RESEARCH ARTICLE. Abstract. Introduction
|
|
|
- Nicholas Chambers
- 6 years ago
- Views:
Transcription
1 DOI: /APJCP New Genetic Variation in the BCR Gene RESEARCH ARTICLE New Genetic Variation in BCR gene of Major B3a2 Breakpoint BCR-ABL Fusion Gene in Patients with Chronic Myelogenous Leukemia in Yogyakarta, Indonesia Tri Agusti Sholikah 1,2, Susanna Hilda Hutajulu 3,6, Dewi Sulistyawati 4, Sumartiningsih Aning 4, Sri Fatmawati 4, Anditta Syifarahmah 5, Kartika Widayati 6, Johan Kurnianda 6, Dewi Kartikawati Paramita 7 * Abstract Background: Polymorphic bases in several exons of the BCR gene have been found in several studies of the BCR-ABL fusion gene. Most of the polymorphisms do not have any implications for the primary structure of the BCR-ABL protein. Nucleotide changes are often located in the area close to the fusion region, and therefore may influence primer annealing. Our previous work failed to amplify 15 of 200 samples from BCR-ABL positive chronic myelogenous leukemia (CML) patients using multiplex PCR, the standard method to detect BCR-ABL transcripts used in our institution. The failure was considered due to problems in primer annealing caused by sequence variations. Sequence analysis of BCR-ABL fusion gene breakpoint types in CML patients has never been hitherto performed in Indonesia. Therefore, the aim of this study was to perform sequence analysis of several samples that did not show amplification using the standard method. Methods: Fifteen samples were qualitatively amplified by two-step PCR using inner primers in the 2 nd PCR to determine the breakpoint type of the BCR-ABL fusion gene. The 2 nd PCR products were used as templates to perform sequence analysis, and the results were compared to those in genbank. Result: Seven and 5 of 15 samples were confirmed as major b3a2 and major b2a2, respectively. One sample featured a combination of b3a2 and b2a2, and 2 samples a combination of b3a2 and b2a2 with an additional fragment at 500bp. Sequence analysis showed 3 sequence variations in the major b3a2 breakpoint. One had been reported earlier (c.3296t>c) but the others (c.3245c>t and c.3359t>c) were novel. Fragments at 500bp were confirmed as b3a2 and similar sequence b3a2 in genbank. Conclusion: This study found two new genetic variations in the BCR gene in BCR-ABL fusion cases. Keywords: CML- BCR-ABL-b3a2- b2a2- major breakpoint Asian Pac J Cancer Prev, 18 (5), Introduction Chronic myelogenous leukemia (CML) is a hematologic malignancy characterized by excessive accumulation of myeloid cells (Huntly et al., 2003; Perrotti et al., 2010). CML is a relatively frequent disease, corresponding to 15% of total leukemia and 7% to 20% of leukemia in adults (Garcia-Manero et al., 2003; Sillaber et al., 2003). In Asia, the incidence of CML is per individuals per year, and higher incidence has been reported in the Western country, which is 1-2 per individuals per year (Garcia-Manero et al., 2003; Baccarani and Dreyling, 2010; Kim et al., 2010 ; Jabbour and Kanterjian, 2014). The median onset of CML patients in Western country is 65 years old. It is 20 years older than the median onset of CML patients in Asia, which is 45 years old (Au et al., 2009; Kim et al., 2010). Etiophatogenesis of CML mostly involves Breakpoint Cluster Region-Abelson Murine Leukemia (BCR-ABL) fusion gene. The BCR-ABL fusion gene is derived from a reciprocal translocation between chromosome 9 and 22, t(9;22)(q34;ql1), known as Philadelphia chromosome. This translocation fuse the 3 part of the ABL gene on chromosome 9 and the 5 part of the BCR gene on chromosome 22, resulting in the formation of the BCR-ABL fusion oncogene (Ohsaka et al., 2002; Sillaber et al., 2003; Druker, 2008). The fusion gene has various sequences. It depends on the breakpoint on BCR gene that fuses with ABL gene. In general, 3 breakpoint cluster regions on BCR gene have been reported, namely major (M-bcr), minor (m-bcr) and micro (µ-bcr) (Smith et al., 2003). Furthermore, several studies has been 1 Biomedical Science Study Program, 3 Departement of Internal Medicine, 4 Molecular Biology Laboratory, 5 Medical Study Program, 7 Departement of Histology and Cell Biology, Faculty of Medicine, Universitas Gadjah Mada, 6 Division of Hematology and Medical Oncology, Department of Internal Medicine, Dr Sardjito Hospital, Yogyakarta, 2 Departement of Histology, Faculty of Medicine, Universitas Sebelas Maret, Surakarta, Indonesia. *For Correspondence: dkparamita@ugm.ac.id Asian Pacific Journal of Cancer Prevention, Vol
2 Tri Agusti Sholikah et al identified polymorphic base of several exons of BCR gene in BCR ABL fusion gene (De V. Meissner et al., 1998; Saußele et al., 2000). However, most of the polymorphisms do not have any implication to the primary structure of BCR-ABL protein (Saußele et al., 2000). Standard diagnosis of CML is done based on detection of the BCR-ABL fusion gene include its breakpoint type (Goh et al., 2006). In our institution, the standard methods to detect BCR-ABL fusion gene and its breakpoint type in CML patients are qualitative multiplex RT-PCR and nested PCR. The amplified product is analyzed by using electrophoresis method. However, the method is not sensitive to detect a small genetic variation. The nucleotide changes in the junctional region may influence the annealing of the primer (De V. Meissner et al., 1998). Between March 2010 and August 2014, 200 blood samples of patients with BCR-ABL positive CML were analyzed in the local laboratory. Fifteen samples could not be amplified by using the method above. The amplification failure was considered as the failure of primer annealing that may be caused by sequence variations. Therefore, this study aims to perform sequence analysis in those 15 samples. Furthermore, sequence analysis of BCR-ABL fusion gene breakpoint types in CML patients was never been reported in Indonesia. Materials and Methods Samples and ethical issue Fifteen cdna samples were included in this study. The cdna samples were derived from peripheral blood of BCR-ABL positive CML patients that clinically confirmed at Dr. Sardjito Hospital Yogyakarta Indonesia between March 2010 and August Patients with other malignancy were excluded. This study was approved by the Ethical Committee of the Faculty of Medicine Universitas Gadjah Mada Yogyakarta Indonesia (No. KE/ KF/1029/EC). Reverse Transcriptase-PCR (RT-PCR) Two-step PCR by using inner primer in the second PCR was performed in this study. The first PCR was done either using a set of primer (specific conventional PCR) or primers to detect several fragments in one reaction (multiplex PCR). Initially, all of the samples were amplified by using specific conventional and nested PCR. Once the samples could not be amplified using first PCR set, then they were amplified using multiplex and nested PCR. Figure 1 showed the algorithm of BCR-ABL fusion gene amplification in this study. Conventional or multiplex reverse trancriptase (RT)-PCR was performed according to previous study (Goh et al., 2006). RT-PCR was performed by using Biorad C1000 thermocycler machine and PCR kit from Roche (cat.# ). K562 cells line carrying b3a2 major breakpoint type was used as positive control. All PCR results were visualized by electrophoresis method using 2% agarose gel. Table 1 and 2 demonstrated the primers to amplify breakpoints of interest. The product of conventional PCR can be seen at 616bp for b3a2 and 552bp for b2a2 and for nested PCR should be at 443bp for b3a2 and 368bp 1344 Asian Pacific Journal of Cancer Prevention, Vol 18 for b2a2. The product for multiplex PCR had to be at 627bp for b3a2; 552bp for b2a2; 429bp for minor e1a2; and 1,167 bp for micro e19a2. The expected band size for micro breakpoint type (e19a2) (sequences on exon 19 of BCR gene and exon 3 of ABL gene) was 206bp for e19a2. Sequence analysis All of PCR products were sequenced. The sequencing results were analyzed by using MEGA 5.1 software. The sequencing results were aligned to BCR-ABL sequence from genbank according to their breakpoint type. Results Conventional RT-PCR was used to detect a specific rearrangement of BCR-ABL transcript. All samples (n=15) amplified by conventional RT-PCR using specific primer for major breakpoint type showed negative results (Figure 2A). All products of the conventional RT-PCR were used as DNA template for the second nested PCR using inner primer. Thirteen of 15 samples showed positive band and 2 samples (no 36 and 140) were negative. Seven samples (no 28, 35, 38, 132, 165, 170 and 185) showed band at 443 bp, which were in similar size with positive control band of K562 cell line. Thus, those samples were confirmed as b3a2 major breakpoint. Two samples (no 137 and 138) had fragment at 368bp, specific band for b2a2 major breakpoint. Four samples (no 147, 158, 162, and 164) showed band at 368bp size and additional band (Figure 2B). In order to clarify the position of the bands of samples no 147, 158, 162, and 164, electrophoresis was Figure 1. Algorythm of PCR for BCR-ABL Transcript Type Identification
3 DOI: /APJCP New Genetic Variation in the BCR Gene Figure 3. Sequence Analysis Results. A. BCR exon 13 of Sample 165. Arrow shows sequence variation at 47 th base, C replaced by T; B. BCR exon 14 of Sample 165. Arrow shows sequence variation at 56 th base, T replaced by C. C. BCR exon 13 of Sample 170. Arrow shows sequence variation at 98 th base, T replaced by C. Figure 2. Electrophoresis Result of Every Stage of PCR. A. Conventional RT-PCR with major primer. All samples showed negative result; B. Nested PCR (a) with major primer. Samples 28, 35, 38, 132, 165, 170 and 185 were confirmed as b3a2 major breakpoint type; Samples 137 and 138 were confirmed as b2a2 major breakpoint type; Samples 147, 158, 162, and 164 were confirmed as b2a2 major breakpoint type with additional band; C. Electrophoresis on 4% agarose gel. Samples 147 and 158 were confirmed as b2a2 major breakpoint type; Samples 162 and 164 were confirmed as a combination of both b3a2 and b2a2 type with additional band at 500 bp size; D. Nested PCR (b) using major primer. Sample 36 was confirmed as a combination of both b3a2 and b2a2 type; Sample 140 was confirmed as b2a2 major breakpoint type: (L) DNA ladder; (K+) positive control from K562 cell line; (K-) negative control. repeated using 4% agarose gel. Samples no 147 and 158 confirmed as b2a2 major breakpoint. Samples162 and 164 had combination of both b3a2 and b2a2 breakpoint with additional band at around 500bp (Figure 2C). Samples no 36 and 140, which gave negative result in conventional and nested PCR were amplified by using multiplex RT-PCR and nested. Nested PCR using primer specific for major breakpoint confirmed that Sample 36 had a combination of b2a2 and b3a2 breakpoint; and Sample 140 had major b2a2 breakpoint type (figure 2D). Results of the RT-PCR are listed in Table 3. All samples were sequenced, and the nucleotides sequence was compared to the ones from genbank as the standard. Sequence analysis found that four samples had b3a2 breakpoint type (BCR-ABL-28, -35, -38, -132, and -185) and five samples had b2a2 major breakpoint type (BCR-ABL-137, -138, -140, -147, and -158) and the sequences were similar to the ones in genbank for each type. Sample no 36, which was confirmed as a combination of b3a2 and b2a2 type, had similar sequences with b3a2 and b2a2 in genbank. Sequences of both b3a2 and b2a2 types in samples number 162 and 164 were also similar to the ones in genbank. Additional band at Table 1. Primer Sequences of Major Breakpoint in Conventional RT-PCR and Nested PCR (Goh et al., 2006). Primer Conventioanl RT-PCR Nested PCR nested (a) Sense 5 -ctacggagaggctgaagaa-3 (BCR exon 11) 5 -gtgcagagtggagggagaac-3 (BCR exon 12) Anti sense 5 -cgtgatgtagttgcttggga-3 (ABL exon 3) 5 -acaccattccccattgtgat-3 (ABL exon 3) Table 2. Primer Sequences in Multiplex RT-PCR and Nested PCR (Goh et al., 2006). Primer Multiplex RT-PCR Nested PCR (b) Major : Sense 5 -gctacggagaggctgaagaa-3 (BCR exon 11) 5 -gtgcagagtggagggagaac-3 (BCR exon 12) 5 caacagtccttcgacagcag-3 (BCR exon 1) Micro : 5 -cttcgacgtcaaagcccttc-3 (BCR exon 19) Major : Antisense 5 -cgtgatgtagttgcttggga-3 (ABL exon 3) 5 -acaccattccccattgtgat-3 (ABL exon 3) Micro : 5 -ctaagacccggagcttttca-3 (ABL exon 3) Asian Pacific Journal of Cancer Prevention, Vol
4 Tri Agusti Sholikah et al Table 3. Breakpoint Type of CML Patients Samples No Sample Code Breakpoint Type 1 BCR-ABL-28 B3a2 2 BCR-ABL-35 B3a2 3 BCR-ABL-36 B3a2 and b2a2 4 BCR-ABL-38 B3a2 5 BCR-ABL-132 B3a2 6 BCR-ABL-137 B2a2 7 BCR-ABL-138 B2a2 8 BCR-ABL-140 B2a2 9 BCR-ABL-147 B2a2 10 BCR-ABL-158 B2a2 11 BCR-ABL-162 B3a2, b2a2 and 500 bp 12 BCR-ABL-164 B3a2, b2a2 and 500 bp 13 BCR-ABL-165 B3a2 14 BCR-ABL-170 B3a2 15 BCR-ABL-185 B3a2 500bp in two samples showed similar sequence with b3a2 breakpoint in genbank. Two samples with b3a2 major breakpoint type (Samples 165 and 170) showed sequences variations. Two sequence variations were found in Sample 165, which are at 47 th base of exon 13 and 56 th base of exon 14. Both are in BCR gene. The 47 th base of exon 13 BCR gene (Cytosine) was replaced by Thymin, c.3245c>t (Genbank accession NM_ ; nomenclature according to Antonarakis, 1998) (Figure 3A). This change was occurred at third nucleotide of the codon, and led to silent mutation. The codon changed from ACC into ACT without any amino acid change, which is Threonin. The second sequence variation was the replacement of Thymine into Cytosine at 56 th base of exon 14 BCR gene, c.3359t>c (Genbank accession NM_ ; nomenclature according to Antonarakis, 1998), corresponding to ACT and ACC (Figure 3B). The change also led to silent mutation, because both codons encode Threonin. Those two sequence variations in Sample 165 had not been reported yet. Further study need to be done to determine whether these new variations are polymorphism in the BCR gene. Sequence variation of Sample 170 was the replacement of Thymine into Cytosine at 98 th base of exon 13 BCR gene, c.3296t>c (Genbank accession NM_ ; nomenclature according to Antonarakis, 1998). This change was also occurred in the third nucleotide of the codon, AAT changed into AAC. Both codons encode Asparagine. Sequence variation of sample 170 had been reported as polymorphism in previous studies by De V. Meissner et al., (1998) and Saußele et al., (2000) (Figure 3C). The results of sequence analysis are listed in Table 4. Table 4. Sequence Analysis Results Sample Code Fragment size (bp) BCR exon 12 BCR exon 13 BCR exon 14 ABL exon 1 ABL exon 2 ABL exon 3 Additional Sequence / sequence variation Conclusion B3a B3a B2a B3a B3a B3a B2a B2a B2a B2a B2a B2a B3a B3a B2a B3a B3a th base of BCR exon 13, C T; 56 th base of BCR exon 14, T C th base of BCR exon 13, T C B3a2; c.3245c>t B3a2; c.3359t>c B3a2; c.3296t>c B3a Asian Pacific Journal of Cancer Prevention, Vol 18
5 Discussion Molecular method in CML diagnosis in Molecular Biology Laboratory, Faculty of Medicine, Universitas Gadjah Mada (UGM)-Dr. Sardjito hospital has been done since The qualitative method used in this study was two-step PCR by using inner primer in the second PCR (nested). The two-step PCR included either conventional RT-PCR and nested PCR or multiplex RT-PCR and nested PCR. The methods used in this study were modified from the ones previously reported (Goh et al., 2006). Since more than 95% of CML patients had major breakpoint type (Pane et al., 2002; Shet et al., 2002; Goh et al., 2006), the initial amplification used in this study is conventional RT-PCR to amplify the major breakpoint. All samples (15 samples) showed negative results. They were further amplified by using nested PCR using inner primer for major breakpoint. Nested PCR was a variant of PCR using a pair of primer annealling to the inside area of the first PCR product. Therefore, the product of nested PCR was shorter than the first PCR products. Nested-PCR was a very sensitive and specific method. The sensitivity of nested-pcr was two to three times higher than the first step PCR (Peres de Rozas et al., 2008). The amplification by nested PCR for major breakpoint found that 13 samples were confirmed as major breakpoint, while the other two were negative. The two negative samples were further confirmed that one sample having a combination of b3a2 and b2a2 major breakpoints and the other one have b2a2 major breakpoint. The samples were success to be amplified after the modification of the cycles from 40 to 30 (data not shown). One of the factors that might influence the amplification failure was cycle number. Too many PCR cycles might increase nonspecific amplification (Roux, 2009). Overall, the result got 20 fragments from RT-PCR, that is 7 samples of major b3a2, 5 samples of major b2a2, 1 sample had a combination of b3a2 and b2a2 types and 2 samples had a combination of b3a2 and b2a2 type with additional fragment at 500bp. All of 20 fragments were sequenced. Sequence analysis showed that all of the fragments with b2a2 type had similar sequences with the b2a2 sequence in genbank. Eight of 10 fragments with b3a2 type had a similar sequence with b3a2 sequence in genbank, while the other two (sample no 165 and 170) had sequence variations. Two new sequence variations in sample no 165 (c.3245c>t and c.3359t>c) never been reported yet, while the other one found in sample no 170 (c.3296t>c) had been reported as a polymorphism in previous studies by De V. Meissner et al.(1998) and Saußele et al. (2000). All of the variations (mutations) found in this study did not cause the change of amino acid. Therefore, they do not have implication in the structure of BCR-ABL protein (Saußele et al., 2000). However, eventhough silent mutationt supposed to be have no effect, several studies showed that silent mutation gave harmful effects, by speeding up or slowing down protein synthesis, or affecting the splicing (Mynampati et al., 2016; Yadegari et al., 2016). In this study, we did not see hematological and clinical data of the patients with new mutations, thus DOI: /APJCP New Genetic Variation in the BCR Gene further study need to be done to see whether those silent mutation have any correlation with disease status of the patients. In this study, the mutation was found in only one patient. Study in the population that represent the case is need to prove whether the mutation have harmfull effect to the disease. In addition, the use of samples derived from bone marrow is important because the origin of the disease in the bone marrow, and in this case is to know whether the mutation is germline or somatic. Other than functional effects, silent mutation showed in the sample 170 was identified in the 8th position before the junctional region of BCR-ABL cdna (De V. Meissner et al., 1998). Due to the location of alteration was close to the fusion region, it might have a significant influence on the annealing of PCR primers, probes for real time PCR, and antisense oligonucleotides. A thymidine/cytosine replacement occurs frequently in BCR exon b2. In order to develop the real time PCR for BCR-ABL transcript, the designed probe should not be at the critical region to avoid underestimation of the number of BCR- ABL transcripts (Saußele et al., 2000). The sequence of additional bands at 500bp size in sample number 162 and 164 were similar to the b3a2 sequence in genbank. Thus, the additional bands were suspected as an unspecific band that could be caused by several conditions, such as too many PCR cycles, too long annealing time, too low annealing temperature, and too long extension time. PCR components could also be the source of the problem, such as too high concentration of primers, dntpmix or water contained impurities, or too high concentration of Mg2+ (Henegariu et al., 1997; Roux, 2009). In conclusion, this study found two new base variations in BCR gene of BCR-ABL fusion gene, namely c.3245c>t in exon 13 and c.3359t>c in exon 14. Both variations were silent mutations. Further study need to be done to explore whether the mutations gave harm effect to the disease. Statement of conflict of Interest The authors declare that they have no conflict of interest. Acknowlegments The work is supported by Research Funding for Academic staffs and students of Master s Degree Program (Dana Masyarakat) from the Faculty of Medicine, Universitas Gadjah Mada, Yogyakarta, Indonesia and by Master s Degree Program Scholarship from Directorate of Higher Study (Beasiswa Pendidikan Pascasarjana Dalam Negeri/BPP-DN Dikti). References Antonarakis SE (1998). Recommendations for a nomenclature system for human gene mutations. Hum Mutat, 11, 1-3. Au WY, Caguioa PB, Chuah C, et al (2009). Chronic myeloid leukemia in Asia. Int J Hematol, 89, Baccarani M, Dreyling M (2010). Chronic myeloid leukemia: ESMO clinical practice guidelines for diagnosis, treatment Asian Pacific Journal of Cancer Prevention, Vol
6 Tri Agusti Sholikah et al and follow-up. Ann Oncol, 21, De V. Meissner R, Dias PMB, Covas DT, et al (1998). A polymorphism in exon b2 of the major breakpoint cluster region (M-bcr) identified in chronic myeloid leukaemia patients. Br J Haematol, 103, Druker BJ (2008). Translation of the Philadelphia chromosome into therapy for CML. Blood, 112, Garcia-Manero G, Faderl S, O Brien S, et al (2003). Chronic myelogenous leukemia: A review and update of therapeutic strategies. Cancer, 98, Goh HG, Hwang JY, Kim SH, et al (2006). Comprehensive analysis of BCR-ABL transcript types in Korean CML patients using a newly developed multiplex RT-PCR. Transl Res, 148, Henegariu O, Heerema NA, Dlouhy SR, Vance GH, Vogt PH (1997). Multiplex PCR: Critical parameters and step-by-step protocol. BioTechniques, 23, Huntly BJP, Bench A, Green AR (2003). Double jeopardy from a single translocation: deletions of the derivative chromosome 9 in chronic myeloid leukemia. Blood, 102, Jabbour EJ, Kanterjian H (2014). CME Information: Chronic myeloid leukemia: 2014 update on diagnosis, monitoring, and management. Am J Hematol, 89, Kim DW, Banavali SD, Bunworasate U, et al (2010). Chronic myeloid leukemia in the Asia-Pacific region: Current practice, challenges and opportunities in the targeted therapy era. Leuk Res, 34, Mynampati BK, Muthukumarappa T, Ghosh S, Ram J (2016). A silent mutation in human alpha-a crystallin gene in patients with age-related nuclear or cortical cataract. Bosn J Basic Med Sci, 20, 1-6. Ohsaka A, Shiina S, Kobayashi M, Kudo H, Kawaguchi R (2002). Philadelphia chromosome-positive chronic myeloid leukemia expressing p190bcr-abl. Intern Med, 41, Pane F, Intrieri M, Quintarelli C, et al (2002). BCR/ABL genes and leukemic phenotype: from molecular mechanisms to clinical correlations. Oncogene, 21, Peres de Rozas AM, Gonzalez J, Aloy N, et al (2008). Standardization of nested-pcr for the detection of pasteurella multocida, staphylococcus aureus, myxomatosis virus, and rabbit haemorrhagic disease virus. Pathology and Hygiene, pp Perrotti D, Jamieson C, Goldman J, Skorski T (2010). Chronic myeloid leukemia : mechanism of blastic transformation. J Clin Invest, 120, Roux KH (2009). Optimization and troubleshooting in PCR. cold spring harbor laboratory Press, 4, pp 1-7. Saußele S, Weißer A, Muller M, et al (2000). Frequent polymorphism in BCR exon b2 identified in BCR-ABL positive and negative individuals using fluorescent hybridization probes. Leukemia, 14, Shet A, Jahagirdar B, Verfaillie C (2002). Chronic myelogenous leukemia: mechanisms underlying disease progression. Leukemia, 16, Sillaber C, Mayerhofer M, Agis H, et al (2003). Chronic myeloid leukemia: Pathophysiology, diagnostic parameters,and current treatment concepts. Wien Klin Wochenschr, 115, Smith DL, Burthem J, Whetton AD (2003). Molecular pathogenesis of chronic myeloid leukaemia. Expert Rev Mol Med, 5, Yadegari M, Biswas A, Akher M, et al (2016). Intron retention resulting from a silent mutation in the VWF gene that structurally influence the 5 slice site. Blood, 128, Asian Pacific Journal of Cancer Prevention, Vol 18
ONE STEP MULTIPLEX RT-PCR FOR BCRlABL GENE IN MALAY PATIENTS DIAGNOSED AS LEUKAEMIA
 ONE STEP MULTIPLEX RT-PCR FOR BCRlABL GENE IN MALAY PATIENTS DIAGNOSED AS LEUKAEMIA 1Rosline H, 1Majdan R, 1Wan Zaidah A, 1Rapiaah M, 1Selamah G, 2A A Baba, 3D M Donald 1Department of Haematology, 2Department
ONE STEP MULTIPLEX RT-PCR FOR BCRlABL GENE IN MALAY PATIENTS DIAGNOSED AS LEUKAEMIA 1Rosline H, 1Majdan R, 1Wan Zaidah A, 1Rapiaah M, 1Selamah G, 2A A Baba, 3D M Donald 1Department of Haematology, 2Department
Molecular Detection of BCR/ABL1 for the Diagnosis and Monitoring of CML
 Molecular Detection of BCR/ABL1 for the Diagnosis and Monitoring of CML Imran Mirza, MD, MS, FRCPC Pathology & Laboratory Medicine Institute Sheikh Khalifa Medical City, Abu Dhabi, UAE. imirza@skmc.ae
Molecular Detection of BCR/ABL1 for the Diagnosis and Monitoring of CML Imran Mirza, MD, MS, FRCPC Pathology & Laboratory Medicine Institute Sheikh Khalifa Medical City, Abu Dhabi, UAE. imirza@skmc.ae
A novel isothermal amplification approach for rapid identification of BCR-ABL fusion genes at onset:
 A novel isothermal amplification approach for rapid identification of BCR-ABL fusion genes at onset: Josh Glason: Sales Manager, DiaSorin Australia Pty Ltd September 6, 2014 This product is not currently
A novel isothermal amplification approach for rapid identification of BCR-ABL fusion genes at onset: Josh Glason: Sales Manager, DiaSorin Australia Pty Ltd September 6, 2014 This product is not currently
Role of FISH in Hematological Cancers
 Role of FISH in Hematological Cancers Thomas S.K. Wan PhD,FRCPath,FFSc(RCPA) Honorary Professor, Department of Pathology & Clinical Biochemistry, Queen Mary Hospital, University of Hong Kong. e-mail: wantsk@hku.hk
Role of FISH in Hematological Cancers Thomas S.K. Wan PhD,FRCPath,FFSc(RCPA) Honorary Professor, Department of Pathology & Clinical Biochemistry, Queen Mary Hospital, University of Hong Kong. e-mail: wantsk@hku.hk
Template for Reporting Results of Monitoring Tests for Patients With Chronic Myelogenous Leukemia (BCR-ABL1+)
 Template for Reporting Results of Monitoring Tests for Patients With Chronic Myelogenous Leukemia (BCR-ABL1+) Version: CMLBiomarkers 1.0.0.2 Protocol Posting Date: June 2017 This biomarker template is
Template for Reporting Results of Monitoring Tests for Patients With Chronic Myelogenous Leukemia (BCR-ABL1+) Version: CMLBiomarkers 1.0.0.2 Protocol Posting Date: June 2017 This biomarker template is
Correspondence should be addressed to Hooman H. Rashidi;
 Case Reports in Hematology Volume 2015, Article ID 458052, 6 pages http://dx.doi.org/10.1155/2015/458052 Case Report Optimal Molecular Methods in Detecting p190 BCR-ABL Fusion Variants in Hematologic Malignancies:
Case Reports in Hematology Volume 2015, Article ID 458052, 6 pages http://dx.doi.org/10.1155/2015/458052 Case Report Optimal Molecular Methods in Detecting p190 BCR-ABL Fusion Variants in Hematologic Malignancies:
BCR-ABL1 Kinase Domain Mutation Status (including educational clinical scenario)
 BCR-ABL1 Kinase Domain Mutation Status 171802 (including educational clinical scenario) Issue date: 23rd February 2018 Closing date: 23rd March 2018 Results for the BCR-ABL1 Kinase Domain Mutation Status
BCR-ABL1 Kinase Domain Mutation Status 171802 (including educational clinical scenario) Issue date: 23rd February 2018 Closing date: 23rd March 2018 Results for the BCR-ABL1 Kinase Domain Mutation Status
Chronic myeloid leukemia (CML)
 Bioscientia Int. Support Services Dept. Konrad-Adenauer-Strasse 17 55218 Ingelheim Germany Phone 49(0)6132-781-203 49(0)6132-781-224 49(0)6132-781-165 Fax 49(0)6132-781-236 int.support@bioscientia.com
Bioscientia Int. Support Services Dept. Konrad-Adenauer-Strasse 17 55218 Ingelheim Germany Phone 49(0)6132-781-203 49(0)6132-781-224 49(0)6132-781-165 Fax 49(0)6132-781-236 int.support@bioscientia.com
Differentiation-induced Changes of Mediterranean Fever Gene (MEFV) Expression in HL-60 Cell
 Differentiation-induced Changes of Mediterranean Fever Gene (MEFV) Expression in HL-60 Cell Wenxin Li Department of Biological Sciences Fordham University Abstract MEFV is a human gene that codes for an
Differentiation-induced Changes of Mediterranean Fever Gene (MEFV) Expression in HL-60 Cell Wenxin Li Department of Biological Sciences Fordham University Abstract MEFV is a human gene that codes for an
Molecular Hematopathology Leukemias I. January 14, 2005
 Molecular Hematopathology Leukemias I January 14, 2005 Chronic Myelogenous Leukemia Diagnosis requires presence of Philadelphia chromosome t(9;22)(q34;q11) translocation BCR-ABL is the result BCR on chr
Molecular Hematopathology Leukemias I January 14, 2005 Chronic Myelogenous Leukemia Diagnosis requires presence of Philadelphia chromosome t(9;22)(q34;q11) translocation BCR-ABL is the result BCR on chr
The Human Major Histocompatibility Complex
 The Human Major Histocompatibility Complex 1 Location and Organization of the HLA Complex on Chromosome 6 NEJM 343(10):702-9 2 Inheritance of the HLA Complex Haplotype Inheritance (Family Study) 3 Structure
The Human Major Histocompatibility Complex 1 Location and Organization of the HLA Complex on Chromosome 6 NEJM 343(10):702-9 2 Inheritance of the HLA Complex Haplotype Inheritance (Family Study) 3 Structure
Leukemia BCR-ABL Fusion Gene Real Time RT-PCR Kit
 Revision No.: ZJ0003 Issue Date: Aug 7 th, 2008 Leukemia BCR-ABL Fusion Gene Real Time RT-PCR Kit Cat. No.: TR-0126-02 For use with ABI Prism 7000/7300/7500/7900(96 well); Smart Cycler II; icycler iq 4/iQ
Revision No.: ZJ0003 Issue Date: Aug 7 th, 2008 Leukemia BCR-ABL Fusion Gene Real Time RT-PCR Kit Cat. No.: TR-0126-02 For use with ABI Prism 7000/7300/7500/7900(96 well); Smart Cycler II; icycler iq 4/iQ
SALSA MLPA probemix P315-B1 EGFR
 SALSA MLPA probemix P315-B1 EGFR Lot B1-0215 and B1-0112. As compared to the previous A1 version (lot 0208), two mutation-specific probes for the EGFR mutations L858R and T709M as well as one additional
SALSA MLPA probemix P315-B1 EGFR Lot B1-0215 and B1-0112. As compared to the previous A1 version (lot 0208), two mutation-specific probes for the EGFR mutations L858R and T709M as well as one additional
Patterns of BCR/ABL Gene Rearrangements by Fluorescence in Situ Hybridization (FISH) in Chronic Myeloid Leukaemia
 Pak J Med Res Vol. 57, No. 4, 2018 Original Article Patterns of BCR/ABL Gene Rearrangements by Fluorescence in Situ Hybridization (FISH) in Chronic Myeloid Leukaemia Khadija Iftikhar, Chaudhry Altaf Hussain,
Pak J Med Res Vol. 57, No. 4, 2018 Original Article Patterns of BCR/ABL Gene Rearrangements by Fluorescence in Situ Hybridization (FISH) in Chronic Myeloid Leukaemia Khadija Iftikhar, Chaudhry Altaf Hussain,
Chapter 4 Cellular Oncogenes ~ 4.6 -
 Chapter 4 Cellular Oncogenes - 4.2 ~ 4.6 - Many retroviruses carrying oncogenes have been found in chickens and mice However, attempts undertaken during the 1970s to isolate viruses from most types of
Chapter 4 Cellular Oncogenes - 4.2 ~ 4.6 - Many retroviruses carrying oncogenes have been found in chickens and mice However, attempts undertaken during the 1970s to isolate viruses from most types of
How to detect the rare BCR ABL (e14a3) transcript: A case report and literature review
 ONCOLOGY LETTERS 14: 5619-5623, 2017 How to detect the rare BCR ABL (e14a3) transcript: A case report and literature review LIN HUI HU, LIAN FANG PU, DONG DONG YANG, CUI ZHANG, HUI PING WANG, YANG YANG
ONCOLOGY LETTERS 14: 5619-5623, 2017 How to detect the rare BCR ABL (e14a3) transcript: A case report and literature review LIN HUI HU, LIAN FANG PU, DONG DONG YANG, CUI ZHANG, HUI PING WANG, YANG YANG
MRD in CML (BCR-ABL1)
 MRD in CML (BCR-ABL1) Moleculaire Biologie en Cytometrie cursus Barbara Denys LAbo Hematologie UZ Gent 6 mei 2011 2008 Universitair Ziekenhuis Gent 1 Myeloproliferative Neoplasms o WHO classification 2008:
MRD in CML (BCR-ABL1) Moleculaire Biologie en Cytometrie cursus Barbara Denys LAbo Hematologie UZ Gent 6 mei 2011 2008 Universitair Ziekenhuis Gent 1 Myeloproliferative Neoplasms o WHO classification 2008:
Major molecular response to imatinib in a patient with chronic myeloid leukemia expressing a novel form of e8a2 BCR-ABL transcript.
 Major molecular response to imatinib in a patient with chronic myeloid leukemia expressing a novel form of e8a2 BCR-ABL transcript. Andrei Tchirkov, Jean-Louis Couderc, Bernard Périssel, Carole Goumy,
Major molecular response to imatinib in a patient with chronic myeloid leukemia expressing a novel form of e8a2 BCR-ABL transcript. Andrei Tchirkov, Jean-Louis Couderc, Bernard Périssel, Carole Goumy,
Hematologic and cytogenetic responses of Imatinib Mesylate and significance of Sokal score in chronic myeloid leukemia patients
 ORIGINAL ALBANIAN MEDICAL RESEARCH JOURNAL Hematologic and cytogenetic responses of Imatinib Mesylate and significance of Sokal score in chronic myeloid leukemia patients Dorina Roko 1, Anila Babameto-Laku
ORIGINAL ALBANIAN MEDICAL RESEARCH JOURNAL Hematologic and cytogenetic responses of Imatinib Mesylate and significance of Sokal score in chronic myeloid leukemia patients Dorina Roko 1, Anila Babameto-Laku
Diagnosis. Chapter 2. Diagnostic laboratory tests. Blood picture and biochemistry. Bone marrow morphology
 Chapter 2 Diagnosis When a patient presents with suspected chronic myeloid leukemia (CML) appropriate assessments are needed to confirm the diagnosis and stage of disease, and to assign a risk score to
Chapter 2 Diagnosis When a patient presents with suspected chronic myeloid leukemia (CML) appropriate assessments are needed to confirm the diagnosis and stage of disease, and to assign a risk score to
"Molecular Monitoring Strategies to Track Response to BCR-ABL Kinase Inhibitors in CML"
 Association of Molecular Pathology USCAP Companion Meeting Sunday, February 12, 2006 7:00 PM Dan Jones, MD, PhD Associate Professor Medical Director, Molecular Diagnostic Laboratory Division of Pathology
Association of Molecular Pathology USCAP Companion Meeting Sunday, February 12, 2006 7:00 PM Dan Jones, MD, PhD Associate Professor Medical Director, Molecular Diagnostic Laboratory Division of Pathology
Bio 111 Study Guide Chapter 17 From Gene to Protein
 Bio 111 Study Guide Chapter 17 From Gene to Protein BEFORE CLASS: Reading: Read the introduction on p. 333, skip the beginning of Concept 17.1 from p. 334 to the bottom of the first column on p. 336, and
Bio 111 Study Guide Chapter 17 From Gene to Protein BEFORE CLASS: Reading: Read the introduction on p. 333, skip the beginning of Concept 17.1 from p. 334 to the bottom of the first column on p. 336, and
Chronic Myelogenous Leukemia with e13a3 (b2a3) Type of BCR-ABL Transcript Having a DNA Breakpoint between ABL exons a2 and a3
 American Journal of Hematology 74:268 272 (2003) Chronic Myelogenous Leukemia with e13a3 (b2a3) Type of BCR-ABL Transcript Having a DNA Breakpoint between ABL exons a2 and a3 Li-Gen Liu, 1,2 Hideo Tanaka,
American Journal of Hematology 74:268 272 (2003) Chronic Myelogenous Leukemia with e13a3 (b2a3) Type of BCR-ABL Transcript Having a DNA Breakpoint between ABL exons a2 and a3 Li-Gen Liu, 1,2 Hideo Tanaka,
Introduction to Genetics
 Introduction to Genetics Table of contents Chromosome DNA Protein synthesis Mutation Genetic disorder Relationship between genes and cancer Genetic testing Technical concern 2 All living organisms consist
Introduction to Genetics Table of contents Chromosome DNA Protein synthesis Mutation Genetic disorder Relationship between genes and cancer Genetic testing Technical concern 2 All living organisms consist
Supplementary Figure 1. Spitzoid Melanoma with PPFIBP1-MET fusion. (a) Histopathology (4x) shows a domed papule with melanocytes extending into the
 Supplementary Figure 1. Spitzoid Melanoma with PPFIBP1-MET fusion. (a) Histopathology (4x) shows a domed papule with melanocytes extending into the deep dermis. (b) The melanocytes demonstrate abundant
Supplementary Figure 1. Spitzoid Melanoma with PPFIBP1-MET fusion. (a) Histopathology (4x) shows a domed papule with melanocytes extending into the deep dermis. (b) The melanocytes demonstrate abundant
[COMPREHENSIVE GENETIC ASSAY PANEL ON
 2014 SN GENELAB AND RESEARCH CENTER DR. SALIL VANIAWALA, PH.D [COMPREHENSIVE GENETIC ASSAY PANEL ON MYELOPROLIFERATIVE NEOPLASMS] SN Genelab presents one of the most comprehensive genetic assay panel for
2014 SN GENELAB AND RESEARCH CENTER DR. SALIL VANIAWALA, PH.D [COMPREHENSIVE GENETIC ASSAY PANEL ON MYELOPROLIFERATIVE NEOPLASMS] SN Genelab presents one of the most comprehensive genetic assay panel for
JAK2 V617F analysis. Indication: monitoring of therapy
 JAK2 V617F analysis BCR-ABL genotyping The exact chromosomal defect in Philadelphia chromosome is a translocation. Parts of two chromosomes, 9 and 22, switch places. The result is a fusion gene, created
JAK2 V617F analysis BCR-ABL genotyping The exact chromosomal defect in Philadelphia chromosome is a translocation. Parts of two chromosomes, 9 and 22, switch places. The result is a fusion gene, created
Iran J Cancer Preven May; 8(3):e2334. Published online 2015 May 27. Research Article
 Iran J Cancer Preven. 2015 May; 8(3):e2334. Published online 2015 May 27. Research Article Characterizing of Four Common BCR-ABL Kinase Domain Mutations (T315I, Y253H, M351T and E255K) in Iranian Chronic
Iran J Cancer Preven. 2015 May; 8(3):e2334. Published online 2015 May 27. Research Article Characterizing of Four Common BCR-ABL Kinase Domain Mutations (T315I, Y253H, M351T and E255K) in Iranian Chronic
SUPPLEMENTARY INFORMATION
 doi: 1.138/nature8645 Physical coverage (x haploid genomes) 11 6.4 4.9 6.9 6.7 4.4 5.9 9.1 7.6 125 Neither end mapped One end mapped Chimaeras Correct Reads (million ns) 1 75 5 25 HCC1187 HCC1395 HCC1599
doi: 1.138/nature8645 Physical coverage (x haploid genomes) 11 6.4 4.9 6.9 6.7 4.4 5.9 9.1 7.6 125 Neither end mapped One end mapped Chimaeras Correct Reads (million ns) 1 75 5 25 HCC1187 HCC1395 HCC1599
Test Name Results Units Bio. Ref. Interval. Positive
 LL - LL-ROHINI (NATIONAL REFERENCE 135091534 Age 36 Years Gender Female 1/9/2017 120000AM 1/9/2017 105316AM 2/9/2017 104147AM Ref By Final LEUKEMIA GENETIC ROFILE ANY SIX MARKERS, CR QUALITATIVE AML ETO
LL - LL-ROHINI (NATIONAL REFERENCE 135091534 Age 36 Years Gender Female 1/9/2017 120000AM 1/9/2017 105316AM 2/9/2017 104147AM Ref By Final LEUKEMIA GENETIC ROFILE ANY SIX MARKERS, CR QUALITATIVE AML ETO
A. ILEA 1* A. M. VLADAREANU 2 E. NICULESCU-MIZIL 3
 Bulletin of the Transilvania University of Braşov Series VI: Medical Sciences Vol. 9 (58) No. 1-2016 THE SIMULTANEOUS OCCURRENCE OF THE BCR-ABL TRANSLOCATION AND THE Jak2V617F MUTATION. DETECTION AND DYNAMICS
Bulletin of the Transilvania University of Braşov Series VI: Medical Sciences Vol. 9 (58) No. 1-2016 THE SIMULTANEOUS OCCURRENCE OF THE BCR-ABL TRANSLOCATION AND THE Jak2V617F MUTATION. DETECTION AND DYNAMICS
AIDS - Knowledge and Dogma. Conditions for the Emergence and Decline of Scientific Theories Congress, July 16/ , Vienna, Austria
 AIDS - Knowledge and Dogma Conditions for the Emergence and Decline of Scientific Theories Congress, July 16/17 2010, Vienna, Austria Reliability of PCR to detect genetic sequences from HIV Juan Manuel
AIDS - Knowledge and Dogma Conditions for the Emergence and Decline of Scientific Theories Congress, July 16/17 2010, Vienna, Austria Reliability of PCR to detect genetic sequences from HIV Juan Manuel
Blast Phase Chronic Myelogenous Leukemia
 Blast Phase Chronic Myelogenous Leukemia Benjamin Powers, MD; and Suman Kambhampati, MD The dramatic improvement in survival with tyrosine kinase inhibitors has not been demonstrated in the advanced blast
Blast Phase Chronic Myelogenous Leukemia Benjamin Powers, MD; and Suman Kambhampati, MD The dramatic improvement in survival with tyrosine kinase inhibitors has not been demonstrated in the advanced blast
Methylation status of SOCS1 and SOCS3 in BCR-ABL negative and. JAK2V617F negative chronic myeloproliferative disorders.
 Methylation status of SOCS1 and SOCS3 in BCR-ABL negative and JAK2V617F negative chronic myeloproliferative disorders. To the Editor BCR-ABL negative Chronic Myeloproliferative Disorders (s) are a heterogeneous
Methylation status of SOCS1 and SOCS3 in BCR-ABL negative and JAK2V617F negative chronic myeloproliferative disorders. To the Editor BCR-ABL negative Chronic Myeloproliferative Disorders (s) are a heterogeneous
Application. Application of molecular biology methods in hematology and oncology. Oncogenes are involved in:
 Application Application of molecular biology methods in hematology and oncology Research on the etiology of cancer Diagnostics / Differential diagnostics Stratifying for treatment Prognosing the outcome
Application Application of molecular biology methods in hematology and oncology Research on the etiology of cancer Diagnostics / Differential diagnostics Stratifying for treatment Prognosing the outcome
MP BCR-ABL1 Testing in Chronic Myelogenous Leukemia and Acute Lymphoblastic Leukemia
 Medical Policy BCBSA Ref. Policy: 2.04.85 Last Review: 10/18/2018 Effective Date: 10/18/2018 Section: Medicine Related Policies 8.01.30 Hematopoietic Cell Transplantation for Chronic Myelogenous Leukemia
Medical Policy BCBSA Ref. Policy: 2.04.85 Last Review: 10/18/2018 Effective Date: 10/18/2018 Section: Medicine Related Policies 8.01.30 Hematopoietic Cell Transplantation for Chronic Myelogenous Leukemia
Structural Variation and Medical Genomics
 Structural Variation and Medical Genomics Andrew King Department of Biomedical Informatics July 8, 2014 You already know about small scale genetic mutations Single nucleotide polymorphism (SNPs) Deletions,
Structural Variation and Medical Genomics Andrew King Department of Biomedical Informatics July 8, 2014 You already know about small scale genetic mutations Single nucleotide polymorphism (SNPs) Deletions,
Monitor TM assay: comparison with traditional RT-qPCR methods
 Preliminary evaluation of the GeneXpert Dx System for CML patients monitoring through the Xpert BCR-ABL Monitor TM assay: comparison with traditional RT-qPCR methods Silvia Calatroni Division of Hematology
Preliminary evaluation of the GeneXpert Dx System for CML patients monitoring through the Xpert BCR-ABL Monitor TM assay: comparison with traditional RT-qPCR methods Silvia Calatroni Division of Hematology
Template for Reporting Results of Monitoring Tests for Patients With Chronic Myelogenous Leukemia (BCR-ABL1+)
 Template for Reporting Results of Monitoring Tests for Patients With Chronic Myelogenous Leukemia (BCR-ABL1+) Template web posting date: July 2015 Authors Todd W. Kelley, MD, FCAP University of Utah and
Template for Reporting Results of Monitoring Tests for Patients With Chronic Myelogenous Leukemia (BCR-ABL1+) Template web posting date: July 2015 Authors Todd W. Kelley, MD, FCAP University of Utah and
Hematopathology Case Study
 Hematopathology Case Study AMP Outreach Course 2009 AMP Annual Meeting John Greg Howe Ph.D. Department of Laboratory Medicine Yale University School of Medicine November 19, 2009 HISTORY Case History An
Hematopathology Case Study AMP Outreach Course 2009 AMP Annual Meeting John Greg Howe Ph.D. Department of Laboratory Medicine Yale University School of Medicine November 19, 2009 HISTORY Case History An
SALSA MLPA KIT P060-B2 SMA
 SALSA MLPA KIT P6-B2 SMA Lot 111, 511: As compared to the previous version B1 (lot 11), the 88 and 96 nt DNA Denaturation control fragments have been replaced (QDX2). Please note that, in contrast to the
SALSA MLPA KIT P6-B2 SMA Lot 111, 511: As compared to the previous version B1 (lot 11), the 88 and 96 nt DNA Denaturation control fragments have been replaced (QDX2). Please note that, in contrast to the
oncogenes-and- tumour-suppressor-genes)
 Special topics in tumor biochemistry oncogenes-and- tumour-suppressor-genes) Speaker: Prof. Jiunn-Jye Chuu E-Mail: jjchuu@mail.stust.edu.tw Genetic Basis of Cancer Cancer-causing mutations Disease of aging
Special topics in tumor biochemistry oncogenes-and- tumour-suppressor-genes) Speaker: Prof. Jiunn-Jye Chuu E-Mail: jjchuu@mail.stust.edu.tw Genetic Basis of Cancer Cancer-causing mutations Disease of aging
Chapter 16 Mutations. Practice Questions:
 Biology 234 J. G. Doheny Chapter 16 Mutations Practice Questions: Answer the following questions with one or two sentences. 1. List the name of one test that can be used to identify mutagens. 2. What is
Biology 234 J. G. Doheny Chapter 16 Mutations Practice Questions: Answer the following questions with one or two sentences. 1. List the name of one test that can be used to identify mutagens. 2. What is
ADx Bone Marrow Report. Patient Information Referring Physician Specimen Information
 ADx Bone Marrow Report Patient Information Referring Physician Specimen Information Patient Name: Specimen: Bone Marrow Site: Left iliac Physician: Accession #: ID#: Reported: 08/19/2014 - CHRONIC MYELOGENOUS
ADx Bone Marrow Report Patient Information Referring Physician Specimen Information Patient Name: Specimen: Bone Marrow Site: Left iliac Physician: Accession #: ID#: Reported: 08/19/2014 - CHRONIC MYELOGENOUS
Nucleic Acid Testing - Oncology. Molecular Diagnosis. Gain/Loss of Nucleic Acid. Objectives. MYCN and Neuroblastoma. Molecular Diagnosis
 Nucleic Acid Testing - Oncology Molecular Diagnosis Nucleic acid based testing in Oncology Gross alterations in DNA content of tumors (ploidy) Gain/Loss of nucleic acids Markers of Clonality Oncogene/Tumor
Nucleic Acid Testing - Oncology Molecular Diagnosis Nucleic acid based testing in Oncology Gross alterations in DNA content of tumors (ploidy) Gain/Loss of nucleic acids Markers of Clonality Oncogene/Tumor
CHAPTER 4 RESULTS. showed that all three replicates had similar growth trends (Figure 4.1) (p<0.05; p=0.0000)
 CHAPTER 4 RESULTS 4.1 Growth Characterization of C. vulgaris 4.1.1 Optical Density Growth study of Chlorella vulgaris based on optical density at 620 nm (OD 620 ) showed that all three replicates had similar
CHAPTER 4 RESULTS 4.1 Growth Characterization of C. vulgaris 4.1.1 Optical Density Growth study of Chlorella vulgaris based on optical density at 620 nm (OD 620 ) showed that all three replicates had similar
Molecular Diagnosis. Nucleic acid based testing in Oncology
 Molecular Diagnosis Nucleic acid based testing in Oncology Objectives Describe uses of NAT in Oncology Diagnosis, Prediction, monitoring. Genetics Screening, presymptomatic testing, diagnostic testing,
Molecular Diagnosis Nucleic acid based testing in Oncology Objectives Describe uses of NAT in Oncology Diagnosis, Prediction, monitoring. Genetics Screening, presymptomatic testing, diagnostic testing,
MPL W515L K mutation
 MPL W515L K mutation BCR-ABL genotyping The exact chromosomal defect in Philadelphia chromosome is a translocation. Parts of two chromosomes, 9 and 22, switch places. The result is a fusion gene, created
MPL W515L K mutation BCR-ABL genotyping The exact chromosomal defect in Philadelphia chromosome is a translocation. Parts of two chromosomes, 9 and 22, switch places. The result is a fusion gene, created
An update on imatinib mesylate therapy in chronic myeloid leukaemia patients in a teaching hospital in Malaysia
 Singapore Med J 2012; 53(1) : 57 An update on imatinib mesylate therapy in chronic myeloid leukaemia patients in a teaching hospital in Malaysia Bee PC 1, MD, MMed, Gan GG 1, MBBS, FRCP, Tai YT 1, MBBS,
Singapore Med J 2012; 53(1) : 57 An update on imatinib mesylate therapy in chronic myeloid leukaemia patients in a teaching hospital in Malaysia Bee PC 1, MD, MMed, Gan GG 1, MBBS, FRCP, Tai YT 1, MBBS,
Identification of Novel Variant of EML4-ALK Fusion Gene in NSCLC: Potential Benefits of the RT-PCR Method
 International journal of Biomedical science ORIGINAL ARTICLE Identification of Novel Variant of EML4-ALK Fusion Gene in NSCLC: Potential Benefits of the RT-PCR Method Martin K. H. Maus 1, 2, Craig Stephens
International journal of Biomedical science ORIGINAL ARTICLE Identification of Novel Variant of EML4-ALK Fusion Gene in NSCLC: Potential Benefits of the RT-PCR Method Martin K. H. Maus 1, 2, Craig Stephens
MRC-Holland MLPA. Description version 19;
 SALSA MLPA probemix P6-B2 SMA Lot B2-712, B2-312, B2-111, B2-511: As compared to the previous version B1 (lot B1-11), the 88 and 96 nt DNA Denaturation control fragments have been replaced (QDX2). SPINAL
SALSA MLPA probemix P6-B2 SMA Lot B2-712, B2-312, B2-111, B2-511: As compared to the previous version B1 (lot B1-11), the 88 and 96 nt DNA Denaturation control fragments have been replaced (QDX2). SPINAL
George R. Honig Junius G. Adams III. Human Hemoglobin. Genetics. Springer-Verlag Wien New York
 George R. Honig Junius G. Adams III Human Hemoglobin Genetics Springer-Verlag Wien New York George R. Honig, M.D., Ph.D. Professor and Head Department of Pediatrics, College of Medicine University of Illinois
George R. Honig Junius G. Adams III Human Hemoglobin Genetics Springer-Verlag Wien New York George R. Honig, M.D., Ph.D. Professor and Head Department of Pediatrics, College of Medicine University of Illinois
Diagnostic challenge: Acute leukemia with biphenotypic blasts and BCR-ABL1 translocation
 Case Study Diagnostic challenge: Acute leukemia with biphenotypic blasts and BCR-ABL1 translocation Ling Wang 1 and Xiangdong Xu 1,2,* 1 Department of Pathology, University of California, San Diego; 2
Case Study Diagnostic challenge: Acute leukemia with biphenotypic blasts and BCR-ABL1 translocation Ling Wang 1 and Xiangdong Xu 1,2,* 1 Department of Pathology, University of California, San Diego; 2
New: P077 BRCA2. This new probemix can be used to confirm results obtained with P045 BRCA2 probemix.
 SALSA MLPA KIT P045-B2 BRCA2/CHEK2 Lot 0410, 0609. As compared to version B1, four reference probes have been replaced and extra control fragments at 100 and 105 nt (X/Y specific) have been included. New:
SALSA MLPA KIT P045-B2 BRCA2/CHEK2 Lot 0410, 0609. As compared to version B1, four reference probes have been replaced and extra control fragments at 100 and 105 nt (X/Y specific) have been included. New:
Role of Paired Box9 (PAX9) (rs ) and Muscle Segment Homeobox1 (MSX1) (581C>T) Gene Polymorphisms in Tooth Agenesis
 EC Dental Science Special Issue - 2017 Role of Paired Box9 (PAX9) (rs2073245) and Muscle Segment Homeobox1 (MSX1) (581C>T) Gene Polymorphisms in Tooth Agenesis Research Article Dr. Sonam Sethi 1, Dr. Anmol
EC Dental Science Special Issue - 2017 Role of Paired Box9 (PAX9) (rs2073245) and Muscle Segment Homeobox1 (MSX1) (581C>T) Gene Polymorphisms in Tooth Agenesis Research Article Dr. Sonam Sethi 1, Dr. Anmol
Activation of cellular proto-oncogenes to oncogenes. How was active Ras identified?
 Dominant Acting Oncogenes Eugene E. Marcantonio, M.D. Ph.D. Oncogenes are altered forms of normal cellular genes called proto-oncogenes that are involved in pathways regulating cell growth, differentiation,
Dominant Acting Oncogenes Eugene E. Marcantonio, M.D. Ph.D. Oncogenes are altered forms of normal cellular genes called proto-oncogenes that are involved in pathways regulating cell growth, differentiation,
RESEARCH ARTICLE. Introduction. Methods Wiley Periodicals, Inc.
 BCR/ABL level at 6 months identifies good risk CML subgroup after failing early molecular response at 3 months following imatinib therapy for CML in chronic phase AJH Dennis (Dong Hwan) Kim, 1 * Nada Hamad,
BCR/ABL level at 6 months identifies good risk CML subgroup after failing early molecular response at 3 months following imatinib therapy for CML in chronic phase AJH Dennis (Dong Hwan) Kim, 1 * Nada Hamad,
Managing Chronic Myeloid Leukemia
 Concise Reference Managing Chronic Myeloid Leukemia Timothy P Hughes, David M Ross, Junia V Melo Extracted from Handbook of Chronic Myeloid Leukemia Published by Springer Healthcare Ltd, 236 Gray s Inn
Concise Reference Managing Chronic Myeloid Leukemia Timothy P Hughes, David M Ross, Junia V Melo Extracted from Handbook of Chronic Myeloid Leukemia Published by Springer Healthcare Ltd, 236 Gray s Inn
Endocrine Surgery. Characteristics of the Germline MEN1 Mutations in Korea: A Literature Review ORIGINAL ARTICLE. The Korean Journal of INTRODUCTION
 ORIGINAL ARTICLE ISSN 1598-1703 (Print) ISSN 2287-6782 (Online) Korean J Endocrine Surg 2014;14:7-11 The Korean Journal of Endocrine Surgery Characteristics of the Germline MEN1 Mutations in Korea: A Literature
ORIGINAL ARTICLE ISSN 1598-1703 (Print) ISSN 2287-6782 (Online) Korean J Endocrine Surg 2014;14:7-11 The Korean Journal of Endocrine Surgery Characteristics of the Germline MEN1 Mutations in Korea: A Literature
Haemoglobinopathies case studies 11 th Annual Sickle Cell and Thalassaemia Conference October 2017
 Haemoglobinopathies case studies 11 th Annual Sickle Cell and Thalassaemia Conference 11 13 October 2017 Chris Lambert Haematology Service Delivery Manager Viapath Laboratories Kings College Hospital HUMAN
Haemoglobinopathies case studies 11 th Annual Sickle Cell and Thalassaemia Conference 11 13 October 2017 Chris Lambert Haematology Service Delivery Manager Viapath Laboratories Kings College Hospital HUMAN
MRC-Holland MLPA. Description version 08; 18 November 2016
 SALSA MLPA probemix P122-D1 NF1 AREA Lot D1-1016. As compared to lot C2-0312, four probes in the NF1 area and one reference probe have been removed, four reference probes have been replaced and several
SALSA MLPA probemix P122-D1 NF1 AREA Lot D1-1016. As compared to lot C2-0312, four probes in the NF1 area and one reference probe have been removed, four reference probes have been replaced and several
LESSON 3.2 WORKBOOK. How do normal cells become cancer cells? Workbook Lesson 3.2
 For a complete list of defined terms, see the Glossary. Transformation the process by which a cell acquires characteristics of a tumor cell. LESSON 3.2 WORKBOOK How do normal cells become cancer cells?
For a complete list of defined terms, see the Glossary. Transformation the process by which a cell acquires characteristics of a tumor cell. LESSON 3.2 WORKBOOK How do normal cells become cancer cells?
Chronic Myeloid Leukaemia
 Chronic Myeloid Leukaemia Molecular Response: What is really important? Jeff Szer The Royal Melbourne Hospital PROBABILITY, % PROBABILITY OF SURVIVAL AFTER MYELOABLATIVE TRANSPLANTS FOR CML IN CHRONIC
Chronic Myeloid Leukaemia Molecular Response: What is really important? Jeff Szer The Royal Melbourne Hospital PROBABILITY, % PROBABILITY OF SURVIVAL AFTER MYELOABLATIVE TRANSPLANTS FOR CML IN CHRONIC
Beta Thalassemia Case Study Introduction to Bioinformatics
 Beta Thalassemia Case Study Sami Khuri Department of Computer Science San José State University San José, California, USA sami.khuri@sjsu.edu www.cs.sjsu.edu/faculty/khuri Outline v Hemoglobin v Alpha
Beta Thalassemia Case Study Sami Khuri Department of Computer Science San José State University San José, California, USA sami.khuri@sjsu.edu www.cs.sjsu.edu/faculty/khuri Outline v Hemoglobin v Alpha
Characterisation of structural variation in breast. cancer genomes using paired-end sequencing on. the Illumina Genome Analyser
 Characterisation of structural variation in breast cancer genomes using paired-end sequencing on the Illumina Genome Analyser Phil Stephens Cancer Genome Project Why is it important to study cancer? Why
Characterisation of structural variation in breast cancer genomes using paired-end sequencing on the Illumina Genome Analyser Phil Stephens Cancer Genome Project Why is it important to study cancer? Why
SALSA MLPA probemix P241-D2 MODY mix 1 Lot D As compared to version D1 (lot D1-0911), one reference probe has been replaced.
 mix P241-D2 MODY mix 1 Lot D2-0413. As compared to version D1 (lot D1-0911), one reference has been replaced. Maturity-Onset Diabetes of the Young (MODY) is a distinct form of non insulin-dependent diabetes
mix P241-D2 MODY mix 1 Lot D2-0413. As compared to version D1 (lot D1-0911), one reference has been replaced. Maturity-Onset Diabetes of the Young (MODY) is a distinct form of non insulin-dependent diabetes
NEXT GENERATION SEQUENCING. R. Piazza (MD, PhD) Dept. of Medicine and Surgery, University of Milano-Bicocca
 NEXT GENERATION SEQUENCING R. Piazza (MD, PhD) Dept. of Medicine and Surgery, University of Milano-Bicocca SANGER SEQUENCING 5 3 3 5 + Capillary Electrophoresis DNA NEXT GENERATION SEQUENCING SOLEXA-ILLUMINA
NEXT GENERATION SEQUENCING R. Piazza (MD, PhD) Dept. of Medicine and Surgery, University of Milano-Bicocca SANGER SEQUENCING 5 3 3 5 + Capillary Electrophoresis DNA NEXT GENERATION SEQUENCING SOLEXA-ILLUMINA
CML CML CML. tyrosine kinase inhibitor CML. 22 t(9;22)(q34;q11) chronic myeloid leukemia CML ABL. BCR-ABL c- imatinib mesylate CML CML BCR-ABL
 1 Key Wordschronic myeloid leukemiaimatinib mesylate tyrosine kinase inhibitor chronic myeloid leukemia CML imatinib mesylate CML CML CML CML Ph 10 1 30 50 3 5 CML α IFNα Ph Ph cytogenetic response CRmajor
1 Key Wordschronic myeloid leukemiaimatinib mesylate tyrosine kinase inhibitor chronic myeloid leukemia CML imatinib mesylate CML CML CML CML Ph 10 1 30 50 3 5 CML α IFNα Ph Ph cytogenetic response CRmajor
Atypical BCR-ABL fusion transcript (e6a2) in pediatric acute lymphoblastic leukemia
 www.edoriumjournals.com CASE REPORT OPEN ACCESS Atypical BCR-ABL fusion transcript (e6a2) in pediatric acute lymphoblastic leukemia Dushyant Kumar, Manoj Kumar Panigrahi, Deepti Dewangan, Sarjana Dutt,
www.edoriumjournals.com CASE REPORT OPEN ACCESS Atypical BCR-ABL fusion transcript (e6a2) in pediatric acute lymphoblastic leukemia Dushyant Kumar, Manoj Kumar Panigrahi, Deepti Dewangan, Sarjana Dutt,
Laboratory Tools for Diagnosis and Monitoring Response in Patients with Chronic Myeloid Leukemia
 in chromosome Laboratory Tools for Diagnosis and Monitoring Response in Patients with Chronic Myeloid Leukemia Tali Tohami PhD 1, Arnon Nagler MD 2,3 and Ninette Amariglio PhD 1 1 Hematology Laboratory
in chromosome Laboratory Tools for Diagnosis and Monitoring Response in Patients with Chronic Myeloid Leukemia Tali Tohami PhD 1, Arnon Nagler MD 2,3 and Ninette Amariglio PhD 1 1 Hematology Laboratory
Test Name Results Units Bio. Ref. Interval. Positive
 LL - LL-ROHINI (NATIONAL REFERENCE 135091533 Age 28 Years Gender Male 1/9/2017 120000AM 1/9/2017 105415AM 4/9/2017 23858M Ref By Final LEUKEMIA DIAGNOSTIC COMREHENSIVE ROFILE, ANY 6 MARKERS t (1;19) (q23
LL - LL-ROHINI (NATIONAL REFERENCE 135091533 Age 28 Years Gender Male 1/9/2017 120000AM 1/9/2017 105415AM 4/9/2017 23858M Ref By Final LEUKEMIA DIAGNOSTIC COMREHENSIVE ROFILE, ANY 6 MARKERS t (1;19) (q23
Template for Reporting Results of Biomarker Testing for Myeloproliferative Neoplasms
 Template for Reporting Results of Biomarker Testing for Myeloproliferative Neoplasms Version: MPNBiomarkers 1.0.0.2 Protocol Posting Date: June 2017 This biomarker template is NOT required for accreditation
Template for Reporting Results of Biomarker Testing for Myeloproliferative Neoplasms Version: MPNBiomarkers 1.0.0.2 Protocol Posting Date: June 2017 This biomarker template is NOT required for accreditation
Chronic myelogenous leukemia (CML) is a slowprogressing
 At a Glance Practical Implications p e148 Author Information p e151 Full text and PDF Web exclusive Patterns of Specific Testing for Patients With Chronic Myelogenous Leukemia Original Research Allison
At a Glance Practical Implications p e148 Author Information p e151 Full text and PDF Web exclusive Patterns of Specific Testing for Patients With Chronic Myelogenous Leukemia Original Research Allison
Reporting TP53 gene analysis results in CLL
 Reporting TP53 gene analysis results in CLL Mutations in TP53 - From discovery to clinical practice in CLL Discovery Validation Clinical practice Variant diversity *Leroy at al, Cancer Research Review
Reporting TP53 gene analysis results in CLL Mutations in TP53 - From discovery to clinical practice in CLL Discovery Validation Clinical practice Variant diversity *Leroy at al, Cancer Research Review
Therapy-related acute myeloid leukemia with germline TP53 mutation (Li-Fraumeni syndrome) SH Chelsey Deel MD Teresa Scordino MD
 Therapy-related acute myeloid leukemia with germline TP53 mutation (Li-Fraumeni syndrome) SH2017-167 Chelsey Deel MD Teresa Scordino MD Clinical History HPI: 44 year old Caucasian female referred for evaluation
Therapy-related acute myeloid leukemia with germline TP53 mutation (Li-Fraumeni syndrome) SH2017-167 Chelsey Deel MD Teresa Scordino MD Clinical History HPI: 44 year old Caucasian female referred for evaluation
BRCA1 alternative splicing in breast tumorogenesis
 Available online at www.scholarsresearchlibrary.com Annals of Biological Research, 2013, 4 (4):139-143 (http://scholarsresearchlibrary.com/archive.html) ISSN 0976-1233 CODEN (USA): ABRNBW BRCA1 alternative
Available online at www.scholarsresearchlibrary.com Annals of Biological Research, 2013, 4 (4):139-143 (http://scholarsresearchlibrary.com/archive.html) ISSN 0976-1233 CODEN (USA): ABRNBW BRCA1 alternative
Molecular Characterization of the NF2 Gene in Korean Patients with Neurofibromatosis Type 2: A Report of Four Novel Mutations
 Korean J Lab Med 2010;30:190-4 DOI 10.3343/kjlm.2010.30.2.190 Original Article Diagnostic Genetics Molecular Characterization of the NF2 Gene in Korean Patients with Neurofibromatosis Type 2: A Report
Korean J Lab Med 2010;30:190-4 DOI 10.3343/kjlm.2010.30.2.190 Original Article Diagnostic Genetics Molecular Characterization of the NF2 Gene in Korean Patients with Neurofibromatosis Type 2: A Report
When to change therapy? Andreas Hochhaus Universitätsklinikum Jena, Germany
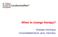 When to change therapy? Andreas Hochhaus Universitätsklinikum Jena, Germany Chromosome 22 Chromosome 9 e1 1b m-bcr M-bcr e1 e2 b1 b5 5 3 BCR ABL 5 3 1a a2 a3 μ -bcr e19 a11 e1a2 b2a2 b3a2 e19a2 p190 bcr-abl
When to change therapy? Andreas Hochhaus Universitätsklinikum Jena, Germany Chromosome 22 Chromosome 9 e1 1b m-bcr M-bcr e1 e2 b1 b5 5 3 BCR ABL 5 3 1a a2 a3 μ -bcr e19 a11 e1a2 b2a2 b3a2 e19a2 p190 bcr-abl
The molecular definition of chromosomal
 Haematologica 1998; 83:593-601 original paper Detection of bcr-abl transcript in chronic myelogenous leukemia patients by reverse-transcription-polymerase chain reaction and capillary electrophoresis GIOVANNI
Haematologica 1998; 83:593-601 original paper Detection of bcr-abl transcript in chronic myelogenous leukemia patients by reverse-transcription-polymerase chain reaction and capillary electrophoresis GIOVANNI
SALSA MLPA probemix P241-D2 MODY mix 1 Lot D2-0716, D As compared to version D1 (lot D1-0911), one reference probe has been replaced.
 mix P241-D2 MODY mix 1 Lot D2-0716, D2-0413. As compared to version D1 (lot D1-0911), one reference has been replaced. Maturity-Onset Diabetes of the Young (MODY) is a distinct form of non insulin-dependent
mix P241-D2 MODY mix 1 Lot D2-0716, D2-0413. As compared to version D1 (lot D1-0911), one reference has been replaced. Maturity-Onset Diabetes of the Young (MODY) is a distinct form of non insulin-dependent
Cytogendies, in situ Hybridization and Molecular Approaches in the Diagnosis of Cancer*
 ANNALS OF CLINICAL AND LABORATORY SCIENCE, Vol. 28, No. 6 Copyright 1998, Institute for Clinical Science, Inc. Cytogendies, in situ Hybridization and Molecular Approaches in the Diagnosis of Cancer* ARMAND
ANNALS OF CLINICAL AND LABORATORY SCIENCE, Vol. 28, No. 6 Copyright 1998, Institute for Clinical Science, Inc. Cytogendies, in situ Hybridization and Molecular Approaches in the Diagnosis of Cancer* ARMAND
SALSA MS-MLPA KIT ME011-A1 Mismatch Repair genes (MMR) Lot 0609, 0408, 0807, 0407
 SALSA MS-MLPA KIT ME011-A1 Mismatch Repair genes (MMR) Lot 0609, 0408, 0807, 0407 The Mismatch Repair (MMR) system is critical for the maintenance of genomic stability. MMR increases the fidelity of DNA
SALSA MS-MLPA KIT ME011-A1 Mismatch Repair genes (MMR) Lot 0609, 0408, 0807, 0407 The Mismatch Repair (MMR) system is critical for the maintenance of genomic stability. MMR increases the fidelity of DNA
Open Access Original Article
 Open Access Original Article Cytogenetic and Molecular Analyses of Philadelphia Chromosome Variants in CML (Chronic Myeloid Leukemia) Patients from Sindh using Karyotyping and RT-PCR Ikram Din Ujjan 1,
Open Access Original Article Cytogenetic and Molecular Analyses of Philadelphia Chromosome Variants in CML (Chronic Myeloid Leukemia) Patients from Sindh using Karyotyping and RT-PCR Ikram Din Ujjan 1,
Oxford BRC Haemato-Molecular Diagnostic Service
 Short User Guide Oxford BRC Haemato-Molecular Diagnostic Service The Oxford University Hospitals NHS trust s department of Haematology provides a comprehensive molecular diagnostic service for a range
Short User Guide Oxford BRC Haemato-Molecular Diagnostic Service The Oxford University Hospitals NHS trust s department of Haematology provides a comprehensive molecular diagnostic service for a range
MRC-Holland MLPA. Description version 30; 06 June 2017
 SALSA MLPA probemix P081-C1/P082-C1 NF1 P081 Lot C1-0517, C1-0114. As compared to the previous B2 version (lot B2-0813, B2-0912), 11 target probes are replaced or added, and 10 new reference probes are
SALSA MLPA probemix P081-C1/P082-C1 NF1 P081 Lot C1-0517, C1-0114. As compared to the previous B2 version (lot B2-0813, B2-0912), 11 target probes are replaced or added, and 10 new reference probes are
Treatment and Survival in Patients with Chronic Myeloid Leukemia in a Chronic Phase in the West of Iran
 DOI:http://dx.doi.org/10.7314/APJCP.2015.16.17.7555 RESEARCH ARTICLE Treatment and Survival in Patients with Chronic Myeloid Leukemia in a Chronic Phase in the West of Iran Mehrdad Payandeh 1&, Masoud
DOI:http://dx.doi.org/10.7314/APJCP.2015.16.17.7555 RESEARCH ARTICLE Treatment and Survival in Patients with Chronic Myeloid Leukemia in a Chronic Phase in the West of Iran Mehrdad Payandeh 1&, Masoud
DETERMINANT VALUE OF THE CYTOGENETIC AND MOLECULAR IMATINIB THERAPEUTIC RESPONSE IN CHRONIC MYELOID LEUKEMIA
 Analele Ştiinţifice ale Universităţii Alexandru Ioan Cuza, Secţiunea Genetică şi Biologie Moleculară, TOM XIV, 2013 DETERMINANT VALUE OF THE CYTOGENETIC AND MOLECULAR IMATINIB THERAPEUTIC RESPONSE IN CHRONIC
Analele Ştiinţifice ale Universităţii Alexandru Ioan Cuza, Secţiunea Genetică şi Biologie Moleculară, TOM XIV, 2013 DETERMINANT VALUE OF THE CYTOGENETIC AND MOLECULAR IMATINIB THERAPEUTIC RESPONSE IN CHRONIC
35 Current Trends in the
 35 Current Trends in the Management of Chronic Myelogenous Leukemia Abstract: CML is a hematopoietic stem cell disease which is characterized by the presence of Philadelphia chromosome (Ph-chromosome)
35 Current Trends in the Management of Chronic Myelogenous Leukemia Abstract: CML is a hematopoietic stem cell disease which is characterized by the presence of Philadelphia chromosome (Ph-chromosome)
MRC-Holland MLPA. Description version 29; 31 July 2015
 SALSA MLPA probemix P081-C1/P082-C1 NF1 P081 Lot C1-0114. As compared to the previous B2 version (lot 0813 and 0912), 11 target probes are replaced or added, and 10 new reference probes are included. P082
SALSA MLPA probemix P081-C1/P082-C1 NF1 P081 Lot C1-0114. As compared to the previous B2 version (lot 0813 and 0912), 11 target probes are replaced or added, and 10 new reference probes are included. P082
BWA alignment to reference transcriptome and genome. Convert transcriptome mappings back to genome space
 Whole genome sequencing Whole exome sequencing BWA alignment to reference transcriptome and genome Convert transcriptome mappings back to genome space genomes Filter on MQ, distance, Cigar string Annotate
Whole genome sequencing Whole exome sequencing BWA alignment to reference transcriptome and genome Convert transcriptome mappings back to genome space genomes Filter on MQ, distance, Cigar string Annotate
Submitted: May 13, 2004; Revised: June 17, 2004; Accepted: June 17, 2004; Published: July 1, 2004.
 Biol. Proced. Online 2004;6(1): 144-148. doi:10.1251/bpo83 Two different point mutations in ABL gene ATP-binding domain conferring Primary Imatinib resistance in a Chronic Myeloid Leukemia (CML) patient:
Biol. Proced. Online 2004;6(1): 144-148. doi:10.1251/bpo83 Two different point mutations in ABL gene ATP-binding domain conferring Primary Imatinib resistance in a Chronic Myeloid Leukemia (CML) patient:
Talpaz M. et al. Dasatinib in Imatinib-resistant Philadelphia chromosomepositive leukemias. N Engl J Med (2006) 354;24:
 References Sprycel Talpaz M. et al. Dasatinib in Imatinib-resistant Philadelphia chromosomepositive leukemias. N Engl J Med (2006) 354;24:2531-2541. National Comprehensive Cancer Network. Clinical Practice
References Sprycel Talpaz M. et al. Dasatinib in Imatinib-resistant Philadelphia chromosomepositive leukemias. N Engl J Med (2006) 354;24:2531-2541. National Comprehensive Cancer Network. Clinical Practice
Chronic Myelogenous Leukemia (Hematology) By DEISSEROTH READ ONLINE
 Chronic Myelogenous Leukemia (Hematology) By DEISSEROTH READ ONLINE If searched for the ebook by DEISSEROTH Chronic Myelogenous Leukemia (Hematology) in pdf format, in that case you come on to correct
Chronic Myelogenous Leukemia (Hematology) By DEISSEROTH READ ONLINE If searched for the ebook by DEISSEROTH Chronic Myelogenous Leukemia (Hematology) in pdf format, in that case you come on to correct
Dr Catherine Woolnough, Hospital Scientist, Chemical Pathology, Royal Prince Alfred Hospital. NSW Health Pathology University of Sydney
 Dr Catherine Woolnough, Hospital Scientist, Chemical Pathology, Royal Prince Alfred Hospital NSW Health Pathology University of Sydney Thyroid Cancer TC incidence rates in NSW Several subtypes - Papillary
Dr Catherine Woolnough, Hospital Scientist, Chemical Pathology, Royal Prince Alfred Hospital NSW Health Pathology University of Sydney Thyroid Cancer TC incidence rates in NSW Several subtypes - Papillary
Cancer and Tyrosine Kinase Inhibition
 Cancer and Tyrosine Kinase Inhibition 1 Motivation (1) In first world countries cancer is the second most common cause of death after cardiovascular diseases. For most tumors, treatment is limited to surgery,
Cancer and Tyrosine Kinase Inhibition 1 Motivation (1) In first world countries cancer is the second most common cause of death after cardiovascular diseases. For most tumors, treatment is limited to surgery,
Chromothripsis: A New Mechanism For Tumorigenesis? i Fellow s Conference Cheryl Carlson 6/10/2011
 Chromothripsis: A New Mechanism For Tumorigenesis? i Fellow s Conference Cheryl Carlson 6/10/2011 Massive Genomic Rearrangement Acquired in a Single Catastrophic Event during Cancer Development Cell 144,
Chromothripsis: A New Mechanism For Tumorigenesis? i Fellow s Conference Cheryl Carlson 6/10/2011 Massive Genomic Rearrangement Acquired in a Single Catastrophic Event during Cancer Development Cell 144,
RNA SEQUENCING AND DATA ANALYSIS
 RNA SEQUENCING AND DATA ANALYSIS Length of mrna transcripts in the human genome 5,000 5,000 4,000 3,000 2,000 4,000 1,000 0 0 200 400 600 800 3,000 2,000 1,000 0 0 2,000 4,000 6,000 8,000 10,000 Length
RNA SEQUENCING AND DATA ANALYSIS Length of mrna transcripts in the human genome 5,000 5,000 4,000 3,000 2,000 4,000 1,000 0 0 200 400 600 800 3,000 2,000 1,000 0 0 2,000 4,000 6,000 8,000 10,000 Length
Therapy-related MDS/AML with KMT2A (MLL) Rearrangement Following Therapy for APL Case 0328
 Therapy-related MDS/AML with KMT2A (MLL) Rearrangement Following Therapy for APL Case 0328 Kenneth N. Holder, Leslie J. Greebon, Gopalrao Velagaleti, Hongxin Fan, Russell A. Higgins Initial Case: Clinical
Therapy-related MDS/AML with KMT2A (MLL) Rearrangement Following Therapy for APL Case 0328 Kenneth N. Holder, Leslie J. Greebon, Gopalrao Velagaleti, Hongxin Fan, Russell A. Higgins Initial Case: Clinical
