Proteins: Proteomics & Protein-Protein Interactions Part I
|
|
|
- Rolf Rose
- 6 years ago
- Views:
Transcription
1 Proteins: Proteomics & Protein-Protein Interactions Part I Jesse Rinehart, PhD Department of Cellular & Molecular Physiology Systems Biology Institute
2 DNA RNA PROTEIN
3 DNA RNA PROTEIN
4 Proteins: Proteomics & Protein-Protein Interactions Overview Techniques & Technologies - Mass Spectrometry - Protein-Protein Interactions - Genetic & Biochemical Strategies - Protein Purification - Quantitative Proteomics Applications - Representative Studies Putting it all together. - Databases & Pathways
5 Proteins: Proteomics & Protein-Protein Interactions Overview Techniques & Technologies - Mass Spectrometry - Protein-Protein Interactions - Genetic & Biochemical Strategies - Protein Purification - Quantitative Proteomics Applications - Representative Studies Putting it all together. - Databases & Pathways
6 Principles of Mass Spectrometry (MS) In a mass spectrum we measure m/z (mass-to-charge) For proteins we measure peptide m/z A sample has to be ionizable in order to be analyzed
7 Basic Components of a Mass Spectrometer
8 Two major ionization techniques enabled the success of mass spectrometry in the life sciences. - Electrospray Ionization (ESI) Fenn, J.B. et al, Science, 1989, 246, Matrix Assisted Laser Desorption Ionization (MALDI) Karas, M.; Hillenkamp, F., Anal. Chem., 1995, 60, 2299 MS based Proteomics is born: - MS to measure weight of large intact proteins - Non-covalently bonded protein complexes can also be measured (ESI only) - Intact peptides measured and sequenced
9 Electrospray Ionization (ESI) F Surface Tension Electrospray Fan T drying gas Sample Electrospray Needle Liquid Filament Rayleigh Instability Breakup Sample Matrix Assisted Laser Desorption Ionization (MALDI)
10 Mass Spectrometry takes the 2002 Nobel Prize in Chemistry Awarded to John B. Fenn & Koichi Tanaka *Fenn - Discovered Electrospray Ionization (ESI) *Pioneering work at Yale University in the Department of Chemical Engineering Tanaka - Discovered Matrix Assisted Laser Desorption Ionization (MALDI)
11 Basic Mass Spectrometer Typical LC-MS Setup Time of Flight (TOF) Mass Analyzers Quadrupole Orbitrap
12 Typical work flow for LC-MS shotgun proteomics Peptide Pool Protein mixture Enz. Digest LC n-uplc LTQ-Orbitrap MS MS isolate & fragment MS/MS peptide peptide peptide peptide fragments
13 Typical work flow for LC-MS shotgun proteomics Protein mixture Enz. Digest Peptide Pool LC n-uplc Trypsin cuts after Lys (K ) & Arg (R) LTQ-Orbitrap MS Proteins and Protein Structure (Branden, C. and Tooze, J. Introduction to Protein Structure)
14 Trypsin digest followed by LC-MS: Examples of Sequence Coverage Band 3 Anion Transporter β-actin
15 Protein mixture Enz. Digest Peptide Pool LC n-uplc LTQ-Orbitrap MS Peptide ions have a mass (m) and a charge (z). 100 Da peptide: +1 = 100 m/z +2 = 50 m/z +3 = 33.3 m/z
16 Protein mixture Enz. Digest Peptide Pool LC n-uplc LTQ-Orbitrap MS Peptide ions are isolated and sequenced
17 Protein mixture Enz. Digest Peptide Pool LC n-uplc LTQ-Orbitrap MS Computational Steps: massive amounts of MS data are read & interpreted. Databases searched to match peptide sequences.
18 Proteomics The study of the expression, location, interaction, function, and structure of all the proteins in a given cell, organelle, tissue, organ, or whole organism. [Study of post-translational modifications (protein phosphorylation, acetylation, glycosylation ) via MS has grown in recent years to dramatically expand the field of Proteomics]
19 A tour of proteomics: Studies with the budding yeast Saccharomyces cerevisiae 2000 & 2001 Uetz et al, A comprehensive analysis of protein-protein interactions in Saccharomyces cerevisiae. Nature. & Ito et al, A comprehensive two-hybrid analysis to explore the yeast protein interactome. PNAS. Large scale yeast two hybrid screens to map proteome wide interactions Washburn, et al. Large-scale analysis of the yeast proteome by multidimensional protein identification technology. Nature Biotechnol. Established the shotgun technology by showing that many proteins in a yeast-cell lysate could be identified in a single experiment Ho, Y. et al. Systematic identification of protein complexes in Saccharomyces cerevisiae by mass spectrometry. Nature. & Gavin, A. C. et al. Functional organization of the yeast proteome by systematic analysis of protein complexes. Nature. Protein protein interaction maps can be obtained by MS; the yeast cell is organized into protein complexes Ghaemmaghami, S. et al. Global analysis of protein expression in yeast. Nature. & Huh, W. K. et al. Global analysis of protein localization in budding yeast. Nature. TAP-Tag and expression studies & GFP-Tag and localization studies 2006 Krogan NJ, et al. Global landscape of protein complexes in the yeast Saccharomyces cerevisiae. Nature. TAP-Tag and Protein-Protein Interaction 2008 de Godoy LM, et al. Comprehensive mass-spectrometry-based proteome quantification of haploid versus diploid yeast. Nature. SILAC based quantitation of an entire proteome Picotti P, et al. Full dynamic range proteome analysis of S. cerevisiae by targeted proteomics. Cell. Towards proteome wide targeted proteomics.
20 The *pace of proteomics is set by a combination of techniques and technological advances. *orders of magnitude behind genomics and transcriptomics Yeast proteome reported in Washburn et al. Nature Biotech 2001: ~82 hours* = 1,484 proteins *estimates from paper: 3 15 X 110 minute runs for each fraction Yeast proteome by Hunter, Colangelo, Rinehart, et al unpublished 2010: One 60 minute run = 1,286 proteins 58,857 MS/MS AB SCIEX TripleTOF 5600
21 Proteins: Proteomics & Protein-Protein Interactions Overview Techniques & Technologies - Mass Spectrometry - Protein-Protein Interactions - Genetic & Biochemical Strategies - Protein Purification - Quantitative Proteomics Applications - Representative Studies Putting it all together. - Databases & Pathways
22 A comprehensive analysis of protein-protein interactions in Saccharomyces cerevisiae. Uetz et al, Nature 2000 Ito et al, PNAS 2001 Yeast Two Hybrid Assay Clone bait and prey constructs and place in separate strains. Mat a Mat Mate a +
23 Uetz et al, Nature 2000 Ito et al, PNAS 2001
24 A comprehensive analysis of protein-protein interactions in Saccharomyces cerevisiae. Uetz et al, Nature 2000 Ito et al, PNAS 2001 Yeast Two Hybrid Assay Advantages: - In vivo assay - Simple Some Disadvantages - Hard to execute on large scale - False positives: a real interaction or possible interaction - Interaction in nucleus (required for GAL system) - Clones are fusion proteins and sometimes partial proteins - Multiple protein complexes not captured
25 Human Two Hybrid Map 8,100 ORFs (~7,200 genes) 10,597 interactions
26
27 Protein-Protein interaction maps: Proteins are represented by nodes and interactions are represented by edges between nodes. node edge Human Interactome in 2010 (~100,000) Bonetta, Nature 2010
28 Protein-Protein interactions: Some examples: - Physical and direct - Physical and indirect - Multi-protein complexes - Scaffolds - Transient Kinase - Kinase & substrate - Metabolic Substrate Enz A Enz B
29 A tour of proteomics: Studies with the budding yeast Saccharomyces cerevisiae 2000 & 2001 Uetz et al, A comprehensive analysis of protein-protein interactions in Saccharomyces cerevisiae. Nature. & Ito et al, A comprehensive two-hybrid analysis to explore the yeast protein interactome. PNAS. Large scale yeast two hybrid screens to map proteome wide interactions Washburn, et al. Large-scale analysis of the yeast proteome by multidimensional protein identification technology. Nature Biotechnol. Established the shotgun technology by showing that many proteins in a yeast-cell lysate could be identified in a single experiment Ho, Y. et al. Systematic identification of protein complexes in Saccharomyces cerevisiae by mass spectrometry. Nature. & Gavin, A. C. et al. Functional organization of the yeast proteome by systematic analysis of protein complexes. Nature. Protein protein interaction maps can be obtained by MS; the yeast cell is organized into protein complexes Ghaemmaghami, S. et al. Global analysis of protein expression in yeast. Nature. & Huh, W. K. et al. Global analysis of protein localization in budding yeast. Nature. TAP-Tag and expression studies & GFP-Tag and localization studies 2006 Krogan NJ, et al. Global landscape of protein complexes in the yeast Saccharomyces cerevisiae. Nature. TAP-Tag and Protein-Protein Interaction 2008 de Godoy LM, et al. Comprehensive mass-spectrometry-based proteome quantification of haploid versus diploid yeast. Nature. SILAC based quantitation of an entire proteome Picotti P, et al. Full dynamic range proteome analysis of S. cerevisiae by targeted proteomics. Cell. Towards proteome wide targeted proteomics.
30 Bait Molecular Handles Cannavo E et al. J. Biol. Chem. 2007
31 Global TAP Tagging in yeast 2003 Ghaemmaghami, S. et al. Global analysis of protein expression in yeast. Nature. & Huh, W. K. et al. Global analysis of protein localization in budding yeast. Nature. TAP-Tag and expression studies & GFP-Tag and localization studies
32 RT: Time (min) TIC MS OT TIC MS OT TIC MS OT Ho, Y. et al. Systematic identification of protein complexes in Saccharomyces cerevisiae by mass spectrometry. Nature. & Gavin, A. C. et al. Functional organization of the yeast proteome by systematic analysis of protein complexes. Nature. Protein protein interaction maps can be obtained by MS; the yeast cell is organized into protein complexes Krogan NJ, et al. Global landscape of protein complexes in the yeast Saccharomyces cerevisiae. Nature. TAP-Tag and Protein-Protein Interaction Collection of tagged bait expression strains TAP bait + Interacting proteins Multiple runs of shot gun MS & SDS-PAGE with MS on individual proteins NL: 7.64E9 TIC MS OT NL: 6.59E9 TIC MS OT NL: 6.74E9 NL: 6.76E9 NL: 7.37E9 TIC MS OT NL: 5.71E9 TIC MS OT NL: 5.71E9 TIC MS OT NL: 5.66E9 Krogan et al. observed 7,123 protein protein interactions: Important aspects: - Tagged the native genes and did not overexpress the fusion proteins - Could immediately validate partners (reciprocal purification in data set) - Complementary MS techniques, deeper coverage of complexes - Authors state, rigorous computational procedures to assign confidence values to our predictions
33 Interconnected complexes 4,562 tagged proteins 2,357 successful purifications Identified 4,087 interacting proteins ~72 % proteome Majority of the yeast proteome is organized into complexes Many complexes are conserved in other species Complexes with little or no interconnectivity Krogan NJ, et al. Nature. 2006
34 Particle Autophagy 2005 Pearson Prentice Hall, Inc. Pearson Prentice Hall, Inc
35 RT: Time (min) TIC MS OT TIC MS OT TIC MS OT Transfect tagged bait IP Bait + Interacting proteins Multiple runs of shot gun LC-MS/MS NL: 7.64E9 TIC MS OT NL: 6.59E9 TIC MS OT NL: 6.74E9 NL: 6.76E9 NL: 7.37E9 TIC MS OT NL: 5.71E9 TIC MS OT NL: 5.71E9 TIC MS OT NL: 5.66E9 ~65 bait proteins LC-MS/MS identifies 2553 proteins Autophagy Interaction Network Data analysis to sort out real interaction from background Authors use CompPASS to identify High-Confidence Interacting Proteins (HCIP) 763 HCIPs identified that compose The Autophagy Interaction Network Behreands et al, Nature 2010
36 Concatenating Data: Supplementing observed protein complexes with the Protein-Protein Interaction Databases: MINT, BioGRID, STRING Behreands et al, Nature 2010
37 A Functional Organization of the Network Phenotype after sirna Less autophagosomes More autophagosomes Behreands et al, Nature 2010
38 Protein-Protein Interaction Databases ,555 interactions + 45,777 proteins
39 Protein interaction networks: Some of the many important aspects: - Parts List - Organization and assembly - Biological function can be inferred However: - Interaction data is largely static Next Step: - How do protein interaction networks change over time?
40
41 Proteins: Proteomics & Protein-Protein Interactions Overview Techniques & Technologies - Mass Spectrometry - Protein-Protein Interactions - Genetic & Biochemical Strategies - Protein Purification - Quantitative Proteomics Applications - Representative Studies Putting it all together. - Databases & Pathways
42 Protein mixture Enz. Digest Peptide Pool LC n-uplc LTQ-Orbitrap MS Computational Steps: massive amounts of MS data are read & interpreted. Databases searched to match peptide sequences.
43 Typical work flow for LC-MS shotgun proteomics Peptide Pool Protein mixture Enz. Digest LC n-uplc LTQ-Orbitrap MS MS isolate & fragment MS/MS peptide peptide peptide peptide fragments
44 Typical work flow for LC-MRM targeted proteomics Peptide Pool LC Protein mixture Enz. Digest n-uplc QQQ MS Multiple Reaction Monitoring (MRM) MRM Spectra Detect Fragment Select Peptide Fragment peptide Select Fragment
45 MS Data is not inherently quantitative, but Rinehart et al., unpublished Red Blood Cell RBC membrane: a native multi-protein complex RBC membrane proteome Coomassie Stained SDS-PAGE (250 ug Protein) ~16 bands Spectrin Spectrin β # peptides (unique) RBC membrane proteome Shotgun Proteomics 1ug Peptides (242 Proteins) Band 3 Band 4.1 β-actin
46 Quantitative Proteomics S.I.L.A.C. - Stable isotope labeling with amino acids in cell culture -Ong S.E. et al. Molecular & Cell Proteomics 2002 Stable isotopes are not radioactive, and they occur naturally in nature. For example, 99% of all carbon in the world is carbon-12 ( 12 C) and 1% is carbon-13 ( 13 C). SILAC reagents have enriched stable isotopes that have been placed into compounds in abundances much greater than their natural abundance. We can obtain labeled compounds with ~95-99% 13 C. Because a mass spectrometer separates ions by mass, we use mass spectrometry to distinguish isotopes in compounds by their mass. Simultaneous comparison in the same MS run is key
47 A tour of proteomics: Studies with the budding yeast Saccharomyces cerevisiae 2000 & 2001 Uetz et al, A comprehensive analysis of protein-protein interactions in Saccharomyces cerevisiae. Nature. & Ito et al, A comprehensive two-hybrid analysis to explore the yeast protein interactome. PNAS. Large scale yeast two hybrid screens to map proteome wide interactions Washburn, et al. Large-scale analysis of the yeast proteome by multidimensional protein identification technology. Nature Biotechnol. Established the shotgun technology by showing that many proteins in a yeast-cell lysate could be identified in a single experiment Ho, Y. et al. Systematic identification of protein complexes in Saccharomyces cerevisiae by mass spectrometry. Nature. & Gavin, A. C. et al. Functional organization of the yeast proteome by systematic analysis of protein complexes. Nature. Protein protein interaction maps can be obtained by MS; the yeast cell is organized into protein complexes Ghaemmaghami, S. et al. Global analysis of protein expression in yeast. Nature. & Huh, W. K. et al. Global analysis of protein localization in budding yeast. Nature. TAP-Tag and expression studies & GFP-Tag and localization studies 2006 Krogan NJ, et al. Global landscape of protein complexes in the yeast Saccharomyces cerevisiae. Nature. TAP-Tag and Protein-Protein Interaction 2008 de Godoy LM, et al. Comprehensive mass-spectrometry-based proteome quantification of haploid versus diploid yeast. Nature. SILAC based quantitation of an entire proteome Picotti P, et al. Full dynamic range proteome analysis of S. cerevisiae by targeted proteomics. Cell. Towards proteome wide targeted proteomics.
48 2008 de Godoy LM, et al. Comprehensive mass-spectrometry-based proteome quantification of haploid versus diploid yeast. Nature. SILAC based quantitation of an entire proteome. S.I.L.A.C. - Stable isotope labeling with amino acids in cell culture -Ong SE et al. Molecular & Cell Proteomics Picotti P, et al. Full dynamic range proteome analysis of S. cerevisiae by targeted proteomics. Cell. Towards proteome wide targeted proteomics. Multiple Reaction Monitoring (MRM) Synthetic peptide Select Peptide Fragment peptide Select Fragment
49 2008 de Godoy LM, et al. Comprehensive mass-spectrometry-based proteome quantification of haploid versus diploid yeast. Nature. 30;455(7217): SILAC based quantitation of an entire proteome Picotti P, et al. Full dynamic range proteome analysis of S. cerevisiae by targeted proteomics. Cell. Towards proteome wide targeted proteomics.
50 2008 de Godoy LM, et al. Comprehensive mass-spectrometry-based proteome quantification of haploid versus diploid yeast. Nature. SILAC based quantitation of an entire proteome. Pheromone signaling is required for mating of haploid cells and is absent from diploid cells Picotti P, et al. Full dynamic range proteome analysis of S. cerevisiae by targeted proteomics. Cell. Towards proteome wide targeted proteomics. Network expression dynamics
51 Disomic Chr. V WT
52 Protein Regulators? Readers Tail Lysine 4 Histone 3 Vermeulen et al., Cell 2010
53 TFIID Active Genes Vermeulen et al., Cell 2010
54 Vermeulen et al., Cell 2010
55 A SILAC approach to study protein phosphorylation dynamics
56 Major technological advances in mass spectrometers and phosphopeptide enrichment Protein mixture TiO 2 Enrichment Flow Through Phosphopeptides Enriched
57 * Phosphopeptide signatures in MS Phosphopeptide P MS * P +80 Da in precursor isolate & fragment P P MS/MS * -98 Da loss of phosphoric acid H 3 PO 4 during fragmentation
58 V H VHMTWTK m/z M T W T K V H M T W T VHMT P WTK m/z * * * * * * * P * +80 Da in precursor - H 3 PO 4, 98 Da K (Threonine changes to 2-aminodehydrobutyric acid, -18 Da)
59 Cell swelling ON! De-phosphorylation activates KCC3 P SILAC Adapted from Rinehart, et al. Cell, 2009
60 Quantitative Proteomics Reveals Dynamics in Signaling Networks Phosphorylation dynamics after EGF stimulation SILAC approach enables dynamic analysis wikimedia.org MS spectra triplets Olsen, et al. Cell, 2006
61 Phosphorylation dynamics after EGF stimulation wikimedia.org Olsen, et al. Cell, 2006
62 Proteins: Proteomics & Protein-Protein Interactions Overview Techniques & Technologies - Mass Spectrometry - Protein-Protein Interactions - Genetic & Biochemical Strategies - Protein Purification - Quantitative Proteomics Applications - Representative Studies Putting it all together. - Databases & Pathways
Introduction to Proteomics 1.0
 Introduction to Proteomics 1.0 CMSP Workshop Pratik Jagtap Managing Director, CMSP Objectives Why are we here? For participants: Learn basics of MS-based proteomics Learn what s necessary for success using
Introduction to Proteomics 1.0 CMSP Workshop Pratik Jagtap Managing Director, CMSP Objectives Why are we here? For participants: Learn basics of MS-based proteomics Learn what s necessary for success using
Proteomics/Peptidomics
 Proteomics/Peptidomics System biology tools and preclinical models for translational research in endometriosis, ESHRE Campus workshop, 4-5 September 2009 E. Waelkens Proteomics: What? Proteins Proteomics
Proteomics/Peptidomics System biology tools and preclinical models for translational research in endometriosis, ESHRE Campus workshop, 4-5 September 2009 E. Waelkens Proteomics: What? Proteins Proteomics
Biological Mass spectrometry in Protein Chemistry
 Biological Mass spectrometry in Protein Chemistry Tuula Nyman Institute of Biotechnology tuula.nyman@helsinki.fi MASS SPECTROMETRY is an analytical technique that identifies the chemical composition of
Biological Mass spectrometry in Protein Chemistry Tuula Nyman Institute of Biotechnology tuula.nyman@helsinki.fi MASS SPECTROMETRY is an analytical technique that identifies the chemical composition of
Quantification with Proteome Discoverer. Bernard Delanghe
 Quantification with Proteome Discoverer Bernard Delanghe Overview: Which approach to use? Proteome Discoverer Quantification Method What When to use Metabolic labeling SILAC Cell culture systems Small
Quantification with Proteome Discoverer Bernard Delanghe Overview: Which approach to use? Proteome Discoverer Quantification Method What When to use Metabolic labeling SILAC Cell culture systems Small
Mass Spectrometry. Mass spectrometer MALDI-TOF ESI/MS/MS. Basic components. Ionization source Mass analyzer Detector
 Mass Spectrometry MALDI-TOF ESI/MS/MS Mass spectrometer Basic components Ionization source Mass analyzer Detector 1 Principles of Mass Spectrometry Proteins are separated by mass to charge ratio (limit
Mass Spectrometry MALDI-TOF ESI/MS/MS Mass spectrometer Basic components Ionization source Mass analyzer Detector 1 Principles of Mass Spectrometry Proteins are separated by mass to charge ratio (limit
Lecture 3. Tandem MS & Protein Sequencing
 Lecture 3 Tandem MS & Protein Sequencing Nancy Allbritton, M.D., Ph.D. Department of Physiology & Biophysics 824-9137 (office) nlallbri@uci.edu Office- Rm D349 Medical Science D Bldg. Tandem MS Steps:
Lecture 3 Tandem MS & Protein Sequencing Nancy Allbritton, M.D., Ph.D. Department of Physiology & Biophysics 824-9137 (office) nlallbri@uci.edu Office- Rm D349 Medical Science D Bldg. Tandem MS Steps:
New Instruments and Services
 New Instruments and Services Liwen Zhang Mass Spectrometry and Proteomics Facility The Ohio State University Summer Workshop 2016 Thermo Orbitrap Fusion http://planetorbitrap.com/orbitrap fusion Thermo
New Instruments and Services Liwen Zhang Mass Spectrometry and Proteomics Facility The Ohio State University Summer Workshop 2016 Thermo Orbitrap Fusion http://planetorbitrap.com/orbitrap fusion Thermo
New Instruments and Services
 New Instruments and Services http://planetorbitrap.com/orbitrap fusion Combining the best of quadrupole, Orbitrap, and ion trap mass analysis in a revolutionary Tribrid architecture, the Orbitrap Fusion
New Instruments and Services http://planetorbitrap.com/orbitrap fusion Combining the best of quadrupole, Orbitrap, and ion trap mass analysis in a revolutionary Tribrid architecture, the Orbitrap Fusion
MS/MS to Targeted Proteomics (MRM)
 MS/MS to Targeted Proteomics (MRM) How it worked on the Human Lens Proteome Jayson Falkner PhD jay@singleorganism.com Genes Show Limited Value in Predicting Diseases With only a few exceptions, what the
MS/MS to Targeted Proteomics (MRM) How it worked on the Human Lens Proteome Jayson Falkner PhD jay@singleorganism.com Genes Show Limited Value in Predicting Diseases With only a few exceptions, what the
REDOX PROTEOMICS. Roman Zubarev.
 REDOX PROTEOMICS Roman Zubarev Roman.Zubarev@ki.se Physiological Chemistry I, Department for Medical Biochemistry & Biophysics, Karolinska Institutet, Stockholm What is (RedOx) Proteomics? Proteomics -
REDOX PROTEOMICS Roman Zubarev Roman.Zubarev@ki.se Physiological Chemistry I, Department for Medical Biochemistry & Biophysics, Karolinska Institutet, Stockholm What is (RedOx) Proteomics? Proteomics -
Advances in Hybrid Mass Spectrometry
 The world leader in serving science Advances in Hybrid Mass Spectrometry ESAC 2008 Claire Dauly Field Marketing Specialist, Proteomics New hybrids instruments LTQ Orbitrap XL with ETD MALDI LTQ Orbitrap
The world leader in serving science Advances in Hybrid Mass Spectrometry ESAC 2008 Claire Dauly Field Marketing Specialist, Proteomics New hybrids instruments LTQ Orbitrap XL with ETD MALDI LTQ Orbitrap
PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System
 Application Note LC/MS PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System Purpose This application note describes an automated workflow
Application Note LC/MS PTM Discovery Method for Automated Identification and Sequencing of Phosphopeptides Using the Q TRAP LC/MS/MS System Purpose This application note describes an automated workflow
Learning Objectives. Overview of topics to be discussed 10/25/2013 HIGH RESOLUTION MASS SPECTROMETRY (HRMS) IN DISCOVERY PROTEOMICS
 HIGH RESOLUTION MASS SPECTROMETRY (HRMS) IN DISCOVERY PROTEOMICS A clinical proteomics perspective Michael L. Merchant, PhD School of Medicine, University of Louisville Louisville, KY Learning Objectives
HIGH RESOLUTION MASS SPECTROMETRY (HRMS) IN DISCOVERY PROTEOMICS A clinical proteomics perspective Michael L. Merchant, PhD School of Medicine, University of Louisville Louisville, KY Learning Objectives
Don t miss a thing on your peptide mapping journey How to get full coverage peptide maps using high resolution accurate mass spectrometry
 Don t miss a thing on your peptide mapping journey How to get full coverage peptide maps using high resolution accurate mass spectrometry Kai Scheffler, PhD BioPharma Support Expert,LSMS Europe The world
Don t miss a thing on your peptide mapping journey How to get full coverage peptide maps using high resolution accurate mass spectrometry Kai Scheffler, PhD BioPharma Support Expert,LSMS Europe The world
Comparison of mass spectrometers performances
 Comparison of mass spectrometers performances Instrument Mass Mass Sensitivity resolution accuracy Quadrupole 1 x 10 3 0.1 Da* 0.5-1.0 pmol DE-MALDI 2 x 10 4 20 ppm 1-10 fmol peptide 1-5 pmol protein Ion
Comparison of mass spectrometers performances Instrument Mass Mass Sensitivity resolution accuracy Quadrupole 1 x 10 3 0.1 Da* 0.5-1.0 pmol DE-MALDI 2 x 10 4 20 ppm 1-10 fmol peptide 1-5 pmol protein Ion
One Gene, Many Proteins. Applications of Mass Spectrometry to Proteomics. Why Proteomics? Raghothama Chaerkady, Ph.D.
 Applications of Mass Spectrometry to Proteomics Raghothama Chaerkady, Ph.D. McKusick-Nathans Institute of Genetic Medicine and the Department of Biological Chemistry Why Proteomics? One Gene, Many Proteins
Applications of Mass Spectrometry to Proteomics Raghothama Chaerkady, Ph.D. McKusick-Nathans Institute of Genetic Medicine and the Department of Biological Chemistry Why Proteomics? One Gene, Many Proteins
2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry
 Dr. Sanjeeva Srivastava 1. Fundamental of Mass Spectrometry Role of MS and basic concepts 2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry 2 1 MS basic concepts Mass spectrometry - technique
Dr. Sanjeeva Srivastava 1. Fundamental of Mass Spectrometry Role of MS and basic concepts 2. Ionization Sources 3. Mass Analyzers 4. Tandem Mass Spectrometry 2 1 MS basic concepts Mass spectrometry - technique
Mass Spectrometry. - Introduction - Ion sources & sample introduction - Mass analyzers - Basics of biomolecule MS - Applications
 - Introduction - Ion sources & sample introduction - Mass analyzers - Basics of biomolecule MS - Applications Adapted from Mass Spectrometry in Biotechnology Gary Siuzdak,, Academic Press 1996 1 Introduction
- Introduction - Ion sources & sample introduction - Mass analyzers - Basics of biomolecule MS - Applications Adapted from Mass Spectrometry in Biotechnology Gary Siuzdak,, Academic Press 1996 1 Introduction
Proteome analysis Cellular proteomics Plasma proteomics
 Proteome analysis Cellular proteomics Plasma proteomics Janne Lehtiö Maria Pernemalm Karolinska Biomics Center Karolinska Institutet / Karolinska University Hospital Janne Lehtio 2009-02-09 1 KBC Proteomics
Proteome analysis Cellular proteomics Plasma proteomics Janne Lehtiö Maria Pernemalm Karolinska Biomics Center Karolinska Institutet / Karolinska University Hospital Janne Lehtio 2009-02-09 1 KBC Proteomics
Ion Source. Mass Analyzer. Detector. intensity. mass/charge
 Proteomics Informatics Overview of spectrometry (Week 2) Ion Source Analyzer Detector Peptide Fragmentation Ion Source Analyzer 1 Fragmentation Analyzer 2 Detector b y Liquid Chromatography (LC)-MS/MS
Proteomics Informatics Overview of spectrometry (Week 2) Ion Source Analyzer Detector Peptide Fragmentation Ion Source Analyzer 1 Fragmentation Analyzer 2 Detector b y Liquid Chromatography (LC)-MS/MS
Fundamentals of Soft Ionization and MS Instrumentation
 Fundamentals of Soft Ionization and MS Instrumentation Ana Varela Coelho varela@itqb.unl.pt Mass Spectrometry Lab Analytical Services Unit Index Mass spectrometers and its components Ionization methods:
Fundamentals of Soft Ionization and MS Instrumentation Ana Varela Coelho varela@itqb.unl.pt Mass Spectrometry Lab Analytical Services Unit Index Mass spectrometers and its components Ionization methods:
Mass Spectrometry Course Árpád Somogyi Chemistry and Biochemistry MassSpectrometry Facility) University of Debrecen, April 12-23, 2010
 Mass Spectrometry Course Árpád Somogyi Chemistry and Biochemistry MassSpectrometry Facility) University of Debrecen, April 12-23, 2010 Introduction, Ionization Methods Mass Analyzers, Ion Activation Methods
Mass Spectrometry Course Árpád Somogyi Chemistry and Biochemistry MassSpectrometry Facility) University of Debrecen, April 12-23, 2010 Introduction, Ionization Methods Mass Analyzers, Ion Activation Methods
Mass Spectrometry Infrastructure
 Mass Spectrometry Infrastructure Todd Williams, Ph.D. Director KU Mass Spectrometry and Analytical Proteomics Laboratory Mass Spectrometry Lab B025 Malott Hall Mission The Mass Spectrometry and analytical
Mass Spectrometry Infrastructure Todd Williams, Ph.D. Director KU Mass Spectrometry and Analytical Proteomics Laboratory Mass Spectrometry Lab B025 Malott Hall Mission The Mass Spectrometry and analytical
Protein Identification and Phosphorylation Site Determination by de novo sequencing using PepFrag TM MALDI-Sequencing kit
 Application Note Tel: +82-54-223-2463 Fax : +82-54-223-2460 http://www.genomine.com venture ldg 306 Pohang techno park Pohang, kyungbuk, Korea(ROK) Protein Identification and Phosphorylation Site Determination
Application Note Tel: +82-54-223-2463 Fax : +82-54-223-2460 http://www.genomine.com venture ldg 306 Pohang techno park Pohang, kyungbuk, Korea(ROK) Protein Identification and Phosphorylation Site Determination
Supporting Information. Lysine Propionylation to Boost Proteome Sequence. Coverage and Enable a Silent SILAC Strategy for
 Supporting Information Lysine Propionylation to Boost Proteome Sequence Coverage and Enable a Silent SILAC Strategy for Relative Protein Quantification Christoph U. Schräder 1, Shaun Moore 1,2, Aaron A.
Supporting Information Lysine Propionylation to Boost Proteome Sequence Coverage and Enable a Silent SILAC Strategy for Relative Protein Quantification Christoph U. Schräder 1, Shaun Moore 1,2, Aaron A.
Mass Spectrometry and Proteomics - Lecture 4 - Matthias Trost Newcastle University
 Mass Spectrometry and Proteomics - Lecture 4 - Matthias Trost Newcastle University matthias.trost@ncl.ac.uk previously Peptide fragmentation Hybrid instruments 117 The Building Blocks of Life DNA RNA Proteins
Mass Spectrometry and Proteomics - Lecture 4 - Matthias Trost Newcastle University matthias.trost@ncl.ac.uk previously Peptide fragmentation Hybrid instruments 117 The Building Blocks of Life DNA RNA Proteins
Metabolomics: quantifying the phenotype
 Metabolomics: quantifying the phenotype Metabolomics Promises Quantitative Phenotyping What can happen GENOME What appears to be happening Bioinformatics TRANSCRIPTOME What makes it happen PROTEOME Systems
Metabolomics: quantifying the phenotype Metabolomics Promises Quantitative Phenotyping What can happen GENOME What appears to be happening Bioinformatics TRANSCRIPTOME What makes it happen PROTEOME Systems
1. Sample Introduction to MS Systems:
 MS Overview: 9.10.08 1. Sample Introduction to MS Systems:...2 1.1. Chromatography Interfaces:...3 1.2. Electron impact: Used mainly in Protein MS hard ionization source...4 1.3. Electrospray Ioniztion:
MS Overview: 9.10.08 1. Sample Introduction to MS Systems:...2 1.1. Chromatography Interfaces:...3 1.2. Electron impact: Used mainly in Protein MS hard ionization source...4 1.3. Electrospray Ioniztion:
Double charge of 33kD peak A1 A2 B1 B2 M2+ M/z. ABRF Proteomics Research Group - Qualitative Proteomics Study Identifier Number 14146
 Abstract The 2008 ABRF Proteomics Research Group Study offers participants the chance to participate in an anonymous study to identify qualitative differences between two protein preparations. We used
Abstract The 2008 ABRF Proteomics Research Group Study offers participants the chance to participate in an anonymous study to identify qualitative differences between two protein preparations. We used
Shotgun Proteomics MS/MS. Protein Mixture. proteolysis. Peptide Mixture. Time. Abundance. Abundance. m/z. Abundance. m/z 2. Abundance.
 Abundance Abundance Abundance Abundance Abundance Shotgun Proteomics Protein Mixture 1 2 3 MS/MS proteolysis m/z 2 3 Time µlc m/z MS 1 m/z Peptide Mixture m/z Block Diagram of a Mass Spectrometer Sample
Abundance Abundance Abundance Abundance Abundance Shotgun Proteomics Protein Mixture 1 2 3 MS/MS proteolysis m/z 2 3 Time µlc m/z MS 1 m/z Peptide Mixture m/z Block Diagram of a Mass Spectrometer Sample
Nature Methods: doi: /nmeth.3177
 Supplementary Figure 1 Characterization of LysargiNase, trypsin and LysN missed cleavages. (a) Proportion of peptides identified in LysargiNase and trypsin digests of MDA-MB-231 cell lysates carrying 0,
Supplementary Figure 1 Characterization of LysargiNase, trypsin and LysN missed cleavages. (a) Proportion of peptides identified in LysargiNase and trypsin digests of MDA-MB-231 cell lysates carrying 0,
LC/MS/MS SOLUTIONS FOR LIPIDOMICS. Biomarker and Omics Solutions FOR DISCOVERY AND TARGETED LIPIDOMICS
 LC/MS/MS SOLUTIONS FOR LIPIDOMICS Biomarker and Omics Solutions FOR DISCOVERY AND TARGETED LIPIDOMICS Lipids play a key role in many biological processes, such as the formation of cell membranes and signaling
LC/MS/MS SOLUTIONS FOR LIPIDOMICS Biomarker and Omics Solutions FOR DISCOVERY AND TARGETED LIPIDOMICS Lipids play a key role in many biological processes, such as the formation of cell membranes and signaling
Methods in Mass Spectrometry. Dr. Noam Tal Laboratory of Mass Spectrometry School of Chemistry, Tel Aviv University
 Methods in Mass Spectrometry Dr. Noam Tal Laboratory of Mass Spectrometry School of Chemistry, Tel Aviv University Sample Engineering Chemistry Biology Life Science Medicine Industry IDF / Police Sample
Methods in Mass Spectrometry Dr. Noam Tal Laboratory of Mass Spectrometry School of Chemistry, Tel Aviv University Sample Engineering Chemistry Biology Life Science Medicine Industry IDF / Police Sample
Biological Mass Spectrometry. April 30, 2014
 Biological Mass Spectrometry April 30, 2014 Mass Spectrometry Has become the method of choice for precise protein and nucleic acid mass determination in a very wide mass range peptide and nucleotide sequencing
Biological Mass Spectrometry April 30, 2014 Mass Spectrometry Has become the method of choice for precise protein and nucleic acid mass determination in a very wide mass range peptide and nucleotide sequencing
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/8/398/rs12/dc1 Supplementary Materials for Quantitative phosphoproteomics reveals new roles for the protein phosphatase PP6 in mitotic cells Scott F. Rusin, Kate
www.sciencesignaling.org/cgi/content/full/8/398/rs12/dc1 Supplementary Materials for Quantitative phosphoproteomics reveals new roles for the protein phosphatase PP6 in mitotic cells Scott F. Rusin, Kate
Protein Analysis using Electrospray Ionization Mass Spectroscopy *
 OpenStax-CNX module: m38341 1 Protein Analysis using Electrospray Ionization Mass Spectroscopy * Wilhelm Kienast Andrew R. Barron This work is produced by OpenStax-CNX and licensed under the Creative Commons
OpenStax-CNX module: m38341 1 Protein Analysis using Electrospray Ionization Mass Spectroscopy * Wilhelm Kienast Andrew R. Barron This work is produced by OpenStax-CNX and licensed under the Creative Commons
RDP Cores Highlights: the CF Analytics Core. Facundo M. Fernández School of Chemistry and Biochemistry Georgia Institute of Technology
 CF@LANTA RDP Cores Highlights: the CF Analytics Core Facundo M. Fernández School of Chemistry and Biochemistry Georgia Institute of Technology CF@LANTA RDP Center The CF@LANTA RDP Center at Emory University
CF@LANTA RDP Cores Highlights: the CF Analytics Core Facundo M. Fernández School of Chemistry and Biochemistry Georgia Institute of Technology CF@LANTA RDP Center The CF@LANTA RDP Center at Emory University
Multiplex Protein Quantitation using itraq Reagents in a Gel-Based Workflow
 Multiplex Protein Quantitation using itraq Reagents in a Gel-Based Workflow Purpose Described herein is a workflow that combines the isobaric tagging reagents, itraq Reagents, with the separation power
Multiplex Protein Quantitation using itraq Reagents in a Gel-Based Workflow Purpose Described herein is a workflow that combines the isobaric tagging reagents, itraq Reagents, with the separation power
New Developments in LC-IMS-MS Proteomic Measurements and Informatic Analyses
 New Developments in LC-IMS-MS Proteomic Measurements and Informatic Analyses Erin Shammel Baker Kristin E. Burnum-Johnson, Xing Zhang, Cameron P. Casey, Yehia M. Ibrahim, Matthew E. Monroe, Tao Liu, Brendan
New Developments in LC-IMS-MS Proteomic Measurements and Informatic Analyses Erin Shammel Baker Kristin E. Burnum-Johnson, Xing Zhang, Cameron P. Casey, Yehia M. Ibrahim, Matthew E. Monroe, Tao Liu, Brendan
Lab 1 - Klinisk Kemisk Diagnos9k
 Lab 1 - Klinisk Kemisk Diagnos9k Mass Spectrometry Quan%fica%on & Protein iden%fica%on by pep%de mass fingerprint melinda.rezeli@bme.lth.se 073-978 0298 Lab 1 - Klinisk Kemisk Diagnos9k MALDI mass spectrometry
Lab 1 - Klinisk Kemisk Diagnos9k Mass Spectrometry Quan%fica%on & Protein iden%fica%on by pep%de mass fingerprint melinda.rezeli@bme.lth.se 073-978 0298 Lab 1 - Klinisk Kemisk Diagnos9k MALDI mass spectrometry
MS/MS Scan Modes. Eötvös University, Budapest April 16, MS/MS Scan Modes. Árpád Somogyi. Product Ion Scan Select. Scan. Precursor Ion Scan Scan
 MS/MS Modes Árpád Somogyi Eötvös University, Budapest April 16, 2012 MS/MS Modes Product Ion Precursor Ion Neutral Loss Δ ed Reaction Monitoring (SRM) 1 modes in a triple quadrupole (QqQ) (one quadrupole
MS/MS Modes Árpád Somogyi Eötvös University, Budapest April 16, 2012 MS/MS Modes Product Ion Precursor Ion Neutral Loss Δ ed Reaction Monitoring (SRM) 1 modes in a triple quadrupole (QqQ) (one quadrupole
SCS Mass Spectrometry Laboratory
 SCS Mass Spectrometry Laboratory Contact Information Staff 31 Noyes Laboratory (8:00-5:00 M-F) 217-333-2545 http://scs.illinois.edu/massspec/ Furong Sun (frs@illinois.edu) Furong Sun Director Training
SCS Mass Spectrometry Laboratory Contact Information Staff 31 Noyes Laboratory (8:00-5:00 M-F) 217-333-2545 http://scs.illinois.edu/massspec/ Furong Sun (frs@illinois.edu) Furong Sun Director Training
NIH Public Access Author Manuscript J Proteome Res. Author manuscript; available in PMC 2014 July 05.
 NIH Public Access Author Manuscript Published in final edited form as: J Proteome Res. 2013 July 5; 12(7): 3071 3086. doi:10.1021/pr3011588. Evaluation and Optimization of Mass Spectrometric Settings during
NIH Public Access Author Manuscript Published in final edited form as: J Proteome Res. 2013 July 5; 12(7): 3071 3086. doi:10.1021/pr3011588. Evaluation and Optimization of Mass Spectrometric Settings during
Phosphorylation of proteins Steve Barnes Feb 19th, 2002 in some cases, proteins are found in a stable, hyperphosphorylated state, e.g.
 Phosphorylation of proteins Steve Barnes Feb 19th, 2002 in some cases, proteins are found in a stable, hyperphosphorylated state, e.g., casein more interestingly, in most other cases, it is a transient
Phosphorylation of proteins Steve Barnes Feb 19th, 2002 in some cases, proteins are found in a stable, hyperphosphorylated state, e.g., casein more interestingly, in most other cases, it is a transient
Flow-Through Electron Capture Dissociation in a novel Branched RF Ion Trap
 Flow-Through Electron Capture Dissociation in a novel Branched RF Ion Trap Takashi Baba, J. Larry Campbell, Yves Le Blanc, Jim. W. Hager and Bruce A. Thomson ASMS, June 18 / 2014 1 2014 AB SCIEX Trapping
Flow-Through Electron Capture Dissociation in a novel Branched RF Ion Trap Takashi Baba, J. Larry Campbell, Yves Le Blanc, Jim. W. Hager and Bruce A. Thomson ASMS, June 18 / 2014 1 2014 AB SCIEX Trapping
The Detection of Allergens in Food Products with LC-MS
 The Detection of Allergens in Food Products with LC-MS Something for the future? Jacqueline van der Wielen Scope of Organisation Dutch Food and Consumer Product Safety Authority: Law enforcement Control
The Detection of Allergens in Food Products with LC-MS Something for the future? Jacqueline van der Wielen Scope of Organisation Dutch Food and Consumer Product Safety Authority: Law enforcement Control
Multidimensional Tracking of GPCR Signaling with Proximity- Labeling. Technical Journal Club Anna Henzi,
 Multidimensional Tracking of GPCR Signaling with Proximity- Labeling Technical Journal Club Anna Henzi, 11.07.2017 Proximity-dependent labeling methods Enzymes create radicals Proximity-dependent biotin
Multidimensional Tracking of GPCR Signaling with Proximity- Labeling Technical Journal Club Anna Henzi, 11.07.2017 Proximity-dependent labeling methods Enzymes create radicals Proximity-dependent biotin
Mass spectra of peptides and proteins - and LC analysis of proteomes Stephen Barnes, PhD
 Mass spectra of peptides and proteins - and LC analysis of proteomes Stephen Barnes, PhD 4-7117 sbarnes@uab.edu Overview A mass spectrum Electrospray MS Analysis of intact proteins Molecular weight calculations
Mass spectra of peptides and proteins - and LC analysis of proteomes Stephen Barnes, PhD 4-7117 sbarnes@uab.edu Overview A mass spectrum Electrospray MS Analysis of intact proteins Molecular weight calculations
Nature Biotechnology: doi: /nbt Supplementary Figure 1
 Supplementary Figure 1 The timeline of the NGAG method for extraction of N-linked glycans and glycosite-containing peptides. The timeline can be changed based on the number of samples. Supplementary Figure
Supplementary Figure 1 The timeline of the NGAG method for extraction of N-linked glycans and glycosite-containing peptides. The timeline can be changed based on the number of samples. Supplementary Figure
Mass Spectrometry and Proteomics. Professor Xudong Yao Bioanalytical Chemistry Spring 2007
 Mass Spectrometry and Proteomics Professor Xudong Yao Bioanalytical Chemistry Spring 2007 Proteomics and -omics Roles of mass spectrometry Comparative proteomics Chemical proteomics Protein, Proteome and
Mass Spectrometry and Proteomics Professor Xudong Yao Bioanalytical Chemistry Spring 2007 Proteomics and -omics Roles of mass spectrometry Comparative proteomics Chemical proteomics Protein, Proteome and
MS1 and MS2 crosstalk in label free quantitation of mass spectrometry data independent acquisitions
 MS1 and MS2 crosstalk in label free quantitation of mass spectrometry data independent acquisitions MS1 528.18 +++ m/z 568.98 ++ m/z 678.34 ++ m/z MS2/SWATH June 9th, 2013 Matthew J. Rardin SIRT3 regulated
MS1 and MS2 crosstalk in label free quantitation of mass spectrometry data independent acquisitions MS1 528.18 +++ m/z 568.98 ++ m/z 678.34 ++ m/z MS2/SWATH June 9th, 2013 Matthew J. Rardin SIRT3 regulated
MALDI-TOF. Introduction. Schematic and Theory of MALDI
 MALDI-TOF Proteins and peptides have been characterized by high pressure liquid chromatography (HPLC) or SDS PAGE by generating peptide maps. These peptide maps have been used as fingerprints of protein
MALDI-TOF Proteins and peptides have been characterized by high pressure liquid chromatography (HPLC) or SDS PAGE by generating peptide maps. These peptide maps have been used as fingerprints of protein
Ionization Methods. Neutral species Charged species. Removal/addition of electron(s) Removal/addition of proton(s)
 Ionization Methods Neutral species Charged species Removal/addition of electron(s) M + e - (M +. )* + 2e - electron ionization Removal/addition of proton(s) M + (Matrix)-H MH + + (Matrix) - chemical ionization
Ionization Methods Neutral species Charged species Removal/addition of electron(s) M + e - (M +. )* + 2e - electron ionization Removal/addition of proton(s) M + (Matrix)-H MH + + (Matrix) - chemical ionization
Development of a Human Cell-Free Expression System to Generate Stable-Isotope-Labeled Protein Standards for Quantitative Mass Spectrometry
 Development of a Human Cell-Free Expression System to Generate Stable-Isotope-Labeled Protein Standards for Quantitative Mass Spectrometry Ryan D. omgarden 1, Derek aerenwald 2, Eric Hommema 1, Scott Peterman
Development of a Human Cell-Free Expression System to Generate Stable-Isotope-Labeled Protein Standards for Quantitative Mass Spectrometry Ryan D. omgarden 1, Derek aerenwald 2, Eric Hommema 1, Scott Peterman
Quantitation of Protein Phosphorylation Using Multiple Reaction Monitoring
 Quantitation of Protein Phosphorylation Using Multiple Reaction Monitoring Application Note Authors Ning Tang, Christine A. Miller and Keith Waddell Agilent Technologies, Inc. Santa Clara, CA USA This
Quantitation of Protein Phosphorylation Using Multiple Reaction Monitoring Application Note Authors Ning Tang, Christine A. Miller and Keith Waddell Agilent Technologies, Inc. Santa Clara, CA USA This
Automating Mass Spectrometry-Based Quantitative Glycomics using Tandem Mass Tag (TMT) Reagents with SimGlycan
 PREMIER Biosoft Automating Mass Spectrometry-Based Quantitative Glycomics using Tandem Mass Tag (TMT) Reagents with SimGlycan Ne uaca2-3galb1-4glc NAcb1 6 Gal NAca -Thr 3 Ne uaca2-3galb1 Ningombam Sanjib
PREMIER Biosoft Automating Mass Spectrometry-Based Quantitative Glycomics using Tandem Mass Tag (TMT) Reagents with SimGlycan Ne uaca2-3galb1-4glc NAcb1 6 Gal NAca -Thr 3 Ne uaca2-3galb1 Ningombam Sanjib
Agilent Protein In-Gel Tryptic Digestion Kit
 Agilent 5188-2749 Protein In-Gel Tryptic Digestion Kit Agilent Protein In-Gel Tryptic Digestion Kit Instructions Kit Contents The Protein In-Gel Tryptic Digestion Kit includes sufficient reagents for approximately
Agilent 5188-2749 Protein In-Gel Tryptic Digestion Kit Agilent Protein In-Gel Tryptic Digestion Kit Instructions Kit Contents The Protein In-Gel Tryptic Digestion Kit includes sufficient reagents for approximately
Protein Post Translational Modification (PTM) & Profiling Core Erol E. Gulcicek
 Protein Post Translational Modification (PTM) & Profiling Core Erol E. Gulcicek Determine sites of Post Translational Modifications and changes in their Levels - Phosphorylation*** - Ubiquitination - Palmitoylation
Protein Post Translational Modification (PTM) & Profiling Core Erol E. Gulcicek Determine sites of Post Translational Modifications and changes in their Levels - Phosphorylation*** - Ubiquitination - Palmitoylation
Department of Chemistry, Université de Montréal, C.P. 6128, Succursale centre-ville, Montréal, Québec, H3C 3J7, Canada.
 Phosphoproteome dynamics of Saccharomyces cerevisiae under heat shock and cold stress Evgeny Kanshin 1,5, Peter Kubiniok 1,2,5, Yogitha Thattikota 1,3, Damien D Amours 1,3 and Pierre Thibault 1,2,4 * 1
Phosphoproteome dynamics of Saccharomyces cerevisiae under heat shock and cold stress Evgeny Kanshin 1,5, Peter Kubiniok 1,2,5, Yogitha Thattikota 1,3, Damien D Amours 1,3 and Pierre Thibault 1,2,4 * 1
SUNY UPSTATE MEDICAL UNIVERSITY PROTEOMICS CORE
 SUNY UPSTATE MEDICAL UNIVERSITY PROTEOMICS CORE Location: SUNY Upstate Medical University Weiskotten Hall Addition, Room 4303 750 East Adams Street Syracuse, NY 13210 Contact Information: David Kakhniashvili,
SUNY UPSTATE MEDICAL UNIVERSITY PROTEOMICS CORE Location: SUNY Upstate Medical University Weiskotten Hall Addition, Room 4303 750 East Adams Street Syracuse, NY 13210 Contact Information: David Kakhniashvili,
Mass Spectrometry at the Laboratory of Food Chemistry. Edwin Bakx Laboratory of Food Chemistry Wageningen University
 Mass Spectrometry at the Wageningen University Mass Spectrometry at the 3 UPLC/CE - ESI - Ion trap MS systems UPLC Thermo Acella with a Velos or VelosPro CE Beckman PA800 with a Thermo VelosPro 1 UPLC-
Mass Spectrometry at the Wageningen University Mass Spectrometry at the 3 UPLC/CE - ESI - Ion trap MS systems UPLC Thermo Acella with a Velos or VelosPro CE Beckman PA800 with a Thermo VelosPro 1 UPLC-
Proteomics of body liquids as a source for potential methods for medical diagnostics Prof. Dr. Evgeny Nikolaev
 Proteomics of body liquids as a source for potential methods for medical diagnostics Prof. Dr. Evgeny Nikolaev Institute for Biochemical Physics, Rus. Acad. Sci., Moscow, Russia. Institute for Energy Problems
Proteomics of body liquids as a source for potential methods for medical diagnostics Prof. Dr. Evgeny Nikolaev Institute for Biochemical Physics, Rus. Acad. Sci., Moscow, Russia. Institute for Energy Problems
LOCALISATION, IDENTIFICATION AND SEPARATION OF MOLECULES. Gilles Frache Materials Characterization Day October 14 th 2016
 LOCALISATION, IDENTIFICATION AND SEPARATION OF MOLECULES Gilles Frache Materials Characterization Day October 14 th 2016 1 MOLECULAR ANALYSES Which focus? LOCALIZATION of molecules by Mass Spectrometry
LOCALISATION, IDENTIFICATION AND SEPARATION OF MOLECULES Gilles Frache Materials Characterization Day October 14 th 2016 1 MOLECULAR ANALYSES Which focus? LOCALIZATION of molecules by Mass Spectrometry
4.2 RESULTS AND DISCUSSION
 phosphorylated proteins on treatment with Erlotinib.This chapter describes the finding of the SILAC experiment. 4.2 RESULTS AND DISCUSSION This is the first global report of its kind using dual strategies
phosphorylated proteins on treatment with Erlotinib.This chapter describes the finding of the SILAC experiment. 4.2 RESULTS AND DISCUSSION This is the first global report of its kind using dual strategies
Supporting information
 Supporting information Figure legends Supplementary Table 1. Specific product ions obtained from fragmentation of lithium adducts in the positive ion mode comparing the different positional isomers of
Supporting information Figure legends Supplementary Table 1. Specific product ions obtained from fragmentation of lithium adducts in the positive ion mode comparing the different positional isomers of
Figure S6. A-J) Annotated UVPD mass spectra for top ten peptides found among the peptides identified by Byonic but not SEQUEST + Percolator.
 Extending Proteome Coverage by Combining MS/MS Methods and a Modified Bioinformatics Platform adapted for Database Searching of Positive and Negative Polarity 193 nm Ultraviolet Photodissociation Mass
Extending Proteome Coverage by Combining MS/MS Methods and a Modified Bioinformatics Platform adapted for Database Searching of Positive and Negative Polarity 193 nm Ultraviolet Photodissociation Mass
Rapid identification and resistance assessment: The future is mass spectrometry
 Rapid identification and resistance assessment: The future is mass spectrometry Dr Sanmarié Schlebusch Director of Microbiology Mater Pathology Brisbane Outline Introduction Plug and play Pre-prep and
Rapid identification and resistance assessment: The future is mass spectrometry Dr Sanmarié Schlebusch Director of Microbiology Mater Pathology Brisbane Outline Introduction Plug and play Pre-prep and
Symposium on Protein Fractionation
 Symposium on Protein Fractionation Proteomic Analysis of Organ-Specific Breast Cancer Metastasis Speaker: Emily I. Chen, Ph.D The Scripps Research Institute Department of Cell Biology CANCER METASTASIS
Symposium on Protein Fractionation Proteomic Analysis of Organ-Specific Breast Cancer Metastasis Speaker: Emily I. Chen, Ph.D The Scripps Research Institute Department of Cell Biology CANCER METASTASIS
MALDI Imaging Drug Imaging Detlev Suckau Head of R&D MALDI Bruker Daltonik GmbH. December 19,
 MALDI Imaging Drug Imaging Detlev Suckau Head of R&D MALDI Bruker Daltonik GmbH December 19, 2014 1 The principle of MALDI imaging Spatially resolved mass spectra are recorded Each mass signal represents
MALDI Imaging Drug Imaging Detlev Suckau Head of R&D MALDI Bruker Daltonik GmbH December 19, 2014 1 The principle of MALDI imaging Spatially resolved mass spectra are recorded Each mass signal represents
Chapter 3. Protein Structure and Function
 Chapter 3 Protein Structure and Function Broad functional classes So Proteins have structure and function... Fine! -Why do we care to know more???? Understanding functional architechture gives us POWER
Chapter 3 Protein Structure and Function Broad functional classes So Proteins have structure and function... Fine! -Why do we care to know more???? Understanding functional architechture gives us POWER
MASS SPECTROMETRY BASED METABOLOMICS. Pavel Aronov. ABRF2010 Metabolomics Research Group March 21, 2010
 MASS SPECTROMETRY BASED METABOLOMICS Pavel Aronov ABRF2010 Metabolomics Research Group March 21, 2010 Types of Experiments in Metabolomics targeted non targeted Number of analyzed metabolites is limited
MASS SPECTROMETRY BASED METABOLOMICS Pavel Aronov ABRF2010 Metabolomics Research Group March 21, 2010 Types of Experiments in Metabolomics targeted non targeted Number of analyzed metabolites is limited
In-Gel Tryptic Digestion Kit
 INSTRUCTIONS In-Gel Tryptic Digestion Kit 3747 N. Meridian Road P.O. Box 117 Rockford, IL 61105 89871 1468.2 Number Description 89871 In-Gel Tryptic Digestion Kit, sufficient reagents for approximately
INSTRUCTIONS In-Gel Tryptic Digestion Kit 3747 N. Meridian Road P.O. Box 117 Rockford, IL 61105 89871 1468.2 Number Description 89871 In-Gel Tryptic Digestion Kit, sufficient reagents for approximately
Structural vs. nonstructural proteins
 Why would you want to study proteins associated with viruses or virus infection? Receptors Mechanism of uncoating How is gene expression carried out, exclusively by viral enzymes? Gene expression phases?
Why would you want to study proteins associated with viruses or virus infection? Receptors Mechanism of uncoating How is gene expression carried out, exclusively by viral enzymes? Gene expression phases?
Mass Spectrometry based metabolomics
 Mass Spectrometry based metabolomics Metabolomics- A realm of small molecules (
Mass Spectrometry based metabolomics Metabolomics- A realm of small molecules (
Sequence Identification And Spatial Distribution of Rat Brain Tryptic Peptides Using MALDI Mass Spectrometric Imaging
 Sequence Identification And Spatial Distribution of Rat Brain Tryptic Peptides Using MALDI Mass Spectrometric Imaging AB SCIEX MALDI TOF/TOF* Systems Patrick Pribil AB SCIEX, Canada MALDI mass spectrometric
Sequence Identification And Spatial Distribution of Rat Brain Tryptic Peptides Using MALDI Mass Spectrometric Imaging AB SCIEX MALDI TOF/TOF* Systems Patrick Pribil AB SCIEX, Canada MALDI mass spectrometric
Tandem mass spectrometry is becoming the
 Fragmentation of Phosphopeptides in an Ion Trap Mass Spectrometer Jon P. DeGnore and Jun Qin Laboratory of Biophysical Chemistry, National Heart, Lung, and Blood Institute, National Institutes of Health,
Fragmentation of Phosphopeptides in an Ion Trap Mass Spectrometer Jon P. DeGnore and Jun Qin Laboratory of Biophysical Chemistry, National Heart, Lung, and Blood Institute, National Institutes of Health,
The Comparison of High Resolution MS with Triple Quadrupole MS for the Analysis of Oligonucleotides
 The Comparison of High Resolution MS with Triple Quadrupole MS for the Analysis of Oligonucleotides Mohammed Abrar Unilabs York Bioanalytical Solutions Outline Introduction Why LC-MS/MS? Limitations of
The Comparison of High Resolution MS with Triple Quadrupole MS for the Analysis of Oligonucleotides Mohammed Abrar Unilabs York Bioanalytical Solutions Outline Introduction Why LC-MS/MS? Limitations of
Mass Spectrometry and Proteomics Xudong Yao
 Mass Spectrometry and Proteomics Xudong Yao Dept of Chemistry University of Connecticut Storrs, CT April 19, 2005 Proteomics and -omics Roles of mass spectrometry Comparative proteomics Gel or non-gel
Mass Spectrometry and Proteomics Xudong Yao Dept of Chemistry University of Connecticut Storrs, CT April 19, 2005 Proteomics and -omics Roles of mass spectrometry Comparative proteomics Gel or non-gel
Chapter 3. Structure of Enzymes. Enzyme Engineering
 Chapter 3. Structure of Enzymes Enzyme Engineering 3.1 Introduction With purified protein, Determining M r of the protein Determining composition of amino acids and the primary structure Determining the
Chapter 3. Structure of Enzymes Enzyme Engineering 3.1 Introduction With purified protein, Determining M r of the protein Determining composition of amino acids and the primary structure Determining the
4th Multidimensional Chromatography Workshop Toronto (January, 2013) Herman C. Lam, Ph.D. Calibration & Validation Group
 4th Multidimensional Chromatography Workshop Toronto (January, 2013) Herman C. Lam, Ph.D. Calibration & Validation Group MDLC for Shotgun Proteomics Introduction General concepts Advantages Challenges
4th Multidimensional Chromatography Workshop Toronto (January, 2013) Herman C. Lam, Ph.D. Calibration & Validation Group MDLC for Shotgun Proteomics Introduction General concepts Advantages Challenges
Mass-Spectrometric Analysis of Lipids (Lipidomics)
 Mass-Spectrometric Analysis of Lipids (Lipidomics) 1. Identification 2. Quantification 3. Metabolism Why to do lipidomics? Biology: Functions of different lipids? Medicine: Diagnostics and Therapy Industry:
Mass-Spectrometric Analysis of Lipids (Lipidomics) 1. Identification 2. Quantification 3. Metabolism Why to do lipidomics? Biology: Functions of different lipids? Medicine: Diagnostics and Therapy Industry:
Enhancing Sequence Coverage in Proteomics Studies by Using a Combination of Proteolytic Enzymes
 Enhancing Sequence Coverage in Proteomics Studies by Using a Combination of Proteolytic Enzymes Dominic Baeumlisberger 2, Christopher Kurz 3, Tabiwang N. Arrey, Marion Rohmer 2, Carola Schiller 3, Thomas
Enhancing Sequence Coverage in Proteomics Studies by Using a Combination of Proteolytic Enzymes Dominic Baeumlisberger 2, Christopher Kurz 3, Tabiwang N. Arrey, Marion Rohmer 2, Carola Schiller 3, Thomas
FOURIER TRANSFORM MASS SPECTROMETRY
 FOURIER TRANSFORM MASS SPECTROMETRY FT-ICR Theory Ion Cyclotron Motion Inward directed Lorentz force causes ions to move in circular orbits about the magnetic field axis Alan G. Marshall, Christopher L.
FOURIER TRANSFORM MASS SPECTROMETRY FT-ICR Theory Ion Cyclotron Motion Inward directed Lorentz force causes ions to move in circular orbits about the magnetic field axis Alan G. Marshall, Christopher L.
Application of a new capillary HPLC- ICP-MS interface to the identification of selenium-containing proteins in selenized yeast
 Application of a new capillary HPLC- ICP-MS interface to the identification of selenium-containing proteins in selenized yeast Application note Food supplements Authors Juliusz Bianga and Joanna Szpunar
Application of a new capillary HPLC- ICP-MS interface to the identification of selenium-containing proteins in selenized yeast Application note Food supplements Authors Juliusz Bianga and Joanna Szpunar
Quantification by Mass Spectrometry
 Quantification by Mass Spectrometry PC219 Lecture 5 30 April, 2010 sguan@cgl.ucsf.edu 1 Applications of Quantitative Proteomics Qualitative protein identification is NOT sufficient to describe a biological
Quantification by Mass Spectrometry PC219 Lecture 5 30 April, 2010 sguan@cgl.ucsf.edu 1 Applications of Quantitative Proteomics Qualitative protein identification is NOT sufficient to describe a biological
Topics. Protein Bioinformatics ( ) How a Mass Spectrometers Measure Mass. Lecture 9: Quantitative Proteomics Tuesday, April 27, 2010
 Protein Bioinformatics (26.655) Lecture 9: Quantitative Proteomics Tuesday, April 27, 21 Robert N. Cole, Ph.D. Mass Spectrometry and Proteomics Facility Johns Hopkins School of Medicine 371 Broadway Research
Protein Bioinformatics (26.655) Lecture 9: Quantitative Proteomics Tuesday, April 27, 21 Robert N. Cole, Ph.D. Mass Spectrometry and Proteomics Facility Johns Hopkins School of Medicine 371 Broadway Research
Applications of HPLC-MALDI-TOF MS/MS Phosphoproteomic Analysis in Oncological Clinical Diagnostics
 Current Proteomics, 2011, 8, 153-167 153 Applications of HPLC-MALDI-TOF MS/MS Phosphoproteomic Analysis in Oncological Clinical Diagnostics Courtney L. Haddock*,1, Barry Holtz 1, Neil Senzer 1,2,3,4 and
Current Proteomics, 2011, 8, 153-167 153 Applications of HPLC-MALDI-TOF MS/MS Phosphoproteomic Analysis in Oncological Clinical Diagnostics Courtney L. Haddock*,1, Barry Holtz 1, Neil Senzer 1,2,3,4 and
Nat Rev Mol Cell Biol (2006) 7, Mass spectrometry and protein analysis. (2006). Domon and Aebersold Science 312,
 Nat Rev Mol Cell Biol (2006) 7, 391-403. Mass spectrometry and protein analysis. (2006). Domon and Aebersold Science 312, 212-217 Affinity Capture / Enrichment Key to success in PTM ID Purified protein
Nat Rev Mol Cell Biol (2006) 7, 391-403. Mass spectrometry and protein analysis. (2006). Domon and Aebersold Science 312, 212-217 Affinity Capture / Enrichment Key to success in PTM ID Purified protein
Study of different types of ubiquitination
 Study of different types of ubiquitination Rudi Beyaert (rudi.beyaert@irc.vib-ugent.be) VIB UGent Center for Inflammation Research Ghent, Belgium VIB Training Novel Proteomics Tools: Identifying PTMs October
Study of different types of ubiquitination Rudi Beyaert (rudi.beyaert@irc.vib-ugent.be) VIB UGent Center for Inflammation Research Ghent, Belgium VIB Training Novel Proteomics Tools: Identifying PTMs October
An Alternative Approach: Top-Down Bioanalysis of Intact Large Molecules Can this be part of the future? Lecture 8, Page 27
 An Alternative Approach: Top-Down Bioanalysis of Intact Large Molecules Can this be part of the future? Lecture 8, Page 27 Top-down HRAM Bioanalysis of Native Proteins/Molecules Relative Abundance 100
An Alternative Approach: Top-Down Bioanalysis of Intact Large Molecules Can this be part of the future? Lecture 8, Page 27 Top-down HRAM Bioanalysis of Native Proteins/Molecules Relative Abundance 100
The peptide samples of C. elegans (15 mg) and S. cerevisiae (20 mg) were prepared as
 Supplementary material and methods The peptide samples of C. elegans (15 mg) and S. cerevisiae (20 mg) were prepared as described recently 1. The dried down peptides were resolubilized to a final concentration
Supplementary material and methods The peptide samples of C. elegans (15 mg) and S. cerevisiae (20 mg) were prepared as described recently 1. The dried down peptides were resolubilized to a final concentration
Mass Spectrometry and Proteomics - Lecture 1 - Matthias Trost Newcastle University
 Mass Spectrometry and Proteomics - Lecture 1 - Matthias Trost Newcastle University matthias.trost@ncl.ac.uk Content Lectures 1-3 The basics of mass measurement Ionisation techniques Mass analysers Detectors
Mass Spectrometry and Proteomics - Lecture 1 - Matthias Trost Newcastle University matthias.trost@ncl.ac.uk Content Lectures 1-3 The basics of mass measurement Ionisation techniques Mass analysers Detectors
Small Molecule Science: Experimental designs for achieving ultra trace analysis
 Small Molecule Science: Experimental designs for achieving ultra trace analysis Michael P. Balogh Principal scientist Waters Corporation 211 Waters Corporation 1 www.cosmoscience.org The Society for Small
Small Molecule Science: Experimental designs for achieving ultra trace analysis Michael P. Balogh Principal scientist Waters Corporation 211 Waters Corporation 1 www.cosmoscience.org The Society for Small
FOURIER TRANSFORM MASS SPECTROMETRY
 FOURIER TRANSFORM MASS SPECTROMETRY FT-ICR Theory Ion Cyclotron Motion Inward directed Lorentz force causes ions to move in circular orbits about the magnetic field axis Alan G. Marshall, Christopher L.
FOURIER TRANSFORM MASS SPECTROMETRY FT-ICR Theory Ion Cyclotron Motion Inward directed Lorentz force causes ions to move in circular orbits about the magnetic field axis Alan G. Marshall, Christopher L.
ABSTRACT. Dr. Catherine Fenselau, Department of Chemistry and Biochemistry. Presented in this work is a novel bottom up proteomics approach to
 ABSTRACT Title of Document: INTEGRATION OF ASP-SPECIFIC MICROWAVE-ACCELERATED ACID HYDROLYSIS INTO PROTEOMIC ANALYSES Stephen J. Swatkoski, Ph.D., 2007 Directed By: Dr. Catherine Fenselau, Department of
ABSTRACT Title of Document: INTEGRATION OF ASP-SPECIFIC MICROWAVE-ACCELERATED ACID HYDROLYSIS INTO PROTEOMIC ANALYSES Stephen J. Swatkoski, Ph.D., 2007 Directed By: Dr. Catherine Fenselau, Department of
Nature Structural & Molecular Biology: doi: /nsmb.3218
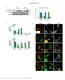 Supplementary Figure 1 Endogenous EGFR trafficking and responses depend on biased ligands. (a) Lysates from HeLa cells stimulated for 2 min. with increasing concentration of ligands were immunoblotted
Supplementary Figure 1 Endogenous EGFR trafficking and responses depend on biased ligands. (a) Lysates from HeLa cells stimulated for 2 min. with increasing concentration of ligands were immunoblotted
Solving practical problems. Maria Kuhtinskaja
 Solving practical problems Maria Kuhtinskaja What does a mass spectrometer do? It measures mass better than any other technique. It can give information about chemical structures. What are mass measurements
Solving practical problems Maria Kuhtinskaja What does a mass spectrometer do? It measures mass better than any other technique. It can give information about chemical structures. What are mass measurements
Dynamic Analysis of HIV-Human Protein-Protein Interactions During Infection
 Dynamic Analysis of HIV-Human Protein-Protein Interactions During Infection Jeffrey Johnson, 1 Shannon Eliuk, 2 Amnon Golan, 1 Tasha Johnson, 1 Vlad Zabrouskov, 2 Nevan Krogan 1 1 UCSF, San Francisco,
Dynamic Analysis of HIV-Human Protein-Protein Interactions During Infection Jeffrey Johnson, 1 Shannon Eliuk, 2 Amnon Golan, 1 Tasha Johnson, 1 Vlad Zabrouskov, 2 Nevan Krogan 1 1 UCSF, San Francisco,
Complexity DNA. Genome RNA. Transcriptome. Protein. Proteome. Metabolites. Metabolome
 DNA Genome Complexity RNA Transcriptome Systems Biology Linking all the components of a cell in a quantitative and temporal manner Protein Proteome Metabolites Metabolome Where are the functional elements?
DNA Genome Complexity RNA Transcriptome Systems Biology Linking all the components of a cell in a quantitative and temporal manner Protein Proteome Metabolites Metabolome Where are the functional elements?
The N-terminal loop of IRAK-4 death domain regulates ordered assembly of the Myddosome signalling scaffold
 The N-terminal loop of IRAK-4 death domain regulates ordered assembly of the Myddosome signalling scaffold Anthony C.G. Dossang * 1, Precious G. Motshwene * 1, Yang Yang* 1, Martyn F. Symmons*, Clare E.
The N-terminal loop of IRAK-4 death domain regulates ordered assembly of the Myddosome signalling scaffold Anthony C.G. Dossang * 1, Precious G. Motshwene * 1, Yang Yang* 1, Martyn F. Symmons*, Clare E.
