Bayesian Inference for Single-cell ClUstering and ImpuTing (BISCUIT) Elham Azizi
|
|
|
- Allen Adams
- 6 years ago
- Views:
Transcription
1 Bayesian Inference for Single-cell ClUstering and ImpuTing (BISCUIT) Elham Azizi BioC 2017: Where Software and Biology Connect
2 Profiling Tumor-Immune Ecosystem in Breast Cancer Immunotherapy treatments successful only in a subset of patients and cancer types Underlying biology determining success is not known Variability in responses suggest a complex immune environment Goal: Unsupervised characterization of tumor-infiltrating immune subpopulations across subtypes of breast cancer, identify impact of environmental cues Understanding the tumor-immune ecosystem can guide development of treatments to activate immune cells against the tumor Pilot Data: Single-cell RNA-seq 9000 CD45+ immune cells from 4 tumors (patients) *figure adapted from Kroemer Nat Med Collaboration with Alexander Rudensky, MSKCC
3 Single-cell RNA-seq reveals heterogeneity in expression Measurement of gene expression at resolution of single cells Break Count matrix Allows characterizing novel cell types and functions based on heterogeneity indrop (Klein et al Cell 2015) PBMC single-cell data (Zheng et al. biorxiv 2016) 3
4 Single-cell RNA-seq data for immune cells from 4 breast cancer tumors MSK1 MSK2 MSK3 MSK4 lymphoidc ells monocytic cells 9000 CD45+ cells from 4 tumors Normalization by library size Unclear structure of cell types Large patient biases Problems of scrna-seq data: Sampling sparse amounts of mrna leads to Drop-outs Amplification differences Cell-type specific capture rates 4
5 Goal: Characterizing cell subpopulations using Single-cell RNA-seq data Count Matrix Genes Cells 2D projection of cells (TSNE) Cluster 1 Cluster 2 Cluster 3 5
6 Problems: Single-cell RNA-seq data involves significant dropouts and library size variation Observed Count Matrix Genes Cells 2D projection of cells (TSNE) Need to Impute dropouts & Normalize data Cluster 1 Cluster 2 Cluster 3 6
7 Common Approach: Normalizing independent of cell types Observed Count Matrix Cells Normalization To mean/median library size Downsampling BASiCS with spike-ins/erccs Genes Problems: Clustering Cells Downstream Analysis Dropouts not resolved Zeros remain zero! Removes biological stochasticity specific to cell type Leads to improper clustering; Biased downstream analysis 7
8 Main Concepts behind Biscuit for Normalization and Imputing 8
9 Problem #1: Imputing Two ideas for imputing expression in Single-cell RNA-seq data A 2D projection of cells (TSNE) 9
10 Problem #1: Imputing Idea 1: Impute dropouts based on cell type A No expression of Gene A in a cell But we observe cells with same type mostly have high expression of Gene A 10
11 Problem #1: Imputing Idea 1: Impute dropouts based on cell type A No expression of Gene A in a cell But we observe cells with same type mostly have high expression of Gene A Impute dropout in Gene A based on similar cells 11
12 Problem #1: Imputing Idea 2: Impute dropouts based on co-expression patterns No significant inference based on similar cells 12
13 Problem #1: Imputing Idea 2: Impute dropouts based on co-expression patterns No significant inference based on similar cells However Gene A always co-expressed with Gene B in cells of same type 13
14 Problem #1: Imputing Idea 2: Impute dropouts based on co-expression patterns No significant inference based on similar cells However Gene A always co-expressed with Gene B in cells of same type Impute dropout in Gene A based on Gene B 14
15 Problem #2: Normalization Normalization of Single-cell RNA-seq data Histogram of library size in example SC dataset From Zeisel, Science 2014 In addition to imputing dropouts, we need to normalize data by library size 15
16 Problem #2: Normalization Problem with Global Normalization Cells with different sizes have very different total number of transcripts Example Housekeeping Gene High chance of Dropouts in smaller cells 0 16
17 Problem #2: Normalization Problem with Global Normalization Dropout not resolved 0 0 Spurious Differential Expression 17
18 Problem #2: Normalization Key: Different normalization for each cell type Chicken and egg problem: Normalize based on cell types but we do not know cell types! 18
19 Approach: Simultaneous inference of clusters and imputing parameters Clustering Cells iterative learning Imputing & Normalization x1 x3 x 0.75 x1 19
20 Approach: Simultaneous inference of clusters and imputing parameters Cell-specific Parameters Parameters characterizing each cluster iterative for Imputing & learning Normalization x1 Assignment of cells to clusters x3 x 0.75 x1 20
21 Modeling Single-cell data using a Bayesian Mixture Model 21
22 Modeling Clusters of Cells using a Bayesian Mixture Model Ideal Count Matrix (normalized) Genes Cells Cluster 1 Cluster 2 Cluster 3 22
23 Modeling Clusters of Cells using a Bayesian Mixture Model Ideal Count Matrix (normalized) Cells Cluster 1 Cluster 2 Cluster 3 Genes One gene Cluster 1 Cluster 2 Cluster 3 Each gene: Mixture of Log-Normal Models 23
24 Modeling Clusters of Cells using a Bayesian Mixture Model Ideal Count Matrix (normalized) Cells Cluster 1 Cluster 2 Cluster 3 Genes Two genes Cluster 1 Cluster 2 Cluster 3 Modeling all genes together: Mixture of Multivariate Log-Normals To also take advantage of co-expression patterns in learning clusters 24
25 Without Technical Variation Generative Model 25
26 Generative Model with Technical Variation Without Technical Variation Latent counts which we want to recover With Technical Variation Observation 26
27 BISCUIT (Bayesian Inference for Single-cell ClUstering and ImpuTing) Data-driven priors to adapt to different datasets Global Distributions of Observed Data Hyper-priors Hyper-parameters Priors set based on observed lib size distribution Cluster-specific parameters Cell-specific parameters Prabhakaran*, Azizi* ICML 2016 Observed gene expression per cell j 27
28 Model Specification Prabhakaran*, Azizi* ICML
29 Inference Algorithm Parallel Sampling from derived conditional posterior distributions: P(parameter data, other parameters) Estimate hyper-priors based on Data Sample hyper-parameters Sampling technical variation parameters Gibbs iterations Sampling cluster-specific parameters Sampling assignment of cells to clusters using Chinese Restaurant Process (CRP) scaling mean, cov per cell Also allows estimating the number of clusters Prabhakaran*, Azizi* ICML
30 Performance on Simulated Data Data simulated from model for 100 cells, 50 genes in 3 clusters Confusion matrices showing true labels and those from MCMC-based methods Boxplots of F-scores obtained in 15 experiments with randomly-generated data Prabhakaran*, Azizi* ICML
31 Model Mismatch: Robustness when counts are not LogNormal Noncentral Student s t Prabhakaran*, Azizi* ICML 2016 (Log) Negative binomial 31
32 Performance on Single-cell Data Zeisel et al., mouse cortex cells, with UMIs Deep coverage gives good ground truth for 7 Cell types Selected 558 genes with largest standard deviation across cells Fit model to log(counts+1) Prabhakaran*, Azizi* ICML 2016 F-score:
33 Comparison to other methods F-score: 0.21 DBscan F-score: 0.91 F-score: 0.79 F-score: 0.74 F-score:
34 Cluster-dependent Imputing & Normalizing Genes Cells Observed Data BISCUIT clusters & parameters Impute & Normalize With a linear transformation I 34
35 Cluster-dependent Imputing & Normalizing Observed Data BISCUIT clusters & parameters Cells Imputed Data Genes Genes Cells Cluster 1 Cluster 2 Cluster 3 35
36 Characterizing tumor-infiltrating immune cells in breast cancer 36
37 Single-cell RNA-seq data for immune cells from 4 breast cancer tumors MSK1 MSK2 MSK3 MSK4 Unclear structure of cell types Unclear structure of cell types Large patient biases Large patient biases 37
38 Single-cell RNA-seq data for immune cells from 4 breast cancer tumors MSK1 MSK2 MSK3 MSK4 Unclear structure of cell types Large patient biases Defined Structure Corrected patient biases 38
39 Impact of environment in immune cell heterogeneity Little known about how tissue microenvironment modulates anti-tumor immunity Collected 50K CD45+ leukocytes from 8 patients Different ranges of tumor grade, ER, PR, Her2, age, two cases of metastases Different environments (tissues): Tumor Peripheral blood Lymphnode Normal (prophylactic mastectomies) Largest single cell immune map based on tissue residence. 39
40 Map of immune cells from 8 breast cancer patients Normalized by BISCUIT 50K cells 82 clusters 82 clusters Higher entropy shows more mixing of patients with Biscuit 40
41 Map of immune cells from 8 breast cancer patients Normalized by BISCUIT Tregs macrophages B cells DCs Cytotoxic T Central memory T monocytes neutrophils 82 clusters T cells NK cells 41
42 Impact of Environments Tissues Tregs Monocytic cells B cells T cells NK cells 42
43 82 clusters Tregs Monocytic cells B cells What are the differences across inferred cell subpopulations? T cells NK cells 43
44 T cell activation: First component of variation Correlated genes enriched for cytokine production & signaling, lymphocyte activation, leukocyte differentiation, ligand receptor interaction 44
45 T cell activation: First component of variation Comparing distribution of cells along the activation component shows tumor is more activated. Tumor Lymphnode Normal Blood Cell Density T cell Activation Component 45
46 Cell Density T cell Activation Component Activation state of each cluster 46 46
47 Activation of monocytic cells: first components of variation 47
48 Covariance patterns identify Treg clusters Markers differentially expressed in mean but differ in covariance patterns 48
49 Different covariance patterns of immunotherapy targets in Tregs across patients Incorporating personalized co-expression of drug targets can broaden the scope of immunotherapy 49
50 Macrophage clusters differ in covariance between M1/M2 markers 50
51 Summary Analyzing single cell data involves computational challenges: dropouts, technical variation dependent on cell types BISCUIT: A bayesian approach for simultaneous clustering and imputing Clusters identified with both mean and gene-gene covariance patterns Incorporating covariance informations improves normalization and imputing Map of tumor-immune ecosystem in breast cancer Single cell data for 50K CD45+ cells from 8 patients analyzed with Biscuit Substantial diversity of immune cell types driven by environment Activation of T cells and monocytic cell types explain most of variation Covariance patterns can be informative in characterization of cell types and development of personalized treatments 51
52 Acknowledgments Dana Pe er Alexander Rudensky Barbara Engelhardt (Princeton) David Blei (Columbia) Pe er Lab Sandhya Prabhakaran Ambrose Carr Andrew Cornish Linas Mazutis Vaidotas Kiseliovas Juozas Nainys Rudensky Lab George Plitas Kasia Konopacki R code: elham.azizi@gmail.com 52
The power of single cells: Building a tumor immune atlas Dana Pe er Department of Biological Science Department of Systems Biology Columbia
 The power of single cells: Building a tumor immune atlas Dana Pe er Department of Biological Science Department of Systems Biology Columbia University The Precision Medicine Initiative Most personalized
The power of single cells: Building a tumor immune atlas Dana Pe er Department of Biological Science Department of Systems Biology Columbia University The Precision Medicine Initiative Most personalized
Single-Cell Sequencing in Cancer. Peter A. Sims, Columbia University G4500: Cellular & Molecular Biology of Cancer October 22, 2018
 Single-Cell Sequencing in Cancer Peter A. Sims, Columbia University G4500: Cellular & Molecular Biology of Cancer October 22, 2018 Lecture will focus on technology for and applications of single-cell RNA-seq
Single-Cell Sequencing in Cancer Peter A. Sims, Columbia University G4500: Cellular & Molecular Biology of Cancer October 22, 2018 Lecture will focus on technology for and applications of single-cell RNA-seq
Research Strategy: 1. Background and Significance
 Research Strategy: 1. Background and Significance 1.1. Heterogeneity is a common feature of cancer. A better understanding of this heterogeneity may present therapeutic opportunities: Intratumor heterogeneity
Research Strategy: 1. Background and Significance 1.1. Heterogeneity is a common feature of cancer. A better understanding of this heterogeneity may present therapeutic opportunities: Intratumor heterogeneity
AbSeq on the BD Rhapsody system: Exploration of single-cell gene regulation by simultaneous digital mrna and protein quantification
 BD AbSeq on the BD Rhapsody system: Exploration of single-cell gene regulation by simultaneous digital mrna and protein quantification Overview of BD AbSeq antibody-oligonucleotide conjugates. High-throughput
BD AbSeq on the BD Rhapsody system: Exploration of single-cell gene regulation by simultaneous digital mrna and protein quantification Overview of BD AbSeq antibody-oligonucleotide conjugates. High-throughput
Supplement to SCnorm: robust normalization of single-cell RNA-seq data
 Supplement to SCnorm: robust normalization of single-cell RNA-seq data Supplementary Note 1: SCnorm does not require spike-ins, since we find that the performance of spike-ins in scrna-seq is often compromised,
Supplement to SCnorm: robust normalization of single-cell RNA-seq data Supplementary Note 1: SCnorm does not require spike-ins, since we find that the performance of spike-ins in scrna-seq is often compromised,
Data Analysis Using Regression and Multilevel/Hierarchical Models
 Data Analysis Using Regression and Multilevel/Hierarchical Models ANDREW GELMAN Columbia University JENNIFER HILL Columbia University CAMBRIDGE UNIVERSITY PRESS Contents List of examples V a 9 e xv " Preface
Data Analysis Using Regression and Multilevel/Hierarchical Models ANDREW GELMAN Columbia University JENNIFER HILL Columbia University CAMBRIDGE UNIVERSITY PRESS Contents List of examples V a 9 e xv " Preface
Aspects of Statistical Modelling & Data Analysis in Gene Expression Genomics. Mike West Duke University
 Aspects of Statistical Modelling & Data Analysis in Gene Expression Genomics Mike West Duke University Papers, software, many links: www.isds.duke.edu/~mw ABS04 web site: Lecture slides, stats notes, papers,
Aspects of Statistical Modelling & Data Analysis in Gene Expression Genomics Mike West Duke University Papers, software, many links: www.isds.duke.edu/~mw ABS04 web site: Lecture slides, stats notes, papers,
Bayesian Prediction Tree Models
 Bayesian Prediction Tree Models Statistical Prediction Tree Modelling for Clinico-Genomics Clinical gene expression data - expression signatures, profiling Tree models for predictive sub-typing Combining
Bayesian Prediction Tree Models Statistical Prediction Tree Modelling for Clinico-Genomics Clinical gene expression data - expression signatures, profiling Tree models for predictive sub-typing Combining
Molecular Profiling of Tumor Microenvironment Alex Chenchik, Ph.D. Cellecta, Inc.
 Molecular Profiling of Tumor Microenvironment Alex Chenchik, Ph.D. Cellecta, Inc. Cellecta, Inc. Founded: April 2006 Headquarters: Mountain View, CA 12 SBIR Grants Custom Service Provider for Functional
Molecular Profiling of Tumor Microenvironment Alex Chenchik, Ph.D. Cellecta, Inc. Cellecta, Inc. Founded: April 2006 Headquarters: Mountain View, CA 12 SBIR Grants Custom Service Provider for Functional
Identification of Tissue Independent Cancer Driver Genes
 Identification of Tissue Independent Cancer Driver Genes Alexandros Manolakos, Idoia Ochoa, Kartik Venkat Supervisor: Olivier Gevaert Abstract Identification of genomic patterns in tumors is an important
Identification of Tissue Independent Cancer Driver Genes Alexandros Manolakos, Idoia Ochoa, Kartik Venkat Supervisor: Olivier Gevaert Abstract Identification of genomic patterns in tumors is an important
Spatially resolved multiparametric single cell analysis. Technical Journal Club 19th September 2017 Christina Müller (Group Speck)
 Spatially resolved multiparametric single cell analysis Technical Journal Club 19th September 2017 Christina Müller (Group Speck) Why spatially resolved multiparametric single cell analysis? Multiparametric
Spatially resolved multiparametric single cell analysis Technical Journal Club 19th September 2017 Christina Müller (Group Speck) Why spatially resolved multiparametric single cell analysis? Multiparametric
Kelvin Chan Feb 10, 2015
 Underestimation of Variance of Predicted Mean Health Utilities Derived from Multi- Attribute Utility Instruments: The Use of Multiple Imputation as a Potential Solution. Kelvin Chan Feb 10, 2015 Outline
Underestimation of Variance of Predicted Mean Health Utilities Derived from Multi- Attribute Utility Instruments: The Use of Multiple Imputation as a Potential Solution. Kelvin Chan Feb 10, 2015 Outline
Advanced Bayesian Models for the Social Sciences
 Advanced Bayesian Models for the Social Sciences Jeff Harden Department of Political Science, University of Colorado Boulder jeffrey.harden@colorado.edu Daniel Stegmueller Department of Government, University
Advanced Bayesian Models for the Social Sciences Jeff Harden Department of Political Science, University of Colorado Boulder jeffrey.harden@colorado.edu Daniel Stegmueller Department of Government, University
Adaptive Trial Design and Incorporation of Biomarkers to Maximize Achievable Objectives. In Early Phase Clinical Studies
 Adaptive Trial Design and Incorporation of Biomarkers to Maximize Achievable Objectives In Early Phase Clinical Studies Exclusive Offer for Attendees! Stay tuned until after the webinar to receive details
Adaptive Trial Design and Incorporation of Biomarkers to Maximize Achievable Objectives In Early Phase Clinical Studies Exclusive Offer for Attendees! Stay tuned until after the webinar to receive details
Ordinal Data Modeling
 Valen E. Johnson James H. Albert Ordinal Data Modeling With 73 illustrations I ". Springer Contents Preface v 1 Review of Classical and Bayesian Inference 1 1.1 Learning about a binomial proportion 1 1.1.1
Valen E. Johnson James H. Albert Ordinal Data Modeling With 73 illustrations I ". Springer Contents Preface v 1 Review of Classical and Bayesian Inference 1 1.1 Learning about a binomial proportion 1 1.1.1
fl/+ KRas;Atg5 fl/+ KRas;Atg5 fl/fl KRas;Atg5 fl/fl KRas;Atg5 Supplementary Figure 1. Gene set enrichment analyses. (a) (b)
 KRas;At KRas;At KRas;At KRas;At a b Supplementary Figure 1. Gene set enrichment analyses. (a) GO gene sets (MSigDB v3. c5) enriched in KRas;Atg5 fl/+ as compared to KRas;Atg5 fl/fl tumors using gene set
KRas;At KRas;At KRas;At KRas;At a b Supplementary Figure 1. Gene set enrichment analyses. (a) GO gene sets (MSigDB v3. c5) enriched in KRas;Atg5 fl/+ as compared to KRas;Atg5 fl/fl tumors using gene set
Outlier Analysis. Lijun Zhang
 Outlier Analysis Lijun Zhang zlj@nju.edu.cn http://cs.nju.edu.cn/zlj Outline Introduction Extreme Value Analysis Probabilistic Models Clustering for Outlier Detection Distance-Based Outlier Detection Density-Based
Outlier Analysis Lijun Zhang zlj@nju.edu.cn http://cs.nju.edu.cn/zlj Outline Introduction Extreme Value Analysis Probabilistic Models Clustering for Outlier Detection Distance-Based Outlier Detection Density-Based
Confounding by indication developments in matching, and instrumental variable methods. Richard Grieve London School of Hygiene and Tropical Medicine
 Confounding by indication developments in matching, and instrumental variable methods Richard Grieve London School of Hygiene and Tropical Medicine 1 Outline 1. Causal inference and confounding 2. Genetic
Confounding by indication developments in matching, and instrumental variable methods Richard Grieve London School of Hygiene and Tropical Medicine 1 Outline 1. Causal inference and confounding 2. Genetic
Advanced Bayesian Models for the Social Sciences. TA: Elizabeth Menninga (University of North Carolina, Chapel Hill)
 Advanced Bayesian Models for the Social Sciences Instructors: Week 1&2: Skyler J. Cranmer Department of Political Science University of North Carolina, Chapel Hill skyler@unc.edu Week 3&4: Daniel Stegmueller
Advanced Bayesian Models for the Social Sciences Instructors: Week 1&2: Skyler J. Cranmer Department of Political Science University of North Carolina, Chapel Hill skyler@unc.edu Week 3&4: Daniel Stegmueller
Michael Hallquist, Thomas M. Olino, Paul A. Pilkonis University of Pittsburgh
 Comparing the evidence for categorical versus dimensional representations of psychiatric disorders in the presence of noisy observations: a Monte Carlo study of the Bayesian Information Criterion and Akaike
Comparing the evidence for categorical versus dimensional representations of psychiatric disorders in the presence of noisy observations: a Monte Carlo study of the Bayesian Information Criterion and Akaike
MISSING DATA AND PARAMETERS ESTIMATES IN MULTIDIMENSIONAL ITEM RESPONSE MODELS. Federico Andreis, Pier Alda Ferrari *
 Electronic Journal of Applied Statistical Analysis EJASA (2012), Electron. J. App. Stat. Anal., Vol. 5, Issue 3, 431 437 e-issn 2070-5948, DOI 10.1285/i20705948v5n3p431 2012 Università del Salento http://siba-ese.unile.it/index.php/ejasa/index
Electronic Journal of Applied Statistical Analysis EJASA (2012), Electron. J. App. Stat. Anal., Vol. 5, Issue 3, 431 437 e-issn 2070-5948, DOI 10.1285/i20705948v5n3p431 2012 Università del Salento http://siba-ese.unile.it/index.php/ejasa/index
Regulatory T Cells Exhibit Distinct Features in Human Breast Cancer
 Resource Regulatory T Cells Exhibit Distinct Features in Human Breast Cancer Graphical Abstract Authors George Plitas, Catherine Konopacki, Kenmin Wu,..., Ekaterina V. Putintseva, Dmitriy M. Chudakov,
Resource Regulatory T Cells Exhibit Distinct Features in Human Breast Cancer Graphical Abstract Authors George Plitas, Catherine Konopacki, Kenmin Wu,..., Ekaterina V. Putintseva, Dmitriy M. Chudakov,
DeconRNASeq: A Statistical Framework for Deconvolution of Heterogeneous Tissue Samples Based on mrna-seq data
 DeconRNASeq: A Statistical Framework for Deconvolution of Heterogeneous Tissue Samples Based on mrna-seq data Ting Gong, Joseph D. Szustakowski April 30, 2018 1 Introduction Heterogeneous tissues are frequently
DeconRNASeq: A Statistical Framework for Deconvolution of Heterogeneous Tissue Samples Based on mrna-seq data Ting Gong, Joseph D. Szustakowski April 30, 2018 1 Introduction Heterogeneous tissues are frequently
Cellecta Overview. Started Operations in 2007 Headquarters: Mountain View, CA
 Cellecta Overview Started Operations in 2007 Headquarters: Mountain View, CA Focus: Development of flexible, scalable, and broadly parallel genetic screening assays to expedite the discovery and characterization
Cellecta Overview Started Operations in 2007 Headquarters: Mountain View, CA Focus: Development of flexible, scalable, and broadly parallel genetic screening assays to expedite the discovery and characterization
Supplemental Information. Aryl Hydrocarbon Receptor Controls. Monocyte Differentiation. into Dendritic Cells versus Macrophages
 Immunity, Volume 47 Supplemental Information Aryl Hydrocarbon Receptor Controls Monocyte Differentiation into Dendritic Cells versus Macrophages Christel Goudot, Alice Coillard, Alexandra-Chloé Villani,
Immunity, Volume 47 Supplemental Information Aryl Hydrocarbon Receptor Controls Monocyte Differentiation into Dendritic Cells versus Macrophages Christel Goudot, Alice Coillard, Alexandra-Chloé Villani,
Chapter 1. Introduction
 Chapter 1 Introduction 1.1 Motivation and Goals The increasing availability and decreasing cost of high-throughput (HT) technologies coupled with the availability of computational tools and data form a
Chapter 1 Introduction 1.1 Motivation and Goals The increasing availability and decreasing cost of high-throughput (HT) technologies coupled with the availability of computational tools and data form a
EXPression ANalyzer and DisplayER
 EXPression ANalyzer and DisplayER Tom Hait Aviv Steiner Igor Ulitsky Chaim Linhart Amos Tanay Seagull Shavit Rani Elkon Adi Maron-Katz Dorit Sagir Eyal David Roded Sharan Israel Steinfeld Yossi Shiloh
EXPression ANalyzer and DisplayER Tom Hait Aviv Steiner Igor Ulitsky Chaim Linhart Amos Tanay Seagull Shavit Rani Elkon Adi Maron-Katz Dorit Sagir Eyal David Roded Sharan Israel Steinfeld Yossi Shiloh
Ch.20 Dynamic Cue Combination in Distributional Population Code Networks. Ka Yeon Kim Biopsychology
 Ch.20 Dynamic Cue Combination in Distributional Population Code Networks Ka Yeon Kim Biopsychology Applying the coding scheme to dynamic cue combination (Experiment, Kording&Wolpert,2004) Dynamic sensorymotor
Ch.20 Dynamic Cue Combination in Distributional Population Code Networks Ka Yeon Kim Biopsychology Applying the coding scheme to dynamic cue combination (Experiment, Kording&Wolpert,2004) Dynamic sensorymotor
RNA-seq Introduction
 RNA-seq Introduction DNA is the same in all cells but which RNAs that is present is different in all cells There is a wide variety of different functional RNAs Which RNAs (and sometimes then translated
RNA-seq Introduction DNA is the same in all cells but which RNAs that is present is different in all cells There is a wide variety of different functional RNAs Which RNAs (and sometimes then translated
Comparison of discrimination methods for the classification of tumors using gene expression data
 Comparison of discrimination methods for the classification of tumors using gene expression data Sandrine Dudoit, Jane Fridlyand 2 and Terry Speed 2,. Mathematical Sciences Research Institute, Berkeley
Comparison of discrimination methods for the classification of tumors using gene expression data Sandrine Dudoit, Jane Fridlyand 2 and Terry Speed 2,. Mathematical Sciences Research Institute, Berkeley
Bayesian Estimation of a Meta-analysis model using Gibbs sampler
 University of Wollongong Research Online Applied Statistics Education and Research Collaboration (ASEARC) - Conference Papers Faculty of Engineering and Information Sciences 2012 Bayesian Estimation of
University of Wollongong Research Online Applied Statistics Education and Research Collaboration (ASEARC) - Conference Papers Faculty of Engineering and Information Sciences 2012 Bayesian Estimation of
Supplementary Table 1: List of parameters used in this study. Parameter Unit Parameter Unit Clinical parameters CD4 T cells Microbial translocation
 Supplementary Table 1: List of parameters used in this study. Left table shows parameters available for all subjects and right table shows parameters available for a smaller subset of subjects. Parameters
Supplementary Table 1: List of parameters used in this study. Left table shows parameters available for all subjects and right table shows parameters available for a smaller subset of subjects. Parameters
MODEL-BASED CLUSTERING IN GENE EXPRESSION MICROARRAYS: AN APPLICATION TO BREAST CANCER DATA
 International Journal of Software Engineering and Knowledge Engineering Vol. 13, No. 6 (2003) 579 592 c World Scientific Publishing Company MODEL-BASED CLUSTERING IN GENE EXPRESSION MICROARRAYS: AN APPLICATION
International Journal of Software Engineering and Knowledge Engineering Vol. 13, No. 6 (2003) 579 592 c World Scientific Publishing Company MODEL-BASED CLUSTERING IN GENE EXPRESSION MICROARRAYS: AN APPLICATION
Supplemental Information. Genomic Characterization of Murine. Monocytes Reveals C/EBPb Transcription. Factor Dependence of Ly6C Cells
 Immunity, Volume 46 Supplemental Information Genomic Characterization of Murine Monocytes Reveals C/EBPb Transcription Factor Dependence of Ly6C Cells Alexander Mildner, Jörg Schönheit, Amir Giladi, Eyal
Immunity, Volume 46 Supplemental Information Genomic Characterization of Murine Monocytes Reveals C/EBPb Transcription Factor Dependence of Ly6C Cells Alexander Mildner, Jörg Schönheit, Amir Giladi, Eyal
TRIPODS Workshop: Models & Machine Learning for Causal I. & Decision Making
 TRIPODS Workshop: Models & Machine Learning for Causal Inference & Decision Making in Medical Decision Making : and Predictive Accuracy text Stavroula Chrysanthopoulou, PhD Department of Biostatistics
TRIPODS Workshop: Models & Machine Learning for Causal Inference & Decision Making in Medical Decision Making : and Predictive Accuracy text Stavroula Chrysanthopoulou, PhD Department of Biostatistics
Welcome. Nanostring Immuno-Oncology Summit. September 21st, FOR RESEARCH USE ONLY. Not for use in diagnostic procedures.
 Welcome Nanostring Immuno-Oncology Summit September 21st, 2017 1 FOR RESEARCH USE ONLY. Not for use in diagnostic procedures. FOR RESEARCH USE ONLY. Not for use in diagnostic procedures. Agenda 4:00-4:30
Welcome Nanostring Immuno-Oncology Summit September 21st, 2017 1 FOR RESEARCH USE ONLY. Not for use in diagnostic procedures. FOR RESEARCH USE ONLY. Not for use in diagnostic procedures. Agenda 4:00-4:30
Introduction to Discrimination in Microarray Data Analysis
 Introduction to Discrimination in Microarray Data Analysis Jane Fridlyand CBMB University of California, San Francisco Genentech Hall Auditorium, Mission Bay, UCSF October 23, 2004 1 Case Study: Van t
Introduction to Discrimination in Microarray Data Analysis Jane Fridlyand CBMB University of California, San Francisco Genentech Hall Auditorium, Mission Bay, UCSF October 23, 2004 1 Case Study: Van t
A COMPARISON OF IMPUTATION METHODS FOR MISSING DATA IN A MULTI-CENTER RANDOMIZED CLINICAL TRIAL: THE IMPACT STUDY
 A COMPARISON OF IMPUTATION METHODS FOR MISSING DATA IN A MULTI-CENTER RANDOMIZED CLINICAL TRIAL: THE IMPACT STUDY Lingqi Tang 1, Thomas R. Belin 2, and Juwon Song 2 1 Center for Health Services Research,
A COMPARISON OF IMPUTATION METHODS FOR MISSING DATA IN A MULTI-CENTER RANDOMIZED CLINICAL TRIAL: THE IMPACT STUDY Lingqi Tang 1, Thomas R. Belin 2, and Juwon Song 2 1 Center for Health Services Research,
Type 1 Diabetes: Islet expressing GAD65 (green) with DAPI (Blue) Islet expressing Insulin (red) in 3D confocal imaging
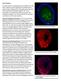 Type 1 Diabetes: Our group has been studying autoimmune diabetes for many years. Recently, we have developed a humanized mouse model of Type 1 Diabetes (T1D). We believe this model will help understand
Type 1 Diabetes: Our group has been studying autoimmune diabetes for many years. Recently, we have developed a humanized mouse model of Type 1 Diabetes (T1D). We believe this model will help understand
The 16th KJC Bioinformatics Symposium Integrative analysis identifies potential DNA methylation biomarkers for pan-cancer diagnosis and prognosis
 The 16th KJC Bioinformatics Symposium Integrative analysis identifies potential DNA methylation biomarkers for pan-cancer diagnosis and prognosis Tieliu Shi tlshi@bio.ecnu.edu.cn The Center for bioinformatics
The 16th KJC Bioinformatics Symposium Integrative analysis identifies potential DNA methylation biomarkers for pan-cancer diagnosis and prognosis Tieliu Shi tlshi@bio.ecnu.edu.cn The Center for bioinformatics
Supplementary Materials Extracting a Cellular Hierarchy from High-dimensional Cytometry Data with SPADE
 Supplementary Materials Extracting a Cellular Hierarchy from High-dimensional Cytometry Data with SPADE Peng Qiu1,4, Erin F. Simonds2, Sean C. Bendall2, Kenneth D. Gibbs Jr.2, Robert V. Bruggner2, Michael
Supplementary Materials Extracting a Cellular Hierarchy from High-dimensional Cytometry Data with SPADE Peng Qiu1,4, Erin F. Simonds2, Sean C. Bendall2, Kenneth D. Gibbs Jr.2, Robert V. Bruggner2, Michael
Modelling Spatially Correlated Survival Data for Individuals with Multiple Cancers
 Modelling Spatially Correlated Survival Data for Individuals with Multiple Cancers Dipak K. Dey, Ulysses Diva and Sudipto Banerjee Department of Statistics University of Connecticut, Storrs. March 16,
Modelling Spatially Correlated Survival Data for Individuals with Multiple Cancers Dipak K. Dey, Ulysses Diva and Sudipto Banerjee Department of Statistics University of Connecticut, Storrs. March 16,
Lecture 13: Finding optimal treatment policies
 MACHINE LEARNING FOR HEALTHCARE 6.S897, HST.S53 Lecture 13: Finding optimal treatment policies Prof. David Sontag MIT EECS, CSAIL, IMES (Thanks to Peter Bodik for slides on reinforcement learning) Outline
MACHINE LEARNING FOR HEALTHCARE 6.S897, HST.S53 Lecture 13: Finding optimal treatment policies Prof. David Sontag MIT EECS, CSAIL, IMES (Thanks to Peter Bodik for slides on reinforcement learning) Outline
Simultaneous Measurement Imputation and Outcome Prediction for Achilles Tendon Rupture Rehabilitation
 Simultaneous Measurement Imputation and Outcome Prediction for Achilles Tendon Rupture Rehabilitation Charles Hamesse 1, Paul Ackermann 2, Hedvig Kjellström 1, and Cheng Zhang 3 1 KTH Royal Institute of
Simultaneous Measurement Imputation and Outcome Prediction for Achilles Tendon Rupture Rehabilitation Charles Hamesse 1, Paul Ackermann 2, Hedvig Kjellström 1, and Cheng Zhang 3 1 KTH Royal Institute of
How many people do you know?: Efficiently estimating personal network size
 How many people do you know?: Efficiently estimating personal network size Tian Zheng Department of Statistics Columbia University April 22nd, 2009 1 / 34 Acknowledgements Collaborators Tyler McCormick
How many people do you know?: Efficiently estimating personal network size Tian Zheng Department of Statistics Columbia University April 22nd, 2009 1 / 34 Acknowledgements Collaborators Tyler McCormick
4. Model evaluation & selection
 Foundations of Machine Learning CentraleSupélec Fall 2017 4. Model evaluation & selection Chloé-Agathe Azencot Centre for Computational Biology, Mines ParisTech chloe-agathe.azencott@mines-paristech.fr
Foundations of Machine Learning CentraleSupélec Fall 2017 4. Model evaluation & selection Chloé-Agathe Azencot Centre for Computational Biology, Mines ParisTech chloe-agathe.azencott@mines-paristech.fr
Practical Bayesian Optimization of Machine Learning Algorithms. Jasper Snoek, Ryan Adams, Hugo LaRochelle NIPS 2012
 Practical Bayesian Optimization of Machine Learning Algorithms Jasper Snoek, Ryan Adams, Hugo LaRochelle NIPS 2012 ... (Gaussian Processes) are inadequate for doing speech and vision. I still think they're
Practical Bayesian Optimization of Machine Learning Algorithms Jasper Snoek, Ryan Adams, Hugo LaRochelle NIPS 2012 ... (Gaussian Processes) are inadequate for doing speech and vision. I still think they're
Biostatistical modelling in genomics for clinical cancer studies
 This work was supported by Entente Cordiale Cancer Research Bursaries Biostatistical modelling in genomics for clinical cancer studies Philippe Broët JE 2492 Faculté de Médecine Paris-Sud In collaboration
This work was supported by Entente Cordiale Cancer Research Bursaries Biostatistical modelling in genomics for clinical cancer studies Philippe Broët JE 2492 Faculté de Médecine Paris-Sud In collaboration
High-Throughput Sequencing Course
 High-Throughput Sequencing Course Introduction Biostatistics and Bioinformatics Summer 2017 From Raw Unaligned Reads To Aligned Reads To Counts Differential Expression Differential Expression 3 2 1 0 1
High-Throughput Sequencing Course Introduction Biostatistics and Bioinformatics Summer 2017 From Raw Unaligned Reads To Aligned Reads To Counts Differential Expression Differential Expression 3 2 1 0 1
Inferring Biological Meaning from Cap Analysis Gene Expression Data
 Inferring Biological Meaning from Cap Analysis Gene Expression Data HRYSOULA PAPADAKIS 1. Introduction This project is inspired by the recent development of the Cap analysis gene expression (CAGE) method,
Inferring Biological Meaning from Cap Analysis Gene Expression Data HRYSOULA PAPADAKIS 1. Introduction This project is inspired by the recent development of the Cap analysis gene expression (CAGE) method,
The effects of Cell Subset Isolation Method on Gene Expression inleukocytes
 Nadejda Beliakova-Bethell, Marta Massanella et al The effects of Cell Subset Isolation Method on Gene Expression inleukocytes The effects of Cell Subset Isolation Method on Gene Expression inleukocytes
Nadejda Beliakova-Bethell, Marta Massanella et al The effects of Cell Subset Isolation Method on Gene Expression inleukocytes The effects of Cell Subset Isolation Method on Gene Expression inleukocytes
CSE Introduction to High-Perfomance Deep Learning ImageNet & VGG. Jihyung Kil
 CSE 5194.01 - Introduction to High-Perfomance Deep Learning ImageNet & VGG Jihyung Kil ImageNet Classification with Deep Convolutional Neural Networks Alex Krizhevsky, Ilya Sutskever, Geoffrey E. Hinton,
CSE 5194.01 - Introduction to High-Perfomance Deep Learning ImageNet & VGG Jihyung Kil ImageNet Classification with Deep Convolutional Neural Networks Alex Krizhevsky, Ilya Sutskever, Geoffrey E. Hinton,
Monocyte subsets in health and disease. Marion Frankenberger
 Monocyte subsets in health and disease Marion Frankenberger main cellular components: Leukocytes Erythrocytes Composition of whole blood Monocytes belong to the cellular components of peripheral blood
Monocyte subsets in health and disease Marion Frankenberger main cellular components: Leukocytes Erythrocytes Composition of whole blood Monocytes belong to the cellular components of peripheral blood
7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans.
 Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
Nature Neuroscience: doi: /nn Supplementary Figure 1. Behavioral training.
 Supplementary Figure 1 Behavioral training. a, Mazes used for behavioral training. Asterisks indicate reward location. Only some example mazes are shown (for example, right choice and not left choice maze
Supplementary Figure 1 Behavioral training. a, Mazes used for behavioral training. Asterisks indicate reward location. Only some example mazes are shown (for example, right choice and not left choice maze
Supplemental Information. Regulatory T Cells Exhibit Distinct. Features in Human Breast Cancer
 Immunity, Volume 45 Supplemental Information Regulatory T Cells Exhibit Distinct Features in Human Breast Cancer George Plitas, Catherine Konopacki, Kenmin Wu, Paula D. Bos, Monica Morrow, Ekaterina V.
Immunity, Volume 45 Supplemental Information Regulatory T Cells Exhibit Distinct Features in Human Breast Cancer George Plitas, Catherine Konopacki, Kenmin Wu, Paula D. Bos, Monica Morrow, Ekaterina V.
ANALYTISCHE STRATEGIE Tissue Imaging. Bernd Bodenmiller Institute of Molecular Life Sciences University of Zurich
 ANALYTISCHE STRATEGIE Tissue Imaging Bernd Bodenmiller Institute of Molecular Life Sciences University of Zurich Quantitative Breast single cancer cell analysis Switzerland Brain Breast Lung Colon-rectum
ANALYTISCHE STRATEGIE Tissue Imaging Bernd Bodenmiller Institute of Molecular Life Sciences University of Zurich Quantitative Breast single cancer cell analysis Switzerland Brain Breast Lung Colon-rectum
MATS: a Bayesian framework for flexible detection of differential alternative splicing from RNA-Seq data
 Nucleic Acids Research Advance Access published February 1, 2012 Nucleic Acids Research, 2012, 1 13 doi:10.1093/nar/gkr1291 MATS: a Bayesian framework for flexible detection of differential alternative
Nucleic Acids Research Advance Access published February 1, 2012 Nucleic Acids Research, 2012, 1 13 doi:10.1093/nar/gkr1291 MATS: a Bayesian framework for flexible detection of differential alternative
SUPPLEMENTARY APPENDIX
 SUPPLEMENTARY APPENDIX 1) Supplemental Figure 1. Histopathologic Characteristics of the Tumors in the Discovery Cohort 2) Supplemental Figure 2. Incorporation of Normal Epidermal Melanocytic Signature
SUPPLEMENTARY APPENDIX 1) Supplemental Figure 1. Histopathologic Characteristics of the Tumors in the Discovery Cohort 2) Supplemental Figure 2. Incorporation of Normal Epidermal Melanocytic Signature
An Introduction to Multiple Imputation for Missing Items in Complex Surveys
 An Introduction to Multiple Imputation for Missing Items in Complex Surveys October 17, 2014 Joe Schafer Center for Statistical Research and Methodology (CSRM) United States Census Bureau Views expressed
An Introduction to Multiple Imputation for Missing Items in Complex Surveys October 17, 2014 Joe Schafer Center for Statistical Research and Methodology (CSRM) United States Census Bureau Views expressed
An application of a pattern-mixture model with multiple imputation for the analysis of longitudinal trials with protocol deviations
 Iddrisu and Gumedze BMC Medical Research Methodology (2019) 19:10 https://doi.org/10.1186/s12874-018-0639-y RESEARCH ARTICLE Open Access An application of a pattern-mixture model with multiple imputation
Iddrisu and Gumedze BMC Medical Research Methodology (2019) 19:10 https://doi.org/10.1186/s12874-018-0639-y RESEARCH ARTICLE Open Access An application of a pattern-mixture model with multiple imputation
Introduction to Machine Learning. Katherine Heller Deep Learning Summer School 2018
 Introduction to Machine Learning Katherine Heller Deep Learning Summer School 2018 Outline Kinds of machine learning Linear regression Regularization Bayesian methods Logistic Regression Why we do this
Introduction to Machine Learning Katherine Heller Deep Learning Summer School 2018 Outline Kinds of machine learning Linear regression Regularization Bayesian methods Logistic Regression Why we do this
Supplemental Information. A Highly Sensitive and Robust Method. for Genome-wide 5hmC Profiling. of Rare Cell Populations
 Molecular ell, Volume 63 Supplemental Information Highly Sensitive and Robust Method for enome-wide hm Profiling of Rare ell Populations Dali Han, Xingyu Lu, lan H. Shih, Ji Nie, Qiancheng You, Meng Michelle
Molecular ell, Volume 63 Supplemental Information Highly Sensitive and Robust Method for enome-wide hm Profiling of Rare ell Populations Dali Han, Xingyu Lu, lan H. Shih, Ji Nie, Qiancheng You, Meng Michelle
Statistical Audit. Summary. Conceptual and. framework. MICHAELA SAISANA and ANDREA SALTELLI European Commission Joint Research Centre (Ispra, Italy)
 Statistical Audit MICHAELA SAISANA and ANDREA SALTELLI European Commission Joint Research Centre (Ispra, Italy) Summary The JRC analysis suggests that the conceptualized multi-level structure of the 2012
Statistical Audit MICHAELA SAISANA and ANDREA SALTELLI European Commission Joint Research Centre (Ispra, Italy) Summary The JRC analysis suggests that the conceptualized multi-level structure of the 2012
Effector T Cells and
 1 Effector T Cells and Cytokines Andrew Lichtman, MD PhD Brigham and Women's Hospital Harvard Medical School 2 Lecture outline Cytokines Subsets of CD4+ T cells: definitions, functions, development New
1 Effector T Cells and Cytokines Andrew Lichtman, MD PhD Brigham and Women's Hospital Harvard Medical School 2 Lecture outline Cytokines Subsets of CD4+ T cells: definitions, functions, development New
Supplementary Methods
 Supplementary Methods Analysis of time course gene expression data. The time course data of the expression level of a representative gene is shown in the below figure. The trajectory of longitudinal expression
Supplementary Methods Analysis of time course gene expression data. The time course data of the expression level of a representative gene is shown in the below figure. The trajectory of longitudinal expression
Nature Getetics: doi: /ng.3471
 Supplementary Figure 1 Summary of exome sequencing data. ( a ) Exome tumor normal sample sizes for bladder cancer (BLCA), breast cancer (BRCA), carcinoid (CARC), chronic lymphocytic leukemia (CLLX), colorectal
Supplementary Figure 1 Summary of exome sequencing data. ( a ) Exome tumor normal sample sizes for bladder cancer (BLCA), breast cancer (BRCA), carcinoid (CARC), chronic lymphocytic leukemia (CLLX), colorectal
Lecture 21. RNA-seq: Advanced analysis
 Lecture 21 RNA-seq: Advanced analysis Experimental design Introduction An experiment is a process or study that results in the collection of data. Statistical experiments are conducted in situations in
Lecture 21 RNA-seq: Advanced analysis Experimental design Introduction An experiment is a process or study that results in the collection of data. Statistical experiments are conducted in situations in
Application Note # MT-111 Concise Interpretation of MALDI Imaging Data by Probabilistic Latent Semantic Analysis (plsa)
 Application Note # MT-111 Concise Interpretation of MALDI Imaging Data by Probabilistic Latent Semantic Analysis (plsa) MALDI imaging datasets can be very information-rich, containing hundreds of mass
Application Note # MT-111 Concise Interpretation of MALDI Imaging Data by Probabilistic Latent Semantic Analysis (plsa) MALDI imaging datasets can be very information-rich, containing hundreds of mass
4. Th1-related gene expression in infected versus mock-infected controls from Fig. 2 with gene annotation.
 List of supplemental information 1. Graph of mouse weight loss during course of infection- Line graphs showing mouse weight data during course of infection days 1 to 10 post-infections (p.i.). 2. Graph
List of supplemental information 1. Graph of mouse weight loss during course of infection- Line graphs showing mouse weight data during course of infection days 1 to 10 post-infections (p.i.). 2. Graph
Introduction to Computational Neuroscience
 Introduction to Computational Neuroscience Lecture 10: Brain-Computer Interfaces Ilya Kuzovkin So Far Stimulus So Far So Far Stimulus What are the neuroimaging techniques you know about? Stimulus So Far
Introduction to Computational Neuroscience Lecture 10: Brain-Computer Interfaces Ilya Kuzovkin So Far Stimulus So Far So Far Stimulus What are the neuroimaging techniques you know about? Stimulus So Far
Bayesian Bi-Cluster Change-Point Model for Exploring Functional Brain Dynamics
 Int'l Conf. Bioinformatics and Computational Biology BIOCOMP'18 85 Bayesian Bi-Cluster Change-Point Model for Exploring Functional Brain Dynamics Bing Liu 1*, Xuan Guo 2, and Jing Zhang 1** 1 Department
Int'l Conf. Bioinformatics and Computational Biology BIOCOMP'18 85 Bayesian Bi-Cluster Change-Point Model for Exploring Functional Brain Dynamics Bing Liu 1*, Xuan Guo 2, and Jing Zhang 1** 1 Department
NEW METHODS FOR SENSITIVITY TESTS OF EXPLOSIVE DEVICES
 NEW METHODS FOR SENSITIVITY TESTS OF EXPLOSIVE DEVICES Amit Teller 1, David M. Steinberg 2, Lina Teper 1, Rotem Rozenblum 2, Liran Mendel 2, and Mordechai Jaeger 2 1 RAFAEL, POB 2250, Haifa, 3102102, Israel
NEW METHODS FOR SENSITIVITY TESTS OF EXPLOSIVE DEVICES Amit Teller 1, David M. Steinberg 2, Lina Teper 1, Rotem Rozenblum 2, Liran Mendel 2, and Mordechai Jaeger 2 1 RAFAEL, POB 2250, Haifa, 3102102, Israel
Manipulating the Tumor Environment
 Manipulating the Tumor Environment Vincenzo Bronte Verona University Hospital vincenzo.bronte@univr.it Escape from immune control can be viewed as one of the «Hallmarks of Cancer» D. Hanahan and R. A.
Manipulating the Tumor Environment Vincenzo Bronte Verona University Hospital vincenzo.bronte@univr.it Escape from immune control can be viewed as one of the «Hallmarks of Cancer» D. Hanahan and R. A.
Supplementary Figure 1: High-throughput profiling of survival after exposure to - radiation. (a) Cells were plated in at least 7 wells in a 384-well
 Supplementary Figure 1: High-throughput profiling of survival after exposure to - radiation. (a) Cells were plated in at least 7 wells in a 384-well plate at cell densities ranging from 25-225 cells in
Supplementary Figure 1: High-throughput profiling of survival after exposure to - radiation. (a) Cells were plated in at least 7 wells in a 384-well plate at cell densities ranging from 25-225 cells in
Mediation Analysis With Principal Stratification
 University of Pennsylvania ScholarlyCommons Statistics Papers Wharton Faculty Research 3-30-009 Mediation Analysis With Principal Stratification Robert Gallop Dylan S. Small University of Pennsylvania
University of Pennsylvania ScholarlyCommons Statistics Papers Wharton Faculty Research 3-30-009 Mediation Analysis With Principal Stratification Robert Gallop Dylan S. Small University of Pennsylvania
Pathology of Inflammatory Breast Cancer (IBC) A rare tumor
 Pathology of Inflammatory Breast Cancer (IBC) A rare tumor Jelle Wesseling 1, John Martens 2, Gabe Sonke 1, Carolien Schröder 3 1: Netherlands Cancer Institute Antoni van Leeuwenhoek Hospital, Amsterdam
Pathology of Inflammatory Breast Cancer (IBC) A rare tumor Jelle Wesseling 1, John Martens 2, Gabe Sonke 1, Carolien Schröder 3 1: Netherlands Cancer Institute Antoni van Leeuwenhoek Hospital, Amsterdam
Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes.
 Supplementary Figure 1 Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes. (a,b) Values of coefficients associated with genomic features, separately
Supplementary Figure 1 Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes. (a,b) Values of coefficients associated with genomic features, separately
Cross-Study Projections of Genomic Biomarkers: An Evaluation in Cancer Genomics
 of Genomic Biomarkers: An Evaluation in Cancer Genomics Joseph E. Lucas 1 *, Carlos M. Carvalho 2, Julia Ling-Yu Chen 1, Jen-Tsan Chi 1, Mike West 3 1 Institute for Genome Sciences and Policy, Duke University,
of Genomic Biomarkers: An Evaluation in Cancer Genomics Joseph E. Lucas 1 *, Carlos M. Carvalho 2, Julia Ling-Yu Chen 1, Jen-Tsan Chi 1, Mike West 3 1 Institute for Genome Sciences and Policy, Duke University,
Data analysis and binary regression for predictive discrimination. using DNA microarray data. (Breast cancer) discrimination. Expression array data
 West Mike of Statistics & Decision Sciences Institute Duke University wwwstatdukeedu IPAM Functional Genomics Workshop November Two group problems: Binary outcomes ffl eg, ER+ versus ER ffl eg, lymph node
West Mike of Statistics & Decision Sciences Institute Duke University wwwstatdukeedu IPAM Functional Genomics Workshop November Two group problems: Binary outcomes ffl eg, ER+ versus ER ffl eg, lymph node
DNA vaccine, peripheral T-cell tolerance modulation 185
 Subject Index Airway hyperresponsiveness (AHR) animal models 41 43 asthma inhibition 45 overview 41 mast cell modulation of T-cells 62 64 respiratory tolerance 40, 41 Tregs inhibition role 44 respiratory
Subject Index Airway hyperresponsiveness (AHR) animal models 41 43 asthma inhibition 45 overview 41 mast cell modulation of T-cells 62 64 respiratory tolerance 40, 41 Tregs inhibition role 44 respiratory
Biomarker Identification in Breast Cancer: Beta-Adrenergic Receptor Signaling and Pathways to Therapeutic Response
 , http://dx.doi.org/10.5936/csbj.201303003 CSBJ Biomarker Identification in Breast Cancer: Beta-Adrenergic Receptor Signaling and Pathways to Therapeutic Response Liana E. Kafetzopoulou a, David J. Boocock
, http://dx.doi.org/10.5936/csbj.201303003 CSBJ Biomarker Identification in Breast Cancer: Beta-Adrenergic Receptor Signaling and Pathways to Therapeutic Response Liana E. Kafetzopoulou a, David J. Boocock
ChIP-seq hands-on. Iros Barozzi, Campus IFOM-IEO (Milan) Saverio Minucci, Gioacchino Natoli Labs
 ChIP-seq hands-on Iros Barozzi, Campus IFOM-IEO (Milan) Saverio Minucci, Gioacchino Natoli Labs Main goals Becoming familiar with essential tools and formats Visualizing and contextualizing raw data Understand
ChIP-seq hands-on Iros Barozzi, Campus IFOM-IEO (Milan) Saverio Minucci, Gioacchino Natoli Labs Main goals Becoming familiar with essential tools and formats Visualizing and contextualizing raw data Understand
How Systems Biology Can Advance Immune Oncology. Birgit Schoeberl, PhD NEDMG Summer Symposium May 31st
 How Systems Biology Can Advance Immune Oncology Birgit Schoeberl, PhD NEDMG Summer Symposium May 31st Immuno-Oncology is One of the Most Promising Avenues in the Battle Against Cancer Conventional Therapy
How Systems Biology Can Advance Immune Oncology Birgit Schoeberl, PhD NEDMG Summer Symposium May 31st Immuno-Oncology is One of the Most Promising Avenues in the Battle Against Cancer Conventional Therapy
Lecture 11: Clustering to discover disease subtypes and stages
 MACHINE LEARNING FOR HEALTHCARE 6.S897, HST.S53 Lecture 11: Clustering to discover disease subtypes and stages Prof. David Sontag MIT EECS, CSAIL, IMES Outline of today s class 1. Overview of clustering
MACHINE LEARNING FOR HEALTHCARE 6.S897, HST.S53 Lecture 11: Clustering to discover disease subtypes and stages Prof. David Sontag MIT EECS, CSAIL, IMES Outline of today s class 1. Overview of clustering
Bayesian Inference Bayes Laplace
 Bayesian Inference Bayes Laplace Course objective The aim of this course is to introduce the modern approach to Bayesian statistics, emphasizing the computational aspects and the differences between the
Bayesian Inference Bayes Laplace Course objective The aim of this course is to introduce the modern approach to Bayesian statistics, emphasizing the computational aspects and the differences between the
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers
 Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
Sawtooth Software. MaxDiff Analysis: Simple Counting, Individual-Level Logit, and HB RESEARCH PAPER SERIES. Bryan Orme, Sawtooth Software, Inc.
 Sawtooth Software RESEARCH PAPER SERIES MaxDiff Analysis: Simple Counting, Individual-Level Logit, and HB Bryan Orme, Sawtooth Software, Inc. Copyright 009, Sawtooth Software, Inc. 530 W. Fir St. Sequim,
Sawtooth Software RESEARCH PAPER SERIES MaxDiff Analysis: Simple Counting, Individual-Level Logit, and HB Bryan Orme, Sawtooth Software, Inc. Copyright 009, Sawtooth Software, Inc. 530 W. Fir St. Sequim,
Digitizing the Proteomes From Big Tissue Biobanks
 Digitizing the Proteomes From Big Tissue Biobanks Analyzing 24 Proteomes Per Day by Microflow SWATH Acquisition and Spectronaut Pulsar Analysis Jan Muntel 1, Nick Morrice 2, Roland M. Bruderer 1, Lukas
Digitizing the Proteomes From Big Tissue Biobanks Analyzing 24 Proteomes Per Day by Microflow SWATH Acquisition and Spectronaut Pulsar Analysis Jan Muntel 1, Nick Morrice 2, Roland M. Bruderer 1, Lukas
International Society of Breast Pathology. Immune Targeting in Breast Cancer. USCAP 2017 Annual Meeting
 International Society of Breast Pathology USCAP 2017 Annual Meeting Immune Targeting in Breast Cancer Ashley Cimino-Mathews, MD Assistant Professor of Pathology and Oncology The Johns Hopkins Hospital
International Society of Breast Pathology USCAP 2017 Annual Meeting Immune Targeting in Breast Cancer Ashley Cimino-Mathews, MD Assistant Professor of Pathology and Oncology The Johns Hopkins Hospital
CyTOF workflow: differential discovery in high-throughput high-dimensional cytometry datasets!
 Workshop!! CyTOF workflow: differential discovery in high-throughput high-dimensional cytometry datasets! Malgorzata Nowicka! University of Zurich! BioC 2017: Where Software and Biology Connect! Boston,
Workshop!! CyTOF workflow: differential discovery in high-throughput high-dimensional cytometry datasets! Malgorzata Nowicka! University of Zurich! BioC 2017: Where Software and Biology Connect! Boston,
Methodology for the VoicesDMV Survey
 M E T R O P O L I T A N H O U S I N G A N D C O M M U N I T I E S P O L I C Y C E N T E R Methodology for the VoicesDMV Survey Timothy Triplett December 2017 Voices of the Community: DC, Maryland, Virginia
M E T R O P O L I T A N H O U S I N G A N D C O M M U N I T I E S P O L I C Y C E N T E R Methodology for the VoicesDMV Survey Timothy Triplett December 2017 Voices of the Community: DC, Maryland, Virginia
ImmunoTools special Award 2014
 ImmunoTools special Award 2014 Davide Ferrari, PhD University of Ferrara, Department of Morphology, Surgery and Experimental Medicine. Section of Pathology, Oncology and Exper. Biology via L. Borsari 46,
ImmunoTools special Award 2014 Davide Ferrari, PhD University of Ferrara, Department of Morphology, Surgery and Experimental Medicine. Section of Pathology, Oncology and Exper. Biology via L. Borsari 46,
Imputation classes as a framework for inferences from non-random samples. 1
 Imputation classes as a framework for inferences from non-random samples. 1 Vladislav Beresovsky (hvy4@cdc.gov) National Center for Health Statistics, CDC 1 Disclaimer: The findings and conclusions in
Imputation classes as a framework for inferences from non-random samples. 1 Vladislav Beresovsky (hvy4@cdc.gov) National Center for Health Statistics, CDC 1 Disclaimer: The findings and conclusions in
Bayesian Joint Modelling of Benefit and Risk in Drug Development
 Bayesian Joint Modelling of Benefit and Risk in Drug Development EFSPI/PSDM Safety Statistics Meeting Leiden 2017 Disclosure is an employee and shareholder of GSK Data presented is based on human research
Bayesian Joint Modelling of Benefit and Risk in Drug Development EFSPI/PSDM Safety Statistics Meeting Leiden 2017 Disclosure is an employee and shareholder of GSK Data presented is based on human research
TITLE: A Data-Driven Approach to Patient Risk Stratification for Acute Respiratory Distress Syndrome (ARDS)
 TITLE: A Data-Driven Approach to Patient Risk Stratification for Acute Respiratory Distress Syndrome (ARDS) AUTHORS: Tejas Prahlad INTRODUCTION Acute Respiratory Distress Syndrome (ARDS) is a condition
TITLE: A Data-Driven Approach to Patient Risk Stratification for Acute Respiratory Distress Syndrome (ARDS) AUTHORS: Tejas Prahlad INTRODUCTION Acute Respiratory Distress Syndrome (ARDS) is a condition
Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor suppressor genes
 Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor suppressor genes Kaifu Chen 1,2,3,4,5,10, Zhong Chen 6,10, Dayong Wu 6, Lili Zhang 7, Xueqiu Lin 1,2,8,
Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor suppressor genes Kaifu Chen 1,2,3,4,5,10, Zhong Chen 6,10, Dayong Wu 6, Lili Zhang 7, Xueqiu Lin 1,2,8,
Comparative Study of K-means, Gaussian Mixture Model, Fuzzy C-means algorithms for Brain Tumor Segmentation
 Comparative Study of K-means, Gaussian Mixture Model, Fuzzy C-means algorithms for Brain Tumor Segmentation U. Baid 1, S. Talbar 2 and S. Talbar 1 1 Department of E&TC Engineering, Shri Guru Gobind Singhji
Comparative Study of K-means, Gaussian Mixture Model, Fuzzy C-means algorithms for Brain Tumor Segmentation U. Baid 1, S. Talbar 2 and S. Talbar 1 1 Department of E&TC Engineering, Shri Guru Gobind Singhji
Introduction to Computational Neuroscience
 Introduction to Computational Neuroscience Lecture 5: Data analysis II Lesson Title 1 Introduction 2 Structure and Function of the NS 3 Windows to the Brain 4 Data analysis 5 Data analysis II 6 Single
Introduction to Computational Neuroscience Lecture 5: Data analysis II Lesson Title 1 Introduction 2 Structure and Function of the NS 3 Windows to the Brain 4 Data analysis 5 Data analysis II 6 Single
Rise of the Machines
 Rise of the Machines Statistical machine learning for observational studies: confounding adjustment and subgroup identification Armand Chouzy, ETH (summer intern) Jason Wang, Celgene PSI conference 2018
Rise of the Machines Statistical machine learning for observational studies: confounding adjustment and subgroup identification Armand Chouzy, ETH (summer intern) Jason Wang, Celgene PSI conference 2018
