidence hsa-mir-155 dbdemc hsa-mir-92a unconfirmed hsa-mir-146a dbdemc hsa-mir-195 dbdemc hsa-mir-122 dbdemc;mir2disease hsa-mir-126 dbdemc;mir2disease hsa-mir-34a dbdemc hsa-mir-29c dbdemc;mir2disease hsa-mir-221 unconfirmed hsa-mir-23a dbdemc hsa-mir-16 dbdemc hsa-mir-143 dbdemc hsa-mir-125b unconfirmed hsa-mir-223 dbdemc hsa-mir-29a dbdemc;mir2disease hsa-mir-214 dbdemc;mir2disease hsa-mir-133a unconfirmed hsa-let-7a dbdemc hsa-mir-29b dbdemc;mir2disease hsa-mir-148a dbdemc hsa-mir-145 dbdemc hsa-mir-200b dbdemc;mir2disease hsa-mir-26a dbdemc;mir2disease hsa-mir-31 dbdemc hsa-mir-199a dbdemc;mir2disease hsa-mir-142 unconfirmed hsa-mir-222 dbdemc hsa-mir-106b dbdemc;mir2disease hsa-mir-1 dbdemc hsa-mir-34c dbdemc hsa-mir-15b dbdemc hsa-mir-182 dbdemc;mir2disease hsa-mir-20a dbdemc;mir2disease hsa-mir-200a dbdemc hsa-mir-17 dbdemc;mir2disease hsa-mir-101 dbdemc;mir2disease hsa-mir-30b dbdemc hsa-let-7c dbdemc hsa-mir-206 dbdemc hsa-mir-181a dbdemc hsa-mir-19a dbdemc hsa-mir-9 dbdemc hsa-mir-196a dbdemc hsa-mir-34b dbdemc hsa-mir-19b dbdemc;mir2disease hsa-mir-183 dbdemc hsa-mir-18a dbdemc hsa-mir-133b unconfirmed hsa-mir-150 dbdemc;mir2disease hsa-let-7b unconfirmed The first column records top 1-25 related mirnas. The second column records the top related mirnas. to be associated with gene mutations, as well as viruses, pathogens, radiation, chemical drugs, autoimmune diseases, etc. [61]. For example, re-expression of mirna-150 induces EBV-positive Burkitt lymphoma differentiation by modulating c-myb in vitro [62]. Besides, the expressions of mirna-21 and mirna-210 in plasma of previously untreated lymphoma patient group were higher than those of the patients treated for 6 or more courses [63]. MCMDA model predicts the top 10 and top 50 mirnas related to lymphoma. As a result, 9 of the top 10 and 39 of the top 50 potential mirnas were confirmed in the dedemc and mir2disease database (See Table 3). Prostate neoplasms is a malignant tumor which originates in the epithelial cells of prostate [64]. Factors that increase the risk of prostate neoplasms include older age, a family history of the disease, race and a diet high in processed meat, red meat or milk products or low in certain vegetables [65]. Up to now, lots of mirnas have been discovered to be associated with prostate neoplasms. For instance, the proto-oncogene ERG is a target of mirna-145 in prostate cancer [66]. MCMDA predicts the top 10 and top 50 potential mirnas which are associated with prostate neoplasms. As a consequence, 9 of the top 10 and 45 of the top 50 6
7 Table 3: Prediction of the top 50 predicted mirnas associated with lymphoma based on known associations in HMDD database mirna Evidence mirna Evidence hsa-mir-30b dbdemc hsa-mir-208a unconfirmed hsa-mir-148a dbdemc hsa-mir-26b dbdemc hsa-mir-373 dbdemc hsa-mir-143 unconfirmed hsa-mir-196a dbdemc hsa-mir-9 dbdemc hsa-mir-23a dbdemc hsa-let-7b dbdemc hsa-mir-206 dbdemc hsa-mir-96 dbdemc hsa-mir-195 dbdemc hsa-let-7d dbdemc hsa-mir-372 unconfirmed hsa-mir-93 dbdemc hsa-mir-199a dbdemc hsa-mir-483 unconfirmed hsa-mir-15b dbdemc hsa-mir-371a unconfirmed hsa-mir-34c unconfirmed hsa-let-7e dbdemc;mir2disease hsa-mir-34b dbdemc hsa-mir-7 dbdemc hsa-mir-183 dbdemc hsa-mir-223 dbdemc hsa-mir-132 dbdemc hsa-mir-106a dbdemc;mir2disease hsa-mir-214 dbdemc hsa-mir-205 dbdemc hsa-mir-182 dbdemc hsa-mir-222 dbdemc hsa-mir-31 unconfirmed hsa-mir-335 dbdemc hsa-mir-133a dbdemc hsa-mir-27a dbdemc hsa-mir-212 dbdemc hsa-mir-181c dbdemc hsa-mir-141 dbdemc hsa-mir-224 dbdemc hsa-mir-142 unconfirmed hsa-mir-27b dbdemc hsa-mir-192 dbdemc hsa-mir-30a dbdemc hsa-mir-429 unconfirmed hsa-mir-370 unconfirmed hsa-mir-451a unconfirmed hsa-mir-1 dbdemc hsa-mir-106b dbdemc hsa-let-7g dbdemc The first column records top 1-25 related mirnas. The second column records the top related mirnas. predicted mirnas were confirmed in the dbdemc and mir2disease database (See Table 4). The result of case studies on the four aforementioned human diseases illustrates that MCMDA achieves excellent prediction performance. Moreover, we prioritized the potential mirnas associated with all the human diseases in HMDD database (See Supplementary Table 1). We hope that the predictions of MCMDA can be verified in future scientific researches. DISCUSSION Nowadays, researchers propose several computational methods to predict the potential associations between mirnas and diseases because computational models could select the most promising mirnas related to human diseases and are less expensive than the traditional experimental methods. In order to predict potential mirna-disease associations, we developed a computational model of MCMDA by analyzing the known mirna-disease associations and implementing the matrix completion algorithm to get the association score of each mirna-disease pair. MCMDA obtained excellent prediction performances based on LOOCV and 5-fold cross validation. In addition, the predicted mirnas associated with four important human diseases: colon neoplasms, kidney neoplasms, lymphoma and prostate neoplasms, were verified by the experimental literatures 7
8 Table 4: Prediction of the top 50 predicted mirnas associated with prostate neoplasms based on known associations in HMDD database mirna Evidence mirna Evidence hsa-mir-146a mir2disease hsa-mir-150 dbdemc hsa-mir-122 unconfirmed hsa-mir-126 dbdemc;mir2disease hsa-mir-155 dbdemc hsa-mir-195 dbdemc;mir2disease hsa-mir-21 dbdemc;mir2disease hsa-mir-29c dbdemc hsa-mir-34a dbdemc;mir2disease hsa-mir-223 dbdemc;mir2disease hsa-mir-16 dbdemc;mir2disease hsa-mir-143 dbdemc;mir2disease hsa-mir-221 dbdemc;mir2disease hsa-mir-23a dbdemc;mir2disease hsa-mir-29a dbdemc hsa-let-7a dbdemc;mir2disease hsa-mir-133a dbdemc hsa-mir-200b unconfirmed hsa-mir-29b dbdemc;mir2disease hsa-mir-214 dbdemc;mir2disease hsa-mir-15a dbdemc;mir2disease hsa-mir-148a mir2disease hsa-mir-26a dbdemc;mir2disease hsa-mir-106b dbdemc hsa-mir-222 dbdemc;mir2disease hsa-mir-34c dbdemc hsa-mir-199a dbdemc;mir2disease hsa-mir-142 unconfirmed hsa-mir-1 dbdemc hsa-mir-31 dbdemc;mir2disease hsa-mir-20a mir2disease hsa-mir-141 mir2disease hsa-mir-17 mir2disease hsa-mir-182 dbdemc;mir2disease hsa-mir-15b dbdemc hsa-mir-200a dbdemc hsa-mir-19a dbdemc hsa-mir-101 dbdemc;mir2disease hsa-mir-19b dbdemc;mir2disease hsa-let-7c dbdemc;mir2disease hsa-mir-206 dbdemc hsa-mir-192 dbdemc hsa-mir-30b dbdemc;mir2disease hsa-mir-181a dbdemc;mir2disease hsa-mir-18a unconfirmed hsa-mir-9 dbdemc hsa-mir-196a dbdemc hsa-mir-34b dbdemc hsa-mir-92a unconfirmed hsa-mir-133b dbdemc The first column records top 1-25 related mirnas. The second column records the top related mirnas. in dbdemc and mir2disease database. The results from cross validation and case studies indicated that MCMDA was effective in predicting potential mirna-disease associations although it only depends on known mirnadisease associations. The reasons why MCMDA achieved excellent performances are as follows. Firstly, MCMDA predicts the mirna-disease associations by using the matrix completion algorithm based on the observation that the mirna-disease matrix is low-rank. MCMDA fills the candidate samples without known associations with 0 and then iteratively updates them with the predictive scores. Besides, MCMDA is based on the known mirnadisease associations in HMDD database. Plenty of known associations guarantee the efficiency of the predictions in MCMDA. Finally, negative associations which are required in some previous models are not needed in MCMDA. Yet, there still exist several limitations in MCMDA. Firstly, MCMDA method is based on the known mirnadisease associations, which means it cannot predict the potential mirnas associated with the new diseases without any known related mirnas and potential diseases associated with new mirnas. Besides, there is no powerful method to find the optimal parameters for MCMDA. Finally, the current mirna-disease associations are insufficient. To be specific, there are merely 5430 known mirna-disease associations within 8
9 Figure 2: Flowchart of MCMDA model to predict the potential mirna-disease associations based on the known associations in HMDD database. 9
10 the possible exploration spaces of 495 mirnas and 383 diseases. The more known associations are confirmed in the future, the more accurate MCMDA model can become. MATERIALS AND METHODS Human mirna-disease associations The known mirna-disease associations were downloaded from HMDD v2.0 database [67] which consisted of 5430 known mirna-disease associations, 495 mirnas, and 383 diseases. We furthermore constructed an adjacency matrix M to represent known mirna-disease associations. For instance, if mirna mi () is reported to be associated with disease d( j) in the database, the value of M(, ij) is 1 and otherwise 0. Ω denotes the set of all the known associations in matrix M which means (, i j) Ω if mi () is associated with d( j). n m represents the number of mirnas in HMDD database and n d represents the number of diseases. MCMDA We developed MCMDA based on the known mirna-disease associations in HMDD database to predict the potential associations (See Figure 2). MCMDA uses the singular value thresholding (SVT) algorithm to accomplish the matrix completion procedure. First, the mirna-disease association matrix M was obtained according to known mirna-disease associations. Here, all the known associations between mirnas and diseases in HMDD database are used as training samples. The matrix completion algorithm is iterative and a n n m d prediction matrix X k (k denotes the iteration times) can be obtained in each iteration. When MCMDA ends, the matrix X n (n denotes the ultimate iteration times) is obtained which records the scores of all the possible mirna-disease pairs. To ensure that the scores of known associations in X n are close to those in M, the following optimization problem needs to be solved. min f ( X) X τ st.. P ( X) = P ( M) Ω Ω (1) where X is a n n m d candidate solution matrix with scores of all the unknown mirna-disease samples, P Ω is the orthogonal projector onto the span of matrices vanishing outside of Ω so that the (i,j) th component of P ( X) Ω is equal to X(, i j) if (, i j) Ω or zero otherwise. f ( X) τ is a nonlinear function of X which can be written as the following form. 1 minτ X + X * 2 X 2 F st.. P ( X) = P ( M) Ω Ω (2) where X * is the nuclear form of the matrix X which is the sum of the singular values of X, X F denotes the Frobernius form of X which is n m d i= 1 n j= 1 2 X(, i j), τ is a thresholding which will be introduced later. According to [68], problem (2) can be optimized using the Lagrangian multiplier method. Specifically, we introduce a Lagrangian multiplier Y and get the Lagrangian function as below: LXY (, ) = f ( X) + < Y, P ( M) P ( X) > (3) τ Ω Ω The singular value decomposition (SVD) of matrix X with rank r, which represents the number of singular values of matrix X, is needed in matrix completion algorithm. * X = UΣV, Σ= diag({ σ } ) i 1 i r (4) where U and V are n r m and r n d matrices. Σ= diag({ σ } ) i 1 i r means that Σ is a r r diagonal matrix with positive singular values { σ i } 1 i r on its main diagonal. For τ 0, we introduce an operator D defined τ as follows: * D ( X) = UD ( Σ) V, D ( Σ ) = diag({ σ τ} ) τ ω ω i + (5) where { σ τ } i + is the positive part of { σ τ } i. In other words, { σ τ } i + is equal to σ i τ if σ τ 0 i or 0 otherwise and it effectively shrinks the singular values of X toward 0. The value of τ is 5 n n m d according to the previous research of matrix completion algorithm [69]. There are two key steps which are special instances of Uzawa s algorithm [70] to find a saddle point of (3) 0 1 n 1 n in each iteration. We introduce { Y, Y,..., Y, Y } which are a series of n n m d matrices to record the intermediate scores of matrices { X 1.. X n }. First, update X with Y: X Then, update Y with X: k = D τ Y k 1 ( ) k k 1 k Y = Y + δ ( M X ) k (7) where Y 0 is a zero matrix [71] and { δ k } k 1 is the step size. It is usually thought that the iteration can converge to an unique solution when 0< δ < 2 [72], specifically, we empirically set the value of { δ } = 1.5 k k 1 according to the excellent performance in previous model [73]. MCMDA applies K.K.T conditions as the stopping criteria which are checked in each iteration to makes sure the scores of the known associations in the prediction matrix are close enough to the original matrix M: k P ( M X ) <ε P ( M ) Ω F Ω F (8) where ε is a stopping tolerance, the value is 10 4 since it proved to be appropriate in restricting the iteration times (6) 10
11 in previous algorithm [71]. If the stopping criteria is met, MCMDA stops iteration immediately and the ultimate matrix X n is obtained. Finally, a parameter maxiter is set which restricts the max iteration times and avoids the infinite loop. Specifically, maxiter is set 500 to ensure that the ultimate matrix has reliable predicted scores. Based on the method mentioned above, the ultimate matrix X n is obtained by above calculation process which can be utilized to predict the potential mirna-disease associations. ACKNOWLEDGMENTS JQL was supported by National Natural Science Foundation of China under Grant No , Natural Science foundation of Guangdong Province under Grant No. 2014A , and Technology Planning Project from Guangdong Province under Grant No. 2014B XC was supported by National Natural Science Foundation of China under Grant No and GYY was supported by National Natural Science Foundation of China under Grant No and ZHY were supported by National Natural Science Foundation of China under Grant No , Pioneer Hundred Talents Program of Chinese Academy of Sciences, and Fundamental Research Funds for the Central Universities under Grant No. 2014YC07. CONFLICTS OF INTEREST The authors declare no conflicts of interest. REFERENCES 1. Llave C, Xie Z, Kasschau KD, Carrington JC. Cleavage of Scarecrow-like mrna targets directed by a class of Arabidopsis mirna. Science. 2002; 297: Eulalio A, Huntzinger E, Izaurralde E. Getting to the Root of mirna-mediated Gene Silencing. Cell. 2008; 132: Telese F, Gamliel A, Skowronska-Krawczyk D, Garcia- Bassets I, Rosenfeld M. Seq-ing Insights into the Epigenetics of Neuronal Gene Regulation. Neuron. 2013; 77: Lu J, Clark AG. Impact of microrna regulation on variation in human gene expression. Genome Res. 2012; 22: Zhu L, Zhao J, Wang J, Hu C, Peng J, Luo R, Zhou C, Liu J, Lin J, Jin Y. MicroRNAs Are Involved in the Regulation of Ovary Development in the Pathogenic Blood Fluke Schistosoma japonicum. Plos Pathogens. 2016; 12:e Fernando TR, Rodriguez-Malave NI, Rao DS. MicroRNAs in B cell development and malignancy. J Hematol Oncol. 2012; 5:7. 7. Lizé M, Pilarski S, Dobbelstein M. E2F1-inducible microrna 449a/b suppresses cell proliferation and promotes apoptosis. Cell Death Differ. 2009; 17: Kim YK. Extracellular micrornas as Biomarkers in Human Disease. Chonnam Med J. 2015; 51: Alaimo S, Giugno R, Pulvirenti A. ncpred: ncrna-disease Association Prediction through Tripartite Network-Based Inference. Front Bioeng Biotechnol. 2014; 2: JM L, RH Z, ST L, CX X, HH J, WJ D, P D, W C, M Y, L C. Down-regulation of fecal mir-143 and mir-145 as potential markers for colorectal cancer. Saudi Med J. 2012; 33: Stahlhut Espinosa CE, Slack FJ. The role of micrornas in cancer. Yale J Biol Med. 2006; 79: Takamizawa J, Konishi H, Yanagisawa K, Tomida S, Osada H, Endoh H, Harano T, Yatabe Y, Nagino M, Nimura Y. Reduced expression of the let-7 micrornas in human lung cancers in association with shortened postoperative survival. Cancer Res. 2004; 64: Yanaihara N, Caplen N, Bowman E, Seike M, Kumamoto K, Yi M, Stephens RM, Okamoto A, Yokota J, Tanaka T. Unique microrna molecular profiles in lung cancer diagnosis and prognosis. Cancer Cell. 2006; 9: Inoue T, Iinuma H, Ogawa E, Inaba T, Fukushima R. Clinicopathological and prognostic significance of microrna-107 and its relationship to DICER1 mrna expression in gastric cancer. Oncol Rep. 2012; 27: Chen X, Yan GY. Semi-supervised learning for potential human microrna-disease associations inference. Sci Rep. 2014; 4: Chen X, Yan CC, Zhang X, You ZH, Deng L, Liu Y, Zhang Y, Dai Q. WBSMDA: Within and Between Score for MiRNA-Disease Association prediction. Sci Rep. 2016; 6: Zeng X, Zhang X, Zou Q. Integrative approaches for predicting microrna function and prioritizing diseaserelated microrna using biological interaction networks. Brief Bioinform. 2016; 17: Chen X. Predicting lncrna-disease associations and constructing lncrna functional similarity network based on the information of mirna. Sci Rep. 2015; 5: Chen X, Yan CC, Zhang X, Li Z, Deng L, Zhang Y, Dai Q. RBMMMDA: predicting multiple types of diseasemicrorna associations. Sci Rep. 2015; 5: Chen X, Yan CC, Luo C, Ji W, Zhang Y, Dai Q. Constructing lncrna functional similarity network based on lncrnadisease associations and disease semantic similarity. Sci Rep. 2015; 5: Chen X, Yan GY. Novel human lncrna-disease association inference based on lncrna expression profiles. Bioinformatics. 2013; 29:
12 22. Chen X, You Z, Yan G, Gong D. IRWRLDA: Improved Random Walk with Restart for LncRNA-Disease Association prediction ; 7: Chen X, Yan CC, Zhang X, You Z-H. Long non-coding RNAs and complex diseases: from experimental results to computational models. Briefings in Bioinformatics. 2016:bbw Chen X, Liu MX, Cui QH, Yan GY. Prediction of Disease-Related Interactions between MicroRNAs and Environmental Factors Based on a Semi-Supervised Classifier. PLoS One. 2012; 7:e Chen X. KATZLDA: KATZ measure for the lncrnadisease association prediction. Sci Rep. 2015; 5: Chen X. mirefrwr: a novel disease-related micrornaenvironmental factor interactions prediction method. Mol Biosyst. 2016; 12: Huang YA, You ZH, Xing C, Chan K, Xin L. Sequencebased prediction of protein-protein interactions using weighted sparse representation model combined with global encoding. Bmc Bioinformatics. 2016; 17: Huang YA, Chen X, You ZH, Huang DS, Chan KC. ILNCSIM: improved lncrna functional similarity calculation model ; 7: Chen X, Huang Y, Wang X, You Z, Chan K. FMLNCSIM: fuzzy measure-based lncrna functional similarity calculation model ; 7: Wong L, You Z-H, Ming Z, Li J, Chen X, Huang Y-A. Detection of interactions between proteins through rotation forest and local phase quantization descriptors. International journal of molecular sciences. 2016; 17: Lan W, Li M, Zhao K, Jin L, Wu FX, Yi P, Wang J. LDAP: a web server for lncrna-disease association prediction. Bioinformatics Pasquier C, Gardès J. Prediction of mirna-disease associations with a vector space model. Sci Rep. 2016; 6: Bandyopadhyay S, Mitra R, Maulik U, Zhang MQ. Development of the human cancer microrna network. Silence. 2010; 1: Jiang Q, Hao Y, Wang G, Juan L, Zhang T, Teng M, Liu Y, Wang Y. Prioritization of disease micrornas through a human phenome-micrornaome network. BMC Syst Biol. 2010; 4:S Shi H, Xu J, Zhang G, Xu L, Li C, Li W, Zheng Z, Wei J, Zheng G, Xia L. Walking the interactome to identify human mirna-disease associations through the functional link between mirna targets and disease genes. BMC Syst Biol. 2013; 7: Mørk S, Pletscherfrankild S, Caro AP, Gorodkin J, Jensen LJ. Protein-driven inference of mirna disease associations. Bioinformatics. 2014; 30: Xu C, Ping Y, Li X, Zhao H, Wang L, Fan H, Xiao Y, Li X. Prioritizing candidate disease mirnas by integrating phenotype associations of multiple diseases with matched mirna and mrna expression profiles. Mol Biosyst. 2014; 10: Xuan P, Han K, Guo M, Guo Y, Li J, Ding J, Liu Y, Dai Q, Li J, Teng Z. Prediction of micrornas Associated with Human Diseases Based on Weighted k Most Similar Neighbors. PLoS One. 2013; 8:e Xuan P, Han K, Guo Y, Li J, Li X, Zhong Y, Zhang Z, Ding J. Prediction of potential disease-associated micrornas based on random walk. Bioinformatics. 2015; 31: Zeng X, Ding N, Rodríguezpatón A, Lin Z, Ju Y. Prediction of MicroRNA disease Associations by Matrix Completion. Current Proteomics. 2016; 13: Chen X, Liu MX, Yan GY. RWRMDA: predicting novel human microrna-disease associations. Mol Biosyst. 2012; 8: Chen X, Yan C, Zhang X, You Z, Huang Y, Yan G. HGIMDA: Heterogeneous Graph Inference for MiRNA- Disease Association prediction ; 7: Xu J, Li CX, Lv JY, Li YS, Xiao Y, Shao TT, Huo X, Li X, Zou Y, Han QL. Prioritizing candidate disease mirnas by topological features in the mirna target-dysregulated network: case study of prostate cancer. Mol Cancer Ther. 2011; 10: Yang Z, Ren F, Liu C, He S, Sun G, Gao Q, Yao L, Zhang Y, Miao R, Cao Y. dbdemc: a database of differentially expressed mirnas in human cancers. BMC genomics. 2010; 11:S Jiang Q, Wang Y, Hao Y, Juan L, Teng M, Zhang X, Li M, Wang G, Liu Y. mir2disease: a manually curated database for microrna deregulation in human disease. Nucleic Acids Res. 2009; 37:D98-D Jiang Q, Hao Y, Wang G, Juan L, Zhang T, Teng M, Liu Y, Wang Y. Prioritization of disease micrornas through a human phenome-micrornaome network. BMC Syst Biol. 2010; 4:S Oved K, Morag A, Pasmanikchor M, Oronkarni V, Shomron N, Rehavi M, Stingl JC, Gurwitz D. Genomewide mirna expression profiling of human lymphoblastoid cell lines identifies tentative SSRI antidepressant response biomarkers. Pharmacogenomics. 2012; 13: Phipps AI, Lindor NM, Jenkins MA, Baron JA, Win AK, Gallinger S, Gryfe R, Newcomb PA. Colon and rectal cancer survival by tumor location and microsatellite instability: the Colon Cancer Family Registry. Dis Colon Rectum. 2013; 56: Liu F, Yuan D, Wei Y, Wang W, Yan L, Wen T, Xu M, Yang J, Li B. Systematic review and meta-analysis of the relationship between EPHX1 polymorphisms and colorectal cancer risk. PLoS One. 2012; 7:e Pita-Fernández S, Pértega-Díaz S, López-Calviño B, Seoane-Pillado T, Gago-García E, Seijo-Bestilleiro R, González-Santamaría P, Pazos-Sierra A. Diagnostic and 12
13 treatment delay, quality of life and satisfaction with care in colorectal cancer patients: a study protocol. Health Qual Life Outcomes. 2013; 11: Chong VH, Abdullah MS, Telisinghe PU, Jalihal A. Colorectal cancer: incidence and trend in Brunei Darussalam. Singapore Med J. 2009; 50: Shi B, Sepplorenzino L, Prisco M, Linsley P, Deangelis T, Baserga R. Micro RNA 145 targets the insulin receptor substrate-1 and inhibits the growth of colon cancer cells. J Biol Chem. 2007; 282: Guo C, Sah JF, Beard L, Willson JKV, Markowitz SD, Guda K. The noncoding RNA, mir-126, suppresses the growth of neoplastic cells by targeting phosphatidylinositol 3-kinase signaling and is frequently lost in colon cancers. Genes Chromosomes Cancer. 2008; 47: Manojlović D, Elaković D, Lj M. [Therapeutic value of transcatheter embolization in malignant tumors of the renal parenchyma]. Srp Arh Celok Lek. 1986; 114: Takehara K, Nishikido M, Koga S, Miyata Y, Harada T, Tamaru N, Kanetake H. Multifocal transitional cell carcinoma associated with renal cell carcinoma in a patient on long-term haemodialysis. Nephrol Dial Transplant. 2002; 17: Duque JLF, Loughlin KR, O Leary MP, Kumar S, Richie JP. Partial nephrectomy: alternative treatment for selected patients with renal cell carcinoma. Urology. 1998; 52: Senanayake U, Das S, Vesely P, Alzoughbi W, Fröhlich LF, Chowdhury P, Leuschner I, Hoefler G, Guertl B. mir-192, mir-194, mir-215, mir-200c and mir-141 are downregulated and their common target ACVR2B is strongly expressed in renal childhood neoplasms. Carcinogenesis. 2012; 33: Liu W, Zabirnyk OH, Shiao YH, Nickerson ML, Khalil S, Anderson LM, Perantoni AO, Phang JM. mir-23b targets proline oxidase, a novel tumor suppressor protein in renal cancer. Oncogene. 2010; 29: Delforoush M, Strese S, Wickström M, Larsson R, Enblad G, Gullbo J. In vitro and in vivo activity of melflufen (J1) in lymphoma. BMC Cancer. 2016; 16: Mcduffie HH, Pahwa P, Karunanayake CP, Spinelli JJ, Dosman JA. Clustering of cancer among families of cases with Hodgkin Lymphoma (HL), Multiple Myeloma (MM), Non-Hodgkin's Lymphoma (NHL), Soft Tissue Sarcoma (STS) and control subjects. BMC Cancer. 2009; 9: Virginia Pascual FA, Pinakeen Patel, Karolina Palucka, Damien Chaussabel, Jacques Banchereau. How the Study of Children with Rheumatic Diseases Identified Interferon alpha and Interleukin 1 as Novel Therapeutic Targets. Immunol Rev. 2008; 223: Chen S, Wang Z, Dai X, Pan J, Ge J, Han X, Wu Z, Zhou X, Zhao T. Re-expression of microrna-150 induces EBVpositive Burkitt lymphoma differentiation by modulating c-myb in vitro. Cancer Sci. 2013; 104: Tian-Tian GE, Liang Y, Rong FU, Wang GJ, Er-Bao R, Wen QU, Wang XM, Liu H, Yu-Hong WU, Song J. Expressions of mir-21, mir-155 and mir-210 in Plasma of Patients withlymphoma and Its Clinical Significance. Zhongguo shi yan xue ye xue za zhi 2012; 20: Gmyrek GA, Walburg M, Webb CP, Yu HM, You X, Vaughan ED, Vande Woude GF, Knudsen BS. Normal and malignant prostate epithelial cells differ in their response to hepatocyte growth factor/scatter factor. Am J Pathol. 2001; 159: Walsh PC, Partin AW. Family history facilitates the early diagnosis of prostate carcinoma. Cancer. 1997; 80: Hart M, Wach S, Nolte E, Szczyrba J, Menon R, Taubert H, Hartmann A, Stoehr R, Wieland W, Grässer FA. The protooncogene ERG is a target of microrna mir-145 in prostate cancer. FEBS J. 2013; 280: Li Y, Qiu C, Tu J, Geng B, Yang J, Jiang T, Cui Q. HMDD v2.0: a database for experimentally supported human microrna and disease associations. Nucleic Acids Res. 2014; 42:D1070-D Bertsekas DP. (1999). Nonlinear programming: Athena scientific Belmont). 69. Candès EJ, Recht B. Exact Matrix Completion via Convex Optimization. Foundations of Computational Mathematics. 2008; 9: Elman HC, Golub GH. Inexact and preconditioned Uzawa algorithms for saddle point problems. Siam J Numer Anal. 2010; 31: Cai JF, Cand, S, Emmanuel J., Shen Z. A Singular Value Thresholding Algorithm for Matrix Completion. Siam Journal on Optimization. 2010; 20: Kanehisa M, Goto S, Hattori M, Aokikinoshita KF, Itoh M, Kawashima S, Katayama T, Araki M, Hirakawa M. From genomics to chemical genomics: new developments in KEGG. Nucleic Acids Res. 2006; 34:D354-D Liao Q, Guan N, Wu C, Zhang Q. Predicting Unknown Interactions Between Known Drugs and Targets via Matrix Completion. Lecture Notes in Computer Science. 2016; 9651:
PRMDA: personalized recommendation-based MiRNA-disease association prediction
 /, 2017, Vol. 8, (No. 49), pp: 85568-85583 PRMDA: personalized recommendation-based MiRNA-disease association prediction Zhu-Hong You 1,*, Luo-Pin Wang 2,*, Xing Chen 3, Shanwen Zhang 1, Xiao-Fang Li 1,
/, 2017, Vol. 8, (No. 49), pp: 85568-85583 PRMDA: personalized recommendation-based MiRNA-disease association prediction Zhu-Hong You 1,*, Luo-Pin Wang 2,*, Xing Chen 3, Shanwen Zhang 1, Xiao-Fang Li 1,
Predicting microrna-disease associations using bipartite local models and hubness-aware regression
 RNA Biology ISSN: 1547-6286 (Print) 1555-8584 (Online) Journal homepage: http://www.tandfonline.com/loi/krnb20 Predicting microrna-disease associations using bipartite local models and hubness-aware regression
RNA Biology ISSN: 1547-6286 (Print) 1555-8584 (Online) Journal homepage: http://www.tandfonline.com/loi/krnb20 Predicting microrna-disease associations using bipartite local models and hubness-aware regression
Genome-wide association study of esophageal squamous cell carcinoma in Chinese subjects identifies susceptibility loci at PLCE1 and C20orf54
 CORRECTION NOTICE Nat. Genet. 42, 759 763 (2010); published online 22 August 2010; corrected online 27 August 2014 Genome-wide association study of esophageal squamous cell carcinoma in Chinese subjects
CORRECTION NOTICE Nat. Genet. 42, 759 763 (2010); published online 22 August 2010; corrected online 27 August 2014 Genome-wide association study of esophageal squamous cell carcinoma in Chinese subjects
Prediction of potential disease-associated micrornas based on random walk
 Bioinformatics, 3(), 25, 85 85 doi:.93/bioinformatics/btv39 Advance Access Publication Date: 23 January 25 Original Paper Data and text mining Prediction of potential disease-associated micrornas based
Bioinformatics, 3(), 25, 85 85 doi:.93/bioinformatics/btv39 Advance Access Publication Date: 23 January 25 Original Paper Data and text mining Prediction of potential disease-associated micrornas based
Downregulation of serum mir-17 and mir-106b levels in gastric cancer and benign gastric diseases
 Brief Communication Downregulation of serum mir-17 and mir-106b levels in gastric cancer and benign gastric diseases Qinghai Zeng 1 *, Cuihong Jin 2 *, Wenhang Chen 2, Fang Xia 3, Qi Wang 3, Fan Fan 4,
Brief Communication Downregulation of serum mir-17 and mir-106b levels in gastric cancer and benign gastric diseases Qinghai Zeng 1 *, Cuihong Jin 2 *, Wenhang Chen 2, Fang Xia 3, Qi Wang 3, Fan Fan 4,
SNMDA: A novel method for predicting microrna disease associations based on sparse neighbourhood
 Received: 18 April 2018 Revised: 25 May 2018 Accepted: 21 June 2018 DOI: 10.1111/jcmm.13799 ORIGINAL ARTICLE SNMDA: A novel method for predicting microrna disease associations based on sparse neighbourhood
Received: 18 April 2018 Revised: 25 May 2018 Accepted: 21 June 2018 DOI: 10.1111/jcmm.13799 ORIGINAL ARTICLE SNMDA: A novel method for predicting microrna disease associations based on sparse neighbourhood
M icrornas (mirnas) are a class of small endogenous single-stranded non-coding RNAs (,22 nt),
 OPEN SUBJECT AREAS: MIRNAS COMPUTATIONAL BIOLOGY AND BIOINFORMATICS SYSTEMS BIOLOGY CANCER GENOMICS Received 7 May 2014 Accepted 13 June 2014 Published 30 June 2014 Correspondence and requests for materials
OPEN SUBJECT AREAS: MIRNAS COMPUTATIONAL BIOLOGY AND BIOINFORMATICS SYSTEMS BIOLOGY CANCER GENOMICS Received 7 May 2014 Accepted 13 June 2014 Published 30 June 2014 Correspondence and requests for materials
Expression of mir-146a-5p in patients with intracranial aneurysms and its association with prognosis
 European Review for Medical and Pharmacological Sciences 2018; 22: 726-730 Expression of mir-146a-5p in patients with intracranial aneurysms and its association with prognosis H.-L. ZHANG 1, L. LI 2, C.-J.
European Review for Medical and Pharmacological Sciences 2018; 22: 726-730 Expression of mir-146a-5p in patients with intracranial aneurysms and its association with prognosis H.-L. ZHANG 1, L. LI 2, C.-J.
Integrating random walk and binary regression to identify novel mirna-disease association
 Niu et al. BMC Bioinformatics (2019) 20:59 https://doi.org/10.1186/s12859-019-2640-9 RESEARCH ARTICLE Open Access Integrating random walk and binary regression to identify novel mirna-disease association
Niu et al. BMC Bioinformatics (2019) 20:59 https://doi.org/10.1186/s12859-019-2640-9 RESEARCH ARTICLE Open Access Integrating random walk and binary regression to identify novel mirna-disease association
A novel method for identifying potential disease-related mirnas via disease-mirna-target heterogeneous network
 Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 A novel method for identifying potential disease-related mirnas via disease-mirna-target
Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 A novel method for identifying potential disease-related mirnas via disease-mirna-target
CircHIPK3 is upregulated and predicts a poor prognosis in epithelial ovarian cancer
 European Review for Medical and Pharmacological Sciences 2018; 22: 3713-3718 CircHIPK3 is upregulated and predicts a poor prognosis in epithelial ovarian cancer N. LIU 1, J. ZHANG 1, L.-Y. ZHANG 1, L.
European Review for Medical and Pharmacological Sciences 2018; 22: 3713-3718 CircHIPK3 is upregulated and predicts a poor prognosis in epithelial ovarian cancer N. LIU 1, J. ZHANG 1, L.-Y. ZHANG 1, L.
Association of mir-21 with esophageal cancer prognosis: a meta-analysis
 Association of mir-21 with esophageal cancer prognosis: a meta-analysis S.-W. Wen 1, Y.-F. Zhang 1, Y. Li 1, Z.-X. Liu 2, H.-L. Lv 1, Z.-H. Li 1, Y.-Z. Xu 1, Y.-G. Zhu 1 and Z.-Q. Tian 1 1 Department of
Association of mir-21 with esophageal cancer prognosis: a meta-analysis S.-W. Wen 1, Y.-F. Zhang 1, Y. Li 1, Z.-X. Liu 2, H.-L. Lv 1, Z.-H. Li 1, Y.-Z. Xu 1, Y.-G. Zhu 1 and Z.-Q. Tian 1 1 Department of
Review Article Long Noncoding RNA H19 in Digestive System Cancers: A Meta-Analysis of Its Association with Pathological Features
 BioMed Research International Volume 2016, Article ID 4863609, 8 pages http://dx.doi.org/10.1155/2016/4863609 Review Article Long Noncoding RNA H19 in Digestive System Cancers: A Meta-Analysis of Its Association
BioMed Research International Volume 2016, Article ID 4863609, 8 pages http://dx.doi.org/10.1155/2016/4863609 Review Article Long Noncoding RNA H19 in Digestive System Cancers: A Meta-Analysis of Its Association
Up-regulation of long non-coding RNA SNHG6 predicts poor prognosis in renal cell carcinoma
 European Review for Medical and Pharmacological Sciences 2018; 22: 8624-8629 Up-regulation of long non-coding RNA SNHG6 predicts poor prognosis in renal cell carcinoma H.-X. AN 1, B. XU 2, Q. WANG 3, Y.-S.
European Review for Medical and Pharmacological Sciences 2018; 22: 8624-8629 Up-regulation of long non-coding RNA SNHG6 predicts poor prognosis in renal cell carcinoma H.-X. AN 1, B. XU 2, Q. WANG 3, Y.-S.
CURRICULUM VITA OF Xiaowen Chen
 Xiaowen Chen College of Bioinformatics Science and Technology Harbin Medical University Harbin, 150086, P. R. China Mobile Phone:+86 13263501862 E-mail: hrbmucxw@163.com Education Experience 2008-2011
Xiaowen Chen College of Bioinformatics Science and Technology Harbin Medical University Harbin, 150086, P. R. China Mobile Phone:+86 13263501862 E-mail: hrbmucxw@163.com Education Experience 2008-2011
Circular RNA_LARP4 is lower expressed and serves as a potential biomarker of ovarian cancer prognosis
 European Review for Medical and Pharmacological Sciences 2018; 22: 7178-7182 Circular RNA_LARP4 is lower expressed and serves as a potential biomarker of ovarian cancer prognosis T. ZOU, P.-L. WANG, Y.
European Review for Medical and Pharmacological Sciences 2018; 22: 7178-7182 Circular RNA_LARP4 is lower expressed and serves as a potential biomarker of ovarian cancer prognosis T. ZOU, P.-L. WANG, Y.
Original Article MicroRNA-27a acts as a novel biomarker in the diagnosis of patients with laryngeal squamous cell carcinoma
 Int J Clin Exp Pathol 2016;9(2):2049-2053 www.ijcep.com /ISSN:1936-2625/IJCEP0014841 Original Article MicroRNA-27a acts as a novel biomarker in the diagnosis of patients with laryngeal squamous cell carcinoma
Int J Clin Exp Pathol 2016;9(2):2049-2053 www.ijcep.com /ISSN:1936-2625/IJCEP0014841 Original Article MicroRNA-27a acts as a novel biomarker in the diagnosis of patients with laryngeal squamous cell carcinoma
EMPEROR'S COLLEGE MTOM COURSE SYLLABUS HERB FORMULAE II
 COURSE DESCRIPTION The second of three courses in the Herb Formulae series. Categories covered in Formulae II include the Tonify Qi and Blood, Regulate Qi, Invigorate the Blood, Stop Bleeding, Stabilize
COURSE DESCRIPTION The second of three courses in the Herb Formulae series. Categories covered in Formulae II include the Tonify Qi and Blood, Regulate Qi, Invigorate the Blood, Stop Bleeding, Stabilize
Expression of mir-1294 is downregulated and predicts a poor prognosis in gastric cancer
 European Review for Medical and Pharmacological Sciences 2018; 22: 5525-5530 Expression of mir-1294 is downregulated and predicts a poor prognosis in gastric cancer Y.-X. SHI, B.-L. YE, B.-R. HU, X.-J.
European Review for Medical and Pharmacological Sciences 2018; 22: 5525-5530 Expression of mir-1294 is downregulated and predicts a poor prognosis in gastric cancer Y.-X. SHI, B.-L. YE, B.-R. HU, X.-J.
Decreased expression of mir-490-3p in osteosarcoma and its clinical significance
 European Review for Medical and Pharmacological Sciences Decreased expression of mir-490-3p in osteosarcoma and its clinical significance B. TANG, C. LIU, Q.-M. ZHANG, M. NI Department of Orthopedics,
European Review for Medical and Pharmacological Sciences Decreased expression of mir-490-3p in osteosarcoma and its clinical significance B. TANG, C. LIU, Q.-M. ZHANG, M. NI Department of Orthopedics,
Original Article Tissue expression level of lncrna UCA1 is a prognostic biomarker for colorectal cancer
 Int J Clin Exp Pathol 2016;9(4):4241-4246 www.ijcep.com /ISSN:1936-2625/IJCEP0012296 Original Article Tissue expression level of lncrna UCA1 is a prognostic biomarker for colorectal cancer Hong Jiang,
Int J Clin Exp Pathol 2016;9(4):4241-4246 www.ijcep.com /ISSN:1936-2625/IJCEP0012296 Original Article Tissue expression level of lncrna UCA1 is a prognostic biomarker for colorectal cancer Hong Jiang,
Identifying Relevant micrornas in Bladder Cancer using Multi-Task Learning
 Identifying Relevant micrornas in Bladder Cancer using Multi-Task Learning Adriana Birlutiu (1),(2), Paul Bulzu (1), Irina Iereminciuc (1), and Alexandru Floares (1) (1) SAIA & OncoPredict, Cluj-Napoca,
Identifying Relevant micrornas in Bladder Cancer using Multi-Task Learning Adriana Birlutiu (1),(2), Paul Bulzu (1), Irina Iereminciuc (1), and Alexandru Floares (1) (1) SAIA & OncoPredict, Cluj-Napoca,
Bootstrapped Integrative Hypothesis Test, COPD-Lung Cancer Differentiation, and Joint mirnas Biomarkers
 Bootstrapped Integrative Hypothesis Test, COPD-Lung Cancer Differentiation, and Joint mirnas Biomarkers Kai-Ming Jiang 1,2, Bao-Liang Lu 1,2, and Lei Xu 1,2,3(&) 1 Department of Computer Science and Engineering,
Bootstrapped Integrative Hypothesis Test, COPD-Lung Cancer Differentiation, and Joint mirnas Biomarkers Kai-Ming Jiang 1,2, Bao-Liang Lu 1,2, and Lei Xu 1,2,3(&) 1 Department of Computer Science and Engineering,
Mir-595 is a significant indicator of poor patient prognosis in epithelial ovarian cancer
 European Review for Medical and Pharmacological Sciences 2017; 21: 4278-4282 Mir-595 is a significant indicator of poor patient prognosis in epithelial ovarian cancer Q.-H. ZHOU 1, Y.-M. ZHAO 2, L.-L.
European Review for Medical and Pharmacological Sciences 2017; 21: 4278-4282 Mir-595 is a significant indicator of poor patient prognosis in epithelial ovarian cancer Q.-H. ZHOU 1, Y.-M. ZHAO 2, L.-L.
RNA-Seq profiling of circular RNAs in human colorectal Cancer liver metastasis and the potential biomarkers
 Xu et al. Molecular Cancer (2019) 18:8 https://doi.org/10.1186/s12943-018-0932-8 LETTER TO THE EDITOR RNA-Seq profiling of circular RNAs in human colorectal Cancer liver metastasis and the potential biomarkers
Xu et al. Molecular Cancer (2019) 18:8 https://doi.org/10.1186/s12943-018-0932-8 LETTER TO THE EDITOR RNA-Seq profiling of circular RNAs in human colorectal Cancer liver metastasis and the potential biomarkers
Lack of association between an insertion/ deletion polymorphism in IL1A and risk of colorectal cancer
 Lack of association between an insertion/ deletion polymorphism in IL1A and risk of colorectal cancer H. Yan 1 *, R. Sun 2 *, X. Pan 3, Z. Li 4, X. Guo 5 and L. Gao 6 1 Department of Hepatobiliary Surgery,
Lack of association between an insertion/ deletion polymorphism in IL1A and risk of colorectal cancer H. Yan 1 *, R. Sun 2 *, X. Pan 3, Z. Li 4, X. Guo 5 and L. Gao 6 1 Department of Hepatobiliary Surgery,
Down-regulation of long non-coding RNA MEG3 serves as an unfavorable risk factor for survival of patients with breast cancer
 European Review for Medical and Pharmacological Sciences 2016; 20: 5143-5147 Down-regulation of long non-coding RNA MEG3 serves as an unfavorable risk factor for survival of patients with breast cancer
European Review for Medical and Pharmacological Sciences 2016; 20: 5143-5147 Down-regulation of long non-coding RNA MEG3 serves as an unfavorable risk factor for survival of patients with breast cancer
instrument. When 13C-UBT positive value is greater than or equal to / - 0.4, the the subject can be 1. Data and methods details are as follows:
 Application of 13 C-urea breath test in screening helicobacter pylori infection during health examination in Chengdu, Sichuan YANG Yan-hua. LIU Yu-ping. CHENG You-fu, SHUAI Ping. LU Qiao. ZHENG Xiao-xia,
Application of 13 C-urea breath test in screening helicobacter pylori infection during health examination in Chengdu, Sichuan YANG Yan-hua. LIU Yu-ping. CHENG You-fu, SHUAI Ping. LU Qiao. ZHENG Xiao-xia,
Overexpression of long-noncoding RNA ZFAS1 decreases survival in human NSCLC patients
 European Review for Medical and Pharmacological Sciences 2016; 20: 5126-5131 Overexpression of long-noncoding RNA ZFAS1 decreases survival in human NSCLC patients F.-M. TIAN 1, F.-Q. MENG 2, X.-B. WANG
European Review for Medical and Pharmacological Sciences 2016; 20: 5126-5131 Overexpression of long-noncoding RNA ZFAS1 decreases survival in human NSCLC patients F.-M. TIAN 1, F.-Q. MENG 2, X.-B. WANG
Prognostic significance of overexpressed long non-coding RNA TUG1 in patients with clear cell renal cell carcinoma
 European Review for Medical and Pharmacological Sciences 2017; 21: 82-86 Prognostic significance of overexpressed long non-coding RNA TUG1 in patients with clear cell renal cell carcinoma P.-Q. WANG, Y.-X.
European Review for Medical and Pharmacological Sciences 2017; 21: 82-86 Prognostic significance of overexpressed long non-coding RNA TUG1 in patients with clear cell renal cell carcinoma P.-Q. WANG, Y.-X.
Expression of long non-coding RNA linc-itgb1 in breast cancer and its influence on prognosis and survival
 European Review for Medical and Pharmacological Sciences 2017; 21: 3397-3401 Expression of long non-coding RNA linc-itgb1 in breast cancer and its influence on prognosis and survival W.-X. LI 1, R.-L.
European Review for Medical and Pharmacological Sciences 2017; 21: 3397-3401 Expression of long non-coding RNA linc-itgb1 in breast cancer and its influence on prognosis and survival W.-X. LI 1, R.-L.
The diagnostic value of determination of serum GOLPH3 associated with CA125, CA19.9 in patients with ovarian cancer
 European Review for Medical and Pharmacological Sciences 2017; 21: 4039-4044 The diagnostic value of determination of serum GOLPH3 associated with CA125, CA19.9 in patients with ovarian cancer H.-Y. FAN
European Review for Medical and Pharmacological Sciences 2017; 21: 4039-4044 The diagnostic value of determination of serum GOLPH3 associated with CA125, CA19.9 in patients with ovarian cancer H.-Y. FAN
Rapid Detection of Milk Protein based on Proteolysis Catalyzed by Trypsinase
 Rapid Detection of Milk Protein based on Proteolysis Catalyzed by Trypsinase Yafeng Chen Institute of Food Quality and Safety, University of Shanghai for Science and Technology Shanghai 93, China Email:cyfxy498@6.com
Rapid Detection of Milk Protein based on Proteolysis Catalyzed by Trypsinase Yafeng Chen Institute of Food Quality and Safety, University of Shanghai for Science and Technology Shanghai 93, China Email:cyfxy498@6.com
Correlation between estrogen receptor β expression and the curative effect of endocrine therapy in breast cancer patients
 1568 Correlation between estrogen receptor β expression and the curative effect of endocrine therapy in breast cancer patients LIYING GUO 1, YU ZHANG 2, WEI ZHANG 3 and DILIMINA YILAMU 1 1 Department of
1568 Correlation between estrogen receptor β expression and the curative effect of endocrine therapy in breast cancer patients LIYING GUO 1, YU ZHANG 2, WEI ZHANG 3 and DILIMINA YILAMU 1 1 Department of
Cancer incidence and patient survival rates among the residents in the Pudong New Area of Shanghai between 2002 and 2006
 Chinese Journal of Cancer Original Article Cancer incidence and patient survival rates among the residents in the Pudong New Area of Shanghai between 2002 and 2006 Xiao-Pan Li 1, Guang-Wen Cao 2, Qiao
Chinese Journal of Cancer Original Article Cancer incidence and patient survival rates among the residents in the Pudong New Area of Shanghai between 2002 and 2006 Xiao-Pan Li 1, Guang-Wen Cao 2, Qiao
MicroRNA expression profiling and functional analysis in prostate cancer. Marco Folini s.c. Ricerca Traslazionale DOSL
 MicroRNA expression profiling and functional analysis in prostate cancer Marco Folini s.c. Ricerca Traslazionale DOSL What are micrornas? For almost three decades, the alteration of protein-coding genes
MicroRNA expression profiling and functional analysis in prostate cancer Marco Folini s.c. Ricerca Traslazionale DOSL What are micrornas? For almost three decades, the alteration of protein-coding genes
EMPEROR'S COLLEGE MTOM COURSE SYLLABUS HERB FORMULAE I
 COURSE DESCRIPTION The first of three courses in the Herb Formulae series. These courses can be taken in any order. The Herb Formulae series analyzes the functions, ingredients, and properties of approximately
COURSE DESCRIPTION The first of three courses in the Herb Formulae series. These courses can be taken in any order. The Herb Formulae series analyzes the functions, ingredients, and properties of approximately
Expression of lncrna TCONS_ in hepatocellular carcinoma and its influence on prognosis and survival
 European Review for Medical and Pharmacological Sciences 2017; 21: 5655-5660 Expression of lncrna TCONS_00027978 in hepatocellular carcinoma and its influence on prognosis and survival Q. CHEN 1, G.-D.
European Review for Medical and Pharmacological Sciences 2017; 21: 5655-5660 Expression of lncrna TCONS_00027978 in hepatocellular carcinoma and its influence on prognosis and survival Q. CHEN 1, G.-D.
Peking University People's Hospital, Peking University Institute of Hematology
 Qian Jiang, M.D. Peking University People's Hospital, Peking University Institute of Hematology No. 11 Xizhimen South Street, Beijing, 100044, China. Phone number: 86-10-66583802 Mobile: 86-13611115100
Qian Jiang, M.D. Peking University People's Hospital, Peking University Institute of Hematology No. 11 Xizhimen South Street, Beijing, 100044, China. Phone number: 86-10-66583802 Mobile: 86-13611115100
mirna Dr. S Hosseini-Asl
 mirna Dr. S Hosseini-Asl 1 2 MicroRNAs (mirnas) are small noncoding RNAs which enhance the cleavage or translational repression of specific mrna with recognition site(s) in the 3 - untranslated region
mirna Dr. S Hosseini-Asl 1 2 MicroRNAs (mirnas) are small noncoding RNAs which enhance the cleavage or translational repression of specific mrna with recognition site(s) in the 3 - untranslated region
GUOPEI LUO, PhD, MD. Shanghai Cancer Center, Shanghai, China. Joint-trained Ph.D. student, from 2008 to 2009, University of California, Irvine
 GUOPEI LUO, PhD, MD EDUCATION Shanghai Cancer Center, Shanghai, China Joint-trained Ph.D. student, from 2008 to 2009, University of California, Irvine Ph.D. student specialized in pancreatic cancer, from
GUOPEI LUO, PhD, MD EDUCATION Shanghai Cancer Center, Shanghai, China Joint-trained Ph.D. student, from 2008 to 2009, University of California, Irvine Ph.D. student specialized in pancreatic cancer, from
Researches on Fermentation Engineering of Polysaccharide of
 13 1 Vol13 No1 1 2009 2 Life Science Research Feb 2009 1a 1b, 2, 1 416000 2 416000 3 410300 : (Cordyceps militaris), :, 6% 1% 25, ; 6% 1% 22, : ; ; ; ; ; : TQ92 : A : 1007-7847(2009)01-0065-06 Researches
13 1 Vol13 No1 1 2009 2 Life Science Research Feb 2009 1a 1b, 2, 1 416000 2 416000 3 410300 : (Cordyceps militaris), :, 6% 1% 25, ; 6% 1% 22, : ; ; ; ; ; : TQ92 : A : 1007-7847(2009)01-0065-06 Researches
Liu Jing and Liu Jing Diagnosis System in Classical TCM Discussions of Six Divisions or Six Confirmations Diagnosis System in Classical TCM Texts
 Liu Jing and Liu Jing Diagnosis System in Classical TCM Discussions of Six Divisions or Six Confirmations Diagnosis System in Classical TCM Texts Liu Jing Bian Zheng system had developed about 1800 years
Liu Jing and Liu Jing Diagnosis System in Classical TCM Discussions of Six Divisions or Six Confirmations Diagnosis System in Classical TCM Texts Liu Jing Bian Zheng system had developed about 1800 years
Clinicopathologic and prognostic relevance of mir-1256 in colorectal cancer: a preliminary clinical study
 European Review for Medical and Pharmacological Sciences 2018; 22: 7704-7709 Clinicopathologic and prognostic relevance of mir-1256 in colorectal cancer: a preliminary clinical study Z.-Y. LIU, L. YANG,
European Review for Medical and Pharmacological Sciences 2018; 22: 7704-7709 Clinicopathologic and prognostic relevance of mir-1256 in colorectal cancer: a preliminary clinical study Z.-Y. LIU, L. YANG,
Original Article CREPT expression correlates with esophageal squamous cell carcinoma histological grade and clinical outcome
 Int J Clin Exp Pathol 2017;10(2):2030-2035 www.ijcep.com /ISSN:1936-2625/IJCEP0009456 Original Article CREPT expression correlates with esophageal squamous cell carcinoma histological grade and clinical
Int J Clin Exp Pathol 2017;10(2):2030-2035 www.ijcep.com /ISSN:1936-2625/IJCEP0009456 Original Article CREPT expression correlates with esophageal squamous cell carcinoma histological grade and clinical
The association between methylenetetrahydrofolate reductase gene C677T polymorphisms and breast cancer risk in Chinese population
 Tumor Biol. (2015) 36:9153 9158 DOI 10.1007/s13277-015-3321-6 EDITORIAL The association between methylenetetrahydrofolate reductase gene C677T polymorphisms and breast cancer risk in Chinese population
Tumor Biol. (2015) 36:9153 9158 DOI 10.1007/s13277-015-3321-6 EDITORIAL The association between methylenetetrahydrofolate reductase gene C677T polymorphisms and breast cancer risk in Chinese population
CORRELATION BETWEEN SURVIVIN OVEREXPRESSION AND CLINICO-PATHOLOGICAL FEATURES OF INVASIVE CERVICAL CANCER: A META-ANALYSIS
 Acta Medica Mediterranea, 2018, 34: 1091 CORRELATION BETWEEN SURVIVIN OVEREXPRESSION AND CLINICO-PATHOLOGICAL FEATURES OF INVASIVE CERVICAL CANCER: A META-ANALYSIS HUI-RONG LI 1, JI-LI BAI 2*, RUI XU 2,
Acta Medica Mediterranea, 2018, 34: 1091 CORRELATION BETWEEN SURVIVIN OVEREXPRESSION AND CLINICO-PATHOLOGICAL FEATURES OF INVASIVE CERVICAL CANCER: A META-ANALYSIS HUI-RONG LI 1, JI-LI BAI 2*, RUI XU 2,
POSITION TITLE Professor. DEGREE (if applicable)
 NAME Wang, Tong era COMMONS USER NAME TOWANG BIOGRAPHICAL SKETCH Provide the following information for the key personnel and other significant contributors in the order listed on Form Page 2. Follow this
NAME Wang, Tong era COMMONS USER NAME TOWANG BIOGRAPHICAL SKETCH Provide the following information for the key personnel and other significant contributors in the order listed on Form Page 2. Follow this
Inferring potential small molecule mirna association based on triple layer heterogeneous network
 https://doi.org/10.1186/s13321-018-0284-9 RESEARCH ARTICLE Open Access Inferring potential small molecule mirna association based on triple layer heterogeneous network Jia Qu 1, Xing Chen 1*, Ya Zhou Sun
https://doi.org/10.1186/s13321-018-0284-9 RESEARCH ARTICLE Open Access Inferring potential small molecule mirna association based on triple layer heterogeneous network Jia Qu 1, Xing Chen 1*, Ya Zhou Sun
Root & Branch Bulk Formula List
 An asterisk * indicates the inclusion of 1 or more granule versions of an herb because of limited availability on the American herbal market. These products are usually animal in nature like E Jiao, Shui
An asterisk * indicates the inclusion of 1 or more granule versions of an herb because of limited availability on the American herbal market. These products are usually animal in nature like E Jiao, Shui
Long noncoding RNA CASC2 inhibits metastasis and epithelial to mesenchymal transition of lung adenocarcinoma via suppressing SOX4
 European Review for Medical and Pharmacological Sciences 2017; 21: 4584-4590 Long noncoding RNA CASC2 inhibits metastasis and epithelial to mesenchymal transition of lung adenocarcinoma via suppressing
European Review for Medical and Pharmacological Sciences 2017; 21: 4584-4590 Long noncoding RNA CASC2 inhibits metastasis and epithelial to mesenchymal transition of lung adenocarcinoma via suppressing
Diagnostic performance of microrna-29a for colorectal cancer: a meta-analysis
 Diagnostic performance of microrna-29a for colorectal cancer: a meta-analysis M.L. Zhi 1 *, Z.J. Liu 2 *, X.Y. Yi 1, L.J. Zhang 1 and Y.X. Bao 1 1 Department of Clinical Laboratory, The Second Affiliated
Diagnostic performance of microrna-29a for colorectal cancer: a meta-analysis M.L. Zhi 1 *, Z.J. Liu 2 *, X.Y. Yi 1, L.J. Zhang 1 and Y.X. Bao 1 1 Department of Clinical Laboratory, The Second Affiliated
Epstein-Barr virus driven promoter hypermethylated genes in gastric cancer
 RESEARCH FUND FOR THE CONTROL OF INFECTIOUS DISEASES Epstein-Barr virus driven promoter hypermethylated genes in gastric cancer J Yu *, KF To, QY Liang K e y M e s s a g e s 1. Somatostatin receptor 1
RESEARCH FUND FOR THE CONTROL OF INFECTIOUS DISEASES Epstein-Barr virus driven promoter hypermethylated genes in gastric cancer J Yu *, KF To, QY Liang K e y M e s s a g e s 1. Somatostatin receptor 1
38 Int'l Conf. Bioinformatics and Computational Biology BIOCOMP'16
 38 Int'l Conf. Bioinformatics and Computational Biology BIOCOMP'16 PGAR: ASD Candidate Gene Prioritization System Using Expression Patterns Steven Cogill and Liangjiang Wang Department of Genetics and
38 Int'l Conf. Bioinformatics and Computational Biology BIOCOMP'16 PGAR: ASD Candidate Gene Prioritization System Using Expression Patterns Steven Cogill and Liangjiang Wang Department of Genetics and
Lack of association between ERCC5 gene polymorphisms and gastric cancer risk in a Chinese population
 Lack of association between ERCC5 gene polymorphisms and gastric cancer risk in a Chinese population J.J. Lu, H.Q. Zhang, P. Mai, X. Ma, X. Chen, Y.X. Yang and L.P. Zhang Gansu Provincial Hospital, Donggang
Lack of association between ERCC5 gene polymorphisms and gastric cancer risk in a Chinese population J.J. Lu, H.Q. Zhang, P. Mai, X. Ma, X. Chen, Y.X. Yang and L.P. Zhang Gansu Provincial Hospital, Donggang
Bayesian Bi-Cluster Change-Point Model for Exploring Functional Brain Dynamics
 Int'l Conf. Bioinformatics and Computational Biology BIOCOMP'18 85 Bayesian Bi-Cluster Change-Point Model for Exploring Functional Brain Dynamics Bing Liu 1*, Xuan Guo 2, and Jing Zhang 1** 1 Department
Int'l Conf. Bioinformatics and Computational Biology BIOCOMP'18 85 Bayesian Bi-Cluster Change-Point Model for Exploring Functional Brain Dynamics Bing Liu 1*, Xuan Guo 2, and Jing Zhang 1** 1 Department
Human leukocyte antigen-b27 alleles in Xinjiang Uygur patients with ankylosing spondylitis
 Human leukocyte antigen-b27 alleles in Xinjiang Uygur patients with ankylosing spondylitis H.-Y. Zou, W.-Z. Yu, Z. Wang, J. He and M. Jiao Institute of Clinical Medicine, Urumqi General Hospital, Lanzhou
Human leukocyte antigen-b27 alleles in Xinjiang Uygur patients with ankylosing spondylitis H.-Y. Zou, W.-Z. Yu, Z. Wang, J. He and M. Jiao Institute of Clinical Medicine, Urumqi General Hospital, Lanzhou
Supplementary Information
 Supplementary Information Table S1. List of mirna gene polymorphisms associated with cancer. mirna Name rs Number Cancer type Effect Ethnicity Study type No. of cases/controls Reference hsa-mir-27a rs11671784
Supplementary Information Table S1. List of mirna gene polymorphisms associated with cancer. mirna Name rs Number Cancer type Effect Ethnicity Study type No. of cases/controls Reference hsa-mir-27a rs11671784
Review Article MicroRNA single-nucleotide polymorphisms and susceptibility to gastric cancer
 Am J Digest Dis 2016;3(1):11-15 www.ajdd.us /ISSN:2329-6992/AJDD0023804 Review Article MicroRNA single-nucleotide polymorphisms and susceptibility to gastric cancer Xinhang Jiang, Xintong Chen, Linhua
Am J Digest Dis 2016;3(1):11-15 www.ajdd.us /ISSN:2329-6992/AJDD0023804 Review Article MicroRNA single-nucleotide polymorphisms and susceptibility to gastric cancer Xinhang Jiang, Xintong Chen, Linhua
Review Article Elevated expression of long noncoding RNA UCA1 can predict a poor prognosis: a meta-analysis
 Int J Clin Exp Med 2016;9(6):9656-9664 www.ijcem.com /ISSN:1940-5901/IJCEM0025465 Review Article Elevated expression of long noncoding RNA UCA1 can predict a poor prognosis: a meta-analysis Tao Chen 1*,
Int J Clin Exp Med 2016;9(6):9656-9664 www.ijcem.com /ISSN:1940-5901/IJCEM0025465 Review Article Elevated expression of long noncoding RNA UCA1 can predict a poor prognosis: a meta-analysis Tao Chen 1*,
Nasopharyngeal carcinoma incidence and mortality in China in 2010
 Chinese Journal of Cancer Original Article Nasopharyngeal carcinoma incidence and mortality in China in 2010 Kuang-Rong Wei 1, Rong-Shou Zheng 2, Si-Wei Zhang 2, Zhi-Heng Liang 1, Zhi-Xiong Ou 1 and Wan-Qing
Chinese Journal of Cancer Original Article Nasopharyngeal carcinoma incidence and mortality in China in 2010 Kuang-Rong Wei 1, Rong-Shou Zheng 2, Si-Wei Zhang 2, Zhi-Heng Liang 1, Zhi-Xiong Ou 1 and Wan-Qing
Original Article Association of osteopontin polymorphism with cancer risk: a meta-analysis
 Int J Clin Exp Med 2015;8(11):20911-20917 www.ijcem.com /ISSN:1940-5901/IJCEM0015120 Original Article Association of osteopontin polymorphism with cancer risk: a meta-analysis Gang Yang 1, Xiaoxing Peng
Int J Clin Exp Med 2015;8(11):20911-20917 www.ijcem.com /ISSN:1940-5901/IJCEM0015120 Original Article Association of osteopontin polymorphism with cancer risk: a meta-analysis Gang Yang 1, Xiaoxing Peng
Evaluation of altered expression of mir-9 and mir-106a as an early diagnostic approach in gastric cancer
 Original Article Evaluation of altered expression of mir-9 and mir-106a as an early diagnostic approach in gastric cancer Khadije Shirmohammadi, Sareh Sohrabi, Zahra Jafarzadeh Samani, Hosein Effatpanah,
Original Article Evaluation of altered expression of mir-9 and mir-106a as an early diagnostic approach in gastric cancer Khadije Shirmohammadi, Sareh Sohrabi, Zahra Jafarzadeh Samani, Hosein Effatpanah,
MicroRNA and Male Infertility: A Potential for Diagnosis
 Review Article MicroRNA and Male Infertility: A Potential for Diagnosis * Abstract MicroRNAs (mirnas) are small non-coding single stranded RNA molecules that are physiologically produced in eukaryotic
Review Article MicroRNA and Male Infertility: A Potential for Diagnosis * Abstract MicroRNAs (mirnas) are small non-coding single stranded RNA molecules that are physiologically produced in eukaryotic
Prostate Disease A Pass By TANG QIAN LI ZHU BIAN
 Prostate Disease A Pass By TANG QIAN LI ZHU BIAN Oxytocin and the Human Prostate in Health and - and PSA is then able to pass into the proliferation within the normal prostate. 4.4. Oxytocin and prostate
Prostate Disease A Pass By TANG QIAN LI ZHU BIAN Oxytocin and the Human Prostate in Health and - and PSA is then able to pass into the proliferation within the normal prostate. 4.4. Oxytocin and prostate
Cancer Cell Research 17 (2018)
 Available at http:// www.cancercellresearch.org ISSN 2161-2609 Prognostic value of MicroRNA-21 in non-small cell lung cancer: a meta-analysis Fei Xu, Leina Ma, Lingxin Feng, Yinjie Wu, Xiaoyan Xu, Zhuang
Available at http:// www.cancercellresearch.org ISSN 2161-2609 Prognostic value of MicroRNA-21 in non-small cell lung cancer: a meta-analysis Fei Xu, Leina Ma, Lingxin Feng, Yinjie Wu, Xiaoyan Xu, Zhuang
Original Article Reduced expression of mir-506 in glioma is associated with advanced tumor progression and unfavorable prognosis
 Int J Clin Exp Pathol 2016;9(1):181-185 www.ijcep.com /ISSN:1936-2625/IJCEP0017823 Original Article Reduced expression of mir-506 in glioma is associated with advanced tumor progression and unfavorable
Int J Clin Exp Pathol 2016;9(1):181-185 www.ijcep.com /ISSN:1936-2625/IJCEP0017823 Original Article Reduced expression of mir-506 in glioma is associated with advanced tumor progression and unfavorable
micrornas (mirna) and Biomarkers
 micrornas (mirna) and Biomarkers Small RNAs Make Big Splash mirnas & Genome Function Biomarkers in Cancer Future Prospects Javed Khan M.D. National Cancer Institute EORTC-NCI-ASCO November 2007 The Human
micrornas (mirna) and Biomarkers Small RNAs Make Big Splash mirnas & Genome Function Biomarkers in Cancer Future Prospects Javed Khan M.D. National Cancer Institute EORTC-NCI-ASCO November 2007 The Human
Low levels of serum mir-99a is a predictor of poor prognosis in breast cancer
 Low levels of serum mir-99a is a predictor of poor prognosis in breast cancer J. Li 1, Z.J. Song 2, Y.Y. Wang 1, Y. Yin 1, Y. Liu 1 and X. Nan 1 1 Tumor Research Department, Shaanxi Provincial Tumor Hospital,
Low levels of serum mir-99a is a predictor of poor prognosis in breast cancer J. Li 1, Z.J. Song 2, Y.Y. Wang 1, Y. Yin 1, Y. Liu 1 and X. Nan 1 1 Tumor Research Department, Shaanxi Provincial Tumor Hospital,
Integrative network analysis: Bridging the gap between Western medicine and traditional Chinese medicine
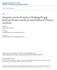 Hong Kong Baptist University HKBU Institutional Repository HKBU Staff Publication 2015 Integrative network analysis: Bridging the gap between Western medicine and traditional Chinese medicine Bing He Hong
Hong Kong Baptist University HKBU Institutional Repository HKBU Staff Publication 2015 Integrative network analysis: Bridging the gap between Western medicine and traditional Chinese medicine Bing He Hong
Up-regulation of serum mir-4262 predicts clinical outcome of patients with acute myeloid leukemia
 European Review for Medical and Pharmacological Sciences 2017; 21: 2172-2176 Up-regulation of serum mir-4262 predicts clinical outcome of patients with acute myeloid leukemia G.-T. HAN 1, Z.-L. SUN 2 1
European Review for Medical and Pharmacological Sciences 2017; 21: 2172-2176 Up-regulation of serum mir-4262 predicts clinical outcome of patients with acute myeloid leukemia G.-T. HAN 1, Z.-L. SUN 2 1
Yi-Long Yang 1, Guo-Yuan Sui 1, Guang-Cong Liu 2, De-Sheng Huang 3, Si-Meng Wang 4 and Lie Wang 1*
 Yang et al. BMC Cancer 2014, 14:956 RESEARCH ARTICLE Open Access The effects of psychological interventions on depression and anxiety among Chinese adults with cancer: a meta-analysis of randomized controlled
Yang et al. BMC Cancer 2014, 14:956 RESEARCH ARTICLE Open Access The effects of psychological interventions on depression and anxiety among Chinese adults with cancer: a meta-analysis of randomized controlled
The updated incidences and mortalities of major cancers in China, 2011
 DOI 10.1186/s40880-015-0042-6 REVIEW Open Access The updated incidences and mortalities of major cancers in China, 2011 Wanqing Chen *, Rongshou Zheng, Hongmei Zeng and Siwei Zhang Abstract Introduction:
DOI 10.1186/s40880-015-0042-6 REVIEW Open Access The updated incidences and mortalities of major cancers in China, 2011 Wanqing Chen *, Rongshou Zheng, Hongmei Zeng and Siwei Zhang Abstract Introduction:
Predicting Breast Cancer Survivability Rates
 Predicting Breast Cancer Survivability Rates For data collected from Saudi Arabia Registries Ghofran Othoum 1 and Wadee Al-Halabi 2 1 Computer Science, Effat University, Jeddah, Saudi Arabia 2 Computer
Predicting Breast Cancer Survivability Rates For data collected from Saudi Arabia Registries Ghofran Othoum 1 and Wadee Al-Halabi 2 1 Computer Science, Effat University, Jeddah, Saudi Arabia 2 Computer
Investigating the role of polymorphisms in mir-146a, -149, and -196a2 in the development of gastric cancer
 Investigating the role of polymorphisms in mir-146a, -149, and -196a2 in the development of gastric cancer Department of Gastrointestinal Surgery, Ren Ji Hospital, School of Medicine, Shanghai Jiao Tong
Investigating the role of polymorphisms in mir-146a, -149, and -196a2 in the development of gastric cancer Department of Gastrointestinal Surgery, Ren Ji Hospital, School of Medicine, Shanghai Jiao Tong
Canqiu Yu 1, Jinwei Chen 2, Li Huang 3*
 A STUDY ON THE ANTITUMOUR EFFECT OF TOTAL FLAVONOIDS FROM PTERIS MULTIFIDA POIR IN H22 TUMOUR-BEARING MICE 459 Canqiu Yu 1, Jinwei Chen 2, Li Huang 3* 1 Department of General Surgery, The Second Xiangya
A STUDY ON THE ANTITUMOUR EFFECT OF TOTAL FLAVONOIDS FROM PTERIS MULTIFIDA POIR IN H22 TUMOUR-BEARING MICE 459 Canqiu Yu 1, Jinwei Chen 2, Li Huang 3* 1 Department of General Surgery, The Second Xiangya
Appropriate Quality regarde as
 A systematic review of antibiotic prescription associated with upper respiratory tract infections in China (Jing Li, MS 1 ) Supplemental Digital Content-Table 1. Quality assessment of included studies
A systematic review of antibiotic prescription associated with upper respiratory tract infections in China (Jing Li, MS 1 ) Supplemental Digital Content-Table 1. Quality assessment of included studies
Profiles of gene expression & diagnosis/prognosis of cancer. MCs in Advanced Genetics Ainoa Planas Riverola
 Profiles of gene expression & diagnosis/prognosis of cancer MCs in Advanced Genetics Ainoa Planas Riverola Gene expression profiles Gene expression profiling Used in molecular biology, it measures the
Profiles of gene expression & diagnosis/prognosis of cancer MCs in Advanced Genetics Ainoa Planas Riverola Gene expression profiles Gene expression profiling Used in molecular biology, it measures the
Comparison of three mathematical prediction models in patients with a solitary pulmonary nodule
 Original Article Comparison of three mathematical prediction models in patients with a solitary pulmonary nodule Xuan Zhang*, Hong-Hong Yan, Jun-Tao Lin, Ze-Hua Wu, Jia Liu, Xu-Wei Cao, Xue-Ning Yang From
Original Article Comparison of three mathematical prediction models in patients with a solitary pulmonary nodule Xuan Zhang*, Hong-Hong Yan, Jun-Tao Lin, Ze-Hua Wu, Jia Liu, Xu-Wei Cao, Xue-Ning Yang From
Expression level of microrna-195 in the serum of patients with gastric cancer and its relationship with the clinicopathological staging of the cancer
 European Review for Medical and Pharmacological Sciences 2016; 20: 1283-1287 Expression level of microrna-195 in the serum of patients with gastric cancer and its relationship with the clinicopathological
European Review for Medical and Pharmacological Sciences 2016; 20: 1283-1287 Expression level of microrna-195 in the serum of patients with gastric cancer and its relationship with the clinicopathological
Clinical impact of serum mir-661 in diagnosis and prognosis of non-small cell lung cancer
 European Review for Medical and Pharmacological Sciences 2017; 21: 5696-5701 Clinical impact of serum mir-661 in diagnosis and prognosis of non-small cell lung cancer G.-H. ZHOU 1, W.-H. YANG 2, B. SUN
European Review for Medical and Pharmacological Sciences 2017; 21: 5696-5701 Clinical impact of serum mir-661 in diagnosis and prognosis of non-small cell lung cancer G.-H. ZHOU 1, W.-H. YANG 2, B. SUN
Open Access Fuzzy Analytic Hierarchy Process-based Chinese Resident Best Fitness Behavior Method Research
 Send Orders for Reprints to reprints@benthamscience.ae The Open Biomedical Engineering Journal, 0, 9, - Open Access Fuzzy Analytic Hierarchy Process-based Chinese Resident Best Fitness Behavior Method
Send Orders for Reprints to reprints@benthamscience.ae The Open Biomedical Engineering Journal, 0, 9, - Open Access Fuzzy Analytic Hierarchy Process-based Chinese Resident Best Fitness Behavior Method
A single nucleotide polymorphism in the promoter region of let-7 family is associated with lung cancer risk in Chinese
 A single nucleotide polymorphism in the promoter region of let-7 family is associated with lung cancer risk in Chinese L.Q. Shen 1 *, Y.Z. Xie 1 *, X.F. Qian 2 *, Z.X. Zhuang 1, Y. Jiao 3 and X.F. Qi 4,5
A single nucleotide polymorphism in the promoter region of let-7 family is associated with lung cancer risk in Chinese L.Q. Shen 1 *, Y.Z. Xie 1 *, X.F. Qian 2 *, Z.X. Zhuang 1, Y. Jiao 3 and X.F. Qi 4,5
Association between downexpression of mir-1301 and poor prognosis in patients with glioma
 European Review for Medical and Pharmacological Sciences 2017; 21: 4298-4303 Association between downexpression of mir-1301 and poor prognosis in patients with glioma Q.-L. BAI, C.-W. HU, X.-R. WANG, G.-F.
European Review for Medical and Pharmacological Sciences 2017; 21: 4298-4303 Association between downexpression of mir-1301 and poor prognosis in patients with glioma Q.-L. BAI, C.-W. HU, X.-R. WANG, G.-F.
Original Article High serum mir-203 predicts the poor prognosis in patients with pancreatic cancer
 Int J Clin Exp Pathol 2017;10(4):4688-4693 www.ijcep.com /ISSN:1936-2625/IJCEP0047128 Original Article High serum mir-203 predicts the poor prognosis in patients with pancreatic cancer Jun Ma, Xiaokun
Int J Clin Exp Pathol 2017;10(4):4688-4693 www.ijcep.com /ISSN:1936-2625/IJCEP0047128 Original Article High serum mir-203 predicts the poor prognosis in patients with pancreatic cancer Jun Ma, Xiaokun
mir-218 tissue expression level is associated with aggressive progression of gastric cancer
 mir-218 tissue expression level is associated with aggressive progression of gastric cancer X.X. Wang 1, S.J. Ge 2, X.L. Wang 2, L.X. Jiang 1, M.F. Sheng 2 and J.J. Ma 2 1 Department of General Surgery,
mir-218 tissue expression level is associated with aggressive progression of gastric cancer X.X. Wang 1, S.J. Ge 2, X.L. Wang 2, L.X. Jiang 1, M.F. Sheng 2 and J.J. Ma 2 1 Department of General Surgery,
Increased long noncoding RNA LINP1 expression and its prognostic significance in human breast cancer
 European Review for Medical and Pharmacological Sciences 2018; 22: 8749-8754 Increased long noncoding RNA LINP1 expression and its prognostic significance in human breast cancer X.-M. LIU 1, B. YANG 2,
European Review for Medical and Pharmacological Sciences 2018; 22: 8749-8754 Increased long noncoding RNA LINP1 expression and its prognostic significance in human breast cancer X.-M. LIU 1, B. YANG 2,
Random walks on mutual microrna-target gene interaction network improve the prediction of disease-associated micrornas
 Le et al. BMC Bioinformatics (2017) 18:479 DOI 10.1186/s12859-017-1924-1 RESEARCH ARTICLE Random walks on mutual microrna-target gene interaction network improve the prediction of disease-associated micrornas
Le et al. BMC Bioinformatics (2017) 18:479 DOI 10.1186/s12859-017-1924-1 RESEARCH ARTICLE Random walks on mutual microrna-target gene interaction network improve the prediction of disease-associated micrornas
Influence of interleukin-18 gene polymorphisms on acute pancreatitis susceptibility in a Chinese population
 Influence of interleukin-18 gene polymorphisms on acute pancreatitis susceptibility in a Chinese population H.B. Gui 1, X.G. Du 2, Z.H. Fu 3 and X.M. Chen 1 1 Department of Emergency, The First Affiliated
Influence of interleukin-18 gene polymorphisms on acute pancreatitis susceptibility in a Chinese population H.B. Gui 1, X.G. Du 2, Z.H. Fu 3 and X.M. Chen 1 1 Department of Emergency, The First Affiliated
Effects of estradiol and progesterone on the growth of HeLa cervical cancer cells
 European Review for Medical harmacological Sciences 2017; 21: 3959-3965 Effects of estradiol and progesterone on the growth of HeLa cervical cancer cells Y. LIU 1, L.-B. TIAN 2, H.-Y. YANG 1, H.-P. ZHANG
European Review for Medical harmacological Sciences 2017; 21: 3959-3965 Effects of estradiol and progesterone on the growth of HeLa cervical cancer cells Y. LIU 1, L.-B. TIAN 2, H.-Y. YANG 1, H.-P. ZHANG
Title:Identification of a novel microrna signature associated with intrahepatic cholangiocarcinoma (ICC) patient prognosis
 Author's response to reviews Title:Identification of a novel microrna signature associated with intrahepatic cholangiocarcinoma (ICC) patient prognosis Authors: Mei-Yin Zhang (zhangmy@sysucc.org.cn) Shu-Hong
Author's response to reviews Title:Identification of a novel microrna signature associated with intrahepatic cholangiocarcinoma (ICC) patient prognosis Authors: Mei-Yin Zhang (zhangmy@sysucc.org.cn) Shu-Hong
Cancer Cell Research 19 (2018)
 Available at http:// www.cancercellresearch.org ISSN 2161-2609 LncRNA RP11-597D13.9 expression and clinical significance in serous Ovarian Cancer based on TCGA database Xiaoyan Gu 1, Pengfei Xu 2, Sujuan
Available at http:// www.cancercellresearch.org ISSN 2161-2609 LncRNA RP11-597D13.9 expression and clinical significance in serous Ovarian Cancer based on TCGA database Xiaoyan Gu 1, Pengfei Xu 2, Sujuan
Downregulation of long non-coding RNA LINC01133 is predictive of poor prognosis in colorectal cancer patients
 European Review for Medical and Pharmacological Sciences 2017; 21: 2103-2107 Downregulation of long non-coding RNA LINC01133 is predictive of poor prognosis in colorectal cancer patients J.-H. ZHANG 1,
European Review for Medical and Pharmacological Sciences 2017; 21: 2103-2107 Downregulation of long non-coding RNA LINC01133 is predictive of poor prognosis in colorectal cancer patients J.-H. ZHANG 1,
Expression and significance of Bmi-1 and Ki67 in colorectal carcinoma tissues
 [Chinese Journal of Cancer 27:12, 568-573; December Expression 2008]; 2008 and significance Sun Yat-sen of University Bmi-1 and Cancer Ki67 in Center colorectal carcinoma tissues Clinical Research Paper
[Chinese Journal of Cancer 27:12, 568-573; December Expression 2008]; 2008 and significance Sun Yat-sen of University Bmi-1 and Cancer Ki67 in Center colorectal carcinoma tissues Clinical Research Paper
Seasonal and geographical dispersal regularity of airborne pollens in China LI Quan-sheng, JIANG Sheng-xue, LI Xin-ze, ZHU Xiao-ming, WEI Qing-yu *
 Med J Chin PLA, Vol. 42, No. 11, November 1, 2017 951 [] [ ] [ ] R562.25[ ] A[] 0177-7402(2017)11-0951-05 [DOI] 10.11855/j.issn.0577-7402.2017.11.03 Seasonal and geographical dispersal regularity of airborne
Med J Chin PLA, Vol. 42, No. 11, November 1, 2017 951 [] [ ] [ ] R562.25[ ] A[] 0177-7402(2017)11-0951-05 [DOI] 10.11855/j.issn.0577-7402.2017.11.03 Seasonal and geographical dispersal regularity of airborne
Long non-coding RNA TUSC7 expression is independently predictive of outcome in glioma
 European Review for Medical and Pharmacological Sciences 2017; 21: 3605-3610 Long non-coding RNA TUSC7 expression is independently predictive of outcome in glioma X.-L. MA 1, W.-D. ZHU 2, L.-X. TIAN 2,
European Review for Medical and Pharmacological Sciences 2017; 21: 3605-3610 Long non-coding RNA TUSC7 expression is independently predictive of outcome in glioma X.-L. MA 1, W.-D. ZHU 2, L.-X. TIAN 2,
High expression of fibroblast activation protein is an adverse prognosticator in gastric cancer.
 Biomedical Research 2017; 28 (18): 7779-7783 ISSN 0970-938X www.biomedres.info High expression of fibroblast activation protein is an adverse prognosticator in gastric cancer. Hu Song 1, Qi-yu Liu 2, Zhi-wei
Biomedical Research 2017; 28 (18): 7779-7783 ISSN 0970-938X www.biomedres.info High expression of fibroblast activation protein is an adverse prognosticator in gastric cancer. Hu Song 1, Qi-yu Liu 2, Zhi-wei
Single Herbs III / Quiz I
 Single Herbs III / Quiz I 1. What herb is good to generate fluids? A. Ren Shen C. Xi Yang Shen B. Tai Zi Shen D. All the Shens 2. What herb is best for Qi collapse? A. Huang Qi C. Dang Shen B. Ren Shen
Single Herbs III / Quiz I 1. What herb is good to generate fluids? A. Ren Shen C. Xi Yang Shen B. Tai Zi Shen D. All the Shens 2. What herb is best for Qi collapse? A. Huang Qi C. Dang Shen B. Ren Shen
Discovering Symptom-herb Relationship by Exploiting SHT Topic Model
 [DOI: 10.2197/ipsjtbio.10.16] Original Paper Discovering Symptom-herb Relationship by Exploiting SHT Topic Model Lidong Wang 1,a) Keyong Hu 1 Xiaodong Xu 2 Received: July 7, 2017, Accepted: August 29,
[DOI: 10.2197/ipsjtbio.10.16] Original Paper Discovering Symptom-herb Relationship by Exploiting SHT Topic Model Lidong Wang 1,a) Keyong Hu 1 Xiaodong Xu 2 Received: July 7, 2017, Accepted: August 29,
Original Article MicroRNA-101 is a novel biomarker for diagnosis and prognosis in breast cancer
 Int J Clin Exp Pathol 2017;10(3):3279-3285 www.ijcep.com /ISSN:1936-2625/IJCEP0022926 Original Article MicroRNA-101 is a novel biomarker for diagnosis and prognosis in breast cancer Na Wei, Qing Ni, Min
Int J Clin Exp Pathol 2017;10(3):3279-3285 www.ijcep.com /ISSN:1936-2625/IJCEP0022926 Original Article MicroRNA-101 is a novel biomarker for diagnosis and prognosis in breast cancer Na Wei, Qing Ni, Min
