Isotope dilution of intracellular amino acids as a tracer of carbon and nitrogen sources of marine planktonic bacteria
|
|
|
- Claribel Butler
- 6 years ago
- Views:
Transcription
1 Vol. 74: , 1991 MARINE ECOLOGY PROGRESS SERIES Mar. Ecol. Prog. Ser. Published August 8 Isotope dilution of intracellular amino acids as a tracer of carbon and nitrogen sources of marine planktonic bacteria Meinhard Simon Limnological Institute, University of Constance, PO Box 5560, W-7750 Konstanz, Germany ABSTRACT- Isotope dilution (ID) of intracellular amino acids In bacterial assemblages in the subarctic Paclflc was measured in order to examine the relat~onshlp between dissolved free anlino aclds (DFAA) and other sources of amino acids for protein synthesis When 3~-an~ino acids were added at trace amounts to <0.8 pm filtrates, ID of single amino aclds vaned greatly. Isotope dilutlon of senne, glyclne, threonine, arginlne, alanine, and tyrosine was 1- to 4-fold, indicating that 25 to 100 O/o of bacterlal biosynthetlc requirements was met by uptake of these amlno acids Vallne, phenylalanine, isoleucine, and leucine exhibited 8- to 50-fold ID Indicating that only a minor fractlon of bacterlal biosynthetic requirements was met by uptake of these amlno acids. These results were consistent wlth measurements of the fractlon of bacterlal biomass productlon supported by uptake of DFAA (8 to 42 '%). Intracellular Isotope dllutlon appeared to be Inversely related to the extracellular concentration of the respective DFAA Glutamate ID was always hlgh (13- to 95-fold) and independent of its concentration. Thls high ID was probably due to substantial ammonium uptake, indicating that In addition to DFAA, ammonium was an Important N-source for amlno acld de novo synthesis and thus for proteln productlon. INTRODUCTION Heterotrophic bacteria are important in the turnover of organic matter in aquatic ecosystems. In pelagic environments they are the predominant group of organisms that assimilate dissolved organic matter (DOM) and transform it into b~omass, making it available to higher trophic levels. Some of the most important DOM components for bacterial growth are dissolved free amino acids (DFAA; Kirchman 1990). As a combined C and N source, they are immediate precursors for protein which comprises 60 '10 of bacterial biomass (Simon & Azam 1989). The importance of DFAA in supporting bacterial growth is usually evaluated by comparing DFAA uptake (DFAA concentrations X turnover of radiolabelled DFAA) with bacterial biomass production. These studies indicate that the fraction of bacterial production supported by uptake of DFAA can vary from <5 to 100 '10 in marine and freshwater environments depending on the substrate supply (J~rgensen 1987, 1990, Fuhrman 1990). To examine the biosynthetic processes of bacterial assemblages in more detail, another approach is to study the contribution of DFAA to intracellular pools, through which all precursors for protein synthesis must pass (Anraku 1980). Amino acids can enter the intracellular pool via 4 possible pathways (Payne 1980): (1) direct uptake of DFAA; (2) uptake of oligopeptides (2 to 5 amino acids) and subsequent intracellular hydrolysis; (3) cell-surface-mediated hydrolysis of dissolved combined amino acids (DCAA) with coupled uptake of single amino acids or oligopeptides; (4) de novo synthesis of amino acids from ammonium and organic carbon. Recently, Simon & Azam (1989) introduced a method by which some of these biosynthetic pathways can be studied in natural assemblages of aquatic bacteria. They measured isotope dilution (ID) of 3H-leucine in the bacterial intracellular pool by comparing the specific activity of 3~-leucine in the bulk water with that in the intracellular bacterial pool. By extending this approach to other amino acids, one can differentiate among the various sources of amino acids contributing to the intracellular pool. High ID would indicate a low contribution of DFAA compared with other O Inter-Research/Prlnted ln Germany /91/0074/0295/$ 03 00
2 296 Mar Ecol. Prog. Ser 74: , 1991 pathways (1). To further differentiate pathways (2), (3), and (4), the IDs of individual amino acids can be compared. If ammonium is the major N source and not DCAA, ID of glutamate should be significantly higher than ID of other amino acids since glutamate is a key metabolite in ammonium uptake together with cuketoglutarate (Brown 1980), which, after transamlnation to glutamate, should dilute the specific activity of 3~-glutamate. I addressed these questions during a study in the subarctic Pacific as part of the NSF-supported SUbarctic Pacific Ecosystem Research (SUPER) program. The overall goal was to examine the role of heterotrophic bacteria in the N budget of these waters, which have high concentrations of inorganic N and P (Anderson et al. 1977, Wheeler & Kokkinakis 1990). The results of this study indicate that ID of single tritiated amino acids in the intracellular pool varied between no dilution and 95-fold ID, depending on the amino acid and extracellular concentratlon. High ID of glutamate suggested that ammonium uptake and de novo synthesis of amino acids was important for bacterial protein production. MATERIALS AND METHODS Experiments were carried out during May 1988 at Station P (145" W, 50' N) in the Gulf of Alaska. Water was collected in 30 1 Teflon-lined GoFlo bottles with Kevlar line. The basic experiments consisted of adding various 3 H - ~ at ~ tracer ~ ~ concentrations s to water samples and measuring uptake rates and isotope dilution of amino acids in the intracellular pool of bacteria (see below). This was compared to bacterial protein, C, and N production rates measured by incorporation of 3 ~ or - 14C-leucine (Leu; see below). Bacterial numbers were counted with the standard acridine orange technique (Hobbie et al. 1977). Amino acid analysis. High performance liquid chromatographic (HPLC) analysis of DFAA after o- phthaldialdehyde pre-column derivatization (Lindroth & Mopper 1979) in water samples and the extracted pool was carried out according to Simon & Azam (1989). A Rainin HPLC system was used. Although most samples were analyzed in the lab after the cruise, some were analyzed on board ship immediately after or within 48 h of the experiment. All analyses gave comparable results irrespective of the time the samples were kept frozen at -20 "C. Replicate analyses usually agreed within 5 Yo. To estimate SA,,, or SA,,, (see below), 1 m1 fractions (flow rate = 1 m1 min-') were collected after separation by HPLC and radioassayed. Isotope dilution. Isotope dilution was measured as described by Simon & Azam (1989) with the following modifications. Water was filtered through 0.8 ym ASP GLU ASU SER GLN HfS GCY ARC A U VAL PHE ILE LEU THR TYR Fig. 1. Mol% of amino acids in the bacterial intracellular pool. Mean of all samples analyzed (N = 7) and standard deviation. Asp: aspartate; Glu: glutamate; Asn: asparagine; Ser. serine; Gin. glutamine; HIS: histidine; Gly: glycine; Thr: threonine; Arg. arginine; Ala alanine; Tyr: tyrosine; Val: valine; Phe. phenylalanine; Ile: isoleucine: Leu: leucine. Gly/Thr and Ala/ Tyr CO-eluted in the HPLC analyses Nuclepore filters by gravity, removing essentially all chlorophyll a and cyanobactena but only 3 O/O of heterotrophic bacteria (Kirchman et al. 1989). A mixture of L- 3~ amino acids (mean specific activity 57 Ci mmol-', NET-250), L-(3,4 3H) glutamate (69.7 Ci mmol-l), or L-(4,5 3 ~ leucine ) (60 Ci mmol-l) were added at flnal concentrations of 0.5 or 10 nm to triplicate 250 m1 samples in polycarbonate bottles. All radiolabels were from New England Nuclear. A subsample was withdrawn at the start of the incubation for HPLC analysis of DFAA concentration and extracellular specific activity. After incubation at in situ temperature in the dark, uptake was stopped after 1.5 to 4 h by rapid filtration through a 0.2 pm, 47 mm Nuclepore filter. Filters were dipped in boiling HPLC-grade water for 4 min to extract the intracellular pool. Extraction times between 30 s and 4 min yielded similar amino acid pool concentrations (Simon & Azam unpubl.). Longer extraction resulted in protein hydrolysis which led to enhanced amino acid concentrations and mol% distributions significantly different from that of the intracellular pool (Fig. 1). Five minute extraction already increased the amlno acid concentration by a factor of 4 and reduced the glutamate mol% to < 20 %. The extracted samples were placed on ice and kept frozen (-20 "C) until HPLC analysis within 4 wk (see above). Isotope dilution (ID) was calculated as ID = SA,,,/SAin,, where SA,,, is specific activity of the amino acld in the bulk water (external pool); SAin, is specific activity of the amino acid in the intracellular pool. Specific activity was calculated as the ratio of the radioactivity coeluting wlth a given amino acid to the amino acid concen-
3 Simon: C and N sources of marine bacter~a 297 tration. Since the specific activity is calculated by 2 variables the standard error of the triplicate determination is ca 50 O/O. Incorporation of amino acids and estimates of bacterial production. Incorporation rates of a mixture of L-3H amino acids, L-3~ glutamate (specifications see above), and L-14C Leu (342 mci mmol-l, New England Nuclear) into the ice-cold trichloroacetic acid (TCA) precipitate were measured according to Kirchman et al. (1985). The amino acid mixture and glutamate were added at 0.5 nm and Leu was added at 10 nm final concentration. Samples were incubated at surface seawater temperature in the dark (triplicates or duplicates and a killed control) and uptake was stopped after 2 to 3 h by filtration onto 0.45 pm cellulose membrane filters. Total DFAA incorporation rates were calculated from ambient DFAA concentrations and the turnover rates of 3~-amino acids. DFAA incorporation rates were converted to C assuming 60 g C (m01 DFAA)~' (50 % of the mean formula weight) and to N by a calculated C:N ratio of 2.6 for the added DFAA mixture. Leucine incorporation (leu,,,,) was converted to bacterial protein production (BPP) according to Simon & Azam (1989): BPP (g C 1-l h-') = leui,,, X 100/7.3 X FW X ID X F where leui,, is exogenous leucine incorporated into the ice-cold TCA precipitate (m011-l h-'), ID is isotope dilution (see above), FWis formula weight of leucine (131.2 g mol-l), Fis fraction of leu,,, in the hot TCA precipitate (0.9). Protein production was further converted to total bacterial C production assuming a C:protein ratio of 0.86 for marine bacteria (Simon & Azam 1989) and to N assuming a C:N ratio of 4 (Nagata 1986, Lee & Fuhrman 1987) Bacterial numbers (10 l~ter ) RESULTS Amino acid isotope dilution in the intracellular pool To examine sources of amino acids for biosynthetic requirements, IDs of various amino acids in the bacterial intracellular pool were measured. Samples for these experiments were taken from the surface and 40 m where bacterial production rates were often at a maximum (Fig. 2). Bacterial numbers in these depths ranged between 5.7 X 10' and 1.0 X 10' cells 1-' (Fig. 2). In situ concentrations of total DFAA were between 5 and 89 nm (Table 1). During pre-filtration and sample manipulation DFAA concentrations in the <0.8 km filtrates increased substantially as compared to the in situ concentrations (Table 1). Serine, glycine+ threonine, arginine, and alanine+tyrosine comprised > 70 % of total DFAA concentrations in situ as well as in the <0.8 pm filtrates. To determine the amino acid concentrations in the bacterial intracellular pool, the volume of the total bacterial assemblage was estimated from the bacterial abundance assuming a mean cell volume of 0.07 pm3 (Lee & Fuhrman 1987) and a water content of 64 O/O (Simon & Azam 1989). Amino acid amounts in the intracellular pool samples were thus divided by the total bacterial pool volume. Pool concentrations ranged from 28 to 135 mm (Table 1). The intracellular pool was dominated by glutamate which comprised 41 m01 O/O of all amino acids (Fig. 1). Other amino acids did not exceed 15 mol% each. Isotope dilution of amino acids in the intracellular pool was rather variable and depended on the amino acid and the extracellular concentration (Table 2) Bacterial production (ng C liter h ) Fig. 2. (A) Bacterial abundance and (B) production on 15 May (o), 17 May (a), and 28 May 1988 (A) at Station P (145'W, 50 N) in the subarct~c Pacific
4 298 Mar Ecol. Prog. Ser 74: , 1991 Table 1 Total DFAA concentration in situ, in <0.8 him filtrates was relatively constant in all experiments (mean = and in the bacterial intracellular pool of the isotope dilution ). experiments. Leucine and glutamate were added at 0.5 and 10 nm final concentration, and the amino acid mixture (AAmix) at 0 5 nm final concentration of each individual amino acid. nd: not determined Comparison of DFAA incorporation and bacterial production I5 May 17 May 28 May 40m Om Om 40m In situ (nm) nd 5 89 l0 < 0.8 pm filtrates Leucine (nm) Glutamate (nm) 126 nd nd 111 AAmix (nm) nd nd Intracellular pool (mm) Radioactivity in the 1 m1 fractions could not be attributed always to a single amino acid, since 2 or 3 amino acids were present in 4 of the 22 l-m1 fractions collected (Table 2). The radiolabels of the 6 amino acids dominating the <0.8 pm filtrate (see above) exhibited 1- to 4.4-fold ID. Their concentrations in < 0.8 pm filtrates were between 30 and 65 nm. Other 3H-amino acids added at 0.5 nm were diluted much higher by non-labelled amino acids (Table 2). Their concentrations in the filtrate always were <4.5 nm except for glutamate which had concentrations between 5.5 and 13 nm. Highest ID occurred with glutamate and valine/ phenylalanine. Mean ID for all amino acids added as a mixture on 28 May was 9.7 at 0 m and 15.5 at 40 m. Interestingly, when 3H-glutamate was added at 10 nm it was diluted by non-labelled glutamate in the intracellular pool nearly as much as when added at 0.5 nm. In contrast, 3H-Leu was diluted 12-fold when added at 0.5 nm but only 2.2-fold when added at 10 nm (Table 2). Isotope dilution of Leu added at 10 nm Incorporation of DFAA into the ice-cold TCA precipitate was compared with total bacterial biomass production in order to estimate the contribution of the DFAA pool to bacterial protein synthesis. On 17 May at 0 m, incorporation of DFAA accounted for 8 '10 of bacterial protein production, 5 O/O of C, and 7.5 % of N production (Table 3). On 28 May at 0 m, respective percentages were34 %, 20 %, and31 %,and at40m42 %, 24 %,and 37 %. This indicates that on these dates and depths the majority of amino acids for bacterial biosynthesic requirements did not come from incorporation of DFAA. On 28 May at 40 m, the incorporation rate of glutamate was also measured to determine how much of the glutamate requirement for protein synthesis was met by incorporation of dissolved free glutamate. Incorporation of glutamate was 4.8 ng (32 pmol) 1-' h-' and accounted for 5 % of total protein production (Table 3). To compare glutamate incorporation with the glutamate requirement, protein production was converted into moles of amino acids by dividing grams protein by 120, which is the average formula weight of amino acids. Thls results In 770 pm01 protein amino acids I-' h-'. Assuming a mol% of 11.5 for glutamate in protein of marine bacteria (Simon & Azam 1989), the glutamate requirement would thus be 89 pm01 1-' h-'. Hence, uptake of free glutamate accounted for 36 '10 of the glutamate requirement for protein synthesis. The requirement for the intracellular glutamate pool is <5 % of that for protein synthesis. Table 2. Isotope dilution of %I-amino acids (specific activity in the bulk waterspecific activity in the bacterial intracellular pool). Amino acids were added as a mixture (AAmix) at 0.5 nm final concentration each or individually as glutamate or leucine at 0.5 or 10 nlm final concentration to 0.8 pm filtrates. Pooled amino acids indicate that their radioactive peaks were not resolved within 1 m1 fractions of the HPLC effluent. nd: not determined Amino acids 15 May 17 May 28 May 40 m 0 m 0 m AAmix Aspartate nd n d Glutamate nd nd 13.3 Serine nd nd 1.O 1.0 Glycine/threonine/histidine nd nd Argin~ne/alanine/tyrosine nd nd Valine/phenylalanine nd nd IsoleucineAeucine nd nd Glutamate 0.5 nm 95.0 nd nd nm 77.6 nd nd n d Leucine 0.5 nm 11.8 nd nd nd 10 nm
5 Simon: C and N sources of marine bacteria 299 Table 3. Bacterial protein, carbon, and nitrogen production and incorporation rates of a mixture of amlno acids into the ice-cold TCA precipitate added at trace amounts (05 nm) C production is 86 '10 and N product~on is 21.5 O/O of protein production. Rates are ng 1-' h-' 17 May 28 May 0 m 0 m 40 m Bacterial production Protein Carbon Nitrogen Incorporation of amino acids Amino acids Amino acid C Amino acid N DISCUSSION The purpose of this study was to examine the role of the DFAA pool versus other sources in supplying amino acids for protein synthesis by the bacterial assemblage in the subarctlc Pacific. The approach was to study the isotope dilution (ID) of 3~-amino acids in the intracellular bacterial pool. It was found that ID varied greatly depending on the amino acid and its extracellular concentration. Highest IDs occurred with glutamate, valine/phenylalanine and isoleucine/leucine (Table 2), indicating that sources other than uptake of the respective DFAA contributed 87 to 98 % to intracellular pool requirements. Concentrations of valine and phenylalanine in the < 0.8 +m filtrates were always < 2.5 nm, those of isoleucine and leucine < 4.5 and < 2 nm, respectively. In contrast to these amino acids, IDs of glycinekhreonine and arginine/alanine/tyrosine were much lower, < 4.4-fold. Serine did not show any ID. Concentrations of serine and glycine/threonine in the <0.8,pm filtrates were > 30 nm, those of arginine and alanine/tyrosine > 14 nm. Thus, ID appeared to vary inversely with the DFAA concentration. ID seemed to be reduced substantially if the concentration was sufficient to maximize DFAA uptake. This conclusion is supported by the experiment in which Leu was added at trace amounts (1.3 nm final concentration) and 10nM. When added at trace amounts ID of Leu was 5-fold higher as compared to the 10 nm addition (Table 2). While preparing the <0.8 pm filtrates DFAA concentrations increased substantially. This probably decreased ID of amino acids since an inverse relationship between ID and extracellular concentration of amino acids was observed. The elevated DFAA concentrations in the < 0.8 pm filtrates thus would lead to ovei-estimating the importance of the DFAA pool for supplying amino acids to the intracellular pool, yielding an upper limit for DFAA as a C and N source. However, the impact of the enhanced amino acid concentration appears to be small. The mean intracellular ID of 4, excluding glutamic acid and valine/phenylalanine, would predict that DFAA were accounting for about 25 % of bacterial protein production. Based on a direct comparison of DFAA incorporation with bacterial production measurements in unfiltered water, DFAA supplied < 8 O/O (17 May) to 42 % (28 May) of bacterial biomass production (Table 3), which is in the range the ID measurements would predict. These percentages are also in the range reported by other studies comparing amino acid uptake and bacterial production (Billen & Fontigny 1987, J~rgensen 1987, 1990, Fuhrman 1990). Isotope dilution of glutamate was always high irrespective of the added glutamate concentration. Glutamate is a key metabolite for ammonium uptake and transamination (Brown 1980). Hence the high ID of glutamate is probably because ammonium uptake by bacteria was high, which is consistent with direct measurements with 15N-ammoniun~ in the subarctic Pacific (Kirchman et al. 1989). The high ID of glutamate also is an indirect indication that other amino acids were synthesized de novo from ammonlum and organic carbon. If only glutamate uptake rate, which explains 36 O/O of the glutamate requirements for protein synthesis, had been measured, the key role of glutamate for ammonium uptake would not have been elucidated. High ID of amino acids also could have occurred if isotopic equilibrium in the intracellular pool had not been reached (Ing & Klug 1982). This would have been the case if the incubation time was substantially shorter than the amino acid pool turnover time. To calculate the pool turnover time the same cell size (0.07 L1m3) and water content (64 '10) as for calculating the pool concentration were assumed (see'results'), giving a pool size of 45 X 10-l8 1 per bacterium. Since this assumes that all cellular water is in the amino acid pool, it results in the largest pool size possible and thus the longest turnover time. Taking 40 mm as the amino acid pool concentration (Table 1) yields a pool size of 2 X 10-l8 m01 total amino acids. The pool turnover time is then the pool size divided by the amount of amino acids taken up for protein synthesis assuming steady state conditions. Converting the protein production rates from g protein into m01 amino acids as above (see last paragraph of 'Results') and per bacterium this corresponds to 6 X 10-l9 moles of total amino acids produced per cell on 28 May. This yields a pool turnover time of 3 h, which is close to the incubation times. Turnover times of individual amino acids were somewhat longer or shorter according to their mol% distribution in the intracellular amino acid pool and bacterial protein.
6 300 Mar Ecol. Prog. S Isotope dilution factors, therefore, might have been affected by the amino acid pool turnover times. The impact, however, was presumably small because ID factors of 1, which were determined for several amino acids, can only occur if isotopic equibrium has been reached. The amino acid pool turnover time in Simon & Azam (1989; Table 4) calculated with the same assumptions as above is 1 h. Simon & Azam showed that isotopic equilibrium of 3H-leucine in the intracellular pool of marine bacteria was reached after 30 min, i.e. is half of the pool turnover time. This is a further indication that the ID factors were affected only slightly by pool turnover times close to incubation times. Isotope dilution factors close to 1 also indicate that a large fraction of the bacterial assemblage was actively incorporating amino acids (Simon 1990). The 2.6-fold ID of Leu added at 10 nm concentrations (Table 2) has implications for estimating protein and biomass production by bacteria with the leucine method (Kirchman et al. 1985, Simon & Azam 1989). The reliability of this method is based on a low ID of Leu (see 'Matenal and Methods'). Simon & Azam (1989) found a 2-fold ID of Leu in the Southern California Bight of the Pacific. The 2.6-fold ID I found in the subarctic Pacific is 1.3-fold higher. Considering the standard deviation and experimental errors involved in this method, this ID is a valuable confirmation of the ID found by Simon & Azam in a rather different environment. This suggests that 2- to 3-fold ID might be valid in marine oligo- to mesotrophic waters if Leu incorporation is maximized and if the incubation time is 3 the turnover time of the intracellular amino acid pool (see above). However, for measuring bacterial protein production in unknown aquatic ecosystems I still suggest determining ID for Leu or using the most conservative assumption (ID=l) and ensuring that leucine incorporation is maximized. Intracellular amino acid concentrations were in the mm range and were comparable to cultured bacteria (Tempest et al. 1970) and to other marine bacterial assemblages (Simon & Azam 1989). They were also comparable to marine algae (Admiraal et al. 1986, Haberstroh & Ahmed 1986). I did not determine the bacterial cell size for estimating the cell volume and water content but assumed a mean cell volume often found in marine bacterial assemblages (Lee & Fuhrman 1987) and calculated the water content according to Simon & Azam (1989). Glutamic acid dominated the intracellular pool by a proportion (41 mol%) also found in other marine bacterial assemblages (Simon & Azam 1989) and dilution cultures of bacterial assemblages growing on ammonium as the major N source (Simon unpubl.). In cultured bacteria grown under glucose or ammonium limitation, the intracellular amino acid pool is even more dominated by glutamate, between 53 and 89 mol% (Tempest et al. 1970). When a culture of Aerobacter aerogenes was grown under phosphorus limitation the glutamate mol%, however, was reduced to 19 O/O and alanine dominated the intracellular amino acid pool, making up 50 % (Tempest et al. 1970). This also holds true for natural aquatic bacterial assemblages. When amino acids are their major nitrogen sources the mol% of glutamate in the intracellular amino acid pool is reduced to < 20 % (Simon unpubl.). The estimates of the contribution of DFAA to bacterial biosynthetic requirements explaining < 37 % of the C and N production suggest that other C and N sources are utilized as well. Potenbally important other sources of amino acids are DCAA and de novo synthesis from other C and N sources such as nucleic acids, alkylamines, mono- and polysaccharides, and ammonium. Although it has been shown that nucleic acids are utilized by aquatic bacterial assemblages (Paul et al. 1987) there IS no information about their relevance in the subarctic Pacific. Addition of alkylamines has little effect on the growth of marine bacterial assemblages (HofIe 1984, krchman 1990). Turnover of proteins (Hollibaugh & Azam 1983) and DCAA (Coffin 1989) by bacterial assemblages has been found to be comparable to turnover of DFAA. Also, bacteria have a preferential uptake for oligopeptides as compared to DFAA (Kirchman & Hodson 1986). Thus, DCAA such as oligopeptides may be another source of amino acids for bacterial growth. Addition of proteins, however, did not stimulate bacterial growth in the subarctic Pacific whereas addition of DFAA did (Kirchman 1990). This observation and the greatly varying ID of the various amino acids suggest that in this study DCAA were not a large source of amino acids for the bacterial intracellular pool. The question of sources for DFAA remains open, however The high glutamate concentration in the intracellular pool and the high ID of glutamate independent of the added concentration suggests that amino acid de novo synthesis from ammonium and organic carbon presumably was the most important source of amino acids in the bacterial intracellular pool. These data are consistent with I5N-ammonium uptake rates measured by Kirchman et al. (1989) at the same site. They measured uptake rates of about 30 ng ammonium-n 1-' h-' by the < 0.8 pm size fraction, comparable to the bacterial N production rates estimated from protein synthesis (Table 3). In conclusion, this study has shown that measuring ID of amino acids in the bacterial intracellular pool is a valuable approach to elucidating the contribution of DFAA and other sources of amino acids to biosynthetic requirements of aquatic bacterial assemblages. Since amino acids are major C and N sources (J~rgensen 1987, 1990, Kirchman 1990), it is important to obtain detailed insight into pathways of amino acids entering
7 Simon: C and N soul the bacterial cell and associated biosynthetic processes. The amino acid mol% composition of the intracellular pool together with ID measurements of amino acids also allows examination of the importance of ammonium uptake for bacterial N supply. This is of special interest in environments with high ammonium concentrations such as the subarctic Pacific (Anderson et al. 1977) but might also be a useful approach in other ecosystenls. Acknowledgements. I am most grateful to D. L. Kirchman who made possible this study and gave valuable comments on drafts of this paper. I also thank 3 anonymous reviewers for constructive criticism on the manuscript This work was supported by a grant from the National Science Foundation (OCE ) to D. L. Kirchman and by Deutsche Forschungsgemeinschaft, Special Collaborative Program SFB-248 'Cycling of Matter in Lake Constance' LITERATURE CITED Admiraal, W., Peletier, H., Laane, R. W. P. M. (1986). Nitrogen metabolism of marine diatoms: excretion, assimilation and cellular pools of free amino acids in seven species with different cell size. J. exp mar Biol. Ecol. 98: Anraku, Y. (1980). Transport and utihzation of amino acids by bacteria. In Payne, J. W. (ed.) Microorganisms and nitrogen sources. Wiley, New York, p Anderson, G. C., Lam, R. K., Booth, B C, Glass, J. M. (1977). A description and numerical analysis of the factor affecting the processes of produchon in the Gulf of Alaska. University of Washington Department of Oceanography Special Report 76 (Ref. M-77-40), Seattle, Washington Billen, G., Fontigny, A. (1987). Dynamics of a Phaeocystisdominated spring bloom in Belgian coastal waters. 11. Bacterioplankton dynamics. Mar Ecol. Prog. Ser 30: Brown. C. M. (1980). Ammonia assimilation and utilization in bacteria and fungi. In: Payne, J. W (ed.) Microorganisms and nitrogen sources. Wiley, New York, p Coffin. R. B. (1989). Bacterial uptake of dissolved free and combined amino acids in estuarine waters. Limnol. Oceanogr 34: Fuhrman, J. A. (1990). Dissolved free amino acid cycling in an estuarine outflow plume. Mar. Ecol. Prog. Ser. 66: Haberstroh, P. R., Ahmed, S I (1986). Resolution by high pressure liquid chromatography of intracellular and extracellular free amino acids of a nitrogen deficient marine diatom, Skeletonema costaturn (Grev.) Cleve, pulsed with nitrate or ammonium. J. exp. mar. Biol. Ecol. 101: Hobble, J. E., Daley, R. J., Jasper, S. (1977). Use of nuclepore filters for counting bacteria by epifluorescence microscopy. Appl. environ. Microbiol. 33: Hofle. M. G. (1984). Degradation of putrescine and cadaverine This article was submitted to the editor in seawater cultures by marine bacteria. Appl. environ. Microbiol Hollibaugh, J. T., Azam, F. (1983). Microbial degradation of dissolved proteins in seawater. Limnol. Oceanogr 28: Jsrgensen, N. 0. G. (1987). Free amino acids in lakes. concentratlons and assimilation rates in relation to phytoplankton and bacterial production. Limnol. Oceanogr Jsrgensen, N. 0 G (1990). Assimilation of free monosaccharides and amino acids relative to bacterial production in eutrophic lake water. Arch. Hydrobiol. Beih Ergebn. Llmnol King. G M, Klug, M. J. (1982). Glucose metabolism in sediments of a eutrophic lake: tracer analysis of uptake and product formation. Appl. environ. Microbiol. 44: firchman, D. L. (1990). Limitation of bacterial growth by dissolved organic matter in the subarctic Pacific. Mar. Ecol. Prog. Ser. 62: Kirchman, D. L., Hodson, R. E. (1986). Metabolic regulation of amino acid uptake in marine waters. Limnol. Oceanogr 31: Kirchman, D. L., Keil, R. G., Wheeler, P. A. (1989). The effect of amino acids on ammonium utilization and regeneration by heterotrophic bacteria in the subarctic Pacific. Deep Sea Res. 36: hrchman, D L., K'Nees, E., Hodson, R. E. (1985). Leucine incorporation and its potential as a measure of protein synthesis by bacteria in natural aquatic systems. Appl. environ. Microbiol. 49: Lee, S., Fuhrman, J. A. (1987). Relationships between biovolume and biomass of naturally derived marine bacterioplankton. Appl. environ. Microbiol. 53: Lindroth, P., Mopper, K. (1979). High performance liquid chromatographic determination of subpicomole amounts of amino acids by precolumn fluorescence derivatization with o-phthaldialdehyde. Analyt. Chem. 51: Nagata, T (1986). Carbon and nitrogen content of natural planktonic bacteria. Appl. environ. Microbiol. 52: Paul, J. H., Jeffrey, W. H., DeFlaun, M. F. (1987). Dynamics of extracellular DNA in the marine environment. Appl. environ. Microbiol. 53: Payne, J. W. (1980). Microorganisms and nitrogen sources: transport and utilization of amino acids, peptides, proteins, and related substrates. Wiley, New York Simon, M. (1990). Improved assessment of bacterial production: comblned measurements of protein synthesis via leucine and cell multiplication via thymidine incorporation. Arch. Hydrobiol. Beih. Ergebn. Limnol. 34: Simon, M,, Azam, F. (1989). Protein content and protein synthesis rates of planktonic marine bacteria. Mar. Ecol. Prog. Ser. 51: Tempest, D. W., Meers, J. M,, Brown, C. M. (1970). Influence of environment on the content and composition of microbial free amino acid pools. J. gen. Microbiol. 64: Wheeler, P. A., Kokkinakis, S. A. (1990). Ammonium recycling limits nitrate use in the oceanic subarctic Pacific. Limnol. Oceanogr 35: Manuscript first received: March 15, 1991 Revised version accepted: June 10, 1991
Utilization of Organic Nitrogen Sources by Two Phytoplankton Species and a Bacterial Isolate in Pure and Mixed Cultures
 APPLIED AND ENVIRONMENTAL MICROBIOLOGY, May 1994, p. 1554-156 Vol. 6, No. 5 99-224/94/$4.+ Copyright C) 1994, American Society for Microbiology Utilization of Organic Nitrogen Sources by Two Phytoplankton
APPLIED AND ENVIRONMENTAL MICROBIOLOGY, May 1994, p. 1554-156 Vol. 6, No. 5 99-224/94/$4.+ Copyright C) 1994, American Society for Microbiology Utilization of Organic Nitrogen Sources by Two Phytoplankton
Ammonium uptake by heterotrophic bacteria in the Delaware estuary and adjacent coastal waters
 Limnol. Oceanogr., 40(5), 1995, 886-897 0 1995, by the American Society of Limnology and Oceanography, Inc. Ammonium uptake by heterotrophic bacteria in the Delaware estuary and adjacent coastal waters
Limnol. Oceanogr., 40(5), 1995, 886-897 0 1995, by the American Society of Limnology and Oceanography, Inc. Ammonium uptake by heterotrophic bacteria in the Delaware estuary and adjacent coastal waters
Upper ocean total organic carbon at BATS Remember DOC = ~98% of the TOC.
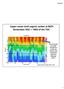 Upper ocean total organic carbon at BATS Remember DOC = ~98% of the TOC. Note the build up in DOC through the spring and summer, with subsequent export the following winter. Figure courtesy of Craig Carlson,
Upper ocean total organic carbon at BATS Remember DOC = ~98% of the TOC. Note the build up in DOC through the spring and summer, with subsequent export the following winter. Figure courtesy of Craig Carlson,
In steady state, new production = carbon export
 In steady state, new production = carbon export Where does primary production go? Export Bacteria Dissolved organic matter Grazing What other components of the biological pump are important? The majority
In steady state, new production = carbon export Where does primary production go? Export Bacteria Dissolved organic matter Grazing What other components of the biological pump are important? The majority
Amino Acids. Amino Acids. Fundamentals. While their name implies that amino acids are compounds that contain an NH. 3 and CO NH 3
 Fundamentals While their name implies that amino acids are compounds that contain an 2 group and a 2 group, these groups are actually present as 3 and 2 respectively. They are classified as α, β, γ, etc..
Fundamentals While their name implies that amino acids are compounds that contain an 2 group and a 2 group, these groups are actually present as 3 and 2 respectively. They are classified as α, β, γ, etc..
1. Describe the relationship of dietary protein and the health of major body systems.
 Food Explorations Lab I: The Building Blocks STUDENT LAB INVESTIGATIONS Name: Lab Overview In this investigation, you will be constructing animal and plant proteins using beads to represent the amino acids.
Food Explorations Lab I: The Building Blocks STUDENT LAB INVESTIGATIONS Name: Lab Overview In this investigation, you will be constructing animal and plant proteins using beads to represent the amino acids.
Release of aminoacids and inorganic nutrients by heterotrophic marine microflagellates
 Vol. 23: 99-106, 1985 MARINE ECOLOGY - PROGRESS SERIES Mar. Ecol. hog. Ser. Published April 25 Release of aminoacids and inorganic nutrients by heterotrophic marine microflagellates Agneta Anderssonl,
Vol. 23: 99-106, 1985 MARINE ECOLOGY - PROGRESS SERIES Mar. Ecol. hog. Ser. Published April 25 Release of aminoacids and inorganic nutrients by heterotrophic marine microflagellates Agneta Anderssonl,
CS612 - Algorithms in Bioinformatics
 Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
Objective: You will be able to explain how the subcomponents of
 Objective: You will be able to explain how the subcomponents of nucleic acids determine the properties of that polymer. Do Now: Read the first two paragraphs from enduring understanding 4.A Essential knowledge:
Objective: You will be able to explain how the subcomponents of nucleic acids determine the properties of that polymer. Do Now: Read the first two paragraphs from enduring understanding 4.A Essential knowledge:
Amino acids-incorporated nanoflowers with an
 Amino acids-incorporated nanoflowers with an intrinsic peroxidase-like activity Zhuo-Fu Wu 1,2,+, Zhi Wang 1,+, Ye Zhang 3, Ya-Li Ma 3, Cheng-Yan He 4, Heng Li 1, Lei Chen 1, Qi-Sheng Huo 3, Lei Wang 1,*
Amino acids-incorporated nanoflowers with an intrinsic peroxidase-like activity Zhuo-Fu Wu 1,2,+, Zhi Wang 1,+, Ye Zhang 3, Ya-Li Ma 3, Cheng-Yan He 4, Heng Li 1, Lei Chen 1, Qi-Sheng Huo 3, Lei Wang 1,*
LAB#23: Biochemical Evidence of Evolution Name: Period Date :
 LAB#23: Biochemical Evidence of Name: Period Date : Laboratory Experience #23 Bridge Worth 80 Lab Minutes If two organisms have similar portions of DNA (genes), these organisms will probably make similar
LAB#23: Biochemical Evidence of Name: Period Date : Laboratory Experience #23 Bridge Worth 80 Lab Minutes If two organisms have similar portions of DNA (genes), these organisms will probably make similar
Biological systems interact, and these systems and their interactions possess complex properties. STOP at enduring understanding 4A
 Biological systems interact, and these systems and their interactions possess complex properties. STOP at enduring understanding 4A Homework Watch the Bozeman video called, Biological Molecules Objective:
Biological systems interact, and these systems and their interactions possess complex properties. STOP at enduring understanding 4A Homework Watch the Bozeman video called, Biological Molecules Objective:
Midterm 1 Last, First
 Midterm 1 BIS 105 Prof. T. Murphy April 23, 2014 There should be 6 pages in this exam. Exam instructions (1) Please write your name on the top of every page of the exam (2) Show all work for full credit
Midterm 1 BIS 105 Prof. T. Murphy April 23, 2014 There should be 6 pages in this exam. Exam instructions (1) Please write your name on the top of every page of the exam (2) Show all work for full credit
Analysis of L- and D-Amino Acids Using UPLC Yuta Mutaguchi 1 and Toshihisa Ohshima 2*
 Analysis of L- and D-Amino Acids Using UPLC Yuta Mutaguchi 1 and Toshihisa Ohshima 2* 1 Department of Biotechnology, Akita Prefectural University, Akita City, Japan; 2 Department of Biomedical Engineering,
Analysis of L- and D-Amino Acids Using UPLC Yuta Mutaguchi 1 and Toshihisa Ohshima 2* 1 Department of Biotechnology, Akita Prefectural University, Akita City, Japan; 2 Department of Biomedical Engineering,
Growth responses of Ulva prolifera to inorganic and organic. nutrients: Implications for macroalgal blooms in the
 1 2 3 Growth responses of Ulva prolifera to inorganic and organic nutrients: Implications for macroalgal blooms in the southern Yellow Sea, China 4 5 Hongmei Li a, Yongyu Zhang a *, Xiurong Han b, Xiaoyong
1 2 3 Growth responses of Ulva prolifera to inorganic and organic nutrients: Implications for macroalgal blooms in the southern Yellow Sea, China 4 5 Hongmei Li a, Yongyu Zhang a *, Xiurong Han b, Xiaoyong
Utilization of dissolved protein and amino acids in the northern Sargasso Sea
 Vol. 18: 293-300,1999 AQUATC MCROBAL ECOLOGY Aquat Microb Ecol Published August 20 Utilization of dissolved protein and amino acids in the northern Sargasso Sea Richard G. Keil*, David L. Kirchman University
Vol. 18: 293-300,1999 AQUATC MCROBAL ECOLOGY Aquat Microb Ecol Published August 20 Utilization of dissolved protein and amino acids in the northern Sargasso Sea Richard G. Keil*, David L. Kirchman University
Biomolecules: amino acids
 Biomolecules: amino acids Amino acids Amino acids are the building blocks of proteins They are also part of hormones, neurotransmitters and metabolic intermediates There are 20 different amino acids in
Biomolecules: amino acids Amino acids Amino acids are the building blocks of proteins They are also part of hormones, neurotransmitters and metabolic intermediates There are 20 different amino acids in
Reactions and amino acids structure & properties
 Lecture 2: Reactions and amino acids structure & properties Dr. Sameh Sarray Hlaoui Common Functional Groups Common Biochemical Reactions AH + B A + BH Oxidation-Reduction A-H + B-OH + energy ª A-B + H
Lecture 2: Reactions and amino acids structure & properties Dr. Sameh Sarray Hlaoui Common Functional Groups Common Biochemical Reactions AH + B A + BH Oxidation-Reduction A-H + B-OH + energy ª A-B + H
9/16/15. Properties of Water. Benefits of Water. More properties of water
 Properties of Water Solid/Liquid Density Water is densest at 4⁰C Ice floats Allows life under the ice Hydrogen bond Ice Hydrogen bonds are stable Liquid water Hydrogen bonds break and re-form Benefits
Properties of Water Solid/Liquid Density Water is densest at 4⁰C Ice floats Allows life under the ice Hydrogen bond Ice Hydrogen bonds are stable Liquid water Hydrogen bonds break and re-form Benefits
Short polymer. Dehydration removes a water molecule, forming a new bond. Longer polymer (a) Dehydration reaction in the synthesis of a polymer
 HO 1 2 3 H HO H Short polymer Dehydration removes a water molecule, forming a new bond Unlinked monomer H 2 O HO 1 2 3 4 H Longer polymer (a) Dehydration reaction in the synthesis of a polymer HO 1 2 3
HO 1 2 3 H HO H Short polymer Dehydration removes a water molecule, forming a new bond Unlinked monomer H 2 O HO 1 2 3 4 H Longer polymer (a) Dehydration reaction in the synthesis of a polymer HO 1 2 3
Utilization of inorganic and organic nitrogen by bacteria in marine systems1
 Limnol. Oceanogr., 31(5), 1986, 998-1009 0 1986, by the American Society of Limnology and Oceanography, Inc. Utilization of inorganic and organic nitrogen by bacteria in marine systems1 Patricia A. Wheeler2
Limnol. Oceanogr., 31(5), 1986, 998-1009 0 1986, by the American Society of Limnology and Oceanography, Inc. Utilization of inorganic and organic nitrogen by bacteria in marine systems1 Patricia A. Wheeler2
Seasonal and vertical difference in negative and positive effects of grazers on heterotrophic bacteria in Lake Biwa
 Limnol. Oceanogr., 45(8), 2000, 1689 1696 2000, by the American Society of Limnology and Oceanography, Inc. Seasonal and vertical difference in negative and positive effects of grazers on heterotrophic
Limnol. Oceanogr., 45(8), 2000, 1689 1696 2000, by the American Society of Limnology and Oceanography, Inc. Seasonal and vertical difference in negative and positive effects of grazers on heterotrophic
2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry
 Dr. Sanjeeva Srivastava 1. Fundamental of Mass Spectrometry Role of MS and basic concepts 2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry 2 1 MS basic concepts Mass spectrometry - technique
Dr. Sanjeeva Srivastava 1. Fundamental of Mass Spectrometry Role of MS and basic concepts 2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry 2 1 MS basic concepts Mass spectrometry - technique
Biological considerations in the measurement of dissolved free amino acids in seawater and implications for chemical and microbiological studies*
 MARINE ECOLOGY - PROGRESS SERIES Mar. Ecol. Prog. Ser. Published August 15 Biological considerations in the measurement of dissolved free amino acids in seawater and implications for chemical and microbiological
MARINE ECOLOGY - PROGRESS SERIES Mar. Ecol. Prog. Ser. Published August 15 Biological considerations in the measurement of dissolved free amino acids in seawater and implications for chemical and microbiological
Proteins are a major component of dissolved organic nitrogen (DON) leached from terrestrially aged Eucalyptus camaldulensis leaves
 Environ. Chem. 216, 13, 877 887 doi:1.171/en165_ac CSIRO 216 Supplementary material Proteins are a major component of dissolved organic nitrogen (DON) leached from terrestrially aged Eucalyptus camaldulensis
Environ. Chem. 216, 13, 877 887 doi:1.171/en165_ac CSIRO 216 Supplementary material Proteins are a major component of dissolved organic nitrogen (DON) leached from terrestrially aged Eucalyptus camaldulensis
Classification of amino acids: -
 Page 1 of 8 P roteinogenic amino acids, also known as standard, normal or primary amino acids are 20 amino acids that are incorporated in proteins and that are coded in the standard genetic code (subunit
Page 1 of 8 P roteinogenic amino acids, also known as standard, normal or primary amino acids are 20 amino acids that are incorporated in proteins and that are coded in the standard genetic code (subunit
9/6/2011. Amino Acids. C α. Nonpolar, aliphatic R groups
 Amino Acids Side chains (R groups) vary in: size shape charge hydrogen-bonding capacity hydrophobic character chemical reactivity C α Nonpolar, aliphatic R groups Glycine (Gly, G) Alanine (Ala, A) Valine
Amino Acids Side chains (R groups) vary in: size shape charge hydrogen-bonding capacity hydrophobic character chemical reactivity C α Nonpolar, aliphatic R groups Glycine (Gly, G) Alanine (Ala, A) Valine
Four Classes of Biological Macromolecules. Biological Macromolecules. Lipids
 Biological Macromolecules Much larger than other par4cles found in cells Made up of smaller subunits Found in all cells Great diversity of func4ons Four Classes of Biological Macromolecules Lipids Polysaccharides
Biological Macromolecules Much larger than other par4cles found in cells Made up of smaller subunits Found in all cells Great diversity of func4ons Four Classes of Biological Macromolecules Lipids Polysaccharides
Zackary Johnson Department of Oceanography
 Zackary Johnson Department of Oceanography http://www.soest.hawaii.edu/oceanography/zij/education/ocn621 Application of Bioenergetics to Biological Oceanography Biochemical parameters indicative of stock
Zackary Johnson Department of Oceanography http://www.soest.hawaii.edu/oceanography/zij/education/ocn621 Application of Bioenergetics to Biological Oceanography Biochemical parameters indicative of stock
Lipids: diverse group of hydrophobic molecules
 Lipids: diverse group of hydrophobic molecules Lipids only macromolecules that do not form polymers li3le or no affinity for water hydrophobic consist mostly of hydrocarbons nonpolar covalent bonds fats
Lipids: diverse group of hydrophobic molecules Lipids only macromolecules that do not form polymers li3le or no affinity for water hydrophobic consist mostly of hydrocarbons nonpolar covalent bonds fats
Amino acid metabolism
 Amino acid metabolism The important reaction commonly employed in the breakdown of an amino acid is always the removal of its -amino group. The product ammonia is excreted after conversion to urea or other
Amino acid metabolism The important reaction commonly employed in the breakdown of an amino acid is always the removal of its -amino group. The product ammonia is excreted after conversion to urea or other
Towards a New Paradigm in Scientific Notation Patterns of Periodicity among Proteinogenic Amino Acids [Abridged Version]
![Towards a New Paradigm in Scientific Notation Patterns of Periodicity among Proteinogenic Amino Acids [Abridged Version] Towards a New Paradigm in Scientific Notation Patterns of Periodicity among Proteinogenic Amino Acids [Abridged Version]](/thumbs/84/90818314.jpg) Earth/matriX: SCIENCE TODAY Towards a New Paradigm in Scientific Notation Patterns of Periodicity among Proteinogenic Amino Acids [Abridged Version] By Charles William Johnson Earth/matriX Editions P.O.
Earth/matriX: SCIENCE TODAY Towards a New Paradigm in Scientific Notation Patterns of Periodicity among Proteinogenic Amino Acids [Abridged Version] By Charles William Johnson Earth/matriX Editions P.O.
Biomass Production and Assimilation of Dissolved Organic Matter by SAR11 Bacteria in the Northwest Atlantic Ocean
 APPLIED AND ENVIRONMENTAL MICROBIOLOGY, June 2005, p. 2979 2986 Vol. 71, No. 6 0099-2240/05/$08.00 0 doi:10.1128/aem.71.6.2979 2986.2005 Copyright 2005, American Society for Microbiology. All Rights Reserved.
APPLIED AND ENVIRONMENTAL MICROBIOLOGY, June 2005, p. 2979 2986 Vol. 71, No. 6 0099-2240/05/$08.00 0 doi:10.1128/aem.71.6.2979 2986.2005 Copyright 2005, American Society for Microbiology. All Rights Reserved.
Page 8/6: The cell. Where to start: Proteins (control a cell) (start/end products)
 Page 8/6: The cell Where to start: Proteins (control a cell) (start/end products) Page 11/10: Structural hierarchy Proteins Phenotype of organism 3 Dimensional structure Function by interaction THE PROTEIN
Page 8/6: The cell Where to start: Proteins (control a cell) (start/end products) Page 11/10: Structural hierarchy Proteins Phenotype of organism 3 Dimensional structure Function by interaction THE PROTEIN
Biology. Lectures winter term st year of Pharmacy study
 Biology Lectures winter term 2008 1 st year of Pharmacy study 3 rd Lecture Chemical composition of living matter chemical basis of life. Atoms, molecules, organic compounds carbohydrates, lipids, proteins,
Biology Lectures winter term 2008 1 st year of Pharmacy study 3 rd Lecture Chemical composition of living matter chemical basis of life. Atoms, molecules, organic compounds carbohydrates, lipids, proteins,
Properties of amino acids in proteins
 Properties of amino acids in proteins one of the primary roles of DNA (but far from the only one!!!) is to code for proteins A typical bacterium builds thousands types of proteins, all from ~20 amino acids
Properties of amino acids in proteins one of the primary roles of DNA (but far from the only one!!!) is to code for proteins A typical bacterium builds thousands types of proteins, all from ~20 amino acids
James H. Rich College of Marine Studies, University of Delaware, Lewes 19958
 Limnol. Oceanogr., 41(4), 1996, 9-64 1996, by the American Society of Limnology and Oceanography, Inc. Concentrations and uptake of neutral monosaccharides along 14OOW in the equatorial Pacific: Contribution
Limnol. Oceanogr., 41(4), 1996, 9-64 1996, by the American Society of Limnology and Oceanography, Inc. Concentrations and uptake of neutral monosaccharides along 14OOW in the equatorial Pacific: Contribution
1. (38 pts.) 2. (25 pts.) 3. (15 pts.) 4. (12 pts.) 5. (10 pts.) Bonus (12 pts.) TOTAL (100 points)
 Moorpark College Chemistry 11 Spring 2010 Instructor: Professor Torres Examination #5: Section Five May 4, 2010 ame: (print) ame: (sign) Directions: Make sure your examination contains TWELVE total pages
Moorpark College Chemistry 11 Spring 2010 Instructor: Professor Torres Examination #5: Section Five May 4, 2010 ame: (print) ame: (sign) Directions: Make sure your examination contains TWELVE total pages
Authors. Abstract. Introduction. Food
 Improved and Simplified Liquid Chromatography/Atmospheric Pressure Chemical Ionization Mass Spectrometry Method for the Analysis of Underivatized Free Ao Acids in Various Foods Application Food Authors
Improved and Simplified Liquid Chromatography/Atmospheric Pressure Chemical Ionization Mass Spectrometry Method for the Analysis of Underivatized Free Ao Acids in Various Foods Application Food Authors
Dynamics of the decline of a phytoplankton bloom after an upwelling event
 Vol. 16: 121-126, 1984 MARINE ECOLOGY - PROGRESS SERIES Mar. Ecol. Prog. Ser. Published February 29 Dynamics of the decline of a phytoplankton bloom after an upwelling event R. G. Barlow Sea Fisheries
Vol. 16: 121-126, 1984 MARINE ECOLOGY - PROGRESS SERIES Mar. Ecol. Prog. Ser. Published February 29 Dynamics of the decline of a phytoplankton bloom after an upwelling event R. G. Barlow Sea Fisheries
Carbon versus iron limitation of bacterial growth in the California upwelling regime
 December 2000 Volume 45 Number 8 Limnol. Oceanogr., 45(8), 2000, 1681 1688 2000, by the American Society of Limnology and Oceanography, Inc. Carbon versus iron limitation of bacterial growth in the California
December 2000 Volume 45 Number 8 Limnol. Oceanogr., 45(8), 2000, 1681 1688 2000, by the American Society of Limnology and Oceanography, Inc. Carbon versus iron limitation of bacterial growth in the California
Proteins are sometimes only produced in one cell type or cell compartment (brain has 15,000 expressed proteins, gut has 2,000).
 Lecture 2: Principles of Protein Structure: Amino Acids Why study proteins? Proteins underpin every aspect of biological activity and therefore are targets for drug design and medicinal therapy, and in
Lecture 2: Principles of Protein Structure: Amino Acids Why study proteins? Proteins underpin every aspect of biological activity and therefore are targets for drug design and medicinal therapy, and in
DISSOLVED ORGANIC MATTER (DOM) AS A DIRECT ORGANIC SUBSIDY TO AQUATIC CONSUMERS
 DISSOLVED ORGANIC MATTER (DOM) AS A DIRECT ORGANIC SUBSIDY TO AQUATIC CONSUMERS STEPHEN B. BAINES ECOLOGY AND EVOLUTION STONY BROOK UNIVERSITY Acknowledgements: Hudson Roditi, Nick Fisher, Jon Cole, Cindy
DISSOLVED ORGANIC MATTER (DOM) AS A DIRECT ORGANIC SUBSIDY TO AQUATIC CONSUMERS STEPHEN B. BAINES ECOLOGY AND EVOLUTION STONY BROOK UNIVERSITY Acknowledgements: Hudson Roditi, Nick Fisher, Jon Cole, Cindy
Dissolved organic matter dynamics in the sea
 Dissolved organic matter dynamics in the sea DOC distributions in the Atlantic Marine Microplankton Ecology OCN 626 Matthew Church Figures from Dennis Hansell DOC distributions in the Pacific Basin scale
Dissolved organic matter dynamics in the sea DOC distributions in the Atlantic Marine Microplankton Ecology OCN 626 Matthew Church Figures from Dennis Hansell DOC distributions in the Pacific Basin scale
Chemistry 121 Winter 17
 Chemistry 121 Winter 17 Introduction to Organic Chemistry and Biochemistry Instructor Dr. Upali Siriwardane (Ph.D. Ohio State) E-mail: upali@latech.edu Office: 311 Carson Taylor Hall ; Phone: 318-257-4941;
Chemistry 121 Winter 17 Introduction to Organic Chemistry and Biochemistry Instructor Dr. Upali Siriwardane (Ph.D. Ohio State) E-mail: upali@latech.edu Office: 311 Carson Taylor Hall ; Phone: 318-257-4941;
Study of Amino Acids in DDGS
 Study of Amino Acids in DDGS Y. Zhang, J. V. Simpson and B. A. Wrenn National Corn-to-Ethanol Research Center Edwardsville, IL 62025 Hans Stein University of Illinois Urbana Champaign Gerald C. Shurson
Study of Amino Acids in DDGS Y. Zhang, J. V. Simpson and B. A. Wrenn National Corn-to-Ethanol Research Center Edwardsville, IL 62025 Hans Stein University of Illinois Urbana Champaign Gerald C. Shurson
The Structure and Function of Macromolecules
 The Structure and Function of Macromolecules Macromolecules are polymers Polymer long molecule consisting of many similar building blocks. Monomer the small building block molecules. Carbohydrates, proteins
The Structure and Function of Macromolecules Macromolecules are polymers Polymer long molecule consisting of many similar building blocks. Monomer the small building block molecules. Carbohydrates, proteins
Metabolism of amino acids. Vladimíra Kvasnicová
 Metabolism of amino acids Vladimíra Kvasnicová Classification of proteinogenic AAs -metabolic point of view 1) biosynthesis in a human body nonessential (are synthesized) essential (must be present in
Metabolism of amino acids Vladimíra Kvasnicová Classification of proteinogenic AAs -metabolic point of view 1) biosynthesis in a human body nonessential (are synthesized) essential (must be present in
Cells. Variation and Function of Cells
 Cells Variation and Function of Cells Plasma Membrane= the skin of a cell, it protects and nourishes the cell while communicating with other cells at the same time. Lipid means fat and they are hydrophobic
Cells Variation and Function of Cells Plasma Membrane= the skin of a cell, it protects and nourishes the cell while communicating with other cells at the same time. Lipid means fat and they are hydrophobic
For questions 1-4, match the carbohydrate with its size/functional group name:
 Chemistry 11 Fall 2013 Examination #5 PRACTICE 1 For the first portion of this exam, select the best answer choice for the questions below and mark the answers on your scantron. Then answer the free response
Chemistry 11 Fall 2013 Examination #5 PRACTICE 1 For the first portion of this exam, select the best answer choice for the questions below and mark the answers on your scantron. Then answer the free response
Chemical Nature of the Amino Acids. Table of a-amino Acids Found in Proteins
 Chemical Nature of the Amino Acids All peptides and polypeptides are polymers of alpha-amino acids. There are 20 a- amino acids that are relevant to the make-up of mammalian proteins (see below). Several
Chemical Nature of the Amino Acids All peptides and polypeptides are polymers of alpha-amino acids. There are 20 a- amino acids that are relevant to the make-up of mammalian proteins (see below). Several
Close coupling between release and uptake of dissolved free amino acids in seawater studied by an isotope dilution approach*
 Vol. 37: 45-52, 1987 MARINE ECOLOGY - PROGRESS SERIES Mar. Ecol. Prog. Ser. Published April 6 Close coupling between release and uptake of dissolved free amino acids in seawater studied by an isotope dilution
Vol. 37: 45-52, 1987 MARINE ECOLOGY - PROGRESS SERIES Mar. Ecol. Prog. Ser. Published April 6 Close coupling between release and uptake of dissolved free amino acids in seawater studied by an isotope dilution
Cells N5 Homework book
 1 Cells N5 Homework book 2 Homework 1 3 4 5 Homework2 Cell Ultrastructure and Membrane 1. Name and give the function of the numbered organelles in the cell below: A E B D C 2. Name 3 structures you might
1 Cells N5 Homework book 2 Homework 1 3 4 5 Homework2 Cell Ultrastructure and Membrane 1. Name and give the function of the numbered organelles in the cell below: A E B D C 2. Name 3 structures you might
Gentilucci, Amino Acids, Peptides, and Proteins. Peptides and proteins are polymers of amino acids linked together by amide bonds CH 3
 Amino Acids Peptides and proteins are polymers of amino acids linked together by amide bonds Aliphatic Side-Chain Amino Acids - - H CH glycine alanine 3 proline valine CH CH 3 - leucine - isoleucine CH
Amino Acids Peptides and proteins are polymers of amino acids linked together by amide bonds Aliphatic Side-Chain Amino Acids - - H CH glycine alanine 3 proline valine CH CH 3 - leucine - isoleucine CH
Introduction to Protein Structure Collection
 Introduction to Protein Structure Collection Teaching Points This collection is designed to introduce students to the concepts of protein structure and biochemistry. Different activities guide students
Introduction to Protein Structure Collection Teaching Points This collection is designed to introduce students to the concepts of protein structure and biochemistry. Different activities guide students
(65 pts.) 27. (10 pts.) 28. (15 pts.) 29. (10 pts.) TOTAL (100 points) Moorpark College Chemistry 11 Spring Instructor: Professor Gopal
 Moorpark College Chemistry 11 Spring 2012 Instructor: Professor Gopal Examination # 5: Section Five May 1, 2012 Name: (print) GOOD LUCK! Directions: Make sure your examination contains TWELVE total pages
Moorpark College Chemistry 11 Spring 2012 Instructor: Professor Gopal Examination # 5: Section Five May 1, 2012 Name: (print) GOOD LUCK! Directions: Make sure your examination contains TWELVE total pages
Solid-phase extraction of dissolved organic matter (SPE-DOM) from river, estuarine and open ocean waters
 Solid-phase extraction of dissolved organic matter (SPE-DOM) from river, estuarine and open ocean waters Gerhard Kattner 1, Thorsten Dittmar 2, Boris Koch 1, and Norbert Hertkorn 3 1 Alfred Wegener Institute
Solid-phase extraction of dissolved organic matter (SPE-DOM) from river, estuarine and open ocean waters Gerhard Kattner 1, Thorsten Dittmar 2, Boris Koch 1, and Norbert Hertkorn 3 1 Alfred Wegener Institute
AMINO ACID METABOLISM. Sri Widia A Jusman Dept. of Biochemistry & Molecular Biology FMUI
 AMINO ACID METABOLISM Sri Widia A Jusman Dept. of Biochemistry & Molecular Biology FMUI Amino acids derived from dietary protein absorbed from intestine through blood taken up by tissues used for biosynthesis
AMINO ACID METABOLISM Sri Widia A Jusman Dept. of Biochemistry & Molecular Biology FMUI Amino acids derived from dietary protein absorbed from intestine through blood taken up by tissues used for biosynthesis
1-To know what is protein 2-To identify Types of protein 3- To Know amino acids 4- To be differentiate between essential and nonessential amino acids
 Amino acids 1-To know what is protein 2-To identify Types of protein 3- To Know amino acids 4- To be differentiate between essential and nonessential amino acids 5-To understand amino acids synthesis Amino
Amino acids 1-To know what is protein 2-To identify Types of protein 3- To Know amino acids 4- To be differentiate between essential and nonessential amino acids 5-To understand amino acids synthesis Amino
Copyright 2008 Pearson Education, Inc., publishing as Pearson Benjamin Cummings
 Concept 5.4: Proteins have many structures, resulting in a wide range of functions Proteins account for more than 50% of the dry mass of most cells Protein functions include structural support, storage,
Concept 5.4: Proteins have many structures, resulting in a wide range of functions Proteins account for more than 50% of the dry mass of most cells Protein functions include structural support, storage,
If you like us, please share us on social media. The latest UCD Hyperlibrary newsletter is now complete, check it out.
 Sign In Forgot Password Register username username password password Sign In If you like us, please share us on social media. The latest UCD Hyperlibrary newsletter is now complete, check it out. ChemWiki
Sign In Forgot Password Register username username password password Sign In If you like us, please share us on social media. The latest UCD Hyperlibrary newsletter is now complete, check it out. ChemWiki
Methionine (Met or M)
 Fig. 5-17 Nonpolar Fig. 5-17a Nonpolar Glycine (Gly or G) Alanine (Ala or A) Valine (Val or V) Leucine (Leu or L) Isoleucine (Ile or I) Methionine (Met or M) Phenylalanine (Phe or F) Polar Trypotphan (Trp
Fig. 5-17 Nonpolar Fig. 5-17a Nonpolar Glycine (Gly or G) Alanine (Ala or A) Valine (Val or V) Leucine (Leu or L) Isoleucine (Ile or I) Methionine (Met or M) Phenylalanine (Phe or F) Polar Trypotphan (Trp
Level and activity of D-amino acids in mouse brain tissue and blood
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 SUPPLEMENTARY INFORMATION Level and activity of D-amino acids in mouse brain tissue and blood Choyce A. Weatherly 1, Siqi Du 1, Curran Parpia 1, Polan T. Santos 2, Adam
1 2 3 4 5 6 7 8 9 10 11 12 13 14 SUPPLEMENTARY INFORMATION Level and activity of D-amino acids in mouse brain tissue and blood Choyce A. Weatherly 1, Siqi Du 1, Curran Parpia 1, Polan T. Santos 2, Adam
Effect of nitrate-n incorporation on the composition of the intracellular amino acid pool of N-deprived Tetraselmis marina
 British Phycological Journal ISSN: 0007-1617 (Print) (Online) Journal homepage: https://www.tandfonline.com/loi/tejp19 Effect of nitrate-n incorporation on the composition of the intracellular amino acid
British Phycological Journal ISSN: 0007-1617 (Print) (Online) Journal homepage: https://www.tandfonline.com/loi/tejp19 Effect of nitrate-n incorporation on the composition of the intracellular amino acid
Mercaptoethanesulfonic acid as the reductive thiol-containing reagent employed for the derivatization of amino acids with o-phthaldialdehyde analysis
 Acta Univ. Sapientiae, Alimentaria, 1 (2008) 49 60 Mercaptoethanesulfonic acid as the reductive thiol-containing reagent employed for the derivatization of amino acids with o-phthaldialdehyde analysis
Acta Univ. Sapientiae, Alimentaria, 1 (2008) 49 60 Mercaptoethanesulfonic acid as the reductive thiol-containing reagent employed for the derivatization of amino acids with o-phthaldialdehyde analysis
EFFICIENT UTILIZATION OF DISSOLVED FREE AMINO ACIDS BY SUSPENDED MARINE BACTERIA
 EFFICIENT UTILIZATION OF DISSOLVED FREE AMINO ACIDS BY SUSPENDED MARINE BACTERIA By: Anthony V. Palumbo, Randolph L. Ferguson, and Parke A. Rublee Palumbo, A.V., R.L. Ferguson, and P.A. Rublee. 1983. Efficient
EFFICIENT UTILIZATION OF DISSOLVED FREE AMINO ACIDS BY SUSPENDED MARINE BACTERIA By: Anthony V. Palumbo, Randolph L. Ferguson, and Parke A. Rublee Palumbo, A.V., R.L. Ferguson, and P.A. Rublee. 1983. Efficient
(30 pts.) 16. (24 pts.) 17. (20 pts.) 18. (16 pts.) 19. (5 pts.) 20. (5 pts.) TOTAL (100 points)
 Moorpark College Chemistry 11 Spring 2009 Instructor: Professor Torres Examination # 5: Section Five April 30, 2009 ame: (print) ame: (sign) Directions: Make sure your examination contains TWELVE total
Moorpark College Chemistry 11 Spring 2009 Instructor: Professor Torres Examination # 5: Section Five April 30, 2009 ame: (print) ame: (sign) Directions: Make sure your examination contains TWELVE total
Nitrogen Metabolism. Overview
 Nitrogen Metabolism Pratt and Cornely Chapter 18 Overview Nitrogen assimilation Amino acid biosynthesis Nonessential aa Essential aa Nucleotide biosynthesis Amino Acid Catabolism Urea Cycle Juicy Steak
Nitrogen Metabolism Pratt and Cornely Chapter 18 Overview Nitrogen assimilation Amino acid biosynthesis Nonessential aa Essential aa Nucleotide biosynthesis Amino Acid Catabolism Urea Cycle Juicy Steak
LC-MS Analysis of Amino Acids on a Novel Mixed-Mode HPLC Column
 Liquid Chromatography Mass Spectrometry SSI-LCMS-022 LC-MS Analysis of Amino Acids on a ovel Mixed-Mode PLC Column LCMS-8040 Background There are four established methods for analyzing amino acids: prelabeled,
Liquid Chromatography Mass Spectrometry SSI-LCMS-022 LC-MS Analysis of Amino Acids on a ovel Mixed-Mode PLC Column LCMS-8040 Background There are four established methods for analyzing amino acids: prelabeled,
For questions 1-4, match the carbohydrate with its size/functional group name:
 Chemistry 11 Fall 2013 Examination #5 PRACTICE 1 ANSWERS For the first portion of this exam, select the best answer choice for the questions below and mark the answers on your scantron. Then answer the
Chemistry 11 Fall 2013 Examination #5 PRACTICE 1 ANSWERS For the first portion of this exam, select the best answer choice for the questions below and mark the answers on your scantron. Then answer the
Proximate composition, amino acid and fatty acid composition of fish maws. Department of Biology, Lingnan Normal University, Zhanjiang, , China
 SUPPLEMENTARY MATERIAL Proximate composition, amino acid and fatty acid composition of fish maws Jing Wen a, Ling Zeng b *, Youhou Xu c, Yulin Sun a, Ziming Chen b and Sigang Fan d a Department of Biology,
SUPPLEMENTARY MATERIAL Proximate composition, amino acid and fatty acid composition of fish maws Jing Wen a, Ling Zeng b *, Youhou Xu c, Yulin Sun a, Ziming Chen b and Sigang Fan d a Department of Biology,
Analysis of free amino acids in tobacco with a new LC/MS/MS. procedure. S. C. Moldoveanu R.J. Reynolds Tobacco Co.
 Analysis of free amino acids in tobacco with a new LC/MS/MS procedure S. C. Moldoveanu R.J. Reynolds Tobacco Co. Jeff Zhu Eurofins Background A considerable number of analytical methods are reported in
Analysis of free amino acids in tobacco with a new LC/MS/MS procedure S. C. Moldoveanu R.J. Reynolds Tobacco Co. Jeff Zhu Eurofins Background A considerable number of analytical methods are reported in
Molecular Biology. general transfer: occurs normally in cells. special transfer: occurs only in the laboratory in specific conditions.
 Chapter 9: Proteins Molecular Biology replication general transfer: occurs normally in cells transcription special transfer: occurs only in the laboratory in specific conditions translation unknown transfer:
Chapter 9: Proteins Molecular Biology replication general transfer: occurs normally in cells transcription special transfer: occurs only in the laboratory in specific conditions translation unknown transfer:
Practice Problems 3. a. What is the name of the bond formed between two amino acids? Are these bonds free to rotate?
 Life Sciences 1a Practice Problems 3 1. Draw the oligopeptide for Ala-Phe-Gly-Thr-Asp. You do not need to indicate the stereochemistry of the sidechains. Denote with arrows the bonds formed between the
Life Sciences 1a Practice Problems 3 1. Draw the oligopeptide for Ala-Phe-Gly-Thr-Asp. You do not need to indicate the stereochemistry of the sidechains. Denote with arrows the bonds formed between the
Biomolecules Amino Acids & Protein Chemistry
 Biochemistry Department Date: 17/9/ 2017 Biomolecules Amino Acids & Protein Chemistry Prof.Dr./ FAYDA Elazazy Professor of Biochemistry and Molecular Biology Intended Learning Outcomes ILOs By the end
Biochemistry Department Date: 17/9/ 2017 Biomolecules Amino Acids & Protein Chemistry Prof.Dr./ FAYDA Elazazy Professor of Biochemistry and Molecular Biology Intended Learning Outcomes ILOs By the end
Feeding Measurements
 Feeding Measurements Measures of Feeding Rates (or specific mortality rate) Clearance Rate Concept Clearance Rate (F) = Volume cleared of prey pred -1 time -1 predator prey A consumer may have different
Feeding Measurements Measures of Feeding Rates (or specific mortality rate) Clearance Rate Concept Clearance Rate (F) = Volume cleared of prey pred -1 time -1 predator prey A consumer may have different
Application Note. Determination of Amino acids by UHPLC with automated OPA- Derivatization by the Autosampler. Summary. Fig. 1.
 Application Note Determination of Amino acids by UHPLC with automated PA- Derivatization by the Autosampler Category Bio Analysis Matrix - Method UHPLC Keywords Proteinogenic Amino acids, Canonical Amino
Application Note Determination of Amino acids by UHPLC with automated PA- Derivatization by the Autosampler Category Bio Analysis Matrix - Method UHPLC Keywords Proteinogenic Amino acids, Canonical Amino
AP Bio. Protiens Chapter 5 1
 Concept.4: Proteins have many structures, resulting in a wide range of functions Proteins account for more than 0% of the dry mass of most cells Protein functions include structural support, storage, transport,
Concept.4: Proteins have many structures, resulting in a wide range of functions Proteins account for more than 0% of the dry mass of most cells Protein functions include structural support, storage, transport,
Amino acid Catabolism
 Enzymatic digestion of dietary proteins in gastrointestinal-tract. Amino acid Catabolism Amino acids: 1. There are 20 different amino acid, they are monomeric constituents of proteins 2. They act as precursors
Enzymatic digestion of dietary proteins in gastrointestinal-tract. Amino acid Catabolism Amino acids: 1. There are 20 different amino acid, they are monomeric constituents of proteins 2. They act as precursors
Moorpark College Chemistry 11 Fall Instructor: Professor Gopal. Examination # 5: Section Five May 7, Name: (print)
 Moorpark College Chemistry 11 Fall 2013 Instructor: Professor Gopal Examination # 5: Section Five May 7, 2013 Name: (print) Directions: Make sure your examination contains TEN total pages (including this
Moorpark College Chemistry 11 Fall 2013 Instructor: Professor Gopal Examination # 5: Section Five May 7, 2013 Name: (print) Directions: Make sure your examination contains TEN total pages (including this
2. Which of the following amino acids is most likely to be found on the outer surface of a properly folded protein?
 Name: WHITE Student Number: Answer the following questions on the computer scoring sheet. 1 mark each 1. Which of the following amino acids would have the highest relative mobility R f in normal thin layer
Name: WHITE Student Number: Answer the following questions on the computer scoring sheet. 1 mark each 1. Which of the following amino acids would have the highest relative mobility R f in normal thin layer
Amino Acids. Review I: Protein Structure. Amino Acids: Structures. Amino Acids (contd.) Rajan Munshi
 Review I: Protein Structure Rajan Munshi BBSI @ Pitt 2005 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2005 Amino Acids Building blocks of proteins 20 amino acids
Review I: Protein Structure Rajan Munshi BBSI @ Pitt 2005 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2005 Amino Acids Building blocks of proteins 20 amino acids
Nitrogen Metabolism. Overview
 Nitrogen Metabolism Pratt and Cornely Chapter 18 Overview Nitrogen assimilation Amino acid biosynthesis Nonessential aa Essential aa Nucleotide biosynthesis Amino Acid Catabolism Urea Cycle Juicy Steak
Nitrogen Metabolism Pratt and Cornely Chapter 18 Overview Nitrogen assimilation Amino acid biosynthesis Nonessential aa Essential aa Nucleotide biosynthesis Amino Acid Catabolism Urea Cycle Juicy Steak
Nitrogen Metabolism. Pratt and Cornely Chapter 18
 Nitrogen Metabolism Pratt and Cornely Chapter 18 Overview Nitrogen assimilation Amino acid biosynthesis Nonessential aa Essential aa Nucleotide biosynthesis Amino Acid Catabolism Urea Cycle Juicy Steak
Nitrogen Metabolism Pratt and Cornely Chapter 18 Overview Nitrogen assimilation Amino acid biosynthesis Nonessential aa Essential aa Nucleotide biosynthesis Amino Acid Catabolism Urea Cycle Juicy Steak
Definitions and Biochemical Pathways History (Literature Review) Use of Lipids to Build Proteins: Lab Mice & Polar Bears
 Definitions and Biochemical Pathways History (Literature Review) Use of Lipids to Build Proteins: Lab Mice & Polar Bears Use of Carbohydrates to Build Proteins: Tilapia Role of Gut Microbiome: Tilapia
Definitions and Biochemical Pathways History (Literature Review) Use of Lipids to Build Proteins: Lab Mice & Polar Bears Use of Carbohydrates to Build Proteins: Tilapia Role of Gut Microbiome: Tilapia
The Structure and Function of Large Biological Molecules Part 4: Proteins Chapter 5
 Key Concepts: The Structure and Function of Large Biological Molecules Part 4: Proteins Chapter 5 Proteins include a diversity of structures, resulting in a wide range of functions Proteins Enzymatic s
Key Concepts: The Structure and Function of Large Biological Molecules Part 4: Proteins Chapter 5 Proteins include a diversity of structures, resulting in a wide range of functions Proteins Enzymatic s
Chapter 4: Information and Knowledge in the Protein Insulin
 Chapter 4: Information and Knowledge in the Protein Insulin This chapter will calculate the information and molecular knowledge in a real protein. The techniques discussed in this chapter to calculate
Chapter 4: Information and Knowledge in the Protein Insulin This chapter will calculate the information and molecular knowledge in a real protein. The techniques discussed in this chapter to calculate
Midterm 2 Results. Standard Deviation:
 Midterm 2 Results High: Low: Mean: Standard Deviation: 97.5% 16% 58% 16.3 Lecture 17 Amino Acid Metabolism Urea Cycle N and S assimilation Last cofactors: THF and SAM Dietary (Exogenous) Proteins Hydrolyzed
Midterm 2 Results High: Low: Mean: Standard Deviation: 97.5% 16% 58% 16.3 Lecture 17 Amino Acid Metabolism Urea Cycle N and S assimilation Last cofactors: THF and SAM Dietary (Exogenous) Proteins Hydrolyzed
Fundamentals of Organic Chemistry CHEM 109 For Students of Health Colleges
 Fundamentals of Organic Chemistry CHEM 109 For Students of Health Colleges Credit hrs.: (2+1) King Saud University College of Science, Chemistry Department CHEM 109 CHAPTER 9. AMINO ACIDS, PEPTIDES AND
Fundamentals of Organic Chemistry CHEM 109 For Students of Health Colleges Credit hrs.: (2+1) King Saud University College of Science, Chemistry Department CHEM 109 CHAPTER 9. AMINO ACIDS, PEPTIDES AND
Nitrogen uptake by heterotrophic bacteria and phytoplankton in the nitrate-rich Thames estuary
 MARINE ECOLOGY PROGRESS SERIES Vol. 203: 13 21, 2000 Published September 18 Mar Ecol Prog Ser Nitrogen uptake by heterotrophic bacteria and phytoplankton in the nitrate-rich Thames estuary Jack J. Middelburg*,
MARINE ECOLOGY PROGRESS SERIES Vol. 203: 13 21, 2000 Published September 18 Mar Ecol Prog Ser Nitrogen uptake by heterotrophic bacteria and phytoplankton in the nitrate-rich Thames estuary Jack J. Middelburg*,
Macromolecules Structure and Function
 Macromolecules Structure and Function Within cells, small organic molecules (monomers) are joined together to form larger molecules (polymers). Macromolecules are large molecules composed of thousands
Macromolecules Structure and Function Within cells, small organic molecules (monomers) are joined together to form larger molecules (polymers). Macromolecules are large molecules composed of thousands
Algal Biofuels Research: Using basic science to maximize fuel output. Jacob Dums, PhD candidate, Heike Sederoff Lab March 9, 2015
 Algal Biofuels Research: Using basic science to maximize fuel output Jacob Dums, PhD candidate, jtdums@ncsu.edu Heike Sederoff Lab March 9, 2015 Outline Research Approach Dunaliella Increase Oil Content
Algal Biofuels Research: Using basic science to maximize fuel output Jacob Dums, PhD candidate, jtdums@ncsu.edu Heike Sederoff Lab March 9, 2015 Outline Research Approach Dunaliella Increase Oil Content
MS/MS Scan Modes. Eötvös University, Budapest April 16, MS/MS Scan Modes. Árpád Somogyi. Product Ion Scan Select. Scan. Precursor Ion Scan Scan
 MS/MS Modes Árpád Somogyi Eötvös University, Budapest April 16, 2012 MS/MS Modes Product Ion Precursor Ion Neutral Loss Δ ed Reaction Monitoring (SRM) 1 modes in a triple quadrupole (QqQ) (one quadrupole
MS/MS Modes Árpád Somogyi Eötvös University, Budapest April 16, 2012 MS/MS Modes Product Ion Precursor Ion Neutral Loss Δ ed Reaction Monitoring (SRM) 1 modes in a triple quadrupole (QqQ) (one quadrupole
Title. Author(s)DEGUCHI, Eisaburo; NIIYAMA, Masayoshi; KAGOTA, Katsu. CitationJapanese Journal of Veterinary Research, 26(3-4): 68. Issue Date
 Title INCORPORATION OF ^N ADMINISTERED TO GERMFREE AND AMINO ACIDS OF HYDROLYZED LIVER AND MUSCLE PROTEINS Author(s)DEGUCHI, Eisaburo; NIIYAMA, Masayoshi; KAGOTA, Katsu CitationJapanese Journal of
Title INCORPORATION OF ^N ADMINISTERED TO GERMFREE AND AMINO ACIDS OF HYDROLYZED LIVER AND MUSCLE PROTEINS Author(s)DEGUCHI, Eisaburo; NIIYAMA, Masayoshi; KAGOTA, Katsu CitationJapanese Journal of
OCN621: Biological Oceanography- Bioenergetics-III
 OCN621: Biological Oceanography- Bioenergetics-III Guangyi Wang POST 103B guangyi@hawaii.edu http://www.soest.hawaii.edu/oceanography/zij/education/ocn621/ dc 1 /dt = growth Standing Stock C 1 Overview
OCN621: Biological Oceanography- Bioenergetics-III Guangyi Wang POST 103B guangyi@hawaii.edu http://www.soest.hawaii.edu/oceanography/zij/education/ocn621/ dc 1 /dt = growth Standing Stock C 1 Overview
AMINO ACIDS STRUCTURE, CLASSIFICATION, PROPERTIES. PRIMARY STRUCTURE OF PROTEINS
 AMINO ACIDS STRUCTURE, CLASSIFICATION, PROPERTIES. PRIMARY STRUCTURE OF PROTEINS Elena Rivneac PhD, Associate Professor Department of Biochemistry and Clinical Biochemistry State University of Medicine
AMINO ACIDS STRUCTURE, CLASSIFICATION, PROPERTIES. PRIMARY STRUCTURE OF PROTEINS Elena Rivneac PhD, Associate Professor Department of Biochemistry and Clinical Biochemistry State University of Medicine
Chapter 5: Structure and Function of Macromolecules AP Biology 2011
 Chapter 5: Structure and Function of Macromolecules AP Biology 2011 1 Macromolecules Fig. 5.1 Carbohydrates Lipids Proteins Nucleic Acids Polymer - large molecule consisting of many similar building blocks
Chapter 5: Structure and Function of Macromolecules AP Biology 2011 1 Macromolecules Fig. 5.1 Carbohydrates Lipids Proteins Nucleic Acids Polymer - large molecule consisting of many similar building blocks
CHM333 LECTURE 6: 1/25/12 SPRING 2012 Professor Christine Hrycyna AMINO ACIDS II: CLASSIFICATION AND CHEMICAL CHARACTERISTICS OF EACH AMINO ACID:
 AMINO ACIDS II: CLASSIFICATION AND CHEMICAL CHARACTERISTICS OF EACH AMINO ACID: - The R group side chains on amino acids are VERY important. o Determine the properties of the amino acid itself o Determine
AMINO ACIDS II: CLASSIFICATION AND CHEMICAL CHARACTERISTICS OF EACH AMINO ACID: - The R group side chains on amino acids are VERY important. o Determine the properties of the amino acid itself o Determine
PROTEINS. Amino acids are the building blocks of proteins. Acid L-form * * Lecture 6 Macromolecules #2 O = N -C -C-O.
 Proteins: Linear polymers of amino acids workhorses of the cell tools, machines & scaffolds Lecture 6 Macromolecules #2 PRTEINS 1 Enzymes catalysts that mediate reactions, increase reaction rate Structural
Proteins: Linear polymers of amino acids workhorses of the cell tools, machines & scaffolds Lecture 6 Macromolecules #2 PRTEINS 1 Enzymes catalysts that mediate reactions, increase reaction rate Structural
