The peptide samples of C. elegans (15 mg) and S. cerevisiae (20 mg) were prepared as
|
|
|
- Victor Flynn
- 5 years ago
- Views:
Transcription
1 Supplementary material and methods The peptide samples of C. elegans (15 mg) and S. cerevisiae (20 mg) were prepared as described recently 1. The dried down peptides were resolubilized to a final concentration of 1 mg/ml in off-gel electrophoresis buffer containing 6.25% glycerol and 1.25% IPG buffer (GE Healthcare). The peptides were separated on ph 3-10 IPG strips (GE Healthcare) with a 3100 OFFGEL fractionator (Agilent) as previously described 2 using a protocol of 1 hour rehydration at maximum 500 V, 50 ma and 200 mw followed by the separation at maximum 8000V, 100 ma and 300 mw until 50 kvh were reached. After iso-electric focusing, the fractions were concentrated and cleaned up by C18 reversed-phase spin columns according to the manufacturer's instructions (Harvard Apparatus). Phosphopeptides in each fraction were isolated as described in 3 and 4, analyzed using LC- MS/MS and database searches (based on the SGD database (20. Oct 2007) for yeast 5, the wormpep183 database for worm 6 and the IPI database v3.23 for human 7 ) were done as described in 1. We made all the data in PhosphoPep searchable by spectral matching through SpectraST 8. ( Specifically, for each distinct phosphopeptide identified in this study, all corresponding MS2 spectra were collapsed into a single consensus spectrum. Unknown query spectra can then be identified by spectral searching against the library of phosphopeptide consensus spectra. SpectraST can be used both as a web interface in PhosphoPep, and as a stand-alone application released as part of the TPP suite of software 1, 9. The identified (phospho)peptides were mapped to all possible proteins/gene products present in the corresponding database. For Table 1 the total phosphorylation sites includes all sites of phosphopeptides with a dcn >= 0.0 as computed by Sequest 10. A phosphorylation sites was considered to have an assigned site if a dcn (between the first and second Sequest output entry) threshold was exceeded 1, 11. In case of the D. melanogaster dataset a dcn >= 0.1 as computed by Sequest 10 and for the S. cerevisiae, C. elegans as well as the human dataset a dcn >= as computed by Sequest 10 was used to define a phosphorylation site as assigned. 1
2 For Supplementary Figure 1A the protein copies per cell were taken from the publication by Ghaemmagami et al 12. For Supplementary Figure 1A and 1B both the all proteins as well as the GO annotations were taken/retrieved from the yeast SGD database 5. For the C. elegans dataset we omitted the GO analysis as for the 2,959 proteins identified only for 348 a GO annotation molecular function and for 373 a GO annotation biological process is given. Also, to our knowledge, so far no dataset is published which accurately predicts or describes protein abundances in C. elegans and therefore an analogous analysis as for Supplementary Figure 1A was omitted as well. 2
3 Supplementary Figure 1A A comparison of the yeast phosphoprotein abundance (blue) with the abundance of most proteins of the yeast proteome (red) as determined by Ghaemmaghami et al 12. Proteins with more than 20,000 copies per cell are not displayed. The distribution of proteins with more than 20,000 copies per cell is nearly identical between the identified phosphoproteome and yeast proteome. The X-axis displays the protein copies per cell, the Y-axis the percentage of protein counts per copies per cell bin (bin size 100) divided by all proteins from the phosphoprotein or Ghaemmaghami et al 12 data set. 3
4 Supplementary Figure 1B Fraction of identified phosphoproteins (blue) assigned to a given biological function according to gene ontology. As comparison all yeast proteins (red) assigned to a given biological function are shown. 4
5 Supplementary References 1. Bodenmiller, B. et al. PhosphoPep--a phosphoproteome resource for systems biology research in Drosophila Kc167 cells. Mol. Syst. Biol. 3, 139 (2007). 2. Heller, M. et al. Added value for tandem mass spectrometry shotgun proteomics data validation through isoelectric focusing of peptides. J. Proteome Res. 4, (2005). 3. Bodenmiller, B., Mueller, L.N., Mueller, M., Domon, B. & Aebersold, R. Reproducible isolation of distinct, overlapping segments of the phosphoproteome. Nat. Methods 4, (2007). 4. Bodenmiller, B. et al. An integrated chemical, mass spectrometric and computational strategy for (quantitative) phosphoproteomics: application to Drosophila melanogaster Kc167 cells. Mol. BioSyst. 3, (2007). 5. Cherry, J.M. et al. Genetic and physical maps of Saccharomyces cerevisiae. Nature 387, (1997). 6. Rogers, A. et al. WormBase Nucleic Acids Res. 36, D (2008). 7. Kersey, P.J. et al. The International Protein Index: an integrated database for proteomics experiments. Proteomics 4, (2004). 8. Lam, H. et al. Development and validation of a spectral library searching method for peptide identification from MS/MS. Proteomics 7, (2007). 9. Keller, A., Eng, J., Zhang, N., Li, X.J. & Aebersold, R. A uniform proteomics MS/MS analysis platform utilizing open XML file formats. Mol. Syst. Biol. 1, (2005). 10. Eng, J.K., McCormack, A.L. & Yates, J.R. An approach to correlate tandem mass spectral data of peptides with amino acid sequences in a protein database. J. Am. Soc. Mass Spectrom. 5, (1994). 11. Beausoleil, S.A., Villen, J., Gerber, S.A., Rush, J. & Gygi, S.P. A probability-based approach for high-throughput protein phosphorylation analysis and site localization. Nat. Biotechnol. 24, (2006). 12. Ghaemmaghami, S. et al. Global analysis of protein expression in yeast. Nature 425, (2003). 5
6 New PhosphoPep help page and tutorial for users Buttons used in PhosphoPep View available KEGG pathways for this protein ( 1 Start cytoscape network with this protein ( 2 View orthologs/homolog information ( 3 Search for protein interaction information in String ( 4 Look up protein information in PeptideAtlas ( 5 Search protein sequence at Scansite ( 6 Importantly, as the current knowledge about cellular pathways is far from complete, only a portion of the phosphoproteins can be placed into pathways. This partial knowledge also applies to the orthologous protein information as well as to the prediction of a kinase for a given phosphorylation site. Scores and numbers used in PhosphoPep PeptideProphet When interpreting tandem mass spectrometry data, it is crucial to determine if an identification is correct. The PeptideProphet computes a probability of a given fragment ion spectrum to be correctly assigned to a peptide sequence by a given database search algorithm and assigns a score accordingly 7, 8. The range of the score is from 0 (worst) to 1 (best). Depending on the dataset or database the probabilities can slightly vary at a given threshold/score. Tryptic Ends As we analyze peptides in our tandem mass spectrometry experiments we have to digest the proteins using a protease. This is often done by using trypsin. Trypsin cleaves after arginine and lysine but exhibits also some unspecific cleavage 9. Two tryptic ends means that both ends were specifically cut by trypsin. Peptide Mass Molecular mass of the (phospho)peptide deltacn The deltacn score (dcn) is a score computed by the Sequest 10 algorithm which we use to interpret tandem mass spectra. The dcn is the difference between the (normalized) cross-correlation parameter of the first- and second-ranked amino acid sequence assigned to a tandem mass spectrum. Simplified, the dcn tells you how much better 6
7 the first (best) database search hit fits to a tandem mass spectrum than the second hit. In the case of phosphopeptides the dcn also correlates to the correctness of the phosphorylation site assignment within the phosphopeptide sequence 11. # Obs Number of times the phosphopeptide was identified in our experiments # Mappings Maps to # of gene models / maps to # of transcripts How to assess the quality of a phosphopeptide identified using tandem mass spectrometry In order to understand the basic methods of peptide identification using tandem mass spectrometry we strongly recommend studying the presentation which you can find under the link * The presentation is easy to understand and represents a nice introduction to proteomics. Of note: As the following text was written for users without any experience in mass spectrometry we attempted to describe each topic in a simplified manner, sometimes at the expense of accuracy. For users who wish to learn more about each topic we propose to read the literature given at the end of this tutorial. When phosphopeptides are analyzed using liquid chromatography tandem mass spectrometry and phosphopeptide sequences are assigned to the resulting spectra using database search algorithms, primarily two types of error can occur. The first type of error is the mis-assignment of the fragment ion spectrum to a peptide sequence 7, 8. The second type of error is the mis-assignment of the site of phosphorylation in an otherwise correctly identified phosphopeptide 11. Here we explain how each of the errors was assed and how the users of PhosphoPep can use the computed scores and some rules to judge if a phosphopeptide was correctly identified and the site correctly assigned. * Proteome Software Inc. 7
8 Is the phosphopeptide correctly identified? As mentioned above, one type of error in the automatic interpretation of tandem mass spectra is the mis-assignment of the fragment ion spectrum to a peptide sequence. This type of error can be estimated by applying statistical models such as the PeptideProphet 8 and/or by using decoy sequence databases 12. All data loaded into PhosphoPep were assessed using both methods and we already applied a stringent cut off on all data. Therefore the false positive content in the case of the fly data is about 2.6 % (for yeast, worm and human this number is similar). This means that if you don t apply any further filter criteria about 1 out of 38 phosphopeptide entries is wrong. For bioinformatic large scale analyses this false positive rate is in most cases very acceptable, but for a biologist who wants to perform follow up experiments this can already be too high and therefore it is desirable to choose your own false positive rate. So how do you choose your own false positive rate? One of the statistical tools to compute the false positive rate, the PeptideProphet 8, computes a score (ranging from 0 (worst) to 1(best)). This score is displayed for every peptide in PhosphoPep 13. As mentioned above, we have already prefiltered the data, therefore the lowest PeptideProphet score you will find is 0.8. The closer the score is to 1.0 the lower is the chance that you pick a wrongly identified phosphopeptide. For example, at a Peptide Prophet cut off of 0.99 approximately 0.2 % of all entries (equal or above this score) are estimated to be false positive assignments (1 out of 500 phosphopeptide entries) for the fly dataset. 8
9 With the button to the left you can choose the Prophet Score cut off on your own. One further criterion which increases the certainty that a phosphopeptide was correctly identified is the # Obs which tells you how often a phosphopeptide was identified in our experiments. The chance that a phosphopeptide which was identified multiple times is wrong is lower than that of a phosphopeptides that was just identified once (but keep in mind that this is only a rule of thumb and exceptions exist) 14. So taken together, if you choose a phosphopeptide for follow up experiments make sure that it has a high PeptideProphet score and was observed multiple times. 9
10 Is the site of phosphorylation correctly assigned? Often phosphopeptides are rich in serine and threonine residues which can sometimes puzzle the algorithm for the automatic interpretation of tandem mass spectra in regards to which serine/threonine(/tyrosine) was phosphorylated 11. Therefore another type of error connected to phosphopeptides identified using tandem mass spectrometry is the misassignment of the site of phosphorylation in an otherwise correctly identified phosphopeptide 11. This error was estimated by comparing the search engine output scores for the potential phosphorylated forms of a peptide, assuming that any hydroxy-amino acid in a phosphopeptide could be phosphorylated. Based on this estimation we highlighted the phosphopeptides either red (high probability of correct assignment) or yellow (low probability of correct assignment) 10, 11, 13. As one typical approach to study protein phosphorylation is to mutate the site of phosphorylation to another amino acid residue it is advisable to ascertain that you choose the correct amino acid. There are several steps you can take in order to ensure that the site of phosphorylation was correctly assigned. 1) Take a look at the dcn value. The first step to determine the certainty in the phosphorylation site assignment is to look at the dcn score. Simplified, the dcn tells you how much better the first (best) database search hit fits to a tandem mass spectrum than the second hit. Now if the first and second hits are the same phosphopeptide but the Sequest algorithm has problems to unequivocally assign the phosphorylation site, the score will be very low, often close to zero. 10
11 Again as a rule of thumb: The higher the dcn score the more certain is the phosphorylation site assignment. Normally, a score of dcn > corresponds to a high certainty that the site is correctly assigned 11. Below a phosphopeptide is shown which was identified several times but the site of phosphorylation could never be assigned with high certainty. As a result the same phosphopeptide exists in several versions in PhosphoPep. Such agglomerations of the same peptide with many different phosphorylation sites are a hint that the site is not well assigned (but keep in mind, some proteins are heavily phosphorylated and therefore the same peptide can exist in different phosphorylation forms). 11
12 2) Take a look at the kinase phosphorylation motif An additional step to take in order to confirm a site of phosphorylation is to look at the possible kinase motif surrounding the phosphorylation site 15. In the example below phosphorylation sites on the protein FUS3, a MAPK, are shown. Here it is not clear whether R.IIDESAADNSEPTGQQS*GMTEY*VATR.W or R.IIDESAADNSEPTGQQSGMT*EY*VATR.W is correct. Knowing that the MAP kinases are activated by the phosphorylation in the TXY motif we can assume that the R.IIDESAADNSEPTGQQSGMT*EY*VATR.W is correct. 3) Predict the motif using Scansite In case you do not have all kinase motifs memorized you can use the Scansite algorithm 6 to search the protein sequence for possible kinase motifs. For this simply click on the button Search protein sequence at Scansite in the Protein Info section. 12
13 4) Check the evolutionary conservation of the site You can also check whether your phosphorylation site of interest is evolutionary conserved which can be an additional indication for the correct assignment of a phosphorylation site. For this click on the button View orthologs/homolog information and a new window will be opened, showing the alignment of the amino acid sequences with the identified phosphorylation sites between yeast, worm fly and human. In the example below, we can conclude that the unassigned phosphothreonine (highlighted in yellow) is correctly assigned and that in the top amino acid sequence either the tyrosine or threonine in the TXY motif should be phosphorylated. 13
14 5) Take a look at the tandem mass spectrum In order to assess whether the phosphorylation site was correctly assigned, it is always advisable to take a look at the tandem mass spectrum of the phosphopeptide. You can open it by clicking on the symbol (The manual interpretation of tandem mass spectra can be, especially in the case of phosphopeptides, difficult. Therefore we recommend to study the following slides which are a nice introduction to this topic. You can find them under the URL * This will open a new window in which the tandem mass spectrum is displayed (See next page). In the upper window you see the tandem mass spectrum in which the fragment ion peaks are assigned with y-ion or b-ion together with a number (ion assignment nomenclature) as well as below the spectrum the amino acid sequence of the phosphopeptide is shown (a phosphoserine is indicated as S[167], a phosphothreonine as T[181] and a phosphotyrosine as Y[243] ). Here you have to look for the following: left and right of the amino acid sequence the fragment ion signals which were found and could be assigned in the tandem mass spectrum are highlighted in red. In our example the question is, if the serine (at position 6) is phosphorylated LSLTDS 167 TETIENNATVK or the adjacent threonine LSLTDST 167 ETIENNATVK at position 7. * Proteome Software Inc. 14
15 Of note, most spectra loaded into the PhosphoPep database are consensus spectra 16, which means that only repeatedly observed peptide fragment ions are shown. Noise signals were removed. Fragment ion confirming the threonine without phosphorylation Fragment ion confirming the phosphorylation site (S167) b 1+ b 2+ # AA # y 1+ y L S L T D S[167] T E T I E N N A T V K Phosphorylation site The inspection of the highlighted ions shows that indeed all peptide fragment ions, including the one corresponding to the phosphorylated serine as well as the non-phosphorylated 15
16 threonine, were identified and assigned, strengthening that the assigned serine phosphorylation is correct. In addition, take a look at the tandem mass spectrum. Here you can see that both assigned fragment ions are rather intense. In addition, the y 11+ fragment ion at m/z 1,300.3 ( ) or the y 11++ fragment ion at m/z which would indicate that threonine 7 phosphorylated, are missing. The same is true for the b 6+ ion at m/z which would indicate that serine 6 is not phosphorylated. Both findings suggest the correct assignment of the phosphorylation site. On the next two pages an example is shown in which the assignment of the correct site of phosphorylation is difficult. It is not clear if the highlighted serine TS*VSEAQNTQPQVANADAK or the highlighted threonine T*SVSEAQNTQPQVANADAK is phosphorylated. First, most peptide fragment ions which could unequivocally distinguish the two phosphorylation sites are outside the recorded m/z range. Second, the y 18++ fragment ion at m/z which could indicate that the serine 2 is phosphorylated, but not threonine 1, is present at low relative intensity in a m/z region crowded with signals, therefore it is not sure whether this is a real fragment ion or noise. 16
17 Fragment ion pointing to the phosphorylation of serine 2 b 1+ b 2+ # AA # y 1+ y T S[167] V S E A Q N T Q P Q V A N A D A K
18 No fragment ion(s) pointing towards phosphorylation of threonine 1 or serine 2 b 1+ b 2+ # AA # y 1+ y T[181] S V S E A Q N T Q P Q V A N A D A K
19 References 1. Kanehisa, M. et al. From genomics to chemical genomics: new developments in KEGG. Nucleic Acids Res. 34, D (2006). 2. Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, (2003). 3. Chen, F., Mackey, A.J., Stoeckert, C.J., Jr. & Roos, D.S. OrthoMCL-DB: querying a comprehensive multi-species collection of ortholog groups. Nucleic Acids Res. 34, D (2006). 4. von Mering, C. et al. STRING 7--recent developments in the integration and prediction of protein interactions. Nucleic Acids Res. 35, D (2007). 5. Desiere, F. et al. Integration with the human genome of peptide sequences obtained by high-throughput mass spectrometry. Genome Biol. 6, R9 (2005). 6. Obenauer, J.C., Cantley, L.C. & Yaffe, M.B. Scansite 2.0: Proteome-wide prediction of cell signaling interactions using short sequence motifs. Nucleic Acids Res. 31, (2003). 7. Keller, A., Eng, J., Zhang, N., Li, X.J. & Aebersold, R. A uniform proteomics MS/MS analysis platform utilizing open XML file formats. Mol. Syst. Biol. 1, (2005). 8. Keller, A., Nesvizhskii, A.I., Kolker, E. & Aebersold, R. Empirical statistical model to estimate the accuracy of peptide identifications made by MS/MS and database search. Anal. Chem. 74, (2002). 9. Picotti, P., Aebersold, R. & Domon, B. The Implications of Proteolytic Background for Shotgun Proteomics. Mol. Cell. Proteomics (2007). 10. Eng, J.K., McCormack, A.L. & Yates, J.R. An approach to correlate tandem mass spectral data of peptides with amino acid sequences in a protein database. J. Am. Soc. Mass Spectrom. 5, (1994). 11. Beausoleil, S.A., Villen, J., Gerber, S.A., Rush, J. & Gygi, S.P. A probability-based approach for high-throughput protein phosphorylation analysis and site localization. Nat. Biotechnol. 24, (2006). 12. Elias, J.E. & Gygi, S.P. Target-decoy search strategy for increased confidence in large-scale protein identifications by mass spectrometry. Nat. Methods 4, (2007). 13. Bodenmiller, B. et al. PhosphoPep--a phosphoproteome resource for systems biology research in Drosophila Kc167 cells. Mol. Syst. Biol. 3, 139 (2007). 14. Reiter, L. et al. Protein identification false discovery rates for very large proteomics datasets generated by tandem mass spectrometry. Submitted (2008). 15. Olsen, J.V. et al. Global, in vivo, and site-specific phosphorylation dynamics in signaling networks. Cell 127, (2006). 16. Lam, H. et al. Development and validation of a spectral library searching method for peptide identification from MS/MS. Proteomics 7, (2007). 19
Lecture 3. Tandem MS & Protein Sequencing
 Lecture 3 Tandem MS & Protein Sequencing Nancy Allbritton, M.D., Ph.D. Department of Physiology & Biophysics 824-9137 (office) nlallbri@uci.edu Office- Rm D349 Medical Science D Bldg. Tandem MS Steps:
Lecture 3 Tandem MS & Protein Sequencing Nancy Allbritton, M.D., Ph.D. Department of Physiology & Biophysics 824-9137 (office) nlallbri@uci.edu Office- Rm D349 Medical Science D Bldg. Tandem MS Steps:
Probability-Based Protein Identification for Post-Translational Modifications and Amino Acid Variants Using Peptide Mass Fingerprint Data
 Probability-Based Protein Identification for Post-Translational Modifications and Amino Acid Variants Using Peptide Mass Fingerprint Data Tong WW, McComb ME, Perlman DH, Huang H, O Connor PB, Costello
Probability-Based Protein Identification for Post-Translational Modifications and Amino Acid Variants Using Peptide Mass Fingerprint Data Tong WW, McComb ME, Perlman DH, Huang H, O Connor PB, Costello
Protein Reports CPTAC Common Data Analysis Pipeline (CDAP)
 Protein Reports CPTAC Common Data Analysis Pipeline (CDAP) v. 05/03/2016 Summary The purpose of this document is to describe the protein reports generated as part of the CPTAC Common Data Analysis Pipeline
Protein Reports CPTAC Common Data Analysis Pipeline (CDAP) v. 05/03/2016 Summary The purpose of this document is to describe the protein reports generated as part of the CPTAC Common Data Analysis Pipeline
Top 10 Tips for Successful Searching ASMS 2003
 Top 10 Tips for Successful Searching I'd like to present our top 10 tips for successful searching with Mascot. Like any hit parade, we will, of course, count them off in reverse order 1 10. Don t specify
Top 10 Tips for Successful Searching I'd like to present our top 10 tips for successful searching with Mascot. Like any hit parade, we will, of course, count them off in reverse order 1 10. Don t specify
Types of Modifications
 Modifications 1 Types of Modifications Post-translational Phosphorylation, acetylation Artefacts Oxidation, acetylation Derivatisation Alkylation of cysteine, ICAT, SILAC Sequence variants Errors, SNP
Modifications 1 Types of Modifications Post-translational Phosphorylation, acetylation Artefacts Oxidation, acetylation Derivatisation Alkylation of cysteine, ICAT, SILAC Sequence variants Errors, SNP
Nature Methods: doi: /nmeth.3177
 Supplementary Figure 1 Characterization of LysargiNase, trypsin and LysN missed cleavages. (a) Proportion of peptides identified in LysargiNase and trypsin digests of MDA-MB-231 cell lysates carrying 0,
Supplementary Figure 1 Characterization of LysargiNase, trypsin and LysN missed cleavages. (a) Proportion of peptides identified in LysargiNase and trypsin digests of MDA-MB-231 cell lysates carrying 0,
Figure S6. A-J) Annotated UVPD mass spectra for top ten peptides found among the peptides identified by Byonic but not SEQUEST + Percolator.
 Extending Proteome Coverage by Combining MS/MS Methods and a Modified Bioinformatics Platform adapted for Database Searching of Positive and Negative Polarity 193 nm Ultraviolet Photodissociation Mass
Extending Proteome Coverage by Combining MS/MS Methods and a Modified Bioinformatics Platform adapted for Database Searching of Positive and Negative Polarity 193 nm Ultraviolet Photodissociation Mass
Molecular Cell, Volume 46. Supplemental Information
 Molecular Cell, Volume 46 Supplemental Information Mapping N-Glycosylation Sites across Seven Evolutionary Distant Species Reveals a Divergent Substrate Proteome Despite a Common Core Machinery Dorota
Molecular Cell, Volume 46 Supplemental Information Mapping N-Glycosylation Sites across Seven Evolutionary Distant Species Reveals a Divergent Substrate Proteome Despite a Common Core Machinery Dorota
User Guide. Protein Clpper. Statistical scoring of protease cleavage sites. 1. Introduction Protein Clpper Analysis Procedure...
 User Guide Protein Clpper Statistical scoring of protease cleavage sites Content 1. Introduction... 2 2. Protein Clpper Analysis Procedure... 3 3. Input and Output Files... 9 4. Contact Information...
User Guide Protein Clpper Statistical scoring of protease cleavage sites Content 1. Introduction... 2 2. Protein Clpper Analysis Procedure... 3 3. Input and Output Files... 9 4. Contact Information...
Mass Spectrometry and Proteomics - Lecture 4 - Matthias Trost Newcastle University
 Mass Spectrometry and Proteomics - Lecture 4 - Matthias Trost Newcastle University matthias.trost@ncl.ac.uk previously Peptide fragmentation Hybrid instruments 117 The Building Blocks of Life DNA RNA Proteins
Mass Spectrometry and Proteomics - Lecture 4 - Matthias Trost Newcastle University matthias.trost@ncl.ac.uk previously Peptide fragmentation Hybrid instruments 117 The Building Blocks of Life DNA RNA Proteins
PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System
 Application Note LC/MS PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System Purpose This application note describes an automated workflow
Application Note LC/MS PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System Purpose This application note describes an automated workflow
Protein Identification and Phosphorylation Site Determination by de novo sequencing using PepFrag TM MALDI-Sequencing kit
 Application Note Tel: +82-54-223-2463 Fax : +82-54-223-2460 http://www.genomine.com venture ldg 306 Pohang techno park Pohang, kyungbuk, Korea(ROK) Protein Identification and Phosphorylation Site Determination
Application Note Tel: +82-54-223-2463 Fax : +82-54-223-2460 http://www.genomine.com venture ldg 306 Pohang techno park Pohang, kyungbuk, Korea(ROK) Protein Identification and Phosphorylation Site Determination
New Developments in LC-IMS-MS Proteomic Measurements and Informatic Analyses
 New Developments in LC-IMS-MS Proteomic Measurements and Informatic Analyses Erin Shammel Baker Kristin E. Burnum-Johnson, Xing Zhang, Cameron P. Casey, Yehia M. Ibrahim, Matthew E. Monroe, Tao Liu, Brendan
New Developments in LC-IMS-MS Proteomic Measurements and Informatic Analyses Erin Shammel Baker Kristin E. Burnum-Johnson, Xing Zhang, Cameron P. Casey, Yehia M. Ibrahim, Matthew E. Monroe, Tao Liu, Brendan
NIH Public Access Author Manuscript J Proteome Res. Author manuscript; available in PMC 2014 July 05.
 NIH Public Access Author Manuscript Published in final edited form as: J Proteome Res. 2013 July 5; 12(7): 3071 3086. doi:10.1021/pr3011588. Evaluation and Optimization of Mass Spectrometric Settings during
NIH Public Access Author Manuscript Published in final edited form as: J Proteome Res. 2013 July 5; 12(7): 3071 3086. doi:10.1021/pr3011588. Evaluation and Optimization of Mass Spectrometric Settings during
Introduction to Proteomics 1.0
 Introduction to Proteomics 1.0 CMSP Workshop Pratik Jagtap Managing Director, CMSP Objectives Why are we here? For participants: Learn basics of MS-based proteomics Learn what s necessary for success using
Introduction to Proteomics 1.0 CMSP Workshop Pratik Jagtap Managing Director, CMSP Objectives Why are we here? For participants: Learn basics of MS-based proteomics Learn what s necessary for success using
Databehandling. 3. Mark e.g. the first fraction (1: 0-45 min, 2: min, 3; min, 4: min, 5: min, 6: min).
 Databehandling Data analysis 1. Choose Open in the Data analysis window. 2. Press the Open folder and choose the desired analysis. Click the + button, so that the Chromatograms line appears. Click the
Databehandling Data analysis 1. Choose Open in the Data analysis window. 2. Press the Open folder and choose the desired analysis. Click the + button, so that the Chromatograms line appears. Click the
Proteome Informatics. Brian C. Searle Creative Commons Attribution
 Proteome Informatics Brian C. Searle searleb@uw.edu Creative Commons Attribution Section structure Class 1 Class 2 Homework 1 Mass spectrometry and de novo sequencing Database searching and E-value estimation
Proteome Informatics Brian C. Searle searleb@uw.edu Creative Commons Attribution Section structure Class 1 Class 2 Homework 1 Mass spectrometry and de novo sequencing Database searching and E-value estimation
4.2 RESULTS AND DISCUSSION
 phosphorylated proteins on treatment with Erlotinib.This chapter describes the finding of the SILAC experiment. 4.2 RESULTS AND DISCUSSION This is the first global report of its kind using dual strategies
phosphorylated proteins on treatment with Erlotinib.This chapter describes the finding of the SILAC experiment. 4.2 RESULTS AND DISCUSSION This is the first global report of its kind using dual strategies
Supplementary Figures and Notes for Digestion and depletion of abundant
 Supplementary Figures and Notes for Digestion and depletion of abundant proteins improves proteomic coverage Bryan R. Fonslow, Benjamin D. Stein, Kristofor J. Webb, Tao Xu, Jeong Choi, Sung Kyu Park, and
Supplementary Figures and Notes for Digestion and depletion of abundant proteins improves proteomic coverage Bryan R. Fonslow, Benjamin D. Stein, Kristofor J. Webb, Tao Xu, Jeong Choi, Sung Kyu Park, and
OncoPPi Portal A Cancer Protein Interaction Network to Inform Therapeutic Strategies
 OncoPPi Portal A Cancer Protein Interaction Network to Inform Therapeutic Strategies 2017 Contents Datasets... 2 Protein-protein interaction dataset... 2 Set of known PPIs... 3 Domain-domain interactions...
OncoPPi Portal A Cancer Protein Interaction Network to Inform Therapeutic Strategies 2017 Contents Datasets... 2 Protein-protein interaction dataset... 2 Set of known PPIs... 3 Domain-domain interactions...
US-HUPO 2011 Agilent lunch seminar. Robert Moritz. Developing a Complete and Reproducible Human SRMAtlas. Duchy of Luxembourg.
 US-HUPO 2011 Agilent lunch seminar Robert Moritz Developing a Complete and Reproducible Human SRMAtlas Human SRMAtlas Duchy of Luxembourg Outline What is the SRMAtlas? Introduction to the ISB PeptideAtlas:
US-HUPO 2011 Agilent lunch seminar Robert Moritz Developing a Complete and Reproducible Human SRMAtlas Human SRMAtlas Duchy of Luxembourg Outline What is the SRMAtlas? Introduction to the ISB PeptideAtlas:
Learning Objectives. Overview of topics to be discussed 10/25/2013 HIGH RESOLUTION MASS SPECTROMETRY (HRMS) IN DISCOVERY PROTEOMICS
 HIGH RESOLUTION MASS SPECTROMETRY (HRMS) IN DISCOVERY PROTEOMICS A clinical proteomics perspective Michael L. Merchant, PhD School of Medicine, University of Louisville Louisville, KY Learning Objectives
HIGH RESOLUTION MASS SPECTROMETRY (HRMS) IN DISCOVERY PROTEOMICS A clinical proteomics perspective Michael L. Merchant, PhD School of Medicine, University of Louisville Louisville, KY Learning Objectives
MS/MS to Targeted Proteomics (MRM)
 MS/MS to Targeted Proteomics (MRM) How it worked on the Human Lens Proteome Jayson Falkner PhD jay@singleorganism.com Genes Show Limited Value in Predicting Diseases With only a few exceptions, what the
MS/MS to Targeted Proteomics (MRM) How it worked on the Human Lens Proteome Jayson Falkner PhD jay@singleorganism.com Genes Show Limited Value in Predicting Diseases With only a few exceptions, what the
Department of Chemistry, Université de Montréal, C.P. 6128, Succursale centre-ville, Montréal, Québec, H3C 3J7, Canada.
 Phosphoproteome dynamics of Saccharomyces cerevisiae under heat shock and cold stress Evgeny Kanshin 1,5, Peter Kubiniok 1,2,5, Yogitha Thattikota 1,3, Damien D Amours 1,3 and Pierre Thibault 1,2,4 * 1
Phosphoproteome dynamics of Saccharomyces cerevisiae under heat shock and cold stress Evgeny Kanshin 1,5, Peter Kubiniok 1,2,5, Yogitha Thattikota 1,3, Damien D Amours 1,3 and Pierre Thibault 1,2,4 * 1
Metabolomic and Proteomics Solutions for Integrated Biology. Christine Miller Omics Market Manager ASMS 2015
 Metabolomic and Proteomics Solutions for Integrated Biology Christine Miller Omics Market Manager ASMS 2015 Integrating Biological Analysis Using Pathways Protein A R HO R Protein B Protein X Identifies
Metabolomic and Proteomics Solutions for Integrated Biology Christine Miller Omics Market Manager ASMS 2015 Integrating Biological Analysis Using Pathways Protein A R HO R Protein B Protein X Identifies
Tandem mass spectrometry is becoming the
 Fragmentation of Phosphopeptides in an Ion Trap Mass Spectrometer Jon P. DeGnore and Jun Qin Laboratory of Biophysical Chemistry, National Heart, Lung, and Blood Institute, National Institutes of Health,
Fragmentation of Phosphopeptides in an Ion Trap Mass Spectrometer Jon P. DeGnore and Jun Qin Laboratory of Biophysical Chemistry, National Heart, Lung, and Blood Institute, National Institutes of Health,
Glycosylation analysis of blood plasma proteins
 Glycosylation analysis of blood plasma proteins Thesis booklet Eszter Tóth Doctoral School of Pharmaceutical Sciences Semmelweis University Supervisor: Károly Vékey DSc Official reviewers: Borbála Dalmadiné
Glycosylation analysis of blood plasma proteins Thesis booklet Eszter Tóth Doctoral School of Pharmaceutical Sciences Semmelweis University Supervisor: Károly Vékey DSc Official reviewers: Borbála Dalmadiné
SimGlycan. A high-throughput glycan and glycopeptide data analysis tool for LC-, MALDI-, ESI- Mass Spectrometry workflows.
 PREMIER Biosoft SimGlycan A high-throughput glycan and glycopeptide data analysis tool for LC-, MALDI-, ESI- Mass Spectrometry workflows SimGlycan software processes and interprets the MS/MS and higher
PREMIER Biosoft SimGlycan A high-throughput glycan and glycopeptide data analysis tool for LC-, MALDI-, ESI- Mass Spectrometry workflows SimGlycan software processes and interprets the MS/MS and higher
Automating Mass Spectrometry-Based Quantitative Glycomics using Tandem Mass Tag (TMT) Reagents with SimGlycan
 PREMIER Biosoft Automating Mass Spectrometry-Based Quantitative Glycomics using Tandem Mass Tag (TMT) Reagents with SimGlycan Ne uaca2-3galb1-4glc NAcb1 6 Gal NAca -Thr 3 Ne uaca2-3galb1 Ningombam Sanjib
PREMIER Biosoft Automating Mass Spectrometry-Based Quantitative Glycomics using Tandem Mass Tag (TMT) Reagents with SimGlycan Ne uaca2-3galb1-4glc NAcb1 6 Gal NAca -Thr 3 Ne uaca2-3galb1 Ningombam Sanjib
Proteins: Proteomics & Protein-Protein Interactions Part I
 Proteins: Proteomics & Protein-Protein Interactions Part I Jesse Rinehart, PhD Department of Cellular & Molecular Physiology Systems Biology Institute DNA RNA PROTEIN DNA RNA PROTEIN Proteins: Proteomics
Proteins: Proteomics & Protein-Protein Interactions Part I Jesse Rinehart, PhD Department of Cellular & Molecular Physiology Systems Biology Institute DNA RNA PROTEIN DNA RNA PROTEIN Proteins: Proteomics
Nature Biotechnology: doi: /nbt Supplementary Figure 1
 Supplementary Figure 1 The timeline of the NGAG method for extraction of N-linked glycans and glycosite-containing peptides. The timeline can be changed based on the number of samples. Supplementary Figure
Supplementary Figure 1 The timeline of the NGAG method for extraction of N-linked glycans and glycosite-containing peptides. The timeline can be changed based on the number of samples. Supplementary Figure
A computational framework for discovery of glycoproteomic biomarkers
 A computational framework for discovery of glycoproteomic biomarkers Haixu Tang, Anoop Mayampurath, Chuan-Yih Yu Indiana University, Bloomington Yehia Mechref, Erwang Song Texas Tech University 1 Goal:
A computational framework for discovery of glycoproteomic biomarkers Haixu Tang, Anoop Mayampurath, Chuan-Yih Yu Indiana University, Bloomington Yehia Mechref, Erwang Song Texas Tech University 1 Goal:
Agilent Protein In-Gel Tryptic Digestion Kit
 Agilent 5188-2749 Protein In-Gel Tryptic Digestion Kit Agilent Protein In-Gel Tryptic Digestion Kit Instructions Kit Contents The Protein In-Gel Tryptic Digestion Kit includes sufficient reagents for approximately
Agilent 5188-2749 Protein In-Gel Tryptic Digestion Kit Agilent Protein In-Gel Tryptic Digestion Kit Instructions Kit Contents The Protein In-Gel Tryptic Digestion Kit includes sufficient reagents for approximately
* Skyline LC SRM. 2 Skyline LC-MS/MS. Skyline University of Washington Windows.
 Proteome Letters 2016 1 89-94 * *E-mail: fmatsuda@ist.osaka-u.ac.jp 565-0871 1-5 2016 4 20 2016 5 26 2016 5 27 LC SRM LC SRM 1 LC-MS/MS LC 1 3 SRM SRM LC-MS Skyline SRM LC-MS LC SRM 2 Skyline Skyline University
Proteome Letters 2016 1 89-94 * *E-mail: fmatsuda@ist.osaka-u.ac.jp 565-0871 1-5 2016 4 20 2016 5 26 2016 5 27 LC SRM LC SRM 1 LC-MS/MS LC 1 3 SRM SRM LC-MS Skyline SRM LC-MS LC SRM 2 Skyline Skyline University
4th Multidimensional Chromatography Workshop Toronto (January, 2013) Herman C. Lam, Ph.D. Calibration & Validation Group
 4th Multidimensional Chromatography Workshop Toronto (January, 2013) Herman C. Lam, Ph.D. Calibration & Validation Group MDLC for Shotgun Proteomics Introduction General concepts Advantages Challenges
4th Multidimensional Chromatography Workshop Toronto (January, 2013) Herman C. Lam, Ph.D. Calibration & Validation Group MDLC for Shotgun Proteomics Introduction General concepts Advantages Challenges
Mass Spectrometry. Mass spectrometer MALDI-TOF ESI/MS/MS. Basic components. Ionization source Mass analyzer Detector
 Mass Spectrometry MALDI-TOF ESI/MS/MS Mass spectrometer Basic components Ionization source Mass analyzer Detector 1 Principles of Mass Spectrometry Proteins are separated by mass to charge ratio (limit
Mass Spectrometry MALDI-TOF ESI/MS/MS Mass spectrometer Basic components Ionization source Mass analyzer Detector 1 Principles of Mass Spectrometry Proteins are separated by mass to charge ratio (limit
In-Gel Tryptic Digestion Kit
 INSTRUCTIONS In-Gel Tryptic Digestion Kit 3747 N. Meridian Road P.O. Box 117 Rockford, IL 61105 89871 1468.2 Number Description 89871 In-Gel Tryptic Digestion Kit, sufficient reagents for approximately
INSTRUCTIONS In-Gel Tryptic Digestion Kit 3747 N. Meridian Road P.O. Box 117 Rockford, IL 61105 89871 1468.2 Number Description 89871 In-Gel Tryptic Digestion Kit, sufficient reagents for approximately
Unsupervised Identification of Isotope-Labeled Peptides
 Unsupervised Identification of Isotope-Labeled Peptides Joshua E Goldford 13 and Igor GL Libourel 124 1 Biotechnology institute, University of Minnesota, Saint Paul, MN 55108 2 Department of Plant Biology,
Unsupervised Identification of Isotope-Labeled Peptides Joshua E Goldford 13 and Igor GL Libourel 124 1 Biotechnology institute, University of Minnesota, Saint Paul, MN 55108 2 Department of Plant Biology,
Chapter 1. Introduction
 Chapter 1 Introduction 1.1 Motivation and Goals The increasing availability and decreasing cost of high-throughput (HT) technologies coupled with the availability of computational tools and data form a
Chapter 1 Introduction 1.1 Motivation and Goals The increasing availability and decreasing cost of high-throughput (HT) technologies coupled with the availability of computational tools and data form a
Nature Structural & Molecular Biology: doi: /nsmb.3218
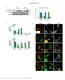 Supplementary Figure 1 Endogenous EGFR trafficking and responses depend on biased ligands. (a) Lysates from HeLa cells stimulated for 2 min. with increasing concentration of ligands were immunoblotted
Supplementary Figure 1 Endogenous EGFR trafficking and responses depend on biased ligands. (a) Lysates from HeLa cells stimulated for 2 min. with increasing concentration of ligands were immunoblotted
Shotgun metaproteomics of the human distal gut microbiota. Present by Lei Chen
 Shotgun metaproteomics of the human distal gut microbiota Present by Lei Chen (lc6@indana.edu) Outline Background What are the goals? Materials and Methods Results Discussion Background The human gastrointestinal
Shotgun metaproteomics of the human distal gut microbiota Present by Lei Chen (lc6@indana.edu) Outline Background What are the goals? Materials and Methods Results Discussion Background The human gastrointestinal
MS/MS Library Creation of Q-TOF LC/MS Data for MassHunter PCDL Manager
 MS/MS Library Creation of Q-TOF LC/MS Data for MassHunter PCDL Manager Quick Start Guide Step 1. Calibrate the Q-TOF LC/MS for low m/z ratios 2 Step 2. Set up a Flow Injection Analysis (FIA) method for
MS/MS Library Creation of Q-TOF LC/MS Data for MassHunter PCDL Manager Quick Start Guide Step 1. Calibrate the Q-TOF LC/MS for low m/z ratios 2 Step 2. Set up a Flow Injection Analysis (FIA) method for
Protein sequence mapping is commonly used to
 Reproducible Microwave-Assisted Acid Hydrolysis of Proteins Using a Household Microwave Oven and Its Combination with LC-ESI MS/MS for Mapping Protein Sequences and Modifications Nan Wang and Liang Li
Reproducible Microwave-Assisted Acid Hydrolysis of Proteins Using a Household Microwave Oven and Its Combination with LC-ESI MS/MS for Mapping Protein Sequences and Modifications Nan Wang and Liang Li
Improve Protein Analysis with the New, Mass Spectrometry- Compatible ProteasMAX Surfactant
 Improve Protein Analysis with the New, Mass Spectrometry- Compatible Surfactant ABSTRACT Incomplete solubilization and digestion and poor peptide recovery are frequent limitations in protein sample preparation
Improve Protein Analysis with the New, Mass Spectrometry- Compatible Surfactant ABSTRACT Incomplete solubilization and digestion and poor peptide recovery are frequent limitations in protein sample preparation
Enhancing Sequence Coverage in Proteomics Studies by Using a Combination of Proteolytic Enzymes
 Enhancing Sequence Coverage in Proteomics Studies by Using a Combination of Proteolytic Enzymes Dominic Baeumlisberger 2, Christopher Kurz 3, Tabiwang N. Arrey, Marion Rohmer 2, Carola Schiller 3, Thomas
Enhancing Sequence Coverage in Proteomics Studies by Using a Combination of Proteolytic Enzymes Dominic Baeumlisberger 2, Christopher Kurz 3, Tabiwang N. Arrey, Marion Rohmer 2, Carola Schiller 3, Thomas
2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry
 Dr. Sanjeeva Srivastava 1. Fundamental of Mass Spectrometry Role of MS and basic concepts 2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry 2 1 MS basic concepts Mass spectrometry - technique
Dr. Sanjeeva Srivastava 1. Fundamental of Mass Spectrometry Role of MS and basic concepts 2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry 2 1 MS basic concepts Mass spectrometry - technique
MSnID Package for Handling MS/MS Identifications
 MSnID Package for Handling MS/MS Identifications Vladislav A. Petyuk November 26, 214 Contents 1 Introduction 1 2 Starting the project 2 3 Reading MS/MS data 2 4 Updating columns 3 5 Basic info on the
MSnID Package for Handling MS/MS Identifications Vladislav A. Petyuk November 26, 214 Contents 1 Introduction 1 2 Starting the project 2 3 Reading MS/MS data 2 4 Updating columns 3 5 Basic info on the
SpliceDB: database of canonical and non-canonical mammalian splice sites
 2001 Oxford University Press Nucleic Acids Research, 2001, Vol. 29, No. 1 255 259 SpliceDB: database of canonical and non-canonical mammalian splice sites M.Burset,I.A.Seledtsov 1 and V. V. Solovyev* The
2001 Oxford University Press Nucleic Acids Research, 2001, Vol. 29, No. 1 255 259 SpliceDB: database of canonical and non-canonical mammalian splice sites M.Burset,I.A.Seledtsov 1 and V. V. Solovyev* The
Quantitative chromatin proteomics reveals a dynamic histone. post-translational modification landscape that defines asexual
 Quantitative chromatin proteomics reveals a dynamic histone post-translational modification landscape that defines asexual and sexual Plasmodium falciparum parasites Nanika Coetzee 1, Simone Sidoli 2,
Quantitative chromatin proteomics reveals a dynamic histone post-translational modification landscape that defines asexual and sexual Plasmodium falciparum parasites Nanika Coetzee 1, Simone Sidoli 2,
Guanghui Han, Mingliang Ye,*, Xinning Jiang, Rui Chen, Jian Ren, Yu Xue, Fangjun Wang, Chunxia Song, Xuebiao Yao, and Hanfa Zou*,
 Anal. Chem. 2009, 81, 5794 5805 Comprehensive and Reliable Phosphorylation Site Mapping of Individual Phosphoproteins by Combination of Multiple Stage Mass Spectrometric Analysis with a Target-Decoy Database
Anal. Chem. 2009, 81, 5794 5805 Comprehensive and Reliable Phosphorylation Site Mapping of Individual Phosphoproteins by Combination of Multiple Stage Mass Spectrometric Analysis with a Target-Decoy Database
Application of a new capillary HPLC- ICP-MS interface to the identification of selenium-containing proteins in selenized yeast
 Application of a new capillary HPLC- ICP-MS interface to the identification of selenium-containing proteins in selenized yeast Application note Food supplements Authors Juliusz Bianga and Joanna Szpunar
Application of a new capillary HPLC- ICP-MS interface to the identification of selenium-containing proteins in selenized yeast Application note Food supplements Authors Juliusz Bianga and Joanna Szpunar
Bioinformatics Laboratory Exercise
 Bioinformatics Laboratory Exercise Biology is in the midst of the genomics revolution, the application of robotic technology to generate huge amounts of molecular biology data. Genomics has led to an explosion
Bioinformatics Laboratory Exercise Biology is in the midst of the genomics revolution, the application of robotic technology to generate huge amounts of molecular biology data. Genomics has led to an explosion
Application Note # ET-17 / MT-99 Characterization of the N-glycosylation Pattern of Antibodies by ESI - and MALDI mass spectrometry
 Bruker Daltonics Application Note # ET-17 / MT-99 Characterization of the N-glycosylation Pattern of Antibodies by ESI - and MALDI mass spectrometry Abstract Analysis of the N-glycosylation pattern on
Bruker Daltonics Application Note # ET-17 / MT-99 Characterization of the N-glycosylation Pattern of Antibodies by ESI - and MALDI mass spectrometry Abstract Analysis of the N-glycosylation pattern on
Proteomics of body liquids as a source for potential methods for medical diagnostics Prof. Dr. Evgeny Nikolaev
 Proteomics of body liquids as a source for potential methods for medical diagnostics Prof. Dr. Evgeny Nikolaev Institute for Biochemical Physics, Rus. Acad. Sci., Moscow, Russia. Institute for Energy Problems
Proteomics of body liquids as a source for potential methods for medical diagnostics Prof. Dr. Evgeny Nikolaev Institute for Biochemical Physics, Rus. Acad. Sci., Moscow, Russia. Institute for Energy Problems
Digitizing the Proteomes From Big Tissue Biobanks
 Digitizing the Proteomes From Big Tissue Biobanks Analyzing 24 Proteomes Per Day by Microflow SWATH Acquisition and Spectronaut Pulsar Analysis Jan Muntel 1, Nick Morrice 2, Roland M. Bruderer 1, Lukas
Digitizing the Proteomes From Big Tissue Biobanks Analyzing 24 Proteomes Per Day by Microflow SWATH Acquisition and Spectronaut Pulsar Analysis Jan Muntel 1, Nick Morrice 2, Roland M. Bruderer 1, Lukas
The 1997 ABRF Mass Spectrometry Committee Collaborative Study: Identification of Phosphopeptides in a Tryptic Digest of Apomyoglobin
 The 1997 ABRF Mass Spectrometry Committee Collaborative Study: Identification of Phosphopeptides in a Tryptic Digest of Apomyoglobin Kristine Swiderek, Farzin Gharahdaghi, Lowell Ericsson, Murray Hackett,
The 1997 ABRF Mass Spectrometry Committee Collaborative Study: Identification of Phosphopeptides in a Tryptic Digest of Apomyoglobin Kristine Swiderek, Farzin Gharahdaghi, Lowell Ericsson, Murray Hackett,
MSSimulator. Simulation of Mass Spectrometry Data. Chris Bielow, Stephan Aiche, Sandro Andreotti, Knut Reinert FU Berlin, Germany
 Chris Bielow Algorithmic Bioinformatics, Institute for Computer Science MSSimulator Chris Bielow, Stephan Aiche, Sandro Andreotti, Knut Reinert FU Berlin, Germany Simulation of Mass Spectrometry Data Motivation
Chris Bielow Algorithmic Bioinformatics, Institute for Computer Science MSSimulator Chris Bielow, Stephan Aiche, Sandro Andreotti, Knut Reinert FU Berlin, Germany Simulation of Mass Spectrometry Data Motivation
Hands-On Ten The BRCA1 Gene and Protein
 Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
Data mining with Ensembl Biomart. Stéphanie Le Gras
 Data mining with Ensembl Biomart Stéphanie Le Gras (slegras@igbmc.fr) Guidelines Genome data Genome browsers Getting access to genomic data: Ensembl/BioMart 2 Genome Sequencing Example: Human genome 2000:
Data mining with Ensembl Biomart Stéphanie Le Gras (slegras@igbmc.fr) Guidelines Genome data Genome browsers Getting access to genomic data: Ensembl/BioMart 2 Genome Sequencing Example: Human genome 2000:
Targeted proteomics reveals strain-specific changes in the mouse insulin and central metabolic pathways after sustained high-fat diet
 Molecular Systems Biology Peer Review Process File Targeted proteomics reveals strain-specific changes in the mouse insulin and central metabolic pathways after sustained high-fat diet Eduard Sabidó, Yibo
Molecular Systems Biology Peer Review Process File Targeted proteomics reveals strain-specific changes in the mouse insulin and central metabolic pathways after sustained high-fat diet Eduard Sabidó, Yibo
Nat Rev Mol Cell Biol (2006) 7, Mass spectrometry and protein analysis. (2006). Domon and Aebersold Science 312,
 Nat Rev Mol Cell Biol (2006) 7, 391-403. Mass spectrometry and protein analysis. (2006). Domon and Aebersold Science 312, 212-217 Affinity Capture / Enrichment Key to success in PTM ID Purified protein
Nat Rev Mol Cell Biol (2006) 7, 391-403. Mass spectrometry and protein analysis. (2006). Domon and Aebersold Science 312, 212-217 Affinity Capture / Enrichment Key to success in PTM ID Purified protein
Moving from targeted towards non-targeted approaches
 Gesundheitsdirektion Moving from targeted towards non-targeted approaches Anton Kaufmann Official Food Control Authority of the Canton of Zurich () Switzerland 2 Overview I From single residue to multi
Gesundheitsdirektion Moving from targeted towards non-targeted approaches Anton Kaufmann Official Food Control Authority of the Canton of Zurich () Switzerland 2 Overview I From single residue to multi
38 Int'l Conf. Bioinformatics and Computational Biology BIOCOMP'16
 38 Int'l Conf. Bioinformatics and Computational Biology BIOCOMP'16 PGAR: ASD Candidate Gene Prioritization System Using Expression Patterns Steven Cogill and Liangjiang Wang Department of Genetics and
38 Int'l Conf. Bioinformatics and Computational Biology BIOCOMP'16 PGAR: ASD Candidate Gene Prioritization System Using Expression Patterns Steven Cogill and Liangjiang Wang Department of Genetics and
SUPPORTING INFORMATION. Lysine Carbonylation is a Previously Unrecognized Contributor. to Peroxidase Activation of Cytochrome c by Chloramine-T
 Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2019 SUPPORTING INFORMATION Lysine Carbonylation is a Previously Unrecognized Contributor to
Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2019 SUPPORTING INFORMATION Lysine Carbonylation is a Previously Unrecognized Contributor to
PREPARED FOR: U.S. Army Medical Research and Materiel Command Fort Detrick, Maryland
 AD Award Number: W81XWH-12-1-0248 TITLE: Targeted Inhibition of Tyrosine Kinase-Mediated Epigenetic Alterations to Prevent Resurgence of Castration-Resistant Prostate Cancer PRINCIPAL INVESTIGATOR: Dr.
AD Award Number: W81XWH-12-1-0248 TITLE: Targeted Inhibition of Tyrosine Kinase-Mediated Epigenetic Alterations to Prevent Resurgence of Castration-Resistant Prostate Cancer PRINCIPAL INVESTIGATOR: Dr.
Sequence Identification And Spatial Distribution of Rat Brain Tryptic Peptides Using MALDI Mass Spectrometric Imaging
 Sequence Identification And Spatial Distribution of Rat Brain Tryptic Peptides Using MALDI Mass Spectrometric Imaging AB SCIEX MALDI TOF/TOF* Systems Patrick Pribil AB SCIEX, Canada MALDI mass spectrometric
Sequence Identification And Spatial Distribution of Rat Brain Tryptic Peptides Using MALDI Mass Spectrometric Imaging AB SCIEX MALDI TOF/TOF* Systems Patrick Pribil AB SCIEX, Canada MALDI mass spectrometric
Shotgun Proteomics MS/MS. Protein Mixture. proteolysis. Peptide Mixture. Time. Abundance. Abundance. m/z. Abundance. m/z 2. Abundance.
 Abundance Abundance Abundance Abundance Abundance Shotgun Proteomics Protein Mixture 1 2 3 MS/MS proteolysis m/z 2 3 Time µlc m/z MS 1 m/z Peptide Mixture m/z Block Diagram of a Mass Spectrometer Sample
Abundance Abundance Abundance Abundance Abundance Shotgun Proteomics Protein Mixture 1 2 3 MS/MS proteolysis m/z 2 3 Time µlc m/z MS 1 m/z Peptide Mixture m/z Block Diagram of a Mass Spectrometer Sample
Biological Mass spectrometry in Protein Chemistry
 Biological Mass spectrometry in Protein Chemistry Tuula Nyman Institute of Biotechnology tuula.nyman@helsinki.fi MASS SPECTROMETRY is an analytical technique that identifies the chemical composition of
Biological Mass spectrometry in Protein Chemistry Tuula Nyman Institute of Biotechnology tuula.nyman@helsinki.fi MASS SPECTROMETRY is an analytical technique that identifies the chemical composition of
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10962 Supplementary Figure 1. Expression of AvrAC-FLAG in protoplasts. Total protein extracted from protoplasts described in Fig. 1a was subjected to anti-flag immunoblot to detect AvrAC-FLAG
doi:10.1038/nature10962 Supplementary Figure 1. Expression of AvrAC-FLAG in protoplasts. Total protein extracted from protoplasts described in Fig. 1a was subjected to anti-flag immunoblot to detect AvrAC-FLAG
Biomolecular Mass Spectrometry
 Lipids ot different than other organic small molecules Carbohydrates Polymers of monosaccharides linked via glycosidic bonds (acetals/ ketals) many different combinationsvery interesting no time ucleic
Lipids ot different than other organic small molecules Carbohydrates Polymers of monosaccharides linked via glycosidic bonds (acetals/ ketals) many different combinationsvery interesting no time ucleic
LC/MS/MS SOLUTIONS FOR LIPIDOMICS. Biomarker and Omics Solutions FOR DISCOVERY AND TARGETED LIPIDOMICS
 LC/MS/MS SOLUTIONS FOR LIPIDOMICS Biomarker and Omics Solutions FOR DISCOVERY AND TARGETED LIPIDOMICS Lipids play a key role in many biological processes, such as the formation of cell membranes and signaling
LC/MS/MS SOLUTIONS FOR LIPIDOMICS Biomarker and Omics Solutions FOR DISCOVERY AND TARGETED LIPIDOMICS Lipids play a key role in many biological processes, such as the formation of cell membranes and signaling
Module 3: Pathway and Drug Development
 Module 3: Pathway and Drug Development Table of Contents 1.1 Getting Started... 6 1.2 Identifying a Dasatinib sensitive cancer signature... 7 1.2.1 Identifying and validating a Dasatinib Signature... 7
Module 3: Pathway and Drug Development Table of Contents 1.1 Getting Started... 6 1.2 Identifying a Dasatinib sensitive cancer signature... 7 1.2.1 Identifying and validating a Dasatinib Signature... 7
Trypsin Mass Spectrometry Grade
 058PR-03 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Trypsin Mass Spectrometry Grade A Chemically Modified, TPCK treated, Affinity Purified
058PR-03 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Trypsin Mass Spectrometry Grade A Chemically Modified, TPCK treated, Affinity Purified
A Method to Determine an E-score Threshold to Identify Potential Cross Reactive Allergens
 A Method to Determine an E-score Threshold to Identify Potential Cross Reactive Allergens Andre Silvanovich Ph. D. Gary Bannon Ph. D. Scott McClain Ph. D. Monsanto Company Regulatory Product Characterization
A Method to Determine an E-score Threshold to Identify Potential Cross Reactive Allergens Andre Silvanovich Ph. D. Gary Bannon Ph. D. Scott McClain Ph. D. Monsanto Company Regulatory Product Characterization
Phosphorylation of proteins Steve Barnes Feb 19th, 2002 in some cases, proteins are found in a stable, hyperphosphorylated state, e.g.
 Phosphorylation of proteins Steve Barnes Feb 19th, 2002 in some cases, proteins are found in a stable, hyperphosphorylated state, e.g., casein more interestingly, in most other cases, it is a transient
Phosphorylation of proteins Steve Barnes Feb 19th, 2002 in some cases, proteins are found in a stable, hyperphosphorylated state, e.g., casein more interestingly, in most other cases, it is a transient
The distribution of log 2 ratio (H/L) for quantified peptides. cleavage sites in each bin of log 2 ratio of quantified. peptides
 Journal: Nature Methods Article Title: Corresponding Author: Protein digestion priority is independent of their abundances Mingliang Ye and Hanfa Zou Supplementary Figure 1 Supplementary Figure 2 The distribution
Journal: Nature Methods Article Title: Corresponding Author: Protein digestion priority is independent of their abundances Mingliang Ye and Hanfa Zou Supplementary Figure 1 Supplementary Figure 2 The distribution
Lipid annotation with MS2Analyzer. Yan Ma 10/24/2013
 Lipid annotation with MS2Analyzer Yan Ma 10/24/2013 Checklist before you start You need to have: 1.A computer with Java environment and Office(2003 or higher) 2.MS/MS spectra in MGF files 3.Latest version
Lipid annotation with MS2Analyzer Yan Ma 10/24/2013 Checklist before you start You need to have: 1.A computer with Java environment and Office(2003 or higher) 2.MS/MS spectra in MGF files 3.Latest version
CEU MASS MEDIATOR USER'S MANUAL Version 2.0, 31 st July 2017
 CEU MASS MEDIATOR USER'S MANUAL Version 2.0, 31 st July 2017 1. Introduction... 2 1.1. System Requirements... 2 2. Peak search... 3 2.1. Simple Search... 3 2.2. Advanced Search... 5 2.3. Batch Search...
CEU MASS MEDIATOR USER'S MANUAL Version 2.0, 31 st July 2017 1. Introduction... 2 1.1. System Requirements... 2 2. Peak search... 3 2.1. Simple Search... 3 2.2. Advanced Search... 5 2.3. Batch Search...
Supplementary Figure S1. Appearacne of new acetyl groups in acetylated lysines using 2,3-13 C 6 pyruvate as a tracer instead of labeled glucose.
 Supplementary Figure S1. Appearacne of new acetyl groups in acetylated lysines using 2,3-13 C 6 pyruvate as a tracer instead of labeled glucose. (a) Relative levels of both new acetylation and all acetylation
Supplementary Figure S1. Appearacne of new acetyl groups in acetylated lysines using 2,3-13 C 6 pyruvate as a tracer instead of labeled glucose. (a) Relative levels of both new acetylation and all acetylation
Double charge of 33kD peak A1 A2 B1 B2 M2+ M/z. ABRF Proteomics Research Group - Qualitative Proteomics Study Identifier Number 14146
 Abstract The 2008 ABRF Proteomics Research Group Study offers participants the chance to participate in an anonymous study to identify qualitative differences between two protein preparations. We used
Abstract The 2008 ABRF Proteomics Research Group Study offers participants the chance to participate in an anonymous study to identify qualitative differences between two protein preparations. We used
One Gene, Many Proteins. Applications of Mass Spectrometry to Proteomics. Why Proteomics? Raghothama Chaerkady, Ph.D.
 Applications of Mass Spectrometry to Proteomics Raghothama Chaerkady, Ph.D. McKusick-Nathans Institute of Genetic Medicine and the Department of Biological Chemistry Why Proteomics? One Gene, Many Proteins
Applications of Mass Spectrometry to Proteomics Raghothama Chaerkady, Ph.D. McKusick-Nathans Institute of Genetic Medicine and the Department of Biological Chemistry Why Proteomics? One Gene, Many Proteins
Quantitation of Protein Phosphorylation Using Multiple Reaction Monitoring
 Quantitation of Protein Phosphorylation Using Multiple Reaction Monitoring Application Note Authors Ning Tang, Christine A. Miller and Keith Waddell Agilent Technologies, Inc. Santa Clara, CA USA This
Quantitation of Protein Phosphorylation Using Multiple Reaction Monitoring Application Note Authors Ning Tang, Christine A. Miller and Keith Waddell Agilent Technologies, Inc. Santa Clara, CA USA This
Characterization of Disulfide Linkages in Proteins by 193 nm Ultraviolet Photodissociation (UVPD) Mass Spectrometry. Supporting Information
 Characterization of Disulfide Linkages in Proteins by 193 nm Ultraviolet Photodissociation (UVPD) Mass Spectrometry M. Montana Quick, Christopher M. Crittenden, Jake A. Rosenberg, and Jennifer S. Brodbelt
Characterization of Disulfide Linkages in Proteins by 193 nm Ultraviolet Photodissociation (UVPD) Mass Spectrometry M. Montana Quick, Christopher M. Crittenden, Jake A. Rosenberg, and Jennifer S. Brodbelt
Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random
 S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
[ APPLICATION NOTE ] High Sensitivity Intact Monoclonal Antibody (mab) HRMS Quantification APPLICATION BENEFITS INTRODUCTION WATERS SOLUTIONS KEYWORDS
![[ APPLICATION NOTE ] High Sensitivity Intact Monoclonal Antibody (mab) HRMS Quantification APPLICATION BENEFITS INTRODUCTION WATERS SOLUTIONS KEYWORDS [ APPLICATION NOTE ] High Sensitivity Intact Monoclonal Antibody (mab) HRMS Quantification APPLICATION BENEFITS INTRODUCTION WATERS SOLUTIONS KEYWORDS](/thumbs/79/80328542.jpg) Yun Wang Alelyunas, Henry Shion, Mark Wrona Waters Corporation, Milford, MA, USA APPLICATION BENEFITS mab LC-MS method which enables users to achieve highly sensitive bioanalysis of intact trastuzumab
Yun Wang Alelyunas, Henry Shion, Mark Wrona Waters Corporation, Milford, MA, USA APPLICATION BENEFITS mab LC-MS method which enables users to achieve highly sensitive bioanalysis of intact trastuzumab
Metabolite identification in metabolomics: Metlin Database and interpretation of MSMS spectra
 Metabolite identification in metabolomics: Metlin Database and interpretation of MSMS spectra Jeevan K. Prasain, PhD Department of Pharmacology and Toxicology, UAB jprasain@uab.edu Outline Introduction
Metabolite identification in metabolomics: Metlin Database and interpretation of MSMS spectra Jeevan K. Prasain, PhD Department of Pharmacology and Toxicology, UAB jprasain@uab.edu Outline Introduction
Supplementary Information
 Supplementary Information A novel mass spectrometric strategy BEMAP reveals Extensive O-linked protein glycosylation in Enterotoxigenic Escherichia coli Anders Boysen, Giuseppe Palmisano, Thøger Jensen
Supplementary Information A novel mass spectrometric strategy BEMAP reveals Extensive O-linked protein glycosylation in Enterotoxigenic Escherichia coli Anders Boysen, Giuseppe Palmisano, Thøger Jensen
SIEVE 2.1 Proteomics Example
 SIEVE 2.1 Proteomics Example Software Overview What is SIEVE? SIEVE is Thermo Scientific s differential software solution. SIEVE will continue to enhance our current product for label-free differential
SIEVE 2.1 Proteomics Example Software Overview What is SIEVE? SIEVE is Thermo Scientific s differential software solution. SIEVE will continue to enhance our current product for label-free differential
Technical Note # TN-31 Redefining MALDI-TOF/TOF Performance
 Bruker Daltonics Technical Note # TN-31 Redefining MALDI-TOF/TOF Performance The new ultraflextreme exceeds all current expectations of MALDI-TOF/TOF technology: A proprietary khz smartbeam-ii TM MALDI
Bruker Daltonics Technical Note # TN-31 Redefining MALDI-TOF/TOF Performance The new ultraflextreme exceeds all current expectations of MALDI-TOF/TOF technology: A proprietary khz smartbeam-ii TM MALDI
RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays
 Supplementary Materials RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays Junhee Seok 1*, Weihong Xu 2, Ronald W. Davis 2, Wenzhong Xiao 2,3* 1 School of Electrical Engineering,
Supplementary Materials RASA: Robust Alternative Splicing Analysis for Human Transcriptome Arrays Junhee Seok 1*, Weihong Xu 2, Ronald W. Davis 2, Wenzhong Xiao 2,3* 1 School of Electrical Engineering,
Protein and peptide separations,
 A Quarterly Technical Newsletter Spring Grace Vydac: Leading the Way in Separations for Proteomics Protein and peptide separations, always important to protein chemists and enzymologists, have recently
A Quarterly Technical Newsletter Spring Grace Vydac: Leading the Way in Separations for Proteomics Protein and peptide separations, always important to protein chemists and enzymologists, have recently
Babu Antharavally, Ryan Bomgarden, and John Rogers Thermo Fisher Scientific, Rockford, IL
 A Versatile High-Recovery Method for Removing Detergents from Low-Concentration Protein or Peptide Samples for Mass Spectrometry Sample Preparation and Analysis Babu Antharavally, Ryan Bomgarden, and John
A Versatile High-Recovery Method for Removing Detergents from Low-Concentration Protein or Peptide Samples for Mass Spectrometry Sample Preparation and Analysis Babu Antharavally, Ryan Bomgarden, and John
Thermo Scientific LipidSearch Software for Lipidomics Workflows. Automated Identification and Relative. Quantitation of Lipids by LC/MS
 Thermo Scientific LipidSearch Software for Lipidomics Workflows Automated Identification and Relative of Lipids by LC/MS The promise of lipidomics Lipidomics is a new field of study crucial for understanding
Thermo Scientific LipidSearch Software for Lipidomics Workflows Automated Identification and Relative of Lipids by LC/MS The promise of lipidomics Lipidomics is a new field of study crucial for understanding
IMPaLA tutorial.
 IMPaLA tutorial http://impala.molgen.mpg.de/ 1. Introduction IMPaLA is a web tool, developed for integrated pathway analysis of metabolomics data alongside gene expression or protein abundance data. It
IMPaLA tutorial http://impala.molgen.mpg.de/ 1. Introduction IMPaLA is a web tool, developed for integrated pathway analysis of metabolomics data alongside gene expression or protein abundance data. It
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/8/398/rs12/dc1 Supplementary Materials for Quantitative phosphoproteomics reveals new roles for the protein phosphatase PP6 in mitotic cells Scott F. Rusin, Kate
www.sciencesignaling.org/cgi/content/full/8/398/rs12/dc1 Supplementary Materials for Quantitative phosphoproteomics reveals new roles for the protein phosphatase PP6 in mitotic cells Scott F. Rusin, Kate
The N-terminal loop of IRAK-4 death domain regulates ordered assembly of the Myddosome signalling scaffold
 The N-terminal loop of IRAK-4 death domain regulates ordered assembly of the Myddosome signalling scaffold Anthony C.G. Dossang * 1, Precious G. Motshwene * 1, Yang Yang* 1, Martyn F. Symmons*, Clare E.
The N-terminal loop of IRAK-4 death domain regulates ordered assembly of the Myddosome signalling scaffold Anthony C.G. Dossang * 1, Precious G. Motshwene * 1, Yang Yang* 1, Martyn F. Symmons*, Clare E.
ISOELECTRIC TRAPPING AND MASS SPECTROMETRY: TOOLS FOR PROTEOMICS. A Dissertation STEPHANIE MARIE COLOGNA
 ISOELECTRIC TRAPPING AND MASS SPECTROMETRY: TOOLS FOR PROTEOMICS A Dissertation by STEPHANIE MARIE COLOGNA Submitted to the Office of Graduate Studies of Texas A&M University in partial fulfillment of
ISOELECTRIC TRAPPING AND MASS SPECTROMETRY: TOOLS FOR PROTEOMICS A Dissertation by STEPHANIE MARIE COLOGNA Submitted to the Office of Graduate Studies of Texas A&M University in partial fulfillment of
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Experimental design and workflow utilized to generate the WMG Protein Atlas.
 Supplementary Figure 1 Experimental design and workflow utilized to generate the WMG Protein Atlas. (a) Illustration of the plant organs and nodule infection time points analyzed. (b) Proteomic workflow
Supplementary Figure 1 Experimental design and workflow utilized to generate the WMG Protein Atlas. (a) Illustration of the plant organs and nodule infection time points analyzed. (b) Proteomic workflow
Contents. 2 Statistics Static reference method Sampling reference set Statistics Sampling Types...
 Department of Medical Protein Research, VIB, B-9000 Ghent, Belgium Department of Biochemistry, Ghent University, B-9000 Ghent, Belgium http://www.computationalproteomics.com icelogo manual Niklaas Colaert
Department of Medical Protein Research, VIB, B-9000 Ghent, Belgium Department of Biochemistry, Ghent University, B-9000 Ghent, Belgium http://www.computationalproteomics.com icelogo manual Niklaas Colaert
Supporting Information MALDI versus ESI the impact of the ion source on peptide identification
 Supporting Information MALDI versus ESI the impact of the ion source on peptide identification Wiebke Maria Nadler, 1 Dietmar Waidelich, 2 Alexander Kerner, 1 Sabrina Hanke, 1 Regina Berg, 3 Andreas Trumpp
Supporting Information MALDI versus ESI the impact of the ion source on peptide identification Wiebke Maria Nadler, 1 Dietmar Waidelich, 2 Alexander Kerner, 1 Sabrina Hanke, 1 Regina Berg, 3 Andreas Trumpp
