IDENTIFICATION AND PARTIAL CHARACTERIZATION OF A SEX SPECIFIC PROTEIN IN KOI CARP (Cyprinus carpio haematopterus)
|
|
|
- Madison Quinn
- 5 years ago
- Views:
Transcription
1 Short communication Acta Veterinaria-Beograd 2017, 67 (2), UDK: : DOI: /acve IDENTIFICATION AND PARTIAL CHARACTERIZATION OF A SEX SPECIFIC PROTEIN IN KOI CARP (Cyprinus carpio haematopterus) POPOVSKI Zoran 1*, KWASEK Karolina 2, WOJNO Michal 2, DABROWSKI Konrad 2, WICK Macdonald 3 1 Faculty of Agriculture and Food Sciences, Ss Cyril and Methodius University, Skopje, R. of Macedonia; 2 School of Environmental and Natural Resources, The Ohio State University, Columbus, OH; 3 Department of Animal Sciences, The Ohio State University, Columbus, OH (Received 11 August 2016, Accepted 22 February 2017) Gender identification of fish species is carried out mainly by examining external morphological characteristics, which in general, it is very complex and not always a reliable approach. Electrophoresis of plasma proteins can be used as an alternative and useful molecular tool for a more precise sex determination. The presence of female specific proteins in the plasma is a starting point for the application of this technique. In this study, reducing discontinuous sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) was applied to analyze plasma proteins of male and female koi carp (Cyprinus carpio haematopterus). Image analyses of electrophoregrams with resolved plasma proteins by SDS-PAGE showed that it is an appropriate technique to discriminate male from female samples. It is based on the presence of apolipoprotein B-100 which can be used as a suitable marker. Further amino acid characterization of apolipoprotein B-100 confirmed that it is a specific protein for female individuals. Key words: koi carp, SDS-PAGE, apolipoprotein, structure INTRODUCTION In the past, the identification of fish gender was carried out mainly by examining the external morphological characteristics. This can have a negative impact on the fish being analyzed. A non-destructive method for gender identification of fish could prevent negative effects. Numerous studies have demonstrated the impact of proteomics on answering key biological questions; especially those that help us understand the vital functions of a living system [1]. Plasma proteins are a suitable proxy of the physiological status. Currently, electrophoresis of sarcoplasmic proteins, plasma proteins, liver proteins and a number of enzymes often have been used by some researchers as an aid in species or gender identification of fish [2]. Limited *Corresponding author: zoran.popovski@karpos.gov.mk 285
2 Acta Veterinaria-Beograd 2017, 67 (2), studies exist on aspects of physiological changes associated with the reproductive cycle and gametogenesis. Using electrophoresis a different profile of plasma proteins appeared at different stages of maturation and attempts to trace the presence of vitellogenin were made [3]. The presence of apolipoproteins as part of plasma high density lipoproteins in female trout was reported without its characterization [4]. Proteomic technology for different applications is robust, quantitatively accurate, and high throughput analytical platform. In the recent years, mass spectrometry-based proteomics has developed into a powerful technology that is able to routinely study complex proteome samples [5]. Therefore, the objective of this study was to identify and characterize a female specific plasma protein in koi carp by a combination of electrophoretic and proteomic analyses. MATERIALS AND METHODS The animal handling procedures were approved on January 7, 2009 by the Institutional Animal Care and Use Committee of Ohio State University, protocol 2008A0221 that followed the federal norms and regulations. Fourteen koi carp (Cyprinus carpio haematopterus), with a mass between 190 and 1,200 g, were obtained from the aquaculture laboratory in the School of Environmental and Natural Resources at Ohio State University, Columbus, OH (Tab. 1). Table 1. Koi carp, sample IDs, sex and weight (grams) Sample ID Sex wgt (grams) 1 Male Female Female Female Male Male Male Female Male 1, Female Female Male Male Female
3 Popovski et al.: Sex specific protein in Koi carp Initial gender determination was done using a not fully reliable morphological characterization based on the common features for gender dimorphism such as pectoral fin and body shape and appearance of special white tubercles on the male head during the breeding season [6]. The blood was taken from the dorsal aorta of live individuals. Blood samples were centrifuged at 1,500 x g for 10 minutes to separate the plasma. The obtained plasma was used for protein electrophoresis. Five hundred ml of plasma were thoroughly mixed in 1 ml of sample buffer (8 M urea / 2 M thiourea, 2 mm DTT, 50 mm Tris, 3% SDS, 0.004% bromophenol blue, ph 6.8) and incubated on ice for 30 min. Samples were centrifuged at 10,000 g for 10 min at room temperature before loading into a 1 mm 12 cm 14 cm discontinuous polyacrylamide slab gel consisting of a 12.5% resolving gel [30:0.8, acrylamide/n,n0- bis(methylene acrylamide)] and a 3% stacking gel. Electrophoretic separation was carried out at a constant voltage of 10 Vcm-1. Following electrophoretic separation, the gels were imaged and analyzed as previously described [7] on a flatbed scanner. Digital images were analyzed using the Total Lab TL120 (Nonlinear Dynamics Inc., Newcastle upon Tyne, U.K.) software. The selected band was digested with sequencing grade trypsin from Promega (Madison WI) or sequencing grade chymotrypsin from Roche T (Indianapolis, IN) using the Multiscreen Solvinert Filter Plates from Millipore (Bedford, MA). Capillary-liquid chromatography tandem mass spectrometry (Cap-LC/MS/MS) was performed on a Thermo Scientific LTQ mass spectrometer equipped with a CaptiveSpray source (Bruker Daltonics Billerica, MA) operated in positive ion mode. The LC system was an UltiMate 3000 system from Thermo Scientific. The scan sequence of the mass spectrometer was based on the TopTen method. The analysis was programmed for a full scan recorded between Da, and a MS/MS scan to generate product ion spectra to determine amino acid sequence in consecutive instrument scans of the ten most abundant peaks in the spectrum. Sequence information from the MS/MS data was processed by converting the.raw files into a merged file (.mgf) using an in-house program, RAW2MZXML_n_MGF_ batch (merge.pl, a Perl script). The resulting mgf files were searched using Mascot Daemon by Matrix Science version (Boston, MA) and the database searched against the most recent SwissProt or NCBI databases. The mass accuracy of the precursor ions were set to 1.8 Da and the fragment mass accuracy was set to 0.8 Da. A decoy database was searched to determine the false discovery rate (FDR) and peptides were filtered according to the FDR and proteins identified required bold red peptides. Protein identifications were checked manually and proteins with a Mascot score of 50 or higher with a minimum of two unique peptides from one protein having a -b or -y ion sequence tag of five residues or better were accepted. 287
4 Acta Veterinaria-Beograd 2017, 67 (2), RESULTS AND DISCUSSION The appearance of a female specific protein in the electrophoregram of plasma proteins profile of female koi carp Cyprinus carpio haematopterus obtained by 12.5% SDS-PAGE is presented in Figure 1. Figure % SDS-PAGE on plasma proteins from of koi carp. Lane designations presented in Table 1. Molecular weight standards in lane (L) where the sizes of bands (in kda) are given close to far right column The bend in the white box was identified as female-specific for Cyprinus carpio haematopteru) The band in the white box was identified as being specific to the female fish and was prepared for primary protein sequence analysis (Suppl. 1). The protein was identified as Apoliprotein B from Larimichthys a genus of ray-finned saltwater fishes commonly known as yellow croakers. Although Apolipoprotein B, in many species, has been reported to normally migrate at approximately 407 kda, the protein identified as Apolipoprotein B, in this study, migrated at kda. In addition, the fragment coverage identified by MASCOT analysis spans a migration corresponding to a molecular weight of a minimum of 212 kda (Table 2). There is no report of the primary sequence of any Apolipoproteins in koi carp. Therefore, this discrepancy in the migration and molecular weight is possibly due to either an inherent difference in the primary protein sequence in Apolipoprotein in koi carp and/or the peptides generated and used to determine the primary sequence in the submitted band were homologous to sequences within an Apolipoprotein-like protein. Regardless, the plasma protein identified as Apolipoprotein in this study, is unique to the female koi carp studied. The protein is absent in the plasma of males. 288
5 Popovski et al.: Sex specific protein in Koi carp Supplementary file 1. Results of primary sequence analysis showing MASCOT search results identifying Apolipoprotein B-100 in the submitted ~200 kda band. 289
6 Acta Veterinaria-Beograd 2017, 67 (2), These kind of studies brought about a new look to taxonomical evaluation. Discrimination of related taxa can be easily made according to their electrophoretic results of plasma proteins [8]. SDS-PAGE can be used as a technique for gender determination of koi carp based on the identification of the presence of a female specific protein in plasma not necessarily followed by proteomics confirmation. Authors contributions PZ designed the experiment, performed an electrophoretic analysis and drafted the paper. KK was involved in fish growing and lab work. WM made preliminary morphological gender identification and provided necessary fish data. DK provided the blood samples and approved the article. WM was involved in all phases of electrophoretic and proteomic analysis and did a proofreading of the text. Declaration of conflicting interests The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article. REFERENCES 1. Surinova S, Schiess R, Huttenhain R, Cerciello F, Wollscheid B, Aebersold R: On the Development of Plasma Protein Biomarkers. J Proteome Res 2011, 10: Yilmaz M, Yilmaz R, Alas A: An electrophoretic taxonomic study on serum proteins of Acanthobrama marmid, Leuciscus cephalus, and Chondrostoma regium. EurAsia J BioSci 2007, 3: Kumar LK, Gopalakrishnan A, Mohindra V, James PB: Identification of female specific protein in seabass, Lates calcarifer (Bloch). Ind J Fish 1999, 46: Babin PJ: Apolipoproteins and the association of egg yolk proteins with plasma high density lipoproteins after ovulation and follicular atresia in the rainbow trout (Salmo gairdneri). J Biol Chem 1987, 262: Zhang H, Alvin Y L, Loriaux P, Wollscheid B, Zhou Y, Watts J, Aebersold R: Mass spectrometric detection of tissue proteins in plasma. Mol. Cell. Proteomics 2007, 6-1: Holmes K, Pitham T, Fletcher N: The world of Koi. Barron s Educational Series, Inc., Hauppauge, NY, Zapata I, Zerby H, Wick M:. Functional proteomic analysis predicts beef tenderness and the tenderness differential. J Agri Food Chem 2009, 57: Theophilus J, Rao PR: Electrophoretic studies on the plasma proteins of the three species of genus channa. Ind. J. Fish 1998, 35:
7 Popovski et al.: Sex specific protein in Koi carp IDENTIFIKACIJA I PARCIJALNA KARAKTERIZACIJA POLNO SPECIFIČNOG PROTEINA AZIJSKOG ŠARANA (Cyprinus carpio haematopterus) POPOVSKI Zoran, KWASEK Karolina, WOJNO Michal, DABROWSKI Konrad, WICK Macdonald Identifikacija pola riba se izvodi prvenstveno na osnovu ispitivanja spoljašnjih morfoloških karakteristika, što predstavlja složen i ne uvek dostupan način. U tom smislu elektroforeza plazma proteina može da se koristi kao precizna i pouzdana alternativna molekularna metoda. Prisustvo proteina plazme koji su specifični za ženski pol predstavlja polaznu osnovu za primenu ove tehnike. U ovoj studiji primenjen je redukujući diskontinuirani natrijum dodecil sulfat-poliakrilamidni gel (SDS-PAGE) u cilju ispitivanja plazma proteina mužijaka i ženke Azijskog šarana (Cyprinus carpio haematopterus). Analiza slike elektroforezograma sa pomoću SDS-PAGE elektroforezom razdvojenim plazma proteinima je pokazala da je metoda odgovarajuća za diskriminaciju uzoraka prema polu. Zasnovana je na prisustvu apolipoproteina B-100 koji u ovom slučaju predstavlja pouzdan marker. Dodatna karakterizacija aminokiselina apolipoproteina B-100 je potvrdila da on predstavlja protein koji je specifičan za ženke. 291
Improve Protein Analysis with the New, Mass Spectrometry- Compatible ProteasMAX Surfactant
 Improve Protein Analysis with the New, Mass Spectrometry- Compatible Surfactant ABSTRACT Incomplete solubilization and digestion and poor peptide recovery are frequent limitations in protein sample preparation
Improve Protein Analysis with the New, Mass Spectrometry- Compatible Surfactant ABSTRACT Incomplete solubilization and digestion and poor peptide recovery are frequent limitations in protein sample preparation
Babu Antharavally, Ryan Bomgarden, and John Rogers Thermo Fisher Scientific, Rockford, IL
 A Versatile High-Recovery Method for Removing Detergents from Low-Concentration Protein or Peptide Samples for Mass Spectrometry Sample Preparation and Analysis Babu Antharavally, Ryan Bomgarden, and John
A Versatile High-Recovery Method for Removing Detergents from Low-Concentration Protein or Peptide Samples for Mass Spectrometry Sample Preparation and Analysis Babu Antharavally, Ryan Bomgarden, and John
Learning Objectives. Overview of topics to be discussed 10/25/2013 HIGH RESOLUTION MASS SPECTROMETRY (HRMS) IN DISCOVERY PROTEOMICS
 HIGH RESOLUTION MASS SPECTROMETRY (HRMS) IN DISCOVERY PROTEOMICS A clinical proteomics perspective Michael L. Merchant, PhD School of Medicine, University of Louisville Louisville, KY Learning Objectives
HIGH RESOLUTION MASS SPECTROMETRY (HRMS) IN DISCOVERY PROTEOMICS A clinical proteomics perspective Michael L. Merchant, PhD School of Medicine, University of Louisville Louisville, KY Learning Objectives
Protein MultiColor Stable, Low Range
 Product Name: DynaMarker Protein MultiColor Stable, Low Range Code No: DM670L Lot No: ******* Size: 200 μl x 3 (DM670 x 3) (120 mini-gel lanes) Storage: 4 C Stability: 12 months at 4 C Storage Buffer:
Product Name: DynaMarker Protein MultiColor Stable, Low Range Code No: DM670L Lot No: ******* Size: 200 μl x 3 (DM670 x 3) (120 mini-gel lanes) Storage: 4 C Stability: 12 months at 4 C Storage Buffer:
Double charge of 33kD peak A1 A2 B1 B2 M2+ M/z. ABRF Proteomics Research Group - Qualitative Proteomics Study Identifier Number 14146
 Abstract The 2008 ABRF Proteomics Research Group Study offers participants the chance to participate in an anonymous study to identify qualitative differences between two protein preparations. We used
Abstract The 2008 ABRF Proteomics Research Group Study offers participants the chance to participate in an anonymous study to identify qualitative differences between two protein preparations. We used
PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System
 Application Note LC/MS PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System Purpose This application note describes an automated workflow
Application Note LC/MS PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System Purpose This application note describes an automated workflow
Unsupervised Identification of Isotope-Labeled Peptides
 Unsupervised Identification of Isotope-Labeled Peptides Joshua E Goldford 13 and Igor GL Libourel 124 1 Biotechnology institute, University of Minnesota, Saint Paul, MN 55108 2 Department of Plant Biology,
Unsupervised Identification of Isotope-Labeled Peptides Joshua E Goldford 13 and Igor GL Libourel 124 1 Biotechnology institute, University of Minnesota, Saint Paul, MN 55108 2 Department of Plant Biology,
Protein Reports CPTAC Common Data Analysis Pipeline (CDAP)
 Protein Reports CPTAC Common Data Analysis Pipeline (CDAP) v. 05/03/2016 Summary The purpose of this document is to describe the protein reports generated as part of the CPTAC Common Data Analysis Pipeline
Protein Reports CPTAC Common Data Analysis Pipeline (CDAP) v. 05/03/2016 Summary The purpose of this document is to describe the protein reports generated as part of the CPTAC Common Data Analysis Pipeline
Characterization of Disulfide Linkages in Proteins by 193 nm Ultraviolet Photodissociation (UVPD) Mass Spectrometry. Supporting Information
 Characterization of Disulfide Linkages in Proteins by 193 nm Ultraviolet Photodissociation (UVPD) Mass Spectrometry M. Montana Quick, Christopher M. Crittenden, Jake A. Rosenberg, and Jennifer S. Brodbelt
Characterization of Disulfide Linkages in Proteins by 193 nm Ultraviolet Photodissociation (UVPD) Mass Spectrometry M. Montana Quick, Christopher M. Crittenden, Jake A. Rosenberg, and Jennifer S. Brodbelt
Nature Biotechnology: doi: /nbt Supplementary Figure 1
 Supplementary Figure 1 The timeline of the NGAG method for extraction of N-linked glycans and glycosite-containing peptides. The timeline can be changed based on the number of samples. Supplementary Figure
Supplementary Figure 1 The timeline of the NGAG method for extraction of N-linked glycans and glycosite-containing peptides. The timeline can be changed based on the number of samples. Supplementary Figure
Proteomics of body liquids as a source for potential methods for medical diagnostics Prof. Dr. Evgeny Nikolaev
 Proteomics of body liquids as a source for potential methods for medical diagnostics Prof. Dr. Evgeny Nikolaev Institute for Biochemical Physics, Rus. Acad. Sci., Moscow, Russia. Institute for Energy Problems
Proteomics of body liquids as a source for potential methods for medical diagnostics Prof. Dr. Evgeny Nikolaev Institute for Biochemical Physics, Rus. Acad. Sci., Moscow, Russia. Institute for Energy Problems
Sequence Identification And Spatial Distribution of Rat Brain Tryptic Peptides Using MALDI Mass Spectrometric Imaging
 Sequence Identification And Spatial Distribution of Rat Brain Tryptic Peptides Using MALDI Mass Spectrometric Imaging AB SCIEX MALDI TOF/TOF* Systems Patrick Pribil AB SCIEX, Canada MALDI mass spectrometric
Sequence Identification And Spatial Distribution of Rat Brain Tryptic Peptides Using MALDI Mass Spectrometric Imaging AB SCIEX MALDI TOF/TOF* Systems Patrick Pribil AB SCIEX, Canada MALDI mass spectrometric
Automating Mass Spectrometry-Based Quantitative Glycomics using Tandem Mass Tag (TMT) Reagents with SimGlycan
 PREMIER Biosoft Automating Mass Spectrometry-Based Quantitative Glycomics using Tandem Mass Tag (TMT) Reagents with SimGlycan Ne uaca2-3galb1-4glc NAcb1 6 Gal NAca -Thr 3 Ne uaca2-3galb1 Ningombam Sanjib
PREMIER Biosoft Automating Mass Spectrometry-Based Quantitative Glycomics using Tandem Mass Tag (TMT) Reagents with SimGlycan Ne uaca2-3galb1-4glc NAcb1 6 Gal NAca -Thr 3 Ne uaca2-3galb1 Ningombam Sanjib
Universal sample preparation method for proteome analysis
 nature methods Universal sample preparation method for proteome analysis Jacek R Wi niewski, Alexandre Zougman, Nagarjuna Nagaraj & Matthias Mann Supplementary figures and text: Supplementary Figure 1
nature methods Universal sample preparation method for proteome analysis Jacek R Wi niewski, Alexandre Zougman, Nagarjuna Nagaraj & Matthias Mann Supplementary figures and text: Supplementary Figure 1
Enhancing Sequence Coverage in Proteomics Studies by Using a Combination of Proteolytic Enzymes
 Enhancing Sequence Coverage in Proteomics Studies by Using a Combination of Proteolytic Enzymes Dominic Baeumlisberger 2, Christopher Kurz 3, Tabiwang N. Arrey, Marion Rohmer 2, Carola Schiller 3, Thomas
Enhancing Sequence Coverage in Proteomics Studies by Using a Combination of Proteolytic Enzymes Dominic Baeumlisberger 2, Christopher Kurz 3, Tabiwang N. Arrey, Marion Rohmer 2, Carola Schiller 3, Thomas
Quantitative chromatin proteomics reveals a dynamic histone. post-translational modification landscape that defines asexual
 Quantitative chromatin proteomics reveals a dynamic histone post-translational modification landscape that defines asexual and sexual Plasmodium falciparum parasites Nanika Coetzee 1, Simone Sidoli 2,
Quantitative chromatin proteomics reveals a dynamic histone post-translational modification landscape that defines asexual and sexual Plasmodium falciparum parasites Nanika Coetzee 1, Simone Sidoli 2,
Technical Note # TN-31 Redefining MALDI-TOF/TOF Performance
 Bruker Daltonics Technical Note # TN-31 Redefining MALDI-TOF/TOF Performance The new ultraflextreme exceeds all current expectations of MALDI-TOF/TOF technology: A proprietary khz smartbeam-ii TM MALDI
Bruker Daltonics Technical Note # TN-31 Redefining MALDI-TOF/TOF Performance The new ultraflextreme exceeds all current expectations of MALDI-TOF/TOF technology: A proprietary khz smartbeam-ii TM MALDI
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/2/4/e1500980/dc1 Supplementary Materials for The crystal structure of human dopamine -hydroxylase at 2.9 Å resolution Trine V. Vendelboe, Pernille Harris, Yuguang
advances.sciencemag.org/cgi/content/full/2/4/e1500980/dc1 Supplementary Materials for The crystal structure of human dopamine -hydroxylase at 2.9 Å resolution Trine V. Vendelboe, Pernille Harris, Yuguang
Nature Methods: doi: /nmeth.3177
 Supplementary Figure 1 Characterization of LysargiNase, trypsin and LysN missed cleavages. (a) Proportion of peptides identified in LysargiNase and trypsin digests of MDA-MB-231 cell lysates carrying 0,
Supplementary Figure 1 Characterization of LysargiNase, trypsin and LysN missed cleavages. (a) Proportion of peptides identified in LysargiNase and trypsin digests of MDA-MB-231 cell lysates carrying 0,
Lecture 3. Tandem MS & Protein Sequencing
 Lecture 3 Tandem MS & Protein Sequencing Nancy Allbritton, M.D., Ph.D. Department of Physiology & Biophysics 824-9137 (office) nlallbri@uci.edu Office- Rm D349 Medical Science D Bldg. Tandem MS Steps:
Lecture 3 Tandem MS & Protein Sequencing Nancy Allbritton, M.D., Ph.D. Department of Physiology & Biophysics 824-9137 (office) nlallbri@uci.edu Office- Rm D349 Medical Science D Bldg. Tandem MS Steps:
AbsoluteIDQ p150 Kit. Targeted Metabolite Identifi cation and Quantifi cation. Bringing our targeted metabolomics expertise to your lab.
 AbsoluteIDQ p150 Kit Targeted Metabolite Identifi cation and Quantifi cation Bringing our targeted metabolomics expertise to your lab. The Biocrates AbsoluteIDQ p150 mass spectrometry Assay Preparation
AbsoluteIDQ p150 Kit Targeted Metabolite Identifi cation and Quantifi cation Bringing our targeted metabolomics expertise to your lab. The Biocrates AbsoluteIDQ p150 mass spectrometry Assay Preparation
NIH Public Access Author Manuscript J Proteome Res. Author manuscript; available in PMC 2014 July 05.
 NIH Public Access Author Manuscript Published in final edited form as: J Proteome Res. 2013 July 5; 12(7): 3071 3086. doi:10.1021/pr3011588. Evaluation and Optimization of Mass Spectrometric Settings during
NIH Public Access Author Manuscript Published in final edited form as: J Proteome Res. 2013 July 5; 12(7): 3071 3086. doi:10.1021/pr3011588. Evaluation and Optimization of Mass Spectrometric Settings during
The Detection of Allergens in Food Products with LC-MS
 The Detection of Allergens in Food Products with LC-MS Something for the future? Jacqueline van der Wielen Scope of Organisation Dutch Food and Consumer Product Safety Authority: Law enforcement Control
The Detection of Allergens in Food Products with LC-MS Something for the future? Jacqueline van der Wielen Scope of Organisation Dutch Food and Consumer Product Safety Authority: Law enforcement Control
Bruker Daltonics. autoflex III smartbeam. The Standard in MALDI-TOF Performance MALDI-TOF/TOF. think forward
 Bruker Daltonics autoflex III smartbeam The Standard in MALDI-TOF Performance think forward MALDI-TOF/TOF Designed for a Routine High Level of Performance The autoflex III smartbeam brings the power of
Bruker Daltonics autoflex III smartbeam The Standard in MALDI-TOF Performance think forward MALDI-TOF/TOF Designed for a Routine High Level of Performance The autoflex III smartbeam brings the power of
The distribution of log 2 ratio (H/L) for quantified peptides. cleavage sites in each bin of log 2 ratio of quantified. peptides
 Journal: Nature Methods Article Title: Corresponding Author: Protein digestion priority is independent of their abundances Mingliang Ye and Hanfa Zou Supplementary Figure 1 Supplementary Figure 2 The distribution
Journal: Nature Methods Article Title: Corresponding Author: Protein digestion priority is independent of their abundances Mingliang Ye and Hanfa Zou Supplementary Figure 1 Supplementary Figure 2 The distribution
Comparison of mass spectrometers performances
 Comparison of mass spectrometers performances Instrument Mass Mass Sensitivity resolution accuracy Quadrupole 1 x 10 3 0.1 Da* 0.5-1.0 pmol DE-MALDI 2 x 10 4 20 ppm 1-10 fmol peptide 1-5 pmol protein Ion
Comparison of mass spectrometers performances Instrument Mass Mass Sensitivity resolution accuracy Quadrupole 1 x 10 3 0.1 Da* 0.5-1.0 pmol DE-MALDI 2 x 10 4 20 ppm 1-10 fmol peptide 1-5 pmol protein Ion
Microalbuminuric Diabetic patients N=18
 ONLINE APPENDIX Table A1 Clinical and laboratory features of healthy subjects, type 2 diabetic patients and NDCKD patients enrolled in the present study. Healthy Subjects N= 2 Normoalbuminuric Diabetic
ONLINE APPENDIX Table A1 Clinical and laboratory features of healthy subjects, type 2 diabetic patients and NDCKD patients enrolled in the present study. Healthy Subjects N= 2 Normoalbuminuric Diabetic
Figure S6. A-J) Annotated UVPD mass spectra for top ten peptides found among the peptides identified by Byonic but not SEQUEST + Percolator.
 Extending Proteome Coverage by Combining MS/MS Methods and a Modified Bioinformatics Platform adapted for Database Searching of Positive and Negative Polarity 193 nm Ultraviolet Photodissociation Mass
Extending Proteome Coverage by Combining MS/MS Methods and a Modified Bioinformatics Platform adapted for Database Searching of Positive and Negative Polarity 193 nm Ultraviolet Photodissociation Mass
Introduction to Proteomics 1.0
 Introduction to Proteomics 1.0 CMSP Workshop Pratik Jagtap Managing Director, CMSP Objectives Why are we here? For participants: Learn basics of MS-based proteomics Learn what s necessary for success using
Introduction to Proteomics 1.0 CMSP Workshop Pratik Jagtap Managing Director, CMSP Objectives Why are we here? For participants: Learn basics of MS-based proteomics Learn what s necessary for success using
Structural vs. nonstructural proteins
 Why would you want to study proteins associated with viruses or virus infection? Receptors Mechanism of uncoating How is gene expression carried out, exclusively by viral enzymes? Gene expression phases?
Why would you want to study proteins associated with viruses or virus infection? Receptors Mechanism of uncoating How is gene expression carried out, exclusively by viral enzymes? Gene expression phases?
Databehandling. 3. Mark e.g. the first fraction (1: 0-45 min, 2: min, 3; min, 4: min, 5: min, 6: min).
 Databehandling Data analysis 1. Choose Open in the Data analysis window. 2. Press the Open folder and choose the desired analysis. Click the + button, so that the Chromatograms line appears. Click the
Databehandling Data analysis 1. Choose Open in the Data analysis window. 2. Press the Open folder and choose the desired analysis. Click the + button, so that the Chromatograms line appears. Click the
Mass Spectrometry. Mass spectrometer MALDI-TOF ESI/MS/MS. Basic components. Ionization source Mass analyzer Detector
 Mass Spectrometry MALDI-TOF ESI/MS/MS Mass spectrometer Basic components Ionization source Mass analyzer Detector 1 Principles of Mass Spectrometry Proteins are separated by mass to charge ratio (limit
Mass Spectrometry MALDI-TOF ESI/MS/MS Mass spectrometer Basic components Ionization source Mass analyzer Detector 1 Principles of Mass Spectrometry Proteins are separated by mass to charge ratio (limit
Supporting Information: Protein Corona Analysis of Silver Nanoparticles Exposed to Fish Plasma
 Supporting Information: Protein Corona Analysis of Silver Nanoparticles Exposed to Fish Plasma Jiejun Gao 1, Lu Lin 2, Alexander Wei 2,*, and Maria S. Sepúlveda 1,* 1 Department of Forestry and Natural
Supporting Information: Protein Corona Analysis of Silver Nanoparticles Exposed to Fish Plasma Jiejun Gao 1, Lu Lin 2, Alexander Wei 2,*, and Maria S. Sepúlveda 1,* 1 Department of Forestry and Natural
Shotgun Proteomics MS/MS. Protein Mixture. proteolysis. Peptide Mixture. Time. Abundance. Abundance. m/z. Abundance. m/z 2. Abundance.
 Abundance Abundance Abundance Abundance Abundance Shotgun Proteomics Protein Mixture 1 2 3 MS/MS proteolysis m/z 2 3 Time µlc m/z MS 1 m/z Peptide Mixture m/z Block Diagram of a Mass Spectrometer Sample
Abundance Abundance Abundance Abundance Abundance Shotgun Proteomics Protein Mixture 1 2 3 MS/MS proteolysis m/z 2 3 Time µlc m/z MS 1 m/z Peptide Mixture m/z Block Diagram of a Mass Spectrometer Sample
New Instruments and Services
 New Instruments and Services Liwen Zhang Mass Spectrometry and Proteomics Facility The Ohio State University Summer Workshop 2016 Thermo Orbitrap Fusion http://planetorbitrap.com/orbitrap fusion Thermo
New Instruments and Services Liwen Zhang Mass Spectrometry and Proteomics Facility The Ohio State University Summer Workshop 2016 Thermo Orbitrap Fusion http://planetorbitrap.com/orbitrap fusion Thermo
Determination of Benzodiazepines in Urine by CE-MS/MS
 Determination of Benzodiazepines in Urine by CE-MS/MS Application ote Forensic Toxicology Authors audimir Lucio do Lago Department of Fundamental Chemistry, Institute of Chemistry University of São Paulo,
Determination of Benzodiazepines in Urine by CE-MS/MS Application ote Forensic Toxicology Authors audimir Lucio do Lago Department of Fundamental Chemistry, Institute of Chemistry University of São Paulo,
2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry
 Dr. Sanjeeva Srivastava 1. Fundamental of Mass Spectrometry Role of MS and basic concepts 2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry 2 1 MS basic concepts Mass spectrometry - technique
Dr. Sanjeeva Srivastava 1. Fundamental of Mass Spectrometry Role of MS and basic concepts 2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry 2 1 MS basic concepts Mass spectrometry - technique
Multiplex Protein Quantitation using itraq Reagents in a Gel-Based Workflow
 Multiplex Protein Quantitation using itraq Reagents in a Gel-Based Workflow Purpose Described herein is a workflow that combines the isobaric tagging reagents, itraq Reagents, with the separation power
Multiplex Protein Quantitation using itraq Reagents in a Gel-Based Workflow Purpose Described herein is a workflow that combines the isobaric tagging reagents, itraq Reagents, with the separation power
Bioanalytical Quantitation of Biotherapeutics Using Intact Protein vs. Proteolytic Peptides by LC-HR/AM on a Q Exactive MS
 Bioanalytical Quantitation of Biotherapeutics Using Intact Protein vs. Proteolytic Peptides by LC-HR/AM on a Q Exactive MS Jenny Chen, Hongxia Wang, Zhiqi Hao, Patrick Bennett, and Greg Kilby Thermo Fisher
Bioanalytical Quantitation of Biotherapeutics Using Intact Protein vs. Proteolytic Peptides by LC-HR/AM on a Q Exactive MS Jenny Chen, Hongxia Wang, Zhiqi Hao, Patrick Bennett, and Greg Kilby Thermo Fisher
Mass Spectrometry and Proteomics - Lecture 4 - Matthias Trost Newcastle University
 Mass Spectrometry and Proteomics - Lecture 4 - Matthias Trost Newcastle University matthias.trost@ncl.ac.uk previously Peptide fragmentation Hybrid instruments 117 The Building Blocks of Life DNA RNA Proteins
Mass Spectrometry and Proteomics - Lecture 4 - Matthias Trost Newcastle University matthias.trost@ncl.ac.uk previously Peptide fragmentation Hybrid instruments 117 The Building Blocks of Life DNA RNA Proteins
[ APPLICATION NOTE ] High Sensitivity Intact Monoclonal Antibody (mab) HRMS Quantification APPLICATION BENEFITS INTRODUCTION WATERS SOLUTIONS KEYWORDS
![[ APPLICATION NOTE ] High Sensitivity Intact Monoclonal Antibody (mab) HRMS Quantification APPLICATION BENEFITS INTRODUCTION WATERS SOLUTIONS KEYWORDS [ APPLICATION NOTE ] High Sensitivity Intact Monoclonal Antibody (mab) HRMS Quantification APPLICATION BENEFITS INTRODUCTION WATERS SOLUTIONS KEYWORDS](/thumbs/79/80328542.jpg) Yun Wang Alelyunas, Henry Shion, Mark Wrona Waters Corporation, Milford, MA, USA APPLICATION BENEFITS mab LC-MS method which enables users to achieve highly sensitive bioanalysis of intact trastuzumab
Yun Wang Alelyunas, Henry Shion, Mark Wrona Waters Corporation, Milford, MA, USA APPLICATION BENEFITS mab LC-MS method which enables users to achieve highly sensitive bioanalysis of intact trastuzumab
New Developments in LC-IMS-MS Proteomic Measurements and Informatic Analyses
 New Developments in LC-IMS-MS Proteomic Measurements and Informatic Analyses Erin Shammel Baker Kristin E. Burnum-Johnson, Xing Zhang, Cameron P. Casey, Yehia M. Ibrahim, Matthew E. Monroe, Tao Liu, Brendan
New Developments in LC-IMS-MS Proteomic Measurements and Informatic Analyses Erin Shammel Baker Kristin E. Burnum-Johnson, Xing Zhang, Cameron P. Casey, Yehia M. Ibrahim, Matthew E. Monroe, Tao Liu, Brendan
O O H. Robert S. Plumb and Paul D. Rainville Waters Corporation, Milford, MA, U.S. INTRODUCTION EXPERIMENTAL. LC /MS conditions
 Simplifying Qual/Quan Analysis in Discovery DMPK using UPLC and Xevo TQ MS Robert S. Plumb and Paul D. Rainville Waters Corporation, Milford, MA, U.S. INTRODUCTION The determination of the drug metabolism
Simplifying Qual/Quan Analysis in Discovery DMPK using UPLC and Xevo TQ MS Robert S. Plumb and Paul D. Rainville Waters Corporation, Milford, MA, U.S. INTRODUCTION The determination of the drug metabolism
Supporting Information
 Supporting Information Development of a High Coverage Pseudotargeted Lipidomics Method Based on Ultra-High Performance Liquid Chromatography-Mass Spectrometry Qiuhui Xuan 1,2#, Chunxiu Hu 1#, Di Yu 1,2,
Supporting Information Development of a High Coverage Pseudotargeted Lipidomics Method Based on Ultra-High Performance Liquid Chromatography-Mass Spectrometry Qiuhui Xuan 1,2#, Chunxiu Hu 1#, Di Yu 1,2,
SimGlycan. A high-throughput glycan and glycopeptide data analysis tool for LC-, MALDI-, ESI- Mass Spectrometry workflows.
 PREMIER Biosoft SimGlycan A high-throughput glycan and glycopeptide data analysis tool for LC-, MALDI-, ESI- Mass Spectrometry workflows SimGlycan software processes and interprets the MS/MS and higher
PREMIER Biosoft SimGlycan A high-throughput glycan and glycopeptide data analysis tool for LC-, MALDI-, ESI- Mass Spectrometry workflows SimGlycan software processes and interprets the MS/MS and higher
The N-terminal loop of IRAK-4 death domain regulates ordered assembly of the Myddosome signalling scaffold
 The N-terminal loop of IRAK-4 death domain regulates ordered assembly of the Myddosome signalling scaffold Anthony C.G. Dossang * 1, Precious G. Motshwene * 1, Yang Yang* 1, Martyn F. Symmons*, Clare E.
The N-terminal loop of IRAK-4 death domain regulates ordered assembly of the Myddosome signalling scaffold Anthony C.G. Dossang * 1, Precious G. Motshwene * 1, Yang Yang* 1, Martyn F. Symmons*, Clare E.
Structural Characterization of Prion-like Conformational Changes of the Neuronal Isoform of Aplysia CPEB
 Structural Characterization of Prion-like Conformational Changes of the Neuronal Isoform of Aplysia CPEB Bindu L. Raveendra, 1,5 Ansgar B. Siemer, 2,6 Sathyanarayanan V. Puthanveettil, 1,3,7 Wayne A. Hendrickson,
Structural Characterization of Prion-like Conformational Changes of the Neuronal Isoform of Aplysia CPEB Bindu L. Raveendra, 1,5 Ansgar B. Siemer, 2,6 Sathyanarayanan V. Puthanveettil, 1,3,7 Wayne A. Hendrickson,
Antoine Bouchoux, Pierre-Emerson Cayemitte, Julien Jardin, Geneviève Gésan-Guiziou, and Bernard Cabane
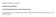 Biophysical Journal, Volume 96 Supplementary Material Casein Micelle Dispersions under Osmotic Stress Antoine Bouchoux, Pierre-Emerson Cayemitte, Julien Jardin, Geneviève Gésan-Guiziou, and Bernard Cabane
Biophysical Journal, Volume 96 Supplementary Material Casein Micelle Dispersions under Osmotic Stress Antoine Bouchoux, Pierre-Emerson Cayemitte, Julien Jardin, Geneviève Gésan-Guiziou, and Bernard Cabane
TECHNICAL BULLETIN. R 2 GlcNAcβ1 4GlcNAcβ1 Asn
 GlycoProfile II Enzymatic In-Solution N-Deglycosylation Kit Product Code PP0201 Storage Temperature 2 8 C TECHNICAL BULLETIN Product Description Glycosylation is one of the most common posttranslational
GlycoProfile II Enzymatic In-Solution N-Deglycosylation Kit Product Code PP0201 Storage Temperature 2 8 C TECHNICAL BULLETIN Product Description Glycosylation is one of the most common posttranslational
SUPPLEMENTAL INFORMATION
 SUPPLEMENTAL INFORMATION EXPERIMENTAL PROCEDURES Tryptic digestion protection experiments - PCSK9 with Ab-3D5 (1:1 molar ratio) in 50 mm Tris, ph 8.0, 150 mm NaCl was incubated overnight at 4 o C. The
SUPPLEMENTAL INFORMATION EXPERIMENTAL PROCEDURES Tryptic digestion protection experiments - PCSK9 with Ab-3D5 (1:1 molar ratio) in 50 mm Tris, ph 8.0, 150 mm NaCl was incubated overnight at 4 o C. The
Metabolomic fingerprinting of serum samples by direct infusion mass spectrometry
 Metabolomic fingerprinting of serum samples by direct infusion mass spectrometry Raúl González-Domínguez * Department of Chemistry, Faculty of Experimental Sciences. University of Huelva, Spain. * Corresponding
Metabolomic fingerprinting of serum samples by direct infusion mass spectrometry Raúl González-Domínguez * Department of Chemistry, Faculty of Experimental Sciences. University of Huelva, Spain. * Corresponding
Rapid, Simple Impurity Characterization with the Xevo TQ Mass Spectrometer
 Robert Plumb, Michael D. Jones, and Marian Twohig Waters Corporation, Milford, MA, USA INTRODUCTION The detection and characterization of impurities and degradation products of an active pharmaceutical
Robert Plumb, Michael D. Jones, and Marian Twohig Waters Corporation, Milford, MA, USA INTRODUCTION The detection and characterization of impurities and degradation products of an active pharmaceutical
Digitizing the Proteomes From Big Tissue Biobanks
 Digitizing the Proteomes From Big Tissue Biobanks Analyzing 24 Proteomes Per Day by Microflow SWATH Acquisition and Spectronaut Pulsar Analysis Jan Muntel 1, Nick Morrice 2, Roland M. Bruderer 1, Lukas
Digitizing the Proteomes From Big Tissue Biobanks Analyzing 24 Proteomes Per Day by Microflow SWATH Acquisition and Spectronaut Pulsar Analysis Jan Muntel 1, Nick Morrice 2, Roland M. Bruderer 1, Lukas
Supporting information
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2014 Supporting information Glycan Reductive Isotope-coded Amino Acid Labeling (GRIAL) for Mass Spectrometry-based
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2014 Supporting information Glycan Reductive Isotope-coded Amino Acid Labeling (GRIAL) for Mass Spectrometry-based
Metabolomics: quantifying the phenotype
 Metabolomics: quantifying the phenotype Metabolomics Promises Quantitative Phenotyping What can happen GENOME What appears to be happening Bioinformatics TRANSCRIPTOME What makes it happen PROTEOME Systems
Metabolomics: quantifying the phenotype Metabolomics Promises Quantitative Phenotyping What can happen GENOME What appears to be happening Bioinformatics TRANSCRIPTOME What makes it happen PROTEOME Systems
4th Multidimensional Chromatography Workshop Toronto (January, 2013) Herman C. Lam, Ph.D. Calibration & Validation Group
 4th Multidimensional Chromatography Workshop Toronto (January, 2013) Herman C. Lam, Ph.D. Calibration & Validation Group MDLC for Shotgun Proteomics Introduction General concepts Advantages Challenges
4th Multidimensional Chromatography Workshop Toronto (January, 2013) Herman C. Lam, Ph.D. Calibration & Validation Group MDLC for Shotgun Proteomics Introduction General concepts Advantages Challenges
Dr. Erin E. Chambers Waters Corporation. Presented by Dr. Diego Rodriguez Cabaleiro Waters Europe Waters Corporation 1
 Development of an SPE-LC/MS/MS Assay for the Simultaneous Quantification of Amyloid Beta Peptides in Cerebrospinal Fluid in Support of Alzheimer s Research Dr. Erin E. Chambers Waters Corporation Presented
Development of an SPE-LC/MS/MS Assay for the Simultaneous Quantification of Amyloid Beta Peptides in Cerebrospinal Fluid in Support of Alzheimer s Research Dr. Erin E. Chambers Waters Corporation Presented
Supplementary Information. Top-down/bottom-up mass spectrometry workflow using dissolvable polyacrylamide gels
 Supplementary Information Top-down/bottom-up mass spectrometry workflow using dissolvable polyacrylamide gels Nobuaki Takemori,,* Ayako Takemori,, Piriya Wongkongkathep, Michael Nshanian, Rachel R. Ogorzalek
Supplementary Information Top-down/bottom-up mass spectrometry workflow using dissolvable polyacrylamide gels Nobuaki Takemori,,* Ayako Takemori,, Piriya Wongkongkathep, Michael Nshanian, Rachel R. Ogorzalek
Analysis of Testosterone, Androstenedione, and Dehydroepiandrosterone Sulfate in Serum for Clinical Research
 Analysis of Testosterone, Androstenedione, and Dehydroepiandrosterone Sulfate in Serum for Clinical Research Dominic Foley, Michelle Wills, and Lisa Calton Waters Corporation, Wilmslow, UK APPLICATION
Analysis of Testosterone, Androstenedione, and Dehydroepiandrosterone Sulfate in Serum for Clinical Research Dominic Foley, Michelle Wills, and Lisa Calton Waters Corporation, Wilmslow, UK APPLICATION
Peptide sequencing using chemically assisted fragmentation (CAF) and Ettan MALDI-ToF Pro mass spectrometry
 application note Ett MALDI-ToF Pro Peptide sequencing using chemically assisted fragmentation (CAF) d Ett MALDI-ToF Pro mass spectrometry Key words: MALDI-ToF, PSD, peptide sequencing, chemically assisted
application note Ett MALDI-ToF Pro Peptide sequencing using chemically assisted fragmentation (CAF) d Ett MALDI-ToF Pro mass spectrometry Key words: MALDI-ToF, PSD, peptide sequencing, chemically assisted
Application Note # LCMS-89 High quantification efficiency in plasma targeted proteomics with a full-capability discovery Q-TOF platform
 Application Note # LCMS-89 High quantification efficiency in plasma targeted proteomics with a full-capability discovery Q-TOF platform Abstract Targeted proteomics for biomarker verification/validation
Application Note # LCMS-89 High quantification efficiency in plasma targeted proteomics with a full-capability discovery Q-TOF platform Abstract Targeted proteomics for biomarker verification/validation
Don t miss a thing on your peptide mapping journey How to get full coverage peptide maps using high resolution accurate mass spectrometry
 Don t miss a thing on your peptide mapping journey How to get full coverage peptide maps using high resolution accurate mass spectrometry Kai Scheffler, PhD BioPharma Support Expert,LSMS Europe The world
Don t miss a thing on your peptide mapping journey How to get full coverage peptide maps using high resolution accurate mass spectrometry Kai Scheffler, PhD BioPharma Support Expert,LSMS Europe The world
Post-Lab Activity STUDENT MANUAL POST-LAB ACTIVITY. Analysis and Interpretation of Results
 STUDENT MANUAL POST-LAB ACTIVITY Post-Lab Activity Analysis and Interpretation of Results Detailed Gel Analysis Does molecular evidence support or refute the theory of evolution? Does your molecular evidence
STUDENT MANUAL POST-LAB ACTIVITY Post-Lab Activity Analysis and Interpretation of Results Detailed Gel Analysis Does molecular evidence support or refute the theory of evolution? Does your molecular evidence
A Definitive Lipidomics Workflow for Human Plasma Utilizing Off-line Enrichment and Class Specific Separation of Phospholipids
 A Definitive Lipidomics Workflow for Human Plasma Utilizing Off-line Enrichment and Class Specific Separation of Phospholipids Jeremy Netto, 1 Stephen Wong, 1 Federico Torta, 2 Pradeep Narayanaswamy, 2
A Definitive Lipidomics Workflow for Human Plasma Utilizing Off-line Enrichment and Class Specific Separation of Phospholipids Jeremy Netto, 1 Stephen Wong, 1 Federico Torta, 2 Pradeep Narayanaswamy, 2
Supplementary material: Materials and suppliers
 Supplementary material: Materials and suppliers Electrophoresis consumables including tris-glycine, acrylamide, SDS buffer and Coomassie Brilliant Blue G-2 dye (CBB) were purchased from Ameresco (Solon,
Supplementary material: Materials and suppliers Electrophoresis consumables including tris-glycine, acrylamide, SDS buffer and Coomassie Brilliant Blue G-2 dye (CBB) were purchased from Ameresco (Solon,
SCS MOLECULAR WEIGHT MARKERS 2,500-17,000 Caltons
 LECTROPHORES/S Revised November 1992 SCS MOLECULAR WEIGHT MARKERS 2,500-17,000 Caltons I NTRODUCTION Electrophoresis in polyacrylamide gels in the presence of sodium dodecyl sulfate (SDS), an anionic detergent,
LECTROPHORES/S Revised November 1992 SCS MOLECULAR WEIGHT MARKERS 2,500-17,000 Caltons I NTRODUCTION Electrophoresis in polyacrylamide gels in the presence of sodium dodecyl sulfate (SDS), an anionic detergent,
Proteomic tools to assess meat authenticity
 Proteomic tools to assess meat authenticity Miguel A. Sentandreu & Enrique Sentandreu Institute of Agrochemistry and Food Technology (CSIC) Spain The problem of meat authentication Clear and reliable information
Proteomic tools to assess meat authenticity Miguel A. Sentandreu & Enrique Sentandreu Institute of Agrochemistry and Food Technology (CSIC) Spain The problem of meat authentication Clear and reliable information
Application Note # ET-17 / MT-99 Characterization of the N-glycosylation Pattern of Antibodies by ESI - and MALDI mass spectrometry
 Bruker Daltonics Application Note # ET-17 / MT-99 Characterization of the N-glycosylation Pattern of Antibodies by ESI - and MALDI mass spectrometry Abstract Analysis of the N-glycosylation pattern on
Bruker Daltonics Application Note # ET-17 / MT-99 Characterization of the N-glycosylation Pattern of Antibodies by ESI - and MALDI mass spectrometry Abstract Analysis of the N-glycosylation pattern on
Glycosylation analysis of blood plasma proteins
 Glycosylation analysis of blood plasma proteins Thesis booklet Eszter Tóth Doctoral School of Pharmaceutical Sciences Semmelweis University Supervisor: Károly Vékey DSc Official reviewers: Borbála Dalmadiné
Glycosylation analysis of blood plasma proteins Thesis booklet Eszter Tóth Doctoral School of Pharmaceutical Sciences Semmelweis University Supervisor: Károly Vékey DSc Official reviewers: Borbála Dalmadiné
LECTURE-15. itraq Clinical Applications HANDOUT. Isobaric Tagging for Relative and Absolute quantitation (itraq) is a quantitative MS
 LECTURE-15 itraq Clinical Applications HANDOUT PREAMBLE Isobaric Tagging for Relative and Absolute quantitation (itraq) is a quantitative MS based method for quantifying proteins subject to various different
LECTURE-15 itraq Clinical Applications HANDOUT PREAMBLE Isobaric Tagging for Relative and Absolute quantitation (itraq) is a quantitative MS based method for quantifying proteins subject to various different
Development of a Bioanalytical Method for Quantification of Amyloid Beta Peptides in Cerebrospinal Fluid
 Development of a Bioanalytical Method for Quantification of Amyloid Beta Peptides in Cerebrospinal Fluid Joanne ( 乔安妮 ) Mather Senior Scientist Waters Corporation Data courtesy of Erin Chambers and Mary
Development of a Bioanalytical Method for Quantification of Amyloid Beta Peptides in Cerebrospinal Fluid Joanne ( 乔安妮 ) Mather Senior Scientist Waters Corporation Data courtesy of Erin Chambers and Mary
Application of LC/Electrospray Ion Trap Mass Spectrometry for Identification and Quantification of Pesticides in Complex Matrices
 Application ote #LCMS-2 esquire series Application of LC/Electrospray Ion Trap Mass Spectrometry for Identification and Quantification of Pesticides in Complex Matrices Introduction The simple monitoring
Application ote #LCMS-2 esquire series Application of LC/Electrospray Ion Trap Mass Spectrometry for Identification and Quantification of Pesticides in Complex Matrices Introduction The simple monitoring
AbsoluteIDQ p180 Kit. Targeted Metabolite Identifi cation and Quantifi cation
 AbsoluteIDQ p180 Kit Targeted Metabolite Identifi cation and Quantifi cation Bringing our targeted metabolomics expertise to your lab. The Biocrates AbsoluteIDQ p180 mass spectrometry Assay Preparation
AbsoluteIDQ p180 Kit Targeted Metabolite Identifi cation and Quantifi cation Bringing our targeted metabolomics expertise to your lab. The Biocrates AbsoluteIDQ p180 mass spectrometry Assay Preparation
Sequence Coverage (%) Profilin-1 P UD 2
 Protein Name Accession Number (Swissprot) Sequence Coverage (%) No. of MS/MS Queries Mascot Score 1 Reference Cytoskeletal proteins Beta-actin P60709 37 14 298 Alpha-actin P68032 33 10 141 20 Beta-actin-like
Protein Name Accession Number (Swissprot) Sequence Coverage (%) No. of MS/MS Queries Mascot Score 1 Reference Cytoskeletal proteins Beta-actin P60709 37 14 298 Alpha-actin P68032 33 10 141 20 Beta-actin-like
Practical experiments / Oil/protein crops
 ASPECTS OF PRODUCT QUALITY IN PLANT PRODUCTION Practical experiments / Oil/protein crops 1. Glucosinolates 2. NIRS for oil / protein / carbohydrate content Analytical methods for crop Pre-requisites quality
ASPECTS OF PRODUCT QUALITY IN PLANT PRODUCTION Practical experiments / Oil/protein crops 1. Glucosinolates 2. NIRS for oil / protein / carbohydrate content Analytical methods for crop Pre-requisites quality
* Skyline LC SRM. 2 Skyline LC-MS/MS. Skyline University of Washington Windows.
 Proteome Letters 2016 1 89-94 * *E-mail: fmatsuda@ist.osaka-u.ac.jp 565-0871 1-5 2016 4 20 2016 5 26 2016 5 27 LC SRM LC SRM 1 LC-MS/MS LC 1 3 SRM SRM LC-MS Skyline SRM LC-MS LC SRM 2 Skyline Skyline University
Proteome Letters 2016 1 89-94 * *E-mail: fmatsuda@ist.osaka-u.ac.jp 565-0871 1-5 2016 4 20 2016 5 26 2016 5 27 LC SRM LC SRM 1 LC-MS/MS LC 1 3 SRM SRM LC-MS Skyline SRM LC-MS LC SRM 2 Skyline Skyline University
SUNY UPSTATE MEDICAL UNIVERSITY PROTEOMICS CORE
 SUNY UPSTATE MEDICAL UNIVERSITY PROTEOMICS CORE Location: SUNY Upstate Medical University Weiskotten Hall Addition, Room 4303 750 East Adams Street Syracuse, NY 13210 Contact Information: David Kakhniashvili,
SUNY UPSTATE MEDICAL UNIVERSITY PROTEOMICS CORE Location: SUNY Upstate Medical University Weiskotten Hall Addition, Room 4303 750 East Adams Street Syracuse, NY 13210 Contact Information: David Kakhniashvili,
Quantification with Proteome Discoverer. Bernard Delanghe
 Quantification with Proteome Discoverer Bernard Delanghe Overview: Which approach to use? Proteome Discoverer Quantification Method What When to use Metabolic labeling SILAC Cell culture systems Small
Quantification with Proteome Discoverer Bernard Delanghe Overview: Which approach to use? Proteome Discoverer Quantification Method What When to use Metabolic labeling SILAC Cell culture systems Small
Application of a new capillary HPLC- ICP-MS interface to the identification of selenium-containing proteins in selenized yeast
 Application of a new capillary HPLC- ICP-MS interface to the identification of selenium-containing proteins in selenized yeast Application note Food supplements Authors Juliusz Bianga and Joanna Szpunar
Application of a new capillary HPLC- ICP-MS interface to the identification of selenium-containing proteins in selenized yeast Application note Food supplements Authors Juliusz Bianga and Joanna Szpunar
Trypsin Mass Spectrometry Grade
 058PR-03 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Trypsin Mass Spectrometry Grade A Chemically Modified, TPCK treated, Affinity Purified
058PR-03 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Trypsin Mass Spectrometry Grade A Chemically Modified, TPCK treated, Affinity Purified
4-Plex itraq Based Quantitative Proteomic Analysis Using an Agilent Accurate -Mass Q-TOF
 4-Plex itraq Based Quantitative Proteomic Analysis Using an Agilent Accurate -Mass Q-TOF Application Note Authors H. C. Harsha, G. S. S. Kumar, and A. Pandey Institute of Bioinformatics Bangalore India
4-Plex itraq Based Quantitative Proteomic Analysis Using an Agilent Accurate -Mass Q-TOF Application Note Authors H. C. Harsha, G. S. S. Kumar, and A. Pandey Institute of Bioinformatics Bangalore India
Identification & Confirmation of Structurally Related Degradation Products of Simvastatin
 Identification & Confirmation of Structurally Related Degradation Products of Simvastatin Power of QTRAP Systems for Identification and Confirmation of Degradation Products Dilip Reddy 1, Chandra Sekar
Identification & Confirmation of Structurally Related Degradation Products of Simvastatin Power of QTRAP Systems for Identification and Confirmation of Degradation Products Dilip Reddy 1, Chandra Sekar
Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random
 S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
One Gene, Many Proteins. Applications of Mass Spectrometry to Proteomics. Why Proteomics? Raghothama Chaerkady, Ph.D.
 Applications of Mass Spectrometry to Proteomics Raghothama Chaerkady, Ph.D. McKusick-Nathans Institute of Genetic Medicine and the Department of Biological Chemistry Why Proteomics? One Gene, Many Proteins
Applications of Mass Spectrometry to Proteomics Raghothama Chaerkady, Ph.D. McKusick-Nathans Institute of Genetic Medicine and the Department of Biological Chemistry Why Proteomics? One Gene, Many Proteins
Systematic analysis of protein-detergent complexes applying dynamic light scattering to optimize solutions for crystallization trials
 Supporting information 1 2 3 Volume 71 (2015) Supporting information for article: 4 5 6 7 8 Systematic analysis of protein-detergent complexes applying dynamic light scattering to optimize solutions for
Supporting information 1 2 3 Volume 71 (2015) Supporting information for article: 4 5 6 7 8 Systematic analysis of protein-detergent complexes applying dynamic light scattering to optimize solutions for
Jose Castro-Perez, Henry Shion, Kate Yu, John Shockcor, Emma Marsden-Edwards, Jeff Goshawk Waters Corporation, Milford, MA, U.S. and Manchester, UK
 HIGH-THRUGHPUT REACTIVE METABLITE SCREEIG FR DICLFEAC BY UPLC AD XEV TQ MS WITH SCAWAVE Jose Castro-Perez, Henry Shion, Kate Yu, John Shockcor, Emma Marsden-Edwards, Jeff Goshawk Waters Corporation, Milford,
HIGH-THRUGHPUT REACTIVE METABLITE SCREEIG FR DICLFEAC BY UPLC AD XEV TQ MS WITH SCAWAVE Jose Castro-Perez, Henry Shion, Kate Yu, John Shockcor, Emma Marsden-Edwards, Jeff Goshawk Waters Corporation, Milford,
An Alternative Approach: Top-Down Bioanalysis of Intact Large Molecules Can this be part of the future? Lecture 8, Page 27
 An Alternative Approach: Top-Down Bioanalysis of Intact Large Molecules Can this be part of the future? Lecture 8, Page 27 Top-down HRAM Bioanalysis of Native Proteins/Molecules Relative Abundance 100
An Alternative Approach: Top-Down Bioanalysis of Intact Large Molecules Can this be part of the future? Lecture 8, Page 27 Top-down HRAM Bioanalysis of Native Proteins/Molecules Relative Abundance 100
Identification of Ginsenosides Using the SCIEX X500R QTOF System
 Identification of Ginsenosides Using the SCIEX X500R QTOF System Wang Sha, Cheng Haiyan, Liu Ting, Li Lijun, Jin Wenhai[Author] SCIEX, Pacific Applications Support Center (Beijing). China Background Ginseng
Identification of Ginsenosides Using the SCIEX X500R QTOF System Wang Sha, Cheng Haiyan, Liu Ting, Li Lijun, Jin Wenhai[Author] SCIEX, Pacific Applications Support Center (Beijing). China Background Ginseng
Fig.S1 ESI-MS spectrum of reaction of ApA and THPTb after 16 h.
 Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2014 Experiment Cleavage of dinucleotides Dinucleotides (ApA, CpC, GpG, UpU) were purchased from
Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2014 Experiment Cleavage of dinucleotides Dinucleotides (ApA, CpC, GpG, UpU) were purchased from
SUPPLEMENTARY MATERIAL
 SUPPLEMENTARY MATERIAL Purification and biochemical properties of SDS-stable low molecular weight alkaline serine protease from Citrullus Colocynthis Muhammad Bashir Khan, 1,3 Hidayatullah khan, 2 Muhammad
SUPPLEMENTARY MATERIAL Purification and biochemical properties of SDS-stable low molecular weight alkaline serine protease from Citrullus Colocynthis Muhammad Bashir Khan, 1,3 Hidayatullah khan, 2 Muhammad
Quantitation of Protein Phosphorylation Using Multiple Reaction Monitoring
 Quantitation of Protein Phosphorylation Using Multiple Reaction Monitoring Application Note Authors Ning Tang, Christine A. Miller and Keith Waddell Agilent Technologies, Inc. Santa Clara, CA USA This
Quantitation of Protein Phosphorylation Using Multiple Reaction Monitoring Application Note Authors Ning Tang, Christine A. Miller and Keith Waddell Agilent Technologies, Inc. Santa Clara, CA USA This
Use of Derivatization for LC- MS/MS Analysis in the Clinical Lab
 Use of Derivatization for LC- MS/MS Analysis in the Clinical Lab Asian Pacific Conference on Chromatography & Mass Spectrometry 2010 15 January 2010 Alan L. Rockwood and Mark M. Kushnir ARUP Laboratories,
Use of Derivatization for LC- MS/MS Analysis in the Clinical Lab Asian Pacific Conference on Chromatography & Mass Spectrometry 2010 15 January 2010 Alan L. Rockwood and Mark M. Kushnir ARUP Laboratories,
LABORATÓRIUMI GYAKORLAT SILLABUSZ SYLLABUS OF A PRACTICAL DEMONSTRATION. financed by the program
 TÁMOP-4.1.1.C-13/1/KONV-2014-0001 projekt Az élettudományi-klinikai felsőoktatás gyakorlatorientált és hallgatóbarát korszerűsítése a vidéki képzőhelyek nemzetközi versenyképességének erősítésére program
TÁMOP-4.1.1.C-13/1/KONV-2014-0001 projekt Az élettudományi-klinikai felsőoktatás gyakorlatorientált és hallgatóbarát korszerűsítése a vidéki képzőhelyek nemzetközi versenyképességének erősítésére program
Probability-Based Protein Identification for Post-Translational Modifications and Amino Acid Variants Using Peptide Mass Fingerprint Data
 Probability-Based Protein Identification for Post-Translational Modifications and Amino Acid Variants Using Peptide Mass Fingerprint Data Tong WW, McComb ME, Perlman DH, Huang H, O Connor PB, Costello
Probability-Based Protein Identification for Post-Translational Modifications and Amino Acid Variants Using Peptide Mass Fingerprint Data Tong WW, McComb ME, Perlman DH, Huang H, O Connor PB, Costello
Research Article A Colorimetric Method for Monitoring Tryptic Digestion Prior to Shotgun Proteomics
 International Journal of Proteomics Volume 24, Article ID 25482, 9 pages http://dx.doi.org/.55/24/25482 Research Article A Colorimetric Method for Monitoring Tryptic Digestion Prior to Shotgun Proteomics
International Journal of Proteomics Volume 24, Article ID 25482, 9 pages http://dx.doi.org/.55/24/25482 Research Article A Colorimetric Method for Monitoring Tryptic Digestion Prior to Shotgun Proteomics
A computational framework for discovery of glycoproteomic biomarkers
 A computational framework for discovery of glycoproteomic biomarkers Haixu Tang, Anoop Mayampurath, Chuan-Yih Yu Indiana University, Bloomington Yehia Mechref, Erwang Song Texas Tech University 1 Goal:
A computational framework for discovery of glycoproteomic biomarkers Haixu Tang, Anoop Mayampurath, Chuan-Yih Yu Indiana University, Bloomington Yehia Mechref, Erwang Song Texas Tech University 1 Goal:
DetergentOUT Tween. DetergentOUT GBS10. OrgoSol DetergentOUT
 252PR 01 G-Biosciences, St Louis, MO. USA 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name DetergentOUT Detergent Removal Systems For the Removal of Detergents
252PR 01 G-Biosciences, St Louis, MO. USA 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name DetergentOUT Detergent Removal Systems For the Removal of Detergents
Supplementary Figures and Notes for Digestion and depletion of abundant
 Supplementary Figures and Notes for Digestion and depletion of abundant proteins improves proteomic coverage Bryan R. Fonslow, Benjamin D. Stein, Kristofor J. Webb, Tao Xu, Jeong Choi, Sung Kyu Park, and
Supplementary Figures and Notes for Digestion and depletion of abundant proteins improves proteomic coverage Bryan R. Fonslow, Benjamin D. Stein, Kristofor J. Webb, Tao Xu, Jeong Choi, Sung Kyu Park, and
New Mass Spectrometry Tools to Transform Metabolomics and Lipidomics
 New Mass Spectrometry Tools to Transform Metabolomics and Lipidomics July.3.13 Ken Miller Vice President of Marketing, Life Sciences Mass Spectrometry 1 The world leader in serving science Omics & the
New Mass Spectrometry Tools to Transform Metabolomics and Lipidomics July.3.13 Ken Miller Vice President of Marketing, Life Sciences Mass Spectrometry 1 The world leader in serving science Omics & the
Technical Report. Abstract: 2. Experiment. 1. Introduction. Hiroshi Tsugawa 1,*, Eiichiro Fukusaki 1
 C146-E217 Technical Report Effectiveness of Metabolomics Research Using Gas Chromatograph / Quadrupole Mass Spectrometer with High-Sensitivity and High-Speed Scanning Hiroshi Tsugawa 1,*, Eiichiro Fukusaki
C146-E217 Technical Report Effectiveness of Metabolomics Research Using Gas Chromatograph / Quadrupole Mass Spectrometer with High-Sensitivity and High-Speed Scanning Hiroshi Tsugawa 1,*, Eiichiro Fukusaki
