SUPPLEMENTARY INFORMATION
|
|
|
- Jeremy Chambers
- 5 years ago
- Views:
Transcription
1 doi: /nature07422 SUPPLEMENTARY INFRMATIN K S(P) R S I M Q(L4) R M 7 6 Sp Q I K R L(L0) L L S E Figure S1a Difference electron density (mfo DFc) for the peptide (Qpeptide), from the first round of refinement after molecular replacement before building the peptide into the model. Contour level 0.08 e/å3. The electron density is clear for the two leucine residues and the main chain, but less so for the glutamine (4) side chain. This and all other structure figures were made using CCP4mg (Potterton, L. et al. Developments in the CCP4 moleculargraphics project. Acta Crystallogr D Biol Crystallogr 60, (2004)). Figure S1b Sequence of the Qpeptide with the numbering scheme relative to the first leucine as 0. The Epeptide has Q(L4) replaced by E Figure S2 A sample of the final refined 2mFo DFc electron density map for the Qpeptide structure, through the βsheet in the subunit, contoured at 0.18 e/å3 1
2 SUPPLEMENTARY INFRMATIN Dataset Q peptide E peptide Space group, unit cell (a, c) P43212, 169.9, Å P43212, 171.2,324.3 Å Wavelength Å Å Rotation range (nearest axis to spindle) 112 (c*) 114 (a*) Resolution range Å (high resolution) ( ) ( ) Rmerge (1.17) (0.96) Rmerge in top intensity bin Rmeas (1.32) (1.06) Rpim (0.42) Number of observations (unique) (96683) (64153) < <Ih> / σ(<ih>) > 8.6 (1.8) 7.8 (1.9) Completeness 100.0% (100.0%) 95.8% (89.1%) Multiplicity 8.8 (9.0) 6.4 (5.0) Wilson <B> 60Å 2 58 Å 2 Refinement Rwork (0.33) 0.24 (0.33) Rfree (0.37) (0.38)) Number of reflections (test set) (9168) (5902) RMSD bonds (angles) 0.007Å (1.1 ) 0.009Å (1.3 ) Number of TLS groups <Biso> (peptides) 79Å 2 (84Å 2 ) 86Å 2 (108Å 2 ) Number of atoms Number of sulphate ions (waters) 34 (26) 34 (26) Supplementary Table S1: Data collection & refinement statistics. Rmerge = Σhl Ihl <Ih> / Σhl <Ih> ; Rmeas = Σhl [nh/( nh1)] 1/2 Ihl <Ih> / Σhl <Ih> R pim = Σhl [1/( nh1)] 1/2 Ihl <Ih> / Σhl <Ih> where nh is the number of observations of reflection h. 2
3 SUPPLEMENTARY INFRMATIN R11 S 4 2 InsP 6 S 4 2 E36 Y53 K57 K61 Y58 E(L4) L(L0) Figure S3 Refined electron density for the two sulphate ions (contoured at 0.18 e/å 3, cropped around the sulphates) which bind to the α subunit, mimicking the binding of InsP6 seen in the inactive structure (thin bonds). All labelled side chains are in the α subunit. Figure S4 Difference electron density (mfo DFc) for the Epeptide from the first round of refinement after molecular replacement before building the peptide into the model, contoured at 0.06 e/å 3. The structure of the Qpeptide is shown with thin bonds. wt core V88D V98S V98F L103S A63W N92W L65S Y62S E100Y R15S/αR21E R15S/αR21S dileucine signal binding (tested for CD4 and LRP9) Supplementary Table S2. : Binding of AP2 core mutants to dileucine sorting signals. The indicated AP2 core mutants were tested for binding to the dileucine signals of CD4 and LRP9 using the biosensorbased membrane mimic assay. ++ +: level of binding displayed by wt core, ++: affinity reduced by less than 5fold; +: 510 fold reduced affinity; : no detectable binding. We did not observe any obvious difference in binding to the dileucine signals of CD4 and LRP9, except that binding to the latter was generally slightly stronger (see table S3 for details). 3
4 SUPPLEMENTARY INFRMATIN Membrane protein L +1 Affinity for wt AP2 core (μm) phosphocd4 Qpeptide SQIKRLL 0.85 phosphocd4 Epeptide SEIKRLL 0.65 Tyrosinase EEKQPLL 0.6 TRP1 EANQPLL 0.75 LimpII DERAPLI 0.9 LRP9 AEDEPLL 0.55 Supplementary Table S3. Binding of AP2 core to a range of different dileucine signals. Binding of AP2 cores to a number of different 'acidic dileucine' motifs displayed in PtdIns4,5P2 containing liposomes was recorded in the biosensorbased membrane mimic assay and the rate constants were determined as described. Irrespective of the type of dileucine signal analysed, the KDs for binding were in the range of µm. Binding experiments to liposomes containing control peptides with mutated dileucine signals (LL replaced by AA) served as negative controls except in the case of LimpII where no peptide was available) showed that there was no binding to acidic dileucine motifs if the L[LI] group is replaced by AA (data not shown). This agrees with in vivo studies ( Letourneur, F. & Klausner, R. D. A novel dileucine motif and a tyrosinebased motif independently mediate lysosomal targeting and endocytosis of CD3 chains. Cell 69, (1992)) and our unpublished data showing that a reporter construct consisting of the lumenal and transmembrane regions of CD8 fused to the cytosolic signal of LRP (cytosolic sequence SHAGSRATEASEDEPLLSS) was endocytosed from the plasma membrane whereas replacing LL with AA prevented uptake of a reporter construct without affecting YxxΦbased endocytosis of transferrin (data not shown). 4
5 SUPPLEMENTARY INFRMATIN Figure S5 Liposome capture and analysis. a : Sequential injection of the indicated types of liposomes on the 4 flowcells of a BIAcore L1 sensor surface. After capture all flowcells were treated with NaH to generate a stable surface for further injections. b: After liposome capture as in a, the ENTHdomain of epsin1 was passed over all 4 flowcells to reveal the amount of PtdIns4,5P2 in the different liposomes. Note that the peak of the sensorgrams differ by less than 20RU demonstrating equal amounts of PtdIns4,5P2 in the 4 different liposome preparations. c: Superimposition of sensorgrams obtained for the binding of varying concentrations of the wt AP2 core to the phosphorylated CD4Q peptide. The plots were further used to calculate the rate constants and KD of the interaction and similar plots were used to derive all KD values mentioned throughout the manuscript. 50kDa 25kDa ! trunkgst "2 trunk µ2 GST contaminant Figure S6 Coomassie brilliant blue stained SDS PAGE Gel showing GST core proteins used for KD determinations shown in Figure3. Lane1 Molecular weight markers, lane2 wt AP2 core, lane3 V88D, lane4 L103S, lane5 R15S/ αr21s lane6 R15S/αR21E #2 5
6 SUPPLEMENTARY INFRMATIN Figure S7 ** * *** * * MIRFILIQNRAGKTRLAKWYMQFDDDEKQKLIEEVHAVVTVRDAKHTNFVEFRNFKIIYRRYAGLYFC MMRFMLLFSRQGKLRLQKWYLATSDKERKKMVRELMQVVLARKPKMCSFLEWRDLKVVYKRYASLYFC MIKAILIFNNHGKPRLSKFYQPYSEDTQQPIIRETFHLVSKRDENVCNFLEGGLLIGGSDNKLIYRHYATLYFV MIKFFLMVNKQGQTRLSKYYEHVDINKRALLETEVSKSCLSRSSEQCSFIEYKDFKLIYRQYAALFVV * * * * ******* ICVDVNDNNLAYLEAIHNFVEVLNEYFHNVCELDLVFNFYKVYTVVDEMFLAGEIRETSQTKVLKQLLMLQSLE CAIEGQDNELITLELIHRYVELLDKYFGSVCELDIIFNFEKAYFILDEFLMGGDVQDTSKKSVLKAIEQADLLQ FCVDSSESELGILDLIQVFVETLDKCFENVCELDLIFHVDKVHNILAEMVMGGMVLETNMNEIVTQIDAQNKLE VGVNDTENEMAIYEFIHNFVEVLDGYFSRVSELDIMFNLDKVHIILDEMVLNGCIVETNRARILAPLLILDKLS a The residues involved in binding the acidic and dihydrophobic parts of acidic dileucine motifs are conserved in all σ subunits from mammalian AP complexes. Mammalian (1142), (1124), (1148) and (1142) were aligned using ClustalW. The resulting output was coloured as follows: deep purple indicates absolute conservation, purple indicates a high degree of conservation and light purple indicates a modest degree of conservation. The results have been plotted on surface representations of in the Q peptide complex in figure 2. residues are numbered beneath the alignment. Coloured asterisks indicate residues involved in binding to the CD4 peptide; blue asterisks mark residues involved in binding to the 4 position peptide residue (Q in this case); green asterisks mark residues involved in binding the LL motif and black asterisks indicate residues contacting other parts of the peptide. Mutations in residues marked with a black arrow abrogate or greatly diminish binding to dileucine peptides. ** * *** * * M. musculus MIRFILIQNRAGKTRLAKWYMQFDDDEKQKLIEEVHAVVTVRDAKHTNFVEFRNFKIIYRRYAGLYFCICV A. thaliana MIRFILLQNRQGKTRLAKYYVPLEESEKHKVEYEVHRLVVNRDAKFTNFVEFRTHKVIYRRYAGLFFSVCV D. discoideum MIHFILIQNRQGKTRLSKWYTPYEDVEKRKLSHEIHKIVNSRETKFTNFVEFRTHRIVYRRYAGLFFSVCV X. laevis MIRFILIQNRAGKTRLAKWYMQFDDDEKQKLIEEVHAVVTVRDAKHTNFVEFRNFKIIYRRYAGLYFCICV D. melanogaster MIRFILIQNRAGKTRLAKWYMNFDDDEKQKLIEEVHAVVTVRDAKHTNFVEFRNFKIVYRRYAGLYFCICV C. elegans MIRFILIQNRAGKTRLAKWYMHFDDDEKQKLIEEVHACVTVRDAKHTNFVEFRNFKIVYRRYAGLYFCICV S. pombe MIQFILIQNRHGKNRLSKYYVPFDDDEKVRLKARIHQLISQRNQKFQANFLEWENSKLVYRRYAGLYFCFCV * ** * ******* M. musculus DVNDNNLAYLEAIHNFVEVLNEYFHNVCELDLVFNFYKVYTVVDEMFLAGEIRETSQTKVLKQLLMLQSLE A. thaliana DITDNELAYLESIHLFVEILDHFFSNVCELDLVFNFHKVYLILDEFILAGELQETSKRAIIERMSELEKLQ D. discoideum DPTDNELFCLEAIHLFVEVLDAYFGNVCELDLVFNFYKVYAIIDEVFLAGELMEPSKHVILQRMEFLDNLP X. laevis DVNDNNLAYLEAIHNFVEVLNEYFHNVCELDLVFNFYKVYTVVDEMFLAGEIRETSQTKVLKQLLMLQSLE D. melanogaster DVNDNNLCYLEAIHNFVEVLNEYFHNVCELDLVFNFYKVYSVVDEMFLAGEIRETSQTKVLKQLLTLNSLE C. elegans DITDNNLYYLEAIHNFVEVLNEYFHNVCELDLVFNFYKVYTVVDEMFLAGEIRETSQTKVLKQLLMLTSLE S. pombe DSTDNDLAILEMIHFFVEILDSFFGNVCELDLIFNFYKVSAILDEIILGGEIGESNKKSVLERIEALEKLE b The residues involved in binding the acidic and dihydrophobic parts of acidic dileucine motifs are conserved in subunits from a variety of species from yeast to mammals. (residues 1142) proteins from various species were aligned using ClustalW and the resulting output coloured and annotated as in a. M. musculus residues are numbered beneath the alignment. ζcp ζcp ** * *** * * MIRFILIQNRAGKTRLAKWYMQFDDDEKQKLIEEVHAVVTVRDAKHTNFVEFRNFKIIYRRYAGLYFCI MMRFMLLFSRQGKLRLQKWYLATSDKERKKMVRELMQVVLARKPKMCSFLEWRDLKVVYKRYASLYFCC MIKAILIFNNHGKPRLSKFYQPYSEDTQQPIIRETFHLVSKRDENVCNFLEGGLLIGGSDNKLIYRHYATLYFVF MIKFFLMVNKQGQTRLSKYYEHVDINKRALLETEVSKSCLSRSSEQCSFIEYKDFKLIYRQYAALFVVV VKAILILDNDGDRLFAKYYDDTYPSVKEQKAFEKNIFNKTHRTDSEIALLEGLTVVYKSSIDLYFYV * ** * * **** ** CVDVNDNNLAYLEAIHNFVEVLNEYFHNVCELDLVFNFYKVYTVVDEMFLAGEIRETSQTKVLKQLLMLQSLE AIEGQDNELITLELIHRYVELLDKYFGSVCELDIIFNFEKAYFILDEFLMGGDVQDTSKKSVLKAIEQADLLQ CVDSSESELGILDLIQVFVETLDKCFENVCELDLIFHVDKVHNILAEMVMGGMVLETNMNEIVTQIDAQNKLE GVNDTENEMAIYEFIHNFVEVLDGYFSRVSELDIMFNLDKVHIILDEMVLNGCIVETNRARILAPLLILDKLS IGSSYENELMLMAVLNCLFDSLSQMLRKNVEKRALLENMEGLFLAVDEIVDGGVILESDPQQVVHRVALRGEDV c The residues involved in binding only the dihydrophobic part of the acidic dileucine motif are conserved between mammalian σ subunits and the homologous ζcp. The sequences of mammalian (1142), (1124), (1148) and (1142) and ζcp (13153) (PDB id 2HF6) were aligned using ClustalW and the resulting output coloured and annotated as in a. 6
7 SUPPLEMENTARY INFRMATIN a b!r21 V98 Q(L4) L103 L(L0) R15 E100 A63 L65 N92 V88 Y62 Figure S8 a Structure of the dileucine peptide binding site showing the peptide in gold. Residues coloured red are those whose mutation strongly inhibits dileucine peptide binding whilst not affecting YxxΦ motif binding (See table S3). Labels for mutated residues are coloured pale blue for and dark blue for α. The protein is displayed as a semitransparent surface coloured red where underlying mutations effect dileucine peptide binding. c "2L103 "2L65 H NH Q(L4) T9 NH #2 Nterminus T8 H H N F7 "2V98 "2F67 NH L(L0) N H H 2 N "2N92 Y6 "2Y62 H "2E89 "2V88 b The same as a with the surface drawn solid. c Schematic diagram of the contacts made by the β2 Nterminus to the LL peptide site on in the inactive IP6liganded structure. The hydrophobic residues Y6 and F7 occupy equivalent sites to the two leucines L+1 and L0 in the peptide, with the main chain running in the opposite direction (compare with figures 2b and 4c). ther interactions are not conserved. CIMPR peptide SDEDLLHI D D E L L Q I L L R K CD4 peptide SQIKRLLS VHS domain of GGA AP2,!2 & " Figure S9 The mechanisms of binding of [ED]xxxL[LI] to /α of AP2 and of DxxLL to the GGA VHS domain are completely different, and are based on different underlying structures. a DxxLL motif in gold binding to the GGA VHS domain (purple) b [ED]xxxL[LI] motif shown in gold binding to (pale blue) and α(dark blue). The peptides are shown in approximately the same orientation 7
Supporting Information
 Supporting Information McCullough et al. 10.1073/pnas.0801567105 A α10 α8 α9 N α7 α6 α5 C β2 β1 α4 α3 α2 α1 C B N C Fig. S1. ALIX Bro1 in complex with the C-terminal CHMP4A helix. (A) Ribbon diagram showing
Supporting Information McCullough et al. 10.1073/pnas.0801567105 A α10 α8 α9 N α7 α6 α5 C β2 β1 α4 α3 α2 α1 C B N C Fig. S1. ALIX Bro1 in complex with the C-terminal CHMP4A helix. (A) Ribbon diagram showing
Table S1. X-ray data collection and refinement statistics
 Table S1. X-ray data collection and refinement statistics Data collection H7.167 Fab-Sh2/H7 complex Beamline SSRL 12-2 Wavelength (Å) 0.97950 Space group I2 1 3 Unit cell parameters (Å, º) a = b = c=207.3,
Table S1. X-ray data collection and refinement statistics Data collection H7.167 Fab-Sh2/H7 complex Beamline SSRL 12-2 Wavelength (Å) 0.97950 Space group I2 1 3 Unit cell parameters (Å, º) a = b = c=207.3,
(B D) Three views of the final refined 2Fo-Fc electron density map of the Vpr (red)-ung2 (green) interacting region, contoured at 1.4σ.
 Supplementary Figure 1 Overall structure of the DDB1 DCAF1 Vpr UNG2 complex. (A) The final refined 2Fo-Fc electron density map, contoured at 1.4σ of Vpr, illustrating well-defined side chains. (B D) Three
Supplementary Figure 1 Overall structure of the DDB1 DCAF1 Vpr UNG2 complex. (A) The final refined 2Fo-Fc electron density map, contoured at 1.4σ of Vpr, illustrating well-defined side chains. (B D) Three
Supplementary Table 1. Data collection and refinement statistics (molecular replacement).
 Supplementary Table 1. Data collection and refinement statistics (molecular replacement). Data set statistics HLA A*0201- ALWGPDPAAA PPI TCR PPI TCR/A2- ALWGPDPAAA PPI TCR/A2- ALWGPDPAAA Space Group P2
Supplementary Table 1. Data collection and refinement statistics (molecular replacement). Data set statistics HLA A*0201- ALWGPDPAAA PPI TCR PPI TCR/A2- ALWGPDPAAA PPI TCR/A2- ALWGPDPAAA Space Group P2
Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site.
 Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
Supplementary Figure 1 Detergent solubilised 5 TMD binds pregnanolone at the Q245 neurosteroid potentiation site. (a) Gel filtration profiles of purified 5 TMD samples at 100 nm, heated beforehand for
Supplementary Information Janssen et al.
 Supplementary Information Janssen et al. Insights into complement convertase formation based on the structure of the factor B CVF complex Bert J.C. Janssen 1, Lucio Gomes 1, Roman I. Koning 2, Dmitri I.
Supplementary Information Janssen et al. Insights into complement convertase formation based on the structure of the factor B CVF complex Bert J.C. Janssen 1, Lucio Gomes 1, Roman I. Koning 2, Dmitri I.
Nature Structural & Molecular Biology: doi: /nsmb.1933
 The structural basis of open channel block in a prokaryotic pentameric ligand-gated ion channel Ricarda J. C. Hilf, Carlo Bertozzi, Iwan Zimmermann, Alwin Reiter, Dirk Trauner and Raimund Dutzler a GLIC
The structural basis of open channel block in a prokaryotic pentameric ligand-gated ion channel Ricarda J. C. Hilf, Carlo Bertozzi, Iwan Zimmermann, Alwin Reiter, Dirk Trauner and Raimund Dutzler a GLIC
FCC2 5CY7 FCC1 5CY8. Actinonin 5CVQ
 Table S1. Data collection and refinement statistics Data collection Apo 5E5D MA 5CPD MAS 5CP0 Actinonin 5CVQ FCC1 5CY8 FCC2 5CY7 FCC 5CWY FCC4 5CVK FCC5 5CWX FCC6 5CVP X-ray source 17A-KEK PAL4A PAL4A
Table S1. Data collection and refinement statistics Data collection Apo 5E5D MA 5CPD MAS 5CP0 Actinonin 5CVQ FCC1 5CY8 FCC2 5CY7 FCC 5CWY FCC4 5CVK FCC5 5CWX FCC6 5CVP X-ray source 17A-KEK PAL4A PAL4A
Supplementary Materials for
 www.sciencemag.org/cgi/content/full/science.aal4326/dc1 Supplementary Materials for Structure of a eukaryotic voltage-gated sodium channel at near-atomic resolution Huaizong Shen, Qiang Zhou, Xiaojing
www.sciencemag.org/cgi/content/full/science.aal4326/dc1 Supplementary Materials for Structure of a eukaryotic voltage-gated sodium channel at near-atomic resolution Huaizong Shen, Qiang Zhou, Xiaojing
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/2/9/e1600292/dc1 Supplementary Materials for Native phasing of x-ray free-electron laser data for a G protein coupled receptor Alexander Batyuk, Lorenzo Galli,
advances.sciencemag.org/cgi/content/full/2/9/e1600292/dc1 Supplementary Materials for Native phasing of x-ray free-electron laser data for a G protein coupled receptor Alexander Batyuk, Lorenzo Galli,
Supplementary Information
 Supplementary Information Two common structural motifs for TCR recognition by staphylococcal enterotoxins Karin Erica Johanna Rödström 1, Paulina Regenthal 1, Christopher Bahl 2, Alex Ford 2, David Baker
Supplementary Information Two common structural motifs for TCR recognition by staphylococcal enterotoxins Karin Erica Johanna Rödström 1, Paulina Regenthal 1, Christopher Bahl 2, Alex Ford 2, David Baker
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/2/4/e1500980/dc1 Supplementary Materials for The crystal structure of human dopamine -hydroxylase at 2.9 Å resolution Trine V. Vendelboe, Pernille Harris, Yuguang
advances.sciencemag.org/cgi/content/full/2/4/e1500980/dc1 Supplementary Materials for The crystal structure of human dopamine -hydroxylase at 2.9 Å resolution Trine V. Vendelboe, Pernille Harris, Yuguang
Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation
 Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation Hang Yu, 1, 2, a) Wei Han, 1, 3, b) Wen Ma, 1, 2 1, 2, 3, c) and Klaus Schulten 1) Beckman
Transient β-hairpin Formation in α-synuclein Monomer Revealed by Coarse-grained Molecular Dynamics Simulation Hang Yu, 1, 2, a) Wei Han, 1, 3, b) Wen Ma, 1, 2 1, 2, 3, c) and Klaus Schulten 1) Beckman
SUPPLEMENTAL MATERIAL. UNC119 is required for G protein trafficking in sensory neurons
 1 SUPPLEMENTAL MATERIAL UNC119 is required for G protein trafficking in sensory neurons Houbin Zhang, Ryan N. Constantine, Sergey Vorobiev, Yang Chen, Jayaraman Seetharaman, Yuanpeng Janet Huang, Rong
1 SUPPLEMENTAL MATERIAL UNC119 is required for G protein trafficking in sensory neurons Houbin Zhang, Ryan N. Constantine, Sergey Vorobiev, Yang Chen, Jayaraman Seetharaman, Yuanpeng Janet Huang, Rong
Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random
 S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
S1 Supplementary Figure 1 (previous page). EM analysis of full-length GCGR. (a) Exemplary tilt pair images of the GCGR mab23 complex acquired for Random Conical Tilt (RCT) reconstruction (left: -50,right:
Supplementary Materials. High affinity binding of phosphatidylinositol-4-phosphate. by Legionella pneumophila DrrA
 Supplementary Materials High affinity binding of phosphatidylinositol-4-phosphate by Legionella pneumophila DrrA Running title: Molecular basis of PtdIns(4)P-binding by DrrA Stefan Schoebel, Wulf Blankenfeldt,
Supplementary Materials High affinity binding of phosphatidylinositol-4-phosphate by Legionella pneumophila DrrA Running title: Molecular basis of PtdIns(4)P-binding by DrrA Stefan Schoebel, Wulf Blankenfeldt,
The antiparasitic drug ivermectin is a novel FXR ligand that regulates metabolism
 Supplementary Information The antiparasitic drug ivermectin is a novel FXR ligand that regulates metabolism Address correspondence to Yong Li (yongli@xmu.edu.cn, Tel: 86-592-218151) GW464 CDCA Supplementary
Supplementary Information The antiparasitic drug ivermectin is a novel FXR ligand that regulates metabolism Address correspondence to Yong Li (yongli@xmu.edu.cn, Tel: 86-592-218151) GW464 CDCA Supplementary
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Design of isolated protein and RNC constructs, and homogeneity of purified RNCs. (a) Schematic depicting the design and nomenclature used for all the isolated proteins and RNCs used
Supplementary Figure 1 Design of isolated protein and RNC constructs, and homogeneity of purified RNCs. (a) Schematic depicting the design and nomenclature used for all the isolated proteins and RNCs used
The clathrin adaptor Numb regulates intestinal cholesterol. absorption through dynamic interaction with NPC1L1
 The clathrin adaptor Numb regulates intestinal cholesterol absorption through dynamic interaction with NPC1L1 Pei-Shan Li 1, Zhen-Yan Fu 1,2, Ying-Yu Zhang 1, Jin-Hui Zhang 1, Chen-Qi Xu 1, Yi-Tong Ma
The clathrin adaptor Numb regulates intestinal cholesterol absorption through dynamic interaction with NPC1L1 Pei-Shan Li 1, Zhen-Yan Fu 1,2, Ying-Yu Zhang 1, Jin-Hui Zhang 1, Chen-Qi Xu 1, Yi-Tong Ma
Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular
 Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular weight markers (M). Supplementary Figure-2. Overlay of
Supplementary Figure-1. SDS PAGE analysis of purified designed carbonic anhydrase enzymes. M1-M4 shown in lanes 1-4, respectively, with molecular weight markers (M). Supplementary Figure-2. Overlay of
Atypical Natural Killer T-cell receptor recognition of CD1d-lipid antigens supplementary Information.
 Atypical Natural Killer T-cell receptor recognition of CD1d-lipid antigens supplementary Information. Supplementary Figure 1. Phenotypic analysis of TRBV25-1 + and TRBV25-1 - CD1d-α-GalCerreactive cells.
Atypical Natural Killer T-cell receptor recognition of CD1d-lipid antigens supplementary Information. Supplementary Figure 1. Phenotypic analysis of TRBV25-1 + and TRBV25-1 - CD1d-α-GalCerreactive cells.
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10962 Supplementary Figure 1. Expression of AvrAC-FLAG in protoplasts. Total protein extracted from protoplasts described in Fig. 1a was subjected to anti-flag immunoblot to detect AvrAC-FLAG
doi:10.1038/nature10962 Supplementary Figure 1. Expression of AvrAC-FLAG in protoplasts. Total protein extracted from protoplasts described in Fig. 1a was subjected to anti-flag immunoblot to detect AvrAC-FLAG
<Supplemental information>
 The Structural Basis of Endosomal Anchoring of KIF16B Kinesin Nichole R. Blatner, Michael I. Wilson, Cai Lei, Wanjin Hong, Diana Murray, Roger L. Williams, and Wonhwa Cho Protein
The Structural Basis of Endosomal Anchoring of KIF16B Kinesin Nichole R. Blatner, Michael I. Wilson, Cai Lei, Wanjin Hong, Diana Murray, Roger L. Williams, and Wonhwa Cho Protein
Organization of ATPases
 The Primary Active Transporter II: The ATPase Objectives: Organization P type with NPA domains Proton pumps of the rotary V type ATPase 1 Organization of P type, solute transport, found in plasma membranes
The Primary Active Transporter II: The ATPase Objectives: Organization P type with NPA domains Proton pumps of the rotary V type ATPase 1 Organization of P type, solute transport, found in plasma membranes
SUPPLEMENTARY INFORMATION
 Supplementary Discussion The HGGG sequence motif forms an oxyanion hole in HLSs. In the case of GID1, the third Gly residue of the sequence motif is replaced with Ser (Ser123 in OsGID1), i.e., the sequence
Supplementary Discussion The HGGG sequence motif forms an oxyanion hole in HLSs. In the case of GID1, the third Gly residue of the sequence motif is replaced with Ser (Ser123 in OsGID1), i.e., the sequence
SUPPLEMENTARY INFORMATION. Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity
 SUPPLEMENTARY INFORMATION Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity Christopher Woods 1, Maturos Malaisree 1, Ben Long 2, Simon McIntosh-Smith
SUPPLEMENTARY INFORMATION Computational Assay of H7N9 Influenza Neuraminidase Reveals R292K Mutation Reduces Drug Binding Affinity Christopher Woods 1, Maturos Malaisree 1, Ben Long 2, Simon McIntosh-Smith
Figure S1. (A) SDS-PAGE separation of GST-fusion proteins purified from E.coli BL21 strain is shown. An equal amount of GST-tag control, LRRK2 LRR
 Figure S1. (A) SDS-PAGE separation of GST-fusion proteins purified from E.coli BL21 strain is shown. An equal amount of GST-tag control, LRRK2 LRR and LRRK2 WD40 GST fusion proteins (5 µg) were loaded
Figure S1. (A) SDS-PAGE separation of GST-fusion proteins purified from E.coli BL21 strain is shown. An equal amount of GST-tag control, LRRK2 LRR and LRRK2 WD40 GST fusion proteins (5 µg) were loaded
Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase. Biol 405 Molecular Medicine
 Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase Biol 405 Molecular Medicine 1998 Crystal structure of phenylalanine hydroxylase solved. The polypeptide consists of three regions: Regulatory
Phenylketonuria (PKU) Structure of Phenylalanine Hydroxylase Biol 405 Molecular Medicine 1998 Crystal structure of phenylalanine hydroxylase solved. The polypeptide consists of three regions: Regulatory
Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.
 Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.5 % SDS-PAGE gel under reducing and non-reducing conditions.
Supplementary Figure 1 Preparation, crystallization and structure determination of EpEX. (a), Purified EpEX and EpEX analyzed on homogenous 12.5 % SDS-PAGE gel under reducing and non-reducing conditions.
Protein Secondary Structure
 Protein Secondary Structure Reading: Berg, Tymoczko & Stryer, 6th ed., Chapter 2, pp. 37-45 Problems in textbook: chapter 2, pp. 63-64, #1,5,9 Directory of Jmol structures of proteins: http://www.biochem.arizona.edu/classes/bioc462/462a/jmol/routines/routines.html
Protein Secondary Structure Reading: Berg, Tymoczko & Stryer, 6th ed., Chapter 2, pp. 37-45 Problems in textbook: chapter 2, pp. 63-64, #1,5,9 Directory of Jmol structures of proteins: http://www.biochem.arizona.edu/classes/bioc462/462a/jmol/routines/routines.html
Nature Structural and Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Mutational analysis of the SA2-Scc1 interaction in vitro and in human cells. (a) Autoradiograph (top) and Coomassie stained gel (bottom) of 35 S-labeled Myc-SA2 proteins (input)
Supplementary Figure 1 Mutational analysis of the SA2-Scc1 interaction in vitro and in human cells. (a) Autoradiograph (top) and Coomassie stained gel (bottom) of 35 S-labeled Myc-SA2 proteins (input)
Amino Acids. Review I: Protein Structure. Amino Acids: Structures. Amino Acids (contd.) Rajan Munshi
 Review I: Protein Structure Rajan Munshi BBSI @ Pitt 2005 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2005 Amino Acids Building blocks of proteins 20 amino acids
Review I: Protein Structure Rajan Munshi BBSI @ Pitt 2005 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2005 Amino Acids Building blocks of proteins 20 amino acids
This PDF file includes: Supplementary Figures 1 to 6 Supplementary Tables 1 to 2 Supplementary Methods Supplementary References
 Structure of the catalytic core of p300 and implications for chromatin targeting and HAT regulation Manuela Delvecchio, Jonathan Gaucher, Carmen Aguilar-Gurrieri, Esther Ortega, Daniel Panne This PDF file
Structure of the catalytic core of p300 and implications for chromatin targeting and HAT regulation Manuela Delvecchio, Jonathan Gaucher, Carmen Aguilar-Gurrieri, Esther Ortega, Daniel Panne This PDF file
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/10/471/eaah5085/dc1 Supplementary Materials for Phosphorylation of the exocyst protein Exo84 by TBK1 promotes insulin-stimulated GLUT4 trafficking Maeran Uhm,
www.sciencesignaling.org/cgi/content/full/10/471/eaah5085/dc1 Supplementary Materials for Phosphorylation of the exocyst protein Exo84 by TBK1 promotes insulin-stimulated GLUT4 trafficking Maeran Uhm,
SUPPLEMENTARY INFORMATION (SI) FIGURES AND TABLES
 SUPPLEMENTARY INFORMATION (SI) FIGURES AND TABLES 1 Title: Discovery of a junctional epitope antibody that stabilizes IL-6 and gp80 protein:protein interaction and modulates its downstream signaling Authors:
SUPPLEMENTARY INFORMATION (SI) FIGURES AND TABLES 1 Title: Discovery of a junctional epitope antibody that stabilizes IL-6 and gp80 protein:protein interaction and modulates its downstream signaling Authors:
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature10913 Supplementary Figure 1 2F o -F c electron density maps of cognate and near-cognate trna Leu 2 in the A site of the 70S ribosome. The maps are contoured at 1.2 sigma and some of
doi:10.1038/nature10913 Supplementary Figure 1 2F o -F c electron density maps of cognate and near-cognate trna Leu 2 in the A site of the 70S ribosome. The maps are contoured at 1.2 sigma and some of
T H E J O U R N A L O F C E L L B I O L O G Y
 T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Krenn et al., http://www.jcb.org/cgi/content/full/jcb.201110013/dc1 Figure S1. Levels of expressed proteins and demonstration that C-terminal
T H E J O U R N A L O F C E L L B I O L O G Y Supplemental material Krenn et al., http://www.jcb.org/cgi/content/full/jcb.201110013/dc1 Figure S1. Levels of expressed proteins and demonstration that C-terminal
CS612 - Algorithms in Bioinformatics
 Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
Spring 2016 Protein Structure February 7, 2016 Introduction to Protein Structure A protein is a linear chain of organic molecular building blocks called amino acids. Introduction to Protein Structure Amine
supplementary information
 DOI: 1.138/ncb1 Control Atg7 / NAC 1 1 1 1 (mm) Control Atg7 / NAC 1 1 1 1 (mm) Lamin B Gstm1 Figure S1 Neither the translocation of into the nucleus nor the induction of antioxidant proteins in autophagydeficient
DOI: 1.138/ncb1 Control Atg7 / NAC 1 1 1 1 (mm) Control Atg7 / NAC 1 1 1 1 (mm) Lamin B Gstm1 Figure S1 Neither the translocation of into the nucleus nor the induction of antioxidant proteins in autophagydeficient
Supplementary Information A Hydrophobic Barrier Deep Within the Inner Pore of the TWIK-1 K2P Potassium Channel Aryal et al.
 Supplementary Information A Hydrophobic Barrier Deep Within the Inner Pore of the TWIK-1 K2P Potassium Channel Aryal et al. Supplementary Figure 1 TWIK-1 stability during MD simulations in a phospholipid
Supplementary Information A Hydrophobic Barrier Deep Within the Inner Pore of the TWIK-1 K2P Potassium Channel Aryal et al. Supplementary Figure 1 TWIK-1 stability during MD simulations in a phospholipid
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature22394 Supplementary Table 1 Observed intermolecular interactions within the GLP-1:GLP-1R TMD interface. Superscripts refer to the Wootten residue numbering system
SUPPLEMENTARY INFORMATION doi:10.1038/nature22394 Supplementary Table 1 Observed intermolecular interactions within the GLP-1:GLP-1R TMD interface. Superscripts refer to the Wootten residue numbering system
Lysosomes and endocytic pathways 9/27/2012 Phyllis Hanson
 Lysosomes and endocytic pathways 9/27/2012 Phyllis Hanson General principles Properties of lysosomes Delivery of enzymes to lysosomes Endocytic uptake clathrin, others Endocytic pathways recycling vs.
Lysosomes and endocytic pathways 9/27/2012 Phyllis Hanson General principles Properties of lysosomes Delivery of enzymes to lysosomes Endocytic uptake clathrin, others Endocytic pathways recycling vs.
Supporting Information
 Supporting Information Guan et al. 10.1073/pnas.1217609110 Fig. S1. Three patterns of reactivity for CD4-induced (CD4i) mabs. The following representative ELISAs show three patterns of reactivity for CD4i
Supporting Information Guan et al. 10.1073/pnas.1217609110 Fig. S1. Three patterns of reactivity for CD4-induced (CD4i) mabs. The following representative ELISAs show three patterns of reactivity for CD4i
klp-18 (RNAi) Control. supplementary information. starting strain: AV335 [emb-27(g48); GFP::histone; GFP::tubulin] bleach
![klp-18 (RNAi) Control. supplementary information. starting strain: AV335 [emb-27(g48); GFP::histone; GFP::tubulin] bleach klp-18 (RNAi) Control. supplementary information. starting strain: AV335 [emb-27(g48); GFP::histone; GFP::tubulin] bleach](/thumbs/91/104639484.jpg) DOI: 10.1038/ncb1891 A. starting strain: AV335 [emb-27(g48); GFP::histone; GFP::tubulin] bleach embryos let hatch overnight transfer to RNAi plates; incubate 5 days at 15 C RNAi food L1 worms adult worms
DOI: 10.1038/ncb1891 A. starting strain: AV335 [emb-27(g48); GFP::histone; GFP::tubulin] bleach embryos let hatch overnight transfer to RNAi plates; incubate 5 days at 15 C RNAi food L1 worms adult worms
Chapter 6. X-ray structure analysis of D30N tethered HIV-1 protease. dimer/saquinavir complex
 Chapter 6 X-ray structure analysis of D30N tethered HIV-1 protease dimer/saquinavir complex 6.1 Introduction: The arrival of HIV protease inhibitors (PIs) in late 1995 marked the beginning of an important
Chapter 6 X-ray structure analysis of D30N tethered HIV-1 protease dimer/saquinavir complex 6.1 Introduction: The arrival of HIV protease inhibitors (PIs) in late 1995 marked the beginning of an important
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature11429 S1a 6 7 8 9 Nlrc4 allele S1b Nlrc4 +/+ Nlrc4 +/F Nlrc4 F/F 9 Targeting construct 422 bp 273 bp FRT-neo-gb-PGK-FRT 3x.STOP S1c Nlrc4 +/+ Nlrc4 F/F casp1
SUPPLEMENTARY INFORMATION doi:10.1038/nature11429 S1a 6 7 8 9 Nlrc4 allele S1b Nlrc4 +/+ Nlrc4 +/F Nlrc4 F/F 9 Targeting construct 422 bp 273 bp FRT-neo-gb-PGK-FRT 3x.STOP S1c Nlrc4 +/+ Nlrc4 F/F casp1
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 The UBL and RING1 interface remains associated in the complex structures of Parkin and pub. a) Asymmetric Unit of crystal structure of UBLR0RBR and pub complex showing UBL (green),
Supplementary Figure 1 The UBL and RING1 interface remains associated in the complex structures of Parkin and pub. a) Asymmetric Unit of crystal structure of UBLR0RBR and pub complex showing UBL (green),
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION TITLE: Structural Basis of Signal Sequence Surveillance and Selection by the SRP-SR Complex AUTHORS and AFFILIATIONS Ottilie von Loeffelholz 1,2, Kèvin Knoops 1,2,6, Aileen Ariosa
SUPPLEMENTARY INFORMATION TITLE: Structural Basis of Signal Sequence Surveillance and Selection by the SRP-SR Complex AUTHORS and AFFILIATIONS Ottilie von Loeffelholz 1,2, Kèvin Knoops 1,2,6, Aileen Ariosa
Studying dynamic protein:protein interactions in vesicle trafficking. David Owen CIMR University of Cambridge
 Studying dynamic protein:protein interactions in vesicle trafficking David Owen CIMR University of Cambridge The importance of vesicle trafficking in mammalian cells Transducing signals small molecules/nutrients
Studying dynamic protein:protein interactions in vesicle trafficking David Owen CIMR University of Cambridge The importance of vesicle trafficking in mammalian cells Transducing signals small molecules/nutrients
Insights into the Giardia intestinalis Enolase and Human Plasminogen interaction
 Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Supplementary Information Insights into the Giardia intestinalis Enolase and Human
Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2017 Supplementary Information Insights into the Giardia intestinalis Enolase and Human
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/4/3/eaaq0762/dc1 Supplementary Materials for Structures of monomeric and oligomeric forms of the Toxoplasma gondii perforin-like protein 1 Tao Ni, Sophie I. Williams,
advances.sciencemag.org/cgi/content/full/4/3/eaaq0762/dc1 Supplementary Materials for Structures of monomeric and oligomeric forms of the Toxoplasma gondii perforin-like protein 1 Tao Ni, Sophie I. Williams,
Catalysis & specificity: Proteins at work
 Catalysis & specificity: Proteins at work Introduction Having spent some time looking at the elements of structure of proteins and DNA, as well as their ability to form intermolecular interactions, it
Catalysis & specificity: Proteins at work Introduction Having spent some time looking at the elements of structure of proteins and DNA, as well as their ability to form intermolecular interactions, it
Thermal shift binding experiments were carried out using Thermofluor 384 ELS system. Protein
 Supplementary Methods Thermal shift assays Thermal shift binding experiments were carried out using Thermofluor 384 ELS system. Protein unfolding was examined by monitoring the fluorescence of ANS (1-anilinonaphthalene-8-
Supplementary Methods Thermal shift assays Thermal shift binding experiments were carried out using Thermofluor 384 ELS system. Protein unfolding was examined by monitoring the fluorescence of ANS (1-anilinonaphthalene-8-
HOMEWORK II and Swiss-PDB Viewer Tutorial DUE 9/26/03 62 points total. The ph at which a peptide has no net charge is its isoelectric point.
 BIOCHEMISTRY I HOMEWORK II and Swiss-PDB Viewer Tutorial DUE 9/26/03 62 points total 1). 8 points total T or F (2 points each; if false, briefly state why it is false) The ph at which a peptide has no
BIOCHEMISTRY I HOMEWORK II and Swiss-PDB Viewer Tutorial DUE 9/26/03 62 points total 1). 8 points total T or F (2 points each; if false, briefly state why it is false) The ph at which a peptide has no
2. Which of the following amino acids is most likely to be found on the outer surface of a properly folded protein?
 Name: WHITE Student Number: Answer the following questions on the computer scoring sheet. 1 mark each 1. Which of the following amino acids would have the highest relative mobility R f in normal thin layer
Name: WHITE Student Number: Answer the following questions on the computer scoring sheet. 1 mark each 1. Which of the following amino acids would have the highest relative mobility R f in normal thin layer
Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels. Supplementary Information
 Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels Craig T. Armstrong, Philip E. Mason, J. L. Ross Anderson and Christopher E. Dempsey
Arginine side chain interactions and the role of arginine as a mobile charge carrier in voltage sensitive ion channels Craig T. Armstrong, Philip E. Mason, J. L. Ross Anderson and Christopher E. Dempsey
Identification of non-ser/thr-pro consensus motifs for Cdk1 and their roles in mitotic regulation of C2H2 zinc finger proteins and Ect2
 SUPPLEMENTARY INFORMATION Identification of non-ser/thr-pro consensus motifs for Cdk1 and their roles in mitotic regulation of C2H2 zinc finger proteins and Ect2 Kazuhiro Suzuki, Kosuke Sako, Kazuhiro
SUPPLEMENTARY INFORMATION Identification of non-ser/thr-pro consensus motifs for Cdk1 and their roles in mitotic regulation of C2H2 zinc finger proteins and Ect2 Kazuhiro Suzuki, Kosuke Sako, Kazuhiro
Supplementary information: Binding of N-methylscopolamine to the extracellular domain of muscarinic acetylcholine receptors
 Supplementary information: Binding of N-methylscopolamine to the extracellular domain of muscarinic acetylcholine receptors Jan Jakubík, Alena Randáková, Pavel Zimčík, Esam E. El-Fakahany, and Vladimír
Supplementary information: Binding of N-methylscopolamine to the extracellular domain of muscarinic acetylcholine receptors Jan Jakubík, Alena Randáková, Pavel Zimčík, Esam E. El-Fakahany, and Vladimír
P450 CYCLE. All P450s follow the same catalytic cycle of;
 P450 CYCLE All P450s follow the same catalytic cycle of; 1. Initial substrate binding 2. First electron reduction 3. Oxygen binding 4. Second electron transfer 5 and 6. Proton transfer/dioxygen cleavage
P450 CYCLE All P450s follow the same catalytic cycle of; 1. Initial substrate binding 2. First electron reduction 3. Oxygen binding 4. Second electron transfer 5 and 6. Proton transfer/dioxygen cleavage
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/6/278/rs11/dc1 Supplementary Materials for In Vivo Phosphoproteomics Analysis Reveals the Cardiac Targets of β-adrenergic Receptor Signaling Alicia Lundby,* Martin
www.sciencesignaling.org/cgi/content/full/6/278/rs11/dc1 Supplementary Materials for In Vivo Phosphoproteomics Analysis Reveals the Cardiac Targets of β-adrenergic Receptor Signaling Alicia Lundby,* Martin
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature19102 Supplementary Discussion Benzothiazepine Binding in Ca V Ab Diltiazem and other benzothiazepines inhibit Ca V 1.2 channels in a frequency-dependent manner consistent with pore block
doi:10.1038/nature19102 Supplementary Discussion Benzothiazepine Binding in Ca V Ab Diltiazem and other benzothiazepines inhibit Ca V 1.2 channels in a frequency-dependent manner consistent with pore block
2013 John Wiley & Sons, Inc. All rights reserved. PROTEIN SORTING. Lecture 10 BIOL 266/ Biology Department Concordia University. Dr. S.
 PROTEIN SORTING Lecture 10 BIOL 266/4 2014-15 Dr. S. Azam Biology Department Concordia University Introduction Membranes divide the cytoplasm of eukaryotic cells into distinct compartments. The endomembrane
PROTEIN SORTING Lecture 10 BIOL 266/4 2014-15 Dr. S. Azam Biology Department Concordia University Introduction Membranes divide the cytoplasm of eukaryotic cells into distinct compartments. The endomembrane
STRUCTURE OF A UBIQUITIN-LOADED HECT LIGASE REVEALS THE MOLECULAR BASIS FOR CATALYTIC PRIMING SUPPLEMENTARY INFORMATION
 STRUCTURE OF A UBIQUITINLOADED ECT LIGASE REVEALS TE MOLECULAR BASIS FOR CATALYTIC PRIMING Elena Maspero 1, Eleonora Valentini 1, Sara Mari 1, Valentina Cecatiello 2, Paolo Soffientini 1, Sebastiano Pasqualato
STRUCTURE OF A UBIQUITINLOADED ECT LIGASE REVEALS TE MOLECULAR BASIS FOR CATALYTIC PRIMING Elena Maspero 1, Eleonora Valentini 1, Sara Mari 1, Valentina Cecatiello 2, Paolo Soffientini 1, Sebastiano Pasqualato
Draw how two amino acids form the peptide bond. Draw in the space below:
 Name Date Period Modeling Protein Folding Draw how two amino acids form the peptide bond. Draw in the space below: What we are doing today: The core idea in life sciences is that there is a fundamental
Name Date Period Modeling Protein Folding Draw how two amino acids form the peptide bond. Draw in the space below: What we are doing today: The core idea in life sciences is that there is a fundamental
MBB 694:407, 115:511. Please use BLOCK CAPITAL letters like this --- A, B, C, D, E. Not lowercase!
 MBB 694:407, 115:511 First Test Severinov/Deis Tue. Sep. 30, 2003 Name Index number (not SSN) Row Letter Seat Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions
MBB 694:407, 115:511 First Test Severinov/Deis Tue. Sep. 30, 2003 Name Index number (not SSN) Row Letter Seat Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:1.138/nature9814 a A SHARPIN FL B SHARPIN ΔNZF C SHARPIN T38L, F39V b His-SHARPIN FL -1xUb -2xUb -4xUb α-his c Linear 4xUb -SHARPIN FL -SHARPIN TF_LV -SHARPINΔNZF -SHARPIN
SUPPLEMENTARY INFORMATION doi:1.138/nature9814 a A SHARPIN FL B SHARPIN ΔNZF C SHARPIN T38L, F39V b His-SHARPIN FL -1xUb -2xUb -4xUb α-his c Linear 4xUb -SHARPIN FL -SHARPIN TF_LV -SHARPINΔNZF -SHARPIN
SDS-Assisted Protein Transport Through Solid-State Nanopores
 Supplementary Information for: SDS-Assisted Protein Transport Through Solid-State Nanopores Laura Restrepo-Pérez 1, Shalini John 2, Aleksei Aksimentiev 2 *, Chirlmin Joo 1 *, Cees Dekker 1 * 1 Department
Supplementary Information for: SDS-Assisted Protein Transport Through Solid-State Nanopores Laura Restrepo-Pérez 1, Shalini John 2, Aleksei Aksimentiev 2 *, Chirlmin Joo 1 *, Cees Dekker 1 * 1 Department
Supplementary Information for Structural basis for the inhibition of Polo-like kinase 1
 Supplementary Information for Structural basis for inhibition of Polo-like kinase 1 Jun Xu, Chen Shen, Tao Wang & Junmin Quan Correspondence authors: Tao Wang, taowang@pkusz.edu.cn; Junmin Quan, quanjm@szpku.edu.cn
Supplementary Information for Structural basis for inhibition of Polo-like kinase 1 Jun Xu, Chen Shen, Tao Wang & Junmin Quan Correspondence authors: Tao Wang, taowang@pkusz.edu.cn; Junmin Quan, quanjm@szpku.edu.cn
Danish Research Institute of Translational Neuroscience DANDRITE, Nordic-EMBL Partnership
 Supplementary Information for Tuning of the Na,K-ATPase by the beta subunit Florian Hilbers 1,2,3, Wojciech Kopec 4, Toke Jost Isaksen 3,5, Thomas Hellesøe Holm 3,5, Karin Lykke- Hartmann 3,5,6, Poul Nissen
Supplementary Information for Tuning of the Na,K-ATPase by the beta subunit Florian Hilbers 1,2,3, Wojciech Kopec 4, Toke Jost Isaksen 3,5, Thomas Hellesøe Holm 3,5, Karin Lykke- Hartmann 3,5,6, Poul Nissen
SUPPORTING INFORMATION FOR. A Computational Approach to Enzyme Design: Using Docking and MM- GBSA Scoring
 SUPPRTING INFRMATIN FR A Computational Approach to Enzyme Design: Predicting ω- Aminotransferase Catalytic Activity Using Docking and MM- GBSA Scoring Sarah Sirin, 1 Rajesh Kumar, 2 Carlos Martinez, 2
SUPPRTING INFRMATIN FR A Computational Approach to Enzyme Design: Predicting ω- Aminotransferase Catalytic Activity Using Docking and MM- GBSA Scoring Sarah Sirin, 1 Rajesh Kumar, 2 Carlos Martinez, 2
Introduction to proteins and protein structure
 Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
Introduction to proteins and protein structure The questions and answers below constitute an introduction to the fundamental principles of protein structure. They are all available at [link]. What are
Supplementary Figure 1. PD-L1 is glycosylated in cancer cells. (a) Western blot analysis of PD-L1 in breast cancer cells. (b) Western blot analysis
 Supplementary Figure 1. PD-L1 is glycosylated in cancer cells. (a) Western blot analysis of PD-L1 in breast cancer cells. (b) Western blot analysis of PD-L1 in ovarian cancer cells. (c) Western blot analysis
Supplementary Figure 1. PD-L1 is glycosylated in cancer cells. (a) Western blot analysis of PD-L1 in breast cancer cells. (b) Western blot analysis of PD-L1 in ovarian cancer cells. (c) Western blot analysis
File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description:
 File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description: Supplementary Figure 1. Schematic of Ras biochemical coupled assay.
File name: Supplementary Information Description: Supplementary figures and supplementary tables. File name: Peer review file Description: Supplementary Figure 1. Schematic of Ras biochemical coupled assay.
Lane: 1. Spectra BR protein ladder 2. PFD 3. TERM 4. 3-way connector 5. 2-way connector
 kda 1 2 3 4 5 26 14 1 7 Lane: 1. Spectra BR protein ladder 2. PFD 3. TERM 4. 3-way connector 5. 2-way connector 5 4 35 25 15 1 Supplementary Figure 1. SDS-PAGE of acterially expressed and purified proteins.
kda 1 2 3 4 5 26 14 1 7 Lane: 1. Spectra BR protein ladder 2. PFD 3. TERM 4. 3-way connector 5. 2-way connector 5 4 35 25 15 1 Supplementary Figure 1. SDS-PAGE of acterially expressed and purified proteins.
Supplementary Fig. 1. Identification of acetylation of K68 of SOD2
 Supplementary Fig. 1. Identification of acetylation of K68 of SOD2 A B H. sapiens 54 KHHAAYVNNLNVTEEKYQEALAK 75 M. musculus 54 KHHAAYVNNLNATEEKYHEALAK 75 X. laevis 55 KHHATYVNNLNITEEKYAEALAK 77 D. rerio
Supplementary Fig. 1. Identification of acetylation of K68 of SOD2 A B H. sapiens 54 KHHAAYVNNLNVTEEKYQEALAK 75 M. musculus 54 KHHAAYVNNLNATEEKYHEALAK 75 X. laevis 55 KHHATYVNNLNITEEKYAEALAK 77 D. rerio
OPTION GROUP: BIOLOGICAL MOLECULES 3 PROTEINS WORKBOOK. Tyrone R.L. John, Chartered Biologist
 NAME: OPTION GROUP: BIOLOGICAL MOLECULES 3 PROTEINS WORKBOOK Tyrone R.L. John, Chartered Biologist 1 Tyrone R.L. John, Chartered Biologist 2 Instructions REVISION CHECKLIST AND ASSESSMENT OBJECTIVES Regular
NAME: OPTION GROUP: BIOLOGICAL MOLECULES 3 PROTEINS WORKBOOK Tyrone R.L. John, Chartered Biologist 1 Tyrone R.L. John, Chartered Biologist 2 Instructions REVISION CHECKLIST AND ASSESSMENT OBJECTIVES Regular
This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is worth 2 points.
 MBB 407/511 Molecular Biology and Biochemistry First Examination - October 1, 2002 Name Social Security Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is
MBB 407/511 Molecular Biology and Biochemistry First Examination - October 1, 2002 Name Social Security Number This exam consists of two parts. Part I is multiple choice. Each of these 25 questions is
SAM Teacher s Guide Four Levels of Protein Structure
 SAM Teacher s Guide Four Levels of Protein Structure Overview Students explore how protein folding creates distinct, functional proteins by examining each of the four different levels of protein structure.
SAM Teacher s Guide Four Levels of Protein Structure Overview Students explore how protein folding creates distinct, functional proteins by examining each of the four different levels of protein structure.
Integrin v 3 targeted therapy for Kaposi s sarcoma with an in vitro evolved antibody 1
 Integrin v 3 targeted therapy for Kaposi s sarcoma with an in vitro evolved antibody 1 CHRISTOPH RADER, 2 MIKHAIL POPKOV, JOHN A. NEVES, AND CARLOS F. BARBAS III 2 Department of Molecular Biology and The
Integrin v 3 targeted therapy for Kaposi s sarcoma with an in vitro evolved antibody 1 CHRISTOPH RADER, 2 MIKHAIL POPKOV, JOHN A. NEVES, AND CARLOS F. BARBAS III 2 Department of Molecular Biology and The
Supplementary Information
 Supplementary Information Archaeal Elp3 catalyzes trna wobble uridine modification at C5 via a radical mechanism Kiruthika Selvadurai, Pei Wang, Joseph Seimetz & Raven H Huang* Department of Biochemistry,
Supplementary Information Archaeal Elp3 catalyzes trna wobble uridine modification at C5 via a radical mechanism Kiruthika Selvadurai, Pei Wang, Joseph Seimetz & Raven H Huang* Department of Biochemistry,
Nature Immunology: doi: /ni.3866
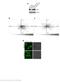 Nature Immunology: doi:10.1038/ni.3866 Supplementary Figure 1 The effect of TIPE2 on chemotaxis. a, The expression of TIPE2 in dhl-60c, dhl-60t, TIPE2-expressing and 15/16Q-expressing dhl-60t neutrophils
Nature Immunology: doi:10.1038/ni.3866 Supplementary Figure 1 The effect of TIPE2 on chemotaxis. a, The expression of TIPE2 in dhl-60c, dhl-60t, TIPE2-expressing and 15/16Q-expressing dhl-60t neutrophils
Supplementary materials
 Supplementary materials Chemical library from ChemBridge 50,240 structurally diverse small molecule compounds dissolved in DMSO Hits Controls: No virus added μ Primary screening at 20 g/ml of compounds
Supplementary materials Chemical library from ChemBridge 50,240 structurally diverse small molecule compounds dissolved in DMSO Hits Controls: No virus added μ Primary screening at 20 g/ml of compounds
Cellular Neurobiology BIPN140. 1st Midterm Exam October 18 th, Tuesday Material covered: Lectures 1-6 & Reading
 Cellular Neurobiology BIPN140 1st Midterm Exam October 18 th, Tuesday Material covered: Lectures 1-6 & Reading Review session October 17 th 3500 Pacitic Hall, 6-8 pm (access code is 127895) Come with questions!
Cellular Neurobiology BIPN140 1st Midterm Exam October 18 th, Tuesday Material covered: Lectures 1-6 & Reading Review session October 17 th 3500 Pacitic Hall, 6-8 pm (access code is 127895) Come with questions!
SUPPLEMENTARY INFORMATION
 Supplementary Figures Supplementary Figure S1. Binding of full-length OGT and deletion mutants to PIP strips (Echelon Biosciences). Supplementary Figure S2. Binding of the OGT (919-1036) fragments with
Supplementary Figures Supplementary Figure S1. Binding of full-length OGT and deletion mutants to PIP strips (Echelon Biosciences). Supplementary Figure S2. Binding of the OGT (919-1036) fragments with
Review II: The Molecules of Life
 Review II: The Molecules of Life Judy Wieber BBSI @ Pitt 2007 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2007 Outline Introduction Proteins Carbohydrates Lipids
Review II: The Molecules of Life Judy Wieber BBSI @ Pitt 2007 Department of Computational Biology University of Pittsburgh School of Medicine May 24, 2007 Outline Introduction Proteins Carbohydrates Lipids
HDL surface lipids mediate CETP binding as revealed by electron microscopy and molecular dynamics simulation
 HDL surface lipids mediate CETP binding as revealed by electron microscopy and molecular dynamics simulation Meng Zhang 1, River Charles 1, Huimin Tong 1, Lei Zhang 1, Mili Patel 2, Francis Wang 1, Matthew
HDL surface lipids mediate CETP binding as revealed by electron microscopy and molecular dynamics simulation Meng Zhang 1, River Charles 1, Huimin Tong 1, Lei Zhang 1, Mili Patel 2, Francis Wang 1, Matthew
Biomembranes structure and function. B. Balen
 Biomembranes structure and function B. Balen All cells are surrounded by membranes Selective barrier But also important for: 1. Compartmentalization 2. Biochemical activities 3. Transport of dissolved
Biomembranes structure and function B. Balen All cells are surrounded by membranes Selective barrier But also important for: 1. Compartmentalization 2. Biochemical activities 3. Transport of dissolved
SUPPLEMENTAL INFORMATION. Evolving Spindlin1 Small Molecule Inhibitors Using Protein Microarrays
 SUPPLEMETAL IFRMATI Evolving Spindlin1 Small Molecule Inhibitors Using Protein Microarrays arkhyun Bae 1, Monica Viviano 2, Xiaonan Su 3, Jie Lu 4, Donghang Cheng 1, Cari Sagum 1, Sabrina Castellano 2,6,
SUPPLEMETAL IFRMATI Evolving Spindlin1 Small Molecule Inhibitors Using Protein Microarrays arkhyun Bae 1, Monica Viviano 2, Xiaonan Su 3, Jie Lu 4, Donghang Cheng 1, Cari Sagum 1, Sabrina Castellano 2,6,
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3076 Supplementary Figure 1 btrcp targets Cep68 for degradation during mitosis. a) Cep68 immunofluorescence in interphase and metaphase. U-2OS cells were transfected with control sirna
DOI: 10.1038/ncb3076 Supplementary Figure 1 btrcp targets Cep68 for degradation during mitosis. a) Cep68 immunofluorescence in interphase and metaphase. U-2OS cells were transfected with control sirna
Supporting Information
 Supporting Information Bauer et al. 10.1073/pnas.1610421113 SI Materials and Methods Immunofluorescence experiments were performed as described (8). Briefly, NUGC-3 cells were treated with 25 μm PK11007
Supporting Information Bauer et al. 10.1073/pnas.1610421113 SI Materials and Methods Immunofluorescence experiments were performed as described (8). Briefly, NUGC-3 cells were treated with 25 μm PK11007
LDLR-related protein 10 (LRP10) regulates amyloid precursor protein (APP) trafficking and processing: evidence for a role in Alzheimer s disease
 Brodeur et al. Molecular Neurodegeneration 2012, 7:31 RESEARCH ARTICLE Open Access LDLR-related protein 10 (LRP10) regulates amyloid precursor protein (APP) trafficking and processing: evidence for a role
Brodeur et al. Molecular Neurodegeneration 2012, 7:31 RESEARCH ARTICLE Open Access LDLR-related protein 10 (LRP10) regulates amyloid precursor protein (APP) trafficking and processing: evidence for a role
Type of file: PDF Title of file for HTML: Supplementary Information Description: Supplementary Figures
 Type of file: PDF Title of file for HTML: Supplementary Information Description: Supplementary Figures Type of file: MOV Title of file for HTML: Supplementary Movie 1 Description: NLRP3 is moving along
Type of file: PDF Title of file for HTML: Supplementary Information Description: Supplementary Figures Type of file: MOV Title of file for HTML: Supplementary Movie 1 Description: NLRP3 is moving along
Drug Receptor Interactions and Pharmacodynamics
 Drug Receptor Interactions and Pharmacodynamics Dr. Raz Mohammed MSc Pharmacology School of Pharmacy 22.10.2017 Lec 6 Pharmacodynamics definition Pharmacodynamics describes the actions of a drug on the
Drug Receptor Interactions and Pharmacodynamics Dr. Raz Mohammed MSc Pharmacology School of Pharmacy 22.10.2017 Lec 6 Pharmacodynamics definition Pharmacodynamics describes the actions of a drug on the
Activities for the α-helix / β-sheet Construction Kit
 Activities for the α-helix / β-sheet Construction Kit The primary sequence of a protein, composed of amino acids, determines the organization of the sequence into the secondary structure. There are two
Activities for the α-helix / β-sheet Construction Kit The primary sequence of a protein, composed of amino acids, determines the organization of the sequence into the secondary structure. There are two
a Anti-Dab2 RGD Merge
 a Anti-Da Merge *** ** Percentage of cells adhered 8 6 4 RGE RAD RGE RAD Supplementary Figure Da are localized at clusters. (a) Endogenous Da is localized at clusters. (), ut not RGE nor RAD peptides can
a Anti-Da Merge *** ** Percentage of cells adhered 8 6 4 RGE RAD RGE RAD Supplementary Figure Da are localized at clusters. (a) Endogenous Da is localized at clusters. (), ut not RGE nor RAD peptides can
SUPPLEMENTARY INFORMATION
 Supplementary Table 1. Cell sphingolipids and S1P bound to endogenous TRAF2. Sphingolipid Cell pmol/mg TRAF2 immunoprecipitate pmol/mg Sphingomyelin 4200 ± 250 Not detected Monohexosylceramide 311 ± 18
Supplementary Table 1. Cell sphingolipids and S1P bound to endogenous TRAF2. Sphingolipid Cell pmol/mg TRAF2 immunoprecipitate pmol/mg Sphingomyelin 4200 ± 250 Not detected Monohexosylceramide 311 ± 18
Ionization of amino acids
 Amino Acids 20 common amino acids there are others found naturally but much less frequently Common structure for amino acid COOH, -NH 2, H and R functional groups all attached to the a carbon Ionization
Amino Acids 20 common amino acids there are others found naturally but much less frequently Common structure for amino acid COOH, -NH 2, H and R functional groups all attached to the a carbon Ionization
The Basics: A general review of molecular biology:
 The Basics: A general review of molecular biology: DNA Transcription RNA Translation Proteins DNA (deoxy-ribonucleic acid) is the genetic material It is an informational super polymer -think of it as the
The Basics: A general review of molecular biology: DNA Transcription RNA Translation Proteins DNA (deoxy-ribonucleic acid) is the genetic material It is an informational super polymer -think of it as the
RECAP FROM MONDAY AND TUESDAY LECTURES
 RECAP FROM MONDAY AND TUESDAY LECTURES Nucleo-cytoplasmic transport factors: interact with the target macromolecule (signal recognition) interact with the NPC (nucleoporin Phe - Gly repeats) NPC cytoplasmic
RECAP FROM MONDAY AND TUESDAY LECTURES Nucleo-cytoplasmic transport factors: interact with the target macromolecule (signal recognition) interact with the NPC (nucleoporin Phe - Gly repeats) NPC cytoplasmic
Supplementary Material
 Supplementary Material Materials and methods Enzyme assay The enzymatic activity of -glucosidase toward salicin was measured with the Miller method (Miller, 1959) using glucose as the standard. A total
Supplementary Material Materials and methods Enzyme assay The enzymatic activity of -glucosidase toward salicin was measured with the Miller method (Miller, 1959) using glucose as the standard. A total
