Supporting Information. to the article. Conical lipids in flat bilayers induce packing defects similar to that induced by positive curvature
|
|
|
- Barrie Robert Ray
- 5 years ago
- Views:
Transcription
1 Supporting Information to the article Conical lipids in flat bilayers induce packing defects similar to that induced by positive curvature Lydie Vamparys 1,2,3, Romain Gautier 4, Stefano Vanni 4, W.F. Drew Bennett 5, D. Peter Tieleman 5, Bruno Antonny 4, Catherine Etchebest 1,2,3, and Patrick F.J. Fuchs 1,2,3 1 INSERM, U665, F Paris, France 2 Univ Paris Diderot, Sorbonne Paris Cité, UMR_S 665, F Paris, France 3 Institut National de la Transfusion Sanguine, F Paris, France 4 IPMC, UMR 7275 CNRS, Université de Nice Sophia Antipolis, F-06560, Valbonne, France 5 Department of Biological Sciences, and Institute for Biocomplexity and Informatics, University of Calgary, Calgary, Alberta, Canada Supplementary Methods Bilayers of different sizes were simulated for different times according to the type of analysis. The choice was motivated by the best balance between convergence of the measured property / computational cost / avoidance of issues due to undulations on big patches. To evaluate the error, we generally divided the trajectory in different numbers of blocks and calculated the standard deviation of the mean of each block (as summarized in Table S1). Hydrophobic thickness Hydrophobic thickness was evaluated from the peak to peak distance of C2 atoms in density plots (see Figure 2c of the main paper, dashed vertical lines). 1
2 Area and volume of DOPC and DOG The area of DOPC and DOG was evaluated by analyzing simulations of 70 lipids at different DOG fraction, as proposed by Edholm and Nagle (1). A similar analysis was performed to assess the volume of DOPC and DOG (see Figures S1 and S2 below). Lateral diffusion Lateral diffusion of DOPC molecules (considering phosphorous atoms of DOPC) was calculated on patches of 128 lipids from the slope of the mean square displacement curve using Einstein law. To correct for the overall motion of each monolayer, the linear momentum was removed at each step for each monolayer separately (2). The trajectory was divided into two blocks. The error was evaluated by calculating the standard deviation over the two leaflets of the two blocks. Potential of mean force Using a procedure similar to ref (3) (note that the force field therein is slightly different to the one used in the present study), we used umbrella sampling to determine the potential of mean force (PMF) for moving a DOG (or DOPC) head group from equilibrium into the center of the bilayer. A harmonic potential was applied to the OH of DOG (or phosphate of DOPC) with respect to the center of mass of the bilayer, with a force constant of 3000 kj mol -1 nm -2. For the DOG and DOPC PMF, 21 independent simulations with a 0.1 nm spacing between the umbrella positions (0 nm to 2 nm), were run for 120 ns (two sets with 60 ns for each of the 21 umbrella windows). The starting conformations of the two sets were obtained by pulling either DOG or DOPC in the two directions. The weighted histogram analysis method was used to calculate the PMFs (4). We thus obtained one PMF for each leaflet and the error was estimated as the standard deviation over the two leaflets. Lateral pressure profile The lateral pressure profile was computed on the patches of 128 lipids as a post-trajectory analysis using a modified version of GROMACS (5, 6). The simulation box was 2
3 basically cut into slices of 0.1 nm along the z direction, and the local pressure was computed for each slice from the local virial as described in refs (5, 6). The local pressure profile is evaluated as follows: π(z) = P L - P N (1), where P L is the lateral component (in the xy plane, P L =(P xx +P yy )/2) of the pressure, and P N is the normal component (along z, P N =P zz ). Recently Ollila noticed that all-atom simulations give non-constant P N as compared to coarse-grained simulations for which sampling is more than two orders of magnitude higher (7). According to this author, one possible explanation could be that the rerun is performed using a long cutoff (2.0 nm) for computing electrostatics, whereas the original simulation runs are performed using PME (the use of a cutoff is due to the difficulty of extracting specific contributions from the reciprocal term of PME). In our simulations, P N was not constant over z but similar in all simulations. As we are only interested in the relative behavior between the different systems, we thus only report the P L component in the presented pressure profiles that we call P(z), as was done previously (5, 6). When P(z) is positive, the bilayer tends to expand whereas when P(z) is negative it tends to shrink. For a bilayer with no spontaneous curvature, the integral of P(z) (i.e. surface tension γ) should be equal to 0 (perfect balance between negative and positive contributions). Bilayer compressibility The area compressibility modulus K A is evaluated from the fluctuations of bilayer area: K A = (Ak B T)/σ 2 A (2), where A is the total bilayer area, k B the Boltzmann constant, T the absolute temperature and σ 2 A the variance of A. To avoid issues related to undulations we used small patches of 70 lipids (almost undulation free) but pushed the simulations up to 800 ns to evaluate the error (by dividing the trajectory in 3 slices - after 100 ns of equilibration and computed the standard deviation of K A over the 3 slices). It has been shown that computing compressibility on small patches overestimates the K A value (7), and on big patches the undulations have to be taken out of the pure area fluctuations (7). Nevertheless we were only interested in the trend on going from a system to another, not in the precise reproduction of experimental values. 3
4 Bending rigidity Bilayer rigidity was evaluated from the undulations of the big patches (1,024 lipids). Using the continuum model of Helfrich (8), it is possible to extract the rigidity (or bending) modulus κ from the power spectrum (see for example refs (9, 10)): 2 U( q) q 4 = kbt / Aκ (3), where A is the total area of the system, U(q) is the discrete Fourier transform of the out-ofplane displacement u(r) of the bilayer surface in Monge representation (r=[r x,r y ]), and < > means averaging over the trajectory (note that the term in q -2 involving microscopic surface tension γ has been omitted since we are only interested in the undulation regime). To get κ from equation (3), we first had to evaluate U(q). To do so, we first discretized the squared bilayer of edge length L into an N N mesh of elementary grid size l=l/(n-1). The phosphorous atom of the upper leaflet with the lowest [x,y] coordinates was chosen arbitrarily as the origin (i.e. r 1 ). For each grid point k and for each leaflet, we computed z(r k ) as the mean of the z coordinate of the three nearest phosphorous atoms. The out-of-plane displacement was computed from the mean z position between the two leaflets: u(r k )=½(z upper (r k )+z lower (r k )). U(q) was then evaluated from u(r k ) using a 2D discrete Fourier transform: U ( 1 2 N iq.rk q ) = 2 u( rk ) e (4), N k= 1 where q is the 2D wave vector, i.e. q=[q x,q y ]=2π[n x,n y ]/L (n x and n y are integers spanning 0 to N-1). Finally, U(q) was converted to U(q), where q is the norm of q. κ has been evaluated from formula (3) using a non-linear fit (so that it enforces the q -4 behavior, i.e. a slope of -4 from the plot U(q) 2 vs q with a log-log scale). Note that we used only the points between 0.33 (the lowest point due to the size of our system, e.g. roughly 19x19 nm for the pure DOPC system) and 1 nm -1 which correspond to the undulation regime. For this analysis, we used one frame every 100 ps from 5 ns to 200 ns. Error was evaluated by dividing the trajectory in 3 blocks and by computing the corresponding standard deviation. 4
5 Packing defect mapping Packing defects were mapped from simulations of 280 lipids, taking one frame per 100 ps from 100 to 200 ns. For each frame, water and ions were taken out and the corresponding pdb file was used in a pipeline of scripts to extract the packing defect areas. The next two sections describe the pipelines used for the ASA method and our new Cartesian methods. Packing defect mapping using 1) the ASA approach This method has been introduced recently by Voth and coworkers (11). We used the same global framework but with different details. Starting from the structure of the bilayer without water and ions, we duplicated the system in all xy directions to avoid issues on the edges. The lipids 1.5 nm away from the central patch were discarded. We then computed the accessible surface area (ASA) with the program NACCESS (12) using the same van der Waals radii as in ref (11). In order to compare our results to this work, we used the same probe radius (0.3 nm). This size was initially chosen because it roughly corresponds to the volume of a big hydrophobic residue such as Phe. All aliphatic carbons from the central (initial) patch with a non-null ASA were then extracted and analyzed for each leaflet separately. All non-null ASA were considered as packing defects and projected on the xy plane and converted to defect radii located at the (x,y) position of the corresponding atom (at this stage, a defect is thus a circle). A distance matrix was then computed between all individual defects, and overlapping defects were merged in a single defect according to the following criterions: i) if two defects overlapped (distance from their center < sum of their radii) they were merged into a single one; ii) if two defects did not overlap but were separated by a distance shorter than a threshold (distance from their center < sum of their radii + threshold), the defects were merged. We chose a threshold of 0.3 nm based on visual inspection of several snapshots (so that no polar atom could be located between the two defects). The area of merged defects was then computed using a grid approach with elementary squares of 0.01 nm 2. Packing defect mapping using 2) the Cartesian scheme We selected a plane that is perpendicular to the membrane normal and created a grid of 0.1 nm resolution. For each grid point, we scanned the normal to the membrane plane starting from the solvent and descending up to 0.1 nm below the nearest glycerol (C2 carbon of sn-2 chain). In the following, we ignore solvent and ion atoms and the analysis on each leaflet is done separately. At each z position of a given grid point, the presence of an eventual 5
6 overlapping atom was evaluated by calculating a 3D distance between the grid point center and the center of any atom of the bilayer. Overlap was assigned when this distance was shorter than the grid point diagonal half-length (0.07 nm) plus the van der Waals radius of the atom (see Figure S1). If a polar head atom was encountered the grid point was discarded. If an aliphatic atom was met, we retained the grid point and defined it as a chemical defect of size 0.01 nm 2. If the first atom met was aliphatic and its z position was 0.1 nm below the sn-2 carbon of the nearest glycerol, the grid point was assigned as a geometrical defect of size 0.01 nm 2. Adjacent elementary defects were then merged, resulting in defects of various sizes. Note that geometrical defects represent a sub-category of chemical defects. The van der Waals radii used for this analysis were taken from the r min values (distance at which the Lennard-Jones is minimal) of the Berger united-atom force field (13) (after the initial observation that classical radii used to compute ASA in NACCESS (12) or VMD (14) were giving poorly discriminated results and to better take into account the relative size of CH 2 and CH 3 ). Sensitivity of packing defect size distributions to different parameters We first checked whether the distributions were sensitive to system size. Too small patches (notably 70 lipids) gave in general distributions shifted to the left. We found that patches of 280 lipids were giving results close to those obtained on patches of 1,024 lipids. We thus performed all the packing defect analyses on systems of 280 lipids. For the ASA method, the results depended on the chosen threshold used for merging defects. We used a threshold of 0.3 nm. A smaller threshold gives distributions shifted to the left and vice-versa. We found that the value of 0.3 nm was the best compromise by visual inspection of various snapshots (two defects should be merged if and only if there is no polar head atom in between). For each distribution in Figure 5 of the main paper, we carefully checked their proper convergence. We found that 1,000 frames (from 100 to 200 ns every 100 ps) were giving enough statistics (e.g. for DMPC the number of defects was 51,149 for the ASA method, 107,744 for chemical defects, and 65,721 for geometrical defects; for DOPC/DOG the number of defects was 71,972 for the ASA method, 109,695 for chemical defects, and 83,423 for geometrical defects). The distributions were also sensitive to the way of binning the corresponding histogram from the raw data. For each method (ASA, chemical or geometrical defects), we imposed (for each 6
7 lipid composition) 101 breaks to get 100 distinct bins (from 0 to 1.2 nm for ASA and geometrical defects, and from 0 to 2.5 nm for chemical defects). We also tested the effect of removing ions. Note that all the patches of 280 lipids used for studying packing defects in the main paper were simulated with 120 mm of ions (Na + / Cl - ). Some additional simulations without ions were thus performed. Removing ions marginally affected the decays and did not change the relative differences between compositions. Last, we tested the effect of temperature. Increasing temperature shifted the distributions to the right (towards larger defects), but again the relative difference between compositions was preserved. The increase of temperature for a particular composition can thus mimic the distribution of another composition at a different temperature (for example DMPC at 333K displays the same distribution as DOPC at 300K). This parameter is important since we performed simulations at a higher temperature (333K) in the accompanying paper. 7
8 Supplementary Figures Figure S1. Scheme of the Cartesian method for detecting packing defects. First a grid with 0.1 nm resolution (in light blue above the bilayer in surface representation) is built and the analysis is done separately on each leaflet (note that the size of each grid point has been scaled up for clarity). For each grid point, the z position is scanned starting outside the bilayer and descending towards the bilayer center. At each z position, the method searches an eventual overlap with any atom of that leaflet. A distance d between the center of the grid point and the center of each atom is calculated. An overlap is assigned when d is shorter than the van der Waals radius (r vdw ) plus the grid point half diagonal length (ldiag 1/2 ). The two possible cases are sketched in top view on the left of the figure (the grid point is represented as a light blue square whereas the atom is represented as a black circle): 1) no overlap is assigned (d > r vdw + ldiag 1/2 ); 2) an overlap is assigned when the atom spans the inscribed or circumscribed circle of the grid point (d < r vdw + ldiag 1/2 ). The snapshot in surface representation was generated using Pymol (15). 8
9 Figure S2. Evaluation of DOPC and DOG area from the simulations at different DOPC/DOG ratios. These areas were extracted from a linear fit to the following equation: AL ( x) x = ADOPC ( x) + ADOG ( x), where A L is the area per total lipid, A DOPC the area per 1-x 1 x DOPC, A DOG the area per DOG, x the mol fraction of DOG (1). DOPC area thus corresponds to the intercept and that of DOG to the slope. We find A DOPC =0.70 nm 2 and A DOG =0.46 nm 2. 9
10 Figure S3. Evaluation of DOPC and DOG volume from the simulations at different DOPC/DOG ratios. These volumes were extracted from a linear fit to the following equation: V L (x)=(1-x)v DOPC (x)+xv DOG (x), where V L is the volume per total lipid, V DOPC the volume per DOPC, V DOG the volume per DOG, x the mol fraction of DOG (1). V L required an additional simulation of pure water (same number of molecules as for the bilayer systems) and was computed as follows: V L =(V bilayer_system -V water )/N, where N is the total number of lipids, V water is the volume of the water box, V bilayer_system is the volume of the whole system containing the bilayer (bilayer+water). V DOPC was computed from the simulation at x=0 and V DOG was extrapolated from the straight line at x=1. We find V DOPC =1.31 nm 3 and V DOG =1.12 nm 3. 10
11 Figure S4. Radial distribution function g(r) of oxygen atoms O4 vs water oxygen atoms OW. O4 atom corresponds to the oxygen that is connecting the glycerol and the phosphate group in DOPC, whereas it is the oxygen of the hydroxyl group in DOG (see Figure 1 of the main article). The top panel shows any O4 atom for both systems. In the bottom panel, the detail is given for DOG and DOPC in the DOPC/DOG mixture. The hydrogen-bond donor and acceptor capacity of O4 in DOG explains that the first peak is higher in DOPC/DOG (red) compared to pure DOPC (black) for which O4 is only acceptor. Inversely, the second peak is lower in DOPC/DOG because DOG is partitioned deeper towards the bilayer center and has thus less second-neighbor OW atoms accessible. 11
12 Figure S5. Snapshots during DOG and DOPC flip-flop. Water: thick blue lines; DOPC: thin grey lines; DOG: thin red lines; DOPC nitrogen: grey balls; DOG hydroxyl: red balls. The restrained lipid is shown in thicker lines. A and B. DOG flip-flop across a pure DOPC bilayer. C and D. DOPC flip-flop across a pure DOPC bilayer. For A and C, the restrained lipids head group is at the bilayer center, and at 0.6 nm from the bilayer center of B and D. The DOG in the center of the bilayer (A) loses its hydration whereas there is a persisting water pore for DOPC (C). The snapshots were generated with VMD (14). 12
13 Figure S6. Distribution of the fraction of area occupied by packing defects within the total DOG/DOPC area for (A) geometric and (B) chemical defects. The mean and standard deviation are reported in Table 2 of the main manuscript. The vertical dotted lines represent the mean of each distribution. These distributions were accumulated from the trajectory of 280 lipids on 3000 snapshots (one every 100 ps). 13
14 Figure S7. Packing defects do not co-localize with DOG. The snapshot has been taken from a simulation of a DOPC/DOG membrane (280 lipids). The aliphatic chains of DOG and DOPC are represented in blue and pale-green spheres, respectively (for clarity no polar head is shown). Packing defects are depicted as magenta spheres (one per grid point). One can clearly see that the defects are located either on DOG or DOPC, and that there is no specific preference of packing defects for DOG. Similar analysis on several snapshots leads to the same conclusion. This snapshot was rendered with Pymol (15). 14
15 Table S1. Overview of the different simulations and type of analysis. The first column indicates the lipid composition, the second the patch size, the third the simulation time used for the analysis (after equilibration), the fourth the number of blocks used for calculating the error (as the standard deviation of the mean of each block), the last column indicates the type of analysis. System Pure DOPC DOPC-10%DOG DOPC-15%DOG DOPC-20%DOG DOPC-25%DOG DOPC-35%DOG Nb lipids Time (ns) Nb blocks Type of analysis Area/lipid Volume/lipid Pure DOPC DOPC-15%DOG Hydrophobic thickness pure DOPC DOPC-15%DOG pure DOPC DOPC-15%DOG pure DOPC DOPC-15%DOG pure DOPC DOPC-15%DOG pure POPC pure DMPC pure DOPC DOPC-15%DOG pure DOPC DOPC-15%DOG Compressibility modulus Density Radial distribution Lateral pressure profile Lateral diffusion Packing defects Bending rigidity modulus 15
16 Supporting References 1. Edholm, O., and J. F. Nagle Areas of Molecules in Membranes Consisting of Mixtures. Biophys J 89: Wohlert, J., and O. Edholm Dynamics in atomistic simulations of phospholipid membranes: Nuclear magnetic resonance relaxation rates and lateral diffusion. J Chem Phys 125: Bennett, W. F. D., and D. P. Tieleman Molecular simulation of rapid translocation of cholesterol, diacylglycerol, and ceramide in model raft and nonraft membranes. J Lipid Res 53: Kumar, S., J. M. Rosenberg, D. Bouzida, R. H. Swendsen, and P. A. Kollman The weighted histogram analysis method for free-energy calculations on biomolecules. I. The method. J Comput Chem 13: Ollila, O. H. S., H. J. Risselada, M. Louhivuori, E. Lindahl, I. Vattulainen, and S. J. Marrink D Pressure Field in Lipid Membranes and Membrane-Protein Complexes. Phys Rev Lett 102: Ollila, S., M. T. Hyvönen, and I. Vattulainen Polyunsaturation in Lipid Membranes: Dynamic Properties and Lateral Pressure Profiles. J Phys Chem B 111: Waheed, Q., and O. Edholm Undulation Contributions to the Area Compressibility in Lipid Bilayer Simulations. Biophys J 97: Helfrich, W Elastic properties of lipid bilayers: theory and possible experiments. Z Naturforsch C 28: Brandt, Erik G., Anthony R. Braun, Jonathan N. Sachs, John F. Nagle, and O. Edholm Interpretation of Fluctuation Spectra in Lipid Bilayer Simulations. Biophys J 100: Lindahl, E., and O. Edholm Mesoscopic Undulations and Thickness Fluctuations in Lipid Bilayers from Molecular Dynamics Simulations. Biophys J 79: Cui, H., E. Lyman, and G. A. Voth Mechanism of Membrane Curvature Sensing by Amphipathic Helix Containing Proteins. Biophys J 100: Hubbard, S. J., and J. M. Thornton NACCESS. Department of Biochemistry and Molecular Biology, University College London. 13. Berger, O., O. Edholm, and F. Jahnig Molecular dynamics simulations of a fluid bilayer of dipalmitoylphosphatidylcholine at full hydration, constant pressure, and constant temperature. Biophys J 72: Humphrey, W., A. Dalke, and K. Schulten VMD: visual molecular dynamics. Journal of molecular graphics 14:33-38, Schrodinger, L The PyMOL Molecular Graphics System, Version 1.3r1. 16
We parameterized a coarse-grained fullerene consistent with the MARTINI coarse-grained force field
 Parameterization of the fullerene coarse-grained model We parameterized a coarse-grained fullerene consistent with the MARTINI coarse-grained force field for lipids 1 and proteins 2. In the MARTINI force
Parameterization of the fullerene coarse-grained model We parameterized a coarse-grained fullerene consistent with the MARTINI coarse-grained force field for lipids 1 and proteins 2. In the MARTINI force
Interactions of Polyethylenimines with Zwitterionic and. Anionic Lipid Membranes
 Interactions of Polyethylenimines with Zwitterionic and Anionic Lipid Membranes Urszula Kwolek, Dorota Jamróz, Małgorzata Janiczek, Maria Nowakowska, Paweł Wydro, Mariusz Kepczynski Faculty of Chemistry,
Interactions of Polyethylenimines with Zwitterionic and Anionic Lipid Membranes Urszula Kwolek, Dorota Jamróz, Małgorzata Janiczek, Maria Nowakowska, Paweł Wydro, Mariusz Kepczynski Faculty of Chemistry,
Penetration of Gold Nanoparticle through Human Skin: Unraveling Its Mechanisms at the Molecular Scale
 Penetration of Gold Nanoparticle through Human Skin: Unraveling Its Mechanisms at the Molecular Scale Rakesh Gupta and Beena Rai* TATA Research Development & Design Centre, TCS Innovation labs, Pune India,
Penetration of Gold Nanoparticle through Human Skin: Unraveling Its Mechanisms at the Molecular Scale Rakesh Gupta and Beena Rai* TATA Research Development & Design Centre, TCS Innovation labs, Pune India,
Supplementary Information
 Coarse-Grained Molecular Dynamics Simulations of Photoswitchable Assembly and Disassembly Xiaoyan Zheng, Dong Wang,* Zhigang Shuai,* MOE Key Laboratory of Organic Optoelectronics and Molecular Engineering,
Coarse-Grained Molecular Dynamics Simulations of Photoswitchable Assembly and Disassembly Xiaoyan Zheng, Dong Wang,* Zhigang Shuai,* MOE Key Laboratory of Organic Optoelectronics and Molecular Engineering,
Supplementary Information: A Critical. Comparison of Biomembrane Force Fields: Structure and Dynamics of Model DMPC, POPC, and POPE Bilayers
 Supplementary Information: A Critical Comparison of Biomembrane Force Fields: Structure and Dynamics of Model DMPC, POPC, and POPE Bilayers Kristyna Pluhackova,, Sonja A. Kirsch, Jing Han, Liping Sun,
Supplementary Information: A Critical Comparison of Biomembrane Force Fields: Structure and Dynamics of Model DMPC, POPC, and POPE Bilayers Kristyna Pluhackova,, Sonja A. Kirsch, Jing Han, Liping Sun,
Coarse grained simulations of Lipid Bilayer Membranes
 Coarse grained simulations of Lipid Bilayer Membranes P. B. Sunil Kumar Department of Physics IIT Madras, Chennai 600036 sunil@iitm.ac.in Atomistic MD: time scales ~ 10 ns length scales ~100 nm 2 To study
Coarse grained simulations of Lipid Bilayer Membranes P. B. Sunil Kumar Department of Physics IIT Madras, Chennai 600036 sunil@iitm.ac.in Atomistic MD: time scales ~ 10 ns length scales ~100 nm 2 To study
Translocation of C 60 and Its Derivatives Across a Lipid Bilayer
 Translocation of C 60 and Its Derivatives Across a Lipid Bilayer Rui Qiao* NANO LETTERS 2007 Vol. 7, No. 3 614-619 Department of Mechanical Engineering, Clemson UniVersity, Clemson, South Carolina 29634
Translocation of C 60 and Its Derivatives Across a Lipid Bilayer Rui Qiao* NANO LETTERS 2007 Vol. 7, No. 3 614-619 Department of Mechanical Engineering, Clemson UniVersity, Clemson, South Carolina 29634
Supporting Information Identification of Amino Acids with Sensitive Nanoporous MoS 2 : Towards Machine Learning-Based Prediction
 Supporting Information Identification of Amino Acids with Sensitive Nanoporous MoS 2 : Towards Machine Learning-Based Prediction Amir Barati Farimani, Mohammad Heiranian, Narayana R. Aluru 1 Department
Supporting Information Identification of Amino Acids with Sensitive Nanoporous MoS 2 : Towards Machine Learning-Based Prediction Amir Barati Farimani, Mohammad Heiranian, Narayana R. Aluru 1 Department
Interaction of Functionalized C 60 Nanoparticles with Lipid Membranes
 Interaction of Functionalized C 60 Nanoparticles with Lipid Membranes Kevin Gasperich Advisor: Dmitry Kopelevich ABSTRACT The recent rapid development of applications for fullerenes has led to an increase
Interaction of Functionalized C 60 Nanoparticles with Lipid Membranes Kevin Gasperich Advisor: Dmitry Kopelevich ABSTRACT The recent rapid development of applications for fullerenes has led to an increase
Bilayer Deformation, Pores & Micellation Induced by Oxidized Lipids
 Supporting Information Bilayer Deformation, Pores & Micellation Induced by Oxidized Lipids Phansiri Boonnoy 1, Viwan Jarerattanachat 1,2, Mikko Karttunen 3*, and Jirasak Wongekkabut 1* 1 Department of
Supporting Information Bilayer Deformation, Pores & Micellation Induced by Oxidized Lipids Phansiri Boonnoy 1, Viwan Jarerattanachat 1,2, Mikko Karttunen 3*, and Jirasak Wongekkabut 1* 1 Department of
Supporting Information: Revisiting partition in. hydrated bilayer systems
 Supporting Information: Revisiting partition in hydrated bilayer systems João T. S. Coimbra, Pedro A. Fernandes, Maria J. Ramos* UCIBIO, REQUIMTE, Departamento de Química e Bioquímica, Faculdade de Ciências,
Supporting Information: Revisiting partition in hydrated bilayer systems João T. S. Coimbra, Pedro A. Fernandes, Maria J. Ramos* UCIBIO, REQUIMTE, Departamento de Química e Bioquímica, Faculdade de Ciências,
Massive oxidation of phospholipid membranes. leads to pore creation and bilayer. disintegration
 Massive oxidation of phospholipid membranes leads to pore creation and bilayer disintegration Lukasz Cwiklik and Pavel Jungwirth Institute of Organic Chemistry and Biochemistry, Academy of Sciences of
Massive oxidation of phospholipid membranes leads to pore creation and bilayer disintegration Lukasz Cwiklik and Pavel Jungwirth Institute of Organic Chemistry and Biochemistry, Academy of Sciences of
Supporting material. Membrane permeation induced by aggregates of human islet amyloid polypeptides
 Supporting material Membrane permeation induced by aggregates of human islet amyloid polypeptides Chetan Poojari Forschungszentrum Jülich GmbH, Institute of Complex Systems: Structural Biochemistry (ICS-6),
Supporting material Membrane permeation induced by aggregates of human islet amyloid polypeptides Chetan Poojari Forschungszentrum Jülich GmbH, Institute of Complex Systems: Structural Biochemistry (ICS-6),
Supplementary information for Effects of Stretching Speed on. Mechanical Rupture of Phospholipid/Cholesterol Bilayers: Molecular
 Supplementary information for Effects of Stretching Speed on Mechanical Rupture of Phospholipid/Cholesterol Bilayers: Molecular Dynamics Simulation Taiki Shigematsu, Kenichiro Koshiyama*, and Shigeo Wada
Supplementary information for Effects of Stretching Speed on Mechanical Rupture of Phospholipid/Cholesterol Bilayers: Molecular Dynamics Simulation Taiki Shigematsu, Kenichiro Koshiyama*, and Shigeo Wada
Gold nanocrystals at DPPC bilayer. Bo Song, Huajun Yuan, Cynthia J. Jameson, Sohail Murad
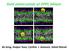 Gold nanocrystals at DPPC bilayer Bo Song, Huajun Yuan, Cynthia J. Jameson, Sohail Murad Coarse-grained mapping strategy of a DPPC lipid molecule. The structure of gold nanocrystals (bare gold nanoparticles)
Gold nanocrystals at DPPC bilayer Bo Song, Huajun Yuan, Cynthia J. Jameson, Sohail Murad Coarse-grained mapping strategy of a DPPC lipid molecule. The structure of gold nanocrystals (bare gold nanoparticles)
Flip-Flop Induced Relaxation Of Bending Energy: Implications For Membrane Remodeling
 Biophysical Journal, Volume 97 Supporting Material Flip-Flop Induced Relaxation Of Bending Energy: Implications For Membrane Remodeling Raphael Jeremy Bruckner, Sheref S. Mansy, Alonso Ricardo, L. Mahadevan,
Biophysical Journal, Volume 97 Supporting Material Flip-Flop Induced Relaxation Of Bending Energy: Implications For Membrane Remodeling Raphael Jeremy Bruckner, Sheref S. Mansy, Alonso Ricardo, L. Mahadevan,
Supporting information for: Quantifying. lateral inhomogeneity of. cholesterol-containing membranes
 Supporting information for: Quantifying lateral inhomogeneity of cholesterol-containing membranes Celsa Díaz-Tejada, Igor Ariz-Extreme, Neha Awasthi, and Jochen S. Hub Georg-August-University Göttingen,
Supporting information for: Quantifying lateral inhomogeneity of cholesterol-containing membranes Celsa Díaz-Tejada, Igor Ariz-Extreme, Neha Awasthi, and Jochen S. Hub Georg-August-University Göttingen,
Supporting Information
 Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 018 Supporting Information Modulating Interactions between Ligand-Coated Nanoparticles and Phase-Separated
Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 018 Supporting Information Modulating Interactions between Ligand-Coated Nanoparticles and Phase-Separated
Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site.
 Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation
 Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation Hang Yu, 1, 2, a) Wei Han, 1, 3, b) Wen Ma, 1, 2 1, 2, 3, c) and Klaus Schulten 1) Beckman
Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation Hang Yu, 1, 2, a) Wei Han, 1, 3, b) Wen Ma, 1, 2 1, 2, 3, c) and Klaus Schulten 1) Beckman
Conformational Flexibility of the Peptide Hormone Ghrelin in Solution and Lipid Membrane Bound: A Molecular Dynamics Study
 Open Access Article The authors, the publisher, and the right holders grant the right to use, reproduce, and disseminate the work in digital form to all users. Journal of Biomolecular Structure & Dynamics,
Open Access Article The authors, the publisher, and the right holders grant the right to use, reproduce, and disseminate the work in digital form to all users. Journal of Biomolecular Structure & Dynamics,
Coarse-grained simulation studies of mesoscopic membrane phenomena
 Coarse-grained simulation studies of mesoscopic membrane phenomena Markus Deserno Department of Physics Carnegie Mellon University with: Ira R. Cooke, Gregoria Illya, Benedict J. Reynwar, Vagelis A. Harmandaris,
Coarse-grained simulation studies of mesoscopic membrane phenomena Markus Deserno Department of Physics Carnegie Mellon University with: Ira R. Cooke, Gregoria Illya, Benedict J. Reynwar, Vagelis A. Harmandaris,
The main biological functions of the many varied types of lipids include: energy storage protection insulation regulation of physiological processes
 Big Idea In the biological sciences, a dehydration synthesis (condensation reaction) is typically defined as a chemical reaction that involves the loss of water from the reacting molecules. This reaction
Big Idea In the biological sciences, a dehydration synthesis (condensation reaction) is typically defined as a chemical reaction that involves the loss of water from the reacting molecules. This reaction
Ethanol Induces the Formation of Water-Permeable Defects in Model. Bilayers of Skin Lipids
 Electronic Supplementary Material (ESI) for Chemical Communications. This journal is The Royal Society of Chemistry 2015 Ethanol Induces the Formation of Water-Permeable Defects in Model Bilayers of Skin
Electronic Supplementary Material (ESI) for Chemical Communications. This journal is The Royal Society of Chemistry 2015 Ethanol Induces the Formation of Water-Permeable Defects in Model Bilayers of Skin
RELEASE OF CONTENT THROUGH MECHANO- SENSITIVE GATES IN PRESSURISED LIPOSOMES. Martti Louhivuori University of Groningen
 RELEASE OF CONTENT THROUGH MECHANO- SENSITIVE GATES IN PRESSURISED LIPOSOMES Martti Louhivuori University of Groningen www.cgmartini.nl MARTINI coarse-grained model water P4 butane Qo Na DPPC cholesterol
RELEASE OF CONTENT THROUGH MECHANO- SENSITIVE GATES IN PRESSURISED LIPOSOMES Martti Louhivuori University of Groningen www.cgmartini.nl MARTINI coarse-grained model water P4 butane Qo Na DPPC cholesterol
Free Volume Properties of Sphingomyelin, DMPC, DPPC, and PLPC Bilayers
 PUBLISHED IN: JOURNAL OF COMPUTATIONAL AND THEORETICAL NANOSCIENCE, VOL. 2, PP. 40-43 (2005). Free Volume Properties of Sphingomyelin, DMPC, DPPC, and PLPC Bilayers M. Kupiainen, E. Falck, S. Ollila, P.
PUBLISHED IN: JOURNAL OF COMPUTATIONAL AND THEORETICAL NANOSCIENCE, VOL. 2, PP. 40-43 (2005). Free Volume Properties of Sphingomyelin, DMPC, DPPC, and PLPC Bilayers M. Kupiainen, E. Falck, S. Ollila, P.
Coarse-grained model for phospholipid/cholesterol bilayer employing inverse Monte Carlo with thermodynamic constraints
 THE JOURNAL OF CHEMICAL PHYSICS 126, 075101 2007 Coarse-grained model for phospholipid/cholesterol bilayer employing inverse Monte Carlo with thermodynamic constraints Teemu Murtola Laboratory of Physics,
THE JOURNAL OF CHEMICAL PHYSICS 126, 075101 2007 Coarse-grained model for phospholipid/cholesterol bilayer employing inverse Monte Carlo with thermodynamic constraints Teemu Murtola Laboratory of Physics,
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. (a) Uncropped version of Fig. 2a. RM indicates that the translation was done in the absence of rough mcirosomes. (b) LepB construct containing the GGPG-L6RL6-
Supplementary Figures Supplementary Figure 1. (a) Uncropped version of Fig. 2a. RM indicates that the translation was done in the absence of rough mcirosomes. (b) LepB construct containing the GGPG-L6RL6-
obtained for the simulations of the E2 conformation of SERCA in a pure POPC lipid bilayer (blue) and in a
 Supplementary Figure S1. Distribution of atoms along the bilayer normal. Normalized density profiles obtained for the simulations of the E2 conformation of SERCA in a pure POPC lipid bilayer (blue) and
Supplementary Figure S1. Distribution of atoms along the bilayer normal. Normalized density profiles obtained for the simulations of the E2 conformation of SERCA in a pure POPC lipid bilayer (blue) and
Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random
 S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels. Supplementary Information
 Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels Craig T. Armstrong, Philip E. Mason, J. L. Ross Anderson and Christopher E. Dempsey
Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels Craig T. Armstrong, Philip E. Mason, J. L. Ross Anderson and Christopher E. Dempsey
Biological Membranes. Lipid Membranes. Bilayer Permeability. Common Features of Biological Membranes. A highly selective permeability barrier
 Biological Membranes Structure Function Composition Physicochemical properties Self-assembly Molecular models Lipid Membranes Receptors, detecting the signals from outside: Light Odorant Taste Chemicals
Biological Membranes Structure Function Composition Physicochemical properties Self-assembly Molecular models Lipid Membranes Receptors, detecting the signals from outside: Light Odorant Taste Chemicals
A: All atom molecular simulation systems
 Cholesterol level affects surface charge of lipid membranes in physiological environment Aniket Magarkar a, Vivek Dhawan b, Paraskevi Kallinteri a, Tapani Viitala c, Mohammed Elmowafy c, Tomasz Róg d,
Cholesterol level affects surface charge of lipid membranes in physiological environment Aniket Magarkar a, Vivek Dhawan b, Paraskevi Kallinteri a, Tapani Viitala c, Mohammed Elmowafy c, Tomasz Róg d,
in-silico Design of Nanoparticles for Transdermal Drug Delivery Application
 Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2018 in-silico Design of Nanoparticles for Transdermal Drug Delivery Application Rakesh Gupta and Beena
Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2018 in-silico Design of Nanoparticles for Transdermal Drug Delivery Application Rakesh Gupta and Beena
Life Sciences 1a. Practice Problems 4
 Life Sciences 1a Practice Problems 4 1. KcsA, a channel that allows K + ions to pass through the membrane, is a protein with four identical subunits that form a channel through the center of the tetramer.
Life Sciences 1a Practice Problems 4 1. KcsA, a channel that allows K + ions to pass through the membrane, is a protein with four identical subunits that form a channel through the center of the tetramer.
Molecular modeling of the pathways of vesicle membrane interaction. Tongtao Yue and Xianren Zhang
 Molecular modeling of the pathways of vesicle membrane interaction Tongtao Yue and Xianren Zhang I. ELECTRONIC SUPPLEMENTARY INFORMATION (ESI): METHODS Dissipative particle dynamics method The dissipative
Molecular modeling of the pathways of vesicle membrane interaction Tongtao Yue and Xianren Zhang I. ELECTRONIC SUPPLEMENTARY INFORMATION (ESI): METHODS Dissipative particle dynamics method The dissipative
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/4/3/eaaq0762/dc1 Supplementary Materials for Structures of monomeric and oligomeric forms of the Toxoplasma gondii perforin-like protein 1 Tao Ni, Sophie I. Williams,
advances.sciencemag.org/cgi/content/full/4/3/eaaq0762/dc1 Supplementary Materials for Structures of monomeric and oligomeric forms of the Toxoplasma gondii perforin-like protein 1 Tao Ni, Sophie I. Williams,
0.5 nm nm acyl tail region (hydrophobic) 1.5 nm. Hydrophobic repulsion organizes amphiphilic molecules: These scales are 5 10xk B T:
 Lecture 31: Biomembranes: The hydrophobic energy scale and membrane behavior 31.1 Reading for Lectures 30-32: PKT Chapter 11 (skip Ch. 10) Upshot of last lecture: Generic membrane lipid: Can be cylindrical
Lecture 31: Biomembranes: The hydrophobic energy scale and membrane behavior 31.1 Reading for Lectures 30-32: PKT Chapter 11 (skip Ch. 10) Upshot of last lecture: Generic membrane lipid: Can be cylindrical
SUPPLEMENTARY INFORMATION. Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity
 SUPPLEMENTARY INFORMATION Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity Christopher Woods 1, Maturos Malaisree 1, Ben Long 2, Simon McIntosh-Smith
SUPPLEMENTARY INFORMATION Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity Christopher Woods 1, Maturos Malaisree 1, Ben Long 2, Simon McIntosh-Smith
SDS-Assisted Protein Transport Through Solid-State Nanopores
 Supplementary Information for: SDS-Assisted Protein Transport Through Solid-State Nanopores Laura Restrepo-Pérez 1, Shalini John 2, Aleksei Aksimentiev 2 *, Chirlmin Joo 1 *, Cees Dekker 1 * 1 Department
Supplementary Information for: SDS-Assisted Protein Transport Through Solid-State Nanopores Laura Restrepo-Pérez 1, Shalini John 2, Aleksei Aksimentiev 2 *, Chirlmin Joo 1 *, Cees Dekker 1 * 1 Department
Simulation of Self-Assembly of Ampiphiles Using Molecular Dynamics
 Simulation of Self-Assembly of Ampiphiles Using Molecular Dynamics Haneesh Kesari, Reza Banki, and Misty Davies Project Final Paper ME346 Stanford University December 15, 2005 1 Introduction Understanding
Simulation of Self-Assembly of Ampiphiles Using Molecular Dynamics Haneesh Kesari, Reza Banki, and Misty Davies Project Final Paper ME346 Stanford University December 15, 2005 1 Introduction Understanding
Amphipathic Lipid Packing Sensor Motifs: Probing Bilayer Defects with Hydrophobic Residues
 Biophysical Journal Volume 104 February 2013 575 584 575 Amphipathic Lipid Packing Sensor Motifs: Probing Bilayer Defects with Hydrophobic Residues Stefano Vanni, Lydie Vamparys, { Romain Gautier, Guillaume
Biophysical Journal Volume 104 February 2013 575 584 575 Amphipathic Lipid Packing Sensor Motifs: Probing Bilayer Defects with Hydrophobic Residues Stefano Vanni, Lydie Vamparys, { Romain Gautier, Guillaume
Physicochemical Properties of Nanoparticles Regulate Translocation across Pulmonary. Surfactant Monolayer and Formation of Lipoprotein Corona
 Physicochemical Properties of Nanoparticles Regulate Translocation across Pulmonary Surfactant Monolayer and Formation of Lipoprotein Corona Supplementary Information Guoqing Hu 1 *, Bao Jiao 1, Xinghua
Physicochemical Properties of Nanoparticles Regulate Translocation across Pulmonary Surfactant Monolayer and Formation of Lipoprotein Corona Supplementary Information Guoqing Hu 1 *, Bao Jiao 1, Xinghua
Document Version Publisher s PDF, also known as Version of Record (includes final page, issue and volume numbers)
 Coarse-grained model for phospholipid/cholesterol bilayer employing inverse Monte Carlo with thermodynamic constraints Murtola, T.; Falck, E.; Karttunen, M.E.J.; Vattulainen, I. Published in: Journal of
Coarse-grained model for phospholipid/cholesterol bilayer employing inverse Monte Carlo with thermodynamic constraints Murtola, T.; Falck, E.; Karttunen, M.E.J.; Vattulainen, I. Published in: Journal of
Fluid Mozaic Model of Membranes
 Replacement for the 1935 Davson Danielli model Provided explanation for Gortner-Grendel lack of lipid and permitted the unit membrane model. Trans membrane protein by labelling Fry & Edidin showed that
Replacement for the 1935 Davson Danielli model Provided explanation for Gortner-Grendel lack of lipid and permitted the unit membrane model. Trans membrane protein by labelling Fry & Edidin showed that
Impact of cholesterol on voids in phospholipid membranes
 JOURNAL OF CHEMICAL PHYSICS VOLUME 121, NUMBER 24 22 DECEMBER 2004 Impact of cholesterol on voids in phospholipid membranes Emma Falck a) Laboratory of Physics and Helsinki Institute of Physics, Helsinki
JOURNAL OF CHEMICAL PHYSICS VOLUME 121, NUMBER 24 22 DECEMBER 2004 Impact of cholesterol on voids in phospholipid membranes Emma Falck a) Laboratory of Physics and Helsinki Institute of Physics, Helsinki
Lipids and Membranes
 Lipids Lipids are hydrophobic or amphiphilic insoluble in water soluble in organic solvents soluble in lipids Lipids are used as energy storage molecules structural components of membranes protective molecules
Lipids Lipids are hydrophobic or amphiphilic insoluble in water soluble in organic solvents soluble in lipids Lipids are used as energy storage molecules structural components of membranes protective molecules
Simulationen von Lipidmembranen
 Simulationen von Lipidmembranen Thomas Stockner thomas.stockner@meduniwien.ac.at Molecular biology Molecular modelling Membranes environment Many cellular functions occur in or around membranes: energy
Simulationen von Lipidmembranen Thomas Stockner thomas.stockner@meduniwien.ac.at Molecular biology Molecular modelling Membranes environment Many cellular functions occur in or around membranes: energy
Supplementary Figure 1. Overview of steps in the construction of photosynthetic protocellular systems
 Supplementary Figure 1 Overview of steps in the construction of photosynthetic protocellular systems (a) The small unilamellar vesicles were made with phospholipids. (b) Three types of small proteoliposomes
Supplementary Figure 1 Overview of steps in the construction of photosynthetic protocellular systems (a) The small unilamellar vesicles were made with phospholipids. (b) Three types of small proteoliposomes
The effect of orientation dynamics in melittin as antimicrobial peptide in lipid bilayer calculated by free energy method
 Journal of Physics: Conference Series PAPER OPEN ACCESS The effect of orientation dynamics in melittin as antimicrobial peptide in lipid bilayer calculated by free energy method To cite this article: Sri
Journal of Physics: Conference Series PAPER OPEN ACCESS The effect of orientation dynamics in melittin as antimicrobial peptide in lipid bilayer calculated by free energy method To cite this article: Sri
Unit 1 Exploring and Understanding Data
 Unit 1 Exploring and Understanding Data Area Principle Bar Chart Boxplot Conditional Distribution Dotplot Empirical Rule Five Number Summary Frequency Distribution Frequency Polygon Histogram Interquartile
Unit 1 Exploring and Understanding Data Area Principle Bar Chart Boxplot Conditional Distribution Dotplot Empirical Rule Five Number Summary Frequency Distribution Frequency Polygon Histogram Interquartile
Irina V. Ionova, Vsevolod A. Livshits, and Derek Marsh
 Phase Diagram of Ternary Cholesterol/Palmitoylsphingomyelin/Palmitoyloleoyl- Phosphatidylcholine Mixtures: Spin-abel EPR Study of ipid-raft Formation Irina V. Ionova, Vsevolod A. ivshits, and Derek Marsh
Phase Diagram of Ternary Cholesterol/Palmitoylsphingomyelin/Palmitoyloleoyl- Phosphatidylcholine Mixtures: Spin-abel EPR Study of ipid-raft Formation Irina V. Ionova, Vsevolod A. ivshits, and Derek Marsh
Adaptable Lipid Matrix Promotes Protein Protein Association in Membranes
 Supporting information Adaptable Lipid Matrix Promotes Protein Protein Association in Membranes Andrey S. Kuznetsov, Anton A. Polyansky,, Markus Fleck, Pavel E. Volynsky, and Roman G. Efremov *,, M. M.
Supporting information Adaptable Lipid Matrix Promotes Protein Protein Association in Membranes Andrey S. Kuznetsov, Anton A. Polyansky,, Markus Fleck, Pavel E. Volynsky, and Roman G. Efremov *,, M. M.
Simulationen von Lipidmembranen
 Simulationen von Lipidmembranen Thomas Stockner Thomas.stockner@meduniwien.ac.at Summary Methods Force Field MD simulations Membrane simulations Application Oxidized lipids Anesthetics Molecular biology
Simulationen von Lipidmembranen Thomas Stockner Thomas.stockner@meduniwien.ac.at Summary Methods Force Field MD simulations Membrane simulations Application Oxidized lipids Anesthetics Molecular biology
TUTORIAL IN SMALL ANGLE X-RAY SCATTERING ANALYSIS
 TUTORIAL IN SMALL ANGLE X-RAY SCATTERING ANALYSIS at the Abdus Salam International Center of Theoretical Physics (ICTP) Heinz Amenitsch Sigrid Bernstorff Michael Rappolt Trieste, 15. May 2006 (14:30-17:15)
TUTORIAL IN SMALL ANGLE X-RAY SCATTERING ANALYSIS at the Abdus Salam International Center of Theoretical Physics (ICTP) Heinz Amenitsch Sigrid Bernstorff Michael Rappolt Trieste, 15. May 2006 (14:30-17:15)
Coarse-Grained Molecular Dynamics for Copolymer- Vesicle Self-Assembly. Case Study: Sterically Stabilized Liposomes.
 Coarse-Grained Molecular Dynamics for Copolymer- Vesicle Self-Assembly. Case Study: Sterically Stabilized Liposomes. Alexander Kantardjiev 1 and Pavletta Shestakova 1 1 Institute of Organic Chemistry with
Coarse-Grained Molecular Dynamics for Copolymer- Vesicle Self-Assembly. Case Study: Sterically Stabilized Liposomes. Alexander Kantardjiev 1 and Pavletta Shestakova 1 1 Institute of Organic Chemistry with
Improved Coarse-Grained Modeling of Cholesterol-Containing Lipid Bilayers
 This is an open access article published under an ACS AuthorChoice License, which permits copying and redistribution of the article or any adaptations for non-commercial purposes. pubs.acs.org/jctc Improved
This is an open access article published under an ACS AuthorChoice License, which permits copying and redistribution of the article or any adaptations for non-commercial purposes. pubs.acs.org/jctc Improved
FIRM. Full Iterative Relaxation Matrix program
 FIRM Full Iterative Relaxation Matrix program FIRM is a flexible program for calculating NOEs and back-calculated distance constraints using the full relaxation matrix approach. FIRM is an interactive
FIRM Full Iterative Relaxation Matrix program FIRM is a flexible program for calculating NOEs and back-calculated distance constraints using the full relaxation matrix approach. FIRM is an interactive
Role of Surface Ligands in Nanoparticle Permeation through a Model. Membrane: A Coarse grained Molecular Dynamics Simulations Study
 Role of Surface Ligands in Nanoparticle Permeation through a Model Membrane: A Coarse grained Molecular Dynamics Simulations Study Bo Song, Huajun Yuan, Cynthia J. Jameson, Sohail Murad * Department of
Role of Surface Ligands in Nanoparticle Permeation through a Model Membrane: A Coarse grained Molecular Dynamics Simulations Study Bo Song, Huajun Yuan, Cynthia J. Jameson, Sohail Murad * Department of
Topic 7b: Biological Membranes
 Topic 7b: Biological Membranes Overview: Why does life need a compartment? Nature of the packaging what is it made of? properties? New types of deformations in 2D Applications: Stretching membranes, forming
Topic 7b: Biological Membranes Overview: Why does life need a compartment? Nature of the packaging what is it made of? properties? New types of deformations in 2D Applications: Stretching membranes, forming
Nature Neuroscience: doi: /nn Supplementary Figure 1. Behavioral training.
 Supplementary Figure 1 Behavioral training. a, Mazes used for behavioral training. Asterisks indicate reward location. Only some example mazes are shown (for example, right choice and not left choice maze
Supplementary Figure 1 Behavioral training. a, Mazes used for behavioral training. Asterisks indicate reward location. Only some example mazes are shown (for example, right choice and not left choice maze
Structure and dynamics of water and lipid molecules in charged anionic DMPG lipid bilayer membranes
 Structure and dynamics of water and lipid molecules in charged anionic DMPG lipid bilayer membranes A. K. Rønnest, G. H. Peters, F. Y. Hansen, H. Taub, and A. Miskowiec Citation: The Journal of Chemical
Structure and dynamics of water and lipid molecules in charged anionic DMPG lipid bilayer membranes A. K. Rønnest, G. H. Peters, F. Y. Hansen, H. Taub, and A. Miskowiec Citation: The Journal of Chemical
Supplementary Information A Hydrophobic Barrier Deep Within the Inner Pore of the TWIK-1 K2P Potassium Channel Aryal et al.
 Supplementary Information A Hydrophobic Barrier Deep Within the Inner Pore of the TWIK-1 K2P Potassium Channel Aryal et al. Supplementary Figure 1 TWIK-1 stability during MD simulations in a phospholipid
Supplementary Information A Hydrophobic Barrier Deep Within the Inner Pore of the TWIK-1 K2P Potassium Channel Aryal et al. Supplementary Figure 1 TWIK-1 stability during MD simulations in a phospholipid
Supplementary information. Membrane curvature enables N-Ras lipid anchor sorting to. liquid-ordered membrane phases
 Supplementary information Membrane curvature enables N-Ras lipid anchor sorting to liquid-ordered membrane phases Jannik Bruun Larsen 1,2,3, Martin Borch Jensen 1,2,3,6, Vikram K. Bhatia 1,2,3,7, Søren
Supplementary information Membrane curvature enables N-Ras lipid anchor sorting to liquid-ordered membrane phases Jannik Bruun Larsen 1,2,3, Martin Borch Jensen 1,2,3,6, Vikram K. Bhatia 1,2,3,7, Søren
MOLECULAR DYNAMICS SIMULATION OF MIXED LIPID BILAYER WITH DPPC AND MPPC: EFFECT OF CONFIGURATIONS IN GEL-PHASE
 MOLECULAR DYNAMICS SIMULATION OF MIXED LIPID BILAYER WITH DPPC AND MPPC: EFFECT OF CONFIGURATIONS IN GEL-PHASE A Thesis Presented to The Academic Faculty by Young Kyoung Kim In Partial Fulfillment of the
MOLECULAR DYNAMICS SIMULATION OF MIXED LIPID BILAYER WITH DPPC AND MPPC: EFFECT OF CONFIGURATIONS IN GEL-PHASE A Thesis Presented to The Academic Faculty by Young Kyoung Kim In Partial Fulfillment of the
Chapter 7: Membranes
 Chapter 7: Membranes Roles of Biological Membranes The Lipid Bilayer and the Fluid Mosaic Model Transport and Transfer Across Cell Membranes Specialized contacts (junctions) between cells What are the
Chapter 7: Membranes Roles of Biological Membranes The Lipid Bilayer and the Fluid Mosaic Model Transport and Transfer Across Cell Membranes Specialized contacts (junctions) between cells What are the
Lectures 16-17: Biological Membranes: Life in Two Dimensions
 Lectures 16-17: Biological Membranes: Life in Two Dimensions Lecturer: Brigita Urbanc Office: 1-909 (E-mail: brigita@drexel.edu) Course website: www.physics.drexel.edu/~brigita/courses/biophys_011-01/
Lectures 16-17: Biological Membranes: Life in Two Dimensions Lecturer: Brigita Urbanc Office: 1-909 (E-mail: brigita@drexel.edu) Course website: www.physics.drexel.edu/~brigita/courses/biophys_011-01/
Lecture 10 More about proteins
 Lecture 10 More about proteins Today we're going to extend our discussion of protein structure. This may seem far-removed from gene cloning, but it is the path to understanding the genes that we are cloning.
Lecture 10 More about proteins Today we're going to extend our discussion of protein structure. This may seem far-removed from gene cloning, but it is the path to understanding the genes that we are cloning.
Chapter 1 Membrane Structure and Function
 Chapter 1 Membrane Structure and Function Architecture of Membranes Subcellular fractionation techniques can partially separate and purify several important biological membranes, including the plasma and
Chapter 1 Membrane Structure and Function Architecture of Membranes Subcellular fractionation techniques can partially separate and purify several important biological membranes, including the plasma and
Dissipative particle dynamics of tension-induced membrane fusion
 Molecular Simulation Vol. 35, No. 7, June 2009, 554 560 Dissipative particle dynamics of tension-induced membrane fusion Andrea Grafmüller a, Julian Shillcock b and Reinhard Lipowsky a * a Theory and Bio-Systems,
Molecular Simulation Vol. 35, No. 7, June 2009, 554 560 Dissipative particle dynamics of tension-induced membrane fusion Andrea Grafmüller a, Julian Shillcock b and Reinhard Lipowsky a * a Theory and Bio-Systems,
Lipids and Membranes
 Lipids and Membranes Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry I Philadelphia University Faculty of pharmacy Biological membranes are composed of lipid bilayers
Lipids and Membranes Presented by Dr. Mohammad Saadeh The requirements for the Pharmaceutical Biochemistry I Philadelphia University Faculty of pharmacy Biological membranes are composed of lipid bilayers
Modeling and Molecular Dynamics of Membrane Proteins
 Modeling and Molecular Dynamics of Membrane Proteins Emad Tajkhorshid Department of Biochemistry, Center for Biophysics and Computational Biology, and Beckman Institute University of Illinois at Urbana-Champaign
Modeling and Molecular Dynamics of Membrane Proteins Emad Tajkhorshid Department of Biochemistry, Center for Biophysics and Computational Biology, and Beckman Institute University of Illinois at Urbana-Champaign
University of Groningen. Lipids out of equilibrium Tieleman, D. Peter; Marrink, Siewert. Published in: Journal of the American Chemical Society
 University of Groningen Lipids out of equilibrium Tieleman, D. Peter; Marrink, Siewert Published in: Journal of the American Chemical Society DOI: 10.1021/ja0624321 IMPORTANT NOTE: You are advised to consult
University of Groningen Lipids out of equilibrium Tieleman, D. Peter; Marrink, Siewert Published in: Journal of the American Chemical Society DOI: 10.1021/ja0624321 IMPORTANT NOTE: You are advised to consult
SUPPORTING INFORMATION. Lysine Carbonylation is a Previously Unrecognized Contributor. to Peroxidase Activation of Cytochrome c by Chloramine-T
 Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2019 SUPPORTING INFORMATION Lysine Carbonylation is a Previously Unrecognized Contributor to
Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2019 SUPPORTING INFORMATION Lysine Carbonylation is a Previously Unrecognized Contributor to
Cholesterol Modulates the Membrane Effects and Spatial Organization of Membrane-Penetrating Ligands for G-Protein Coupled Receptors
 12046 J. Phys. Chem. B 2010, 114, 12046 12057 Cholesterol Modulates the Membrane Effects and Spatial Organization of Membrane-Penetrating Ligands for G-Protein Coupled Receptors George Khelashvili,* Sayan
12046 J. Phys. Chem. B 2010, 114, 12046 12057 Cholesterol Modulates the Membrane Effects and Spatial Organization of Membrane-Penetrating Ligands for G-Protein Coupled Receptors George Khelashvili,* Sayan
The Fusion of Membranes and Vesicles: Pathway and Energy Barriers from Dissipative Particle Dynamics
 The Fusion of Membranes and Vesicles: Pathway and Energy Barriers from Dissipative Particle Dynamics Andrea Grafmüller 1 Theory and Bio-Systems, Max Planck Institute for Colloids and Interfaces, Potsdam,
The Fusion of Membranes and Vesicles: Pathway and Energy Barriers from Dissipative Particle Dynamics Andrea Grafmüller 1 Theory and Bio-Systems, Max Planck Institute for Colloids and Interfaces, Potsdam,
G protein coupled receptor interactions with cholesterol deep in the membrane
 G protein coupled receptor interactions with cholesterol deep in the membrane Samuel Genheden a,b, Jonathan W. Essex a,c, & Anthony G. Lee c,d * a School of Chemistry, University of Southampton, Southampton,
G protein coupled receptor interactions with cholesterol deep in the membrane Samuel Genheden a,b, Jonathan W. Essex a,c, & Anthony G. Lee c,d * a School of Chemistry, University of Southampton, Southampton,
Phase Behavior of a Phospholipid/Fatty Acid/Water Mixture Studied in Atomic Detail
 Published on Web 01/19/2006 Phase Behavior of a Phospholipid/Fatty Acid/Water Mixture Studied in Atomic Detail Volker Knecht,*, Alan E. Mark,, and Siewert-Jan Marrink Contribution from the Max Planck Institute
Published on Web 01/19/2006 Phase Behavior of a Phospholipid/Fatty Acid/Water Mixture Studied in Atomic Detail Volker Knecht,*, Alan E. Mark,, and Siewert-Jan Marrink Contribution from the Max Planck Institute
Molecular Structure and Permeability at the Interface between Phase-Separated Membrane Domains
 SUPPORTING INFORMATION Molecular Structure and Permeability at the Interface between Phase-Separated Membrane Domains Rodrigo M. Cordeiro Universidade Federal do ABC, Avenida dos Estados 5001, CEP 09210-580,
SUPPORTING INFORMATION Molecular Structure and Permeability at the Interface between Phase-Separated Membrane Domains Rodrigo M. Cordeiro Universidade Federal do ABC, Avenida dos Estados 5001, CEP 09210-580,
Physics of Cellular Materials: Biomembranes
 Physics of Cellular Materials: Biomembranes Tom Chou 1 1 Dept. of Biomathematics, UCL, Los ngeles, C 90095-1766 (Dated: December 6, 2002) Here I will review the mathematics and statistical physics associated
Physics of Cellular Materials: Biomembranes Tom Chou 1 1 Dept. of Biomathematics, UCL, Los ngeles, C 90095-1766 (Dated: December 6, 2002) Here I will review the mathematics and statistical physics associated
MARTINI Coarse-Grained Model of Triton TX-100 in Pure DPPC. Monolayer and Bilayer Interfaces. Supporting Information
 MARTINI Coarse-Grained Model of Triton TX-100 in Pure DPPC Monolayer and Bilayer Interfaces. Antonio Pizzirusso a, Antonio De Nicola* a, Giuseppe Milano a a Dipartimento di Chimica e Biologia, Università
MARTINI Coarse-Grained Model of Triton TX-100 in Pure DPPC Monolayer and Bilayer Interfaces. Antonio Pizzirusso a, Antonio De Nicola* a, Giuseppe Milano a a Dipartimento di Chimica e Biologia, Università
Physical Cell Biology Lecture 10: membranes elasticity and geometry. Hydrophobicity as an entropic effect
 Physical Cell Biology Lecture 10: membranes elasticity and geometry Phillips: Chapter 5, Chapter 11 and Pollard Chapter 13 Hydrophobicity as an entropic effect 1 Self-Assembly of Lipid Structures Lipid
Physical Cell Biology Lecture 10: membranes elasticity and geometry Phillips: Chapter 5, Chapter 11 and Pollard Chapter 13 Hydrophobicity as an entropic effect 1 Self-Assembly of Lipid Structures Lipid
Membranes & Membrane Proteins
 School on Biomolecular Simulations Membranes & Membrane Proteins Vani Vemparala The Institute of Mathematical Sciences Chennai November 13 2007 JNCASR, Bangalore Cellular Environment Plasma membrane extracellular
School on Biomolecular Simulations Membranes & Membrane Proteins Vani Vemparala The Institute of Mathematical Sciences Chennai November 13 2007 JNCASR, Bangalore Cellular Environment Plasma membrane extracellular
Under the Influence of Alcohol: The Effect of Ethanol and Methanol on Lipid Bilayers
 Biophysical Journal Volume 90 February 2006 1121 1135 1121 Under the Influence of Alcohol: The Effect of Ethanol and Methanol on Lipid Bilayers Michael Patra,* y Emppu Salonen, z Emma Terama, z Ilpo Vattulainen,
Biophysical Journal Volume 90 February 2006 1121 1135 1121 Under the Influence of Alcohol: The Effect of Ethanol and Methanol on Lipid Bilayers Michael Patra,* y Emppu Salonen, z Emma Terama, z Ilpo Vattulainen,
Structure of Dipalmitoylphosphatidylcholine/Cholesterol Bilayer at Low and High Cholesterol Concentrations: Molecular Dynamics Simulation
 Biophysical Journal Volume 77 October 1999 2075 2089 2075 Structure of Dipalmitoylphosphatidylcholine/Cholesterol Bilayer at Low and High Cholesterol Concentrations: Molecular Dynamics Simulation Alexander
Biophysical Journal Volume 77 October 1999 2075 2089 2075 Structure of Dipalmitoylphosphatidylcholine/Cholesterol Bilayer at Low and High Cholesterol Concentrations: Molecular Dynamics Simulation Alexander
Phase Behavior of Model Lipid Bilayers
 J. Phys. Chem. B 2005, 109, 6553-6563 6553 Phase Behavior of Model Lipid Bilayers Marieke Kranenburg and Berend Smit*,, The Van t Hoff Institute for Molecular Sciences, UniVersity of Amsterdam, Nieuwe
J. Phys. Chem. B 2005, 109, 6553-6563 6553 Phase Behavior of Model Lipid Bilayers Marieke Kranenburg and Berend Smit*,, The Van t Hoff Institute for Molecular Sciences, UniVersity of Amsterdam, Nieuwe
Reading for lecture 6
 Reading for lecture 6 1. Lipids and Lipid Bilayers 2. Membrane Proteins Voet and Voet, Chapter 11 Alberts et al Chapter 6 Jones, R.A.L, Soft Condensed Matter 195pp Oxford University Press, ISBN 0-19-850590-6
Reading for lecture 6 1. Lipids and Lipid Bilayers 2. Membrane Proteins Voet and Voet, Chapter 11 Alberts et al Chapter 6 Jones, R.A.L, Soft Condensed Matter 195pp Oxford University Press, ISBN 0-19-850590-6
Distribution of Amino Acids in a Lipid Bilayer from Computer Simulations
 Biophysical Journal Volume 94 May 2008 3393 3404 3393 Distribution of Amino Acids in a Lipid Bilayer from Computer Simulations Justin L. MacCallum, W. F. Drew Bennett, and D. Peter Tieleman Department
Biophysical Journal Volume 94 May 2008 3393 3404 3393 Distribution of Amino Acids in a Lipid Bilayer from Computer Simulations Justin L. MacCallum, W. F. Drew Bennett, and D. Peter Tieleman Department
Lessons of Slicing Membranes: Interplay of Packing, Free Area, and Lateral Diffusion in Phospholipid/Cholesterol Bilayers
 1076 Biophysical Journal Volume 87 August 2004 1076 1091 Lessons of Slicing Membranes: Interplay of Packing, Free Area, and Lateral Diffusion in Phospholipid/Cholesterol Bilayers Emma Falck,* Michael Patra,
1076 Biophysical Journal Volume 87 August 2004 1076 1091 Lessons of Slicing Membranes: Interplay of Packing, Free Area, and Lateral Diffusion in Phospholipid/Cholesterol Bilayers Emma Falck,* Michael Patra,
Chemical Surface Transformation 1
 Chemical Surface Transformation 1 Chemical reactions at Si H surfaces (inorganic and organic) can generate very thin films (sub nm thickness up to µm): inorganic layer formation by: thermal conversion:
Chemical Surface Transformation 1 Chemical reactions at Si H surfaces (inorganic and organic) can generate very thin films (sub nm thickness up to µm): inorganic layer formation by: thermal conversion:
Nanoparticle Permeation Induces Water Penetration, Ion Transport, and Lipid Flip-Flop
 pubs.acs.org/langmuir Nanoparticle Permeation Induces Water Penetration, Ion Transport, and Lipid Flip-Flop Bo Song, Huajun Yuan, Sydney V. Pham, Cynthia J. Jameson, and Sohail Murad* Department of Chemical
pubs.acs.org/langmuir Nanoparticle Permeation Induces Water Penetration, Ion Transport, and Lipid Flip-Flop Bo Song, Huajun Yuan, Sydney V. Pham, Cynthia J. Jameson, and Sohail Murad* Department of Chemical
level and yield the relevant dynamical and structural information.
 Lecture 1 ATOMIC LEVEL SIMULATIONS OF MEMBRANES A fundamental understanding of the properties of biological membranes and membrane proteins from the atomic point of view is undoubtedly of great biochemical,
Lecture 1 ATOMIC LEVEL SIMULATIONS OF MEMBRANES A fundamental understanding of the properties of biological membranes and membrane proteins from the atomic point of view is undoubtedly of great biochemical,
SUPPLEMENTARY INFORMATION
 High-speed atomic force microscopy shows that annexin V stabilizes membranes on the second timescale Atsushi Miyagi, Chris Chipot, Martina Rangl & Simon Scheuring Supplementary Movies: Supplementary Movie
High-speed atomic force microscopy shows that annexin V stabilizes membranes on the second timescale Atsushi Miyagi, Chris Chipot, Martina Rangl & Simon Scheuring Supplementary Movies: Supplementary Movie
Interaction Between Amyloid-b (1 42) Peptide and Phospholipid Bilayers: A Molecular Dynamics Study
 Biophysical Journal Volume 96 February 2009 785 797 785 Interaction Between Amyloid-b (1 42) Peptide and Phospholipid Bilayers: A Molecular Dynamics Study Charles H. Davis and Max L. Berkowitz * Department
Biophysical Journal Volume 96 February 2009 785 797 785 Interaction Between Amyloid-b (1 42) Peptide and Phospholipid Bilayers: A Molecular Dynamics Study Charles H. Davis and Max L. Berkowitz * Department
Lipid Bilayers Are Excellent For Cell Membranes
 Lipid Bilayers Are Excellent For Cell Membranes ydrophobic interaction is the driving force Self-assembly in water Tendency to close on themselves Self-sealing (a hole is unfavorable) Extensive: up to
Lipid Bilayers Are Excellent For Cell Membranes ydrophobic interaction is the driving force Self-assembly in water Tendency to close on themselves Self-sealing (a hole is unfavorable) Extensive: up to
H 2 O. Liquid, solid, and vapor coexist in the same environment
 Water H 2 O Liquid, solid, and vapor coexist in the same environment WATER MOLECULES FORM HYDROGEN BONDS Water is a fundamental requirement for life, so it is important to understand the structural and
Water H 2 O Liquid, solid, and vapor coexist in the same environment WATER MOLECULES FORM HYDROGEN BONDS Water is a fundamental requirement for life, so it is important to understand the structural and
A proposed mechanism for tear film breakup: a molecular level view by employing in-silico approach
 Journal for Modeling in Ophthalmology 2018; 1:19-23 Meeting highlights article: ARVO 2017 A proposed mechanism for tear film breakup: a molecular level view by employing in-silico approach Lukasz Cwiklik
Journal for Modeling in Ophthalmology 2018; 1:19-23 Meeting highlights article: ARVO 2017 A proposed mechanism for tear film breakup: a molecular level view by employing in-silico approach Lukasz Cwiklik
Four-Scale Description of Membrane Sculpting by BAR Domains
 2806 Biophysical Journal Volume 95 September 2008 2806 2821 Four-Scale Description of Membrane Sculpting by BAR Domains Anton Arkhipov, Ying Yin, and Klaus Schulten Department of Physics and Beckman Institute,
2806 Biophysical Journal Volume 95 September 2008 2806 2821 Four-Scale Description of Membrane Sculpting by BAR Domains Anton Arkhipov, Ying Yin, and Klaus Schulten Department of Physics and Beckman Institute,
Week 5 Section. Junaid Malek, M.D.
 Week 5 Section Junaid Malek, M.D. HIV: Anatomy Membrane (partiallystolen from host cell) 2 Glycoproteins (proteins modified by added sugar) 2 copies of RNA Capsid HIV Genome Encodes: Structural Proteins
Week 5 Section Junaid Malek, M.D. HIV: Anatomy Membrane (partiallystolen from host cell) 2 Glycoproteins (proteins modified by added sugar) 2 copies of RNA Capsid HIV Genome Encodes: Structural Proteins
