Assignment 5: Integrative epigenomics analysis
|
|
|
- Samantha Bryan
- 5 years ago
- Views:
Transcription
1 Assignment 5: Integrative epigenomics analysis Due date: Friday, 2/24 10am. Note: no late assignments will be accepted. Introduction CpG islands (CGIs) are important regulatory regions in the genome. What are their epigenetic profiles? Let s find out! We obtained several epigenomic datasets from the Roadmap Epigenome Project from a human brain germinal matrix sample: BGM_WGBS.bed: methylation data from whole genome bisulfite sequencing (WGBS) BGM_MeDIP.bed: methylation data from methylated DNA immunoprecipitation sequencing (MeDIP-seq) BGM_MRE.bed: methylation data from methyl-sensitive restriction enzyme sequencing (MRE-seq) You are also provided with some annotation files: CGI.bed: locations of CGIs. CpG.bed: locations of CpGs. refgene.bed: locations of RefSeq genes. Please see the appendix for a description of the files. All files are located in /home/assignments/assignment5/. All files are based on the hg19 reference genome assembly. To make things easier, we have decided to focus on chromosome 21. (The full datasets can be found in the Roadmap Epigenome Project flagship paper.) From these data we want to obtain an epigenomic description of CGIs. Now you are ready to explore epigenomics data! A few notes before you get started In this assignment, you will use the powerful command-line tool bedtools to help with some of the analysis. bedtools (v2.25.0) is already installed on the server. To print the bedtools help menu, type $ bedtools --help All plots must have informative axis labels, titles, etc. In this assignment, you will be writing scripts that produce output files. Unlike previous assignments, you re going to run the same scripts on different inputs. In order to not overwrite the output file, your script cannot hardcode the output filename. Instead, you will use Python to dynamically name the output filenames based on the input file names. Here s some example code: import os 1
2 basename = os.path.splitext(os.path.basename(<input_filename>))[0] plotname = basename + _methylation_distribution.png Part 1 Analyzing DNA methylation data Part 1.0 First, we are interested in answering some typical epigenomics questions. Write a script called analyze_wgbs_methylation.py that takes a WGBS bed file and 1. Calculates the methylation level for each CpG using the formula: # C base calls CpG methylation level = # C base calls + # T base calls 2. Writes a bed-like file of CpG methylation. The columns will be 1) chromosome, 2) start, 3) stop, and 4) methylation level. Do not output CpGs that have 0X coverage. Save the file as <WGBS bed basename>_cpg_methylation.bed. 3. Plots the distribution, e.g., with a histogram, of CpG methylation levels as <WGBS bed basename>_methylation_distribution.png. 4. Plots the distribution of read coverage for all CpGs for coverages between 0X and 100X as <WGBS bed basename>_cpg_coverage_distribution.png. 5. Calculates and print the fraction of CpGs that have 0X coverage. The usage of the script will be: $ python3 analyze_wgbs_methylation.py <WGBS bed> Run analyze_wgbs_methylation.py on the given WGBS data. Paste the command in your README. Copy your output files to your submissions directory. Question 1 What does DNA methylation look like across chromosome 21? Question 2 What does the CpG coverage look like across chromosome 21? Question 2.1 What fraction of the CpGs have 0X coverage? Part 1.1 Using bedtools, create a bed file with the average CpG methylation level in each CGI. This file should be in bed format (i.e., have the columns 1) chromosome, 2) start, 3) stop, 4) CGI name, and 5) average CpG methylation in CGI). Name your final file with the average CpG methylation level in each CGI WGBS_CGI_methylation.bed. Paste the commands to generate this file in your README. Hint: use the bedtools intersect and bedtools groupby subcommands. Part 1.2 2
3 Write a script called analyze_cgi_methylation.py that takes the average CGI methylation bed file and plots the distribution of average CGI methylation levels. Save the plot as <average CGI methylation bed basename>_distribution.png. The usage of the script will be: $ python3 analyze_cgi_methylation.py <average CGI methylation bed> Run analyze_cgi_methylation.py on WGBS_CGI_methylation.bed. Save the output figure as WGBS_CGI_methylation_distribution.png. Paste the command in your README. Question 3 What does DNA methylation look like for CpGs in CGIs? How does it compare to all the CpGs on chromosome 21? Part Lastly, we want to explore the CpG methylation profiles in promoter-cgis versus nonpromoter-cgis. Write a script called generate_promoters.py that takes a bed file of gene coordinates and creates a bed file of their promoters. The columns will be 1) chromosome, 2) start, 3) stop, 4) gene name, and 5) strand. The usage of the script will be: $ python3 generate_promoters.py <bed of gene coordinates> Run generate_promoters.py on refgene.bed. Save the output file as refgene_promoters.bed. In your README, justify how you defined promoter and paste your command for creating this file. Generate a bed file of promoter-cgis called promoter_cgi.bed and a bed file of non-promoter- CGIs called non_promoter_cgi.bed. Promoter-CGIs are defined as CGIs that overlap with a promoter. In your README, justify your overlapping criteria and paste your commands for creating these files. Hint: use bedtools intersect. Calculate the average CpG methylation level for each promoter-cgi and non-promoter-cgi. Save these files as average_promoter_cgi_methylation.bed and average_non_promoter_cgi_methylation.bed, respectively. Paste your commands for creating these files in your README. Hint: see your commands for part 1.1. Run analyze_cgi_methylation.py on average_promoter_cgi_methylation.bed and average_non_promoter_cgi_methylation.bed. Save the output figures as average_promoter_cgi_methylation.png and average_non_promoter_cgi_methylation.png, respectively. Paste your commands for creating these files in your README. 3
4 Question 4 How do the DNA methylation profiles of promoter-cgis and non-promoter-cgis differ? Part Use this modified nuc count script (/home/assignments/assignment5/nuc_count_multisequence_fasta.py) to calculate the frequency of CpGs in promoter-cgi and non-promoter-cgis fastas. In your README, paste the CpG frequency for both the promoter-cgis and non-promoter-cgis and your commands to calculate the frequencies. Hint: use bedtools getfasta to convert a bed file to a fasta file. Use this chromosome 21 fasta file: /home/assignments/assignment5/hg19_chr21.fa. Question 5 What is a possible biological explanation for the difference in CpG frequencies? Interpret your results from parts and 1.3.1: what are the simple rules for describing regulation by DNA methylation in promoters? Part 2 Comparing CGI MeDIP-seq, MRE-seq, and WGBS methylation level. The script bed_reads_rkpm.pl has already been written for you. This script takes a bed file of features, e.g., CGIs, and a bed file of reads and calculates the RKPM methylation score. The RPKM methylation score is defined as: RPKM methylation score = # of reads that overlap feature size of feature in kb # of mapped reads in millions The output is a bed-like file: 1 Start coordinate of feature (0-based, inclusive) End coordinate of feature (0-based, exclusive) RPK (reads per kb) methylation score RPKM methylation score Usage: $ perl bed_reads_rpkm.pl <feature bed> <reads bed> > <RPKM methylation levels> Note: this is a script written in the Perl programming language, so it is invoked by typing perl <perl script>. 4
5 Use bed_reads_rkpm.pl to calculate the CGI RPKM methylation scores using the MeDIP-seq and MRE-seq data. Save the output files as MeDIP_CGI_RPKM.bed and MRE_CGI_RPKM.bed, respectively. Paste the commands in your README. Write a script called compare_methylome_technologies.py that 1. Creates the following scatter plots a. MeDIP-seq RPKM vs. MRE-seq RPKM. Save the plot as <MeDIP-seq RPKM bed basename>_vs_<mre-seq RPKM bed basename>.png. b. MeDIP-seq RPKM vs. WGBS average DNA methylation level. Save the plot as <MeDIP-seq RPKM bed basename>_vs_<wgbs methylation level bed basename>.png. c. MRE-seq RPKM vs. WGBS average DNA methylation level. Save the plot as <MRE-seq RPKM bed basename>_vs_<wgbs methylation level bed basename>.png. 2. Calculate the correlation for each comparison. Print the correlation to stdout. Hint: use scipy.stats. The usage of the script will be: $ python3 compare_methylome_technologies.py <MeDIP-seq RPKM bed> <MRE-seq RPKM bed> <WGBS methylation level bed> Run compare_methylome_technologies.py on MeDIP_CGI_RPKM.bed, MRE_CGI_RPKM.bed, and WGBS_CGI_methylation.bed. Save the output figures as MeDIP_CGI_RPKM_vs_MRE_CGI_RPKM.png, MeDIP_CGI_RPKM_vs_WGBS_CGI_methylation.png, and MRE_CGI_RPKM_vs_WGBS_CGI_methylation.png. Paste the command to generate the figures, the correlations, and justify which correlation statistic you used in your README. Question 6 How do MeDIP-seq and methylation correlate? How do MRE-seq and methylation correlate? How do MeDIP-seq and MRE-seq correlate? There is at least one outlier. In your README, list their locations and explain the potential cause(s) for the outlier(s). (Hint: look at the CGIs in a genome browser.) Explain why (or why not) the outlier(s) should be removed. If you removed them, recreate the scatter plots and recalculate correlations. Paste the correlations in your README. Save the output figures as MeDIP_CGI_RPKM_vs_MRE_CGI_RPKM_outliers_removed.png, MeDIP_CGI_RPKM_vs_WGBS_CGI_methylation_outliers_removed.png, and MRE_CGI_RPKM_vs_WGBS_CGI_methylation_outliers_removed.png What to turn in All scripts that you wrote analyze_wgbs_methylation.py analyze_cgi_methylation.py generate_promoters.py compare_methylome_technologies.py All output files 5
6 BGM_WGBS_CpG_methylation.bed BGM_WGBS_methylation_distribution.png BGM_WGBS_CpG_coverage_distribution.png WGBS_CGI_methylation.bed WGBS_CGI_methylation_distribution.png refgene_promoters.bed promoter_cgi.bed non_promoter_cgi.bed average_promoter_cgi_methylation.bed average_non_promoter_cgi_methylation.bed average_promoter_cgi_methylation.png average_non_promoter_cgi_methylation.png MeDIP_CGI_RPKM.bed MRE_CGI_RPKM.bed MeDIP_CGI_RPKM_vs_MRE_CGI_RPKM.png (and possibly MeDIP_CGI_RPKM_vs_MRE_CGI_RPKM_outliers_removed.png) MeDIP_CGI_RPKM_vs_WGBS_CGI_methylation.png (and possibly MeDIP_CGI_RPKM_vs_WGBS_CGI_methylation_outliers_removed.png) MRE_CGI_RPKM_vs_WGBS_CGI_methylation.png (and possibly MRE_CGI_RPKM_vs_WGBS_CGI_methylation_outliers_removed.png) Turn in a README.txt file with your commands and answers the above questions. These files should be in your assignment5/submission folder. Extra credit Going the distance (again!) Examine H3K4me3, the histone mark for promoters. The H3K4me4 ChIP-seq dataset is located here: /home/assignments/assignment5/bgm_h3k4me3.bed. Use bed_reads_rkpm.pl to calculate the ChIP-seq RPKM score for promoter-cgis and nonpromoter-cgis using the H3K4me4 ChIP-seq data. Save the output files as H3K4me3_RPKM_promoter_CGI.bed and H3K4me3_RPKM_non_promoter_CGI.bed, respectively. Paste the commands in your README. Write a script called analyze_h3k4me3_scores.py that creates one figure with a boxplot of the H3K4me3 RPKM score distribution for promoter-cgis and a boxplot of the H3K4me3 RPKM score distribution for non-promoter-cgis. Save the plot as <basename of first bed of RPKM scores>_and_<basename of second bed of RPKM scores>.png. The usage of the script will be: $ python3 analyze_h3k4me3_scores.py <first bed of RPKM scores> <second bed of RPKM scores> Run analyze_h3k4me3_scores.py on H3K4me3_RPKM_promoter_CGI.bed and H3K4me3_RPKM_non_promoter_CGI.bed. Save the output figure as 6
7 H3K4me3_RPKM_promoter_CGI_and_H3K4me3_RPKM_non_promoter_CGI.png. Paste the commands in your README. Question EC.1 How does the H3K4me3 signal differ in promoter-cgis and non-promoter-cgis? Question EC.2 What are some better alternatives to model MeDIP-seq data and MRE-seq data instead of using RPKM? Explain. Question EC.3 What would be a better way to compare H3K4me3 values instead of using boxplots? Explain. What to to turn in for extra credit All scripts that you wrote analyze_h3k4me3_scores.py All output files H3K4me3_RPKM_promoter_CGI.bed H3K4me3_RPKM_non_promoter_CGI.bed H3K4me3_RPKM_promoter_CGI_and_H3K4me3_RPKM_non_promoter_CGI.png Add your commands and answers to the extra credit questions in the README.txt. 7
8 Appendix: Description of input files This appendix describes the data files in /home/assignments/assignment5/. All coordinates are based on the hg19 reference genome. The data has been reduced to chromosome 21. Fetal brain sample files: BGM_WGBS.bed: contains methylation data from WGBS in bed format. Each row contains the coordinate of a CpG with at least 1X coverage, the number of times the C was sequenced as C and the number of times the C was sequenced as T. (Note: Reads that mapped to either of the two strands in the CpG were combined.) From these numbers, one can estimate the level of methylation for each CpG, as well as its coverage. The file is in a bed-variant format: 1 Start coordinate of CpG (0-based, inclusive) End coordinate of CpG (0-based, exclusive) # of C basecalls for CpG 12 4 # of T basecalls for CpG 9 BGM_MeDIP.bed: contains methylation data from MeDIP-seq in bed format. Each row contains the coordinate of a mapped MeDIP-seq read, the read, the mapping quality, and the strand the read aligned to: 1 Start coordinate of MeDIP-seq read (0-based, inclusive) 2 End coordinate of MeDIP-seq read (0-based, exclusive) Sequence of MeDIP-seq read CAGAACTTGAGTGTGTAAGCTCCCAGAAAGAAGAGAAACACATTG AAGTGATTTCAACCTTCTCCACAGCCTTTC 4 Mapping quality 23 5 Strand that read mapped to + 8
9 BGM_MRE.bed: contains methylation data from MRE-seq in bed format. Each row contains the coordinate of a mapped MRE-seq read, the read, the mapping quality, and the strand the read aligned to. 1 Start coordinate of MRE-seq read (0-based, inclusive) 2 End coordinate of MRE-seq read (0-based, exclusive) Sequence of ChIP-seq read CATCTCGGCTCACTGCGAGCTCAGCCTCCTGGCTTCGTGCCATTCTCC TGCCTCAGCCTCTCTAGTAGCGGG 4 Mapping quality 37 5 Strand that read mapped to + BGM_H3K4me3.bed (for extra credit): contains H3K4me3 ChIP-seq data in bed format. Each row contains the coordinate of a mapped ChIP-seq read, the read, the mapping quality, and the strand the read aligned to: 1 Start coordinate of ChIP-seq read (0-based, inclusive) 2 End coordinate of ChIP-seq read (0-based, exclusive) Sequence of ChIP-seq read GAGGTAGATCATCTTGGTCCAATCAGACTGAAATGCCTTGAGGCTAGA TTTCAGTCTTTGTGGCAGGTGGGGGAA 4 Mapping quality 25 5 Strand that read mapped to + Annotation files: CGI.bed: contains locations of CGIs in bed format. Each row contains the coordinate and name of a CGI: 9
10 1 Start coordinate of CGI (0-based, inclusive) End coordinate of CGI (0-based, exclusive) CGI name (derived from the # of CpGs in CGI) (not unique) CpG: Number of CpGs in the CGI 285 CpG.bed: contains locations of CpGs in bed format. Each row contains the coordinate of a CpG: 1 Start coordinate of CpG (0-based, inclusive) End coordinate of CpG (0-based, exclusive) refgene.bed: contains locations of refseq genes in bed format. Each row contains the coordinate of a gene, the gene name, the number of exons in the gene, and the strand of the gene: 1 Start coordinate, i.e., leftmost coordinate, of gene (0-based, inclusive) 2 End coordinate, i.e., rightmost coordinate, of gene read (0-based, exclusive) Gene name MIR364 4 Number of exons in gene 1 5 Strand + 10
Use Case 9: Coordinated Changes of Epigenomic Marks Across Tissue Types. Epigenome Informatics Workshop Bioinformatics Research Laboratory
 Use Case 9: Coordinated Changes of Epigenomic Marks Across Tissue Types Epigenome Informatics Workshop Bioinformatics Research Laboratory 1 Introduction Active or inactive states of transcription factor
Use Case 9: Coordinated Changes of Epigenomic Marks Across Tissue Types Epigenome Informatics Workshop Bioinformatics Research Laboratory 1 Introduction Active or inactive states of transcription factor
Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq
 Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne ChIP-seq against histone variants: Biological
Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne ChIP-seq against histone variants: Biological
Measuring DNA Methylation with the MinION. Winston Timp Department of Biomedical Engineering Johns Hopkins University 12/1/16
 Measuring DNA Methylation with the MinION Winston Timp Department of Biomedical Engineering Johns Hopkins University 12/1/16 Epigenetics: Modern Modern Definition of epigenetics involves heritable changes
Measuring DNA Methylation with the MinION Winston Timp Department of Biomedical Engineering Johns Hopkins University 12/1/16 Epigenetics: Modern Modern Definition of epigenetics involves heritable changes
ChromHMM Tutorial. Jason Ernst Assistant Professor University of California, Los Angeles
 ChromHMM Tutorial Jason Ernst Assistant Professor University of California, Los Angeles Talk Outline Chromatin states analysis and ChromHMM Accessing chromatin state annotations for ENCODE2 and Roadmap
ChromHMM Tutorial Jason Ernst Assistant Professor University of California, Los Angeles Talk Outline Chromatin states analysis and ChromHMM Accessing chromatin state annotations for ENCODE2 and Roadmap
Accessing and Using ENCODE Data Dr. Peggy J. Farnham
 1 William M Keck Professor of Biochemistry Keck School of Medicine University of Southern California How many human genes are encoded in our 3x10 9 bp? C. elegans (worm) 959 cells and 1x10 8 bp 20,000
1 William M Keck Professor of Biochemistry Keck School of Medicine University of Southern California How many human genes are encoded in our 3x10 9 bp? C. elegans (worm) 959 cells and 1x10 8 bp 20,000
Nature Methods: doi: /nmeth.3115
 Supplementary Figure 1 Analysis of DNA methylation in a cancer cohort based on Infinium 450K data. RnBeads was used to rediscover a clinically distinct subgroup of glioblastoma patients characterized by
Supplementary Figure 1 Analysis of DNA methylation in a cancer cohort based on Infinium 450K data. RnBeads was used to rediscover a clinically distinct subgroup of glioblastoma patients characterized by
Hands-On Ten The BRCA1 Gene and Protein
 Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
Hands-On Ten The BRCA1 Gene and Protein Objective: To review transcription, translation, reading frames, mutations, and reading files from GenBank, and to review some of the bioinformatics tools, such
Supplemental Figure S1. Tertiles of FKBP5 promoter methylation and internal regulatory region
 Supplemental Figure S1. Tertiles of FKBP5 promoter methylation and internal regulatory region methylation in relation to PSS and fetal coupling. A, PSS values for participants whose placentas showed low,
Supplemental Figure S1. Tertiles of FKBP5 promoter methylation and internal regulatory region methylation in relation to PSS and fetal coupling. A, PSS values for participants whose placentas showed low,
Data mining with Ensembl Biomart. Stéphanie Le Gras
 Data mining with Ensembl Biomart Stéphanie Le Gras (slegras@igbmc.fr) Guidelines Genome data Genome browsers Getting access to genomic data: Ensembl/BioMart 2 Genome Sequencing Example: Human genome 2000:
Data mining with Ensembl Biomart Stéphanie Le Gras (slegras@igbmc.fr) Guidelines Genome data Genome browsers Getting access to genomic data: Ensembl/BioMart 2 Genome Sequencing Example: Human genome 2000:
Results. Abstract. Introduc4on. Conclusions. Methods. Funding
 . expression that plays a role in many cellular processes affecting a variety of traits. In this study DNA methylation was assessed in neuronal tissue from three pigs (frontal lobe) and one great tit (whole
. expression that plays a role in many cellular processes affecting a variety of traits. In this study DNA methylation was assessed in neuronal tissue from three pigs (frontal lobe) and one great tit (whole
Comparative DNA methylome analysis of endometrial carcinoma reveals complex and distinct deregulation of cancer promoters and enhancers
 Zhang et al. BMC Genomics 2014, 15:868 RESEARCH ARTICLE Open Access Comparative DNA methylome analysis of endometrial carcinoma reveals complex and distinct deregulation of cancer promoters and enhancers
Zhang et al. BMC Genomics 2014, 15:868 RESEARCH ARTICLE Open Access Comparative DNA methylome analysis of endometrial carcinoma reveals complex and distinct deregulation of cancer promoters and enhancers
Measuring DNA Methylation with the MinION
 Measuring DNA Methylation with the MinION Winston Timp Department of Biomedical Engineering Johns Hopkins University Epigenetics: Modern Modern Definition of epigenetics involves heritable changes other
Measuring DNA Methylation with the MinION Winston Timp Department of Biomedical Engineering Johns Hopkins University Epigenetics: Modern Modern Definition of epigenetics involves heritable changes other
High Throughput Sequence (HTS) data analysis. Lei Zhou
 High Throughput Sequence (HTS) data analysis Lei Zhou (leizhou@ufl.edu) High Throughput Sequence (HTS) data analysis 1. Representation of HTS data. 2. Visualization of HTS data. 3. Discovering genomic
High Throughput Sequence (HTS) data analysis Lei Zhou (leizhou@ufl.edu) High Throughput Sequence (HTS) data analysis 1. Representation of HTS data. 2. Visualization of HTS data. 3. Discovering genomic
Variant Classification. Author: Mike Thiesen, Golden Helix, Inc.
 Variant Classification Author: Mike Thiesen, Golden Helix, Inc. Overview Sequencing pipelines are able to identify rare variants not found in catalogs such as dbsnp. As a result, variants in these datasets
Variant Classification Author: Mike Thiesen, Golden Helix, Inc. Overview Sequencing pipelines are able to identify rare variants not found in catalogs such as dbsnp. As a result, variants in these datasets
Tutorial: RNA-Seq Analysis Part II: Non-Specific Matches and Expression Measures
 : RNA-Seq Analysis Part II: Non-Specific Matches and Expression Measures March 15, 2013 CLC bio Finlandsgade 10-12 8200 Aarhus N Denmark Telephone: +45 70 22 55 09 Fax: +45 70 22 55 19 www.clcbio.com support@clcbio.com
: RNA-Seq Analysis Part II: Non-Specific Matches and Expression Measures March 15, 2013 CLC bio Finlandsgade 10-12 8200 Aarhus N Denmark Telephone: +45 70 22 55 09 Fax: +45 70 22 55 19 www.clcbio.com support@clcbio.com
SUPPLEMENTAL INFORMATION
 SUPPLEMENTAL INFORMATION GO term analysis of differentially methylated SUMIs. GO term analysis of the 458 SUMIs with the largest differential methylation between human and chimp shows that they are more
SUPPLEMENTAL INFORMATION GO term analysis of differentially methylated SUMIs. GO term analysis of the 458 SUMIs with the largest differential methylation between human and chimp shows that they are more
CNV PCA Search Tutorial
 CNV PCA Search Tutorial Release 8.1 Golden Helix, Inc. March 18, 2014 Contents 1. Data Preparation 2 A. Join Log Ratio Data with Phenotype Information.............................. 2 B. Activate only
CNV PCA Search Tutorial Release 8.1 Golden Helix, Inc. March 18, 2014 Contents 1. Data Preparation 2 A. Join Log Ratio Data with Phenotype Information.............................. 2 B. Activate only
Chip Seq Peak Calling in Galaxy
 Chip Seq Peak Calling in Galaxy Chris Seward PowerPoint by Pei-Chen Peng Chip-Seq Peak Calling in Galaxy Chris Seward 2018 1 Introduction This goals of the lab are as follows: 1. Gain experience using
Chip Seq Peak Calling in Galaxy Chris Seward PowerPoint by Pei-Chen Peng Chip-Seq Peak Calling in Galaxy Chris Seward 2018 1 Introduction This goals of the lab are as follows: 1. Gain experience using
DNA Sequence Bioinformatics Analysis with the Galaxy Platform
 DNA Sequence Bioinformatics Analysis with the Galaxy Platform University of São Paulo, Brazil 28 July - 1 August 2014 Dave Clements Johns Hopkins University Robson Francisco de Souza University of São
DNA Sequence Bioinformatics Analysis with the Galaxy Platform University of São Paulo, Brazil 28 July - 1 August 2014 Dave Clements Johns Hopkins University Robson Francisco de Souza University of São
Epigenetics. Jenny van Dongen Vrije Universiteit (VU) Amsterdam Boulder, Friday march 10, 2017
 Epigenetics Jenny van Dongen Vrije Universiteit (VU) Amsterdam j.van.dongen@vu.nl Boulder, Friday march 10, 2017 Epigenetics Epigenetics= The study of molecular mechanisms that influence the activity of
Epigenetics Jenny van Dongen Vrije Universiteit (VU) Amsterdam j.van.dongen@vu.nl Boulder, Friday march 10, 2017 Epigenetics Epigenetics= The study of molecular mechanisms that influence the activity of
A fully Bayesian approach for the analysis of Whole-Genome Bisulfite Sequencing Data
 A fully Bayesian approach for the analysis of Whole-Genome Bisulfite Sequencing Data Leonardo Bottolo 1,2,3 1 Department of Medical Genetics, University of Cambridge, UK 2 The Alan Turing Institute, London,
A fully Bayesian approach for the analysis of Whole-Genome Bisulfite Sequencing Data Leonardo Bottolo 1,2,3 1 Department of Medical Genetics, University of Cambridge, UK 2 The Alan Turing Institute, London,
BIMM 143. RNA sequencing overview. Genome Informatics II. Barry Grant. Lecture In vivo. In vitro.
 RNA sequencing overview BIMM 143 Genome Informatics II Lecture 14 Barry Grant http://thegrantlab.org/bimm143 In vivo In vitro In silico ( control) Goal: RNA quantification, transcript discovery, variant
RNA sequencing overview BIMM 143 Genome Informatics II Lecture 14 Barry Grant http://thegrantlab.org/bimm143 In vivo In vitro In silico ( control) Goal: RNA quantification, transcript discovery, variant
EPIGENOMICS PROFILING SERVICES
 EPIGENOMICS PROFILING SERVICES Chromatin analysis DNA methylation analysis RNA-seq analysis Diagenode helps you uncover the mysteries of epigenetics PAGE 3 Integrative epigenomics analysis DNA methylation
EPIGENOMICS PROFILING SERVICES Chromatin analysis DNA methylation analysis RNA-seq analysis Diagenode helps you uncover the mysteries of epigenetics PAGE 3 Integrative epigenomics analysis DNA methylation
Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types.
 Supplementary Figure 1 Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types. (a) Pearson correlation heatmap among open chromatin profiles of different
Supplementary Figure 1 Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types. (a) Pearson correlation heatmap among open chromatin profiles of different
Supplementary Fig.S1. MeDIP RPKM distribution of 5kb windows. Relative expression of DNMT. Percentage of genome. (fold change)
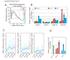 Supplementary Fig.S1 Percentage of genome 10% 8% 6% 4% 2% 0 MeDIP RPKM distribution of 5kb windows 0 0. 0.5 0.75 >1 MeDIP RPKM value EC Relative expression of DNMT (fold change) 12 6 4 3 2 1 EC DNMT1 DNMT2
Supplementary Fig.S1 Percentage of genome 10% 8% 6% 4% 2% 0 MeDIP RPKM distribution of 5kb windows 0 0. 0.5 0.75 >1 MeDIP RPKM value EC Relative expression of DNMT (fold change) 12 6 4 3 2 1 EC DNMT1 DNMT2
General Instructions:
 CSCE 110: Programming I Spring 2019 Lab 4 General Instructions: Lab is due online by 11:59 pm of the due date. The assignment must be typed, not handwritten or scanned. Label your Python programs q.py,
CSCE 110: Programming I Spring 2019 Lab 4 General Instructions: Lab is due online by 11:59 pm of the due date. The assignment must be typed, not handwritten or scanned. Label your Python programs q.py,
MODULE 4: SPLICING. Removal of introns from messenger RNA by splicing
 Last update: 05/10/2017 MODULE 4: SPLICING Lesson Plan: Title MEG LAAKSO Removal of introns from messenger RNA by splicing Objectives Identify splice donor and acceptor sites that are best supported by
Last update: 05/10/2017 MODULE 4: SPLICING Lesson Plan: Title MEG LAAKSO Removal of introns from messenger RNA by splicing Objectives Identify splice donor and acceptor sites that are best supported by
Large conserved domains of low DNA methylation maintained by Dnmt3a
 Supplementary information Large conserved domains of low DNA methylation maintained by Dnmt3a Mira Jeong# 1, Deqiang Sun # 2, Min Luo# 1, Yun Huang 3, Grant A. Challen %1, Benjamin Rodriguez 2, Xiaotian
Supplementary information Large conserved domains of low DNA methylation maintained by Dnmt3a Mira Jeong# 1, Deqiang Sun # 2, Min Luo# 1, Yun Huang 3, Grant A. Challen %1, Benjamin Rodriguez 2, Xiaotian
Vega: Variational Segmentation for Copy Number Detection
 Vega: Variational Segmentation for Copy Number Detection Sandro Morganella Luigi Cerulo Giuseppe Viglietto Michele Ceccarelli Contents 1 Overview 1 2 Installation 1 3 Vega.RData Description 2 4 Run Vega
Vega: Variational Segmentation for Copy Number Detection Sandro Morganella Luigi Cerulo Giuseppe Viglietto Michele Ceccarelli Contents 1 Overview 1 2 Installation 1 3 Vega.RData Description 2 4 Run Vega
A Quick-Start Guide for rseqdiff
 A Quick-Start Guide for rseqdiff Yang Shi (email: shyboy@umich.edu) and Hui Jiang (email: jianghui@umich.edu) 09/05/2013 Introduction rseqdiff is an R package that can detect differential gene and isoform
A Quick-Start Guide for rseqdiff Yang Shi (email: shyboy@umich.edu) and Hui Jiang (email: jianghui@umich.edu) 09/05/2013 Introduction rseqdiff is an R package that can detect differential gene and isoform
ONLINE. Online supplementary information S1 (Box) Method. Supplementary data. Online links
 ONLINE Online supplementary information S1 (Box) Method The of the human or mouse genomes were downloaded from the UCSC Genome Browser using the Tables feature. The d component of each genome was comprehensively
ONLINE Online supplementary information S1 (Box) Method The of the human or mouse genomes were downloaded from the UCSC Genome Browser using the Tables feature. The d component of each genome was comprehensively
Exercises: Differential Methylation
 Exercises: Differential Methylation Version 2018-04 Exercises: Differential Methylation 2 Licence This manual is 2014-18, Simon Andrews. This manual is distributed under the creative commons Attribution-Non-Commercial-Share
Exercises: Differential Methylation Version 2018-04 Exercises: Differential Methylation 2 Licence This manual is 2014-18, Simon Andrews. This manual is distributed under the creative commons Attribution-Non-Commercial-Share
An epigenetic approach to understanding (and predicting?) environmental effects on gene expression
 www.collaslab.com An epigenetic approach to understanding (and predicting?) environmental effects on gene expression Philippe Collas University of Oslo Institute of Basic Medical Sciences Stem Cell Epigenetics
www.collaslab.com An epigenetic approach to understanding (and predicting?) environmental effects on gene expression Philippe Collas University of Oslo Institute of Basic Medical Sciences Stem Cell Epigenetics
Processing, integrating and analysing chromatin immunoprecipitation followed by sequencing (ChIP-seq) data
 Processing, integrating and analysing chromatin immunoprecipitation followed by sequencing (ChIP-seq) data Bioinformatics methods, models and applications to disease Alex Essebier ChIP-seq experiment To
Processing, integrating and analysing chromatin immunoprecipitation followed by sequencing (ChIP-seq) data Bioinformatics methods, models and applications to disease Alex Essebier ChIP-seq experiment To
STAT1 regulates microrna transcription in interferon γ stimulated HeLa cells
 CAMDA 2009 October 5, 2009 STAT1 regulates microrna transcription in interferon γ stimulated HeLa cells Guohua Wang 1, Yadong Wang 1, Denan Zhang 1, Mingxiang Teng 1,2, Lang Li 2, and Yunlong Liu 2 Harbin
CAMDA 2009 October 5, 2009 STAT1 regulates microrna transcription in interferon γ stimulated HeLa cells Guohua Wang 1, Yadong Wang 1, Denan Zhang 1, Mingxiang Teng 1,2, Lang Li 2, and Yunlong Liu 2 Harbin
Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63.
 Supplementary Figure Legends Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63. A. Screenshot of the UCSC genome browser from normalized RNAPII and RNA-seq ChIP-seq data
Supplementary Figure Legends Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63. A. Screenshot of the UCSC genome browser from normalized RNAPII and RNA-seq ChIP-seq data
PREPARED FOR: U.S. Army Medical Research and Materiel Command Fort Detrick, Maryland
 AD Award Number: W81XWH-12-1-0298 TITLE: MTHFR Functional Polymorphism C677T and Genomic Instability in the Etiology of Idiopathic Autism in Simplex Families PRINCIPAL INVESTIGATOR: Xudong Liu, PhD CONTRACTING
AD Award Number: W81XWH-12-1-0298 TITLE: MTHFR Functional Polymorphism C677T and Genomic Instability in the Etiology of Idiopathic Autism in Simplex Families PRINCIPAL INVESTIGATOR: Xudong Liu, PhD CONTRACTING
Table S1. Total and mapped reads produced for each ChIP-seq sample
 Tale S1. Total and mapped reads produced for each ChIP-seq sample Sample Total Reads Mapped Reads Col- H3K27me3 rep1 125662 1334323 (85.76%) Col- H3K27me3 rep2 9176437 7986731 (87.4%) atmi1a//c H3K27m3
Tale S1. Total and mapped reads produced for each ChIP-seq sample Sample Total Reads Mapped Reads Col- H3K27me3 rep1 125662 1334323 (85.76%) Col- H3K27me3 rep2 9176437 7986731 (87.4%) atmi1a//c H3K27m3
1. To review research methods and the principles of experimental design that are typically used in an experiment.
 Your Name: Section: 36-201 INTRODUCTION TO STATISTICAL REASONING Computer Lab Exercise Lab #7 (there was no Lab #6) Treatment for Depression: A Randomized Controlled Clinical Trial Objectives: 1. To review
Your Name: Section: 36-201 INTRODUCTION TO STATISTICAL REASONING Computer Lab Exercise Lab #7 (there was no Lab #6) Treatment for Depression: A Randomized Controlled Clinical Trial Objectives: 1. To review
Tutorial. ChIP Sequencing. Sample to Insight. September 15, 2016
 ChIP Sequencing September 15, 2016 Sample to Insight CLC bio, a QIAGEN Company Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 www.clcbio.com support-clcbio@qiagen.com ChIP Sequencing
ChIP Sequencing September 15, 2016 Sample to Insight CLC bio, a QIAGEN Company Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 www.clcbio.com support-clcbio@qiagen.com ChIP Sequencing
Histone Modifications Are Associated with Transcript Isoform Diversity in Normal and Cancer Cells
 Histone Modifications Are Associated with Transcript Isoform Diversity in Normal and Cancer Cells Ondrej Podlaha 1, Subhajyoti De 2,3,4, Mithat Gonen 5, Franziska Michor 1 * 1 Department of Biostatistics
Histone Modifications Are Associated with Transcript Isoform Diversity in Normal and Cancer Cells Ondrej Podlaha 1, Subhajyoti De 2,3,4, Mithat Gonen 5, Franziska Michor 1 * 1 Department of Biostatistics
EXPression ANalyzer and DisplayER
 EXPression ANalyzer and DisplayER Tom Hait Aviv Steiner Igor Ulitsky Chaim Linhart Amos Tanay Seagull Shavit Rani Elkon Adi Maron-Katz Dorit Sagir Eyal David Roded Sharan Israel Steinfeld Yossi Shiloh
EXPression ANalyzer and DisplayER Tom Hait Aviv Steiner Igor Ulitsky Chaim Linhart Amos Tanay Seagull Shavit Rani Elkon Adi Maron-Katz Dorit Sagir Eyal David Roded Sharan Israel Steinfeld Yossi Shiloh
Session 6: Integration of epigenetic data. Peter J Park Department of Biomedical Informatics Harvard Medical School July 18-19, 2016
 Session 6: Integration of epigenetic data Peter J Park Department of Biomedical Informatics Harvard Medical School July 18-19, 2016 Utilizing complimentary datasets Frequent mutations in chromatin regulators
Session 6: Integration of epigenetic data Peter J Park Department of Biomedical Informatics Harvard Medical School July 18-19, 2016 Utilizing complimentary datasets Frequent mutations in chromatin regulators
Supplementary Figure 1. Efficiency of Mll4 deletion and its effect on T cell populations in the periphery. Nature Immunology: doi: /ni.
 Supplementary Figure 1 Efficiency of Mll4 deletion and its effect on T cell populations in the periphery. Expression of Mll4 floxed alleles (16-19) in naive CD4 + T cells isolated from lymph nodes and
Supplementary Figure 1 Efficiency of Mll4 deletion and its effect on T cell populations in the periphery. Expression of Mll4 floxed alleles (16-19) in naive CD4 + T cells isolated from lymph nodes and
Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor suppressor genes
 Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor suppressor genes Kaifu Chen 1,2,3,4,5,10, Zhong Chen 6,10, Dayong Wu 6, Lili Zhang 7, Xueqiu Lin 1,2,8,
Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor suppressor genes Kaifu Chen 1,2,3,4,5,10, Zhong Chen 6,10, Dayong Wu 6, Lili Zhang 7, Xueqiu Lin 1,2,8,
Genome-Wide Localization of Protein-DNA Binding and Histone Modification by a Bayesian Change-Point Method with ChIP-seq Data
 Genome-Wide Localization of Protein-DNA Binding and Histone Modification by a Bayesian Change-Point Method with ChIP-seq Data Haipeng Xing, Yifan Mo, Will Liao, Michael Q. Zhang Clayton Davis and Geoffrey
Genome-Wide Localization of Protein-DNA Binding and Histone Modification by a Bayesian Change-Point Method with ChIP-seq Data Haipeng Xing, Yifan Mo, Will Liao, Michael Q. Zhang Clayton Davis and Geoffrey
Analyzing the Loss of Allele-Specific Methylation in Human Colorectal Adenoma with Bisulfite Sequencing
 Research Collection Master Thesis Analyzing the Loss of Allele-Specific Methylation in Human Colorectal Adenoma with Bisulfite Sequencing Author(s): Machlab, Dania Publication Date: 2015 Permanent Link:
Research Collection Master Thesis Analyzing the Loss of Allele-Specific Methylation in Human Colorectal Adenoma with Bisulfite Sequencing Author(s): Machlab, Dania Publication Date: 2015 Permanent Link:
P. Tang ( 鄧致剛 ); PJ Huang ( 黄栢榕 ) g( ); g ( ) Bioinformatics Center, Chang Gung University.
 Databases and Tools for High Throughput Sequencing Analysis P. Tang ( 鄧致剛 ); PJ Huang ( 黄栢榕 ) g( ); g ( ) Bioinformatics Center, Chang Gung University. HTseq Platforms Applications on Biomedical Sciences
Databases and Tools for High Throughput Sequencing Analysis P. Tang ( 鄧致剛 ); PJ Huang ( 黄栢榕 ) g( ); g ( ) Bioinformatics Center, Chang Gung University. HTseq Platforms Applications on Biomedical Sciences
Module 3: Pathway and Drug Development
 Module 3: Pathway and Drug Development Table of Contents 1.1 Getting Started... 6 1.2 Identifying a Dasatinib sensitive cancer signature... 7 1.2.1 Identifying and validating a Dasatinib Signature... 7
Module 3: Pathway and Drug Development Table of Contents 1.1 Getting Started... 6 1.2 Identifying a Dasatinib sensitive cancer signature... 7 1.2.1 Identifying and validating a Dasatinib Signature... 7
Here are the various choices. All of them are found in the Analyze menu in SPSS, under the sub-menu for Descriptive Statistics :
 Descriptive Statistics in SPSS When first looking at a dataset, it is wise to use descriptive statistics to get some idea of what your data look like. Here is a simple dataset, showing three different
Descriptive Statistics in SPSS When first looking at a dataset, it is wise to use descriptive statistics to get some idea of what your data look like. Here is a simple dataset, showing three different
Integrated analysis of sequencing data
 Integrated analysis of sequencing data How to combine *-seq data M. Defrance, M. Thomas-Chollier, C. Herrmann, D. Puthier, J. van Helden *ChIP-seq, RNA-seq, MeDIP-seq, Transcription factor binding ChIP-seq
Integrated analysis of sequencing data How to combine *-seq data M. Defrance, M. Thomas-Chollier, C. Herrmann, D. Puthier, J. van Helden *ChIP-seq, RNA-seq, MeDIP-seq, Transcription factor binding ChIP-seq
Nature Structural & Molecular Biology: doi: /nsmb.2419
 Supplementary Figure 1 Mapped sequence reads and nucleosome occupancies. (a) Distribution of sequencing reads on the mouse reference genome for chromosome 14 as an example. The number of reads in a 1 Mb
Supplementary Figure 1 Mapped sequence reads and nucleosome occupancies. (a) Distribution of sequencing reads on the mouse reference genome for chromosome 14 as an example. The number of reads in a 1 Mb
Visualizing Biopolymers and Their Building Blocks
 Visualizing Biopolymers and Their Building Blocks By Sharlene Denos (UIUC) & Kathryn Hafner (Danville High) Living things are primarily composed of carbon-based (organic) polymers. These are made up many
Visualizing Biopolymers and Their Building Blocks By Sharlene Denos (UIUC) & Kathryn Hafner (Danville High) Living things are primarily composed of carbon-based (organic) polymers. These are made up many
Heintzman, ND, Stuart, RK, Hon, G, Fu, Y, Ching, CW, Hawkins, RD, Barrera, LO, Van Calcar, S, Qu, C, Ching, KA, Wang, W, Weng, Z, Green, RD,
 Heintzman, ND, Stuart, RK, Hon, G, Fu, Y, Ching, CW, Hawkins, RD, Barrera, LO, Van Calcar, S, Qu, C, Ching, KA, Wang, W, Weng, Z, Green, RD, Crawford, GE, Ren, B (2007) Distinct and predictive chromatin
Heintzman, ND, Stuart, RK, Hon, G, Fu, Y, Ching, CW, Hawkins, RD, Barrera, LO, Van Calcar, S, Qu, C, Ching, KA, Wang, W, Weng, Z, Green, RD, Crawford, GE, Ren, B (2007) Distinct and predictive chromatin
Run Time Tester Requirements Document
 Run Time Tester Requirements Document P. Sherwood 7 March 2004 Version: 0.4 Status: Draft After review 2 March 2004 Contents 1 Introduction 2 1.1 Scope of the Requirements Document.....................
Run Time Tester Requirements Document P. Sherwood 7 March 2004 Version: 0.4 Status: Draft After review 2 March 2004 Contents 1 Introduction 2 1.1 Scope of the Requirements Document.....................
OECD QSAR Toolbox v.4.2. An example illustrating RAAF scenario 6 and related assessment elements
 OECD QSAR Toolbox v.4.2 An example illustrating RAAF scenario 6 and related assessment elements Outlook Background Objectives Specific Aims Read Across Assessment Framework (RAAF) The exercise Workflow
OECD QSAR Toolbox v.4.2 An example illustrating RAAF scenario 6 and related assessment elements Outlook Background Objectives Specific Aims Read Across Assessment Framework (RAAF) The exercise Workflow
Computer Science 101 Project 2: Predator Prey Model
 Computer Science 101 Project 2: Predator Prey Model Real-life situations usually are complicated and difficult to model exactly because of the large number of variables present in real systems. Computer
Computer Science 101 Project 2: Predator Prey Model Real-life situations usually are complicated and difficult to model exactly because of the large number of variables present in real systems. Computer
7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans.
 Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
SUPPLEMENTARY FIGURES: Supplementary Figure 1
 SUPPLEMENTARY FIGURES: Supplementary Figure 1 Supplementary Figure 1. Glioblastoma 5hmC quantified by paired BS and oxbs treated DNA hybridized to Infinium DNA methylation arrays. Workflow depicts analytic
SUPPLEMENTARY FIGURES: Supplementary Figure 1 Supplementary Figure 1. Glioblastoma 5hmC quantified by paired BS and oxbs treated DNA hybridized to Infinium DNA methylation arrays. Workflow depicts analytic
Unit 1 Outline Science Practices. Part 1 - The Scientific Method. Screencasts found at: sciencepeek.com. 1. List the steps of the scientific method.
 Screencasts found at: sciencepeek.com Part 1 - The Scientific Method 1. List the steps of the scientific method. 2. What is an observation? Give an example. Quantitative or Qualitative Data? 35 grams?
Screencasts found at: sciencepeek.com Part 1 - The Scientific Method 1. List the steps of the scientific method. 2. What is an observation? Give an example. Quantitative or Qualitative Data? 35 grams?
Two-Way Independent ANOVA
 Two-Way Independent ANOVA Analysis of Variance (ANOVA) a common and robust statistical test that you can use to compare the mean scores collected from different conditions or groups in an experiment. There
Two-Way Independent ANOVA Analysis of Variance (ANOVA) a common and robust statistical test that you can use to compare the mean scores collected from different conditions or groups in an experiment. There
Hour 2: lm (regression), plot (scatterplots), cooks.distance and resid (diagnostics) Stat 302, Winter 2016 SFU, Week 3, Hour 1, Page 1
 Agenda for Week 3, Hr 1 (Tuesday, Jan 19) Hour 1: - Installing R and inputting data. - Different tools for R: Notepad++ and RStudio. - Basic commands:?,??, mean(), sd(), t.test(), lm(), plot() - t.test()
Agenda for Week 3, Hr 1 (Tuesday, Jan 19) Hour 1: - Installing R and inputting data. - Different tools for R: Notepad++ and RStudio. - Basic commands:?,??, mean(), sd(), t.test(), lm(), plot() - t.test()
ImageJ plugin for semiautomatic measurement of roentgenological attachment level
 ImageJ plugin for semiautomatic measurement of roentgenological attachment level User manual and installation instructions v. 1.1, June 2015 The plugin is written by Gerald R. Torgersen 1 based on the
ImageJ plugin for semiautomatic measurement of roentgenological attachment level User manual and installation instructions v. 1.1, June 2015 The plugin is written by Gerald R. Torgersen 1 based on the
DNA-seq Bioinformatics Analysis: Copy Number Variation
 DNA-seq Bioinformatics Analysis: Copy Number Variation Elodie Girard elodie.girard@curie.fr U900 institut Curie, INSERM, Mines ParisTech, PSL Research University Paris, France NGS Applications 5C HiC DNA-seq
DNA-seq Bioinformatics Analysis: Copy Number Variation Elodie Girard elodie.girard@curie.fr U900 institut Curie, INSERM, Mines ParisTech, PSL Research University Paris, France NGS Applications 5C HiC DNA-seq
Figure S2. Distribution of acgh probes on all ten chromosomes of the RIL M0022
 96 APPENDIX B. Supporting Information for chapter 4 "changes in genome content generated via segregation of non-allelic homologs" Figure S1. Potential de novo CNV probes and sizes of apparently de novo
96 APPENDIX B. Supporting Information for chapter 4 "changes in genome content generated via segregation of non-allelic homologs" Figure S1. Potential de novo CNV probes and sizes of apparently de novo
The Epigenome Tools 2: ChIP-Seq and Data Analysis
 The Epigenome Tools 2: ChIP-Seq and Data Analysis Chongzhi Zang zang@virginia.edu http://zanglab.com PHS5705: Public Health Genomics March 20, 2017 1 Outline Epigenome: basics review ChIP-seq overview
The Epigenome Tools 2: ChIP-Seq and Data Analysis Chongzhi Zang zang@virginia.edu http://zanglab.com PHS5705: Public Health Genomics March 20, 2017 1 Outline Epigenome: basics review ChIP-seq overview
Supplementary Material for IPred - Integrating Ab Initio and Evidence Based Gene Predictions to Improve Prediction Accuracy
 1 SYSTEM REQUIREMENTS 1 Supplementary Material for IPred - Integrating Ab Initio and Evidence Based Gene Predictions to Improve Prediction Accuracy Franziska Zickmann and Bernhard Y. Renard Research Group
1 SYSTEM REQUIREMENTS 1 Supplementary Material for IPred - Integrating Ab Initio and Evidence Based Gene Predictions to Improve Prediction Accuracy Franziska Zickmann and Bernhard Y. Renard Research Group
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Heatmap of GO terms for differentially expressed genes. The terms were hierarchically clustered using the GO term enrichment beta. Darker red, higher positive
Supplementary Figures Supplementary Figure 1. Heatmap of GO terms for differentially expressed genes. The terms were hierarchically clustered using the GO term enrichment beta. Darker red, higher positive
EXTENDED DATA FORMATTING GUIDE
 EXTENDED DATA FORMATTING GUIDE Nature paper but are included online within the full-text HTML and at the end of the online PDF. Nature separate panel (for example, see Extended Data Figure 2, panel d)
EXTENDED DATA FORMATTING GUIDE Nature paper but are included online within the full-text HTML and at the end of the online PDF. Nature separate panel (for example, see Extended Data Figure 2, panel d)
Quantification of DNA Methylation and Epimutations from Bisulfite Sequencing Data the BEAT package
 Quantification of DNA Methylation and Epimutations from Bisulfite Sequencing Data the BEAT package Kemal Akman 1, Achim Tresch Max-Planck-Institute for Plant Breeding Research Cologne, Germany 1 akman@mpipz.mpg.de
Quantification of DNA Methylation and Epimutations from Bisulfite Sequencing Data the BEAT package Kemal Akman 1, Achim Tresch Max-Planck-Institute for Plant Breeding Research Cologne, Germany 1 akman@mpipz.mpg.de
Back to the Basics: Methyl-Seq 101
 Back to the Basics: Methyl-Seq 101 Presented By: Alex Siebold, Ph.D. October 9, 2013 Field Applications Scientist Agilent Technologies Life Sciences & Diagnostics Group Life Sciences & Diagnostics Group
Back to the Basics: Methyl-Seq 101 Presented By: Alex Siebold, Ph.D. October 9, 2013 Field Applications Scientist Agilent Technologies Life Sciences & Diagnostics Group Life Sciences & Diagnostics Group
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Frequency of alternative-cassette-exon engagement with the ribosome is consistent across data from multiple human cell types and from mouse stem cells. Box plots showing AS frequency
Supplementary Figure 1 Frequency of alternative-cassette-exon engagement with the ribosome is consistent across data from multiple human cell types and from mouse stem cells. Box plots showing AS frequency
Not IN Our Genes - A Different Kind of Inheritance.! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014
 Not IN Our Genes - A Different Kind of Inheritance! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014 Epigenetics in Mainstream Media Epigenetics *Current definition:
Not IN Our Genes - A Different Kind of Inheritance! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014 Epigenetics in Mainstream Media Epigenetics *Current definition:
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 U1 inhibition causes a shift of RNA-seq reads from exons to introns. (a) Evidence for the high purity of 4-shU-labeled RNAs used for RNA-seq. HeLa cells transfected with control
Supplementary Figure 1 U1 inhibition causes a shift of RNA-seq reads from exons to introns. (a) Evidence for the high purity of 4-shU-labeled RNAs used for RNA-seq. HeLa cells transfected with control
Nature Immunology: doi: /ni Supplementary Figure 1. Characteristics of SEs in T reg and T conv cells.
 Supplementary Figure 1 Characteristics of SEs in T reg and T conv cells. (a) Patterns of indicated transcription factor-binding at SEs and surrounding regions in T reg and T conv cells. Average normalized
Supplementary Figure 1 Characteristics of SEs in T reg and T conv cells. (a) Patterns of indicated transcription factor-binding at SEs and surrounding regions in T reg and T conv cells. Average normalized
GridMAT-MD: A Grid-based Membrane Analysis Tool for use with Molecular Dynamics
 GridMAT-MD: A Grid-based Membrane Analysis Tool for use with Molecular Dynamics William J. Allen, Justin A. Lemkul, and David R. Bevan Department of Biochemistry, Virginia Tech User s Guide Version 1.0.2
GridMAT-MD: A Grid-based Membrane Analysis Tool for use with Molecular Dynamics William J. Allen, Justin A. Lemkul, and David R. Bevan Department of Biochemistry, Virginia Tech User s Guide Version 1.0.2
Benchmark Dose Modeling Cancer Models. Allen Davis, MSPH Jeff Gift, Ph.D. Jay Zhao, Ph.D. National Center for Environmental Assessment, U.S.
 Benchmark Dose Modeling Cancer Models Allen Davis, MSPH Jeff Gift, Ph.D. Jay Zhao, Ph.D. National Center for Environmental Assessment, U.S. EPA Disclaimer The views expressed in this presentation are those
Benchmark Dose Modeling Cancer Models Allen Davis, MSPH Jeff Gift, Ph.D. Jay Zhao, Ph.D. National Center for Environmental Assessment, U.S. EPA Disclaimer The views expressed in this presentation are those
Dr Rick Tearle Senior Applications Specialist, EMEA Complete Genomics Complete Genomics, Inc.
 Dr Rick Tearle Senior Applications Specialist, EMEA Complete Genomics Topics Overview of Data Processing Pipeline Overview of Data Files 2 DNA Nano-Ball (DNB) Read Structure Genome : acgtacatgcattcacacatgcttagctatctctcgccag
Dr Rick Tearle Senior Applications Specialist, EMEA Complete Genomics Topics Overview of Data Processing Pipeline Overview of Data Files 2 DNA Nano-Ball (DNB) Read Structure Genome : acgtacatgcattcacacatgcttagctatctctcgccag
PSSV User Manual (V2.1)
 PSSV User Manual (V2.1) 1. Introduction A novel pattern-based probabilistic approach, PSSV, is developed to identify somatic structural variations from WGS data. Specifically, discordant and concordant
PSSV User Manual (V2.1) 1. Introduction A novel pattern-based probabilistic approach, PSSV, is developed to identify somatic structural variations from WGS data. Specifically, discordant and concordant
Instructions for the ECN201 Project on Least-Cost Nutritionally-Adequate Diets
 Instructions for the ECN201 Project on Least-Cost Nutritionally-Adequate Diets John P. Burkett October 15, 2015 1 Overview For this project, each student should (a) determine her or his nutritional needs,
Instructions for the ECN201 Project on Least-Cost Nutritionally-Adequate Diets John P. Burkett October 15, 2015 1 Overview For this project, each student should (a) determine her or his nutritional needs,
Discovery of Novel Human Gene Regulatory Modules from Gene Co-expression and
 Discovery of Novel Human Gene Regulatory Modules from Gene Co-expression and Promoter Motif Analysis Shisong Ma 1,2*, Michael Snyder 3, and Savithramma P Dinesh-Kumar 2* 1 School of Life Sciences, University
Discovery of Novel Human Gene Regulatory Modules from Gene Co-expression and Promoter Motif Analysis Shisong Ma 1,2*, Michael Snyder 3, and Savithramma P Dinesh-Kumar 2* 1 School of Life Sciences, University
Part-II: Statistical analysis of ChIP-seq data
 Part-II: Statistical analysis of ChIP-seq data Outline ChIP-seq data, features, detailed modeling aspects (today). Other ChIP-seq related problems - overview (next lecture). IDR (next lecture) Stat 877
Part-II: Statistical analysis of ChIP-seq data Outline ChIP-seq data, features, detailed modeling aspects (today). Other ChIP-seq related problems - overview (next lecture). IDR (next lecture) Stat 877
A Practical Guide to Integrative Genomics by RNA-seq and ChIP-seq Analysis
 A Practical Guide to Integrative Genomics by RNA-seq and ChIP-seq Analysis Jian Xu, Ph.D. Children s Research Institute, UTSW Introduction Outline Overview of genomic and next-gen sequencing technologies
A Practical Guide to Integrative Genomics by RNA-seq and ChIP-seq Analysis Jian Xu, Ph.D. Children s Research Institute, UTSW Introduction Outline Overview of genomic and next-gen sequencing technologies
Titrations in Cytobank
 The Premier Platform for Single Cell Analysis (1) Titrations in Cytobank To analyze data from a titration in Cytobank, the first step is to upload your own FCS files or clone an experiment you already
The Premier Platform for Single Cell Analysis (1) Titrations in Cytobank To analyze data from a titration in Cytobank, the first step is to upload your own FCS files or clone an experiment you already
MethylMix An R package for identifying DNA methylation driven genes
 MethylMix An R package for identifying DNA methylation driven genes Olivier Gevaert May 3, 2016 Stanford Center for Biomedical Informatics Department of Medicine 1265 Welch Road Stanford CA, 94305-5479
MethylMix An R package for identifying DNA methylation driven genes Olivier Gevaert May 3, 2016 Stanford Center for Biomedical Informatics Department of Medicine 1265 Welch Road Stanford CA, 94305-5479
OUTLIER SUBJECTS PROTOCOL (art_groupoutlier)
 OUTLIER SUBJECTS PROTOCOL (art_groupoutlier) Paul K. Mazaika 2/23/2009 Outlier subjects are a problem in fmri data sets for clinical populations. This protocol and program are a method to identify outlier
OUTLIER SUBJECTS PROTOCOL (art_groupoutlier) Paul K. Mazaika 2/23/2009 Outlier subjects are a problem in fmri data sets for clinical populations. This protocol and program are a method to identify outlier
Problem 3: Simulated Rheumatoid Arthritis Data
 Problem 3: Simulated Rheumatoid Arthritis Data Michael B Miller Michael Li Gregg Lind Soon-Young Jang The plan
Problem 3: Simulated Rheumatoid Arthritis Data Michael B Miller Michael Li Gregg Lind Soon-Young Jang The plan
Peak-calling for ChIP-seq and ATAC-seq
 Peak-calling for ChIP-seq and ATAC-seq Shamith Samarajiwa CRUK Autumn School in Bioinformatics 2017 University of Cambridge Overview Peak-calling: identify enriched (signal) regions in ChIP-seq or ATAC-seq
Peak-calling for ChIP-seq and ATAC-seq Shamith Samarajiwa CRUK Autumn School in Bioinformatics 2017 University of Cambridge Overview Peak-calling: identify enriched (signal) regions in ChIP-seq or ATAC-seq
Metadata of the chapter that will be visualized online
 Metadata of the chapter that will be visualized online ChapterTitle Chapter Sub-Title Advanced Analysis of Human Plasma Circulating DNA Sequences Produced by Parallel Tagged Sequencing on the 454 Platform
Metadata of the chapter that will be visualized online ChapterTitle Chapter Sub-Title Advanced Analysis of Human Plasma Circulating DNA Sequences Produced by Parallel Tagged Sequencing on the 454 Platform
LAB ASSIGNMENT 4 INFERENCES FOR NUMERICAL DATA. Comparison of Cancer Survival*
 LAB ASSIGNMENT 4 1 INFERENCES FOR NUMERICAL DATA In this lab assignment, you will analyze the data from a study to compare survival times of patients of both genders with different primary cancers. First,
LAB ASSIGNMENT 4 1 INFERENCES FOR NUMERICAL DATA In this lab assignment, you will analyze the data from a study to compare survival times of patients of both genders with different primary cancers. First,
ChIP-seq analysis. J. van Helden, M. Defrance, C. Herrmann, D. Puthier, N. Servant, M. Thomas-Chollier, O.Sand
 ChIP-seq analysis J. van Helden, M. Defrance, C. Herrmann, D. Puthier, N. Servant, M. Thomas-Chollier, O.Sand Tuesday : quick introduction to ChIP-seq and peak-calling (Presentation + Practical session)
ChIP-seq analysis J. van Helden, M. Defrance, C. Herrmann, D. Puthier, N. Servant, M. Thomas-Chollier, O.Sand Tuesday : quick introduction to ChIP-seq and peak-calling (Presentation + Practical session)
Nature Genetics: doi: /ng Supplementary Figure 1. Assessment of sample purity and quality.
 Supplementary Figure 1 Assessment of sample purity and quality. (a) Hematoxylin and eosin staining of formaldehyde-fixed, paraffin-embedded sections from a human testis biopsy collected concurrently with
Supplementary Figure 1 Assessment of sample purity and quality. (a) Hematoxylin and eosin staining of formaldehyde-fixed, paraffin-embedded sections from a human testis biopsy collected concurrently with
Supervised Learner for the Prediction of Hi-C Interaction Counts and Determination of Influential Features. Tyler Yue Lab
 Supervised Learner for the Prediction of Hi-C Interaction Counts and Determination of Influential Features Tyler Derr @ Yue Lab tsd5037@psu.edu Background Hi-C is a chromosome conformation capture (3C)
Supervised Learner for the Prediction of Hi-C Interaction Counts and Determination of Influential Features Tyler Derr @ Yue Lab tsd5037@psu.edu Background Hi-C is a chromosome conformation capture (3C)
sirna count per 50 kb small RNAs matching the direct strand Repeat length (bp) per 50 kb repeats in the chromosome
 Qi et al. 26-3-2564C Qi et al., Figure S1 sirna count per 5 kb small RNAs matching the direct strand sirna count per 5 kb small RNAs matching the complementary strand Repeat length (bp) per 5 kb repeats
Qi et al. 26-3-2564C Qi et al., Figure S1 sirna count per 5 kb small RNAs matching the direct strand sirna count per 5 kb small RNAs matching the complementary strand Repeat length (bp) per 5 kb repeats
IPUMS â CPS Extraction and Analysis
 Minnesota Population Center Training and Development IPUMS â CPS Extraction and Analysis Exercise 2 OBJECTIVE: Gain an understanding of how the IPUMS dataset is structured and how it can be leveraged to
Minnesota Population Center Training and Development IPUMS â CPS Extraction and Analysis Exercise 2 OBJECTIVE: Gain an understanding of how the IPUMS dataset is structured and how it can be leveraged to
Package NarrowPeaks. August 3, Version Date Type Package
 Package NarrowPeaks August 3, 2013 Version 1.5.0 Date 2013-02-13 Type Package Title Analysis of Variation in ChIP-seq using Functional PCA Statistics Author Pedro Madrigal , with contributions
Package NarrowPeaks August 3, 2013 Version 1.5.0 Date 2013-02-13 Type Package Title Analysis of Variation in ChIP-seq using Functional PCA Statistics Author Pedro Madrigal , with contributions
The Effectiveness of Captopril
 Lab 7 The Effectiveness of Captopril In the United States, pharmaceutical manufacturers go through a very rigorous process in order to get their drugs approved for sale. This process is designed to determine
Lab 7 The Effectiveness of Captopril In the United States, pharmaceutical manufacturers go through a very rigorous process in order to get their drugs approved for sale. This process is designed to determine
cn.mops - Mixture of Poissons for CNV detection in NGS data Günter Klambauer Institute of Bioinformatics, Johannes Kepler University Linz
 Software Manual Institute of Bioinformatics, Johannes Kepler University Linz cn.mops - Mixture of Poissons for CNV detection in NGS data Günter Klambauer Institute of Bioinformatics, Johannes Kepler University
Software Manual Institute of Bioinformatics, Johannes Kepler University Linz cn.mops - Mixture of Poissons for CNV detection in NGS data Günter Klambauer Institute of Bioinformatics, Johannes Kepler University
Preliminary Report on Simple Statistical Tests (t-tests and bivariate correlations)
 Preliminary Report on Simple Statistical Tests (t-tests and bivariate correlations) After receiving my comments on the preliminary reports of your datasets, the next step for the groups is to complete
Preliminary Report on Simple Statistical Tests (t-tests and bivariate correlations) After receiving my comments on the preliminary reports of your datasets, the next step for the groups is to complete
