Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation
|
|
|
- Carmel Walton
- 5 years ago
- Views:
Transcription
1 Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation Hang Yu, 1, 2, a) Wei Han, 1, 3, b) Wen Ma, 1, 2 1, 2, 3, c) and Klaus Schulten 1) Beckman Institute for Advanced Science and Technology, University of Illinois at Urbana-Champaign, Urbana, Illinois 61801, United States 2) Center for Biophysics and Computational Biology, University of Illinois at Urbana-Champaign, Urbana, Illinois 61801, United States 3) Department of Physics, University of Illinois at Urbana-Champaign, Urbana, Illinois 61801, United States (Dated: 16 November 2015) a) Contributed equally to this work b) Contributed equally to this work; Current address: School of Chemical Biology and Biotechnology, Peking University Shenzhen Graduate School, Shenzhen, Guangdong, , China c) Author to whom correspondence should be addressed. Electronic mail: kschulte@ks.uiuc.edu
2 FIG. 1. Secondary structure content and conformations of initial structures. A, B. Secondary structure content in starting structures for α-synuclein simulations (TABLE I). Percentage of α-helix, β-sheet (A) and turn (B) for 10 starting structures of WT α-synuclein simulations are shown in blue diamonds, red squares and green triangles, respectively. Secondary structure contents were analyzed using VMD 1. C. Conformations of the ten initial structures employed for simulations in the present study. Shown are the backbone atoms of the β-hairpin region (region 38-53) in cartoon (gold) and licorice representations (carbon atoms in light blue, nitrogen in dark blue, oxygen in red and hydrogen in white). 2
3 FIG. 2. Individual and accumulated contribution of the first hundred principal components from PCA analysis of simulated WT α-synuclein. Plotted in red dashed line and black dots are the individual and accumulated contributions of structural variation, respectively, of each principal component. 3
4 FIG. 3. Representative structures of the five most populated conformational clusters for WT α-synuclein. N-terminus, NAC region and C-terminus are colored blue, pink and red, respectively. The same coloring scheme is applied to further figures of this study. Cluster 2, representing 5.10 % of all conformations, exhibits the formed β-hairpin (see green circle); however, the largest cluster, cluster 1, also exhibits a β-hairpin motif (red circle), but one not yet formed well. 4
5 FIG. 4. Hairpin RMSD and representative structures of WT α-synuclein and its A30P, A53T mutants. Plotted are the RMSD values of the β-hairpin region (region 38-53) of WT α-synuclein and its A30P, A53T mutants (in red, purple and green, respectively) over simulation time. Representative structures of the WT α-synuclein and its A30P, A53T mutants are shown on the right. The center structure from the largest cluster of the WT α-synuclein β-hairpin region (region 38-53) was selected as a representative structure for WT α-synuclein. Representative structures of A30P, A53T mutant α-synuclein were obtained with the same procedure in the case of mutant simulations. Backbone atoms of the β-hairpin region (region 38-53) were selected as reference atoms for alignment and RMSD calculation. Individual simulations are separated by dotted lines. The cutoffs defining folding (3.25 Å) and unfolding (5.00 Å) are shown as black lines. A β-hairpin folding event is defined as a transition of the β-hairpin region (region 38-53) from an unfolded state (RMSD > 5.00 Å) to a folded state (RMSD < 3.25 Å). 5
6 FIG. 5. Residual secondary structure content in WT α-synuclein and its largest cluster, cluster 1. Fractions of α-helix, β-sheet and turn are shown in the top, middle and bottom panels, respectively. The traces for WT α-synuclein and its cluster 1 are colored red and blue, respectively. The x-axis giving residue numbers is also shown as a chain of blue, gold, pink and red bars as defined in FIG. 1. 6
7 FIG. 6. Hairpin RMSD values arising in simulations of the isolated β-hairpin region (region 38-53) for WT α-synuclein and its G47V mutant. The RMSD values are plotted over simulation time in gold (WT) and blue (G47V), respectively. Backbone atoms of the β-hairpin region (region 38-53) were selected as reference atoms for alignment and RMSD calculation. Individual simulations are separated by dotted lines. The cutoffs defining folding (3.25 Å) and unfolding (5.00 Å) are shown as black lines. A β-hairpin folding event is defined as a transition of the β-hairpin region (region 38-53) from an unfolded state (RMSD > 5.00 Å) to a folded state (RMSD < 3.25 Å). 7
8 FIG. 7. Contact map of WT α-synuclein. Contacts between β-hairpin structure (region 38-53) and C-terminus (region ) are circled in green (see also FIG. 3, green circle). The probability of contact between residues i and j within the α-synuclein monomer was calculated as the probability for the centers of mass of the two residues to be within a distance of 8.5 Å. Free energy (G) is calculated from residual contact probabilities (P) as G = -k BT ln(p). Plotted are the contact probabilities from most favorable contact to least favorable contact in terms of relative free energy (kcal/mol). Colors are defined here, as seen in the color bar on the right, in terms of relative free energy (kcal/mol) colored from dark blue to yellow. The x-axis of the contact map is shown as a chain of blue, gold, pink and red bars denoting the positions of N-terminus, β-hairpin region, NAC region and C-terminus on the map, respectively, as defined in FIG. 1. Contacts between adjacent residues are removed for clarity. 8
9 9 FIG. 8. Stability of interactions between C-terminus and β-hairpin structure in WT α-synuclein and its A30P and A53T mutants. Starting from the center structure of the conformational ensemble containing the β-hairpins (region 38-53) in contact with the C-terminus (region , FIG. 1B), ten 200-ns simulations were performed for WT α-synuclein and its A30P and A53T mutants (TABLE II, InteractionWT, InteractionA30P and InteractionA53T). Final frames of the 10 simulations are shown in cartoon representation. β-hairpin and C-terminus are shown in gold and red, respectively; the remainder of α-synuclein is shown in grey. The interactions between β-hairpin structure and C-terminus are maintained in most cases (WT: 10/10, A30P: 9/10, A53T: 9/10). Circled in green are the simulations in which the interactions between β-hairpin structure and C-terminus got lost.
10 FIG. 9. Ramachandran plot of glycine 47 in the isolated β-hairpin region. This plot illustrates residual structural conformations through backbone dihedral angles φ and ψ. Distributions of these angles indicate favorable conformations of glycine 47. Plotted are the dihedral angles φ and ψ according to if the angles belong to the most favorable conformation or to the least favorable conformation. Colors are defined here, as seen in the color bar on the right, in terms of relative free energy (kcal/mol) colored from dark red to white. Free energy (G) is calculated from angle distribution probabilities (P) as G = -k BT ln(p). Regions on the Ramachandran plot favoring α-helices and left-hand-α-helical conformations are denoted as α and L α, respectively. 10
11 FIG. 10. Ramachandran plot of valine 47 in the isolated β-hairpin region with G47V mutation. This plot illustrates residual structural conformations through backbone dihedral angles φ and ψ. Distributions of these angles indicate favorable conformations of valine 47. Plotted are the dihedral angles φ and ψ for the valine side group at amino acid sequence position 47. Colors are defined as in FIG. S9. Regions on the Ramachandran plot favoring α-helices and β-sheet conformations are denoted as α and β, respectively. 11
12 12 FIG. 11. Residual secondary structure content in isolated β-hairpin region (region 38-53) for WT α-synuclein in water and in the presence of salt. Fractions of α-helix, β-sheet and turn are shown in the top, middle and bottom panels, respectively. The traces for isolated β-hairpin region (region 38-53) for WT α-synuclein in water and in the presence of 0.15M NaCl physiological salt concentration are colored orange and blue, respectively.
13 FIG. 12. Hairpin RMSD values arising in simulations of the isolated β-hairpin region (region 38-53) for WT α-synuclein in water and in the presence of salt. The RMSD values are plotted over simulation time in blue (in the presence of 0.15M NaCl physiological salt concentration). Backbone atoms of the β-hairpin region (region 38-53) were selected as reference atoms for alignment and RMSD calculation. Individual simulations are separated by dotted lines. The cutoffs defining folding (3.25 Å) and unfolding (5.00 Å) are shown as black lines. A β-hairpin folding event is defined as a transition of the β-hairpin region (region 38-53) from an unfolded state (RMSD > 5.00 Å) to a folded state (RMSD < 3.25 Å). 13
14 14 FIG. 13. Secondary structure content and auto-correlation time of end-to-end distance for WT α-synuclein simulations. A, B. Secondary structure content for α-synuclein simulations (TABLE I). Percentage of α-helix, β-sheet (A) and turn (B) averaged over 10 WT α-synuclein simulations are shown in blue diamonds, red squares and green triangles, respectively. Standard deviations are shown in error bars. C. Auto-correlation time of the WT α-synuclein end-to-end distance. For each of the 10 WT α-synuclein simulations, the end-to-end distance was defined as the distance between C α atoms of residue 1 and 140. The distance was monitored in each simulation over time and fitted into the Python ACOR package 2 to obtain the auto-correlation time.
15 FIG. 14. Secondary structure content and auto-correlation time of end-to-end distance for simulated α-synuclein A30P mutant. A, B. Secondary structure content for α-synuclein A30P mutant (TABLE I). Percentage of α-helix, β-sheet (A) and turn (B) averaged over 10 α-synuclein A30P mutant simulations are shown in blue diamonds, red squares and green triangles, respectively. Standard deviations are shown in error bars. C. Auto-correlation time of the α-synuclein A30P mutant end-to-end distance. 15
16 FIG. 15. Secondary structure content and auto-correlation time of end-to-end distance for α-synuclein simulated A53T mutant. A, B. Secondary structure content for α-synuclein A53T mutant (TABLE I). Percentage of α-helix, β-sheet (A) and turn (B) averaged over 10 α-synuclein A53T mutant simulations are shown in blue diamonds, red squares and green triangles, respectively. Standard deviations are shown in error bars. C. Auto-correlation time of the α-synuclein A53T mutant end-to-end distance. 16
17 17 1 W. Humphrey, A. Dalke, and K. Schulten, VMD Visual Molecular Dynamics, J. Mol. Graphics 14, (1996). 2 A. Sokal, Monte Carlo methods in statistical mechanics: foundations and new algorithms (Springer, 1997).
CS612 - Algorithms in Bioinformatics
 Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site.
 Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/4/3/eaaq0762/dc1 Supplementary Materials for Structures of monomeric and oligomeric forms of the Toxoplasma gondii perforin-like protein 1 Tao Ni, Sophie I. Williams,
advances.sciencemag.org/cgi/content/full/4/3/eaaq0762/dc1 Supplementary Materials for Structures of monomeric and oligomeric forms of the Toxoplasma gondii perforin-like protein 1 Tao Ni, Sophie I. Williams,
Table S1: Kinetic parameters of drug and substrate binding to wild type and HIV-1 protease variants. Data adapted from Ref. 6 in main text.
 Dynamical Network of HIV-1 Protease Mutants Reveals the Mechanism of Drug Resistance and Unhindered Activity Rajeswari Appadurai and Sanjib Senapati* BJM School of Biosciences and Department of Biotechnology,
Dynamical Network of HIV-1 Protease Mutants Reveals the Mechanism of Drug Resistance and Unhindered Activity Rajeswari Appadurai and Sanjib Senapati* BJM School of Biosciences and Department of Biotechnology,
Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular
 Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular weight markers (M). Supplementary Figure-2. Overlay of
Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular weight markers (M). Supplementary Figure-2. Overlay of
Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random
 S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is worth 2 points.
 MBB 407/511 Molecular Biology and Biochemistry First Examination - October 1, 2002 Name Social Security Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is
MBB 407/511 Molecular Biology and Biochemistry First Examination - October 1, 2002 Name Social Security Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is
SUPPLEMENTARY INFORMATION. Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity
 SUPPLEMENTARY INFORMATION Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity Christopher Woods 1, Maturos Malaisree 1, Ben Long 2, Simon McIntosh-Smith
SUPPLEMENTARY INFORMATION Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity Christopher Woods 1, Maturos Malaisree 1, Ben Long 2, Simon McIntosh-Smith
Secondary Structure. by hydrogen bonds
 Secondary Structure In the previous protein folding activity, you created a hypothetical 15-amino acid protein and learned that basic principles of chemistry determine how each protein spontaneously folds
Secondary Structure In the previous protein folding activity, you created a hypothetical 15-amino acid protein and learned that basic principles of chemistry determine how each protein spontaneously folds
SDS-Assisted Protein Transport Through Solid-State Nanopores
 Supplementary Information for: SDS-Assisted Protein Transport Through Solid-State Nanopores Laura Restrepo-Pérez 1, Shalini John 2, Aleksei Aksimentiev 2 *, Chirlmin Joo 1 *, Cees Dekker 1 * 1 Department
Supplementary Information for: SDS-Assisted Protein Transport Through Solid-State Nanopores Laura Restrepo-Pérez 1, Shalini John 2, Aleksei Aksimentiev 2 *, Chirlmin Joo 1 *, Cees Dekker 1 * 1 Department
Secondary Structure North 72nd Street, Wauwatosa, WI Phone: (414) Fax: (414) dmoleculardesigns.com
 Secondary Structure In the previous protein folding activity, you created a generic or hypothetical 15-amino acid protein and learned that basic principles of chemistry determine how each protein spontaneously
Secondary Structure In the previous protein folding activity, you created a generic or hypothetical 15-amino acid protein and learned that basic principles of chemistry determine how each protein spontaneously
Lecture 10 More about proteins
 Lecture 10 More about proteins Today we're going to extend our discussion of protein structure. This may seem far-removed from gene cloning, but it is the path to understanding the genes that we are cloning.
Lecture 10 More about proteins Today we're going to extend our discussion of protein structure. This may seem far-removed from gene cloning, but it is the path to understanding the genes that we are cloning.
Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.
 Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.5 % SDS-PAGE gel under reducing and non-reducing conditions.
Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.5 % SDS-PAGE gel under reducing and non-reducing conditions.
Unveiling transient protein-protein interactions that modulate inhibition of alpha-synuclein aggregation
 Supplementary information Unveiling transient protein-protein interactions that modulate inhibition of alpha-synuclein aggregation by beta-synuclein, a pre-synaptic protein that co-localizes with alpha-synuclein.
Supplementary information Unveiling transient protein-protein interactions that modulate inhibition of alpha-synuclein aggregation by beta-synuclein, a pre-synaptic protein that co-localizes with alpha-synuclein.
Introduction to Protein Structure Collection
 Introduction to Protein Structure Collection Teaching Points This collection is designed to introduce students to the concepts of protein structure and biochemistry. Different activities guide students
Introduction to Protein Structure Collection Teaching Points This collection is designed to introduce students to the concepts of protein structure and biochemistry. Different activities guide students
Copyright Mark Brandt, Ph.D. 46
 Examples of tein Structures tein types teins fall into three general classes, based on their overall three-dimensional structure and on their functional role: fibrous, membrane, and globular. Fibrous proteins
Examples of tein Structures tein types teins fall into three general classes, based on their overall three-dimensional structure and on their functional role: fibrous, membrane, and globular. Fibrous proteins
Supporting Information Identification of Amino Acids with Sensitive Nanoporous MoS 2 : Towards Machine Learning-Based Prediction
 Supporting Information Identification of Amino Acids with Sensitive Nanoporous MoS 2 : Towards Machine Learning-Based Prediction Amir Barati Farimani, Mohammad Heiranian, Narayana R. Aluru 1 Department
Supporting Information Identification of Amino Acids with Sensitive Nanoporous MoS 2 : Towards Machine Learning-Based Prediction Amir Barati Farimani, Mohammad Heiranian, Narayana R. Aluru 1 Department
Activities for the α-helix / β-sheet Construction Kit
 Activities for the α-helix / β-sheet Construction Kit The primary sequence of a protein, composed of amino acids, determines the organization of the sequence into the secondary structure. There are two
Activities for the α-helix / β-sheet Construction Kit The primary sequence of a protein, composed of amino acids, determines the organization of the sequence into the secondary structure. There are two
BCH Graduate Survey of Biochemistry
 BCH 5045 Graduate Survey of Biochemistry Instructor: Charles Guy Producer: Ron Thomas Director: Glen Graham Lecture 10 Slide sets available at: http://hort.ifas.ufl.edu/teach/guyweb/bch5045/index.html
BCH 5045 Graduate Survey of Biochemistry Instructor: Charles Guy Producer: Ron Thomas Director: Glen Graham Lecture 10 Slide sets available at: http://hort.ifas.ufl.edu/teach/guyweb/bch5045/index.html
(B D) Three views of the final refined 2Fo-Fc electron density map of the Vpr (red)-ung2 (green) interacting region, contoured at 1.4σ.
 Supplementary Figure 1 Overall structure of the DDB1 DCAF1 Vpr UNG2 complex. (A) The final refined 2Fo-Fc electron density map, contoured at 1.4σ of Vpr, illustrating well-defined side chains. (B D) Three
Supplementary Figure 1 Overall structure of the DDB1 DCAF1 Vpr UNG2 complex. (A) The final refined 2Fo-Fc electron density map, contoured at 1.4σ of Vpr, illustrating well-defined side chains. (B D) Three
Protein Secondary Structure
 Protein Secondary Structure Reading: Berg, Tymoczko & Stryer, 6th ed., Chapter 2, pp. 37-45 Problems in textbook: chapter 2, pp. 63-64, #1,5,9 Directory of Jmol structures of proteins: http://www.biochem.arizona.edu/classes/bioc462/462a/jmol/routines/routines.html
Protein Secondary Structure Reading: Berg, Tymoczko & Stryer, 6th ed., Chapter 2, pp. 37-45 Problems in textbook: chapter 2, pp. 63-64, #1,5,9 Directory of Jmol structures of proteins: http://www.biochem.arizona.edu/classes/bioc462/462a/jmol/routines/routines.html
BIO 311C Spring Lecture 15 Friday 26 Feb. 1
 BIO 311C Spring 2010 Lecture 15 Friday 26 Feb. 1 Illustration of a Polypeptide amino acids peptide bonds Review Polypeptide (chain) See textbook, Fig 5.21, p. 82 for a more clear illustration Folding and
BIO 311C Spring 2010 Lecture 15 Friday 26 Feb. 1 Illustration of a Polypeptide amino acids peptide bonds Review Polypeptide (chain) See textbook, Fig 5.21, p. 82 for a more clear illustration Folding and
HOMEWORK II and Swiss-PDB Viewer Tutorial DUE 9/26/03 62 points total. The ph at which a peptide has no net charge is its isoelectric point.
 BIOCHEMISTRY I HOMEWORK II and Swiss-PDB Viewer Tutorial DUE 9/26/03 62 points total 1). 8 points total T or F (2 points each; if false, briefly state why it is false) The ph at which a peptide has no
BIOCHEMISTRY I HOMEWORK II and Swiss-PDB Viewer Tutorial DUE 9/26/03 62 points total 1). 8 points total T or F (2 points each; if false, briefly state why it is false) The ph at which a peptide has no
Draw how two amino acids form the peptide bond. Draw in the space below:
 Name Date Period Modeling Protein Folding Draw how two amino acids form the peptide bond. Draw in the space below: What we are doing today: The core idea in life sciences is that there is a fundamental
Name Date Period Modeling Protein Folding Draw how two amino acids form the peptide bond. Draw in the space below: What we are doing today: The core idea in life sciences is that there is a fundamental
Amino Acids. Review I: Protein Structure. Amino Acids: Structures. Amino Acids (contd.) Rajan Munshi
 Review I: Protein Structure Rajan Munshi BBSI @ Pitt 2005 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2005 Amino Acids Building blocks of proteins 20 amino acids
Review I: Protein Structure Rajan Munshi BBSI @ Pitt 2005 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2005 Amino Acids Building blocks of proteins 20 amino acids
Adaptable Lipid Matrix Promotes Protein Protein Association in Membranes
 Supporting information Adaptable Lipid Matrix Promotes Protein Protein Association in Membranes Andrey S. Kuznetsov, Anton A. Polyansky,, Markus Fleck, Pavel E. Volynsky, and Roman G. Efremov *,, M. M.
Supporting information Adaptable Lipid Matrix Promotes Protein Protein Association in Membranes Andrey S. Kuznetsov, Anton A. Polyansky,, Markus Fleck, Pavel E. Volynsky, and Roman G. Efremov *,, M. M.
Introduction to proteins and protein structure
 Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
Supplementary Material
 Supplementary Material Materials and methods Enzyme assay The enzymatic activity of -glucosidase toward salicin was measured with the Miller method (Miller, 1959) using glucose as the standard. A total
Supplementary Material Materials and methods Enzyme assay The enzymatic activity of -glucosidase toward salicin was measured with the Miller method (Miller, 1959) using glucose as the standard. A total
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Design of isolated protein and RNC constructs, and homogeneity of purified RNCs. (a) Schematic depicting the design and nomenclature used for all the isolated proteins and RNCs used
Supplementary Figure 1 Design of isolated protein and RNC constructs, and homogeneity of purified RNCs. (a) Schematic depicting the design and nomenclature used for all the isolated proteins and RNCs used
List of Figures. List of Tables
 Supporting Information for: Signaling Domain of Sonic Hedgehog as Cannibalistic Calcium-Regulated Zinc-Peptidase Rocio Rebollido-Rios 1, Shyam Bandari 3, Christoph Wilms 1, Stanislav Jakuschev 1, Andrea
Supporting Information for: Signaling Domain of Sonic Hedgehog as Cannibalistic Calcium-Regulated Zinc-Peptidase Rocio Rebollido-Rios 1, Shyam Bandari 3, Christoph Wilms 1, Stanislav Jakuschev 1, Andrea
Supporting material. Membrane permeation induced by aggregates of human islet amyloid polypeptides
 Supporting material Membrane permeation induced by aggregates of human islet amyloid polypeptides Chetan Poojari Forschungszentrum Jülich GmbH, Institute of Complex Systems: Structural Biochemistry (ICS-6),
Supporting material Membrane permeation induced by aggregates of human islet amyloid polypeptides Chetan Poojari Forschungszentrum Jülich GmbH, Institute of Complex Systems: Structural Biochemistry (ICS-6),
Molecular Graphics Perspective of Protein Structure and Function
 Molecular Graphics Perspective of Protein Structure and Function VMD Highlights > 20,000 registered Users Platforms: Unix (16 builds) Windows MacOS X Display of large biomolecules and simulation trajectories
Molecular Graphics Perspective of Protein Structure and Function VMD Highlights > 20,000 registered Users Platforms: Unix (16 builds) Windows MacOS X Display of large biomolecules and simulation trajectories
Insights into the Giardia intestinalis Enolase and Human Plasminogen interaction
 Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Supplementary Information Insights into the Giardia intestinalis Enolase and Human
Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Supplementary Information Insights into the Giardia intestinalis Enolase and Human
Rapid Characterization of Allosteric Networks. with Ensemble Normal Mode Analysis
 Supporting Information Rapid Characterization of Allosteric Networks with Ensemble Normal Mode Analysis Xin-Qiu Yao 1, Lars Skjærven 2, and Barry J. Grant 1* 1 Department of Computational Medicine and
Supporting Information Rapid Characterization of Allosteric Networks with Ensemble Normal Mode Analysis Xin-Qiu Yao 1, Lars Skjærven 2, and Barry J. Grant 1* 1 Department of Computational Medicine and
Supplementary Information A Hydrophobic Barrier Deep Within the Inner Pore of the TWIK-1 K2P Potassium Channel Aryal et al.
 Supplementary Information A Hydrophobic Barrier Deep Within the Inner Pore of the TWIK-1 K2P Potassium Channel Aryal et al. Supplementary Figure 1 TWIK-1 stability during MD simulations in a phospholipid
Supplementary Information A Hydrophobic Barrier Deep Within the Inner Pore of the TWIK-1 K2P Potassium Channel Aryal et al. Supplementary Figure 1 TWIK-1 stability during MD simulations in a phospholipid
Bioinformatics for molecular biology
 Bioinformatics for molecular biology Structural bioinformatics tools, predictors, and 3D modeling Structural Biology Review Dr Research Scientist Department of Microbiology, Oslo University Hospital -
Bioinformatics for molecular biology Structural bioinformatics tools, predictors, and 3D modeling Structural Biology Review Dr Research Scientist Department of Microbiology, Oslo University Hospital -
Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase. Biol 405 Molecular Medicine
 Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase Biol 405 Molecular Medicine 1998 Crystal structure of phenylalanine hydroxylase solved. The polypeptide consists of three regions: Regulatory
Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase Biol 405 Molecular Medicine 1998 Crystal structure of phenylalanine hydroxylase solved. The polypeptide consists of three regions: Regulatory
Multiple-Choice Questions Answer ALL 20 multiple-choice questions on the Scantron Card in PENCIL
 Multiple-Choice Questions Answer ALL 20 multiple-choice questions on the Scantron Card in PENCIL For Questions 1-10 choose ONE INCORRECT answer. 1. Which ONE of the following statements concerning the
Multiple-Choice Questions Answer ALL 20 multiple-choice questions on the Scantron Card in PENCIL For Questions 1-10 choose ONE INCORRECT answer. 1. Which ONE of the following statements concerning the
SUPPORTING INFORMATION FOR. A Computational Approach to Enzyme Design: Using Docking and MM- GBSA Scoring
 SUPPRTING INFRMATIN FR A Computational Approach to Enzyme Design: Predicting ω- Aminotransferase Catalytic Activity Using Docking and MM- GBSA Scoring Sarah Sirin, 1 Rajesh Kumar, 2 Carlos Martinez, 2
SUPPRTING INFRMATIN FR A Computational Approach to Enzyme Design: Predicting ω- Aminotransferase Catalytic Activity Using Docking and MM- GBSA Scoring Sarah Sirin, 1 Rajesh Kumar, 2 Carlos Martinez, 2
Biology 2E- Zimmer Protein structure- amino acid kit
 Biology 2E- Zimmer Protein structure- amino acid kit Name: This activity will use a physical model to investigate protein shape and develop key concepts that govern how proteins fold into their final three-dimensional
Biology 2E- Zimmer Protein structure- amino acid kit Name: This activity will use a physical model to investigate protein shape and develop key concepts that govern how proteins fold into their final three-dimensional
Lecture 15. Membrane Proteins I
 Lecture 15 Membrane Proteins I Introduction What are membrane proteins and where do they exist? Proteins consist of three main classes which are classified as globular, fibrous and membrane proteins. A
Lecture 15 Membrane Proteins I Introduction What are membrane proteins and where do they exist? Proteins consist of three main classes which are classified as globular, fibrous and membrane proteins. A
Supplementary Materials. High affinity binding of phosphatidylinositol-4-phosphate. by Legionella pneumophila DrrA
 Supplementary Materials High affinity binding of phosphatidylinositol-4-phosphate by Legionella pneumophila DrrA Running title: Molecular basis of PtdIns(4)P-binding by DrrA Stefan Schoebel, Wulf Blankenfeldt,
Supplementary Materials High affinity binding of phosphatidylinositol-4-phosphate by Legionella pneumophila DrrA Running title: Molecular basis of PtdIns(4)P-binding by DrrA Stefan Schoebel, Wulf Blankenfeldt,
Hemoglobin & Sickle Cell Anemia Exercise
 Name StarBiochem Hemoglobin & Sickle Cell Anemia Exercise Learning Objectives In this exercise, you will use StarBiochem, a protein 3D viewer, to explore the structure of the normal hemoglobin protein
Name StarBiochem Hemoglobin & Sickle Cell Anemia Exercise Learning Objectives In this exercise, you will use StarBiochem, a protein 3D viewer, to explore the structure of the normal hemoglobin protein
Interactions of Polyethylenimines with Zwitterionic and. Anionic Lipid Membranes
 Interactions of Polyethylenimines with Zwitterionic and Anionic Lipid Membranes Urszula Kwolek, Dorota Jamróz, Małgorzata Janiczek, Maria Nowakowska, Paweł Wydro, Mariusz Kepczynski Faculty of Chemistry,
Interactions of Polyethylenimines with Zwitterionic and Anionic Lipid Membranes Urszula Kwolek, Dorota Jamróz, Małgorzata Janiczek, Maria Nowakowska, Paweł Wydro, Mariusz Kepczynski Faculty of Chemistry,
Catalysis & specificity: Proteins at work
 Catalysis & specificity: Proteins at work Introduction Having spent some time looking at the elements of structure of proteins and DNA, as well as their ability to form intermolecular interactions, it
Catalysis & specificity: Proteins at work Introduction Having spent some time looking at the elements of structure of proteins and DNA, as well as their ability to form intermolecular interactions, it
obtained for the simulations of the E2 conformation of SERCA in a pure POPC lipid bilayer (blue) and in a
 Supplementary Figure S1. Distribution of atoms along the bilayer normal. Normalized density profiles obtained for the simulations of the E2 conformation of SERCA in a pure POPC lipid bilayer (blue) and
Supplementary Figure S1. Distribution of atoms along the bilayer normal. Normalized density profiles obtained for the simulations of the E2 conformation of SERCA in a pure POPC lipid bilayer (blue) and
Supplementary Materials for
 www.sciencemag.org/cgi/content/full/science.aal4326/dc1 Supplementary Materials for Structure of a eukaryotic voltage-gated sodium channel at near-atomic resolution Huaizong Shen, Qiang Zhou, Xiaojing
www.sciencemag.org/cgi/content/full/science.aal4326/dc1 Supplementary Materials for Structure of a eukaryotic voltage-gated sodium channel at near-atomic resolution Huaizong Shen, Qiang Zhou, Xiaojing
Nature Structural & Molecular Biology: doi: /nsmb.1933
 The structural basis of open channel block in a prokaryotic pentameric ligand-gated ion channel Ricarda J. C. Hilf, Carlo Bertozzi, Iwan Zimmermann, Alwin Reiter, Dirk Trauner and Raimund Dutzler a GLIC
The structural basis of open channel block in a prokaryotic pentameric ligand-gated ion channel Ricarda J. C. Hilf, Carlo Bertozzi, Iwan Zimmermann, Alwin Reiter, Dirk Trauner and Raimund Dutzler a GLIC
H C. C α. Proteins perform a vast array of biological function including: Side chain
 Topics The topics: basic concepts of molecular biology elements on Python overview of the field biological databases and database searching sequence alignments phylogenetic trees microarray data analysis
Topics The topics: basic concepts of molecular biology elements on Python overview of the field biological databases and database searching sequence alignments phylogenetic trees microarray data analysis
Term Definition Example Amino Acids
 Name 1. What are some of the functions that proteins have in a living organism. 2. Define the following and list two amino acids that fit each description. Term Definition Example Amino Acids Hydrophobic
Name 1. What are some of the functions that proteins have in a living organism. 2. Define the following and list two amino acids that fit each description. Term Definition Example Amino Acids Hydrophobic
Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels. Supplementary Information
 Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels Craig T. Armstrong, Philip E. Mason, J. L. Ross Anderson and Christopher E. Dempsey
Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels Craig T. Armstrong, Philip E. Mason, J. L. Ross Anderson and Christopher E. Dempsey
Supplementary Table 1. Data collection and refinement statistics (molecular replacement).
 Supplementary Table 1. Data collection and refinement statistics (molecular replacement). Data set statistics HLA A*0201- ALWGPDPAAA PPI TCR PPI TCR/A2- ALWGPDPAAA PPI TCR/A2- ALWGPDPAAA Space Group P2
Supplementary Table 1. Data collection and refinement statistics (molecular replacement). Data set statistics HLA A*0201- ALWGPDPAAA PPI TCR PPI TCR/A2- ALWGPDPAAA PPI TCR/A2- ALWGPDPAAA Space Group P2
Review II: The Molecules of Life
 Review II: The Molecules of Life Judy Wieber BBSI @ Pitt 2007 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2007 Outline Introduction Proteins Carbohydrates Lipids
Review II: The Molecules of Life Judy Wieber BBSI @ Pitt 2007 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2007 Outline Introduction Proteins Carbohydrates Lipids
Supporting Information
 Supporting Information Mechanism of inactivation of -aminobutyric acid aminotransferase by (1S,3S)-3-amino-4-difluoromethylenyl-1- cyclopentanoic acid (CPP-115) Hyunbeom Lee, 1, Emma H. Doud, 1,2 Rui Wu,
Supporting Information Mechanism of inactivation of -aminobutyric acid aminotransferase by (1S,3S)-3-amino-4-difluoromethylenyl-1- cyclopentanoic acid (CPP-115) Hyunbeom Lee, 1, Emma H. Doud, 1,2 Rui Wu,
in-silico Design of Nanoparticles for Transdermal Drug Delivery Application
 Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2018 in-silico Design of Nanoparticles for Transdermal Drug Delivery Application Rakesh Gupta and Beena
Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2018 in-silico Design of Nanoparticles for Transdermal Drug Delivery Application Rakesh Gupta and Beena
Proteins and their structure
 Proteins and their structure Proteins are the most abundant biological macromolecules, occurring in all cells and all parts of cells. Proteins also occur in great variety; thousands of different kinds,
Proteins and their structure Proteins are the most abundant biological macromolecules, occurring in all cells and all parts of cells. Proteins also occur in great variety; thousands of different kinds,
SAM Teacher s Guide Four Levels of Protein Structure
 SAM Teacher s Guide Four Levels of Protein Structure Overview Students explore how protein folding creates distinct, functional proteins by examining each of the four different levels of protein structure.
SAM Teacher s Guide Four Levels of Protein Structure Overview Students explore how protein folding creates distinct, functional proteins by examining each of the four different levels of protein structure.
Table S1. X-ray data collection and refinement statistics
 Table S1. X-ray data collection and refinement statistics Data collection H7.167 Fab-Sh2/H7 complex Beamline SSRL 12-2 Wavelength (Å) 0.97950 Space group I2 1 3 Unit cell parameters (Å, º) a = b = c=207.3,
Table S1. X-ray data collection and refinement statistics Data collection H7.167 Fab-Sh2/H7 complex Beamline SSRL 12-2 Wavelength (Å) 0.97950 Space group I2 1 3 Unit cell parameters (Å, º) a = b = c=207.3,
Guided Inquiry Skills Lab. Additional Lab 1 Making Models of Macromolecules. Problem. Introduction. Skills Focus. Materials.
 Additional Lab 1 Making Models of Macromolecules Guided Inquiry Skills Lab Problem How do monomers join together to form polymers? Introduction A small number of elements make up most of the mass of your
Additional Lab 1 Making Models of Macromolecules Guided Inquiry Skills Lab Problem How do monomers join together to form polymers? Introduction A small number of elements make up most of the mass of your
Green Segment Contents
 Green Segment Contents Parts Reference Guide Green Segment 1 8 2 6 3 4 5 7 1. Amino Acid Side Chain Chart shows the properties and atomic structure of side chains. 2. Amino Acid Side Chains affect protein
Green Segment Contents Parts Reference Guide Green Segment 1 8 2 6 3 4 5 7 1. Amino Acid Side Chain Chart shows the properties and atomic structure of side chains. 2. Amino Acid Side Chains affect protein
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. (a) Uncropped version of Fig. 2a. RM indicates that the translation was done in the absence of rough mcirosomes. (b) LepB construct containing the GGPG-L6RL6-
Supplementary Figures Supplementary Figure 1. (a) Uncropped version of Fig. 2a. RM indicates that the translation was done in the absence of rough mcirosomes. (b) LepB construct containing the GGPG-L6RL6-
The Basics: A general review of molecular biology:
 The Basics: A general review of molecular biology: DNA Transcription RNA Translation Proteins DNA (deoxy-ribonucleic acid) is the genetic material It is an informational super polymer -think of it as the
The Basics: A general review of molecular biology: DNA Transcription RNA Translation Proteins DNA (deoxy-ribonucleic acid) is the genetic material It is an informational super polymer -think of it as the
Gold nanocrystals at DPPC bilayer. Bo Song, Huajun Yuan, Cynthia J. Jameson, Sohail Murad
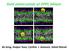 Gold nanocrystals at DPPC bilayer Bo Song, Huajun Yuan, Cynthia J. Jameson, Sohail Murad Coarse-grained mapping strategy of a DPPC lipid molecule. The structure of gold nanocrystals (bare gold nanoparticles)
Gold nanocrystals at DPPC bilayer Bo Song, Huajun Yuan, Cynthia J. Jameson, Sohail Murad Coarse-grained mapping strategy of a DPPC lipid molecule. The structure of gold nanocrystals (bare gold nanoparticles)
Proteins consist of joined amino acids They are joined by a Also called an Amide Bond
 Lecture Two: Peptide Bond & Protein Structure [Chapter 2 Berg, Tymoczko & Stryer] (Figures in Red are for the 7th Edition) (Figures in Blue are for the 8th Edition) Proteins consist of joined amino acids
Lecture Two: Peptide Bond & Protein Structure [Chapter 2 Berg, Tymoczko & Stryer] (Figures in Red are for the 7th Edition) (Figures in Blue are for the 8th Edition) Proteins consist of joined amino acids
Macromolecules Cut & Paste
 Macromolecules Cut & Paste Adapted from http://mrswords.weebly.com/uploads/1/5/2/4/15244382/ch_6-3_life_molecules_cut-out_lab.pdf INTRODUCTION Many of the molecules in living cells are so large that they
Macromolecules Cut & Paste Adapted from http://mrswords.weebly.com/uploads/1/5/2/4/15244382/ch_6-3_life_molecules_cut-out_lab.pdf INTRODUCTION Many of the molecules in living cells are so large that they
Crystal Structure of the Subtilisin Carlsberg: OMTKY3 Complex
 John Clizer & Greg Ralph Crystal Structure of the Subtilisin Carlsberg: OMTKY3 Complex The turkey ovomucoid third domain (OMTKY3) is considered to be one of the most studied protein inhibitors. 1 Ovomucin
John Clizer & Greg Ralph Crystal Structure of the Subtilisin Carlsberg: OMTKY3 Complex The turkey ovomucoid third domain (OMTKY3) is considered to be one of the most studied protein inhibitors. 1 Ovomucin
The role of Ca² + ions in the complex assembling of protein Z and Z-dependent protease inhibitor: A structure and dynamics investigation
 www.bioinformation.net Hypothesis Volume 8(9) The role of Ca² + ions in the complex assembling of protein Z and Z-dependent protease inhibitor: A structure and dynamics investigation Zahra Karimi 1 *,
www.bioinformation.net Hypothesis Volume 8(9) The role of Ca² + ions in the complex assembling of protein Z and Z-dependent protease inhibitor: A structure and dynamics investigation Zahra Karimi 1 *,
Lipid Bilayers Are Excellent For Cell Membranes
 Lipid Bilayers Are Excellent For Cell Membranes ydrophobic interaction is the driving force Self-assembly in water Tendency to close on themselves Self-sealing (a hole is unfavorable) Extensive: up to
Lipid Bilayers Are Excellent For Cell Membranes ydrophobic interaction is the driving force Self-assembly in water Tendency to close on themselves Self-sealing (a hole is unfavorable) Extensive: up to
Interaction Between Amyloid-b (1 42) Peptide and Phospholipid Bilayers: A Molecular Dynamics Study
 Biophysical Journal Volume 96 February 2009 785 797 785 Interaction Between Amyloid-b (1 42) Peptide and Phospholipid Bilayers: A Molecular Dynamics Study Charles H. Davis and Max L. Berkowitz * Department
Biophysical Journal Volume 96 February 2009 785 797 785 Interaction Between Amyloid-b (1 42) Peptide and Phospholipid Bilayers: A Molecular Dynamics Study Charles H. Davis and Max L. Berkowitz * Department
Biochemistry - I. Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture 1 Amino Acids I
 Biochemistry - I Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture 1 Amino Acids I Hello, welcome to the course Biochemistry 1 conducted by me Dr. S Dasgupta,
Biochemistry - I Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture 1 Amino Acids I Hello, welcome to the course Biochemistry 1 conducted by me Dr. S Dasgupta,
Structural biology of viruses
 Structural biology of viruses Biophysical Chemistry 1, Fall 2010 Coat proteins DNA/RNA packaging Reading assignment: Chap. 15 Virus particles self-assemble from coat monomers Virus Structure and Function
Structural biology of viruses Biophysical Chemistry 1, Fall 2010 Coat proteins DNA/RNA packaging Reading assignment: Chap. 15 Virus particles self-assemble from coat monomers Virus Structure and Function
Nature Neuroscience: doi: /nn Supplementary Figure 1. Neuron class-specific arrangements of Khc::nod::lacZ label in dendrites.
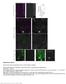 Supplementary Figure 1 Neuron class-specific arrangements of Khc::nod::lacZ label in dendrites. Staining with fluorescence antibodies to detect GFP (Green), β-galactosidase (magenta/white). (a, b) Class
Supplementary Figure 1 Neuron class-specific arrangements of Khc::nod::lacZ label in dendrites. Staining with fluorescence antibodies to detect GFP (Green), β-galactosidase (magenta/white). (a, b) Class
Chem Lecture 2 Protein Structure
 Chem 452 - Lecture 2 Protein Structure 110923 Proteins are the workhorses of a living cell and involve themselves in nearly all of the activities that take place in a cell. Their wide range of structures
Chem 452 - Lecture 2 Protein Structure 110923 Proteins are the workhorses of a living cell and involve themselves in nearly all of the activities that take place in a cell. Their wide range of structures
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10913 Supplementary Figure 1 2F o -F c electron density maps of cognate and near-cognate trna Leu 2 in the A site of the 70S ribosome. The maps are contoured at 1.2 sigma and some of
doi:10.1038/nature10913 Supplementary Figure 1 2F o -F c electron density maps of cognate and near-cognate trna Leu 2 in the A site of the 70S ribosome. The maps are contoured at 1.2 sigma and some of
Supplementary information: Binding of N-methylscopolamine to the extracellular domain of muscarinic acetylcholine receptors
 Supplementary information: Binding of N-methylscopolamine to the extracellular domain of muscarinic acetylcholine receptors Jan Jakubík, Alena Randáková, Pavel Zimčík, Esam E. El-Fakahany, and Vladimír
Supplementary information: Binding of N-methylscopolamine to the extracellular domain of muscarinic acetylcholine receptors Jan Jakubík, Alena Randáková, Pavel Zimčík, Esam E. El-Fakahany, and Vladimír
Biological Molecules
 The Chemical Building Blocks of Life Chapter 3 Biological molecules consist primarily of -carbon bonded to carbon, or -carbon bonded to other molecules. Carbon can form up to 4 covalent bonds. Carbon may
The Chemical Building Blocks of Life Chapter 3 Biological molecules consist primarily of -carbon bonded to carbon, or -carbon bonded to other molecules. Carbon can form up to 4 covalent bonds. Carbon may
Practice Problems 3. a. What is the name of the bond formed between two amino acids? Are these bonds free to rotate?
 Life Sciences 1a Practice Problems 3 1. Draw the oligopeptide for Ala-Phe-Gly-Thr-Asp. You do not need to indicate the stereochemistry of the sidechains. Denote with arrows the bonds formed between the
Life Sciences 1a Practice Problems 3 1. Draw the oligopeptide for Ala-Phe-Gly-Thr-Asp. You do not need to indicate the stereochemistry of the sidechains. Denote with arrows the bonds formed between the
The Chemical Building Blocks of Life. Chapter 3
 The Chemical Building Blocks of Life Chapter 3 Biological Molecules Biological molecules consist primarily of -carbon bonded to carbon, or -carbon bonded to other molecules. Carbon can form up to 4 covalent
The Chemical Building Blocks of Life Chapter 3 Biological Molecules Biological molecules consist primarily of -carbon bonded to carbon, or -carbon bonded to other molecules. Carbon can form up to 4 covalent
Judy Wieber. Department of Computational Biology. May 27, 2008
 Review II: The Molecules of Life Judy Wieber BBSI @ Pitt 2008 Department of Computational Biology University it of Pittsburgh School of Medicine i May 27, 2008 Outline Introduction Proteins Carbohydrates
Review II: The Molecules of Life Judy Wieber BBSI @ Pitt 2008 Department of Computational Biology University it of Pittsburgh School of Medicine i May 27, 2008 Outline Introduction Proteins Carbohydrates
WHY MACROMOLECULAR STRUCTURE?
 WHY MACROMOLECULAR STRUCTURE? Why Do Scientists Study Structure? Biological function operates through structure. The genetic code is expressed through structure. 20 th century physicists developed the
WHY MACROMOLECULAR STRUCTURE? Why Do Scientists Study Structure? Biological function operates through structure. The genetic code is expressed through structure. 20 th century physicists developed the
Supplementary materials
 Supplementary materials Chemical library from ChemBridge 50,240 structurally diverse small molecule compounds dissolved in DMSO Hits Controls: No virus added μ Primary screening at 20 g/ml of compounds
Supplementary materials Chemical library from ChemBridge 50,240 structurally diverse small molecule compounds dissolved in DMSO Hits Controls: No virus added μ Primary screening at 20 g/ml of compounds
Protein Statistics. Scott Clark University of California: Davis Oregon State University. August 24, 2006
 Protein Statistics Scott Clark University of California: Davis Oregon State University August 24, 2006 Abstract Proteins are vital for the functionality of any organic system. The folding of proteins,
Protein Statistics Scott Clark University of California: Davis Oregon State University August 24, 2006 Abstract Proteins are vital for the functionality of any organic system. The folding of proteins,
Hemoglobin & Sickle Cell Anemia Exercise
 Name StarBiochem Hemoglobin & Sickle Cell Anemia Exercise Background Hemoglobin is the protein in red blood cells responsible for carrying oxygen from the lungs to the rest of the body and for returning
Name StarBiochem Hemoglobin & Sickle Cell Anemia Exercise Background Hemoglobin is the protein in red blood cells responsible for carrying oxygen from the lungs to the rest of the body and for returning
Amino acids & Protein Structure Chemwiki: Chapter , with most emphasis on 16.3, 16.4 and 16.6
 Amino acids & Protein Structure Chemwiki: Chapter 16. 16.1, 16.3-16.9 with most emphasis on 16.3, 16.4 and 16.6 1 1. Most jobs (except information storage) in cells are performed by proteins. 2. Proteins
Amino acids & Protein Structure Chemwiki: Chapter 16. 16.1, 16.3-16.9 with most emphasis on 16.3, 16.4 and 16.6 1 1. Most jobs (except information storage) in cells are performed by proteins. 2. Proteins
Supplementary material for:
 Supplementary material for: CHOLESTEROL SENSITIVITY OF KIR2.1 IS CONTROLLED BY A BELT OF RESIDUES AROUND THE CYTOSOLIC PORE Avia Rosenhouse-Dantsker 1, Diomedes E. Logothetis 2 and Irena Levitan 1 1 Department
Supplementary material for: CHOLESTEROL SENSITIVITY OF KIR2.1 IS CONTROLLED BY A BELT OF RESIDUES AROUND THE CYTOSOLIC PORE Avia Rosenhouse-Dantsker 1, Diomedes E. Logothetis 2 and Irena Levitan 1 1 Department
Insulin mrna to Protein Kit
 Insulin mrna to Protein Kit A 3DMD Paper BioInformatics and Mini-Toober Folding Activity Student Handout www.3dmoleculardesigns.com Insulin mrna to Protein Kit Contents Becoming Familiar with the Data...
Insulin mrna to Protein Kit A 3DMD Paper BioInformatics and Mini-Toober Folding Activity Student Handout www.3dmoleculardesigns.com Insulin mrna to Protein Kit Contents Becoming Familiar with the Data...
Supplementary Information Janssen et al.
 Supplementary Information Janssen et al. Insights into complement convertase formation based on the structure of the factor B CVF complex Bert J.C. Janssen 1, Lucio Gomes 1, Roman I. Koning 2, Dmitri I.
Supplementary Information Janssen et al. Insights into complement convertase formation based on the structure of the factor B CVF complex Bert J.C. Janssen 1, Lucio Gomes 1, Roman I. Koning 2, Dmitri I.
Ch5: Macromolecules. Proteins
 Ch5: Macromolecules Proteins Essential Knowledge 4.A.1 The subcomponents of biological molecules and their sequence determine the properties of that molecule A. Structure and function of polymers are derived
Ch5: Macromolecules Proteins Essential Knowledge 4.A.1 The subcomponents of biological molecules and their sequence determine the properties of that molecule A. Structure and function of polymers are derived
Protein structure. Dr. Mamoun Ahram Summer semester,
 Protein structure Dr. Mamoun Ahram Summer semester, 2017-2018 Overview of proteins Proteins have different structures and some have repeating inner structures, other do not. A protein may have gazillion
Protein structure Dr. Mamoun Ahram Summer semester, 2017-2018 Overview of proteins Proteins have different structures and some have repeating inner structures, other do not. A protein may have gazillion
Figure 1.1 PHITS geometry for PTB irradiations with: broad beam, upper panel; mono energetic beams, lower panel. Pictures of the setups and of the
 Figure 1.1 PHITS geometry for PTB irradiations with: broad beam, upper panel; mono energetic beams, lower panel. Pictures of the setups and of the PMMA ring holder with 7 containers are also shown. Relative
Figure 1.1 PHITS geometry for PTB irradiations with: broad beam, upper panel; mono energetic beams, lower panel. Pictures of the setups and of the PMMA ring holder with 7 containers are also shown. Relative
Life Sciences 1a. Practice Problems 4
 Life Sciences 1a Practice Problems 4 1. KcsA, a channel that allows K + ions to pass through the membrane, is a protein with four identical subunits that form a channel through the center of the tetramer.
Life Sciences 1a Practice Problems 4 1. KcsA, a channel that allows K + ions to pass through the membrane, is a protein with four identical subunits that form a channel through the center of the tetramer.
Nature Getetics: doi: /ng.3471
 Supplementary Figure 1 Summary of exome sequencing data. ( a ) Exome tumor normal sample sizes for bladder cancer (BLCA), breast cancer (BRCA), carcinoid (CARC), chronic lymphocytic leukemia (CLLX), colorectal
Supplementary Figure 1 Summary of exome sequencing data. ( a ) Exome tumor normal sample sizes for bladder cancer (BLCA), breast cancer (BRCA), carcinoid (CARC), chronic lymphocytic leukemia (CLLX), colorectal
!"#$%&' (#%) /&'(2+"( /&3&4,, ! " #$% - &'()!% *-sheet -(!-Helix - &'(&') +,(-. - &'()&+) /&%.(0&+(! - &'(1&2%( Basic amino acids
 Basic amino acids pk ~ 10.5 pk ~ 12.5 pk ~ 6.0 Polar 25!"#$%&' (#%)! " #$% - &'()!% *-sheet -(!-Helix - &'(&') +,(-. - &'()&+) /&%.(0&+(! - &'(1&2%( /&'(2+"( /&3&4,, :++55 ('&.! 6($.(" 40 > 3&4,, ('&.!
Basic amino acids pk ~ 10.5 pk ~ 12.5 pk ~ 6.0 Polar 25!"#$%&' (#%)! " #$% - &'()!% *-sheet -(!-Helix - &'(&') +,(-. - &'()&+) /&%.(0&+(! - &'(1&2%( /&'(2+"( /&3&4,, :++55 ('&.! 6($.(" 40 > 3&4,, ('&.!
S system H system /T 0 ph = log([h + ]) E = mc 2 S = klnw G = H T S ph = pk a + log([a ]/[HA]) K a = [H + ][A ]/[HA] G = RTlnK eq e iπ + 1 = 0
![S system H system /T 0 ph = log([h + ]) E = mc 2 S = klnw G = H T S ph = pk a + log([a ]/[HA]) K a = [H + ][A ]/[HA] G = RTlnK eq e iπ + 1 = 0 S system H system /T 0 ph = log([h + ]) E = mc 2 S = klnw G = H T S ph = pk a + log([a ]/[HA]) K a = [H + ][A ]/[HA] G = RTlnK eq e iπ + 1 = 0](/thumbs/94/121313803.jpg) Biochemistry 463, Summer II Your Name: University of Maryland, College Park Your SID #: Biochemistry and Physiology Prof. Jason Kahn Exam I (100 points total) July 27, 2009 You have 80 minutes for this
Biochemistry 463, Summer II Your Name: University of Maryland, College Park Your SID #: Biochemistry and Physiology Prof. Jason Kahn Exam I (100 points total) July 27, 2009 You have 80 minutes for this
University of Illinois at Urbana-Champaign NIH Resource for Macromolecular Modeling and Bioinformatics Beckman Institute.
 University of Illinois at Urbana-Champaign NIH Resource for Macromolecular Modeling and Bioinformatics Beckman Institute Aquaporins VMD Developers: John Stone Dan Wright John Eargle Fatemeh Khalili Elizabeth
University of Illinois at Urbana-Champaign NIH Resource for Macromolecular Modeling and Bioinformatics Beckman Institute Aquaporins VMD Developers: John Stone Dan Wright John Eargle Fatemeh Khalili Elizabeth
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature22394 Supplementary Table 1 Observed intermolecular interactions within the GLP-1:GLP-1R TMD interface. Superscripts refer to the Wootten residue numbering system
SUPPLEMENTARY INFORMATION doi:10.1038/nature22394 Supplementary Table 1 Observed intermolecular interactions within the GLP-1:GLP-1R TMD interface. Superscripts refer to the Wootten residue numbering system
Lecture 4: 8/26. CHAPTER 4 Protein Three Dimensional Structure
 Lecture 4: 8/26 CHAPTER 4 Protein Three Dimensional Structure Summary of the Lecture 3 There are 20 amino acids and only the L isomer amino acid exist in proteins Each amino acid consists of a central
Lecture 4: 8/26 CHAPTER 4 Protein Three Dimensional Structure Summary of the Lecture 3 There are 20 amino acids and only the L isomer amino acid exist in proteins Each amino acid consists of a central
ATP-independent reversal of a membrane protein aggregate by a chloroplast SRP
 ATP-independent reversal of a membrane protein aggregate by a chloroplast SRP Peera Jaru-Ampornpan 1, Kuang Shen 1,3, Vinh Q. Lam 1,3, Mona Ali 2, Sebastian Doniach 2, Tony Z. Jia 1, Shu-ou Shan 1 Supplementary
ATP-independent reversal of a membrane protein aggregate by a chloroplast SRP Peera Jaru-Ampornpan 1, Kuang Shen 1,3, Vinh Q. Lam 1,3, Mona Ali 2, Sebastian Doniach 2, Tony Z. Jia 1, Shu-ou Shan 1 Supplementary
WHAT IS A PROTEIN? OBJECTIVES The objective of this worksheet is to understand the structure and function of proteins. PART A: Understanding Proteins
 WHAT IS A PROTEIN? OBJECTIVES The objective of this worksheet is to understand the structure and function of proteins PART A: Understanding Proteins As you may already know proteins are an essential part
WHAT IS A PROTEIN? OBJECTIVES The objective of this worksheet is to understand the structure and function of proteins PART A: Understanding Proteins As you may already know proteins are an essential part
