SUPPLEMENTARY INFORMATION
|
|
|
- Dayna Matthews
- 5 years ago
- Views:
Transcription
1 Glucose is a ligand for the oxysterol receptor LXR Supplementary Figure 1 Schematic representation of pathways influencing glucose fate in the liver. Glucose induces insulin secretion, suppressing hepatic gluconeogenesis and through LXR activating SREBP-1c expression and lipogenesis. Glucose can also bind directly to LXR to induce SREBP-1c expression, suppress hepatic glucose output, and increase ChREBP expression. ChREBP activitity is modulated by glucose metabolites, further increasing lipogenesis. For clarity, glycogen metabolism is not included in the diagram. Glucose mm : HepG2 LXRα β actin 293T LXRα β actin Supplementary Figure 2 Glucose concentration does not influence the levels of LXR protein expressed in transfected cells. HepG2 or 293T cells were transfected with Flag-tagged hlxrα and grown in media with the indicated glucose concentration. Extracts were prepared 48 h post-transfection and analyzed by Western blot. 1
2 Supplementary Figure 3 Coactivator recruitment dose response curves for D-glucose and LXR agonists. Efficacy is relative to GW3965. Error bars represent s.d. vehicle GW µm D-glucose 30 mm PK µg/ml: Kd 28 Kd 17 Kd 14 Kd 7 Kd Supplementary Figure 4 Glucose shields LXRα from protease digestion. Purified histidinetagged hlxrα LBD was incubated with the indicated concentration of proteinase K and either vehicle or ligands for 15 minutes at room temperature. Digestion products were analyzed by SDS PAGE and stained with Comassie blue. Arrows point to the most prominent proteolytic fragments protected by the presence of GW3965 or D-glucose. 2
3 Supplementary Figure 5 Glucose shifts the melting temperature of LXRβ. Purified hlxrβ LBD alone or incubated with either D-glucose or GW3965 at the indicated concentration was analyzed by differential scanning calorimetry (DSC). This technique is used to determine thermal transition temperatures for samples in solution; ligand binding generally stabilizes the protein and the T m of the protein-ligand complex is higher than that of the protein in the absence of ligand. Both glucose and GW3965 increased the melting point of the protein, indicating that they are binding directly to LXRβ. DSC was performed on a Microcal VP-DSC differential scanning calorimeter, in a temperature range that covered C, at a scanning rate equal to 90 C/h. 400 µg of hlxrβ LBD in a total cell volume of 0.4 ml were used in each thermal scan. Subsequent scans containing D-glucose and GW3965 were separated with buffer only samples to avoid cross-contamination within the sample cell. In experiments with added ligand, the reference cell contained an equal concentration of ligand to confirm the absence of any significant contribution to the T m shift. Results were analyzed with Origin 7.0 software. Thermograms display excess heat capacity vs. temperature and melting temperatures were calculated from an intrinsic software feature. Proteins used for protease protection and differential scanning calorimetry were made in-house due to the large amount of protein that these assays require. Briefly, the ligand binding domain ( ) of human LXRβ was cloned into a vector for expression in baculovirus which contains 3
4 an N-terminal hexa-his tag with a TEV protease cleavage site. Lysis of insect cells was achieved by sonication. After removal of cell debris by centrifugation, the resulting lysate containing soluble LXRβ LBD was purified by Ni-NTA chromatography followed S200 gel filtration chromatography. LXRβ-containing fractions were pooled and the protein was concentrated to ~10 mg/ml. The purity of the protein was evaluated to be higher than 95% based on SDS PAGE. Human LXRα( ) was cloned into the same vector and processed identically. Identity of purified proteins was checked by standard LC MS/MS methods after trypsin digestion, and their functional activity evaluated by a SPA assay with a reference compound. Supplementary Figure 6 Glucose and other LXR ligands suppress gluconeogenic genes under conditions of maximal induction. HepG2 cells were pre-treated for 8 h with 1 mm 8- Br-cAMP and 1 µm dexamethasone prior to addition of ligands or insulin (100 nm). Insulin has no effect on the expression of ABCG1, but appears to work together with LXR ligands to suppress gluconeogenesis. Error bars represent s.d. 4
5 Supplementary Figure 7 LXR is required for glucose modulation of cholesterol efflux genes. In cells pretreated for 48 h with sirna against hlxrα, neither D-glucose nor GW3965 stimulate expression of ABCA1, ABCG1, or CETP (see Methods). Error bars represent s.d. 5
6 Supplementary Figure 8 Inhibition of glucose uptake suppresses induction of LXR targets by glucose and diminishes the response to GW3965. Cells were pre-treated for 16 h with cytochalasin B (CcB, 5µM) prior to ligand addition (see Methods). Error bars represent s.d. 6
7 Supplementary Figure 9 Glucose can regulate LXR targets in the absence of insulin secretion. Insulin-depleted STZ-injected mice fasted overnight were challenged orally with GW3965 (50 mpk), or refed with a glucose or sucrose diet and sacrificed 6 h later. Hepatic gene expression was assessed by qrt-pcr. Note that expression of direct LXR targets (e.g. SPα, SREBP-1c) correlates with circulating glucose, not insulin. Error bars represent s.d. *p<0.05, **p<0.001 treatment vs. fasted. 7
8 Supplementary Figure 10 Glucose induces expression of insulin-independent LXR targets and SREBP-1c in intestine. Sucrose has a milder effect. Wild-type mice fasted overnight were challenged orally with GW3965 (50 mpk), or refed with a glucose or sucrose diet and sacrificed 6 h later. Hepatic gene expression was assessed by qrt-pcr. Error bars represent s.d. *p<0.05, **p<0.001 treatment vs. fasted. 8
9 Materials and Methods Constructs Expression constructs for chimeric NR LBDs were generated by fusing the LBDs to the DBD of GAL4 and inserting them into pcdna3. Assays were validated using known ligands. The ptk-2xlxre-abca1-luc reporter and pcdna3 expression plasmids for LXRα/β have been described 22. Cell culture and treatments HepG2 cells were cultured in DMEM with 10% FBS. For transcriptional assays, cells were plated in 384 well plates in DMEM with varying glucose concentration (0, 2, 25 mm) with 10% fetal bovine lipoprotein deficient serum (FBLDS, Intracel) in the presence of 7.5 μm lovastatin and 100 μm mevalonic acid. For gene expression analysis, cells were plated in 48 well plates in the above conditions, and treated for 16 h with DMSO, 1 μm GW3965, 5 μm 22-R-HC, or 20 mm D-glucose. Induction of gluconeogenic genes was achieved using 8-Br-cMAP (1 mm) and dexamethasone (1 µm) for 8 h prior to test compound or insulin (100 nm) addition; cells were harvested 6 h after ligand addition. For sirna experiments, HepG2 cells were transfected with an hlxrα (Dharmacon, SMART pool M ) or control sirna pool (Dharmacon, sicontrol non-targeting sirna pool, D , designed to have >4 mismatches with all known human and mouse genes) using DharmaFECT 4; 48 h later, ligands were added, and cells were harvested 24 h later. To check for knockdown, nuclear extracts were analyzed by Western blot using an antibody against LXR (Calbiochem). In glucose uptake inhibition experiments, cells were pre-treated with 5 µm cytochalasin B for 16 h before addition of ligands for 8 h. Transcriptional assays HepG2 cells were transfected using FuGENE 6 (Roche). 24 h later, ligands were added as 12- point dose response curves starting at 10 μm for GW3965 and T , 30 μm for 22-R-HC and 22-S-HC, and 20 mm for carbohydrates. 16 h later, luciferase activity was read and normalized to an internal pcmv-gfp control. Fluorescence-resonance energy transfer assay The purity and identity of commercial proteins used in binding assays was verified by mass fingerprinting using trypsin and MALDI-TOF mass spectrometry. Purity was greater than 85% as evaluated by SDS PAGE, with no detectable protease, DNase, or RNase activity. Functional activity of all protein batches received was tested using coactivator recruitment and SPA assays with reference compounds to insure that results with standards were within published values. In all cases, fractional occupancy using a 12-point dose response to labeled T was calculated to verify protein quality prior to its use in FRET or SPA assays. The top dose of labeled T used was greater than 400 times its K d, ensuring receptor saturation. FRET assay was performed in 384 well plates in 20 μl of volume. A mix of 5 nm His-LXRαLBD or His-LXRβLBD (Roche), 5 nm biotinylated SRC-1 peptide (Biotin- CPSSHSSLTERHKILHRLLQEGSPSC-OH), Europium-labeled anti-his antibody (Perkin Elmer) and allophycocyanin-labeled streptavidin (Prozyme) was prepared. Ligands were tested in 12-point dose response curves starting at 10 μm for GW3965 and T , 30 μm for 22-R- HC and 22-S-HC, and 20 mm for glycolytic intermediates. Plates were incubated for 3 h at rt 9
10 and FRET read on an AnalystGT (Molecular Devices). In combination experiments, compounds were added simultaneously. Scintillation proximity assay SPA was performed as in Janowski et al 23. Scintillant-filled beads (Amersham) were diluted in SPA buffer (10 mm K 2 HPO 4, 10 mm KH 2 PO 4, 2 mm EDTA, 50 mm NaCl, 1 mm DTT, 2 mm CHAPS, 10% glycerol) to a concentration of 5 mg/ml. Assay was performed in 384 well plates (Packard) in 20 μl containing beads (0.2 mg per well) and His-LXRαLBD (120 ng per well) or His-LXRβLBD (50 ng per well) (Roche). Amount of protein did not deplete ligand concentration. Binding of [ 3 H]-glucose was tested as a 12-point dilution curve starting at 70 mm. Displacement assays used 25 nm [ 3 H]-T (Amersham) or 20 mm [ 3 H]-glucose (Amersham). Ligands were tested in 12-point dose response curves starting at 12 μm for GW3965 and T , 35 μm for 22-R-HC and 22-S-HC, and 70 mm for carbohydrates. In combination experiments, ligands were added simultaneously. Plates were shaken for 3 h at room temperature and read in a Packard Topcount at 1 min per well. Wells devoid of competitor represented 100% binding. Non-specific binding was measured by leaving LXR protein out of the SPA reaction. To calculate the fraction of LXR protein that bound ligand, we used the following formula from the law of mass action: Fractional occupancy = [Receptor- Agonist]/[Receptor], where [Receptor-Agonist] is the concentration of the receptor-agonist complex, and [Receptor] is the total concentration of the receptor. In our assays, the fraction of LXRs that bind the ligand is 95-99%. Generation of K d and K i values Dose response and competition curves were generated by nonlinear regression analysis using GraphPad Prism and the apparent equilibrium association/dissociation constants (K d and K i values) were determined using the method described by DeBlasi and colleagues 24. Animal experiments 10-week-old male C57BL/6 mice were maintained on standard chow diet. Mice fasted for 24 h were refed with sucrose and D-glucose diets lacking cholesterol (Research Diets Inc., D , D ) or gavaged with GW mpk, 6 h before sacrifice. Age-matched mice were rendered insulin-deficient via injection of streptozotocin (100 µg/kg IP) on 3 consecutive days. STZ-treated mice were used once hyperglycemia (glucose > 350 mg/dl) became evident (6 days). All experiments were carried out with strict awareness of the light cycle in place to avoid variation in gene expression arising from potential circadian regulation. Experiments were approved by GNF s IACUC. Gene expression analysis RNA was analyzed by TaqMan qrt-pcr using the one-step Superscript III platinum reagent (Invitrogen). Samples were run in triplicate as multiplexed reactions with a normalizing internal control (36B4). Probe and primer sequences are available on request. Statistical analysis was performed by using a two-tailed Student s t test. 10
11 Supplementary Table 1 Structure-activity relationship of glycolysis derivatives on LXR/RXR transactivation assay LXRα LXRβ Compound K d, μm EC 50, μm Efficacy K d, μm EC 50, μm Efficacy GW ± ± T ± ± (R)-HC 4.05 ± ± (S)-HC D-(+)-Glucose 1770 ± ± D-Glucose-6-P - No fit No fit D-(-)-Fructose - No fit No fit 0.10 D-Fructose-6-P - No fit No fit D-Fructose-1,6-P DL-Glyceraldehyde-3-P D-(-)-3-Phosphoglycerate Phosphoenolpyruvate Pyruvate K d values are presented ± S.D. EC 50, effective concentration for 50% maximal activation; Efficacy, maximal fold activation relative to GW3965; -, below detection; No fit, no plateau at 20 mm. 11
12 Supplementary Table 2 Structure-activity relationship of glycolysis derivatives on coactivator recruitment assay LXRα LXRβ Compound K d, μm EC 50, μm Efficacy K d, μm EC 50, μm Efficacy GW ± ± T ± ± (R)-HC 1.12 ± ± (S)-HC D-(+)-Glucose 1931 ± ± L-Glucose 1410 ± ± D-Glucose-6-P 1765 ± ± D-(-)-Fructose D-Fructose-6-P D-Fructose-1,6-P DL-Glyceraldehyde-3-P D-(-)-3-Phosphoglycerate Phosphoenolpyruvate Pyruvate K d values are presented ± S.D. EC 50, effective concentration for 50% maximal activation; Efficacy, maximal fold activation relative to GW3965; -, below detection. 12
13 Supplementary Table 3 Structure-activity relationship of glycolysis derivatives on SPA displacement assay LXRα LXRβ Compound K i, μm EC 50, μm Efficacy K i, μm EC 50, μm Efficacy GW ± ± T ± ± (R)-HC ± ± (S)-HC ± ± D-(+)-Glucose 1801 ± ± L-Glucose 1745 ± ± D-Glucose 6-P 2967 ± ± D-(-)-Fructose D-Fructose 6-P D-Fructose 1,6-P DL-Glyceraldehyde 3-P D-(-)-3-Phosphoglycerate Phosphoenolpyruvate Pyruvate K i values are presented ± S.D. EC 50, effective concentration for 50% maximal displacement of [ H]-T ; Efficacy, maximal activity relative to GW3965; -, below detection
2.5. AMPK activity
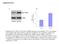 Supplement Fig. A 3 B phos-ampk 2.5 * Control AICAR AMPK AMPK activity (Absorbance at 45 nm) 2.5.5 Control AICAR Supplement Fig. Effects of AICAR on AMPK activation in macrophages. J774. macrophages were
Supplement Fig. A 3 B phos-ampk 2.5 * Control AICAR AMPK AMPK activity (Absorbance at 45 nm) 2.5.5 Control AICAR Supplement Fig. Effects of AICAR on AMPK activation in macrophages. J774. macrophages were
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION FOR Liver X Receptor α mediates hepatic triglyceride accumulation through upregulation of G0/G1 Switch Gene 2 (G0S2) expression I: SUPPLEMENTARY METHODS II: SUPPLEMENTARY FIGURES
SUPPLEMENTARY INFORMATION FOR Liver X Receptor α mediates hepatic triglyceride accumulation through upregulation of G0/G1 Switch Gene 2 (G0S2) expression I: SUPPLEMENTARY METHODS II: SUPPLEMENTARY FIGURES
Supplementary Table 1. Metabolic parameters in GFP and OGT-treated mice
 Supplementary Table 1. Metabolic parameters in GFP and OGT-treated mice Fasted Refed GFP OGT GFP OGT Liver G6P (mmol/g) 0.03±0.01 0.04±0.02 0.60±0.04 0.42±0.10 A TGs (mg/g of liver) 20.08±5.17 16.29±0.8
Supplementary Table 1. Metabolic parameters in GFP and OGT-treated mice Fasted Refed GFP OGT GFP OGT Liver G6P (mmol/g) 0.03±0.01 0.04±0.02 0.60±0.04 0.42±0.10 A TGs (mg/g of liver) 20.08±5.17 16.29±0.8
The antiparasitic drug ivermectin is a novel FXR ligand that regulates metabolism
 Supplementary Information The antiparasitic drug ivermectin is a novel FXR ligand that regulates metabolism Address correspondence to Yong Li (yongli@xmu.edu.cn, Tel: 86-592-218151) GW464 CDCA Supplementary
Supplementary Information The antiparasitic drug ivermectin is a novel FXR ligand that regulates metabolism Address correspondence to Yong Li (yongli@xmu.edu.cn, Tel: 86-592-218151) GW464 CDCA Supplementary
SUPPLEMENTARY INFORMATION
 Supplementary Figures Supplementary Figure S1. Binding of full-length OGT and deletion mutants to PIP strips (Echelon Biosciences). Supplementary Figure S2. Binding of the OGT (919-1036) fragments with
Supplementary Figures Supplementary Figure S1. Binding of full-length OGT and deletion mutants to PIP strips (Echelon Biosciences). Supplementary Figure S2. Binding of the OGT (919-1036) fragments with
HEK293FT cells were transiently transfected with reporters, N3-ICD construct and
 Supplementary Information Luciferase reporter assay HEK293FT cells were transiently transfected with reporters, N3-ICD construct and increased amounts of wild type or kinase inactive EGFR. Transfections
Supplementary Information Luciferase reporter assay HEK293FT cells were transiently transfected with reporters, N3-ICD construct and increased amounts of wild type or kinase inactive EGFR. Transfections
Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells
 Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells (b). TRIM33 was immunoprecipitated, and the amount of
Supplementary Figure 1.TRIM33 binds β-catenin in the nucleus. a & b, Co-IP of endogenous TRIM33 with β-catenin in HT-29 cells (a) and HEK 293T cells (b). TRIM33 was immunoprecipitated, and the amount of
Supplemental information
 Carcinoemryonic antigen-related cell adhesion molecule 6 (CEACAM6) promotes EGF receptor signaling of oral squamous cell carcinoma metastasis via the complex N-glycosylation y Chiang et al. Supplemental
Carcinoemryonic antigen-related cell adhesion molecule 6 (CEACAM6) promotes EGF receptor signaling of oral squamous cell carcinoma metastasis via the complex N-glycosylation y Chiang et al. Supplemental
Supplementary Figure 1. PAQR3 knockdown inhibits SREBP-2 processing in CHO-7 cells CHO-7 cells were transfected with control sirna or a sirna
 Supplementary Figure 1. PAQR3 knockdown inhibits SREBP-2 processing in CHO-7 cells CHO-7 cells were transfected with control sirna or a sirna targeted for hamster PAQR3. At 24 h after the transfection,
Supplementary Figure 1. PAQR3 knockdown inhibits SREBP-2 processing in CHO-7 cells CHO-7 cells were transfected with control sirna or a sirna targeted for hamster PAQR3. At 24 h after the transfection,
Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS)
 Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS) and their exosomes (EXO) in resting (REST) and activated
Supplementary Figure S1. Venn diagram analysis of mrna microarray data and mirna target analysis. (a) Western blot analysis of T lymphoblasts (CLS) and their exosomes (EXO) in resting (REST) and activated
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature12652 Supplementary Figure 1. PRDM16 interacts with endogenous EHMT1 in brown adipocytes. Immunoprecipitation of PRDM16 complex by flag antibody (M2) followed by Western blot analysis
doi:10.1038/nature12652 Supplementary Figure 1. PRDM16 interacts with endogenous EHMT1 in brown adipocytes. Immunoprecipitation of PRDM16 complex by flag antibody (M2) followed by Western blot analysis
Electron micrograph of phosphotungstanic acid-stained exosomes derived from murine
 1 SUPPLEMENTARY INFORMATION SUPPLEMENTARY FIGURES Supplementary Figure 1. Physical properties of murine DC-derived exosomes. a, Electron micrograph of phosphotungstanic acid-stained exosomes derived from
1 SUPPLEMENTARY INFORMATION SUPPLEMENTARY FIGURES Supplementary Figure 1. Physical properties of murine DC-derived exosomes. a, Electron micrograph of phosphotungstanic acid-stained exosomes derived from
Supplementary Figure 1. Normal T lymphocyte populations in Dapk -/- mice. (a) Normal thymic development in Dapk -/- mice. Thymocytes from WT and Dapk
 Supplementary Figure 1. Normal T lymphocyte populations in Dapk -/- mice. (a) Normal thymic development in Dapk -/- mice. Thymocytes from WT and Dapk -/- mice were stained for expression of CD4 and CD8.
Supplementary Figure 1. Normal T lymphocyte populations in Dapk -/- mice. (a) Normal thymic development in Dapk -/- mice. Thymocytes from WT and Dapk -/- mice were stained for expression of CD4 and CD8.
Supporting Information Table of content
 Supporting Information Table of content Supporting Information Fig. S1 Supporting Information Fig. S2 Supporting Information Fig. S3 Supporting Information Fig. S4 Supporting Information Fig. S5 Supporting
Supporting Information Table of content Supporting Information Fig. S1 Supporting Information Fig. S2 Supporting Information Fig. S3 Supporting Information Fig. S4 Supporting Information Fig. S5 Supporting
Use of a camp BRET Sensor to Characterize a Novel Regulation of camp by the Sphingosine-1-phosphate/G 13 Pathway
 Use of a camp BRET Sensor to Characterize a Novel Regulation of camp by the Sphingosine-1-phosphate/G 13 Pathway SUPPLEMENTAL DATA Characterization of the CAMYEL sensor and calculation of intracellular
Use of a camp BRET Sensor to Characterize a Novel Regulation of camp by the Sphingosine-1-phosphate/G 13 Pathway SUPPLEMENTAL DATA Characterization of the CAMYEL sensor and calculation of intracellular
Supplementary Information for. Inhibitors of Hedgehog Acyltransferase Block Sonic Hedgehog Signaling. Marilyn D. Resh 1,2,6*
 Supplementary Information for Inhibitors of Hedgehog Acyltransferase Block Sonic Hedgehog Signaling Elissaveta Petrova 1,5, Jessica Rios-Esteves 1,2, Ouathek Ouerfelli 3, J. Fraser Glickman 4, and Marilyn
Supplementary Information for Inhibitors of Hedgehog Acyltransferase Block Sonic Hedgehog Signaling Elissaveta Petrova 1,5, Jessica Rios-Esteves 1,2, Ouathek Ouerfelli 3, J. Fraser Glickman 4, and Marilyn
Optimization of a LanthaScreen Kinase assay for ZAP70
 Optimization of a LanthaScreen Kinase assay for ZAP70 Overview This protocol describes how to develop a LanthaScreen kinase assay designed to detect and characterize kinase inhibitors. The development
Optimization of a LanthaScreen Kinase assay for ZAP70 Overview This protocol describes how to develop a LanthaScreen kinase assay designed to detect and characterize kinase inhibitors. The development
SUPPLEMENTARY INFORMATION
 Supplementary Table 1. Cell sphingolipids and S1P bound to endogenous TRAF2. Sphingolipid Cell pmol/mg TRAF2 immunoprecipitate pmol/mg Sphingomyelin 4200 ± 250 Not detected Monohexosylceramide 311 ± 18
Supplementary Table 1. Cell sphingolipids and S1P bound to endogenous TRAF2. Sphingolipid Cell pmol/mg TRAF2 immunoprecipitate pmol/mg Sphingomyelin 4200 ± 250 Not detected Monohexosylceramide 311 ± 18
Supplementary Information. Cryptochrome Mediates Circadian Regulation of camp. Signalling and Hepatic Gluconeogenesis
 Supplementary Information Cryptochrome Mediates Circadian Regulation of camp Signalling and Hepatic Gluconeogenesis Eric E. Zhang 1,2*, Yi Liu 3,5*, Renaud Dentin 3, Pagkapol Y. Pongsawakul 1, Andrew C.
Supplementary Information Cryptochrome Mediates Circadian Regulation of camp Signalling and Hepatic Gluconeogenesis Eric E. Zhang 1,2*, Yi Liu 3,5*, Renaud Dentin 3, Pagkapol Y. Pongsawakul 1, Andrew C.
Supplementary Fig. 1. Identification of acetylation of K68 of SOD2
 Supplementary Fig. 1. Identification of acetylation of K68 of SOD2 A B H. sapiens 54 KHHAAYVNNLNVTEEKYQEALAK 75 M. musculus 54 KHHAAYVNNLNATEEKYHEALAK 75 X. laevis 55 KHHATYVNNLNITEEKYAEALAK 77 D. rerio
Supplementary Fig. 1. Identification of acetylation of K68 of SOD2 A B H. sapiens 54 KHHAAYVNNLNVTEEKYQEALAK 75 M. musculus 54 KHHAAYVNNLNATEEKYHEALAK 75 X. laevis 55 KHHATYVNNLNITEEKYAEALAK 77 D. rerio
Supplementary Fig. 1. GPRC5A post-transcriptionally down-regulates EGFR expression. (a) Plot of the changes in steady state mrna levels versus
 Supplementary Fig. 1. GPRC5A post-transcriptionally down-regulates EGFR expression. (a) Plot of the changes in steady state mrna levels versus changes in corresponding proteins between wild type and Gprc5a-/-
Supplementary Fig. 1. GPRC5A post-transcriptionally down-regulates EGFR expression. (a) Plot of the changes in steady state mrna levels versus changes in corresponding proteins between wild type and Gprc5a-/-
Data Sheet TIGIT / NFAT Reporter - Jurkat Cell Line Catalog #60538
 Data Sheet TIGIT / NFAT Reporter - Jurkat Cell Line Catalog #60538 Background: TIGIT is a co-inhibitory receptor that is highly expressed in Natural Killer (NK) cells, activated CD4+, CD8+ and regulatory
Data Sheet TIGIT / NFAT Reporter - Jurkat Cell Line Catalog #60538 Background: TIGIT is a co-inhibitory receptor that is highly expressed in Natural Killer (NK) cells, activated CD4+, CD8+ and regulatory
Islet viability assay and Glucose Stimulated Insulin Secretion assay RT-PCR and Western Blot
 Islet viability assay and Glucose Stimulated Insulin Secretion assay Islet cell viability was determined by colorimetric (3-(4,5-dimethylthiazol-2-yl)-2,5- diphenyltetrazolium bromide assay using CellTiter
Islet viability assay and Glucose Stimulated Insulin Secretion assay Islet cell viability was determined by colorimetric (3-(4,5-dimethylthiazol-2-yl)-2,5- diphenyltetrazolium bromide assay using CellTiter
T H E J O U R N A L O F C E L L B I O L O G Y
 Supplemental material Chairoungdua et al., http://www.jcb.org/cgi/content/full/jcb.201002049/dc1 T H E J O U R N A L O F C E L L B I O L O G Y Figure S1. Expression of CD9 and CD82 inhibits Wnt/ -catenin
Supplemental material Chairoungdua et al., http://www.jcb.org/cgi/content/full/jcb.201002049/dc1 T H E J O U R N A L O F C E L L B I O L O G Y Figure S1. Expression of CD9 and CD82 inhibits Wnt/ -catenin
TFEB-mediated increase in peripheral lysosomes regulates. Store Operated Calcium Entry
 TFEB-mediated increase in peripheral lysosomes regulates Store Operated Calcium Entry Luigi Sbano, Massimo Bonora, Saverio Marchi, Federica Baldassari, Diego L. Medina, Andrea Ballabio, Carlotta Giorgi
TFEB-mediated increase in peripheral lysosomes regulates Store Operated Calcium Entry Luigi Sbano, Massimo Bonora, Saverio Marchi, Federica Baldassari, Diego L. Medina, Andrea Ballabio, Carlotta Giorgi
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells
 MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells Margaret S Ebert, Joel R Neilson & Phillip A Sharp Supplementary figures and text: Supplementary Figure 1. Effect of sponges on
Optimization of a LanthaScreen Kinase assay for NTRK1 (TRKA)
 Optimization of a LanthaScreen Kinase assay for NTRK1 (TRKA) Overview This protocol describes how to develop a LanthaScreen kinase assay designed to detect and characterize kinase inhibitors. The development
Optimization of a LanthaScreen Kinase assay for NTRK1 (TRKA) Overview This protocol describes how to develop a LanthaScreen kinase assay designed to detect and characterize kinase inhibitors. The development
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:1.138/nature9814 a A SHARPIN FL B SHARPIN ΔNZF C SHARPIN T38L, F39V b His-SHARPIN FL -1xUb -2xUb -4xUb α-his c Linear 4xUb -SHARPIN FL -SHARPIN TF_LV -SHARPINΔNZF -SHARPIN
SUPPLEMENTARY INFORMATION doi:1.138/nature9814 a A SHARPIN FL B SHARPIN ΔNZF C SHARPIN T38L, F39V b His-SHARPIN FL -1xUb -2xUb -4xUb α-his c Linear 4xUb -SHARPIN FL -SHARPIN TF_LV -SHARPINΔNZF -SHARPIN
Supplementary Material
 Supplementary Material Nuclear import of purified HIV-1 Integrase. Integrase remains associated to the RTC throughout the infection process until provirus integration occurs and is therefore one likely
Supplementary Material Nuclear import of purified HIV-1 Integrase. Integrase remains associated to the RTC throughout the infection process until provirus integration occurs and is therefore one likely
Over-expression of MKP-3 and knockdown of MKP-3 and FOXO1 in primary rat. day, cells were transduced with adenoviruses expressing GFP, MKP-3 or shgfp,
 SUPPLEMENTAL METHODS Over-expression of MKP-3 and knockdown of MKP-3 and FOXO1 in primary rat hepatocytes Primary rat hepatocytes were seeded as described in experimental procedures. The next day, cells
SUPPLEMENTAL METHODS Over-expression of MKP-3 and knockdown of MKP-3 and FOXO1 in primary rat hepatocytes Primary rat hepatocytes were seeded as described in experimental procedures. The next day, cells
SUPPLEMENTAL INFORMATION
 SUPPLEMENTAL INFORMATION EXPERIMENTAL PROCEDURES Tryptic digestion protection experiments - PCSK9 with Ab-3D5 (1:1 molar ratio) in 50 mm Tris, ph 8.0, 150 mm NaCl was incubated overnight at 4 o C. The
SUPPLEMENTAL INFORMATION EXPERIMENTAL PROCEDURES Tryptic digestion protection experiments - PCSK9 with Ab-3D5 (1:1 molar ratio) in 50 mm Tris, ph 8.0, 150 mm NaCl was incubated overnight at 4 o C. The
Supporting Online Material Material and Methods References Supplemental Figures S1, S2, and S3
 Supporting Online Material Material and Methods References Supplemental Figures S1, S2, and S3 Sarbassov et al. 1 Material and Methods Materials Reagents were obtained from the following sources: protein
Supporting Online Material Material and Methods References Supplemental Figures S1, S2, and S3 Sarbassov et al. 1 Material and Methods Materials Reagents were obtained from the following sources: protein
hexahistidine tagged GRP78 devoid of the KDEL motif (GRP78-His) on SDS-PAGE. This
 SUPPLEMENTAL FIGURE LEGEND Fig. S1. Generation and characterization of. (A) Coomassie staining of soluble hexahistidine tagged GRP78 devoid of the KDEL motif (GRP78-His) on SDS-PAGE. This protein was expressed
SUPPLEMENTAL FIGURE LEGEND Fig. S1. Generation and characterization of. (A) Coomassie staining of soluble hexahistidine tagged GRP78 devoid of the KDEL motif (GRP78-His) on SDS-PAGE. This protein was expressed
Supplementary data Supplementary Figure 1 Supplementary Figure 2
 Supplementary data Supplementary Figure 1 SPHK1 sirna increases RANKL-induced osteoclastogenesis in RAW264.7 cell culture. (A) RAW264.7 cells were transfected with oligocassettes containing SPHK1 sirna
Supplementary data Supplementary Figure 1 SPHK1 sirna increases RANKL-induced osteoclastogenesis in RAW264.7 cell culture. (A) RAW264.7 cells were transfected with oligocassettes containing SPHK1 sirna
The clathrin adaptor Numb regulates intestinal cholesterol. absorption through dynamic interaction with NPC1L1
 The clathrin adaptor Numb regulates intestinal cholesterol absorption through dynamic interaction with NPC1L1 Pei-Shan Li 1, Zhen-Yan Fu 1,2, Ying-Yu Zhang 1, Jin-Hui Zhang 1, Chen-Qi Xu 1, Yi-Tong Ma
The clathrin adaptor Numb regulates intestinal cholesterol absorption through dynamic interaction with NPC1L1 Pei-Shan Li 1, Zhen-Yan Fu 1,2, Ying-Yu Zhang 1, Jin-Hui Zhang 1, Chen-Qi Xu 1, Yi-Tong Ma
HCC1937 is the HCC1937-pcDNA3 cell line, which was derived from a breast cancer with a mutation
 SUPPLEMENTARY INFORMATION Materials and Methods Human cell lines and culture conditions HCC1937 is the HCC1937-pcDNA3 cell line, which was derived from a breast cancer with a mutation in exon 20 of BRCA1
SUPPLEMENTARY INFORMATION Materials and Methods Human cell lines and culture conditions HCC1937 is the HCC1937-pcDNA3 cell line, which was derived from a breast cancer with a mutation in exon 20 of BRCA1
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3461 In the format provided by the authors and unedited. Supplementary Figure 1 (associated to Figure 1). Cpeb4 gene-targeted mice develop liver steatosis. a, Immunoblot displaying CPEB4
DOI: 10.1038/ncb3461 In the format provided by the authors and unedited. Supplementary Figure 1 (associated to Figure 1). Cpeb4 gene-targeted mice develop liver steatosis. a, Immunoblot displaying CPEB4
Table S1. Primer sequences used for qrt-pcr. CACCATTGGCAATGAGCGGTTC AGGTCTTTGCGGATGTCCACGT ACTB AAGTCCATGTGCTGGCAGCACT ATCACCACTCCGAAGTCCGTCT LCOR
 Table S1. Primer sequences used for qrt-pcr. ACTB LCOR KLF6 CTBP1 CDKN1A CDH1 ATF3 PLAU MMP9 TFPI2 CACCATTGGCAATGAGCGGTTC AGGTCTTTGCGGATGTCCACGT AAGTCCATGTGCTGGCAGCACT ATCACCACTCCGAAGTCCGTCT CGGCTGCAGGAAAGTTTACA
Table S1. Primer sequences used for qrt-pcr. ACTB LCOR KLF6 CTBP1 CDKN1A CDH1 ATF3 PLAU MMP9 TFPI2 CACCATTGGCAATGAGCGGTTC AGGTCTTTGCGGATGTCCACGT AAGTCCATGTGCTGGCAGCACT ATCACCACTCCGAAGTCCGTCT CGGCTGCAGGAAAGTTTACA
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11700 Figure 1: RIP3 as a potential Sirt2 interacting protein. Transfected Flag-tagged Sirt2 was immunoprecipitated from cells and eluted from the Sepharose beads using Flag peptide.
doi:10.1038/nature11700 Figure 1: RIP3 as a potential Sirt2 interacting protein. Transfected Flag-tagged Sirt2 was immunoprecipitated from cells and eluted from the Sepharose beads using Flag peptide.
HIV-1 Virus-like Particle Budding Assay Nathan H Vande Burgt, Luis J Cocka * and Paul Bates
 HIV-1 Virus-like Particle Budding Assay Nathan H Vande Burgt, Luis J Cocka * and Paul Bates Department of Microbiology, Perelman School of Medicine at the University of Pennsylvania, Philadelphia, USA
HIV-1 Virus-like Particle Budding Assay Nathan H Vande Burgt, Luis J Cocka * and Paul Bates Department of Microbiology, Perelman School of Medicine at the University of Pennsylvania, Philadelphia, USA
Supplementary Figure 1. MAT IIα is Acetylated at Lysine 81.
 IP: Flag a Mascot PTM Modified Mass Error Position Gene Names Score Score Sequence m/z [ppm] 81 MAT2A;AMS2;MATA2 35.6 137.28 _AAVDYQK(ac)VVR_ 595.83-2.28 b Pre-immu After-immu Flag- WT K81R WT K81R / Flag
IP: Flag a Mascot PTM Modified Mass Error Position Gene Names Score Score Sequence m/z [ppm] 81 MAT2A;AMS2;MATA2 35.6 137.28 _AAVDYQK(ac)VVR_ 595.83-2.28 b Pre-immu After-immu Flag- WT K81R WT K81R / Flag
ACC ELOVL MCAD. CPT1α 1.5 *** 0.5. Reverbα *** *** 0.5. Fasted. Refed
 Supplementary Figure A 8 SREBPc 6 5 FASN ELOVL6.5.5.5 ACC.5.5 CLOCK.5.5 CRY.5.5 PPARα.5.5 ACSL CPTα.5.5.5.5 MCAD.5.5 PEPCK.5.5 G6Pase 5.5.5.5 BMAL.5.5 Reverbα.5.5 Reverbβ.5.5 PER.5.5 PER B Fasted Refed
Supplementary Figure A 8 SREBPc 6 5 FASN ELOVL6.5.5.5 ACC.5.5 CLOCK.5.5 CRY.5.5 PPARα.5.5 ACSL CPTα.5.5.5.5 MCAD.5.5 PEPCK.5.5 G6Pase 5.5.5.5 BMAL.5.5 Reverbα.5.5 Reverbβ.5.5 PER.5.5 PER B Fasted Refed
SUPPLEMENT. Materials and methods
 SUPPLEMENT Materials and methods Cell culture and reagents Cell media and reagents were from Invitrogen unless otherwise indicated. Antibiotics and Tet-certified serum were from Clontech. In experiments
SUPPLEMENT Materials and methods Cell culture and reagents Cell media and reagents were from Invitrogen unless otherwise indicated. Antibiotics and Tet-certified serum were from Clontech. In experiments
Table S1. Sequence of human and mouse primers used for RT-qPCR measurements.
 Table S1. Sequence of human and mouse primers used for RT-qPCR measurements. Ca9, carbonic anhydrase IX; Ndrg1, N-myc downstream regulated gene 1; L28, ribosomal protein L28; Hif1a, hypoxia inducible factor
Table S1. Sequence of human and mouse primers used for RT-qPCR measurements. Ca9, carbonic anhydrase IX; Ndrg1, N-myc downstream regulated gene 1; L28, ribosomal protein L28; Hif1a, hypoxia inducible factor
Supplementary Data Table of Contents:
 Supplementary Data Table of Contents: - Supplementary Methods - Supplementary Figures S1(A-B) - Supplementary Figures S2 (A-B) - Supplementary Figures S3 - Supplementary Figures S4(A-B) - Supplementary
Supplementary Data Table of Contents: - Supplementary Methods - Supplementary Figures S1(A-B) - Supplementary Figures S2 (A-B) - Supplementary Figures S3 - Supplementary Figures S4(A-B) - Supplementary
p47 negatively regulates IKK activation by inducing the lysosomal degradation of polyubiquitinated NEMO
 Supplementary Information p47 negatively regulates IKK activation by inducing the lysosomal degradation of polyubiquitinated NEMO Yuri Shibata, Masaaki Oyama, Hiroko Kozuka-Hata, Xiao Han, Yuetsu Tanaka,
Supplementary Information p47 negatively regulates IKK activation by inducing the lysosomal degradation of polyubiquitinated NEMO Yuri Shibata, Masaaki Oyama, Hiroko Kozuka-Hata, Xiao Han, Yuetsu Tanaka,
Supplementary Figure 1. mrna targets were found in exosomes and absent in free-floating supernatant. Serum exosomes and exosome-free supernatant were
 Supplementary Figure 1. mrna targets were found in exosomes and absent in free-floating supernatant. Serum exosomes and exosome-free supernatant were separated via ultracentrifugation and lysed to analyze
Supplementary Figure 1. mrna targets were found in exosomes and absent in free-floating supernatant. Serum exosomes and exosome-free supernatant were separated via ultracentrifugation and lysed to analyze
A Hepatocyte Growth Factor Receptor (Met) Insulin Receptor hybrid governs hepatic glucose metabolism SUPPLEMENTARY FIGURES, LEGENDS AND METHODS
 A Hepatocyte Growth Factor Receptor (Met) Insulin Receptor hybrid governs hepatic glucose metabolism Arlee Fafalios, Jihong Ma, Xinping Tan, John Stoops, Jianhua Luo, Marie C. DeFrances and Reza Zarnegar
A Hepatocyte Growth Factor Receptor (Met) Insulin Receptor hybrid governs hepatic glucose metabolism Arlee Fafalios, Jihong Ma, Xinping Tan, John Stoops, Jianhua Luo, Marie C. DeFrances and Reza Zarnegar
Supplementary Figure S1 Supplementary Figure S2
 Supplementary Figure S A) The blots shown in Figure B were qualified by using Gel-Pro analyzer software (Rockville, MD, USA). The ratio of LC3II/LC3I to actin was then calculated. The data are represented
Supplementary Figure S A) The blots shown in Figure B were qualified by using Gel-Pro analyzer software (Rockville, MD, USA). The ratio of LC3II/LC3I to actin was then calculated. The data are represented
General Laboratory methods Plasma analysis: Gene Expression Analysis: Immunoblot analysis: Immunohistochemistry:
 General Laboratory methods Plasma analysis: Plasma insulin (Mercodia, Sweden), leptin (duoset, R&D Systems Europe, Abingdon, United Kingdom), IL-6, TNFα and adiponectin levels (Quantikine kits, R&D Systems
General Laboratory methods Plasma analysis: Plasma insulin (Mercodia, Sweden), leptin (duoset, R&D Systems Europe, Abingdon, United Kingdom), IL-6, TNFα and adiponectin levels (Quantikine kits, R&D Systems
Protocol for Gene Transfection & Western Blotting
 The schedule and the manual of basic techniques for cell culture Advanced Protocol for Gene Transfection & Western Blotting Schedule Day 1 26/07/2008 Transfection Day 3 28/07/2008 Cell lysis Immunoprecipitation
The schedule and the manual of basic techniques for cell culture Advanced Protocol for Gene Transfection & Western Blotting Schedule Day 1 26/07/2008 Transfection Day 3 28/07/2008 Cell lysis Immunoprecipitation
Supplemental Figure 1. Western blot analysis indicated that MIF was detected in the fractions of
 Supplemental Figure Legends Supplemental Figure 1. Western blot analysis indicated that was detected in the fractions of plasma membrane and cytosol but not in nuclear fraction isolated from Pkd1 null
Supplemental Figure Legends Supplemental Figure 1. Western blot analysis indicated that was detected in the fractions of plasma membrane and cytosol but not in nuclear fraction isolated from Pkd1 null
Supplementary Figure 1. PD-L1 is glycosylated in cancer cells. (a) Western blot analysis of PD-L1 in breast cancer cells. (b) Western blot analysis
 Supplementary Figure 1. PD-L1 is glycosylated in cancer cells. (a) Western blot analysis of PD-L1 in breast cancer cells. (b) Western blot analysis of PD-L1 in ovarian cancer cells. (c) Western blot analysis
Supplementary Figure 1. PD-L1 is glycosylated in cancer cells. (a) Western blot analysis of PD-L1 in breast cancer cells. (b) Western blot analysis of PD-L1 in ovarian cancer cells. (c) Western blot analysis
Supplementary Figure 1 Role of Raf-1 in TLR2-Dectin-1-mediated cytokine expression
 Supplementary Figure 1 Supplementary Figure 1 Role of Raf-1 in TLR2-Dectin-1-mediated cytokine expression. Quantitative real-time PCR of indicated mrnas in DCs stimulated with TLR2-Dectin-1 agonist zymosan
Supplementary Figure 1 Supplementary Figure 1 Role of Raf-1 in TLR2-Dectin-1-mediated cytokine expression. Quantitative real-time PCR of indicated mrnas in DCs stimulated with TLR2-Dectin-1 agonist zymosan
Requires Signaling though Akt2 Independent of the. Transcription Factors FoxA2, FoxO1, and SREBP1c
 Cell Metabolism, Volume 14 Supplemental Information Postprandial Hepatic Lipid Metabolism Requires Signaling though Akt2 Independent of the Transcription Factors FoxA2, FoxO1, and SREBP1c Min Wan, Karla
Cell Metabolism, Volume 14 Supplemental Information Postprandial Hepatic Lipid Metabolism Requires Signaling though Akt2 Independent of the Transcription Factors FoxA2, FoxO1, and SREBP1c Min Wan, Karla
Supplemental Figure 1 ELISA scheme to measure plasma total, mature and furin-cleaved
 1 Supplemental Figure Legends Supplemental Figure 1 ELISA scheme to measure plasma total, mature and furin-cleaved PCSK9 concentrations. 4 Plasma mature and furin-cleaved PCSK9s were measured by a sandwich
1 Supplemental Figure Legends Supplemental Figure 1 ELISA scheme to measure plasma total, mature and furin-cleaved PCSK9 concentrations. 4 Plasma mature and furin-cleaved PCSK9s were measured by a sandwich
Optimization of a LanthaScreen Kinase assay for CSF1R (FMS)
 Optimization of a LanthaScreen Kinase assay for CSF1R (FMS) Overview This protocol describes how to develop a LanthaScreen kinase assay designed to detect and characterize kinase inhibitors. The development
Optimization of a LanthaScreen Kinase assay for CSF1R (FMS) Overview This protocol describes how to develop a LanthaScreen kinase assay designed to detect and characterize kinase inhibitors. The development
Supplementary Information
 Supplementary Information Supplementary Figure 1. CD4 + T cell activation and lack of apoptosis after crosslinking with anti-cd3 + anti-cd28 + anti-cd160. (a) Flow cytometry of anti-cd160 (5D.10A11) binding
Supplementary Information Supplementary Figure 1. CD4 + T cell activation and lack of apoptosis after crosslinking with anti-cd3 + anti-cd28 + anti-cd160. (a) Flow cytometry of anti-cd160 (5D.10A11) binding
Supplementary Figure 1. Establishment of prostacyclin-secreting hmscs. (a) PCR showed the integration of the COX-1-10aa-PGIS transgene into the
 Supplementary Figure 1. Establishment of prostacyclin-secreting hmscs. (a) PCR showed the integration of the COX-1-10aa-PGIS transgene into the genomic DNA of hmscs (PGI2- hmscs). Native hmscs and plasmid
Supplementary Figure 1. Establishment of prostacyclin-secreting hmscs. (a) PCR showed the integration of the COX-1-10aa-PGIS transgene into the genomic DNA of hmscs (PGI2- hmscs). Native hmscs and plasmid
Ginkgo biloba leaf extract induces DNA damage by inhibiting topoisomerase II activity in human hepatic cells
 Ginkgo biloba leaf extract induces DNA damage by inhibiting topoisomerase II activity in human hepatic cells Zhuhong Zhang 1, Si Chen 2, Hu Mei 3, Jiekun Xuan 2, Xiaoqing Guo 1, Letha Couch 2, Vasily N.
Ginkgo biloba leaf extract induces DNA damage by inhibiting topoisomerase II activity in human hepatic cells Zhuhong Zhang 1, Si Chen 2, Hu Mei 3, Jiekun Xuan 2, Xiaoqing Guo 1, Letha Couch 2, Vasily N.
The Schedule and the Manual of Basic Techniques for Cell Culture
 The Schedule and the Manual of Basic Techniques for Cell Culture 1 Materials Calcium Phosphate Transfection Kit: Invitrogen Cat.No.K2780-01 Falcon tube (Cat No.35-2054:12 x 75 mm, 5 ml tube) Cell: 293
The Schedule and the Manual of Basic Techniques for Cell Culture 1 Materials Calcium Phosphate Transfection Kit: Invitrogen Cat.No.K2780-01 Falcon tube (Cat No.35-2054:12 x 75 mm, 5 ml tube) Cell: 293
supplementary information
 Figure S1 Nucleotide binding status of RagA mutants. Wild type and mutant forms of MycRagA was transfected into HEK293 cells and the transfected cells were labeled with 32 Pphosphate. MycRagA was immunoprecipitated
Figure S1 Nucleotide binding status of RagA mutants. Wild type and mutant forms of MycRagA was transfected into HEK293 cells and the transfected cells were labeled with 32 Pphosphate. MycRagA was immunoprecipitated
SUPPLEMENTARY INFORMATION
 Complete but curtailed T-cell response to very-low-affinity antigen Dietmar Zehn, Sarah Y. Lee & Michael J. Bevan Supp. Fig. 1: TCR chain usage among endogenous K b /Ova reactive T cells. C57BL/6 mice
Complete but curtailed T-cell response to very-low-affinity antigen Dietmar Zehn, Sarah Y. Lee & Michael J. Bevan Supp. Fig. 1: TCR chain usage among endogenous K b /Ova reactive T cells. C57BL/6 mice
Figures S1-S5, Figure Legends, Table S1 List of primers used in the study
 Insulin receptor alternative splicing is regulated by insulin signaling and modulates beta cell survival Pushkar Malakar,4, Lital Chartarifsky,4, Ayat Hija, Gil Leibowitz 3, Benjamin Glaser 3, Yuval Dor,
Insulin receptor alternative splicing is regulated by insulin signaling and modulates beta cell survival Pushkar Malakar,4, Lital Chartarifsky,4, Ayat Hija, Gil Leibowitz 3, Benjamin Glaser 3, Yuval Dor,
Validation & Assay Performance Summary
 Validation & Assay Performance Summary LanthaScreen IGF-1R GripTite Cells Cat. no. K1834 Modification Detected: Phosphorylation of Multiple Tyr Residues on IGF-1R LanthaScreen Cellular Assay Validation
Validation & Assay Performance Summary LanthaScreen IGF-1R GripTite Cells Cat. no. K1834 Modification Detected: Phosphorylation of Multiple Tyr Residues on IGF-1R LanthaScreen Cellular Assay Validation
Cells and reagents. Synaptopodin knockdown (1) and dynamin knockdown (2)
 Supplemental Methods Cells and reagents. Synaptopodin knockdown (1) and dynamin knockdown (2) podocytes were cultured as described previously. Staurosporine, angiotensin II and actinomycin D were all obtained
Supplemental Methods Cells and reagents. Synaptopodin knockdown (1) and dynamin knockdown (2) podocytes were cultured as described previously. Staurosporine, angiotensin II and actinomycin D were all obtained
Nature Methods: doi: /nmeth Supplementary Figure 1
 Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
Supplementary Figure 1 Subtiligase-catalyzed ligations with ubiquitin thioesters and 10-mer biotinylated peptides. (a) General scheme for ligations between ubiquitin thioesters and 10-mer, biotinylated
Supporting Information
 Supporting Information Burford et al. 1.173/pnas.1339311 SI Materials and Methods β-arrestin Recruitment Assay. PathHunter human osteosarcoma cells (U2OS) expressing either μ-opioid receptors (U2OS- OPRM1)
Supporting Information Burford et al. 1.173/pnas.1339311 SI Materials and Methods β-arrestin Recruitment Assay. PathHunter human osteosarcoma cells (U2OS) expressing either μ-opioid receptors (U2OS- OPRM1)
Figure S1. Generation of inducible PTEN deficient mice and the BMMCs (A) B6.129 Pten loxp/loxp mice were mated with B6.
 Figure S1. Generation of inducible PTEN deficient mice and the BMMCs (A) B6.129 Pten loxp/loxp mice were mated with B6.129-Gt(ROSA)26Sor tm1(cre/ert2)tyj /J mice. To induce deletion of the Pten locus,
Figure S1. Generation of inducible PTEN deficient mice and the BMMCs (A) B6.129 Pten loxp/loxp mice were mated with B6.129-Gt(ROSA)26Sor tm1(cre/ert2)tyj /J mice. To induce deletion of the Pten locus,
Integrin CD11b negatively regulates TLR-triggered inflammatory responses by. activating Syk and promoting MyD88 and TRIF degradation via cbl-b
 Integrin CD11b negatively regulates TLR-triggered inflammatory responses by activating Syk and promoting MyD88 and TRIF degradation via cbl-b Chaofeng Han, Jing Jin, Sheng Xu, Haibo Liu, Nan Li, and Xuetao
Integrin CD11b negatively regulates TLR-triggered inflammatory responses by activating Syk and promoting MyD88 and TRIF degradation via cbl-b Chaofeng Han, Jing Jin, Sheng Xu, Haibo Liu, Nan Li, and Xuetao
Universal sample preparation method for proteome analysis
 nature methods Universal sample preparation method for proteome analysis Jacek R Wi niewski, Alexandre Zougman, Nagarjuna Nagaraj & Matthias Mann Supplementary figures and text: Supplementary Figure 1
nature methods Universal sample preparation method for proteome analysis Jacek R Wi niewski, Alexandre Zougman, Nagarjuna Nagaraj & Matthias Mann Supplementary figures and text: Supplementary Figure 1
MTC-TT and TPC-1 cell lines were cultured in RPMI medium (Gibco, Breda, The Netherlands)
 Supplemental data Materials and Methods Cell culture MTC-TT and TPC-1 cell lines were cultured in RPMI medium (Gibco, Breda, The Netherlands) supplemented with 15% or 10% (for TPC-1) fetal bovine serum
Supplemental data Materials and Methods Cell culture MTC-TT and TPC-1 cell lines were cultured in RPMI medium (Gibco, Breda, The Netherlands) supplemented with 15% or 10% (for TPC-1) fetal bovine serum
SUPPLEMENTARY INFORMATION
 a c e doi:10.1038/nature10407 b d f Supplementary Figure 1. SERCA2a complex analysis. (a) Two-dimensional SDS-PAGE gels of SERCA2a complexes. A silver-stained SDSPAGE gel is shown, which reveals a 12 kda
a c e doi:10.1038/nature10407 b d f Supplementary Figure 1. SERCA2a complex analysis. (a) Two-dimensional SDS-PAGE gels of SERCA2a complexes. A silver-stained SDSPAGE gel is shown, which reveals a 12 kda
(A) PCR primers (arrows) designed to distinguish wild type (P1+P2), targeted (P1+P2) and excised (P1+P3)14-
 1 Supplemental Figure Legends Figure S1. Mammary tumors of ErbB2 KI mice with 14-3-3σ ablation have elevated ErbB2 transcript levels and cell proliferation (A) PCR primers (arrows) designed to distinguish
1 Supplemental Figure Legends Figure S1. Mammary tumors of ErbB2 KI mice with 14-3-3σ ablation have elevated ErbB2 transcript levels and cell proliferation (A) PCR primers (arrows) designed to distinguish
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature11429 S1a 6 7 8 9 Nlrc4 allele S1b Nlrc4 +/+ Nlrc4 +/F Nlrc4 F/F 9 Targeting construct 422 bp 273 bp FRT-neo-gb-PGK-FRT 3x.STOP S1c Nlrc4 +/+ Nlrc4 F/F casp1
SUPPLEMENTARY INFORMATION doi:10.1038/nature11429 S1a 6 7 8 9 Nlrc4 allele S1b Nlrc4 +/+ Nlrc4 +/F Nlrc4 F/F 9 Targeting construct 422 bp 273 bp FRT-neo-gb-PGK-FRT 3x.STOP S1c Nlrc4 +/+ Nlrc4 F/F casp1
SUPPLEMENTARY INFORMATION. Supplementary Figures S1-S9. Supplementary Methods
 SUPPLEMENTARY INFORMATION SUMO1 modification of PTEN regulates tumorigenesis by controlling its association with the plasma membrane Jian Huang 1,2#, Jie Yan 1,2#, Jian Zhang 3#, Shiguo Zhu 1, Yanli Wang
SUPPLEMENTARY INFORMATION SUMO1 modification of PTEN regulates tumorigenesis by controlling its association with the plasma membrane Jian Huang 1,2#, Jie Yan 1,2#, Jian Zhang 3#, Shiguo Zhu 1, Yanli Wang
Nature Neuroscience: doi: /nn Supplementary Figure 1
 Supplementary Figure 1 EGFR inhibition activates signaling pathways (a-b) EGFR inhibition activates signaling pathways (a) U251EGFR cells were treated with erlotinib (1µM) for the indicated times followed
Supplementary Figure 1 EGFR inhibition activates signaling pathways (a-b) EGFR inhibition activates signaling pathways (a) U251EGFR cells were treated with erlotinib (1µM) for the indicated times followed
Essential Medium, containing 10% fetal bovine serum, 100 U/ml penicillin and 100 µg/ml streptomycin. Huvec were cultured in
 Supplemental data Methods Cell culture media formulations A-431 and U-87 MG cells were maintained in Dulbecco s Modified Eagle s Medium. FaDu cells were cultured in Eagle's Minimum Essential Medium, containing
Supplemental data Methods Cell culture media formulations A-431 and U-87 MG cells were maintained in Dulbecco s Modified Eagle s Medium. FaDu cells were cultured in Eagle's Minimum Essential Medium, containing
(Stratagene, La Jolla, CA) (Supplemental Fig. 1A). A 5.4-kb EcoRI fragment
 SUPPLEMENTAL INFORMATION Supplemental Methods Generation of RyR2-S2808D Mice Murine genomic RyR2 clones were isolated from a 129/SvEvTacfBR λ-phage library (Stratagene, La Jolla, CA) (Supplemental Fig.
SUPPLEMENTAL INFORMATION Supplemental Methods Generation of RyR2-S2808D Mice Murine genomic RyR2 clones were isolated from a 129/SvEvTacfBR λ-phage library (Stratagene, La Jolla, CA) (Supplemental Fig.
Application Note. Introduction
 Simultaneously Measuring Oxidation of Exogenous and Endogenous Fatty Acids Using the XF Palmitate-BSA FAO Substrate with the Agilent Seahorse XF Cell Mito Stress Test Application Note Introduction The
Simultaneously Measuring Oxidation of Exogenous and Endogenous Fatty Acids Using the XF Palmitate-BSA FAO Substrate with the Agilent Seahorse XF Cell Mito Stress Test Application Note Introduction The
Figure S1 Time-dependent down-modulation of HER3 by EZN No Treatment. EZN-3920, 2 μm. Time, h
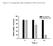 Figure S1 Time-dependent down-modulation of HER3 by EZN-392 HE ER3 mrna A, %Contr rol 12 No Treatment EZN-392, 2 μm 1 8 6 4 2 2 8 24 Time, h Figure S2. Specific target down-modulation by HER3 (EZN-392)
Figure S1 Time-dependent down-modulation of HER3 by EZN-392 HE ER3 mrna A, %Contr rol 12 No Treatment EZN-392, 2 μm 1 8 6 4 2 2 8 24 Time, h Figure S2. Specific target down-modulation by HER3 (EZN-392)
CHAPTER 4 RESULTS. showed that all three replicates had similar growth trends (Figure 4.1) (p<0.05; p=0.0000)
 CHAPTER 4 RESULTS 4.1 Growth Characterization of C. vulgaris 4.1.1 Optical Density Growth study of Chlorella vulgaris based on optical density at 620 nm (OD 620 ) showed that all three replicates had similar
CHAPTER 4 RESULTS 4.1 Growth Characterization of C. vulgaris 4.1.1 Optical Density Growth study of Chlorella vulgaris based on optical density at 620 nm (OD 620 ) showed that all three replicates had similar
Tel: ; Fax: ;
 Tel.: +98 216 696 9291; Fax: +98 216 696 9291; E-mail: mrasadeghi@pasteur.ac.ir Tel: +98 916 113 7679; Fax: +98 613 333 6380; E-mail: abakhshi_e@ajums.ac.ir A Soluble Chromatin-bound MOI 0 1 5 0 1 5 HDAC2
Tel.: +98 216 696 9291; Fax: +98 216 696 9291; E-mail: mrasadeghi@pasteur.ac.ir Tel: +98 916 113 7679; Fax: +98 613 333 6380; E-mail: abakhshi_e@ajums.ac.ir A Soluble Chromatin-bound MOI 0 1 5 0 1 5 HDAC2
Supplementary Information
 Supplementary Information Akt regulates hepatic metabolism by suppressing a Foxo1 dependent global inhibition of adaptation to nutrient intake Mingjian Lu 1, Min Wan 1, Karla F. Leavens 1, Qingwei Chu
Supplementary Information Akt regulates hepatic metabolism by suppressing a Foxo1 dependent global inhibition of adaptation to nutrient intake Mingjian Lu 1, Min Wan 1, Karla F. Leavens 1, Qingwei Chu
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11095 Supplementary Table 1. Summary of the binding between Angptls and various Igdomain containing receptors as determined by flow cytometry analysis. The results were summarized from
doi:10.1038/nature11095 Supplementary Table 1. Summary of the binding between Angptls and various Igdomain containing receptors as determined by flow cytometry analysis. The results were summarized from
Supplementary information. The Light Intermediate Chain 2 Subpopulation of Dynein Regulates Mitotic. Spindle Orientation
 Supplementary information The Light Intermediate Chain 2 Subpopulation of Dynein Regulates Mitotic Spindle Orientation Running title: Dynein LICs distribute mitotic functions. Sagar Mahale a, d, *, Megha
Supplementary information The Light Intermediate Chain 2 Subpopulation of Dynein Regulates Mitotic Spindle Orientation Running title: Dynein LICs distribute mitotic functions. Sagar Mahale a, d, *, Megha
Supplementary Figure 1 IL-27 IL
 Tim-3 Supplementary Figure 1 Tc0 49.5 0.6 Tc1 63.5 0.84 Un 49.8 0.16 35.5 0.16 10 4 61.2 5.53 10 3 64.5 5.66 10 2 10 1 10 0 31 2.22 10 0 10 1 10 2 10 3 10 4 IL-10 28.2 1.69 IL-27 Supplementary Figure 1.
Tim-3 Supplementary Figure 1 Tc0 49.5 0.6 Tc1 63.5 0.84 Un 49.8 0.16 35.5 0.16 10 4 61.2 5.53 10 3 64.5 5.66 10 2 10 1 10 0 31 2.22 10 0 10 1 10 2 10 3 10 4 IL-10 28.2 1.69 IL-27 Supplementary Figure 1.
(a) Significant biological processes (upper panel) and disease biomarkers (lower panel)
 Supplementary Figure 1. Functional enrichment analyses of secretomic proteins. (a) Significant biological processes (upper panel) and disease biomarkers (lower panel) 2 involved by hrab37-mediated secretory
Supplementary Figure 1. Functional enrichment analyses of secretomic proteins. (a) Significant biological processes (upper panel) and disease biomarkers (lower panel) 2 involved by hrab37-mediated secretory
Supplementary Figure 1: si-craf but not si-braf sensitizes tumor cells to radiation.
 Supplementary Figure 1: si-craf but not si-braf sensitizes tumor cells to radiation. (a) Embryonic fibroblasts isolated from wildtype (WT), BRAF -/-, or CRAF -/- mice were irradiated (6 Gy) and DNA damage
Supplementary Figure 1: si-craf but not si-braf sensitizes tumor cells to radiation. (a) Embryonic fibroblasts isolated from wildtype (WT), BRAF -/-, or CRAF -/- mice were irradiated (6 Gy) and DNA damage
AlphaScreen TNFα Binding Assay Kit: A Homogeneous, Sensitive and High-Throughput Assay for Screening TNFα Receptors
 AlphaScreen TNFα Binding Assay Kit: A Homogeneous, Sensitive and High-Throughput Assay for Screening TNFα Receptors Bouchard N., Legault M. and Wenham D. PerkinElmer BioSignal, 1744 William, suite 3, Montréal,
AlphaScreen TNFα Binding Assay Kit: A Homogeneous, Sensitive and High-Throughput Assay for Screening TNFα Receptors Bouchard N., Legault M. and Wenham D. PerkinElmer BioSignal, 1744 William, suite 3, Montréal,
SUPPLEMENTARY FIGURES AND TABLE
 SUPPLEMENTARY FIGURES AND TABLE Supplementary Figure S1: Characterization of IRE1α mutants. A. U87-LUC cells were transduced with the lentiviral vector containing the GFP sequence (U87-LUC Tet-ON GFP).
SUPPLEMENTARY FIGURES AND TABLE Supplementary Figure S1: Characterization of IRE1α mutants. A. U87-LUC cells were transduced with the lentiviral vector containing the GFP sequence (U87-LUC Tet-ON GFP).
Protocol for A-549 VIM RFP (ATCC CCL-185EMT) TGFβ1 EMT Induction and Drug Screening
 Protocol for A-549 VIM RFP (ATCC CCL-185EMT) TGFβ1 EMT Induction and Drug Screening Introduction: Vimentin (VIM) intermediate filament (IF) proteins are associated with EMT in lung cancer and its metastatic
Protocol for A-549 VIM RFP (ATCC CCL-185EMT) TGFβ1 EMT Induction and Drug Screening Introduction: Vimentin (VIM) intermediate filament (IF) proteins are associated with EMT in lung cancer and its metastatic
Data Sheet PD-1 / NFAT Reporter - Jurkat Cell Line Catalog #: 60535
 Data Sheet PD-1 / NFAT Reporter - Jurkat Cell Line Catalog #: 60535 Product Description Recombinant Jurkat T cell expressing firefly luciferase gene under the control of NFAT response elements with constitutive
Data Sheet PD-1 / NFAT Reporter - Jurkat Cell Line Catalog #: 60535 Product Description Recombinant Jurkat T cell expressing firefly luciferase gene under the control of NFAT response elements with constitutive
Defective Hepatic Autophagy in Obesity Promotes ER Stress and Causes Insulin Resistance
 Cell Metabolism, Volume 11 Supplemental Information Defective Hepatic Autophagy in Obesity Promotes ER Stress and Causes Insulin Resistance Ling Yang, Ping Li, Suneng Fu, Ediz S. Calay, and Gökhan S. Hotamisligil
Cell Metabolism, Volume 11 Supplemental Information Defective Hepatic Autophagy in Obesity Promotes ER Stress and Causes Insulin Resistance Ling Yang, Ping Li, Suneng Fu, Ediz S. Calay, and Gökhan S. Hotamisligil
Supplementary information
 Supplementary information Human Cytomegalovirus MicroRNA mir-us4-1 Inhibits CD8 + T Cell Response by Targeting ERAP1 Sungchul Kim, Sanghyun Lee, Jinwook Shin, Youngkyun Kim, Irini Evnouchidou, Donghyun
Supplementary information Human Cytomegalovirus MicroRNA mir-us4-1 Inhibits CD8 + T Cell Response by Targeting ERAP1 Sungchul Kim, Sanghyun Lee, Jinwook Shin, Youngkyun Kim, Irini Evnouchidou, Donghyun
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/8/407/ra127/dc1 Supplementary Materials for Loss of FTO in adipose tissue decreases Angptl4 translation and alters triglyceride metabolism Chao-Yung Wang,* Shian-Sen
www.sciencesignaling.org/cgi/content/full/8/407/ra127/dc1 Supplementary Materials for Loss of FTO in adipose tissue decreases Angptl4 translation and alters triglyceride metabolism Chao-Yung Wang,* Shian-Sen
Supplementary information. MARCH8 inhibits HIV-1 infection by reducing virion incorporation of envelope glycoproteins
 Supplementary information inhibits HIV-1 infection by reducing virion incorporation of envelope glycoproteins Takuya Tada, Yanzhao Zhang, Takayoshi Koyama, Minoru Tobiume, Yasuko Tsunetsugu-Yokota, Shoji
Supplementary information inhibits HIV-1 infection by reducing virion incorporation of envelope glycoproteins Takuya Tada, Yanzhao Zhang, Takayoshi Koyama, Minoru Tobiume, Yasuko Tsunetsugu-Yokota, Shoji
a surface permeabilized
 a surface permeabilized RAW 64.7 P388D1 J774 b CD11b + Ly-6G - Blood Monocytes WT Supplementary Figure 1. Cell surface expression on macrophages and DCs. (a) RAW64.7, P388D1, and J774 cells were subjected
a surface permeabilized RAW 64.7 P388D1 J774 b CD11b + Ly-6G - Blood Monocytes WT Supplementary Figure 1. Cell surface expression on macrophages and DCs. (a) RAW64.7, P388D1, and J774 cells were subjected
SUPPLEMENTAL MATERIAL. Supplementary Methods
 SUPPLEMENTAL MATERIAL Supplementary Methods Culture of cardiomyocytes, fibroblasts and cardiac microvascular endothelial cells The isolation and culturing of neonatal rat ventricular cardiomyocytes was
SUPPLEMENTAL MATERIAL Supplementary Methods Culture of cardiomyocytes, fibroblasts and cardiac microvascular endothelial cells The isolation and culturing of neonatal rat ventricular cardiomyocytes was
Supplementary Fig. S1. Schematic diagram of minigenome segments.
 open reading frame 1565 (segment 5) 47 (-) 3 5 (+) 76 101 125 149 173 197 221 246 287 open reading frame 890 (segment 8) 60 (-) 3 5 (+) 172 Supplementary Fig. S1. Schematic diagram of minigenome segments.
open reading frame 1565 (segment 5) 47 (-) 3 5 (+) 76 101 125 149 173 197 221 246 287 open reading frame 890 (segment 8) 60 (-) 3 5 (+) 172 Supplementary Fig. S1. Schematic diagram of minigenome segments.
