ChIP-seq analysis. J. van Helden, M. Defrance, C. Herrmann, D. Puthier, N. Servant, M. Thomas-Chollier, O.Sand
|
|
|
- Opal Peters
- 6 years ago
- Views:
Transcription
1 ChIP-seq analysis J. van Helden, M. Defrance, C. Herrmann, D. Puthier, N. Servant, M. Thomas-Chollier, O.Sand Tuesday : quick introduction to ChIP-seq and peak-calling (Presentation + Practical session) Functional annotation of peaks Motif analysis of ChIP-seq peaks Peak quality assessment Interpretation of ChIP-seq/ChIP-exo data
2 Datasets used estrogen-receptor (ESR1) is a key factor in breast cancer developement goal of the study: understand the dependency of ESR1 binding on presence of co-factors, in particular GATA3, which is mutated in breast cancers approaches: GATA3 silencing (sirna), ChIP-seq on ESR1 in wt vs. sigata3 conditions, chromatin profiling
3 Datasets used ESR1 ChIP-seq in WT & sigata3 conditions ( 3 replicates = 6 datasets) H3K4me1 in WT & sigata3 conditions (1 replicate = 2 datasets) Input dataset in MCF-7 (1 replicate = 1 dataset) p300 before estrogen stimulation GATA3/FOXA1 ChIP-seq before/after estrogen stimulation microarray expression data, etc...
4 Chromatin more than just sequence - histone modifications - transcription factor binding - DNA methylation [ Wasserman & Sandelin, Nat.Rev.Gen (2004) ] Carl Herrmann Cancer Regulatory Genomics
5 Histone modifications Carl Herrmann Cancer Regulatory Genomics histones are subject to post-translational modifications at their N-terminal tail Lysine methylation Lysine/arginine acethylation Serine phosphorylation ubiquitylation
6 Chromatin binding proteins Histone modifying enzymes [Wikipedia] Carl Herrmann Cancer Regulatory Genomics
7 Experimental identification of binding sites Carl Herrmann Cancer Regulatory Genomics Chromatin immunoprecipitation (ChIP) followed by sequencing (ChIP-seq) hybridization on array (mostly tilling arrays) ChIP-chip PCR/qPCR main challenge : quality/specificity of the antibodies
8 ChIP-seq signal for transcription factors ChIP seq on DNA binding TF read densities on +/- strand We expect to see a typical strand asymetry in read densities ChIP peak recognition pattern
9 ChIP-seq signal for transcription factors treatment read density (=WIG) input read density (=WIG) peak (=BED) aligned reads + strand (=BAM) aligned reads - strand (=BAM) (this is the data you are going to manipulate...)
10 ChIP-seq signal for transcription factors ChIP seq on DNA binding TF read densities on +/- strand Binding of several TF as complexes tend to blur this asymmetry
11 ChIP-seq signal for histone marks ChIP seq on histone modifications read densities on +/- strand The strand asymmetry is completely lost when considering ChIP datasets for diffuse histone modifications
12 Real example of ChIP-seq signal H3K4me1 H3K4me1 reads
13 Real example of ChIP-seq signal ESR1 input H3K4me1 ESR1 reads H3K4me1 reads
14 Keys aspects of peak finding Treating the reads Modelling noise levels Scaling datasets Detecting enriched/peak regions Dealing with replicates ( Exercices)
15 From aligned reads to binding sites Tag shifting vs. extension positive/negative strand read peaks do not represent the true location of the binding site reads can be shifted by d/2 where d is the band size (MACS) increased resolution reads can be elongated to a size of d (FindPeaks, PeakSeq,...) d can be estimate from the data (MACS) or given as input parameter example of MACS model building using top enriched regions
16 From aligned reads to binding sites d/2 Tag shifting shifted position initial position read densities on +/- strand Each tag is shifted by d/2 (i.e. towards the middle of the IP fragment) where d represent the fragment length
17 From aligned reads to binding sites Tag elongation read densities on +/- strand Each tag is computationaly extended in 3' to a total length of d
18 Modelling noise levels ChIP-seq dataset (=treatment) signal = + background noise How do we estimate the noise?
19 Modelling noise levels noise is not uniform (chromatin conformation, local biases, mappability) input dataset is mandatory for reliable local estimation! (although some algorithms do not require it :-( ) chr1:114,720, ,746, kb treatment input?
20 Modelling noise levels the mappability is related to the uniqueness of the k-mers at a particular position of the genome repetitive regions low uniqueness low mappability chr1:114,720, ,746, kb k=35 k=50 k=100 Longer reads more uniquely mapped reads
21 Modelling noise levels random distribution of reads in a window of size w modelled using a theoretical distribution Poisson distribution 1 parameter : λ = expected number of reads in window k P ( X =k )=e λk k! Binomial distribution 2 parameters: p = probability to start a read at a particular position n = number of positions in the window ~ window size (assumes no duplicates!) np = expected number of reads in window P ( X =k )=C kn p k (1 p) n k
22 Modelling noise levels Negative Binomial distribution 2 parameters: p = probability to start a read at a particular position r = number of successes NB distribution can have arbitrarily large variance
23 Scaling unequal datasets treatment (=signal + noise) and input (=noise) datasets generally do not have the same sequencing depth need for normalization input dataset should model the noise level in the treatment dataset naïve approach : upscale/downscale the smaller/larger dataset Input : N reads ChIP-seq dataset M > N reads scale by library size : M M' = N Problem : signal influences scaling factor More signal (but equal noise) artificial noise over-estimation
24 Scaling unequal datasets by library size input 1 area ~ number of reads = treatment area ~ number of reads = = 18 Scaling by library size : upscale input by 18/10 = treatment 10 estimated noise level 1 Noise level is over-estimated! 10
25 Scaling unequal datasets by library size input 1 area ~ number of reads = treatment area ~ number of reads = = 23 Scaling by library size : upscale input by 23/10 = treatment 10 estimated noise level 1 10
26 Scaling unequal datasets by library size input 1 area ~ number of reads = treatment area ~ number of reads = = 23 Scaling by library size : upscale input by 23/10 = treatment 10 estimated noise level 1 10
27 Scaling unequal datasets more advanced : linear regression by exclusing peak regions (PeakSeq) read counts in 1Mb regions in input and treatment all regions excluding enriched (=signal) regions
28 Defining peaks Determining enriched regions sliding window across the genome at each location, evaluate the enrichement of the signal wrt. expected background based on the distribution retain regions with P-values below threshold evaluate FDR Pval < 1e-20 Pval ~ 0.6
29
30 Some methods separate the tag densities into different strands and take advantage of tag asymmetry Most consider merged densities and look for enrichment
31 Tag shift Tag extension Tags unchanged
32 MACS [Zhang et al. Genome Biol. 2008] Step 1 : estimating fragment length d slide a window of size BANDWIDTH retain top regions with MFOLD enrichment of treatment vs. input plot average +/- strand read densities estimate d enrichment > MFOLD treatment control
33 MACS [Zhang et al. Genome Biol. 2008] Step 2 : identification of local noise parameter slide a window of size 2*d across treatment and input estimate parameter λlocal of Poisson distribution 1 kb 5 kb 10 kb full genome estimate λ over diff. ranges take the max
34 MACS [Zhang et al. Genome Biol. 2008] Step 3 : identification of enriched/peak regions determine regions with P-values < PVALUE determine summit position inside enriched regions as max density P-val = 1e-30
35 MACS [Zhang et al. Genome Biol. 2008] Step 4 : estimating FDR positive peaks (P-values) swap treatment and input; call negative peaks (P-value) FDR(p) = # negative peaks with Pval < p # positive peaks with Pval < p increasing P-value FDR = 2/25=0.08
36 Differential enrichment analysis ChIP-seq performed under two different conditions Question : what are the differentially enriched/bound peak regions? application to histone modifications DNA methylation Nair et al., PLOS One 2012
37 Differential enrichment analysis ChIPDiff (Xu. et al., Bioinformatics 2008) statistical model : HMM (1 = not enriched; 2 = enriched in samplea; 3 = enriched in sample B) does not need pre-defined peaks/regions command line executable ChIPnorm (Nair et al., PLOS One 2012) quantile normalization of enriched-significant bins in both samples requires signal and input datasets for both samples MATLAB program MAnorm common peaks MAnorm (Shao et al., Genome Biology 2012) MA based normalization of regions containing common peaks applied to all regions requires a priori defined peaks for each library MATLAB/R program all peaks DiffReps (Shen et al. PLOS One, 2013) perl executables
38 Important questions to aks (beforehand...) Nature Methods 2012
39 Important questions to aks (beforehand...) sequencing depth? library complexity (insufficient depth insufficient complexity) saturation of ChIP peaks not achieved choice of peak caller narrow peaks (TF) or broad peaks (histone modification) sensitivity / specificity reproducibility (across replicates) Irreproducible Discovery Range (IDR) TF H3K36me3
40 Important questions to aks (beforehand...) single vs PE? PE improves mapping in low-complexity regions improves library complexity but is generally an (financial) overkill remove redundant reads? for deeply sequenced libraries, redundant reads are not always artefacts...
41 Functional annotation of ChIP-peaks how do we go from peaks to functions?
42 Interesting questions to ask where do peaks localize? proximal to TSS? distal (= enhancer) regions? what are the closest genes (potential targets)? is there a functional enrichment (e.g. GO categories) in genes/regions bound?
43 Positional biases where do peaks localize? Proximal promoter? Intergenic regions? Intronic regions?
44 Positional biases Distance to TSS :
45 Peaks Genes Functions collect sets of genes compute over-represented functional annotations Gene Ontology Phenotypic annotations Biological Pathways Typical tools DAVID [Huang et al., NAR 2009] Babelomics [Medina et al., NAR 2010]
46 Peaks Genes Functions 5kb 5kb Drawbacks restricting to proximal regions discards a large number of binding events "nearest gene" approach introduces bias towards genes with large intergenic regions e.g. : "multicellular organism development" : 14% of the genes, but 33% of the genome associated
47 Genes Regions Peaks Idea : assign functional annotation to genomic regions use statistics to avoid biases assign to each gene a regulatory domain basal (-5kb/+1kb from TSS) extended (up to nearest basal region ; max 1Mb) each domain is annotated to the functional terms of the corresponding gene "Functional domains" "GREAT improves functional interpretation of cis-regulatory regions" McLean et al. Nat. Biotech. (2010)
48 Genes Regions Peaks term A term B Given that 60% of the genome is annotated to A, would I randomly expect 3 or more peaks to fall into region A? Given that 15% of the genome is annotated to B, would I randomly expect 3 or more peaks to fall into region B? "GREAT improves functional interpretation of cis-regulatory regions" McLean et al. Nat. Biotech. (2010) p > 0.5 p = 0.07
49
50 GREAT vs. proximal peaks GREAT Proximal 2kb peaks Best GO term P-val MGI expression P-val Best GO-term p300 limb Embryonic limb morphogenesis 1E-27 TS19 limb 7E-49 Skeletal system 4E-06 development p300 forebrain CNS development 8E-36 TS17 forebrain 6E-41 Forebrain development p300 midbrain CNS development 1E-12 TS 15 CNS 1E-14 none more specific terms with higher significance more peaks/genes taken into account P-val MGI expression P-val 2E-04 TS19 limb 3E-05 TS22 forebrain 3E-07 none
Peak-calling for ChIP-seq and ATAC-seq
 Peak-calling for ChIP-seq and ATAC-seq Shamith Samarajiwa CRUK Autumn School in Bioinformatics 2017 University of Cambridge Overview Peak-calling: identify enriched (signal) regions in ChIP-seq or ATAC-seq
Peak-calling for ChIP-seq and ATAC-seq Shamith Samarajiwa CRUK Autumn School in Bioinformatics 2017 University of Cambridge Overview Peak-calling: identify enriched (signal) regions in ChIP-seq or ATAC-seq
ChIP-seq data analysis
 ChIP-seq data analysis Harri Lähdesmäki Department of Computer Science Aalto University November 24, 2017 Contents Background ChIP-seq protocol ChIP-seq data analysis Transcriptional regulation Transcriptional
ChIP-seq data analysis Harri Lähdesmäki Department of Computer Science Aalto University November 24, 2017 Contents Background ChIP-seq protocol ChIP-seq data analysis Transcriptional regulation Transcriptional
Computational aspects of ChIP-seq. John Marioni Research Group Leader European Bioinformatics Institute European Molecular Biology Laboratory
 Computational aspects of ChIP-seq John Marioni Research Group Leader European Bioinformatics Institute European Molecular Biology Laboratory ChIP-seq Using highthroughput sequencing to investigate DNA
Computational aspects of ChIP-seq John Marioni Research Group Leader European Bioinformatics Institute European Molecular Biology Laboratory ChIP-seq Using highthroughput sequencing to investigate DNA
Accessing and Using ENCODE Data Dr. Peggy J. Farnham
 1 William M Keck Professor of Biochemistry Keck School of Medicine University of Southern California How many human genes are encoded in our 3x10 9 bp? C. elegans (worm) 959 cells and 1x10 8 bp 20,000
1 William M Keck Professor of Biochemistry Keck School of Medicine University of Southern California How many human genes are encoded in our 3x10 9 bp? C. elegans (worm) 959 cells and 1x10 8 bp 20,000
Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq
 Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne ChIP-seq against histone variants: Biological
Computational Analysis of UHT Sequences Histone modifications, CAGE, RNA-Seq Philipp Bucher Wednesday January 21, 2009 SIB graduate school course EPFL, Lausanne ChIP-seq against histone variants: Biological
The Epigenome Tools 2: ChIP-Seq and Data Analysis
 The Epigenome Tools 2: ChIP-Seq and Data Analysis Chongzhi Zang zang@virginia.edu http://zanglab.com PHS5705: Public Health Genomics March 20, 2017 1 Outline Epigenome: basics review ChIP-seq overview
The Epigenome Tools 2: ChIP-Seq and Data Analysis Chongzhi Zang zang@virginia.edu http://zanglab.com PHS5705: Public Health Genomics March 20, 2017 1 Outline Epigenome: basics review ChIP-seq overview
Processing, integrating and analysing chromatin immunoprecipitation followed by sequencing (ChIP-seq) data
 Processing, integrating and analysing chromatin immunoprecipitation followed by sequencing (ChIP-seq) data Bioinformatics methods, models and applications to disease Alex Essebier ChIP-seq experiment To
Processing, integrating and analysing chromatin immunoprecipitation followed by sequencing (ChIP-seq) data Bioinformatics methods, models and applications to disease Alex Essebier ChIP-seq experiment To
Nature Genetics: doi: /ng Supplementary Figure 1
 Supplementary Figure 1 Expression deviation of the genes mapped to gene-wise recurrent mutations in the TCGA breast cancer cohort (top) and the TCGA lung cancer cohort (bottom). For each gene (each pair
Supplementary Figure 1 Expression deviation of the genes mapped to gene-wise recurrent mutations in the TCGA breast cancer cohort (top) and the TCGA lung cancer cohort (bottom). For each gene (each pair
ChIP-seq hands-on. Iros Barozzi, Campus IFOM-IEO (Milan) Saverio Minucci, Gioacchino Natoli Labs
 ChIP-seq hands-on Iros Barozzi, Campus IFOM-IEO (Milan) Saverio Minucci, Gioacchino Natoli Labs Main goals Becoming familiar with essential tools and formats Visualizing and contextualizing raw data Understand
ChIP-seq hands-on Iros Barozzi, Campus IFOM-IEO (Milan) Saverio Minucci, Gioacchino Natoli Labs Main goals Becoming familiar with essential tools and formats Visualizing and contextualizing raw data Understand
Part-II: Statistical analysis of ChIP-seq data
 Part-II: Statistical analysis of ChIP-seq data Outline ChIP-seq data, features, detailed modeling aspects (today). Other ChIP-seq related problems - overview (next lecture). IDR (next lecture) Stat 877
Part-II: Statistical analysis of ChIP-seq data Outline ChIP-seq data, features, detailed modeling aspects (today). Other ChIP-seq related problems - overview (next lecture). IDR (next lecture) Stat 877
Patterns of Histone Methylation and Chromatin Organization in Grapevine Leaf. Rachel Schwope EPIGEN May 24-27, 2016
 Patterns of Histone Methylation and Chromatin Organization in Grapevine Leaf Rachel Schwope EPIGEN May 24-27, 2016 What does H3K4 methylation do? Plant of interest: Vitis vinifera Culturally important
Patterns of Histone Methylation and Chromatin Organization in Grapevine Leaf Rachel Schwope EPIGEN May 24-27, 2016 What does H3K4 methylation do? Plant of interest: Vitis vinifera Culturally important
Breast cancer. Risk factors you cannot change include: Treatment Plan Selection. Inferring Transcriptional Module from Breast Cancer Profile Data
 Breast cancer Inferring Transcriptional Module from Breast Cancer Profile Data Breast Cancer and Targeted Therapy Microarray Profile Data Inferring Transcriptional Module Methods CSC 177 Data Warehousing
Breast cancer Inferring Transcriptional Module from Breast Cancer Profile Data Breast Cancer and Targeted Therapy Microarray Profile Data Inferring Transcriptional Module Methods CSC 177 Data Warehousing
Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor suppressor genes
 Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor suppressor genes Kaifu Chen 1,2,3,4,5,10, Zhong Chen 6,10, Dayong Wu 6, Lili Zhang 7, Xueqiu Lin 1,2,8,
Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor suppressor genes Kaifu Chen 1,2,3,4,5,10, Zhong Chen 6,10, Dayong Wu 6, Lili Zhang 7, Xueqiu Lin 1,2,8,
7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans.
 Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
Supplementary Figure 1 7SK ChIRP-seq is specifically RNA dependent and conserved between mice and humans. Regions targeted by the Even and Odd ChIRP probes mapped to a secondary structure model 56 of the
Discovery of Novel Human Gene Regulatory Modules from Gene Co-expression and
 Discovery of Novel Human Gene Regulatory Modules from Gene Co-expression and Promoter Motif Analysis Shisong Ma 1,2*, Michael Snyder 3, and Savithramma P Dinesh-Kumar 2* 1 School of Life Sciences, University
Discovery of Novel Human Gene Regulatory Modules from Gene Co-expression and Promoter Motif Analysis Shisong Ma 1,2*, Michael Snyder 3, and Savithramma P Dinesh-Kumar 2* 1 School of Life Sciences, University
EPIGENOMICS PROFILING SERVICES
 EPIGENOMICS PROFILING SERVICES Chromatin analysis DNA methylation analysis RNA-seq analysis Diagenode helps you uncover the mysteries of epigenetics PAGE 3 Integrative epigenomics analysis DNA methylation
EPIGENOMICS PROFILING SERVICES Chromatin analysis DNA methylation analysis RNA-seq analysis Diagenode helps you uncover the mysteries of epigenetics PAGE 3 Integrative epigenomics analysis DNA methylation
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers
 Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
Analysis of Massively Parallel Sequencing Data Application of Illumina Sequencing to the Genetics of Human Cancers Gordon Blackshields Senior Bioinformatician Source BioScience 1 To Cancer Genetics Studies
Integrated analysis of sequencing data
 Integrated analysis of sequencing data How to combine *-seq data M. Defrance, M. Thomas-Chollier, C. Herrmann, D. Puthier, J. van Helden *ChIP-seq, RNA-seq, MeDIP-seq, Transcription factor binding ChIP-seq
Integrated analysis of sequencing data How to combine *-seq data M. Defrance, M. Thomas-Chollier, C. Herrmann, D. Puthier, J. van Helden *ChIP-seq, RNA-seq, MeDIP-seq, Transcription factor binding ChIP-seq
DNA Sequence Bioinformatics Analysis with the Galaxy Platform
 DNA Sequence Bioinformatics Analysis with the Galaxy Platform University of São Paulo, Brazil 28 July - 1 August 2014 Dave Clements Johns Hopkins University Robson Francisco de Souza University of São
DNA Sequence Bioinformatics Analysis with the Galaxy Platform University of São Paulo, Brazil 28 July - 1 August 2014 Dave Clements Johns Hopkins University Robson Francisco de Souza University of São
Genome-Wide Localization of Protein-DNA Binding and Histone Modification by a Bayesian Change-Point Method with ChIP-seq Data
 Genome-Wide Localization of Protein-DNA Binding and Histone Modification by a Bayesian Change-Point Method with ChIP-seq Data Haipeng Xing, Yifan Mo, Will Liao, Michael Q. Zhang Clayton Davis and Geoffrey
Genome-Wide Localization of Protein-DNA Binding and Histone Modification by a Bayesian Change-Point Method with ChIP-seq Data Haipeng Xing, Yifan Mo, Will Liao, Michael Q. Zhang Clayton Davis and Geoffrey
High Throughput Sequence (HTS) data analysis. Lei Zhou
 High Throughput Sequence (HTS) data analysis Lei Zhou (leizhou@ufl.edu) High Throughput Sequence (HTS) data analysis 1. Representation of HTS data. 2. Visualization of HTS data. 3. Discovering genomic
High Throughput Sequence (HTS) data analysis Lei Zhou (leizhou@ufl.edu) High Throughput Sequence (HTS) data analysis 1. Representation of HTS data. 2. Visualization of HTS data. 3. Discovering genomic
Session 6: Integration of epigenetic data. Peter J Park Department of Biomedical Informatics Harvard Medical School July 18-19, 2016
 Session 6: Integration of epigenetic data Peter J Park Department of Biomedical Informatics Harvard Medical School July 18-19, 2016 Utilizing complimentary datasets Frequent mutations in chromatin regulators
Session 6: Integration of epigenetic data Peter J Park Department of Biomedical Informatics Harvard Medical School July 18-19, 2016 Utilizing complimentary datasets Frequent mutations in chromatin regulators
Genome-wide Association Studies (GWAS) Pasieka, Science Photo Library
 Lecture 5 Genome-wide Association Studies (GWAS) Pasieka, Science Photo Library Chi-squared test to evaluate whether the odds ratio is different from 1. Corrected for multiple testing Source: wikipedia.org
Lecture 5 Genome-wide Association Studies (GWAS) Pasieka, Science Photo Library Chi-squared test to evaluate whether the odds ratio is different from 1. Corrected for multiple testing Source: wikipedia.org
Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression. Chromatin Array of nuc
 Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression Chromatin Array of nuc 1 Transcriptional control in Eukaryotes: Chromatin undergoes structural changes
Transcriptional control in Eukaryotes: (chapter 13 pp276) Chromatin structure affects gene expression Chromatin Array of nuc 1 Transcriptional control in Eukaryotes: Chromatin undergoes structural changes
Supplementary Figure 1 IL-27 IL
 Tim-3 Supplementary Figure 1 Tc0 49.5 0.6 Tc1 63.5 0.84 Un 49.8 0.16 35.5 0.16 10 4 61.2 5.53 10 3 64.5 5.66 10 2 10 1 10 0 31 2.22 10 0 10 1 10 2 10 3 10 4 IL-10 28.2 1.69 IL-27 Supplementary Figure 1.
Tim-3 Supplementary Figure 1 Tc0 49.5 0.6 Tc1 63.5 0.84 Un 49.8 0.16 35.5 0.16 10 4 61.2 5.53 10 3 64.5 5.66 10 2 10 1 10 0 31 2.22 10 0 10 1 10 2 10 3 10 4 IL-10 28.2 1.69 IL-27 Supplementary Figure 1.
Raymond Auerbach PhD Candidate, Yale University Gerstein and Snyder Labs August 30, 2012
 Elucidating Transcriptional Regulation at Multiple Scales Using High-Throughput Sequencing, Data Integration, and Computational Methods Raymond Auerbach PhD Candidate, Yale University Gerstein and Snyder
Elucidating Transcriptional Regulation at Multiple Scales Using High-Throughput Sequencing, Data Integration, and Computational Methods Raymond Auerbach PhD Candidate, Yale University Gerstein and Snyder
EXPression ANalyzer and DisplayER
 EXPression ANalyzer and DisplayER Tom Hait Aviv Steiner Igor Ulitsky Chaim Linhart Amos Tanay Seagull Shavit Rani Elkon Adi Maron-Katz Dorit Sagir Eyal David Roded Sharan Israel Steinfeld Yossi Shiloh
EXPression ANalyzer and DisplayER Tom Hait Aviv Steiner Igor Ulitsky Chaim Linhart Amos Tanay Seagull Shavit Rani Elkon Adi Maron-Katz Dorit Sagir Eyal David Roded Sharan Israel Steinfeld Yossi Shiloh
STAT1 regulates microrna transcription in interferon γ stimulated HeLa cells
 CAMDA 2009 October 5, 2009 STAT1 regulates microrna transcription in interferon γ stimulated HeLa cells Guohua Wang 1, Yadong Wang 1, Denan Zhang 1, Mingxiang Teng 1,2, Lang Li 2, and Yunlong Liu 2 Harbin
CAMDA 2009 October 5, 2009 STAT1 regulates microrna transcription in interferon γ stimulated HeLa cells Guohua Wang 1, Yadong Wang 1, Denan Zhang 1, Mingxiang Teng 1,2, Lang Li 2, and Yunlong Liu 2 Harbin
CTCF-Mediated Functional Chromatin Interactome in Pluripotent Cells
 SUPPLEMENTARY INFORMATION CTCF-Mediated Functional Chromatin Interactome in Pluripotent Cells Lusy Handoko 1,*, Han Xu 1,*, Guoliang Li 1,*, Chew Yee Ngan 1, Elaine Chew 1, Marie Schnapp 1, Charlie Wah
SUPPLEMENTARY INFORMATION CTCF-Mediated Functional Chromatin Interactome in Pluripotent Cells Lusy Handoko 1,*, Han Xu 1,*, Guoliang Li 1,*, Chew Yee Ngan 1, Elaine Chew 1, Marie Schnapp 1, Charlie Wah
A Practical Guide to Integrative Genomics by RNA-seq and ChIP-seq Analysis
 A Practical Guide to Integrative Genomics by RNA-seq and ChIP-seq Analysis Jian Xu, Ph.D. Children s Research Institute, UTSW Introduction Outline Overview of genomic and next-gen sequencing technologies
A Practical Guide to Integrative Genomics by RNA-seq and ChIP-seq Analysis Jian Xu, Ph.D. Children s Research Institute, UTSW Introduction Outline Overview of genomic and next-gen sequencing technologies
Heintzman, ND, Stuart, RK, Hon, G, Fu, Y, Ching, CW, Hawkins, RD, Barrera, LO, Van Calcar, S, Qu, C, Ching, KA, Wang, W, Weng, Z, Green, RD,
 Heintzman, ND, Stuart, RK, Hon, G, Fu, Y, Ching, CW, Hawkins, RD, Barrera, LO, Van Calcar, S, Qu, C, Ching, KA, Wang, W, Weng, Z, Green, RD, Crawford, GE, Ren, B (2007) Distinct and predictive chromatin
Heintzman, ND, Stuart, RK, Hon, G, Fu, Y, Ching, CW, Hawkins, RD, Barrera, LO, Van Calcar, S, Qu, C, Ching, KA, Wang, W, Weng, Z, Green, RD, Crawford, GE, Ren, B (2007) Distinct and predictive chromatin
Nature Genetics: doi: /ng Supplementary Figure 1. Assessment of sample purity and quality.
 Supplementary Figure 1 Assessment of sample purity and quality. (a) Hematoxylin and eosin staining of formaldehyde-fixed, paraffin-embedded sections from a human testis biopsy collected concurrently with
Supplementary Figure 1 Assessment of sample purity and quality. (a) Hematoxylin and eosin staining of formaldehyde-fixed, paraffin-embedded sections from a human testis biopsy collected concurrently with
Not IN Our Genes - A Different Kind of Inheritance.! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014
 Not IN Our Genes - A Different Kind of Inheritance! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014 Epigenetics in Mainstream Media Epigenetics *Current definition:
Not IN Our Genes - A Different Kind of Inheritance! Christopher Phiel, Ph.D. University of Colorado Denver Mini-STEM School February 4, 2014 Epigenetics in Mainstream Media Epigenetics *Current definition:
RNA-seq Introduction
 RNA-seq Introduction DNA is the same in all cells but which RNAs that is present is different in all cells There is a wide variety of different functional RNAs Which RNAs (and sometimes then translated
RNA-seq Introduction DNA is the same in all cells but which RNAs that is present is different in all cells There is a wide variety of different functional RNAs Which RNAs (and sometimes then translated
Statistical analysis of ChIP-seq data
 Statistical analysis of ChIP-seq data Sündüz Keleş Department of Statistics Department of Biostatistics and Medical Informatics University of Wisconsin, Madison Feb 9, 2012 BMI 776 (Spring 12) 02/09/2012
Statistical analysis of ChIP-seq data Sündüz Keleş Department of Statistics Department of Biostatistics and Medical Informatics University of Wisconsin, Madison Feb 9, 2012 BMI 776 (Spring 12) 02/09/2012
Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types.
 Supplementary Figure 1 Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types. (a) Pearson correlation heatmap among open chromatin profiles of different
Supplementary Figure 1 Comparison of open chromatin regions between dentate granule cells and other tissues and neural cell types. (a) Pearson correlation heatmap among open chromatin profiles of different
Statistical Assessment of the Global Regulatory Role of Histone. Acetylation in Saccharomyces cerevisiae. (Support Information)
 Statistical Assessment of the Global Regulatory Role of Histone Acetylation in Saccharomyces cerevisiae (Support Information) Authors: Guo-Cheng Yuan, Ping Ma, Wenxuan Zhong and Jun S. Liu Linear Relationship
Statistical Assessment of the Global Regulatory Role of Histone Acetylation in Saccharomyces cerevisiae (Support Information) Authors: Guo-Cheng Yuan, Ping Ma, Wenxuan Zhong and Jun S. Liu Linear Relationship
Histones modifications and variants
 Histones modifications and variants Dr. Institute of Molecular Biology, Johannes Gutenberg University, Mainz www.imb.de Lecture Objectives 1. Chromatin structure and function Chromatin and cell state Nucleosome
Histones modifications and variants Dr. Institute of Molecular Biology, Johannes Gutenberg University, Mainz www.imb.de Lecture Objectives 1. Chromatin structure and function Chromatin and cell state Nucleosome
MIR retrotransposon sequences provide insulators to the human genome
 Supplementary Information: MIR retrotransposon sequences provide insulators to the human genome Jianrong Wang, Cristina Vicente-García, Davide Seruggia, Eduardo Moltó, Ana Fernandez- Miñán, Ana Neto, Elbert
Supplementary Information: MIR retrotransposon sequences provide insulators to the human genome Jianrong Wang, Cristina Vicente-García, Davide Seruggia, Eduardo Moltó, Ana Fernandez- Miñán, Ana Neto, Elbert
Exploring chromatin regulation by ChIP-Sequencing
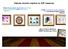 Exploring chromatin regulation by ChIP-Sequencing From datasets quality assessment, enrichment patterns identification and multi-profiles integration to the reconstitution of gene regulatory wires describing
Exploring chromatin regulation by ChIP-Sequencing From datasets quality assessment, enrichment patterns identification and multi-profiles integration to the reconstitution of gene regulatory wires describing
cis-regulatory enrichment analysis in human, mouse and fly
 cis-regulatory enrichment analysis in human, mouse and fly Zeynep Kalender Atak, PhD Laboratory of Computational Biology VIB-KU Leuven Center for Brain & Disease Research Laboratory of Computational Biology
cis-regulatory enrichment analysis in human, mouse and fly Zeynep Kalender Atak, PhD Laboratory of Computational Biology VIB-KU Leuven Center for Brain & Disease Research Laboratory of Computational Biology
Nature Structural & Molecular Biology: doi: /nsmb.2419
 Supplementary Figure 1 Mapped sequence reads and nucleosome occupancies. (a) Distribution of sequencing reads on the mouse reference genome for chromosome 14 as an example. The number of reads in a 1 Mb
Supplementary Figure 1 Mapped sequence reads and nucleosome occupancies. (a) Distribution of sequencing reads on the mouse reference genome for chromosome 14 as an example. The number of reads in a 1 Mb
Recent advances in ChIP-seq analysis: from quality management to whole-genome annotation
 Briefings in Bioinformatics Advance Access published March 15, 2016 Briefings in Bioinformatics, 2016, 1 12 doi: 10.1093/bib/bbw023 Paper Recent advances in ChIP-seq analysis: from quality management to
Briefings in Bioinformatics Advance Access published March 15, 2016 Briefings in Bioinformatics, 2016, 1 12 doi: 10.1093/bib/bbw023 Paper Recent advances in ChIP-seq analysis: from quality management to
An epigenetic approach to understanding (and predicting?) environmental effects on gene expression
 www.collaslab.com An epigenetic approach to understanding (and predicting?) environmental effects on gene expression Philippe Collas University of Oslo Institute of Basic Medical Sciences Stem Cell Epigenetics
www.collaslab.com An epigenetic approach to understanding (and predicting?) environmental effects on gene expression Philippe Collas University of Oslo Institute of Basic Medical Sciences Stem Cell Epigenetics
Supervised Learner for the Prediction of Hi-C Interaction Counts and Determination of Influential Features. Tyler Yue Lab
 Supervised Learner for the Prediction of Hi-C Interaction Counts and Determination of Influential Features Tyler Derr @ Yue Lab tsd5037@psu.edu Background Hi-C is a chromosome conformation capture (3C)
Supervised Learner for the Prediction of Hi-C Interaction Counts and Determination of Influential Features Tyler Derr @ Yue Lab tsd5037@psu.edu Background Hi-C is a chromosome conformation capture (3C)
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Heatmap of GO terms for differentially expressed genes. The terms were hierarchically clustered using the GO term enrichment beta. Darker red, higher positive
Supplementary Figures Supplementary Figure 1. Heatmap of GO terms for differentially expressed genes. The terms were hierarchically clustered using the GO term enrichment beta. Darker red, higher positive
Table S1. Total and mapped reads produced for each ChIP-seq sample
 Tale S1. Total and mapped reads produced for each ChIP-seq sample Sample Total Reads Mapped Reads Col- H3K27me3 rep1 125662 1334323 (85.76%) Col- H3K27me3 rep2 9176437 7986731 (87.4%) atmi1a//c H3K27m3
Tale S1. Total and mapped reads produced for each ChIP-seq sample Sample Total Reads Mapped Reads Col- H3K27me3 rep1 125662 1334323 (85.76%) Col- H3K27me3 rep2 9176437 7986731 (87.4%) atmi1a//c H3K27m3
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature16931 Contents 1 Nomenclature 3 2 Fertility of homozygous B6 H/H and heterozygous B6 B6/H mice 3 3 DSB hotspot maps 3 3.1 DSB hotspot caller............................... 3 3.2 Calling
doi:10.1038/nature16931 Contents 1 Nomenclature 3 2 Fertility of homozygous B6 H/H and heterozygous B6 B6/H mice 3 3 DSB hotspot maps 3 3.1 DSB hotspot caller............................... 3 3.2 Calling
cn.mops - Mixture of Poissons for CNV detection in NGS data Günter Klambauer Institute of Bioinformatics, Johannes Kepler University Linz
 Software Manual Institute of Bioinformatics, Johannes Kepler University Linz cn.mops - Mixture of Poissons for CNV detection in NGS data Günter Klambauer Institute of Bioinformatics, Johannes Kepler University
Software Manual Institute of Bioinformatics, Johannes Kepler University Linz cn.mops - Mixture of Poissons for CNV detection in NGS data Günter Klambauer Institute of Bioinformatics, Johannes Kepler University
Nature Immunology: doi: /ni Supplementary Figure 1. Characteristics of SEs in T reg and T conv cells.
 Supplementary Figure 1 Characteristics of SEs in T reg and T conv cells. (a) Patterns of indicated transcription factor-binding at SEs and surrounding regions in T reg and T conv cells. Average normalized
Supplementary Figure 1 Characteristics of SEs in T reg and T conv cells. (a) Patterns of indicated transcription factor-binding at SEs and surrounding regions in T reg and T conv cells. Average normalized
GeneOverlap: An R package to test and visualize
 GeneOverlap: An R package to test and visualize gene overlaps Li Shen Contact: li.shen@mssm.edu or shenli.sam@gmail.com Icahn School of Medicine at Mount Sinai New York, New York http://shenlab-sinai.github.io/shenlab-sinai/
GeneOverlap: An R package to test and visualize gene overlaps Li Shen Contact: li.shen@mssm.edu or shenli.sam@gmail.com Icahn School of Medicine at Mount Sinai New York, New York http://shenlab-sinai.github.io/shenlab-sinai/
Yue Wei 1, Rui Chen 2, Carlos E. Bueso-Ramos 3, Hui Yang 1, and Guillermo Garcia-Manero 1
 Genome-wide CHIP-Seq Analysis of Histone Methylation Reveals Modulators of NF- B Signaling And the Histone Demethylase JMJD3 Implicated in Myelodysplastic Syndrome Yue Wei 1, Rui Chen 2, Carlos E. Bueso-Ramos
Genome-wide CHIP-Seq Analysis of Histone Methylation Reveals Modulators of NF- B Signaling And the Histone Demethylase JMJD3 Implicated in Myelodysplastic Syndrome Yue Wei 1, Rui Chen 2, Carlos E. Bueso-Ramos
Package NarrowPeaks. August 3, Version Date Type Package
 Package NarrowPeaks August 3, 2013 Version 1.5.0 Date 2013-02-13 Type Package Title Analysis of Variation in ChIP-seq using Functional PCA Statistics Author Pedro Madrigal , with contributions
Package NarrowPeaks August 3, 2013 Version 1.5.0 Date 2013-02-13 Type Package Title Analysis of Variation in ChIP-seq using Functional PCA Statistics Author Pedro Madrigal , with contributions
SUPPLEMENTAL INFORMATION
 SUPPLEMENTAL INFORMATION GO term analysis of differentially methylated SUMIs. GO term analysis of the 458 SUMIs with the largest differential methylation between human and chimp shows that they are more
SUPPLEMENTAL INFORMATION GO term analysis of differentially methylated SUMIs. GO term analysis of the 458 SUMIs with the largest differential methylation between human and chimp shows that they are more
TITLE: Identification of Estrogen Receptor Beta Binding Sites in the Human Genomes
 AD Award Number: W81XWH-09-1-0297 TITLE: Identification of Estrogen Receptor Beta Binding Sites in the Human Genomes PRINCIPAL INVESTIGATOR: Thien Le CONTRACTING ORGANIZATION: University of Chicago, Chicago,
AD Award Number: W81XWH-09-1-0297 TITLE: Identification of Estrogen Receptor Beta Binding Sites in the Human Genomes PRINCIPAL INVESTIGATOR: Thien Le CONTRACTING ORGANIZATION: University of Chicago, Chicago,
Chip Seq Peak Calling in Galaxy
 Chip Seq Peak Calling in Galaxy Chris Seward PowerPoint by Pei-Chen Peng Chip-Seq Peak Calling in Galaxy Chris Seward 2018 1 Introduction This goals of the lab are as follows: 1. Gain experience using
Chip Seq Peak Calling in Galaxy Chris Seward PowerPoint by Pei-Chen Peng Chip-Seq Peak Calling in Galaxy Chris Seward 2018 1 Introduction This goals of the lab are as follows: 1. Gain experience using
Differential peak calling of ChIP-seq signals with replicates with THOR
 Published online 2 August 2016 Nucleic Acids Research, 2016, Vol. 44, No. 20 e153 doi: 10.1093/nar/gkw680 Differential peak calling of ChIP-seq signals with replicates with THOR Manuel Allhoff 1,2,3, Kristin
Published online 2 August 2016 Nucleic Acids Research, 2016, Vol. 44, No. 20 e153 doi: 10.1093/nar/gkw680 Differential peak calling of ChIP-seq signals with replicates with THOR Manuel Allhoff 1,2,3, Kristin
Assignment 5: Integrative epigenomics analysis
 Assignment 5: Integrative epigenomics analysis Due date: Friday, 2/24 10am. Note: no late assignments will be accepted. Introduction CpG islands (CGIs) are important regulatory regions in the genome. What
Assignment 5: Integrative epigenomics analysis Due date: Friday, 2/24 10am. Note: no late assignments will be accepted. Introduction CpG islands (CGIs) are important regulatory regions in the genome. What
ChromHMM Tutorial. Jason Ernst Assistant Professor University of California, Los Angeles
 ChromHMM Tutorial Jason Ernst Assistant Professor University of California, Los Angeles Talk Outline Chromatin states analysis and ChromHMM Accessing chromatin state annotations for ENCODE2 and Roadmap
ChromHMM Tutorial Jason Ernst Assistant Professor University of California, Los Angeles Talk Outline Chromatin states analysis and ChromHMM Accessing chromatin state annotations for ENCODE2 and Roadmap
Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63.
 Supplementary Figure Legends Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63. A. Screenshot of the UCSC genome browser from normalized RNAPII and RNA-seq ChIP-seq data
Supplementary Figure Legends Supplementary Figure S1. Gene expression analysis of epidermal marker genes and TP63. A. Screenshot of the UCSC genome browser from normalized RNAPII and RNA-seq ChIP-seq data
Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes.
 Supplementary Figure 1 Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes. (a,b) Values of coefficients associated with genomic features, separately
Supplementary Figure 1 Relationship between genomic features and distributions of RS1 and RS3 rearrangements in breast cancer genomes. (a,b) Values of coefficients associated with genomic features, separately
Statistical analysis of RIM data (retroviral insertional mutagenesis) Bioinformatics and Statistics The Netherlands Cancer Institute Amsterdam
 Statistical analysis of RIM data (retroviral insertional mutagenesis) Lodewyk Wessels Bioinformatics and Statistics The Netherlands Cancer Institute Amsterdam Viral integration Viral integration Viral
Statistical analysis of RIM data (retroviral insertional mutagenesis) Lodewyk Wessels Bioinformatics and Statistics The Netherlands Cancer Institute Amsterdam Viral integration Viral integration Viral
Gene expression analysis. Roadmap. Microarray technology: how it work Applications: what can we do with it Preprocessing: Classification Clustering
 Gene expression analysis Roadmap Microarray technology: how it work Applications: what can we do with it Preprocessing: Image processing Data normalization Classification Clustering Biclustering 1 Gene
Gene expression analysis Roadmap Microarray technology: how it work Applications: what can we do with it Preprocessing: Image processing Data normalization Classification Clustering Biclustering 1 Gene
cn.mops - Mixture of Poissons for CNV detection in NGS data Günter Klambauer Institute of Bioinformatics, Johannes Kepler University Linz
 Software Manual Institute of Bioinformatics, Johannes Kepler University Linz cn.mops - Mixture of Poissons for CNV detection in NGS data Günter Klambauer Institute of Bioinformatics, Johannes Kepler University
Software Manual Institute of Bioinformatics, Johannes Kepler University Linz cn.mops - Mixture of Poissons for CNV detection in NGS data Günter Klambauer Institute of Bioinformatics, Johannes Kepler University
Supplementary Figure 1. Efficiency of Mll4 deletion and its effect on T cell populations in the periphery. Nature Immunology: doi: /ni.
 Supplementary Figure 1 Efficiency of Mll4 deletion and its effect on T cell populations in the periphery. Expression of Mll4 floxed alleles (16-19) in naive CD4 + T cells isolated from lymph nodes and
Supplementary Figure 1 Efficiency of Mll4 deletion and its effect on T cell populations in the periphery. Expression of Mll4 floxed alleles (16-19) in naive CD4 + T cells isolated from lymph nodes and
DNA-seq Bioinformatics Analysis: Copy Number Variation
 DNA-seq Bioinformatics Analysis: Copy Number Variation Elodie Girard elodie.girard@curie.fr U900 institut Curie, INSERM, Mines ParisTech, PSL Research University Paris, France NGS Applications 5C HiC DNA-seq
DNA-seq Bioinformatics Analysis: Copy Number Variation Elodie Girard elodie.girard@curie.fr U900 institut Curie, INSERM, Mines ParisTech, PSL Research University Paris, France NGS Applications 5C HiC DNA-seq
Introduction to Systems Biology of Cancer Lecture 2
 Introduction to Systems Biology of Cancer Lecture 2 Gustavo Stolovitzky IBM Research Icahn School of Medicine at Mt Sinai DREAM Challenges High throughput measurements: The age of omics Systems Biology
Introduction to Systems Biology of Cancer Lecture 2 Gustavo Stolovitzky IBM Research Icahn School of Medicine at Mt Sinai DREAM Challenges High throughput measurements: The age of omics Systems Biology
Transcript-indexed ATAC-seq for immune profiling
 Transcript-indexed ATAC-seq for immune profiling Technical Journal Club 22 nd of May 2018 Christina Müller Nature Methods, Vol.10 No.12, 2013 Nature Biotechnology, Vol.32 No.7, 2014 Nature Medicine, Vol.24,
Transcript-indexed ATAC-seq for immune profiling Technical Journal Club 22 nd of May 2018 Christina Müller Nature Methods, Vol.10 No.12, 2013 Nature Biotechnology, Vol.32 No.7, 2014 Nature Medicine, Vol.24,
Mechanisms of alternative splicing regulation
 Mechanisms of alternative splicing regulation The number of mechanisms that are known to be involved in splicing regulation approximates the number of splicing decisions that have been analyzed in detail.
Mechanisms of alternative splicing regulation The number of mechanisms that are known to be involved in splicing regulation approximates the number of splicing decisions that have been analyzed in detail.
Dominic J Smiraglia, PhD Department of Cancer Genetics. DNA methylation in prostate cancer
 Dominic J Smiraglia, PhD Department of Cancer Genetics DNA methylation in prostate cancer Overarching theme Epigenetic regulation allows the genome to be responsive to the environment Sets the tone for
Dominic J Smiraglia, PhD Department of Cancer Genetics DNA methylation in prostate cancer Overarching theme Epigenetic regulation allows the genome to be responsive to the environment Sets the tone for
The Insulator Binding Protein CTCF Positions 20 Nucleosomes around Its Binding Sites across the Human Genome
 The Insulator Binding Protein CTCF Positions 20 Nucleosomes around Its Binding Sites across the Human Genome Yutao Fu 1, Manisha Sinha 2,3, Craig L. Peterson 3, Zhiping Weng 1,4,5 * 1 Bioinformatics Program,
The Insulator Binding Protein CTCF Positions 20 Nucleosomes around Its Binding Sites across the Human Genome Yutao Fu 1, Manisha Sinha 2,3, Craig L. Peterson 3, Zhiping Weng 1,4,5 * 1 Bioinformatics Program,
Nature Biotechnology: doi: /nbt.1904
 Supplementary Information Comparison between assembly-based SV calls and array CGH results Genome-wide array assessment of copy number changes, such as array comparative genomic hybridization (acgh), is
Supplementary Information Comparison between assembly-based SV calls and array CGH results Genome-wide array assessment of copy number changes, such as array comparative genomic hybridization (acgh), is
Nature Immunology: doi: /ni Supplementary Figure 1. Transcriptional program of the TE and MP CD8 + T cell subsets.
 Supplementary Figure 1 Transcriptional program of the TE and MP CD8 + T cell subsets. (a) Comparison of gene expression of TE and MP CD8 + T cell subsets by microarray. Genes that are 1.5-fold upregulated
Supplementary Figure 1 Transcriptional program of the TE and MP CD8 + T cell subsets. (a) Comparison of gene expression of TE and MP CD8 + T cell subsets by microarray. Genes that are 1.5-fold upregulated
Single-strand DNA library preparation improves sequencing of formalin-fixed and paraffin-embedded (FFPE) cancer DNA
 www.impactjournals.com/oncotarget/ Oncotarget, Supplementary Materials 2016 Single-strand DNA library preparation improves sequencing of formalin-fixed and paraffin-embedded (FFPE) DNA Supplementary Materials
www.impactjournals.com/oncotarget/ Oncotarget, Supplementary Materials 2016 Single-strand DNA library preparation improves sequencing of formalin-fixed and paraffin-embedded (FFPE) DNA Supplementary Materials
Measuring DNA Methylation with the MinION. Winston Timp Department of Biomedical Engineering Johns Hopkins University 12/1/16
 Measuring DNA Methylation with the MinION Winston Timp Department of Biomedical Engineering Johns Hopkins University 12/1/16 Epigenetics: Modern Modern Definition of epigenetics involves heritable changes
Measuring DNA Methylation with the MinION Winston Timp Department of Biomedical Engineering Johns Hopkins University 12/1/16 Epigenetics: Modern Modern Definition of epigenetics involves heritable changes
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. Pan-cancer analysis of global and local DNA methylation variation a) Variations in global DNA methylation are shown as measured by averaging the genome-wide
Supplementary Figures Supplementary Figure 1. Pan-cancer analysis of global and local DNA methylation variation a) Variations in global DNA methylation are shown as measured by averaging the genome-wide
Genomic structural variation
 Genomic structural variation Mario Cáceres The new genomic variation DNA sequence differs across individuals much more than researchers had suspected through structural changes A huge amount of structural
Genomic structural variation Mario Cáceres The new genomic variation DNA sequence differs across individuals much more than researchers had suspected through structural changes A huge amount of structural
Genome-wide relationship between histone H3 lysine 4 mono- and tri-methylation and transcription factor binding
 Letter Genome-wide relationship between histone H3 lysine 4 mono- and tri-methylation and transcription factor binding A. Gordon Robertson, 1,4 Mikhail Bilenky, 1,4 Angela Tam, 1 Yongjun Zhao, 1 Thomas
Letter Genome-wide relationship between histone H3 lysine 4 mono- and tri-methylation and transcription factor binding A. Gordon Robertson, 1,4 Mikhail Bilenky, 1,4 Angela Tam, 1 Yongjun Zhao, 1 Thomas
Figure S1, Beyer et al.
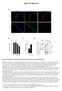 Figure S1, eyer et al. Pax7 Myogenin si sitrl Hoechst T = 72h 14 1.8.6.4.2 12 1 8 6 4 2 24h 48h 96h diff. sitrl siset1 212 72h diff. b1 td r t Se km MyH Vinculin Myogenin β-ctin Vinculin MW b1 ka td r
Figure S1, eyer et al. Pax7 Myogenin si sitrl Hoechst T = 72h 14 1.8.6.4.2 12 1 8 6 4 2 24h 48h 96h diff. sitrl siset1 212 72h diff. b1 td r t Se km MyH Vinculin Myogenin β-ctin Vinculin MW b1 ka td r
REVIEWERS' COMMENTS: Reviewer #1 (Remarks to the Author):
 Editorial Note: this manuscript has been previously reviewed at another journal that is not operating a transparent peer review scheme. This document only contains reviewer comments and rebuttal letters
Editorial Note: this manuscript has been previously reviewed at another journal that is not operating a transparent peer review scheme. This document only contains reviewer comments and rebuttal letters
ChipSeq. Technique and science. The genome wide dynamics of the binding of ldb1 complexes during erythroid differentiation
 Center for Biomics ChipSeq Technique and science The genome wide dynamics of the binding of ldb1 complexes during erythroid differentiation Wilfred van IJcken Sequencing Seminar Illumina September 8 Scheme
Center for Biomics ChipSeq Technique and science The genome wide dynamics of the binding of ldb1 complexes during erythroid differentiation Wilfred van IJcken Sequencing Seminar Illumina September 8 Scheme
Allelic reprogramming of the histone modification H3K4me3 in early mammalian development
 Allelic reprogramming of the histone modification H3K4me3 in early mammalian development 张戈 Method and material STAR ChIP seq (small-scale TELP-assisted rapid ChIP seq) 200 mouse embryonic stem cells PWK/PhJ
Allelic reprogramming of the histone modification H3K4me3 in early mammalian development 张戈 Method and material STAR ChIP seq (small-scale TELP-assisted rapid ChIP seq) 200 mouse embryonic stem cells PWK/PhJ
ChIPSeq. Technique and science. The genome wide dynamics of the binding of ldb1 complexes during erythroid differentiation
 Center for Biomics ChIPSeq Technique and science The genome wide dynamics of the binding of ldb1 complexes during erythroid differentiation Wilfred van IJcken EU Sequencing Seminar Illumina June 16 Genomics
Center for Biomics ChIPSeq Technique and science The genome wide dynamics of the binding of ldb1 complexes during erythroid differentiation Wilfred van IJcken EU Sequencing Seminar Illumina June 16 Genomics
Sudin Bhattacharya Institute for Integrative Toxicology
 Beyond the AHRE: the Role of Epigenomics in Gene Regulation by the AHR (or, Varied Applications of Computational Modeling in Toxicology and Ingredient Safety) Sudin Bhattacharya Institute for Integrative
Beyond the AHRE: the Role of Epigenomics in Gene Regulation by the AHR (or, Varied Applications of Computational Modeling in Toxicology and Ingredient Safety) Sudin Bhattacharya Institute for Integrative
Inferring Biological Meaning from Cap Analysis Gene Expression Data
 Inferring Biological Meaning from Cap Analysis Gene Expression Data HRYSOULA PAPADAKIS 1. Introduction This project is inspired by the recent development of the Cap analysis gene expression (CAGE) method,
Inferring Biological Meaning from Cap Analysis Gene Expression Data HRYSOULA PAPADAKIS 1. Introduction This project is inspired by the recent development of the Cap analysis gene expression (CAGE) method,
Supplemental Information. A Highly Sensitive and Robust Method. for Genome-wide 5hmC Profiling. of Rare Cell Populations
 Molecular ell, Volume 63 Supplemental Information Highly Sensitive and Robust Method for enome-wide hm Profiling of Rare ell Populations Dali Han, Xingyu Lu, lan H. Shih, Ji Nie, Qiancheng You, Meng Michelle
Molecular ell, Volume 63 Supplemental Information Highly Sensitive and Robust Method for enome-wide hm Profiling of Rare ell Populations Dali Han, Xingyu Lu, lan H. Shih, Ji Nie, Qiancheng You, Meng Michelle
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature19360 Supplementary Tables Supplementary Table 1. Number of monoclonal reads in each sample Sample Number of cells Total reads Aligned reads Monoclonal reads
SUPPLEMENTARY INFORMATION doi:10.1038/nature19360 Supplementary Tables Supplementary Table 1. Number of monoclonal reads in each sample Sample Number of cells Total reads Aligned reads Monoclonal reads
Regulation of Gene Expression in Eukaryotes
 Ch. 19 Regulation of Gene Expression in Eukaryotes BIOL 222 Differential Gene Expression in Eukaryotes Signal Cells in a multicellular eukaryotic organism genetically identical differential gene expression
Ch. 19 Regulation of Gene Expression in Eukaryotes BIOL 222 Differential Gene Expression in Eukaryotes Signal Cells in a multicellular eukaryotic organism genetically identical differential gene expression
SUPPLEMENTARY FIGURE 1: f-i
 SUPPLEMENTARY FIGURE 1: Comparisons of the biological replicates of ChIP-seq, Input and Bisulfite-seq. (a) Density plot of MeCP2 ChIP-seq genome coverage (calculated using tiled 15 bp windows) shows high
SUPPLEMENTARY FIGURE 1: Comparisons of the biological replicates of ChIP-seq, Input and Bisulfite-seq. (a) Density plot of MeCP2 ChIP-seq genome coverage (calculated using tiled 15 bp windows) shows high
Supplemental Figure S1. Tertiles of FKBP5 promoter methylation and internal regulatory region
 Supplemental Figure S1. Tertiles of FKBP5 promoter methylation and internal regulatory region methylation in relation to PSS and fetal coupling. A, PSS values for participants whose placentas showed low,
Supplemental Figure S1. Tertiles of FKBP5 promoter methylation and internal regulatory region methylation in relation to PSS and fetal coupling. A, PSS values for participants whose placentas showed low,
Analysis of the peroxisome proliferator-activated receptor-β/δ (PPARβ/δ) cistrome reveals novel co-regulatory role of ATF4
 Khozoie et al. BMC Genomics 2012, 13:665 RESEARCH ARTICLE Open Access Analysis of the peroxisome proliferator-activated receptor-β/δ (PPARβ/δ) cistrome reveals novel co-regulatory role of ATF4 Combiz Khozoie
Khozoie et al. BMC Genomics 2012, 13:665 RESEARCH ARTICLE Open Access Analysis of the peroxisome proliferator-activated receptor-β/δ (PPARβ/δ) cistrome reveals novel co-regulatory role of ATF4 Combiz Khozoie
Obstacles and challenges in the analysis of microrna sequencing data
 Obstacles and challenges in the analysis of microrna sequencing data (mirna-seq) David Humphreys Genomics core Dr Victor Chang AC 1936-1991, Pioneering Cardiothoracic Surgeon and Humanitarian The ABCs
Obstacles and challenges in the analysis of microrna sequencing data (mirna-seq) David Humphreys Genomics core Dr Victor Chang AC 1936-1991, Pioneering Cardiothoracic Surgeon and Humanitarian The ABCs
H3K4 demethylase KDM5B regulates global dynamics of transcription elongation and alternative splicing in embryonic stem cells
 Nucleic Acids Research, 2017 1 doi: 10.1093/nar/gkx251 H3K4 demethylase KDM5B regulates global dynamics of transcription elongation and alternative splicing in embryonic stem cells Runsheng He 1,2 and
Nucleic Acids Research, 2017 1 doi: 10.1093/nar/gkx251 H3K4 demethylase KDM5B regulates global dynamics of transcription elongation and alternative splicing in embryonic stem cells Runsheng He 1,2 and
PREPARED FOR: U.S. Army Medical Research and Materiel Command Fort Detrick, Maryland
 AD Award Number: W81XWH-12-1-0298 TITLE: MTHFR Functional Polymorphism C677T and Genomic Instability in the Etiology of Idiopathic Autism in Simplex Families PRINCIPAL INVESTIGATOR: Xudong Liu, PhD CONTRACTING
AD Award Number: W81XWH-12-1-0298 TITLE: MTHFR Functional Polymorphism C677T and Genomic Instability in the Etiology of Idiopathic Autism in Simplex Families PRINCIPAL INVESTIGATOR: Xudong Liu, PhD CONTRACTING
Chromosome-Wide Analysis of Parental Allele-Specific Chromatin and DNA Methylation
 MOLECULAR AND CELLULAR BIOLOGY, Apr. 2011, p. 1757 1770 Vol. 31, No. 8 0270-7306/11/$12.00 doi:10.1128/mcb.00961-10 Copyright 2011, American Society for Microbiology. All Rights Reserved. Chromosome-Wide
MOLECULAR AND CELLULAR BIOLOGY, Apr. 2011, p. 1757 1770 Vol. 31, No. 8 0270-7306/11/$12.00 doi:10.1128/mcb.00961-10 Copyright 2011, American Society for Microbiology. All Rights Reserved. Chromosome-Wide
SUPPLEMENTARY INFORMATION
 doi:.38/nature8975 SUPPLEMENTAL TEXT Unique association of HOTAIR with patient outcome To determine whether the expression of other HOX lincrnas in addition to HOTAIR can predict patient outcome, we measured
doi:.38/nature8975 SUPPLEMENTAL TEXT Unique association of HOTAIR with patient outcome To determine whether the expression of other HOX lincrnas in addition to HOTAIR can predict patient outcome, we measured
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/8/375/ra41/dc1 Supplementary Materials for Actin cytoskeletal remodeling with protrusion formation is essential for heart regeneration in Hippo-deficient mice
www.sciencesignaling.org/cgi/content/full/8/375/ra41/dc1 Supplementary Materials for Actin cytoskeletal remodeling with protrusion formation is essential for heart regeneration in Hippo-deficient mice
BWA alignment to reference transcriptome and genome. Convert transcriptome mappings back to genome space
 Whole genome sequencing Whole exome sequencing BWA alignment to reference transcriptome and genome Convert transcriptome mappings back to genome space genomes Filter on MQ, distance, Cigar string Annotate
Whole genome sequencing Whole exome sequencing BWA alignment to reference transcriptome and genome Convert transcriptome mappings back to genome space genomes Filter on MQ, distance, Cigar string Annotate
Supplemental Figure 1. Genes showing ectopic H3K9 dimethylation in this study are DNA hypermethylated in Lister et al. study.
 mc mc mc mc SUP mc mc Supplemental Figure. Genes showing ectopic HK9 dimethylation in this study are DNA hypermethylated in Lister et al. study. Representative views of genes that gain HK9m marks in their
mc mc mc mc SUP mc mc Supplemental Figure. Genes showing ectopic HK9 dimethylation in this study are DNA hypermethylated in Lister et al. study. Representative views of genes that gain HK9m marks in their
Genome-Wide Analysis of Histone Modifications in Human Endometrial Stromal Cells
 ORIGINAL RESEARCH Genome-Wide Analysis of Histone Modifications in Human Endometrial Stromal Cells Isao Tamura, Yasuyuki Ohkawa, Tetsuya Sato, Mikita Suyama, Kosuke Jozaki, Maki Okada, Lifa Lee, Ryo Maekawa,
ORIGINAL RESEARCH Genome-Wide Analysis of Histone Modifications in Human Endometrial Stromal Cells Isao Tamura, Yasuyuki Ohkawa, Tetsuya Sato, Mikita Suyama, Kosuke Jozaki, Maki Okada, Lifa Lee, Ryo Maekawa,
