BWA alignment to reference transcriptome and genome. Convert transcriptome mappings back to genome space
|
|
|
- Albert Chambers
- 5 years ago
- Views:
Transcription
1 Whole genome sequencing Whole exome sequencing BWA alignment to reference transcriptome and genome Convert transcriptome mappings back to genome space genomes Filter on MQ, distance, Cigar string Annotate read pairs exomes Structural variant algorithm run over genomic alignments Filter on gene names Tumour vs normal comparison Candidate somatic pseudogenes Visual inspection in IGV, PCR validation Supplementary Figure 1. Bioinformatic analysis of whole-genome shotgun sequencing and exome pull-down sequencing data.
2 r La d T N T N T N T N T N T N T N T N de de La d La d de r r PCR validations of pseudogenes identified by bioinformatics algorithm T N T N T N T N T N T N T N Germline pseudogene Example of a somatic pseudogene (CAST from Supplementary Figure 3b) Note the band present in the tumour (T) lane, but absent from normal (N) Supplementary Figure 2. PCR validations of somatic pseudogenes. An example gel is shown of PCR validations of somatic pseudogenes. Each PCR was performed on tumour (T) and matched normal (N) DNA, and run on an agarose gel in adjacent lanes. Somatic pseudogenes are recognized by the presence of a band in the tumour lane that is absent from the matched normal DNA, whereas germline pseudogenes have bands present in PCRs from tumour and normal DNA.
3 a CENPF exons Split in read Insert between paired reads Linked to chr20: 54Mb Mapping position of read CENPF: Somatic pseudogene Intergenic Intergenic chr20: 54,081,176 chr20: 54,081,217 TT...TT AA...AA Exon Exon 13 chr1: 214,837,429 chr1: 214,818,324 PolyA tail b chr18: 22,136,016 CAST: Somatic pseudogene chr5: 96,089, Not mapped Not mapped Exon 29 PCR Supplementary Figure 3. Somatic pseudogenes. (a) A somatic CENPF pseudogene in a non-small cell lung cancer. Sequencing reads from high-coverage whole genome shotgun sequencing of the tumour reveal a series of split reads (red) crossing the four canonical exon-exon splice junctions in the gene. In addition, read pairs map to adjacent exons with an insert size larger than expected (light brown). At either end of the gene, read-pairs linking to chr20 could be identified, revealing that the CENPF pseudogene is inserted into an intergenic region with an intact polya tail and a target-site deletion of 40bp. (b) A somatic CAST pseudogene in a colorectal cancer. The 5 insertion point was confirmed as somatic by PCR and capillary sequencing across an exon-exon junction and insertion site.
4 Tail-to-tail inverted rearrangement on chromosome 2 Chr2: 6Mb HDDC2: Somatic pseudogene Chr2: 36Mb chr2: 6,819,517 chr2: 36,569,513 Exon Exon 1 A T chr6: 125,597,042 chr6: 125,623,178 2bp microhomology (AA) 1bp non-templated sequence Supplementary Figure 4. A somatic pseudogene was identified in which the 5 and 3 insertion points were ~30Mb apart on chromosome 2 and in an inverted orientation. This suggests that the pseudogene was inserted in the breakpoint of a genomic rearrangement during DNA repair.
5 a PSD3: Intron 15 KTN1: Somatic pseudogene PSD3: Intron 15 chr8: 18,405,785 chr14: 56,150,945 chr14: 56,106,712 chr8: 18,405,779 TTTTCTT AAAAGAA TTT...TTT AAA...AAA Exon TTTTCTT AAAAGAA chr14: 56,151,124 chr14: 56,104,006 Target site duplication Inverted rearrangement 100bp Target site duplication b ESR1 intron 3 RCOR1: Somatic pseudogene ESR1 intron 3 chr6: 152,238,254 chr14: 103,173,781 chr14: 103,193,405 chr6: 152,238,239 AAAGACAACAGGGGCT TTTCTGTTGTCCCCGA Exon Exon 12 A..A T..T AAAGACAACAGGGGCT TTTCTGTTGTCCCCGA chr14: 103,180,941 chr14: 103,196,898 Target site duplication Inverted rearrangement Target site duplication Supplementary Figure 5. Examples of somatic pseudogenes with internal inverted rearrangements.
6 100 1% Negative Positive 80 Frequency % 0% 2% 9.1% 0% 9.7% 19% 0% 0% 0% 0% 0% 18% 11% 0% 0% 0% 0% Lung Colorectal Cholangiocarcinoma Cell lines Gastric Chondrosarcoma Breast Renal Pancreatic Ovarian Osteosarcoma Myeloma Meningioma Melanoma Ewings Chordoma Ependymoma ALL Adenoid cystic Supplementary Figure 6. Distribution of samples positive (blue) or negative (light brown) for somatic pseudogenes across different tissue types.
7 a HDAC1 somatic pseudogene in a lung cancer Intergenic insertion ATAACTTTT TATTGAAAA T...T A...A Intergenic insertion Exon ATAACTTTT TATTGAAAA RASA2 intron 1 T...T A...A HDAC1 somatic pseudogene in a colorectal cancer Exon RASA2 intron 1 b Insertion into pre-mirna5694 TTTAGACCATTTCT AAATCTGGTAAAGA AKR1C1 somatic pseudogene in a lung cancer T...T A...A Exon Insertion into pre-mirna5694 TTTAGACCATTTCT AAATCTGGTAAAGA Not mapped AKR1C3 somatic pseudogene in a lung cancer Exon Not mapped 3kb upstream of TIPIN FMO3 somatic pseudogene in a lung cancer GCATTCTTTT CGTAAGTTTT T...T A...A Exon GCATTCTTTT CGTAAGTTTT Supplementary Figure 7. Recurrence of somatic pseudogenes. (a) Three somatic pseudogenes involving genes encoding enzymes that metabolise chemicals in cigarette smoke (AKR1C1, AKR1C3 and FMO3) observed in lung cancers from smokers. (b) Two somatic HDAC1 pseudogenes, one full-length in a colorectal cancer and one 5 truncated in a lung cancer. The different insertion points were confirmed by PCR.
8 a KTN1: Somatic pseudogene chr14: 56,150,945 chr8: 18,405,785 TTTTCTT TTT...TTT AAAAGAA AAA...AAA Exon 44 chr14: 56,151,124 PSD3: Exon 16 chr14: 56,106, chr8: 18,405,779 TTTTCTT AAAAGAA chr14: 56,104,006 Insertion point PSD3: Exon 15 Coverage PD7354c RNA reads Coverage PD7354h RNA reads Mates map to KTN1 exon 44 PD7354h b PD9061a (control) Supplementary Figure 8. Expression of a KTN1 pseudogene inserted into the last intron of PSD3 in a primary squamous cell lung cancer. (A) Two RNA samples were available from different time-points during the evolution of this cancer the somatic pseudogene was present in genomic DNA at both time-points. In the RNA-sequencing data at both time-points (PD7354c and PD7354h), a cluster of read-pairs maps to the PSD3 intron immediately adjacent to the insertion point (vertical green line). We find four read-pairs (in orange) which align with one end in the PSD3 intron and the other end in the KTN1 3 UTR, shown in the zoomed-in image. (B) No such expression is seen in another cancer (PD9061a).
9 a MLL Nonsense Frameshift Splice In frame indel Missense Silent Other tumour types zfcxxc PHD Bromo FYRN PHD HC5HC2H Head and neck cancers b FYRC SET MGA Other tumour types Tbox Lung adenocarcinomas Nonsense Frameshift Splice In frame indel Missense Silent Supplementary Figure 9. Patterns of mutations observed in (a) MLL and (b) MGA from a compendium of 7,651 publicly available exomes from cancers and matched normal samples. MGA shows a statistically significant excess of nonsense mutations (q=4x10-8) by Poisson regression (reference 31, main text) based on 58 nonsense mutations compared to 58 synonymous mutations. This indicates it is a likely tumour suppressor gene.
10 RNA sequencing coverage KIF18A KIAA UTR Exon 14 KIF18A somatic pseudogene BIN3 3 UTR RNA-sequencing reads across insertion points KIAA1967 BIN3 Supplementary Figure 10. RNA-seq of the KIF18A somatic pseudogene. Multiple read pairs support expression of the junctions between KIAA1967-KIF18A and BIN3-KIF18A.
11 Supplementary Table 1 Sample Type Cancer type Data type Read length (bp) Insert size (bp) Depth Matched normal depth PD4226a Primary Breast Exome % at >30x 78% at >30x PD6037a Primary Cholangiocarcinoma Exome % at >30x 54% at > 30x PD6368a Primary Chondrosarcoma Exome % at >30x 83% at >30x PD7261a Primary Colorectal Exome % at >30x 70% at > 30x PD9061a Primary Colorectal Exome % at >30x 26% at > 30x PD6022a Primary Gastric Exome % at >30x 60% at >30x PD6377a Primary Gastric Low depth genome x PCR only PD6384a Primary Gastric Low depth genome x PCR only PD6388a Primary Gastric Low depth genome x PCR only PD7354c Primary Lung Genome x 31.9x PD7354h Primary Lung Genome x 31.9x PD7354k Primary Lung Genome x 31.9x PD7354r Primary Lung Genome x 31.9x PD7355a Primary Lung Genome x 31.4x PD7356c Primary Lung Genome x 33.3x PD7356i Primary Lung Genome x 33.3x PD4861b Primary Lung Low depth genome x 2.0x PD4864b Primary Lung Low depth genome x 2.0x LB771-HNC Cell line Head & neck Exome % at >30x 84% at >30x NCI-H2009 Cell line Lung Exome % at >30x 81% at > 30x NCI-H2087 Cell line Lung Exome % at >30x 80% at > 30x
12 Supplementary Table 2 Tumour type Exome (positive) Genome (positive) Total (positive) Prevalence Adenoid cystic 46 (0) 0 46 (0) 0 % Acute lymphoblastic leukaemia 58 (0) 0 58 (0) 0 % Ependymoma 11(0) 0 11 (0) 0 % Breast 101 (1) 69 (0) 170 (1) 1 % Cholangiocarcinoma 9 (1) 0 9 (1) 11 % Chondrosarcoma 50 (1) 0 50 (1) 2 % Chordoma 25 (0) 0 25 (0) 0 % Colorectal 11 (2) 0 11 (2) 18 % Ewing's sarcoma 29 (0) 0 29 (0) 0 % Gastric 14 (1) 30 (3) 44 (4) 9 % Lung 0 27 (5) 27 (5) 19 % Melanoma 12 (0) 0 12 (0) 0 % Meningioma 3 (0) 0 3 (0) 0 % Myeloma 15 (0) 0 15 (0) 0 % Osteosarcoma 59 (0) 15 (0) 59 (0) 0 % Ovarian 0 18 (0) 18 (0) 0 % Pancreatic 0 11 (0) 11 (0) 0 % Renal 16 (0) 0 16 (0) 0 % Cell lines 31 (3) 0 31 (3) 14% TOTAL 490 (9) 170 (8) 660 (17) 3 %
13 Supplementary Table 3 Sample Gene Exons involved Insertion site 5' Insertion Site Pseudogene into 5' IS Microhomology / NTS 3' Insertion Site TCGA SSRP Intergenic chr17:34,610,045 chr11:57,102,120 4bp microhomology (ATCA) chr17:34,610,035 TCGA KRT6A 1-4; 4-9 In intron 9 of CTIF chr18:46,376,578 chr12:52,880,960 polya tail chr18:46,376,566 TCGA NOL7 1-8 Intergenic chr19:21,080,183 chr6:13,615,594 7bp NTS (CAGGCTCT) chr19:21,079,840 TCGA KRT6A 1-9 Intergenic chr14:55,576,932 chr12:52,887,038 5bp NTS (GCGAG) Not mapped TCGA CNIH4 1-4 In intron 4 of DDO chr6:110,725,517 chr1:224,544,589 1bp NTS (C) chr6:110,725,506
14 Pseudogene into 3' IS Microhomology / NTS Target site duplication Internal rearrangement Microhomology / NTS chr11:57,093,466 polya tail ATCAAGCTCTT chr12:52,884,672 1 bp microhomology (G) GGCCTTCCTTTTT Inverted exons 1;4 1bp microhomology to chr6:13,621,127 polya tail 200bp duplication Not mapped Not mapped Not mapped chr1:224,563,688 polya tail TCCTTTTGTCTT
15 Supplementary Table 4 Pseudogene Data type Depth Insertion junctions splice junctions KRT6A Exome 81% at >30x KRT6A Genome 41x KIF18A Exome 81% at >30x 9 90 KIF18A Genome 41x C9orf41 Exome 73% at >30x C9orf41 Genome 46x PTPN12 Exome 73% at >30x PTPN12 Genome 46x IBTK Exome 73% at >30x IBTK Genome 46x ARPC5 Exome 69% at >30x 4 55 ARPC5 Genome 40x 21 23
SUPPLEMENTARY INFORMATION
 doi: 1.138/nature8645 Physical coverage (x haploid genomes) 11 6.4 4.9 6.9 6.7 4.4 5.9 9.1 7.6 125 Neither end mapped One end mapped Chimaeras Correct Reads (million ns) 1 75 5 25 HCC1187 HCC1395 HCC1599
doi: 1.138/nature8645 Physical coverage (x haploid genomes) 11 6.4 4.9 6.9 6.7 4.4 5.9 9.1 7.6 125 Neither end mapped One end mapped Chimaeras Correct Reads (million ns) 1 75 5 25 HCC1187 HCC1395 HCC1599
Characterisation of structural variation in breast. cancer genomes using paired-end sequencing on. the Illumina Genome Analyser
 Characterisation of structural variation in breast cancer genomes using paired-end sequencing on the Illumina Genome Analyser Phil Stephens Cancer Genome Project Why is it important to study cancer? Why
Characterisation of structural variation in breast cancer genomes using paired-end sequencing on the Illumina Genome Analyser Phil Stephens Cancer Genome Project Why is it important to study cancer? Why
COMPUTATIONAL OPTIMISATION OF TARGETED DNA SEQUENCING FOR CANCER DETECTION
 COMPUTATIONAL OPTIMISATION OF TARGETED DNA SEQUENCING FOR CANCER DETECTION Pierre Martinez, Nicholas McGranahan, Nicolai Juul Birkbak, Marco Gerlinger, Charles Swanton* SUPPLEMENTARY INFORMATION SUPPLEMENTARY
COMPUTATIONAL OPTIMISATION OF TARGETED DNA SEQUENCING FOR CANCER DETECTION Pierre Martinez, Nicholas McGranahan, Nicolai Juul Birkbak, Marco Gerlinger, Charles Swanton* SUPPLEMENTARY INFORMATION SUPPLEMENTARY
SUPPLEMENTARY INFORMATION. Intron retention is a widespread mechanism of tumor suppressor inactivation.
 SUPPLEMENTARY INFORMATION Intron retention is a widespread mechanism of tumor suppressor inactivation. Hyunchul Jung 1,2,3, Donghoon Lee 1,4, Jongkeun Lee 1,5, Donghyun Park 2,6, Yeon Jeong Kim 2,6, Woong-Yang
SUPPLEMENTARY INFORMATION Intron retention is a widespread mechanism of tumor suppressor inactivation. Hyunchul Jung 1,2,3, Donghoon Lee 1,4, Jongkeun Lee 1,5, Donghyun Park 2,6, Yeon Jeong Kim 2,6, Woong-Yang
RNA SEQUENCING AND DATA ANALYSIS
 RNA SEQUENCING AND DATA ANALYSIS Length of mrna transcripts in the human genome 5,000 5,000 4,000 3,000 2,000 4,000 1,000 0 0 200 400 600 800 3,000 2,000 1,000 0 0 2,000 4,000 6,000 8,000 10,000 Length
RNA SEQUENCING AND DATA ANALYSIS Length of mrna transcripts in the human genome 5,000 5,000 4,000 3,000 2,000 4,000 1,000 0 0 200 400 600 800 3,000 2,000 1,000 0 0 2,000 4,000 6,000 8,000 10,000 Length
Using the Bravo Liquid-Handling System for Next Generation Sequencing Sample Prep
 Using the Bravo Liquid-Handling System for Next Generation Sequencing Sample Prep Tom Walsh, PhD Division of Medical Genetics University of Washington Next generation sequencing Sanger sequencing gold
Using the Bravo Liquid-Handling System for Next Generation Sequencing Sample Prep Tom Walsh, PhD Division of Medical Genetics University of Washington Next generation sequencing Sanger sequencing gold
Introduction. Introduction
 Introduction We are leveraging genome sequencing data from The Cancer Genome Atlas (TCGA) to more accurately define mutated and stable genes and dysregulated metabolic pathways in solid tumors. These efforts
Introduction We are leveraging genome sequencing data from The Cancer Genome Atlas (TCGA) to more accurately define mutated and stable genes and dysregulated metabolic pathways in solid tumors. These efforts
RNA SEQUENCING AND DATA ANALYSIS
 RNA SEQUENCING AND DATA ANALYSIS Download slides and package http://odin.mdacc.tmc.edu/~rverhaak/package.zip http://odin.mdacc.tmc.edu/~rverhaak/rna-seqlecture.zip Overview Introduction into the topic
RNA SEQUENCING AND DATA ANALYSIS Download slides and package http://odin.mdacc.tmc.edu/~rverhaak/package.zip http://odin.mdacc.tmc.edu/~rverhaak/rna-seqlecture.zip Overview Introduction into the topic
MSI positive MSI negative
 Pritchard et al. 2014 Supplementary Figure 1 MSI positive MSI negative Hypermutated Median: 673 Average: 659.2 Non-Hypermutated Median: 37.5 Average: 43.6 Supplementary Figure 1: Somatic Mutation Burden
Pritchard et al. 2014 Supplementary Figure 1 MSI positive MSI negative Hypermutated Median: 673 Average: 659.2 Non-Hypermutated Median: 37.5 Average: 43.6 Supplementary Figure 1: Somatic Mutation Burden
Supplementary Tables. Supplementary Figures
 Supplementary Files for Zehir, Benayed et al. Mutational Landscape of Metastatic Cancer Revealed from Prospective Clinical Sequencing of 10,000 Patients Supplementary Tables Supplementary Table 1: Sample
Supplementary Files for Zehir, Benayed et al. Mutational Landscape of Metastatic Cancer Revealed from Prospective Clinical Sequencing of 10,000 Patients Supplementary Tables Supplementary Table 1: Sample
Variant Classification. Author: Mike Thiesen, Golden Helix, Inc.
 Variant Classification Author: Mike Thiesen, Golden Helix, Inc. Overview Sequencing pipelines are able to identify rare variants not found in catalogs such as dbsnp. As a result, variants in these datasets
Variant Classification Author: Mike Thiesen, Golden Helix, Inc. Overview Sequencing pipelines are able to identify rare variants not found in catalogs such as dbsnp. As a result, variants in these datasets
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers
 Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes.
 Supplementary Figure 1 Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes. (a,b) Values of coefficients associated with genomic features, separately
Supplementary Figure 1 Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes. (a,b) Values of coefficients associated with genomic features, separately
Mobile element insertions are frequent in oesophageal adenocarcinomas and can mislead paired-end sequencing analysis
 Paterson et al. BMC Genomics (2015) 16:473 DOI 10.1186/s12864-015-1685-z RESEARCH ARTICLE Mobile element insertions are frequent in oesophageal adenocarcinomas and can mislead paired-end sequencing analysis
Paterson et al. BMC Genomics (2015) 16:473 DOI 10.1186/s12864-015-1685-z RESEARCH ARTICLE Mobile element insertions are frequent in oesophageal adenocarcinomas and can mislead paired-end sequencing analysis
Session 4 Rebecca Poulos
 The Cancer Genome Atlas (TCGA) & International Cancer Genome Consortium (ICGC) Session 4 Rebecca Poulos Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 20
The Cancer Genome Atlas (TCGA) & International Cancer Genome Consortium (ICGC) Session 4 Rebecca Poulos Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 20
NGS in tissue and liquid biopsy
 NGS in tissue and liquid biopsy Ana Vivancos, PhD Referencias So, why NGS in the clinics? 2000 Sanger Sequencing (1977-) 2016 NGS (2006-) ABIPrism (Applied Biosystems) Up to 2304 per day (96 sequences
NGS in tissue and liquid biopsy Ana Vivancos, PhD Referencias So, why NGS in the clinics? 2000 Sanger Sequencing (1977-) 2016 NGS (2006-) ABIPrism (Applied Biosystems) Up to 2304 per day (96 sequences
TITLE: - Whole Genome Sequencing of High-Risk Families to Identify New Mutational Mechanisms of Breast Cancer Predisposition
 AD Award Number: W81XWH-13-1-0336 TITLE: - Whole Genome Sequencing of High-Risk Families to Identify New Mutational Mechanisms of Breast Cancer Predisposition PRINCIPAL INVESTIGATOR: Mary-Claire King,
AD Award Number: W81XWH-13-1-0336 TITLE: - Whole Genome Sequencing of High-Risk Families to Identify New Mutational Mechanisms of Breast Cancer Predisposition PRINCIPAL INVESTIGATOR: Mary-Claire King,
Whole Genome and Transcriptome Analysis of Anaplastic Meningioma. Patrick Tarpey Cancer Genome Project Wellcome Trust Sanger Institute
 Whole Genome and Transcriptome Analysis of Anaplastic Meningioma Patrick Tarpey Cancer Genome Project Wellcome Trust Sanger Institute Outline Anaplastic meningioma compared to other cancers Whole genomes
Whole Genome and Transcriptome Analysis of Anaplastic Meningioma Patrick Tarpey Cancer Genome Project Wellcome Trust Sanger Institute Outline Anaplastic meningioma compared to other cancers Whole genomes
Session 4 Rebecca Poulos
 The Cancer Genome Atlas (TCGA) & International Cancer Genome Consortium (ICGC) Session 4 Rebecca Poulos Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 28
The Cancer Genome Atlas (TCGA) & International Cancer Genome Consortium (ICGC) Session 4 Rebecca Poulos Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 28
6.3 DNA Mutations. SBI4U Ms. Ho-Lau
 6.3 DNA Mutations SBI4U Ms. Ho-Lau DNA Mutations Gene expression can be affected by errors that occur during DNA replication. Some errors are repaired, but others can become mutations (changes in the nucleotide
6.3 DNA Mutations SBI4U Ms. Ho-Lau DNA Mutations Gene expression can be affected by errors that occur during DNA replication. Some errors are repaired, but others can become mutations (changes in the nucleotide
The Cancer Genome Atlas & International Cancer Genome Consortium
 The Cancer Genome Atlas & International Cancer Genome Consortium Session 3 Dr Jason Wong Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 31 st July 2014 1
The Cancer Genome Atlas & International Cancer Genome Consortium Session 3 Dr Jason Wong Prince of Wales Clinical School Introductory bioinformatics for human genomics workshop, UNSW 31 st July 2014 1
Supplementary Figure 1. Estimation of tumour content
 Supplementary Figure 1. Estimation of tumour content a, Approach used to estimate the tumour content in S13T1/T2, S6T1/T2, S3T1/T2 and S12T1/T2. Tissue and tumour areas were evaluated by two independent
Supplementary Figure 1. Estimation of tumour content a, Approach used to estimate the tumour content in S13T1/T2, S6T1/T2, S3T1/T2 and S12T1/T2. Tissue and tumour areas were evaluated by two independent
Hands-On Ten The BRCA1 Gene and Protein
 Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
Supplementary results
 doi:1.138/nature946 Supplementary results Effects of rearrangements on protein coding genes Genomic instability drives cancer development through effects on genes, such as copy number changes, internal
doi:1.138/nature946 Supplementary results Effects of rearrangements on protein coding genes Genomic instability drives cancer development through effects on genes, such as copy number changes, internal
TOWARDS ACCURATE GERMLINE AND SOMATIC INDEL DISCOVERY WITH MICRO-ASSEMBLY. Giuseppe Narzisi, PhD Bioinformatics Scientist
 TOWARDS ACCURATE GERMLINE AND SOMATIC INDEL DISCOVERY WITH MICRO-ASSEMBLY Giuseppe Narzisi, PhD Bioinformatics Scientist July 29, 2014 Micro-Assembly Approach to detect INDELs 2 Outline 1 Detecting INDELs:
TOWARDS ACCURATE GERMLINE AND SOMATIC INDEL DISCOVERY WITH MICRO-ASSEMBLY Giuseppe Narzisi, PhD Bioinformatics Scientist July 29, 2014 Micro-Assembly Approach to detect INDELs 2 Outline 1 Detecting INDELs:
Identifying Mutations Responsible for Rare Disorders Using New Technologies
 Identifying Mutations Responsible for Rare Disorders Using New Technologies Jacek Majewski, Department of Human Genetics, McGill University, Montreal, QC Canada Mendelian Diseases Clear mode of inheritance
Identifying Mutations Responsible for Rare Disorders Using New Technologies Jacek Majewski, Department of Human Genetics, McGill University, Montreal, QC Canada Mendelian Diseases Clear mode of inheritance
MODULE 3: TRANSCRIPTION PART II
 MODULE 3: TRANSCRIPTION PART II Lesson Plan: Title S. CATHERINE SILVER KEY, CHIYEDZA SMALL Transcription Part II: What happens to the initial (premrna) transcript made by RNA pol II? Objectives Explain
MODULE 3: TRANSCRIPTION PART II Lesson Plan: Title S. CATHERINE SILVER KEY, CHIYEDZA SMALL Transcription Part II: What happens to the initial (premrna) transcript made by RNA pol II? Objectives Explain
Bio 111 Study Guide Chapter 17 From Gene to Protein
 Bio 111 Study Guide Chapter 17 From Gene to Protein BEFORE CLASS: Reading: Read the introduction on p. 333, skip the beginning of Concept 17.1 from p. 334 to the bottom of the first column on p. 336, and
Bio 111 Study Guide Chapter 17 From Gene to Protein BEFORE CLASS: Reading: Read the introduction on p. 333, skip the beginning of Concept 17.1 from p. 334 to the bottom of the first column on p. 336, and
DNA-seq Bioinformatics Analysis: Copy Number Variation
 DNA-seq Bioinformatics Analysis: Copy Number Variation Elodie Girard elodie.girard@curie.fr U900 institut Curie, INSERM, Mines ParisTech, PSL Research University Paris, France NGS Applications 5C HiC DNA-seq
DNA-seq Bioinformatics Analysis: Copy Number Variation Elodie Girard elodie.girard@curie.fr U900 institut Curie, INSERM, Mines ParisTech, PSL Research University Paris, France NGS Applications 5C HiC DNA-seq
Nature Genetics: doi: /ng Supplementary Figure 1. TCGA data set on HNSCCs reanalyzed in this study.
 Supplementary Figure 1 TCGA data set on HNSCCs reanalyzed in this study. Summary of the TCGA dataset on HNSCCs re-analyzed in this study and the respective numbers of samples available within each. Supplementary
Supplementary Figure 1 TCGA data set on HNSCCs reanalyzed in this study. Summary of the TCGA dataset on HNSCCs re-analyzed in this study and the respective numbers of samples available within each. Supplementary
DMD Genetics: complicated, complex and critical to understand
 DMD Genetics: complicated, complex and critical to understand Stanley Nelson, MD Professor of Human Genetics, Pathology and Laboratory Medicine, and Psychiatry Co Director, Center for Duchenne Muscular
DMD Genetics: complicated, complex and critical to understand Stanley Nelson, MD Professor of Human Genetics, Pathology and Laboratory Medicine, and Psychiatry Co Director, Center for Duchenne Muscular
Nature Genetics: doi: /ng Supplementary Figure 1. Somatic coding mutations identified by WES/WGS for 83 ATL cases.
 Supplementary Figure 1 Somatic coding mutations identified by WES/WGS for 83 ATL cases. (a) The percentage of targeted bases covered by at least 2, 10, 20 and 30 sequencing reads (top) and average read
Supplementary Figure 1 Somatic coding mutations identified by WES/WGS for 83 ATL cases. (a) The percentage of targeted bases covered by at least 2, 10, 20 and 30 sequencing reads (top) and average read
CITATION FILE CONTENT/FORMAT
 CITATION For any resultant publications using please cite: Matthew A. Field, Vicky Cho, T. Daniel Andrews, and Chris C. Goodnow (2015). "Reliably detecting clinically important variants requires both combined
CITATION For any resultant publications using please cite: Matthew A. Field, Vicky Cho, T. Daniel Andrews, and Chris C. Goodnow (2015). "Reliably detecting clinically important variants requires both combined
Nature Biotechnology: doi: /nbt.1904
 Supplementary Information Comparison between assembly-based SV calls and array CGH results Genome-wide array assessment of copy number changes, such as array comparative genomic hybridization (acgh), is
Supplementary Information Comparison between assembly-based SV calls and array CGH results Genome-wide array assessment of copy number changes, such as array comparative genomic hybridization (acgh), is
Reporting TP53 gene analysis results in CLL
 Reporting TP53 gene analysis results in CLL Mutations in TP53 - From discovery to clinical practice in CLL Discovery Validation Clinical practice Variant diversity *Leroy at al, Cancer Research Review
Reporting TP53 gene analysis results in CLL Mutations in TP53 - From discovery to clinical practice in CLL Discovery Validation Clinical practice Variant diversity *Leroy at al, Cancer Research Review
Understanding genetics, mutation and other details. Stanley F. Nelson, MD 6/29/18
 Understanding genetics, mutation and other details Stanley F. Nelson, MD 6/29/18 1 6 11 16 21 Duchenne muscular dystrophy 26 31 36 41 46 51 56 61 66 71 76 81 86 91 96 600 500 400 300 200 100 0 Duchenne/Becker
Understanding genetics, mutation and other details Stanley F. Nelson, MD 6/29/18 1 6 11 16 21 Duchenne muscular dystrophy 26 31 36 41 46 51 56 61 66 71 76 81 86 91 96 600 500 400 300 200 100 0 Duchenne/Becker
Breast and ovarian cancer in Serbia: the importance of mutation detection in hereditary predisposition genes using NGS
 Breast and ovarian cancer in Serbia: the importance of mutation detection in hereditary predisposition genes using NGS dr sc. Ana Krivokuća Laboratory for molecular genetics Institute for Oncology and
Breast and ovarian cancer in Serbia: the importance of mutation detection in hereditary predisposition genes using NGS dr sc. Ana Krivokuća Laboratory for molecular genetics Institute for Oncology and
Analyse de données de séquençage haut débit
 Analyse de données de séquençage haut débit Vincent Lacroix Laboratoire de Biométrie et Biologie Évolutive INRIA ERABLE 9ème journée ITS 21 & 22 novembre 2017 Lyon https://its.aviesan.fr Sequencing is
Analyse de données de séquençage haut débit Vincent Lacroix Laboratoire de Biométrie et Biologie Évolutive INRIA ERABLE 9ème journée ITS 21 & 22 novembre 2017 Lyon https://its.aviesan.fr Sequencing is
genomics for systems biology / ISB2020 RNA sequencing (RNA-seq)
 RNA sequencing (RNA-seq) Module Outline MO 13-Mar-2017 RNA sequencing: Introduction 1 WE 15-Mar-2017 RNA sequencing: Introduction 2 MO 20-Mar-2017 Paper: PMID 25954002: Human genomics. The human transcriptome
RNA sequencing (RNA-seq) Module Outline MO 13-Mar-2017 RNA sequencing: Introduction 1 WE 15-Mar-2017 RNA sequencing: Introduction 2 MO 20-Mar-2017 Paper: PMID 25954002: Human genomics. The human transcriptome
a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation,
 Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
Supplementary Information Supplementary Figures Supplementary Figure 1. a) List of KMTs targeted in the shrna screen. The official symbol, KMT designation, gene ID and specifities are provided. Those highlighted
Chromothripsis: A New Mechanism For Tumorigenesis? i Fellow s Conference Cheryl Carlson 6/10/2011
 Chromothripsis: A New Mechanism For Tumorigenesis? i Fellow s Conference Cheryl Carlson 6/10/2011 Massive Genomic Rearrangement Acquired in a Single Catastrophic Event during Cancer Development Cell 144,
Chromothripsis: A New Mechanism For Tumorigenesis? i Fellow s Conference Cheryl Carlson 6/10/2011 Massive Genomic Rearrangement Acquired in a Single Catastrophic Event during Cancer Development Cell 144,
NGS in Cancer Pathology After the Microscope: From Nucleic Acid to Interpretation
 NGS in Cancer Pathology After the Microscope: From Nucleic Acid to Interpretation Michael R. Rossi, PhD, FACMG Assistant Professor Division of Cancer Biology, Department of Radiation Oncology Department
NGS in Cancer Pathology After the Microscope: From Nucleic Acid to Interpretation Michael R. Rossi, PhD, FACMG Assistant Professor Division of Cancer Biology, Department of Radiation Oncology Department
New Drug development and Personalized Therapy in The Era of Molecular Medicine
 New Drug development and Personalized Therapy in The Era of Molecular Medicine Ramesh K. Ramanathan MD Virginia G. Piper Cancer Center Translational Genomics Research Institute Scottsdale, AZ Clinical
New Drug development and Personalized Therapy in The Era of Molecular Medicine Ramesh K. Ramanathan MD Virginia G. Piper Cancer Center Translational Genomics Research Institute Scottsdale, AZ Clinical
MODULE 4: SPLICING. Removal of introns from messenger RNA by splicing
 Last update: 05/10/2017 MODULE 4: SPLICING Lesson Plan: Title MEG LAAKSO Removal of introns from messenger RNA by splicing Objectives Identify splice donor and acceptor sites that are best supported by
Last update: 05/10/2017 MODULE 4: SPLICING Lesson Plan: Title MEG LAAKSO Removal of introns from messenger RNA by splicing Objectives Identify splice donor and acceptor sites that are best supported by
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Frequency of alternative-cassette-exon engagement with the ribosome is consistent across data from multiple human cell types and from mouse stem cells. Box plots showing AS frequency
Supplementary Figure 1 Frequency of alternative-cassette-exon engagement with the ribosome is consistent across data from multiple human cell types and from mouse stem cells. Box plots showing AS frequency
Nature Medicine: doi: /nm.4439
 Figure S1. Overview of the variant calling and verification process. This figure expands on Fig. 1c with details of verified variants identification in 547 additional validation samples. Somatic variants
Figure S1. Overview of the variant calling and verification process. This figure expands on Fig. 1c with details of verified variants identification in 547 additional validation samples. Somatic variants
PSSV User Manual (V1.0)
 PSSV User Manual (V1.0) 1. Introduction A novel pattern-based probabilistic approach, PSSV, is developed to identify somatic structural variations from WGS data. Specifically, discordant and concordant
PSSV User Manual (V1.0) 1. Introduction A novel pattern-based probabilistic approach, PSSV, is developed to identify somatic structural variations from WGS data. Specifically, discordant and concordant
CRISPR/Cas9 Enrichment and Long-read WGS for Structural Variant Discovery
 CRISPR/Cas9 Enrichment and Long-read WGS for Structural Variant Discovery PacBio CoLab Session October 20, 2017 For Research Use Only. Not for use in diagnostics procedures. Copyright 2017 by Pacific Biosciences
CRISPR/Cas9 Enrichment and Long-read WGS for Structural Variant Discovery PacBio CoLab Session October 20, 2017 For Research Use Only. Not for use in diagnostics procedures. Copyright 2017 by Pacific Biosciences
Single-strand DNA library preparation improves sequencing of formalin-fixed and paraffin-embedded (FFPE) cancer DNA
 www.impactjournals.com/oncotarget/ Oncotarget, Supplementary Materials 2016 Single-strand DNA library preparation improves sequencing of formalin-fixed and paraffin-embedded (FFPE) DNA Supplementary Materials
www.impactjournals.com/oncotarget/ Oncotarget, Supplementary Materials 2016 Single-strand DNA library preparation improves sequencing of formalin-fixed and paraffin-embedded (FFPE) DNA Supplementary Materials
Raymond Auerbach PhD Candidate, Yale University Gerstein and Snyder Labs August 30, 2012
 Elucidating Transcriptional Regulation at Multiple Scales Using High-Throughput Sequencing, Data Integration, and Computational Methods Raymond Auerbach PhD Candidate, Yale University Gerstein and Snyder
Elucidating Transcriptional Regulation at Multiple Scales Using High-Throughput Sequencing, Data Integration, and Computational Methods Raymond Auerbach PhD Candidate, Yale University Gerstein and Snyder
Advance Your Genomic Research Using Targeted Resequencing with SeqCap EZ Library
 Advance Your Genomic Research Using Targeted Resequencing with SeqCap EZ Library Marilou Wijdicks International Product Manager Research For Life Science Research Only. Not for Use in Diagnostic Procedures.
Advance Your Genomic Research Using Targeted Resequencing with SeqCap EZ Library Marilou Wijdicks International Product Manager Research For Life Science Research Only. Not for Use in Diagnostic Procedures.
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 U1 inhibition causes a shift of RNA-seq reads from exons to introns. (a) Evidence for the high purity of 4-shU-labeled RNAs used for RNA-seq. HeLa cells transfected with control
Supplementary Figure 1 U1 inhibition causes a shift of RNA-seq reads from exons to introns. (a) Evidence for the high purity of 4-shU-labeled RNAs used for RNA-seq. HeLa cells transfected with control
CDH1 truncating alterations were detected in all six plasmacytoid-variant bladder tumors analyzed by whole-exome sequencing.
 Supplementary Figure 1 CDH1 truncating alterations were detected in all six plasmacytoid-variant bladder tumors analyzed by whole-exome sequencing. Whole-exome sequencing of six plasmacytoid-variant bladder
Supplementary Figure 1 CDH1 truncating alterations were detected in all six plasmacytoid-variant bladder tumors analyzed by whole-exome sequencing. Whole-exome sequencing of six plasmacytoid-variant bladder
PSSV User Manual (V2.1)
 PSSV User Manual (V2.1) 1. Introduction A novel pattern-based probabilistic approach, PSSV, is developed to identify somatic structural variations from WGS data. Specifically, discordant and concordant
PSSV User Manual (V2.1) 1. Introduction A novel pattern-based probabilistic approach, PSSV, is developed to identify somatic structural variations from WGS data. Specifically, discordant and concordant
Supplementary Figure 1. Spitzoid Melanoma with PPFIBP1-MET fusion. (a) Histopathology (4x) shows a domed papule with melanocytes extending into the
 Supplementary Figure 1. Spitzoid Melanoma with PPFIBP1-MET fusion. (a) Histopathology (4x) shows a domed papule with melanocytes extending into the deep dermis. (b) The melanocytes demonstrate abundant
Supplementary Figure 1. Spitzoid Melanoma with PPFIBP1-MET fusion. (a) Histopathology (4x) shows a domed papule with melanocytes extending into the deep dermis. (b) The melanocytes demonstrate abundant
Analysis with SureCall 2.1
 Analysis with SureCall 2.1 Danielle Fletcher Field Application Scientist July 2014 1 Stages of NGS Analysis Primary analysis, base calling Control Software FASTQ file reads + quality 2 Stages of NGS Analysis
Analysis with SureCall 2.1 Danielle Fletcher Field Application Scientist July 2014 1 Stages of NGS Analysis Primary analysis, base calling Control Software FASTQ file reads + quality 2 Stages of NGS Analysis
Supplementary Figure 1
 Supplementary Figure 1 Supplementary Fig. 1: Quality assessment of formalin-fixed paraffin-embedded (FFPE)-derived DNA and nuclei. (a) Multiplex PCR analysis of unrepaired and repaired bulk FFPE gdna from
Supplementary Figure 1 Supplementary Fig. 1: Quality assessment of formalin-fixed paraffin-embedded (FFPE)-derived DNA and nuclei. (a) Multiplex PCR analysis of unrepaired and repaired bulk FFPE gdna from
Introduction to genetic variation. He Zhang Bioinformatics Core Facility 6/22/2016
 Introduction to genetic variation He Zhang Bioinformatics Core Facility 6/22/2016 Outline Basic concepts of genetic variation Genetic variation in human populations Variation and genetic disorders Databases
Introduction to genetic variation He Zhang Bioinformatics Core Facility 6/22/2016 Outline Basic concepts of genetic variation Genetic variation in human populations Variation and genetic disorders Databases
Ambient temperature regulated flowering time
 Ambient temperature regulated flowering time Applications of RNAseq RNA- seq course: The power of RNA-seq June 7 th, 2013; Richard Immink Overview Introduction: Biological research question/hypothesis
Ambient temperature regulated flowering time Applications of RNAseq RNA- seq course: The power of RNA-seq June 7 th, 2013; Richard Immink Overview Introduction: Biological research question/hypothesis
Section Chapter 14. Go to Section:
 Section 12-3 Chapter 14 Go to Section: Content Objectives Write these Down! I will be able to identify: The origin of genetic differences among organisms. The possible kinds of different mutations. The
Section 12-3 Chapter 14 Go to Section: Content Objectives Write these Down! I will be able to identify: The origin of genetic differences among organisms. The possible kinds of different mutations. The
of TERT, MLL4, CCNE1, SENP5, and ROCK1 on tumor development were discussed.
 Supplementary Note The potential association and implications of HBV integration at known and putative cancer genes of TERT, MLL4, CCNE1, SENP5, and ROCK1 on tumor development were discussed. Human telomerase
Supplementary Note The potential association and implications of HBV integration at known and putative cancer genes of TERT, MLL4, CCNE1, SENP5, and ROCK1 on tumor development were discussed. Human telomerase
Nature Getetics: doi: /ng.3471
 Supplementary Figure 1 Summary of exome sequencing data. ( a ) Exome tumor normal sample sizes for bladder cancer (BLCA), breast cancer (BRCA), carcinoid (CARC), chronic lymphocytic leukemia (CLLX), colorectal
Supplementary Figure 1 Summary of exome sequencing data. ( a ) Exome tumor normal sample sizes for bladder cancer (BLCA), breast cancer (BRCA), carcinoid (CARC), chronic lymphocytic leukemia (CLLX), colorectal
DNA is the genetic material that provides instructions for what our bodies look like and how they function. DNA is packaged into structures called
 DNA is the genetic material that provides instructions for what our bodies look like and how they function. DNA is packaged into structures called chromosomes. We have 23 pairs of chromosomes (for a total
DNA is the genetic material that provides instructions for what our bodies look like and how they function. DNA is packaged into structures called chromosomes. We have 23 pairs of chromosomes (for a total
CNV detection. Introduction and detection in NGS data. G. Demidov 1,2. NGSchool2016. Centre for Genomic Regulation. CNV detection. G.
 Introduction and detection in NGS data 1,2 1 Genomic and Epigenomic Variation in Disease group, Centre for Genomic Regulation 2 Universitat Pompeu Fabra NGSchool2016 methods: methods Outline methods: methods
Introduction and detection in NGS data 1,2 1 Genomic and Epigenomic Variation in Disease group, Centre for Genomic Regulation 2 Universitat Pompeu Fabra NGSchool2016 methods: methods Outline methods: methods
Accel-Amplicon Panels
 Accel-Amplicon Panels Amplicon sequencing has emerged as a reliable, cost-effective method for ultra-deep targeted sequencing. This highly adaptable approach is especially applicable for in-depth interrogation
Accel-Amplicon Panels Amplicon sequencing has emerged as a reliable, cost-effective method for ultra-deep targeted sequencing. This highly adaptable approach is especially applicable for in-depth interrogation
Genomic structural variation
 Genomic structural variation Mario Cáceres The new genomic variation DNA sequence differs across individuals much more than researchers had suspected through structural changes A huge amount of structural
Genomic structural variation Mario Cáceres The new genomic variation DNA sequence differs across individuals much more than researchers had suspected through structural changes A huge amount of structural
The search for cis-regulatory driver mutations in cancer genomes
 /, Vol. 6, No. 32 The search for cis-regulatory driver mutations in cancer genomes Rebecca C. Poulos 1, Mathew A. Sloane 1, Luke B. Hesson 1, Jason W. H. Wong 1 1 Prince of Wales Clinical School and Lowy
/, Vol. 6, No. 32 The search for cis-regulatory driver mutations in cancer genomes Rebecca C. Poulos 1, Mathew A. Sloane 1, Luke B. Hesson 1, Jason W. H. Wong 1 1 Prince of Wales Clinical School and Lowy
SVIM: Structural variant identification with long reads DAVID HELLER MAX PLANCK INSTITUTE FOR MOLECULAR GENETICS, BERLIN JUNE 2O18, SMRT LEIDEN
 SVIM: Structural variant identification with long reads DAVID HELLER MAX PLANCK INSTITUTE FOR MOLECULAR GENETICS, BERLIN JUNE 2O18, SMRT LEIDEN Structural variation (SV) Variants larger than 50bps Affect
SVIM: Structural variant identification with long reads DAVID HELLER MAX PLANCK INSTITUTE FOR MOLECULAR GENETICS, BERLIN JUNE 2O18, SMRT LEIDEN Structural variation (SV) Variants larger than 50bps Affect
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Heatmap of GO terms for differentially expressed genes. The terms were hierarchically clustered using the GO term enrichment beta. Darker red, higher positive
Supplementary Figures Supplementary Figure 1. Heatmap of GO terms for differentially expressed genes. The terms were hierarchically clustered using the GO term enrichment beta. Darker red, higher positive
The mutations that drive cancer. Paul Edwards. Department of Pathology and Cancer Research UK Cambridge Institute, University of Cambridge
 The mutations that drive cancer Paul Edwards Department of Pathology and Cancer Research UK Cambridge Institute, University of Cambridge Previously on Cancer... hereditary predisposition Normal Cell Slightly
The mutations that drive cancer Paul Edwards Department of Pathology and Cancer Research UK Cambridge Institute, University of Cambridge Previously on Cancer... hereditary predisposition Normal Cell Slightly
Supplemental Figure 1. Small RNA size distribution from different soybean tissues.
 Supplemental Figure 1. Small RNA size distribution from different soybean tissues. The size of small RNAs was plotted versus frequency (percentage) among total sequences (A, C, E and G) or distinct sequences
Supplemental Figure 1. Small RNA size distribution from different soybean tissues. The size of small RNAs was plotted versus frequency (percentage) among total sequences (A, C, E and G) or distinct sequences
Protein Domain-Centric Approach to Study Cancer Somatic Mutations from High-throughput Sequencing Studies
 Protein Domain-Centric Approach to Study Cancer Somatic Mutations from High-throughput Sequencing Studies Dr. Maricel G. Kann Assistant Professor Dept of Biological Sciences UMBC 2 The term protein domain
Protein Domain-Centric Approach to Study Cancer Somatic Mutations from High-throughput Sequencing Studies Dr. Maricel G. Kann Assistant Professor Dept of Biological Sciences UMBC 2 The term protein domain
CHR POS REF OBS ALLELE BUILD CLINICAL_SIGNIFICANCE
 CHR POS REF OBS ALLELE BUILD CLINICAL_SIGNIFICANCE is_clinical dbsnp MITO GENE chr1 13273 G C heterozygous - - -. - DDX11L1 chr1 949654 A G Homozygous 52 - - rs8997 - ISG15 chr1 1021346 A G heterozygous
CHR POS REF OBS ALLELE BUILD CLINICAL_SIGNIFICANCE is_clinical dbsnp MITO GENE chr1 13273 G C heterozygous - - -. - DDX11L1 chr1 949654 A G Homozygous 52 - - rs8997 - ISG15 chr1 1021346 A G heterozygous
Supplementary Appendix
 Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Yatsenko AN, Georgiadis AP, Röpke A, et al. X-linked TEX11
Supplementary Appendix This appendix has been provided by the authors to give readers additional information about their work. Supplement to: Yatsenko AN, Georgiadis AP, Röpke A, et al. X-linked TEX11
MEDICAL GENOMICS LABORATORY. Next-Gen Sequencing and Deletion/Duplication Analysis of NF1 Only (NF1-NG)
 Next-Gen Sequencing and Deletion/Duplication Analysis of NF1 Only (NF1-NG) Ordering Information Acceptable specimen types: Fresh blood sample (3-6 ml EDTA; no time limitations associated with receipt)
Next-Gen Sequencing and Deletion/Duplication Analysis of NF1 Only (NF1-NG) Ordering Information Acceptable specimen types: Fresh blood sample (3-6 ml EDTA; no time limitations associated with receipt)
Supplementary Figure 1. Spectrum and signatures of substitutions.
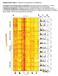 Supplementary Figure 1. Spectrum and signatures of substitutions. a. Heatmaps of trinucleotide context of substitutions. Each square represents a substitution in a specific trinucleotide context, normalised
Supplementary Figure 1. Spectrum and signatures of substitutions. a. Heatmaps of trinucleotide context of substitutions. Each square represents a substitution in a specific trinucleotide context, normalised
Tumor suppressor genes D R. S H O S S E I N I - A S L
 Tumor suppressor genes 1 D R. S H O S S E I N I - A S L What is a Tumor Suppressor Gene? 2 A tumor suppressor gene is a type of cancer gene that is created by loss-of function mutations. In contrast to
Tumor suppressor genes 1 D R. S H O S S E I N I - A S L What is a Tumor Suppressor Gene? 2 A tumor suppressor gene is a type of cancer gene that is created by loss-of function mutations. In contrast to
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature17676 Supplementary Methods 1. Sample selection, pathology review and clinical data collection Internal Review Boards of each participating institution approved collection and use of
doi:10.1038/nature17676 Supplementary Methods 1. Sample selection, pathology review and clinical data collection Internal Review Boards of each participating institution approved collection and use of
UNIVERSITY OF TORINO DEPARTMENT OF ONCOLOGY. Giorgio V. Scagliotti University of Torino Dipartment of Oncology
 Giorgio V. Scagliotti University of Torino Dipartment of Oncology giorgio.scagliotti@unito.it Molecular landscape of MM not fully characterized to allow personalized treatment Recurrent genetic alterations
Giorgio V. Scagliotti University of Torino Dipartment of Oncology giorgio.scagliotti@unito.it Molecular landscape of MM not fully characterized to allow personalized treatment Recurrent genetic alterations
6/12/2018. Disclosures. Clinical Genomics The CLIA Lab Perspective. Outline. COH HopeSeq Heme Panels
 Clinical Genomics The CLIA Lab Perspective Disclosures Raju K. Pillai, M.D. Hematopathologist / Molecular Pathologist Director, Pathology Bioinformatics City of Hope National Medical Center, Duarte, CA
Clinical Genomics The CLIA Lab Perspective Disclosures Raju K. Pillai, M.D. Hematopathologist / Molecular Pathologist Director, Pathology Bioinformatics City of Hope National Medical Center, Duarte, CA
Welcome to the Genetic Code: An Overview of Basic Genetics. October 24, :00pm 3:00pm
 Welcome to the Genetic Code: An Overview of Basic Genetics October 24, 2016 12:00pm 3:00pm Course Schedule 12:00 pm 2:00 pm Principles of Mendelian Genetics Introduction to Genetics of Complex Disease
Welcome to the Genetic Code: An Overview of Basic Genetics October 24, 2016 12:00pm 3:00pm Course Schedule 12:00 pm 2:00 pm Principles of Mendelian Genetics Introduction to Genetics of Complex Disease
TRANSLATION: 3 Stages to translation, can you guess what they are?
 TRANSLATION: Translation: is the process by which a ribosome interprets a genetic message on mrna to place amino acids in a specific sequence in order to synthesize polypeptide. 3 Stages to translation,
TRANSLATION: Translation: is the process by which a ribosome interprets a genetic message on mrna to place amino acids in a specific sequence in order to synthesize polypeptide. 3 Stages to translation,
Circular RNAs (circrnas) act a stable mirna sponges
 Circular RNAs (circrnas) act a stable mirna sponges cernas compete for mirnas Ancestal mrna (+3 UTR) Pseudogene RNA (+3 UTR homolgy region) The model holds true for all RNAs that share a mirna binding
Circular RNAs (circrnas) act a stable mirna sponges cernas compete for mirnas Ancestal mrna (+3 UTR) Pseudogene RNA (+3 UTR homolgy region) The model holds true for all RNAs that share a mirna binding
AVENIO family of NGS oncology assays ctdna and Tumor Tissue Analysis Kits
 AVENIO family of NGS oncology assays ctdna and Tumor Tissue Analysis Kits Accelerating clinical research Next-generation sequencing (NGS) has the ability to interrogate many different genes and detect
AVENIO family of NGS oncology assays ctdna and Tumor Tissue Analysis Kits Accelerating clinical research Next-generation sequencing (NGS) has the ability to interrogate many different genes and detect
Sequencing Cancer Genomes and Transcriptomes
 Sequencing Cancer Genomes and Transcriptomes Dr. Richard Moore Sequencing Group Leader Genome Sciences Centre, BC Cancer Agency Capacity at the GSC 11 Illumina GA II 2 AB SOLiD 8 3730 xls Paired and Single
Sequencing Cancer Genomes and Transcriptomes Dr. Richard Moore Sequencing Group Leader Genome Sciences Centre, BC Cancer Agency Capacity at the GSC 11 Illumina GA II 2 AB SOLiD 8 3730 xls Paired and Single
iplex genotyping IDH1 and IDH2 assays utilized the following primer sets (forward and reverse primers along with extension primers).
 Supplementary Materials Supplementary Methods iplex genotyping IDH1 and IDH2 assays utilized the following primer sets (forward and reverse primers along with extension primers). IDH1 R132H and R132L Forward:
Supplementary Materials Supplementary Methods iplex genotyping IDH1 and IDH2 assays utilized the following primer sets (forward and reverse primers along with extension primers). IDH1 R132H and R132L Forward:
Multiplex target enrichment using DNA indexing for ultra-high throughput variant detection
 Multiplex target enrichment using DNA indexing for ultra-high throughput variant detection Dr Elaine Kenny Neuropsychiatric Genetics Research Group Institute of Molecular Medicine Trinity College Dublin
Multiplex target enrichment using DNA indexing for ultra-high throughput variant detection Dr Elaine Kenny Neuropsychiatric Genetics Research Group Institute of Molecular Medicine Trinity College Dublin
Introduction to Cancer Bioinformatics and cancer biology. Anthony Gitter Cancer Bioinformatics (BMI 826/CS 838) January 20, 2015
 Introduction to Cancer Bioinformatics and cancer biology Anthony Gitter Cancer Bioinformatics (BMI 826/CS 838) January 20, 2015 Why cancer bioinformatics? Devastating disease, no cure on the horizon Major
Introduction to Cancer Bioinformatics and cancer biology Anthony Gitter Cancer Bioinformatics (BMI 826/CS 838) January 20, 2015 Why cancer bioinformatics? Devastating disease, no cure on the horizon Major
MRC-Holland MLPA. Description version 06; 23 December 2016
 SALSA MLPA probemix P417-B2 BAP1 Lot B2-1216. As compared to version B1 (lot B1-0215), two reference probes have been added and two target probes have a minor change in length. The BAP1 (BRCA1 associated
SALSA MLPA probemix P417-B2 BAP1 Lot B2-1216. As compared to version B1 (lot B1-0215), two reference probes have been added and two target probes have a minor change in length. The BAP1 (BRCA1 associated
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 Multiple samples from five patients (P4, P8, P14, P15 and P17) with Barrett s esophagus and adjacent EAC show that the poor overlap is not a result of sampling bias. Bar graphs showing
Supplementary Figure 1 Multiple samples from five patients (P4, P8, P14, P15 and P17) with Barrett s esophagus and adjacent EAC show that the poor overlap is not a result of sampling bias. Bar graphs showing
Supplementary Methods
 Supplementary Methods Short Read Preprocessing Reads are preprocessed differently according to how they will be used: detection of the variant in the tumor, discovery of an artifact in the normal or for
Supplementary Methods Short Read Preprocessing Reads are preprocessed differently according to how they will be used: detection of the variant in the tumor, discovery of an artifact in the normal or for
Nature Genetics: doi: /ng Supplementary Figure 1. Mutational signatures in BCC compared to melanoma.
 Supplementary Figure 1 Mutational signatures in BCC compared to melanoma. (a) The effect of transcription-coupled repair as a function of gene expression in BCC. Tumor type specific gene expression levels
Supplementary Figure 1 Mutational signatures in BCC compared to melanoma. (a) The effect of transcription-coupled repair as a function of gene expression in BCC. Tumor type specific gene expression levels
Studying Alternative Splicing
 Studying Alternative Splicing Meelis Kull PhD student in the University of Tartu supervisor: Jaak Vilo CS Theory Days Rõuge 27 Overview Alternative splicing Its biological function Studying splicing Technology
Studying Alternative Splicing Meelis Kull PhD student in the University of Tartu supervisor: Jaak Vilo CS Theory Days Rõuge 27 Overview Alternative splicing Its biological function Studying splicing Technology
MEDICAL GENOMICS LABORATORY. Peripheral Nerve Sheath Tumor Panel by Next-Gen Sequencing (PNT-NG)
 Peripheral Nerve Sheath Tumor Panel by Next-Gen Sequencing (PNT-NG) Ordering Information Acceptable specimen types: Blood (3-6ml EDTA; no time limitations associated with receipt) Saliva (OGR-575 DNA Genotek;
Peripheral Nerve Sheath Tumor Panel by Next-Gen Sequencing (PNT-NG) Ordering Information Acceptable specimen types: Blood (3-6ml EDTA; no time limitations associated with receipt) Saliva (OGR-575 DNA Genotek;
Computer Science, Biology, and Biomedical Informatics (CoSBBI) Outline. Molecular Biology of Cancer AND. Goals/Expectations. David Boone 7/1/2015
 Goals/Expectations Computer Science, Biology, and Biomedical (CoSBBI) We want to excite you about the world of computer science, biology, and biomedical informatics. Experience what it is like to be a
Goals/Expectations Computer Science, Biology, and Biomedical (CoSBBI) We want to excite you about the world of computer science, biology, and biomedical informatics. Experience what it is like to be a
The Cancer Genome Atlas
 The Cancer Genome Atlas July 14, 2011 Kenna M. Shaw, Ph.D. Deputy Director The Cancer Genome Atlas Program TCGA: Core Objectives Launched in 2006 as a pilot and expanded in 2009, the goals of TCGA are
The Cancer Genome Atlas July 14, 2011 Kenna M. Shaw, Ph.D. Deputy Director The Cancer Genome Atlas Program TCGA: Core Objectives Launched in 2006 as a pilot and expanded in 2009, the goals of TCGA are
Cancer Informatics Lecture
 Cancer Informatics Lecture Mayo-UIUC Computational Genomics Course June 22, 2018 Krishna Rani Kalari Ph.D. Associate Professor 2017 MFMER 3702274-1 Outline The Cancer Genome Atlas (TCGA) Genomic Data Commons
Cancer Informatics Lecture Mayo-UIUC Computational Genomics Course June 22, 2018 Krishna Rani Kalari Ph.D. Associate Professor 2017 MFMER 3702274-1 Outline The Cancer Genome Atlas (TCGA) Genomic Data Commons
Investigating rare diseases with Agilent NGS solutions
 Investigating rare diseases with Agilent NGS solutions Chitra Kotwaliwale, Ph.D. 1 Rare diseases affect 350 million people worldwide 7,000 rare diseases 80% are genetic 60 million affected in the US, Europe
Investigating rare diseases with Agilent NGS solutions Chitra Kotwaliwale, Ph.D. 1 Rare diseases affect 350 million people worldwide 7,000 rare diseases 80% are genetic 60 million affected in the US, Europe
Correspondence to Nature Genetics: Exploring pediatric cancer mutation information using ProteinPaint
 SUPPLEMENTARY INFORMATION FOR Correspondence to Nature Genetics: Exploring pediatric cancer mutation information using ProteinPaint Xin Zhou 1, Michael Edmonson 1, Mark R. Wilkinson 1, Aman Patel 1, Gang
SUPPLEMENTARY INFORMATION FOR Correspondence to Nature Genetics: Exploring pediatric cancer mutation information using ProteinPaint Xin Zhou 1, Michael Edmonson 1, Mark R. Wilkinson 1, Aman Patel 1, Gang
Computational Systems Biology: Biology X
 Bud Mishra Room 1002, 715 Broadway, Courant Institute, NYU, New York, USA L#4:(October-0-4-2010) Cancer and Signals 1 2 1 2 Evidence in Favor Somatic mutations, Aneuploidy, Copy-number changes and LOH
Bud Mishra Room 1002, 715 Broadway, Courant Institute, NYU, New York, USA L#4:(October-0-4-2010) Cancer and Signals 1 2 1 2 Evidence in Favor Somatic mutations, Aneuploidy, Copy-number changes and LOH
