AINTEGUMENTA Contributes to Organ Polarity and Regulates Growth of Lateral Organs in Combination with YABBY Genes 1
|
|
|
- Mavis Waters
- 5 years ago
- Views:
Transcription
1 AINTEGUMENTA Contributes to Organ Polarity and Regulates Growth of Lateral Organs in Combination with YABBY Genes 1 Staci Nole-Wilson 2 and Beth A. Krizek* Department of Biological Sciences, University of South Carolina, Columbia, South Carolina Lateral organs in flowering plants display polarity along their adaxial-abaxial axis with distinct cell types forming at different positions along this axis. Members of three classes of transcription factors in Arabidopsis (Arabidopsis thaliana; the Class III homeodomain/leucine zipper [HD-ZIP] proteins, KANADI proteins, and YABBY proteins) are expressed in either the adaxial or abaxial domain of organ primordia where they confer these respective identities. Little is known about the factors that act upstream of these polarity-determining genes to regulate their expression. We have investigated the relationship between AINTEGUMENTA (ANT), a gene that promotes initiation and growth of lateral organ primordia, and polarity genes. Although ant single mutants do not display any obvious defects in organ polarity, loss of ANT activity in combination with mutations in one or more YABBY genes results in polarity defects greater than those observed in the yabby mutants alone. Our results suggest that ANT acts in combination with the YABBY gene FILAMENTOUS FLOWER (FIL) to promote organ polarity by upregulating the expression of the adaxial-specifying HD-ZIP gene PHABULOSA. Furthermore, we show that ANT acts with FIL to up-regulate expression of the floral homeotic gene APETALA3. Our work defines new roles for ANT in the development of lateral organs. In flowering plants, leaves and floral organs are produced on the periphery of apical meristems. These lateral organs possess an inherent asymmetry with regard to the meristem in that their adaxial side is adjacent and close to the meristem, while their abaxial side is located further from the meristem. This asymmetry gives rise to a polarity that is readily apparent at the cellular and whole organ level and which can have important functional consequences. For example, cells within the adaxial region of a leaf are specialized for light capture, while those in the abaxial region are specialized for gas exchange. In addition, outgrowth of the leaf lamina is dependent on the juxtaposition of cells with adaxial and abaxial identities (Waites and Hudson, 1995). A similar mechanism may be responsible for the outgrowth of flattened floral organs such as sepals and petals. Members of three classes of transcription factors contribute to the establishment of adaxial and abaxial cell fates in lateral organs of Arabidopsis (Arabidopsis thaliana; for review, see Engstrom et al., 2004). Class III homeodomain/leu zipper (HD-ZIP) proteins specify 1 This work was supported by the U.S. Department of Energy (grant no. 98ER20312). 2 Present address: Department of Genetics, North Carolina State University, Raleigh, NC * Corresponding author; krizek@sc.edu; fax The author responsible for the distribution of materials integral to the finding presented in this article in accordance with the policy described in the Instructions for Authors ( is: Beth A. Krizek (krizek@sc.edu). Article, publication date, and citation information can be found at adaxial identity, while KANADI and YABBY proteins specify abaxial identity. Three HD-ZIP genes, PHABU- LOSA (PHB), PHAVOLUTA (PHV), and REVOLUTA (REV), are expressed in the adaxial domain of lateral organ primordia (McConnell et al., 2001; Emery et al., 2003). Dominant mutations in PHB or PHV result in transformation of abaxial cells to adaxial fates and the production of radially symmetric lateral organs (McConnell and Barton, 1998; McConnell et al., 2001). phb phv rev triple mutants produce just a single abaxialized and radialized cotyledon in the most severe case (Emery et al., 2003). At least three members of the KANADI gene family (KAN1, KAN2, and KAN3) redundantly specify abaxial identity (Eshed et al., 2001; Kerstetter et al., 2001). Transformation of abaxial cell types into adaxial cell types is observed with increasing severity in kan double and triple mutants, while ectopic expression of KAN genes results in the development of abaxial tissues in adaxial regions and the radialization of lateral organs (Eshed et al., 2001, 2004; Kerstetter et al., 2001). KAN1 is expressed in the abaxial domain of developing lateral organs, complementary to the expression of PHB-like genes in the adaxial domain (Kerstetter et al., 2001). Members of the YABBY gene family also contribute to the specification of abaxial identity. Three YABBY genes, FILAMENTOUS FLOWER (FIL), YABBY2 (YAB2), and YABBY3 (YAB3), are expressed in the abaxial half of all lateral organ primordia (Siegfried et al., 1999). Ectopic expression of these YABBY genes can convert some adaxial cell types into abaxial cells (Sawa et al., 1999; Siegfried et al., 1999). Although loss of both FIL and YAB3 activity does not result in conversion of abaxial cells into adaxial cells, Plant Physiology, July 2006, Vol. 141, pp , Ó 2006 American Society of Plant Biologists 977
2 Nole-Wilson and Krizek increased adaxialization of lateral organs occurs in kan1/1 kan2 plants upon loss of FIL and YAB3 activity (Eshed et al., 2004). KANADI proteins are members of the GARP family of transcriptional regulators, while YABBYproteins are zinc finger-containing proteins with an HMG box-like domain (Bowman and Smyth, 1999; Sawa et al., 1999; Eshed et al., 2001; Kerstetter et al., 2001). It has been proposed that a meristem-derived signal is responsible for the establishment of adaxial-abaxial polarity in lateral organs (for review, see Bowman et al., 2002). Cells closest to the meristem, those in the adaxial domain, perceive this signal, while cells further away do not. In response, PHB-like proteins are activated in cells of the adaxial domain. Because of the antagonism between PHB-like and KANADI genes, KANADI gene expression is subsequently inhibited in the adaxial region and becomes restricted to the abaxial domain (Eshed et al., 2004). Thus, an initial asymmetry due to the meristem-derived signal is thought to be maintained by antagonism between the PHB-like and KANADI genes and results in restriction of PHBlike gene expression to the adaxial domain and KANADI gene expression to the abaxial domain. Additionally, the mutual repression between the PHB-like and KANADI genes leads to the abaxial specific expression of the YABBY genes (Eshed et al., 2004). We are interested in the relationship between factors that promote the initiation and growth of lateral organ primordia, such as AINTEGUMENTA (ANT), and factors that act within lateral organ primordia to establish their polarity, such as PHB-like, KANADI, and YABBY proteins. ANT expression is up-regulated in leaf and flower founder cells in apical meristems and is one of the earliest markers of lateral organ specification (Elliott et al., 1996; Long and Barton, 2000). It has been suggested that YABBY genes could initially be activated by proteins that promote primordia initiation (such as ANT), while other factors act later to restrict YABBY gene expression to abaxial regions (Bowman, 2000). ANT encodes a transcription factor of the APETALA2/ ethylene-responsive element binding factor family that binds to 5#-gCAC(A/G)N(A/T)TcCC(a/g)ANG (c/t)-3# DNA sequences (Nole-Wilson and Krizek, 2000). Although we find that ANT can bind in vitro to such a sequence within the FIL and YAB3 promoters, ANT is not required for normal levels of FIL or YAB3 expression. Characterization of fil-8 ant-4 and fil-8 yab3-2 ant-4 double and triple mutants does suggest a role for ANT in the establishment of adaxial-abaxial polarity in leaves and floral organs. RESULTS ANT Binds to a Conserved Element in the FIL and YAB3 Promoters FIL and YAB3 are expressed in largely overlapping domains and share sequence similarity within an approximately 300-bp 5# regulatory region, part of which is shown in Figure 1A (Siegfried et al., 1999; Watanabe and Okada, 2003). Deletion analysis of the FIL promoter identified two cis-acting regulatory elements required for proper FIL expression (Watanabe and Okada, 2003). A region proximal to the FIL coding sequence (21,742 to 21,547) is required for expression in both adaxial and abaxial domains, while a 12-bp (21,748 to 21,737) sequence is required for the abaxialspecific expression of the FIL gene (Fig. 1A). A sequence with similarity to the ANT consensus binding site is present within the former FIL region and within the YAB3 promoter. The putative ANT binding sites in the FIL and YAB3 promoters match the in vitrodetermined ANT consensus binding site in 10 of 14 conserved positions (Fig. 1A). Gel mobility shifts revealed that ANT binds in vitro to fragments of both promoters that contain these sequences (Fig. 1B). The observed binding to either of these sites is weaker than that of ANT to the consensus binding site (BS15; Fig. 1B). Binding to the FIL and YAB3 promoter sites was competed by unlabeled BS15 but not by a mutated version of this binding site (data not shown). FIL and YAB3 Expression Is Normal in ant Flowers To probe the potential role of ANT in FIL and YAB3 regulation, we examined the expression of FIL and YAB3 in an ant mutant background. If ANT is a positive regulator of FIL and/or YAB3, we might expect FIL and YAB3 expression to be reduced in ant mutants. FIL expression was examined in ant-4 flowers by in situ hybridization. A similar level and pattern of FIL expression was observed in Landsberg erecta (Ler) and ant-4 flowers (Fig. 2, A D). YAB3 mrna was examined in Ler and ant-4 inflorescences by real-time reverse transcription (RT)-PCR. Similar levels of YAB3 mrna were present in both genotypes (Fig. 2E). These results suggest that ANT activity is not required for activation of YAB3 or FIL in flowers. fil ant and fil yab3 ant Plants Are Reduced in Size To gain insight into the relationship between ANT and the two YABBY genes, we generated fil-8 ant-4 double mutants and fil-8 yab3-2 ant-4 triple mutants. The double and triple mutant plants were dwarfed and exhibited severe alterations in organ development during both vegetative and reproductive development (Figs. 3, A D, and 5, A and B). While the leaves of yab3-2, fil-8, and ant-4 single mutants were not dramatically different in size from those of wild type (Kumaran et al., 2002), the leaves of fil-8 yab3-2, fil-8 ant-4, and fil-8 yab3-2 ant-4 were considerably smaller than those of wild type (Fig. 3, E H; Table I). The decreased leaf area of the double and triple mutants resulted from reductions in both length and width (Table I). In the triple fil-8 yab3-2 ant-4 mutant, there was a dramatic reduction in lamina expansion such that the petioles of fil-8 yab3-2 ant-4 leaves were often not clearly distinguishable from the lamina (Fig. 3, H and L). 978 Plant Physiol. Vol. 141, 2006
3 Regulation of Organ Growth and Polarity by AINTEGUMENTA Figure 1. ANT binds to a sequence within the FIL and YAB3 promoters. A, Alignment of a conserved element within the 5# regulatory region of FIL and YAB3 to the ANT consensus site. The putative ANT binding site is underlined. Nucleotides shared between the ANT consensus binding site and the FIL and YAB3 promoters are shown in bold. A putative Kruppel binding site (21,748 to 21,737), required for repression of FIL in the adaxial domain, is boxed. Numbers indicate positions relative to the start codons. B, Gel shift showing binding of ANT to the consensus binding site (BS15), the YAB3 promoter site, and the FIL promoter site. The YAB3 promoter fragment is a 106-bp sequence corresponding to nucleotides 21,561 to 21,456 and the FIL promoter fragment is a 131-bp sequence corresponding to nucleotides 21,763 to 21,633. Lanes 2 and 5 contain no protein. Increasing amounts of ANT protein are shown in lanes 3 and 4 (and 6 and 7). The same amount of ANT protein was used in lanes 1, 4, and 7. To determine the basis for the smaller leaf size in fil-8 ant-4 and fil-8 yab3-2 ant-4 plants, the size of mature leaf epidermal cells was examined using scanning electron microscopy (SEM). Epidermal cells were larger in fil-8 ant-4 and fil-8 yab3-2 ant-4 plants compared to Ler (Fig. 4, A, B, and I L). This indicates that the smaller leaf blades of the double and triple mutants are due to the presence of fewer cells. Similarly, the reduced height of the double and triple mutant plants (Fig. 5A; Table II) was due to fewer cells in the stem (data not shown). fil ant and fil yab ant Mutants Show Disruptions in Leaf Polarity A juxtaposition of adaxial and abaxial cell types is thought to be required for leaf blade expansion (for review, see Bowman, 2000; Bowman et al., 2002). To determine whether the reduced expansion of leaf blades in fil-8 ant-4 and fil-8 yab3-2 ant-4 plants might be a consequence of reduced leaf polarity, epidermal cell morphologies were examined. The adaxial and abaxial surfaces of wild-type leaves are distinct. The adaxial epidermis is flat, while the abaxial epidermis is undulating (Fig. 4, A and B). In addition, adaxial epidermal cells are fairly uniform in size, while abaxial epidermal cells are variably sized and puzzle shaped (Fig. 4, A and B). The adaxial and abaxial surfaces of ant-4 and fil-8 leaves are normal (Fig. 4, C F). The adaxial surface of fil-8 yab3-2 leaves is normal (Fig. 4G). However, the abaxial surface of fil-8 yab3-2 leaves was altered slightly from wild type, indicating a partial loss of abaxial identity (Fig. 4H; Siegfried et al., 1999). More dramatic changes in adaxial and abaxial identities were observed in fil-8 ant-4 and fil-8 yab3-2 ant-4 plants. Adaxial epidermal cells of fil-8 ant-4 leaves were variable in size and sometimes puzzle shaped, slightly resembling abaxial epidermal cells (Fig. 4I). In addition, the abaxial surface was flatter than wild type and the cells larger than wild type (Fig. 4J). This suggests a partial loss of both adaxial and abaxial identities in fil-8 ant-4 plants. Thus, at least some of the reduced growth of fil-8 ant-4 leaves may result from a loss of polarity. The more dramatic reduction in lamina expansion in the triple mutant was correlated with a more complete loss of polarity. In fil-8 yab3-2 ant-4 leaves, adaxial and abaxial epidermal cells Plant Physiol. Vol. 141,
4 Nole-Wilson and Krizek with the other identity (i.e. replacement of abaxial cell fates with adaxial identities in kan mutants). fil-8 ant-4 and fil-8 yab3-2 ant-4 leaves also exhibit alterations in their vascular patterning. Vascular tissue in wild-type leaves exhibits a reticulate pattern with minor veins branching from the major vein (Fig. 3I). There was a marked decrease in vascular branching in the leaves of fil-8 ant-4 plants (Fig. 3K). This phenotype is similar to that reported previously for yab3-1 fil-5 leaves (Siegfried et al., 1999) and fil-8 yab3-2 (Fig. 3J). fil-8 yab3-2 ant-4 leaves often have just a single vein running the length of the leaf (Fig. 3L). In some cases, one to several shorter veins branch from this central vein. fil ant and fil yab ant Mutants Show Disruptions in Floral Organ Identity and Polarity Figure 2. FIL and YAB3 expression in wild-type and ant-4 plants. A, FIL mrna is present in floral primordia and sepal primordia in this transverse Ler inflorescence section. B, FIL mrna in an ant-4 inflorescence. C, FIL expression in a stage 8 Ler flower. D, FIL expression in a stage 8 ant-4 flower. Size bars correspond to 50 mm in A to D. Bottom section, Relative expression levels (compared to ACTIN2) of YAB3 in Ler and ant-4 inflorescences. The average of two experiments is shown. The bars show SD. closely resembled each other with neither the adaxial or abaxial surface displaying its characteristic appearance (Fig. 4, K and L). Loss of both adaxial and abaxial identities distinguishes the fil ant and fil yab ant mutants from mutations in either the KAN or PHBlike genes, where there is replacement of one identity fil-8 ant-4 and fil-8 yab3-2 ant-4 plants exhibit inflorescence defects similar to those observed in fil-8 plants with the inflorescence meristem switching between the production of flowers and filaments (Sawa et al., 1999). In fil-8 ant-4, around 12 flowers were produced before the inflorescence meristem started to produce filaments (Fig. 6A). This is similar to the number of individual flowers initiated by the inflorescence meristem of fil-8 plants prior to filament production. Fewer flowers were produced prior to filament production in fil-8 yab3-2 ant-4 plants (Fig. 5J). After producing some filaments, fil-8 ant-4 and fil-8 yab3-2 ant-4 inflorescence meristems switched to producing a mixture of flower-like structures and filaments (Fig. 6). This was subsequently followed by termination of the inflorescence meristem. After termination of the primary inflorescence, secondary and axillary inflorescences grew out, resulting in the production of short and bushy fil-8 ant-4 and fil-8 yab3-2 ant-4 plants (Fig. 5). Figure 3. Wild-type, fil-8 yab3-2, fil-8 ant-4, and fil-8 yab3-2 ant-4 leaves. A to D, Mature rosettes just prior to bolting: Ler (A), fil-8 yab3-2 (B), fil-8 ant-4 (C), and fil-8 yab3-2 ant-4 (D). The pictures in A to D are taken at the same magnification. Insets in B and D show closer views. E to H, Fully expanded leaves from Ler (E), fil-8 yab3-2 (F), fil-8 ant-4 (G), and fil-8 yab3-2 ant-4 (H) plants. The pictures in E to H are taken at the same magnification. I to L, Vascular patterns of Ler (I), fil-8 yab3-2 (J), fil-8 ant-4 (K), and fil-8 yab3-2 ant-4 (L) fully expanded leaves. The pictures in I to L are taken at the same magnification. Leaves shown in E to L were from position 5 or 6 of the rosette. Size bars correspond to 1 mm in I to L. 980 Plant Physiol. Vol. 141, 2006
5 Regulation of Organ Growth and Polarity by AINTEGUMENTA Table I. Leaf size in wild-type and mutant plants The data are indicated as averages 6 SD. *, Values that are significantly different from wild type (Student s t test, P, 0.01). Leaf Area Leaf Width Leaf Length mm 2 mm mm Ler ant * * fil fil-8 yab * * * fil-8 ant * * * fil-8 yab3-2 ant * * * The flowers produced by fil-8 ant-4 and fil-8 yab3-2 ant-4 plants were much smaller than flowers of wild type, fil-8, or fil-8 yab3-2 (Fig. 5B). In addition, they exhibited loss of floral identity as demonstrated by the presence of flowers with subtending leaves and by a loss of floral organ identity. fil-8 ant-4 and fil-8 yab3-2 ant-4 flowers typically consisted of narrow, flat, green organs; filaments; and carpelloid organs (Figs. 5, G and I, and 6, B and C). Because these organs lack most recognizable features of floral organs, we examined their development and cell types by SEM to better characterize them. Because fil-8 ant-4 and fil-8 yab3-2 ant-4 flowers were quite similar, we present a detailed SEM analysis of just fil-8 ant-4 flowers. fil-8 ant-4 flowers typically produced two or three whorls of floral organs with variable numbers and positions of organs within each whorl (Fig. 6, D F). Flowers arising later on the inflorescence typically produced a fewer number of organs. The flat, outermost organs from early arising fil-8 ant-4 flowers had epidermal cells resembling those of sepals (Fig. 6G). In later-arising flowers, these organs became thinner and more pointed. SEM analysis showed that these laterarising outer whorl organs were mosaics containing both leaf-like and sepal-like cells (Fig. 6H). Filamentous organs present in the outer two whorls of fil-8 ant-4 flowers were variable in appearance. Those in the outermost whorl were typically dark green and had epidermal cells resembling those of sepals (Fig. 6I), while filaments in the second whorl were light green or white in color with more regular cells in files (Fig. 6J). In addition, filaments in the outer whorl tended to be thicker than those in the second whorl. The flat innermost organs of fil-8 ant-4 flowers resembled unfused or partially fused carpels. Style-like cells were found at the top of these organs (Fig. 6, K and L), and valve-like ovary cells were present along the rest of their length. Stigmatic papillae were sometimes present, although not necessarily at the apex of these organs (Fig. 6C). The carpel-like organs arose as distinct primordia rather than the fused ring of tissue that develops in wild-type flowers (Fig. 6D). These primordia gave rise to either distinct organs (Fig. 6E) or partially fused organs (Fig. 6F). Internal tissues present within a normal gynoecium (septum, transmitting tract, placenta, and ovules) were almost completely lacking in fil-8 ant-4 flowers (Fig. 6, B and C). In a few early arising fil-8 ant-4 flowers, narrow white organs, and/or yellow stamen-like organs were present (Figs. 5H and 6M). SEM examination indicated that petal epidermal cells were present on the surface of some of these white organs (Fig. 6N). In some cases, these organs exhibited polarity defects, as petals cells with both adaxial and abaxial morphologies were present on the abaxial surface of these organs (Fig. 6O). Epidermal cells characteristic of stamens were present on the stamen-like organs (Fig. 6, P and Q). A small amount of internal carpel tissue was occasionally present in early arising fil-8 ant-4 flowers (Fig. 6P). fil-8 yab3-2 ant-4 flowers exhibit a slightly more severe phenotype than fil-8 ant-4 flowers in that organs with Figure 4. Adaxial and abaxial surfaces of leaves from wild-type, ant-4, fil-8, fil-8 yab3-2, fil-8 ant-4, and fil-8 yab3-2 ant-4 plants. Shown are adaxial leaf surfaces of Ler (A), ant-4 (C), fil-8 (E), fil-8 yab3-2 (G), fil-8 ant-4 (I), and fil-8 yab3-2 ant-4 (K). Also shown are abaxial leaf surfaces of Ler (B), ant-4 (D), fil-8 (F), fil-8 yab3-2 (H), fil-8 ant-4 (J), and fil-8 yab3-2 ant-4 (L). Two cells with similar morphologies in K and L are noted with *. Size bars correspond to 100 mm. Plant Physiol. Vol. 141,
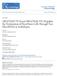
6 Nole-Wilson and Krizek Figure 5. Flowers from wild-type, ant-4, fil-8, fil-8 yab3-2, fil-8 ant-4, and fil-8 yab3-2 ant-4 plants. A, Side view of fil-8, fil-8 yab3-2, fil-8 ant-4, and fil-8 yab3-2 ant-4 plants of approximately 3 weeks of age. B, A single image showing the side views of fil-8, fil-8 yab3-2, fil-8 ant-4, and fil-8 yab3-2 ant-4 flowers (from left to right) and the relative sizes of the flowers. C, Ler flower. D, ant-4 flower. E, fil-8 flower. F, fil-8 yab3-2 flower. G, fil-8 ant-4 flower. H, Early arising fil-8 ant-4 flower. Arrow points to a staminoid organ and arrowhead points to a white petaloid organ. I, fil-8 yab3-2 ant-4 flower. J, fil-8 yab3-2 ant-4 inflorescence meristem that has initiated filament production. Arrow points to the inflorescence meristem. petal and stamen-like cells were never observed in fil-8 yab3-2 ant-4 flowers. fil-8 ant-4 and fil-8 yab3-2 ant-4 flowers have a more severe phenotype than fil-8 yab3-2 flowers (Fig. 5, F, G, and I). fil-8 yab3-2 flowers consist of radialized or flat sepal-like organs, no petals, small stamen filaments lacking anthers, and a carpel with a larger style and no replum (Siegfried et al., 1999; Kumaran et al., 2002). In comparison, fil-8 ant-4 and fil-8 yab3-2 ant-4 flowers consisted of fewer floral organs and show increased radialization of outer whorl organs and loss of carpel adaxial tissue. fil ant Flowers Show Altered Expression of Floral Organ Identity Genes Because of the greatly reduced floral organ identity in fil-8 ant-4 flowers, we examined the expression patterns of the floral homeotic genes APETALA3 (AP3) and AGAMOUS (AG) by in situ hybridization. AP3 is a B class floral homeotic gene involved in the specification of petal and stamen identities, and AG is a C function gene involved in the specification of stamen and carpel identities (Yanofsky et al., 1990; Jack et al., 1992). AP3 expression is greatly reduced or absent in young fil-8 ant-4 flowers as compared with wild type (Fig. 7, A D). When AP3 mrna was detected in young fil-8 ant-4 flowers, it was present in a normal spatial pattern (Fig. 7B). AP3 mrna was only rarely observed in organs of older fil-8 ant-4 flowers (Fig. 7, E and F). AP3 mrna was often observed in a few cells located between the inner and outer whorls of fil-8 ant-4 flowers (Fig. 7G). Weak patches of AP3 expression were rarely observed in filaments produced in place of flowers (Fig. 7H). In wild-type plants, AG mrna was first detected in the center of the floral meristem of stage 3 flowers (Fig. 7I). In fil-8 ant-4 plants, AG was misexpressed in the inflorescence meristem (Fig. 7J). AG mrna was also detected in stage one and two floral meristems, earlier than the first appearance of AG mrna in Ler flowers (data not shown). AG was expressed at high levels in the center of young fil-8 ant-4 floral meristems, similar to the pattern seen in young Ler stage 3 floral meristems (Fig. 7, K and L). In some cases, this AG expression domain was broader in stage 4 fil-8 ant-4 flowers than stage 4 Ler flowers and extended into the outermost organ primordia (Fig. 7M). In Ler flowers, AG is expressed throughout developing stamens and carpels until late stages of flower development (Fig. 7N). In older fil-8 ant-4 flowers, AG mrna was detected on the inner surface of carpel-like organs (Fig. 7O). AG mrna was also present in the center of filamentous structures produced by the inflorescence meristem (Fig. 7P). fil ant Flowers Show Reduced Floral Expression of the Adaxial Cell Fate Regulator PHB The radialization of fil-8 ant-4 and fil-8 yab3-2 ant-4 floral organs suggests that these organs have lost adaxial-abaxial polarity. To further investigate this possibility, we examined the expression of the adaxial cell fate regulator PHB in fil-8 ant-4 flowers. PHB mrna is present in the inflorescence meristem and throughout young floral meristems of Ler flowers (Fig. 8A). In stage 4 flowers, PHB mrna was detected in the center of the floral meristem and in the adaxial half of developing sepal primordia (Fig. 8B). The Table II. Height of wild-type and mutant plants The data are indicated as averages 6 SD. *, Values that are significantly different from wild type (Student s t test, P, 0.01). Height cm Ler ant fil fil-8 yab * fil-8 ant * fil-8 yab3-2 ant * 982 Plant Physiol. Vol. 141, 2006
7 Regulation of Organ Growth and Polarity by AINTEGUMENTA Figure 6. SEM analyses of fil-8 ant-4 flowers. A, fil-8 ant-4 inflorescence meristem producing a mixture of flowers (white arrow) and filamentous structures (black arrow). B, fil-8 ant-4 flower. C, fil-8 ant-4 flower. Stigmatic papillae are indicated with an arrow. D, Stage 7 fil-8 ant-4 flower showing three whorls of organ primordia. E, fil-8 ant-4 flower with two visible whorls of organs. The inner whorl contains three unfused organ primordia. F, Stage 8 fil-8 ant-4 flower with two visible whorls of organs. A black arrow points to a region of fusion between the inner whorl organ primordia. G, An outer whorl organ with sepal-like cells. H, An outer whorl organs with leaf-like (Le) and sepal-like (Se) cells. I, An outer whorl filamentous organ with sepal-like (Se) cells. J, An inner whorl filamentous organ (arrow). K, An inner whorl carpel-like organ with ovary valve-like (Ov) cells at the base and style-like (Sy) cells at the top. L, Close-up of organ shown in K. M, An early arising fil-8 ant-4 flower. A black arrow indicates a petal-like organ. N, Close-up of the petal-like organ in M showing cells with petal morphologies. O, Close-up of the organ in M. Petal cells with adaxial (black arrows) and abaxial (white arrows) morphologies are present on the abaxial surface of this fil-8 ant-4 organ. The adaxial petal epidermal cells are conical in shape with epicuticular thickenings oriented along the cone axis. The abaxial epidermal petal cells are flatter with more zigzagged epicuticular thickenings. P, Staminoid organ (black arrow) present in an early arising fil-8 ant-4 flower. The white arrow points to adaxial carpel tissue. Q, Close-up of the anther-like region of the staminoid organ in P. Size bars correspond to 20 mm in H, L, O, and Q; 50 mm in D to G, I, J, and N; 100 mm in A and K; 200 mm in M and P; and 500 mm in B and C. expression pattern and levels of PHB mrna were similar in fil-8 and ant-4 single mutants as compared with Ler (Fig. 8, C F). In fil-8 ant-4 plants, PHB mrna was typically present at lower levels in the inflorescence meristem and stage 1 and 2 floral meristems as compared with wild type (Fig. 8G). PHB mrna was usually absent from the outer whorl organs of fil-8 ant-4 flowers and was present in reduced amounts in the floral meristem of young stage 3 and 4 fil-8 ant-4 flowers (Fig. 8H). DISCUSSION ANT is an important regulator of lateral organ development. ANT expression marks cells that will leave the meristem to form lateral organs, and it is required for proper initiation and growth of lateral organs. Mutations in ANT result in the production of fewer and smaller floral organs (Elliott et al., 1996; Klucher et al., 1996). Despite its importance in lateral organ development, little is known about the genes regulated by this transcription factor. Our results here suggest that ANT acts with FIL to regulate organogenesis and to up-regulate genes establishing organ polarity and those specifying organ identity. ANT Regulates Organ Polarity While lateral organs in ant single mutants exhibit normal polarity, defects along the adaxial/abaxial axis were visible at the whole organ, cellular, and molecular level in fil ant and fil yab ant plants. fil yab ant leaves showed dramatic reductions in lamina growth and loss of both adaxial and abaxial epidermal cell identities. fil ant and fil yab ant floral organs were severely radialized with many floral organs replaced by filaments or very narrow organs. In some floral organs, adaxial cell types were found in abaxial positions. These defects are more severe than those observed in either fil or fil yab mutants. To investigate whether the role of ANT in polarity establishment involves regulation of known adaxial and/or abaxial identity factors, we examined the Plant Physiol. Vol. 141,
8 Nole-Wilson and Krizek Figure 7. AP3 and AG expression in wild-type and fil-8 ant-4 flowers. AP3 expression is shown in A to H, and AG expression is shown in I to P. Size bars correspond to 50 mm in A to P. A, AP3 mrna in a Ler stage 3 flower. B, AP3 mrna in a stage 3 fil-8 ant-4 flower. C, Arrow points to region of AP3 expression in a stage 3 fil-8 ant-4 flower. D, No AP3 mrna was detected in this fil-8 ant-4 flower. E, AP3 mrna is detected in the second and third whorls of stage 5 (left) and stage 8 (right) flowers. F, AP3 mrna is detected in a stamen-like organ of a fil-8 ant-4 flower. G, AP3 mrna is present in a few cells located between the outer and inner whorl in older fil-8 ant-4 flowers. H, AP3 mrna was rarely detected in filaments (arrow) initiated by the inflorescence meristem of fil-8 ant-4 plants. I, AG expression is first detected in a stage 3 flower in wild type. J, AG is expressed in the inflorescence meristem of this fil-8 ant-4 plant. K, AG mrna is detected in the central part of the floral meristem in a stage 3 Ler flower. L, AG expression in a stage 3 fil-8 ant-4 flower. M, AG expression in the outer organ primordia (arrow) of a stage 4 fil-8 ant-4 flower. N, AG is expressed in the stamens and carpels of a stage 8 Ler flower. O, AG mrna is detected on the adaxial surface of the central carpel-like organs in a fil-8 ant-4 flower. P, AG mrna is detected in the central region of filamentous structures (arrow) produced by a fil-8 ant-4 inflorescence meristem. expression of FIL, YAB3, and PHB in ant mutants. While the expression of FIL, YAB3, and PHB are normal in ant-4 flowers, PHB expression was reduced in fil ant double mutants. This suggests that ANT and FIL together are required for normal levels of PHB mrna. Examination of the PHB promoter revealed a sequence matching the ANT consensus binding site in 12 of 14 conserved positions including one gap (S. Nole-Wilson and R. Franks, personal communication). We were unable to detect binding of ANT to this site in vitro, suggesting that ANT is not a direct regulator of PHB expression, its role in PHB regulation involves additional factors, and/or that ANT binds to a different DNA sequence within the context of the PHB promoter. It is curious that an ANT binding site is present within a conserved region of the FIL and YAB3 promoters but that ANT is not required for FIL or YAB3 expression. Seven AINTEGUMENTA-like (AIL) genes are present within the Arabidopsis genome, and several of these genes are expressed in young floral primordia in overlapping domains with ANT (Nole- Wilson et al., 2005). Sequence conservation within the DNA-binding APETALA2 repeat regions of AIL proteins suggest that they may have similar DNA-binding specificities. It is possible that redundancy within the AIL family masks any effect of loss of ANT activity on FIL and YAB3 expression. It will be necessary to generate double, triple, and perhaps higher order mutants within members of the AIL gene family to investigate this possibility. As both adaxial and abaxial identities are partially lost in fil yab ant mutants and ANT is expressed throughout lateral organs, our results suggest that ANT is a positive regulator of both genes specifying adaxial fates and those specifying abaxial fates. ANT may function as a general activator of PHB-like and YABBY genes throughout organ primordia with their region-specific expression resulting from mutually repressive interactions between the PHB-like and KAN genes. Supporting our view that ANT is a positive regulator of genes specifying either adaxial or abaxial fates, preliminary examination of ant rev double mutants revealed enhanced carpel phenotypes 984 Plant Physiol. Vol. 141, 2006
9 Regulation of Organ Growth and Polarity by AINTEGUMENTA Figure 8. PHB expression in wild-type, fil-8, ant-4, and fil-8 ant-4 flowers. Size bars correspond to 50 mm in A to H. A, PHB expression in a wild-type inflorescence. B, PHB is expressed in the adaxial region of sepal primordia (arrows) and in the floral meristem dome in a stage 4 Ler flower. C, PHB expression in a fil-8 inflorescence. D, PHB expression in a stage 3 fil-8 flower. E, PHB expression in an ant-4 inflorescence. F, PHB expression in a stage 4 ant-4 flower. G, PHB mrna is present at reduced levels in fil-8 ant-4 inflorescences. H, PHB is expressed in the floral meristem dome but not in the outer whorl organ primordia of this stage 3 fil-8 ant-4 flower. St 2/4, Stage 2 and stage 4 flowers; IM, inflorescence stem. including enhanced loss of some adaxial tissues relative to either single mutant (S. Nole-Wilson and R. Franks, personal communication). ANT May Act with SEUSS and LEUNIG to Regulate Expression of PHB SEUSS (SEU) and LEUNIG (LUG) have been proposed to promote polarity along the adaxial/abaxial axis in petals by positively regulating PHB and FIL expression (Franks et al., 2006). Similar effects on leaf lamina/petal blade expansion and vascular development are observed in ag seu lug petals as reported here for fil yab ant leaves (Franks et al., 2006). These similarities are particularly intriguing as SEU, LUG, and ANT share other functions during flower development. All three proteins act as negative regulators of the floral homeotic gene AG (Liu and Meyerowitz, 1995; Krizek et al., 2000; Franks et al., 2002). Interestingly, FIL also acts as an AG repressor in whorls one and two (Chen et al., 1999). Similar carpel growth defects result from the combined loss of LUG and ANT or the combined loss of FIL and ANT. In both fil ant and lug ant flowers, the inner whorl consists of unfused or partially fused valve-like structures with style cells at their tips and an almost complete loss of adaxial tissues (placenta, ovules, and septa; Krizek et al., 2000; Liu et al., 2000). The similarities in these phenotypes suggest that ANT, FIL, SEU, and LUG have overlapping and partially redundant functions. These proteins might form a complex in which the SEU-LUG transcriptional corepressor (Sridhar et al., 2004) is recruited to promoter sequences via interaction with either of the DNA-binding proteins ANT or FIL (Nole-Wilson and Krizek, 2000; Kanaya et al., 2002). The Antirrhinum LUG ortholog STYLOSA has been shown to physically interact with YABBY proteins (Navarro et al., 2004). ANT and FIL could share some functions such that severe phenotypes only result in the absence of both ANT and FIL. ANT and YABBY Genes Promote Lamina Expansion and Floral Organ Identity Besides contributing to the specification of abaxial identity, YABBY genes are important regulators of lamina expansion. Polar expression of YABBY genes has been proposed to regulate signaling events between the adaxial and abaxial domains that control cell division in each domain and expansion of the leaf lamina (Eshed et al., 2004). Although ant mutants show only slight reductions in leaf area, the narrower lamina of fil yab ant leaves as compared with fil yab leaves indicates that ANT plays an important growth promotion role in leaves as well as flowers. Consistent with a role in lamina expansion, ANT expression in developing leaves becomes refined to the central and marginal regions in a pattern that is quite similar to the Solanum tuberosum YABBY gene StYABBY1 (Long and Barton, 2000; Eshed et al., 2004). While petals and stamens are present in fil and ant single mutants, organs with petal or stamen characteristics are rare in fil ant double mutants. The loss of floral organ identify in fil ant double mutants was correlated with altered floral homeotic gene expression. AP3 expression was reduced in fil ant flowers while the pattern of AG expression was altered. Thus, ANTacts as a positive regulator of the class B gene AP3 and acts to prevent AG expression in inflorescences and flowers prior to stage 3. A role for ANT in AG repression has been noted previously (Krizek et al., 2000; Liu et al., 2000). Similar losses in floral organ identity have been observed in other genotypes with polarity defects. For example, kan1 kan2 fil yab flowers consist of carpels and radialized organs that lack Plant Physiol. Vol. 141,
10 Nole-Wilson and Krizek cell types characteristic of sepals, petals, or stamens (Eshed et al., 2004). These results suggest a connection between the establishment of polarity and the specification of organ identity during flower development. Further studies will be needed to probe this relationship and to better understand the diverse processes that ANT regulates during lateral organ development. MATERIALS AND METHODS Protein Expression Full-length ANT lackingastopcodonwasclonedintopqe12 (Qiagen) and expressed by induction with 1 mm isopropyl-b-d-thiogalactoside in XL1-Blue MRF# Tet cells (Stratagene) at 30 C. Cells were harvested between 6 and 8 h after induction. ANT was purified using Ni-NTA (Qiagen) under denaturing conditions according to the manufacturer s instructions. ANT refolded upon dilution in the DNA-binding reactions. Gel Mobility Shift Assays Gel mobility shift assays were carried out as described previously (Nole- Wilson and Krizek, 2000), except that binding reactions were incubated for 4 h at roomtemperatureorovernight at 4 C. The YAB3 andfil binding sites were created by PCR amplification of Columbia genomic sequence using YAB-12 (5#-CTCGAGATTAAGTGTGAAAACAACTGAT-3#) and YAB-13 (5#-GAATT- CAAAGGACGCAAAGTTCGATG-3#) or FIL-3 (5#-TACACTCGAGTTAAG- GAATGACAACAACGGG-3#) and YAB-9 (5#-TACCGGATCCGAATTCGCA- GTTCCCAATGGA-3#), respectively. These fragments correspond to nucleotides at positions 1,561 to 1,456 upstream of the YAB3 start codon and nucleotides at positions 1,763 to 1,633 upstream of the FIL start codon. The PCR products were cloned into pcrscript (Stratagene) and the probes were prepared as described previously (Nole-Wilson and Krizek, 2000). Plant Growth Conditions Arabidopsis (Arabidopsis thaliana) ecotype Ler was used as the wild type. Plants were grown in a soil mixture of Fafard 4P:perlite:vermiculite in a ratio of 4:1:1 under continuous light ( mmol m 22 s 21 ) at a temperature of 22 C. Plants were fertilized once at 1 to 2 weeks postgermination. Real-Time RT-PCR Inflorescences were collected from 3- to 4-week-old Ler and ant-4 plants grown on soil under continuous light at 22 C. Total RNA was extracted and DNase treated as described previously (Nole-Wilson et al., 2005). Approximately 5 mg of total RNA was reverse transcribed using the Superscript First- Strand Synthesis system for RT-PCR (Invitrogen). Real-time RT-PCR was performed as described previously except that ACTIN2 (ACT2) was used for normalization purposes (Nole-Wilson et al., 2005). The ACT2 primers were ACT-3 (5#-CCTTTGTTGCTGTTGACT-3#) and ACT-4 (5#-GAACAAGACTT- CTGGGCATCT-3#). The YAB3 primers were YAB3-10 (5#-GCGGAGGGCA- GAATATAAAC-3#) and YAB3-11 (5#-CACTGATCTTCCGTTGCGA-3#). In Situ Hybridization Inflorescences were fixed, embedded, sectioned, hybridized, and washed as described previously (Krizek, 1999). Digoxigenin-labeled RNA probes were synthesized by in vitro transcription using T7 RNA polymerase (AP3, AG, and FIL probes) or T3 RNA polymerase (PHB probe) and the appropriate linearized plasmids. The AP3, AG, and FIL plasmids used for probe production have been described before (Yanofsky et al., 1990; Jack et al., 1992; Siegfried et al., 1999). The PHB probe corresponds to nucleotides 609 to 2,559 of PHB and was made after linearization of PHB/pBSKS with XbaI. Generation of fil-8 ant-4 and fil-8 yab3-2 ant-4 Plants yab3-2 fil-8/1 seeds were obtained from John Bowman. These alleles have been described previously (Kumaran et al., 1999, 2002). Both yab3-2 and fil-8 mutants are Ds insertion lines in the Ler background. YAB3 transcripts are not detectable in yab3-2 (Kumaran et al., 2002). ant-4 seeds were obtained from Charles Gasser (Baker et al., 1997). ant-4 is in the Ler background and contains a T-to-A transversion at nucleotide 1,335, altering the donor splice site of the fourth intron. These alleles were chosen as they are all strong alleles in the Ler background. The double and triple mutants were generated by pollinating putative fil-8 yab3-2/1 plants with ant-4 pollen. F 2 and subsequent generations were observed for segregation of plants with novel phenotypes. fil-8 ant-4 double mutants and fil-8 yab3-2 ant-4 triple mutants were confirmed by PCR genotyping. PCR Genotyping yab3-2 Green leaf tissue was prepared as described previously (Klimyuk et al., 1993) and subjected to PCR. PCR reactions using YAB35I-F (5#-GCCCTCCTC- TCTCTCTTACTC-3#) and YAB35I-R2 (5#-TCTGACCGTCACCGTCTTGA-3#) verified the absence of the wild-type YAB3 allele and PCR reactions using Ds5I-1 (5#-CCGTTTACCGTTTTGTATATCCCG-3#) and YAB35I-R2 verified the presence of the yab3-2 mutant allele. fil-8 Green leaf tissue was prepared as described above and subjected to PCR. PCR reactions using AFO-FW3 (5#-AGATTCCTAAAGCACCACCC-3#) and YAB1R (5#-GATACGTTGGATCTCCTCCC-3#) verified the absence of the wild-type FIL allele and PCR reactions using AFO-FW3 and Ds5I-1 verified the presence of the fil-8 allele. ant-4 DNA was isolated from green leaf tissue by grinding in 200 mm Tris, ph 7.5, 250 mm NaCl, 25mM EDTA, 0.5% SDS, and precipitation with isopropanol. The DNA was then PCR amplified using ANT-6 (5#-TCAAGGATC- CACTTTTGGACAACGAACTTCT-3#) and ANT-33 (5#-TCTTGGATCCTG- CAACATATTCTTGTCTAGT-3#). The PCR product was gel purified and cloned into the BamHI site of pgem3z (Promega). Sequencing of multiple clones confirmed the presence of only the ant-4 allele in putative fil-8 ant-4 and fil-8 yab3-2 ant-4 plants. Leaf Area and Plant Height Measurements For each genotype, the largest rosette leaf was removed from six different plants at the time of bolting. Leaf surface area was measured with a LI-COR LI-3000 portable area meter. The length and width of rosette leaves were measured using an ocular micrometer or a ruler. Plant heights were determined for six different plants of each genotype at the time when their primary inflorescences were starting to senesce. Bolting time was approximately the same for all genotypes. Leaf Vascular Staining Leaves were fixed overnight at room temperature in a 3:1 solution of ethanol:acetic acid. The tissue was mounted in 70% ethanol and examined using a dissecting microscope with illumination from below. SEM Tissue for SEM was fixed, dried, dissected, and coated as described previously (Krizek, 1999). SEM analysis was performed on a FEI XL30 ESEM (Hillsboro). ACKNOWLEDGMENTS We thank John Bowman for the fil-8 yab3-2/1 seeds and the FIL in situ plasmid, Charles Gasser for the ant-4 seeds, David Lincoln for help with leaf area measurements, John Herr for assistance with the xylem staining, and Mike Prigge and Steve Clark for the PHB in situ plasmid. We also thank Bob 986 Plant Physiol. Vol. 141, 2006
11 Regulation of Organ Growth and Polarity by AINTEGUMENTA Franks for sharing unpublished data and providing valuable comments on the manuscript. Received January 2, 2006; revised May 5, 2006; accepted May 12, 2006; published May 19, LITERATURE CITED Baker SC, Robinson-Beers K, Villanueva JM, Gaiser JC, Gasser CS (1997) Interactions among genes regulating ovule development in Arabidopsis thaliana. Genetics 145: Bowman JL (2000) The YABBY gene family and abaxial cell fate. Curr Opin Plant Biol 3: Bowman JL, Eshed Y, Baum SF (2002) Establishment of polarity in angiosperm lateral organs. Trends Genet 18: Bowman JL, Smyth DR (1999) CRABS CLAW, a gene that regulates carpel and nectary development in Arabidopsis, encodes a novel protein with zinc finger and helix-loop-helix domains. Development 126: Chen Q, Atkinson A, Otsuga D, Christensen T, Reynolds L, Drews GN (1999) The Arabidopsis FILAMENTOUS FLOWER gene is required for flower formation. Development 126: Elliott RC, Betzner AS, Huttner E, Oakes MP, Tucker WQJ, Gerentes D, Perez P, Smyth DR (1996) AINTEGUMENTA,anAPETALA2-likegeneof Arabidopsis with pleiotropic roles in ovule development and floral organ growth. Plant Cell 8: Emery JF, Floyd SK, Alvarez J, Eshed Y, Hawker NP, Izhaki A, Baum SF, Bowman JL (2003) Radial patterning of Arabidopsis shoots by class III HD-ZIP and KANADI genes. Curr Biol 13: Engstrom EM, Izhaki A, Bowman JL (2004) Promoter bashing, micro- RNAs, and KNOX genes. New insights, regulators, and targets-ofregulation in the establishment of lateral organ polarity in Arabidopsis. Plant Physiol 135: Eshed Y, Baum SF, Perea JV, Bowman JL (2001) Establishment of polarity in lateral organs of plants. Curr Biol 11: Eshed Y, Izhaki A, Baum SF, Floyd SK, Bowman JL (2004) Asymmetric leaf development and blade expansion in Arabidopsis are mediated by KANADI and YABBY activities. Development 131: Franks RG, Liu Z, Fischer RL (2006) SEUSS and LEUNIG regulate cell proliferation, vascular development andorganpolarityinarabidopsis petals. Planta (in press) Franks RG, Wang C, Levin JZ, Liu Z (2002) SEUSS, a member of a novel family of plant regulatory proteins, represses floral homeotic gene expression with LEUNIG. Development129: Jack T, Brockman LL, Meyerowitz EM (1992) The homeotic gene APETALA3 of Arabidopsis thaliana encodes a MADS box and is expressed in petals and stamens. Cell 68: Kanaya E, Nakajima N, Okada K (2002) Non-sequence-specific DNA binding by the FILAMENTOUS FLOWER protein from Arabidopsis thaliana in reduced by EDTA. J Biol Chem 227: Kerstetter RA, Bollman K, Taylor RA, Bomblies K, Poethig RS (2001) KANADI regulates organ polarity in Arabidopsis. Nature 411: Klimyuk VI, Carroll BJ, Thomas CM, Jones JDG (1993) Alkali treatment for rapid preparation of plant material for reliable PCR analysis. Plant J 3: Klucher KM, Chow H, Reiser L, Fischer RL (1996) The AINTEGUMENTA gene of Arabidopsis required for ovule and female gametophyte development is related to the floral homeotic gene APETALA2. PlantCell8: Krizek BA (1999) Ectopic expression of AINTEGUMENTA in Arabidopsis plants results in increased growth of floral organs. Dev Genet 25: Krizek BA, Prost V, Macias A (2000) AINTEGUMENTA promotes petal identity and acts as a negative regulator of AGAMOUS. Plant Cell 12: Kumaran MK, Bowman JL, Sundaresan V (2002) YABBY polarity genes mediate the repression of KNOX homeobox genes in Arabidopsis. Plant Cell 14: Kumaran MK, Ye D, Yang W-C, Griffith ME, Chaudhury AM, Sundaresan V (1999) Molecular cloning of ABNORMAL FLORAL ORGANS: agene required for flower development in Arabidopsis. Sex Plant Reprod 12: Liu Z, Franks RG, Klink VP (2000) Regulation of gynoecium marginal tissue formation by LEUNIG and AINTEGUMENTA. PlantCell12: Liu Z, Meyerowitz EM (1995) LEUNIG regulates AGAMOUS expression in Arabidopsis flowers. Development 121: Long J, Barton MK (2000) Initiation of axillary and floral meristems in Arabidopsis. Dev Biol 218: McConnell JR, Barton MK (1998) Leaf polarity and meristem formation in Arabidopsis. Development 125: McConnell JR, Emery J, Eshed Y, Bao N, Bowman J, Barton MK (2001) Role of PHABULOSA and PHAVOLUTA in determining radial patterning in shoots. Nature 411: Navarro C, Efremova N, Golz JF, Rubiera R, Kuckenberg M, Castillo R, Tietz O, Saedler H, Schwarz-Sommer Z (2004) Molecular and genetic interactions between STYLOSA and GRAMINIFOLIA in the control of Antirrhinum vegetative and reproductive development. Development 131: Nole-Wilson S, Krizek BA (2000) DNA binding properties of the Arabidopsis floral development protein AINTEGUMENTA. Nucleic Acids Res 28: Nole-Wilson S, Tranby T, Krizek BA (2005) AINTEGUMENTA-like (AIL) genes are expressed in young tissues and may specify meristematic or division-competent states. Plant Mol Biol 57: Sawa S, Watanabe K, Goto K, Kanaya E, Morita EM, Okada K (1999) FILAMENTOUS FLOWER, a meristem and organ identity gene of Arabidopsis, encodes a protein with a zinc finger and HMG-related domains. Genes Dev 13: Siegfried KR, Eshed Y, Baum SF, Otsuga D, Drews GN, Bowman JL (1999) Members of the YABBY gene family specify abaxial cell fate in Arabidopsis. Development 126: Sridhar VV, Surendrarao A, Gonzalez D, Conlan RS, Liu Z (2004) Transcriptional repression of target genes by LEUNIG and SEUSS, two interacting regulatory proteins for Arabidopsis flower development. Proc Natl Acad Sci USA 101: Waites R, Hudson A (1995) phantastica: a gene required for dorsoventrality of leaves in Antirrhinum majus. Development121: Watanabe K, Okada K (2003) Two discrete cis elements control the abaxial side-specific expression of the FILAMENTOUS FLOWER gene in Arabidopsis. Plant Cell 15: Yanofsky MF, Ma H, Bowman JL, Drews GN, Feldman KA, Meyerowitz EM (1990) The protein encoded by the Arabidopsis homeotic gene AGAMOUS resembles transcription factors. Nature 346: Plant Physiol. Vol. 141,
Supplemental Figure S1. The number of hydathodes is reduced in the as2-1 rev-1
 Supplemental Data Supplemental Figure S1. The number of hydathodes is reduced in the as2-1 rev-1 and kan1-11 kan2-5 double mutants. A, The numbers of hydathodes in different leaves of Col-0, as2-1 rev-1,
Supplemental Data Supplemental Figure S1. The number of hydathodes is reduced in the as2-1 rev-1 and kan1-11 kan2-5 double mutants. A, The numbers of hydathodes in different leaves of Col-0, as2-1 rev-1,
FILAMENTOUS FLOWER Controls the Formation and Development of Arabidopsis Inflorescences and Floral Meristems
 The Plant Cell, Vol. 11, 69 86, January 1999, www.plantcell.org 1999 American Society of Plant Physiologists FILAMENTOUS FLOWER Controls the Formation and Development of Arabidopsis Inflorescences and
The Plant Cell, Vol. 11, 69 86, January 1999, www.plantcell.org 1999 American Society of Plant Physiologists FILAMENTOUS FLOWER Controls the Formation and Development of Arabidopsis Inflorescences and
Asymmetric leaf development and blade expansion in Arabidopsis are mediated by KANADI and YABBY activities
 Research article 2997 Asymmetric leaf development and blade expansion in Arabidopsis are mediated by KANADI and YABBY activities Yuval Eshed 1,2, *,, Anat Izhaki 1, *, Stuart F. Baum 1,, Sandra K. Floyd
Research article 2997 Asymmetric leaf development and blade expansion in Arabidopsis are mediated by KANADI and YABBY activities Yuval Eshed 1,2, *,, Anat Izhaki 1, *, Stuart F. Baum 1,, Sandra K. Floyd
Regulation of Floral Organ Identity. Dr. Chloe Diamond Mara
 Regulation of Floral Organ Identity Dr. Chloe Diamond Mara Flower Development Angiosperms (flowering plants) are the most widespread group of land plants Flowers are the reproductive organs that consist
Regulation of Floral Organ Identity Dr. Chloe Diamond Mara Flower Development Angiosperms (flowering plants) are the most widespread group of land plants Flowers are the reproductive organs that consist
Arabidopsis: Flower Development and Patterning
 Arabidopsis: Flower Development and Patterning John L Bowman, University of California, Davis, California, USA The development of flowers and floral organs is directed by genetic programmes likely to be
Arabidopsis: Flower Development and Patterning John L Bowman, University of California, Davis, California, USA The development of flowers and floral organs is directed by genetic programmes likely to be
Regulation of Floral-Organ- Type by SUPERMAN
 Regulation of Floral-Organ- Type by SUPERMAN 1. Need for regulators of the organ-identity genes. 2. The Superman mutant phenotype-predicting the role of SUPERMAN. 3. Testing our hypothesis of the role
Regulation of Floral-Organ- Type by SUPERMAN 1. Need for regulators of the organ-identity genes. 2. The Superman mutant phenotype-predicting the role of SUPERMAN. 3. Testing our hypothesis of the role
Testing the ABC floral-organ identity model: expression of A and C function genes
 Objectives: Testing the ABC floral-organ identity model: expression of A and C function genes To test the validity of the ABC model for floral organ identity we will: 1. Use the model to make predictions
Objectives: Testing the ABC floral-organ identity model: expression of A and C function genes To test the validity of the ABC model for floral organ identity we will: 1. Use the model to make predictions
RABBIT EARS is a second-whorl repressor of AGAMOUS that maintains spatial boundaries in Arabidopsis flowers
 The Plant Journal (2006) 45, 369 383 doi: 10.1111/j.1365-313X.2005.02633.x RABBIT EARS is a second-whorl repressor of AGAMOUS that maintains spatial boundaries in Arabidopsis flowers Beth A. Krizek 1,*,
The Plant Journal (2006) 45, 369 383 doi: 10.1111/j.1365-313X.2005.02633.x RABBIT EARS is a second-whorl repressor of AGAMOUS that maintains spatial boundaries in Arabidopsis flowers Beth A. Krizek 1,*,
A genetic framework for fruit patterning in Arabidopsis thaliana
 Research article 4687 A genetic framework for fruit patterning in Arabidopsis thaliana José R. Dinneny 1,2, Detlef Weigel 2,3 and Martin F. Yanofsky 1, * 1 Division of Biological Sciences, University of
Research article 4687 A genetic framework for fruit patterning in Arabidopsis thaliana José R. Dinneny 1,2, Detlef Weigel 2,3 and Martin F. Yanofsky 1, * 1 Division of Biological Sciences, University of
Supplemental Data. Müller-Xing et al. (2014). Plant Cell /tpc
 Supplemental Figure 1. Phenotypes of iclf (clf-28 swn-7 CLF pro :CLF-GR) plants. A, Late rescue of iclf plants by renewed DEX treatment; senescent inflorescence with elongated siliques (arrow; 90 DAG,
Supplemental Figure 1. Phenotypes of iclf (clf-28 swn-7 CLF pro :CLF-GR) plants. A, Late rescue of iclf plants by renewed DEX treatment; senescent inflorescence with elongated siliques (arrow; 90 DAG,
Involvement of CUP-SHAPED COTYLEDON Genes in Gynoecium and Ovule Development in Arabidopsis thaliana
 Plant CellPhysiol. 41(1): 60-67 (2000) JSPP 2000 Involvement of CUP-SHAPED COTYLEDON Genes in Gynoecium and Ovule Development in Arabidopsis thaliana Tetsuya Ishida ', Mitsuhiro Aida 2, Shinobu Takada
Plant CellPhysiol. 41(1): 60-67 (2000) JSPP 2000 Involvement of CUP-SHAPED COTYLEDON Genes in Gynoecium and Ovule Development in Arabidopsis thaliana Tetsuya Ishida ', Mitsuhiro Aida 2, Shinobu Takada
The Arabidopsis homeotic genes APETALA3 and PISTILLATA are sufficient to provide the B class organ identity function
 University of South Carolina Scholar Commons Faculty Publications Biological Sciences, Department of 1-1-1996 The Arabidopsis homeotic genes APETALA3 and PISTILLATA are sufficient to provide the B class
University of South Carolina Scholar Commons Faculty Publications Biological Sciences, Department of 1-1-1996 The Arabidopsis homeotic genes APETALA3 and PISTILLATA are sufficient to provide the B class
The Role of Lipids in Flowering Development of Arabidopsis Enhanced pah1pah2 Plants. Toshiro Ito 1 & Lee Lishi 2
 The Role of Lipids in Flowering Development of Arabidopsis Enhanced pah1pah2 Plants Toshiro Ito 1 & Lee Lishi 2 Department of Biological Sciences, Faculty of Science, National University of Singapore,
The Role of Lipids in Flowering Development of Arabidopsis Enhanced pah1pah2 Plants Toshiro Ito 1 & Lee Lishi 2 Department of Biological Sciences, Faculty of Science, National University of Singapore,
Floral Organ Mutants and the Study of Organ Morphogenesis
 Floral Organ Mutants and the Study of Organ Morphogenesis Objectives: 1. How does one use mutants to understand floral organ morphogenesis? 2. What are the phenotypes of some floral organ mutants? 3. What
Floral Organ Mutants and the Study of Organ Morphogenesis Objectives: 1. How does one use mutants to understand floral organ morphogenesis? 2. What are the phenotypes of some floral organ mutants? 3. What
UFO and LEAFY in Arabidopsis
 Development 128, 2735-2746 (2001) Printed in Great Britain The Company of Biologists Limited 2001 DEV0360 2735 The ASK1 gene regulates B function gene expression in cooperation with UFO and LEAFY in Arabidopsis
Development 128, 2735-2746 (2001) Printed in Great Britain The Company of Biologists Limited 2001 DEV0360 2735 The ASK1 gene regulates B function gene expression in cooperation with UFO and LEAFY in Arabidopsis
Activation of the Arabidopsis B Class Homeotic Genes by APETALA1
 The Plant Cell, Vol. 13, 739 753, April 2001, www.plantcell.org 2001 American Society of Plant Physiologists RESEARCH ARTICLE Activation of the Arabidopsis B Class Homeotic Genes by APETALA1 Medard Ng
The Plant Cell, Vol. 13, 739 753, April 2001, www.plantcell.org 2001 American Society of Plant Physiologists RESEARCH ARTICLE Activation of the Arabidopsis B Class Homeotic Genes by APETALA1 Medard Ng
Beth A. Krizek & Marcie Eaddy
 AINTEGUMENTA-LIKE6 regulates cellular differentiation in flowers Beth A. Krizek & Marcie Eaddy Plant Molecular Biology An International Journal on Molecular Biology, Molecular Genetics and Biochemistry
AINTEGUMENTA-LIKE6 regulates cellular differentiation in flowers Beth A. Krizek & Marcie Eaddy Plant Molecular Biology An International Journal on Molecular Biology, Molecular Genetics and Biochemistry
CRABS CLAW and SPATULA, two Arabidopsis genes that control carpel
 Development 126, 2377-2386 (1999) Printed in Great Britain The Company of Biologists Limited 1999 DEV0225 2377 CRABS CLAW and SPATULA, two Arabidopsis genes that control carpel development in parallel
Development 126, 2377-2386 (1999) Printed in Great Britain The Company of Biologists Limited 1999 DEV0225 2377 CRABS CLAW and SPATULA, two Arabidopsis genes that control carpel development in parallel
CLAVATA1, a regulator of meristem and flower development in Arabidopsis
 Development 119, 397-418 (1993) Printed in Great Britain The Company of Biologists Limited 1993 397 CLAVATA1, a regulator of meristem and flower development in Arabidopsis Steven E. Clark, Mark P. Running
Development 119, 397-418 (1993) Printed in Great Britain The Company of Biologists Limited 1993 397 CLAVATA1, a regulator of meristem and flower development in Arabidopsis Steven E. Clark, Mark P. Running
Genes Directing Flower Development in Arabidopsis
 The Plant Cell, Vol. 1,37-52, January 1989, 1989 American Society of Plant Physiologists Genes Directing Flower Development in Arabidopsis John L. Bowman, David R. Smyth, 1 and Elliot M. Meyerowitz 2 Division
The Plant Cell, Vol. 1,37-52, January 1989, 1989 American Society of Plant Physiologists Genes Directing Flower Development in Arabidopsis John L. Bowman, David R. Smyth, 1 and Elliot M. Meyerowitz 2 Division
BELL1 and AGAMOUS genes promote ovule identity in Arabidopsis thaliana
 The Plant Journal (1999) 18(3), 329 336 SHORT COMMUNICATION BELL1 and AGAMOUS genes promote ovule identity in Arabidopsis thaliana Tamara L. Western and George W. Haughn* Botany Department, University
The Plant Journal (1999) 18(3), 329 336 SHORT COMMUNICATION BELL1 and AGAMOUS genes promote ovule identity in Arabidopsis thaliana Tamara L. Western and George W. Haughn* Botany Department, University
A genetic and molecular model for flower development in Arabidopsis thaliana
 Development Supplement I, 1991, 157-167 Printed in Great Britain The Company of Biologists Limited 1991 157 A genetic and molecular model for flower development in Arabidopsis thaliana ELLIOT M. MEYEROWITZ*,
Development Supplement I, 1991, 157-167 Printed in Great Britain The Company of Biologists Limited 1991 157 A genetic and molecular model for flower development in Arabidopsis thaliana ELLIOT M. MEYEROWITZ*,
Genetic Specification of floral organ identity. Initiating floral development. Deciding when to initiate flowering - induced mutations -in Nature
 Genetic Specification of floral organ identity Initiating floral development Deciding when to initiate flowering - induced mutations -in Nature Flower structure of rabidopsis Stamens arpels Petals Sepals
Genetic Specification of floral organ identity Initiating floral development Deciding when to initiate flowering - induced mutations -in Nature Flower structure of rabidopsis Stamens arpels Petals Sepals
Supplemental Data. Wu et al. (2010). Plant Cell /tpc
 Supplemental Figure 1. FIM5 is preferentially expressed in stamen and mature pollen. The expression data of FIM5 was extracted from Arabidopsis efp browser (http://www.bar.utoronto.ca/efp/development/),
Supplemental Figure 1. FIM5 is preferentially expressed in stamen and mature pollen. The expression data of FIM5 was extracted from Arabidopsis efp browser (http://www.bar.utoronto.ca/efp/development/),
Determination of Arabidopsis Floral Meristem ldentity by AGA MOUS
 The Plant Cell, Vol. 9, 393-408, March 1997 O 1997 American Society of Plant Physiologists Determination of Arabidopsis loral Meristem ldentity by AGA MOUS Yukiko Mizukamil and Hong Ma2 Cold Spring Harbor
The Plant Cell, Vol. 9, 393-408, March 1997 O 1997 American Society of Plant Physiologists Determination of Arabidopsis loral Meristem ldentity by AGA MOUS Yukiko Mizukamil and Hong Ma2 Cold Spring Harbor
Ectopic Expression of AINTEGUMENTA in Arabidopsis Plants Results in Increased Growth of Floral Organs
 DEVELOPMENTAL GENETICS 25:224 236 (1999) Ectopic Expression of AINTEGUMENTA in Arabidopsis Plants Results in Increased Growth of Floral Organs BETH ALLYN KRIZEK* Department of Biological Sciences, University
DEVELOPMENTAL GENETICS 25:224 236 (1999) Ectopic Expression of AINTEGUMENTA in Arabidopsis Plants Results in Increased Growth of Floral Organs BETH ALLYN KRIZEK* Department of Biological Sciences, University
The Flower - what is it? 1/31/18. Magnoliophyta - Flowering Plants. Magnoliophyta - Flowering Plants. Magnoliophyta - Flowering Plants
 - what is it? Floral structure will be examined in lab next Mon/Tues save space in your notes! Introduction to Angiosperms "angio-" = vessel; so "angiosperm" means "vessel for the seed [seed encased in
- what is it? Floral structure will be examined in lab next Mon/Tues save space in your notes! Introduction to Angiosperms "angio-" = vessel; so "angiosperm" means "vessel for the seed [seed encased in
Signals Derived from YABBY Gene Activities in Organ Primordia Regulate Growth and Partitioning of Arabidopsis Shoot Apical Meristems W
 The Plant Cell, Vol. 20: 1217 1230, May 2008, www.plantcell.org ª 2008 American Society of Plant Biologists Signals Derived from YABBY Gene Activities in Organ Primordia Regulate Growth and Partitioning
The Plant Cell, Vol. 20: 1217 1230, May 2008, www.plantcell.org ª 2008 American Society of Plant Biologists Signals Derived from YABBY Gene Activities in Organ Primordia Regulate Growth and Partitioning
Genetics of Floral Development An Evo-Devo Approach
 Genetics of Floral Development An Evo-Devo Approach Biology 317 Kelsey Galimba 8.5.2013 Why are flowering plants so diverse? Why are flowering plants so diverse? http://www.botanicalgarden.ubc.ca/potd/
Genetics of Floral Development An Evo-Devo Approach Biology 317 Kelsey Galimba 8.5.2013 Why are flowering plants so diverse? Why are flowering plants so diverse? http://www.botanicalgarden.ubc.ca/potd/
The Ff.010 Gene Product Regulates the Expression Domain of Homeotic Genes AP3 and PI in Arabidopsis Flowers
 The Plant Cell, Vol. 3, 1221-1237, November 1991 1991 American Society of Plant Physiologists The Ff.010 Gene Product Regulates the Expression Domain of Homeotic Genes AP3 and PI in Arabidopsis Flowers
The Plant Cell, Vol. 3, 1221-1237, November 1991 1991 American Society of Plant Physiologists The Ff.010 Gene Product Regulates the Expression Domain of Homeotic Genes AP3 and PI in Arabidopsis Flowers
Flower Morphology. Flower Structure. Name
 right 84 83 82 81 80 79 78 77 76 75 74 73 72 71 70 69 68 67 score 100 98.8 97.6 96.4 95.2 94.0 92.9 91.7 90.5 89.3 88.1 86.9 85.7 84.5 83.3 82.1 81.0 79.8 Flower Morphology Name You are already familiar
right 84 83 82 81 80 79 78 77 76 75 74 73 72 71 70 69 68 67 score 100 98.8 97.6 96.4 95.2 94.0 92.9 91.7 90.5 89.3 88.1 86.9 85.7 84.5 83.3 82.1 81.0 79.8 Flower Morphology Name You are already familiar
CLAVATA3 is a specific regulator of shoot and floral meristem development
 Development 2, 20572067 (995) Printed in Great Britain The Company of Biologists Limited 995 2057 CLAVATA3 is a specific regulator of shoot and floral meristem development affecting the same processes
Development 2, 20572067 (995) Printed in Great Britain The Company of Biologists Limited 995 2057 CLAVATA3 is a specific regulator of shoot and floral meristem development affecting the same processes
Ribosomal proteins promote leaf adaxial identity
 RESEARCH ARTICLE 1325 Development 135, 1325-1334 (2008) doi:10.1242/dev.017913 Ribosomal proteins promote leaf adaxial identity Yao Yao*, Qihua Ling*, Hua Wang and Hai Huang Establishing abaxial-adaxial
RESEARCH ARTICLE 1325 Development 135, 1325-1334 (2008) doi:10.1242/dev.017913 Ribosomal proteins promote leaf adaxial identity Yao Yao*, Qihua Ling*, Hua Wang and Hai Huang Establishing abaxial-adaxial
Redundantly in the Temporal Regulation of Floral Meristem Termination in Arabidopsis thaliana W
 The Plant Cell, Vol. 20: 901 919, April 2008, www.plantcell.org ª 2008 American Society of Plant Biologists REBELOTE, SQUINT, andultrapetala1 Function Redundantly in the Temporal Regulation of Floral Meristem
The Plant Cell, Vol. 20: 901 919, April 2008, www.plantcell.org ª 2008 American Society of Plant Biologists REBELOTE, SQUINT, andultrapetala1 Function Redundantly in the Temporal Regulation of Floral Meristem
Plant Terminology. Floral Symmetry
 Plant Terminology Parts of a Flower Pedicel--the stalk of an individual flower Calyx--outermost whorl of a flower Sepal--one member of the calyx Corolla--second whorl of a flower Petal--one member of the
Plant Terminology Parts of a Flower Pedicel--the stalk of an individual flower Calyx--outermost whorl of a flower Sepal--one member of the calyx Corolla--second whorl of a flower Petal--one member of the
HANABA TARANU Is a GATA Transcription Factor That Regulates Shoot Apical Meristem and Flower Development in Arabidopsis W
 The Plant Cell, Vol. 16, 2586 2600, October 2004, www.plantcell.org ª 2004 American Society of Plant Biologists HANABA TARANU Is a GATA Transcription Factor That Regulates Shoot Apical Meristem and Flower
The Plant Cell, Vol. 16, 2586 2600, October 2004, www.plantcell.org ª 2004 American Society of Plant Biologists HANABA TARANU Is a GATA Transcription Factor That Regulates Shoot Apical Meristem and Flower
The NGATHA Distal Organ Development Genes Are Essential for Style Specification in Arabidopsis W
 The Plant Cell, Vol. 21: 1373 1393, May 2009, www.plantcell.org ã 2009 American Society of Plant Biologists The NGATHA Distal Organ Development Genes Are Essential for Style Specification in Arabidopsis
The Plant Cell, Vol. 21: 1373 1393, May 2009, www.plantcell.org ã 2009 American Society of Plant Biologists The NGATHA Distal Organ Development Genes Are Essential for Style Specification in Arabidopsis
A LEAFY co-regulator encoded by UNUSUAL FLORAL ORGANS Ilha Lee, Diana S. Wolfe, Ove Nilsson and Detlef Weigel
 Research Paper 95 A LEAFY co-regulator encoded by UNUSUAL FLORAL ORGANS Ilha Lee, Diana S. Wolfe, Ove Nilsson and Detlef Weigel Background: Development of petals and stamens in Arabidopsis flowers requires
Research Paper 95 A LEAFY co-regulator encoded by UNUSUAL FLORAL ORGANS Ilha Lee, Diana S. Wolfe, Ove Nilsson and Detlef Weigel Background: Development of petals and stamens in Arabidopsis flowers requires
Flower Morphology. Flower Structure
 wrong 0 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 right 84 83 82 81 80 79 78 77 76 75 74 73 72 71 70 69 68 67 66 65 64 score 100 98.8 97.6 96.4 95.2 94.0 92.9 91.7 90.5 89.3 88.1 86.9 85.7 84.5
wrong 0 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 right 84 83 82 81 80 79 78 77 76 75 74 73 72 71 70 69 68 67 66 65 64 score 100 98.8 97.6 96.4 95.2 94.0 92.9 91.7 90.5 89.3 88.1 86.9 85.7 84.5
The SEP4 Gene of Arabidopsis thaliana Functions in Floral Organ and Meristem Identity
 Current Biology, Vol. 14, 1935 1940, November 9, 2004, 2004 Elsevier Ltd. All rights reserved. DOI 10.1016/j.cub.2004.10.028 The SEP4 Gene of Arabidopsis thaliana Functions in Floral Organ and Meristem
Current Biology, Vol. 14, 1935 1940, November 9, 2004, 2004 Elsevier Ltd. All rights reserved. DOI 10.1016/j.cub.2004.10.028 The SEP4 Gene of Arabidopsis thaliana Functions in Floral Organ and Meristem
From model plants to crops: the MADS box family of gene controlling flower development in Crocus (Crocus sativus L.)
 From model plants to crops: the MADS box family of gene controlling flower development in Crocus (Crocus sativus L.) A. Tsaftaris 1, 2, K. Pasentsis 1, A. Kalivas 2, A. Polidoros 1 1 Institute of Agobiotechnology
From model plants to crops: the MADS box family of gene controlling flower development in Crocus (Crocus sativus L.) A. Tsaftaris 1, 2, K. Pasentsis 1, A. Kalivas 2, A. Polidoros 1 1 Institute of Agobiotechnology
Arabidopsis PRC1 core component AtRING1 regulates stem cell-determining carpel development mainly through repression of class I KNOX genes
 Chen et al. BMC Biology (2016) 14:112 DOI 10.1186/s12915-016-0336-4 RESEARCH ARTICLE Open Access Arabidopsis PRC1 core component AtRING1 regulates stem cell-determining carpel development mainly through
Chen et al. BMC Biology (2016) 14:112 DOI 10.1186/s12915-016-0336-4 RESEARCH ARTICLE Open Access Arabidopsis PRC1 core component AtRING1 regulates stem cell-determining carpel development mainly through
Specification of Arabidopsis floral meristem identity by repression of flowering time genes
 RESEARCH ARTICLE 1901 Development 134, 1901-1910 (2007) doi:10.1242/dev.003103 Specification of Arabidopsis floral meristem identity by repression of flowering time genes Chang Liu 1, *, Jing Zhou 1, *,
RESEARCH ARTICLE 1901 Development 134, 1901-1910 (2007) doi:10.1242/dev.003103 Specification of Arabidopsis floral meristem identity by repression of flowering time genes Chang Liu 1, *, Jing Zhou 1, *,
ARGONAUTE10 and ARGONAUTE1 Regulate the Termination of Floral Stem Cells Through Two MicroRNAs in Arabidopsis
 University of Kentucky UKnowledge Plant and Soil Sciences Faculty Publications Plant and Soil Sciences 3-31-2011 ARGONAUTE10 and ARGONAUTE1 Regulate the Termination of Floral Stem Cells Through Two MicroRNAs
University of Kentucky UKnowledge Plant and Soil Sciences Faculty Publications Plant and Soil Sciences 3-31-2011 ARGONAUTE10 and ARGONAUTE1 Regulate the Termination of Floral Stem Cells Through Two MicroRNAs
Chapter 38. Plant Reproduction. AP Biology
 Chapter 38. Plant Reproduction 1 Animal vs. Plant life cycle Animal multicellular 2n Plant multicellular sporophyte 2n gametes 1n spores 1n unicellular gametes 1n multicellular gametophyte 1n 2 Alternation
Chapter 38. Plant Reproduction 1 Animal vs. Plant life cycle Animal multicellular 2n Plant multicellular sporophyte 2n gametes 1n spores 1n unicellular gametes 1n multicellular gametophyte 1n 2 Alternation
HALF FILLED promotes reproductive tract development and fertilization efficiency in Arabidopsis thaliana
 RESEARCH ARTICLE 2999 Development 138, 2999-3009 (2011) doi:10.1242/dev.067793 2011. Published by The Company of Biologists Ltd HALF FILLED promotes reproductive tract development and fertilization efficiency
RESEARCH ARTICLE 2999 Development 138, 2999-3009 (2011) doi:10.1242/dev.067793 2011. Published by The Company of Biologists Ltd HALF FILLED promotes reproductive tract development and fertilization efficiency
The Flower, Pollination, and Seeds
 The Flower, Pollination, and Seeds Class 9 th Chapters 6,7,8 1 The Flower A complete or a perfect flower, has all the four Whorls. If, even one whorl is missing, it is an Incomplete Flower. The fourth
The Flower, Pollination, and Seeds Class 9 th Chapters 6,7,8 1 The Flower A complete or a perfect flower, has all the four Whorls. If, even one whorl is missing, it is an Incomplete Flower. The fourth
Chapter 38. Plant Reproduction. AP Biology
 Chapter 38. Plant Reproduction 1 Animal vs. Plant life cycle Animal multicellular 2n Plant multicellular sporophyte 2n gametes 1n spores 1n unicellular gametes 1n multicellular gametophyte 1n 2 Alternation
Chapter 38. Plant Reproduction 1 Animal vs. Plant life cycle Animal multicellular 2n Plant multicellular sporophyte 2n gametes 1n spores 1n unicellular gametes 1n multicellular gametophyte 1n 2 Alternation
KNAT1 and ERECTA Regulate Inflorescence Architecture in Arabidopsis
 The Plant Cell, Vol. 14, 547 558, March 2002, www.plantcell.org 2002 American Society of Plant Biologists RESEARCH ARTICLE KNAT1 and ERECTA Regulate Inflorescence Architecture in Arabidopsis Scott J. Douglas,
The Plant Cell, Vol. 14, 547 558, March 2002, www.plantcell.org 2002 American Society of Plant Biologists RESEARCH ARTICLE KNAT1 and ERECTA Regulate Inflorescence Architecture in Arabidopsis Scott J. Douglas,
Peony Flower Anatomy I
 Peony Flower Anatomy I Don Hollingsworth, APS Director Maryville, Missouri What Makes a Peony Flower Luxurious? Rich luxury of the flowers explains why peonies are wanted, why loved and why known in history
Peony Flower Anatomy I Don Hollingsworth, APS Director Maryville, Missouri What Makes a Peony Flower Luxurious? Rich luxury of the flowers explains why peonies are wanted, why loved and why known in history
ACURIOUS malformation in one of the flowers on a raceme of
 ON AN ABNORMALITY IN PURPUREA By VIOLET L. ANDERSON. DIGITALIS Quain Student of Botany, University College, London. (With 6 figures in the text.) ACURIOUS malformation in one of the flowers on a raceme
ON AN ABNORMALITY IN PURPUREA By VIOLET L. ANDERSON. DIGITALIS Quain Student of Botany, University College, London. (With 6 figures in the text.) ACURIOUS malformation in one of the flowers on a raceme
Supplemental Information. Spatial Auxin Signaling. Controls Leaf Flattening in Arabidopsis
 Current Biology, Volume 27 Supplemental Information Spatial Auxin Signaling Controls Leaf Flattening in Arabidopsis Chunmei Guan, Binbin Wu, Ting Yu, Qingqing Wang, Naden T. Krogan, Xigang Liu, and Yuling
Current Biology, Volume 27 Supplemental Information Spatial Auxin Signaling Controls Leaf Flattening in Arabidopsis Chunmei Guan, Binbin Wu, Ting Yu, Qingqing Wang, Naden T. Krogan, Xigang Liu, and Yuling
Flowering Plant Reproduction
 Lab Exercise Flowering Plant Reproduction Objectives - To be able to identify the parts of a flower - Be able to distinguish between dicots and monocots based on flower morphology - Become familiar with
Lab Exercise Flowering Plant Reproduction Objectives - To be able to identify the parts of a flower - Be able to distinguish between dicots and monocots based on flower morphology - Become familiar with
Flowers, Fruit and Seeds Notes Flower Structure and Reproduction Taken from
 Flowers, Fruit and Seeds Notes Flower Structure and Reproduction Taken from http://www.biologycorner.com/worksheets/flower_coloring.html Flowers are the plant's reproductive structures. Angiosperms are
Flowers, Fruit and Seeds Notes Flower Structure and Reproduction Taken from http://www.biologycorner.com/worksheets/flower_coloring.html Flowers are the plant's reproductive structures. Angiosperms are
Functional Diversification of the Two C-Class MADS Box Genes OSMADS3 and OSMADS58 in Oryza sativa W OA
 The Plant Cell, Vol. 18, 15 28, January 2006, www.plantcell.org ª 2005 American Society of Plant Biologists RESEARCH ARTICLES Functional Diversification of the Two C-Class MADS Box Genes OSMADS3 and OSMADS58
The Plant Cell, Vol. 18, 15 28, January 2006, www.plantcell.org ª 2005 American Society of Plant Biologists RESEARCH ARTICLES Functional Diversification of the Two C-Class MADS Box Genes OSMADS3 and OSMADS58
Gwyneth C. Ingram,a Justin Goodrich,a Mark D. Wilkinson,b Rüdiger Simon,a George W. Haughn,b and Enrico S. Coena,
 The Plant Cell, Vol. 7, 1501-1510, September 1995 O 1995 American Society of Plant Physiologists Parallels between UNUSUAL FLORAL ORGANS and FMBRATA, Genes Controlling Flower Development in Arabidopsis
The Plant Cell, Vol. 7, 1501-1510, September 1995 O 1995 American Society of Plant Physiologists Parallels between UNUSUAL FLORAL ORGANS and FMBRATA, Genes Controlling Flower Development in Arabidopsis
Expression of a mutant maize gene in the ventral leaf epidermis is sufficient to signal a switch of the leaf s dorsoventral axis
 Development 129, 4581-4589 (2002) Printed in Great Britain The Company of Biologists Limited 2002 DEV0435 4581 Expression of a mutant maize gene in the ventral leaf epidermis is sufficient to signal a
Development 129, 4581-4589 (2002) Printed in Great Britain The Company of Biologists Limited 2002 DEV0435 4581 Expression of a mutant maize gene in the ventral leaf epidermis is sufficient to signal a
Genetic Separation of Third and Fourth Whorl Functions of AGAMOUS
 The Plant Cell, Vol. 7, 1249-1258, August 1995 O 1995 American Society of Plant Physiologists Genetic Separation of Third and Fourth Whorl Functions of AGAMOUS Leslie E. Sieburth, Mark P. Running, and
The Plant Cell, Vol. 7, 1249-1258, August 1995 O 1995 American Society of Plant Physiologists Genetic Separation of Third and Fourth Whorl Functions of AGAMOUS Leslie E. Sieburth, Mark P. Running, and
Reproductive Development and Structure
 Reproductive Development and Structure Bởi: OpenStaxCollege Sexual reproduction takes place with slight variations in different groups of plants. Plants have two distinct stages in their lifecycle: the
Reproductive Development and Structure Bởi: OpenStaxCollege Sexual reproduction takes place with slight variations in different groups of plants. Plants have two distinct stages in their lifecycle: the
Plant Life Cycles. Plant life cycles alternate between. producing gametes. Life cycle phases look different among various
 Plant Life Cycles Plant life cycles alternate between two cycles: Producing spores and producing gametes A two phase life cycle is called alternation of generations Diploid phase Haploid phase Alternates
Plant Life Cycles Plant life cycles alternate between two cycles: Producing spores and producing gametes A two phase life cycle is called alternation of generations Diploid phase Haploid phase Alternates
Arabidopsis gynoecium structure in the wild type and in ettin mutants
 Development 121, 1519-1532 (1995) Printed in Great Britain The Company of Biologists Limited 1995 1519 Arabidopsis gynoecium structure in the wild type and in ettin mutants R. Allen Sessions* and Patricia
Development 121, 1519-1532 (1995) Printed in Great Britain The Company of Biologists Limited 1995 1519 Arabidopsis gynoecium structure in the wild type and in ettin mutants R. Allen Sessions* and Patricia
Introduction. Copyright 2002 Pearson Education, Inc., publishing as Benjamin Cummings
 Introduction It has been said that an oak is an acorn s way of making more acorns. In a Darwinian view of life, the fitness of an organism is measured only by its ability to replace itself with healthy,
Introduction It has been said that an oak is an acorn s way of making more acorns. In a Darwinian view of life, the fitness of an organism is measured only by its ability to replace itself with healthy,
Kingdom Plantae, Part II - Gymnosperms and Angiosperms
 Kingdom Plantae, Part II - Gymnosperms and Angiosperms I. Introduction Reproduction in the seed plants (Gymnosperms and Angiosperms) has been greatly influenced by the requirements of a terrestrial existence.
Kingdom Plantae, Part II - Gymnosperms and Angiosperms I. Introduction Reproduction in the seed plants (Gymnosperms and Angiosperms) has been greatly influenced by the requirements of a terrestrial existence.
AINTEGUMENTA-like (AIL) genes are expressed in young tissues and may specify meristematic or division-competent states
 Plant Molecular Biology (2005) 57:613 628 Ó Springer 2005 DOI 10.1007/s11103-005-0955-6 AINTEGUMENTA-like (AIL) genes are expressed in young tissues and may specify meristematic or division-competent states
Plant Molecular Biology (2005) 57:613 628 Ó Springer 2005 DOI 10.1007/s11103-005-0955-6 AINTEGUMENTA-like (AIL) genes are expressed in young tissues and may specify meristematic or division-competent states
PLENA and FARINELLI: redundancy and regulatory interactions between two Antirrhinum MADS-box factors controlling flower development
 The EMBO Journal Vol.18 No.14 pp.4023 4034, 1999 PLENA and FARINELLI: redundancy and regulatory interactions between two Antirrhinum MADS-box factors controlling flower development Brendan Davies 1, Patrick
The EMBO Journal Vol.18 No.14 pp.4023 4034, 1999 PLENA and FARINELLI: redundancy and regulatory interactions between two Antirrhinum MADS-box factors controlling flower development Brendan Davies 1, Patrick
AINTEGUMENTA, an A PETA LAP-like Gene of Arabidopsis with Pleiotropic Roles in Ovule Development and Floral Organ Growth
 The Plant Cell, Vol. 8, 155-168, February 1996 O 1996 American Society of Plant Physiologists AINTEGUMENTA, an A PETA LAP-like Gene of Arabidopsis with Pleiotropic Roles in Ovule Development and Floral
The Plant Cell, Vol. 8, 155-168, February 1996 O 1996 American Society of Plant Physiologists AINTEGUMENTA, an A PETA LAP-like Gene of Arabidopsis with Pleiotropic Roles in Ovule Development and Floral
An Enhancer Trap Line Associated with a D-Class Cyclin Gene in Arabidopsis 1
 An Enhancer Trap Line Associated with a D-Class Cyclin Gene in Arabidopsis 1 Kankshita Swaminathan, Yingzhen Yang, Natasha Grotz, Lauren Campisi, and Thomas Jack* Department of Biological Sciences, Dartmouth
An Enhancer Trap Line Associated with a D-Class Cyclin Gene in Arabidopsis 1 Kankshita Swaminathan, Yingzhen Yang, Natasha Grotz, Lauren Campisi, and Thomas Jack* Department of Biological Sciences, Dartmouth
LABORATORY 2: Flowers
 LABORATORY 2: Flowers INTRODUCTION The goal of this laboratory exercise is to familiarize you with flowers, their structure, variation, and importance to the plant. By the end of today s laboratory exercise
LABORATORY 2: Flowers INTRODUCTION The goal of this laboratory exercise is to familiarize you with flowers, their structure, variation, and importance to the plant. By the end of today s laboratory exercise
Supplemental Data. Beck et al. (2010). Plant Cell /tpc
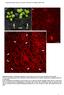 Supplemental Figure 1. Phenotypic comparison of the rosette leaves of four-week-old mpk4 and Col-0 plants. A mpk4 vs Col-0 plants grown in soil. Note the extreme dwarfism of the mpk4 plants (white arrows)
Supplemental Figure 1. Phenotypic comparison of the rosette leaves of four-week-old mpk4 and Col-0 plants. A mpk4 vs Col-0 plants grown in soil. Note the extreme dwarfism of the mpk4 plants (white arrows)
Patterns of Cell Division, Cell Differentiation and Cell Elongation in Epidermis and Cortex of Arabidopsis pedicels in the Wild Type and in erecta
 Patterns of Cell Division, Cell Differentiation and Cell Elongation in Epidermis and Cortex of Arabidopsis pedicels in the Wild Type and in erecta Mark G. R. Bundy, Olivia A. Thompson, Matthew T. Sieger,
Patterns of Cell Division, Cell Differentiation and Cell Elongation in Epidermis and Cortex of Arabidopsis pedicels in the Wild Type and in erecta Mark G. R. Bundy, Olivia A. Thompson, Matthew T. Sieger,
Early Flower Development in Arabídopsis
 The Plant Cell, Vol. 2, 755-767, August 1990 O 1990 American Society of Plant Physiologists Early Flower Development in Arabídopsis David R. Smyth,' John L. Bowman, and Elliot M. Meyerowitz* Division of
The Plant Cell, Vol. 2, 755-767, August 1990 O 1990 American Society of Plant Physiologists Early Flower Development in Arabídopsis David R. Smyth,' John L. Bowman, and Elliot M. Meyerowitz* Division of
3/18/2012. Chapter 36. Flower Parts. Flower Parts. Reproduction in Angiosperms
 Chapter 36 Reproduction in Angiosperms Bryophytes >450mya 360 mya Fig. 27-4, p. 584 Lily Flower Flower Parts Sepals cover and protect flower parts in bud Collectively calyx Petals Can attract animal pollinators
Chapter 36 Reproduction in Angiosperms Bryophytes >450mya 360 mya Fig. 27-4, p. 584 Lily Flower Flower Parts Sepals cover and protect flower parts in bud Collectively calyx Petals Can attract animal pollinators
Chapter 38 Angiosperm Reproduction and Biotechnology
 Chapter 38 Angiosperm Reproduction and Biotechnology Concept 38.1 Pollination enables gametes to come together within a flower Diploid (2n) sporophytes produce spores by meiosis; these grow into haploid
Chapter 38 Angiosperm Reproduction and Biotechnology Concept 38.1 Pollination enables gametes to come together within a flower Diploid (2n) sporophytes produce spores by meiosis; these grow into haploid
Open Flower. Juvenile leaf Flowerbud. Carpel 35 NA NA NA NA 61 NA 95 NA NA 15 NA 41 3 NA
 PaxDB Root Juvenile leaf Flowerbud Open Flower Carpel Mature Pollen Silique Seed Sec3a Sec3b Sec5a Sec5b Sec6 Sec8 Sec10a/b Sec15a Sec15b Exo84a Exo84b Exo84c Exo70A1 Exo70A2 Exo70A3 49 47 8 75 104 79
PaxDB Root Juvenile leaf Flowerbud Open Flower Carpel Mature Pollen Silique Seed Sec3a Sec3b Sec5a Sec5b Sec6 Sec8 Sec10a/b Sec15a Sec15b Exo84a Exo84b Exo84c Exo70A1 Exo70A2 Exo70A3 49 47 8 75 104 79
Plant Reproduction. In a nutshell
 Plant Reproduction In a nutshell 2007-2008 Plant Diversity mosses ferns conifers flowering plants Bryophytes non-vascular land plants Pteridophytes seedless vascular plants Gymnosperm pollen & naked seeds
Plant Reproduction In a nutshell 2007-2008 Plant Diversity mosses ferns conifers flowering plants Bryophytes non-vascular land plants Pteridophytes seedless vascular plants Gymnosperm pollen & naked seeds
The Homeotic Protein AGAMOUS Controls Late Stamen Development by Regulating a Jasmonate Biosynthetic Gene in Arabidopsis W
 The Plant Cell, Vol. 19: 3516 3529, November 2007, www.plantcell.org ª 2007 American Society of Plant Biologists The Homeotic Protein AGAMOUS Controls Late Stamen Development by Regulating a Jasmonate
The Plant Cell, Vol. 19: 3516 3529, November 2007, www.plantcell.org ª 2007 American Society of Plant Biologists The Homeotic Protein AGAMOUS Controls Late Stamen Development by Regulating a Jasmonate
Operation Flower Dissection
 Operation Flower Dissection Classroom Activity: K-4 Time: One to two 50-minute class periods Overview: In this activity, students will observe the similarities and differences between flowers of different
Operation Flower Dissection Classroom Activity: K-4 Time: One to two 50-minute class periods Overview: In this activity, students will observe the similarities and differences between flowers of different
Sex Determination in the Monoecious Species Cucumber Is Confined to Specific Floral Whorls
 The Plant Cell, Vol. 13, 481 493, March 2001, www.plantcell.org 2001 American Society of Plant Physiologists Sex Determination in the Monoecious Species Cucumber Is Confined to Specific Floral Whorls Martin
The Plant Cell, Vol. 13, 481 493, March 2001, www.plantcell.org 2001 American Society of Plant Physiologists Sex Determination in the Monoecious Species Cucumber Is Confined to Specific Floral Whorls Martin
Chapter 31: Plant Reproduction
 Chapter 31: Plant Reproduction Plants and Pollinators Pollen had evolved by 390 million years ago Sperm packed inside a nutritious package Transferred first by wind currents Later transferred by insects
Chapter 31: Plant Reproduction Plants and Pollinators Pollen had evolved by 390 million years ago Sperm packed inside a nutritious package Transferred first by wind currents Later transferred by insects
The AINTEGUMENTA Gene of Arabidopsis Required for Ovule and Female Gametophyte Development 1s Related to the Floral Homeotic Gene APETALA2
 The Plant Cell, Vol. 8, 137-153, February 1996 0 1996 American Society of Plant Physiologists RESEARCH ARTICLE The AINTEGUMENTA Gene of Arabidopsis Required for Ovule and Female Gametophyte Development
The Plant Cell, Vol. 8, 137-153, February 1996 0 1996 American Society of Plant Physiologists RESEARCH ARTICLE The AINTEGUMENTA Gene of Arabidopsis Required for Ovule and Female Gametophyte Development
The NGATHA Genes Direct Style Development in the Arabidopsis Gynoecium C W
 The Plant Cell, Vol. 21: 1394 1409, May 2009, www.plantcell.org ã 2009 American Society of Plant Biologists The NGATHA Genes Direct Style Development in the Arabidopsis Gynoecium C W Marina Trigueros,
The Plant Cell, Vol. 21: 1394 1409, May 2009, www.plantcell.org ã 2009 American Society of Plant Biologists The NGATHA Genes Direct Style Development in the Arabidopsis Gynoecium C W Marina Trigueros,
Turning floral organs into leaves, leaves into floral organs Koji Goto*, Junko Kyozuka and John L Bowman
 449 Turning floral organs into leaves, leaves into floral organs Koji Goto*, Junko Kyozuka and John L Bowman The development of the floral organs is specified by the combinations of three classes of gene
449 Turning floral organs into leaves, leaves into floral organs Koji Goto*, Junko Kyozuka and John L Bowman The development of the floral organs is specified by the combinations of three classes of gene
Introduction. Copyright 2002 Pearson Education, Inc., publishing as Benjamin Cummings
 Introduction It has been said that an oak is an acorn s way of making more acorns. In a Darwinian view of life, the fitness of an organism is measured only by its ability to replace itself with healthy,
Introduction It has been said that an oak is an acorn s way of making more acorns. In a Darwinian view of life, the fitness of an organism is measured only by its ability to replace itself with healthy,
Lab sect. (TA name/time): BIOLOGY 317 Spring First Hourly Exam 4/22/10
 Name: Lab sect. (TA name/time): BIOLOGY 317 Spring 2011 First Hourly Exam 4/22/10 1) (24 pts) Match the letter of the family given on the right with the characteristics for a plant described on the left.
Name: Lab sect. (TA name/time): BIOLOGY 317 Spring 2011 First Hourly Exam 4/22/10 1) (24 pts) Match the letter of the family given on the right with the characteristics for a plant described on the left.
The PRETTY FEW SEEDS2 gene encodes an Arabidopsis homeodomain protein that regulates ovule development
 First posted online on 19 January 2005 as 10.1242/dev.01654 Access the most recent epress version at online http://dev.biologists.org/lookup/doi/10.1242/dev.01654 publication date 19 January 2005 841 The
First posted online on 19 January 2005 as 10.1242/dev.01654 Access the most recent epress version at online http://dev.biologists.org/lookup/doi/10.1242/dev.01654 publication date 19 January 2005 841 The
Signaling in the Nitrogen Assimilation Pathway of Arabidopsis Thaliana
 Biochemistry: Signaling in the Nitrogen Assimilation Pathway of Arabidopsis Thaliana 38 CAMERON E. NIENABER ʻ04 Abstract Long recognized as essential plant nutrients and metabolites, inorganic and organic
Biochemistry: Signaling in the Nitrogen Assimilation Pathway of Arabidopsis Thaliana 38 CAMERON E. NIENABER ʻ04 Abstract Long recognized as essential plant nutrients and metabolites, inorganic and organic
Supplemental Data. Wang et al. (2013). Plant Cell /tpc
 Supplemental Data. Wang et al. (2013). Plant Cell 10.1105/tpc.112.108993 Supplemental Figure 1. 3-MA Treatment Reduces the Growth of Seedlings. Two-week-old Nicotiana benthamiana seedlings germinated on
Supplemental Data. Wang et al. (2013). Plant Cell 10.1105/tpc.112.108993 Supplemental Figure 1. 3-MA Treatment Reduces the Growth of Seedlings. Two-week-old Nicotiana benthamiana seedlings germinated on
Anatomy of a New Sepaloid Mutant Flower in Sugarbeet 1
 January-June, 1991 Anatomy of a New SepaJoid Mutant Flower in Sugarbeet 23 Anatomy of a New Sepaloid Mutant Flower in Sugarbeet 1 J. C. Theurer, S. A. Owens 3, and F. W. Ewers 4 2Sugarbeet, Bean and Cereal
January-June, 1991 Anatomy of a New SepaJoid Mutant Flower in Sugarbeet 23 Anatomy of a New Sepaloid Mutant Flower in Sugarbeet 1 J. C. Theurer, S. A. Owens 3, and F. W. Ewers 4 2Sugarbeet, Bean and Cereal
NOTES: CH 38 Plant Reproduction
 NOTES: CH 38 Plant Reproduction *Modifications in reproduction were key adaptations enabling plants to spread into a variety of terrestrial habitats. * Water has been replaced by wind and animals as a
NOTES: CH 38 Plant Reproduction *Modifications in reproduction were key adaptations enabling plants to spread into a variety of terrestrial habitats. * Water has been replaced by wind and animals as a
PETAL LOSS, a trihelix transcription factor gene, regulates perianth
 Research article 4035 PETAL LOSS, a trihelix transcription factor gene, regulates perianth architecture in the Arabidopsis flower Philip B. Brewer, Paul A. Howles*, Kristen Dorian, Megan E. Griffith, Tetsuya
Research article 4035 PETAL LOSS, a trihelix transcription factor gene, regulates perianth architecture in the Arabidopsis flower Philip B. Brewer, Paul A. Howles*, Kristen Dorian, Megan E. Griffith, Tetsuya
A SIX-STAMENED FLOWER IN ZEA MAYS L.
 130 A SIX-STAMENED FLOWER IN ZEA MAYS L. BY B. C. Department of Botany, University of Leeds (With 2 figures in the text) During the examination of the tassel of a maize plant, a number of flowers were
130 A SIX-STAMENED FLOWER IN ZEA MAYS L. BY B. C. Department of Botany, University of Leeds (With 2 figures in the text) During the examination of the tassel of a maize plant, a number of flowers were
Divergence of Function and Regulation of Class B Floral Organ ldentity Genes
 The Plant Cell, Vol. 9, 559-570, April 1997 O 1997 American Society of Plant Physiologists Divergence of Function and Regulation of Class B Floral Organ ldentity Genes Alon Samach,a Susanne E. Kohalmi,b.l
The Plant Cell, Vol. 9, 559-570, April 1997 O 1997 American Society of Plant Physiologists Divergence of Function and Regulation of Class B Floral Organ ldentity Genes Alon Samach,a Susanne E. Kohalmi,b.l
plant reproduction Alternation of Generations chapter 38
 Alternation of Generations Haploid (n) plant reproduction chapter 38 Diploid (2n) Sporangium Spore dispersal Spore (n) Young Mature (n) ARCHEGONIUM ANTHERIDIUM Sperm Mature Sorus Sporangium sporophyte
Alternation of Generations Haploid (n) plant reproduction chapter 38 Diploid (2n) Sporangium Spore dispersal Spore (n) Young Mature (n) ARCHEGONIUM ANTHERIDIUM Sperm Mature Sorus Sporangium sporophyte
Termination of Stem Cell Maintenance in Arabidopsis Floral Meristems by Interactions
 Cell, Vol. 105, 805 814, June 15, 2001, Copyright 2001 by Cell Press Termination of Stem Cell Maintenance in Arabidopsis Floral Meristems by Interactions between WUSCHEL and AGAMOUS Michael Lenhard, 2
Cell, Vol. 105, 805 814, June 15, 2001, Copyright 2001 by Cell Press Termination of Stem Cell Maintenance in Arabidopsis Floral Meristems by Interactions between WUSCHEL and AGAMOUS Michael Lenhard, 2
The early inflorescence of Arabidopsis thaliana demonstrates positional effects in floral organ growth and meristem patterning
 Plant Reproduction (2018) 31:171 191 https://doi.org/10.1007/s00497-017-0320-3 ORIGINAL ARTICLE The early inflorescence of Arabidopsis thaliana demonstrates positional effects in floral organ growth and
Plant Reproduction (2018) 31:171 191 https://doi.org/10.1007/s00497-017-0320-3 ORIGINAL ARTICLE The early inflorescence of Arabidopsis thaliana demonstrates positional effects in floral organ growth and
Seed Plants Lab. Learning Objectives. Procedure and Questions
 Seed Plants Lab Learning Objectives Define the terms (meanings of the names) angiosperm and gymnosperm State what type of cells create eggs and what type of cells create sperm in gymnosperms and angiosperms
Seed Plants Lab Learning Objectives Define the terms (meanings of the names) angiosperm and gymnosperm State what type of cells create eggs and what type of cells create sperm in gymnosperms and angiosperms
2014 Pearson Education, Inc. 1
 1 Stamen Anther Filament Stigma Carpel Style Ovary Petal Sepal Ovule 2 A B Sepals Petals Stamens Carpels C A + B gene activity B + C gene activity C gene activity Carpel Petal (a) A schematic diagram of
1 Stamen Anther Filament Stigma Carpel Style Ovary Petal Sepal Ovule 2 A B Sepals Petals Stamens Carpels C A + B gene activity B + C gene activity C gene activity Carpel Petal (a) A schematic diagram of
W.4.1 Write opinion pieces on topics or texts, supporting a point of view with reasons and information.
 Flower Dissection Lesson Overview Flowers use pollination as a mechanism for reproduction and survival. Students will learn about pollination and how each structure plays a role in this process. They will
Flower Dissection Lesson Overview Flowers use pollination as a mechanism for reproduction and survival. Students will learn about pollination and how each structure plays a role in this process. They will
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/1171320/dc1 Supporting Online Material for A Frazzled/DCC-Dependent Transcriptional Switch Regulates Midline Axon Guidance Long Yang, David S. Garbe, Greg J. Bashaw*
www.sciencemag.org/cgi/content/full/1171320/dc1 Supporting Online Material for A Frazzled/DCC-Dependent Transcriptional Switch Regulates Midline Axon Guidance Long Yang, David S. Garbe, Greg J. Bashaw*
